Chapter 26 RNA metabolism
|
|
|
- Gary Warner
- 7 years ago
- Views:
Transcription
1 Chapter 26 RNA metabolism 26.0 Intro RNA appears to be very similar to DNA has the 2' OH on the ribose has U instead of T Most RNA functions as single stand so folds back on itself Gives more structural diversity RNA functions in information storage, information transmission and as enzyme. Suspect it may have been The molecule in primitive cell Will see RNA interaction with protein for most functions RNA will have both structural and catalytic role in these structure All RNA except RNA of RNA viruses derived from DNA. DNA sequence copied onto RNA through process of transcription 3 major kinds of RNA Messenger mrna carries genetic information from nucleus to ribosome for translation Transfer trna associates specific AA with specific triplets on mrna Ribosomal rrna integral structural part of ribosome Also lots of RNA that does not fit above categories Regulatory, catalytic functions or precursors In mammalian cells there are more of these than m,t,&r! Transcription more selective than replication will only transcribe one gene or one group of genes, not entire DNA regulation of this process described in chapter 28 Sum of all RNA synthesized in cell is called transcriptome From last chapter know that in mammalian cells the amount of DNA that actually codes for protein is small, so might expect transcriptome to be small. But you would be wrong. It is actually quite large. A wide range of special function RNA s are made, and we really don t know what all of them do!
2 2 Here will concentrate on synthesis of RNA from DNA and processing and turnover of RNA molecules Will also look at RNA template/ DNA copy system 26.1 DNA Dependent RNA synthesis Will compare to DNA that we just did similar chemical mechanism polarity use of template initiation, elongation, termination stages different does NOT require a primer uses a limited segment of DNA only 1 strand of DNA is the template A. RNA synthesized by RNA polymerases DNA dependent RNA polymerase Isolated by 4 different groups in 1960's Needs DNA template 4 NTP s Mg Protein also binds Zn! Diagram figure 26-1a Add units to the 3' end (same mech as DNA, the OH acts as a nucloephile to attach the á PO 4 so releases PPi So 5' 3' polarity (NMP) n + NTP (NMP) n+1 + PPi Requires DNA for activity Most active on double stranded DNA Most active when bound to double stand DNA, yet only 1 of 2 strands copied in 3' 5' direction using template No primer required RNA polymerase binds at specific sequences called promoters 5'PPP of first residue not cleaved but remains Only about 8 bp of RNA are bound to DNA before peels off and DNA duplex reforms
3 3 E coli RNA polymerase keep about 17 bp of DNA unwound for transcription Proceeds at a rate of nucleotides/sec (DNA Pol III 1000/sec DNA Pol I 16-20/sec) DNA can t unwind this fast so observe + supercoiled ahead and - supercoiled behind (26.1B,C) So need topoisomerases to relieve strain 2 strand of DNA different functions Template strand - the one RNA codes from Non-template strand or coding strand - identical in sequence to RNA except for T/U switch Either stand can be template strand Regulatory sequences, by convention are the sequence in the nontemplate (coding) strand DNA dependent RNA polymerase in E coli 5 core subunits á2ââ ù 390,000 MW th A 6 unit ó Part of a family of similar units Bind transiently to core, to direct binding to different promoters 70 Most common ó is ó (70,000 MW) 4 5 No proofreading ability. 1 mistake in 10 to 10 Not as critical Many RNA s made from 1 DNA RNA s are used for a while then degraded, so not permanent Have observed RNA polymerase pausing after incorporating Mismatch, and reaction can be reversed, But don t know if this is real or artifact B. RNA synthesis initiated at promoters RNA is initiated at specific sites called promoters many differ promoters, so are of intense research In E coli promoter region runs from about 70 bp upstream to 30 bp downstream from actual initiation site 70 Promoter for ó has features in figure short similar sequenced at -10 and -35 (10 and 35 before the transcribed sequence)
4 4 Not always the same but get a consensus -10 is TATAAT -35 is TTGACA Highly expressed genes may have additional AT-rich upstream recognition element called UP in -40 to -60 Bound by á subunit of polyermase Efficiency that bind and initiate transcription determined by how close sequence is to consensus, if they are spaced correctly, and how far they are from actual start of transcription A change of 1 base in -10 and -35 can drop efficiency times Overall initiation broken into 2 phases Figure 26-6 Binding - binding of RNA polymerase to promoter on DNA Directed by ó factor First a closed complex (DNA is intact) Then DNA is unwound in -10 region to form open complex Initiation Movement to the actual start place of transcription Start of transcription ó drops off randomly after elongation begins Is replaced with NusA protein NusA only comes off when transcription is complete Other ó factors Table 26-1 other promoter than one bound by ó 70 in previous chapter talked about heat shock proteins 32 these are made when ó is expressed Use of different ó used to coordinate expression of different sets of genes C. Transcription is Regulated at several levels Transcription regulated so only those genes needed by cell are expressed But most control is targeted at regulation of RNA polymerase binding, Additional regulation occur in all steps of transcription including: initiation, elongation and termination Control mechanisms 1. Different ó bind to different promoters 2. binding of regulatory proteins Activators - bind to a DNA to increase binding of polymerase camp receptor protein (CRP) expressed when cell doesn t have glucose as a source of E, Turn on genes needed to utilize other sugars
5 5 Repressors - bind to DNA to block synthesis of RNA Lac repressor. if lactose not available, block expression of these genes (will see in chapter 28) D. Specific Sequences Signal Termination of RNA Synthesis RNA synthesis is highly processive - Protein can t fall off till job is done Certain DNA sequence pause in RNA synthesis If pause is long enough, will terminate not well understood in eukaryotes so will look at E coli 2 mechanisms ñ-independent (Figure 26-7a) RNA contain self complementary sequences Hairpin form based before end of strand End of message has 3 conserved A s that are transcribed to U s Polymerase hits poly A it pauses When pauses hairpin forms Hairpin disrupts RNA/polymerase binding and complex falls off ñ - dependent Figure 26-7b No poly A Usually a CA rich sequence called rut Rho utilization element ñ protein binds to RNA at a specific site Move 5' 3' on RNA until it reach polymerase that is paused In releases the RNA polymerase with use of ATP Mech not known E. Eukaryotic cells have three kinds of Nuclear RNA polymerases More complicated system 3 different complexes - share several components RNA polymerase I (pol I) - only for synthesis of preribosomal RNA 1 long piece that contains 3 smaller pieces. The the pieces are 3 different ribosomal RNA s Called 18s 5.8s and 28S more later in chapter Great variety in sequence of promoters between species RNA polymerase II (Pol II) primarily for synthesis of mrna s And some specialized RNA s Recognizes thousands of promoters Common feature is TATA box at -30 and Inr sequence at +1 See figure 26-8
6 6 RNA polymerase III (Pol III) makes trna s, 5S rrna and some specialized RNA s 5S is a fourth rrna and 5.8s Promoters well characterized Some control points are inside the gene rather than at the starting end! F. RNA Polymerase II requires many other proteins Polymerase itself 12 subunits (prokaryote was only 5) Largest subunit, RBP1, high degree of homology with bacterial â Carboxy terminal tail with many repeats of sequence YSPTSPS 27 repeats, 18 exact in yeast 52 repeats, 21 exact in mouse and human Give domain special name CTD (carboxyl-terminal domain) Separated from main body with unstructured region Will see several special roles for this region RBP2 similar to bacterial â RBP 3 and RBP 11 - some similarity to bacterial á Whole host of other proteins called transcription factors (TF...) To form active transcription complex General Transcription factors - required for all pol II promoters TFIIA -transcription factor A for Pol II etc Highly conserved for all eukaryotes Overall transcription split into many phases Assembly Initiation Elongation Termination Keep an eye on figure 26-9 and table 26-2 as discuss details In process many proteins may be present in preassembled complexes I. Assembly TBP (TATA binding protein) binds to TATA box (If no TATA box TBP comes in with TFIID, but this process not well understood and not in figure 26-9) Bind TFIIB to TBP - this enlarges extend of DNA binding Bind TFIIA not essential but tightens binding at weaker sites Bind TFIIF and RNA polymerase II (This factor helps make more specific to site) Finally TFIIE and TFIIH bind to make complete closed complex H is a helicase - starts unwinding DNA near initiation site Requires ATP
7 7 Now get open complex Didn t show you all the details actually >30 peptides involved! II. Initiation TFIIH also acts as a kinase Phosphorylates several sites on Pol II in CTD This activates pol II to start transcription CTD can also be phosphorylated by other enzymes like CDK9) Phosphorylation may have other effects as well in elongation As first nucleotides synthesized TFIIE and TFIIH are released factors marks entry into elongation III. Elongation TFIIF stays attached Other factors - elongation factors - bind to enhance activity (Don t confuse with protein synthesis elongation factors!) Some of these factors bind to phosphates on CTD tail Keep elongation from pausing Some factors involved in post transriptional processing IV. Termination When transcription is complete and complex terminated, PolII is dephosphorylated so can begin another transcript Not understood V. Regulation Elaborate, involved many proteins with initiation complex Will study details in chapter 28 Some proteins interact with transcription factors Some with PolII itself VI. TFIIH - Many functions Have seen several activities Helicase to open DNA and kinase to phosphorylate Pol II Is a complex and some subunits of complex are also used in excision repair of DNA If transcription hits a damaged DNA TFIIH initiates repair mech Lack of TFIIH associated with some human diseases Xeroderma pigmentosum and Cockaynes syndrome
8 8 G. DNA-dependent RNA polymerase can be selectively inhibited Inhibited by actinomycin D in both prokaryotes and eukaryotes Figure inserts between 2 GC pairs in DNA prevents movement of polymerase this is only thing it does so can be used as diagnostic for things linked to RNA synthesis Acridine is the same Rifampicin - binds to â subunit of bacterial RNA polymerase Prevents release of promoter Sometimes used as an antibiotic Mushroom Amanita phalloides makes á-amanitin Stops mrna synthesis in animal by blocking RNA pol II can also block pol III in higher conc, but does not block RNA pol I or bacterial RNA polymerase, or the mushrooms own pol II 26.2 RNA processing newly synthesized RNA call Primary Transcript Many RNA molecules in bacteria and almost all RNA in eukaryotes need to be processed after synthesis -fair amount of process in both pro and eukaryotic trna - cleavage of ends and lots of base and sugar modification - eukaryotic mrna -cutting out regions and adding caps at both ends Also the final processing - want to destroy mrna s after they have been used! Some of the enzymes that do this processing are RNA s not proteins call these ribozymes Newly synthesized RNA molecule called primary transcript Most processing done in eukaryotic mrna, pro and eukary t-rna special function RNA s also processed Let s look at overview Figure Start with Eukaryotic mrna 5' end Splice 3' end Elaborate complexs perform each processing step, sometimes all happening at once, Most all interacting with that phophylated CTD tail on polymerase. Even tied with proteins that bind to RNA in nucleus and transport into cytosol and delivered to ribosome then go to pro and eu t-rna
9 9 A. Eukaryotic mrna s capped at 5' end 5' cap - 7-methyl guanine linked to 5' terminal residue (See figure 26-12) Protects from ribonucleases Used for specific binding interactions in ribosome (next chapter) G added first using GTP Then methylated st nd Additional methylations on 1 and 2 sugar 2' OH s Processing done very early After first 20 or 30 residues added All enzymes needed bound to CTD domain Once cap is added Release from enzymes but bound to CTD by cap-binding complex(cbc) B. Both Introns and Exons are transcribed from DNA to RNA in bacteria RNA is collinear with proteins sequence and protein can be synthesized directly from RNA (in fact Proteins synthesis may be initiated before the mrna is off the DNA) ( no separate nucleus so this is easy) Genes in eukaryotes contain introns - noncoding regions in RNA in vertebrates almost all proteins EXCEPT Histones have introns bacteria have a few introns In transcription entire DNA sequence introns and exons are transcribed by RNA polymerase Introns then cut out and exons spliced together Weren t discovered until late 1970's when tried to hybridize RNA back to its DNA Most eukaryotic exons <1000 bases, many in range (30-60 AA s) introns vary from 50 to 20,000 nucleotides In higher eukaryotes may have more bases in introns than exons C. RNA Catalyzed Splicing 4 types of introns I & II share key characteristics but some differences in details of splicing I some nuclear, mitochondrial, and chloroplast r, t, and mrna II usually mitochondrial or chloroplast mrna in fungi algae and plants Some I and II found in rare bacterial splicing events Neither required ATP for splicing
10 10 Both use similar mech - figure a transesterification reaction OH of one ribose attacks phosphate of another, no E is lost so that s why don t need ATP Type I mech Figure need GTP, but not for energy (can use GMP, or GDP!) Uses 3'OH of ribose as nucleophile Forms a 3'-5' bond with the 5' end of the intron The 3'OH on the exon then attaches the splice point And get the exon spliced out Type II mech (figure 26-15) Uses a 2' OH from an A inside the exon to start it off End up with a lariat NO ENZYMES REQUIRED - self splicing! Third type of splicing - requires protein - Most common mechanism Called Spliceosomal intron Because splicing is done in a protein complex called spliceosome Figure a&b Largest class Nuclear mrna s Ends up with lariats (so resembles type II) But requires RNA-protein complexes called Small nuclear ribonucleoproteins snrnp s or snurps Each snrpp contains has its own RNA Called snrna for small nuclear snrna of residue 5 snrna s (U1,U2,U4,U5,U6) are abundant in nucleus And several proteins Both protein and RNA parts are highly conserved Spliceosomal introns usually GU at 5' end AG at 3' end U1snRNA has sequence complimentary to 5' splice site of many introns U1snRNP binds to this region Then add U2, U4,U5,and U6 snrnp s to make Spliceosome Contains the 5 RNA s and about 50 proteins
11 11 Need ATP so assemble spliceosome, but not for splicing reaction A less common spliceosome contains U11 and U12 instead of U1 and U2 snrnp s 5' AU...AC(3') sequence Some splicing components tethered to CTD? Figure 26-16c First 5' junction and spliceosome tethered to CTD Then 3' end comes through and splice removed After splicing intron remains in nucleus and is degraded Fourth type of splicing Used in certain trna s Requires a splicing endonuclease Requires ATP Endonuclease cleaves at both end of intron 2 ends are joined like in a ligase with ATP for E Type III splicing with spliceosome limited to eukaryotes Type I & II splicing seen in some bacteria and viruses Splicing more common in archaea than in bacteria D. Eukaryotic mrna also has a distinctive 3' end 80 to 250 residues of A Binding site for specific proteins Some protein help protect from degradation Some used in interaction with ribosome Some prokariots also get a poly A tail But in prokariots this stimulates degradation! Mech Figure RNA extended past site where put on tail Cleaved at addition site by an endonuclease Endonuclease part of large enzyme complex associated with CTD Site marked by sequence AAUAA 10 to 30 residues on 5' side (upstream) G-U rich region residues downstream Cleavage leave a 3' OH open Polyadenylate polymerase then used ATP to add A tail Does not require a template Does require a primer
12 12 Overall processing summarized in figure E. Multiple Products are derived form 1 gene via differential splicing Splicing seem to be a pretty wasteful process don t really know why at this point some cells have used to their advantage Most eukaryotic mrnas produce a single mature RNA Some can be processed differently to make different mature RNA and different proteins COMPLEX TRANSCRIPTS Figure Processing to obtains different mrna 1 way differentiates for attachment of poly A tail 26-19a Used in variable domain of immunoglobins Another way alternate splicing patterns 26-19b This is observed in 3 different myosins found in fruit flies Both are used in production of two different hormones from rat thyroid and brain (figure 26-20) (calcitonin & calcitoningene-related peptide CGRP) Many, perhaps most genes in mammalian genomes use alternative splicing Not used as much in lower eukariots F. rrna and trna also undergo processing rrna of both prokaryote and eukaryotes made from large precursors that must be cleaved Also many methyl groups added and bases modified Figure Long pieces called pre-ribosomal RNA s Bacteria 5S,16S and 23S and a few trna s all come from one 30S Does anybody know what S means? RNA of about 6,500 bases RNA removed at both ends and in between 16S RNA 11 base modifications 10 bases methylated 23S RNA 11 bases modified 12 methylated Ecoli has 7 such preribosomal RNA sequences scattered throughout the genone See figure rrna sequences the same
13 13 Eukaryotes In between sequences differ Cleaved by RNAses, not by splicing RNA synthesized using RNA Polymerase I 45S preribosomal RNA processed in nucleolus Figure To make 18,28 and 5.8 rrna s Ribosome also need a 5S rrna Entirely separate gene and uses RNA polymerase III Ribosome assembled in complex in Nucleolus Includes all nucleases required for cleavages All enzymes required for base modifications Up to 200 base modifications Each relies on a different snorna- Protein complex (snornp) Typically 1 snorna & 4-5 proteins Yeast complex 170 nonribosomal proteins 70 snorna s, 78 ribosomal proteins Humans have more base modifications so even more snorna s SnoRNA (figure 26-25) Small nucleolar RNA s snornas Guide modification, cleavage and some protein interaction nucleotides Typically an intron from another gene Contains sequence that is complementary to site on rrna where interaction occurs t-rna s Figure Most cells have distinct trna s (20 AA s, 64 anticodons) Eukaryotic have multiple copies of many t-rna genes Usually have to take off RNA from both ends and eukaryotes may have introns to splice out Sometimes may have 2 or more trnas on a single RNA that need to be separated
14 14 RNAse P removes 5' end Found in all organisms Has both protein and RNA Can function in bacteria without protein! 3' end removed by other endonucleases including RNaseD Further processing Amino acid will be attached to a CCA sequence on th 3' end many bacterial and ALL eukaryotic don t have this sequence Must be added by trna nucleotidyl transferase Has separate sites for all 3 bases Binds of RNA and adds No template needed Base modification Methylation, deamindation, reduction of some bases Examples figure Occur at characteristic positions in structure Sometimes need for structure Inosine will be important in next chapter as the wobble base G. Special-function RNA processing The variety and number of special function RNA s is rapidly expanding (Was not even in 4th edition of text) Many of these RNA s require processing snrna s SnRNA s synthesized as large precursor by RNA polymerase II Ribonucleases remove excess from ends 11 base modifications snorna Some snorna are introns of other genes Again precursor is larger than actual product As splicing occurs, sno protein binds and ribonuclese cuts the excess RNA off at both ends Micro RNA s (mirna) (new 5th edition) ~22 nucleotides long Complementary to sequence in particular regions of mrna Regulate expression by cleaving or suppressing translation Involved in gene regulation (Chapter 28) Present in all multicelluar organisms May be 1% of human genome And targets on 1/3 of the mrna s Figure 26-27
15 15 Synthesized as large precusor Will not worry about details of processing H. Some Event in RNA metabolism are catalyzed by RNA enzymes 3 best studied Self splicing of type I introns RNase P Hammerhead ribozyme Do transesterification or phosphodiester bond cleavage Self splicing group I intron 400 bases RNAse P 337 bases Hammerhead ribozyme total of 41 bases in 2 strands (Critical portion of larger enzyme but can function alone) activity depend on structure Heat or solvent denature and destroy activity I. Group I introns Had Mechanism earlier Figure Secondary structure shown Enzymatic properties Acts like enzyme Binding of GTP can be saturated, has a Km of 30ìM Can be competitively inhibited by inhibitor (3' deoxyg) Very precise in activity Uses it s own RNA to make alignments internal sequence guide Originally thought that self splicing technically not an enzyme because can t be reused But intron from Teterahymena can guide other splicing events II. Characteristic of other Ribozymes RNAse P 337 nucleotides 17,500 MW protein (13%) (~ 160AA s) 111,000 MW RNA (330X 337) (86%) 156 AA s Under some conditions RNA can work alone RNA can recognize shape of pre-trna and CCA sequence (Implies reaction is second in processing?) Protein stabilizes RNA and facilitates function Can cleave CCA from diverse sequences
16 16 Hammerhead ribozyme From a plant virusoid Small RNA associated with a plant virus Promotes self splicing reaction I. Cellular mrna degradation want RNA synthesis to balance RNA degradation part of control on protein levels Rate of RNA degradation vary greatly for different genes Protein needed briefly, RNA may last a minute or a few seconds Protein needed continuously may be stable over several cell lifetimes! In eukaryotes average lifetime about 3 hours Pool usually exchanged about 10x / generation Half life of bacterial mrna about 1.5 min in bacteria In E coli usually start with one or more cuts by endoribonuclease Followed by 3' 5' exoribonuclease In lower eukaryotes Shorten poly A tail Decap 5' end Degrade 5' 3' All eukaryotes have complex of up to 10 3' 5' exoribonucleases called exosome Usually involved in processing 3' ends of r- and t- and special function RNA s like sn and sno May also be major degredation pathway in higher Eukaryotes Some bacterial mrna have a hairpin loop in ñ-independent terminator adds stability Similar hairpins also seen in more stable primary transcripts I. Polynucleotide Phosphorylase A curiosity discovered in 1955 could synthesize RNA randomely from 5' diphosphates (NOT TRI)
17 17 Needed no template can t use NTP s Reaction is reversible so think is really and endonuclease Can be used to synthesize your own random RNA to your specifications 26.3 RNA-Dependent Synthesis of RNA and DNA So far DNA has been the template there are some cases where RNA is the template In particular RNA viruses So not just DNA to RNA to Protein but RNA to RNA and RNA to DNA A. Reverse Transcriptase produced DNA from viral RNA Figure Retrovirus life cycle Certain RNA viruses that infect animal cells carry a RNA dependent DNA polymerase or reverse transcriptase in the virus particle Genome is a single stranded RNA, usually about 10,000 nucleotides long st 1 step in infection is to copy viral RNA into DNA Then degrade RNA and replace with DNA Then often incorporate DNA into host DNA When host transcribes this gene then start making viral particles These are the retroviruses Predicted to exist in 1960's Detected in 1970's Typically have 3 genes Figure Gag, pol, and env Single transcript carries gag and pol, LTR s & ø (LTR Long terminal repeat) Translated as a single protein Then cleaved into 6 proteins with different functions Protein from gag portion make up core of viral particle Pol gene contains protease to cleave the long peptide Integrase for integrating DNA into host Reverse transcriptase Env gene is viral coat protein
18 At end of RNA are long terminal repeats (LTR s) of a few hundred bases Used to help integrate DNA into host genome Contain promoters for viral genes Reverse Transcriptase Many reverse transcriptases have 2 subunits, á&â â is 90,000 á is 65,000 fragment of beta Like many other RNA and DNA polyerases, reverse transcriptase contain Zn 2+ Works best with its own RNA but can be used with others Does 3 different reactions 1 Synthesis of DNA on RNA template 2. degradation of RNA template 3. DNA synthesis on DNA template DNA/RNA synthase and RNAse different parts of enzyme For DNA synthesis needs a cellular trna primer Is a cellular trna carried in virus, that came from previously infected cell trna 3' end complimentary with a sequence in viral RNA DNA synthesized in 5'-3' direction (like everybody else) Reverse transcriptase does not have a 3'-5' proofreading Error rate 1/20,000 bases Very high error rate seems typical for RNA viruses (Error rate for HIV virus is even worse!) That is why RNA viruses have faster viral evolution Reverse transcriptases useful for biotech industry If you have a protein you want to express You isolate the RNA on the ribosome as the protein is being synthesized Use the reverse transcriptase to make back into DNA for your work B. Retroviruses cause cancer and AIDS most retroviruses do not kill host, but integrate into host genome and get expressed when host divides Some viruses - RNA tumor viruses - contain an oncogene Cause cell to grow abnormally Human immunodeficiency virus (HIV) causes acquired immune deficiency syndrome (AIDS) - a retrovirus 18
19 A fairly standard retrovirus with a few extra genes Not standard because kill host cells (T lymphocytes) instead of making tumors By killing T lymphocytes kills immune system Not standard because reverse transcriptase 10x less accurate than most 1 or more errors in very copy! Any 2 viral copies are different Makes hard to make a vaccine because coat protein is always changing C. Transposons, Retroviruses and Introns - a Common origin? Some Transposons (See DNA chapter) have structure very similar to retroviruses Sometimes transposons closely resemble retroviruses called retrotransposons Code for an enzyme homologous to reverse transcriptase Flanked by LTR s Transpose from one location to another using RNA intermediate ( RNA RNA/DNA DNA/DNA) Probably using their reverse transcriptase Most eukaryotic transposons use this mech Prokaryotic use a different mech (DNA intermediate) Retrotransposons lack an env gene so no viral particle A defective virus caught in a cell!? (poor thing) Sequence comparisons suggest that reverse transcriptase is ancient enzyme May predate multicellular organisms Many Group I and Group II introns are also genetically mobile! Spliced out intron codes for a DNA endonuclease Allows these intron to be introduced into homologous genes that don t have introns Process called homing (figure 26-37) Group I homing is DNA based Group II homing is RNA based Suggest introns are artifact of ancient retrovirus and transposons??? 19
20 20 D. Telomerase is a specialized reverse transcriptase Telomere - a specialized structure at the ends of linear eukaryotic chromosomes generally many copies of short oligonucleotide sequence T G on one 3' strand C A on 5' x and y between 1 and 4 x y y x Range in length form a few dozen base pairs (ciliated protozoa) to 10 of thousands in mammals TG side longer than other side so has region of single stranded DNA Can be up to a few hundred bases on 3' end of strand Ends of DNA not easily replicated by cellular polymerase DNA replication needs template and primer Look back at figure synthesis of okazaki fragments How do you replicate the 3' strand and an end? Need to get an RNA primer RNA primer then cleaved off This would shorten the end every time!! (The 5' end is OK because Okazaki fragment formed in interior then DNA out to end) This problem solved by telomerase that adds telomeres to end of DNA Uses an unusual mech Is a RNA/protein complex RNA part 150 nucleotides Contains about 1.5 copies of a CyA x repeat Figure Its RNA base pairs with 3' end of DNA Uses its own RNA as template for making DNA to fill in missing end on 3' DNA Thus is a reverse transcriptase But unlike others can only use its own RNA as a short template After extension of TxG y strand, CyA x synthesized by polymerases Single stranded region protected by specific binding protein in lower Eukariots (espcially if < a few 100 bp) In higher eukaryots >1000 bp Single strand end sequestered in T-loop Figure 26-38b Loop bound by protein TRF1 & TRF2 to protect from nucleases and repair enzymes
21 21 Telomeres and aging Protozoans can lose telomerase activity If this happens genes get shorter and shorter Eventually cell line dies Also happens in cell lines cultured from humans Germ-lines - cell lines that reproduce forever Contain telomerase activity Try to culture somatic cells They lack telomerase activity They die quickly Fibroblast See a link between age of individual cell is take from and Length of telomere If introduce telomerase activity into the cell line Cells live much longer Definitely an area for future research E. Some Viral RNA s are replicated by RNA dependent RNA polymerase some E. coli bacteriophages, and some eukariotic viruses contain RNA genomes RNA serves as mrna for viral proteins RNA synthesized by an RNA dependent RNA polymerase Never a DNA intermediate, that is why not called a retrovirus Protein is 210,000 MW Has 4 subunits One subunit 65,000 MW Active site of replication Product of viral rep gene 3 other units normally part of e. coli protein synthesis! Tu and Ts factors used in carrying trna to ribosome S1 normal part of 30S ribosome? help rep protein find and binds viral RNA?? Makes RNA complementary to viral RNA Looks like a normal polymerase except requirement or RNA template No proofreading ability so high error rate Will not replicate DNA Will not replicate other RNA s, specific for it s own
22 F. RNA Synthesis offers clues to evolution interesting speculations but few hard facts for teaching but it is an interesting read 22
2013 W. H. Freeman and Company. 26 RNA Metabolism
 2013 W. H. Freeman and Company 26 RNA Metabolism CHAPTER 26 RNA Metabolism Key topics: Transcription: DNA-dependent synthesis of RNA Capping and splicing: RNA processing Overview of RNA Function Ribonucleic
2013 W. H. Freeman and Company 26 RNA Metabolism CHAPTER 26 RNA Metabolism Key topics: Transcription: DNA-dependent synthesis of RNA Capping and splicing: RNA processing Overview of RNA Function Ribonucleic
Structure and Function of DNA
 Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Sample Questions for Exam 3
 Sample Questions for Exam 3 1. All of the following occur during prometaphase of mitosis in animal cells except a. the centrioles move toward opposite poles. b. the nucleolus can no longer be seen. c.
Sample Questions for Exam 3 1. All of the following occur during prometaphase of mitosis in animal cells except a. the centrioles move toward opposite poles. b. the nucleolus can no longer be seen. c.
2. The number of different kinds of nucleotides present in any DNA molecule is A) four B) six C) two D) three
 Chem 121 Chapter 22. Nucleic Acids 1. Any given nucleotide in a nucleic acid contains A) two bases and a sugar. B) one sugar, two bases and one phosphate. C) two sugars and one phosphate. D) one sugar,
Chem 121 Chapter 22. Nucleic Acids 1. Any given nucleotide in a nucleic acid contains A) two bases and a sugar. B) one sugar, two bases and one phosphate. C) two sugars and one phosphate. D) one sugar,
DNA Replication & Protein Synthesis. This isn t a baaaaaaaddd chapter!!!
 DNA Replication & Protein Synthesis This isn t a baaaaaaaddd chapter!!! The Discovery of DNA s Structure Watson and Crick s discovery of DNA s structure was based on almost fifty years of research by other
DNA Replication & Protein Synthesis This isn t a baaaaaaaddd chapter!!! The Discovery of DNA s Structure Watson and Crick s discovery of DNA s structure was based on almost fifty years of research by other
4. DNA replication Pages: 979-984 Difficulty: 2 Ans: C Which one of the following statements about enzymes that interact with DNA is true?
 Chapter 25 DNA Metabolism Multiple Choice Questions 1. DNA replication Page: 977 Difficulty: 2 Ans: C The Meselson-Stahl experiment established that: A) DNA polymerase has a crucial role in DNA synthesis.
Chapter 25 DNA Metabolism Multiple Choice Questions 1. DNA replication Page: 977 Difficulty: 2 Ans: C The Meselson-Stahl experiment established that: A) DNA polymerase has a crucial role in DNA synthesis.
Complex multicellular organisms are produced by cells that switch genes on and off during development.
 Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Control of Gene Expression
 Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
1 Mutation and Genetic Change
 CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Transcription: RNA Synthesis, Processing & Modification
 Transcription: RNA Synthesis, Processing & Modification 1 Central dogma DNA RNA Protein Reverse transcription 2 Transcription The process of making RNA from DNA Produces all type of RNA mrna, trna, rrna,
Transcription: RNA Synthesis, Processing & Modification 1 Central dogma DNA RNA Protein Reverse transcription 2 Transcription The process of making RNA from DNA Produces all type of RNA mrna, trna, rrna,
Central Dogma. Lecture 10. Discussing DNA replication. DNA Replication. DNA mutation and repair. Transcription
 Central Dogma transcription translation DNA RNA Protein replication Discussing DNA replication (Nucleus of eukaryote, cytoplasm of prokaryote) Recall Replication is semi-conservative and bidirectional
Central Dogma transcription translation DNA RNA Protein replication Discussing DNA replication (Nucleus of eukaryote, cytoplasm of prokaryote) Recall Replication is semi-conservative and bidirectional
Lecture Series 7. From DNA to Protein. Genotype to Phenotype. Reading Assignments. A. Genes and the Synthesis of Polypeptides
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Chapter 6 DNA Replication
 Chapter 6 DNA Replication Each strand of the DNA double helix contains a sequence of nucleotides that is exactly complementary to the nucleotide sequence of its partner strand. Each strand can therefore
Chapter 6 DNA Replication Each strand of the DNA double helix contains a sequence of nucleotides that is exactly complementary to the nucleotide sequence of its partner strand. Each strand can therefore
Module 3 Questions. 7. Chemotaxis is an example of signal transduction. Explain, with the use of diagrams.
 Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Protein Synthesis How Genes Become Constituent Molecules
 Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Transcription and Translation of DNA
 Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Name Class Date. Figure 13 1. 2. Which nucleotide in Figure 13 1 indicates the nucleic acid above is RNA? a. uracil c. cytosine b. guanine d.
 13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
From DNA to Protein. Proteins. Chapter 13. Prokaryotes and Eukaryotes. The Path From Genes to Proteins. All proteins consist of polypeptide chains
 Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION. Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.
 Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Name Date Period. 2. When a molecule of double-stranded DNA undergoes replication, it results in
 DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
Molecular Genetics. RNA, Transcription, & Protein Synthesis
 Molecular Genetics RNA, Transcription, & Protein Synthesis Section 1 RNA AND TRANSCRIPTION Objectives Describe the primary functions of RNA Identify how RNA differs from DNA Describe the structure and
Molecular Genetics RNA, Transcription, & Protein Synthesis Section 1 RNA AND TRANSCRIPTION Objectives Describe the primary functions of RNA Identify how RNA differs from DNA Describe the structure and
Appendix C DNA Replication & Mitosis
 K.Muma Bio 6 Appendix C DNA Replication & Mitosis Study Objectives: Appendix C: DNA replication and Mitosis 1. Describe the structure of DNA and where it is found. 2. Explain complimentary base pairing:
K.Muma Bio 6 Appendix C DNA Replication & Mitosis Study Objectives: Appendix C: DNA replication and Mitosis 1. Describe the structure of DNA and where it is found. 2. Explain complimentary base pairing:
13.4 Gene Regulation and Expression
 13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
AP BIOLOGY 2009 SCORING GUIDELINES
 AP BIOLOGY 2009 SCORING GUIDELINES Question 4 The flow of genetic information from DNA to protein in eukaryotic cells is called the central dogma of biology. (a) Explain the role of each of the following
AP BIOLOGY 2009 SCORING GUIDELINES Question 4 The flow of genetic information from DNA to protein in eukaryotic cells is called the central dogma of biology. (a) Explain the role of each of the following
Basic Principles of Transcription and Translation
 The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific traits by dictating the synthesis of
The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific traits by dictating the synthesis of
GENE REGULATION. Teacher Packet
 AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
RNA: Transcription and Processing
 8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
BCH401G Lecture 39 Andres
 BCH401G Lecture 39 Andres Lecture Summary: Ribosome: Understand its role in translation and differences between translation in prokaryotes and eukaryotes. Translation: Understand the chemistry of this
BCH401G Lecture 39 Andres Lecture Summary: Ribosome: Understand its role in translation and differences between translation in prokaryotes and eukaryotes. Translation: Understand the chemistry of this
DNA Replication in Prokaryotes
 OpenStax-CNX module: m44488 1 DNA Replication in Prokaryotes OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
OpenStax-CNX module: m44488 1 DNA Replication in Prokaryotes OpenStax College This work is produced by OpenStax-CNX and licensed under the Creative Commons Attribution License 3.0 By the end of this section,
RNA & Protein Synthesis
 RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
Transcription in prokaryotes. Elongation and termination
 Transcription in prokaryotes Elongation and termination After initiation the σ factor leaves the scene. Core polymerase is conducting the elongation of the chain. The core polymerase contains main nucleotide
Transcription in prokaryotes Elongation and termination After initiation the σ factor leaves the scene. Core polymerase is conducting the elongation of the chain. The core polymerase contains main nucleotide
Lecture 6. Regulation of Protein Synthesis at the Translational Level
 Regulation of Protein Synthesis (6.1) Lecture 6 Regulation of Protein Synthesis at the Translational Level Comparison of EF-Tu-GDP and EF-Tu-GTP conformations EF-Tu-GDP EF-Tu-GTP Next: Comparison of GDP
Regulation of Protein Synthesis (6.1) Lecture 6 Regulation of Protein Synthesis at the Translational Level Comparison of EF-Tu-GDP and EF-Tu-GTP conformations EF-Tu-GDP EF-Tu-GTP Next: Comparison of GDP
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA
 CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
Chapter 17: From Gene to Protein
 AP Biology Reading Guide Fred and Theresa Holtzclaw Julia Keller 12d Chapter 17: From Gene to Protein 1. What is gene expression? Gene expression is the process by which DNA directs the synthesis of proteins
AP Biology Reading Guide Fred and Theresa Holtzclaw Julia Keller 12d Chapter 17: From Gene to Protein 1. What is gene expression? Gene expression is the process by which DNA directs the synthesis of proteins
Chem 465 Biochemistry II
 Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Formation of the ribosomal initiation complex for bacterial protein synthesis does not require: A) EF-Tu. B) formylmethionyl
Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Formation of the ribosomal initiation complex for bacterial protein synthesis does not require: A) EF-Tu. B) formylmethionyl
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College
 Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
DNA (genetic information in genes) RNA (copies of genes) proteins (functional molecules) directionality along the backbone 5 (phosphate) to 3 (OH)
 DNA, RNA, replication, translation, and transcription Overview Recall the central dogma of biology: DNA (genetic information in genes) RNA (copies of genes) proteins (functional molecules) DNA structure
DNA, RNA, replication, translation, and transcription Overview Recall the central dogma of biology: DNA (genetic information in genes) RNA (copies of genes) proteins (functional molecules) DNA structure
Specific problems. The genetic code. The genetic code. Adaptor molecules match amino acids to mrna codons
 Tutorial II Gene expression: mrna translation and protein synthesis Piergiorgio Percipalle, PhD Program Control of gene transcription and RNA processing mrna translation and protein synthesis KAROLINSKA
Tutorial II Gene expression: mrna translation and protein synthesis Piergiorgio Percipalle, PhD Program Control of gene transcription and RNA processing mrna translation and protein synthesis KAROLINSKA
MUTATION, DNA REPAIR AND CANCER
 MUTATION, DNA REPAIR AND CANCER 1 Mutation A heritable change in the genetic material Essential to the continuity of life Source of variation for natural selection New mutations are more likely to be harmful
MUTATION, DNA REPAIR AND CANCER 1 Mutation A heritable change in the genetic material Essential to the continuity of life Source of variation for natural selection New mutations are more likely to be harmful
Viruses. Viral components: Capsid. Chapter 10: Viruses. Viral components: Nucleic Acid. Viral components: Envelope
 Viruses Chapter 10: Viruses Lecture Exam #3 Wednesday, November 22 nd (This lecture WILL be on Exam #3) Dr. Amy Rogers Office Hours: MW 9-10 AM Too small to see with a light microscope Visible with electron
Viruses Chapter 10: Viruses Lecture Exam #3 Wednesday, November 22 nd (This lecture WILL be on Exam #3) Dr. Amy Rogers Office Hours: MW 9-10 AM Too small to see with a light microscope Visible with electron
The Nucleus: DNA, Chromatin And Chromosomes
 The Nucleus: DNA, Chromatin And Chromosomes Professor Alfred Cuschieri Department of Anatomy, University of Malta. Objectives By the end of this unit the student should be able to: 1. List the major structural
The Nucleus: DNA, Chromatin And Chromosomes Professor Alfred Cuschieri Department of Anatomy, University of Malta. Objectives By the end of this unit the student should be able to: 1. List the major structural
a. Ribosomal RNA rrna a type ofrna that combines with proteins to form Ribosomes on which polypeptide chains of proteins are assembled
 Biology 101 Chapter 14 Name: Fill-in-the-Blanks Which base follows the next in a strand of DNA is referred to. as the base (1) Sequence. The region of DNA that calls for the assembly of specific amino
Biology 101 Chapter 14 Name: Fill-in-the-Blanks Which base follows the next in a strand of DNA is referred to. as the base (1) Sequence. The region of DNA that calls for the assembly of specific amino
2006 7.012 Problem Set 3 KEY
 2006 7.012 Problem Set 3 KEY Due before 5 PM on FRIDAY, October 13, 2006. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Which reaction is catalyzed by each
2006 7.012 Problem Set 3 KEY Due before 5 PM on FRIDAY, October 13, 2006. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Which reaction is catalyzed by each
Translation Study Guide
 Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Transfection-Transfer of non-viral genetic material into eukaryotic cells. Infection/ Transduction- Transfer of viral genetic material into cells.
 Transfection Key words: Transient transfection, Stable transfection, transfection methods, vector, plasmid, origin of replication, reporter gene/ protein, cloning site, promoter and enhancer, signal peptide,
Transfection Key words: Transient transfection, Stable transfection, transfection methods, vector, plasmid, origin of replication, reporter gene/ protein, cloning site, promoter and enhancer, signal peptide,
trna Processing and Modification
 trna Processing and Modification RNA POL III - TRANSCRIPTS 5S RNA, trna, repetitive Sequenzen (Alu-typ), versch. kleine stabile RNAs (7SL - RNA vom signal recognition particle (SRP)), U6 RNA 5S RNA nicht
trna Processing and Modification RNA POL III - TRANSCRIPTS 5S RNA, trna, repetitive Sequenzen (Alu-typ), versch. kleine stabile RNAs (7SL - RNA vom signal recognition particle (SRP)), U6 RNA 5S RNA nicht
Lecture 26: Overview of deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) structure
 Lecture 26: Overview of deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) structure Nucleic acids play an important role in the storage and expression of genetic information. They are divided into
Lecture 26: Overview of deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) structure Nucleic acids play an important role in the storage and expression of genetic information. They are divided into
2. True or False? The sequence of nucleotides in the human genome is 90.9% identical from one person to the next. False (it s 99.
 1. True or False? A typical chromosome can contain several hundred to several thousand genes, arranged in linear order along the DNA molecule present in the chromosome. True 2. True or False? The sequence
1. True or False? A typical chromosome can contain several hundred to several thousand genes, arranged in linear order along the DNA molecule present in the chromosome. True 2. True or False? The sequence
Given these characteristics of life, which of the following objects is considered a living organism? W. X. Y. Z.
 Cell Structure and Organization 1. All living things must possess certain characteristics. They are all composed of one or more cells. They can grow, reproduce, and pass their genes on to their offspring.
Cell Structure and Organization 1. All living things must possess certain characteristics. They are all composed of one or more cells. They can grow, reproduce, and pass their genes on to their offspring.
The sequence of bases on the mrna is a code that determines the sequence of amino acids in the polypeptide being synthesized:
 Module 3F Protein Synthesis So far in this unit, we have examined: How genes are transmitted from one generation to the next Where genes are located What genes are made of How genes are replicated How
Module 3F Protein Synthesis So far in this unit, we have examined: How genes are transmitted from one generation to the next Where genes are located What genes are made of How genes are replicated How
Coding sequence the sequence of nucleotide bases on the DNA that are transcribed into RNA which are in turn translated into protein
 Assignment 3 Michele Owens Vocabulary Gene: A sequence of DNA that instructs a cell to produce a particular protein Promoter a control sequence near the start of a gene Coding sequence the sequence of
Assignment 3 Michele Owens Vocabulary Gene: A sequence of DNA that instructs a cell to produce a particular protein Promoter a control sequence near the start of a gene Coding sequence the sequence of
13.2 Ribosomes & Protein Synthesis
 13.2 Ribosomes & Protein Synthesis Introduction: *A specific sequence of bases in DNA carries the directions for forming a polypeptide, a chain of amino acids (there are 20 different types of amino acid).
13.2 Ribosomes & Protein Synthesis Introduction: *A specific sequence of bases in DNA carries the directions for forming a polypeptide, a chain of amino acids (there are 20 different types of amino acid).
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
AP Biology TEST #5 - Chapters 11-14, 16 - REVIEW SHEET
 NAME: AP Biology TEST #5 - Chapters 11-14, 16 - REVIEW SHEET 1. Griffith's experiments showing the transformation of R strain pneumococcus bacteria to S strain pneumococcus bacteria in the presence of
NAME: AP Biology TEST #5 - Chapters 11-14, 16 - REVIEW SHEET 1. Griffith's experiments showing the transformation of R strain pneumococcus bacteria to S strain pneumococcus bacteria in the presence of
From DNA to Protein
 Nucleus Control center of the cell contains the genetic library encoded in the sequences of nucleotides in molecules of DNA code for the amino acid sequences of all proteins determines which specific proteins
Nucleus Control center of the cell contains the genetic library encoded in the sequences of nucleotides in molecules of DNA code for the amino acid sequences of all proteins determines which specific proteins
Translation. Translation: Assembly of polypeptides on a ribosome
 Translation Translation: Assembly of polypeptides on a ribosome Living cells devote more energy to the synthesis of proteins than to any other aspect of metabolism. About a third of the dry mass of a cell
Translation Translation: Assembly of polypeptides on a ribosome Living cells devote more energy to the synthesis of proteins than to any other aspect of metabolism. About a third of the dry mass of a cell
Copyright 1999 2003 by Mark Brandt, Ph.D.
 Central dogma of molecular biology The term central dogma of molecular biology is patterned after religious terminology. owever, it refers to a process that is subject to the changes in understanding that
Central dogma of molecular biology The term central dogma of molecular biology is patterned after religious terminology. owever, it refers to a process that is subject to the changes in understanding that
Recombinant DNA Technology
 Recombinant DNA Technology Dates in the Development of Gene Cloning: 1965 - plasmids 1967 - ligase 1970 - restriction endonucleases 1972 - first experiments in gene splicing 1974 - worldwide moratorium
Recombinant DNA Technology Dates in the Development of Gene Cloning: 1965 - plasmids 1967 - ligase 1970 - restriction endonucleases 1972 - first experiments in gene splicing 1974 - worldwide moratorium
Nucleotides and Nucleic Acids
 Nucleotides and Nucleic Acids Brief History 1 1869 - Miescher Isolated nuclein from soiled bandages 1902 - Garrod Studied rare genetic disorder: Alkaptonuria; concluded that specific gene is associated
Nucleotides and Nucleic Acids Brief History 1 1869 - Miescher Isolated nuclein from soiled bandages 1902 - Garrod Studied rare genetic disorder: Alkaptonuria; concluded that specific gene is associated
Cellular Respiration Worksheet 1. 1. What are the 3 phases of the cellular respiration process? Glycolysis, Krebs Cycle, Electron Transport Chain.
 Cellular Respiration Worksheet 1 1. What are the 3 phases of the cellular respiration process? Glycolysis, Krebs Cycle, Electron Transport Chain. 2. Where in the cell does the glycolysis part of cellular
Cellular Respiration Worksheet 1 1. What are the 3 phases of the cellular respiration process? Glycolysis, Krebs Cycle, Electron Transport Chain. 2. Where in the cell does the glycolysis part of cellular
1.5 page 3 DNA Replication S. Preston 1
 AS Unit 1: Basic Biochemistry and Cell Organisation Name: Date: Topic 1.5 Nucleic Acids and their functions Page 3 l. DNA Replication 1. Go through PowerPoint 2. Read notes p2 and then watch the animation
AS Unit 1: Basic Biochemistry and Cell Organisation Name: Date: Topic 1.5 Nucleic Acids and their functions Page 3 l. DNA Replication 1. Go through PowerPoint 2. Read notes p2 and then watch the animation
Outline. interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work?
 Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Basic Concepts of DNA, Proteins, Genes and Genomes
 Basic Concepts of DNA, Proteins, Genes and Genomes Kun-Mao Chao 1,2,3 1 Graduate Institute of Biomedical Electronics and Bioinformatics 2 Department of Computer Science and Information Engineering 3 Graduate
Basic Concepts of DNA, Proteins, Genes and Genomes Kun-Mao Chao 1,2,3 1 Graduate Institute of Biomedical Electronics and Bioinformatics 2 Department of Computer Science and Information Engineering 3 Graduate
Thymine = orange Adenine = dark green Guanine = purple Cytosine = yellow Uracil = brown
 1 DNA Coloring - Transcription & Translation Transcription RNA, Ribonucleic Acid is very similar to DNA. RNA normally exists as a single strand (and not the double stranded double helix of DNA). It contains
1 DNA Coloring - Transcription & Translation Transcription RNA, Ribonucleic Acid is very similar to DNA. RNA normally exists as a single strand (and not the double stranded double helix of DNA). It contains
RNA and Protein Synthesis
 Name lass Date RN and Protein Synthesis Information and Heredity Q: How does information fl ow from DN to RN to direct the synthesis of proteins? 13.1 What is RN? WHT I KNOW SMPLE NSWER: RN is a nucleic
Name lass Date RN and Protein Synthesis Information and Heredity Q: How does information fl ow from DN to RN to direct the synthesis of proteins? 13.1 What is RN? WHT I KNOW SMPLE NSWER: RN is a nucleic
DNA, RNA, Protein synthesis, and Mutations. Chapters 12-13.3
 DNA, RNA, Protein synthesis, and Mutations Chapters 12-13.3 1A)Identify the components of DNA and explain its role in heredity. DNA s Role in heredity: Contains the genetic information of a cell that can
DNA, RNA, Protein synthesis, and Mutations Chapters 12-13.3 1A)Identify the components of DNA and explain its role in heredity. DNA s Role in heredity: Contains the genetic information of a cell that can
Lecture 8. Protein Trafficking/Targeting. Protein targeting is necessary for proteins that are destined to work outside the cytoplasm.
 Protein Trafficking/Targeting (8.1) Lecture 8 Protein Trafficking/Targeting Protein targeting is necessary for proteins that are destined to work outside the cytoplasm. Protein targeting is more complex
Protein Trafficking/Targeting (8.1) Lecture 8 Protein Trafficking/Targeting Protein targeting is necessary for proteins that are destined to work outside the cytoplasm. Protein targeting is more complex
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
Control of Gene Expression
 Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
The Steps. 1. Transcription. 2. Transferal. 3. Translation
 Protein Synthesis Protein synthesis is simply the "making of proteins." Although the term itself is easy to understand, the multiple steps that a cell in a plant or animal must go through are not. In order
Protein Synthesis Protein synthesis is simply the "making of proteins." Although the term itself is easy to understand, the multiple steps that a cell in a plant or animal must go through are not. In order
Basic attributes of genetic processes (replication, transcription, translation)
 411-3 2008 Lecture notes I. First general topic in the course will be mutation (in broadest sense, any change to an organismʼs genetic material). Intimately intertwined with this is the process of DNA
411-3 2008 Lecture notes I. First general topic in the course will be mutation (in broadest sense, any change to an organismʼs genetic material). Intimately intertwined with this is the process of DNA
Activity 7.21 Transcription factors
 Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
Bio 102 Practice Problems Recombinant DNA and Biotechnology
 Bio 102 Practice Problems Recombinant DNA and Biotechnology Multiple choice: Unless otherwise directed, circle the one best answer: 1. Which of the following DNA sequences could be the recognition site
Bio 102 Practice Problems Recombinant DNA and Biotechnology Multiple choice: Unless otherwise directed, circle the one best answer: 1. Which of the following DNA sequences could be the recognition site
Answer: 2. Uracil. Answer: 2. hydrogen bonds. Adenine, Cytosine and Guanine are found in both RNA and DNA.
 Answer: 2. Uracil Adenine, Cytosine and Guanine are found in both RNA and DNA. Thymine is found only in DNA; Uracil takes its (Thymine) place in RNA molecules. Answer: 2. hydrogen bonds The complementary
Answer: 2. Uracil Adenine, Cytosine and Guanine are found in both RNA and DNA. Thymine is found only in DNA; Uracil takes its (Thymine) place in RNA molecules. Answer: 2. hydrogen bonds The complementary
CHAPTER 40 The Mechanism of Protein Synthesis
 CHAPTER 40 The Mechanism of Protein Synthesis Problems: 2,3,6,7,9,13,14,15,18,19,20 Initiation: Locating the start codon. Elongation: Reading the codons (5 3 ) and synthesizing protein amino carboxyl.
CHAPTER 40 The Mechanism of Protein Synthesis Problems: 2,3,6,7,9,13,14,15,18,19,20 Initiation: Locating the start codon. Elongation: Reading the codons (5 3 ) and synthesizing protein amino carboxyl.
Cells & Cell Organelles
 Cells & Cell Organelles The Building Blocks of Life H Biology Types of cells bacteria cells Prokaryote - no organelles Eukaryotes - organelles animal cells plant cells Cell size comparison Animal cell
Cells & Cell Organelles The Building Blocks of Life H Biology Types of cells bacteria cells Prokaryote - no organelles Eukaryotes - organelles animal cells plant cells Cell size comparison Animal cell
PRACTICE TEST QUESTIONS
 PART A: MULTIPLE CHOICE QUESTIONS PRACTICE TEST QUESTIONS DNA & PROTEIN SYNTHESIS B 1. One of the functions of DNA is to A. secrete vacuoles. B. make copies of itself. C. join amino acids to each other.
PART A: MULTIPLE CHOICE QUESTIONS PRACTICE TEST QUESTIONS DNA & PROTEIN SYNTHESIS B 1. One of the functions of DNA is to A. secrete vacuoles. B. make copies of itself. C. join amino acids to each other.
Micro RNAs: potentielle Biomarker für das. Blutspenderscreening
 Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
CHAPTER 30: PROTEIN SYNTHESIS
 CHAPTER 30: PROTEIN SYNTHESIS (Translation) Translation: mrna protein LECTURE TOPICS Complexity, stages, rate, accuracy Amino acid activation [trna charging] trnas and translating the Genetic Code - Amino
CHAPTER 30: PROTEIN SYNTHESIS (Translation) Translation: mrna protein LECTURE TOPICS Complexity, stages, rate, accuracy Amino acid activation [trna charging] trnas and translating the Genetic Code - Amino
mrna EDITING Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A. Copyright 2005
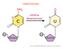 mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
To be able to describe polypeptide synthesis including transcription and splicing
 Thursday 8th March COPY LO: To be able to describe polypeptide synthesis including transcription and splicing Starter Explain the difference between transcription and translation BATS Describe and explain
Thursday 8th March COPY LO: To be able to describe polypeptide synthesis including transcription and splicing Starter Explain the difference between transcription and translation BATS Describe and explain
Gene Switches Teacher Information
 STO-143 Gene Switches Teacher Information Summary Kit contains How do bacteria turn on and turn off genes? Students model the action of the lac operon that regulates the expression of genes essential for
STO-143 Gene Switches Teacher Information Summary Kit contains How do bacteria turn on and turn off genes? Students model the action of the lac operon that regulates the expression of genes essential for
Recombinant DNA and Biotechnology
 Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Gene Regulation -- The Lac Operon
 Gene Regulation -- The Lac Operon Specific proteins are present in different tissues and some appear only at certain times during development. All cells of a higher organism have the full set of genes:
Gene Regulation -- The Lac Operon Specific proteins are present in different tissues and some appear only at certain times during development. All cells of a higher organism have the full set of genes:
Genetics Module B, Anchor 3
 Genetics Module B, Anchor 3 Key Concepts: - An individual s characteristics are determines by factors that are passed from one parental generation to the next. - During gamete formation, the alleles for
Genetics Module B, Anchor 3 Key Concepts: - An individual s characteristics are determines by factors that are passed from one parental generation to the next. - During gamete formation, the alleles for
Viral Infection: Receptors
 Viral Infection: Receptors Receptors: Identification of receptors has come from expressing the gene for the receptor in a cell to which a virus does not normally bind -OR- By blocking virus attachment
Viral Infection: Receptors Receptors: Identification of receptors has come from expressing the gene for the receptor in a cell to which a virus does not normally bind -OR- By blocking virus attachment
CCR Biology - Chapter 8 Practice Test - Summer 2012
 Name: Class: Date: CCR Biology - Chapter 8 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. What did Hershey and Chase know
Name: Class: Date: CCR Biology - Chapter 8 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. What did Hershey and Chase know
Eukaryotes. www.njctl.org PSI Biology Eukaryotes & Gene Expression
 Eukaryotes The Eukaryotic Cell Classwork 1. Identify two characteristics that are shared by all cells. 2. Suppose you are investigating a cell that contains a nucleus. Would you categorize this cell as
Eukaryotes The Eukaryotic Cell Classwork 1. Identify two characteristics that are shared by all cells. 2. Suppose you are investigating a cell that contains a nucleus. Would you categorize this cell as
Protein Synthesis. Page 41 Page 44 Page 47 Page 42 Page 45 Page 48 Page 43 Page 46 Page 49. Page 41. DNA RNA Protein. Vocabulary
 Protein Synthesis Vocabulary Transcription Translation Translocation Chromosomal mutation Deoxyribonucleic acid Frame shift mutation Gene expression Mutation Point mutation Page 41 Page 41 Page 44 Page
Protein Synthesis Vocabulary Transcription Translation Translocation Chromosomal mutation Deoxyribonucleic acid Frame shift mutation Gene expression Mutation Point mutation Page 41 Page 41 Page 44 Page
DNA. Discovery of the DNA double helix
 DNA Replication DNA Discovery of the DNA double helix A. 1950 s B. Rosalind Franklin - X-ray photo of DNA. C. Watson and Crick - described the DNA molecule from Franklin s X-ray. What is DNA? Question:
DNA Replication DNA Discovery of the DNA double helix A. 1950 s B. Rosalind Franklin - X-ray photo of DNA. C. Watson and Crick - described the DNA molecule from Franklin s X-ray. What is DNA? Question:
Biological Sciences Initiative. Human Genome
 Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Chapter 5: Organization and Expression of Immunoglobulin Genes
 Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
RNA Structure and folding
 RNA Structure and folding Overview: The main functional biomolecules in cells are polymers DNA, RNA and proteins For RNA and Proteins, the specific sequence of the polymer dictates its final structure
RNA Structure and folding Overview: The main functional biomolecules in cells are polymers DNA, RNA and proteins For RNA and Proteins, the specific sequence of the polymer dictates its final structure
Lecture 4. Polypeptide Synthesis Overview
 Initiation of Protein Synthesis (4.1) Lecture 4 Polypeptide Synthesis Overview Polypeptide synthesis proceeds sequentially from N Terminus to C terminus. Amino acids are not pre-positioned on a template.
Initiation of Protein Synthesis (4.1) Lecture 4 Polypeptide Synthesis Overview Polypeptide synthesis proceeds sequentially from N Terminus to C terminus. Amino acids are not pre-positioned on a template.
Chapter 11: Molecular Structure of DNA and RNA
 Chapter 11: Molecular Structure of DNA and RNA Student Learning Objectives Upon completion of this chapter you should be able to: 1. Understand the major experiments that led to the discovery of DNA as
Chapter 11: Molecular Structure of DNA and RNA Student Learning Objectives Upon completion of this chapter you should be able to: 1. Understand the major experiments that led to the discovery of DNA as
3120-1 - Page 1. Name:
 Name: 1) Which series is arranged in correct order according to decreasing size of structures? A) DNA, nucleus, chromosome, nucleotide, nitrogenous base B) chromosome, nucleus, nitrogenous base, nucleotide,
Name: 1) Which series is arranged in correct order according to decreasing size of structures? A) DNA, nucleus, chromosome, nucleotide, nitrogenous base B) chromosome, nucleus, nitrogenous base, nucleotide,
