mirnaselect pep-mir Cloning and Expression Vector
|
|
|
- Daniel Farmer
- 10 years ago
- Views:
Transcription
1 Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect pep-mir Cloning and Expression Vector (Part No. MIR-EXP): One tube 2. mirnaselect pep-mir Null Control Vector (Part No. MIR-NULL): One tube Background MicroRNAs (mirnas) are nucleotide RNA molecules that regulate the stability or translational efficiency of target mrnas. These regulatory RNAs function by acting as sequence-specific guides which recruit a large protein complex known as the RNA-induced silencing complex (RISC) to target mrnas which are subsequently silenced. Diverse functions have been attributed to mirnas including the regulation of cellular differentiation, proliferation, and apoptosis. Moreover, significant evidence has accumulated implicating a fundamental role for mirnas in the development of cancer. mirnas are initially transcribed as long precursor transcripts known as primary micrornas (primirnas). Within these transcripts, the mature mirna sequences are found in ~60 80 nucleotide hairpin structures. Mature mirnas are generated from pri-mirnas by sequential processing (Figure 1). PrimiRNAs are initially recognized in the nucleus by the microprocessor complex which includes as core components the RNase-III enzyme Drosha and its obligate partner DGCR8. This complex excises the hairpin structure containing the mature mirna sequence. The liberated hairpins, referred to as precursor mirnas (pre-mirnas), are recognized by the nuclear export factor exportin 5 which transports them to the cytoplasm. There, the RNase-III enzyme Dicer performs a second cleavage to generate a doublestranded nucleotide RNA molecule. The RISC then associates with this RNA duplex and unwinds it. Generally, only one strand is stably incorporated into the RISC; the other is discarded and rapidly degraded. mirnas guide the RISC to target messages that are subsequently cleaved or translationally silenced. Synthetic mirna molecules based on predicted mature mirna sequence are sometimes used. Despite their optimized design criteria, synthetic mirnas underscore the importance of primary mirna in its native expressed form. The primary mirna contains critical biological components involved in mature mirna expression and cellular processing, and is often processed into several mature mirna molecules. Cell Biolabs mirnaselect Cloning and Expression Vector is designed to clone and express an individual mirna precursor in its native context while preserving putative hairpin structures to ensure biologically relevant interactions with endogenous processing machinery and regulatory partners, leading to properly cleaved micrornas. Individual mirna precursor from any species can be cloned between BamHI and Nhe I sites (Figure 2).
2 Figure 1. mirna Biogenesis and function The mirnaselect pep-mir cloning and expression vector contains the following features: mirna Processing mirna stem loop precursor in its native context is cloned between BamHI and Nhe I sites. To preserve the putative hairpin structure and proper endogenous processing, mirna stem loop sequence is flanked by its native intron sequence. EF-1α Promoter - ensures a high level of expression in mammalian cells Puromycin resistance marker - to monitor cells positive for expression and stable selection SV40 polyadenylation signal - enables efficient termination of transcription puc origin - for high copy replication and maintenance of the plasmid in E. coli Ampicillin resistance gene - for selection in E. coli Figure 2. Schematic representation of pep-mir cloning and expression vector.
3 Cell Biolabs mirnaselect pep-mir Null Control Vector is similar to the pep-mir vector except it does not contain any mirna precursor or the BamHI and NheI cloning sites (Figure 3). Figure 3. Schematic representation of pep-mir null control vector. mirna precursor cloning sites: BamHI-1.1 kb -NheI 1 tcgaggatcc ctttaagtct tctgaggagc atggacctgc tagactgcaa gctgctgcat 61 tctgaggagc agggacctgc taaactgcaa gctgccgcag gcggagattt ttgcctgttt 121 ggtttatttt gttaccactg agtccccagc tctgagaaca gagcgaggca tgtgatcgat 181 gcatcgtaac tgttgagcaa atgaatgctg agctgtgatg gcaccgacgt gtccggggag 241 aggacgcacc cggctgtgtg gacatgtgcc cagggcccag gacagcgcca cggaagagga 301 cacacccggc tgtgtggaca tgtgcccagg gcccgggaca gcgccacgga agaggacgca 361 cccggctgtg tgcacatgtg cccagggccc gggacagcgc cacggaagag gacgcacccg 421 gctgtgtgca catgtgccca gggcccggga cagcgccacg gaagaggacg cacccggctg 481 tgtgcacatg tgcccagggc ccgggacagc gccacggaag aggacgcacc cggctgtgtg 541 gacatgtgcc cagggcccgg gacagcgcca cggaagagga cgcacaggac agcgccacgg 601 aagaggacgc acccggctgt gtgcacatgt gcccagggcc cgggacagcg ccacggaaga 661 ggacgcaccc ggctgtgtgc acatgtgccc agggcccggg acagcgccat ggaagaggac 721 gcacccggct gtgtgcacat gtgcccaggg cccgggacag cgccacggaa gaggacgcac 781 ccggctgtgt gcacatgtgc ccagggcccg ggacagcgcc acggaagagg atgcacccgg 841 ctgtgtggac atgtgcccag ggcccgggac agcgccacgg aagaggacgc acccggctgt 901 gtggacatgt gcccagggcc cgggacagcg ccacggaaga ggacgcaccc ggctgtgtgg 961 acatgtgccc agggcccggg acagcgccac ggaagaggac gcacaggaca gcgccacgga 1021 agaggacgca cccggctgtg tggcagggga ggttcccaag agcaggctca gggctctggg 1081 gtctgagtga gggctctcca gccaggctag ctcga Note: pep-mir Cloning and Expression Vector should NOT be used as a transfection control vector, because the vector itself contains 1.1 kb of mir-941 precursor sequence between the BamHI and NheI sites. For transfection control, we recommend pep-mir Null Control Vector. mirna Precursor Cloning All of our premade human mirna precursor clones are based on the following design, and the resulting overexpression of the mature mirna is confirmed by Northern blot. Here we use human let-7a-2 mirna as an example:
4 1. Download desired mirna stem loop sequence from Sanger s mirna database: Homo sapiens let-7a-2 stem-loop structure uu g u uagaa ua a agg gag uag agguuguauaguu u c u c ucc uuc auc uccgacaugucaa a g a -u g c --uag gg a Homo sapiens let-7a-2 stem-loop sequence AGGUUGAGGUAGUAGGUUGUAUAGUUUAGAAUUACAUCAAGGGAGAUAACUGUACAGCCUCCUAGCUUUCCU 2. Blast search mirna stem loop sequence: > ref NT_ Hs11_34054 Homo sapiens chromosome 11 genomic contig, reference assembly Length= Query 1 AGGTTGAGGTAGTAGGTTGTATAGTTTAGAATTACATCAAGGGAGATAACTGTACAGCCT 60 Sbjct AGGTTGAGGTAGTAGGTTGTATAGTTTAGAATTACATCAAGGGAGATAACTGTACAGCCT Query 61 CCTAGCTTTCCT 72 Sbjct CCTAGCTTTCCT PCR and Cloning: 1) Add 100 base native flank sequence to both upstream and downstream of the mirna stem loop. Human let-7a-2 mirna precursor sequence including the 100 base flank sequences on both ends of the stem loop: let-7a-2 stem-loop sequence is underlined. GCCCAAATAGGTGACAGCACGATGAATCATTATAAGACTAACTTGTAATTTCCCTGCTTAAGAAATG GTAGTTTTCCAGCCATTGTGACTGCATGCTCCCAGGTTGAGGTAGTAGGTTGTATAGTTTAGAATTA CATCAAGGGAGATAACTGTACAGCCTCCTAGCTTTCCTTGGGTCTTGCACTAAACAACATGGTGAGA ACGATCATGATTCCTCCAGGCCTTTTCTCCCTATGAAAGGTAAGATTGGGTACGATTATTTTATGGT ATTT 2) Design PCR primer including BamHI site at forward primer with four extra bases and NheI site at reverse primer. Forward PCR Primer: tcga-ggatcc (BamHI)-21 nt Reverse PCR Primer: tcga-gctagc (NheI)-21 nt For human let-7a-2 mirna precursor: Forward PCR Primer: tcga-ggatcc-gcccaaataggtgacagcacg Reverse PCR Primer: tcga-gctagc-aaataccataaaataatcgta
5 3) PCR the mirna precursor from genomic DNA and clone into the BamHI/NheI sites of the expression vector. PCR Product of let-7a-2 precursor: let-7a-2 stem-loop sequence is underlined. 1 tcgaggatcc gcccaaatag gtgacagcac gatgaatcat tataagacta acttgtaatt 61 tccctgctta agaaatggta gttttccagc cattgtgact gcatgctccc aggttgaggt 121 agtaggttgt atagtttaga attacatcaa gggagataac tgtacagcct cctagctttc 181 cttgggtctt gcactaaaca acatggtgag aacgatcatg attcctccag gccttttctc 241 cctatgaaag gtaagattgg gtacgattat tttatggtat ttgctagctc ga 4) Validate the insert by DNA sequencing. Forward Sequencing Primer: tcctcagccg tcgcttcatg Reverse Sequencing Primer: gtgtggggaa actccatcgc Methods 1) Bacterial culture: the mirna cloning and expression vector and the null control vector are provided as bacterial glycerol stocks. Individual colonies can be obtained by culturing in an LBampicillin plate. 2) Plasmid isolation: we recommend EndoFree Plasmid Kits (QIAGEN). 3) Transfection into target cells: we recommend Lipofectamine 2000 (Invitrogen). 4) Stable selection: 48 hrs post-transfection, select stable clones in 1-10 µg/ml Puromycincontaining medium. References 1. microrna sequences listed in Sanger s mirbase ( 2. John, B., C. Sander and D. S. Marks (2006) Methods Mol Biol 342: Johnson, S. M., H. Grosshans, J. Shingara, M. Byrom, R. Jarvis, A. Cheng, E. Labourier, K. L. Reinert, D. Brown and F. J. Slack (2005) Cell 120: Kim, V. N. (2005) Mol Cells 19: Lee, R. C., R. L. Feinbaum and V. Ambros (1993) Cell 75: Lee, Y., K. Jeon, J. T. Lee, S. Kim and V. N. Kim (2002) Embo J 21: Yi, R., Y. Qin, I. G. Macara and B. R. Cullen (2003) Genes Dev 17: Warranty These products are warranted to perform as described in their labeling and in Cell Biolabs literature when used in accordance with their instructions. THERE ARE NO WARRANTIES THAT EXTEND BEYOND THIS EXPRESSED WARRANTY AND CELL BIOLABS DISCLAIMS ANY IMPLIED WARRANTY OF MERCHANTABILITY OR WARRANTY OF FITNESS FOR PARTICULAR PURPOSE. CELL BIOLABS s sole obligation and purchaser s exclusive remedy for breach of this warranty shall be, at the option of CELL BIOLABS, to repair or replace the products. In no event shall CELL BIOLABS be liable for any proximate, incidental or consequential damages in connection with the products. This product is for RESEARCH USE ONLY; not for use in diagnostic procedures. Contact Information Cell Biolabs, Inc.
6 7758 Arjons Drive San Diego, CA Worldwide: USA Toll-Free: CBL : Cell Biolabs, Inc. - All rights reserved. No part of these works may be reproduced in any form without permissions in writing.
mirnaselect pegp-mir Cloning and Expression Vector
 Product Data Sheet mirnaselect pegp-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-GP-C STORAGE: -80ºC QUANTITY: 100 µl of bacterial glycerol stock Components 1. mirnaselect pegp-mir Cloning
Product Data Sheet mirnaselect pegp-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-GP-C STORAGE: -80ºC QUANTITY: 100 µl of bacterial glycerol stock Components 1. mirnaselect pegp-mir Cloning
RAPAd mirna Adenoviral Expression System Catalog #: VPK-253
 DATENBLATT RAPAd mirna Adenoviral Expression System Catalog #: VPK-253 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction MicroRNAs (mirnas) are 18 24 nucleotide RNA molecules that
DATENBLATT RAPAd mirna Adenoviral Expression System Catalog #: VPK-253 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction MicroRNAs (mirnas) are 18 24 nucleotide RNA molecules that
Micro RNAs: potentielle Biomarker für das. Blutspenderscreening
 Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation
 OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
Transfection-Transfer of non-viral genetic material into eukaryotic cells. Infection/ Transduction- Transfer of viral genetic material into cells.
 Transfection Key words: Transient transfection, Stable transfection, transfection methods, vector, plasmid, origin of replication, reporter gene/ protein, cloning site, promoter and enhancer, signal peptide,
Transfection Key words: Transient transfection, Stable transfection, transfection methods, vector, plasmid, origin of replication, reporter gene/ protein, cloning site, promoter and enhancer, signal peptide,
Outline. interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work?
 Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Mir-X mirna First-Strand Synthesis Kit User Manual
 User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
岑 祥 股 份 有 限 公 司 技 術 專 員 費 軫 尹 20100803
 技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
Description: Molecular Biology Services and DNA Sequencing
 Description: Molecular Biology s and DNA Sequencing DNA Sequencing s Single Pass Sequencing Sequence data only, for plasmids or PCR products Plasmid DNA or PCR products Plasmid DNA: 20 100 ng/μl PCR Product:
Description: Molecular Biology s and DNA Sequencing DNA Sequencing s Single Pass Sequencing Sequence data only, for plasmids or PCR products Plasmid DNA or PCR products Plasmid DNA: 20 100 ng/μl PCR Product:
restriction enzymes 350 Home R. Ward: Spring 2001
 restriction enzymes 350 Home Restriction Enzymes (endonucleases): molecular scissors that cut DNA Properties of widely used Type II restriction enzymes: recognize a single sequence of bases in dsdna, usually
restriction enzymes 350 Home Restriction Enzymes (endonucleases): molecular scissors that cut DNA Properties of widely used Type II restriction enzymes: recognize a single sequence of bases in dsdna, usually
Dicer Substrate RNAi Design
 INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
Outline. MicroRNA Bioinformatics. microrna biogenesis. short non-coding RNAs not considered in this lecture. ! Introduction
 Outline MicroRNA Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology (CMB) Karolinska Institutet! Introduction! microrna target site prediction! Useful resources 2 short non-coding RNAs
Outline MicroRNA Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology (CMB) Karolinska Institutet! Introduction! microrna target site prediction! Useful resources 2 short non-coding RNAs
PART 3.3: MicroRNA and Cancer
 BIBM 2010 Tutorial: Epigenomics and Cancer PART 3.3: MicroRNA and Cancer Dec 18, 2010 Sun Kim at Indiana University Outline of Part 3.3 Background on microrna Role of microrna in cancer MicroRNA pathway
BIBM 2010 Tutorial: Epigenomics and Cancer PART 3.3: MicroRNA and Cancer Dec 18, 2010 Sun Kim at Indiana University Outline of Part 3.3 Background on microrna Role of microrna in cancer MicroRNA pathway
Name Class Date. Figure 13 1. 2. Which nucleotide in Figure 13 1 indicates the nucleic acid above is RNA? a. uracil c. cytosine b. guanine d.
 13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
Lentivector-based MicroRNA Precursor Constructs
 Lentivector-based MicroRNA Precursor Constructs Cat. # PMIRHxxxPA-1 User Manual Grow bacterial stock on receipt Make plasmid DNA for experiments (ver. 2008-02-28-001) A limited-use label license covers
Lentivector-based MicroRNA Precursor Constructs Cat. # PMIRHxxxPA-1 User Manual Grow bacterial stock on receipt Make plasmid DNA for experiments (ver. 2008-02-28-001) A limited-use label license covers
MatureBayes: A Probabilistic Algorithm for Identifying the Mature mirna within Novel Precursors
 MatureBayes: A Probabilistic Algorithm for Identifying the Mature mirna within Novel Precursors Katerina Gkirtzou 1,2, Ioannis Tsamardinos 1,2, Panagiotis Tsakalides 1,2, Panayiota Poirazi 3 * 1 Computer
MatureBayes: A Probabilistic Algorithm for Identifying the Mature mirna within Novel Precursors Katerina Gkirtzou 1,2, Ioannis Tsamardinos 1,2, Panagiotis Tsakalides 1,2, Panayiota Poirazi 3 * 1 Computer
EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES
 C GUGAAG EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES MicroRNA-adapted shrna (shrna mir ) for increased, specific and consistent knockdown. MicroRNA PROCESSING PATHWAY UTILIZED FOR shrna mir Developed
C GUGAAG EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES MicroRNA-adapted shrna (shrna mir ) for increased, specific and consistent knockdown. MicroRNA PROCESSING PATHWAY UTILIZED FOR shrna mir Developed
Comprehensive mirna Research Technologies
 Comprehensive mirna Research Technologies Sample & Assay Technologies QIAGEN solutions for advancing microrna research In the last few years, the identification of microrna (mirna) and the recognition
Comprehensive mirna Research Technologies Sample & Assay Technologies QIAGEN solutions for advancing microrna research In the last few years, the identification of microrna (mirna) and the recognition
2013 W. H. Freeman and Company. 26 RNA Metabolism
 2013 W. H. Freeman and Company 26 RNA Metabolism CHAPTER 26 RNA Metabolism Key topics: Transcription: DNA-dependent synthesis of RNA Capping and splicing: RNA processing Overview of RNA Function Ribonucleic
2013 W. H. Freeman and Company 26 RNA Metabolism CHAPTER 26 RNA Metabolism Key topics: Transcription: DNA-dependent synthesis of RNA Capping and splicing: RNA processing Overview of RNA Function Ribonucleic
NIH Mammalian Gene Collection (MGC)
 USER GUIDE NIH Mammalian Gene Collection (MGC) Catalog number FL1002 Revision date 28 November 2011 Publication Part number 25-0610 MAN0000351 For Research Use Only. Not for diagnostic procedures. ii Table
USER GUIDE NIH Mammalian Gene Collection (MGC) Catalog number FL1002 Revision date 28 November 2011 Publication Part number 25-0610 MAN0000351 For Research Use Only. Not for diagnostic procedures. ii Table
pmod2-puro A plasmid containing a synthetic Puromycin resistance gene Catalog # pmod2-puro For research use only Version # 11H29-MM
 pmod2-puro A plasmid containing a synthetic Puromycin resistance gene Catalog # pmod2-puro For research use only Version # 11H29-MM PrOduct information content: - 20 mg of lyophilized pmod2-puro plasmid
pmod2-puro A plasmid containing a synthetic Puromycin resistance gene Catalog # pmod2-puro For research use only Version # 11H29-MM PrOduct information content: - 20 mg of lyophilized pmod2-puro plasmid
STOP. Before using this product, please read the Limited Use License statement below:
 STOP Before using this product, please read the Limited Use License statement below: Important Limited Use License information for pdrive5lucia-rgfap The purchase of the pdrive5lucia-rgfap vector conveys
STOP Before using this product, please read the Limited Use License statement below: Important Limited Use License information for pdrive5lucia-rgfap The purchase of the pdrive5lucia-rgfap vector conveys
Taq98 Hot Start 2X Master Mix
 Taq98 Hot Start 2X Master Mix Optimized for 98C Denaturation Lucigen Corporation 2905 Parmenter St, Middleton, WI 53562 USA Toll Free: (888) 575-9695 (608) 831-9011 FAX: (608) 831-9012 [email protected]
Taq98 Hot Start 2X Master Mix Optimized for 98C Denaturation Lucigen Corporation 2905 Parmenter St, Middleton, WI 53562 USA Toll Free: (888) 575-9695 (608) 831-9011 FAX: (608) 831-9012 [email protected]
Supplementary Figure 1.
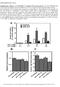 Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
RNAi Shooting the Messenger!
 RNAi Shooting the Messenger! Bronya Keats, Ph.D. Department of Genetics Louisiana State University Health Sciences Center New Orleans Email: [email protected] RNA interference (RNAi) A mechanism by which
RNAi Shooting the Messenger! Bronya Keats, Ph.D. Department of Genetics Louisiana State University Health Sciences Center New Orleans Email: [email protected] RNA interference (RNAi) A mechanism by which
CytoSelect 48-Well Cell Adhesion Assay (ECM Array, Fluorometric Format)
 Product Manual CytoSelect 48-Well Cell Adhesion Assay (ECM Array, Fluorometric Format) Catalog Number CBA-071 CBA-071-5 48 assays 5 x 48 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures
Product Manual CytoSelect 48-Well Cell Adhesion Assay (ECM Array, Fluorometric Format) Catalog Number CBA-071 CBA-071-5 48 assays 5 x 48 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures
RNAi History, Mechanism and Application
 DRUG DISCOVERY AND DEVELOPMENT RNAi History, Mechanism and Application Shuo Gu The History of RNAi About the author: Dr. Shuo Gu is currently a Postdoctoral Scholar at Stanford University, School of Medicine.
DRUG DISCOVERY AND DEVELOPMENT RNAi History, Mechanism and Application Shuo Gu The History of RNAi About the author: Dr. Shuo Gu is currently a Postdoctoral Scholar at Stanford University, School of Medicine.
pcas-guide System Validation in Genome Editing
 pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
Rapid GST Inclusion Body Solubilization and Renaturation Kit
 Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
A role of microrna in the regulation of telomerase? Yuan Ming Yeh, Pei Rong Huang, and Tzu Chien V. Wang
 A role of microrna in the regulation of telomerase? Yuan Ming Yeh, Pei Rong Huang, and Tzu Chien V. Wang Department of Molecular and Cellular Biology, Chang Gung University, Kwei San, Tao Yuan 333, Taiwan
A role of microrna in the regulation of telomerase? Yuan Ming Yeh, Pei Rong Huang, and Tzu Chien V. Wang Department of Molecular and Cellular Biology, Chang Gung University, Kwei San, Tao Yuan 333, Taiwan
DNA Fingerprinting. Unless they are identical twins, individuals have unique DNA
 DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
Functional RNAs; RNA catalysts, mirna,
 Functional RNAs; RNA catalysts, mirna, srna, RNAi... RNAs have many functions rrna (ribosomal RNA) trna (transfer RNA) mrna (Messenger RNA) snrna (including snorna) ) (Small nuclear RNA- splicing) Other
Functional RNAs; RNA catalysts, mirna, srna, RNAi... RNAs have many functions rrna (ribosomal RNA) trna (transfer RNA) mrna (Messenger RNA) snrna (including snorna) ) (Small nuclear RNA- splicing) Other
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
Pure-IP Western Blot Detection Kit
 Product Manual Pure-IP Western Blot Detection Kit Catalog Number PRB-5002 20 blots FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction The technique of immunoprecipitation (IP) is used
Product Manual Pure-IP Western Blot Detection Kit Catalog Number PRB-5002 20 blots FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction The technique of immunoprecipitation (IP) is used
Biotechnology: DNA Technology & Genomics
 Chapter 20. Biotechnology: DNA Technology & Genomics 2003-2004 The BIG Questions How can we use our knowledge of DNA to: diagnose disease or defect? cure disease or defect? change/improve organisms? What
Chapter 20. Biotechnology: DNA Technology & Genomics 2003-2004 The BIG Questions How can we use our knowledge of DNA to: diagnose disease or defect? cure disease or defect? change/improve organisms? What
Transcription and Translation of DNA
 Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
MGC premier Expression-Ready cdna clones TCH1103, TCM1104, TCR1105, TCB1106, TCH1203, TCM1204, TCR1205, TCB1206, TCH1303, TCM1304, TCR1305
 MGC premier Expression-Ready cdna clones TCH1103, TCM1104, TCR1105, TCB1106, TCH1203, TCM1204, TCR1205, TCB1206, TCH1303, TCM1304, TCR1305 The MGC premier Expression-Ready collection has the high quality,
MGC premier Expression-Ready cdna clones TCH1103, TCM1104, TCR1105, TCB1106, TCH1203, TCM1204, TCR1205, TCB1206, TCH1303, TCM1304, TCR1305 The MGC premier Expression-Ready collection has the high quality,
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals
 Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Essentials of Real Time PCR. About Sequence Detection Chemistries
 Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION. Professor Bharat Patel Office: Science 2, 2.36 Email: [email protected].
 Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: [email protected] What is Gene Expression & Gene Regulation? 1. Gene Expression
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: [email protected] What is Gene Expression & Gene Regulation? 1. Gene Expression
Biology for the Nanotechnology Classroom. Erinn Mee [email protected] 10/9/2015
 Biology for the Nanotechnology Classroom Erinn Mee [email protected] 10/9/2015 Size and Scale http://learn.genetics.utah.edu/content/cells/scale/scale.html Visualizing Bacteria Escherichia coli LM 1000x
Biology for the Nanotechnology Classroom Erinn Mee [email protected] 10/9/2015 Size and Scale http://learn.genetics.utah.edu/content/cells/scale/scale.html Visualizing Bacteria Escherichia coli LM 1000x
CCR Biology - Chapter 9 Practice Test - Summer 2012
 Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Global MicroRNA Amplification Kit
 Global MicroRNA Amplification Kit Store kit at -20 C on receipt (ver. 3-060901) A limited-use label license covers this product. By use of this product, you accept the terms and conditions outlined in
Global MicroRNA Amplification Kit Store kit at -20 C on receipt (ver. 3-060901) A limited-use label license covers this product. By use of this product, you accept the terms and conditions outlined in
Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr
 User Manual Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories,
User Manual Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories,
HCS604.03 Exercise 1 Dr. Jones Spring 2005. Recombinant DNA (Molecular Cloning) exercise:
 HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
Recombinant DNA and Biotechnology
 Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Design of conditional gene targeting vectors - a recombineering approach
 Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College
 Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
Recombinant DNA Unit Exam
 Recombinant DNA Unit Exam Question 1 Restriction enzymes are extensively used in molecular biology. Below are the recognition sites of two of these enzymes, BamHI and BclI. a) BamHI, cleaves after the
Recombinant DNA Unit Exam Question 1 Restriction enzymes are extensively used in molecular biology. Below are the recognition sites of two of these enzymes, BamHI and BclI. a) BamHI, cleaves after the
13.4 Gene Regulation and Expression
 13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
2. The number of different kinds of nucleotides present in any DNA molecule is A) four B) six C) two D) three
 Chem 121 Chapter 22. Nucleic Acids 1. Any given nucleotide in a nucleic acid contains A) two bases and a sugar. B) one sugar, two bases and one phosphate. C) two sugars and one phosphate. D) one sugar,
Chem 121 Chapter 22. Nucleic Acids 1. Any given nucleotide in a nucleic acid contains A) two bases and a sugar. B) one sugar, two bases and one phosphate. C) two sugars and one phosphate. D) one sugar,
Relative Quantification of mirna Target mrnas by Real-Time qpcr. 1 Introduction. Gene Expression Application Note No. 4
 Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
CytoSelect LDH Cytotoxicity Assay Kit
 Product Manual CytoSelect LDH Cytotoxicity Assay Kit Catalog Number CBA-241 960 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction The measurement and monitoring of cell cytotoxicity
Product Manual CytoSelect LDH Cytotoxicity Assay Kit Catalog Number CBA-241 960 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction The measurement and monitoring of cell cytotoxicity
RNA: Transcription and Processing
 8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
Considerable evidence now indicates that small noncoding
 MicroRNAs and small interfering RNAs can inhibit mrna expression by similar mechanisms Yan Zeng*, Rui Yi, and Bryan R. Cullen* *Howard Hughes Medical Institute and Department of Molecular Genetics and
MicroRNAs and small interfering RNAs can inhibit mrna expression by similar mechanisms Yan Zeng*, Rui Yi, and Bryan R. Cullen* *Howard Hughes Medical Institute and Department of Molecular Genetics and
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
TOOLS sirna and mirna. User guide
 TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
RNA Interference. - Localization - Labeling - Expression Profiling - In Vivo Delivery - Transfection
 RNA Interference - Localization - Labeling - Expression Profiling - In Vivo Delivery - Transfection Rna interference x RNAi Pathways RNA interference x Tools and Multiplexing RNAi Tools pri-mirna Application
RNA Interference - Localization - Labeling - Expression Profiling - In Vivo Delivery - Transfection Rna interference x RNAi Pathways RNA interference x Tools and Multiplexing RNAi Tools pri-mirna Application
User Manual/Hand book. qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse)
 User Manual/Hand book qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse) Kit Components Cat. No. MA003...Human Whole Genome mirna qpcr Profiling Kit (-20 C) The following components are sufficient
User Manual/Hand book qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse) Kit Components Cat. No. MA003...Human Whole Genome mirna qpcr Profiling Kit (-20 C) The following components are sufficient
Lezioni Dipartimento di Oncologia Farmacologia Molecolare. RNA interference. Giovanna Damia 29 maggio 2006
 Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Profiling of non-coding RNA classes Gunter Meister
 Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Chapter 5: Organization and Expression of Immunoglobulin Genes
 Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
MicroRNA formation. 4th International Symposium on Non-Surgical Contraceptive Methods of Pet Population Control
 MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Central Dogma. Lecture 10. Discussing DNA replication. DNA Replication. DNA mutation and repair. Transcription
 Central Dogma transcription translation DNA RNA Protein replication Discussing DNA replication (Nucleus of eukaryote, cytoplasm of prokaryote) Recall Replication is semi-conservative and bidirectional
Central Dogma transcription translation DNA RNA Protein replication Discussing DNA replication (Nucleus of eukaryote, cytoplasm of prokaryote) Recall Replication is semi-conservative and bidirectional
Interplay between microrna and Hepatitis B virus replication. A Thesis. Submitted to the Faculty. Drexel University. Yi Guo
 i Interplay between microrna and Hepatitis B virus replication A Thesis Submitted to the Faculty of Drexel University by Yi Guo in partial fulfillment of the requirements for the degree of Master of Science
i Interplay between microrna and Hepatitis B virus replication A Thesis Submitted to the Faculty of Drexel University by Yi Guo in partial fulfillment of the requirements for the degree of Master of Science
Control of Gene Expression
 Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
THE ENZYMES. Department of Microbiology, Immunology, and Molecular Genetics, Molecular Biology Institute University of California
 VOLUME THIRTY TWO THE ENZYMES Eukaryotic RNases and their Partners in RNA Degradation and Biogenesis, Part B Edited by FENG GUO Department of Biological Chemistry, David Geffen School of Medicine, Molecular
VOLUME THIRTY TWO THE ENZYMES Eukaryotic RNases and their Partners in RNA Degradation and Biogenesis, Part B Edited by FENG GUO Department of Biological Chemistry, David Geffen School of Medicine, Molecular
LightSwitch Luciferase Assay System
 Constructs, Assays & Services Assay System Promoter GoClones Synthetic Response Elements Pathway Cell Lines 3 UTR GoClones mirna Mimics & Inhibitors Synthetic mirna Targets Transfection Reagents Assay
Constructs, Assays & Services Assay System Promoter GoClones Synthetic Response Elements Pathway Cell Lines 3 UTR GoClones mirna Mimics & Inhibitors Synthetic mirna Targets Transfection Reagents Assay
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA
 CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved
 microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
Recombinant DNA & Genetic Engineering. Tools for Genetic Manipulation
 Recombinant DNA & Genetic Engineering g Genetic Manipulation: Tools Kathleen Hill Associate Professor Department of Biology The University of Western Ontario Tools for Genetic Manipulation DNA, RNA, cdna
Recombinant DNA & Genetic Engineering g Genetic Manipulation: Tools Kathleen Hill Associate Professor Department of Biology The University of Western Ontario Tools for Genetic Manipulation DNA, RNA, cdna
GENE REGULATION. Teacher Packet
 AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
Name Date Period. 2. When a molecule of double-stranded DNA undergoes replication, it results in
 DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
One Step mirna Cloning Vector G0760 pfbaavmu6mcs CMVeGFP SV40pA
 One Step mirna Cloning Vector G0760 pfbaavmu6mcs CMVeGFP SV40pA Plasmid Features: Coordinates Feature 194-348 Tn7L 385-511 SV40pA (complementary-outside ITRs) 689-818 AAV2 ITR (130bp) 877-1188 EcoRI mu6
One Step mirna Cloning Vector G0760 pfbaavmu6mcs CMVeGFP SV40pA Plasmid Features: Coordinates Feature 194-348 Tn7L 385-511 SV40pA (complementary-outside ITRs) 689-818 AAV2 ITR (130bp) 877-1188 EcoRI mu6
Expression and Purification of Recombinant Protein in bacteria and Yeast. Presented By: Puspa pandey, Mohit sachdeva & Ming yu
 Expression and Purification of Recombinant Protein in bacteria and Yeast Presented By: Puspa pandey, Mohit sachdeva & Ming yu DNA Vectors Molecular carriers which carry fragments of DNA into host cell.
Expression and Purification of Recombinant Protein in bacteria and Yeast Presented By: Puspa pandey, Mohit sachdeva & Ming yu DNA Vectors Molecular carriers which carry fragments of DNA into host cell.
A Novel Method to Detect Functional MicroRNA Targets
 doi:10.1016/j.jmb.2006.02.063 J. Mol. Biol. (2006) 358, 983 996 A Novel Method to Detect Functional MicroRNA Targets Sergei Vatolin*, Kapila Navaratne and Robert J. Weil Brain Tumor Institute, The Cleveland
doi:10.1016/j.jmb.2006.02.063 J. Mol. Biol. (2006) 358, 983 996 A Novel Method to Detect Functional MicroRNA Targets Sergei Vatolin*, Kapila Navaratne and Robert J. Weil Brain Tumor Institute, The Cleveland
From DNA to Protein
 Nucleus Control center of the cell contains the genetic library encoded in the sequences of nucleotides in molecules of DNA code for the amino acid sequences of all proteins determines which specific proteins
Nucleus Control center of the cell contains the genetic library encoded in the sequences of nucleotides in molecules of DNA code for the amino acid sequences of all proteins determines which specific proteins
QuickTiter FeLV Core Antigen ELISA Kit (FeLV p27)
 Product Manual QuickTiter FeLV Core Antigen ELISA Kit (FeLV p27) Catalog Numbers VPK-155 96 wells FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Feline leukemia virus (FeLV) is
Product Manual QuickTiter FeLV Core Antigen ELISA Kit (FeLV p27) Catalog Numbers VPK-155 96 wells FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Feline leukemia virus (FeLV) is
mrna EDITING Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A. Copyright 2005
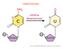 mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
10 µg lyophilized plasmid DNA (store lyophilized plasmid at 20 C)
 TECHNICAL DATA SHEET BIOLUMINESCENCE RESONANCE ENERGY TRANSFER RENILLA LUCIFERASE FUSION PROTEIN EXPRESSION VECTOR Product: prluc-c Vectors Catalog number: Description: Amount: The prluc-c vectors contain
TECHNICAL DATA SHEET BIOLUMINESCENCE RESONANCE ENERGY TRANSFER RENILLA LUCIFERASE FUSION PROTEIN EXPRESSION VECTOR Product: prluc-c Vectors Catalog number: Description: Amount: The prluc-c vectors contain
