Alternative Splicing in Higher Plants. Just Adding to Proteomic Diversity or an Additional Layer of Regulation?
|
|
|
- Holly Williamson
- 7 years ago
- Views:
Transcription
1 in Higher Plants Just Adding to Proteomic Diversity or an Additional Layer of Regulation?
2 Alternative splicing is nearly ubiquitous in eukaryotes It has been found in plants, flies, worms, mammals, etc. Note: Saccharomyces cerevisiae is an intron poor species (95% of genes lack introns) Alternative splicing is rare in this species Alternative splicing has recently expanded as an area of research with the advent of high throughput sequencing efforts 2 divergent roles Proteomic diversity Post-transcriptional regulation
3 RNA Polymerase II mediated transcription of a gene Alternative splicing is nearly ubiquitous in eukaryotes Alternative splicing is defined as different isoforms of mrna that can originate from a common locus mrna is generated from RNA polymerase II transcription of a gene A schema of eukaryotic transcription is given below RNA Polymerase II Pre-mRNA Gene X 5 3 DNA
4 Pre-mRNA processing in a mature mrna Pre-mRNA generated by RNA polymerase II must processed to be used for translation These steps include: Removal of introns and ligation of the exons Capping of the 5 end of the mrna Poly A tail Pre-mRNA 5 3 Intron removal (splicing) 5 capping Addition of the poly A tail AAAAAAAAAAAAAAAAA Mature mrna is ready for translation
5 Altenative splicing is commonly defined as alternatively processed mrna structure(s) relative to a predominant isoform originating from a common template (locus) The predominant isoform is the one most commonly observed Note Predominance is dependent upon the context (spatial, temporal) of expression Alternative splicing is commonly detected by: Aligning ESTs to the genome (or each other) and looking for structural diversity (genome wide) RT-PCR/RACE of a particular locus and looking for alternate isoforms (locus specific) Microarrays that cover the locus can be used to detect alternate isoforms (not common in plants)
6 Some points to consider There are two commonly recognized hypotheses about why alternative splicing is employed in eukaryotes Proteomic diversity Post-transcriptional regulation Alternative splicing was first postulated in 1978 by Walter Gilbert as a way to create novel isoforms (mrnas) from a common template (the DNA template for the gene) In order to detect alternative splicing, one needs the sequence of both the processed or mature mrna and the original genomic sequence Alignment of the mrna to the genomic sequence is performed in silico using alignment programs (not by hand!!!) If two or more mrnas are derived from the same locus but have differing structures, then this gene is said to be alternatively spliced Thus alternative splicing is defined by different variations or isoforms of the mrna that come from a shared gene
7 Genome wide alternative splicing analyses began in earnest in the late 1990s Requirements: - Genome sequence (spanning entire gene-space) - EST collections - Alignment/mapping strategy (e.g. PASA) - Detection of alternative isoforms (coordinate based strategy after mapping) First genomes that were analyzed using a genome-wide strategy were human and mouse Initial rates of genome wide analysis was 10-20% of genes were alternatively spliced Subsequent rates of alternative splicing have expanded to >60% of alternative splicing in human Humans have > 7million ESTs currently and mouse has >4 million ESTs As EST numbers increase, alternative splicing rates in a genome are also found to increase.
8 Interest in alternative splicing analyses in humans became greater with the annotation of the genome and the relatively low gene number The complexity of human development is difficult to explain using only 30,000 genes Humans have had a recent expansion of the transposable element Alu in the genome - 4% of human protein coding genes contain at least one Alu - Comparative analyses among primates have showed that Alu elements in introns can be converted into an exon (SLOWLY) - Thus far, all trancribed human genes with an Alu element are alternatively spliced
9 Phenotypic effects of alternative splicing are most commonly noted in human disease Cystic fibrosis is commonly associated with SNPs that reduce splicing of exons 9 and 12 Recent multivariate analysis has supported the intuitive idea that longer genes with higher numbers of introns are more likely to lead to splicing defects Proposed that between 15 and 60% of hereditary diseases are due to splicing mutations Diseases known to be caused by splicing disorders include: cancer, neurodegenerative diseases, ataxia, retardation, hypercholesterolemia, Its direct role in phenotypic variation in other species is less clear and has been analyzed in a more circumscribed fasion
10 Generating a cdna library Leaf Grind-up Leaf tissue Extraction Leaf mrna set AAAAAAAAAAAAAAAAA AAAAAAAAAAAAAAAAA AAAAAAAAAAAAAAAAA AAAAAAAAAAAAAAAAA AAAAAAAAAAAAAAAAA AAAAAAAAAAAAAAAAA AAAAAAAAAAAAAAAAA AAAAAAAAAAAAAAAAA Plasmid cdna generation Cloning into a plasmid Double stranded cdnas cdna library
11 Sequencing a cdna library Sequence top strand cdna Both strands of the cdna have been sequenced and you can identify the original mrna sequence Sequence bottom strand Repeat this procedure thousands of times from a cdna library to get sample of which genes are expressed and what their structure is
12 Expressed Sequence Tag (EST) sequence Sequence top strand This sequence of one end of a cdna is often called an EST or expressed sequence tag. cdna Expressed sequence tags (ESTs) are often bp in length ESTs reflect a partial sequence of a gene (usually) given that genes are commonly >1000bp in length ESTs can be used to identify areas of the genomic sequence where there are genes more intensive strategies are used to identify the full length of the gene
13 Aligning cdnas to the genomic sequence cdna Sequence both strands Generate a consensus sequence cdna consensus sequence Now use the consensus cdna sequence to align to the genomic sequence cdna Genomic DNA sequence Note that the alignment of the cdna to the genomic sequence is not contiguous, rather the cdna sequence appears to be spliced blocks of disparate genomic sequence
14 Quick refresher of intron splicing Higher eukaryotes use the following dinucleotide pairs in splicing U2 based spliceosome GT AG (99%) GC AG (1%) U12 based minor spliceosome AT AC (<0.1%) The dinucleotides are the two nucleotides in the intron that abut the exon There is a branch point site is upstream of the 3 acceptor in the intron Donor BPS Acceptor 5 Exon GT GC AT A AG AC 3 Exon
15 Consensus sequence for the acceptor site (32 bp around the splice site) Note the polypyrimidine tract upstream Note the AG dinucleotide is invariant Note the sequence downstream of the AG is a randomized mixture (reflective of the coding sequence)
16 Consensus sequence for the donor site (32 bp around the splice site) Note the polypyrimidine tract downstream Note the GT dinucleotide is nearly invariant (GC is observed) Note the sequence upstream of the AG is a randomized mixture (reflective of the coding sequence)
17 Alternative splicing analyses have a common set of alternative splicing isoforms: Alternate acceptor Alternate donor Alternate Terminal Exon Skipped/Retained exon Alternate initiation (Initiation within an Intron) Alternate polyadenylation (Termination within an Intron) Retained Intron
18 Note: The alternate acceptor has two different acceptor sites that are utilized The first five exons are identical.but the 3 exons after the fifth exon are quite different Asmbl_9392 has an upstream acceptor that incorporates a stop codon prematurely Asmbl_9391 has a downstream acceptor site that extends the coding sequence
19 Note: The alternate donor sites for the first exon are quite different Asmbl_17039 utilizes a downstream donor that incorporates a stop codon prematurely Asmbl_17038 utilizes an upstream donor site that extends the coding sequence
20 Note: The alternate terminal exon has 3 terminal exon(s) that are completely distinct (i.e. there is no overlap) Asmbl_18191 utilizes a upstream splice site which has a different termination codon and transcriptional termination/polyadenylation site when compared with asmbl_18190
21 Note: The skipped/retained exon has differential inclusion of an internal exon Here the third exon of asmbl_2859 is not included in its cognate pair of asmbl_2858
22 Note: Alternate initiation (Initiation within an Intron) has two distinct initiation sites for transcription, and more narrowly, the initiation of one isoform is contained wholly within the intron of its cognate pair Here transcription of asmbl_11897 is initiated within the fourth intron of asmbl_11896 which leads to very different N-terminal proteins.
23 Note: Alternate polyadenylation (termination within an intron) has two distinct initiation sites for polyadenylation, and more narrowly, the termination of transcription of one isoform is contained wholly within the intron of its cognate pair Here transcription of asmbl_10496 is terminated within the sixth intron of asmbl_10495 which leads to slightly different C-terminal proteins
24 Note: The retained intron isoform possesses an intron that is not spliced out in one isoform when compared to its cognate pair Here, the fourth intron for asmbl_17706 is not spliced out of asmbl_17707 which leads to a premature stop codon being introduced and very distinct C- terminal proteins.
25 Example: Alternative Splicing in rice Rice FL-cDNA Consortium, 2003 >28,000 FLcDNAs produces 13.8% alternatively spliced loci Campbell et al., ,156,705 ESTs and FL-cDNAs 15.7% of all annotated rice genes are alternatively spliced The three most frequent classes are alternate acceptor, alternate donor and retained intron Wang et al., ,816 ESTs and FL-cDNAs 21.2% of the transcribed genes are alternatively spliced Retained intron was the most frequent alternative splicing class
26 Isoform Variability at a Gene How many isoforms can a locus have? Wang and Brendel found that alternative splicing at a locus creates a finite number of isoforms ~60% of the transcribed loci in rice exhibit only a single isoform (no alternative splicing) Campbell found similar results: 68.3% of rice loci have only one isoform (no alternative splicing) For those loci that are alternatively spliced, 57.3% have only two isoforms 2 loci in rice have 17 different isoforms in rice and one Arabidopsis locus has 21 different isoforms
27 Analysis in human for the alternate acceptors looked at the rates of retaining frame If the alternate acceptors are separated by a bp count evenly divisible by 3bp (e.g. 3, 6, 9, etc) then the frame of translation will be maintained The authors found that the acceptors separated by 3bp are the most frequently occurring which is also observed in rice and Arabidopsis The separation by 3bp will introduce a single amino acid in the isoform using the proximal acceptor (red) when compared to the isoform using the distal acceptor (grey) The motif identifed is called NAGNAG where the N stands for any nucleotide Donor Proxmial Distal 5 Exon NAGNAG 3 Exon
28 Donor Proxmial Distal 5 Exon NAGNAG 3 Exon Using the proximal donor site: 5 Exon NAG 3 Exon Using the distal donor site 5 Exon 3 Exon Note how this difference in acceptor sites will alter the length of the mrna by 3bp and alter the translated product by 1 amino acid
29 Human analysis revealed that the NAGNAG was most commonly the sequence CAGCAG Below the CAGCAG motif is also observed in rice when looking at the 3bp difference between alternative acceptors. Note the CAGCAG motif is the most common Proximal Distal
30 For alternate acceptor, alternate donor, retained intron and skipped exon classes where the difference in sequence length is not divisible by 3, what is the effect? Frameshift in the translated product! Frameshifting of the mrna can lead to potentially new motifs being translated OR the incorporation of premature termination codons (PTCs) The introduction of PTCs into a mrna has been shown to induce non-sense mediated (NMD) decay pathways This proposed NMD degradation offers a post-transcriptional regulatory role
31 The rule in mammalian systems has been that NMD pathways are induced when there is a PTC >50bp upstream of an exon/exon junction Degradation occurs at the time of translation where the PTC is detected in the ribosome relative to the exon junctions Recent work in mammalian systems suggests that over one-third of all alternatively spliced transcripts meet the criteria for NMD This phenomenon of high rates of alternative splicing leading to NMD is called Regulated Unproductive Splicing and Translation (RUST)
32 Wang and Brendel found: 62% of alternative splicing isoforms in Arabidopsis will alter the frame of translation 56% of alternative splicing isoforms in rice will alter the frame of translation For these alternative splicing isoforms that have a frameshift: - In Arabidopsis, 42% of these frameshifted events are found to introduce a premature stop codon >50bp upstream of the last exon-exon junction and are NMD candidates -In rice, this percentage is 36% This level of producing NMD candidates suggests that RUST may be present in higher plants and be linked with alternative splicing
33 Conclusions: Alternative Splicing is a common feature of plant genomes As the size of EST collections have expanded, rates of alternative splicing increase as well Alternative splicing generally creates a few isoforms at any give locus restricting its role in expanding proteomic diversity Higher plants show evidence of alternative splicing being linked with NMD similar to mammalian systems
Gene Models & Bed format: What they represent.
 GeneModels&Bedformat:Whattheyrepresent. Gene models are hypotheses about the structure of transcripts produced by a gene. Like all models, they may be correct, partly correct, or entirely wrong. Typically,
GeneModels&Bedformat:Whattheyrepresent. Gene models are hypotheses about the structure of transcripts produced by a gene. Like all models, they may be correct, partly correct, or entirely wrong. Typically,
1 Mutation and Genetic Change
 CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
Human Genome Organization: An Update. Genome Organization: An Update
 Human Genome Organization: An Update Genome Organization: An Update Highlights of Human Genome Project Timetable Proposed in 1990 as 3 billion dollar joint venture between DOE and NIH with 15 year completion
Human Genome Organization: An Update Genome Organization: An Update Highlights of Human Genome Project Timetable Proposed in 1990 as 3 billion dollar joint venture between DOE and NIH with 15 year completion
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
DNA Replication & Protein Synthesis. This isn t a baaaaaaaddd chapter!!!
 DNA Replication & Protein Synthesis This isn t a baaaaaaaddd chapter!!! The Discovery of DNA s Structure Watson and Crick s discovery of DNA s structure was based on almost fifty years of research by other
DNA Replication & Protein Synthesis This isn t a baaaaaaaddd chapter!!! The Discovery of DNA s Structure Watson and Crick s discovery of DNA s structure was based on almost fifty years of research by other
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
Introduction to Genome Annotation
 Introduction to Genome Annotation AGCGTGGTAGCGCGAGTTTGCGAGCTAGCTAGGCTCCGGATGCGA CCAGCTTTGATAGATGAATATAGTGTGCGCGACTAGCTGTGTGTT GAATATATAGTGTGTCTCTCGATATGTAGTCTGGATCTAGTGTTG GTGTAGATGGAGATCGCGTAGCGTGGTAGCGCGAGTTTGCGAGCT
Introduction to Genome Annotation AGCGTGGTAGCGCGAGTTTGCGAGCTAGCTAGGCTCCGGATGCGA CCAGCTTTGATAGATGAATATAGTGTGCGCGACTAGCTGTGTGTT GAATATATAGTGTGTCTCTCGATATGTAGTCTGGATCTAGTGTTG GTGTAGATGGAGATCGCGTAGCGTGGTAGCGCGAGTTTGCGAGCT
Bioinformatics Resources at a Glance
 Bioinformatics Resources at a Glance A Note about FASTA Format There are MANY free bioinformatics tools available online. Bioinformaticists have developed a standard format for nucleotide and protein sequences
Bioinformatics Resources at a Glance A Note about FASTA Format There are MANY free bioinformatics tools available online. Bioinformaticists have developed a standard format for nucleotide and protein sequences
To be able to describe polypeptide synthesis including transcription and splicing
 Thursday 8th March COPY LO: To be able to describe polypeptide synthesis including transcription and splicing Starter Explain the difference between transcription and translation BATS Describe and explain
Thursday 8th March COPY LO: To be able to describe polypeptide synthesis including transcription and splicing Starter Explain the difference between transcription and translation BATS Describe and explain
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION. Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.
 Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals
 Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Protein Synthesis How Genes Become Constituent Molecules
 Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Lecture Series 7. From DNA to Protein. Genotype to Phenotype. Reading Assignments. A. Genes and the Synthesis of Polypeptides
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Module 3 Questions. 7. Chemotaxis is an example of signal transduction. Explain, with the use of diagrams.
 Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
MUTATION, DNA REPAIR AND CANCER
 MUTATION, DNA REPAIR AND CANCER 1 Mutation A heritable change in the genetic material Essential to the continuity of life Source of variation for natural selection New mutations are more likely to be harmful
MUTATION, DNA REPAIR AND CANCER 1 Mutation A heritable change in the genetic material Essential to the continuity of life Source of variation for natural selection New mutations are more likely to be harmful
Structure and Function of DNA
 Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
GENE REGULATION. Teacher Packet
 AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
Luísa Romão. Instituto Nacional de Saúde Dr. Ricardo Jorge Av. Padre Cruz, 1649-016 Lisboa, Portugal. Cooper et al (2009) Cell 136: 777
 Luísa Romão Instituto Nacional de Saúde Dr. Ricardo Jorge Av. Padre Cruz, 1649-016 Lisboa, Portugal Cooper et al (2009) Cell 136: 777 PTC = nonsense or stop codon = UAA, UAG, UGA PTCs can arise in a variety
Luísa Romão Instituto Nacional de Saúde Dr. Ricardo Jorge Av. Padre Cruz, 1649-016 Lisboa, Portugal Cooper et al (2009) Cell 136: 777 PTC = nonsense or stop codon = UAA, UAG, UGA PTCs can arise in a variety
Lecture 3: Mutations
 Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
Transcription and Translation of DNA
 Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Biological Sciences Initiative. Human Genome
 Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
13.4 Gene Regulation and Expression
 13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
Coding sequence the sequence of nucleotide bases on the DNA that are transcribed into RNA which are in turn translated into protein
 Assignment 3 Michele Owens Vocabulary Gene: A sequence of DNA that instructs a cell to produce a particular protein Promoter a control sequence near the start of a gene Coding sequence the sequence of
Assignment 3 Michele Owens Vocabulary Gene: A sequence of DNA that instructs a cell to produce a particular protein Promoter a control sequence near the start of a gene Coding sequence the sequence of
Overview of Eukaryotic Gene Prediction
 Overview of Eukaryotic Gene Prediction CBB 231 / COMPSCI 261 W.H. Majoros What is DNA? Nucleus Chromosome Telomere Centromere Cell Telomere base pairs histones DNA (double helix) DNA is a Double Helix
Overview of Eukaryotic Gene Prediction CBB 231 / COMPSCI 261 W.H. Majoros What is DNA? Nucleus Chromosome Telomere Centromere Cell Telomere base pairs histones DNA (double helix) DNA is a Double Helix
AP BIOLOGY 2009 SCORING GUIDELINES
 AP BIOLOGY 2009 SCORING GUIDELINES Question 4 The flow of genetic information from DNA to protein in eukaryotic cells is called the central dogma of biology. (a) Explain the role of each of the following
AP BIOLOGY 2009 SCORING GUIDELINES Question 4 The flow of genetic information from DNA to protein in eukaryotic cells is called the central dogma of biology. (a) Explain the role of each of the following
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
BioBoot Camp Genetics
 BioBoot Camp Genetics BIO.B.1.2.1 Describe how the process of DNA replication results in the transmission and/or conservation of genetic information DNA Replication is the process of DNA being copied before
BioBoot Camp Genetics BIO.B.1.2.1 Describe how the process of DNA replication results in the transmission and/or conservation of genetic information DNA Replication is the process of DNA being copied before
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE Q5B
 INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
Sample Questions for Exam 3
 Sample Questions for Exam 3 1. All of the following occur during prometaphase of mitosis in animal cells except a. the centrioles move toward opposite poles. b. the nucleolus can no longer be seen. c.
Sample Questions for Exam 3 1. All of the following occur during prometaphase of mitosis in animal cells except a. the centrioles move toward opposite poles. b. the nucleolus can no longer be seen. c.
RNA & Protein Synthesis
 RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
mrna EDITING Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A. Copyright 2005
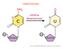 mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
2. The number of different kinds of nucleotides present in any DNA molecule is A) four B) six C) two D) three
 Chem 121 Chapter 22. Nucleic Acids 1. Any given nucleotide in a nucleic acid contains A) two bases and a sugar. B) one sugar, two bases and one phosphate. C) two sugars and one phosphate. D) one sugar,
Chem 121 Chapter 22. Nucleic Acids 1. Any given nucleotide in a nucleic acid contains A) two bases and a sugar. B) one sugar, two bases and one phosphate. C) two sugars and one phosphate. D) one sugar,
Discovery and Quantification of RNA with RNASeq Roderic Guigó Serra Centre de Regulació Genòmica (CRG) roderic.guigo@crg.cat
 Bioinformatique et Séquençage Haut Débit, Discovery and Quantification of RNA with RNASeq Roderic Guigó Serra Centre de Regulació Genòmica (CRG) roderic.guigo@crg.cat 1 RNA Transcription to RNA and subsequent
Bioinformatique et Séquençage Haut Débit, Discovery and Quantification of RNA with RNASeq Roderic Guigó Serra Centre de Regulació Genòmica (CRG) roderic.guigo@crg.cat 1 RNA Transcription to RNA and subsequent
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS)
 Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
From DNA to Protein. Proteins. Chapter 13. Prokaryotes and Eukaryotes. The Path From Genes to Proteins. All proteins consist of polypeptide chains
 Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
Frequently Asked Questions Next Generation Sequencing
 Frequently Asked Questions Next Generation Sequencing Import These Frequently Asked Questions for Next Generation Sequencing are some of the more common questions our customers ask. Questions are divided
Frequently Asked Questions Next Generation Sequencing Import These Frequently Asked Questions for Next Generation Sequencing are some of the more common questions our customers ask. Questions are divided
Recombinant DNA and Biotechnology
 Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Bio 102 Practice Problems Genetic Code and Mutation
 Bio 102 Practice Problems Genetic Code and Mutation Multiple choice: Unless otherwise directed, circle the one best answer: 1. Beadle and Tatum mutagenized Neurospora to find strains that required arginine
Bio 102 Practice Problems Genetic Code and Mutation Multiple choice: Unless otherwise directed, circle the one best answer: 1. Beadle and Tatum mutagenized Neurospora to find strains that required arginine
CCR Biology - Chapter 9 Practice Test - Summer 2012
 Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
A Primer of Genome Science THIRD
 A Primer of Genome Science THIRD EDITION GREG GIBSON-SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc. Publishers Sunderland, Massachusetts USA Contents Preface xi 1 Genome Projects:
A Primer of Genome Science THIRD EDITION GREG GIBSON-SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc. Publishers Sunderland, Massachusetts USA Contents Preface xi 1 Genome Projects:
Genetics Module B, Anchor 3
 Genetics Module B, Anchor 3 Key Concepts: - An individual s characteristics are determines by factors that are passed from one parental generation to the next. - During gamete formation, the alleles for
Genetics Module B, Anchor 3 Key Concepts: - An individual s characteristics are determines by factors that are passed from one parental generation to the next. - During gamete formation, the alleles for
Central Dogma. Lecture 10. Discussing DNA replication. DNA Replication. DNA mutation and repair. Transcription
 Central Dogma transcription translation DNA RNA Protein replication Discussing DNA replication (Nucleus of eukaryote, cytoplasm of prokaryote) Recall Replication is semi-conservative and bidirectional
Central Dogma transcription translation DNA RNA Protein replication Discussing DNA replication (Nucleus of eukaryote, cytoplasm of prokaryote) Recall Replication is semi-conservative and bidirectional
GenBank, Entrez, & FASTA
 GenBank, Entrez, & FASTA Nucleotide Sequence Databases First generation GenBank is a representative example started as sort of a museum to preserve knowledge of a sequence from first discovery great repositories,
GenBank, Entrez, & FASTA Nucleotide Sequence Databases First generation GenBank is a representative example started as sort of a museum to preserve knowledge of a sequence from first discovery great repositories,
Genomes and SNPs in Malaria and Sickle Cell Anemia
 Genomes and SNPs in Malaria and Sickle Cell Anemia Introduction to Genome Browsing with Ensembl Ensembl The vast amount of information in biological databases today demands a way of organising and accessing
Genomes and SNPs in Malaria and Sickle Cell Anemia Introduction to Genome Browsing with Ensembl Ensembl The vast amount of information in biological databases today demands a way of organising and accessing
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation
 OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
Concluding lesson. Student manual. What kind of protein are you? (Basic)
 Concluding lesson Student manual What kind of protein are you? (Basic) Part 1 The hereditary material of an organism is stored in a coded way on the DNA. This code consists of four different nucleotides:
Concluding lesson Student manual What kind of protein are you? (Basic) Part 1 The hereditary material of an organism is stored in a coded way on the DNA. This code consists of four different nucleotides:
European Medicines Agency
 European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
Lecture 6: Single nucleotide polymorphisms (SNPs) and Restriction Fragment Length Polymorphisms (RFLPs)
 Lecture 6: Single nucleotide polymorphisms (SNPs) and Restriction Fragment Length Polymorphisms (RFLPs) Single nucleotide polymorphisms or SNPs (pronounced "snips") are DNA sequence variations that occur
Lecture 6: Single nucleotide polymorphisms (SNPs) and Restriction Fragment Length Polymorphisms (RFLPs) Single nucleotide polymorphisms or SNPs (pronounced "snips") are DNA sequence variations that occur
RNA: Transcription and Processing
 8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
a. Ribosomal RNA rrna a type ofrna that combines with proteins to form Ribosomes on which polypeptide chains of proteins are assembled
 Biology 101 Chapter 14 Name: Fill-in-the-Blanks Which base follows the next in a strand of DNA is referred to. as the base (1) Sequence. The region of DNA that calls for the assembly of specific amino
Biology 101 Chapter 14 Name: Fill-in-the-Blanks Which base follows the next in a strand of DNA is referred to. as the base (1) Sequence. The region of DNA that calls for the assembly of specific amino
Standards, Guidelines and Best Practices for RNA-Seq V1.0 (June 2011) The ENCODE Consortium
 Standards, Guidelines and Best Practices for RNA-Seq V1.0 (June 2011) The ENCODE Consortium I. Introduction: Sequence based assays of transcriptomes (RNA-seq) are in wide use because of their favorable
Standards, Guidelines and Best Practices for RNA-Seq V1.0 (June 2011) The ENCODE Consortium I. Introduction: Sequence based assays of transcriptomes (RNA-seq) are in wide use because of their favorable
The sequence of bases on the mrna is a code that determines the sequence of amino acids in the polypeptide being synthesized:
 Module 3F Protein Synthesis So far in this unit, we have examined: How genes are transmitted from one generation to the next Where genes are located What genes are made of How genes are replicated How
Module 3F Protein Synthesis So far in this unit, we have examined: How genes are transmitted from one generation to the next Where genes are located What genes are made of How genes are replicated How
Name: Date: Period: DNA Unit: DNA Webquest
 Name: Date: Period: DNA Unit: DNA Webquest Part 1 History, DNA Structure, DNA Replication DNA History http://www.dnaftb.org/dnaftb/1/concept/index.html Read the text and answer the following questions.
Name: Date: Period: DNA Unit: DNA Webquest Part 1 History, DNA Structure, DNA Replication DNA History http://www.dnaftb.org/dnaftb/1/concept/index.html Read the text and answer the following questions.
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College
 Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
Becker Muscular Dystrophy
 Muscular Dystrophy A Case Study of Positional Cloning Described by Benjamin Duchenne (1868) X-linked recessive disease causing severe muscular degeneration. 100 % penetrance X d Y affected male Frequency
Muscular Dystrophy A Case Study of Positional Cloning Described by Benjamin Duchenne (1868) X-linked recessive disease causing severe muscular degeneration. 100 % penetrance X d Y affected male Frequency
2006 7.012 Problem Set 3 KEY
 2006 7.012 Problem Set 3 KEY Due before 5 PM on FRIDAY, October 13, 2006. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Which reaction is catalyzed by each
2006 7.012 Problem Set 3 KEY Due before 5 PM on FRIDAY, October 13, 2006. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Which reaction is catalyzed by each
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
How To Understand How Gene Expression Is Regulated
 What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
RETRIEVING SEQUENCE INFORMATION. Nucleotide sequence databases. Database search. Sequence alignment and comparison
 RETRIEVING SEQUENCE INFORMATION Nucleotide sequence databases Database search Sequence alignment and comparison Biological sequence databases Originally just a storage place for sequences. Currently the
RETRIEVING SEQUENCE INFORMATION Nucleotide sequence databases Database search Sequence alignment and comparison Biological sequence databases Originally just a storage place for sequences. Currently the
Protein Synthesis. Page 41 Page 44 Page 47 Page 42 Page 45 Page 48 Page 43 Page 46 Page 49. Page 41. DNA RNA Protein. Vocabulary
 Protein Synthesis Vocabulary Transcription Translation Translocation Chromosomal mutation Deoxyribonucleic acid Frame shift mutation Gene expression Mutation Point mutation Page 41 Page 41 Page 44 Page
Protein Synthesis Vocabulary Transcription Translation Translocation Chromosomal mutation Deoxyribonucleic acid Frame shift mutation Gene expression Mutation Point mutation Page 41 Page 41 Page 44 Page
Chapter 5: Organization and Expression of Immunoglobulin Genes
 Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID papillary renal cell carcinoma (translocation-associated) PRCC Human This gene
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID papillary renal cell carcinoma (translocation-associated) PRCC Human This gene
PrimePCR Assay Validation Report
 Gene Information Gene Name sorbin and SH3 domain containing 2 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SORBS2 Human Arg and c-abl represent the mammalian
Gene Information Gene Name sorbin and SH3 domain containing 2 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SORBS2 Human Arg and c-abl represent the mammalian
Gene mutation and molecular medicine Chapter 15
 Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
Next Generation Sequencing
 Next Generation Sequencing Technology and applications 10/1/2015 Jeroen Van Houdt - Genomics Core - KU Leuven - UZ Leuven 1 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977
Next Generation Sequencing Technology and applications 10/1/2015 Jeroen Van Houdt - Genomics Core - KU Leuven - UZ Leuven 1 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977
Recombinant DNA Technology
 Recombinant DNA Technology Stephen B. Gruber, MD, PhD Division of Molecular Medicine and Genetics November 4, 2002 Learning Objectives Know the basics of gene structure, function and regulation. Be familiar
Recombinant DNA Technology Stephen B. Gruber, MD, PhD Division of Molecular Medicine and Genetics November 4, 2002 Learning Objectives Know the basics of gene structure, function and regulation. Be familiar
Algorithms in Computational Biology (236522) spring 2007 Lecture #1
 Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
Chapter 17: From Gene to Protein
 AP Biology Reading Guide Fred and Theresa Holtzclaw Julia Keller 12d Chapter 17: From Gene to Protein 1. What is gene expression? Gene expression is the process by which DNA directs the synthesis of proteins
AP Biology Reading Guide Fred and Theresa Holtzclaw Julia Keller 12d Chapter 17: From Gene to Protein 1. What is gene expression? Gene expression is the process by which DNA directs the synthesis of proteins
Name Date Period. 2. When a molecule of double-stranded DNA undergoes replication, it results in
 DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
Transcription in prokaryotes. Elongation and termination
 Transcription in prokaryotes Elongation and termination After initiation the σ factor leaves the scene. Core polymerase is conducting the elongation of the chain. The core polymerase contains main nucleotide
Transcription in prokaryotes Elongation and termination After initiation the σ factor leaves the scene. Core polymerase is conducting the elongation of the chain. The core polymerase contains main nucleotide
Name Class Date. Figure 13 1. 2. Which nucleotide in Figure 13 1 indicates the nucleic acid above is RNA? a. uracil c. cytosine b. guanine d.
 13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
Activity 7.21 Transcription factors
 Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
New Technologies for Sensitive, Low-Input RNA-Seq. Clontech Laboratories, Inc.
 New Technologies for Sensitive, Low-Input RNA-Seq Clontech Laboratories, Inc. Outline Introduction Single-Cell-Capable mrna-seq Using SMART Technology SMARTer Ultra Low RNA Kit for the Fluidigm C 1 System
New Technologies for Sensitive, Low-Input RNA-Seq Clontech Laboratories, Inc. Outline Introduction Single-Cell-Capable mrna-seq Using SMART Technology SMARTer Ultra Low RNA Kit for the Fluidigm C 1 System
AP BIOLOGY 2010 SCORING GUIDELINES (Form B)
 AP BIOLOGY 2010 SCORING GUIDELINES (Form B) Question 2 Certain human genetic conditions, such as sickle cell anemia, result from single base-pair mutations in DNA. (a) Explain how a single base-pair mutation
AP BIOLOGY 2010 SCORING GUIDELINES (Form B) Question 2 Certain human genetic conditions, such as sickle cell anemia, result from single base-pair mutations in DNA. (a) Explain how a single base-pair mutation
Transfection-Transfer of non-viral genetic material into eukaryotic cells. Infection/ Transduction- Transfer of viral genetic material into cells.
 Transfection Key words: Transient transfection, Stable transfection, transfection methods, vector, plasmid, origin of replication, reporter gene/ protein, cloning site, promoter and enhancer, signal peptide,
Transfection Key words: Transient transfection, Stable transfection, transfection methods, vector, plasmid, origin of replication, reporter gene/ protein, cloning site, promoter and enhancer, signal peptide,
Genetics Lecture Notes 7.03 2005. Lectures 1 2
 Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Shouguo Gao Ph. D Department of Physics and Comprehensive Diabetes Center
 Computational Challenges in Storage, Analysis and Interpretation of Next-Generation Sequencing Data Shouguo Gao Ph. D Department of Physics and Comprehensive Diabetes Center Next Generation Sequencing
Computational Challenges in Storage, Analysis and Interpretation of Next-Generation Sequencing Data Shouguo Gao Ph. D Department of Physics and Comprehensive Diabetes Center Next Generation Sequencing
Outline. MicroRNA Bioinformatics. microrna biogenesis. short non-coding RNAs not considered in this lecture. ! Introduction
 Outline MicroRNA Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology (CMB) Karolinska Institutet! Introduction! microrna target site prediction! Useful resources 2 short non-coding RNAs
Outline MicroRNA Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology (CMB) Karolinska Institutet! Introduction! microrna target site prediction! Useful resources 2 short non-coding RNAs
CCR Biology - Chapter 8 Practice Test - Summer 2012
 Name: Class: Date: CCR Biology - Chapter 8 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. What did Hershey and Chase know
Name: Class: Date: CCR Biology - Chapter 8 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. What did Hershey and Chase know
Complex multicellular organisms are produced by cells that switch genes on and off during development.
 Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Lab # 12: DNA and RNA
 115 116 Concepts to be explored: Structure of DNA Nucleotides Amino Acids Proteins Genetic Code Mutation RNA Transcription to RNA Translation to a Protein Figure 12. 1: DNA double helix Introduction Long
115 116 Concepts to be explored: Structure of DNA Nucleotides Amino Acids Proteins Genetic Code Mutation RNA Transcription to RNA Translation to a Protein Figure 12. 1: DNA double helix Introduction Long
Hidden Markov Models in Bioinformatics. By Máthé Zoltán Kőrösi Zoltán 2006
 Hidden Markov Models in Bioinformatics By Máthé Zoltán Kőrösi Zoltán 2006 Outline Markov Chain HMM (Hidden Markov Model) Hidden Markov Models in Bioinformatics Gene Finding Gene Finding Model Viterbi algorithm
Hidden Markov Models in Bioinformatics By Máthé Zoltán Kőrösi Zoltán 2006 Outline Markov Chain HMM (Hidden Markov Model) Hidden Markov Models in Bioinformatics Gene Finding Gene Finding Model Viterbi algorithm
Searching Nucleotide Databases
 Searching Nucleotide Databases 1 When we search a nucleic acid databases, Mascot always performs a 6 frame translation on the fly. That is, 3 reading frames from the forward strand and 3 reading frames
Searching Nucleotide Databases 1 When we search a nucleic acid databases, Mascot always performs a 6 frame translation on the fly. That is, 3 reading frames from the forward strand and 3 reading frames
12.1 The Role of DNA in Heredity
 12.1 The Role of DNA in Heredity Only in the last 50 years have scientists understood the role of DNA in heredity. That understanding began with the discovery of DNA s structure. In 1952, Rosalind Franklin
12.1 The Role of DNA in Heredity Only in the last 50 years have scientists understood the role of DNA in heredity. That understanding began with the discovery of DNA s structure. In 1952, Rosalind Franklin
DNA, RNA, Protein synthesis, and Mutations. Chapters 12-13.3
 DNA, RNA, Protein synthesis, and Mutations Chapters 12-13.3 1A)Identify the components of DNA and explain its role in heredity. DNA s Role in heredity: Contains the genetic information of a cell that can
DNA, RNA, Protein synthesis, and Mutations Chapters 12-13.3 1A)Identify the components of DNA and explain its role in heredity. DNA s Role in heredity: Contains the genetic information of a cell that can
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA
 CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
Translation Study Guide
 Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Advances in RainDance Sequence Enrichment Technology and Applications in Cancer Research. March 17, 2011 Rendez-Vous Séquençage
 Advances in RainDance Sequence Enrichment Technology and Applications in Cancer Research March 17, 2011 Rendez-Vous Séquençage Presentation Overview Core Technology Review Sequence Enrichment Application
Advances in RainDance Sequence Enrichment Technology and Applications in Cancer Research March 17, 2011 Rendez-Vous Séquençage Presentation Overview Core Technology Review Sequence Enrichment Application
Gene and Chromosome Mutation Worksheet (reference pgs. 239-240 in Modern Biology textbook)
 Name Date Per Look at the diagrams, then answer the questions. Gene Mutations affect a single gene by changing its base sequence, resulting in an incorrect, or nonfunctional, protein being made. (a) A
Name Date Per Look at the diagrams, then answer the questions. Gene Mutations affect a single gene by changing its base sequence, resulting in an incorrect, or nonfunctional, protein being made. (a) A
Gene Finding CMSC 423
 Gene Finding CMSC 423 Finding Signals in DNA We just have a long string of A, C, G, Ts. How can we find the signals encoded in it? Suppose you encountered a language you didn t know. How would you decipher
Gene Finding CMSC 423 Finding Signals in DNA We just have a long string of A, C, G, Ts. How can we find the signals encoded in it? Suppose you encountered a language you didn t know. How would you decipher
Basic Concepts of DNA, Proteins, Genes and Genomes
 Basic Concepts of DNA, Proteins, Genes and Genomes Kun-Mao Chao 1,2,3 1 Graduate Institute of Biomedical Electronics and Bioinformatics 2 Department of Computer Science and Information Engineering 3 Graduate
Basic Concepts of DNA, Proteins, Genes and Genomes Kun-Mao Chao 1,2,3 1 Graduate Institute of Biomedical Electronics and Bioinformatics 2 Department of Computer Science and Information Engineering 3 Graduate
Outline. interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work?
 Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Lezioni Dipartimento di Oncologia Farmacologia Molecolare. RNA interference. Giovanna Damia 29 maggio 2006
 Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
Question 4 /29 points. Total /100 points
 MIT Department of Biology 7.28, Spring 2005 - Molecular Biology 7.28 Spring 2005 Exam Three Question 1 Question 2 Question 3 /30 points /20 points /21 points Question 4 /29 points Total /100 points 1 Question
MIT Department of Biology 7.28, Spring 2005 - Molecular Biology 7.28 Spring 2005 Exam Three Question 1 Question 2 Question 3 /30 points /20 points /21 points Question 4 /29 points Total /100 points 1 Question
The NF1-gene a hotspot for de novo Alu- and L1-insertion?
 The NF1-gene a hotspot for de novo Alu- and L1-insertion? Katharina Wimmer Division Humangenetik, Medizinische Universität Innsbruck Molekulare Diagnostik 2013 Zürich, 1. März, 2013 2 Die im Vortrag vorgestellten
The NF1-gene a hotspot for de novo Alu- and L1-insertion? Katharina Wimmer Division Humangenetik, Medizinische Universität Innsbruck Molekulare Diagnostik 2013 Zürich, 1. März, 2013 2 Die im Vortrag vorgestellten
SeqScape Software Version 2.5 Comprehensive Analysis Solution for Resequencing Applications
 Product Bulletin Sequencing Software SeqScape Software Version 2.5 Comprehensive Analysis Solution for Resequencing Applications Comprehensive reference sequence handling Helps interpret the role of each
Product Bulletin Sequencing Software SeqScape Software Version 2.5 Comprehensive Analysis Solution for Resequencing Applications Comprehensive reference sequence handling Helps interpret the role of each
Control of Gene Expression
 Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Bio 102 Practice Problems Recombinant DNA and Biotechnology
 Bio 102 Practice Problems Recombinant DNA and Biotechnology Multiple choice: Unless otherwise directed, circle the one best answer: 1. Which of the following DNA sequences could be the recognition site
Bio 102 Practice Problems Recombinant DNA and Biotechnology Multiple choice: Unless otherwise directed, circle the one best answer: 1. Which of the following DNA sequences could be the recognition site
Gene Switches A Model
 Gene Switches A Model Abstract Conceptually, how genetic switches function and their role in the process of evolution, can be difficult for students to visualize. Gene Switches A Model attempts to make
Gene Switches A Model Abstract Conceptually, how genetic switches function and their role in the process of evolution, can be difficult for students to visualize. Gene Switches A Model attempts to make
