Data Analytics. Sequence Exosome RNAs
|
|
|
- Marvin Stevenson
- 10 years ago
- Views:
Transcription
1 Sequence Exosome RNAs The Exo-NGS service provides the exosome researcher with a comprehensive, expert service to isolate and identify exosome-associated RNA biomarkers - leveraging the throughput and scalability of Illumina s MiSeq and HiSeq next-generation sequencing platforms. Most exosomal RNAs are less than 300 nucleotides in length, thus small RNA libraries are ideal for this application. This service is a turnkey solution that is tailored for researchers who are interested in identifying novel exosome RNA biomarkers or understanding the abundance of such biomarkers in the exosomes of their model cellular systems or patient biofluids. Input Sample Requirements Sequence Read Quality Assessment Read Quality Score Good Fair Biofluid Poor Volume Serum 500 l - 1ml Plasma 500 l - 1ml Cell Media 5ml - 10ml Urine 5ml - 10ml Spinal Fluid 5ml - 10ml Ascites Fluid 500 l - 1ml Other Inquire Sequence Read Quality Across all Bases Position in Read (bp) Building Exosome RNA Library Exosomes isolated from samples Exo-RNAs purified with chromatography Illumina bar-codes added and amplified Library size-selection and PAGE purification High sensitivity library QC with Bioanalyzer Multiplexed NGS runs performed Two rounds of quality checks on sequence data are performed. The first round analyzes raw sequencing data generated by the Illumina platform, and library adaptors and Ns are trimmed. The second data check is performed on the trimmed sequence data to ensure read quality before genome mapping. Compare exo-rna sequence profiles across patient samples. SBI and Maverix Biomics have teamed up to provide a complete analytics solution on deep sequencing data of exosome-associated RNAs. The analysis service includes library sequence quality control metrics, data analysis for relative RNA abundance and identity, differential expression analysis and visualization of the data in a cloud-based, private UCSC Genome Browser. Simplify and accelerate your exosome RNA biomarker discovery with the advanced bioinformatics analysis included in SBI s Exo-NGS service. Genome Browser Data Mining Sequencing data uploaded Access to public databases Analyze with accepted analytics Visualize your data automatically Compare to known ENCODE data Put your data into context. Publication-ready figures automatically generated with the Exo-NGS service. Data Analytics Primary Data Deliverables Raw sequencing reads in FASTA format Sequencing read quality values Analyzed Data Small RNA Workflow Relative abundance of each RNA type Table of counts of mature micrornas RNA Type Charts Expression Heatmaps 25 RNA Type Antisense RNA CDBox HAcaBox lincrna LINE LTR microrna Other ncrna pirna RefSeq exons RefSeq introns rrna scarna SINE Tandem repeat trna
2 White Paper Exosome RNA-seq Analysis The Maverix Analytic Platform facilitates discovery of small non-coding RNAs and biomarkers. Maverix Biomics, Inc S. Amphlett Blvd, Suite 214, San Mateo CA
3 Table of Contents Overview 3 Introduction 3 Approach 5 Analysis 6 Case Study 7 Summary 11 References 11 2
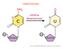
4 Overview Thirty years ago, the extracellular vesicles known as exosomes were identified while studying transferrin/receptor recycling in reticulocytes [1] but only in the last 5-6 years have studies ignited significant interest by elucidating their role in pathogenesis, cell-cell communication, drug, vaccine and gene-vector delivery, and as reservoirs of biomarkers [2]. With research expanding into next-generation sequencing technologies, advances have accelerated in the areas of intercellular communication and disease-related biomarker discovery. RNA-seq facilitates this research and the Maverix Analytic Platform has an analysis kit available to accelerate your discovery. Introduction Exosomes are small extracellular vesicles ( nm in diameter) of endocytic origin. These nanovesicles are formed by inward budding of late endosomes to produce multivesicular endosomes (MVEs), sometimes referred to as multivesicular bodies, and then released into the environment by fusion of the MVEs with the plasma membrane [3]. Figure 1. Release of microvesicles and exosomes. Microvesicles (MVs) bud directly from the plasma membrane, whereas exosomes are represented by small vesicles of different sizes that are formed as the intraluminal vesicles by budding into early endosomes and multivesicular endosomes (MVEs) and are released by fusion of MVEs with the plasma membrane. Other MVEs fuse with lysosomes. Red spots symbolize clathrin associated with vesicles at the plasma membrane (clathrin-coated vesicles [CCV]) or bilayered clathrin coats at endosomes. Membrane-associated and transmembrane proteins on vesicles are represented as triangles and rectangles, respectively. Arrows represent proposed directions of protein and lipid transport between organelles and between MVEs and the plasma membrane for exosome secretion. Illustration from Raposo and Stoorvogel, 2013 [3]. 3
5 Exosomes have been isolated from various sources, including amniotic fluid, bile, blood, breast milk, cerebrospinal fluid, malignant ascites fluid, saliva, semen and urine [3]. Their ability to be easily sampled from a patient s body fluids by relatively noninvasive methods makes them a valuable reagent. Exosomes are released by most cell types, with studies demonstrating various roles in antigen presentation, cell cell communication, immune response, and dissemination of infectious agents. Initially, the molecular components of exosomes were thought to be merely cell debris, but subsequent studies revealed otherwise. Proteins enriched in exosomes are derived from the endosomes, plasma membrane and the cytosol and not from the nucleus, mitochondria, or endoplasmic reticulum. Some common protein families identified in exosomes include chaperones, cytoskeletal proteins, ESCRT proteins, tetraspanin proteins, trimeric G protein subunits, and other proteins involved in transport and fusion. Much of the molecular content of exosomes is cell-specific in nature. The exosomal proteins, mrnas, micrornas, and lipids identified over the past several years have been curated in the extracellular vesicle database ExoCarta ( [4]. Exosome small RNA repeat RNA trf vrna SRP-RNA Y-RNA rrna mirna pirna snrna snorna Description LINEs, LTRs, and simple repeat sequences trna fragments vault RNA, part of the vault ribonucleoprotein complex signal recognition particle RNA ncrna component of the Ro ribonucleoprotein particle ribosomal RNA (<200nt) microrna piwi-interacting RNA small nuclear RNA small nucleolar RNA Table 1. Exosomal small RNAs. Various small non-coding RNAs have been identified in exosomes, many of which play roles in the conveyance of genetically encoded messages between cells [5]. Next-generation sequencing (NGS) has allowed researchers to focus on small non-coding RNAs (ncrnas) in exosomes. ExoCarta catalogues the small ncrnas known as micrornas (mirnas), which are typically 22 nucleotides in length. mirnas are derived from longer stem-loop precursors and are known to function as transcriptional and post-transcriptional regulators of gene expression. In addition to mirnas, many small RNAs have been isolated from exosomes (see Table 1) and are presumed to play roles in communication between cells, potentially modifying the function of target cells [5]. The resurgence of interest in exosomes is accelerating 4
6 the understanding of how these small RNAs facilitate communication between cells and organs, act through gene regulatory functions, and play important roles in a wide range of physiological and pathological processes. Approach RNA-seq is an NGS method that uses high-throughput sequencing technology to sequence the RNA content of an organism, tissue or cell. RNA-seq allows a researcher to define and analyze the transcriptome, which represents the full RNA complement and includes mrna, rrna, trna and other ncrnas. To focus on small ncrnas, size fractionation is performed prior to the ligation of adaptors and conversion to cdna in the subsequent library preparation steps. System Biosciences (SBI) and Maverix Biomics have teamed up to provide a combined solution for the isolation, purification, analysis and visualization of exosome RNA NGS data. SBI has engineered tools and NGS services to accelerate the study of exosomes and exosome RNA biomarkers, including a simple one-step process for isolating exosomes from biofluids, followed by RNA purification and RNA-seq. The sequence reads can then be analyzed using the Exosome Small RNA-seq Analysis Kit on the cloud-based Maverix Analytic Platform, which provides small RNA analysis and data visualization. The small RNA-seq analysis kit facilitates identification of novel small ncrnas, biomarker discovery, transcription start site detection, and the study of small RNA regulatory pathways. Figure 2. The Exosome Small RNA-seq Analysis Kit on the Maverix Analytic Platform. The Maverix Analytic Platform allows researchers to easily and quickly upload their sequence data and launch an analysis kit that uses peer-reviewed open-source algorithms that have been highly cited in life sciences journal articles and are now widely accepted as the gold standard for NGS analysis. Table 2 lists the open-source software utilized in the Exosome Small RNA-seq Analysis Kit. Other software used in the analysis includes the licensed UCSC Genome Browser and utilities [6], as well as Maverix Biomics in-house developed applications. 5
7 Analysis The Exosome Small RNA-seq Analysis Kit initiates with a data quality check of the input sequence using FastQC, an open-source quality control (QC) tool for high-throughput sequence data [7]. FastQC runs analyses of the uploaded raw sequence reads that reveal the quality of the data and inform the subsequent preprocessing steps in the analysis. Following QC, the analysis moves to preprocessing of the RNA-seq reads to improve the quality of data input for read mapping. The open-source tools used are FastqMcf, part of the EA-utils package [8], and PRINSEQ [9]. Data preprocessing detects and removes N s at the ends of reads, trims sequencing adapters, and filters reads for quality and length. FastQC is then re-run to analyze the trimmed reads, allowing a before and after comparison. The summary report generated provides a quality assurance check to validate the processed set of input data used in the subsequent read mapping step. Software Analysis Kit Step Citation FastQC Data quality control 7 ea-utils Data preprocessing 8 PRINSEQ Data preprocessing 9 Bowtie Sequence mapping 10 SAMtools Alignment processing, Genome browser track generation 11 Picard Alignment processing, Genome browser track generation 12 R Statistics and visualization generation 13 Table 2. Open-source software used in the Exosome Small RNA-seq Analysis Kit. List of the opensource software and tools used in the analysis, including the associated analysis step and the software citation. The improved set of sequence reads are mapped to the reference genome using Bowtie, an ultrafast, memory-efficient short read aligner [10], followed by the generation of a mapping summary report for review. Using the open-source software SAMtools [11] and Picard [12], expression analyses are carried out, including computation of read coverage, determination of ncrna abundance and differential expression analysis across samples when applicable. Expression statistics are calculated and visualized using R, a software environment for statistical computing and graphics [13]. The Exosome Small RNA-seq analysis produces a summarization of results, including expression statistics and chromosome distribution, as well as genome browser tracks of read alignment and read coverage for analysis in a genomic context. The Maverix Analytic Platform allows researchers to visualize their data and analytic results using an integrated UCSC Genome Browser, automatically configured for their specific organism of interest. With access to data in the UCSC Genome Browser directly from the platform, researchers can add custom tracks to the browser, securely surf their data, and easily share or publish their results. 6
8 Case Study Exosomes have been isolated from many body fluids including breast milk, which is a complex liquid containing immunological components that can impact the development of the offspring s immune system. In their 2012 publication [14], Zhou, et al. investigated the transcriptome of exosomes in human breast milk with a particular focus on mirnas whose levels are frequently elevated in diseased states. In their study, exosomes were isolated from the breast milk of four women when their infants were 60 days old using SBI's ExoQuick exosome precipitation reagent. Four exosomal small RNA libraries were constructed and each was individually sequenced, with a total of ~86.37 million 36-nt raw reads generated. Figure 3. Sequencing read mapping rate. The charts show the percentage and number of reads, respectively, for trimmed, mapped and unmapped reads for each of the four samples. 7
9 Taking the small RNA sequence data from this study (deposited by the authors to NCBI s Gene Expression Omnibus under accession GSE32253), we launched the Exosome Small RNA-seq Analysis Kit on the Maverix Analytic Platform using the raw sequence reads as input. Figure 3 shows the sequencing read mapping rate, with trimmed, mapped and unmapped reads displayed as a percentage of reads and as total read counts. The trimmed reads are the set of reads filtered out during the data preprocessing step of the analysis. The remaining reads were used as input for the mapping step and are displayed in the chart as either mapped or unmapped reads. Following read mapping, data analytics were undertaken, with expression analysis that included determination of small ncrna and repeat element abundance level. Moving beyond the mirna analysis undertaken in the original publication, the Exosome Small RNA-seq Analysis Kit identifies and maps mirnas, trnas, small rrnas, repeat elements, antisense transcripts and a variety of small ncrnas. Abundance levels are calculated and an expression summary chart is generated for ease of visualization. Figure 4. Exosome small RNA expression summary. Small RNAs, including trna, rrna, mirna, snorna and other ncrnas, as well as antisense transcripts and repeat sequences are displayed in a pie chart for each of the four breast milk exosome samples. 8
10 Another category of results output from the analysis kit are heat maps, which display differential expression data. The heat maps can expedite insights into which transcripts, repeat elements and small RNAs are enriched in exosomes. These graphical representations of data, where the individual values are represented as colors, provide a mechanism for visual analysis of samples side-by-side. The color variations indicate the enrichment levels, facilitating the identification of differential expression between samples or conditions, and accelerating discovery of biomarkers. The heat map navigation makes it easy to explore your results. Hovering over a region of interest reveals a tooltip with the RNA type, name, and chromosomal location with coordinates (Fig. 5). Clicking on a region of the heat map will take you to the associated locus in the UCSC Genome Browser, making it easy to identify regions of interest on the heat map then jump directly to the genomic context for further analysis. Figure 5. Heat map visualization of small RNA expression. The heat map on the left displays the four samples side-by-side (columns 1-4) for a visual comparison of expression levels by color. The fifth column in the heat map distinguishes components by chromosome location, where chromosomes are classified by unique colors. Hovering over a region of the heat map brings up the tooltip as shown, with information about the mapped exosomal component, including name, chromosomal position, and differential expression values for each sample. Clicking on a region of interest on the heat map, or alternatively clicking on the component name in the tooltip, will bring up the associated region in the integrated UCSC Genome Browser for visualization of exosome components in a genomic context. The final output from the analysis includes genome browser tracks that display the mapped reads and read coverage data. Browser tracks can be visualized in the UCSC Genome Browser directly within the Maverix Analytic Platform (Fig. 6). Viewing data in the genome browser allows for analysis in a genomic context and makes sharing and collaborating easy and secure. Once the analysis kit has provided its output, the RNA-seq reads from each of the four samples can be viewed together in the browser to facilitate visual comparisons of read lengths, coverage, and differential expression. 9
11 A B C Figure 6. RNA expression in the genome browser. Exosomal RNA expression viewed as mapped reads and read coverage via browser tracks visualized in the UCSC Genome Browser. Examples of small RNAs identified in breast milk exosomes include mirna (A), trna (B), as well as rrna and snorna (C). 10
12 Summary The study by Zhou, et al. proposed that exosomal mirnas are transferable genetic material from mother to infant, and are essential for the development of the immune system in infants [14]. We used the breast milk dataset for benchmarking of the Exosome Small RNA-seq Analysis Kit in the case study reported in this white paper. Recent exosome studies focus on identification of protein, lipids, mrna, and micrornas [4], the latter of which were the focus of Zhou, et al. in the human breast milk samples. Our analysis, carried out via the Exosome Small RNA-seq Analysis Kit, expands the number and type of small RNAs identified from the dataset. The analytic results show that there are a broad range of small ncrnas, transcripts and repeat elements within the exosomes, many of which may represent novel immune-related components in the breast milk. Utilizing the combined solution offered by SBI and Maverix Biomics, researchers have access to a comprehensive solution from isolation and purification of exosomes, through cloud-based analysis and visualization of exosome-associated RNA biomarkers. With the analysis results available for visualization within the integrated UCSC Genome Browser, the Maverix Analytic Platform makes it easy to download graphs and charts as images and create publication-ready figures from the genome browser views. References 1. Endocytosis and intracellular processing of transferrin and colloidal gold-transferrin in rat reticulocytes: demonstration of a pathway for receptor shedding. Harding C, Heuser J, Stahl P. Eur J Cell Biol Nov;35(2): Vesiclepedia: a compendium for extracellular vesicles with continuous community annotation. Kalra H, et al. PLoS Biol. 2012;10(12):e doi: /journal.pbio Extracellular vesicles: exosomes, microvesicles, and friends. Raposo G, Stoorvogel W. J Cell Biol Feb 18;200(4): doi: /jcb ExoCarta 2012: database of exosomal proteins, RNA and lipids. Mathivanan S, Fahner CJ, Reid GE, Simpson RJ. Nucleic Acids Res Jan;40(Database issue):d doi: /nar/gkr Deep sequencing of RNA from immune cell-derived vesicles uncovers the selective incorporation of small non-coding RNA biotypes with potential regulatory functions. Nolte-'t Hoen EN, Buermans HP, Waasdorp M, Stoorvogel W, Wauben MH, 't Hoen PA. Nucleic Acids Res Oct;40(18): doi: /nar/gks The UCSC genome browser and associated tools. Kuhn RM, Haussler D, Kent WJ. Brief Bioinform Mar;14(2): doi: /bib/bbs FastQC: A quality control tool for high throughput sequence data. Simon Andrews
13 8. ea-utils : "Command-line tools for processing biological sequencing data". Erik Aronesty, 2011; 9. Quality control and preprocessing of metagenomic datasets. Schmieder R, Edwards R. Bioinformatics Mar 15;27(6): doi: /bioinformatics/btr Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Langmead B, Trapnell C, Pop M, Salzberg SL. Genome Biol. 2009;10(3):R25. doi: / gb r The Sequence Alignment/Map format and SAMtools. Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R; 1000 Genome Project Data Processing Subgroup. Bioinformatics Aug 15;25(16): doi: /bioinformatics/btp Picard: A set of tools (in Java) for working with next generation sequencing data in the BAM format R: A language and environment for statistical computing. R Development Core Team (2008). R Foundation for Statistical Computing, Vienna, Austria. ISBN , Immune-related micrornas are abundant in breast milk exosomes. Zhou Q, Li M, Wang X, Li Q, Wang T, Zhu Q, Zhou X, Wang X, Gao X, Li X. Int J Biol Sci. 2012;8(1): doi: /ijbs
14 Discover the Unexpected 1670 S. Amphlett Blvd, Suite 214, San Mateo CA Maverix Biomics, Inc. All Rights Reserved.
PreciseTM Whitepaper
 Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
How-To: SNP and INDEL detection
 How-To: SNP and INDEL detection April 23, 2014 Lumenogix NGS SNP and INDEL detection Mutation Analysis Identifying known, and discovering novel genomic mutations, has been one of the most popular applications
How-To: SNP and INDEL detection April 23, 2014 Lumenogix NGS SNP and INDEL detection Mutation Analysis Identifying known, and discovering novel genomic mutations, has been one of the most popular applications
Analysis of ChIP-seq data in Galaxy
 Analysis of ChIP-seq data in Galaxy November, 2012 Local copy: https://galaxy.wi.mit.edu/ Joint project between BaRC and IT Main site: http://main.g2.bx.psu.edu/ 1 Font Conventions Bold and blue refers
Analysis of ChIP-seq data in Galaxy November, 2012 Local copy: https://galaxy.wi.mit.edu/ Joint project between BaRC and IT Main site: http://main.g2.bx.psu.edu/ 1 Font Conventions Bold and blue refers
AGILENT S BIOINFORMATICS ANALYSIS SOFTWARE
 ACCELERATING PROGRESS IS IN OUR GENES AGILENT S BIOINFORMATICS ANALYSIS SOFTWARE GENESPRING GENE EXPRESSION (GX) MASS PROFILER PROFESSIONAL (MPP) PATHWAY ARCHITECT (PA) See Deeper. Reach Further. BIOINFORMATICS
ACCELERATING PROGRESS IS IN OUR GENES AGILENT S BIOINFORMATICS ANALYSIS SOFTWARE GENESPRING GENE EXPRESSION (GX) MASS PROFILER PROFESSIONAL (MPP) PATHWAY ARCHITECT (PA) See Deeper. Reach Further. BIOINFORMATICS
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS)
 Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
Standards, Guidelines and Best Practices for RNA-Seq V1.0 (June 2011) The ENCODE Consortium
 Standards, Guidelines and Best Practices for RNA-Seq V1.0 (June 2011) The ENCODE Consortium I. Introduction: Sequence based assays of transcriptomes (RNA-seq) are in wide use because of their favorable
Standards, Guidelines and Best Practices for RNA-Seq V1.0 (June 2011) The ENCODE Consortium I. Introduction: Sequence based assays of transcriptomes (RNA-seq) are in wide use because of their favorable
FlipFlop: Fast Lasso-based Isoform Prediction as a Flow Problem
 FlipFlop: Fast Lasso-based Isoform Prediction as a Flow Problem Elsa Bernard Laurent Jacob Julien Mairal Jean-Philippe Vert September 24, 2013 Abstract FlipFlop implements a fast method for de novo transcript
FlipFlop: Fast Lasso-based Isoform Prediction as a Flow Problem Elsa Bernard Laurent Jacob Julien Mairal Jean-Philippe Vert September 24, 2013 Abstract FlipFlop implements a fast method for de novo transcript
GeneSifter: Next Generation Data Management and Analysis for Next Generation Sequencing
 for Next Generation Sequencing Dale Baskin, N. Eric Olson, Laura Lucas, Todd Smith 1 Abstract Next generation sequencing technology is rapidly changing the way laboratories and researchers approach the
for Next Generation Sequencing Dale Baskin, N. Eric Olson, Laura Lucas, Todd Smith 1 Abstract Next generation sequencing technology is rapidly changing the way laboratories and researchers approach the
G E N OM I C S S E RV I C ES
 GENOMICS SERVICES THE NEW YORK GENOME CENTER NYGC is an independent non-profit implementing advanced genomic research to improve diagnosis and treatment of serious diseases. capabilities. N E X T- G E
GENOMICS SERVICES THE NEW YORK GENOME CENTER NYGC is an independent non-profit implementing advanced genomic research to improve diagnosis and treatment of serious diseases. capabilities. N E X T- G E
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
Profiling of non-coding RNA classes Gunter Meister
 Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Lecture 8. Protein Trafficking/Targeting. Protein targeting is necessary for proteins that are destined to work outside the cytoplasm.
 Protein Trafficking/Targeting (8.1) Lecture 8 Protein Trafficking/Targeting Protein targeting is necessary for proteins that are destined to work outside the cytoplasm. Protein targeting is more complex
Protein Trafficking/Targeting (8.1) Lecture 8 Protein Trafficking/Targeting Protein targeting is necessary for proteins that are destined to work outside the cytoplasm. Protein targeting is more complex
Frequently Asked Questions Next Generation Sequencing
 Frequently Asked Questions Next Generation Sequencing Import These Frequently Asked Questions for Next Generation Sequencing are some of the more common questions our customers ask. Questions are divided
Frequently Asked Questions Next Generation Sequencing Import These Frequently Asked Questions for Next Generation Sequencing are some of the more common questions our customers ask. Questions are divided
CRAC: An integrated approach to analyse RNA-seq reads Additional File 3 Results on simulated RNA-seq data.
 : An integrated approach to analyse RNA-seq reads Additional File 3 Results on simulated RNA-seq data. Nicolas Philippe and Mikael Salson and Thérèse Commes and Eric Rivals February 13, 2013 1 Results
: An integrated approach to analyse RNA-seq reads Additional File 3 Results on simulated RNA-seq data. Nicolas Philippe and Mikael Salson and Thérèse Commes and Eric Rivals February 13, 2013 1 Results
Control of Gene Expression
 Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Using Illumina BaseSpace Apps to Analyze RNA Sequencing Data
 Using Illumina BaseSpace Apps to Analyze RNA Sequencing Data The Illumina TopHat Alignment and Cufflinks Assembly and Differential Expression apps make RNA data analysis accessible to any user, regardless
Using Illumina BaseSpace Apps to Analyze RNA Sequencing Data The Illumina TopHat Alignment and Cufflinks Assembly and Differential Expression apps make RNA data analysis accessible to any user, regardless
Protein Synthesis How Genes Become Constituent Molecules
 Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Micro RNAs: potentielle Biomarker für das. Blutspenderscreening
 Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Introduction to NGS data analysis
 Introduction to NGS data analysis Jeroen F. J. Laros Leiden Genome Technology Center Department of Human Genetics Center for Human and Clinical Genetics Sequencing Illumina platforms Characteristics: High
Introduction to NGS data analysis Jeroen F. J. Laros Leiden Genome Technology Center Department of Human Genetics Center for Human and Clinical Genetics Sequencing Illumina platforms Characteristics: High
17 July 2014 WEB-SERVER MANUAL. Contact: Michael Hackenberg ([email protected])
 WEB-SERVER MANUAL Contact: Michael Hackenberg ([email protected]) 1 1 Introduction srnabench is a free web-server tool and standalone application for processing small- RNA data obtained from next generation
WEB-SERVER MANUAL Contact: Michael Hackenberg ([email protected]) 1 1 Introduction srnabench is a free web-server tool and standalone application for processing small- RNA data obtained from next generation
The RNA strategy. RNA as a tool and target in human disease diagnosis and therapy.
 The RNA strategy RNA as a tool and target in human disease diagnosis and therapy. The Laboratory of RNA Biology and Biotechnology at the Centre for Integrative Biology (CIBIO) of the University of Trento,
The RNA strategy RNA as a tool and target in human disease diagnosis and therapy. The Laboratory of RNA Biology and Biotechnology at the Centre for Integrative Biology (CIBIO) of the University of Trento,
Bioinformatics Resources at a Glance
 Bioinformatics Resources at a Glance A Note about FASTA Format There are MANY free bioinformatics tools available online. Bioinformaticists have developed a standard format for nucleotide and protein sequences
Bioinformatics Resources at a Glance A Note about FASTA Format There are MANY free bioinformatics tools available online. Bioinformaticists have developed a standard format for nucleotide and protein sequences
Single-Cell DNA Sequencing with the C 1. Single-Cell Auto Prep System. Reveal hidden populations and genetic diversity within complex samples
 DATA Sheet Single-Cell DNA Sequencing with the C 1 Single-Cell Auto Prep System Reveal hidden populations and genetic diversity within complex samples Single-cell sensitivity Discover and detect SNPs,
DATA Sheet Single-Cell DNA Sequencing with the C 1 Single-Cell Auto Prep System Reveal hidden populations and genetic diversity within complex samples Single-cell sensitivity Discover and detect SNPs,
Methods, tools, and pipelines for analysis of Ion PGM Sequencer mirna and gene expression data
 WHITE PAPER Ion RNA-Seq Methods, tools, and pipelines for analysis of Ion PGM Sequencer mirna and gene expression data Introduction High-resolution measurements of transcriptional activity and organization
WHITE PAPER Ion RNA-Seq Methods, tools, and pipelines for analysis of Ion PGM Sequencer mirna and gene expression data Introduction High-resolution measurements of transcriptional activity and organization
BIOL 3200 Spring 2015 DNA Subway and RNA-Seq Data Analysis
 BIOL 3200 Spring 2015 DNA Subway and RNA-Seq Data Analysis By the end of this lab students should be able to: Describe the uses for each line of the DNA subway program (Red/Yellow/Blue/Green) Describe
BIOL 3200 Spring 2015 DNA Subway and RNA-Seq Data Analysis By the end of this lab students should be able to: Describe the uses for each line of the DNA subway program (Red/Yellow/Blue/Green) Describe
Module 1. Sequence Formats and Retrieval. Charles Steward
 The Open Door Workshop Module 1 Sequence Formats and Retrieval Charles Steward 1 Aims Acquaint you with different file formats and associated annotations. Introduce different nucleotide and protein databases.
The Open Door Workshop Module 1 Sequence Formats and Retrieval Charles Steward 1 Aims Acquaint you with different file formats and associated annotations. Introduce different nucleotide and protein databases.
Shouguo Gao Ph. D Department of Physics and Comprehensive Diabetes Center
 Computational Challenges in Storage, Analysis and Interpretation of Next-Generation Sequencing Data Shouguo Gao Ph. D Department of Physics and Comprehensive Diabetes Center Next Generation Sequencing
Computational Challenges in Storage, Analysis and Interpretation of Next-Generation Sequencing Data Shouguo Gao Ph. D Department of Physics and Comprehensive Diabetes Center Next Generation Sequencing
Name Date Period. 2. When a molecule of double-stranded DNA undergoes replication, it results in
 DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
InSyBio BioNets: Utmost efficiency in gene expression data and biological networks analysis
 InSyBio BioNets: Utmost efficiency in gene expression data and biological networks analysis WHITE PAPER By InSyBio Ltd Konstantinos Theofilatos Bioinformatician, PhD InSyBio Technical Sales Manager August
InSyBio BioNets: Utmost efficiency in gene expression data and biological networks analysis WHITE PAPER By InSyBio Ltd Konstantinos Theofilatos Bioinformatician, PhD InSyBio Technical Sales Manager August
New Technologies for Sensitive, Low-Input RNA-Seq. Clontech Laboratories, Inc.
 New Technologies for Sensitive, Low-Input RNA-Seq Clontech Laboratories, Inc. Outline Introduction Single-Cell-Capable mrna-seq Using SMART Technology SMARTer Ultra Low RNA Kit for the Fluidigm C 1 System
New Technologies for Sensitive, Low-Input RNA-Seq Clontech Laboratories, Inc. Outline Introduction Single-Cell-Capable mrna-seq Using SMART Technology SMARTer Ultra Low RNA Kit for the Fluidigm C 1 System
Guide for Data Visualization and Analysis using ACSN
 Guide for Data Visualization and Analysis using ACSN ACSN contains the NaviCell tool box, the intuitive and user- friendly environment for data visualization and analysis. The tool is accessible from the
Guide for Data Visualization and Analysis using ACSN ACSN contains the NaviCell tool box, the intuitive and user- friendly environment for data visualization and analysis. The tool is accessible from the
Viruses. Viral components: Capsid. Chapter 10: Viruses. Viral components: Nucleic Acid. Viral components: Envelope
 Viruses Chapter 10: Viruses Lecture Exam #3 Wednesday, November 22 nd (This lecture WILL be on Exam #3) Dr. Amy Rogers Office Hours: MW 9-10 AM Too small to see with a light microscope Visible with electron
Viruses Chapter 10: Viruses Lecture Exam #3 Wednesday, November 22 nd (This lecture WILL be on Exam #3) Dr. Amy Rogers Office Hours: MW 9-10 AM Too small to see with a light microscope Visible with electron
Services. Updated 05/31/2016
 Updated 05/31/2016 Services 1. Whole exome sequencing... 2 2. Whole Genome Sequencing (WGS)... 3 3. 16S rrna sequencing... 4 4. Customized gene panels... 5 5. RNA-Seq... 6 6. qpcr... 7 7. HLA typing...
Updated 05/31/2016 Services 1. Whole exome sequencing... 2 2. Whole Genome Sequencing (WGS)... 3 3. 16S rrna sequencing... 4 4. Customized gene panels... 5 5. RNA-Seq... 6 6. qpcr... 7 7. HLA typing...
RETRIEVING SEQUENCE INFORMATION. Nucleotide sequence databases. Database search. Sequence alignment and comparison
 RETRIEVING SEQUENCE INFORMATION Nucleotide sequence databases Database search Sequence alignment and comparison Biological sequence databases Originally just a storage place for sequences. Currently the
RETRIEVING SEQUENCE INFORMATION Nucleotide sequence databases Database search Sequence alignment and comparison Biological sequence databases Originally just a storage place for sequences. Currently the
Using Galaxy for NGS Analysis. Daniel Blankenberg Postdoctoral Research Associate The Galaxy Team http://usegalaxy.org
 Using Galaxy for NGS Analysis Daniel Blankenberg Postdoctoral Research Associate The Galaxy Team http://usegalaxy.org Overview NGS Data Galaxy tools for NGS Data Galaxy for Sequencing Facilities Overview
Using Galaxy for NGS Analysis Daniel Blankenberg Postdoctoral Research Associate The Galaxy Team http://usegalaxy.org Overview NGS Data Galaxy tools for NGS Data Galaxy for Sequencing Facilities Overview
RNA-Seq Tutorial 1. John Garbe Research Informatics Support Systems, MSI March 19, 2012
 RNA-Seq Tutorial 1 John Garbe Research Informatics Support Systems, MSI March 19, 2012 Tutorial 1 RNA-Seq Tutorials RNA-Seq experiment design and analysis Instruction on individual software will be provided
RNA-Seq Tutorial 1 John Garbe Research Informatics Support Systems, MSI March 19, 2012 Tutorial 1 RNA-Seq Tutorials RNA-Seq experiment design and analysis Instruction on individual software will be provided
Accelerate genomic breakthroughs in microbiology. Gain deeper insights with powerful bioinformatic tools.
 Accelerate genomic breakthroughs in microbiology. Gain deeper insights with powerful bioinformatic tools. Empowering microbial genomics. Extensive methods. Expansive possibilities. In microbiome studies
Accelerate genomic breakthroughs in microbiology. Gain deeper insights with powerful bioinformatic tools. Empowering microbial genomics. Extensive methods. Expansive possibilities. In microbiome studies
ISTEP+: Biology I End-of-Course Assessment Released Items and Scoring Notes
 ISTEP+: Biology I End-of-Course Assessment Released Items and Scoring Notes Page 1 of 22 Introduction Indiana students enrolled in Biology I participated in the ISTEP+: Biology I Graduation Examination
ISTEP+: Biology I End-of-Course Assessment Released Items and Scoring Notes Page 1 of 22 Introduction Indiana students enrolled in Biology I participated in the ISTEP+: Biology I Graduation Examination
2007 7.013 Problem Set 1 KEY
 2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
CHAPTER 40 The Mechanism of Protein Synthesis
 CHAPTER 40 The Mechanism of Protein Synthesis Problems: 2,3,6,7,9,13,14,15,18,19,20 Initiation: Locating the start codon. Elongation: Reading the codons (5 3 ) and synthesizing protein amino carboxyl.
CHAPTER 40 The Mechanism of Protein Synthesis Problems: 2,3,6,7,9,13,14,15,18,19,20 Initiation: Locating the start codon. Elongation: Reading the codons (5 3 ) and synthesizing protein amino carboxyl.
BIOINF 525 Winter 2016 Foundations of Bioinformatics and Systems Biology http://tinyurl.com/bioinf525-w16
 Course Director: Dr. Barry Grant (DCM&B, [email protected]) Description: This is a three module course covering (1) Foundations of Bioinformatics, (2) Statistics in Bioinformatics, and (3) Systems
Course Director: Dr. Barry Grant (DCM&B, [email protected]) Description: This is a three module course covering (1) Foundations of Bioinformatics, (2) Statistics in Bioinformatics, and (3) Systems
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
The Steps. 1. Transcription. 2. Transferal. 3. Translation
 Protein Synthesis Protein synthesis is simply the "making of proteins." Although the term itself is easy to understand, the multiple steps that a cell in a plant or animal must go through are not. In order
Protein Synthesis Protein synthesis is simply the "making of proteins." Although the term itself is easy to understand, the multiple steps that a cell in a plant or animal must go through are not. In order
Comparing Methods for Identifying Transcription Factor Target Genes
 Comparing Methods for Identifying Transcription Factor Target Genes Alena van Bömmel (R 3.3.73) Matthew Huska (R 3.3.18) Max Planck Institute for Molecular Genetics Folie 1 Transcriptional Regulation TF
Comparing Methods for Identifying Transcription Factor Target Genes Alena van Bömmel (R 3.3.73) Matthew Huska (R 3.3.18) Max Planck Institute for Molecular Genetics Folie 1 Transcriptional Regulation TF
Translation Study Guide
 Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Quick Hit Activity Using UIL Science Contests For Formative and Summative Assessments of Pre-AP and AP Biology Students
 Quick Hit Activity Using UIL Science Contests For Formative and Summative Assessments of Pre-AP and AP Biology Students Activity Title: Quick Hit Goal of Activity: To perform formative and summative assessments
Quick Hit Activity Using UIL Science Contests For Formative and Summative Assessments of Pre-AP and AP Biology Students Activity Title: Quick Hit Goal of Activity: To perform formative and summative assessments
Delivering the power of the world s most successful genomics platform
 Delivering the power of the world s most successful genomics platform NextCODE Health is bringing the full power of the world s largest and most successful genomics platform to everyday clinical care NextCODE
Delivering the power of the world s most successful genomics platform NextCODE Health is bringing the full power of the world s largest and most successful genomics platform to everyday clinical care NextCODE
Bioruptor NGS: Unbiased DNA shearing for Next-Generation Sequencing
 STGAAC STGAACT GTGCACT GTGAACT STGAAC STGAACT GTGCACT GTGAACT STGAAC STGAAC GTGCAC GTGAAC Wouter Coppieters Head of the genomics core facility GIGA center, University of Liège Bioruptor NGS: Unbiased DNA
STGAAC STGAACT GTGCACT GTGAACT STGAAC STGAACT GTGCACT GTGAACT STGAAC STGAAC GTGCAC GTGAAC Wouter Coppieters Head of the genomics core facility GIGA center, University of Liège Bioruptor NGS: Unbiased DNA
RNA Express. Introduction 3 Run RNA Express 4 RNA Express App Output 6 RNA Express Workflow 12 Technical Assistance
 RNA Express Introduction 3 Run RNA Express 4 RNA Express App Output 6 RNA Express Workflow 12 Technical Assistance ILLUMINA PROPRIETARY 15052918 Rev. A February 2014 This document and its contents are
RNA Express Introduction 3 Run RNA Express 4 RNA Express App Output 6 RNA Express Workflow 12 Technical Assistance ILLUMINA PROPRIETARY 15052918 Rev. A February 2014 This document and its contents are
A Tutorial in Genetic Sequence Classification Tools and Techniques
 A Tutorial in Genetic Sequence Classification Tools and Techniques Jake Drew Data Mining CSE 8331 Southern Methodist University [email protected] www.jakemdrew.com Sequence Characters IUPAC nucleotide
A Tutorial in Genetic Sequence Classification Tools and Techniques Jake Drew Data Mining CSE 8331 Southern Methodist University [email protected] www.jakemdrew.com Sequence Characters IUPAC nucleotide
Ingenuity Pathway Analysis (IPA )
 ProductProfile Ingenuity Pathway Analysis (IPA ) For the analysis and interpretation of omics data IPA is a web-based software application for the analysis, integration, and interpretation of data derived
ProductProfile Ingenuity Pathway Analysis (IPA ) For the analysis and interpretation of omics data IPA is a web-based software application for the analysis, integration, and interpretation of data derived
Analysis and Integration of Big Data from Next-Generation Genomics, Epigenomics, and Transcriptomics
 Analysis and Integration of Big Data from Next-Generation Genomics, Epigenomics, and Transcriptomics Christopher Benner, PhD Director, Integrative Genomics and Bioinformatics Core (IGC) idash Webinar,
Analysis and Integration of Big Data from Next-Generation Genomics, Epigenomics, and Transcriptomics Christopher Benner, PhD Director, Integrative Genomics and Bioinformatics Core (IGC) idash Webinar,
BME 42-620 Engineering Molecular Cell Biology. Lecture 02: Structural and Functional Organization of
 BME 42-620 Engineering Molecular Cell Biology Lecture 02: Structural and Functional Organization of Eukaryotic Cells BME42-620 Lecture 02, September 01, 2011 1 Outline A brief review of the previous lecture
BME 42-620 Engineering Molecular Cell Biology Lecture 02: Structural and Functional Organization of Eukaryotic Cells BME42-620 Lecture 02, September 01, 2011 1 Outline A brief review of the previous lecture
Core Facility Genomics
 Core Facility Genomics versatile genome or transcriptome analyses based on quantifiable highthroughput data ascertainment 1 Topics Collaboration with Harald Binder and Clemens Kreutz Project: Microarray
Core Facility Genomics versatile genome or transcriptome analyses based on quantifiable highthroughput data ascertainment 1 Topics Collaboration with Harald Binder and Clemens Kreutz Project: Microarray
ProteinQuest user guide
 ProteinQuest user guide 1. Introduction... 3 1.1 With ProteinQuest you can... 3 1.2 ProteinQuest basic version 4 1.3 ProteinQuest extended version... 5 2. ProteinQuest dictionaries... 6 3. Directions for
ProteinQuest user guide 1. Introduction... 3 1.1 With ProteinQuest you can... 3 1.2 ProteinQuest basic version 4 1.3 ProteinQuest extended version... 5 2. ProteinQuest dictionaries... 6 3. Directions for
Challenges associated with analysis and storage of NGS data
 Challenges associated with analysis and storage of NGS data Gabriella Rustici Research and training coordinator Functional Genomics Group [email protected] Next-generation sequencing Next-generation sequencing
Challenges associated with analysis and storage of NGS data Gabriella Rustici Research and training coordinator Functional Genomics Group [email protected] Next-generation sequencing Next-generation sequencing
School of Nursing. Presented by Yvette Conley, PhD
 Presented by Yvette Conley, PhD What we will cover during this webcast: Briefly discuss the approaches introduced in the paper: Genome Sequencing Genome Wide Association Studies Epigenomics Gene Expression
Presented by Yvette Conley, PhD What we will cover during this webcast: Briefly discuss the approaches introduced in the paper: Genome Sequencing Genome Wide Association Studies Epigenomics Gene Expression
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals
 Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Nazneen Aziz, PhD. Director, Molecular Medicine Transformation Program Office
 2013 Laboratory Accreditation Program Audioconferences and Webinars Implementing Next Generation Sequencing (NGS) as a Clinical Tool in the Laboratory Nazneen Aziz, PhD Director, Molecular Medicine Transformation
2013 Laboratory Accreditation Program Audioconferences and Webinars Implementing Next Generation Sequencing (NGS) as a Clinical Tool in the Laboratory Nazneen Aziz, PhD Director, Molecular Medicine Transformation
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
GenBank, Entrez, & FASTA
 GenBank, Entrez, & FASTA Nucleotide Sequence Databases First generation GenBank is a representative example started as sort of a museum to preserve knowledge of a sequence from first discovery great repositories,
GenBank, Entrez, & FASTA Nucleotide Sequence Databases First generation GenBank is a representative example started as sort of a museum to preserve knowledge of a sequence from first discovery great repositories,
mrna NGS Data Analysis Report
 mrna NGS Data Analysis Report Project: Test Project (Ref code: 00001) Customer: Test customer Company/Institute: Exiqon Date: Monday, June 29, 2015 Performed by: XploreRNA Exiqon A/S Company Reg. No. (CVR)
mrna NGS Data Analysis Report Project: Test Project (Ref code: 00001) Customer: Test customer Company/Institute: Exiqon Date: Monday, June 29, 2015 Performed by: XploreRNA Exiqon A/S Company Reg. No. (CVR)
MicroRNA formation. 4th International Symposium on Non-Surgical Contraceptive Methods of Pet Population Control
 MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
Next Generation Sequencing: Adjusting to Big Data. Daniel Nicorici, Dr.Tech. Statistikot Suomen Lääketeollisuudessa 29.10.2013
 Next Generation Sequencing: Adjusting to Big Data Daniel Nicorici, Dr.Tech. Statistikot Suomen Lääketeollisuudessa 29.10.2013 Outline Human Genome Project Next-Generation Sequencing Personalized Medicine
Next Generation Sequencing: Adjusting to Big Data Daniel Nicorici, Dr.Tech. Statistikot Suomen Lääketeollisuudessa 29.10.2013 Outline Human Genome Project Next-Generation Sequencing Personalized Medicine
Special report. Chronic Lymphocytic Leukemia (CLL) Genomic Biology 3020 April 20, 2006
 Special report Chronic Lymphocytic Leukemia (CLL) Genomic Biology 3020 April 20, 2006 Gene And Protein The gene that causes the mutation is CCND1 and the protein NP_444284 The mutation deals with the cell
Special report Chronic Lymphocytic Leukemia (CLL) Genomic Biology 3020 April 20, 2006 Gene And Protein The gene that causes the mutation is CCND1 and the protein NP_444284 The mutation deals with the cell
LifeScope Genomic Analysis Software 2.5
 USER GUIDE LifeScope Genomic Analysis Software 2.5 Graphical User Interface DATA ANALYSIS METHODS AND INTERPRETATION Publication Part Number 4471877 Rev. A Revision Date November 2011 For Research Use
USER GUIDE LifeScope Genomic Analysis Software 2.5 Graphical User Interface DATA ANALYSIS METHODS AND INTERPRETATION Publication Part Number 4471877 Rev. A Revision Date November 2011 For Research Use
SEQUENCING. From Sample to Sequence-Ready
 SEQUENCING From Sample to Sequence-Ready ACCESS ARRAY SYSTEM HIGH-QUALITY LIBRARIES, NOT ONCE, BUT EVERY TIME The highest-quality amplicons more sensitive, accurate, and specific Full support for all major
SEQUENCING From Sample to Sequence-Ready ACCESS ARRAY SYSTEM HIGH-QUALITY LIBRARIES, NOT ONCE, BUT EVERY TIME The highest-quality amplicons more sensitive, accurate, and specific Full support for all major
Bioinformatics Unit Department of Biological Services. Get to know us
 Bioinformatics Unit Department of Biological Services Get to know us Domains of Activity IT & programming Microarray analysis Sequence analysis Bioinformatics Team Biostatistical support NGS data analysis
Bioinformatics Unit Department of Biological Services Get to know us Domains of Activity IT & programming Microarray analysis Sequence analysis Bioinformatics Team Biostatistical support NGS data analysis
Molecular Genetics. RNA, Transcription, & Protein Synthesis
 Molecular Genetics RNA, Transcription, & Protein Synthesis Section 1 RNA AND TRANSCRIPTION Objectives Describe the primary functions of RNA Identify how RNA differs from DNA Describe the structure and
Molecular Genetics RNA, Transcription, & Protein Synthesis Section 1 RNA AND TRANSCRIPTION Objectives Describe the primary functions of RNA Identify how RNA differs from DNA Describe the structure and
UGENE Quick Start Guide
 Quick Start Guide This document contains a quick introduction to UGENE. For more detailed information, you can find the UGENE User Manual and other special manuals in project website: http://ugene.unipro.ru.
Quick Start Guide This document contains a quick introduction to UGENE. For more detailed information, you can find the UGENE User Manual and other special manuals in project website: http://ugene.unipro.ru.
Control of Gene Expression
 Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
Non-invasive prenatal detection of chromosome aneuploidies using next generation sequencing: First steps towards clinical application
 Non-invasive prenatal detection of chromosome aneuploidies using next generation sequencing: First steps towards clinical application PD Dr. rer. nat. Markus Stumm Zentrum für Pränataldiagnostik Kudamm-199
Non-invasive prenatal detection of chromosome aneuploidies using next generation sequencing: First steps towards clinical application PD Dr. rer. nat. Markus Stumm Zentrum für Pränataldiagnostik Kudamm-199
SICKLE CELL ANEMIA & THE HEMOGLOBIN GENE TEACHER S GUIDE
 AP Biology Date SICKLE CELL ANEMIA & THE HEMOGLOBIN GENE TEACHER S GUIDE LEARNING OBJECTIVES Students will gain an appreciation of the physical effects of sickle cell anemia, its prevalence in the population,
AP Biology Date SICKLE CELL ANEMIA & THE HEMOGLOBIN GENE TEACHER S GUIDE LEARNING OBJECTIVES Students will gain an appreciation of the physical effects of sickle cell anemia, its prevalence in the population,
Review of the Cell and Its Organelles
 Biology Learning Centre Review of the Cell and Its Organelles Tips for most effective learning of this material: Memorize the names and structures over several days. This will help you retain what you
Biology Learning Centre Review of the Cell and Its Organelles Tips for most effective learning of this material: Memorize the names and structures over several days. This will help you retain what you
Data Analysis for Ion Torrent Sequencing
 IFU022 v140202 Research Use Only Instructions For Use Part III Data Analysis for Ion Torrent Sequencing MANUFACTURER: Multiplicom N.V. Galileilaan 18 2845 Niel Belgium Revision date: August 21, 2014 Page
IFU022 v140202 Research Use Only Instructions For Use Part III Data Analysis for Ion Torrent Sequencing MANUFACTURER: Multiplicom N.V. Galileilaan 18 2845 Niel Belgium Revision date: August 21, 2014 Page
Algorithms in Computational Biology (236522) spring 2007 Lecture #1
 Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
Version 5.0 Release Notes
 Version 5.0 Release Notes 2011 Gene Codes Corporation Gene Codes Corporation 775 Technology Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074 (fax) www.genecodes.com
Version 5.0 Release Notes 2011 Gene Codes Corporation Gene Codes Corporation 775 Technology Drive, Ann Arbor, MI 48108 USA 1.800.497.4939 (USA) +1.734.769.7249 (elsewhere) +1.734.769.7074 (fax) www.genecodes.com
mrna EDITING Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A. Copyright 2005
 mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
Single-Cell Whole Genome Sequencing on the C1 System: a Performance Evaluation
 PN 100-9879 A1 TECHNICAL NOTE Single-Cell Whole Genome Sequencing on the C1 System: a Performance Evaluation Introduction Cancer is a dynamic evolutionary process of which intratumor genetic and phenotypic
PN 100-9879 A1 TECHNICAL NOTE Single-Cell Whole Genome Sequencing on the C1 System: a Performance Evaluation Introduction Cancer is a dynamic evolutionary process of which intratumor genetic and phenotypic
Next Generation Sequencing
 Next Generation Sequencing Technology and applications 10/1/2015 Jeroen Van Houdt - Genomics Core - KU Leuven - UZ Leuven 1 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977
Next Generation Sequencing Technology and applications 10/1/2015 Jeroen Van Houdt - Genomics Core - KU Leuven - UZ Leuven 1 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977
Identification of rheumatoid arthritis and osteoarthritis patients by transcriptome-based rule set generation
 Identification of rheumatoid arthritis and osterthritis patients by transcriptome-based rule set generation Bering Limited Report generated on September 19, 2014 Contents 1 Dataset summary 2 1.1 Project
Identification of rheumatoid arthritis and osterthritis patients by transcriptome-based rule set generation Bering Limited Report generated on September 19, 2014 Contents 1 Dataset summary 2 1.1 Project
BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS
 BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS 1. The Technology Strategy sets out six areas where technological developments are required to push the frontiers of knowledge
BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS 1. The Technology Strategy sets out six areas where technological developments are required to push the frontiers of knowledge
Go where the biology takes you. Genome Analyzer IIx Genome Analyzer IIe
 Go where the biology takes you. Genome Analyzer IIx Genome Analyzer IIe Go where the biology takes you. To published results faster With proven scalability To the forefront of discovery To limitless applications
Go where the biology takes you. Genome Analyzer IIx Genome Analyzer IIe Go where the biology takes you. To published results faster With proven scalability To the forefront of discovery To limitless applications
Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Biochemistry
 Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Biochemistry The Master Degree in Medical Laboratory Sciences /Clinical Biochemistry, is awarded by the Faculty of Graduate Studies
Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Biochemistry The Master Degree in Medical Laboratory Sciences /Clinical Biochemistry, is awarded by the Faculty of Graduate Studies
An Introduction to Next-Generation Sequencing for in vitro Fertilization
 An Introduction to Next-Generation Sequencing for in vitro Fertilization www.illumina.com/ivfprimer Table of Contents Part I. Welcome to Next-Generation Sequencing 3 NGS for in vitro Fertilization 3 Part
An Introduction to Next-Generation Sequencing for in vitro Fertilization www.illumina.com/ivfprimer Table of Contents Part I. Welcome to Next-Generation Sequencing 3 NGS for in vitro Fertilization 3 Part
trna Processing and Modification
 trna Processing and Modification RNA POL III - TRANSCRIPTS 5S RNA, trna, repetitive Sequenzen (Alu-typ), versch. kleine stabile RNAs (7SL - RNA vom signal recognition particle (SRP)), U6 RNA 5S RNA nicht
trna Processing and Modification RNA POL III - TRANSCRIPTS 5S RNA, trna, repetitive Sequenzen (Alu-typ), versch. kleine stabile RNAs (7SL - RNA vom signal recognition particle (SRP)), U6 RNA 5S RNA nicht
Data Analysis & Management of High-throughput Sequencing Data. Quoclinh Nguyen Research Informatics Genomics Core / Medical Research Institute
 Data Analysis & Management of High-throughput Sequencing Data Quoclinh Nguyen Research Informatics Genomics Core / Medical Research Institute Current Issues Current Issues The QSEQ file Number files per
Data Analysis & Management of High-throughput Sequencing Data Quoclinh Nguyen Research Informatics Genomics Core / Medical Research Institute Current Issues Current Issues The QSEQ file Number files per
Dr Alexander Henzing
 Horizon 2020 Health, Demographic Change & Wellbeing EU funding, research and collaboration opportunities for 2016/17 Innovate UK funding opportunities in omics, bridging health and life sciences Dr Alexander
Horizon 2020 Health, Demographic Change & Wellbeing EU funding, research and collaboration opportunities for 2016/17 Innovate UK funding opportunities in omics, bridging health and life sciences Dr Alexander
Cells & Cell Organelles
 Cells & Cell Organelles The Building Blocks of Life H Biology Types of cells bacteria cells Prokaryote - no organelles Eukaryotes - organelles animal cells plant cells Cell size comparison Animal cell
Cells & Cell Organelles The Building Blocks of Life H Biology Types of cells bacteria cells Prokaryote - no organelles Eukaryotes - organelles animal cells plant cells Cell size comparison Animal cell
Next generation DNA sequencing technologies. theory & prac-ce
 Next generation DNA sequencing technologies theory & prac-ce Outline Next- Genera-on sequencing (NGS) technologies overview NGS applica-ons NGS workflow: data collec-on and processing the exome sequencing
Next generation DNA sequencing technologies theory & prac-ce Outline Next- Genera-on sequencing (NGS) technologies overview NGS applica-ons NGS workflow: data collec-on and processing the exome sequencing
How Sequencing Experiments Fail
 How Sequencing Experiments Fail v1.0 Simon Andrews [email protected] Classes of Failure Technical Tracking Library Contamination Biological Interpretation Something went wrong with a machine
How Sequencing Experiments Fail v1.0 Simon Andrews [email protected] Classes of Failure Technical Tracking Library Contamination Biological Interpretation Something went wrong with a machine
Basic processing of next-generation sequencing (NGS) data
 Basic processing of next-generation sequencing (NGS) data Getting from raw sequence data to expression analysis! 1 Reminder: we are measuring expression of protein coding genes by transcript abundance
Basic processing of next-generation sequencing (NGS) data Getting from raw sequence data to expression analysis! 1 Reminder: we are measuring expression of protein coding genes by transcript abundance
GeneProf and the new GeneProf Web Services
 GeneProf and the new GeneProf Web Services Florian Halbritter [email protected] Stem Cell Bioinformatics Group (Simon R. Tomlinson) [email protected] December 10, 2012 Florian Halbritter
GeneProf and the new GeneProf Web Services Florian Halbritter [email protected] Stem Cell Bioinformatics Group (Simon R. Tomlinson) [email protected] December 10, 2012 Florian Halbritter
RNAseq / ChipSeq / Methylseq and personalized genomics
 RNAseq / ChipSeq / Methylseq and personalized genomics 7711 Lecture Subhajyo) De, PhD Division of Biomedical Informa)cs and Personalized Biomedicine, Department of Medicine University of Colorado School
RNAseq / ChipSeq / Methylseq and personalized genomics 7711 Lecture Subhajyo) De, PhD Division of Biomedical Informa)cs and Personalized Biomedicine, Department of Medicine University of Colorado School
