genes, widespread transcription
|
|
|
- Shonda Daniels
- 8 years ago
- Views:
Transcription
1 Leading Edge Essay TUF Love for Junk DNA Aarron T. Willingham 1 and Thomas R. Gingeras 1, * 1 Affymetrix, Inc., Santa Clara, CA 95051, USA *Contact: tom_gingeras@affymetrix.com DOI /j.cell The widespread occurrence of noncoding (nc) RNAs unannotated eukaryotic transcripts with reduced protein coding potential suggests that they are functionally important. Study of ncrnas is increasing our understanding of the organization and regulation of genomes. Over the past five years, researchers working with various organisms and using multiple technologies to explore genomewide gene expression have converged on the same surprising conclusion: is widespread throughout the genome and many-fold higher than existing genome annotations would predict. The burgeoning number of these transcripts of unknown function, or TUFs (Cheng et al., 2005), highlights a remarkably complex al architecture that includes alternative splice isoforms for almost all proteincoding genes, widespread of antisense RNAs, and abundant noncoding RNAs (ncrnas) with important biological functions. By some estimates, TUFs could rival protein-coding transcripts in number (Cawley et al., 2004). Such al diversity may explain how the relatively similar numbers of proteincoding genes estimated for fruit fly (13,985; BDGP release 4), nematode worm (21,009; Wormbase release 150), and human (23,341; NCBI release 36) result in the remarkable phenotypic differences observed among these species. Widespread Transcription Although recent evidence indicates that the complexity of transcripts produced by the human genome and the underlying al architecture is striking, these observations are consistent with much earlier reports. Decades ago, several studies discovered evidence for widespread (see Table S1 available with this article online). In the 1970s, studies involving sea urchin embryos found that heterogeneous nuclear RNA had 10-fold more nucleotide complexity than cytoplasmic mrna associated with polysomes. Studies of lampbrush chromosomes in newt oocytes estimated that empirically measured was an order of magnitude greater than necessary to produce mrnas for the oocyte. An abundance of nonpolyadenylated polysome-associated RNAs (?30% of cytoplasmic RNAs) was first identified in HeLa cells and subsequently observed in a variety of plant and animal cells. Furthermore, a 10-fold greater complexity of nuclear versus cytoplasmic polyadenylated (polya) RNA was also found in human cells, and 5 cap structures outnumber polya segments 3:1 in hamster ovary cells. These original studies arrived at the common conclusion that the complexity of transcripts made by most organisms seemed to be inexplicably larger than expected. Further insight into the nature of this al diversity awaited advancements in our understanding of primary structure of genomes and development of new analytical technologies. Genomic tiling arrays represent an unbiased and sensitive tool for global studies of. Such a technology offers independence from current limited genome annotations while enabling detection of rare transcripts. Using genome tiling microarrays, widespread along chromosomes and across the human genome has been observed, including significant antisense products, and may be an order of magnitude greater than annotation-based predictions (reviewed in Johnson et al., 2005). In 2002, in a systematic analysis of across human chromosomes 21 and 22, our group observed about an order of magnitude more al activity than could be accounted for by predicted protein-coding genes, suggesting that a significant portion of transcribed cytoplasmic polya RNA may indeed be noncoding. A subsequent human genomewide mapping effort of polya RNA from human liver reached similar conclusions and identified >10,000 new transcripts, many of which have homology to genomic sequences in other mammalian species. Tiling array-based whole-genome mapping in the model plant Arabidopsis found that >50% of observed was intergenic and that?30% of annotated genes had associated antisense, some of which was tissue-specific. Furthermore, about 20% of annotated pseudogenes were expressed, suggesting that examples of pseudogene-mediated regulation of gene activity may be common (Hirotsune et al., 2003). In Drosophila,?40% of probes in intronic and intergenic areas detected RNA expression, much of which changed in a developmentally coordinated manner. Furthermore, alternative splicing was observed in?40% of known genes, yielding over 5000 new splice forms. Recently, even the small, well-characterized yeast genome has yielded a more complex transcriptome than expected, with overlapping and differential expression levels even within the same gene (David et al., 2006). Together, these studies provide several observations about transcriptomes: (1) the widespread incidence of unannotated transcripts Cell 125, June 30, Elsevier Inc. 1215
2 with limited protein-coding capacity, often expressed at low levels; (2) a large degree of overlapping, evidenced in part by the presence of abundant antisense ; and (3) most coding genes have alternative splice forms. Combining chromatin immunoprecipitation (ChIP) and tiling microarrays has allowed unbiased, often high-resolution mapping of factor binding sites across the genome, and in the process has lent considerable unbiased empirical support to the existence of widespread unannotated. The binding sites of the NFκB family member RelA/p65 were mapped across human chromosome 22, with nearly 28% of the binding sites shown to lie >50 kb away from known protein-coding genes (Martone et al., 2003). Mapping of DNA binding sites for Sp1, cmyc, and p53 factors on human chromosomes 21 and 22 revealed that almost half of the mapped binding sites potentially correlate with ncrnas, and a significant subset of the noncoding transcripts showed al responsiveness to retinoic acid (Cawley et al., 2004). Recently, a whole-genome map of RNA polymerase II preinitiation complex binding sites in human primary fibroblasts identified?10,000 active promoters, with approximately 13% corresponding to unannotated loci (Kim et al., 2005). Several factors could account for the substantially lower percent of observed binding sites that may correlate with noncoding, such as the highly conservative thresholds used to identify binding sites and the low occupancy by RNA polymerase for low abundance transcripts. Taken together, these results suggest that widespread of TUFs is initiated and regulated by molecular mechanisms similar to those modulating protein-coding RNAs. Widespread unannotated has been confirmed by other experimental means, including mapping 5 ends of transcripts using cap analysis of gene expression (CAGE), 3 ends with serial analysis of gene expression (SAGE), massively parallel signature sequencing (MPSS), and high-throughput fulllength cdna cloning and sequencing methodologies (reviewed in Mattick and Makunin, 2006). These sequencing-based approaches provide strand specificity for transcripts and can detect transcripts originating from repetitive regions of the genome, whereas tiling arrays often have repetitive regions excluded and may use labeling protocols that are strand-insensitive. Expression analysis by SAGE tags found at least 15,000 uncharacterized 3 termini used in the genome, suggesting that new isoforms and genes numbered in the many thousands. In the functional annotation of the mouse genome (FANTOM) project, large-scale sequencing and manual annotation of full-length cdnas characterized 102,281 transcripts, including 32,129 proteincoding transcripts (2222 of which encode new proteins) and 34,030 ncrnas (Carninci et al., 2005). This study also concluded that the total number of transcripts is at least an order of magnitude larger than current estimates of genes present in the mouse genome. Furthermore, only about 40% of characterized ncrna sequences were identified in previous FANTOM cdna collections, suggesting that the number of unannotated transcripts could continue to grow. Sequencing of 3 million human CAGE tags also confirmed that human cells have a similar level of al diversity. Analysis of human gene expression by MPSS found that >65% of signature sequences do not overlap with annotated transcripts; rather, 38% map to introns, 21% are antisense to known exons, and 5% map to intergenic areas. Intergenic transcripts are expressed at low levels on arrays, and it may be that the cloning and sequencing methodologies of MPSS require greater analysis to detect transcripts with such low abundance. Antisense is also observed in cdna sequencing, corroborating findings in microarray studies. Transcription from the opposite strand of a protein-encoding locus produces RNAs that may hybridize with DNA or RNA and could interfere with, translation, or mrna stability of the sense product. Systematic cdna cloning of small RNAs from C. elegans identified 700 distinct endogenous sir- NAs, possibly processed from larger antisense transcripts, precisely complementary to protein-coding regions from more than 500 different genes; such small RNAs may act as sirnas in plants (reviewed in Sontheimer and Carthew, 2005). Genomewide surveys of expressed mouse and human sequences and cdna sequencing experiments identified a surprisingly large number of transcripts antisense to protein-coding genes. This suggests that the majority of sense transcripts (e.g., 72% of mouse genes) may have an antisense partner (Katayama et al., 2005). Noncoding RNAs and Their Functions A key question hangs like an ominous cloud over these observations of widespread : are these transcripts biologically functional, or are they the al noise of a less than precise set of biological processes? Recent experiments in mice in which megabase gene desert regions have been deleted underscore the relevance of this question. Deletion of 1.5 Mb and 0.8 Mb genomic intervals, which together contain 1243 noncoding sequences conserved between rodent and primate, resulted in viable mice with no obvious deleterious phenotypes (Nobrega et al., 2004). However, if history is our guide, then the answer to this question may be complex (see Figure 1). It has taken more than 35 years to identify and characterize seven functional classes of ncrnas: ribosomal (r), transfer (t), small nuclear (sn), antisense (AS), small nucleolar (sno), micro (mi) and Piwi-interacting (pi). With the exception of perhaps the rrna and trna functional classes, the other ncrna classes are believed to 1216 Cell 125, June 30, Elsevier Inc.
3 Figure 1. Discovery Timeline of Noncoding RNAs This timeline highlights the discoveries of the different functional classes of noncoding RNAs (ncrnas) and the emergence of evidence for widespread throughout the genome. Widespread produces of a plethora of unannotated RNA species, most of which are characterized by reduced protein coding potential. Examples of the many regulatory roles of ncrnas in cellular processes are provided. Diversity in the size and function of ncrnas coupled with the abundance of noncoding suggest that many ncrnas still await discovery. (See Table S1 for full citations for timeline events). be incomplete. Many other functions for noncoding transcripts have been identified, including al activation, gene silencing, imprinting, dosage compensation, translational silencing, modulation of protein function, and binding as riboswitches to regulatory metabolites (reviewed in Kiss, 2002; Mattick and Makunin, 2006; Zamore and Haley, 2005). One of the best characterized emerging classes of ncrnas are the micrornas (mirnas) cloned over a decade ago in C. elegans and now recognized as a large conserved family of?22-nucleotide regulatory RNAs essential for a variety of cellular processes (reviewed in Esquela- Kerscher and Slack, 2006; Zamore and Haley, 2005). The differential expression patterns of mirnas determine cell fate and correct differentiation during development (for example, through regulation of Notch signaling and Hox gene expression). MicroRNAs can act as tumor suppressors and oncogenes (for example, by regulating Bcl2, Ras, Myc, and E2F), and they can regulate cellular proliferation and apoptosis. Surprisingly, mirna expression profiles appear to reflect more accurately the developmental lineage and differentiation state of tumors than do mrna profiles. Hundreds of mirnas are estimated to be present in the human genome, and computational analysis suggests that more than 20% 30% of human genes are regulated by mirnas. Microarray experiments support this view, revealing mirnamediated downregulation of large numbers of target mrnas. In addition, mirnas suppress initiation of protein translation, promote mrna degradation and turnover, and initiate al silencing. However, the function of the vast majority of mirnas is as yet unknown. Small nucleolar RNAs (snornas), another abundant group of RNAs, reveal an ever-increasing retinue of cellular functions for ncrnas (reviewed in Kiss, 2002). Originally appreciated for their processing of rrnas and involvement in the postal 2 -O-methylation and pseudouridylation of target rrnas and snrnas, the identification of many orphan snornas lacking complementarity to rrnas, snrnas, and trnas suggests a broad range of RNA substrates for snornas. Furthermore, snor- NAs are known to accumulate (and potentially function) outside of the nucleolus. Given that most snor- NAs are processed from the introns of other genes, their expression is inextricably linked to of their host gene. For example, a brain-specific snorna called HBII-52 modulates alternate splicing of the transcript encoding the serotonin receptor. Patients with Prader-Willi syndrome who lack HBII-52 have different serotonin Cell 125, June 30, Elsevier Inc. 1217
4 receptor splice forms, resulting in an alteration in serotonin efficacy (Kishore and Stamm, 2006). The interaction of ncrnas with proteins is another way in which ncrnas can exert their effects in the cell. For example, steroid receptor RNA activator (SRA) is a al coactivator whose action requires chromatin remodeling and recruitment of histone deacetylases (Zhao et al., 2004). SRA facilitates establishment of the al preinitiation complex by interacting with nuclear receptors, probably forming a scaffold for the binding of coactivators (such as the DEAD box protein p72/p68) and corepressors (such as SHARP). A screen for evolutionarily conserved functional ncrnas identified a noncoding repressor of the NFAT factor (NRON) (Willingham et al., 2005). The demonstrated interaction of NRON with nuclear import factors immediately suggests that NRON in a complex with importin-β specifically downregulates the nuclear import of NFAT, a hypothesis supported by studies of NFAT translocation. Indeed, given the complicated networks of nuclear-cytoplasmic transport and the seemingly limited number of available importin-β family members, such ncrna-mediated regulation of specific cargo proteins may emerge as a common biological strategy for dealing with the complexity of intracellular trafficking. Activation of the heat shock factor (HSF1) is mediated by an RNP complex, which includes the translation elongation factor eef1a and a new ncrna called HSR1 (Shamovsky et al., 2006). Conserved from rodents to humans, HSR1 is essential for heat shock response activation and may act as an RNA thermosensor. The identification of a specific class of small RNAs with a potential role in development has very recently been described. The Argonaute Piwi subfamily proteins have been shown to be important in germline development (Cox et al., 1998). Girard et al. (2006) and Aravin et al. (2006) have now identified and characterized a highly abundant class of small RNAs called pirnas and showed them to be associated with the Piwi proteins MIWI (Girard et al., 2006) and MILI (Aravin et al., 2006) in murine testis. The slightly longer lengths (26 31 nucleotides) and testes-specific expression of these pirnas perhaps hint at the start of a growing number of developmental stage-specific or tissue-specific members of new classes of small RNAs. Finally, the burgeoning number of putative ncrna genes has been cited as a mechanism for increasing the complexity of gene regulatory networks and, together with alternative gene splicing, significantly increases the variety and complexity of the transcriptome. The finding that the rising ratio of noncoding to protein-coding DNA correlates with increasing organismal complexity supports this notion (Mattick and Makunin, 2006). With the limited number of currently characterized ncrnas acting in so many cellular processes and the demonstrated amount of unannotated present in the genome, one can easily appreciate that ncrnas probably function in more cellular pathways than their protein-coding brethren. Implications of Noncoding Transcription Noncoding RNAs possess several properties that make them attractive as regulatory factors. For example, mirnas are 1000-fold smaller than most mrnas. With their 20 nucleotides, mirnas can perform the task of a protein domain composed of 100 amino acids. Considering the effects of compensatory base pair changes on structure and target hybridization, such RNAs could evolve much more readily than protein domains. The strong innate affinity of RNA for both DNA and RNA permits a diversity of stable mismatch-based secondary structure motifs such as stems, bulges, and loops. These elements may be stronger determinants of an RNA s function than the primary sequence itself, and would help to explain the lack of evolutionary conservation observed among many ncrnas. Evolutionary conservation of even well-characterized and biologically important ncrnas appears to be inconsistent at best. The let7 mirna is conserved from worms to humans (Zamore and Haley, 2005), yet the dosage compensation ncrna XIST appears to be rapidly evolving even among rodents (Nesterova et al., 2001). Furthermore, of the?34,000 putative ncrnas identified in mouse, only 3% 4% or so have limited human sequence conservation (Carninci et al., 2005). Only about 7% 20% (depending on the study) of human unannotated transcripts appear to be conserved over most of their lengths with their mouse counterparts; thus, the majority are species-specific (Johnson et al., 2005). Perhaps the criteria used to determine evolutionary conservation should be reevaluated. For example, it could be that the lengths of the conserved sequences are short and discontinuous, resembling islands of conservation within a larger nonconserved sequence. If such patches of conservation turn out to be biologically important for unannotated ncrnas (as they are for mirnas), the number of such conserved regions in the genome would escalate considerably. Given the tolerance of RNA for mismatches and its probable reliance on secondary structure for bioactivity, ncrnas are likely to have a greater degree of plasticity than mrnas. Often multiple mir- NAs must be deleted to obtain a phenotype (Abbott et al., 2005) an illustration of functional redundancy and regulatory complexity that may extend to other ncrnas. Noncoding RNAs have been implicated in a number of diseases, including B cell neoplasia, lung cancer, autism, DiGeorge syndrome, prostate cancer, and schizophrenia (see RNAdb, / rnadb/). Considering the abundance of empirically observed ncrnas, it is highly likely that many more will be linked to human diseases. In addition, mirnas linked to cancer may prove valuable therapeutic targets (Esquela-Kerscher and Slack, 2006) Cell 125, June 30, Elsevier Inc.
5 There are several strategies for investigating the function of new noncoding transcripts. Starting with a large number of putative mouse ncrnas identified by the FANTOM project, Ravasi et al. used expression analysis tools such as microarrays, PCR, and Northern blots combined with lipopolysaccharide treatment to validate the regulated expression of a subset of these ncrnas (Ravasi et al., 2006). In a separate study but starting with the same set of mouse ncrnas, Schultz and colleagues identified a subset of 512 ncrnas apparently conserved during human evolution; they then used sirnas to disrupt these conserved ncrnas and screened for phenotypes using a panel of cellular assays (Willingham et al., 2005). Such detailed expression analyses and sirna-mediated phenotypic screenings highlight a path for the followup characterization of newly mapped al dark matter. Disruption of an essential RNA processing pathway such as Dicer cleavage or rrna maturation should have dramatic al repercussions, which could be detected with tiling arrays. Such an experiment was conducted in yeast by depleting Rpp1, an essential protein required for both RNase P-mediated maturation of trnas and RNase MRP, which processes rrna. This study revealed 74 new ncrnas, many of which were antisense to known protein coding genes (Samanta et al., 2006). Future efforts could focus on: (1) combined use of RNase footprinting and tiling arrays for genomewide identification of protected RNAs; (2) purification of RNA-associated proteins such as helicases and identification of their associated RNAs by tiling array; (3) classification of subcellular and structural fractionations of RNAs using tiling arrays; and (4) development of higher-density tiling arrays that allow high-resolution genomewide surveys of, permitting investigation of rare tissues, primary cells derived from tissues, developmental time-courses, and other low abundance samples. How much of the nonredundant genome is transcribed? Based on published data, estimates range from 10% to 60%. However, this may be an underestimate given the limited number of cells and differentiation states surveyed thus far. Furthermore, arraybased transcript mapping relies on conservative thresholds which select for the highest 2% 5% of probes, yielding a low false positive rate of a few percent. Given the low expression levels of many TUFs, these thresholds are likely to be underestimates of the true amount of unannotated. Indeed, unpublished data from our lab suggest that 75% of RACE products selected from random regions of the human genome and hybridized to microarrays reveal the presence of complex transcripts. Analyzing the distribution of transposable elements that are counter-selected when they occur within functional al units (FTUs) implies that >50% of the genome consists of FTUs, and one-third of these are likely to be ncrnas (Semon and Duret, 2004). Weighing these factors together, we suggest that all of the non-repeat portions of the human genome are transcribed. This may seem an excessive estimate, yet recent data in yeast imply that more that 85% of its genome is transcribed (David et al., 2006). Furthermore, large-scale cdna sequencing and annotation in the mouse has shown that 62% of the mouse genome is transcribed (Carninci et al., 2005), and there are estimates that 90% of the human genome is transcribed (Wong et al., 2001). The overall architecture of transcribed portions of the genome is highly complex. Indeed, the landscape of most transcriptomes is a lattice-like network of overlapping in which the same genomic sequences often serve as portions of separately regulated transcripts, making the boundaries and indeed the concept of the term gene less useful than it once was. Supplemental Data The Supplemental Data for this article, including Table S1, can be found online at 125/7/1215/DC1/. Acknowledgments We thank Philip Kapranov for helpful discussions. References Abbott, A.L., Alvarez-Saavedra, E., Miska, E.A., Lau, N.C., Bartel, D.P., Horvitz, H.R., and Ambros, V. (2005). Dev. Cell 9, Aravin, A., Gaidatzis, D., Pfeffer, S., Lagos- Quintana, M., Landgraf, P., Iovino, N., Morris, P., Brownstein, M.J., Kuramochi-Miyagawa, S., Nakano, T., et al. (2006). Nature. Published online June 4, / nature Carninci, P., Kasukawa, T., Katayama, S., Gough, J., Frith, M.C., Maeda, N., Oyama, R., Ravasi, T., Lenhard, B., Wells, C., et al. (2005). Science 309, Cawley, S., Bekiranov, S., Ng, H.H., Kapranov, P., Sekinger, E.A., Kampa, D., Piccolboni, A., Sementchenko, V., Cheng, J., Williams, A.J., et al. (2004). Cell 116, Cheng, J., Kapranov, P., Drenkow, J., Dike, S., Brubaker, S., Patel, S., Long, J., Stern, D., Tammana, H., Helt, G., et al. (2005). Science 308, Cox, D.N., Chao, A., Baker, J., Chang, L., Qiao, D., and Lin, H. (1998). Genes Dev. 12, David, L., Huber, W., Granovskaia, M., Toedling, J., Palm, C.J., Bofkin, L., Jones, T., Davis, R.W., and Steinmetz, L.M. (2006). Proc. Natl. Acad. Sci. USA 103, Esquela-Kerscher, A., and Slack, F.J. (2006). Nat. Rev. Cancer 6, Girard, A., Sachidanandam, R., Hannon, G.J., and Carmell, M.A. (2006). Nature. Published online June 4, /nature Hirotsune, S., Yoshida, N., Chen, A., Garrett, L., Sugiyama, F., Takahashi, S., Yagami, K., Wynshaw-Boris, A., and Yoshiki, A. (2003). Nature 423, Johnson, J.M., Edwards, S., Shoemaker, D., and Schadt, E.E. (2005). Trends Genet. 21, Katayama, S., Tomaru, Y., Kasukawa, T., Waki, K., Nakanishi, M., Nakamura, M., Nishida, H., Yap, C.C., Suzuki, M., Kawai, J., et al. (2005). Science 309, Kim, T.H., Barrera, L.O., Zheng, M., Qu, C., Singer, M.A., Richmond, T.A., Wu, Y., Green, R.D., and Ren, B. (2005). Nature 436, Kishore, S., and Stamm, S. (2006). Science 311, Kiss, T. (2002). Cell 109, Martone, R., Euskirchen, G., Bertone, P., Hartman, S., Royce, T.E., Luscombe, N.M., Rinn, J.L., Nelson, F.K., Miller, P., Gerstein, M., et al. (2003). Proc. Natl. Acad. Sci. USA 100, Cell 125, June 30, Elsevier Inc. 1219
6 Mattick, J.S., and Makunin, I.V. (2006). Hum. Mol. Genet. 15 (Suppl 1), R17 R29. Nesterova, T.B., Slobodyanyuk, S.Y., Elisaphenko, E.A., Shevchenko, A.I., Johnston, C., Pavlova, M.E., Rogozin, I.B., Kolesnikov, N.N., Brockdorff, N., and Zakian, S.M. (2001). Genome Res. 11, Nobrega, M.A., Zhu, Y., Plajzer-Frick, I., Afzal, V., and Rubin, E.M. (2004). Nature 431, Ravasi, T., Suzuki, H., Pang, K.C., Katayama, S., Furuno, M., Okunishi, R., Fukuda, S., Ru, K., Frith, M.C., Gongora, M.M., et al. (2006). Genome Res. 16, Samanta, M.P., Tongprasit, W., Sethi, H., Chin, C.S., and Stolc, V. (2006). Proc. Natl. Acad. Sci. USA 103, Semon, M., and Duret, L. (2004). Trends Genet. 20, Shamovsky, I., Ivannikov, M., Kandel, E.S., Gershon, D., and Nudler, E. (2006). Nature 440, Sontheimer, E.J., and Carthew, R.W. (2005). Cell 122, Willingham, A.T., Orth, A.P., Batalov, S., Peters, E.C., Wen, B.G., Aza-Blanc, P., Hogenesch, J.B., and Schultz, P.G. (2005). Science 309, Wong, G.K., Passey, D.A., and Yu, J. (2001). Genome Res. 11, Zamore, P.D., and Haley, B. (2005). Science 309, Zhao, X., Patton, J.R., Davis, S.L., Florence, B., Ames, S.J., and Spanjaard, R.A. (2004). Mol. Cell 15, Cell 125, June 30, Elsevier Inc.
7 Supplemental Data TUF Love for Junk DNA Aarron T. Willingham and Thomas R. Gingeras Table S1. timeline date category references abundant cytoplasmic poly(a)- RNA found in human cells (subsequently observed in a variety of plant and animal cells) nucleic acid hybridization reassociation kinetics (Cot curves) find 10- fold more hnrna than mrna in amphibian oogenesis, levels are ~10-fold greater than expected for mrnas alone Cot curves show 10-fold greater complexity of nuclear vs. cytoplasmic poly(a)+ RNA 1974 widespread 1975 widespread 1980 widespread 1980 widespread 5` capped RNAs outnumber poly(a)+ 3-to widespread tiling array experiments find widespread unannotated 2002 widespread sequencing of full-length cdnas and SAGE tags finds widespread ncrnas and antisense (noncoding cdnas average 1800nt) whole genome mapping for Arabidopsis and Drosophila widespread widespread whole genome mapping for human 2004 widespread chromatin immunoprecipitation (ChIP) studies of mammalian promoters find many ncrnas are regulated by factors and signaling molecules fine-mapping of transcriptome for 30% of human genome at 5bp resolution upwards of 70% of sense transcripts are found to have an antisense partner fine-mapping of transcriptome for 100% of human genome at 5bp resolution ribosomal RNAs (~ nt) transfer RNAs (~80nt) small nuclear RNAs (splicing) (~ nt) 2004 widespread 2005 widespread 2005 widespread 2006 widespread (Milcarek et al., 1974; Snider and Morrison- Bogorad, 1992) (Hough et al., 1975) (Varley et al., 1980) (Holland et al., 1980) (Salditt-Georgieff et al., 1981) (Kapranov et al., 2002) (Chen et al., 2002; Okazaki et al., 2002; Saha et al., 2002) (Stolc et al., 2004; Yamada et al., 2003) (Bertone et al., 2004) (Cawley et al., 2004) (Cheng et al., 2005) (Katayama et al., 2005) manuscript in preparation 1958 ncrnas (Crick, 1958) 1958 ncrnas (Hoagland et al., 1958) 1977 ncrnas (Benecke and Penman, 1977; Goldstein et al.,
8 1977) endogenous antisense in prokaryotes 1981 ncrnas (Rosen et al., 1981; Tomizawa and Itoh, 1981) endogenous antisense in eukaryotes 1986 ncrnas (Adeniyi-Jones and Zasloff, 1985; Nepveu and Marcu, 1986; Spencer et al., 1986; Williams and Fried, 1986) cosuppression in plants (later shown to be RNAi-based) 1990 ncrnas (Napoli et al., 1990; van der Krol et al., 1990) Xist, a ncrna required for dosage compensation (16,500nt) small nucleolar RNAs (~60-300nt) first mirna (lin-4) (~22nt) steroid receptor RNA activator (SRA) (875nt) Tsix, antisense regulator of Xist (40,000nt) Air, antisense RNA required for autosomal gene imprinting (~108,000nt) mirnas found to be a large class of ncrnas (~22nt) 1992 ncrnas (Brockdorff et al., 1992; Brown et al., 1992) 1992 ncrnas (Leverette et al., 1992) 1993 ncrnas (Lee et al., 1993) 1999 ncrnas (Lanz et al., 1999) 1999 ncrnas (Lee et al., 1999) 2000 ncrnas (Lyle et al., 2000) 2001 ncrnas (Lee and Ambros, 2001) small RNAs required for heterochromatin formation 2002 ncrnas (Reinhart and Bartel, 2002; Volpe et al., 2002) endogenous riboswitches (in 5` UTR of protein-coding mrnas) TRE ncrnas activate by recruiting histone modifying factors pirnas, an abundant testes specific class of small RNAs with a role in germline development 2002 ncrnas (Mironov et al., 2002; Winkler et al., 2002) 2006 ncrnas (Sanchez-Elsner et al., 2006) 2006 ncrnas (Aravin et al., 2006; Girard et al., 2006) Additional noncoding RNA data available from RNAdb (Pang et al., 2005) and Rfam (Griffiths-Jones et al., 2005) online databases.
9 References: Adeniyi-Jones, S., and Zasloff, M. (1985). Transcription, processing and nuclear transport of a B1 Alu RNA species complementary to an intron of the murine alphafetoprotein gene. Nature 317, Aravin, A., Gaidatzis, D., Pfeffer, S., Lagos-Quintana, M., Landgraf, P., Iovino, N., Morris, P., Brownstein, M. J., Kuramochi-Miyagawa, S., Nakano, T., et al. (2006). A novel class of small RNAs bind to MILI protein in mouse testes. Nature doi: /nature Benecke, B. J., and Penman, S. (1977). A new class of small nuclear RNA molecules synthesized by a type I RNA polymerase in HeLa cells. Cell 12, Bertone, P., Stolc, V., Royce, T. E., Rozowsky, J. S., Urban, A. E., Zhu, X., Rinn, J. L., Tongprasit, W., Samanta, M., Weissman, S., et al. (2004). Global identification of human transcribed sequences with genome tiling arrays. Science 306, Brockdorff, N., Ashworth, A., Kay, G. F., McCabe, V. M., Norris, D. P., Cooper, P. J., Swift, S., and Rastan, S. (1992). The product of the mouse Xist gene is a 15 kb inactive X-specific transcript containing no conserved ORF and located in the nucleus. Cell 71, Brown, C. J., Hendrich, B. D., Rupert, J. L., Lafreniere, R. G., Xing, Y., Lawrence, J., and Willard, H. F. (1992). The human XIST gene: analysis of a 17 kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus. Cell 71, Cawley, S., Bekiranov, S., Ng, H. H., Kapranov, P., Sekinger, E. A., Kampa, D., Piccolboni, A., Sementchenko, V., Cheng, J., Williams, A. J., et al. (2004). Unbiased mapping of factor binding sites along human chromosomes 21 and 22 points to widespread regulation of noncoding RNAs. Cell 116, Chen, J., Sun, M., Lee, S., Zhou, G., Rowley, J. D., and Wang, S. M. (2002). Identifying novel transcripts and novel genes in the human genome by using novel SAGE tags. Proc Natl Acad Sci U S A 99, Cheng, J., Kapranov, P., Drenkow, J., Dike, S., Brubaker, S., Patel, S., Long, J., Stern, D., Tammana, H., Helt, G., et al. (2005). Transcriptional maps of 10 human chromosomes at 5-nucleotide resolution. Science 308, Crick, F. H. (1958). On protein synthesis. Symp Soc Exp Biol 12, Girard, A., Sachidanandam, R., Hannon, G. J., and Carmell, M. A. (2006). A germlinespecific class of small RNAs binds mammalian Piwi proteins. Nature doi: /nature Goldstein, L., Wise, G. E., and Ko, C. (1977). Small nuclear RNA localization during mitosis. An electron microscope study. J Cell Biol 73,
10 Griffiths-Jones, S., Moxon, S., Marshall, M., Khanna, A., Eddy, S. R., and Bateman, A. (2005). Rfam: annotating non-coding RNAs in complete genomes. Nucleic Acids Res 33 Database Issue, D Hoagland, M. B., Stephenson, M. L., Scott, J. F., Hecht, L. I., and Zamecnik, P. C. (1958). A soluble ribonucleic acid intermediate in protein synthesis. J Biol Chem 231, Holland, C. A., Mayrand, S., and Pederson, T. (1980). Sequence complexity of nuclear and messenger RNA in HeLa cells. J Mol Biol 138, Hough, B. R., Smith, M. J., Britten, R. J., and Davidson, E. H. (1975). Sequence complexity of heterogeneous nuclear RNA in sea urchin embryos. Cell 5, Kapranov, P., Cawley, S. E., Drenkow, J., Bekiranov, S., Strausberg, R. L., Fodor, S. P., and Gingeras, T. R. (2002). Large-scale al activity in chromosomes 21 and 22. Science 296, Katayama, S., Tomaru, Y., Kasukawa, T., Waki, K., Nakanishi, M., Nakamura, M., Nishida, H., Yap, C. C., Suzuki, M., Kawai, J., et al. (2005). Antisense in the mammalian transcriptome. Science 309, Lanz, R. B., McKenna, N. J., Onate, S. A., Albrecht, U., Wong, J., Tsai, S. Y., Tsai, M. J., and O'Malley, B. W. (1999). A steroid receptor coactivator, SRA, functions as an RNA and is present in an SRC-1 complex. Cell 97, Lee, J. T., Davidow, L. S., and Warshawsky, D. (1999). Tsix, a gene antisense to Xist at the X-inactivation centre. Nat Genet 21, Lee, R. C., and Ambros, V. (2001). An extensive class of small RNAs in Caenorhabditis elegans. Science 294, Lee, R. C., Feinbaum, R. L., and Ambros, V. (1993). The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, Leverette, R. D., Andrews, M. T., and Maxwell, E. S. (1992). Mouse U14 snrna is a processed intron of the cognate hsc70 heat shock pre-messenger RNA. Cell 71, Lyle, R., Watanabe, D., te Vruchte, D., Lerchner, W., Smrzka, O. W., Wutz, A., Schageman, J., Hahner, L., Davies, C., and Barlow, D. P. (2000). The imprinted antisense RNA at the Igf2r locus overlaps but does not imprint Mas1. Nat Genet 25, Milcarek, C., Price, R., and Penman, S. (1974). The metabolism of a poly(a) minus mrna fraction in HeLa cells. Cell 3, Mironov, A. S., Gusarov, I., Rafikov, R., Lopez, L. E., Shatalin, K., Kreneva, R. A., Perumov, D. A., and Nudler, E. (2002). Sensing small molecules by nascent RNA: a mechanism to control in bacteria. Cell 111, Napoli, C., Lemieux, C., and Jorgensen, R. (1990). Introduction of a Chimeric Chalcone Synthase Gene into Petunia Results in Reversible Co-Suppression of Homologous Genes in trans. Plant Cell 2,
11 Nepveu, A., and Marcu, K. B. (1986). Intragenic pausing and anti-sense within the murine c-myc locus. Embo J 5, Okazaki, Y., Furuno, M., Kasukawa, T., Adachi, J., Bono, H., Kondo, S., Nikaido, I., Osato, N., Saito, R., Suzuki, H., et al. (2002). Analysis of the mouse transcriptome based on functional annotation of 60,770 full-length cdnas. Nature 420, Pang, K. C., Stephen, S., Engstrom, P. G., Tajul-Arifin, K., Chen, W., Wahlestedt, C., Lenhard, B., Hayashizaki, Y., and Mattick, J. S. (2005). RNAdb--a comprehensive mammalian noncoding RNA database. Nucleic Acids Res 33, D Reinhart, B. J., and Bartel, D. P. (2002). Small RNAs correspond to centromere heterochromatic repeats. Science 297, Rosen, J., Ryder, T., Ohtsubo, H., and Ohtsubo, E. (1981). Role of RNA transcripts in replication incompatibility and copy number control in antibiotic resistance plasmid derivatives. Nature 290, Saha, S., Sparks, A. B., Rago, C., Akmaev, V., Wang, C. J., Vogelstein, B., Kinzler, K. W., and Velculescu, V. E. (2002). Using the transcriptome to annotate the genome. Nat Biotechnol 20, Salditt-Georgieff, M., Harpold, M. M., Wilson, M. C., and Darnell, J. E., Jr. (1981). Large heterogeneous nuclear ribonucleic acid has three times as many 5' caps as polyadenylic acid segments, and most caps do not enter polyribosomes. Mol Cell Biol 1, Sanchez-Elsner, T., Gou, D., Kremmer, E., and Sauer, F. (2006). Noncoding RNAs of trithorax response elements recruit Drosophila Ash1 to Ultrabithorax. Science 311, Snider, B. J., and Morrison-Bogorad, M. (1992). Brain non-adenylated mrnas. Brain Res Brain Res Rev 17, Spencer, C. A., Gietz, R. D., and Hodgetts, R. B. (1986). Overlapping units in the dopa decarboxylase region of Drosophila. Nature 322, Stolc, V., Gauhar, Z., Mason, C., Halasz, G., van Batenburg, M. F., Rifkin, S. A., Hua, S., Herreman, T., Tongprasit, W., Barbano, P. E., et al. (2004). A gene expression map for the euchromatic genome of Drosophila melanogaster. Science 306, Tomizawa, J., and Itoh, T. (1981). Plasmid ColE1 incompatibility determined by interaction of RNA I with primer transcript. Proc Natl Acad Sci U S A 78, van der Krol, A. R., Mur, L. A., Beld, M., Mol, J. N., and Stuitje, A. R. (1990). Flavonoid genes in petunia: addition of a limited number of gene copies may lead to a suppression of gene expression. Plant Cell 2, Varley, J. M., Macgregor, H. C., and Erba, H. P. (1980). Satellite DNA is transcribed on lampbrush chromosomes. Nature 283, Volpe, T. A., Kidner, C., Hall, I. M., Teng, G., Grewal, S. I., and Martienssen, R. A. (2002). Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297,
12 Williams, T., and Fried, M. (1986). A mouse locus at which from both DNA strands produces mrnas complementary at their 3' ends. Nature 322, Winkler, W., Nahvi, A., and Breaker, R. R. (2002). Thiamine derivatives bind messenger RNAs directly to regulate bacterial gene expression. Nature 419, Yamada, K., Lim, J., Dale, J. M., Chen, H., Shinn, P., Palm, C. J., Southwick, A. M., Wu, H. C., Kim, C., Nguyen, M., et al. (2003). Empirical analysis of al activity in the Arabidopsis genome. Science 302,
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
Novel Transcribed Regions in the Human Genome
 Novel Transcribed Regions in the Human Genome J. ROZOWSKY,* J. WU, Z. LIAN, U. NAGALAKSHMI, J.O. KORBEL,* P. KAPRANOV,** D. ZHENG,* S. DYKE,** P. NEWBURGER, P. MILLER,, T.R. GINGERAS,** S. WEISSMAN, M.
Novel Transcribed Regions in the Human Genome J. ROZOWSKY,* J. WU, Z. LIAN, U. NAGALAKSHMI, J.O. KORBEL,* P. KAPRANOV,** D. ZHENG,* S. DYKE,** P. NEWBURGER, P. MILLER,, T.R. GINGERAS,** S. WEISSMAN, M.
Micro RNAs: potentielle Biomarker für das. Blutspenderscreening
 Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Outline. interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work?
 Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Profiling of non-coding RNA classes Gunter Meister
 Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
Functional and Biomedical Aspects of Genome Research
 Functional and Biomedical Aspects of Genome Research 20 11 35 Vorlesung SS 04 Bartsch, Jockusch & Schmitt-John Mi. 9:15-10:00, in W7-135 13 Functional RNAs Thomas Schmitt-John micro RNAs small interfering
Functional and Biomedical Aspects of Genome Research 20 11 35 Vorlesung SS 04 Bartsch, Jockusch & Schmitt-John Mi. 9:15-10:00, in W7-135 13 Functional RNAs Thomas Schmitt-John micro RNAs small interfering
V22: involvement of micrornas in GRNs
 What are micrornas? V22: involvement of micrornas in GRNs How can one identify micrornas? What is the function of micrornas? Elisa Izaurralde, MPI Tübingen Huntzinger, Izaurralde, Nat. Rev. Genet. 12,
What are micrornas? V22: involvement of micrornas in GRNs How can one identify micrornas? What is the function of micrornas? Elisa Izaurralde, MPI Tübingen Huntzinger, Izaurralde, Nat. Rev. Genet. 12,
Standards, Guidelines and Best Practices for RNA-Seq V1.0 (June 2011) The ENCODE Consortium
 Standards, Guidelines and Best Practices for RNA-Seq V1.0 (June 2011) The ENCODE Consortium I. Introduction: Sequence based assays of transcriptomes (RNA-seq) are in wide use because of their favorable
Standards, Guidelines and Best Practices for RNA-Seq V1.0 (June 2011) The ENCODE Consortium I. Introduction: Sequence based assays of transcriptomes (RNA-seq) are in wide use because of their favorable
Protein Synthesis How Genes Become Constituent Molecules
 Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Control of Gene Expression
 Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals
 Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
mirnaselect pep-mir Cloning and Expression Vector
 Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
Control of Gene Expression
 Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Basic Concepts of DNA, Proteins, Genes and Genomes
 Basic Concepts of DNA, Proteins, Genes and Genomes Kun-Mao Chao 1,2,3 1 Graduate Institute of Biomedical Electronics and Bioinformatics 2 Department of Computer Science and Information Engineering 3 Graduate
Basic Concepts of DNA, Proteins, Genes and Genomes Kun-Mao Chao 1,2,3 1 Graduate Institute of Biomedical Electronics and Bioinformatics 2 Department of Computer Science and Information Engineering 3 Graduate
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION. Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.
 Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
PART 3.3: MicroRNA and Cancer
 BIBM 2010 Tutorial: Epigenomics and Cancer PART 3.3: MicroRNA and Cancer Dec 18, 2010 Sun Kim at Indiana University Outline of Part 3.3 Background on microrna Role of microrna in cancer MicroRNA pathway
BIBM 2010 Tutorial: Epigenomics and Cancer PART 3.3: MicroRNA and Cancer Dec 18, 2010 Sun Kim at Indiana University Outline of Part 3.3 Background on microrna Role of microrna in cancer MicroRNA pathway
Name Class Date. Figure 13 1. 2. Which nucleotide in Figure 13 1 indicates the nucleic acid above is RNA? a. uracil c. cytosine b. guanine d.
 13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
Dicer Substrate RNAi Design
 INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
Functional RNAs; RNA catalysts, mirna,
 Functional RNAs; RNA catalysts, mirna, srna, RNAi... RNAs have many functions rrna (ribosomal RNA) trna (transfer RNA) mrna (Messenger RNA) snrna (including snorna) ) (Small nuclear RNA- splicing) Other
Functional RNAs; RNA catalysts, mirna, srna, RNAi... RNAs have many functions rrna (ribosomal RNA) trna (transfer RNA) mrna (Messenger RNA) snrna (including snorna) ) (Small nuclear RNA- splicing) Other
DNA Replication & Protein Synthesis. This isn t a baaaaaaaddd chapter!!!
 DNA Replication & Protein Synthesis This isn t a baaaaaaaddd chapter!!! The Discovery of DNA s Structure Watson and Crick s discovery of DNA s structure was based on almost fifty years of research by other
DNA Replication & Protein Synthesis This isn t a baaaaaaaddd chapter!!! The Discovery of DNA s Structure Watson and Crick s discovery of DNA s structure was based on almost fifty years of research by other
Complex multicellular organisms are produced by cells that switch genes on and off during development.
 Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
13.4 Gene Regulation and Expression
 13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
Dark matter in the genome: evidence of widespread transcription detected by microarray tiling experiments
 Review TRENDS in Genetics Vol.21 No.2 February 2005 Dark matter in the genome: evidence of widespread transcription detected by microarray tiling experiments Jason M. Johnson 1, Stephen Edwards 1, Daniel
Review TRENDS in Genetics Vol.21 No.2 February 2005 Dark matter in the genome: evidence of widespread transcription detected by microarray tiling experiments Jason M. Johnson 1, Stephen Edwards 1, Daniel
Sample Questions for Exam 3
 Sample Questions for Exam 3 1. All of the following occur during prometaphase of mitosis in animal cells except a. the centrioles move toward opposite poles. b. the nucleolus can no longer be seen. c.
Sample Questions for Exam 3 1. All of the following occur during prometaphase of mitosis in animal cells except a. the centrioles move toward opposite poles. b. the nucleolus can no longer be seen. c.
Outline. MicroRNA Bioinformatics. microrna biogenesis. short non-coding RNAs not considered in this lecture. ! Introduction
 Outline MicroRNA Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology (CMB) Karolinska Institutet! Introduction! microrna target site prediction! Useful resources 2 short non-coding RNAs
Outline MicroRNA Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology (CMB) Karolinska Institutet! Introduction! microrna target site prediction! Useful resources 2 short non-coding RNAs
Structure and Function of DNA
 Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
THE ENZYMES. Department of Microbiology, Immunology, and Molecular Genetics, Molecular Biology Institute University of California
 VOLUME THIRTY TWO THE ENZYMES Eukaryotic RNases and their Partners in RNA Degradation and Biogenesis, Part B Edited by FENG GUO Department of Biological Chemistry, David Geffen School of Medicine, Molecular
VOLUME THIRTY TWO THE ENZYMES Eukaryotic RNases and their Partners in RNA Degradation and Biogenesis, Part B Edited by FENG GUO Department of Biological Chemistry, David Geffen School of Medicine, Molecular
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
Translation Study Guide
 Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
MicroRNA Mike needs help to degrade all the mrna transcripts! Aaron Arvey ISMB 2010
 Target mrna abundance dilutes microrna and sirna activity MicroRNA Mike needs help to degrade all the mrna transcripts! Aaron Arvey ISMB 2010 Target mrna abundance dilutes microrna and sirna activity Erik
Target mrna abundance dilutes microrna and sirna activity MicroRNA Mike needs help to degrade all the mrna transcripts! Aaron Arvey ISMB 2010 Target mrna abundance dilutes microrna and sirna activity Erik
Human Genome Organization: An Update. Genome Organization: An Update
 Human Genome Organization: An Update Genome Organization: An Update Highlights of Human Genome Project Timetable Proposed in 1990 as 3 billion dollar joint venture between DOE and NIH with 15 year completion
Human Genome Organization: An Update Genome Organization: An Update Highlights of Human Genome Project Timetable Proposed in 1990 as 3 billion dollar joint venture between DOE and NIH with 15 year completion
Lezioni Dipartimento di Oncologia Farmacologia Molecolare. RNA interference. Giovanna Damia 29 maggio 2006
 Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
岑 祥 股 份 有 限 公 司 技 術 專 員 費 軫 尹 20100803
 技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
Understanding the dynamics and function of cellular networks
 Understanding the dynamics and function of cellular networks Cells are complex systems functionally diverse elements diverse interactions that form networks signal transduction-, gene regulatory-, metabolic-
Understanding the dynamics and function of cellular networks Cells are complex systems functionally diverse elements diverse interactions that form networks signal transduction-, gene regulatory-, metabolic-
Control of Gene Expression
 Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
New Technologies for Sensitive, Low-Input RNA-Seq. Clontech Laboratories, Inc.
 New Technologies for Sensitive, Low-Input RNA-Seq Clontech Laboratories, Inc. Outline Introduction Single-Cell-Capable mrna-seq Using SMART Technology SMARTer Ultra Low RNA Kit for the Fluidigm C 1 System
New Technologies for Sensitive, Low-Input RNA-Seq Clontech Laboratories, Inc. Outline Introduction Single-Cell-Capable mrna-seq Using SMART Technology SMARTer Ultra Low RNA Kit for the Fluidigm C 1 System
Title: Surveying Genome to Identify Origins of DNA Replication In Silico
 Title: Surveying Genome to Identify Origins of DNA Replication In Silico Abstract: DNA replication origins are the bases to realize the process of chromosome replication and analyze the progress of cell
Title: Surveying Genome to Identify Origins of DNA Replication In Silico Abstract: DNA replication origins are the bases to realize the process of chromosome replication and analyze the progress of cell
MicroRNA formation. 4th International Symposium on Non-Surgical Contraceptive Methods of Pet Population Control
 MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
Discovery and Quantification of RNA with RNASeq Roderic Guigó Serra Centre de Regulació Genòmica (CRG) roderic.guigo@crg.cat
 Bioinformatique et Séquençage Haut Débit, Discovery and Quantification of RNA with RNASeq Roderic Guigó Serra Centre de Regulació Genòmica (CRG) roderic.guigo@crg.cat 1 RNA Transcription to RNA and subsequent
Bioinformatique et Séquençage Haut Débit, Discovery and Quantification of RNA with RNASeq Roderic Guigó Serra Centre de Regulació Genòmica (CRG) roderic.guigo@crg.cat 1 RNA Transcription to RNA and subsequent
RNAi Shooting the Messenger!
 RNAi Shooting the Messenger! Bronya Keats, Ph.D. Department of Genetics Louisiana State University Health Sciences Center New Orleans Email: bkeats@lsuhsc.edu RNA interference (RNAi) A mechanism by which
RNAi Shooting the Messenger! Bronya Keats, Ph.D. Department of Genetics Louisiana State University Health Sciences Center New Orleans Email: bkeats@lsuhsc.edu RNA interference (RNAi) A mechanism by which
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation
 OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
Biological Sciences Initiative. Human Genome
 Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Five-year relative survival rates. Cancer. Age-adjusted cancer death rates. Proteomic Technologies for Cancer Biomarker Discovery 2010/3/22
 Cancer Five-year relative survival rates Basal lamina Underlyig tissue Normal tissue Carcinoma Invasive carcinoma 1 http://www.cancer.org/docroot/home/index.asp 2 Proteomic Technologies for Cancer Biomarker
Cancer Five-year relative survival rates Basal lamina Underlyig tissue Normal tissue Carcinoma Invasive carcinoma 1 http://www.cancer.org/docroot/home/index.asp 2 Proteomic Technologies for Cancer Biomarker
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS)
 Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
FlipFlop: Fast Lasso-based Isoform Prediction as a Flow Problem
 FlipFlop: Fast Lasso-based Isoform Prediction as a Flow Problem Elsa Bernard Laurent Jacob Julien Mairal Jean-Philippe Vert September 24, 2013 Abstract FlipFlop implements a fast method for de novo transcript
FlipFlop: Fast Lasso-based Isoform Prediction as a Flow Problem Elsa Bernard Laurent Jacob Julien Mairal Jean-Philippe Vert September 24, 2013 Abstract FlipFlop implements a fast method for de novo transcript
1 Mutation and Genetic Change
 CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
Introduction to Genome Annotation
 Introduction to Genome Annotation AGCGTGGTAGCGCGAGTTTGCGAGCTAGCTAGGCTCCGGATGCGA CCAGCTTTGATAGATGAATATAGTGTGCGCGACTAGCTGTGTGTT GAATATATAGTGTGTCTCTCGATATGTAGTCTGGATCTAGTGTTG GTGTAGATGGAGATCGCGTAGCGTGGTAGCGCGAGTTTGCGAGCT
Introduction to Genome Annotation AGCGTGGTAGCGCGAGTTTGCGAGCTAGCTAGGCTCCGGATGCGA CCAGCTTTGATAGATGAATATAGTGTGCGCGACTAGCTGTGTGTT GAATATATAGTGTGTCTCTCGATATGTAGTCTGGATCTAGTGTTG GTGTAGATGGAGATCGCGTAGCGTGGTAGCGCGAGTTTGCGAGCT
Lecture Series 7. From DNA to Protein. Genotype to Phenotype. Reading Assignments. A. Genes and the Synthesis of Polypeptides
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
A Primer of Genome Science THIRD
 A Primer of Genome Science THIRD EDITION GREG GIBSON-SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc. Publishers Sunderland, Massachusetts USA Contents Preface xi 1 Genome Projects:
A Primer of Genome Science THIRD EDITION GREG GIBSON-SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc. Publishers Sunderland, Massachusetts USA Contents Preface xi 1 Genome Projects:
Human Genome and Human Genome Project. Louxin Zhang
 Human Genome and Human Genome Project Louxin Zhang A Primer to Genomics Cells are the fundamental working units of every living systems. DNA is made of 4 nucleotide bases. The DNA sequence is the particular
Human Genome and Human Genome Project Louxin Zhang A Primer to Genomics Cells are the fundamental working units of every living systems. DNA is made of 4 nucleotide bases. The DNA sequence is the particular
micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved
 microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
Transcription and Translation of DNA
 Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
GENE REGULATION. Teacher Packet
 AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
Basic Principles of Transcription and Translation
 The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific traits by dictating the synthesis of
The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific traits by dictating the synthesis of
Non-coding RNA. John S. Mattick* and Igor V. Makunin INTRODUCTION. EXPANSION OF ncrnas AND RNA METABOLISM IN EUKARYOTES
 doi:10.1093/hmg/ddl046 R17 R29 Non-coding RNA John S. Mattick* and Igor V. Makunin Australian Research Council Centre for Functional and Applied Genomics, Institute for Molecular Bioscience, University
doi:10.1093/hmg/ddl046 R17 R29 Non-coding RNA John S. Mattick* and Igor V. Makunin Australian Research Council Centre for Functional and Applied Genomics, Institute for Molecular Bioscience, University
Chapter 5: Organization and Expression of Immunoglobulin Genes
 Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Yang-Ming University, 2009 microrna Biology and Application
 Yang-Ming University, 2009 microrna Biology and Application 3/03 microrna biogenesis and functions Woan-Yuh Tarn 3/10 micrornas and development Woan- Yuh Tarn 3/17 micrornas and nervous system Jeng-Ya
Yang-Ming University, 2009 microrna Biology and Application 3/03 microrna biogenesis and functions Woan-Yuh Tarn 3/10 micrornas and development Woan- Yuh Tarn 3/17 micrornas and nervous system Jeng-Ya
RNA: Transcription and Processing
 8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
From DNA to Protein. Proteins. Chapter 13. Prokaryotes and Eukaryotes. The Path From Genes to Proteins. All proteins consist of polypeptide chains
 Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
!!!!!!!!!!!!!!!!!!!!!!!!!!
 Figure S7 cdl-1 3'UTR gld-1 binding site (4..10) let-7 binding site (326..346) Figure Legends Figure S1. gld-1 genetically interacts with nhl-2 and vig-1 during germline development. (A) nhl-2(ok818) enhances
Figure S7 cdl-1 3'UTR gld-1 binding site (4..10) let-7 binding site (326..346) Figure Legends Figure S1. gld-1 genetically interacts with nhl-2 and vig-1 during germline development. (A) nhl-2(ok818) enhances
mrna EDITING Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A. Copyright 2005
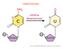 mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
Genome and DNA Sequence Databases. BME 110/BIOL 181 CompBio Tools Todd Lowe March 31, 2009
 Genome and DNA Sequence Databases BME 110/BIOL 181 CompBio Tools Todd Lowe March 31, 2009 Admin Reading: Chapters 1 & 2 Notes available in PDF format on-line (see class calendar page): http://www.soe.ucsc.edu/classes/bme110/spring09/bme110-calendar.html
Genome and DNA Sequence Databases BME 110/BIOL 181 CompBio Tools Todd Lowe March 31, 2009 Admin Reading: Chapters 1 & 2 Notes available in PDF format on-line (see class calendar page): http://www.soe.ucsc.edu/classes/bme110/spring09/bme110-calendar.html
RNA & Protein Synthesis
 RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
http://www.springer.com/3-540-23372-5
 http://www.springer.com/3-540-23372-5 Chromatin Remodeling Factors and DNA Replication Patrick Varga-Weisz Abstract Chromatin structures have to be precisely duplicated during DNA replication to maintain
http://www.springer.com/3-540-23372-5 Chromatin Remodeling Factors and DNA Replication Patrick Varga-Weisz Abstract Chromatin structures have to be precisely duplicated during DNA replication to maintain
Current Motif Discovery Tools and their Limitations
 Current Motif Discovery Tools and their Limitations Philipp Bucher SIB / CIG Workshop 3 October 2006 Trendy Concepts and Hypotheses Transcription regulatory elements act in a context-dependent manner.
Current Motif Discovery Tools and their Limitations Philipp Bucher SIB / CIG Workshop 3 October 2006 Trendy Concepts and Hypotheses Transcription regulatory elements act in a context-dependent manner.
AP BIOLOGY 2009 SCORING GUIDELINES
 AP BIOLOGY 2009 SCORING GUIDELINES Question 4 The flow of genetic information from DNA to protein in eukaryotic cells is called the central dogma of biology. (a) Explain the role of each of the following
AP BIOLOGY 2009 SCORING GUIDELINES Question 4 The flow of genetic information from DNA to protein in eukaryotic cells is called the central dogma of biology. (a) Explain the role of each of the following
How To Understand How Gene Expression Is Regulated
 What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
CCR Biology - Chapter 9 Practice Test - Summer 2012
 Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
2013 W. H. Freeman and Company. 26 RNA Metabolism
 2013 W. H. Freeman and Company 26 RNA Metabolism CHAPTER 26 RNA Metabolism Key topics: Transcription: DNA-dependent synthesis of RNA Capping and splicing: RNA processing Overview of RNA Function Ribonucleic
2013 W. H. Freeman and Company 26 RNA Metabolism CHAPTER 26 RNA Metabolism Key topics: Transcription: DNA-dependent synthesis of RNA Capping and splicing: RNA processing Overview of RNA Function Ribonucleic
trna Processing and Modification
 trna Processing and Modification RNA POL III - TRANSCRIPTS 5S RNA, trna, repetitive Sequenzen (Alu-typ), versch. kleine stabile RNAs (7SL - RNA vom signal recognition particle (SRP)), U6 RNA 5S RNA nicht
trna Processing and Modification RNA POL III - TRANSCRIPTS 5S RNA, trna, repetitive Sequenzen (Alu-typ), versch. kleine stabile RNAs (7SL - RNA vom signal recognition particle (SRP)), U6 RNA 5S RNA nicht
2. The number of different kinds of nucleotides present in any DNA molecule is A) four B) six C) two D) three
 Chem 121 Chapter 22. Nucleic Acids 1. Any given nucleotide in a nucleic acid contains A) two bases and a sugar. B) one sugar, two bases and one phosphate. C) two sugars and one phosphate. D) one sugar,
Chem 121 Chapter 22. Nucleic Acids 1. Any given nucleotide in a nucleic acid contains A) two bases and a sugar. B) one sugar, two bases and one phosphate. C) two sugars and one phosphate. D) one sugar,
Module 3 Questions. 7. Chemotaxis is an example of signal transduction. Explain, with the use of diagrams.
 Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Special report. Chronic Lymphocytic Leukemia (CLL) Genomic Biology 3020 April 20, 2006
 Special report Chronic Lymphocytic Leukemia (CLL) Genomic Biology 3020 April 20, 2006 Gene And Protein The gene that causes the mutation is CCND1 and the protein NP_444284 The mutation deals with the cell
Special report Chronic Lymphocytic Leukemia (CLL) Genomic Biology 3020 April 20, 2006 Gene And Protein The gene that causes the mutation is CCND1 and the protein NP_444284 The mutation deals with the cell
Gene Models & Bed format: What they represent.
 GeneModels&Bedformat:Whattheyrepresent. Gene models are hypotheses about the structure of transcripts produced by a gene. Like all models, they may be correct, partly correct, or entirely wrong. Typically,
GeneModels&Bedformat:Whattheyrepresent. Gene models are hypotheses about the structure of transcripts produced by a gene. Like all models, they may be correct, partly correct, or entirely wrong. Typically,
Name Date Period. 2. When a molecule of double-stranded DNA undergoes replication, it results in
 DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
Analysis of gene expression data. Ulf Leser and Philippe Thomas
 Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
RETRIEVING SEQUENCE INFORMATION. Nucleotide sequence databases. Database search. Sequence alignment and comparison
 RETRIEVING SEQUENCE INFORMATION Nucleotide sequence databases Database search Sequence alignment and comparison Biological sequence databases Originally just a storage place for sequences. Currently the
RETRIEVING SEQUENCE INFORMATION Nucleotide sequence databases Database search Sequence alignment and comparison Biological sequence databases Originally just a storage place for sequences. Currently the
Algorithms in Computational Biology (236522) spring 2007 Lecture #1
 Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
RNA and Protein Synthesis
 Name lass Date RN and Protein Synthesis Information and Heredity Q: How does information fl ow from DN to RN to direct the synthesis of proteins? 13.1 What is RN? WHT I KNOW SMPLE NSWER: RN is a nucleic
Name lass Date RN and Protein Synthesis Information and Heredity Q: How does information fl ow from DN to RN to direct the synthesis of proteins? 13.1 What is RN? WHT I KNOW SMPLE NSWER: RN is a nucleic
Genetics Module B, Anchor 3
 Genetics Module B, Anchor 3 Key Concepts: - An individual s characteristics are determines by factors that are passed from one parental generation to the next. - During gamete formation, the alleles for
Genetics Module B, Anchor 3 Key Concepts: - An individual s characteristics are determines by factors that are passed from one parental generation to the next. - During gamete formation, the alleles for
Translation. Translation: Assembly of polypeptides on a ribosome
 Translation Translation: Assembly of polypeptides on a ribosome Living cells devote more energy to the synthesis of proteins than to any other aspect of metabolism. About a third of the dry mass of a cell
Translation Translation: Assembly of polypeptides on a ribosome Living cells devote more energy to the synthesis of proteins than to any other aspect of metabolism. About a third of the dry mass of a cell
Specific problems. The genetic code. The genetic code. Adaptor molecules match amino acids to mrna codons
 Tutorial II Gene expression: mrna translation and protein synthesis Piergiorgio Percipalle, PhD Program Control of gene transcription and RNA processing mrna translation and protein synthesis KAROLINSKA
Tutorial II Gene expression: mrna translation and protein synthesis Piergiorgio Percipalle, PhD Program Control of gene transcription and RNA processing mrna translation and protein synthesis KAROLINSKA
CompleteⅡ 1st strand cdna Synthesis Kit
 Instruction Manual CompleteⅡ 1st strand cdna Synthesis Kit Catalog # GM30401, GM30402 Green Mountain Biosystems. LLC Web: www.greenmountainbio.com Tel: 800-942-1160 Sales: Sales@ greenmountainbio.com Support:
Instruction Manual CompleteⅡ 1st strand cdna Synthesis Kit Catalog # GM30401, GM30402 Green Mountain Biosystems. LLC Web: www.greenmountainbio.com Tel: 800-942-1160 Sales: Sales@ greenmountainbio.com Support:
Chapter 6 DNA Replication
 Chapter 6 DNA Replication Each strand of the DNA double helix contains a sequence of nucleotides that is exactly complementary to the nucleotide sequence of its partner strand. Each strand can therefore
Chapter 6 DNA Replication Each strand of the DNA double helix contains a sequence of nucleotides that is exactly complementary to the nucleotide sequence of its partner strand. Each strand can therefore
Central Dogma. Lecture 10. Discussing DNA replication. DNA Replication. DNA mutation and repair. Transcription
 Central Dogma transcription translation DNA RNA Protein replication Discussing DNA replication (Nucleus of eukaryote, cytoplasm of prokaryote) Recall Replication is semi-conservative and bidirectional
Central Dogma transcription translation DNA RNA Protein replication Discussing DNA replication (Nucleus of eukaryote, cytoplasm of prokaryote) Recall Replication is semi-conservative and bidirectional
Analytical Study of Hexapod mirnas using Phylogenetic Methods
 Analytical Study of Hexapod mirnas using Phylogenetic Methods A.K. Mishra and H.Chandrasekharan Unit of Simulation & Informatics, Indian Agricultural Research Institute, New Delhi, India akmishra@iari.res.in,
Analytical Study of Hexapod mirnas using Phylogenetic Methods A.K. Mishra and H.Chandrasekharan Unit of Simulation & Informatics, Indian Agricultural Research Institute, New Delhi, India akmishra@iari.res.in,
Trasposable elements: P elements
 Trasposable elements: P elements In 1938 Marcus Rhodes provided the first genetic description of an unstable mutation, an allele of a gene required for the production of pigment in maize. This instability
Trasposable elements: P elements In 1938 Marcus Rhodes provided the first genetic description of an unstable mutation, an allele of a gene required for the production of pigment in maize. This instability
GenBank, Entrez, & FASTA
 GenBank, Entrez, & FASTA Nucleotide Sequence Databases First generation GenBank is a representative example started as sort of a museum to preserve knowledge of a sequence from first discovery great repositories,
GenBank, Entrez, & FASTA Nucleotide Sequence Databases First generation GenBank is a representative example started as sort of a museum to preserve knowledge of a sequence from first discovery great repositories,
To be able to describe polypeptide synthesis including transcription and splicing
 Thursday 8th March COPY LO: To be able to describe polypeptide synthesis including transcription and splicing Starter Explain the difference between transcription and translation BATS Describe and explain
Thursday 8th March COPY LO: To be able to describe polypeptide synthesis including transcription and splicing Starter Explain the difference between transcription and translation BATS Describe and explain
Recombinant DNA and Biotechnology
 Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
From DNA to Protein
 Nucleus Control center of the cell contains the genetic library encoded in the sequences of nucleotides in molecules of DNA code for the amino acid sequences of all proteins determines which specific proteins
Nucleus Control center of the cell contains the genetic library encoded in the sequences of nucleotides in molecules of DNA code for the amino acid sequences of all proteins determines which specific proteins
Discovery & Modeling of Genomic Regulatory Networks with Big Data
 Discovery & Modeling of Genomic Regulatory Networks with Big Data Hamid Bolouri Division of Human Biology Fred Hutchinson Cancer Research Center labs.fhcrc.org/bolouri I have no financial relationships
Discovery & Modeling of Genomic Regulatory Networks with Big Data Hamid Bolouri Division of Human Biology Fred Hutchinson Cancer Research Center labs.fhcrc.org/bolouri I have no financial relationships
Core Facility Genomics
 Core Facility Genomics versatile genome or transcriptome analyses based on quantifiable highthroughput data ascertainment 1 Topics Collaboration with Harald Binder and Clemens Kreutz Project: Microarray
Core Facility Genomics versatile genome or transcriptome analyses based on quantifiable highthroughput data ascertainment 1 Topics Collaboration with Harald Binder and Clemens Kreutz Project: Microarray
Coding sequence the sequence of nucleotide bases on the DNA that are transcribed into RNA which are in turn translated into protein
 Assignment 3 Michele Owens Vocabulary Gene: A sequence of DNA that instructs a cell to produce a particular protein Promoter a control sequence near the start of a gene Coding sequence the sequence of
Assignment 3 Michele Owens Vocabulary Gene: A sequence of DNA that instructs a cell to produce a particular protein Promoter a control sequence near the start of a gene Coding sequence the sequence of
Next Generation Sequencing
 Next Generation Sequencing Technology and applications 10/1/2015 Jeroen Van Houdt - Genomics Core - KU Leuven - UZ Leuven 1 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977
Next Generation Sequencing Technology and applications 10/1/2015 Jeroen Van Houdt - Genomics Core - KU Leuven - UZ Leuven 1 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977
Mitochondrial DNA Analysis
 Mitochondrial DNA Analysis Lineage Markers Lineage markers are passed down from generation to generation without changing Except for rare mutation events They can help determine the lineage (family tree)
Mitochondrial DNA Analysis Lineage Markers Lineage markers are passed down from generation to generation without changing Except for rare mutation events They can help determine the lineage (family tree)
CHAPTER 40 The Mechanism of Protein Synthesis
 CHAPTER 40 The Mechanism of Protein Synthesis Problems: 2,3,6,7,9,13,14,15,18,19,20 Initiation: Locating the start codon. Elongation: Reading the codons (5 3 ) and synthesizing protein amino carboxyl.
CHAPTER 40 The Mechanism of Protein Synthesis Problems: 2,3,6,7,9,13,14,15,18,19,20 Initiation: Locating the start codon. Elongation: Reading the codons (5 3 ) and synthesizing protein amino carboxyl.
