ACCEPTED. reduced expression of sars. Jan Oscarsson*, Anna Kanth, Karin Tegmark-Wisell and Staffan Arvidson
|
|
|
- Monica Manning
- 4 years ago
- Views:
Transcription
1 JB Accepts, published online ahead of print on September 00 J. Bacteriol. doi:0./jb.00-0 Copyright 00, American Society for Microbiology and/or the Listed Authors/Institutions. All Rights Reserved. 0 0 SarA is basically a repressor of hla ( -hemolysin) transcription in Staphylococcus aureus: its apparent role as an activator of hla in the prototype strain depends on reduced expression of sars. Jan Oscarsson*, Anna Kanth, Karin Tegmark-Wisell and Staffan Arvidson Department of Microbiology, Tumor and Cell Biology (MTC), Box 0, Karolinska Institutet, S- Stockholm, Sweden. Running title: regulation of hla by sara in S. aureus *Corresponding author: Jan Oscarsson, Department of Microbiology, Tumor and Cell Biology, Karolinska Institutet, S- Stockholm, Sweden. Phone (), Fax (), Mail: jan.oscarsson@ki.se.present address: Clinical Bacteriology Laboratory, Huddinge University Hospital, S- Stockholm, Sweden Key Words: Staphylococcus aureus, hla ( -hemolysin), regulation, sara, sars, Downloaded from on October, 0 by guest
2 0 0 Abstract In most Staphylococcus aureus strains inactivation of sara increases hla transcription, indicating that sara is a repressor. However, in S. aureus NCTC and its derivatives, used for most studies of hla regulation, inactivation of sara resulted in decreased hla transcription. The disparate phenotype of strain seems associated with its rsbu mutation leading to B deficiency. This has now been verified by the demonstration that sara repressed hla transcription in an rsbu + derivative of strain - (SH000). That sara could act as a repressor of hla in an - background was confirmed by the observation that inactivation of sara in an agr sars rot triple mutant dramatically increased hla transcription to wild-type levels. However, the apparent role of sara as an activator of hla in - was not a result of the rsbu mutation alone as inactivation of sara in another rsbu mutant, strain V, lead to increased hla transcription. Northern blot analysis revealed much higher levels of sars mrna in strain V than in -, which was likely due to the mutation in the sars activator, tcar, in -, which was not found in strain V. On the other hand, the relative increase in sars transcription upon inactivation of sara was -fold higher in - relative to strain V. Because of this, inactivation of sara in - means a net increase in repressor activity, whereas in strain V inactivation of sara means a net decrease in repressor activity, and therefore enhanced hla transcription. Introduction Staphylococcous aureus is a common human pathogen, which also colonizes the nares and skin of about / of all healthy people. The type of infection caused by this organism ranges from superficial cutaneous infections to life-threatening bacteremias. The pathogenesis of S. aureus is Downloaded from on October, 0 by guest
3 0 0 very complex and virulence depends on the production of an array of extracellular toxins and enzymes, and adhesins, which are regulated by a number of global regulators, e.g. agr (accessory gene regulator) and the sara (staphylococcal accessory regulator) family of regulators (reviewed in (,, 0)). In addition, the alternative sigma factor, sigma B ( B ), seems to be involved in virulence gene expression in addition to its role in regulating stress responses and intermediary metabolism (,,, ). Notably, among the genes controlled by sigma B are the virulence regulators sara and sars (, 0, ). The activity of B is regulated by the anti-sigma factor, RsbW, which binds B thereby inhibiting its association with the RNA-polymerase (, ). The ability of RsbW to bind B depends on the phosphorylation status of RsbV, which is determined by the activity of the phosphatase RsbU (,, 0). Most S. aureus strains produce -hemolysin ( -toxin; hla), which is a kda pore-forming protein that can lyse a wide range of human cells, including lymphocytes and keratinocytes (,, ). In addition, -hemolysin may induce apoptosis in T lymphocytes (), and has also been shown to be required for biofilm formation (0). Similar to most secreted toxins and enzymes in S. aureus, expression of -hemolysin (hla) is positively regulated by agr (,, ). The effector molecule of the agr-dependent regulation is RNAIII, which is transcribed from the agr P promoter at the end of the exponential phase of growth in response to autoactivation of the agr P promoter. Autoactivation is achieved via accumulation of a secreted pheromone, AIP, encoded by the agr operon (0,, ). RNAIII appears to increase -hemolysin expression by three different routes, via a direct interaction with the hla transcript that stimulates translation (), and via rot and sars-dependent pathways respectively. The rot gene locus was first recognized as a suppressor mutation, which partially restored -hemolysin (hla) expression in an Downloaded from on October, 0 by guest
4 0 0 agr null mutant (). Gene array methodology has revealed that rot acts as a global regulator, affecting the transcription of many virulence genes, e.g. hlb, spa, geh, and sspa (). The central role of rot in S. aureus virulence gene regulation was recently demonstrated in a rabbit endocarditis model of infection (). Based on the observation that inactivation of rot had no effect on hla expression in an agr + background, and that transcription of rot was not affected by the agr mutation, it has been suggested that RNAIII acts by sequestering Rot, thereby preventing it from binding to target gene promoters (, ). In a recent study some evidence was presented that RNAIII may even promote degradation of Rot via a clpxp-dependent route (). As rot stimulates transcription of sars (, ), which is a strong repressor of hla (), the suppressive effect of rot on hla transcription is probably indirect, mediated by sars. Whether rot is also a direct repressor of hla transcription is not known. Transcription of hla is also positively regulated by saesr, which encodes a two-component signal transduction system (). Transcription of the sae locus was clearly reduced in both agr and sara mutants during post-exponential phase of growth, indicating that sae is downstream of both RNAIII and sara in the regulatory pathway (). However, the level of hla transcription was approximately ten times higher in the sara relative to the agr mutant (), indicating that sara and sae are also epistatic. In the original S. aureus sara mutant isolated by Cheung and coworkers () production of hemolysin was increased in comparison to the parental strain. The same results were obtained when sara was inactivated in a number of other clinical S. aureus strains (, ). However, when the sara mutation was transferred to the prototype S. aureus strain NTCC it resulted in reduced -hemolysin production, indicating that sara acted as an activator of hla transcription (,, ). Most further studies of hla regulation in S. aureus have been carried out with strains Downloaded from on October, 0 by guest
5 0 0 derived from. In this strain, the sara-dependent stimulation of hla transcription seems to be mediated by sars, which is negatively regulated by sara and acts as a repressor of hla (). However, it has been reported that sara also enhances transcription of hla via agr. This effect seems to be direct, as sara has been shown to bind to the agr promoter region (,,, ), but might also be mediated by sart and saru (, ). Although the reason for the disparate phenotype of strain is not known, it seems associated with the B -deficiency of this strain and its derivatives (), because of a mutation in the rsbu gene (). In addition, strain - has a mutation in the tcar gene locus that could indirectly affect hla expression as tcar stimulates transcription of sars (a repressor of hla) (). Restoration of the B activity in strain derivatives dramatically decreased hla expression relative to the parental strains (, ). This was likely sara-independent, as the overall levels of sara transcription in strains - and SH000 (-, rsbu + ) were very similar (). However, other studies, using rsbu + derivatives of -, revealed increased sara expression during post- exponential phase of growth, from the B dependent promoter (P) (, ). Interestingly, preliminary experiments in our laboratory revealed that inactivation of sara in SH000 (- rsbu + ) resulted in increased hemolysin production on rabbit blood agar, suggesting that sara acts as a repressor of hla in an rsbu + background contrary to the parental rsbu deficient strain, -. The present study was undertaken to understand why sara acts as an activator of hla transcription in -, while being a repressor in the rsbu + derivative (SH000) and in most clinical strains. Materials and Methods Downloaded from on October, 0 by guest
6 0 0 Bacterial strains, plasmids and cultivation conditions Bacterial strains and plasmids used in this study are listed in Table. S. aureus strains were grown on Nutrient agar (NA)-plates (Difco). For screening of protease and -hemolysin production NA-plates supplemented with casein or rabbit erythrocytes were used (, ). S. aureus strains were precultured in Tryptic Soy Broth (TSB) (Difco) for - hours. When required 0 µg ml - tetracycline, 0 µg ml - kanamycin, µg ml - erythromycin or µg ml - lincomycin was added to the culture media. Cells from precultures were centrifuged and inoculated in 00 ml of Brain Heart Infusion (BHI) (Difco) in -liter baffled flasks (culture/flask volume ratio :0) to give an optical density at 00nm (OD 00 ) of 0. and incubated on a rotary shaker (0 r.p.m) at C. For induction of xyla promoter constructs xylose was added to the culture media to a final concentration of 0.%. Escherichia coli strains were cultivated on Luria-Bertani (LB) (Difco) agar plates or in LB medium, when required supplemented with 00 µg ml - ampicillin. Construction of strains and plasmids All plasmids were constructed and cloned in E. coli DH, as described (0), and subsequently transferred to the restriction deficient S. aureus strain RN0 by electroporation () before transfer to appropriate S. aureus strains using phage (). For construction of the sars allelic replacement mutant, WA0, PCR fragments (0 and 0 bp) flanking the sars gene, were synthesized using the primer pairs, sarh-forward ( - TATATAGCATGCCAAATACGCCCGACAACACTC- ) and sarh-reversed ( - TATATACCGCGGCATATCGACTTCAGGCTTGAC- ), and spaf ( - CTGTTAGAGCTCTCAATAATTTAAAAAAGC- ) and sarh-back ( - Downloaded from on October, 0 by guest
7 0 0 TATATAGAGCTCGTTATCAGCTTTCGGTGCTTG- ), respectively. Forward primers contained SphI and SacI restriction sites and reverse primers contained SacII and SacI restriction sites (underlined sequences). The PCR fragments were inserted in two steps at either side of the ermb cassette in the plasmid pkt. In the resulting construct (paw) a tetracycline resistance cassette from plasmid pt was inserted between the ApaI and AatII sites to generate paw. This plasmid was then transferred to S. aureus RN0 by electroporation, and integration in the sars locus was confirmed by PCR analysis. The sars::ermb allele was transferred to S. aureus - via -mediated transduction in order to obtain the double crossover that replaced the sars gene by the ermb cassette. Erythromycin/lincomycin resistant transductants were screened for loss of tetracycline resistance and the sars::ermb allele in one selected clone, WA0, was verified by PCR analysis. The hla::ermb allele of strain DU00 was moved to KT00, generating the derivative WA. The rot::tet allele from strain PM rot::tet was transferred to WA0, generating WA. The sara::km allele from PC was transferred to SH000, V, WA, WA0, WA, and PM rot::tet, generating the derivatives KT00, WA, WA00, WA0, WA, and WA0. The sars::ermb allele from WA0 was transferred to SH000, PC, and KT00, resulting in the derivatives WA0, WA0, and WA0. The construct pkt0, containing the sars gene under the control of the xyla promoter, was transferred to WA0 resulting in the strain WA. The construct pkt0, containing the sara gene under the control of the xyla promoter, was transferred to WA0, generating WA. The construct pik, containing sigb under the control of the xyla promoter, was transferred to strain WA0, generating WA0. Sequence analysis of the tcar operon Downloaded from on October, 0 by guest
8 0 0 The tcar gene locus of strain V was amplified by PCR and sequenced in both directions using the oligonucleotide primers tcar ( - CAATGATTGCTGAGGACAGCTGATAAAAATAGTAATTAG- ) and tcar ( - CAATTCTGTTCCTCTTCAGCTGTTATATGTATATCAC- ), which were based on the tcar sequence of strain COL (GeneBank accession no. AY00). DNA sequence analysis was performed using the ABI PRISM BigDye terminator Cycle Sequencing kit v. (Applied Biosystems), and the Applied Biosystem DNA sequencer. Our nucleotide sequence data for the tcar locus in strain V have been deposited in the EMBL database as entry AM0. Northern blot analysis Total S. aureus RNA was prepared using the FAST RNA-blue kit (Bio 0) according to instructions of the manufacturer. Concentration of RNA was determined by measuring the absorbance at 0nm. Samples containing 0µg of total RNA were analyzed by Northern blotting as described previously (). For Northern hybridization, internal fragments of hla (nucleotides [nt] to ; GeneBank accession number X0), RNAIII (nt 0-; X), sara (nt to 0; U), sars (nt 0-; locus SACOL00: CP0000), and s rrna (nt -0; X) were amplified by PCR, radio labeled with ( - P)-dCTP (Amersham) using a random prime labeling kit (Roche Molecular Biochemicals) and used as probes. Radioactivity was detected by a radioisotope imaging system (Phosphorimager SI; Molecular Dynamics), and quantified using the ImageQuant software (Molecular Dynamics). Each experiment was repeated three times. Analysis of -hemolysin in culture supernatants Downloaded from on October, 0 by guest
9 0 0 For Western blot analysis of -hemolysin, S. aureus culture supernatants were harvested at postexponential phase (h) and an amount of supernatant corresponding to a bacterial density of 0. OD 00 units (equal to ml OD 00 = 0.) was separated in % SDS-PAGE and transferred to polyvinylidene difluoride (PVDF) based membranes. Membranes were incubated with monoclonal mouse anti- -hemolysin antibodies (), and bound antibodies were subsequently detected using horseradish peroxidase (HRP)-conjugated sheep anti-mouse antibodies (Amersham). RESULTS Whether sara acts as a repressor or activator of hla in strain - depends on the rsbu activity Inactivation of sara in several clinical S. aureus strains resulted in increased -hemolysin production, indicating that sara is a repressor of hla (, ). However, in strain - (RN0) sara appeared to act as an activator of hla transcription, as inactivation of sara resulted in decreased hla expression (,, ). It has been suggested that the disparate phenotype of - could be due to the rsbu mutation of this strain, resulting in B - deficiency (). To analyze this further, we inactivated the sara gene locus of SH000 (- rsbu + ) by allelic replacement, generating strain KT00. As indicated by slightly increased hemolysis on rabbit blood agar (Fig. A) expression of hla seemed to be upregulated in KT00 relative to SH000. Immunoblotting and inactivation of the hla locus in KT00 confirmed that the increased zone of hemolysis was indeed due to production of -hemolysin (Fig. B and C). The enhanced -hemolysin production of KT00 (SH000 sara) was accompanied by elevated (- to -fold) hla mrna Downloaded from on October, 0 by guest
10 0 0 levels in KT00 relative to SH000, during post-exponential phase of growth (Fig. ), indicating that sara acts as a repressor of hla transcription in SH000. Taken together, our findings suggest that the apparent role of sara as an activator of hla transcription in strain - depends on the rsbu activity. The immunoblot analysis (Fig. C) also revealed partial degradation of -hemolysin in KT00, likely a result of derepressed protease expression in the absence of SarA, consistent with previous observations (, 0). The kda protein band in supernatants from SH000 (Fig. C) represents the extracellular fraction of protein A, as indicated by its reactivity with non-specific IgG, and by its pronounced upregulation in an agr mutant (data not shown). The lack of protein A in KT00 was probably due to increased production of proteases, which is consistent with previous results showing that protein A is degraded by extracellular proteases (). The reduced protein A production in strain WA0 (SH000 sars) is in agreement with sars being an activator of spa (). Whether sara acts as a repressor or activator of hla depends on the relative activity of sars To test whether the effect of rsbu on hla regulation by sara in - is a general phenomenon, we analyzed another natural B deficient S. aureus strain, V, which has an IS element inserted in the rsbu gene (). In this strain, inactivation of sara resulted in a - to -fold increase in hla transcription during post-exponential phase of growth (h and h) (Fig. A), indicating that it is not B activity per se that determines whether sara acts as a repressor or activator of hla transcription in a given strain, but other factors are also involved. As a result of the B deficiency strains - and V had low levels of the B -dependent sara P transcript during postexponential phase of growth (). As sara-dependent regulation of hla transcription is partly mediated by sars, which is a repressor of hla (), the level of sars mrna was determined in Downloaded from on October, 0 by guest 0
11 0 0 -, SH000 and V, and in their corresponding sara mutants (PC, KT00 and WA). In agreement with previous findings (), sars mrna levels were almost not detectable in - (Fig. and Fig. B). On the other hand, we observed a - to -fold increase in sars mrna levels in SH000 relative to - (Fig, ). This is consistent with the observation that transcription of sars is enhanced by B (, ). However, we did not detect the B -dependent sars transcript (P) in SH000, but only increased levels of the A -dependent sars transcript (P, same as the sars transcript that was upregulated in the - sara mutant, PC ()) (Fig. ). This indicates that the upregulation of sars in SH000 is not a direct effect of the B activity. As shown in Fig. B, the level of sars transcription was much higher in strain V than in -. A possible explanation would be that the tcar gene locus, which encodes an activator of sars transcription, is inactive in - (). DNA sequence analysis (see Materials and Methods) confirmed that the tcar allele in strain V was intact and identical to that of strain COL (GenBank accession no. AY00). Consistent with previous results (), inactivation of sara in - (strain PC) resulted in highly elevated (more than 0-fold) sars transcription during post-exponential phase of growth (Fig. and Fig. B). Interestingly, inactivation of sara in SH000 or in V only resulted in an approximately twofold increase in sars transcription (Fig. and Fig. B). Therefore, whether inactivation of sara will increase or decrease hla transcription seems to depend on the relative levels of sara and sars expression in the parental strain, and to what extent sars is upregulated in a cognate sara mutant. The reason why sara appears as an activator of hla transcription in - would thus be that the levels of both sara and sars are low (essentially no repression of hla transcription), and that inactivation of sara results in a very dramatic increase in sars transcription and therefore suppression of hla. To confirm that sara is basically acting as a repressor of hla transcription even in -, a Downloaded from on October, 0 by guest
12 0 0 plasmid carrying sara behind a xylose-inducible promoter (pkt0) was transferred to a sars mutant derived from strain -, resulting in WA. As shown in Fig., induction of sara expression with 0.0% xylose, which did not affect bacterial growth (data not shown), and only very slightly reduced RNAIII production at h and h, completely repressed hla transcription. This is in agreement with the observation that SarA bound to the hla promoter in vitro (, ) and repressed hla transcription in an in vitro assay (), supporting a model in which sara can also suppress hla transcription in a direct way. High hla transcription can be obtained in an agr mutant if sara, sars and rot are inactivated Our present results indicate that hla transcription is suppressed by three regulators, sara, sars and rot. As Rot, which is an activator of sars transcription (), is supposed to be neutralized by RNAIII (), the relative impact of all three repressors was analyzed in an agr mutant. For this, we constructed agr sars sara and agr sara rot triple mutants (WA and WA0), and an agr sars rot sara quadruple mutant (WA0). As shown in Fig. A, hla mrna levels were equally low, relative to the parental strain (RN0), in WA (agr sars sara), WA0 (agr sara rot), and in an agr sars rot triple mutant (WA0). Interestingly, the quadruple mutant (WA0) had clearly increased (up to -fold) hla mrna levels relative to the triple mutants, during post-exponential phase of growth (h and h) (Fig. A). These results indicate that sara, sars and rot are equally potent and sufficient to suppress hla transcription. As rot appeared to suppress hla transcription independent of sars and sara this regulatory effect might be direct. It can also be concluded that RNAIII per se was not required for high transcription of hla in the absence of repression by sara, sars and rot, as the level of hla transcription was very similar in Downloaded from on October, 0 by guest
13 0 0 the quadruple mutant (WA0) and the parental strain, RN0. However, it must be pointed out that the quadruple mutant produced much less -hemolysin than the parental strain (Fig. B and data not shown), confirming that RNAIII is still essential for translation of the hla mrna (). The B -mediated reduction in hla transcription in strain SH000 is independent of sara, sars and rot Considering that both sara and sars can act as repressors of hla transcription, the reason for the low hla mrna levels in strain SH000 would be the increased B -dependent expression of these regulators during post-exponential phase of growth. To further investigate this, the relative impact of sara and sars on hla transcription in SH000 was analyzed by comparing the hla mrna levels in sara and sars single and sara sars double mutants, derived from SH000 (Fig., and data not shown). As hla transcription remained very low in all mutants compared to -, we concluded that sara and sars were not the sole factors responsible for the low hla transcription in SH000. In agreement with this, induction of B expression in an - sara sars double mutant, containing a xylose-inducible sigb gene (strain WA0), resulted in severe suppression of hla transcription (Fig. ), confirming that sara and sars were not required for B - dependent repression of hla. Notably, RNAIII production was impaired (- to -fold) upon induction of B in WA0 (Fig. ), suggesting that the reduction in hla transcription was the result of low RNAIII levels. A similar reduction of RNAIII levels was observed in SH000 relative to its parental strain ()(Fig. ). As RNAIII was not required for transcription of hla in derivatives of - lacking sara, sars and rot (Fig. A), the impact of the reduced RNAIII levels in SH000 on hla transcription was analyzed by construction of a sara sars rot triple Downloaded from on October, 0 by guest
14 0 0 mutant of SH000 (WA00). As hla transcription was hardly upregulated in WA00 relative to SH000 (Fig. ), we concluded that the B -mediated reduction in hla transcription in SH000 was independent of sara, sars and rot, or reduced RNAIII levels. Notably, although inactivation of sars in SH000 derivatives had no marked effect on hla mrna levels (Fig. and data not shown), these strains (WA0 and WA0) were completely non-hemolytic on rabbit blood agar (Fig. A) and lacked detectable -hemolysin (Fig. C and data not shown). This is consistent with the reduced (up to -fold) RNAIII levels in these strains (Fig. and data not shown), as RNAIII is required for translation of the hla mrna (). As complementation of WA0 with pkt0, encoding sars, restored RNAIII expression to parental (SH000) levels (Fig. ), we concluded that sars enhanced RNAIII production in SH000, in contrast to in -. Discussion The present study was undertaken to find an explanation to why sara appears to be an activator of hla transcription in the prototype strain NTCC and its derivatives, while being a repressor of hla in most other S. aureus strains (, ). Our results clearly show that the role of sara as an activator of hla transcription in - was associated with the rsbu mutation in this strain, which leads to B deficiency, as sara acted as a repressor of hla in an rsbu + derivative of - (SH000) (Fig. and Fig. ). However, this was not a direct effect of the B deficiency, as inactivation of sara in another B deficient strain, V, resulted in increased hla transcription (Fig. A). The possibility that strains V and - respond differently to sara inactivation because of their different rsbu mutations can not be excluded. Our results indicate that whether inactivation of sara leads to increased or decreased hla transcription depends on the relative Downloaded from on October, 0 by guest
15 0 0 activity of sara and sars, respectively, i.e., their basal levels of expression, and how much sars transcription is elevated when sara is inactivated. In the parental strain -, the basal postexponential levels of sara and sars mrnas are low due to the impaired B activity ( B enhances sara and sars transcription (, )) and the mutation in tcar (tcar stimulates sars transcription ()). In strain V, which was found to have an intact tcar locus, the level of sars mrna was much higher than in strain - (Fig. B), while the post-exponential level of sara expression was essentially the same in these strains (). We assume that the level of sara in these strains is sufficient to partly repress hla transcription, whereas the suppressive effect of sars is only significant in strain V, which has a lower level of hla mrna than - (our unpublished data). However, inactivation of sara resulted in about -fold higher relative increase in sars transcription in - than in V (Fig. and Fig. B). A likely explanation why this lead to reduced hla transcription in strain -, and increased hla transcription in strain V, would be that the loss of repression of hla due to inactivation of sara in - was counteracted by the massive increase in sars, whereas the level of repression of hla in response to the increase in sars transcription in the V sara mutant (WA) was less than the derepression of hla due to loss of sara activity. Our present findings strongly suggest that sara is basically a repressor of hla transcription in S. aureus strains, including - and its derivatives. Repression of hla transcription by sara seems to be independent of agr (RNAIII), sars and rot. As indicated in Fig., transcription of hla is negatively controlled by three regulators, sara, sars and rot, all of which had to be inactivated or neutralized to achieve full hla transcription. It has been reported that hla transcription is also negatively controlled by sart (). However, it is not clear whether this Downloaded from on October, 0 by guest
16 0 0 effect is direct or mediated by sars, which is stimulated by sart (). On the other hand, as sart is strongly repressed by both sara and agr (), the possible direct repression of hla by sart seems not to be of major importance, as very similar hla mrna levels were obtained in the wildtype (sart repressed by both sara and agr) and in an agr sara sars rot quadruple mutant (Fig. A). The positive effect of RNAIII on hla transcription seems mainly be due to its capability of downregulating sars transcription via neutralization of Rot and repression of sart, which are both required for sars transcription (,,, ). This is consistent with the observation that RNAIII was not required for hla transcription in the absence of sara, sars and rot (Fig. A). Interestingly, inactivation of these regulators in strain SH000 (-, rsbu + ) did not result in elevated hla transcription (Fig. ), indicating that other B -dependent factor(s) (indicated by X with an asterisk in Fig. ) are responsible for the low hla transcription in this strain. The observed requirement of an intact sars allele for RNAIII production in SH000 (Fig. and Fig. ) might be explained by sars acting as a negative regulator of a B -dependent repressor of agr (RNAIII) expression. This and additional B -dependent factors involved in the regulation of hla transcription might be found among the hypothetical proteins identified in a micro array based analysis of the B regulon (). Although the much studied and genetically well characterized strain - and its derivatives seem not representative of most clinical S. aureus isolates because of the mutations in rsbu and tcar, it is still very useful in exploration of the regulatory networks governing virulence gene expression if compared to other strains. ACKNOWLEDGEMENTS We are grateful to Agneta Wahlquist, Lena Norenius and Caroline Harlos for valuable technical assistance. We wish to thank Drs. Simon Foster and Tim Foster for sending us the strains Downloaded from on October, 0 by guest
17 0 0 0 SH000 and DU00, respectively. This work was financed by funds from the Swedish Research Council (VR), the Swedish Foundation for Strategic Research (SSF), and by the Swedish Society for Medical Research (SSMF). REFERENCES. Arvidson, S. 00. Virulence gene regulation and their role in pathogenesis of disease, p. -. In D. Ala'Aldeen and K. Hiramatsu (ed.), Staphylococcus aureus, molecular and clinical aspects. Horwood Publishing, Chichester.. Arvidson, S., and K. Tegmark. 00. Regulation of virulence determinants in Staphylococcus aureus. Int. J. Med. Microbiol. :-0.. Benson, A. K., and W. G. Haldenwang.. Bacillus subtilis B is regulated by a binding protein (RsbW) that blocks its association with core RNA polymerase. Proc. Natl. Acad. Sci. USA 0:0-.. Bischoff, M., P. Dunman, J. Kormanec, D. Macapagal, E. Murphy, W. Mounts, B. Berger-Bächi, and S. Projan. 00. Microarray-based analysis of the Staphylococcus aureus sigmab regulon. J. Bacteriol. :0-0.. Björklind, A., and S. Arvidson. 0. Mutants of Staphylococcus aureus affected in the regulation of exoprotein synthesis. FEMS Microbiol. Lett. :0-0.. Björklind, A., and S. Arvidson.. Occurrence of an extracellular serineproteinase among Staphylococcus aureus strains. Acta. Path. Microbiol. Scand. :-0.. Blevins, J., A. Gillaspy, T. Rechtin, B. Hurlburt, and M. Smeltzer.. The Staphylococcal accessory regulator (sar) represses transcription of the Staphylococcus aureus collagen adhesin gene (cna) in an agr-independent manner. Mol. Microbiol. :-.. Blevins, J. S., K. E. Beenken, M. O. Elasri, B. K. Hurlburt, and M. S. Smeltzer. 00. Strain-dependent differences in the regulatory roles of sara and agr in Staphylococcus aureus. Infect. Immun. 0:0-0.. Bronner, S., H. Monteil, and G. Prevost. 00. Regulation of virulence determinants in Staphylococcus aureus: complexity and applications. FEMS Microbiol. Rev. : Caiazza, N. C., and G. A. O'Toole. 00. Alpha-toxin is required for biofilm formation by Staphylococcus aureus. J Bacteriol :-.. Chan, P. F., and S. J. Foster.. Role of SarA in virulence determinant production and environmental signal transduction in Staphylococcus aureus. J. Bacteriol. 0:-.. Chan, P. F., S. J. Foster, I. Ingham, and M. O. Clements.. The Staphylococcus aureus alternative sigma factor B controls the environmental stress response but not starvation survival or pathogenicity in a mouse abscess model. J. Bacteriol. 0:0-0. Downloaded from on October, 0 by guest
18 Cheung, A. L., M. G. Bayer, and J. H. Heinrichs.. sar genetic determinants necessary for transcription of RNAII and RNAIII in agr locus of Staphylococcus aureus. J. Bacteriol. :-.. Cheung, A. L., J. M. Koomey, C. A. Butler, S. J. Projan, and V. A. Fischetti.. Regulation of exoprotein expression in Staphylococcus aureus by a locus (sar) distinct from agr. Proc. Natl. Acad. Sci. USA :-.. Cheung, A. L., K. Schmidt, B. Bateman, and A. C. Manna. 00. SarS, a SarA homolog repressible by agr, is an activator of protein A synthesis in Staphylococcus aureus. Infect. Immun. :-.. Cheung, A. L., and P. Ying.. Regulation of alpha- and beta-hemolysins by the sar locus of Staphylococcus aureus. J. Bacteriol. :0-.. Chien, Y., and A. L. Cheung.. Molecular interactions between two global regulators, sar and agr, in Staphylococcus aureus. J. Biol. Chem. :-.. Chien, Y., A. Manna, and A. Cheung.. SarA level is a determinant of agr activation in Staphylococcus aureus. Mol. Microbiol. 0:-00.. Chien, Y. T., A. C. Manna, S. J. Projan, and A. L. Cheung.. SarA, a global regulator of virulence determinants in Staphylococcus aureus, binds to a conserved motif essential for sar-dependent gene regulation. J. Biol. Chem. :-. 0. Deora, R., T. Tseng, and T. K. Misra.. Alternative transcription factor B of Staphylococcus aureus: characterization and role in transcription of the global regulatory locus sar. J. Bacteriol. :-.. Dinges, M. M., P. M. Orwin, and P. M. Schlievert Exotoxins of Staphylococcus aureus. Clin. Microbiol. Rev. :-.. Dufour, A., and W. G. Haldenwang.. Interactions between a Bacillus subtilis antisigma factor (RsbW) and its antagonist (RsbV). J. Bacteriol. :-0.. Frees, D., K. Sorensen, and H. Ingmer. 00. Global virulence regulation in Staphylococcus aureus: pinpointing the roles of ClpP and ClpX in the sar/agr regulatory network. Infect. Immun. : Giachino, P., S. Engelmann, and M. Bischoff. 00. B Activity Depends on RsbU in Staphylococcus aureus. J. Bacteriol. :-.. Giraudo, A. T., A. L. Cheung, and R. Nagel.. The sae locus of Staphylococcus aureus controls exoprotein synthesis at the transcriptional level. Arch. Microbiol. :-.. Hanahan, D.. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. :-0.. Heinrichs, J. H., M. G. Bayer, and A. L. Cheung.. Characterization of the sar locus and its interaction with agr in Staphylococcus aureus. J. Bacteriol. :-.. Horsburgh, M. J., J. L. Aish, I. J. White, L. Shaw, J. K. Lithgow, and S. J. Foster. 00. B modulates virulence determinant expression and stress resistance: charecterization of a functional rsbu strain derived from Staphylococcus aureus. J. Bacteriol. :-.. Janzon, L., S. Löfdahl, and S. Arvidson.. Evidence for a coordinate transcriptional control of alpha-toxin and protein A in Staphylococcus aureus. FEMS Microbiol. Lett. :-. Downloaded from on October, 0 by guest
19 Ji, G., R. C. Beavis, and R. P. Novick.. Cell density control of staphylococcal virulence mediated by an octapeptide pheromone. Proc. Natl. Acad. Sci. USA :0-0.. Jonas, D., I. Walev, T. Berger, M. Liebetrau, M. Palmer, and S. Bhakdi.. Novel path to apoptosis: small transmembrane pores created by staphylococcal alpha-toxin in T lymphocytes evoke internucleosomal DNA degradation. Infect. Immun. :0-.. Karlsson, A., and S. Arvidson. 00. Variation in extracellular protease production among clinical isolates of Staphylococcus aureus due to different levels of expression of the protease repressor sara. Infect. Immun. 0:-.. Karlsson, A., P. Saravia-Otten, K. Tegmark, E. Morfeldt, and S. Arvidson. 00. Decreased amounts of cellwall-associated protein A and fibronectin-binding proteins in Staphylococcus aureus sara mutants due to up-regulation of extracellular proteases. Infect. Immun. :-.. Karlsson-Kanth, A., K. Tegmark-Wisell, S. Arvidson, and J. Oscarsson. 00. Natural human isolates of Staphylococcus aureus selected for high production of proteases and alpha-hemolysin are sigma B-deficient. Int. J. Med. Microbiol. :-.. Khan, S. A., G. K. Adler, and R. P. Novick.. Functional origin of replication of pt plasmid DNA is contained within a -base-pair segment. Proc. Natl. Acad. Sci. USA :0-.. Kreiswirth, B., S. Löfdahl, M. J. Betley, M. O'Reilly, M. P. Schleivert, M. S. Bergdoll, and R. P. Novick.. The toxic shock syndrome exotoxin structural gene is not detectably transmitted by a prophage. Nature 0:0-.. Kullik, I., and P. Giachino.. The alternative sigma factor B in Staphylococcus aureus: regulation of the sigb operon in response to growth and heat shock. Arch. Microbiol. :-.. Kullik, I., P. Giachino, and T. Fuchs.. Deletion of the alternative sigma factor B in Staphylococcus aureus reveals its function as a global regulator of virulence genes. J. Bacteriol. 0:-0.. Lina, G., S. Jarraud, G. Ji, T. Greenland, A. Pedraza, J. Etienne, R. P. Novick, and F. Vandenesch.. Transmembrane topology and histidine protein kinase activity of AgrC, the agr signal receptor in Staphylococcus aureus. Mol. Microbiol. :-. 0. Lindsay, J., and S. Foster.. Interactive regulatory pathways control virulence determinant production and stability in response to environmental conditions in Staphylococcus aureus. Mol. Gen. Genet. :-.. Manna, A. C., and A. L. Cheung. 00. saru, a sara homolog, is repressed by SarT and regulates virulence genes in Staphylococcus aureus. Infect. Immun. :-.. Mayville, P., G. Ji, R. Beavis, H. Yang, M. Goger, R. P. Novick, and T. W. Muir.. Structure-activity analysis of synthetic autoinducing thiolactone peptides from Staphylococcus aureus responsible for virulence. Proc. Natl. Acad. Sci. USA :-.. McCallum, N., M. Bischoff, H. Maki, A. Wada, and B. Berger-Bächi. 00. TcaR, a putative MarR-like regulator of sars expression. J. Bacteriol. :-.. McNamara, P. J., and A. S. Bayer. 00. A rot mutation restores parental virulence to an agr-null Staphylococcus aureus strain in a rabbit model of endocarditis. Infect. Immun. :0-0. Downloaded from on October, 0 by guest
20 McNamara, P. J., K. C. Milligan-Monroe, S. Khalili, and R. A. Proctor Identification, cloning, and initial characterization of rot, a locus encoding a regulator of virulence factor expression in Staphylococcus aureus. J. Bacteriol. :-0.. Miyazaki, E., J. M. Chen, C. Ko, and W. R. Bishai.. The Staphylococcus aureus rsbw (orf) gene encodes an anti-sigma factor of SigB. J. Bacteriol. :-.. Morfeldt, E., L. Janzon, S. Arvidson, and S. Löfdahl.. Cloning of a chromosomal locus (exp) which regulates the expression of several exoprotein genes in Staphylococcus aureus. Mol. Gen. Genet. :-0.. Morfeldt, E., D. Taylor, A. von Gabain, and S. Arvidson.. Activation of alphatoxin translation in Staphylococcus aureus by the trans-encoded antisense RNA, RNAIII. EMBO J. :-.. Morfeldt, E., K. Tegmark, and S. Arvidson.. Transcriptional control of the agrdependent virulence gene regulator, RNAIII, in Staphylococcus aureus. Mol. Microbiol. :-. 0. Novick, R. P. 00. Autoinduction and signal transduction in the regulation of staphylococcal virulence. Mol. Microbiol. :-.. Novick, R. P.. Genetic systems in staphylococci. Methods Enzymol. 0:-.. Novick, R. P.. Properties of a cryptic high-frequency transducing phage in Staphylococcus aureus. Virology :-.. Novick, R. P., and D. Jiang. 00. The staphylococcal saers system coordinates environmental signals with agr quorum sensing. Microbiology :0-.. Novick, R. P., H. F. Ross, S. J. Projan, J. Kornblum, B. Kreiswirth, and S. Moghazeh.. Synthesis of staphylococcal virulence factors is controlled by a regulatory RNA molecule. EMBO J. :-.. O'Reilly, M., J. C. de Azavedo, S. Kennedy, and T. J. Foster.. Inactivation of the alpha-haemolysin gene of Staphylococcus aureus - by site-directed mutagenesis and studies on the expression of its haemolysins. Microb. Pathog. :-.. Oscarsson, J., C. Harlos, and S. Arvidson. 00. Regulatory role of proteins binding to the spa (protein A) and sars (staphylococcal accessory regulator) in Staphylococcus aureus NTCC -. Int. J. Med. Microbiol. :-.. Oscarsson, J., K. Tegmark-Wisell, and S. Arvidson. 00. Coordinated and differential control of aureolysin (aur) and serine protease (sspa) transcription in Staphylococcus aureus by sara, rot and agr (RNAIII). Int. J. Med. Microbiol. :in press.. Recsei, P., B. Kreiswirth, M. O'Reilly, P. Schlievert, A. Gruss, and R. P. Novick.. Regulation of exoprotein gene expression in Staphylococcus aureus by agr. Mol. Gen. Genet. 0:-.. Said-Salim, B., P. M. Dunman, F. M. McAleese, D. Macapagal, E. Murphy, P. J. McNamara, S. Arvidson, T. J. Foster, S. J. Projan, and B. N. Kreiswirth. 00. Global regulation of Staphylococcus aureus genes by Rot. J. Bacteriol. : Sambrook, J. E., E. F. Fritsch, and T. Maniatis.. Molecular cloning: a laboratory manual, nd ed ed. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.. Schenk, S., and R. A. Laddaga.. Improved methods for electroporation of Staphylococcus aureus. FEMS Microbiol. Lett. :-.. Schmidt, K. A., A. C. Manna, and A. L. Cheung. 00. SarT influences sars expression in Staphylococcus aureus. Infect. Immun. :-. Downloaded from on October, 0 by guest 0
21 0 0. Schmidt, K. A., A. C. Manna, S. Gill, and A. L. Cheung. 00. SarT, a repressor of alpha-hemolysin in Staphylococcus aureus. Infect. Immun. :-.. Sterba, K. M., S. G. Mackintosh, J. S. Blevins, B. K. Hurlburt, and M. S. Smeltzer. 00. Characterization of Staphylococcus aureus SarA binding sites. J. Bacteriol. :0-.. Söderquist, B., P. Colque-Navarro, L. Blomqvist, P. Olcén, H. Holmberg, and R. Möllby.. Staphylococcal -toxin in septicemic patients; detection in serum, antibody response and production in isolated strains. Serodiagn. Immunother. Infect. Dis. :-.. Tegmark, K Regulation of virulence gene expression in Staphylococcus aureus. Karolinska Institutet, Stockholm.. Tegmark, K., A. Karlsson, and S. Arvidson Identification and characterization of SarH, a new global regulator of virulence gene expression in Staphylococcus aureus. Mol. Microbiol. :-0.. Voelker, U., A. Dufour, and W. G. Haldenwang.. The Bacillus subtilis rsbu gene product is necessary for RsbX-dependent regulation of B. J. Bacteriol. :-.. Walev, I., E. Martin, D. Jonas, M. Mohamadzadeh, W. Muller-Klieser, L. Kunz, and S. Bhakdi.. Staphylococcal alpha-toxin kills human keratinocytes by permeabilizing the plasma membrane for monovalent ions. Infect. Immun. :-. 0. Wise, A., and C. Price.. Four additional genes in the sigb operon of Bacillus subtilis that control activity of the general stress factor B in response to enviromental signals. J. Bacteriol. :-. FIGURE LEGENDS Downloaded from on October, 0 by guest
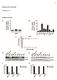
22 FIG.. A. Hemolysin production by strains - (rsbu) and SH000 (rsbu + ) and their corresponding sara (PC and KT00), sars (WA0 and WA0), and sara sars (WA0 and WA0) mutant derivatives, grown on a rabbit blood agar plate. B. Hemolysin production by KT00 (SH000 sara) and WA (SH000 sara hla), grown on a rabbit blood agar plate. C. Immunoblot analysis of -hemolysin in culture supernatants of PC (- sara) (), - (), SH000 ( and ), KT00 (SH000 sara) () and WA0 (SH000 sars) (). Downloaded from on October, 0 by guest
23 FIG.. Northern blot analysis of sars, sara, RNAIII, s rrna and hla in strain SH000 (rsbu + ) and its derivatives WA0 (sars) and KT00 (sara), and in strain - (rsbu) and its derivative, PC (sara). The main sara transcript seen is the sigma B-dependent sara P transcript, which is markedly enhanced in SH000. The lower panel (hla) shows a reduced exposure. Samples were taken at the specified time points during growth of a representative culture. Downloaded from on October, 0 by guest
24 FIG.. A. Northern blot analysis of hla and RNAIII in strain V and WA (V sara). B. Northern blot analysis of sars and s rrna in strain PC (- sara), -, WA (V sara), and V. Samples were taken at the specified time points during growth. FIG.. Northern blot analysis of hla, RNAIII and s rrna in WA (- sars, containing pkt0 with the xylose-inducible sara). Samples were taken at the indicated time points during growth in liquid medium with or without 0.0% xylose (final concentration). Downloaded from on October, 0 by guest
25 0 FIG.. A. Northern blot analysis of hla and s rrna in WA0 (agr sars rot sara), WA0 (agr sars rot), WA0 (agr rot sara), WA (agr sars sara), and RN0 (wt), grown in liquid cultures with samples taken at the indicated time points. B. Hemolysin production by strains from left to right, grown on a rabbit blood agar plate, row : - (wt) and WA (agr sars); row : PM rot::tet (agr rot) and WA0 (agr sars rot sara); row : PM (agr) and WA0 (agr sars rot). Downloaded from on October, 0 by guest
26 FIG.. Northern blot analysis of RNAIII and hla in WA0 (- sara sars, containing pik with the xylose-inducible sigb). Samples were taken at the indicated time points during growth in liquid medium with or without 0.0% xylose (final concentration). FIG.. Northern blot analysis of hla and s rrna in WA00 (sars sara rot triple mutant of SH000) and -. Samples were taken at the indicated time points during growth of a representative culture. Downloaded from on October, 0 by guest 0 FIG.. Effect of complementation of the sars mutant derived from SH000. Northern blot analysis of RNAIII in WA (SH000 sars with pkt0 carrying sars), WA0 (SH000
27 0 sars), and SH000. Samples were taken at the indicated time points during growth of a representative culture. sara sars RNAIII rot sart hla Rot FIG.. Schematic overview of regulatory interactions involving sara, sars, sart, rot, agr (RNAIII) and B in the control of hla transcription. Arrows indicate stimulation and bars repression. An asterisk (*) indicates genes known to be upregulated by B. In exponential phase of growth hla transcription is suppressed by sara, sars and rot (, )(Fig. ). Transcription of sars is stimulated by rot (, ) and sart (), and is suppressed by sara (, ). Transcription of sart is repressed by sara and agr (RNAIII) (). RNAIII expression is enhanced by sara (,, ). In post-exponential phase RNAIII is supposed to neutralize Rot activity (, ), whereas transcription of sara and sars is B -dependent (,, )(Fig. ). In addition, B seems to suppress hla transcription via mechanisms not involving sara, sars and rot (referred to as X) (Fig. ). For further explanation see text. X Downloaded from on October, 0 by guest
28 TABLE. Bacterial strains and plasmids used in this study Strain / plasmid Relevant characteristics Source E.coli DH S. aureus E.coli strain used for propagation of all plasmid constructs () - Prototype S. aureus strain, rsbu () DU00 -, hla::ermb (Em r ) () KS0 Clinical strain () KT00 SH000, sara::km (Km r ) This study NA KS0, sara::km (Km r ) () PC -, sara::km (Km r ) () PM RN0, agr null () PM rot::tet RN0, agr null, rot::tet (Tc r ) P.J. McNamara RN0 Restriction deficient mutant of - () Downloaded from on October, 0 by guest RN0 Laboratory S. aureus strain that is derived from -, () rsbu SH000 - with functional rsbu () V High protease producing strain from which the four ATCC
29 major staphylococcal proteases (sspa, sspb, aur and scp) were originally characterized WA0 -, sars::ermb (Em r ) This study WA0 SH000, sars::ermb (Em r ) This study WA0 -, sara::km, sars::ermb (Km r Em r ) This study WA0 -, sara::km, sars::ermb (pik) (Km r Em r Tc r ) This study WA0 SH000, sara::km, sars::ermb (Km r Em r ) This study WA WA0 (pkt0) (Tc r ) This study WA V, sara::km (Km r ) This study WA SH000, sars::ermb, rot::tet (Em r Tc r ) This study WA00 SH000, sars::ermb, sara::km, rot::tet (Em r Km r Tc r ) This study WA0 RN0, agr null, sars::ermb, rot::tet (Em r Tc r ) () WA0 RN0, agr null, sars::ermb, rot::tet, sara::km (Em r Tc r Km r ) This study WA KT00, hla::ermb (Em r ) This study WA RN0, agr null, sars::ermb (Em r ) () WA RN0, agr null, sars::ermb, sara::km (Em r Km r ) This study Downloaded from on October, 0 by guest WA0 RN0, agr null, rot::tet, sara::km (Tc r Km r ) This study WA -, sars::ermb (pkt0) (Em r Tc r ) This study Plasmids paw pkt with upstream and downstream fragments of the This study r
30 sars locus inserted at either side of ermb (Em r ) paw paw with Tc r cassette from pt (Tc r Em r ) This study pgem-t Easy E. coli cloning vector for PCR products (Amp r ) Promega pkt pgem-t Easy with ermb of Tn (Amp r Em r ) () pik pkt0 pkt0 S. aureus plasmid with sigb under control of xyla promoter (Tc r ) pspt with xylr and the sars under control of xyla promoter (Tc r ) S. aureus plasmid with sara under control of xyla promoter (Tc r ) () () () pspt Shuttle vector (Tc r ) () pt A tetracycline-resistant plasmid from S. aureus (Tc r ) () Amp r = resistance to ampicillin, Km r = resistance to kanamycin/neomycin, Em r = resistance to erythromycin/lincomycin, Tc r = resistance to tetracycline Downloaded from on October, 0 by guest 0
How To Understand How Gene Expression Is Regulated
 What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
Module 3 Questions. 7. Chemotaxis is an example of signal transduction. Explain, with the use of diagrams.
 Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College
 Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
Gene Transcription in Prokaryotes
 Gene Transcription in Prokaryotes Operons: in prokaryotes, genes that encode protein participating in a common pathway are organized together. This group of genes, arranged in tandem, is called an OPERON.
Gene Transcription in Prokaryotes Operons: in prokaryotes, genes that encode protein participating in a common pathway are organized together. This group of genes, arranged in tandem, is called an OPERON.
HCS604.03 Exercise 1 Dr. Jones Spring 2005. Recombinant DNA (Molecular Cloning) exercise:
 HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
Separate Mechanisms Activate B of Bacillus subtilis in Response to Environmental and Metabolic Stresses
 JOURNAL OF BACTERIOLOGY, July 1995, p. 3771 3780 Vol. 177, No. 13 0021-9193/95/$04.00 0 Copyright 1995, American Society for Microbiology Separate Mechanisms Activate B of Bacillus subtilis in Response
JOURNAL OF BACTERIOLOGY, July 1995, p. 3771 3780 Vol. 177, No. 13 0021-9193/95/$04.00 0 Copyright 1995, American Society for Microbiology Separate Mechanisms Activate B of Bacillus subtilis in Response
Effects of Antibiotics on Bacterial Growth and Protein Synthesis: Student Laboratory Manual
 Effects of Antibiotics on Bacterial Growth and Protein Synthesis: Student Laboratory Manual I. Purpose...1 II. Introduction...1 III. Inhibition of Bacterial Growth Protocol...2 IV. Inhibition of in vitro
Effects of Antibiotics on Bacterial Growth and Protein Synthesis: Student Laboratory Manual I. Purpose...1 II. Introduction...1 III. Inhibition of Bacterial Growth Protocol...2 IV. Inhibition of in vitro
2.1.2 Characterization of antiviral effect of cytokine expression on HBV replication in transduced mouse hepatocytes line
 i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
Recombinant DNA Technology
 Recombinant DNA Technology Dates in the Development of Gene Cloning: 1965 - plasmids 1967 - ligase 1970 - restriction endonucleases 1972 - first experiments in gene splicing 1974 - worldwide moratorium
Recombinant DNA Technology Dates in the Development of Gene Cloning: 1965 - plasmids 1967 - ligase 1970 - restriction endonucleases 1972 - first experiments in gene splicing 1974 - worldwide moratorium
Transfection-Transfer of non-viral genetic material into eukaryotic cells. Infection/ Transduction- Transfer of viral genetic material into cells.
 Transfection Key words: Transient transfection, Stable transfection, transfection methods, vector, plasmid, origin of replication, reporter gene/ protein, cloning site, promoter and enhancer, signal peptide,
Transfection Key words: Transient transfection, Stable transfection, transfection methods, vector, plasmid, origin of replication, reporter gene/ protein, cloning site, promoter and enhancer, signal peptide,
Recombinant DNA and Biotechnology
 Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Bacillus Subtilis Expression Vectors. Product Information and Instructions November 2005
 Bacillus Subtilis Expression Vectors Product Information and Instructions November 2005 1 Content 1. Introduction... 3 2. The pht Vectors...4 2.1. Vector Map pht01...4 2.2. Vector Map pht43...5 2.3. Location
Bacillus Subtilis Expression Vectors Product Information and Instructions November 2005 1 Content 1. Introduction... 3 2. The pht Vectors...4 2.1. Vector Map pht01...4 2.2. Vector Map pht43...5 2.3. Location
Milestones of bacterial genetic research:
 Milestones of bacterial genetic research: 1944 Avery's pneumococcal transformation experiment shows that DNA is the hereditary material 1946 Lederberg & Tatum describes bacterial conjugation using biochemical
Milestones of bacterial genetic research: 1944 Avery's pneumococcal transformation experiment shows that DNA is the hereditary material 1946 Lederberg & Tatum describes bacterial conjugation using biochemical
Genetics Lecture Notes 7.03 2005. Lectures 1 2
 Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA
 CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
Gene Expression Assays
 APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
Chapter 18: Applications of Immunology
 Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
The Need for a PARP in vivo Pharmacodynamic Assay
 The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D., Chief Scientific Officer, Trevigen, Inc., Gaithersburg, MD For further infomation, please contact: William Booth, Ph.D. Tel: +44 (0)1235
The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D., Chief Scientific Officer, Trevigen, Inc., Gaithersburg, MD For further infomation, please contact: William Booth, Ph.D. Tel: +44 (0)1235
Bacterial Transformation and Plasmid Purification. Chapter 5: Background
 Bacterial Transformation and Plasmid Purification Chapter 5: Background History of Transformation and Plasmids Bacterial methods of DNA transfer Transformation: when bacteria take up DNA from their environment
Bacterial Transformation and Plasmid Purification Chapter 5: Background History of Transformation and Plasmids Bacterial methods of DNA transfer Transformation: when bacteria take up DNA from their environment
BCOR101 Midterm II Wednesday, October 26, 2005
 BCOR101 Midterm II Wednesday, October 26, 2005 Name Key Please show all of your work. 1. A donor strain is trp+, pro+, met+ and a recipient strain is trp-, pro-, met-. The donor strain is infected with
BCOR101 Midterm II Wednesday, October 26, 2005 Name Key Please show all of your work. 1. A donor strain is trp+, pro+, met+ and a recipient strain is trp-, pro-, met-. The donor strain is infected with
Name Class Date. Figure 13 1. 2. Which nucleotide in Figure 13 1 indicates the nucleic acid above is RNA? a. uracil c. cytosine b. guanine d.
 13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
A and B are not absolutely linked. They could be far enough apart on the chromosome that they assort independently.
 Name Section 7.014 Problem Set 5 Please print out this problem set and record your answers on the printed copy. Answers to this problem set are to be turned in to the box outside 68-120 by 5:00pm on Friday
Name Section 7.014 Problem Set 5 Please print out this problem set and record your answers on the printed copy. Answers to this problem set are to be turned in to the box outside 68-120 by 5:00pm on Friday
1 Mutation and Genetic Change
 CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
Transcription in prokaryotes. Elongation and termination
 Transcription in prokaryotes Elongation and termination After initiation the σ factor leaves the scene. Core polymerase is conducting the elongation of the chain. The core polymerase contains main nucleotide
Transcription in prokaryotes Elongation and termination After initiation the σ factor leaves the scene. Core polymerase is conducting the elongation of the chain. The core polymerase contains main nucleotide
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION. Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.
 Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
EU Reference Laboratory for E. coli Department of Veterinary Public Health and Food Safety Unit of Foodborne Zoonoses Istituto Superiore di Sanità
 Identification and characterization of Verocytotoxin-producing Escherichia coli (VTEC) by Real Time PCR amplification of the main virulence genes and the genes associated with the serogroups mainly associated
Identification and characterization of Verocytotoxin-producing Escherichia coli (VTEC) by Real Time PCR amplification of the main virulence genes and the genes associated with the serogroups mainly associated
KMS-Specialist & Customized Biosimilar Service
 KMS-Specialist & Customized Biosimilar Service 1. Polyclonal Antibody Development Service KMS offering a variety of Polyclonal Antibody Services to fit your research and production needs. we develop polyclonal
KMS-Specialist & Customized Biosimilar Service 1. Polyclonal Antibody Development Service KMS offering a variety of Polyclonal Antibody Services to fit your research and production needs. we develop polyclonal
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation
 OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
T Cell Maturation,Activation and Differentiation
 T Cell Maturation,Activation and Differentiation Positive Selection- In thymus, permits survival of only those T cells whose TCRs recognize self- MHC molecules (self-mhc restriction) Negative Selection-
T Cell Maturation,Activation and Differentiation Positive Selection- In thymus, permits survival of only those T cells whose TCRs recognize self- MHC molecules (self-mhc restriction) Negative Selection-
Appendix 2 Molecular Biology Core Curriculum. Websites and Other Resources
 Appendix 2 Molecular Biology Core Curriculum Websites and Other Resources Chapter 1 - The Molecular Basis of Cancer 1. Inside Cancer http://www.insidecancer.org/ From the Dolan DNA Learning Center Cold
Appendix 2 Molecular Biology Core Curriculum Websites and Other Resources Chapter 1 - The Molecular Basis of Cancer 1. Inside Cancer http://www.insidecancer.org/ From the Dolan DNA Learning Center Cold
DNA Fingerprinting. Unless they are identical twins, individuals have unique DNA
 DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
Genetic Instability of the Global Regulator agr Explains the Phenotype of the xpr Mutation in Staphylococcus aureus KSI9051
 JOURNAL OF BACTERIOLOGY, May 1998, p. 2609 2615 Vol. 180, No. 10 0021-9193/98/$04.00 0 Copyright 1998, American Society for Microbiology Genetic Instability of the Global Regulator agr Explains the Phenotype
JOURNAL OF BACTERIOLOGY, May 1998, p. 2609 2615 Vol. 180, No. 10 0021-9193/98/$04.00 0 Copyright 1998, American Society for Microbiology Genetic Instability of the Global Regulator agr Explains the Phenotype
Studies on regulatory networks governing virulence gene transcription in Staphylococcus aureus
 From Department of Microbiology, Tumor and Cell Biology (MTC) Karolinska Institutet, Stockholm, Sweden Studies on regulatory networks governing virulence gene transcription in Staphylococcus aureus Erik
From Department of Microbiology, Tumor and Cell Biology (MTC) Karolinska Institutet, Stockholm, Sweden Studies on regulatory networks governing virulence gene transcription in Staphylococcus aureus Erik
Gene Regulation -- The Lac Operon
 Gene Regulation -- The Lac Operon Specific proteins are present in different tissues and some appear only at certain times during development. All cells of a higher organism have the full set of genes:
Gene Regulation -- The Lac Operon Specific proteins are present in different tissues and some appear only at certain times during development. All cells of a higher organism have the full set of genes:
QUANTITATIVE RT-PCR. A = B (1+e) n. A=amplified products, B=input templates, n=cycle number, and e=amplification efficiency.
 QUANTITATIVE RT-PCR Application: Quantitative RT-PCR is used to quantify mrna in both relative and absolute terms. It can be applied for the quantification of mrna expressed from endogenous genes, and
QUANTITATIVE RT-PCR Application: Quantitative RT-PCR is used to quantify mrna in both relative and absolute terms. It can be applied for the quantification of mrna expressed from endogenous genes, and
Green Fluorescent Protein (GFP): Genetic Transformation, Synthesis and Purification of the Recombinant Protein
 Green Fluorescent Protein (GFP): Genetic Transformation, Synthesis and Purification of the Recombinant Protein INTRODUCTION Green Fluorescent Protein (GFP) is a novel protein produced by the bioluminescent
Green Fluorescent Protein (GFP): Genetic Transformation, Synthesis and Purification of the Recombinant Protein INTRODUCTION Green Fluorescent Protein (GFP) is a novel protein produced by the bioluminescent
Actions of Hormones on Target Cells Page 1. Actions of Hormones on Target Cells Page 2. Goals/ What You Need to Know Goals What You Need to Know
 Actions of Hormones on Target Cells Graphics are used with permission of: Pearson Education Inc., publishing as Benjamin Cummings (http://www.aw-bc.com) Page 1. Actions of Hormones on Target Cells Hormones
Actions of Hormones on Target Cells Graphics are used with permission of: Pearson Education Inc., publishing as Benjamin Cummings (http://www.aw-bc.com) Page 1. Actions of Hormones on Target Cells Hormones
Microarray Technology
 Microarrays And Functional Genomics CPSC265 Matt Hudson Microarray Technology Relatively young technology Usually used like a Northern blot can determine the amount of mrna for a particular gene Except
Microarrays And Functional Genomics CPSC265 Matt Hudson Microarray Technology Relatively young technology Usually used like a Northern blot can determine the amount of mrna for a particular gene Except
Transmission of genetic variation: conjugation. Transmission of genetic variation: conjugation
 Transmission of genetic variation: conjugation Transmission of genetic variation: conjugation Bacterial Conjugation is genetic recombination in which there is a transfer of DNA from a living donor bacterium
Transmission of genetic variation: conjugation Transmission of genetic variation: conjugation Bacterial Conjugation is genetic recombination in which there is a transfer of DNA from a living donor bacterium
Antibody Function & Structure
 Antibody Function & Structure Specifically bind to antigens in both the recognition phase (cellular receptors) and during the effector phase (synthesis and secretion) of humoral immunity Serology: the
Antibody Function & Structure Specifically bind to antigens in both the recognition phase (cellular receptors) and during the effector phase (synthesis and secretion) of humoral immunity Serology: the
Making the switch to a safer CAR-T cell therapy
 Making the switch to a safer CAR-T cell therapy HaemaLogiX 2015 Technical Journal Club May 24 th 2016 Christina Müller - chimeric antigen receptor = CAR - CAR T cells are generated by lentiviral transduction
Making the switch to a safer CAR-T cell therapy HaemaLogiX 2015 Technical Journal Club May 24 th 2016 Christina Müller - chimeric antigen receptor = CAR - CAR T cells are generated by lentiviral transduction
Complex multicellular organisms are produced by cells that switch genes on and off during development.
 Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Trasposable elements: P elements
 Trasposable elements: P elements In 1938 Marcus Rhodes provided the first genetic description of an unstable mutation, an allele of a gene required for the production of pigment in maize. This instability
Trasposable elements: P elements In 1938 Marcus Rhodes provided the first genetic description of an unstable mutation, an allele of a gene required for the production of pigment in maize. This instability
Medical Microbiology Culture Media :
 Lecture 3 Dr. Ismail I. Daood Medical Microbiology Culture Media : Culture media are used for recognition and identification (diagnosis) of microorganisms. The media are contained in plates (Petri dishes),
Lecture 3 Dr. Ismail I. Daood Medical Microbiology Culture Media : Culture media are used for recognition and identification (diagnosis) of microorganisms. The media are contained in plates (Petri dishes),
Structure and Function of DNA
 Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Innate Immunity. Insects rely solely on an innate immune system for defense against infection
 Innate Immunity Insects rely solely on an innate immune system for defense against infection The importance of mammalian innate immunity has only recently become appreciated The innate immune response
Innate Immunity Insects rely solely on an innate immune system for defense against infection The importance of mammalian innate immunity has only recently become appreciated The innate immune response
Recombinant DNA Unit Exam
 Recombinant DNA Unit Exam Question 1 Restriction enzymes are extensively used in molecular biology. Below are the recognition sites of two of these enzymes, BamHI and BclI. a) BamHI, cleaves after the
Recombinant DNA Unit Exam Question 1 Restriction enzymes are extensively used in molecular biology. Below are the recognition sites of two of these enzymes, BamHI and BclI. a) BamHI, cleaves after the
Control of Gene Expression
 Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Understanding the immune response to bacterial infections
 Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
4. DNA replication Pages: 979-984 Difficulty: 2 Ans: C Which one of the following statements about enzymes that interact with DNA is true?
 Chapter 25 DNA Metabolism Multiple Choice Questions 1. DNA replication Page: 977 Difficulty: 2 Ans: C The Meselson-Stahl experiment established that: A) DNA polymerase has a crucial role in DNA synthesis.
Chapter 25 DNA Metabolism Multiple Choice Questions 1. DNA replication Page: 977 Difficulty: 2 Ans: C The Meselson-Stahl experiment established that: A) DNA polymerase has a crucial role in DNA synthesis.
Design of conditional gene targeting vectors - a recombineering approach
 Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
LAB 16 Rapid Colony Transformation of E. coli with Plasmid DNA
 LAB 16 Rapid Colony Transformation of E. coli with Plasmid DNA Objective: In this laboratory investigation, plasmids containing fragments of foreign DNA will be used to transform Escherichia coli cells,
LAB 16 Rapid Colony Transformation of E. coli with Plasmid DNA Objective: In this laboratory investigation, plasmids containing fragments of foreign DNA will be used to transform Escherichia coli cells,
Essentials of Real Time PCR. About Sequence Detection Chemistries
 Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Becker Muscular Dystrophy
 Muscular Dystrophy A Case Study of Positional Cloning Described by Benjamin Duchenne (1868) X-linked recessive disease causing severe muscular degeneration. 100 % penetrance X d Y affected male Frequency
Muscular Dystrophy A Case Study of Positional Cloning Described by Benjamin Duchenne (1868) X-linked recessive disease causing severe muscular degeneration. 100 % penetrance X d Y affected male Frequency
Supplemental Information. McBrayer et al. Supplemental Data
 1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
Induction of Enzyme Activity in Bacteria:The Lac Operon. Preparation for Laboratory: Web Tutorial - Lac Operon - submit questions
 Induction of Enzyme Activity in Bacteria:The Lac Operon Preparation for Laboratory: Web Tutorial - Lac Operon - submit questions I. Background: For the last week you explored the functioning of the enzyme
Induction of Enzyme Activity in Bacteria:The Lac Operon Preparation for Laboratory: Web Tutorial - Lac Operon - submit questions I. Background: For the last week you explored the functioning of the enzyme
Troubleshooting the Single-step PCR Site-directed Mutagenesis Procedure Intended to Create a Non-functional rop Gene in the pbr322 Plasmid
 Troubleshooting the Single-step PCR Site-directed Mutagenesis Procedure Intended to Create a Non-functional rop Gene in the pbr322 Plasmid Lina Jew Department of Microbiology & Immunology, University of
Troubleshooting the Single-step PCR Site-directed Mutagenesis Procedure Intended to Create a Non-functional rop Gene in the pbr322 Plasmid Lina Jew Department of Microbiology & Immunology, University of
Using chromosomal laci Q1 to control. high copy number plasmids in Escherichia coli. Weickert; Gene 223; 1998 : 221 231
 Using chromosomal laci Q1 to control expression of genes on high copy number plasmids in Escherichia coli Christopher B Glascock Michael J Christopher B. Glascock, Michael J. Weickert; Gene 223; 1998 :
Using chromosomal laci Q1 to control expression of genes on high copy number plasmids in Escherichia coli Christopher B Glascock Michael J Christopher B. Glascock, Michael J. Weickert; Gene 223; 1998 :
Recombinant DNA & Genetic Engineering. Tools for Genetic Manipulation
 Recombinant DNA & Genetic Engineering g Genetic Manipulation: Tools Kathleen Hill Associate Professor Department of Biology The University of Western Ontario Tools for Genetic Manipulation DNA, RNA, cdna
Recombinant DNA & Genetic Engineering g Genetic Manipulation: Tools Kathleen Hill Associate Professor Department of Biology The University of Western Ontario Tools for Genetic Manipulation DNA, RNA, cdna
Molecular Biology. Yeast Transformation. Yeast Plasmids. Gene Disruption, tagging. Cloning by Complementation. Epistasis
 Molecular Biology Yeast Transformation Yeast Plasmids Gene Disruption, tagging Cloning by Complementation Epistasis Transformation Transformation: introduction of DNA 1978, ca 1000x less efficient than
Molecular Biology Yeast Transformation Yeast Plasmids Gene Disruption, tagging Cloning by Complementation Epistasis Transformation Transformation: introduction of DNA 1978, ca 1000x less efficient than
Lecture 3: Mutations
 Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
Gene Switches Teacher Information
 STO-143 Gene Switches Teacher Information Summary Kit contains How do bacteria turn on and turn off genes? Students model the action of the lac operon that regulates the expression of genes essential for
STO-143 Gene Switches Teacher Information Summary Kit contains How do bacteria turn on and turn off genes? Students model the action of the lac operon that regulates the expression of genes essential for
2006 7.012 Problem Set 6 KEY
 2006 7.012 Problem Set 6 KEY ** Due before 5 PM on WEDNESDAY, November 22, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You create an artificial
2006 7.012 Problem Set 6 KEY ** Due before 5 PM on WEDNESDAY, November 22, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You create an artificial
Chapter 43: The Immune System
 Name Period Our students consider this chapter to be a particularly challenging and important one. Expect to work your way slowly through the first three concepts. Take particular care with Concepts 43.2
Name Period Our students consider this chapter to be a particularly challenging and important one. Expect to work your way slowly through the first three concepts. Take particular care with Concepts 43.2
The role of IBV proteins in protection: cellular immune responses. COST meeting WG2 + WG3 Budapest, Hungary, 2015
 The role of IBV proteins in protection: cellular immune responses COST meeting WG2 + WG3 Budapest, Hungary, 2015 1 Presentation include: Laboratory results Literature summary Role of T cells in response
The role of IBV proteins in protection: cellular immune responses COST meeting WG2 + WG3 Budapest, Hungary, 2015 1 Presentation include: Laboratory results Literature summary Role of T cells in response
Gene Mapping Techniques
 Gene Mapping Techniques OBJECTIVES By the end of this session the student should be able to: Define genetic linkage and recombinant frequency State how genetic distance may be estimated State how restriction
Gene Mapping Techniques OBJECTIVES By the end of this session the student should be able to: Define genetic linkage and recombinant frequency State how genetic distance may be estimated State how restriction
GeneCopoeia Genome Editing Tools for Safe Harbor Integration in. Mice and Humans. Ed Davis, Liuqing Qian, Ruiqing li, Junsheng Zhou, and Jinkuo Zhang
 G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
GENE REGULATION. Teacher Packet
 AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE Q5B
 INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
Chapter 5: Organization and Expression of Immunoglobulin Genes
 Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Custom Antibody Services
 prosci-inc.com Custom Antibody Services High Performance Antibodies and More Broad Antibody Catalog Extensive Antibody Services CUSTOM ANTIBODY SERVICES Established in 1998, ProSci Incorporated is a leading
prosci-inc.com Custom Antibody Services High Performance Antibodies and More Broad Antibody Catalog Extensive Antibody Services CUSTOM ANTIBODY SERVICES Established in 1998, ProSci Incorporated is a leading
MAB Solut. MABSolys Génopole Campus 1 5 rue Henri Desbruères 91030 Evry Cedex. www.mabsolut.com. is involved at each stage of your project
 Mabsolus-2015-UK:Mise en page 1 03/07/15 14:13 Page1 Services provider Department of MABSolys from conception to validation MAB Solut is involved at each stage of your project Creation of antibodies Production
Mabsolus-2015-UK:Mise en page 1 03/07/15 14:13 Page1 Services provider Department of MABSolys from conception to validation MAB Solut is involved at each stage of your project Creation of antibodies Production
Question 4 /29 points. Total /100 points
 MIT Department of Biology 7.28, Spring 2005 - Molecular Biology 7.28 Spring 2005 Exam Three Question 1 Question 2 Question 3 /30 points /20 points /21 points Question 4 /29 points Total /100 points 1 Question
MIT Department of Biology 7.28, Spring 2005 - Molecular Biology 7.28 Spring 2005 Exam Three Question 1 Question 2 Question 3 /30 points /20 points /21 points Question 4 /29 points Total /100 points 1 Question
Gene mutation and molecular medicine Chapter 15
 Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
Lecture 13: DNA Technology. DNA Sequencing. DNA Sequencing Genetic Markers - RFLPs polymerase chain reaction (PCR) products of biotechnology
 Lecture 13: DNA Technology DNA Sequencing Genetic Markers - RFLPs polymerase chain reaction (PCR) products of biotechnology DNA Sequencing determine order of nucleotides in a strand of DNA > bases = A,
Lecture 13: DNA Technology DNA Sequencing Genetic Markers - RFLPs polymerase chain reaction (PCR) products of biotechnology DNA Sequencing determine order of nucleotides in a strand of DNA > bases = A,
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
Bacterial Transformation with Green Fluorescent Protein. Table of Contents Fall 2012
 Bacterial Transformation with Green Fluorescent Protein pglo Version Table of Contents Bacterial Transformation Introduction..1 Laboratory Exercise...3 Important Laboratory Practices 3 Protocol...... 4
Bacterial Transformation with Green Fluorescent Protein pglo Version Table of Contents Bacterial Transformation Introduction..1 Laboratory Exercise...3 Important Laboratory Practices 3 Protocol...... 4
Lab 10: Bacterial Transformation, part 2, DNA plasmid preps, Determining DNA Concentration and Purity
 Lab 10: Bacterial Transformation, part 2, DNA plasmid preps, Determining DNA Concentration and Purity Today you analyze the results of your bacterial transformation from last week and determine the efficiency
Lab 10: Bacterial Transformation, part 2, DNA plasmid preps, Determining DNA Concentration and Purity Today you analyze the results of your bacterial transformation from last week and determine the efficiency
Modeling and Simulation of Gene Regulatory Networks
 Modeling and Simulation of Gene Regulatory Networks Hidde de Jong INRIA Grenoble - Rhône-Alpes Hidde.de-Jong@inria.fr http://ibis.inrialpes.fr INRIA Grenoble - Rhône-Alpes and IBIS IBIS: systems biology
Modeling and Simulation of Gene Regulatory Networks Hidde de Jong INRIA Grenoble - Rhône-Alpes Hidde.de-Jong@inria.fr http://ibis.inrialpes.fr INRIA Grenoble - Rhône-Alpes and IBIS IBIS: systems biology
Genetics 301 Sample Final Examination Spring 2003
 Genetics 301 Sample Final Examination Spring 2003 50 Multiple Choice Questions-(Choose the best answer) 1. A cross between two true breeding lines one with dark blue flowers and one with bright white flowers
Genetics 301 Sample Final Examination Spring 2003 50 Multiple Choice Questions-(Choose the best answer) 1. A cross between two true breeding lines one with dark blue flowers and one with bright white flowers
restriction enzymes 350 Home R. Ward: Spring 2001
 restriction enzymes 350 Home Restriction Enzymes (endonucleases): molecular scissors that cut DNA Properties of widely used Type II restriction enzymes: recognize a single sequence of bases in dsdna, usually
restriction enzymes 350 Home Restriction Enzymes (endonucleases): molecular scissors that cut DNA Properties of widely used Type II restriction enzymes: recognize a single sequence of bases in dsdna, usually
The Effects of Glycerol, Glucose, Galactose, Lactose and Glucose with Galactose on the Induction of β-galactosidase in Escherichia coli
 The Effects of Glycerol, Glucose, Galactose, Lactose and Glucose with Galactose on the Induction of β-galactosidase in Escherichia coli VICKY CHAN, LISA F. DREOLINI, KERRY A. FLINTOFF, SONJA J. LLOYD,
The Effects of Glycerol, Glucose, Galactose, Lactose and Glucose with Galactose on the Induction of β-galactosidase in Escherichia coli VICKY CHAN, LISA F. DREOLINI, KERRY A. FLINTOFF, SONJA J. LLOYD,
Cellular Implants as mini-factories for the production of therapeutic antibodies against Altzheimer s disease. rapport research week Jonas Schiffmann
 Cellular Implants as mini-factories for the production of therapeutic antibodies against Altzheimer s disease rapport research week Jonas Schiffmann Supported by Aurélien Lathuilière, Bernard Schneider
Cellular Implants as mini-factories for the production of therapeutic antibodies against Altzheimer s disease rapport research week Jonas Schiffmann Supported by Aurélien Lathuilière, Bernard Schneider
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How to construct transgenic mice
 How to construct transgenic mice Sandra Beer-Hammer Autumn School 2010 Bad Schandau Methods additional genetic information transgenic mouse line gene inactivation gene-deficient knockout mouse line Jak2
How to construct transgenic mice Sandra Beer-Hammer Autumn School 2010 Bad Schandau Methods additional genetic information transgenic mouse line gene inactivation gene-deficient knockout mouse line Jak2
Diabetes and Drug Development
 Diabetes and Drug Development Metabolic Disfunction Leads to Multiple Diseases Hypertension ( blood pressure) Metabolic Syndrome (Syndrome X) LDL HDL Lipoproteins Triglycerides FFA Hyperinsulinemia Insulin
Diabetes and Drug Development Metabolic Disfunction Leads to Multiple Diseases Hypertension ( blood pressure) Metabolic Syndrome (Syndrome X) LDL HDL Lipoproteins Triglycerides FFA Hyperinsulinemia Insulin
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
FACULTY OF MEDICAL SCIENCE
 Doctor of Philosophy Program in Microbiology FACULTY OF MEDICAL SCIENCE Naresuan University 171 Doctor of Philosophy Program in Microbiology The time is critical now for graduate education and research
Doctor of Philosophy Program in Microbiology FACULTY OF MEDICAL SCIENCE Naresuan University 171 Doctor of Philosophy Program in Microbiology The time is critical now for graduate education and research
DNA Sequence Analysis
 DNA Sequence Analysis Two general kinds of analysis Screen for one of a set of known sequences Determine the sequence even if it is novel Screening for a known sequence usually involves an oligonucleotide
DNA Sequence Analysis Two general kinds of analysis Screen for one of a set of known sequences Determine the sequence even if it is novel Screening for a known sequence usually involves an oligonucleotide
Rapid GST Inclusion Body Solubilization and Renaturation Kit
 Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
June 09, 2009 Random Mutagenesis
 Why Mutagenesis? Analysis of protein function June 09, 2009 Random Mutagenesis Analysis of protein structure Protein engineering Analysis of structure-function relationship Analysis of the catalytic center
Why Mutagenesis? Analysis of protein function June 09, 2009 Random Mutagenesis Analysis of protein structure Protein engineering Analysis of structure-function relationship Analysis of the catalytic center
mirnaselect pep-mir Cloning and Expression Vector
 Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
How Does a Doctor Test for AIDS?
 Edvo-Kit #S-70 How Does a Doctor Test for AIDS? S-70 Experiment Objective: The Human Immunodefi ciency Virus (HIV) is an infectious agent that causes Acquired Immunodefi ciency Syndrome (AIDS) in humans.
Edvo-Kit #S-70 How Does a Doctor Test for AIDS? S-70 Experiment Objective: The Human Immunodefi ciency Virus (HIV) is an infectious agent that causes Acquired Immunodefi ciency Syndrome (AIDS) in humans.
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
RNA Viruses. A Practical Approac h. Alan J. Cann
 RNA Viruses A Practical Approac h Alan J. Cann List of protocols page xiii Abbreviations xvii Investigation of RNA virus genome structure 1 A j. Easton, A.C. Marriott and C.R. Pringl e 1 Introduction-the
RNA Viruses A Practical Approac h Alan J. Cann List of protocols page xiii Abbreviations xvii Investigation of RNA virus genome structure 1 A j. Easton, A.C. Marriott and C.R. Pringl e 1 Introduction-the
pcas-guide System Validation in Genome Editing
 pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
GENE CLONING AND RECOMBINANT DNA TECHNOLOGY
 GENE CLONING AND RECOMBINANT DNA TECHNOLOGY What is recombinant DNA? DNA from 2 different sources (often from 2 different species) are combined together in vitro. Recombinant DNA forms the basis of cloning.
GENE CLONING AND RECOMBINANT DNA TECHNOLOGY What is recombinant DNA? DNA from 2 different sources (often from 2 different species) are combined together in vitro. Recombinant DNA forms the basis of cloning.
MICROBIOLOGY LETTERS. Introduction RESEARCH LETTER. Ellen H. James, Andrew M. Edwards & Sivaramesh Wigneshweraraj. Abstract
 RESEARCH LETTER Transcriptional downregulation of agr expression in Staphylococcus aureus during growth in human serum can be overcome by constitutively active mutant forms of the sensor kinase AgrC Ellen
RESEARCH LETTER Transcriptional downregulation of agr expression in Staphylococcus aureus during growth in human serum can be overcome by constitutively active mutant forms of the sensor kinase AgrC Ellen
AMES TEST: Bacterial Reverse Mutation Assay
 AMES TEST: Bacterial Reverse Mutation Assay 1. Introduction The bacteria reversed mutation assay (Ames Test) is used to evaluate the mutagenic properties of test articles. The test uses amino acid-dependent
AMES TEST: Bacterial Reverse Mutation Assay 1. Introduction The bacteria reversed mutation assay (Ames Test) is used to evaluate the mutagenic properties of test articles. The test uses amino acid-dependent
