letters to nature ... Processing of primary micrornas by the Microprocessor complex
|
|
|
- Sophia Phelps
- 8 years ago
- Views:
Transcription
1 ... Processing of primary micrornas by the Microprocessor complex Ahmet M. Denli 1, Bastiaan B. J. Tops 2, Ronald H. A. Plasterk 2, René F. Ketting 2 & Gregory J. Hannon 1 letters to nature 1 Cold Spring Harbor Laboratory, Watson School of Biological Sciences, 1 Bungtown Road, Cold Spring Harbor, New York 11724, USA 2 The Hubrecht Laboratory Centre for Biomedical Genetics, Uppsalalaan 8, 3584 CT Utrecht, The Netherlands... Mature micrornas (mirnas) are generated via a two-step processing pathway to yield,22-nucleotide small RNAs that regulate gene expression at the post-transcriptional level 1. Initial cleavage is catalysed by Drosha, a nuclease of the RNase III family, which acts on primary mirna transcripts (pri-mirnas) in the nucleus 2. Here we show that Drosha exists in a multiprotein complex, the Microprocessor, and begin the process of deconstructing that complex into its constituent components. Along with Drosha, the Microprocessor also contains Pasha (partner of Drosha), a double-stranded RNA binding protein. Suppression of Pasha expression in Drosophila cells or Caenorhabditis elegans interferes with pri-mirna processing, leading to an accumulation of pri-mirnas and a reduction in mature mirnas. Finally, depletion or mutation of pash-1 in C. elegans causes de-repression of a let-7 reporter and the appearance of phenotypic defects overlapping those observed upon examination of worms with lesions in Dicer (dcr-1) or Drosha (drsh-1). Considered together, these results indicate a role for Pasha in mirna maturation and mirna-mediated gene regulation. mirnas are a class of small, non-coding RNAs that enter the RNA interference (RNAi) pathway to regulate the expression of protein-encoding genes at the post-transcriptional level 3. mirna production begins with the synthesis of pri-mirnas, ranging in size from several hundred nucleotides (nt) to several kilobases. These are recognized and cleaved into precursor mirnas (pre-mirnas) in the nucleus by an RNase III family nuclease, Drosha 2 ; the premirnas are short, hairpin RNAs of approximately 70 nt, bearing the 2-nucleotide 3 0 overhang that is a signature of RNase III-mediated cleavage. Pre-miRNAs are exported to the cytoplasm by a RanGTP/ exportin 5-dependent mechanism 4 6, with their characteristic overhang contributing to their entry into this export pathway 5.Oncein the cytoplasm, pre-mirnas are recognized and processed into their mature,,22-nt form by Dicer 7 9, with the 3 0 overhang again playing a role in specifying cleavage Mature mirnas enter RISC (RNAinduced silencing complex) in an asymmetric fashion such that for most mirnas, only one strand is enabled to recognize and repress the expression of target genes 13. The outcome of this recognition is either endonucleolytic cleavage of the targeted messenger RNA 14,15 or interference with protein synthesis by a mechanism that remains unclear With the ultimate goal of addressing the mechanisms by which pri-mirnas are tagged for entry into the RNAi pathway and processed at specific sites, we have undertaken a biochemical characterization of Drosha and its associated factors. Previously, Kim and colleagues showed 2 that epitope-tagged, human Drosha protein was capable of releasing pre-mirnas from pri-mirnas in vitro and contributed to mirna maturation in vivo. Processing was dependent upon the presence of a doublestranded region around the cleavage position. We asked whether a pri-mirna processing activity also existed in Drosophila cells. Although they are absent from Schizosaccharomyces pombe and Arabidopsis, homologues of mammalian Drosha are present in Drosophila and C. elegans 19. Indeed, S2 cell extracts contain an activity that can recognize a pri-mirna, in this case pri-bantam, and cleave this into discrete products that can be identified as prebantam and the regions of the pri-mirna that flank that mature sequence; pre-bantam and the 5 0 and 3 0 flanks are 60, 104 and 112 nt, respectively (Fig. 1a; see Supplementary Fig. S1 for sequences). Immunoprecipitates obtained using an affinity-purified Drosha antibody were capable of generating pre-bantam from pribantam (Fig. 1b). Examination of the supernatants showed that a substantial fraction of pri-mirna processing activity was depleted from the extract by immunoprecipitation (data not shown). Additional pri-mirnas from both human and Drosophila were similarly processed by extracts and immunoprecipitates (data not shown). Considered together, these results implicate Drosha and/or Figure 1 Pri-miRNA processing is conserved in Drosophila. a, Pri-miRNA processing activity was assessed in Drosophila S2 cell extracts using uniformly labelled pri-bantam mirna. Positions of the processed pre-bantam and the two flanking RNAs derived from excision of pre-bantam are indicated. b, Immunoprecipitation experiments with S2 extracts test whether processing activity associates with Drosha. c, To assess the accuracy of pri-mirna processing, cleavage sites were mapped using end-labelled substrates. Pre-bantam or pre-mir30a was 3 0 labelled by ligation to 32 P-pCp, and pri-mir30a was 5 0 labelled by capping with 32 P-GTP. Sizes of cleavage products were as predicted (see Supplementary Fig. S1). RNA size markers, indicated in bases, are shown at the right. d, The 5 0 end of processed pre-mir30a was mapped by primer extension. Control extension of a synthetic pre-mir30a RNA is shown for reference along with a 23 bp 5 0 -end-labelled DNA marker. e, Western blots with anti-drosha antibody across fractions of a Superose 6 column show that Drosha from S2 cells exists in a,500 kda complex. Fraction numbers are given below the panel. NATURE VOL NOVEMBER
2 Figure 2 Pasha, a nuclear dsrna binding domain (dsrbd) protein, associates with Drosha. a, Immunoprecipitates from S2 cell extracts with antibodies directed against endogenous Drosha or Pasha proteins were examined by western blotting with anti- Drosha antibody. To assess the degree of co-association, supernatants were also probed. The immunoprecipitates were derived from extract equivalent to,40 of what was loaded in the supernatant lanes. b, Immunoprecipitations were prepared from S2 cells that were transfected with either a GFP expression construct or a construct expressing T7 epitope-tagged Pasha. c, Immunoprecipitates were incubated with uniformly labelled pribantam and tested for processing activity. its associated factors as the major source of pri-mirna processing activity in Drosophila extracts. The accuracy of pri-mirna processing by our immunoaffinitypurified Drosha was assessed in two ways. First, we examined the processing of two pri-mirna substrates that were labelled at their 5 0 or 3 0 termini (Fig. 1c). Cleavage of pri-bantam or pri-mir30a yielded the expected end fragments. The end of pre-mir30a has previously been mapped by primer extension of in vivo and in vitro processed transcripts 1. Upon analysis of pri-mir30a, processed using immunoaffinity-purified Drosophila Drosha, we find two prominent extension products. The smaller (denoted with an asterisk in Fig. 1d) is consistent with the expected pre-mir30a product, based both on a labelled DNA marker and on primer extension of a synthetic version of the expected pre-mir30a product. A larger product (denoted by X in Fig. 1d), differing from the expected product by 2 3 nt, is of unknown origin. However, it should be noted that many nucleases, including Dicer, are known to make multiple cleavages within their substrates. In contrast, Drosha does not operate on fully duplexed substrates 20 (Supplementary Fig. S2). To investigate how Drosha selects its substrates and how the enzyme determines its specific cleavage sites, we sought to characterize native complexes containing the enzyme. Biochemical fractionation indicates that Drosha is present within an,500 kda complex, which presumably contains additional protein components (Fig. 1e). Given its role in microrna metabolism, we have dubbed this complex the Microprocessor. Recent, genome-wide two-hybrid analysis of Drosophila has generated a substantial list of candidate protein protein interactions 21. In examining the list, we noted a potential interaction between Drosha and a double-stranded RNA (dsrna) binding protein, originally dubbed CG1800 (candidate gene 1800). This protein consists of two domains, a WW domain near the amino terminus and a dsrna binding domain near the carboxyl terminus. Notably, WW domains interact with proline-rich domains, such as the one present near the N terminus of Drosha. Along with Drosha, CG1800 is conserved in C. elegans and mammals, where it was called DGCR8 22. Like Drosha, CG1800 homologues are absent from the Arabidopsis and S. pombe genomes. Results from the published two-hybrid studies, as well as evidence of physical and genetic interaction presented below, caused us to designate CG1800 as Pasha (partner of Drosha). Consistent with the possibility that they might both be present in a single complex, Drosha and Pasha are predominantly nuclear Figure 3 Drosha and Pasha co-exist in a Microprocessor complex. a, Combined nuclear and cytoplasmic extracts from S2 cells were loaded onto a Superose 6 column, and fractions were subjected to western blotting with Drosha or Pasha antibodies (lower panel). The Drosha antibody was used to recover Drosha complexes from each fraction and these were tested for their ability to process pri-bantam (upper panel). Positions of pre-bantam and the 5 0 and 3 0 flanks are indicated. b, Extracts prepared as in a were run on a Q sepharose FF column. Drosha and Pasha fractionation profiles were determined by western blotting (lower panels). Pri-miRNA processing activity on pri-bantam was examined using Drosha immunoprecipitates from each fraction. Direct analysis of processing activity in each fraction showed a similar processing activity profile, albeit with weaker activity than exhibited by the immunoprecipitates. c, Drosha peak fractions from a Q-sepharose column were pooled and loaded onto a phenyl HP column. Fractionation of Drosha and Pasha and processing activity were followed as in b. 232 NATURE VOL NOVEMBER
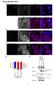
3 proteins (Supplementary Fig. S3; data not shown). Furthermore, Pasha antiserum co-immunoprecipitates Drosha, essentially depleting it from whole cell extracts (Fig. 2a). This interaction was not disrupted by treatment of extracts or immunoprecipitates with RNase (data not shown). Analysis of the reciprocal immunoprecipitate required the use of a T7-tagged Pasha, since the antibody heavy chain interferes with detection of this protein in western blots with the rabbit anti-pasha antibody. In T7-Pasha-expressing cells, a Drosha antibody co-immunoprecipitates tagged Pasha (Fig. 2b). Drosha Pasha complexes are functional since both Drosha and Pasha immunoprecipitates are able to process pri-mirna into premirna in vitro with similar efficiencies (Fig. 2c). To test whether Drosha and Pasha coexist in the 500 kda Microprocessor, we examined chromatographic profiles from S2 cell extracts. Both Figure 4 Depletion of Pasha reduces pri-mirna processing. A, Pri-miR2a levels were analysed following transfection of Drosophila S2 cells with two different dsrnas homologous to Pasha by semiquantitative RT PCR. Error bar indicates standard deviation from the mean. B, Total RNA extracted from S2 cells transfected with either a control dsrna (luc) or either of two Pasha dsrnas was examined for the level of mature mir2a by northern blotting. U6 snrna was used as a control to ensure equal loading. C, C. elegans (rrf-3 mutant) were treated with Pasha (pash-1), Dicer (dcr-1) or control dsrnas (L4440), and total RNA was prepared. Levels of pre- and mature let-7 were assessed by northern blotting. Sizes of selected RNA markers are indicated in bases at the left. D, Phenotypic analysis of Pasha (pash-1) and Drosha (drsh-1) inc. elegans was performed using RNAi and genetic mutants. RNAi against pash-1 leads to vulva (100%) and alae (10/17) defects, similar to those published for dcr-1 mutants 8,9 (a and b, d and e, g and h,andj and k). The observed defects include protrusion and bursting of the vulva, and gaps in, or absence of the alae (asterisks). Occasionally duplicated alae are detected, as shown in e. The drsh-1(tm0654) allele does not result in obvious defects in these structures (alae indicated by arrow heads); however this mutant may be partially rescued by maternal contribution of Drosha. i and l, both drsh-1(tm0654) (10 out of 10) and pash-1(pk2083) (14 out of 14) result in defects in the let-7-mediated silencing of a previously described transgene (pkis2084) 24.Thisdefectis similar to that observed in a dcr-1(s2795) mutant(f: 15 out of 15). In wild-type adults, the lacz expression is silenced (c: 0outof57showinglacZ expression). NATURE VOL NOVEMBER
4 proteins along with pri-mirna processing activity peaked in an,500 kda fraction (Fig. 3a). Similar co-fractionation was also seen if complexes were serially chromatographed on Q sepharose FF and phenyl HP columns (Fig. 3b, c) or on Q sepharose FF and mono S (data not shown). To examine the functional connection between Pasha and mirna metabolism, we asked whether suppression or mutation of Pasha had any affect on pri-mirna processing or mirna function. In Drosophila S2 cells, Pasha dsrnas caused a modest suppression of Pasha mrna (Supplemental Fig. S4). This resulted in both an accumulation of pri-mir2a and a reduction in levels of mature mir2a (Fig. 4a, b). Similar effects on pri-mir30a were seen upon transfection of mammalian cells with two different human Pasha sirnas (Supplementary Fig. S5). Similarly, treatment of C. elegans with pash- 1 dsrna, by feeding of RNAi-hypersensitive rrf-3 mutant worms, resulted in an accumulation of pri-let-7 with a decrease in the mature species (Fig. 4c; Supplementary Fig. S6). Notably, similar effects were seen upon suppression of Dicer (dcr-1), although in this case the pre-mirna also accumulated. A comparison of these two results suggests that Pasha acts in pri-mirna metabolism upstream of premirna production, consistent with its placement by biochemistry as a component of the Microprocessor. Precisely why pri-mirnas also accumulate in worms treated with Dicer dsrna but not in worms treated with control dsrnas is at present unclear. For studies of the biological function of Drosha and Pasha in vivo, we again turned to C. elegans. A deletion allele of Drosha, drsh-1(tm0654) causes a sterile phenotype, with no other visible defects. Specifically, we looked at the presence of alae and the structure of the vulva (Fig. 4D, g and h), as these structures are often affected by lesions in mirna pathway genes like dcr-1 (Fig. 4D, d and e) 8,9. Most probably, such defects are not observed in drsh-1(tm0654) animals because of a strong maternal rescue. Defects in the alae and vulva structures are easily detected in pash-1 RNAi knock-down animals and to a lesser degree in a pash- 1(pk2083) nonsense mutant (Fig. 4D, j and k; data not shown). Typical defects include protrusion or bursting of the vulva, and gaps or absence of the alae. In addition, we used a more sensitive and specific assay to directly monitor the activity of the let-7 mirna. The assay is based on a lacz reporter that is silenced by let-7 through sequences in the 3 0 UTR 23. This results in detectable lacz staining in all larval stages because let-7 is not yet expressed in these stages, but an absence of lacz in the let-7 expressing adult (Fig. 4D, c). As a control, we used dcr-1 mutant animals. Indeed, in this mutant background lacz is reactivated in the adult (Fig. 4D, f) 24. We then asked whether the adult-specific silencing of this reporter requires drsh-1 and/or pash-1. Both pash-1(pk2083) and drsh-1(tm0654) lead to re-expression of the reporter in the adult stage (Fig. 4D, l and i), indicating that in vivo, both Drosha and Pasha proteins are required for the function or synthesis of the let-7 mirna. Considered together, our results indicate that Pasha and Drosha are components of a multiprotein machine, the Microprocessor, which converts pri-mirnas into pre-mirnas. Although the experiments presented here do not permit us to assign a definitive function to Pasha within the Microprocessor, it is reasonable to speculate that Pasha might have one of several roles. For example, it could help in identifying primary mirna transcripts, facilitating delivery of these to Drosha for cleavage. Indeed, Pasha immunoprecipitates contain pri-mirnas (Supplementary Fig. S7). Alternatively, Pasha could help to orient the pri-mirna in the Microprocessor, contributing to the specific positioning of the Drosha cleavage site. A parallel study shows an association between DGCR8 and human Drosha, and presents biochemical and genetic evidence that these proteins cooperate to determine the specificity of Drosha cleavage 25. On the basis of the results presented here, Pasha joins RDE-4, Hyl-1 and R2D to form a growing list of dsrna-binding proteins that play important yet distinct roles in the RNAi pathway. A 234 Methods Drosophila cell culture and extract preparation Drosophila S2 cells were cultured as described previously 29. Extracts were prepared in two different ways. The first method was mainly used for chromatographic studies. Cells were washed with cold PBS and then lysed in buffer 5A (10 mm HEPES ph 7.0, 10 mm NaCl, 1.5 mm MgCl 2, 0.75 mm DTT, 0.2 mm PMSF, protease inhibitors (Roche)) for 30 min and dounce homogenized. Nuclei were spun down, and the supernatant was saved as cytoplasmic extract. The nuclei were resuspended in buffer 5B (10 mm Tris ph 8.0, 500 mm NaCl, 1.5 mm MgCl 2, 20% glycerol, 0.75 mm DTT, 0.2 mm PMSF, protease inhibitors (Roche)) and extracted for 40 min on ice. The extracted nuclei were spun down and the supernatant saved as the nuclear fraction. Nuclear and cytoplasmic fractions were combined and spun at 20,000 g for 30 min (S10). The combined extract was used in most experiments. The second method was principally for immunoprecipitations and involved lysis of cells in buffer DmLB10 (25 mm Tris ph 7.5, 150 mm NaCl, 1.5 mm MgCl 2, 1% Triton X-100, 1 mm DTT, 0.2 mm PMSF, protease inhibitors (Roche)) and a 20,000 g 30 min spin. Immunofluorescence A modification of the protocol in ref. 30 was used. S2 cells were grown on slides coated with Concanavalin A (Sigma). Pri-miRNA processing assay DNA templates for transcription of Drosha substrates were reverse transcribed with the Superscript II Kit (Invitrogen) from total RNA prepared with Trizol (Invitrogen) using primers that flank the pre-mirnas at least 50 bases. Primer sequences as follows: mir30a fwd and rev (see ref. 1); mir2a-1 T7 fwd: 5 0 -TAATACGACTCACTTAGGGATTCTGAA CTCCCGCAGTCAAGTATCC-3 0, mir2a-1 rev: 5 0 -GTGTGAATTATGTGGCGGGAGG TATT-3 0 ; mir-bantam T7-fwd: 5 0 -TAATACGACTCACTTAGGGCGCTCAGATGCAGA TGTTGTTGAT-3 0, mir-bantam rev: 5 0 -GATCGGTCGGCATAAGTTCAAAGC-3 0. Pri-miRNAs were transcribed and uniformly labelled with 32 P (Perkin Elmer) using the Riboprobe Kit (Promega). 5 0 and 3 0 labelling of cold-transcribed (Megascript kit, Ambion) RNA was done using guanylyltransferase (Ambion) and T4 DNA ligase (Amersham), respectively. A typical pri-mirna processing reaction contained 1 ml 10xD (100 mm Tris ph 8.0, 60 mm MgCl 2, 5 mm DTT), 1 ml Rnasin (Promega), 0.2 ml 32 P-labelled pri-mirna, 5 ml extract, 2.8 ml H 2 O. Reactions were allowed to proceed for 2 h at 30 8C. Chromatography AKTA FPLC was used for column chromatography. All the columns and media were purchased from Amersham Biosciences. Buffers for ion exchange chromatography: IEX A : 10 mm Tris ph 7.5, 100 mm NaCl, 1.5 mm MgCl 2, 4% glycerol, 0.75 mm DTT; IEX B : 10 mm Tris ph 7.5, 1 M NaCl, 1.5 mm MgCl 2, 4% glycerol, 0.75 mm DTT; for hydrophobic interaction chromatography HIC A : 10 mm Tris ph 7.5, 1 M NaCl, 1.5 mm MgCl 2, 4% glycerol, 0.75 mm DTT; HIC B : 10 mm Tris ph 7.5, 10 mm NaCl, 1.5 mm MgCl 2, 4% glycerol, 0.75 mm DTT. For size chromatography, buffer IEX A was used. C. elegans lacz reporter assays pkis2084 was generated by X-ray mediated integration of the extrachromosomal transgene of strain CT5 23.NotethatpkIs2084 often drives lacz expression in a few (4 5) seam cells in the head region in a wild-type background. Adult animals are scored positive for a defect in let-7-mediated silencing only if the staining extends towards the tail. As control animals, either wild-type or rrf-3(pk1426) mutant animals were used. Animals injected with dsrna against a gene not related to mirnas (mut-7) were also used as a control, with identical results. Animals carrying pkis2084 were crossed with animals carrying drsh-1(tm0654), dcr-1(s2795) or pash-1(pk2083). Progeny of injected animals and progeny of animals homozygous for the transgene and heterozygous for tm0654, pk2083 or s2795 were fixed and stained with X-gal. The genotype of stained animals was verified by PCR followed by restriction digest or sequencing. pash-1(pk2083) was obtained by target-selected mutagenesis in C. elegans (E. Cuppen, unpublished results), and introduces a PsiI site. pk2083 changes the TTA codon for L527 into TAA, introducing a premature stop. Received 9 August; accepted 16 September 2004; doi: /nature Published online 7 November Lee, Y., Jeon, K., Lee, J. T., Kim, S. & Kim, V. N. MicroRNA maturation: stepwise processing and subcellular localization. EMBO J. 21, (2002). 2. Lee,Y. et al. The nuclear RNase III Drosha initiates microrna processing. Nature 425, (2003). 3. Carrington, J. C. & Ambros, V. Role of micrornas in plant and animal development. Science 301, (2003). 4. Yi, R., Qin, Y., Macara, I. G. & Cullen, B. R. Exportin-5 mediates the nuclear export of pre-micrornas and short hairpin RNAs. Genes Dev. 17, (2003). 5. Lund, E., Guttinger, S., Calado, A., Dahlberg, J. E. & Kutay, U. Nuclear export of microrna precursors. Science 303, (2004). 6. Bohnsack, M. T., Czaplinski, K. & Gorlich, D. Exportin-5 is a RanGTP-dependent dsrna-binding protein that mediates nuclear export of pre-mirnas. RNA 10, (2004). 7. Hutvagner, G. et al. A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science 293, (2001). 8. Grishok, A. et al. Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 106, (2001). 9. Ketting, R. F. et al. Dicer functions in RNA interference and in synthesis of small RNA involved in developmental timing in C. elegans. Genes Dev. 15, (2001). 10. Song, J. J. et al. The crystal structure of the Argonaute2 PAZ domain reveals an RNA binding motif in RNAi effector complexes. Nature Struct. Biol. 10, (2003). 11. Zhang, H., Kolb, F. A., Jaskiewicz, L., Westhof, E. & Filipowicz, W. Single processing center models for human Dicer and bacterial RNase III. Cell 118, (2004). NATURE VOL NOVEMBER
5 12. Siolas, D. et al. Synthetic shrnas as highly potent RNAi triggers. Nature Biotechnol. (submitted). 13. Schwarz, D. S. et al. Asymmetry in the assembly of the RNAi enzyme complex. Cell 115, (2003). 14. Llave, C., Xie, Z., Kasschau, K. D. & Carrington, J. C. Cleavage of Scarecrow-like mrna targets directed by a class of Arabidopsis mirna. Science 297, (2002). 15. Yekta, S., Shih, I. H. & Bartel, D. P. MicroRNA-directed cleavage of HOXB8 mrna. Science 304, (2004). 16. Olsen, P. H. & Ambros, V. Thelin-4 regulatory RNAcontrols developmental timing in Caenorhabditis elegans by blocking LIN-14 protein synthesis after the initiation of translation. Dev. Biol. 216, (1999). 17. Zeng, Y., Wagner, E. J. & Cullen, B. R. Both natural and designed micro RNAs can inhibit the expression of cognate mrnas when expressed in human cells. Mol. Cell 9, (2002). 18. Doench, J. G., Petersen, C. P. & Sharp, P. A. sirnas can function as mirnas. Genes Dev. 17, (2003). 19. Wu, H., Xu, H., Miraglia, L. J. & Crooke, S. T. Human RNase III is a 160-kDa protein involved in preribosomal RNA processing. J. Biol. Chem. 275, (2000). 20. Bernstein, E., Caudy, A. A., Hammond, S. M. & Hannon, G. J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409, (2001). 21. Giot, L. et al. A protein interaction map of Drosophila melanogaster. Science 302, (2003). 22. Shiohama, A., Sasaki, T., Noda, S., Minoshima, S. & Shimizu, N. Molecular cloning and expression analysis of a novel gene DGCR8 located in the DiGeorge syndrome chromosomal region. Biochem. Biophys. Res. Commun. 304, (2003). 23. Reinhart, B. J. et al. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 403, (2000). 24. Caudy, A. A. et al. A micrococcal nuclease homologue in RNAi effector complexes. Nature 425, (2003). 25. Gregory, R. I. et al. The Microprocessor complex mediates the genesis of micrornas. Nature doi: /nature03120 (this issue). 26. Han, M. H., Goud, S., Song, L. & Fedoroff, N. The Arabidopsis double-stranded RNA-binding protein HYL1 plays a role in microrna-mediated gene regulation. Proc. Natl Acad. Sci. USA 101, (2004). 27. Liu, Q. et al. R2D2, a bridge between the initiation and effector steps of the Drosophila RNAi pathway. Science 301, (2003). 28. Tabara, H., Yigit, E., Siomi, H. & Mello, C. C. The dsrna binding protein RDE-4 interacts with RDE- 1, DCR-1, and a DExH-box helicase to direct RNAi in C. elegans. Cell 109, (2002). 29. Caudy, A. A., Myers, M., Hannon, G. J. & Hammond, S. M. Fragile X-related protein and VIG associate with the RNA interference machinery. Genes Dev. 16, (2002). 30. Spector, D. L. & Smith, H. C. Redistribution of U-snRNPs during mitosis. Exp. Cell Res. 163, (1986). implicated in mirna processing. Here we show that human Drosha is a component of two multi-protein complexes. The larger complex contains multiple classes of RNA-associated proteins including RNA helicases, proteins that bind doublestranded RNA, novel heterogeneous nuclear ribonucleoproteins and the Ewing s sarcoma family of proteins. The smaller complex is composed of Drosha and the double-stranded-rna-binding protein, DGCR8, the product of a gene deleted in DiGeorge syndrome. In vivo knock-down and in vitro reconstitution studies revealed that both components of this smaller complex, termed Microprocessor, are necessary and sufficient in mediating the genesis of mirnas from the primary mirna transcript. Supplementary Information accompanies the paper on Acknowledgements We thank F. Rivas for critical reading of the manuscript. Strain BC3825 was obtained from the C. elegans Genetics Center, and drsh-1(tm0654) was obtained from the NBPin Japan (Mitani laboratory). We thank E. Cuppen and his group for help in target-selected mutagenesis. A.M.D. is a David Koch Fellow of the Watson School of Biological Sciences. G.J.H. is supported by an Innovator Award from the US Army Breast Cancer Research Program. This work was also supported by a grant from the NIH (G.J.H.) and by a VENI fellowship from the Netherlands Organisation for Scientific Research (R.F.K.). Competing interests statement The authors declare that they have no competing financial interests. Correspondence and requests for materials should be addressed to G.H. (hannon@cshl.org).... The Microprocessor complex mediates the genesis of micrornas Richard I. Gregory*, Kai-ping Yan*, Govindasamy Amuthan, Thimmaiah Chendrimada, Behzad Doratotaj, Neil Cooch & Ramin Shiekhattar The Wistar Institute, 3601 Spruce Street, Philadelphia, Pennsylvania 19104, USA * These authors contributed equally to this work... MicroRNAs (mirnas) are a growing family of small non-protein-coding regulatory genes that regulate the expression of homologous target-gene transcripts. They have been implicated in the control of cell death and proliferation in flies 1,2, haematopoietic lineage differentiation in mammals 3, neuronal patterning in nematodes 4 and leaf and flower development in plants 5 8. mirnas are processed by the RNA-mediated interference machinery. Drosha is an RNase III enzyme that was recently Figure 1 Isolation of Drosha-containing complexes. a, Fractions of the immunoaffinity eluate from M2 anti-flag beads resolved by SDS PAGE (4 12%). Flag-Drosha was revealed by silver staining and western blotting with anti-flag antibodies. Molecular masses of marker proteins (left) and the polypeptide masses of associated subunits (right) are indicated. b, Silver staining of Superose 6 gel-filtration fractions. Top, fraction number; bottom, molecular mass markers. DAP, Drosha-associated proteins having different molecular masses; asterisks, contaminating polypeptides; IP, immunoprecipitation. NATURE VOL NOVEMBER
Chapter. Processing of primary micrornas by the Microprocessor complex
 Chapter 5 Processing of primary micrornas by the Microprocessor complex Abstract Mature micrornas (mirnas) are generated via a two-step processing pathway to yield ~22-nucleotide small RNAs that regulate
Chapter 5 Processing of primary micrornas by the Microprocessor complex Abstract Mature micrornas (mirnas) are generated via a two-step processing pathway to yield ~22-nucleotide small RNAs that regulate
mirnaselect pep-mir Cloning and Expression Vector
 Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
Functional and Biomedical Aspects of Genome Research
 Functional and Biomedical Aspects of Genome Research 20 11 35 Vorlesung SS 04 Bartsch, Jockusch & Schmitt-John Mi. 9:15-10:00, in W7-135 13 Functional RNAs Thomas Schmitt-John micro RNAs small interfering
Functional and Biomedical Aspects of Genome Research 20 11 35 Vorlesung SS 04 Bartsch, Jockusch & Schmitt-John Mi. 9:15-10:00, in W7-135 13 Functional RNAs Thomas Schmitt-John micro RNAs small interfering
Outline. interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work?
 Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
岑 祥 股 份 有 限 公 司 技 術 專 員 費 軫 尹 20100803
 技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
Micro RNAs: potentielle Biomarker für das. Blutspenderscreening
 Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
RNAi History, Mechanism and Application
 DRUG DISCOVERY AND DEVELOPMENT RNAi History, Mechanism and Application Shuo Gu The History of RNAi About the author: Dr. Shuo Gu is currently a Postdoctoral Scholar at Stanford University, School of Medicine.
DRUG DISCOVERY AND DEVELOPMENT RNAi History, Mechanism and Application Shuo Gu The History of RNAi About the author: Dr. Shuo Gu is currently a Postdoctoral Scholar at Stanford University, School of Medicine.
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation
 OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
Dicer Substrate RNAi Design
 INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
Five-year relative survival rates. Cancer. Age-adjusted cancer death rates. Proteomic Technologies for Cancer Biomarker Discovery 2010/3/22
 Cancer Five-year relative survival rates Basal lamina Underlyig tissue Normal tissue Carcinoma Invasive carcinoma 1 http://www.cancer.org/docroot/home/index.asp 2 Proteomic Technologies for Cancer Biomarker
Cancer Five-year relative survival rates Basal lamina Underlyig tissue Normal tissue Carcinoma Invasive carcinoma 1 http://www.cancer.org/docroot/home/index.asp 2 Proteomic Technologies for Cancer Biomarker
sirna and mirna: an insight into RISCs
 Review TRENDS in Biochemical Sciences Vol.30 No.2 February 2005 sirna and mirna: an insight into s Guiliang Tang Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical
Review TRENDS in Biochemical Sciences Vol.30 No.2 February 2005 sirna and mirna: an insight into s Guiliang Tang Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical
RNAi Shooting the Messenger!
 RNAi Shooting the Messenger! Bronya Keats, Ph.D. Department of Genetics Louisiana State University Health Sciences Center New Orleans Email: bkeats@lsuhsc.edu RNA interference (RNAi) A mechanism by which
RNAi Shooting the Messenger! Bronya Keats, Ph.D. Department of Genetics Louisiana State University Health Sciences Center New Orleans Email: bkeats@lsuhsc.edu RNA interference (RNAi) A mechanism by which
THE ENZYMES. Department of Microbiology, Immunology, and Molecular Genetics, Molecular Biology Institute University of California
 VOLUME THIRTY TWO THE ENZYMES Eukaryotic RNases and their Partners in RNA Degradation and Biogenesis, Part B Edited by FENG GUO Department of Biological Chemistry, David Geffen School of Medicine, Molecular
VOLUME THIRTY TWO THE ENZYMES Eukaryotic RNases and their Partners in RNA Degradation and Biogenesis, Part B Edited by FENG GUO Department of Biological Chemistry, David Geffen School of Medicine, Molecular
Functional RNAs; RNA catalysts, mirna,
 Functional RNAs; RNA catalysts, mirna, srna, RNAi... RNAs have many functions rrna (ribosomal RNA) trna (transfer RNA) mrna (Messenger RNA) snrna (including snorna) ) (Small nuclear RNA- splicing) Other
Functional RNAs; RNA catalysts, mirna, srna, RNAi... RNAs have many functions rrna (ribosomal RNA) trna (transfer RNA) mrna (Messenger RNA) snrna (including snorna) ) (Small nuclear RNA- splicing) Other
Profiling of non-coding RNA classes Gunter Meister
 Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Considerable evidence now indicates that small noncoding
 MicroRNAs and small interfering RNAs can inhibit mrna expression by similar mechanisms Yan Zeng*, Rui Yi, and Bryan R. Cullen* *Howard Hughes Medical Institute and Department of Molecular Genetics and
MicroRNAs and small interfering RNAs can inhibit mrna expression by similar mechanisms Yan Zeng*, Rui Yi, and Bryan R. Cullen* *Howard Hughes Medical Institute and Department of Molecular Genetics and
MicroRNAs: SMALL RNAs WITH A BIG ROLE IN GENE REGULATION
 MicroRNAs: SMALL RNAs WITH A BIG ROLE IN GENE REGULATION Lin He and Gregory J. Hannon MicroRNAs are a family of small, non-coding RNAs that regulate gene expression in a sequence-specific manner. The two
MicroRNAs: SMALL RNAs WITH A BIG ROLE IN GENE REGULATION Lin He and Gregory J. Hannon MicroRNAs are a family of small, non-coding RNAs that regulate gene expression in a sequence-specific manner. The two
pcas-guide System Validation in Genome Editing
 pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
Anti-ATF6 α antibody, mouse monoclonal (1-7)
 Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
In vitro analysis of pri-mirna processing. by Drosha-DGCR8 complex. (Narry Kim s lab)
 In vitro analysis of pri-mirna processing by Drosha-DGCR8 complex (Narry Kim s lab) 1-1. Preparation of radiolabeled pri-mirna transcript The RNA substrate for a cropping reaction can be prepared by in
In vitro analysis of pri-mirna processing by Drosha-DGCR8 complex (Narry Kim s lab) 1-1. Preparation of radiolabeled pri-mirna transcript The RNA substrate for a cropping reaction can be prepared by in
Classic Immunoprecipitation
 292PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Classic Immunoprecipitation Utilizes Protein A/G Agarose for Antibody Binding (Cat.
292PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Classic Immunoprecipitation Utilizes Protein A/G Agarose for Antibody Binding (Cat.
The microrna world: small is mighty
 534 The microrna world: small is mighty Peter Nelson 1, Marianthi Kiriakidou 1,2, Anup Sharma 1, Elsa Maniataki 1 and Zissimos Mourelatos 1 1 Department of Pathology, University of Pennsylvania School
534 The microrna world: small is mighty Peter Nelson 1, Marianthi Kiriakidou 1,2, Anup Sharma 1, Elsa Maniataki 1 and Zissimos Mourelatos 1 1 Department of Pathology, University of Pennsylvania School
RAPAd mirna Adenoviral Expression System Catalog #: VPK-253
 DATENBLATT RAPAd mirna Adenoviral Expression System Catalog #: VPK-253 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction MicroRNAs (mirnas) are 18 24 nucleotide RNA molecules that
DATENBLATT RAPAd mirna Adenoviral Expression System Catalog #: VPK-253 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction MicroRNAs (mirnas) are 18 24 nucleotide RNA molecules that
mirnaselect pegp-mir Cloning and Expression Vector
 Product Data Sheet mirnaselect pegp-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-GP-C STORAGE: -80ºC QUANTITY: 100 µl of bacterial glycerol stock Components 1. mirnaselect pegp-mir Cloning
Product Data Sheet mirnaselect pegp-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-GP-C STORAGE: -80ºC QUANTITY: 100 µl of bacterial glycerol stock Components 1. mirnaselect pegp-mir Cloning
PART 3.3: MicroRNA and Cancer
 BIBM 2010 Tutorial: Epigenomics and Cancer PART 3.3: MicroRNA and Cancer Dec 18, 2010 Sun Kim at Indiana University Outline of Part 3.3 Background on microrna Role of microrna in cancer MicroRNA pathway
BIBM 2010 Tutorial: Epigenomics and Cancer PART 3.3: MicroRNA and Cancer Dec 18, 2010 Sun Kim at Indiana University Outline of Part 3.3 Background on microrna Role of microrna in cancer MicroRNA pathway
V22: involvement of micrornas in GRNs
 What are micrornas? V22: involvement of micrornas in GRNs How can one identify micrornas? What is the function of micrornas? Elisa Izaurralde, MPI Tübingen Huntzinger, Izaurralde, Nat. Rev. Genet. 12,
What are micrornas? V22: involvement of micrornas in GRNs How can one identify micrornas? What is the function of micrornas? Elisa Izaurralde, MPI Tübingen Huntzinger, Izaurralde, Nat. Rev. Genet. 12,
!!!!!!!!!!!!!!!!!!!!!!!!!!
 Figure S7 cdl-1 3'UTR gld-1 binding site (4..10) let-7 binding site (326..346) Figure Legends Figure S1. gld-1 genetically interacts with nhl-2 and vig-1 during germline development. (A) nhl-2(ok818) enhances
Figure S7 cdl-1 3'UTR gld-1 binding site (4..10) let-7 binding site (326..346) Figure Legends Figure S1. gld-1 genetically interacts with nhl-2 and vig-1 during germline development. (A) nhl-2(ok818) enhances
Mir-X mirna First-Strand Synthesis Kit User Manual
 User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
Mirnas andaminas - Interrelated Physiology of Biogenesis and Networking
 The Open Applied Informatics Journal, 2008, 2, 9-13 9 In Silico Evidence of the Relationship Between mirnas and sirnas L. Montanucci 1, P. Fariselli *,1, P.L. Martelli 1, I. Rossi 1,2 and R. Casadio 1
The Open Applied Informatics Journal, 2008, 2, 9-13 9 In Silico Evidence of the Relationship Between mirnas and sirnas L. Montanucci 1, P. Fariselli *,1, P.L. Martelli 1, I. Rossi 1,2 and R. Casadio 1
Post-transcriptional control of gene expression
 Post-transcriptional control of gene expression Possible post-transcriptional controls on gene expression Only a few of these controls are likely to be important for any one gene Mecanismo de processamento
Post-transcriptional control of gene expression Possible post-transcriptional controls on gene expression Only a few of these controls are likely to be important for any one gene Mecanismo de processamento
Biogenesis, Size and Function of Small RNAs
 srnas of Plants Small, Non-coding RNAs of Plants Regulatory RNAs that act through gene silencing Two classes of small RNAs (srnas) o microrna (mirnas) Encoded by genes in the genome o small interfering
srnas of Plants Small, Non-coding RNAs of Plants Regulatory RNAs that act through gene silencing Two classes of small RNAs (srnas) o microrna (mirnas) Encoded by genes in the genome o small interfering
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
2013 W. H. Freeman and Company. 26 RNA Metabolism
 2013 W. H. Freeman and Company 26 RNA Metabolism CHAPTER 26 RNA Metabolism Key topics: Transcription: DNA-dependent synthesis of RNA Capping and splicing: RNA processing Overview of RNA Function Ribonucleic
2013 W. H. Freeman and Company 26 RNA Metabolism CHAPTER 26 RNA Metabolism Key topics: Transcription: DNA-dependent synthesis of RNA Capping and splicing: RNA processing Overview of RNA Function Ribonucleic
Lezioni Dipartimento di Oncologia Farmacologia Molecolare. RNA interference. Giovanna Damia 29 maggio 2006
 Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Comprehensive mirna Research Technologies
 Comprehensive mirna Research Technologies Sample & Assay Technologies QIAGEN solutions for advancing microrna research In the last few years, the identification of microrna (mirna) and the recognition
Comprehensive mirna Research Technologies Sample & Assay Technologies QIAGEN solutions for advancing microrna research In the last few years, the identification of microrna (mirna) and the recognition
Outline. MicroRNA Bioinformatics. microrna biogenesis. short non-coding RNAs not considered in this lecture. ! Introduction
 Outline MicroRNA Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology (CMB) Karolinska Institutet! Introduction! microrna target site prediction! Useful resources 2 short non-coding RNAs
Outline MicroRNA Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology (CMB) Karolinska Institutet! Introduction! microrna target site prediction! Useful resources 2 short non-coding RNAs
Control of Gene Expression
 Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
MicroRNA formation. 4th International Symposium on Non-Surgical Contraceptive Methods of Pet Population Control
 MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
Global MicroRNA Amplification Kit
 Global MicroRNA Amplification Kit Store kit at -20 C on receipt (ver. 3-060901) A limited-use label license covers this product. By use of this product, you accept the terms and conditions outlined in
Global MicroRNA Amplification Kit Store kit at -20 C on receipt (ver. 3-060901) A limited-use label license covers this product. By use of this product, you accept the terms and conditions outlined in
Many roads to maturity: microrna biogenesis pathways and their regulation
 Many roads to maturity: microrna biogenesis pathways and their regulation Julia Winter 1,3, Stephanie Jung 1,3, Sarina Keller 1, Richard I. Gregory 2 and Sven Diederichs 1,4 MicroRNAs are important regulators
Many roads to maturity: microrna biogenesis pathways and their regulation Julia Winter 1,3, Stephanie Jung 1,3, Sarina Keller 1, Richard I. Gregory 2 and Sven Diederichs 1,4 MicroRNAs are important regulators
sirnas can function as mirnas
 RESEARCH COMMUNICATION sirnas can function as mirnas John G. Doench, 1,3 Christian P. Petersen, 1,3 and Phillip A. Sharp 1,2,4 1 Center for Cancer Research, Department of Biology, and 2 McGovern Institute
RESEARCH COMMUNICATION sirnas can function as mirnas John G. Doench, 1,3 Christian P. Petersen, 1,3 and Phillip A. Sharp 1,2,4 1 Center for Cancer Research, Department of Biology, and 2 McGovern Institute
In silico evidence of the relationship between mirnas and sirnas
 In silico evidence of the relationship between mirnas and sirnas In silico evidence of the relationship between mirnas and sirnas Ludovica Montanucci 1, Piero Fariselli 1*, Pier Luigi Martelli 1, Ivan
In silico evidence of the relationship between mirnas and sirnas In silico evidence of the relationship between mirnas and sirnas Ludovica Montanucci 1, Piero Fariselli 1*, Pier Luigi Martelli 1, Ivan
A MicroRNA in a Multiple-Turnover RNAi Enzyme Complex
 A MicroRNA in a Multiple-Turnover RNAi Enzyme Complex György Hutvágner and Phillip D. Zamore* Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Lazare Research
A MicroRNA in a Multiple-Turnover RNAi Enzyme Complex György Hutvágner and Phillip D. Zamore* Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Lazare Research
Characterization of a highly variable eutherian microrna gene
 Characterization of a highly variable eutherian microrna gene HRISTO B. HOUBAVIY, 1 LUCAS DENNIS, 2,3 RUDOLF JAENISCH, 2,3 and PHILLIP A. SHARP 1,2 1 Center for Cancer Research and 2 Department of Biology,
Characterization of a highly variable eutherian microrna gene HRISTO B. HOUBAVIY, 1 LUCAS DENNIS, 2,3 RUDOLF JAENISCH, 2,3 and PHILLIP A. SHARP 1,2 1 Center for Cancer Research and 2 Department of Biology,
Name Class Date. Figure 13 1. 2. Which nucleotide in Figure 13 1 indicates the nucleic acid above is RNA? a. uracil c. cytosine b. guanine d.
 13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
Report. RNA Secondary Structural Determinants of mirna Precursor Processing in Arabidopsis
 Current Biology 20, 37 41, January 12, 2010 ª2010 Elsevier Ltd All rights reserved DOI 10.1016/j.cub.2009.10.076 RNA Secondary Structural Determinants of mirna Precursor Processing in Arabidopsis Report
Current Biology 20, 37 41, January 12, 2010 ª2010 Elsevier Ltd All rights reserved DOI 10.1016/j.cub.2009.10.076 RNA Secondary Structural Determinants of mirna Precursor Processing in Arabidopsis Report
EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES
 C GUGAAG EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES MicroRNA-adapted shrna (shrna mir ) for increased, specific and consistent knockdown. MicroRNA PROCESSING PATHWAY UTILIZED FOR shrna mir Developed
C GUGAAG EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES MicroRNA-adapted shrna (shrna mir ) for increased, specific and consistent knockdown. MicroRNA PROCESSING PATHWAY UTILIZED FOR shrna mir Developed
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals
 Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
CHAPTER 40 The Mechanism of Protein Synthesis
 CHAPTER 40 The Mechanism of Protein Synthesis Problems: 2,3,6,7,9,13,14,15,18,19,20 Initiation: Locating the start codon. Elongation: Reading the codons (5 3 ) and synthesizing protein amino carboxyl.
CHAPTER 40 The Mechanism of Protein Synthesis Problems: 2,3,6,7,9,13,14,15,18,19,20 Initiation: Locating the start codon. Elongation: Reading the codons (5 3 ) and synthesizing protein amino carboxyl.
Dicer-Substrate sirna Technology
 As featured in BioRadiations 120 Dicer-Substrate sirna Technology Advances in sirna Designs Improve Gene-Specific Silencing Steve Kulisch, Teresa Esch, Christina Whitman-Guliaev, and Teresa Rubio, Bio-Rad
As featured in BioRadiations 120 Dicer-Substrate sirna Technology Advances in sirna Designs Improve Gene-Specific Silencing Steve Kulisch, Teresa Esch, Christina Whitman-Guliaev, and Teresa Rubio, Bio-Rad
Viruses and micrornas
 Viruses and micrornas Bryan R Cullen The discovery of RNA interference and cellular micrornas (mirnas) has not only affected how biological research is conducted but also revealed an entirely new level
Viruses and micrornas Bryan R Cullen The discovery of RNA interference and cellular micrornas (mirnas) has not only affected how biological research is conducted but also revealed an entirely new level
Trasposable elements: P elements
 Trasposable elements: P elements In 1938 Marcus Rhodes provided the first genetic description of an unstable mutation, an allele of a gene required for the production of pigment in maize. This instability
Trasposable elements: P elements In 1938 Marcus Rhodes provided the first genetic description of an unstable mutation, an allele of a gene required for the production of pigment in maize. This instability
TOOLS sirna and mirna. User guide
 TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
User Manual/Hand book. qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse)
 User Manual/Hand book qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse) Kit Components Cat. No. MA003...Human Whole Genome mirna qpcr Profiling Kit (-20 C) The following components are sufficient
User Manual/Hand book qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse) Kit Components Cat. No. MA003...Human Whole Genome mirna qpcr Profiling Kit (-20 C) The following components are sufficient
Expression analysis of Drosophila melanogaster micrornas
 Expression analysis of Drosophila melanogaster micrornas Dissertation zur Erlangung des Doktorgrades der Mathematisch-Naturwissenschaftlichen Fakultäten der Georg-August-Universität zu Göttingen vorgelegt
Expression analysis of Drosophila melanogaster micrornas Dissertation zur Erlangung des Doktorgrades der Mathematisch-Naturwissenschaftlichen Fakultäten der Georg-August-Universität zu Göttingen vorgelegt
Construction of an Artificial MicroRNA Expression Vector for Simultaneous Inhibition of Multiple Genes in Mammalian Cells
 Int. J. Mol. Sci. 2009, 10, 2158-2168; doi:10.3390/ijms10052158 OPEN ACCESS Communication International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Construction of an Artificial
Int. J. Mol. Sci. 2009, 10, 2158-2168; doi:10.3390/ijms10052158 OPEN ACCESS Communication International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Construction of an Artificial
2.1.2 Characterization of antiviral effect of cytokine expression on HBV replication in transduced mouse hepatocytes line
 i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
RevertAid Premium First Strand cdna Synthesis Kit
 RevertAid Premium First Strand cdna Synthesis Kit #K1651, #K1652 CERTIFICATE OF ANALYSIS #K1651 Lot QUALITY CONTROL RT-PCR using 100 fg of control GAPDH RNA and GAPDH control primers generated a prominent
RevertAid Premium First Strand cdna Synthesis Kit #K1651, #K1652 CERTIFICATE OF ANALYSIS #K1651 Lot QUALITY CONTROL RT-PCR using 100 fg of control GAPDH RNA and GAPDH control primers generated a prominent
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
RNA Viruses. A Practical Approac h. Alan J. Cann
 RNA Viruses A Practical Approac h Alan J. Cann List of protocols page xiii Abbreviations xvii Investigation of RNA virus genome structure 1 A j. Easton, A.C. Marriott and C.R. Pringl e 1 Introduction-the
RNA Viruses A Practical Approac h Alan J. Cann List of protocols page xiii Abbreviations xvii Investigation of RNA virus genome structure 1 A j. Easton, A.C. Marriott and C.R. Pringl e 1 Introduction-the
Transcription and Translation of DNA
 Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Protein Synthesis How Genes Become Constituent Molecules
 Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Chapter 2 Biogenesis and Physiology of MicroRNAs
 Chapter 2 Biogenesis and Physiology of MicroRNAs Carlos A. Melo and Sonia A. Melo Abstract MicroRNAs (mirnas) are small noncoding RNAs 17 25 nucleotides long that control gene expression by promoting degradation
Chapter 2 Biogenesis and Physiology of MicroRNAs Carlos A. Melo and Sonia A. Melo Abstract MicroRNAs (mirnas) are small noncoding RNAs 17 25 nucleotides long that control gene expression by promoting degradation
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION. Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.
 Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
SUPPLEMENTARY DATA 1
 SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
Application Guide... 2
 Protocol for GenomePlex Whole Genome Amplification from Formalin-Fixed Parrafin-Embedded (FFPE) tissue Application Guide... 2 I. Description... 2 II. Product Components... 2 III. Materials to be Supplied
Protocol for GenomePlex Whole Genome Amplification from Formalin-Fixed Parrafin-Embedded (FFPE) tissue Application Guide... 2 I. Description... 2 II. Product Components... 2 III. Materials to be Supplied
Origins and Mechanisms of mirnas and sirnas
 Leading Edge Review Origins and Mechanisms of mirnas and sirnas Richard W. Carthew 1, * and Erik J. Sontheimer 1, * 1 Department of Biochemistry, Molecular Biology, and Cell Biology, Northwestern University,
Leading Edge Review Origins and Mechanisms of mirnas and sirnas Richard W. Carthew 1, * and Erik J. Sontheimer 1, * 1 Department of Biochemistry, Molecular Biology, and Cell Biology, Northwestern University,
Molecular Cloning, Product Brochure
 , Product Brochure Interest in any of the products, request or order them at Bio-Connect. Bio-Connect B.V. T NL +31 (0)26 326 44 50 T BE +32 (0)2 503 03 48 Begonialaan 3a F NL +31 (0)26 326 44 51 F BE
, Product Brochure Interest in any of the products, request or order them at Bio-Connect. Bio-Connect B.V. T NL +31 (0)26 326 44 50 T BE +32 (0)2 503 03 48 Begonialaan 3a F NL +31 (0)26 326 44 51 F BE
micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved
 microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
Principles of micro-rna production and maturation
 REVIEW (2006) 25, 6156 6162 & 2006 Nature Publishing Group All rights reserved 0950-9232/06 $30.00 www.nature.com/onc Department of Pharmacology, University of Minnesota, Minneapolis, MN, USA Micro-RNAs
REVIEW (2006) 25, 6156 6162 & 2006 Nature Publishing Group All rights reserved 0950-9232/06 $30.00 www.nature.com/onc Department of Pharmacology, University of Minnesota, Minneapolis, MN, USA Micro-RNAs
Functional characterisation of microrna-containing Argonaute protein complexes
 Dissertation zur Erlangung des Doktorgrades der Fakultät für Biologie der Ludwig-Maximilians-Universität München Functional characterisation of microrna-containing Argonaute protein complexes vorgelegt
Dissertation zur Erlangung des Doktorgrades der Fakultät für Biologie der Ludwig-Maximilians-Universität München Functional characterisation of microrna-containing Argonaute protein complexes vorgelegt
2. True or False? The sequence of nucleotides in the human genome is 90.9% identical from one person to the next. False (it s 99.
 1. True or False? A typical chromosome can contain several hundred to several thousand genes, arranged in linear order along the DNA molecule present in the chromosome. True 2. True or False? The sequence
1. True or False? A typical chromosome can contain several hundred to several thousand genes, arranged in linear order along the DNA molecule present in the chromosome. True 2. True or False? The sequence
Lecture Series 7. From DNA to Protein. Genotype to Phenotype. Reading Assignments. A. Genes and the Synthesis of Polypeptides
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
PCR was carried out in a reaction volume of 20 µl using the ABI AmpliTaq GOLD kit (ABI,
 Supplemental Text/Tables PCR Amplification and Sequencing PCR was carried out in a reaction volume of 20 µl using the ABI AmpliTaq GOLD kit (ABI, Foster City, CA). Each PCR reaction contained 20 ng genomic
Supplemental Text/Tables PCR Amplification and Sequencing PCR was carried out in a reaction volume of 20 µl using the ABI AmpliTaq GOLD kit (ABI, Foster City, CA). Each PCR reaction contained 20 ng genomic
RNAi: an ever-growing puzzle
 196 Review TRENDS in Biochemical Sciences Vol.28 No.4 April 2003 RNAi: an ever-growing puzzle Ahmet M. Denli and Gregory J. Hannon Cold Spring Harbor Laboratory, Watson School of Biological Sciences, 1
196 Review TRENDS in Biochemical Sciences Vol.28 No.4 April 2003 RNAi: an ever-growing puzzle Ahmet M. Denli and Gregory J. Hannon Cold Spring Harbor Laboratory, Watson School of Biological Sciences, 1
Identification of Novel microrna Regulatory Proteins in Neurons Using RNAi-Based Screening
 Identification of Novel microrna Regulatory Proteins in Neurons Using RNAi-Based Screening Gabriele Siegel and Gerhard Schratt, PhD University of Heidelberg Interdisciplinary Center for Neurosciences,
Identification of Novel microrna Regulatory Proteins in Neurons Using RNAi-Based Screening Gabriele Siegel and Gerhard Schratt, PhD University of Heidelberg Interdisciplinary Center for Neurosciences,
Molecular and Cell Biology Laboratory (BIOL-UA 223) Instructor: Ignatius Tan Phone: 212-998-8295 Office: 764 Brown Email: ignatius.tan@nyu.
 Molecular and Cell Biology Laboratory (BIOL-UA 223) Instructor: Ignatius Tan Phone: 212-998-8295 Office: 764 Brown Email: ignatius.tan@nyu.edu Course Hours: Section 1: Mon: 12:30-3:15 Section 2: Wed: 12:30-3:15
Molecular and Cell Biology Laboratory (BIOL-UA 223) Instructor: Ignatius Tan Phone: 212-998-8295 Office: 764 Brown Email: ignatius.tan@nyu.edu Course Hours: Section 1: Mon: 12:30-3:15 Section 2: Wed: 12:30-3:15
Transcription in prokaryotes. Elongation and termination
 Transcription in prokaryotes Elongation and termination After initiation the σ factor leaves the scene. Core polymerase is conducting the elongation of the chain. The core polymerase contains main nucleotide
Transcription in prokaryotes Elongation and termination After initiation the σ factor leaves the scene. Core polymerase is conducting the elongation of the chain. The core polymerase contains main nucleotide
MatureBayes: A Probabilistic Algorithm for Identifying the Mature mirna within Novel Precursors
 MatureBayes: A Probabilistic Algorithm for Identifying the Mature mirna within Novel Precursors Katerina Gkirtzou 1,2, Ioannis Tsamardinos 1,2, Panagiotis Tsakalides 1,2, Panayiota Poirazi 3 * 1 Computer
MatureBayes: A Probabilistic Algorithm for Identifying the Mature mirna within Novel Precursors Katerina Gkirtzou 1,2, Ioannis Tsamardinos 1,2, Panagiotis Tsakalides 1,2, Panayiota Poirazi 3 * 1 Computer
From DNA to Protein. Proteins. Chapter 13. Prokaryotes and Eukaryotes. The Path From Genes to Proteins. All proteins consist of polypeptide chains
 Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
Translation Study Guide
 Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Essentials of Real Time PCR. About Sequence Detection Chemistries
 Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Yang-Ming University, 2009 microrna Biology and Application
 Yang-Ming University, 2009 microrna Biology and Application 3/03 microrna biogenesis and functions Woan-Yuh Tarn 3/10 micrornas and development Woan- Yuh Tarn 3/17 micrornas and nervous system Jeng-Ya
Yang-Ming University, 2009 microrna Biology and Application 3/03 microrna biogenesis and functions Woan-Yuh Tarn 3/10 micrornas and development Woan- Yuh Tarn 3/17 micrornas and nervous system Jeng-Ya
Engineering of Yellow Mosaic Virus Resistance (YMVR) in Blackgram. Project ID: 1 April 2000 to 31 August 2004. Project Duration:
 Engineering of Yellow Mosaic Virus Resistance (YMVR) in Blackgram ID: Duration: Coordinator in Switzerland: PS1 1 April 2000 to 31 August 2004 Prof. Thomas Hohn University of Basel Botanisches Institut
Engineering of Yellow Mosaic Virus Resistance (YMVR) in Blackgram ID: Duration: Coordinator in Switzerland: PS1 1 April 2000 to 31 August 2004 Prof. Thomas Hohn University of Basel Botanisches Institut
Control of Gene Expression
 Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
A Novel Method to Detect Functional MicroRNA Targets
 doi:10.1016/j.jmb.2006.02.063 J. Mol. Biol. (2006) 358, 983 996 A Novel Method to Detect Functional MicroRNA Targets Sergei Vatolin*, Kapila Navaratne and Robert J. Weil Brain Tumor Institute, The Cleveland
doi:10.1016/j.jmb.2006.02.063 J. Mol. Biol. (2006) 358, 983 996 A Novel Method to Detect Functional MicroRNA Targets Sergei Vatolin*, Kapila Navaratne and Robert J. Weil Brain Tumor Institute, The Cleveland
Biosafety considerations of new dsrna molecules
 Biosafety considerations of new dsrna molecules GM-free Europe Conference 2015 May 6-8 Berlin, Germany Dr. Sarah Agapito Current GMOs Plant transformed to produce new protein A G C U G G C A A U G U C
Biosafety considerations of new dsrna molecules GM-free Europe Conference 2015 May 6-8 Berlin, Germany Dr. Sarah Agapito Current GMOs Plant transformed to produce new protein A G C U G G C A A U G U C
RNA: Transcription and Processing
 8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
8 RNA: Transcription and Processing WORKING WITH THE FIGURES 1. In Figure 8-3, why are the arrows for genes 1 and 2 pointing in opposite directions? The arrows for genes 1 and 2 indicate the direction
Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr
 User Manual Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories,
User Manual Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories,
Sickle cell anemia: Altered beta chain Single AA change (#6 Glu to Val) Consequence: Protein polymerizes Change in RBC shape ---> phenotypes
 Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
Site-Directed Nucleases and Cisgenesis Maria Fedorova, Ph.D.
 Site-Directed Nucleases and Cisgenesis Maria Fedorova, Ph.D. Regulatory Strategy Lead Enabling Technologies DuPont-Pioneer, USA 1 New Plant Breeding Techniques 2007 New Techniques Working Group established
Site-Directed Nucleases and Cisgenesis Maria Fedorova, Ph.D. Regulatory Strategy Lead Enabling Technologies DuPont-Pioneer, USA 1 New Plant Breeding Techniques 2007 New Techniques Working Group established
PreciseTM Whitepaper
 Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
RT-PCR: Two-Step Protocol
 RT-PCR: Two-Step Protocol We will provide both one-step and two-step protocols for RT-PCR. We recommend the twostep protocol for this class. In the one-step protocol, the components of RT and PCR are mixed
RT-PCR: Two-Step Protocol We will provide both one-step and two-step protocols for RT-PCR. We recommend the twostep protocol for this class. In the one-step protocol, the components of RT and PCR are mixed
GeneCopoeia Genome Editing Tools for Safe Harbor Integration in. Mice and Humans. Ed Davis, Liuqing Qian, Ruiqing li, Junsheng Zhou, and Jinkuo Zhang
 G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
Chromatin Immunoprecipitation (ChIP)
 Chromatin Immunoprecipitation (ChIP) Day 1 A) DNA shearing 1. Samples Dissect tissue (One Mouse OBs) of interest and transfer to an eppendorf containing 0.5 ml of dissecting media (on ice) or PBS but without
Chromatin Immunoprecipitation (ChIP) Day 1 A) DNA shearing 1. Samples Dissect tissue (One Mouse OBs) of interest and transfer to an eppendorf containing 0.5 ml of dissecting media (on ice) or PBS but without
Notch 1 -dependent regulation of cell fate in colorectal cancer
 Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
Complex multicellular organisms are produced by cells that switch genes on and off during development.
 Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
