Identification of Novel microrna Regulatory Proteins in Neurons Using RNAi-Based Screening
|
|
|
- Griselda Jacobs
- 8 years ago
- Views:
Transcription
1 Identification of Novel microrna Regulatory Proteins in Neurons Using RNAi-Based Screening Gabriele Siegel and Gerhard Schratt, PhD University of Heidelberg Interdisciplinary Center for Neurosciences, SFB 488 Junior Group University Hospital of Heidelberg, Institute for Neuroanatomy Heidelberg, Germany
2
3 Identification of Novel microrna Regulatory Proteins in Neurons Using RNAi-Based Screening 29 Introduction The proper functioning of the brain relies on the precise connection of billions of neurons. During the processes of learning and memory formation, specific synapses have to be modulated while neighboring contacts need to stay unchanged. This synaptic plasticity requires de novo protein synthesis in the participating synaptic compartments (Sutton and Schuman, 2006; Bramham and Wells, 2007). The local translation of preexisting mrnas in dendrites is one way to ensure tight spatiotemporal regulation of protein expression in a highly synapsespecific manner. During the past few years, a number of studies have assigned an important role for the regulation of local mrna translation in neurons to micrornas (mirnas), a class of small noncoding RNAs (Bicker and Schratt, 2008). mirnas exert a repressive effect on gene expression by binding to partially complementary sequences within the 3 untranslated region (3 UTR) of target mrnas. This repression leads to an inhibition of productive translation from the respective transcripts (Bartel, 2004). Owing to their mode of action, mirnas represent an excellent way to regulate gene expression posttranscriptionally in a tight spatial and temporal manner. Consequently, they are involved in a great variety of cellular processes, including differentiation, metabolism, and cell death. In the brain, mirnas play a crucial role at different stages of neuronal development and maturation (Fiore et al., 2008). In order to achieve repression of translation, mirnas recruit a multiprotein complex to the target mrna: the so-called mirna-associated RNA-inducedsilencing complex (mirisc). Previous studies identified several important regulators of mirna function, among which are either core components of the mirisc (e.g., Ago, GW182, Rck/p54, MOV10) (Chu and Rana, 2006; Chendrimada et al., 2007; Peters and Meister, 2007; Banerjee et al., 2009) or other regulatory proteins not directly associated with RISC (e.g., Dnd1, HuR) (Bhattacharyya et al., 2006; Kedde et al., 2007). Strikingly, all studies that screened systematically for mirna regulators were performed in nonneuronal cells, leaving open the possibility that critical neuron-specific regulators of mirisc function remain to be identified. This possibility is particularly intriguing because such regulators could couple synaptic stimulation to mirna-dependent control of local translation. In this chapter, we describe a large-scale screening approach for testing the involvement of neuronal RNA-binding proteins (RBPs) in mirisc function. We present evidence that neuronal mirisc function relies not only on some of the RBPs previously identified in nonneuronal systems, but also on additional proteins that could allow mirna function to adapt to the special needs of this highly specialized cell type to adapt. Setting Up a Screening Approach for Regulators of mirna Function in Neurons In order to identify mirna regulators in neurons, we set up a large-scale RNA interference (RNAi) screening experiment. We postulated that the knockdown of important effector proteins in neurons should relieve mirna-mediated translational inhibition. We used the brain-specific mir-134 as a paradigm, since we had recently demonstrated an important role for this mirna in the regulation of local mrna translation in neurons. In mature hippocampal neurons, mir-134 localizes to dendrites, where it inhibits the translation of the Lim-domaincontaining kinase 1 (LimK1) mrna, thereby acting as a negative regulator of dendritic spine size (Schratt et al., 2006). The very same mirna was later shown to regulate dendritic outgrowth by fine-tuning protein levels of the translational repressor Pumilio2 (Pum2) (Fiore et al., 2009). For the RNAi screening experiment, we decided to analyze the function of mir-134 using a Luciferase reporter assay. We selected a reporter gene containing an mir 134 target 3 UTR downstream of the Luciferase coding sequence. The reporter-transcript we used harbors the 3 UTR of a newly identified mir-134 target gene that, in comparison with the other mir-134 targets (Limk1 and Pum2), is more strongly repressed by mir-134, making this a suitable readout for a large-scale RNAi screen. The mirisc is formed by a few core components, such as Argonaute proteins and RCK/p54, and a number of associated factors, e.g., the fragile X mental retardation protein (FMRP) and the Vasa intronic gene (VIG) protein (Caudy et al., 2002), all of which are RBPs. In addition, RBPs like Dnd1 and HuR have been shown to interact with the 3 UTR of the target mrnas, thereby modulating mirna activity in an indirect way (Bhattacharyya et al., 2006; Kedde et al., 2007). RBPs appear to be the key players of the mirna effector machinery. Therefore, to study regulators of mirna function in neurons, we decided to focus on this class of proteins. Using small interfering RNA (sirna) technology, we planned to knock
4 30 down each neuronal RBP individually and assess its importance for mirnamediated translational repression in neurons using the described Luciferase reporter assay. Based on a previous in situ hybridization study (McKee et al., 2005), we listed all RBPs that had been shown to be expressed in postnatal mouse forebrain (~300) and ordered a custom sirna library with three individual sirnas for each candidate gene (Ambion, Austin, TX). In initial experiments, the functionality of our approach was proven by knockdown of the RISC protein GW182 in neuronal cultures, which efficiently interfered with mirna-134 mediated repression of the Luciferase reporter. For the actual screening experiments, 5 days in vitro (DIV) mouse primary cortical neurons were transfected with the mir-134 responsive Luciferasereporter construct, together with the mature mir-134 duplex and an sirna targeting one of the RBP genes. Luciferase assays were performed 2 d after transfection. Each condition was transfected in duplicates, and the entire screen was repeated twice. Hit Validation mir-134 For hit selection, we applied a threshold where at least two out of three different sirnas for each targeted gene were able to relieve the mir-134 mediated repression by at least 50% in all three runs. Applying this threshold, the screen identified 10 RBPs that are required for mir-134 mediated repression of the target gene. Among the hits are known components of the mirna pathway (GW182, RCK/p54) as well as RBPs that have not yet been implicated in mirna function and therefore might be important specifically in the neuronal system. Figure 1. Flowchart of the RNAi screening experiment to identify regulators of mir-134 function. To test the efficacy of the sirnas, we cloned constructs for overexpression of a green fluorescent protein (GFP) tagged version of the candidate RBPs. Those constructs have been cotransfected, together with the sirnas, into
5 Identification of Novel microrna Regulatory Proteins in Neurons Using RNAi-Based Screening 31 harboring the mir-134 binding site but also for the reportertranscript with the mutated binding site. Therefore, the identified RBPs appeared not only to affect mirna-134 mediated repression but to play a more general role in terms of translation regulation. This finding is not too surprising, however, because previous studies have shown that the mirisc components GW182, Ago2, and RCK/p54 behave in a similar fashion. Figure 2. Knockdown of critical RISC proteins leads to the release of mir-134 dependent repression of reporter gene expression. Top: Schematic of the mirna-associated RISC complex assembled on a target mrna. RISC components that have been shown to be required for mirna-mediated repression are indicated. Bottom: Results from a Luciferase reporter assay in cortical neurons using a GW182-siRNA in the context of mir-134 duplex RNA. Bars represent the mean of three independent experiments. Hek293 cells, and their efficacy was assessed by immunoblotting using a GFP-specific antibody. Next, we decided to validate the positive hits using several variations of the reporter assay. First, we verified whether the observed effects were indeed the result of impaired mir-134 function or whether the sirnas affect 3 UTR dependent translation in a more general fashion. To test this hypothesis, we repeated the Luciferase assay for the identified candidates using a Luciferase reporter construct with a mutated mir-134 binding site that does not show significant repression by mir-134. By doing so, we could discriminate between effects that result from a specific interaction with mir-134 and more general effects on mrna translation that might arise by reason of either binding motifs for the candidate RBPs in the Luciferase reporter mrna or direct contacts to the cap structure. Our results indicated that knockdown of the candidate RBPs leads to an increased expression of the Luciferase reporter not only for the construct Next, we investigated whether the identified RBPs are general regulators of mirna function in neurons, or whether some are specific for the studied mir- 134 target interaction. To this end, we studied the effect of the candidate sirnas in Luciferase reporter assays with different combinations of synaptic mirnas and targets. We tested the two other validated mir- 134 targets, LimK1 and Pum2, which allowed us to determine whether the RBP modulates mir-134 function in the context of a specific target gene (i.e., via direct interaction with the 3 UTR) or whether it is able to modulate mir-134 activity in general (i.e., by contributing to the mir- 134 specific RISC). We found that the knockdown of our candidate RBPs interfered with mir-134 mediated translational repression independently of the target reporter construct. Thus, we concluded that the identified RBPs seem to play a more general role in mir-134 dependent translational control. mir-138 Finally, we analyzed whether or not the RBPs also regulate the function of other dendritic mirnas. The brain-enriched mir-138 is present at synaptic sites and functions, similar to mir-134, as a negative regulator of dendritic spine size. This effect is mediated, at least in part, by translational downregulation of its target, acyl-protein-thioesterase 1 (APT 1) (Siegel et al., 2009). In order to test whether our candidate RBPs are involved in the regulation of mir-138, we tested the respective sirnas in the context of mir- 138 mediated repression of APT 1. For all RBPs tested, the knockdown showed very similar effects
6 32 on mir-138 and mir-134 function. Therefore, we concluded that the identified RBPs likely regulate the function of neuronal mirnas in a more general way rather than in an mirna-specific or target-specific manner. This finding, however, does not exclude the possibility that the function of the candidate RBPs may be modified in the context of a specific mirna target interaction. We reasoned that RBPs that are able to regulate mirna function at the level of the mature mirna likely reside in the cytoplasm. To gain insight into the subcellular localization of our candidate RBPs, we transfected the described GFP-fusion proteins into hippocampal neurons and assessed the distribution of the GFP-signal using confocal microscopy. The results revealed that the majority of the candidate RBPs showed a pronounced or exclusive signal within the nucleus, matching their described biological function during mrna splicing or nuclear export. This observation raises the question: Why were these RBPs required for mirna function in our RNAi screening setup? Because we transfected mature mirna duplex, which bypasses processing, we can rule out the option that the nuclear localized RBPs affect mirna function by simply blocking the first mirna processing steps in the nucleus. Since we know that most of the candidate RBPs are involved in RNA export or RNA splicing, a possible scenario could be that these RBPs play a significant role in the proper transport or processing of mrnas encoding important regulator proteins of mirna function, e.g., Argonaute proteins or GW182. Alternatively, these RBPs might be present in the cytoplasm at very low levels, which we were unable to detect. Additional experiments beyond the scope of this study are needed to discriminate between these possibilities. Experimental Outlook Interestingly, a subset of our candidate RBPs was expressed throughout the whole neuron, including the most distal dendritic branches. These candidates are particularly promising because they might play a direct role in the regulation of mirna function Figure 3. GFP-fusion proteins of two selected RBPs identified in the screen were expressed in hippocampal neurons. Whereas the RBP in panel A is expressed ubiquitously within the neuron, the RBP shown in B is restricted to the neuronal nucleus.
7 Identification of Novel microrna Regulatory Proteins in Neurons Using RNAi-Based Screening 33 at the synapse. Therefore, we will focus on this set of candidates in future studies investigating the underlying mechanism and biological significance of neuronal function. As a first step in our experimental efforts, we are interested in finding out whether the candidate RBPs directly associate with RISC. To this end, we will overexpress the RISC core protein Ago2 in Hek293 cells, together with the GFP-tagged RBPs, and perform co-immunoprecipitation experiments to reveal possible interactions. In addition, we will study a direct interaction of the RBPs with either the mirna and/or the target transcript by performing pull-down experiments. On a functional level, we will assess the importance of the candidate proteins for neuron maturation and dendritic spine plasticity. In the Luciferase experiments, knockdown of the candidate RBPs comparably impaired mir-134 and mir-138 function. Both mirnas are known to negatively regulate the size of dendritic spines and, in the case of mir-134, promote dendritic outgrowth (Schratt et al., 2006; Fiore et al., 2009; Siegel et al., 2009). Since the selected candidates have been shown to localize to dendrites and synaptic sites, one can surmise that they regulate mir-134 and mir-138 function in the context of spine maturation and dendritic branching. Thus, we plan to knock down the RBPs in neurons under conditions of elevated mir-134/mir-138 activity to see whether we can counteract the neuronal phenotypes of mir-134/ mir-138 overexpression. Conclusions We believe that it is feasible to perform a largescale RNAi screening experiment in primary mouse neurons. We have already identified a small number of RBPs necessary for mirna-mediated translational repression in neurons that so far had not been associated with mirna function in other cell types. Thus, it is likely that neuron-specific mirna regulators exist. They could prove to be critically involved in the modulation of mirnaregulated local protein synthesis at synaptic sites and might present a new way to couple changes in neuronal activity to the regulation of protein levels at the synapse. Acknowledgments This work was supported by the Deutsche Forschungsgemeinschaft (DFG/SFB488). G. Siegel is a recipient of an Excellence Initiative completion grant from the Graduate Academy of Heidelberg. References Banerjee S, Neveu P, Kosik KS (2009) A coordinated local translational control point at the synapse involving relief from silencing and MOV10 degradation. Neuron 64: Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116: Bhattacharyya SN, Habermacher R, Martine U, Closs EI, Filipowicz W (2006) Relief of micrornamediated translational repression in human cells subjected to stress. Cell 125: Bicker S, Schratt G (2008) micrornas: tiny regulators of synapse function in development and disease. J Cell Mol Med 12: Bramham CR, Wells DG: Dendritic mrna: transport, translation and function (2007) Nat Rev Neurosci 8: Caudy AA, Myers M, Hannon GJ, Hammond SM (2002) Fragile X related protein and VIG associate with the RNA interference machinery. Genes Dev 16: Chendrimada TP, Finn KJ, Ji X, Baillat D, Gregory RI, Liebhaber SA, Pasquinelli AE, Shiekhattar R (2007) MicroRNA silencing through RISC recruitment of eif6. Nature 447: Chu CY, Rana TM (2006) Translation repression in human cells by microrna-induced gene silencing requires RCK/p54. PLoS Biol 4:e210. Fiore R, Siegel G, Schratt G (2008) MicroRNA function in neuronal development, plasticity and disease. Biochim Biophys Acta 1779: Fiore R, Khudayberdiev S, Christensen M, Siegel G, Flavell SW, Kim TK, Greenberg ME, Schratt G (2009) Mef2-mediated transcription of the mir cluster regulates activitydependent dendritogenesis by fine-tuning Pumilio2 protein levels. EMBO J 28: Kedde M, Strasser MJ, Boldajipour B, Oude Vrielink JA, Slanchev K, le Sage C, Nagel R, Voorhoeve PM, van Duijse J, Orom UA, Lund AH, Perrakis A, Raz E, Agami R (2007) RNA-binding protein Dnd1 inhibits microrna access to target mrna. Cell 131: McKee AE, Minet E, Stern C, Riahi S, Stiles CD, Silver PA (2005) A genome-wide in situ hybridization map of RNA-binding proteins reveals anatomically restricted expression in the developing mouse brain. BMC Dev Biol 5:14. Peters L, Meister G (2007) Argonaute proteins: mediators of RNA silencing. Mol Cell 26:
8 34 Schratt GM, Tuebing F, Nigh EA, Kane CG, Sabatini ME, Kiebler M, Greenberg ME (2006) A brain-specific microrna regulates dendritic spine development. Nature 439: Siegel G, Obernosterer G, Fiore R, Oehmen M, Bicker S, Christensen M, Khudayberdiev S, Leuschner PF, Busch CJ, Kane C, Hübel K, Dekker F, Hedberg C, Rengarajan B, Drepper C, Waldmann H, Kauppinen S, Greenberg ME, Draguhn A, Rehmsmeier M, et al (2009) A functional screen implicates microrna-138- dependent regulation of the depalmitoylation enzyme APT1 in dendritic spine morphogenesis. Nat Cell Biol 11: Sutton MA, Schuman EM (2006) Dendritic protein synthesis, synaptic plasticity, and memory. Cell 127:49-58.
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
Outline. interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work?
 Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Micro RNAs: potentielle Biomarker für das. Blutspenderscreening
 Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation
 OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
The role of micrornas in synaptic development and function
 BMB reports Mini Review The role of micrornas in synaptic development and function Rachel Corbin #, Katherine Olsson-Carter # & Frank Slack * Department of Molecular, Cellular and Developmental Biology,
BMB reports Mini Review The role of micrornas in synaptic development and function Rachel Corbin #, Katherine Olsson-Carter # & Frank Slack * Department of Molecular, Cellular and Developmental Biology,
V22: involvement of micrornas in GRNs
 What are micrornas? V22: involvement of micrornas in GRNs How can one identify micrornas? What is the function of micrornas? Elisa Izaurralde, MPI Tübingen Huntzinger, Izaurralde, Nat. Rev. Genet. 12,
What are micrornas? V22: involvement of micrornas in GRNs How can one identify micrornas? What is the function of micrornas? Elisa Izaurralde, MPI Tübingen Huntzinger, Izaurralde, Nat. Rev. Genet. 12,
Functional and Biomedical Aspects of Genome Research
 Functional and Biomedical Aspects of Genome Research 20 11 35 Vorlesung SS 04 Bartsch, Jockusch & Schmitt-John Mi. 9:15-10:00, in W7-135 13 Functional RNAs Thomas Schmitt-John micro RNAs small interfering
Functional and Biomedical Aspects of Genome Research 20 11 35 Vorlesung SS 04 Bartsch, Jockusch & Schmitt-John Mi. 9:15-10:00, in W7-135 13 Functional RNAs Thomas Schmitt-John micro RNAs small interfering
岑 祥 股 份 有 限 公 司 技 術 專 員 費 軫 尹 20100803
 技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals
 Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
A role of microrna in the regulation of telomerase? Yuan Ming Yeh, Pei Rong Huang, and Tzu Chien V. Wang
 A role of microrna in the regulation of telomerase? Yuan Ming Yeh, Pei Rong Huang, and Tzu Chien V. Wang Department of Molecular and Cellular Biology, Chang Gung University, Kwei San, Tao Yuan 333, Taiwan
A role of microrna in the regulation of telomerase? Yuan Ming Yeh, Pei Rong Huang, and Tzu Chien V. Wang Department of Molecular and Cellular Biology, Chang Gung University, Kwei San, Tao Yuan 333, Taiwan
PART 3.3: MicroRNA and Cancer
 BIBM 2010 Tutorial: Epigenomics and Cancer PART 3.3: MicroRNA and Cancer Dec 18, 2010 Sun Kim at Indiana University Outline of Part 3.3 Background on microrna Role of microrna in cancer MicroRNA pathway
BIBM 2010 Tutorial: Epigenomics and Cancer PART 3.3: MicroRNA and Cancer Dec 18, 2010 Sun Kim at Indiana University Outline of Part 3.3 Background on microrna Role of microrna in cancer MicroRNA pathway
Profiling of non-coding RNA classes Gunter Meister
 Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Dicer Substrate RNAi Design
 INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
mirnaselect pep-mir Cloning and Expression Vector
 Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
SUPPLEMENTARY DATA 1
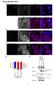 SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
Outline. MicroRNA Bioinformatics. microrna biogenesis. short non-coding RNAs not considered in this lecture. ! Introduction
 Outline MicroRNA Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology (CMB) Karolinska Institutet! Introduction! microrna target site prediction! Useful resources 2 short non-coding RNAs
Outline MicroRNA Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology (CMB) Karolinska Institutet! Introduction! microrna target site prediction! Useful resources 2 short non-coding RNAs
MicroRNA formation. 4th International Symposium on Non-Surgical Contraceptive Methods of Pet Population Control
 MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
LightSwitch Luciferase Assay System
 Constructs, Assays & Services Assay System Promoter GoClones Synthetic Response Elements Pathway Cell Lines 3 UTR GoClones mirna Mimics & Inhibitors Synthetic mirna Targets Transfection Reagents Assay
Constructs, Assays & Services Assay System Promoter GoClones Synthetic Response Elements Pathway Cell Lines 3 UTR GoClones mirna Mimics & Inhibitors Synthetic mirna Targets Transfection Reagents Assay
RNAi Shooting the Messenger!
 RNAi Shooting the Messenger! Bronya Keats, Ph.D. Department of Genetics Louisiana State University Health Sciences Center New Orleans Email: bkeats@lsuhsc.edu RNA interference (RNAi) A mechanism by which
RNAi Shooting the Messenger! Bronya Keats, Ph.D. Department of Genetics Louisiana State University Health Sciences Center New Orleans Email: bkeats@lsuhsc.edu RNA interference (RNAi) A mechanism by which
MicroRNA Mike needs help to degrade all the mrna transcripts! Aaron Arvey ISMB 2010
 Target mrna abundance dilutes microrna and sirna activity MicroRNA Mike needs help to degrade all the mrna transcripts! Aaron Arvey ISMB 2010 Target mrna abundance dilutes microrna and sirna activity Erik
Target mrna abundance dilutes microrna and sirna activity MicroRNA Mike needs help to degrade all the mrna transcripts! Aaron Arvey ISMB 2010 Target mrna abundance dilutes microrna and sirna activity Erik
How To Understand How Gene Expression Is Regulated
 What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION. Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.
 Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Understanding the immune response to bacterial infections
 Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Biogenesis, Size and Function of Small RNAs
 srnas of Plants Small, Non-coding RNAs of Plants Regulatory RNAs that act through gene silencing Two classes of small RNAs (srnas) o microrna (mirnas) Encoded by genes in the genome o small interfering
srnas of Plants Small, Non-coding RNAs of Plants Regulatory RNAs that act through gene silencing Two classes of small RNAs (srnas) o microrna (mirnas) Encoded by genes in the genome o small interfering
Functional RNAs; RNA catalysts, mirna,
 Functional RNAs; RNA catalysts, mirna, srna, RNAi... RNAs have many functions rrna (ribosomal RNA) trna (transfer RNA) mrna (Messenger RNA) snrna (including snorna) ) (Small nuclear RNA- splicing) Other
Functional RNAs; RNA catalysts, mirna, srna, RNAi... RNAs have many functions rrna (ribosomal RNA) trna (transfer RNA) mrna (Messenger RNA) snrna (including snorna) ) (Small nuclear RNA- splicing) Other
CHAPTER 40 The Mechanism of Protein Synthesis
 CHAPTER 40 The Mechanism of Protein Synthesis Problems: 2,3,6,7,9,13,14,15,18,19,20 Initiation: Locating the start codon. Elongation: Reading the codons (5 3 ) and synthesizing protein amino carboxyl.
CHAPTER 40 The Mechanism of Protein Synthesis Problems: 2,3,6,7,9,13,14,15,18,19,20 Initiation: Locating the start codon. Elongation: Reading the codons (5 3 ) and synthesizing protein amino carboxyl.
Control of Gene Expression
 Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
MeCP2 Suppresses Nuclear MicroRNA Processing and Dendritic Growth by Regulating the DGCR8/Drosha Complex
 Article MeCP2 Suppresses Nuclear MicroRNA Processing and Dendritic Growth by Regulating the DGCR8/Drosha Complex Tian-Lin Cheng, 1,4 Zhizhi Wang, 2 Qiuming Liao, 1 Ying Zhu, 1 Wen-Hao Zhou, 3 Wenqing Xu,
Article MeCP2 Suppresses Nuclear MicroRNA Processing and Dendritic Growth by Regulating the DGCR8/Drosha Complex Tian-Lin Cheng, 1,4 Zhizhi Wang, 2 Qiuming Liao, 1 Ying Zhu, 1 Wen-Hao Zhou, 3 Wenqing Xu,
Relative Quantification of mirna Target mrnas by Real-Time qpcr. 1 Introduction. Gene Expression Application Note No. 4
 Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
TOOLS sirna and mirna. User guide
 TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
Considerable evidence now indicates that small noncoding
 MicroRNAs and small interfering RNAs can inhibit mrna expression by similar mechanisms Yan Zeng*, Rui Yi, and Bryan R. Cullen* *Howard Hughes Medical Institute and Department of Molecular Genetics and
MicroRNAs and small interfering RNAs can inhibit mrna expression by similar mechanisms Yan Zeng*, Rui Yi, and Bryan R. Cullen* *Howard Hughes Medical Institute and Department of Molecular Genetics and
Five-year relative survival rates. Cancer. Age-adjusted cancer death rates. Proteomic Technologies for Cancer Biomarker Discovery 2010/3/22
 Cancer Five-year relative survival rates Basal lamina Underlyig tissue Normal tissue Carcinoma Invasive carcinoma 1 http://www.cancer.org/docroot/home/index.asp 2 Proteomic Technologies for Cancer Biomarker
Cancer Five-year relative survival rates Basal lamina Underlyig tissue Normal tissue Carcinoma Invasive carcinoma 1 http://www.cancer.org/docroot/home/index.asp 2 Proteomic Technologies for Cancer Biomarker
Notch 1 -dependent regulation of cell fate in colorectal cancer
 Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
Luísa Romão. Instituto Nacional de Saúde Dr. Ricardo Jorge Av. Padre Cruz, 1649-016 Lisboa, Portugal. Cooper et al (2009) Cell 136: 777
 Luísa Romão Instituto Nacional de Saúde Dr. Ricardo Jorge Av. Padre Cruz, 1649-016 Lisboa, Portugal Cooper et al (2009) Cell 136: 777 PTC = nonsense or stop codon = UAA, UAG, UGA PTCs can arise in a variety
Luísa Romão Instituto Nacional de Saúde Dr. Ricardo Jorge Av. Padre Cruz, 1649-016 Lisboa, Portugal Cooper et al (2009) Cell 136: 777 PTC = nonsense or stop codon = UAA, UAG, UGA PTCs can arise in a variety
RNAi History, Mechanism and Application
 DRUG DISCOVERY AND DEVELOPMENT RNAi History, Mechanism and Application Shuo Gu The History of RNAi About the author: Dr. Shuo Gu is currently a Postdoctoral Scholar at Stanford University, School of Medicine.
DRUG DISCOVERY AND DEVELOPMENT RNAi History, Mechanism and Application Shuo Gu The History of RNAi About the author: Dr. Shuo Gu is currently a Postdoctoral Scholar at Stanford University, School of Medicine.
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
THE ENZYMES. Department of Microbiology, Immunology, and Molecular Genetics, Molecular Biology Institute University of California
 VOLUME THIRTY TWO THE ENZYMES Eukaryotic RNases and their Partners in RNA Degradation and Biogenesis, Part B Edited by FENG GUO Department of Biological Chemistry, David Geffen School of Medicine, Molecular
VOLUME THIRTY TWO THE ENZYMES Eukaryotic RNases and their Partners in RNA Degradation and Biogenesis, Part B Edited by FENG GUO Department of Biological Chemistry, David Geffen School of Medicine, Molecular
Control of Gene Expression
 Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
S1 Text. Modeling deterministic single-cell microrna-p53-mdm2 network Figure 2 Figure 2
 S1 Text. Modeling deterministic single-cell microrna-p53-mdm2 network The schematic diagram of the microrna-p53-mdm2 oscillator is illustrated in Figure 2. The interaction scheme among the mrnas and the
S1 Text. Modeling deterministic single-cell microrna-p53-mdm2 network The schematic diagram of the microrna-p53-mdm2 oscillator is illustrated in Figure 2. The interaction scheme among the mrnas and the
Dicer-Substrate sirna Technology
 As featured in BioRadiations 120 Dicer-Substrate sirna Technology Advances in sirna Designs Improve Gene-Specific Silencing Steve Kulisch, Teresa Esch, Christina Whitman-Guliaev, and Teresa Rubio, Bio-Rad
As featured in BioRadiations 120 Dicer-Substrate sirna Technology Advances in sirna Designs Improve Gene-Specific Silencing Steve Kulisch, Teresa Esch, Christina Whitman-Guliaev, and Teresa Rubio, Bio-Rad
13.4 Gene Regulation and Expression
 13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
GENE REGULATION. Teacher Packet
 AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
Competition between Small RNAs: A Quantitative View
 1712 Biophysical Journal Volume 102 April 2012 1712 1721 Competition between Small RNAs: A Quantitative View Adiel Loinger, D Yael Shemla, D Itamar Simon, Hanah Margalit, and Ofer Biham * Racah Institute
1712 Biophysical Journal Volume 102 April 2012 1712 1721 Competition between Small RNAs: A Quantitative View Adiel Loinger, D Yael Shemla, D Itamar Simon, Hanah Margalit, and Ofer Biham * Racah Institute
Evidence that micrornas are associated with translating messenger RNAs in human cells
 Nature Publishing Group http://www.nature.com/nsmb Evidence that micrornas are associated with translating messenger RNAs in human cells Patricia A Maroney, Yang Yu, Jesse Fisher & Timothy W Nilsen MicroRNAs
Nature Publishing Group http://www.nature.com/nsmb Evidence that micrornas are associated with translating messenger RNAs in human cells Patricia A Maroney, Yang Yu, Jesse Fisher & Timothy W Nilsen MicroRNAs
Chapter 5: Organization and Expression of Immunoglobulin Genes
 Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Activity 7.21 Transcription factors
 Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
Neurotrophic factors and Their receptors
 Neurotrophic factors and Their receptors Huang Shu-Hong Institute of neurobiology 1 For decades, scientists believed that brain cells of the central nervous system could not regrow following damage due
Neurotrophic factors and Their receptors Huang Shu-Hong Institute of neurobiology 1 For decades, scientists believed that brain cells of the central nervous system could not regrow following damage due
Lecture Series 7. From DNA to Protein. Genotype to Phenotype. Reading Assignments. A. Genes and the Synthesis of Polypeptides
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
User Manual/Hand book. qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse)
 User Manual/Hand book qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse) Kit Components Cat. No. MA003...Human Whole Genome mirna qpcr Profiling Kit (-20 C) The following components are sufficient
User Manual/Hand book qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse) Kit Components Cat. No. MA003...Human Whole Genome mirna qpcr Profiling Kit (-20 C) The following components are sufficient
School of Nursing. Presented by Yvette Conley, PhD
 Presented by Yvette Conley, PhD What we will cover during this webcast: Briefly discuss the approaches introduced in the paper: Genome Sequencing Genome Wide Association Studies Epigenomics Gene Expression
Presented by Yvette Conley, PhD What we will cover during this webcast: Briefly discuss the approaches introduced in the paper: Genome Sequencing Genome Wide Association Studies Epigenomics Gene Expression
Mirnas andaminas - Interrelated Physiology of Biogenesis and Networking
 The Open Applied Informatics Journal, 2008, 2, 9-13 9 In Silico Evidence of the Relationship Between mirnas and sirnas L. Montanucci 1, P. Fariselli *,1, P.L. Martelli 1, I. Rossi 1,2 and R. Casadio 1
The Open Applied Informatics Journal, 2008, 2, 9-13 9 In Silico Evidence of the Relationship Between mirnas and sirnas L. Montanucci 1, P. Fariselli *,1, P.L. Martelli 1, I. Rossi 1,2 and R. Casadio 1
Lezioni Dipartimento di Oncologia Farmacologia Molecolare. RNA interference. Giovanna Damia 29 maggio 2006
 Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Product: Expression Arrest TM egfp control shrna vector
 Product: Expression Arrest TM egfp control vector Catalog #: RHS1702 Product Description The laboratory of Dr. Greg Hannon at Cold Spring Harbor Laboratory (CSHL) has created an RNAi Clone Library comprised
Product: Expression Arrest TM egfp control vector Catalog #: RHS1702 Product Description The laboratory of Dr. Greg Hannon at Cold Spring Harbor Laboratory (CSHL) has created an RNAi Clone Library comprised
Comprehensive mirna Research Technologies
 Comprehensive mirna Research Technologies Sample & Assay Technologies QIAGEN solutions for advancing microrna research In the last few years, the identification of microrna (mirna) and the recognition
Comprehensive mirna Research Technologies Sample & Assay Technologies QIAGEN solutions for advancing microrna research In the last few years, the identification of microrna (mirna) and the recognition
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Identifying microrna targets: computational and biochemical approaches
 Identifying microrna targets: computational and biochemical approaches Iddo Ben-Dov Nephrology and Hypertension Hadassah Hebrew University Medical Center T. Tuschl A Short History of a Short RNA The intellectual
Identifying microrna targets: computational and biochemical approaches Iddo Ben-Dov Nephrology and Hypertension Hadassah Hebrew University Medical Center T. Tuschl A Short History of a Short RNA The intellectual
In silico evidence of the relationship between mirnas and sirnas
 In silico evidence of the relationship between mirnas and sirnas In silico evidence of the relationship between mirnas and sirnas Ludovica Montanucci 1, Piero Fariselli 1*, Pier Luigi Martelli 1, Ivan
In silico evidence of the relationship between mirnas and sirnas In silico evidence of the relationship between mirnas and sirnas Ludovica Montanucci 1, Piero Fariselli 1*, Pier Luigi Martelli 1, Ivan
EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES
 C GUGAAG EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES MicroRNA-adapted shrna (shrna mir ) for increased, specific and consistent knockdown. MicroRNA PROCESSING PATHWAY UTILIZED FOR shrna mir Developed
C GUGAAG EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES MicroRNA-adapted shrna (shrna mir ) for increased, specific and consistent knockdown. MicroRNA PROCESSING PATHWAY UTILIZED FOR shrna mir Developed
MicroRNAs: SMALL RNAs WITH A BIG ROLE IN GENE REGULATION
 MicroRNAs: SMALL RNAs WITH A BIG ROLE IN GENE REGULATION Lin He and Gregory J. Hannon MicroRNAs are a family of small, non-coding RNAs that regulate gene expression in a sequence-specific manner. The two
MicroRNAs: SMALL RNAs WITH A BIG ROLE IN GENE REGULATION Lin He and Gregory J. Hannon MicroRNAs are a family of small, non-coding RNAs that regulate gene expression in a sequence-specific manner. The two
Recombinant DNA and Biotechnology
 Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
An Architectonic Perspective on Autism
 An Architectonic Perspective on Autism Clarence E. Schutt Princeton University QuickTime and a TIFF (Uncompressed) decompressor are needed to see this picture. St. Peter s College March 5, 2007 QuickTime
An Architectonic Perspective on Autism Clarence E. Schutt Princeton University QuickTime and a TIFF (Uncompressed) decompressor are needed to see this picture. St. Peter s College March 5, 2007 QuickTime
The Need for a PARP in vivo Pharmacodynamic Assay
 The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D., Chief Scientific Officer, Trevigen, Inc., Gaithersburg, MD For further infomation, please contact: William Booth, Ph.D. Tel: +44 (0)1235
The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D., Chief Scientific Officer, Trevigen, Inc., Gaithersburg, MD For further infomation, please contact: William Booth, Ph.D. Tel: +44 (0)1235
Protein Synthesis How Genes Become Constituent Molecules
 Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Module 3 Questions. 7. Chemotaxis is an example of signal transduction. Explain, with the use of diagrams.
 Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Thermo Scientific Dharmacon ON-TARGETplus sirna The Standard for sirna Specificity
 Thermo Scientific Dharmacon sirna The Standard for sirna Specificity Reduce off-targets up to 90% with the market-leading sirna Combine higher target specificity with guaranteed silencing Increase confidence
Thermo Scientific Dharmacon sirna The Standard for sirna Specificity Reduce off-targets up to 90% with the market-leading sirna Combine higher target specificity with guaranteed silencing Increase confidence
A MicroRNA in a Multiple-Turnover RNAi Enzyme Complex
 A MicroRNA in a Multiple-Turnover RNAi Enzyme Complex György Hutvágner and Phillip D. Zamore* Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Lazare Research
A MicroRNA in a Multiple-Turnover RNAi Enzyme Complex György Hutvágner and Phillip D. Zamore* Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Lazare Research
Name Class Date. Figure 13 1. 2. Which nucleotide in Figure 13 1 indicates the nucleic acid above is RNA? a. uracil c. cytosine b. guanine d.
 13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
Chapter 2 Biogenesis and Physiology of MicroRNAs
 Chapter 2 Biogenesis and Physiology of MicroRNAs Carlos A. Melo and Sonia A. Melo Abstract MicroRNAs (mirnas) are small noncoding RNAs 17 25 nucleotides long that control gene expression by promoting degradation
Chapter 2 Biogenesis and Physiology of MicroRNAs Carlos A. Melo and Sonia A. Melo Abstract MicroRNAs (mirnas) are small noncoding RNAs 17 25 nucleotides long that control gene expression by promoting degradation
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Gene Models & Bed format: What they represent.
 GeneModels&Bedformat:Whattheyrepresent. Gene models are hypotheses about the structure of transcripts produced by a gene. Like all models, they may be correct, partly correct, or entirely wrong. Typically,
GeneModels&Bedformat:Whattheyrepresent. Gene models are hypotheses about the structure of transcripts produced by a gene. Like all models, they may be correct, partly correct, or entirely wrong. Typically,
HiPerFect Transfection Reagent Handbook
 Fifth Edition October 2010 HiPerFect Transfection Reagent Handbook For transfection of eukaryotic cells with sirna and mirna Sample & Assay Technologies QIAGEN Sample and Assay Technologies QIAGEN is the
Fifth Edition October 2010 HiPerFect Transfection Reagent Handbook For transfection of eukaryotic cells with sirna and mirna Sample & Assay Technologies QIAGEN Sample and Assay Technologies QIAGEN is the
Functional Analysis. mircury LNA microrna Mimics
 Functional Analysis mircury LNA microrna Mimics Instruction manual v1.0 April 2014 Table of Contents Product summary......................................................... 3 Content..................................................................
Functional Analysis mircury LNA microrna Mimics Instruction manual v1.0 April 2014 Table of Contents Product summary......................................................... 3 Content..................................................................
sirna Duplexes & RNAi Explorer
 P r o d u c t S p e c i f i c a t i o n s Guaranteed RNAi Explorer kit with Fl/Dabcyl Molecular Beacon Catalog No.:27-6402-01 Guaranteed RNAi Explorer kit with Fluorescein/Tamra TaqMan Catalog No.:27-6402-01
P r o d u c t S p e c i f i c a t i o n s Guaranteed RNAi Explorer kit with Fl/Dabcyl Molecular Beacon Catalog No.:27-6402-01 Guaranteed RNAi Explorer kit with Fluorescein/Tamra TaqMan Catalog No.:27-6402-01
Positive Feedback and Bistable Systems. Copyright 2008: Sauro
 Positive Feedback and Bistable Systems 1 Copyright 2008: Sauro Non-Hysteretic Switches; Ultrasensitivity; Memoryless Switches Output These systems have no memory, that is, once the input signal is removed,
Positive Feedback and Bistable Systems 1 Copyright 2008: Sauro Non-Hysteretic Switches; Ultrasensitivity; Memoryless Switches Output These systems have no memory, that is, once the input signal is removed,
Many roads to maturity: microrna biogenesis pathways and their regulation
 Many roads to maturity: microrna biogenesis pathways and their regulation Julia Winter 1,3, Stephanie Jung 1,3, Sarina Keller 1, Richard I. Gregory 2 and Sven Diederichs 1,4 MicroRNAs are important regulators
Many roads to maturity: microrna biogenesis pathways and their regulation Julia Winter 1,3, Stephanie Jung 1,3, Sarina Keller 1, Richard I. Gregory 2 and Sven Diederichs 1,4 MicroRNAs are important regulators
Inhibition of MicroRNA Sensitizes 3D Breast Cancer Microtissues to Radiation Therapy
 CASE STUDY High Content Image Analysis Inhibition of MicroRNA Sensitizes 3D Breast Cancer Microtissues to Radiation Therapy Dr. Nataša Anastasov leads the Personalized Radiation Therapy Group within the
CASE STUDY High Content Image Analysis Inhibition of MicroRNA Sensitizes 3D Breast Cancer Microtissues to Radiation Therapy Dr. Nataša Anastasov leads the Personalized Radiation Therapy Group within the
TECHNICAL INSIGHTS TECHNOLOGY ALERT
 TECHNICAL INSIGHTS TECHNOLOGY ALERT UNLOCKING BREAST CANCER TUMORS' DRUG RESISTANCE A troubling phenomenon in breast cancer treatment is that some patients tumors become resistant to treatments over time.
TECHNICAL INSIGHTS TECHNOLOGY ALERT UNLOCKING BREAST CANCER TUMORS' DRUG RESISTANCE A troubling phenomenon in breast cancer treatment is that some patients tumors become resistant to treatments over time.
The microrna world: small is mighty
 534 The microrna world: small is mighty Peter Nelson 1, Marianthi Kiriakidou 1,2, Anup Sharma 1, Elsa Maniataki 1 and Zissimos Mourelatos 1 1 Department of Pathology, University of Pennsylvania School
534 The microrna world: small is mighty Peter Nelson 1, Marianthi Kiriakidou 1,2, Anup Sharma 1, Elsa Maniataki 1 and Zissimos Mourelatos 1 1 Department of Pathology, University of Pennsylvania School
Control of Gene Expression
 Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Analyzing microrna Data and Integrating mirna with Gene Expression Data in Partek Genomics Suite 6.6
 Analyzing microrna Data and Integrating mirna with Gene Expression Data in Partek Genomics Suite 6.6 Overview This tutorial outlines how microrna data can be analyzed within Partek Genomics Suite. Additionally,
Analyzing microrna Data and Integrating mirna with Gene Expression Data in Partek Genomics Suite 6.6 Overview This tutorial outlines how microrna data can be analyzed within Partek Genomics Suite. Additionally,
Supplemental Fig. S1. The schematic diagrams of the expression constructs used in this study.
 1 Supplemental data Supplemental Fig. S1. The schematic diagrams of the expression constructs used in this study. Supplemental Fig. S2. Ingenuity Pathway Analysis (IPA) of the 56 putative caspase substrates
1 Supplemental data Supplemental Fig. S1. The schematic diagrams of the expression constructs used in this study. Supplemental Fig. S2. Ingenuity Pathway Analysis (IPA) of the 56 putative caspase substrates
Thermo Scientific Dharmacon sirna Libraries
 Thermo Scientific Dharmacon sirna Libraries Scientific innovation in sirna potency, specificity and delivery Superior sirna in comprehensive collections for reliable screens Market leading products for
Thermo Scientific Dharmacon sirna Libraries Scientific innovation in sirna potency, specificity and delivery Superior sirna in comprehensive collections for reliable screens Market leading products for
Chapter 8. Summary and Perspectives
 Chapter 8 Summary and Perspectives 131 Chapter 8 Summary Overexpression of the multidrug resistance protein MRP1 confer multidrug resistance (MDR) to cancer cells. The contents of this thesis describe
Chapter 8 Summary and Perspectives 131 Chapter 8 Summary Overexpression of the multidrug resistance protein MRP1 confer multidrug resistance (MDR) to cancer cells. The contents of this thesis describe
Mathematical Modeling of microrna-mediated Mechanisms of Translation Repression
 Mathematical Modeling of microrna-mediated Mechanisms of Translation Repression 11 Andrei Zinovyev, Nadya Morozova, Alexander N. Gorban, and Annick Harel-Belan Abstract MicroRNAs can affect the protein
Mathematical Modeling of microrna-mediated Mechanisms of Translation Repression 11 Andrei Zinovyev, Nadya Morozova, Alexander N. Gorban, and Annick Harel-Belan Abstract MicroRNAs can affect the protein
J. Cell Sci. 128: doi:10.1242/jcs.164566: Supplementary Material
 Figure S1. Microtubule and microfilament drug treatments severely interfere with mitotic spindle morphology, length and positioning. (A) GFP-fused lamin-a/c, LAP2 or BAF1 localizes to the spindle and all
Figure S1. Microtubule and microfilament drug treatments severely interfere with mitotic spindle morphology, length and positioning. (A) GFP-fused lamin-a/c, LAP2 or BAF1 localizes to the spindle and all
The RNA strategy. RNA as a tool and target in human disease diagnosis and therapy.
 The RNA strategy RNA as a tool and target in human disease diagnosis and therapy. The Laboratory of RNA Biology and Biotechnology at the Centre for Integrative Biology (CIBIO) of the University of Trento,
The RNA strategy RNA as a tool and target in human disease diagnosis and therapy. The Laboratory of RNA Biology and Biotechnology at the Centre for Integrative Biology (CIBIO) of the University of Trento,
Biological Sciences Initiative. Human Genome
 Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Complex multicellular organisms are produced by cells that switch genes on and off during development.
 Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Sickle cell anemia: Altered beta chain Single AA change (#6 Glu to Val) Consequence: Protein polymerizes Change in RBC shape ---> phenotypes
 Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
Discovery & Modeling of Genomic Regulatory Networks with Big Data
 Discovery & Modeling of Genomic Regulatory Networks with Big Data Hamid Bolouri Division of Human Biology Fred Hutchinson Cancer Research Center labs.fhcrc.org/bolouri I have no financial relationships
Discovery & Modeling of Genomic Regulatory Networks with Big Data Hamid Bolouri Division of Human Biology Fred Hutchinson Cancer Research Center labs.fhcrc.org/bolouri I have no financial relationships
How To Identify A Novel Pathway Leading To Myocardial Infarction
 Press Release Embargo: 10 November 2013 at 1800 London time / 1300 US Eastern Time Genetic research identifies novel pathway leading to myocardial infarction Starting with a severely affected family, a
Press Release Embargo: 10 November 2013 at 1800 London time / 1300 US Eastern Time Genetic research identifies novel pathway leading to myocardial infarction Starting with a severely affected family, a
Protein-responsive ribozyme switches in eukaryotic cells
 Protein-responsive ribozyme switches in eukaryotic cells Andrew B. Kennedy, James V. Vowles, Leo d Espaux, and Christina D. Smolke Presented by Marianne Linz and Jennifer Thornton March 11, 2015 Synthetic
Protein-responsive ribozyme switches in eukaryotic cells Andrew B. Kennedy, James V. Vowles, Leo d Espaux, and Christina D. Smolke Presented by Marianne Linz and Jennifer Thornton March 11, 2015 Synthetic
Technical Note: Analysis of microrna function using mircury LNA TM microrna Inhibitors
 Technical note: Analysis of microrna function using mircury LNA microrna Inhibitors Introduction The study of micrornas has come a long way since the discovery of these small RNAs some fifteen years ago.
Technical note: Analysis of microrna function using mircury LNA microrna Inhibitors Introduction The study of micrornas has come a long way since the discovery of these small RNAs some fifteen years ago.
Determinants of targeting by endogenous and exogenous micrornas and sirnas
 Determinants of targeting by endogenous and exogenous micrornas and sirnas CYDNEY B. NIELSEN, 1,3 NOAM SHOMRON, 1,3 RICKARD SANDBERG, 1 ERAN HORNSTEIN, 2,4 JACOB KITZMAN, 1,5 and CHRISTOPHER B. BURGE 1
Determinants of targeting by endogenous and exogenous micrornas and sirnas CYDNEY B. NIELSEN, 1,3 NOAM SHOMRON, 1,3 RICKARD SANDBERG, 1 ERAN HORNSTEIN, 2,4 JACOB KITZMAN, 1,5 and CHRISTOPHER B. BURGE 1
How to measure mirna expression. Matt Barter
 How to measure mirna expression Matt Barter Expansion of microrna research Pubmed publications 2000 2001 2002 2003 2004 2005 2006 2007 2008 2009 2010 2011 2012 6000 5000 4000 3000 * 2000 1000 0 Year Micro-managers
How to measure mirna expression Matt Barter Expansion of microrna research Pubmed publications 2000 2001 2002 2003 2004 2005 2006 2007 2008 2009 2010 2011 2012 6000 5000 4000 3000 * 2000 1000 0 Year Micro-managers
Activation and effector functions of HMI
 Activation and effector functions of HMI Hathairat Thananchai, DPhil Department of Microbiology Faculty of Medicine Chiang Mai University 25 August 2015 ว ตถ ประสงค หล งจากช วโมงบรรยายน แล วน กศ กษาสามารถ
Activation and effector functions of HMI Hathairat Thananchai, DPhil Department of Microbiology Faculty of Medicine Chiang Mai University 25 August 2015 ว ตถ ประสงค หล งจากช วโมงบรรยายน แล วน กศ กษาสามารถ
Innate Immunity. Insects rely solely on an innate immune system for defense against infection
 Innate Immunity Insects rely solely on an innate immune system for defense against infection The importance of mammalian innate immunity has only recently become appreciated The innate immune response
Innate Immunity Insects rely solely on an innate immune system for defense against infection The importance of mammalian innate immunity has only recently become appreciated The innate immune response
Core Facility Genomics
 Core Facility Genomics versatile genome or transcriptome analyses based on quantifiable highthroughput data ascertainment 1 Topics Collaboration with Harald Binder and Clemens Kreutz Project: Microarray
Core Facility Genomics versatile genome or transcriptome analyses based on quantifiable highthroughput data ascertainment 1 Topics Collaboration with Harald Binder and Clemens Kreutz Project: Microarray
Masters research projects. 1. Adapting Granger causality for use on EEG data.
 Masters research projects 1. Adapting Granger causality for use on EEG data. Background. Granger causality is a concept introduced in the field of economy to determine which variables influence, or cause,
Masters research projects 1. Adapting Granger causality for use on EEG data. Background. Granger causality is a concept introduced in the field of economy to determine which variables influence, or cause,
Supplementary Figure 1.
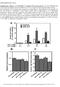 Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
