Report. RNA Secondary Structural Determinants of mirna Precursor Processing in Arabidopsis
|
|
|
- Russell Tate
- 8 years ago
- Views:
Transcription
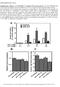
1 Current Biology 20, 37 41, January 12, 2010 ª2010 Elsevier Ltd All rights reserved DOI /j.cub RNA Secondary Structural Determinants of mirna Precursor Processing in Arabidopsis Report Liang Song, 1,2,3 Michael J. Axtell, 1,2,3 and Nina V. Fedoroff 1,2,3,4, * 1 The Huck Institutes of the Life Sciences 2 Department of Biology 3 Plant Biology Graduate Program Pennsylvania State University, University Park, PA 16802, USA We also analyzed plants overexpressing pri-mir167a, whose microrna (mirna) is cleaved from the 5 0 arm of the hairpin, unlike pri-mir171a, whose mirna is cleaved from the hairpin s 3 0 arm. As reported by others [5, 6], 35S:MIR167a plants have shorter siliques as a result of reduced fertility (Figure S1D). We used silique length to quantify the effect of overexpressing pri-mir167a structural variants. Summary MicroRNAs (mirnas) are excised from hairpin structures within primary mirnas (pri-mirnas). Most animal primirnas are processed by two cleavages, the first at a loopdistal site w11 nucleotides (nt) from the end of the hairpin and the second w22 nt beyond the first [1 3]. To identify RNA structural determinants of mirna processing in plants, we analyzed the functional consequences of changing the secondary structure of the lower (loop-distal), middle (mirna:mirna*), and upper (loop-proximal) stems of the hairpin in two different pri-mirnas. Closing bulges immediately below the loop-distal cleavage sites increased the accumulation of accurately cleaved precursor mirnas but decreased the abundance of the mature mirnas. A primirna variant with an unpaired lower stem was not processed, and variants with a perfectly paired middle or upper stem were processed normally. Bioinformatic analysis of pri-mirna structures, together with physical mapping of initial cleavage sites and in vitro processing of pri-mirna, reveals that the first, loop-distal cleavage is often at a distance of w15 nt from an unpaired region. Hence, a common determinant of the rate and location of the initial pri-mirna cleavage is an imperfectly base-paired duplex of w15 nt between the mirna:mirna* duplex and either a less structured region of the lower stem or its end. Results and Discussion Overexpression of Primary MicroRNAs 171a and 167a Causes Developmental Abnormalities Overexpression of mir171, which is both deeply conserved and highly expressed, from a cauliflower mosaic virus 35S promoter (35S:MIR171a) causes multiple developmental defects (see Figure S1A available online). The predominant defects are fewer cauline and rosette leaves and reduced shoot branching (Figures S1A and S1B). These defects are not attributable to mir171-triggered secondary small interfering RNAs (sirnas), because they occurred in the sgs3 background, which is deficient in secondary sirna production (Figure S1A) [4]. Moreover, the phenotypes were positively correlated with the amount of mature mir171 (Figure S1C). We used the number of cauline leaves as a quantifiable trait in analyzing the RNA secondary structural variants described below. *Correspondence: nvf1@psu.edu 4 Present address: US Department of State, 2101 C Street NW, Washington, DC 20520, USA Phenotypic Analysis of Plants Expressing Primary mirna Structural Variants We closed bulges and repaired mismatches in pri-mir171a to make two variants: MIR171a-Closed Bulges Lower stem (MIR171a-CBL), which lacked bulges immediately below the loop-distal cleavage site, and MIR171a-Closed Bulges Middle stem (MIR171a-CBM), which lacked the middle-stem bulge in the mirna:mirna* duplex (Figure 1A). All T1 MIR171a overexpressers had fewer cauline leaves than vector-transformed controls (Figure 1B), indicating that no variants were completely defective in mir171a function. 35S:MIR171a-WT plants had significantly fewer cauline leaves than 35S:MIR171a-CBL plants in two independent transformations (p < 0.01, Mann- Whitney test; Figure 1B; Figure S2A). The phenotypic abnormalities of 35S:MIR171a-WT plants were less severe than those of 35S:MIR171a-CBM plants; however, the difference was not statistically significant in one of two independent transformations (p > 0.05, Figures 1B; p < 0.01, Figure S2A). These observations suggest that bulges in the lower stem immediately below the loop-distal DCL1 cleavage site are important for mirna processing, whereas the bulge in the mirna sequence may be less important. We made two similar variants of pri-mir167a, MIR167a-CBL and MIR167a-CBM, as well as two additional ones: MIR167a- Abolished Lower stem (MIR167a-AL) and MIR167a-Closed Bulges Upper stem (MIR167a-CBU) (Figure 1C). More of the T1 plants developed shorter siliques than vector-transformed controls for all 35S:MIR167a variants except 35S:MIR167a-AL, indicating that the lower stem of the hairpin is critical for processing. However, as observed for pri-mir171a, the abnormalities of 35S:MIR167a-CBL plants were less severe than those of 35S:MIR167a-WT plants in three independent transformations (p < 0.001, Figure 1D; p < 0.05, Figure S2B), suggesting inefficient silencing of mirna targets in the CBL lines. By contrast, the severity of the silique phenotype was similar in the 35S:MIR167a-CBU, 35S:MIR167a-CBM, and 35S:MIR167a-WT T1 plants (p > 0.05, Figure 1D). Thus, both increasing and decreasing base pairing in the lower stem immediately below the loop-distal cleavage site appears to interfere with mirna biogenesis, whereas eliminating bulges in the upper and middle stems does not. The difference between 35S:MIR167a-CBM and 35S:MIR171a-CBM suggests that the consequence of increasing base pairing in the middle stem may vary among MIRNAs. In summary, the results of the phenotypic analyses identify the secondary structure of the lower stem as critical for mirna biogenesis, whereas the secondary structure of the upper and middle stems is less or not important.
2 Current Biology Vol 20 No 1 38 Figure 1. The Importance of the Lower Stem for Arabidopsis Primary mirna Function (A) A diagram of pri-mir171a structural variants. Mutated nucleotides (nt) are indicated by red arrowheads. The following abbreviations are used: CBL, Closed Bulges Lower stem; CBM, Closed Bulges Middle stem. Red nucleotides represent mature mir171a, and blue nucleotides represent mir171a*. (B) Top: distribution of cauline leaf number in the indicated T1 transgenic plants. The numbers of plants analyzed are shown in parentheses. Bottom: pairwise p value cutoffs for significant differences in cauline leaf number distributions in 35S:MIR171a T1 populations (Mann-Whitney test). (C) As in (A) for pri-mir167a structural variants. The following abbreviations are used: AL, Abolished Lower stem; CBU, Closed Bulges Upper stem. (D) As in (B) for the silique length phenotype of pri-mir167a-overexpressing plants. The Lower Stem Is Critical for mirna Processing We then focused on plants overexpressing primary mirnas (pri-mirnas) with altered lower stems. The abundance of the pri-, pre-, and mature mirnas was examined. 35S:MIRNA- CBL plants accumulated less pri-mirna than 35S:MIRNA- WT plants for both MIRNAs (Figure 2A). Plants expressing 35S:MIR167a-AL and 35S:MIR167a-WT accumulated similar amounts of pri-mirna (Figure 2A). Thus, complete disruption of the lower stem did not affect pri-mirna abundance, whereas eliminating lower-stem mismatches reduced its accumulation. RNA blots detected species with sizes consistent with pre-mir171a and pre-mir167a specifically in samples derived from CBL plants (Figures 2B and 2C). Hence, 35S:MIRNA-CBL plants accumulated less pri-mirna and more precursor mirna (pre-mirna) than 35S:MIRNA-WT plants, suggesting that DCL1 cleaves the more extensively basepaired substrate more efficiently than the mispaired substrate. However, RNA blots also showed that 35S:MIR171a-CBL and 35S:MIR167a-CBL plants accumulated less mature mirna than their wild-type counterparts (Figures 2B and 2C), indicating either that the second cleavage that releases mirna: mirna* from the pre-mirna is slower or that processing accuracy has decreased [7]. 35S:MIR167a-AL and vector control plants accumulated similar amounts of mir167 and mir167a* (Figure 2C), suggesting that pri-mir167a-al cannot be processed. pri-mirna secondary structural changes could affect cleavage accuracy, rate, or both. Animal Dicer processes pre-mirnas to release the mirna:mirna* duplex; the cleavage site is determined by a measurement of w22 nt [8]. If DCL1 cleaves plant pre-mirnas by a similar mechanism, the identity of the released mirna:mirna* duplex would be determined solely by the position of the first DCL1 cleavage site in pri-mirna. To assess the accuracy of the first DCL1 cleavage in MIRNA structural variants, we used 5 0 RNA ligase-mediated rapid amplification of cdna ends (5 0 - RLM-RACE) to detect 3 0 remnants of cleaved pri-mirnas RLM-RACE results directly support the previous conclusions that both perfectly basepaired CBL variants are cleaved more efficiently than the wild-type pri-mirnas and that primir167a-al is not processed by DCL1 (Figure S3). The most frequently occurring 5 0 ends reflect accurate cleavage at the loop-distal site in all samples except for 35S:MIR167a-AL (Figures 3A and 3B), indicating that processing accuracy at the loop-distal cleavage site was not affected in the CBL variants. This implies that the 35S:MIRNA-CBL lines accumulate less mirna and mirna* because they process pre-mirna less efficiently. We also detected more random single cleavage sites in 35S:MIRNA-WT than in 35S:MIRNA-CBL samples (Figures 3A and 3B). Because the wild-type transgenes are highly expressed but processed less efficiently than the CBL variants, the unique 5 0 ends are probably due to nonspecific degradation of the transgenic pri-mirna. Similarly, the random distribution of unique 5 0 ends in pri-mir167a-al suggests that DCL1 was unable to process this precursor and that the remnants derived from random cleavage and degradation (Figure 3A). The results obtained with both CBL variants suggest that unpaired residues immediately below the loop-distal DCL1 cleavage site reduce the rate of the loop-distal cleavage but are necessary for the efficient loop-proximal cleavage to liberate the mirna:mirna* duplex. In plants, both pri-mirna and pre-mirna are processed by DCL1 [9, 10]. The increased base pairing in the lower stem may alter the affinity of the DCL1 complex for the RNA substrates, accelerating the first cleavage but perhaps retarding rearrangement of the complex for the second cleavage. Alternatively, the more highly duplexed CBL variants may be cleaved by the DCL4-DRB4 complex rather than the DCL1-HYL1-SE complex. Biogenesis of some newly evolved mirnas depends on DCL4 instead of DCL1, possibly because of the long, near-perfectly basepaired structure of the pri-mirna hairpins [11] and the greater affinity of DRB4 than HYL1 for double-stranded RNA [12].
3 Plant mirna Biogenesis 39 Figure 2. CBL Variants of MIR167a and MIR171a Accumulate Precursor mirna (A) Reverse transcriptase-polymerase chain reaction to detect unprocessed transgenic primir171a (top) or unprocessed transgenic primir167a (bottom) in the indicated transgenic lines. Ubq1 was used as a loading control. (B) RNA blots for the indicated species in the indicated transgenic lines. The expected mobility of pre-mir171a is indicated by solid arrowheads; mature mir171 or mir171a* is indicated by hollow arrowheads. mir159 was used as a loading control. The blot in the first three panels was reprobed twice, and the blot in the last two panels was reprobed once with indicated oligos. (C) As in (B) for MIR167a variants. RNA Secondary Structural Motifs in pri-mirna Processing A second cluster of cleavage sites was observed downstream from the loop-distal cleavage site in pri-mir171a constructs, which had a longer hairpin than the endogenous pri-mir171a because of sequences introduced during cloning (Figure 3B). The transgene-specific cluster occurred in a stem region nt from a multibranched loop (Figure 3B). In vitro processing of pri-mir171a with the long hairpin by a recombinant DCL1-HYL1-SE complex released an approximately 40 nt 5 0 fragment and an artificial mirna (Figure 3C). This is consistent with the extra cleavage sites identified by 5 0 -RLM-RACE, indicating that the extended hairpin contains sufficient structural information for an extra DCL1 cleavage. Because all of the frequently occurring 5 0 ends of pri-mir167a and pri-mir171a are located at a distance of about 15 nt from a less-structured region of the hairpin, we infer that the initial DCL1 cleavage is determined by the presence of a relatively unstructured segment in the pri-mirna hairpin at a distance of w15 nt from the cleavage site. To ask whether such RNA secondary structural motifs are common in Arabidopsis pri-mirna hairpins, we analyzed a filtered set of 82 Arabidopsis MIRNA loci selected from mirbase 13.0 [13]. These 82 loci were those with strong experimental evidence of canonical MIRNA processing (see Supplemental Experimental Procedures). The length of the stem regions within these hairpins was highly diverse, with heterogeneity apparent within all stem regions (Figure S4). This size heterogeneity suggests that, by contrast to what is observed in animals, pri-mirna processing is not always initiated by a consistent distance from the end of the lower stem [3]. To identify common secondary structural features, we counted the number of hairpins in which the stem ends or a big loop (defined as a loop contained within a single stem arm that contains no less than three consecutive unpaired nucleotides) begins at each position in the predicted hairpins. A clear peak of stem ends and/ or big loops is observed in the lower stem nt from the loop-distal cleavage site, adjacent to a valley in the distribution at nt (Figure 4). The frequencies of stem ends and big loops observed at positions in the lower stem are highly unlikely to have occurred by chance (p < 0.001). In 41 of the 82 analyzed pri-mirna hairpins, the loopdistal DCL1 cleavage site resides w15 nt from an unpaired region in the lower stem. Stem ends and big loops are more evenly distributed in the upper stem, indicating that the upper stem lacks such a high-frequency secondary structural motif. Conclusions We have demonstrated that the lower stem is critical for mirna biogenesis. This is consistent with observations described in the accompanying papers by Werner et al. and Mateos et al. in this issue of Current Biology [14, 15]. We also discovered that closing bulges immediately below the loop-distal DCL1 cleavage site has opposite impacts on the efficiency of first and second DCL1 cleavages. We speculate that during evolution, mutations in the lower stem immediately below the initial DCL1 cleavage site may fine-tune the amount of mirnas. Our analyses also suggest that the first DCL1 cleavage in mirna processing often occurs at a distance of w15 nt from either the end of the hairpin or an internal unstructured region. In agreement with this +15 rule, Werner et al. [14] show that changing the position of bulged nucleotides in the lower stem of pri-mir172a changes the position of the lower DCL1 cleavage site. The +15 mechanism may be important in the evolution of MIRNAs. Transcripts of evolutionarily young MIRNA genes derived from inverted gene duplication have long hairpins [11, 16]. These transcripts are processed by DCL1 or DCL4 into heterogeneous small RNAs whose ends are roughly in 21 nt phase with the initial DCL cleavage site [11, 16]. Random mutations, particularly short indels, may introduce internal loops and therefore new initial DCL cleavage sites, changing the sequences of small RNAs liberated from the hairpins. However, the +15 rule is not sufficient to completely specify DCL1 processing accuracy, because aberrant cleavages in the pri-mir171a s extended lower hairpin occur at roughly equal frequency at distances of 14, 15, and 16 nt from the less-structured internal loop, whereas canonical cleavages occur at
4 Current Biology Vol 20 No 1 40 Figure 3. CBL Variants Are Accurately Processed at the Loop-Distal Cleavage Site (A) 5 0 positions and frequencies of the cleavage sites in primir167a. Identified sites are indicated by red arrowheads, and the number of times each was observed is given by the adjacent number. Red nucleotides represent mature microrna (mirna), and blue nucleotides represent mirna*. (B) As in (A) for pri-mir171a variants. Magenta and green bars represent the positions of probes in (C) that are complementary to the nucleotides in the 5 0 arm of the hairpin. (C) In vitro processing of pri-mir171a by a recombinant DCL1-HYL1-SE complex. RNA substrate has the same sequence as the transgenic pri-mir171a-wt shown in (B) except for two extra Gs at the 5 0 end introduced by the T7 promoter. The panels are of the same blot hybridized with different oligonucleotide probes to detect the 5 0 fragment released by the initial DCL1 cleavage (magenta arrowhead) and the artificial mirna (green arrowhead) liberated by two DCL1 cleavages in the extended hairpin. precisely +15 and +16 nt in pri-mir171a and pri-mir167a, respectively. Moreover, computational analysis indicates that there are accurately processed pri-mirnas that do not conform to the +15 rule. Supplemental Information Supplemental Information includes Supplemental Experimental Procedures, four figures, and three tables and can be found with this article online at doi: /j.cub Figure 4. Unpaired Regions in the Lower Stem Are Frequently Located w15 Nucleotides from the mirna: mirna* Duplex Frequencies of primary mirnas in which the hairpin ends or a big loop (R3 unpaired nt within the stem) begins at the indicated hairpin positions either in the 5 0 (top) or 3 0 (bottom) arm of the hairpin. Blue, pink, and yellow bars represent helix ends and big loops in the lower, middle, and upper stems, respectively. p values are estimated by 100,000 simulated random distributions of features in each region.
5 Plant mirna Biogenesis 41 Acknowledgments We thank H. Ma, T. Nakagawa, and R. Singer for sharing seeds and plasmids. We also thank D. Weigel, J. Carrington, J. Palatnik, and S. Werner for sharing data prior to publication. This study was supported by award MCB to N.V.F. and by award MCB to M.J.A. from the United States National Science Foundation. Received: July 19, 2009 Revised: October 3, 2009 Accepted: October 29, 2009 Published online: December 14, 2009 Note Added in Proof While this manuscript was in press, a paper by Cuperus et al., discussing the importance of the loop-distal region in the MIR390 hairpin for mirna biogenesis, was accepted for publication. The citation details are as follows: Cuperus, J.T., Montgomery, T.A., Fahlgren, N., Burke, R.T., Townsend, T., Sullivan, C.M., and Carrington, J.C. (2009). Identification of MIR390a precursor processing-defective mutants in Arabidopsis by direct genome sequencing. Proc. Natl. Acad. Sci. USA 106. Published online December 14, /pnas References 1. Lee, Y., Jeon, K., Lee, J.T., Kim, S., and Kim, V.N. (2002). MicroRNA maturation: Stepwise processing and subcellular localization. EMBO J. 21, Lee, R.C., Feinbaum, R.L., and Ambros, V. (1993). The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, Han, J., Lee, Y., Yeom, K.H., Nam, J.W., Heo, I., Rhee, J.K., Sohn, S.Y., Cho, Y., Zhang, B.T., and Kim, V.N. (2006). Molecular basis for the recognition of primary micrornas by the Drosha-DGCR8 complex. Cell 125, Peragine, A., Yoshikawa, M., Wu, G., Albrecht, H.L., and Poethig, R.S. (2004). SGS3 and SGS2/SDE1/RDR6 are required for juvenile development and the production of trans-acting sirnas in Arabidopsis. Genes Dev. 18, Ru, P., Xu, L., Ma, H., and Huang, H. (2006). Plant fertility defects induced by the enhanced expression of microrna167. Cell Res. 16, Wu, M.F., Tian, Q., and Reed, J.W. (2006). Arabidopsis microrna167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development 133, Mi, S., Cai, T., Hu, Y., Chen, Y., Hodges, E., Ni, F., Wu, L., Li, S., Zhou, H., Long, C., et al. (2008). Sorting of small RNAs into Arabidopsis argonaute complexes is directed by the 5 0 terminal nucleotide. Cell 133, Jinek, M., and Doudna, J.A. (2009). A three-dimensional view of the molecular machinery of RNA interference. Nature 457, Kurihara, Y., and Watanabe, Y. (2004). Arabidopsis micro-rna biogenesis through Dicer-like 1 protein functions. Proc. Natl. Acad. Sci. USA 101, Park, W., Li, J., Song, R., Messing, J., and Chen, X. (2002). CARPEL FACTORY, a Dicer homolog, and HEN1, a novel protein, act in microrna metabolism in Arabidopsis thaliana. Curr. Biol. 12, Rajagopalan, R., Vaucheret, H., Trejo, J., and Bartel, D.P. (2006). A diverse and evolutionarily fluid set of micrornas in Arabidopsis thaliana. Genes Dev. 20, Pouch-Pélissier, M.N., Pélissier, T., Elmayan, T., Vaucheret, H., Boko, D., Jantsch, M.F., and Deragon, J.M. (2008). SINE RNA induces severe developmental defects in Arabidopsis thaliana and interacts with HYL1 (DRB1), a key member of the DCL1 complex. PLoS Genet. 4, e Griffiths-Jones, S., Saini, H.K., van Dongen, S., and Enright, A.J. (2008). mirbase: Tools for microrna genomics. Nucleic Acids Res. 36(database issue), D154 D Werner, S., Wollmann, H., Schneeberger, K., and Weigel, D. (2010). Structure determinants for accurate processing of mir172a in Arabidopsis thaliana. Curr. Biol. 20, this issue, Mateos, J.L., Bologna, N.G., Chorostecki, U., and Palatnik, J.F. (2010). Identification of microrna processing determinants by random mutagenesis of Arabidopsis MIR172a precursor. Curr. Biol. 20, this issue, Allen, E., Xie, Z., Gustafson, A.M., Sung, G.H., Spatafora, J.W., and Carrington, J.C. (2004). Evolution of microrna genes by inverted duplication of target gene sequences in Arabidopsis thaliana. Nat. Genet. 36,
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
Analytical Study of Hexapod mirnas using Phylogenetic Methods
 Analytical Study of Hexapod mirnas using Phylogenetic Methods A.K. Mishra and H.Chandrasekharan Unit of Simulation & Informatics, Indian Agricultural Research Institute, New Delhi, India akmishra@iari.res.in,
Analytical Study of Hexapod mirnas using Phylogenetic Methods A.K. Mishra and H.Chandrasekharan Unit of Simulation & Informatics, Indian Agricultural Research Institute, New Delhi, India akmishra@iari.res.in,
mirnaselect pep-mir Cloning and Expression Vector
 Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
Outline. interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work?
 Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Micro RNAs: potentielle Biomarker für das. Blutspenderscreening
 Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Outline. MicroRNA Bioinformatics. microrna biogenesis. short non-coding RNAs not considered in this lecture. ! Introduction
 Outline MicroRNA Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology (CMB) Karolinska Institutet! Introduction! microrna target site prediction! Useful resources 2 short non-coding RNAs
Outline MicroRNA Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology (CMB) Karolinska Institutet! Introduction! microrna target site prediction! Useful resources 2 short non-coding RNAs
Biogenesis, Size and Function of Small RNAs
 srnas of Plants Small, Non-coding RNAs of Plants Regulatory RNAs that act through gene silencing Two classes of small RNAs (srnas) o microrna (mirnas) Encoded by genes in the genome o small interfering
srnas of Plants Small, Non-coding RNAs of Plants Regulatory RNAs that act through gene silencing Two classes of small RNAs (srnas) o microrna (mirnas) Encoded by genes in the genome o small interfering
PART 3.3: MicroRNA and Cancer
 BIBM 2010 Tutorial: Epigenomics and Cancer PART 3.3: MicroRNA and Cancer Dec 18, 2010 Sun Kim at Indiana University Outline of Part 3.3 Background on microrna Role of microrna in cancer MicroRNA pathway
BIBM 2010 Tutorial: Epigenomics and Cancer PART 3.3: MicroRNA and Cancer Dec 18, 2010 Sun Kim at Indiana University Outline of Part 3.3 Background on microrna Role of microrna in cancer MicroRNA pathway
Functional RNAs; RNA catalysts, mirna,
 Functional RNAs; RNA catalysts, mirna, srna, RNAi... RNAs have many functions rrna (ribosomal RNA) trna (transfer RNA) mrna (Messenger RNA) snrna (including snorna) ) (Small nuclear RNA- splicing) Other
Functional RNAs; RNA catalysts, mirna, srna, RNAi... RNAs have many functions rrna (ribosomal RNA) trna (transfer RNA) mrna (Messenger RNA) snrna (including snorna) ) (Small nuclear RNA- splicing) Other
MicroRNA formation. 4th International Symposium on Non-Surgical Contraceptive Methods of Pet Population Control
 MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
Plant RNAi mechanisms: lessons from silent transgenes. Institut Jean-Pierre Bourgin, INRA Versailles
 Plant RNAi mechanisms: lessons from silent transgenes Institut Jean-Pierre Bourgin, INRA Versailles Plants encode two types of small RNAs: mirna and sirna MIR genes endoir NAT pairs TAS and PolIV loci
Plant RNAi mechanisms: lessons from silent transgenes Institut Jean-Pierre Bourgin, INRA Versailles Plants encode two types of small RNAs: mirna and sirna MIR genes endoir NAT pairs TAS and PolIV loci
V22: involvement of micrornas in GRNs
 What are micrornas? V22: involvement of micrornas in GRNs How can one identify micrornas? What is the function of micrornas? Elisa Izaurralde, MPI Tübingen Huntzinger, Izaurralde, Nat. Rev. Genet. 12,
What are micrornas? V22: involvement of micrornas in GRNs How can one identify micrornas? What is the function of micrornas? Elisa Izaurralde, MPI Tübingen Huntzinger, Izaurralde, Nat. Rev. Genet. 12,
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation
 OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
SoMART: a web server for plant mirna, tasirna and target gene analysis
 The Plant Journal (2012) doi: 10.1111/j.1365-313X.2012.04922.x TECHNICAL ADVANCE SoMART: a web server for plant mirna, tasirna and target gene analysis Feng Li 1,2,*, Ryan Orban 1,2, and Barbara Baker
The Plant Journal (2012) doi: 10.1111/j.1365-313X.2012.04922.x TECHNICAL ADVANCE SoMART: a web server for plant mirna, tasirna and target gene analysis Feng Li 1,2,*, Ryan Orban 1,2, and Barbara Baker
Functional and Biomedical Aspects of Genome Research
 Functional and Biomedical Aspects of Genome Research 20 11 35 Vorlesung SS 04 Bartsch, Jockusch & Schmitt-John Mi. 9:15-10:00, in W7-135 13 Functional RNAs Thomas Schmitt-John micro RNAs small interfering
Functional and Biomedical Aspects of Genome Research 20 11 35 Vorlesung SS 04 Bartsch, Jockusch & Schmitt-John Mi. 9:15-10:00, in W7-135 13 Functional RNAs Thomas Schmitt-John micro RNAs small interfering
Arabidopsis lyrata Small RNAs: Transient MIRNA and Small Interfering RNA Loci within the Arabidopsis Genus W OA
 The Plant Cell, Vol. 22: 1090 1103, April 2010, www.plantcell.org ã 2010 American Society of Plant Biologists Arabidopsis lyrata Small RNAs: Transient MIRNA and Small Interfering RNA Loci within the Arabidopsis
The Plant Cell, Vol. 22: 1090 1103, April 2010, www.plantcell.org ã 2010 American Society of Plant Biologists Arabidopsis lyrata Small RNAs: Transient MIRNA and Small Interfering RNA Loci within the Arabidopsis
CCR Biology - Chapter 9 Practice Test - Summer 2012
 Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Seminars in Cell & Developmental Biology
 Seminars in Cell & Developmental Biology 21 (2010) 790 797 Contents lists available at ScienceDirect Seminars in Cell & Developmental Biology journal homepage: www.elsevier.com/locate/semcdb Review Expression
Seminars in Cell & Developmental Biology 21 (2010) 790 797 Contents lists available at ScienceDirect Seminars in Cell & Developmental Biology journal homepage: www.elsevier.com/locate/semcdb Review Expression
岑 祥 股 份 有 限 公 司 技 術 專 員 費 軫 尹 20100803
 技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
Considerable evidence now indicates that small noncoding
 MicroRNAs and small interfering RNAs can inhibit mrna expression by similar mechanisms Yan Zeng*, Rui Yi, and Bryan R. Cullen* *Howard Hughes Medical Institute and Department of Molecular Genetics and
MicroRNAs and small interfering RNAs can inhibit mrna expression by similar mechanisms Yan Zeng*, Rui Yi, and Bryan R. Cullen* *Howard Hughes Medical Institute and Department of Molecular Genetics and
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals
 Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Dicer Substrate RNAi Design
 INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
Prediction, validation and functional analysis of mirna targets in Arabidopsis thaliana
 Prediction, validation and functional analysis of mirna targets in Arabidopsis thaliana Dissertation to obtain the academic title Doctor of Natural Sciences (Dr. rer. nat.) at the Faculty of Biology in
Prediction, validation and functional analysis of mirna targets in Arabidopsis thaliana Dissertation to obtain the academic title Doctor of Natural Sciences (Dr. rer. nat.) at the Faculty of Biology in
RNAi Shooting the Messenger!
 RNAi Shooting the Messenger! Bronya Keats, Ph.D. Department of Genetics Louisiana State University Health Sciences Center New Orleans Email: bkeats@lsuhsc.edu RNA interference (RNAi) A mechanism by which
RNAi Shooting the Messenger! Bronya Keats, Ph.D. Department of Genetics Louisiana State University Health Sciences Center New Orleans Email: bkeats@lsuhsc.edu RNA interference (RNAi) A mechanism by which
MicroRNAs: something important between the genes Allison C Mallory 1 and Hervé Vaucheret 2
 MicroRNAs: something important between the genes Allison C Mallory 1 and Hervé Vaucheret 2 Non-coding small endogenous RNAs, of 21 24 nucleotides in length, have recently emerged as important regulators
MicroRNAs: something important between the genes Allison C Mallory 1 and Hervé Vaucheret 2 Non-coding small endogenous RNAs, of 21 24 nucleotides in length, have recently emerged as important regulators
Evolution and Functional Diversification of MIRNA Genes
 This article is a Plant Cell Advance Online Publication. The date of its first appearance online is the official date of publication. The article has been edited and the authors have corrected proofs,
This article is a Plant Cell Advance Online Publication. The date of its first appearance online is the official date of publication. The article has been edited and the authors have corrected proofs,
MatureBayes: A Probabilistic Algorithm for Identifying the Mature mirna within Novel Precursors
 MatureBayes: A Probabilistic Algorithm for Identifying the Mature mirna within Novel Precursors Katerina Gkirtzou 1,2, Ioannis Tsamardinos 1,2, Panagiotis Tsakalides 1,2, Panayiota Poirazi 3 * 1 Computer
MatureBayes: A Probabilistic Algorithm for Identifying the Mature mirna within Novel Precursors Katerina Gkirtzou 1,2, Ioannis Tsamardinos 1,2, Panagiotis Tsakalides 1,2, Panayiota Poirazi 3 * 1 Computer
scientificreport Enhanced microrna accumulation through stemloop-adjacent introns scientific report
 scientificreport Enhanced microrna accumulation through stemloop-adjacent introns Rebecca Schwab,2,4, Corinna Speth 2, Sascha Laubinger 2 & Olivier Voinnet,3+ Institut de Biologie Moléculaire des Plantes,
scientificreport Enhanced microrna accumulation through stemloop-adjacent introns Rebecca Schwab,2,4, Corinna Speth 2, Sascha Laubinger 2 & Olivier Voinnet,3+ Institut de Biologie Moléculaire des Plantes,
Comparative epigenomic analysis between model plant and crop
 The 19 th Crucifer Genetics Workshop and Brassica 2014 Genetic Improvement of Brassicaceae Crops in the Era of Genomics Comparative epigenomic analysis between model plant and crop Xiaofeng Cao Institute
The 19 th Crucifer Genetics Workshop and Brassica 2014 Genetic Improvement of Brassicaceae Crops in the Era of Genomics Comparative epigenomic analysis between model plant and crop Xiaofeng Cao Institute
mirnaselect pegp-mir Cloning and Expression Vector
 Product Data Sheet mirnaselect pegp-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-GP-C STORAGE: -80ºC QUANTITY: 100 µl of bacterial glycerol stock Components 1. mirnaselect pegp-mir Cloning
Product Data Sheet mirnaselect pegp-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-GP-C STORAGE: -80ºC QUANTITY: 100 µl of bacterial glycerol stock Components 1. mirnaselect pegp-mir Cloning
Site-Directed Nucleases and Cisgenesis Maria Fedorova, Ph.D.
 Site-Directed Nucleases and Cisgenesis Maria Fedorova, Ph.D. Regulatory Strategy Lead Enabling Technologies DuPont-Pioneer, USA 1 New Plant Breeding Techniques 2007 New Techniques Working Group established
Site-Directed Nucleases and Cisgenesis Maria Fedorova, Ph.D. Regulatory Strategy Lead Enabling Technologies DuPont-Pioneer, USA 1 New Plant Breeding Techniques 2007 New Techniques Working Group established
Profiling of non-coding RNA classes Gunter Meister
 Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Mirnas andaminas - Interrelated Physiology of Biogenesis and Networking
 The Open Applied Informatics Journal, 2008, 2, 9-13 9 In Silico Evidence of the Relationship Between mirnas and sirnas L. Montanucci 1, P. Fariselli *,1, P.L. Martelli 1, I. Rossi 1,2 and R. Casadio 1
The Open Applied Informatics Journal, 2008, 2, 9-13 9 In Silico Evidence of the Relationship Between mirnas and sirnas L. Montanucci 1, P. Fariselli *,1, P.L. Martelli 1, I. Rossi 1,2 and R. Casadio 1
Five-year relative survival rates. Cancer. Age-adjusted cancer death rates. Proteomic Technologies for Cancer Biomarker Discovery 2010/3/22
 Cancer Five-year relative survival rates Basal lamina Underlyig tissue Normal tissue Carcinoma Invasive carcinoma 1 http://www.cancer.org/docroot/home/index.asp 2 Proteomic Technologies for Cancer Biomarker
Cancer Five-year relative survival rates Basal lamina Underlyig tissue Normal tissue Carcinoma Invasive carcinoma 1 http://www.cancer.org/docroot/home/index.asp 2 Proteomic Technologies for Cancer Biomarker
CRAC: An integrated approach to analyse RNA-seq reads Additional File 3 Results on simulated RNA-seq data.
 : An integrated approach to analyse RNA-seq reads Additional File 3 Results on simulated RNA-seq data. Nicolas Philippe and Mikael Salson and Thérèse Commes and Eric Rivals February 13, 2013 1 Results
: An integrated approach to analyse RNA-seq reads Additional File 3 Results on simulated RNA-seq data. Nicolas Philippe and Mikael Salson and Thérèse Commes and Eric Rivals February 13, 2013 1 Results
micrornas Contents Introduction
 micrornas Contents Introduction... 1 Structure and Function of mirnas... 2 Plant mirnas... 4 mirna Biosynthesis... 5 Evolution of mirnas... 6 Conclusions... 9 References and Resources... 9 Introduction
micrornas Contents Introduction... 1 Structure and Function of mirnas... 2 Plant mirnas... 4 mirna Biosynthesis... 5 Evolution of mirnas... 6 Conclusions... 9 References and Resources... 9 Introduction
Mir-X mirna First-Strand Synthesis Kit User Manual
 User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
RNA Structure and folding
 RNA Structure and folding Overview: The main functional biomolecules in cells are polymers DNA, RNA and proteins For RNA and Proteins, the specific sequence of the polymer dictates its final structure
RNA Structure and folding Overview: The main functional biomolecules in cells are polymers DNA, RNA and proteins For RNA and Proteins, the specific sequence of the polymer dictates its final structure
AmiRzyn: PERL Centered Artificial MicroRNA Designing Aid
 Abstract International Research Journal of Biological Sciences ISSN 2278-3202 AmiRzyn: PERL Centered Artificial MicroRNA Designing Aid Joseph Baby 1, Nair Vrundha M 1,2 and Chandy Susheel G. 2 1 Interdisciplinary
Abstract International Research Journal of Biological Sciences ISSN 2278-3202 AmiRzyn: PERL Centered Artificial MicroRNA Designing Aid Joseph Baby 1, Nair Vrundha M 1,2 and Chandy Susheel G. 2 1 Interdisciplinary
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE Q5B
 INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
Lecture 3: Mutations
 Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
RNA Viruses. A Practical Approac h. Alan J. Cann
 RNA Viruses A Practical Approac h Alan J. Cann List of protocols page xiii Abbreviations xvii Investigation of RNA virus genome structure 1 A j. Easton, A.C. Marriott and C.R. Pringl e 1 Introduction-the
RNA Viruses A Practical Approac h Alan J. Cann List of protocols page xiii Abbreviations xvii Investigation of RNA virus genome structure 1 A j. Easton, A.C. Marriott and C.R. Pringl e 1 Introduction-the
Biological Sciences Initiative. Human Genome
 Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Center for Genome Research and Biocomputing, Department of Botany and Plant Pathology, Oregon State University, Corvallis, Oregon 97331 b
 The Plant Cell, Vol. 19: 926 942, March 2007, www.plantcell.org ª 2007 American Society of Plant Biologists Genome-Wide Analysis of the RNA-DEPENDENT RNA POLYMERASE6/DICER-LIKE4 Pathway in Arabidopsis
The Plant Cell, Vol. 19: 926 942, March 2007, www.plantcell.org ª 2007 American Society of Plant Biologists Genome-Wide Analysis of the RNA-DEPENDENT RNA POLYMERASE6/DICER-LIKE4 Pathway in Arabidopsis
Chapter 6 DNA Replication
 Chapter 6 DNA Replication Each strand of the DNA double helix contains a sequence of nucleotides that is exactly complementary to the nucleotide sequence of its partner strand. Each strand can therefore
Chapter 6 DNA Replication Each strand of the DNA double helix contains a sequence of nucleotides that is exactly complementary to the nucleotide sequence of its partner strand. Each strand can therefore
missing LINKS: mirnas and plant development Christine Hunter and R Scott Poethig
 372 missing LINKS: mirnas and plant development Christine Hunter and R Scott Poethig The discovery of hundreds of plant micro RNAs (mirnas) has triggered much speculation about their potential roles in
372 missing LINKS: mirnas and plant development Christine Hunter and R Scott Poethig The discovery of hundreds of plant micro RNAs (mirnas) has triggered much speculation about their potential roles in
Arabidopsis. A Practical Approach. Edited by ZOE A. WILSON Plant Science Division, School of Biological Sciences, University of Nottingham
 Arabidopsis A Practical Approach Edited by ZOE A. WILSON Plant Science Division, School of Biological Sciences, University of Nottingham OXPORD UNIVERSITY PRESS List of Contributors Abbreviations xv xvu
Arabidopsis A Practical Approach Edited by ZOE A. WILSON Plant Science Division, School of Biological Sciences, University of Nottingham OXPORD UNIVERSITY PRESS List of Contributors Abbreviations xv xvu
RETRIEVING SEQUENCE INFORMATION. Nucleotide sequence databases. Database search. Sequence alignment and comparison
 RETRIEVING SEQUENCE INFORMATION Nucleotide sequence databases Database search Sequence alignment and comparison Biological sequence databases Originally just a storage place for sequences. Currently the
RETRIEVING SEQUENCE INFORMATION Nucleotide sequence databases Database search Sequence alignment and comparison Biological sequence databases Originally just a storage place for sequences. Currently the
RAPAd mirna Adenoviral Expression System Catalog #: VPK-253
 DATENBLATT RAPAd mirna Adenoviral Expression System Catalog #: VPK-253 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction MicroRNAs (mirnas) are 18 24 nucleotide RNA molecules that
DATENBLATT RAPAd mirna Adenoviral Expression System Catalog #: VPK-253 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction MicroRNAs (mirnas) are 18 24 nucleotide RNA molecules that
Trasposable elements: P elements
 Trasposable elements: P elements In 1938 Marcus Rhodes provided the first genetic description of an unstable mutation, an allele of a gene required for the production of pigment in maize. This instability
Trasposable elements: P elements In 1938 Marcus Rhodes provided the first genetic description of an unstable mutation, an allele of a gene required for the production of pigment in maize. This instability
Essentials of Real Time PCR. About Sequence Detection Chemistries
 Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
GeneCopoeia Genome Editing Tools for Safe Harbor Integration in. Mice and Humans. Ed Davis, Liuqing Qian, Ruiqing li, Junsheng Zhou, and Jinkuo Zhang
 G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
Recombinant DNA and Biotechnology
 Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Gene mutation and molecular medicine Chapter 15
 Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
Design high specificity CRISPR-Cas9 grnas: principles and tools. Heidi Huang, PhD
 Design high specificity CRISPR-Cas9 grnas: principles and tools Heidi Huang, PhD Webinar Agenda 1 2 3 4 Introduction of CRISPR-Cas9 grna Design Resources and Services Q&A 2 What is CRISPR? CRISPR Clustered
Design high specificity CRISPR-Cas9 grnas: principles and tools Heidi Huang, PhD Webinar Agenda 1 2 3 4 Introduction of CRISPR-Cas9 grna Design Resources and Services Q&A 2 What is CRISPR? CRISPR Clustered
Supplementary Figure 1.
 Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
The RNA strategy. RNA as a tool and target in human disease diagnosis and therapy.
 The RNA strategy RNA as a tool and target in human disease diagnosis and therapy. The Laboratory of RNA Biology and Biotechnology at the Centre for Integrative Biology (CIBIO) of the University of Trento,
The RNA strategy RNA as a tool and target in human disease diagnosis and therapy. The Laboratory of RNA Biology and Biotechnology at the Centre for Integrative Biology (CIBIO) of the University of Trento,
!!!!!!!!!!!!!!!!!!!!!!!!!!
 Figure S7 cdl-1 3'UTR gld-1 binding site (4..10) let-7 binding site (326..346) Figure Legends Figure S1. gld-1 genetically interacts with nhl-2 and vig-1 during germline development. (A) nhl-2(ok818) enhances
Figure S7 cdl-1 3'UTR gld-1 binding site (4..10) let-7 binding site (326..346) Figure Legends Figure S1. gld-1 genetically interacts with nhl-2 and vig-1 during germline development. (A) nhl-2(ok818) enhances
sirna and mirna: an insight into RISCs
 Review TRENDS in Biochemical Sciences Vol.30 No.2 February 2005 sirna and mirna: an insight into s Guiliang Tang Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical
Review TRENDS in Biochemical Sciences Vol.30 No.2 February 2005 sirna and mirna: an insight into s Guiliang Tang Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical
THE ENZYMES. Department of Microbiology, Immunology, and Molecular Genetics, Molecular Biology Institute University of California
 VOLUME THIRTY TWO THE ENZYMES Eukaryotic RNases and their Partners in RNA Degradation and Biogenesis, Part B Edited by FENG GUO Department of Biological Chemistry, David Geffen School of Medicine, Molecular
VOLUME THIRTY TWO THE ENZYMES Eukaryotic RNases and their Partners in RNA Degradation and Biogenesis, Part B Edited by FENG GUO Department of Biological Chemistry, David Geffen School of Medicine, Molecular
Becker Muscular Dystrophy
 Muscular Dystrophy A Case Study of Positional Cloning Described by Benjamin Duchenne (1868) X-linked recessive disease causing severe muscular degeneration. 100 % penetrance X d Y affected male Frequency
Muscular Dystrophy A Case Study of Positional Cloning Described by Benjamin Duchenne (1868) X-linked recessive disease causing severe muscular degeneration. 100 % penetrance X d Y affected male Frequency
Specific Gene Silencing by Artificial MicroRNAs in Physcomitrella patens: An Alternative to Targeted Gene Knockouts 1[C][W][OA]
![Specific Gene Silencing by Artificial MicroRNAs in Physcomitrella patens: An Alternative to Targeted Gene Knockouts 1[C][W][OA] Specific Gene Silencing by Artificial MicroRNAs in Physcomitrella patens: An Alternative to Targeted Gene Knockouts 1[C][W][OA]](/thumbs/24/3567909.jpg) Breakthrough Technologies Specific Gene Silencing by Artificial MicroRNAs in Physcomitrella patens: An Alternative to Targeted Gene Knockouts 1[C][W][OA] Basel Khraiwesh, Stephan Ossowski, Detlef Weigel,
Breakthrough Technologies Specific Gene Silencing by Artificial MicroRNAs in Physcomitrella patens: An Alternative to Targeted Gene Knockouts 1[C][W][OA] Basel Khraiwesh, Stephan Ossowski, Detlef Weigel,
The Human Genome Project
 The Human Genome Project Brief History of the Human Genome Project Physical Chromosome Maps Genetic (or Linkage) Maps DNA Markers Sequencing and Annotating Genomic DNA What Have We learned from the HGP?
The Human Genome Project Brief History of the Human Genome Project Physical Chromosome Maps Genetic (or Linkage) Maps DNA Markers Sequencing and Annotating Genomic DNA What Have We learned from the HGP?
Forensic DNA Testing Terminology
 Forensic DNA Testing Terminology ABI 310 Genetic Analyzer a capillary electrophoresis instrument used by forensic DNA laboratories to separate short tandem repeat (STR) loci on the basis of their size.
Forensic DNA Testing Terminology ABI 310 Genetic Analyzer a capillary electrophoresis instrument used by forensic DNA laboratories to separate short tandem repeat (STR) loci on the basis of their size.
European Medicines Agency
 European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
Biosafety considerations of new dsrna molecules
 Biosafety considerations of new dsrna molecules GM-free Europe Conference 2015 May 6-8 Berlin, Germany Dr. Sarah Agapito Current GMOs Plant transformed to produce new protein A G C U G G C A A U G U C
Biosafety considerations of new dsrna molecules GM-free Europe Conference 2015 May 6-8 Berlin, Germany Dr. Sarah Agapito Current GMOs Plant transformed to produce new protein A G C U G G C A A U G U C
In silico evidence of the relationship between mirnas and sirnas
 In silico evidence of the relationship between mirnas and sirnas In silico evidence of the relationship between mirnas and sirnas Ludovica Montanucci 1, Piero Fariselli 1*, Pier Luigi Martelli 1, Ivan
In silico evidence of the relationship between mirnas and sirnas In silico evidence of the relationship between mirnas and sirnas Ludovica Montanucci 1, Piero Fariselli 1*, Pier Luigi Martelli 1, Ivan
Genetics Module B, Anchor 3
 Genetics Module B, Anchor 3 Key Concepts: - An individual s characteristics are determines by factors that are passed from one parental generation to the next. - During gamete formation, the alleles for
Genetics Module B, Anchor 3 Key Concepts: - An individual s characteristics are determines by factors that are passed from one parental generation to the next. - During gamete formation, the alleles for
Sickle cell anemia: Altered beta chain Single AA change (#6 Glu to Val) Consequence: Protein polymerizes Change in RBC shape ---> phenotypes
 Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
Biotechnology: DNA Technology & Genomics
 Chapter 20. Biotechnology: DNA Technology & Genomics 2003-2004 The BIG Questions How can we use our knowledge of DNA to: diagnose disease or defect? cure disease or defect? change/improve organisms? What
Chapter 20. Biotechnology: DNA Technology & Genomics 2003-2004 The BIG Questions How can we use our knowledge of DNA to: diagnose disease or defect? cure disease or defect? change/improve organisms? What
TOOLS sirna and mirna. User guide
 TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
RNA Silencing, mechanism and applications
 RA Silencing, mechanism and applications The Central Dogma of Molecular Biology There is life outside proteincoding genes There are approximately 70.000 gene predictions for the Human genome But the estimated
RA Silencing, mechanism and applications The Central Dogma of Molecular Biology There is life outside proteincoding genes There are approximately 70.000 gene predictions for the Human genome But the estimated
MicroRNAs and Their Regulatory Roles in Plants
 Annu. Rev. lant Biol. 2006. 57:19 53 The Annual Review of lant Biology is online at plant.annualreviews.org doi: 10.1146/ annurev.arplant.57.032905.105218 opyright c 2006 by Annual Reviews. All rights
Annu. Rev. lant Biol. 2006. 57:19 53 The Annual Review of lant Biology is online at plant.annualreviews.org doi: 10.1146/ annurev.arplant.57.032905.105218 opyright c 2006 by Annual Reviews. All rights
Name Class Date. Figure 13 1. 2. Which nucleotide in Figure 13 1 indicates the nucleic acid above is RNA? a. uracil c. cytosine b. guanine d.
 13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13.4 Gene Regulation and Expression
 13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
RNAi History, Mechanism and Application
 DRUG DISCOVERY AND DEVELOPMENT RNAi History, Mechanism and Application Shuo Gu The History of RNAi About the author: Dr. Shuo Gu is currently a Postdoctoral Scholar at Stanford University, School of Medicine.
DRUG DISCOVERY AND DEVELOPMENT RNAi History, Mechanism and Application Shuo Gu The History of RNAi About the author: Dr. Shuo Gu is currently a Postdoctoral Scholar at Stanford University, School of Medicine.
EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES
 C GUGAAG EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES MicroRNA-adapted shrna (shrna mir ) for increased, specific and consistent knockdown. MicroRNA PROCESSING PATHWAY UTILIZED FOR shrna mir Developed
C GUGAAG EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES MicroRNA-adapted shrna (shrna mir ) for increased, specific and consistent knockdown. MicroRNA PROCESSING PATHWAY UTILIZED FOR shrna mir Developed
Genetomic Promototypes
 Genetomic Promototypes Mirkó Palla and Dana Pe er Department of Mechanical Engineering Clarkson University Potsdam, New York and Department of Genetics Harvard Medical School 77 Avenue Louis Pasteur Boston,
Genetomic Promototypes Mirkó Palla and Dana Pe er Department of Mechanical Engineering Clarkson University Potsdam, New York and Department of Genetics Harvard Medical School 77 Avenue Louis Pasteur Boston,
DNA Fingerprinting. Unless they are identical twins, individuals have unique DNA
 DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
Functional diversity of mirna in plants
 Plant Science 172 (2007) 423 432 Review Functional diversity of mirna in plants Tongwen Yang a,b,1, Lingui Xue a, Lizhe An a,c, * www.elsevier.com/locate/plantsci a Key Laboratory of Arid and Grassland
Plant Science 172 (2007) 423 432 Review Functional diversity of mirna in plants Tongwen Yang a,b,1, Lingui Xue a, Lizhe An a,c, * www.elsevier.com/locate/plantsci a Key Laboratory of Arid and Grassland
www.mpiz-koeln.mpg.de Abt. Entwicklungsbiologie de Pflanzen
 WEB ADDRESS: www.mpiz-koeln.mpg.de Forschung Abt. Entwicklungsbiologie de Pflanzen George Coupland How is the transition from vegetative growth to flowering controlled? - How is it regulated by environmental
WEB ADDRESS: www.mpiz-koeln.mpg.de Forschung Abt. Entwicklungsbiologie de Pflanzen George Coupland How is the transition from vegetative growth to flowering controlled? - How is it regulated by environmental
Design of conditional gene targeting vectors - a recombineering approach
 Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
Control of Gene Expression
 Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
mirnapath: a database of mirnas, target genes and metabolic pathways
 mirnapath: a database of mirnas, target genes and metabolic pathways A.O. Chiromatzo 1, T.Y.K. Oliveira 1, G. Pereira 1, A.Y. Costa 2, C.A.E. Montesco 3, D.E. Gras 1, F. Yosetake 4, J.B. Vilar 5, M. Cervato
mirnapath: a database of mirnas, target genes and metabolic pathways A.O. Chiromatzo 1, T.Y.K. Oliveira 1, G. Pereira 1, A.Y. Costa 2, C.A.E. Montesco 3, D.E. Gras 1, F. Yosetake 4, J.B. Vilar 5, M. Cervato
Lezioni Dipartimento di Oncologia Farmacologia Molecolare. RNA interference. Giovanna Damia 29 maggio 2006
 Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Innovations in Molecular Epidemiology
 Innovations in Molecular Epidemiology Molecular Epidemiology Measure current rates of active transmission Determine whether recurrent tuberculosis is attributable to exogenous reinfection Determine whether
Innovations in Molecular Epidemiology Molecular Epidemiology Measure current rates of active transmission Determine whether recurrent tuberculosis is attributable to exogenous reinfection Determine whether
Unique expression, processing regulation, and regulatory network of peach (Prunus persica) mirnas
 Zhu et al. BMC Plant Biology 2012, 12:149 RESEARCH ARTICLE Open Access Unique expression, processing regulation, and regulatory network of peach (Prunus persica) mirnas Hong Zhu 1,3, Rui Xia 1,2,3, Bingyu
Zhu et al. BMC Plant Biology 2012, 12:149 RESEARCH ARTICLE Open Access Unique expression, processing regulation, and regulatory network of peach (Prunus persica) mirnas Hong Zhu 1,3, Rui Xia 1,2,3, Bingyu
Phased, Secondary, Small Interfering RNAs in Posttranscriptional Regulatory Networks OPEN
 The Plant Cell, Vol. 25: 2400 2415, July 2013, www.plantcell.org ã 2013 American Society of Plant Biologists. All rights reserved. REVIEW Phased, Secondary, Small Interfering RNAs in Posttranscriptional
The Plant Cell, Vol. 25: 2400 2415, July 2013, www.plantcell.org ã 2013 American Society of Plant Biologists. All rights reserved. REVIEW Phased, Secondary, Small Interfering RNAs in Posttranscriptional
"Supplemental Research Data" http://www.genome.org/cgi/content/full/gr.6539108/dc1
 Xenopus microrna genes are predominantly located within introns and are differentially expressed in adult frog tissues via post-transcriptional regulation Guo-Qing Tang and E. Stuart Maxwell Genome Res.
Xenopus microrna genes are predominantly located within introns and are differentially expressed in adult frog tissues via post-transcriptional regulation Guo-Qing Tang and E. Stuart Maxwell Genome Res.
Genetics Test Biology I
 Genetics Test Biology I Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Avery s experiments showed that bacteria are transformed by a. RNA. c. proteins.
Genetics Test Biology I Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Avery s experiments showed that bacteria are transformed by a. RNA. c. proteins.
MicroRNAs: SMALL RNAs WITH A BIG ROLE IN GENE REGULATION
 MicroRNAs: SMALL RNAs WITH A BIG ROLE IN GENE REGULATION Lin He and Gregory J. Hannon MicroRNAs are a family of small, non-coding RNAs that regulate gene expression in a sequence-specific manner. The two
MicroRNAs: SMALL RNAs WITH A BIG ROLE IN GENE REGULATION Lin He and Gregory J. Hannon MicroRNAs are a family of small, non-coding RNAs that regulate gene expression in a sequence-specific manner. The two
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
Basic Concepts of DNA, Proteins, Genes and Genomes
 Basic Concepts of DNA, Proteins, Genes and Genomes Kun-Mao Chao 1,2,3 1 Graduate Institute of Biomedical Electronics and Bioinformatics 2 Department of Computer Science and Information Engineering 3 Graduate
Basic Concepts of DNA, Proteins, Genes and Genomes Kun-Mao Chao 1,2,3 1 Graduate Institute of Biomedical Electronics and Bioinformatics 2 Department of Computer Science and Information Engineering 3 Graduate
Genetics Lecture Notes 7.03 2005. Lectures 1 2
 Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
RNA and Protein Synthesis
 Name lass Date RN and Protein Synthesis Information and Heredity Q: How does information fl ow from DN to RN to direct the synthesis of proteins? 13.1 What is RN? WHT I KNOW SMPLE NSWER: RN is a nucleic
Name lass Date RN and Protein Synthesis Information and Heredity Q: How does information fl ow from DN to RN to direct the synthesis of proteins? 13.1 What is RN? WHT I KNOW SMPLE NSWER: RN is a nucleic
SUGAR BEET (Beta vulgaris L.) PROMOTERS FOR DIRECTED TISSUE- SPECIFIC ROOT TRANSCRIPTION
 SUGAR BEET (Beta vulgaris L.) PROMOTERS FOR DIRECTED TISSUE- SPECIFIC ROOT TRANSCRIPTION Senthilkumar Padmanaban, Haiyan Li, David P. Puthoff and Ann C. Smigocki USDA-ARS Molecular Plant Pathology Laboratory,
SUGAR BEET (Beta vulgaris L.) PROMOTERS FOR DIRECTED TISSUE- SPECIFIC ROOT TRANSCRIPTION Senthilkumar Padmanaban, Haiyan Li, David P. Puthoff and Ann C. Smigocki USDA-ARS Molecular Plant Pathology Laboratory,
LightSwitch Luciferase Assay System
 Constructs, Assays & Services Assay System Promoter GoClones Synthetic Response Elements Pathway Cell Lines 3 UTR GoClones mirna Mimics & Inhibitors Synthetic mirna Targets Transfection Reagents Assay
Constructs, Assays & Services Assay System Promoter GoClones Synthetic Response Elements Pathway Cell Lines 3 UTR GoClones mirna Mimics & Inhibitors Synthetic mirna Targets Transfection Reagents Assay
2. The number of different kinds of nucleotides present in any DNA molecule is A) four B) six C) two D) three
 Chem 121 Chapter 22. Nucleic Acids 1. Any given nucleotide in a nucleic acid contains A) two bases and a sugar. B) one sugar, two bases and one phosphate. C) two sugars and one phosphate. D) one sugar,
Chem 121 Chapter 22. Nucleic Acids 1. Any given nucleotide in a nucleic acid contains A) two bases and a sugar. B) one sugar, two bases and one phosphate. C) two sugars and one phosphate. D) one sugar,
micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved
 microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
Basic Concepts Recombinant DNA Use with Chapter 13, Section 13.2
 Name Date lass Master 19 Basic oncepts Recombinant DN Use with hapter, Section.2 Formation of Recombinant DN ut leavage Splicing opyright lencoe/mcraw-hill, a division of he Mcraw-Hill ompanies, Inc. Bacterial
Name Date lass Master 19 Basic oncepts Recombinant DN Use with hapter, Section.2 Formation of Recombinant DN ut leavage Splicing opyright lencoe/mcraw-hill, a division of he Mcraw-Hill ompanies, Inc. Bacterial
