FiberApp: An Open-Source Software for Tracking and Analyzing Polymers, Filaments, Biomacromolecules, and Fibrous Objects
|
|
|
- Aubrie Edwards
- 10 years ago
- Views:
Transcription
1 This is an open access article published under a Creative Commons Attribution (CC-BY) License, which permits unrestricted use, distribution and reproduction in any medium, provided the author and source are cited. pubs.acs.org/macromolecules FiberApp: An Open-Source Software for Tracking and Analyzing Polymers, Filaments, Biomacromolecules, and Fibrous Objects Ivan Usov and Raffaele Mezzenga* Department of Health Science & Technology, ETH Zurich, Schmelzbergstrasse 9, LFO E23, 8092 Zurich, Switzerland *S Supporting Information ABSTRACT: Biological semiflexible polymers and filaments such as collagen, fibronectin, actin, microtubules, coiled-coil proteins, DNA, sirna, amyloid fibrils, etc., are ubiquitous in nature. In biology, these systems have a direct relation to critical processes ranging from the movement of actin or assembly of viruses at cellular interfaces to the growth of amyloid plaques in neurodegenerative diseases. In technology and applied sciences, synthetic macromolecules or fibrous objects such as carbon nanotubes are involved in countless applications. Accessing their intrinsic properties at the single molecule level, such as their molecular conformations or intrinsic stiffness, is central to the understanding of these systems, their properties, and the design of related applications. In this we introduce FiberApp a new tracking and analysis software based on a cascade of algorithms describing structural and topological features of objects characterized by a very high length-to-width aspect ratio, generally described as fiber-like objects. The program operates on images from any microscopic source (atomic force or transmission electron microscopy, optical, fluorescence, confocal, etc.), acquiring the spatial coordinates of objects by a semiautomated tracking procedure based on A* pathfinding algorithm followed by the application of active contour models and generating virtually any statistical, topological, and graphical output derivable from these coordinates. Demonstrative features of the software include statistical polymer physics analysis of fiber conformations, height, bond and pair correlation functions, mean-squared end-to-end distance and midpoint displacement, 2D order parameter, excess kurtosis, fractal exponent, height profile and its discrete Fourier transform, orientation, length, height, curvature, and kink angle distributions, providing an unprecedented structural description of filamentous synthetic and biological objects. INTRODUCTION In modern natural and applied sciences, image processing plays a pivotal role in gaining qualitative and quantitative information from investigated complex systems. A subtype of signal processing, image processing operates on 2D or 3D arrays of numerical values by means of morphological operators, filters of various types, and other standard signal-processing tools. Image processing has a large impact, for example, in neurosciences, via neuron tracing in 3D stacks of fluorescence microscopy images. 1,2 The extraction of neuron s morphology via accurate localization of soma, axon, and dendrites 3,4 is a step in reconstruction of neuronal cells network; that is of great importance for understanding how the brain works. 5 The image processing is also very helpful in cell biology with segmenting cytoskeletal actin filaments that are involved in cell mobility and division as well as in carrying intracellular transport functions. 6 From elongation rates over time of these filaments, an estimation of kinetic and equilibrium constants for polymerization and depolymerization reactions allows clarification of detailed actin functionality in cells. 7,8 DNA is another important object, for which image processing has been traditionally widely used. It has become a classical system to apply polymer physics concepts 9,10 and to explore statistical properties and scaling behavior of macromolecules in both linear 11 and circular conformations. 12 In the case of amyloid fibrils, understanding the correlations between molecular structure, mechanical properties, and morphology, as obtained from processing atomic force and transmission electron microscopy (AFM, TEM) imaging data, is crucial for elucidating critical steps in the progression of neurodegenerative disorders, such as Alzheimer s, Parkinson s, or sickle cell disease. 21 Particularly, some recent investigations suggest that disease-specific prion fibrils exhibit higher elastic modulus than prion fibrils without disease specificity. 22 By exploiting the stochastic optical reconstruction microscopy, which is essentially a fluorescence microscopy technique based on highaccuracy localization of photoswitchable fluorophores, 23 the image processing can also be used in determining the exchange pathways of monomers within individual supramolecular fibers. 24 In order to test the accuracy of tracking procedures and verify the correctness of computational tools, artificial Received: November 9, 2014 Revised: January 26, 2015 XXXX American Chemical Society A
2 generation of synthetic images with curvilinear objects, that follow a certain shape model, can also be used. 25,26 Simulating images with adjustable size, resolution, level of noise, and system parameters (branching, length, thickness, flexibility, etc.) can greatly help in understanding the limitations of available algorithms and improve their efficiency. 27 Hence, gaining access to quantitative and qualitative information via image processing and statistical image analysis is central to a very vast landscape of fields, spanning from in biomedical research to nanotechnology and biomaterials science. An overarching necessity in all above-mentioned fields is the efficient detection of curvilinear objects in images of various types (2D and 3D), acquired primarily by fluorescence microscopy, confocal laser scanning microscopy, atomic force microscopy, and scanning and transmission electron microscopy (SEM and TEM). This general task has been considered by a vast number of scientists and researchers. The algorithms have been developed based on estimation of the first- and second-order directional derivatives, 28 active contour models that work by energy minimization of external (image intensities, or gradients) and internal (contour tension and elasticity) forces, 29 unbiased detector of curvilinear structures, 30 edge and ridge detection based on the optimization of a Cannylike criterion, 31 or linear feature detection using multiple directional nonmaximum suppression. 32 Some of these methods were widely adapted in medical research and neurosurgery for diagnosis of vascular diseases and planning vascular interventions from magnetic resonance angiographic (MRA) images and, particularly, in neuroscience and neurology, where investigation of crucial neuronal functions provides valuable information about the brain activity and possible applications in treatment of neurodegenerative diseases. An additional complication appearing in neuronal tracing is a separate detection of soma and branching points of dendrites, which is normally treated separately from axon tracing. 1 Particularly, an imaging data obtained in these research fields is represented as plane image stacks in 3D that depict fluorescently labeled cultured neurons. The minimal element of such images is called voxel (a combination of words volume and pixel ), which is an image intensity value on a regular grid in three-dimensional space. Generally, the above-mentioned algorithms with a readaptation to 3D can be used independently or in combination with the auxiliary pre- and postprocessing of recorded data. Some of them are based on an object thinning (skeletonization) method, primarily via morphological operations of erosion and dilation, that are an iterative removal of voxels from the surface of the object until the last thin line remains. 4,36 40 However, often they may not be robust enough on noisy images, which may severely disturb the quality of segmentation, and additional image preprocessing by noise filtering is highly required. Alternative methods employ the minimal-cost path calculation (Dijkstra s ora* 41 ) between two points defined by the user or, in an automatic way, through an appropriate additional utility algorithm. 39,42 In a general 2D case, to facilitate the minimal-cost pathfinding algorithm, or as a standalone method, the eigenvalues of Hessian matrix applied to an image are used to compute for each voxel the match to the object and the corresponding eigenvector for its local direction. 2,43,44 Other methods include voxel scooping 45 and model-based splines, 26,44 which were initially introduced as snakes. 29 More relevant to macromolecules, fibrillar structures, and polymers, various image processing methods are applied to quantify bending properties, network geometry, and basic morphological characteristics. Identification of the 3D gel microstructure, 46 in particular of collagen fibrils forming a matrix, 47 can be performed via the binarization and thresholding voxels of an image, followed by classification of branches 48 or by nucleation and local maximum points identification. 49 The search of centerline for linear, fiber-like objects like actin filaments and microtubules can be implemented in a similar manner, 50 or by minimal cumulative path cost estimation in tracking of outer tips of microtubules, 51 and quite promising model-based approaches. 25 Various methods have been applied for segmenting DNA 11,52,53 and amyloid fibrils 54,55 adsorbed on surface and visualized by AFM or TEM. Statistical and scaling properties of these objects can be derived from the tracked contours and from methods for simulation and measurement of amyloid fibril width, length, and height distributions. 27 Some macromolecules like DNA or carrageenan polysaccharides may exhibit a circular conformation and can be well described by a closed contour. 12,56,57 The intrachain bond correlation functions and statistical behavior of these systems are different compared to linear polymers The tracking approach based on the snakes model 29 provides a direct method to address these complications. These closed snakes can be used to obtain spatial description of circular objects in 2D visualized by AFM or EM, with a follow-up analysis of their particular features. In this context, the development of computational tools for the entire set of linear nonbranched and circular objects is of great importance in nanoscience. There are several examples of programming applications that can be found in the recent literature. Most among them are plug-ins for the Java-based image processing program ImageJ: NeuronJ (neurite tracing and analysis in fluorescence microscopy images), 43 Neurite- Tracer (automated quantification of neurite outgrowth), 39 JFilament (segmentation and tracking of cytoskeletal filaments), 25 and actin filament polymerization depolymerization tracker. 7 Furthermore, MATLAB-based or standalone applications are also available: NeuronMetrics (semiautomated processing of cultured neuron images), 38 NeuronCyto (neurite outgrowth measurement based on image segmentation with topological dependence), 3 V3D-Neuron (3D reconstruction of neurites), 61 DNA Trace, 62 and Simple Neurite Tracer (reconstruction, visualization and analysis of neuronal processes). 42 Yet, none of these softwares are fully rooted on polymer physics concepts to analyze filamentous objects, and a comprehensive polymeric description of filamentous objects is yet to be provided within a single program application, which calls for immediate developments. Over the past years an increase in acquisition modes of experimental visualization techniques and a growth of their quality and resolution have been remarkably rapid. It is noticeable, especially, in AFM imaging with more accurate cantilevers and possibilities to scan very-high-resolution images of the size pixels. With the parallel progress in computer speed and memory volumes, more precise image analysis tools and improved data handling have become highly desirable. In this we present a solution to abovementioned application problems raised in nanoscience a project with the open-source code called FiberApp, written in MATLAB programming language, in which cascading algorithms based on statistical polymer physics concepts have been implemented to yield the most accurate conformational, B
3 Figure 1. A few illustrative examples of systems that can be analyzed by FiberApp software. (A) Bovine serum albumin (BSA) flexible (left) and rigid (right) amyloid fibrils. 63,64 The inset images depict magnified areas of AFM images with point positioning along tracked contours with low and high intrinsic stiffness, respectively. (B) Simulated worm-like chain fibrils, generated directly in FiberApp software, providing a theoretical benchmark model against real systems. 66 (C) Functionalized multiwalled carbon nanotubes (the figure is adapted from Li et al. 67 ). The inset image reflects the feasibility of tracking for hollow objects. (D) Circular DNA adsorbed on APTES-modified mica (with carrageenan fibrils in the background). Image courtesy of Larissa Schefer. (E) β-lactoglobulin fibrils that form 2D liquid crystalline domains (left) 65 and circles (right), pointed by white arrows, at liquid interfaces (figure adapted from Jordens et al. 66 ). (F) Wavy lysozyme amyloid fibrils with subpersistence-length complex scaling behavior (figure adapted from Lara et al. 69 ). (G) Nanoclusters of Fe 3 O 4 nanoparticles with β-lactoglobulin fibrils that can align in the presence of a magnetic field (figure adapted from Bolisetty et al. 70 ). (H) Linear ι-carrageenan polysaccharide chains, prior (left) and after addition of salt (right), forming secondary structures and looped conformation. 57 Image courtesy of Larissa Schefer. The inset images demonstrate the possibility of closed-contour tracking. (I) TEMPO-oxidized wood cellulose nanofibrils with areas of different intrinsic stiffness (kinks), originating from the harsh mechanical treatment during sample preparation. 71 In the inset image it is demonstrated a concept of tracking with special masks that define contour segments with high curvature and low intrinsic stiffness. topological, and structural analysis available to date to describe DESCRIPTION OF THE SOFTWARE fibrous and filamentous biological and synthetic objects. The Typical Systems of Interest Handled by FiberApp input images, on which FiberApp works, may have been Software. The concept of representing fiber-like objects solely generated by any microscopy technique, essentially any TIF by their midline contours, tracked in microscopy images, is image, although most of the examples which will be presented applicable to a broad variety of systems in nanoscience. A few are based on high-resolution atomic force microscopy and types of such objects and their features, which have previously scanning and transmission electron microscopy imaging. We been investigated with the help of FiberApp software, are exemplified in Figure 1. Figure 1A shows the application of employ the most suitable and promising tracking method based FiberApp to bovine serum albumin (BSA) flexible (Figure 1A, on active contour models, reinforced by the additional utility left) and rigid (Figure 1A, right) amyloid fibrils, which are A* pathfinding algorithm for a faster semiautomatic segmentation process, allowing the tracking of thousands of objects in a temperature (90 C) and in an acidic environment (ph 2) for formed upon incubating the protein solution at an elevated short time. The potential to track linear and circular fiber-like several days. This analysis allowed the identification of six objects and, independently, the possibility to define segments distinct polymorphic types of BSA fibrils, which coexist with of heterogeneous stiffness along their contours using mask four types of intermediates. Furthermore, the handedness elements are only some of the key features that broaden the inversion, upon a level change in their hierarchical organization, was resolved. scope of possible applications of the program to a context much 63,64 Another example is the statistical analysis of simulated worm-like chain (WLC) contours with known and wider than that of classic statistical polymer physics. Moreover, predefined structural parameters (Figure 1B). 65 These artificial the ability of FiberApp to simulate in silico statistical data and images of simulated WLC fibrils, with the same resolution as artificial images of fiber-like objects serves as a powerful tool for the AFM imaging data subjected to the tracking procedure, can prediction and rationalization of experimental results. help to rule out possible artifacts that originate from tracking C
4 Figure 2. General scheme of the workflow of FiberApp software. The possible input data types are given at the left side of the figure (blue parallelograms). The main internal data types of the program Image and Fiber Data (green rectangles) define a framework of cyclic tracking procedure, where fiber data can be collected from, potentially, several images. The program s core functionalities are organized according to their specifications and the areas of implementation (orange rectangles). The functions for the fiber data analysis result in forms of output given at the right side of the figure (violet parallelograms). algorithms and, hence, be useful for analyzing the real fibril images. 66 Rigidity and mechanical properties of functionalized multiwalled carbon nanotubes with sulfonic groups are also accessible via FiberApp (Figure 1C). 67 Topological characteristics of DNA, 57 which is also an extensively used system for the application of polymer physics concepts in general, as handled within FiberApp, are shown in Figure 1D. 10,68 Another example, which consists of the liquid crystalline alignment of β-lactoglobulin fibrils at liquid interfaces, where the coexistence of two-dimensional isotropic and nematic phases can occur, is resolved and quantified by a length-scale-dependent order parameter S 2D calculated on all tracked contours (Figure 1E, left). 65 The same polar β-lactoglobulin fibrils at the air water interface do not possess the characteristic Gaussian curvature distribution for 2D WLC but instead have an excess of large curvature values (fat-tailed curvature distribution), which can lead to ring-like fibril conformations (Figure 1E, right). 66 In the case of wavy lysozyme fibrils, prepared by incubation of the protein solution at ph 2 and 60 C for several days, polymer physics concepts were applied via FiberApp on tracked contours in order to study the subpersistence-length complex scaling behavior and the periodicity in height profiles (Figure 1F). 69 Another class of objects handled by the software is a system of magnetic-responsive Fe 3 O 4 nanoparticle-modified protein spherical nanoclusters that can align in the presence of a magnetic field (Figure 1G). 70 The system of ι-carrageenan polysaccharide chains in the random coil conformation (Figure 1H, left) that undergoes a coil helix transition at high ionic strength coupled with an increase in thickness and rigidity of polymers is another example of biomacromolecules which can be handled by the software (Figure 1H, right). Along with the formation of the secondary structure, ι-carrageenan chains also form looped structures that coexist with linear polymer chains. 57 TEMPO-oxidized wood cellulose nanofibrils with kinks provide an example of heterogeneous stiffness along the contours, which causes an additional complication for the tracking procedure. FiberApp solves this problem by employing masks that define kink areas, so that the contour affinity to bend depends on whether the contour segment is inside or outside the mask (Figure 1I). This is essential for the tracking algorithm to correctly follow the nanocellulose fibrils 71 or DNA with protein-induced bending. 72,73 There is also the possibility to apply more sophisticated theories for chains with bends or sections of a different flexibility. 68 In the case of nanocellulose fibrils, the detailed statistical investigation of the kink angle distribution provided convincing evidence that the commonly accepted model of the structure as built of alternating hard crystalline and soft amorphous regions of cellulose chains is not appropriate and supported a process-induced kink formation generated during the preparation treatment. 71 Other examples of supramolecular fibrous aggregates, not shown in Figure 1, with periodic height changes accessible via FiberApp, are offered by self-assembled glycyrrhizic acid forming twisted ribbons 74 or ILQINS hexapeptides forming helical ribbons. 75 Description of the Workflow with FiberApp Software. Together with a brief description of A* pathfinding and active contour models based tracking procedures, modified and adapted to the needs of the software, we provide a complete workflow scheme for processing AFM and other types of microscopy images, as shown in Figure 2. The code is optimized for working with modern superhigh-resolution microscopy images ( pixels and virtually more) minimizing the number of errors upon tracking of the objects. Functions in the Image Preprocessing group can be used prior to fiber tracking in order to remove uneven illumination from the image and allow correct extraction of z coordinates. There are two major steps over the general workflow: Fiber Tracking and Data Analysis. With the first step, each fiberlike object in the image is fully characterized by its contour, which is a sequence of points ordered along the fiber s middle line from one end to the other. These contours have a constant distance between the point projections on the image plane, the step size Δs, which has a typical value of about one to several pixels size of the image. Thus, the coordinates x i, y i (in the image plane), and z i (height) of all the points covered by the contour provide a full spatial description of the fiber s central line. The initial positioning of a contour is manually defined with the help of a modified A* pathfinding algorithm. 41 For that, the image is considered to be a weighted graph of moving costs, where image pixels are represented by graph vertices. The cost of movement between two adjacent pixels is inversely proportional to the starting pixel intensity, and this value is stored as a graph edge between two corresponding graph D
5 Figure 3. Cryo-SEM images with tracked contours of nanocellulose fibrils. (A) A low stiffness parameter was used to perform tracking, which leads to an abrupt contour with undesirable fluctuations along it due to noise in the image. (B) The opposite case, a large stiffness parameter, suppresses the noise influence but does not allow an accurate fit in the vicinities of kinks. (C) The special masks (green squares) can be used to provide the contour with a heterogeneous stiffness, by making contour segments inside the masks softer (or stiffer) in comparison to the outer parts. Figure 4. (A) AFM images with BSA fibrils of type F 1 1 (bottom), R 1 0 (middle), and R 1 r (top). The scale bar and the color bar apply to all AFM images. (B) Height profiles of longitudinal sections along fibrils with respective color identifications from the panel A. The figures in panels A and B are readapted from Usov et al. 63 (C) Length distributions of flexible thin F 1 1 and flexible thick F 2 1 classes of BSA fibrils acquired after 40 h of incubation. (D) Length distributions of rigid thin R 1 0 +R 1 r and rigid thick R 2 0 +R 2 r classes of BSA fibrils acquired after 100 h of incubation. The solid lines represent the best-fits for histograms with a log-normal distribution function. The figures in panels C and D are readapted from Usov et al. 64 (E) Average height distributions of ι-carrageenan polysaccharide chains in random coil and ordered helical conformations and double-stranded DNA. The figure in panel E is reproduced from Schefer et al. 57 (F, G) Periodicity analysis and the pitch size estimation based on Fourier transform and autocorrelation function of the height profiles of supramolecular aggregates at different concentrations. The inset figure depicts an AFM image of analyzed fibrils. The figures in panels F and G are readapted from Saha et al. 74 (H) Cryo-TEM image of twisted lysozyme amyloid fibrils and a tracked contour shown as a red line. Image courtesy of Stephan Handschin. (I) Noisy pixel intensity profile along the contour. (J) Height autocorrelation function with clear peaks defining the periodicity of lysozyme amyloid fibrils. vertices. The algorithm then heuristically searches for the lowest cost path between two points that are manually defined by a user. Hence, the path will preferably go along a bright ridge of a fiber-like object in the image, which is a good first approximation of the object s midline position. The contour is, afterward, iteratively and automatically deformed according to the active contour model. 25,29 These contours are parametric curves that adapt to the image features in order to minimize the total contour energy, which is expressed as the sum of an external energy increasing with the positional offset of the contour from the brightest pixels of the image and an internal energy increasing with the growth of the total curvature along the contour. Functions in Tracking with Masks provide a possibility to use special mask elements that can be placed to define areas of the contour with different bending properties (Figure 3). For example, the curvature-penalizing parameter can be locally reduced, so that the contour can more easily bend there and exhibit a kink. In other words, it allows contours to have a heterogeneous stiffness (e.g., low stiffness inside mask areas, large stiffness outside). In this example, the masks are used to force the contour to bend in the kink areas and also define an angle of contour deviation (kink angle) for further analysis. The E
6 resulting coordinates of contours and masks after the tracking procedure can then be stored in a separate project file and provide the basis for further data analysis. Furthermore, the tool WLC Generator is used for creating both synthetic contour coordinates that follow WLC statistics and images with fibrils scattered on them, while functions in the Data Manipulation set provide a simple solution for combining and splitting statistics from different images or fiber types. In what follows, we describe in detail the second step of the workflow with FiberApp the basics of fiber data analysis that can be performed and typical results output that can be generated. The modulated internal architecture of FiberApp allows incorporating new processing methods with ease, thus making it a flexible platform for future developments. The technical details of processing algorithms, together with specifications of A* pathfinding and active contour models based tracking procedures, are discussed in detail in the Supporting Information. SPECIFICATIONS AND OUTPUTS OF THE IMPLEMENTED FIBER DATA PROCESSING METHODS Basic Morphological Parameters of Fiber-like Objects and Periodicity along Their Contours. The methods in this section provide a primary morphological characterization of the objects accessible by the software and include the height profile, the height and length distributions, the height autocorrelation function, and the height discrete Fourier transform analysis. The height profile is a simple test to briefly estimate the average height and check the presence of repeating patterns along the contour. It also can serve in distinguishing between different families of fiber-like objects. For example, Figure 4B represents the longitudinal sections of the BSA fibrils displayed in Figure 4A, by which three distinct types of thin BSA fibrils (flexible left-handed F 1 1, rigid without periodicity R 0 1, and rigid right-handed R r 1 ) based on their average height, existence of periodical structure along their contours and visual appearance, can be determined. 63 This method can be used to plot height profiles not only along curved objects but also along linear cross sections. The length distribution can provide information about nucleation and growth rates of fibrils, when distributions of samples, taken at different incubation times, are compared. Furthermore, in the system of BSA fibrils the statistical results indicate that the conversion rate of the polymorphic transformation F 1 1 R 0 r 1 +R 1 is slightly lower than that of the transformation F 1 2 R 0 2 +R r 2. This arises from the fact that the total length ratio of flexible classes L(F 1 1 )/L(F 1 2 ) 0.27 at 40 h of incubation is higher than the ratio of rigid classes L(R R r 1 )/L(R R r 2 ) 0.2 at 100 h. 64 The shape of length distribution for amyloid, and cellulose fibrils, 71 as well as many other parameters in biology 76 fits well to the log-normal probability density function f(l): A f ( L) = Lσ 2π e L 2 2 (ln μ) /2σ where L is the total length, μ and σ are the mean value and the standard deviation of the length natural logarithm, respectively, and A is a normalizing constant (Figure 4C,D). By plotting histograms of the average height values, extracted mainly from AFM images, it then becomes possible to analyze and discriminate between different families of fiber-like objects. (1) An example is shown in Figure 4E, where the height distributions are plotted for ι-carrageenan polysaccharide chains in random coil and ordered helical conformations together with the double-stranded DNA. From this accurate analysis, it was possible to resolve a long-standing question whether the ordered helical conformation of carrageenans exists as a single or double helix. The final conclusion that can be drawn states that thicker strands consist only of a single polymer chain. 57 Discrete Fourier transform (DFT) of height profiles provides a way to estimate a pitch size of objects with periodical height variations as in twisted or helical ribbons. Fast Fourier transform (FFT) is a rapid method to calculate DFT and has the form 2 πik ( 1)[( n 1)/ N] n A = h e k = 1,..., N k N n= 1 where h n is the height of the nth point along the contour, N is taken as a maximal number of points in a contour among all fiber-like objects, and i is the imaginary unit. Amplitudes A k of the DFT describes the presence of a frequency component k in the height profile. One clear peak in the graph is the signature of a distinct periodical structure, whose value in nanometers can be estimated via 1000/(peak position). The second way to study periodical patterns is to calculate the height autocorrelation function R xx. The discrete autocorrelation function has the general form R xx ( m) = 1 N m N 1 m n= 0 xx n n+ m where m is the lag, N is the total number of points along a contour, and x identifies either the discrete height h n or twist angle θ n profile. The twist (helix) angle profile is obtained via the height profile s maxima and minima peaks position. The angle values span linearly from 0 to π within the distance between maxima and the consequent minima height peaks and from π to 2π until the next maxima to complete the period. Hence, for sake of simplicity, the twist angle and height profiles have same periodicities (note that often it is assumed that a full period for the angle should have a doubled value in comparison to the height). A presence of periodicity in the resulting curve is a signature of the periodical pattern in height/twist angle profiles with the same period. Both the above-mentioned methods were applied to yield consistent values for the periodicity (p = 8 9 nm) of selfassembled fibrillar supramolecular structures of glycyrrhizic acid (Figure 4F,G). 74 We note here that the information on the pitch size is also essential for estimation of the mechanical properties of fiber-like objects in helical ribbons conformation. 77 Moreover, Figure 4H shows a Cryo-TEM image of twisted lysozyme amyloid fibrils with a tracked contour (red line). Despite the fact that the height profile along the contour is very noisy (Figure 4I), the Height Autocorrelation Function results in the clear peaks that define the periodicity of these lysozyme amyloid fibrils (Figure 4J) showing a great robustness of the approach. Persistence Length Estimation Methods. The persistence length is a basic property of polymers, which quantifies their rigidity and is formally defined via the bond correlation function in 3D as the length over which angular correlations in the tangent direction decrease by e times. 78 There are several approaches of the persistence length estimation for WLC (2) (3) F
7 Figure 5. Overview of the persistence length estimation methods for discrete contours that are employed in FiberApp and their practical applications. Schematic representations of methods: (A) mean-squared end-to-end distance (MSED) versus internal contour length; (B) bond correlation function (BCF); (C) mean-squared midpoint displacement (MSMD) versus internal contour length. (D) AFM image of ι-carrageenan polysaccharide chains in random coil and ordered helical conformations. The red arrows point toward the thick helix segments. (E) AFM image of BSA fibrils on mica. The arrows of the same color point to fibrils of a certain type: F 1 1 (cyan), F 2 1 (violet), R 1 0 (green), and R 2 0 (orange). (F) Plot of MSED versus internal contour length for separate helix and random coil segments of the ι-carrageenan polysaccharides and values of corresponding persistence lengths extracted from fits. (G) Plot of BCF for BSA fibrils of types F 1 1 and F 2 1 with resulting persistence lengths. (H) Plot of MSMD versus internal contour length for BSA fibrils of types R 1 0 and R 2 0 with resulting persistence lengths. The figures in panels D and F are readapted from Schefer et al. 57 The graphs in panels G and H are readapted from Usov et al. 63 discrete contours, consisting of N points and, accordingly, N 1 segments (Figure 5A C). However, there is evidence that some systems like DNA at short length scale 52,79 or simulated bottle-brush polymers when the chain length of the macromolecules tends to infinity 80 do not necessarily follow the standard statistics of WLC but can be well described by the alternative generalized theory of semiflexible polymers. Here, we assume a WLC behavior for investigated systems of fiberlike objects at all length scales. One of the most practical and widely used methods for the persistence length estimation is to calculate the mean-squared end-to-end distance (MSED) between contour segments (Figure 5A). This characteristic for a WLC model in 2D has the following theoretical dependence: R 2 =4λ[l 2λ(1 e l/2λ )], 9 where λ is the persistence length and R is the direct distance between any pair of segments along a contour separated by an arc length l. Employing this approach, a subtle increase in persistence length of ι-carrageenan chains upon a coil helix transition from 22.6 to 26.4 nm could be detected, indicating an increase in rigidity of segments in helical conformation (Figure 5D,F) and yielding a valuable direct indication of the coil helix transition on a single chain level. 57 The bond correlation function (BCF) is the most general way to evaluate the persistence length (Figure 5B). For WLC in 2D it corresponds to cos θ =e l/2λ, 81 where θ is the angle between tangent directions of any two segments along a fibril contour separated by an arc length l. A different method that can be successfully applied only to very stiff fiber-like objects (l < λ) is the mean-squared midpoint displacement (MSMD) as depicted in Figure 5C. The equation, describing the behavior of a midpoint deviation has the form u x 2 = l 3 /48λ, 21,55 where u x 2 is the mean-squared midpoint displacement between any pair of segments along a contour, separated by an arc length l. This expression is derived with an assumption that these deviations are small in comparison to the corresponding arc lengths ( u x l). Both methods were successfully applied for the persistence length estimation of BSA fibrils. The flexible fibrils were analyzed by means of BCF (Figure 5E,G), while rigid fibrils were via the MSMD method (Figure 5E,H). The obtained results are consistent with MSED analysis for all classes (λ MSED is equal to 160 nm for F 1 1, 150 nm for F 2 1, 1280 G
8 Figure 6. (A) Cryo-TEM image of lysozyme amyloid fibrils. (B) Averaged end-to-end distance versus the internal contour length plot with the scaling exponent ν values for sinus model and lysozyme fibrils at three different length scales. (C) Pair correlation function plot with corresponding values for the scaling exponent α. The figures in panels A, B, and C are readapted from Lara et al. 69 (D) AFM image of BSA amyloid fibrils of types F 1 1 and R 1 0. (E, F) Averaged end-to-end distance versus the internal contour length plots (blue axis) and scaling exponent propagations (red axis) for BSA fibrils of types (E) F 1 1 and (F) R 1 0. The figures in panels E and F are readapted from Usov et al. 63 Figure 7. AFM image of β-lactoglobulin fibrils (A) at the air water interface with ring configurations and (B) deposited onto mica from the bulk solution. (C, D) Normalized probability distributions of absolute curvature values extracted from the generated WLC data (violet), the generated WLC data after applying the tracking procedure (blue), real fibrils from panels A and B, respectively (green). In both scenarios real fibrils exhibit an excess of curvature upon being adsorbed to a surface, which is clearly visible in the inset images. The figures in panels A, B, C, and D are reproduced from Jordens et al. 66 (E) Probability density functions of the angles between segments of discrete WLCs with identical contour length L = 700 nm, persistence length λ = 50 nm, and discretization step size Δs = 1 nm. The segments are separated by a distance 1 nm (green), 25 nm (blue), 50 nm (violet), and 500 nm (red). (F) Excess kurtosis as a function of the separation distance between segments of WLCs. nm for R 1 0, and 2550 nm for R 2 0 ) 63 and give access to bending rigidities D of the corresponding fibrils as a characteristic of mechanical properties through the expression D = λk B T. 82 Scaling Behavior and Pair Correlation Function. Based on polymer physics concepts, the average end-to-end distance is a power-law function of the internal contour length l and scales according to the relation R(l) l v, 83 where ν is a scaling exponent. When l is small in comparison to the persistence length (l < λ), the scaling exponent approaches the value 1, indicating rod-like statistical behavior. For the contour lengths above the persistence length l λ the expected scaling exponent might vary, depending on temperature, quality of solvent, excluded volume interaction, and other parameters, and for the important case of self-avoiding random walk (SAW) chains in a good solvent this is equal to 3/4 in 2D and 3/5 in 3D. 10 The 2D pair-correlation function g(r) is defined as g(r) = s(r) /πr 2, 83 where s(r) is the length of the contour segment H
9 Figure 8. (A) AFM image of β-lactoglobulin fibrils adsorbed at the air water interface. (B, C) Orientation distribution of contour segments within areas highlighted in panel A. The 2D order parameter within fibril nematic domains is relatively large (panel B, S 2D = 0.3) in comparison to the corresponding value in the isotropic region (panel C, S 2D = 0.037). (D, E) AFM images of β-lactoglobulin fibrils adsorbed at the air water interface after a short period (10 min) and a long period (60 min) of adsorption times, respectively. (F, G) Decompositions of 2D order parameter S image 2D of the images from the panels D and G, respectively, into S align 2D and S rand 2D with weight factors a and b. The images in panels A, D, E, F, and G are readapted from Jordens et al. 65 incorporated in a circle of radius r, where the average is taken by centering the circles at the contour points with all possible positions of the circle. In the rod-like limit (l < λ) the pair correlation function behaves as g(r) r α, which together with the approximation 2r R(s) s v results in a relationship between two exponents α =1/ν Using FiberApp, the estimation of exponents for sinusoidal lysozyme amyloid fibrils below the persistence length (2.5 μm) was carried out by both approaches (Figure 6A C), with the observable progressive changes in the values of ν and α at multiple length scales. 69 An alternative way of presenting scaling exponents, based on the analysis of two BSA fibril classes F 1 1 and R 0 1 (Figure 6D), is illustrated in Figure 6E,F. A local linear fit of the log log plot with the end-to-end distance versus the internal contour length dependence reflects the scaling exponent at this particular position. The resulting curve is relatively smooth, and in such a case, it can accurately express the evolution of the scaling exponent value with contour length. For different systems the scaling exponent can indicate whether the fiber-like objects are in equilibrated or trapped configurations due to the interaction with a substrate. 9,84 For example, in the case of the flexible BSA fibrils F 1 1, the scaling exponent reached the plateau of 3/4 expected for SAW in 2D, indicating the equilibrated state on mica. For the rigid fibrils R 0 1, on the other hand, the processing length is not long enough compared to the persistence length to draw any conclusion and the scaling exponent does not attain any plateau value, i.e., is not stationary around the 3/4 expected value. 63 Curvature Distribution and Excess Kurtosis. The analysis involving the curvature estimation for polar and chiral fibrils can shed light into important aspects of interaction with a surface upon their adsorption. The probability density function p(κ) for the curvature κ of WLC in 2D is equal to p(κ) = (2λΔs/π) 1/2 e λδsκ2 and is described by a Gaussian distribution of κ. 25,85 It was shown that the fibrils exposed to an inhomogeneous environment, such as an air water interface or a mica substrate (Figure 7A,B), do not possess the same Gaussian distribution as WLCs. In all cases, there is an evidence of an excess of curvature, which can also lead to ring-like conformations of these fibrils (Figure 7A). This spontaneous curvature can be easily detected in the fat tails of the real fibrils curvature distributions in comparison to the synthetic WLCs data, which were subjected to the same tracking procedure in the corresponding simulated images, but still retained the Gaussian shape of curvature distribution (Figure 7C,D). The simulated images of WLCs contours were generated using parameters from the corresponding AFM images, by duly taking into account the finite tip size effect. 86,87 The equilibration of the fiber-like objects adsorbed on a substrate can also be verified using the excess kurtosis of the angle θ between consecutive segments of a tracked discrete contour. 11,53 The excess kurtosis is defined as K = θ 4 / θ 2 2 3, and if the chains are fully equilibrated on a substrate in 2D, then the distribution of θ is Gaussian and K = 0, which is a feature of the WLC behavior. 88 However, it is only possible to apply this method on separation distances below the persistence length (l < λ) because upon increasing the distance between segments the excess kurtosis value starts to decrease down to K = 1.2, and the probability density function of the angle θ transforms from Gaussian into uniform distribution shape (Figure 7E). The corresponding evolution of the excess kurtosis is depicted in Figure 7F for a population of 7000 generated WLC with identical contour length L = 700 nm, persistence length λ = 50 nm, and step size Δs = 1 nm. Orientation Distribution and 2D Order Parameter. The orientation distribution and the length-scale-dependent 2D order parameter can be used for quantifying the alignment of fiber-like objects in different systems and under various conditions. Figure 8A shows the AFM image of β-lactoglobulin fibrils adsorbed at the air water interface with the coexistence of strongly aligned nematic domains and completely randomly oriented fibrils, which form an isotropic phase in analogy to liquid crystals. The orientation distribution of segments belonging to all tracked contours inside box-size-dependent domains exhibits a clear peak along one direction (Figure 8B), while randomly oriented fibrils possess a uniform distribution in all possible directions (Figure 8C). It is possible to quantify the order of the assembly by assigning a particular number for each distribution, i.e., via a 2D order parameter. I
10 This is defined as S 2D =2 cos 2 θ n 1, where θ is the angle between the nth segment and the local director in the chosen area. The values of the 2D order parameter for the abovementioned areas are very different: 0.3 and 0.037, respectively (Figure 8B,C). This measure was used to quantify, for example, the alignment of the magnetic-responsive clusters containing hybrids of Fe 3 O 4 nanoparticles with β-lactoglobulin amyloid fibrils upon application of a magnetic field (Figure 1G). 70 In the case of β-lactoglobulin fibrils adsorbed at the air water interface we introduced an additional feature, which is the length scale dependence of the 2D order parameter, i.e., S 2D (d). For calculating this object, we divided the whole image into square blocks of a certain size d. Calculating and averaging S 2D values for all blocks results in one mean number, which is parametric with d, yielding the length scale dependent S 2D (d). 65 This function S 2D (d) is further expressed as the sum of the weighted components S align 2D (d) and S rand 2D (d), corresponding to the alignment of the nematic and isotropic components, respectively 65 image 2D align 2D rand 2D S () d = as () d + (1 a) S () d where a is the relative surface fraction of the aligned (nematic) domains. It can further be shown that decrease of the order with the box size d is an exponential function 65 align /2Λ 2D S ( d) = b + (1 b)e d where the offset b accounts the finite size of the image and Λ is the characteristic length of the nematic order decay, in analogy to the relationship between the persistence length and tangent correlations along the contour. The component S rand 2D (d) can be evaluated from simulated images of randomly oriented fibrils that are generated using all relevant parameters from the original AFM images. Applying this analysis to the tracked contours of β- lactoglobulin amyloid fibrils at the air water interface after different adsorption times (Figure 8D G), it is possible to quantify isotropic nematic transition in a reliable and rigorous manner. 65 From 2D Projection Traces to 3D Structural Parameters. Much of the discussion above refers to objects with a 2D conformation because either adsorbed on a substrate (AFM, TEM) or with a pseudo-2d conformation when confined within a thin layer of suspension (Cryo-TEM), for which a 2D polymer physics analysis provides an accurate description. It is however interesting to extract also information for objects, which span a truly 3D conformation, but for which only their projection traces on a plane are accessible. This is the case, for example for fibrils visualized by confocal microscopy or fluorescence-based stochastical optical reconstruction microscopy (STORM). In order to access the persistence length of fibrillar structures distributed randomly in 3D, one should carefully consider how the statistical properties of WLC projections change from a 3D space into a 2D plane. When a chain is projected onto the xy plane from the z direction, the mean-square of the projected end-to-end distance modifies into R 2 proj = (2/3) R 2 3D. 9 The observable projected internal contour length l proj also differs from the actual one, l 3D.To understand how, let l 3D be decomposed in N segments of identical length ΔS so that l 3D = NΔS; similarly, l proj = N i i i ΔS proj, where ΔS proj are the individual projections of the various ΔS i on the projection plane. Then we can rewrite l proj = N[(1/N) N i i ΔS proj ], or l proj = N ΔS proj, where ΔS proj is (4) (5) nothing else than the average projection of ΔS i on the projection plane. By averaging over all possible directions of segments ΔS placed at an angle θ from z, we derive ΔS proj = (2/π) π/2 0 ΔS sin θ dθ = (2/π)ΔS, and then l proj = N ΔS proj = N(2/π)ΔS = (2/π)l 3D. By applying the WLC model in 3D expressing R 2 3D vs l 3D for the actual fibril, R 2 3D =2λ[l 3D λ(1 e l3d/λ )], but using only the observable quantities R 2 proj and l proj, one gets R 2 proj =(2λ/3)[πl proj 2λ(1 e πlproj/2λ )], where λ is the real persistence length of WLC in 3D. This illustrates well that in the case of randomly oriented WLC fibrils in 3D their structural parameters, such as the persistence length, are easily accessible through a statistical analysis of their projections traces. Beside the persistence length, other important structural parameters can be extracted from the projection traces. For example, because fibrils are rather rigid, R 2 3D l v 3D, with the scaling exponent ν typically ranging between the very flexible random walk value, ν = 1/2, and the fully rigid chain value, ν = 1, the fractal exponent m =1/v ranges between 1 and 2 and is thus preserved upon projection on a surface of Euclidean dimension two, 89 implying that the scaling and fractal exponents for the real fibril and its projection are identical and thus accessible via the observable R 2 proj l v proj. OUTLOOK AND CONCLUSIONS Given the paramount significance that elongated, semiflexible colloidal objects and biological macromolecules embody in medicine, biology, materials science, and nanotechnology, there is an increasing urgent need to develop semiautomated procedures for the processing of a large amount of microscopy images and generating statistical significant output on the structural features and molecular conformations of these filamentous objects. The newly developed FiberApp open source code is deeply rooted in statistical polymer physics and is meant to bridge the gap currently existing between the highresolution imaging process of fiber-like, filamentous, and macromolecular objects and their structural analysis at the single molecule level. We anticipate that this code will serve a vast community of scientists working on very diverse disciplines and may pave the way to new approaches to the study of biological and synthetic polymers such as microtubules, actin, amyloid fibrils, collagen, nanocellulose, silk fibroin, and the many types of supramolecular fibers found in current applied and fundamental sciences. ASSOCIATED CONTENT *S Supporting Information Technical details of processing algorithms as well as specifications of A* pathfinding and active contour models based tracking procedures; FiberApp software. This material is available free of charge via the Internet at AUTHOR INFORMATION Corresponding Author * [email protected] (R.M.). Notes The authors declare no competing financial interest. J
11 Biographies Ivan Usov was born in 1988 in Cheboksary, Russia. In 2011, under the guidance of Prof. A. Muzafarov, he received his Master s degree with Honors on specialty Physics of Condensed Matter from Moscow State University, Russia. He then joined the group of Prof. R. Mezzenga at ETH Zurich to pursue a PhD degree on statistical analysis of fibrous objects and filaments. His research interests include quantitative analysis, chirality, and mechanical properties of natural fibrillar nanostructures. Raffaele Mezzenga received his PhD in the field of polymer physics from EPFL Lausanne. He was a postdoc at University of California, Santa Barbara, in He then moved to Nestle Research Center to work on the self-assembly of surfactants, natural amphiphiles, and liquid crystals. In 2005 he was hired as Associate Professor in the Physics Department of the University of Fribourg, and he then joined ETH Zurich on 2009 as Full Professor. His research focuses on the fundamental understanding of self-assembly processes in polymers, liquid crystals, food, and biological colloidal systems. Prof. Mezzenga has coauthored more than 200 publications and is recipient of several international awards, among which are the 2004 Swiss Science National Foundation Professorship, the 2011 Young Scientist Research Award (AOCS), the 2011 John Dillon Medal (APS), and the 2013 Biomacromolecules/Macromolecules Young Investigator Award (ACS). ACKNOWLEDGMENTS Authors acknowledge support from the Swiss National Science Foundation (SNF ) and are grateful to Larissa Schefer and Sophia Jordens for inspiring discussions and valuable assistance in software testing. The critical reading of the manuscript by Kathleen Smith and Ilya Savchenko is kindly acknowledged. REFERENCES (1) Meijering, E. Cytometry, Part A 2010, 77, (2) Fan, J.; Zhou, X.; Dy, J. G.; Zhang, Y.; Wong, S. T. C. Neuroinformatics 2009, 7, (3) Yu, W.; Lee, H. K.; Hariharan, S.; Bu, W.; Ahmed, S. Cytometry, Part A 2009, 75, (4) Leandro, J. J.; Cesar, R. M., Jr.; Costa Lda, F. J. Neurosci. Methods 2009, 177, (5) Machens, C. K. Science 2012, 338, (6) Fujiwara, I.; Vavylonis, D.; Pollard, T. D. Proc. Natl. Acad. Sci. U. S. A. 2007, 104, (7) Kuhn, J. R.; Pollard, T. D. Biophys. J. 2005, 88, (8) Fujiwara, I.; Takahashi, S.; Tadakuma, H.; Funatsu, T.; Ishiwata, S. Nat. Cell Biol. 2002, 4, (9) Rivetti, C.; Guthold, M.; Bustamante, C. J. Mol. Biol. 1996, 264, (10) Valle, F.; Favre, M.; De Los Rios, P.; Rosa, A.; Dietler, G. Phys. Rev. Lett. 2005, 95, (11) Faas, F. G. A.; Rieger, B.; van Vliet, L. J.; Cherny, D. I. Biophys. J. 2009, 97, (12) Witz, G.; Rechendorff, K.; Adamcik, J.; Dietler, G. Phys. Rev. Lett. 2008, 101, (13) Schleeger, M.; VandenAkker, C. C.; Deckert-Gaudig, T.; Deckert, V.; Velikov, K. P.; Koenderink, G.; Bonn, M. Polymer 2013, 54, (14) Knowles, T. P.; Fitzpatrick, A. W.; Meehan, S.; Mott, H. R.; Vendruscolo, M.; Dobson, C. M.; Welland, M. E. Science 2007, 318, (15) Volpatti, L. R.; Knowles, T. P. J. J. Polym. Sci., Part B: Polym. Phys. 2014, 52, (16) Adamcik, J.; Lara, C.; Usov, I.; Jeong, J. S.; Ruggeri, F. S.; Dietler, G.; Lashuel, H. A.; Hamley, I. W.; Mezzenga, R. Nanoscale 2012, 4, (17) Cherny, I.; Gazit, E. Angew. Chem., Int. Ed. 2008, 47, (18) Knowles, T. P. J.; Buehler, M. J. Nat. Nanotechnol. 2011, 6, (19) Dobson, C. M. Nature 2003, 426, (20) Selkoe, D. J. Nature 2003, 426, (21) Wang, J. C.; Turner, M. S.; Agarwal, G.; Kwong, S.; Josephs, R.; Ferrone, F. A.; Briehl, R. W. J. Mol. Biol. 2002, 315, (22) Yoon, G.; Kab Kim, Y.; Eom, K.; Na, S. Appl. Phys. Lett. 2013, 102, (23) Rust, M. J.; Bates, M.; Zhuang, X. Nat. Methods 2006, 3, (24) Albertazzi, L.; van der Zwaag, D.; Leenders, C. M. A.; Fitzner, R.; van der Hofstad, R. W.; Meijer, E. W. Science 2014, 344, (25) Smith, M. B.; Li, H.; Shen, T.; Huang, X.; Yusuf, E.; Vavylonis, D. Cytoskeleton 2010, 67, (26) Vasilkoski, Z.; Stepanyants, A. J. Neurosci. Methods 2009, 178, (27) Hall, D. Anal. Biochem. 2012, 421, (28) Haralick, R. M.; Watson, L. T.; Laffey, T. J. Int. J. Rob. Res. 1983, 2, (29) Kass, M.; Witkin, A.; Terzopoulos, D. Int. J. Comput. Vis. 1988, 1, (30) Steger, C. IEEE Trans. Pattern Anal. Mach. Intell. 1998, 20, (31) Jacob, M.; Unser, M. Pattern Anal. Mach. Intell. 2004, 26, (32) Sun, C.; Vallotton, P. J. Microsc. 2009, 234, (33) Frangi, A. F.; Niessen, W. J.; Hoogeveen, R. M.; van Walsum, T.; Viergever, M. A. IEEE Trans. Med. Imaging 1999, 18, (34) Lorigo, L. M.; Faugeras, O. D.; Grimson, W. E.; Keriven, R.; Kikinis, R.; Nabavi, A.; Westin, C. F. Med. Image Anal. 2001, 5, (35) Sato, Y.; Nakajima, S.; Shiraga, N.; Atsumi, H.; Yoshida, S.; Koller, T.; Gerig, G.; Kikinis, R. Med. Image Anal. 1998, 2, K
12 (36) Yuan, X.; Trachtenberg, J. T.; Potter, S. M.; Roysam, B. Neuroinformatics 2009, 7, (37) Broser, P. J.; Erdogan, S.; Grinevich, V.; Osten, P.; Sakmann, B.; Wallace, D. J. J. Neurosci. Methods 2008, 169, (38) Narro, M. L.; Yang, F.; Kraft, R.; Wenk, C.; Efrat, A.; Restifo, L. L. Brain Res. 2007, 1138, (39) Pool, M.; Thiemann, J.; Bar-Or, A.; Fournier, A. E. J. Neurosci. Methods 2008, 168, (40) Janoos, F.; Mosaliganti, K.; Xu, X.; Machiraju, R.; Huang, K.; Wong, S. T. C. Med. Image Anal. 2009, 13, (41) Hart, P.; Nilsson, N.; Raphael, B. IEEE Trans. Syst. Sci. Cybern. 1968, 4, (42) Longair, M. H.; Baker, D. A.; Armstrong, J. D. Bioinformatics 2011, 27, (43) Meijering, E.; Jacob, M.; Sarria, J.-C. F.; Steiner, P.; Hirling, H.; Unser, M. Cytometry, Part A 2004, 58, (44) Schmitt, S.; Evers, J. F.; Duch, C.; Scholz, M.; Obermayer, K. Neuroimage 2004, 23, (45) Rodriguez, A.; Ehlenberger, D. B.; Hof, P. R.; Wearne, S. L. J. Neurosci. Methods 2009, 184, (46) Nisslert, R.; Kvarnstro m, M.; Loreń, N.; Nydeń, M.; Rudemo, M. J. Microsc. 2007, 225, (47) Altendorf, H.; Decencie re, E.; Jeulin, D.; De sa Peixoto, P.; Deniset-Besseau, A.; Angelini, E.; Mosser, G.; Schanne-Klein, M.-C. J. Microsc. 2012, 247, (48) Wu, J.; Rajwa, B.; Filmer, D. L.; Hoffmann, C. M.; Yuan, B.; Chiang, C.; Sturgis, J.; Robinson, J. P. J. Microsc. 2003, 210, (49) Stein, A. M.; Vader, D. A.; Jawerth, L. M.; Weitz, D. A.; Sander, L. M. J. Microsc. 2008, 232, (50) Brangwynne, C. P.; Koenderink, G. H.; Barry, E.; Dogic, Z.; MacKintosh, F. C.; Weitz, D. A. Biophys. J. 2007, 93, (51) Hadjidemetriou, S.; Toomre, D.; Duncan, J. Med. Image Anal. 2008, 12, (52) Wiggins, P. A.; van der Heijden, T.; Moreno-Herrero, F.; Spakowitz, A.; Phillips, R.; Widom, J.; Dekker, C.; Nelson, P. C. Nat. Nanotechnol. 2006, 1, (53) Moukhtar, J.; Faivre-Moskalenko, C.; Milani, P.; Audit, B.; Vaillant, C.; Fontaine, E.; Mongelard, F.; Lavorel, G.; St-Jean, P.; Bouvet, P.; Argoul, F.; Arneodo, A. J. Phys. Chem. B 2010, 114, (54) Adamcik, J.; Jung, J.-M.; Flakowski, J.; De Los Rios, P.; Dietler, G.; Mezzenga, R. Nat. Nanotechnol. 2010, 5, (55) Smith, J. F.; Knowles, T. P. J.; Dobson, C. M.; Macphee, C. E.; Welland, M. E. Proc. Natl. Acad. Sci. U. S. A. 2006, 103, (56) Witz, G.; Rechendorff, K.; Adamcik, J.; Dietler, G. Phys. Rev. Lett. 2011, 106, (57) Schefer, L.; Adamcik, J.; Mezzenga, R. Angew. Chem., Int. Ed. 2014, 53, (58) Drube, F.; Alim, K.; Witz, G.; Dietler, G.; Frey, E. Nano Lett. 2010, 10, (59) Sakaue, T.; Witz, G.; Dietler, G.; Wada, H. EPL 2010, 91, (60) Timoshenko, E. G.; Kuznetsov, Y. A.; Connolly, R. J. Chem. Phys. 2002, 116, (61) Peng, H.; Ruan, Z.; Long, F.; Simpson, J. H.; Myers, E. W. Nat. Biotechnol. 2010, 28, (62) Mikhaylov, A.; Sekatskii, S. K.; Dietler, G. J. Adv. Microsc. Res. 2013, 8, (63) Usov, I.; Adamcik, J.; Mezzenga, R. ACS Nano 2013, 7, (64) Usov, I.; Adamcik, J.; Mezzenga, R. Faraday Discuss. 2013, 166, (65) Jordens, S.; Isa, L.; Usov, I.; Mezzenga, R. Nat. Commun. 2013, 4, (66) Jordens, S.; Riley, E. E.; Usov, I.; Isa, L.; Olmsted, P. D.; Mezzenga, R. ACS Nano 2014, 8, (67) Li, C.; Mezzenga, R. Langmuir 2012, 28, (68) Rivetti, C.; Walker, C.; Bustamante, C. J. Mol. Biol. 1998, 280, (69) Lara, C.; Usov, I.; Adamcik, J.; Mezzenga, R. Phys. Rev. Lett. 2011, 107, (70) Bolisetty, S.; Vallooran, J. J.; Adamcik, J.; Mezzenga, R. ACS Nano 2013, 7, (71) Usov, I.; Nystro m, G.; Adamcik, J.; Handschin, S.; Bergstro m, L.; Mezzenga, R., submitted. (72) Lu, Y.; Weers, B. D.; Stellwagen, N. C. Biophys. J. 2003, 85, (73) Dame, R. T.; van Mameren, J.; Luijsterburg, M. S.; Mysiak, M. E.; Janicíjevic, A.; Pazdzior, G.; van der Vliet, P. C.; Wyman, C.; Wuite, G. J. L. Nucleic Acids Res. 2005, 33, e68. (74) Saha, A.; Adamcik, J.; Bolisetty, S.; Handschin, S.; Mezzenga, R., submitted. (75) Lara, C.; Reynolds, N. P.; Berryman, J. T.; Xu, A.; Zhang, A.; Mezzenga, R. J. Am. Chem. Soc. 2014, 136, (76) McGeoch, C. C. A Guide to Experimental Algorithmics; Cambridge University Press: Cambridge, UK, 2012; p 171. (77) Usov, I.; Mezzenga, R. ACS Nano 2014, 8, (78) Rubinstein, M.; Colby, R. H. Polymer Physics; Oxford University Press: New York, 2003; pp (79) Wiggins, P. A.; Nelson, P. C. Phys. Rev. E: Stat., Nonlinear, Soft Matter Phys. 2006, 73, (80) Hsu, H.-P.; Paul, W.; Binder, K. Macromolecules 2010, 43, (81) Doi, M.; Edwards, S. F. The Theory of Polymer Dynamics; Oxford University Press: New York, 1986; p 317. (82) Manning, G. S. Phys. Rev. A 1986, 34, (83) De Gennes, P. G. Scaling Concepts in Polymer Physics; Cornell University Press: Ithaca, NY, (84) Mu cke, N.; Klenin, K.; Kirmse, R.; Bussiek, M.; Herrmann, H.; Hafner, M.; Langowski, J. PLoS One 2009, 4, e7756. (85) Rappaport, S. M.; Medalion, S.; Rabin, Y. arxiv: , (86) Villarrubia, J. S. Surf. Sci. 1994, 321, (87) Villarrubia, J. S. J. Res. Natl. Inst. Stand. Technol. 1997, 102, (88) Lamour, G.; Yip, C. K.; Li, H.; Gsponer, J. ACS Nano 2014, 8, (89) Mandelbrot, B. B. The Fractal Geometry of Nature; W.H. Freeman and Company: San Francisco, L
FiberApp: an Open-source Software for Tracking and Analyzing Biomacromolecules, Polymers, Filaments and Fibrous Objects
 Supporting Information for FiberApp: an Open-source Software for Tracking and Analyzing Biomacromolecules, Polymers, Filaments and Fibrous Objects Ivan Usov and Raffaele Mezzenga ETH Zurich, Department
Supporting Information for FiberApp: an Open-source Software for Tracking and Analyzing Biomacromolecules, Polymers, Filaments and Fibrous Objects Ivan Usov and Raffaele Mezzenga ETH Zurich, Department
High flexibility of DNA on short length scales probed by atomic force microscopy
 High flexibility of DNA on short length scales probed by atomic force microscopy Wiggins P. A. et al. Nature Nanotechnology (2006) presented by Anja Schwäger 23.01.2008 Outline Theory/Background Elasticity
High flexibility of DNA on short length scales probed by atomic force microscopy Wiggins P. A. et al. Nature Nanotechnology (2006) presented by Anja Schwäger 23.01.2008 Outline Theory/Background Elasticity
Real-time PCR: Understanding C t
 APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
Reflection and Refraction
 Equipment Reflection and Refraction Acrylic block set, plane-concave-convex universal mirror, cork board, cork board stand, pins, flashlight, protractor, ruler, mirror worksheet, rectangular block worksheet,
Equipment Reflection and Refraction Acrylic block set, plane-concave-convex universal mirror, cork board, cork board stand, pins, flashlight, protractor, ruler, mirror worksheet, rectangular block worksheet,
Lecture 19: Proteins, Primary Struture
 CPS260/BGT204.1 Algorithms in Computational Biology November 04, 2003 Lecture 19: Proteins, Primary Struture Lecturer: Pankaj K. Agarwal Scribe: Qiuhua Liu 19.1 The Building Blocks of Protein [1] Proteins
CPS260/BGT204.1 Algorithms in Computational Biology November 04, 2003 Lecture 19: Proteins, Primary Struture Lecturer: Pankaj K. Agarwal Scribe: Qiuhua Liu 19.1 The Building Blocks of Protein [1] Proteins
Clustering & Visualization
 Chapter 5 Clustering & Visualization Clustering in high-dimensional databases is an important problem and there are a number of different clustering paradigms which are applicable to high-dimensional data.
Chapter 5 Clustering & Visualization Clustering in high-dimensional databases is an important problem and there are a number of different clustering paradigms which are applicable to high-dimensional data.
The accurate calibration of all detectors is crucial for the subsequent data
 Chapter 4 Calibration The accurate calibration of all detectors is crucial for the subsequent data analysis. The stability of the gain and offset for energy and time calibration of all detectors involved
Chapter 4 Calibration The accurate calibration of all detectors is crucial for the subsequent data analysis. The stability of the gain and offset for energy and time calibration of all detectors involved
Chapter 8. Low energy ion scattering study of Fe 4 N on Cu(100)
 Low energy ion scattering study of 4 on Cu(1) Chapter 8. Low energy ion scattering study of 4 on Cu(1) 8.1. Introduction For a better understanding of the reconstructed 4 surfaces one would like to know
Low energy ion scattering study of 4 on Cu(1) Chapter 8. Low energy ion scattering study of 4 on Cu(1) 8.1. Introduction For a better understanding of the reconstructed 4 surfaces one would like to know
Collagen I Self-Assembly: Revealing the Developing Structures that Generate Turbidity. Supporting Material
 Collagen I Self-Assembly: Revealing the Developing Structures that Generate Turbidity Supporting Material Jieling Zhu and Laura J. Kaufman* Department of Chemistry, Columbia University, New York, NY 10027
Collagen I Self-Assembly: Revealing the Developing Structures that Generate Turbidity Supporting Material Jieling Zhu and Laura J. Kaufman* Department of Chemistry, Columbia University, New York, NY 10027
4. Which carbohydrate would you find as part of a molecule of RNA? a. Galactose b. Deoxyribose c. Ribose d. Glucose
 1. How is a polymer formed from multiple monomers? a. From the growth of the chain of carbon atoms b. By the removal of an OH group and a hydrogen atom c. By the addition of an OH group and a hydrogen
1. How is a polymer formed from multiple monomers? a. From the growth of the chain of carbon atoms b. By the removal of an OH group and a hydrogen atom c. By the addition of an OH group and a hydrogen
2.500 Threshold. 2.000 1000e - 001. Threshold. Exponential phase. Cycle Number
 application note Real-Time PCR: Understanding C T Real-Time PCR: Understanding C T 4.500 3.500 1000e + 001 4.000 3.000 1000e + 000 3.500 2.500 Threshold 3.000 2.000 1000e - 001 Rn 2500 Rn 1500 Rn 2000
application note Real-Time PCR: Understanding C T Real-Time PCR: Understanding C T 4.500 3.500 1000e + 001 4.000 3.000 1000e + 000 3.500 2.500 Threshold 3.000 2.000 1000e - 001 Rn 2500 Rn 1500 Rn 2000
International Year of Light 2015 Tech-Talks BREGENZ: Mehmet Arik Well-Being in Office Applications Light Measurement & Quality Parameters
 www.led-professional.com ISSN 1993-890X Trends & Technologies for Future Lighting Solutions ReviewJan/Feb 2015 Issue LpR 47 International Year of Light 2015 Tech-Talks BREGENZ: Mehmet Arik Well-Being in
www.led-professional.com ISSN 1993-890X Trends & Technologies for Future Lighting Solutions ReviewJan/Feb 2015 Issue LpR 47 International Year of Light 2015 Tech-Talks BREGENZ: Mehmet Arik Well-Being in
Atomic Force Microscope and Magnetic Force Microscope Background Information
 Atomic Force Microscope and Magnetic Force Microscope Background Information Lego Building Instructions There are several places to find the building instructions for building the Lego models of atomic
Atomic Force Microscope and Magnetic Force Microscope Background Information Lego Building Instructions There are several places to find the building instructions for building the Lego models of atomic
Avizo Inspect New software for industrial inspection and materials R&D
 Avizo Inspect New software for industrial inspection and materials R&D Reduce your design cycle, inspection times, and meet higher-level quality standards at a lower cost. Avizo Inspect software streamlines
Avizo Inspect New software for industrial inspection and materials R&D Reduce your design cycle, inspection times, and meet higher-level quality standards at a lower cost. Avizo Inspect software streamlines
Helices From Readily in Biological Structures
 The α Helix and the β Sheet Are Common Folding Patterns Although the overall conformation each protein is unique, there are only two different folding patterns are present in all proteins, which are α
The α Helix and the β Sheet Are Common Folding Patterns Although the overall conformation each protein is unique, there are only two different folding patterns are present in all proteins, which are α
CALIBRATION OF A ROBUST 2 DOF PATH MONITORING TOOL FOR INDUSTRIAL ROBOTS AND MACHINE TOOLS BASED ON PARALLEL KINEMATICS
 CALIBRATION OF A ROBUST 2 DOF PATH MONITORING TOOL FOR INDUSTRIAL ROBOTS AND MACHINE TOOLS BASED ON PARALLEL KINEMATICS E. Batzies 1, M. Kreutzer 1, D. Leucht 2, V. Welker 2, O. Zirn 1 1 Mechatronics Research
CALIBRATION OF A ROBUST 2 DOF PATH MONITORING TOOL FOR INDUSTRIAL ROBOTS AND MACHINE TOOLS BASED ON PARALLEL KINEMATICS E. Batzies 1, M. Kreutzer 1, D. Leucht 2, V. Welker 2, O. Zirn 1 1 Mechatronics Research
Chemical Synthesis. Overview. Chemical Synthesis of Nanocrystals. Self-Assembly of Nanocrystals. Example: Cu 146 Se 73 (PPh 3 ) 30
 Chemical Synthesis Spontaneous organization of molecules into stable, structurally well-defined aggregates at the nanometer length scale. Overview The 1-100 nm nanoscale length is in between traditional
Chemical Synthesis Spontaneous organization of molecules into stable, structurally well-defined aggregates at the nanometer length scale. Overview The 1-100 nm nanoscale length is in between traditional
Template-based Eye and Mouth Detection for 3D Video Conferencing
 Template-based Eye and Mouth Detection for 3D Video Conferencing Jürgen Rurainsky and Peter Eisert Fraunhofer Institute for Telecommunications - Heinrich-Hertz-Institute, Image Processing Department, Einsteinufer
Template-based Eye and Mouth Detection for 3D Video Conferencing Jürgen Rurainsky and Peter Eisert Fraunhofer Institute for Telecommunications - Heinrich-Hertz-Institute, Image Processing Department, Einsteinufer
Common Core Unit Summary Grades 6 to 8
 Common Core Unit Summary Grades 6 to 8 Grade 8: Unit 1: Congruence and Similarity- 8G1-8G5 rotations reflections and translations,( RRT=congruence) understand congruence of 2 d figures after RRT Dilations
Common Core Unit Summary Grades 6 to 8 Grade 8: Unit 1: Congruence and Similarity- 8G1-8G5 rotations reflections and translations,( RRT=congruence) understand congruence of 2 d figures after RRT Dilations
Physics 441/2: Transmission Electron Microscope
 Physics 441/2: Transmission Electron Microscope Introduction In this experiment we will explore the use of transmission electron microscopy (TEM) to take us into the world of ultrasmall structures. This
Physics 441/2: Transmission Electron Microscope Introduction In this experiment we will explore the use of transmission electron microscopy (TEM) to take us into the world of ultrasmall structures. This
Numerical Analysis of Independent Wire Strand Core (IWSC) Wire Rope
 Numerical Analysis of Independent Wire Strand Core (IWSC) Wire Rope Rakesh Sidharthan 1 Gnanavel B K 2 Assistant professor Mechanical, Department Professor, Mechanical Department, Gojan engineering college,
Numerical Analysis of Independent Wire Strand Core (IWSC) Wire Rope Rakesh Sidharthan 1 Gnanavel B K 2 Assistant professor Mechanical, Department Professor, Mechanical Department, Gojan engineering college,
Microscopy and Nanoindentation. Combining Orientation Imaging. to investigate localized. deformation behaviour. Felix Reinauer
 Combining Orientation Imaging Microscopy and Nanoindentation to investigate localized deformation behaviour Felix Reinauer René de Kloe Matt Nowell Introduction Anisotropy in crystalline materials Presentation
Combining Orientation Imaging Microscopy and Nanoindentation to investigate localized deformation behaviour Felix Reinauer René de Kloe Matt Nowell Introduction Anisotropy in crystalline materials Presentation
Part-Based Recognition
 Part-Based Recognition Benedict Brown CS597D, Fall 2003 Princeton University CS 597D, Part-Based Recognition p. 1/32 Introduction Many objects are made up of parts It s presumably easier to identify simple
Part-Based Recognition Benedict Brown CS597D, Fall 2003 Princeton University CS 597D, Part-Based Recognition p. 1/32 Introduction Many objects are made up of parts It s presumably easier to identify simple
Improved predictive modeling of white LEDs with accurate luminescence simulation and practical inputs
 Improved predictive modeling of white LEDs with accurate luminescence simulation and practical inputs TracePro Opto-Mechanical Design Software s Fluorescence Property Utility TracePro s Fluorescence Property
Improved predictive modeling of white LEDs with accurate luminescence simulation and practical inputs TracePro Opto-Mechanical Design Software s Fluorescence Property Utility TracePro s Fluorescence Property
8.2 Elastic Strain Energy
 Section 8. 8. Elastic Strain Energy The strain energy stored in an elastic material upon deformation is calculated below for a number of different geometries and loading conditions. These expressions for
Section 8. 8. Elastic Strain Energy The strain energy stored in an elastic material upon deformation is calculated below for a number of different geometries and loading conditions. These expressions for
Big Ideas in Mathematics
 Big Ideas in Mathematics which are important to all mathematics learning. (Adapted from the NCTM Curriculum Focal Points, 2006) The Mathematics Big Ideas are organized using the PA Mathematics Standards
Big Ideas in Mathematics which are important to all mathematics learning. (Adapted from the NCTM Curriculum Focal Points, 2006) The Mathematics Big Ideas are organized using the PA Mathematics Standards
Steffen Lindert, René Staritzbichler, Nils Wötzel, Mert Karakaş, Phoebe L. Stewart, and Jens Meiler
 Structure 17 Supplemental Data EM-Fold: De Novo Folding of α-helical Proteins Guided by Intermediate-Resolution Electron Microscopy Density Maps Steffen Lindert, René Staritzbichler, Nils Wötzel, Mert
Structure 17 Supplemental Data EM-Fold: De Novo Folding of α-helical Proteins Guided by Intermediate-Resolution Electron Microscopy Density Maps Steffen Lindert, René Staritzbichler, Nils Wötzel, Mert
TCOM 370 NOTES 99-4 BANDWIDTH, FREQUENCY RESPONSE, AND CAPACITY OF COMMUNICATION LINKS
 TCOM 370 NOTES 99-4 BANDWIDTH, FREQUENCY RESPONSE, AND CAPACITY OF COMMUNICATION LINKS 1. Bandwidth: The bandwidth of a communication link, or in general any system, was loosely defined as the width of
TCOM 370 NOTES 99-4 BANDWIDTH, FREQUENCY RESPONSE, AND CAPACITY OF COMMUNICATION LINKS 1. Bandwidth: The bandwidth of a communication link, or in general any system, was loosely defined as the width of
Introduction to X-Ray Powder Diffraction Data Analysis
 Introduction to X-Ray Powder Diffraction Data Analysis Center for Materials Science and Engineering at MIT http://prism.mit.edu/xray An X-ray diffraction pattern is a plot of the intensity of X-rays scattered
Introduction to X-Ray Powder Diffraction Data Analysis Center for Materials Science and Engineering at MIT http://prism.mit.edu/xray An X-ray diffraction pattern is a plot of the intensity of X-rays scattered
DOING PHYSICS WITH MATLAB COMPUTATIONAL OPTICS RAYLEIGH-SOMMERFELD DIFFRACTION INTEGRAL OF THE FIRST KIND
 DOING PHYSICS WITH MATLAB COMPUTATIONAL OPTICS RAYLEIGH-SOMMERFELD DIFFRACTION INTEGRAL OF THE FIRST KIND THE THREE-DIMENSIONAL DISTRIBUTION OF THE RADIANT FLUX DENSITY AT THE FOCUS OF A CONVERGENCE BEAM
DOING PHYSICS WITH MATLAB COMPUTATIONAL OPTICS RAYLEIGH-SOMMERFELD DIFFRACTION INTEGRAL OF THE FIRST KIND THE THREE-DIMENSIONAL DISTRIBUTION OF THE RADIANT FLUX DENSITY AT THE FOCUS OF A CONVERGENCE BEAM
Solving Simultaneous Equations and Matrices
 Solving Simultaneous Equations and Matrices The following represents a systematic investigation for the steps used to solve two simultaneous linear equations in two unknowns. The motivation for considering
Solving Simultaneous Equations and Matrices The following represents a systematic investigation for the steps used to solve two simultaneous linear equations in two unknowns. The motivation for considering
Raman spectroscopy Lecture
 Raman spectroscopy Lecture Licentiate course in measurement science and technology Spring 2008 10.04.2008 Antti Kivioja Contents - Introduction - What is Raman spectroscopy? - The theory of Raman spectroscopy
Raman spectroscopy Lecture Licentiate course in measurement science and technology Spring 2008 10.04.2008 Antti Kivioja Contents - Introduction - What is Raman spectroscopy? - The theory of Raman spectroscopy
Atomic Force Microscopy Observation and Characterization of a CD Stamper, Lycopodium Spores, and Step-Height Standard Diffraction Grating
 Atomic Force Microscopy Observation and Characterization of a CD Stamper, Lycopodium Spores, and Step-Height Standard Diffraction Grating Michael McMearty and Frit Miot Special Thanks to Brendan Cross
Atomic Force Microscopy Observation and Characterization of a CD Stamper, Lycopodium Spores, and Step-Height Standard Diffraction Grating Michael McMearty and Frit Miot Special Thanks to Brendan Cross
1 Introduction. 1.1 Historical Perspective
 j1 1 Introduction 1.1 Historical Perspective The invention of scanning probe microscopy is considered one of the major advances in materials science since 1950 [1, 2]. Scanning probe microscopy includes
j1 1 Introduction 1.1 Historical Perspective The invention of scanning probe microscopy is considered one of the major advances in materials science since 1950 [1, 2]. Scanning probe microscopy includes
J. P. Oakley and R. T. Shann. Department of Electrical Engineering University of Manchester Manchester M13 9PL U.K. Abstract
 A CURVATURE SENSITIVE FILTER AND ITS APPLICATION IN MICROFOSSIL IMAGE CHARACTERISATION J. P. Oakley and R. T. Shann Department of Electrical Engineering University of Manchester Manchester M13 9PL U.K.
A CURVATURE SENSITIVE FILTER AND ITS APPLICATION IN MICROFOSSIL IMAGE CHARACTERISATION J. P. Oakley and R. T. Shann Department of Electrical Engineering University of Manchester Manchester M13 9PL U.K.
Physics 9e/Cutnell. correlated to the. College Board AP Physics 1 Course Objectives
 Physics 9e/Cutnell correlated to the College Board AP Physics 1 Course Objectives Big Idea 1: Objects and systems have properties such as mass and charge. Systems may have internal structure. Enduring
Physics 9e/Cutnell correlated to the College Board AP Physics 1 Course Objectives Big Idea 1: Objects and systems have properties such as mass and charge. Systems may have internal structure. Enduring
Algebra 1 2008. Academic Content Standards Grade Eight and Grade Nine Ohio. Grade Eight. Number, Number Sense and Operations Standard
 Academic Content Standards Grade Eight and Grade Nine Ohio Algebra 1 2008 Grade Eight STANDARDS Number, Number Sense and Operations Standard Number and Number Systems 1. Use scientific notation to express
Academic Content Standards Grade Eight and Grade Nine Ohio Algebra 1 2008 Grade Eight STANDARDS Number, Number Sense and Operations Standard Number and Number Systems 1. Use scientific notation to express
Jiří Matas. Hough Transform
 Hough Transform Jiří Matas Center for Machine Perception Department of Cybernetics, Faculty of Electrical Engineering Czech Technical University, Prague Many slides thanks to Kristen Grauman and Bastian
Hough Transform Jiří Matas Center for Machine Perception Department of Cybernetics, Faculty of Electrical Engineering Czech Technical University, Prague Many slides thanks to Kristen Grauman and Bastian
Encoders for Linear Motors in the Electronics Industry
 Technical Information Encoders for Linear Motors in the Electronics Industry The semiconductor industry and automation technology increasingly require more precise and faster machines in order to satisfy
Technical Information Encoders for Linear Motors in the Electronics Industry The semiconductor industry and automation technology increasingly require more precise and faster machines in order to satisfy
Modelling, Extraction and Description of Intrinsic Cues of High Resolution Satellite Images: Independent Component Analysis based approaches
 Modelling, Extraction and Description of Intrinsic Cues of High Resolution Satellite Images: Independent Component Analysis based approaches PhD Thesis by Payam Birjandi Director: Prof. Mihai Datcu Problematic
Modelling, Extraction and Description of Intrinsic Cues of High Resolution Satellite Images: Independent Component Analysis based approaches PhD Thesis by Payam Birjandi Director: Prof. Mihai Datcu Problematic
MetaMorph Software Basic Analysis Guide The use of measurements and journals
 MetaMorph Software Basic Analysis Guide The use of measurements and journals Version 1.0.2 1 Section I: How Measure Functions Operate... 3 1. Selected images... 3 2. Thresholding... 3 3. Regions of interest...
MetaMorph Software Basic Analysis Guide The use of measurements and journals Version 1.0.2 1 Section I: How Measure Functions Operate... 3 1. Selected images... 3 2. Thresholding... 3 3. Regions of interest...
2.2 Creaseness operator
 2.2. Creaseness operator 31 2.2 Creaseness operator Antonio López, a member of our group, has studied for his PhD dissertation the differential operators described in this section [72]. He has compared
2.2. Creaseness operator 31 2.2 Creaseness operator Antonio López, a member of our group, has studied for his PhD dissertation the differential operators described in this section [72]. He has compared
Optical Design Tools for Backlight Displays
 Optical Design Tools for Backlight Displays Introduction Backlights are used for compact, portable, electronic devices with flat panel Liquid Crystal Displays (LCDs) that require illumination from behind.
Optical Design Tools for Backlight Displays Introduction Backlights are used for compact, portable, electronic devices with flat panel Liquid Crystal Displays (LCDs) that require illumination from behind.
Classification of Fingerprints. Sarat C. Dass Department of Statistics & Probability
 Classification of Fingerprints Sarat C. Dass Department of Statistics & Probability Fingerprint Classification Fingerprint classification is a coarse level partitioning of a fingerprint database into smaller
Classification of Fingerprints Sarat C. Dass Department of Statistics & Probability Fingerprint Classification Fingerprint classification is a coarse level partitioning of a fingerprint database into smaller
Image Segmentation and Registration
 Image Segmentation and Registration Dr. Christine Tanner ([email protected]) Computer Vision Laboratory, ETH Zürich Dr. Verena Kaynig, Machine Learning Laboratory, ETH Zürich Outline Segmentation
Image Segmentation and Registration Dr. Christine Tanner ([email protected]) Computer Vision Laboratory, ETH Zürich Dr. Verena Kaynig, Machine Learning Laboratory, ETH Zürich Outline Segmentation
Prentice Hall Algebra 2 2011 Correlated to: Colorado P-12 Academic Standards for High School Mathematics, Adopted 12/2009
 Content Area: Mathematics Grade Level Expectations: High School Standard: Number Sense, Properties, and Operations Understand the structure and properties of our number system. At their most basic level
Content Area: Mathematics Grade Level Expectations: High School Standard: Number Sense, Properties, and Operations Understand the structure and properties of our number system. At their most basic level
Analecta Vol. 8, No. 2 ISSN 2064-7964
 EXPERIMENTAL APPLICATIONS OF ARTIFICIAL NEURAL NETWORKS IN ENGINEERING PROCESSING SYSTEM S. Dadvandipour Institute of Information Engineering, University of Miskolc, Egyetemváros, 3515, Miskolc, Hungary,
EXPERIMENTAL APPLICATIONS OF ARTIFICIAL NEURAL NETWORKS IN ENGINEERING PROCESSING SYSTEM S. Dadvandipour Institute of Information Engineering, University of Miskolc, Egyetemváros, 3515, Miskolc, Hungary,
Near-field scanning optical microscopy (SNOM)
 Adviser: dr. Maja Remškar Institut Jožef Stefan January 2010 1 2 3 4 5 6 Fluorescence Raman and surface enhanced Raman 7 Conventional optical microscopy-limited resolution Two broad classes of techniques
Adviser: dr. Maja Remškar Institut Jožef Stefan January 2010 1 2 3 4 5 6 Fluorescence Raman and surface enhanced Raman 7 Conventional optical microscopy-limited resolution Two broad classes of techniques
The Scheduled MRM Algorithm Enables Intelligent Use of Retention Time During Multiple Reaction Monitoring
 The Scheduled MRM Algorithm Enables Intelligent Use of Retention Time During Multiple Reaction Monitoring Delivering up to 2500 MRM Transitions per LC Run Christie Hunter 1, Brigitte Simons 2 1 AB SCIEX,
The Scheduled MRM Algorithm Enables Intelligent Use of Retention Time During Multiple Reaction Monitoring Delivering up to 2500 MRM Transitions per LC Run Christie Hunter 1, Brigitte Simons 2 1 AB SCIEX,
Structural Integrity Analysis
 Structural Integrity Analysis 1. STRESS CONCENTRATION Igor Kokcharov 1.1 STRESSES AND CONCENTRATORS 1.1.1 Stress An applied external force F causes inner forces in the carrying structure. Inner forces
Structural Integrity Analysis 1. STRESS CONCENTRATION Igor Kokcharov 1.1 STRESSES AND CONCENTRATORS 1.1.1 Stress An applied external force F causes inner forces in the carrying structure. Inner forces
DYNAMIC LIGHT SCATTERING COMMON TERMS DEFINED
 DYNAMIC LIGHT SCATTERING COMMON TERMS DEFINED Abstract: There are a number of sources of information that give a mathematical description of the terms used in light scattering. However, these will not
DYNAMIC LIGHT SCATTERING COMMON TERMS DEFINED Abstract: There are a number of sources of information that give a mathematical description of the terms used in light scattering. However, these will not
NEW YORK STATE TEACHER CERTIFICATION EXAMINATIONS
 NEW YORK STATE TEACHER CERTIFICATION EXAMINATIONS TEST DESIGN AND FRAMEWORK September 2014 Authorized for Distribution by the New York State Education Department This test design and framework document
NEW YORK STATE TEACHER CERTIFICATION EXAMINATIONS TEST DESIGN AND FRAMEWORK September 2014 Authorized for Distribution by the New York State Education Department This test design and framework document
Morphological segmentation of histology cell images
 Morphological segmentation of histology cell images A.Nedzved, S.Ablameyko, I.Pitas Institute of Engineering Cybernetics of the National Academy of Sciences Surganova, 6, 00 Minsk, Belarus E-mail [email protected]
Morphological segmentation of histology cell images A.Nedzved, S.Ablameyko, I.Pitas Institute of Engineering Cybernetics of the National Academy of Sciences Surganova, 6, 00 Minsk, Belarus E-mail [email protected]
Galaxy Morphological Classification
 Galaxy Morphological Classification Jordan Duprey and James Kolano Abstract To solve the issue of galaxy morphological classification according to a classification scheme modelled off of the Hubble Sequence,
Galaxy Morphological Classification Jordan Duprey and James Kolano Abstract To solve the issue of galaxy morphological classification according to a classification scheme modelled off of the Hubble Sequence,
Real-Time PCR Vs. Traditional PCR
 Real-Time PCR Vs. Traditional PCR Description This tutorial will discuss the evolution of traditional PCR methods towards the use of Real-Time chemistry and instrumentation for accurate quantitation. Objectives
Real-Time PCR Vs. Traditional PCR Description This tutorial will discuss the evolution of traditional PCR methods towards the use of Real-Time chemistry and instrumentation for accurate quantitation. Objectives
Java Modules for Time Series Analysis
 Java Modules for Time Series Analysis Agenda Clustering Non-normal distributions Multifactor modeling Implied ratings Time series prediction 1. Clustering + Cluster 1 Synthetic Clustering + Time series
Java Modules for Time Series Analysis Agenda Clustering Non-normal distributions Multifactor modeling Implied ratings Time series prediction 1. Clustering + Cluster 1 Synthetic Clustering + Time series
IN current film media, the increase in areal density has
 IEEE TRANSACTIONS ON MAGNETICS, VOL. 44, NO. 1, JANUARY 2008 193 A New Read Channel Model for Patterned Media Storage Seyhan Karakulak, Paul H. Siegel, Fellow, IEEE, Jack K. Wolf, Life Fellow, IEEE, and
IEEE TRANSACTIONS ON MAGNETICS, VOL. 44, NO. 1, JANUARY 2008 193 A New Read Channel Model for Patterned Media Storage Seyhan Karakulak, Paul H. Siegel, Fellow, IEEE, Jack K. Wolf, Life Fellow, IEEE, and
Measuring Line Edge Roughness: Fluctuations in Uncertainty
 Tutor6.doc: Version 5/6/08 T h e L i t h o g r a p h y E x p e r t (August 008) Measuring Line Edge Roughness: Fluctuations in Uncertainty Line edge roughness () is the deviation of a feature edge (as
Tutor6.doc: Version 5/6/08 T h e L i t h o g r a p h y E x p e r t (August 008) Measuring Line Edge Roughness: Fluctuations in Uncertainty Line edge roughness () is the deviation of a feature edge (as
The Olympus stereology system. The Computer Assisted Stereological Toolbox
 The Olympus stereology system The Computer Assisted Stereological Toolbox CAST is a Computer Assisted Stereological Toolbox for PCs running Microsoft Windows TM. CAST is an interactive, user-friendly,
The Olympus stereology system The Computer Assisted Stereological Toolbox CAST is a Computer Assisted Stereological Toolbox for PCs running Microsoft Windows TM. CAST is an interactive, user-friendly,
Introduction to acoustic imaging
 Introduction to acoustic imaging Contents 1 Propagation of acoustic waves 3 1.1 Wave types.......................................... 3 1.2 Mathematical formulation.................................. 4 1.3
Introduction to acoustic imaging Contents 1 Propagation of acoustic waves 3 1.1 Wave types.......................................... 3 1.2 Mathematical formulation.................................. 4 1.3
5-Axis Test-Piece Influence of Machining Position
 5-Axis Test-Piece Influence of Machining Position Michael Gebhardt, Wolfgang Knapp, Konrad Wegener Institute of Machine Tools and Manufacturing (IWF), Swiss Federal Institute of Technology (ETH), Zurich,
5-Axis Test-Piece Influence of Machining Position Michael Gebhardt, Wolfgang Knapp, Konrad Wegener Institute of Machine Tools and Manufacturing (IWF), Swiss Federal Institute of Technology (ETH), Zurich,
18.2 Protein Structure and Function: An Overview
 18.2 Protein Structure and Function: An Overview Protein: A large biological molecule made of many amino acids linked together through peptide bonds. Alpha-amino acid: Compound with an amino group bonded
18.2 Protein Structure and Function: An Overview Protein: A large biological molecule made of many amino acids linked together through peptide bonds. Alpha-amino acid: Compound with an amino group bonded
Understanding Poles and Zeros
 MASSACHUSETTS INSTITUTE OF TECHNOLOGY DEPARTMENT OF MECHANICAL ENGINEERING 2.14 Analysis and Design of Feedback Control Systems Understanding Poles and Zeros 1 System Poles and Zeros The transfer function
MASSACHUSETTS INSTITUTE OF TECHNOLOGY DEPARTMENT OF MECHANICAL ENGINEERING 2.14 Analysis and Design of Feedback Control Systems Understanding Poles and Zeros 1 System Poles and Zeros The transfer function
A Game of Numbers (Understanding Directivity Specifications)
 A Game of Numbers (Understanding Directivity Specifications) José (Joe) Brusi, Brusi Acoustical Consulting Loudspeaker directivity is expressed in many different ways on specification sheets and marketing
A Game of Numbers (Understanding Directivity Specifications) José (Joe) Brusi, Brusi Acoustical Consulting Loudspeaker directivity is expressed in many different ways on specification sheets and marketing
Polymers: Introduction
 Chapter Outline: Polymer Structures Hydrocarbon and Polymer Molecules Chemistry of Polymer Molecules Molecular Weight and Shape Molecular Structure and Configurations Copolymers Polymer Crystals Optional
Chapter Outline: Polymer Structures Hydrocarbon and Polymer Molecules Chemistry of Polymer Molecules Molecular Weight and Shape Molecular Structure and Configurations Copolymers Polymer Crystals Optional
AN EXPERT SYSTEM TO ANALYZE HOMOGENEITY IN FUEL ELEMENT PLATES FOR RESEARCH REACTORS
 AN EXPERT SYSTEM TO ANALYZE HOMOGENEITY IN FUEL ELEMENT PLATES FOR RESEARCH REACTORS Cativa Tolosa, S. and Marajofsky, A. Comisión Nacional de Energía Atómica Abstract In the manufacturing control of Fuel
AN EXPERT SYSTEM TO ANALYZE HOMOGENEITY IN FUEL ELEMENT PLATES FOR RESEARCH REACTORS Cativa Tolosa, S. and Marajofsky, A. Comisión Nacional de Energía Atómica Abstract In the manufacturing control of Fuel
Indiana's Academic Standards 2010 ICP Indiana's Academic Standards 2016 ICP. map) that describe the relationship acceleration, velocity and distance.
 .1.1 Measure the motion of objects to understand.1.1 Develop graphical, the relationships among distance, velocity and mathematical, and pictorial acceleration. Develop deeper understanding through representations
.1.1 Measure the motion of objects to understand.1.1 Develop graphical, the relationships among distance, velocity and mathematical, and pictorial acceleration. Develop deeper understanding through representations
Understand the Sketcher workbench of CATIA V5.
 Chapter 1 Drawing Sketches in Learning Objectives the Sketcher Workbench-I After completing this chapter you will be able to: Understand the Sketcher workbench of CATIA V5. Start a new file in the Part
Chapter 1 Drawing Sketches in Learning Objectives the Sketcher Workbench-I After completing this chapter you will be able to: Understand the Sketcher workbench of CATIA V5. Start a new file in the Part
The Scientific Data Mining Process
 Chapter 4 The Scientific Data Mining Process When I use a word, Humpty Dumpty said, in rather a scornful tone, it means just what I choose it to mean neither more nor less. Lewis Carroll [87, p. 214] In
Chapter 4 The Scientific Data Mining Process When I use a word, Humpty Dumpty said, in rather a scornful tone, it means just what I choose it to mean neither more nor less. Lewis Carroll [87, p. 214] In
Data Exploration Data Visualization
 Data Exploration Data Visualization What is data exploration? A preliminary exploration of the data to better understand its characteristics. Key motivations of data exploration include Helping to select
Data Exploration Data Visualization What is data exploration? A preliminary exploration of the data to better understand its characteristics. Key motivations of data exploration include Helping to select
Lecture 14. Point Spread Function (PSF)
 Lecture 14 Point Spread Function (PSF), Modulation Transfer Function (MTF), Signal-to-noise Ratio (SNR), Contrast-to-noise Ratio (CNR), and Receiver Operating Curves (ROC) Point Spread Function (PSF) Recollect
Lecture 14 Point Spread Function (PSF), Modulation Transfer Function (MTF), Signal-to-noise Ratio (SNR), Contrast-to-noise Ratio (CNR), and Receiver Operating Curves (ROC) Point Spread Function (PSF) Recollect
HW 10. = 3.3 GPa (483,000 psi)
 HW 10 Problem 15.1 Elastic modulus and tensile strength of poly(methyl methacrylate) at room temperature [20 C (68 F)]. Compare these with the corresponding values in Table 15.1. Figure 15.3 is accurate;
HW 10 Problem 15.1 Elastic modulus and tensile strength of poly(methyl methacrylate) at room temperature [20 C (68 F)]. Compare these with the corresponding values in Table 15.1. Figure 15.3 is accurate;
Diffusione e perfusione in risonanza magnetica. E. Pagani, M. Filippi
 Diffusione e perfusione in risonanza magnetica E. Pagani, M. Filippi DW-MRI DIFFUSION-WEIGHTED MRI Principles Diffusion results from a microspic random motion known as Brownian motion THE RANDOM WALK How
Diffusione e perfusione in risonanza magnetica E. Pagani, M. Filippi DW-MRI DIFFUSION-WEIGHTED MRI Principles Diffusion results from a microspic random motion known as Brownian motion THE RANDOM WALK How
1 Example of Time Series Analysis by SSA 1
 1 Example of Time Series Analysis by SSA 1 Let us illustrate the 'Caterpillar'-SSA technique [1] by the example of time series analysis. Consider the time series FORT (monthly volumes of fortied wine sales
1 Example of Time Series Analysis by SSA 1 Let us illustrate the 'Caterpillar'-SSA technique [1] by the example of time series analysis. Consider the time series FORT (monthly volumes of fortied wine sales
AP Physics B Ch. 23 and Ch. 24 Geometric Optics and Wave Nature of Light
 AP Physics B Ch. 23 and Ch. 24 Geometric Optics and Wave Nature of Light Name: Period: Date: MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Reflection,
AP Physics B Ch. 23 and Ch. 24 Geometric Optics and Wave Nature of Light Name: Period: Date: MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Reflection,
A disaccharide is formed when a dehydration reaction joins two monosaccharides. This covalent bond is called a glycosidic linkage.
 CH 5 Structure & Function of Large Molecules: Macromolecules Molecules of Life All living things are made up of four classes of large biological molecules: carbohydrates, lipids, proteins, and nucleic
CH 5 Structure & Function of Large Molecules: Macromolecules Molecules of Life All living things are made up of four classes of large biological molecules: carbohydrates, lipids, proteins, and nucleic
Customer Training Material. Lecture 4. Meshing in Mechanical. Mechanical. ANSYS, Inc. Proprietary 2010 ANSYS, Inc. All rights reserved.
 Lecture 4 Meshing in Mechanical Introduction to ANSYS Mechanical L4-1 Chapter Overview In this chapter controlling meshing operations is described. Topics: A. Global Meshing Controls B. Local Meshing Controls
Lecture 4 Meshing in Mechanical Introduction to ANSYS Mechanical L4-1 Chapter Overview In this chapter controlling meshing operations is described. Topics: A. Global Meshing Controls B. Local Meshing Controls
ALLEN Mouse Brain Atlas
 TECHNICAL WHITE PAPER: QUALITY CONTROL STANDARDS FOR HIGH-THROUGHPUT RNA IN SITU HYBRIDIZATION DATA GENERATION Consistent data quality and internal reproducibility are critical concerns for high-throughput
TECHNICAL WHITE PAPER: QUALITY CONTROL STANDARDS FOR HIGH-THROUGHPUT RNA IN SITU HYBRIDIZATION DATA GENERATION Consistent data quality and internal reproducibility are critical concerns for high-throughput
OPRE 6201 : 2. Simplex Method
 OPRE 6201 : 2. Simplex Method 1 The Graphical Method: An Example Consider the following linear program: Max 4x 1 +3x 2 Subject to: 2x 1 +3x 2 6 (1) 3x 1 +2x 2 3 (2) 2x 2 5 (3) 2x 1 +x 2 4 (4) x 1, x 2
OPRE 6201 : 2. Simplex Method 1 The Graphical Method: An Example Consider the following linear program: Max 4x 1 +3x 2 Subject to: 2x 1 +3x 2 6 (1) 3x 1 +2x 2 3 (2) 2x 2 5 (3) 2x 1 +x 2 4 (4) x 1, x 2
In mathematics, there are four attainment targets: using and applying mathematics; number and algebra; shape, space and measures, and handling data.
 MATHEMATICS: THE LEVEL DESCRIPTIONS In mathematics, there are four attainment targets: using and applying mathematics; number and algebra; shape, space and measures, and handling data. Attainment target
MATHEMATICS: THE LEVEL DESCRIPTIONS In mathematics, there are four attainment targets: using and applying mathematics; number and algebra; shape, space and measures, and handling data. Attainment target
Environmental Remote Sensing GEOG 2021
 Environmental Remote Sensing GEOG 2021 Lecture 4 Image classification 2 Purpose categorising data data abstraction / simplification data interpretation mapping for land cover mapping use land cover class
Environmental Remote Sensing GEOG 2021 Lecture 4 Image classification 2 Purpose categorising data data abstraction / simplification data interpretation mapping for land cover mapping use land cover class
Analyzing LASIK Optical Data Using Zernike Functions
 MATLAB Digest Analyzing LASIK Optical Data Using Zernike Functions By Paul Fricker Researchers in fields as diverse as optometry, astronomy, and photonics face a common challenge: how to accurately measure
MATLAB Digest Analyzing LASIK Optical Data Using Zernike Functions By Paul Fricker Researchers in fields as diverse as optometry, astronomy, and photonics face a common challenge: how to accurately measure
K'NEX DNA Models. Developed by Dr. Gary Benson Department of Biomathematical Sciences Mount Sinai School of Medicine
 KNEX DNA Models Introduction Page 1 of 11 All photos by Kevin Kelliher. To download an Acrobat pdf version of this website Click here. K'NEX DNA Models Developed by Dr. Gary Benson Department of Biomathematical
KNEX DNA Models Introduction Page 1 of 11 All photos by Kevin Kelliher. To download an Acrobat pdf version of this website Click here. K'NEX DNA Models Developed by Dr. Gary Benson Department of Biomathematical
7. DYNAMIC LIGHT SCATTERING 7.1 First order temporal autocorrelation function.
 7. DYNAMIC LIGHT SCATTERING 7. First order temporal autocorrelation function. Dynamic light scattering (DLS) studies the properties of inhomogeneous and dynamic media. A generic situation is illustrated
7. DYNAMIC LIGHT SCATTERING 7. First order temporal autocorrelation function. Dynamic light scattering (DLS) studies the properties of inhomogeneous and dynamic media. A generic situation is illustrated
MA 323 Geometric Modelling Course Notes: Day 02 Model Construction Problem
 MA 323 Geometric Modelling Course Notes: Day 02 Model Construction Problem David L. Finn November 30th, 2004 In the next few days, we will introduce some of the basic problems in geometric modelling, and
MA 323 Geometric Modelling Course Notes: Day 02 Model Construction Problem David L. Finn November 30th, 2004 In the next few days, we will introduce some of the basic problems in geometric modelling, and
Video Camera Image Quality in Physical Electronic Security Systems
 Video Camera Image Quality in Physical Electronic Security Systems Video Camera Image Quality in Physical Electronic Security Systems In the second decade of the 21st century, annual revenue for the global
Video Camera Image Quality in Physical Electronic Security Systems Video Camera Image Quality in Physical Electronic Security Systems In the second decade of the 21st century, annual revenue for the global
Multiple Optimization Using the JMP Statistical Software Kodak Research Conference May 9, 2005
 Multiple Optimization Using the JMP Statistical Software Kodak Research Conference May 9, 2005 Philip J. Ramsey, Ph.D., Mia L. Stephens, MS, Marie Gaudard, Ph.D. North Haven Group, http://www.northhavengroup.com/
Multiple Optimization Using the JMP Statistical Software Kodak Research Conference May 9, 2005 Philip J. Ramsey, Ph.D., Mia L. Stephens, MS, Marie Gaudard, Ph.D. North Haven Group, http://www.northhavengroup.com/
OBJECT TRACKING USING LOG-POLAR TRANSFORMATION
 OBJECT TRACKING USING LOG-POLAR TRANSFORMATION A Thesis Submitted to the Gradual Faculty of the Louisiana State University and Agricultural and Mechanical College in partial fulfillment of the requirements
OBJECT TRACKING USING LOG-POLAR TRANSFORMATION A Thesis Submitted to the Gradual Faculty of the Louisiana State University and Agricultural and Mechanical College in partial fulfillment of the requirements
Keysight Technologies How to Choose your MAC Lever. Technical Overview
 Keysight Technologies How to Choose your MAC Lever Technical Overview Introduction Atomic force microscopy (AFM) is a sub-nanometer scale imaging and measurement tool that can be used to determine a sample
Keysight Technologies How to Choose your MAC Lever Technical Overview Introduction Atomic force microscopy (AFM) is a sub-nanometer scale imaging and measurement tool that can be used to determine a sample
Palmprint Recognition. By Sree Rama Murthy kora Praveen Verma Yashwant Kashyap
 Palmprint Recognition By Sree Rama Murthy kora Praveen Verma Yashwant Kashyap Palm print Palm Patterns are utilized in many applications: 1. To correlate palm patterns with medical disorders, e.g. genetic
Palmprint Recognition By Sree Rama Murthy kora Praveen Verma Yashwant Kashyap Palm print Palm Patterns are utilized in many applications: 1. To correlate palm patterns with medical disorders, e.g. genetic
Compensation Basics - Bagwell. Compensation Basics. C. Bruce Bagwell MD, Ph.D. Verity Software House, Inc.
 Compensation Basics C. Bruce Bagwell MD, Ph.D. Verity Software House, Inc. 2003 1 Intrinsic or Autofluorescence p2 ac 1,2 c 1 ac 1,1 p1 In order to describe how the general process of signal cross-over
Compensation Basics C. Bruce Bagwell MD, Ph.D. Verity Software House, Inc. 2003 1 Intrinsic or Autofluorescence p2 ac 1,2 c 1 ac 1,1 p1 In order to describe how the general process of signal cross-over
Diagrams and Graphs of Statistical Data
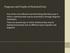 Diagrams and Graphs of Statistical Data One of the most effective and interesting alternative way in which a statistical data may be presented is through diagrams and graphs. There are several ways in
Diagrams and Graphs of Statistical Data One of the most effective and interesting alternative way in which a statistical data may be presented is through diagrams and graphs. There are several ways in
Summarizing and Displaying Categorical Data
 Summarizing and Displaying Categorical Data Categorical data can be summarized in a frequency distribution which counts the number of cases, or frequency, that fall into each category, or a relative frequency
Summarizing and Displaying Categorical Data Categorical data can be summarized in a frequency distribution which counts the number of cases, or frequency, that fall into each category, or a relative frequency
Disulfide Bonds at the Hair Salon
 Disulfide Bonds at the Hair Salon Three Alpha Helices Stabilized By Disulfide Bonds! In order for hair to grow 6 inches in one year, 9 1/2 turns of α helix must be produced every second!!! In some proteins,
Disulfide Bonds at the Hair Salon Three Alpha Helices Stabilized By Disulfide Bonds! In order for hair to grow 6 inches in one year, 9 1/2 turns of α helix must be produced every second!!! In some proteins,
Grade 6 Mathematics Assessment. Eligible Texas Essential Knowledge and Skills
 Grade 6 Mathematics Assessment Eligible Texas Essential Knowledge and Skills STAAR Grade 6 Mathematics Assessment Mathematical Process Standards These student expectations will not be listed under a separate
Grade 6 Mathematics Assessment Eligible Texas Essential Knowledge and Skills STAAR Grade 6 Mathematics Assessment Mathematical Process Standards These student expectations will not be listed under a separate
Basic 3D reconstruction in Imaris 7.6.1
 Basic 3D reconstruction in Imaris 7.6.1 Task The aim of this tutorial is to understand basic Imaris functionality by performing surface reconstruction of glia cells in culture, in order to visualize enclosed
Basic 3D reconstruction in Imaris 7.6.1 Task The aim of this tutorial is to understand basic Imaris functionality by performing surface reconstruction of glia cells in culture, in order to visualize enclosed
The Fourier Analysis Tool in Microsoft Excel
 The Fourier Analysis Tool in Microsoft Excel Douglas A. Kerr Issue March 4, 2009 ABSTRACT AD ITRODUCTIO The spreadsheet application Microsoft Excel includes a tool that will calculate the discrete Fourier
The Fourier Analysis Tool in Microsoft Excel Douglas A. Kerr Issue March 4, 2009 ABSTRACT AD ITRODUCTIO The spreadsheet application Microsoft Excel includes a tool that will calculate the discrete Fourier
Section A. Index. Section A. Planning, Budgeting and Forecasting Section A.2 Forecasting techniques... 1. Page 1 of 11. EduPristine CMA - Part I
 Index Section A. Planning, Budgeting and Forecasting Section A.2 Forecasting techniques... 1 EduPristine CMA - Part I Page 1 of 11 Section A. Planning, Budgeting and Forecasting Section A.2 Forecasting
Index Section A. Planning, Budgeting and Forecasting Section A.2 Forecasting techniques... 1 EduPristine CMA - Part I Page 1 of 11 Section A. Planning, Budgeting and Forecasting Section A.2 Forecasting
