Supporting Information
|
|
|
- Kelly Potter
- 8 years ago
- Views:
Transcription
1 Supporting Information Wiley-VCH Weinheim, Germany
2 Blockage of sirna Binding to a p19 Viral Suppressor of RNA Silencing by Cysteine Alkylation Selena M. Sagan, Roger Koukiekolo, Elisabeth Rodgers, Natalie K. Goto, John Paul Pezacki* Codon Optimization of CIRV p19 CIRV p19 codon optimization for mammalian and bacterial expression was done manually using codon usage tables for Homo sapiens and Escherichia coli obtained from the Genbank Codon Usage Database at and the Carnation Italian Ringspot virus genome (Genbank Accession #: NC_35). Chosen codons had a frequency score >1 for both Homo sapiens and Escherichia coli. Sequence of Codon Optimized CIRV p19 [ ] S.M. Sagan, Dr. R. Koukiekolo, E. Rodgers, Prof. J.P. Pezacki, The Steacie Institute for Molecular Sciences, National Research Council of Canada, 1 Sussex Drive, Ottawa, K1A R6 (Canada) Fax: (+1) John.Pezacki@nrc-cnrc.gc.ca [ ] These authors contributed equally to the work. Prof. N.K. Goto, Prof. J.P. Pezacki, Department of Chemistry, University of Ottawa, Ottawa, Canada, K1N 6N5 Selena M. Sagan, Prof. Natalie K. Goto, Prof. John Paul Pezacki, Department of Biochemistry, Microbiology & Immunology/Ottawa Institute of Systems Biology, University of Ottawa, Ottawa, Canada K1H 8M5 1
3 atggaacgcgctatccaaggcaacgacactcgcgaacaagctaacggtgaacgctgggatggcggctccggcggtatcacttctccattcaaactgcctg acgaaagcccaagctggactgagtggcgcctgtataacgatgagaccaattccaatcaagataatccactgggtttcaaggaaagctggggtttcgggaaa gttgtctttaagcgctatctgcgctacgaccgcaccgaagcttccctgcaccgcgtcctgggctcttggaccggcgattccgttaactatgcagcatctcgcttt ctgggtgccaaccaggtcggctgtacctatagcattcgctttcgcggcgttagcgtcaccatttctggcgggtcccgcactctgcagcatctgtgtgagatgg caattcgctctaagcaagaactgctgcagctgaccccagtcgaagtggaaagcaatgtgtcccgcggctgccctgaaggtattgaaaccttcaagaaagaa agcgagtaa sirna Sequences All sirnas were duplex RNA unless indicated and were purchased from Dharmacon (Lafayette, CO). sirna sequences used in this study are as follows: Cy3-GL2 19-nt sirna 5 -Cy3- UACGCGGAAUACUUCGAdTdT-3 ; unlabeled GL2 21-nt sirna 5 - CGUACGCGGAAUACUUCGAdTdT-3 ; Cy3-GL2 21-nt sirna 5 -Cy3- CGUACGCGGAAUACUUCGAdTdT-3 ; Cy3-GL2 21-nt single-stranded sirna 5 -Cy3- CGUACGCGGAAUACUUCGAdTdT-3 ; Cy3-GL2 23-nt sirna 5 -Cy3- CGUACGCGGAAUACUUCGAAAdTdT-3 ; Cy3-GL2 25-nt sirna 5 -Cy3- UCACGUACGCGGAAUACUUCGAAdTdT-3 ; Cy3-GL2 28-nt sirna 5 -Cy3- ACAUCACGUACGCGGAAUACUUCGAAdTdT-3 ; Cy3-GL3 21-nt sirna 5 -Cy3- CUUACGCUGAGUACUUCGAdTdT-3 ; Cy3-CSK-2 21-nt sirna 5 -Cy3- CUACCGCAUCAUGUACCAUdTdT-3 ; Cy nt sirna 5 - GGUCUCGUAGACCGUGCACdTdT-3. 2
4 A 6 Relative fluorescence intensity sirna (nm) 4 21-nt GL2 sirna B Relative fluorescence intensity C sirna (nm) 4 21-nt CSK-2 sirna Relative fluorescence intensity sirna (nm) 4 21-nt 331 sirna Figure S1. Plots of relative fluorescence vs. sirna concentration for: (A) 21-nt GL2 sirna (K d = 74 nm), (B) 21-nt CSK-2 sirna (K d =57 nm), and (C) 21-nt 331 sirna (K d ~ 12 nm). Each point represents the mean fluoresence of three experiments, with the error bars showing the standard deviation from the mean. Curves obtained from fitting these points to Equation [1] to obtain the indicated K d values are also shown. 3
5 A B Figure S2. (A) Fluorescence measurements for the binding activity of GL2 sirna to CIRV p19-h in the presence of a representative sampling of the small molecule library used to identify compounds 1 and 2. Relative reduction in sirna binding by each compound was measured upon addition of 1 µm to a well where sirna was already bound to p19. Each measurement was performed in triplicate. (B) The decrease in fluorescence measured when an equivalent amount of unlabeled sirna (CSK-2) was added to the mixture of p19 bound with labeled sirna (Cy3- CSK-2). Means ± SE are presented (the error bar for Cy3-CSK-2 is too small to be visualized). Screen hits were selected based on statistical analyses of fluorescence intensity differences between compound treated and mock treated wells (each an average of three measurements). Unlabeled sirna that reduces the fluorescence signal by competitive binding to p19 served as a positive control. Two-tailed P-values of t-tests for the screening data were determined and P- values <.5 was set as the threshold for significance. Compounds 1 and 2, corresponding to entries 61 and 67 above, respectively, both showed P-values <.1. 4
6 Figure S3. Concentration-dependent inhibition of the CSK-2 sirna:p19 interaction by compound 1; with 1 added after the saturating amounts of sirna were added to arrayed CIRV p19-h. Log (IC 5 ) = /-.6 M. The curve was determined bit fitting to a sigmoidal dose-response (variable slope) equation. 5
7 Figure S4. Relative fluorescence intensity obtained from N-ethyl maleimide treated and untreated CIRV p19-h binding to fluorescently tagged GL2 21-nt sirna. Each bar represents the average of three measurements with the error provided by the standard deviation from the mean. (The magnitude of the error is too small to be visualized for the N-ethyl maleimide treated samples.) 6
8 A B C Figure S5. MALDI-TOF spectra for p19 treated with compound 1. A) p19 alone; B) p19 treated with a 1:1 molar ratio of 1; inset shows an expanded mass spectrum for p19 treated with a 1:1 ratio of 1. The peak with m/z = corresponds to unmodified p19 and the peak with m/z = corresponds to p19 doubly modified by 1, within experimental error. 7
9 A B Figure S6. MALDI-TOF spectra for p19 treated with compound 3. A) p19 alone; B) p19 treated with 1:1 ratio of 3:p19. The peak with m/z = 4144 in A) corresponds to unmodified p19 and the peak with m/z = corresponds to p19 containing four modifications by 3 (two modifications per p19 monomer), within experimental error. 8
標 準 納 期 2 週 間 を 誇 るKINOMEscan 受 託 試 験 : 報 告 書 から 抽 出 できる 更 なるデータ? 日 本 事 業 担 当 大 嶺 青 河 DiscoveRx Corporation
 標 準 納 期 2 週 間 を 誇 るKINOMEscan 受 託 試 験 : 報 告 書 から 抽 出 できる 更 なるデータ? 日 本 事 業 担 当 大 嶺 青 河 DiscoveRx Corporation DiscoveRx Corporation Fremont, CA San Francisco, CA San Diego, CA DiscoveRx Corporation 3 KINOMEscan
標 準 納 期 2 週 間 を 誇 るKINOMEscan 受 託 試 験 : 報 告 書 から 抽 出 できる 更 なるデータ? 日 本 事 業 担 当 大 嶺 青 河 DiscoveRx Corporation DiscoveRx Corporation Fremont, CA San Francisco, CA San Diego, CA DiscoveRx Corporation 3 KINOMEscan
*Corresponding author: Yoshitaka Nishiyama, E-mail: nishiyama@molbiol.saitama-u.ac.jp
 SUPPLEMENTAL DATA Elongation Factor G Is a Critical Target during Oxidative Damage to the Translation System of Escherichia coli Takanori Nagano 1, Kouji Kojima 1,7, Toru Hisabori 2, Hidenori Hayashi 3,
SUPPLEMENTAL DATA Elongation Factor G Is a Critical Target during Oxidative Damage to the Translation System of Escherichia coli Takanori Nagano 1, Kouji Kojima 1,7, Toru Hisabori 2, Hidenori Hayashi 3,
ProteinPilot Report for ProteinPilot Software
 ProteinPilot Report for ProteinPilot Software Detailed Analysis of Protein Identification / Quantitation Results Automatically Sean L Seymour, Christie Hunter SCIEX, USA Pow erful mass spectrometers like
ProteinPilot Report for ProteinPilot Software Detailed Analysis of Protein Identification / Quantitation Results Automatically Sean L Seymour, Christie Hunter SCIEX, USA Pow erful mass spectrometers like
岑 祥 股 份 有 限 公 司 技 術 專 員 費 軫 尹 20100803
 技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
Molecular Spectroscopy
 Molecular Spectroscopy UV-Vis Spectroscopy Absorption Characteristics of Some Common Chromophores UV-Vis Spectroscopy Absorption Characteristics of Aromatic Compounds UV-Vis Spectroscopy Effect of extended
Molecular Spectroscopy UV-Vis Spectroscopy Absorption Characteristics of Some Common Chromophores UV-Vis Spectroscopy Absorption Characteristics of Aromatic Compounds UV-Vis Spectroscopy Effect of extended
Real-time PCR: Understanding C t
 APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
Data Analysis on the ABI PRISM 7700 Sequence Detection System: Setting Baselines and Thresholds. Overview. Data Analysis Tutorial
 Data Analysis on the ABI PRISM 7700 Sequence Detection System: Setting Baselines and Thresholds Overview In order for accuracy and precision to be optimal, the assay must be properly evaluated and a few
Data Analysis on the ABI PRISM 7700 Sequence Detection System: Setting Baselines and Thresholds Overview In order for accuracy and precision to be optimal, the assay must be properly evaluated and a few
Essentials of Real Time PCR. About Sequence Detection Chemistries
 Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Application of Automated Data Collection to Surface-Enhanced Raman Scattering (SERS)
 Application Note: 52020 Application of Automated Data Collection to Surface-Enhanced Raman Scattering (SERS) Timothy O. Deschaines, Ph.D., Thermo Fisher Scientific, Madison, WI, USA Key Words Array Automation
Application Note: 52020 Application of Automated Data Collection to Surface-Enhanced Raman Scattering (SERS) Timothy O. Deschaines, Ph.D., Thermo Fisher Scientific, Madison, WI, USA Key Words Array Automation
ph: Measurement and Uses
 ph: Measurement and Uses One of the most important properties of aqueous solutions is the concentration of hydrogen ion. The concentration of H + (or H 3 O + ) affects the solubility of inorganic and organic
ph: Measurement and Uses One of the most important properties of aqueous solutions is the concentration of hydrogen ion. The concentration of H + (or H 3 O + ) affects the solubility of inorganic and organic
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual
 Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Next Generation Sequencing
 Next Generation Sequencing DNA sequence represents a single format onto which a broad range of biological phenomena can be projected for high-throughput data collection Over the past three years, massively
Next Generation Sequencing DNA sequence represents a single format onto which a broad range of biological phenomena can be projected for high-throughput data collection Over the past three years, massively
Project 5: Scoville Heat Value of Foods HPLC Analysis of Capsaicinoids
 Willamette University Chemistry Department 2013 Project 5: HPLC Analysis of Capsaicinoids LABORATORY REPORT: Formal Writing Exercises PRE-LAB ASSIGNMENT Read the entire laboratory project and section 28C
Willamette University Chemistry Department 2013 Project 5: HPLC Analysis of Capsaicinoids LABORATORY REPORT: Formal Writing Exercises PRE-LAB ASSIGNMENT Read the entire laboratory project and section 28C
Introduction to Proteomics
 Introduction to Proteomics Åsa Wheelock, Ph.D. Division of Respiratory Medicine & Karolinska Biomics Center asa.wheelock@ki.se In: Systems Biology and the Omics Cascade, Karolinska Institutet, June 9-13,
Introduction to Proteomics Åsa Wheelock, Ph.D. Division of Respiratory Medicine & Karolinska Biomics Center asa.wheelock@ki.se In: Systems Biology and the Omics Cascade, Karolinska Institutet, June 9-13,
Bioruptor NGS: Unbiased DNA shearing for Next-Generation Sequencing
 STGAAC STGAACT GTGCACT GTGAACT STGAAC STGAACT GTGCACT GTGAACT STGAAC STGAAC GTGCAC GTGAAC Wouter Coppieters Head of the genomics core facility GIGA center, University of Liège Bioruptor NGS: Unbiased DNA
STGAAC STGAACT GTGCACT GTGAACT STGAAC STGAACT GTGCACT GTGAACT STGAAC STGAAC GTGCAC GTGAAC Wouter Coppieters Head of the genomics core facility GIGA center, University of Liège Bioruptor NGS: Unbiased DNA
Turning HTE Agilent LC/MS Data to Knowledge in 1hr
 With permission Reaction Analytics Inc. 2011 Page 1 Turning HTE Agilent LC/MS Data to Knowledge in 1hr Neal Sach With permission Reaction Analytics Inc. 2011 Page 2 Contents Acquiring Data Fast demo method
With permission Reaction Analytics Inc. 2011 Page 1 Turning HTE Agilent LC/MS Data to Knowledge in 1hr Neal Sach With permission Reaction Analytics Inc. 2011 Page 2 Contents Acquiring Data Fast demo method
Application Note # LCMS-62 Walk-Up Ion Trap Mass Spectrometer System in a Multi-User Environment Using Compass OpenAccess Software
 Application Note # LCMS-62 Walk-Up Ion Trap Mass Spectrometer System in a Multi-User Environment Using Compass OpenAccess Software Abstract Presented here is a case study of a walk-up liquid chromatography
Application Note # LCMS-62 Walk-Up Ion Trap Mass Spectrometer System in a Multi-User Environment Using Compass OpenAccess Software Abstract Presented here is a case study of a walk-up liquid chromatography
A new concept for isotope ratio monitoring LC/MS. A Wide Range of Applications
 A new concept for isotope ratio monitoring LC/MS A Wide Range of Applications Andreas Hilkert, Dieter Juchelka, Michael Krummen Thermo Electron (Bremen) GmbH Overview Introduction in Isotope Ratio Monitoring-LC/MS
A new concept for isotope ratio monitoring LC/MS A Wide Range of Applications Andreas Hilkert, Dieter Juchelka, Michael Krummen Thermo Electron (Bremen) GmbH Overview Introduction in Isotope Ratio Monitoring-LC/MS
Determining the Identity of an Unknown Weak Acid
 Purpose The purpose of this experiment is to observe and measure a weak acid neutralization and determine the identity of an unknown acid by titration. Introduction The purpose of this exercise is to identify
Purpose The purpose of this experiment is to observe and measure a weak acid neutralization and determine the identity of an unknown acid by titration. Introduction The purpose of this exercise is to identify
Thermodynamics of Mixing
 Thermodynamics of Mixing Dependence of Gibbs energy on mixture composition is G = n A µ A + n B µ B and at constant T and p, systems tend towards a lower Gibbs energy The simplest example of mixing: What
Thermodynamics of Mixing Dependence of Gibbs energy on mixture composition is G = n A µ A + n B µ B and at constant T and p, systems tend towards a lower Gibbs energy The simplest example of mixing: What
2.500 Threshold. 2.000 1000e - 001. Threshold. Exponential phase. Cycle Number
 application note Real-Time PCR: Understanding C T Real-Time PCR: Understanding C T 4.500 3.500 1000e + 001 4.000 3.000 1000e + 000 3.500 2.500 Threshold 3.000 2.000 1000e - 001 Rn 2500 Rn 1500 Rn 2000
application note Real-Time PCR: Understanding C T Real-Time PCR: Understanding C T 4.500 3.500 1000e + 001 4.000 3.000 1000e + 000 3.500 2.500 Threshold 3.000 2.000 1000e - 001 Rn 2500 Rn 1500 Rn 2000
Introduction To Real Time Quantitative PCR (qpcr)
 Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
ASMS Regulated Bioanalysis Interest Group (RBIG) Workshop. Antibody-Drug Conjugates (ADC) A Complex Problem in Regulated Bioanalysis.
 ASMS Regulated Bioanalysis Interest Group (RBIG) Workshop Antibody-Drug Conjugates (ADC) A Complex Problem in Regulated Bioanalysis June 17, 2014 Presiding: Dr. Keyang Xu (Genentech) and Dr. Fabio Garofolo
ASMS Regulated Bioanalysis Interest Group (RBIG) Workshop Antibody-Drug Conjugates (ADC) A Complex Problem in Regulated Bioanalysis June 17, 2014 Presiding: Dr. Keyang Xu (Genentech) and Dr. Fabio Garofolo
Introduction. Preparation of Template DNA
 Procedures and Recommendations for DNA Sequencing at the Plant-Microbe Genomics Facility Ohio State University Biological Sciences Building Room 420, 484 W. 12th Ave., Columbus OH 43210 Telephone: 614/247-6204;
Procedures and Recommendations for DNA Sequencing at the Plant-Microbe Genomics Facility Ohio State University Biological Sciences Building Room 420, 484 W. 12th Ave., Columbus OH 43210 Telephone: 614/247-6204;
MicroRNA formation. 4th International Symposium on Non-Surgical Contraceptive Methods of Pet Population Control
 MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
Pesticide Analysis by Mass Spectrometry
 Pesticide Analysis by Mass Spectrometry Purpose: The purpose of this assignment is to introduce concepts of mass spectrometry (MS) as they pertain to the qualitative and quantitative analysis of organochlorine
Pesticide Analysis by Mass Spectrometry Purpose: The purpose of this assignment is to introduce concepts of mass spectrometry (MS) as they pertain to the qualitative and quantitative analysis of organochlorine
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
Supporting Information. Minimum active structure of insulin-like. peptide 5 (INSL5)
 Supporting Information Minimum active structure of insulin-like peptide 5 (INSL5) Alessia Belgi 1,2, Ross A.D. Bathgate *1,2,3, Martina Kocan *4, Nitin Patil 1,5, Suode Zhang 1, Geoffrey W. Tregear 1,2,
Supporting Information Minimum active structure of insulin-like peptide 5 (INSL5) Alessia Belgi 1,2, Ross A.D. Bathgate *1,2,3, Martina Kocan *4, Nitin Patil 1,5, Suode Zhang 1, Geoffrey W. Tregear 1,2,
1 st day Basic Training Course
 DATES AND LOCATIONS 13-14 April 2015 Princeton Marriott at Forrestal, 100 College Road East, Princeton NJ 08540, New Jersey 16-17 April 2015 Hotel Nikko San Francisco 222 Mason Street, San Francisco, CA
DATES AND LOCATIONS 13-14 April 2015 Princeton Marriott at Forrestal, 100 College Road East, Princeton NJ 08540, New Jersey 16-17 April 2015 Hotel Nikko San Francisco 222 Mason Street, San Francisco, CA
Determination of Molar Mass by Freezing-Point Depression
 DETERMINATION OF MOLAR MASS BY FREEZING-POINT DEPRESSION 141 Determination of Molar Mass by Freezing-Point Depression OBJECTIVES: Gain familiarity with colligative properties of nonelectrolyte solutions
DETERMINATION OF MOLAR MASS BY FREEZING-POINT DEPRESSION 141 Determination of Molar Mass by Freezing-Point Depression OBJECTIVES: Gain familiarity with colligative properties of nonelectrolyte solutions
SELDI-TOF Mass Spectrometry Protein Data By Huong Thi Dieu La
 SELDI-TOF Mass Spectrometry Protein Data By Huong Thi Dieu La References Alejandro Cruz-Marcelo, Rudy Guerra, Marina Vannucci, Yiting Li, Ching C. Lau, and Tsz-Kwong Man. Comparison of algorithms for pre-processing
SELDI-TOF Mass Spectrometry Protein Data By Huong Thi Dieu La References Alejandro Cruz-Marcelo, Rudy Guerra, Marina Vannucci, Yiting Li, Ching C. Lau, and Tsz-Kwong Man. Comparison of algorithms for pre-processing
Tutorial for Proteomics Data Submission. Katalin F. Medzihradszky Robert J. Chalkley UCSF
 Tutorial for Proteomics Data Submission Katalin F. Medzihradszky Robert J. Chalkley UCSF Why Have Guidelines? Large-scale proteomics studies create huge amounts of data. It is impossible/impractical to
Tutorial for Proteomics Data Submission Katalin F. Medzihradszky Robert J. Chalkley UCSF Why Have Guidelines? Large-scale proteomics studies create huge amounts of data. It is impossible/impractical to
Core Facility Genomics
 Core Facility Genomics versatile genome or transcriptome analyses based on quantifiable highthroughput data ascertainment 1 Topics Collaboration with Harald Binder and Clemens Kreutz Project: Microarray
Core Facility Genomics versatile genome or transcriptome analyses based on quantifiable highthroughput data ascertainment 1 Topics Collaboration with Harald Binder and Clemens Kreutz Project: Microarray
FIDA for Rapid Detection of Protein Based Biomarkers
 FIDA for Rapid Detection of Protein Based Biomarkers Guisheng Zhuang 1, Nicklas N. Poulsen 1, Nina Z. Andersen 1, Jesper Østergaard 1,2, Jørgen Schøller 2 and Henrik Jensen 1,2 1 Department of Pharmacy,
FIDA for Rapid Detection of Protein Based Biomarkers Guisheng Zhuang 1, Nicklas N. Poulsen 1, Nina Z. Andersen 1, Jesper Østergaard 1,2, Jørgen Schøller 2 and Henrik Jensen 1,2 1 Department of Pharmacy,
Review of Chemical Equilibrium 7.51 September 1999. free [A] (µm)
![Review of Chemical Equilibrium 7.51 September 1999. free [A] (µm) Review of Chemical Equilibrium 7.51 September 1999. free [A] (µm)](/thumbs/40/21435474.jpg) Review of Chemical Equilibrium 7.51 September 1999 Equilibrium experiments study how the concentration of reaction products change as a function of reactant concentrations and/or reaction conditions. For
Review of Chemical Equilibrium 7.51 September 1999 Equilibrium experiments study how the concentration of reaction products change as a function of reactant concentrations and/or reaction conditions. For
Analysis of gene expression data. Ulf Leser and Philippe Thomas
 Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
Mass Frontier 7.0 Quick Start Guide
 Mass Frontier 7.0 Quick Start Guide The topics in this guide briefly step you through key features of the Mass Frontier application. Editing a Structure Working with Spectral Trees Building a Library Predicting
Mass Frontier 7.0 Quick Start Guide The topics in this guide briefly step you through key features of the Mass Frontier application. Editing a Structure Working with Spectral Trees Building a Library Predicting
Appendix 5 Overview of requirements in English
 Appendix 5 Overview of requirements in English This document is a translation of Appendix 4 (Bilag 4) section 2. This translation is meant as a service for the bidder and in case of any differences between
Appendix 5 Overview of requirements in English This document is a translation of Appendix 4 (Bilag 4) section 2. This translation is meant as a service for the bidder and in case of any differences between
UV-Visible Spectroscopy
 UV-Visible Spectroscopy UV-Visible Spectroscopy What is UV-Visible Spectroscopy? Molecular spectroscopy that involves study of the interaction of Ultra violet (UV)-Visible radiation with molecules What
UV-Visible Spectroscopy UV-Visible Spectroscopy What is UV-Visible Spectroscopy? Molecular spectroscopy that involves study of the interaction of Ultra violet (UV)-Visible radiation with molecules What
Calculating Nucleic Acid or Protein Concentration Using the GloMax Multi+ Microplate Instrument
 Calculating Nucleic Acid or Protein Concentration Using the GloMax Multi+ Microplate Instrument Technical Note INTRODUCTION Direct measurements of nucleic acid samples at OD 260 or protein samples at OD
Calculating Nucleic Acid or Protein Concentration Using the GloMax Multi+ Microplate Instrument Technical Note INTRODUCTION Direct measurements of nucleic acid samples at OD 260 or protein samples at OD
How To Understand Enzyme Kinetics
 Chapter 12 - Reaction Kinetics In the last chapter we looked at enzyme mechanisms. In this chapter we ll see how enzyme kinetics, i.e., the study of enzyme reaction rates, can be useful in learning more
Chapter 12 - Reaction Kinetics In the last chapter we looked at enzyme mechanisms. In this chapter we ll see how enzyme kinetics, i.e., the study of enzyme reaction rates, can be useful in learning more
QPCR Applications using Stratagene s Mx Real-Time PCR Platform
 QPCR Applications using Stratagene s Mx Real-Time PCR Platform Dan Schoeffner, Ph.D Field Applications Scientist Dan.Schoeffner@Stratagene.com Tech. Services 800-894-1304 Polymerase Chain Reaction Melt
QPCR Applications using Stratagene s Mx Real-Time PCR Platform Dan Schoeffner, Ph.D Field Applications Scientist Dan.Schoeffner@Stratagene.com Tech. Services 800-894-1304 Polymerase Chain Reaction Melt
using ms based proteomics
 quantification using ms based proteomics lennart martens Computational Omics and Systems Biology Group Department of Medical Protein Research, VIB Department of Biochemistry, Ghent University Ghent, Belgium
quantification using ms based proteomics lennart martens Computational Omics and Systems Biology Group Department of Medical Protein Research, VIB Department of Biochemistry, Ghent University Ghent, Belgium
Thermo Scientific Dharmacon ON-TARGETplus sirna The Standard for sirna Specificity
 Thermo Scientific Dharmacon sirna The Standard for sirna Specificity Reduce off-targets up to 90% with the market-leading sirna Combine higher target specificity with guaranteed silencing Increase confidence
Thermo Scientific Dharmacon sirna The Standard for sirna Specificity Reduce off-targets up to 90% with the market-leading sirna Combine higher target specificity with guaranteed silencing Increase confidence
Guidance for Industry
 Guidance for Industry Q2B Validation of Analytical Procedures: Methodology November 1996 ICH Guidance for Industry Q2B Validation of Analytical Procedures: Methodology Additional copies are available from:
Guidance for Industry Q2B Validation of Analytical Procedures: Methodology November 1996 ICH Guidance for Industry Q2B Validation of Analytical Procedures: Methodology Additional copies are available from:
F321 THE STRUCTURE OF ATOMS. ATOMS Atoms consist of a number of fundamental particles, the most important are... in the nucleus of an atom
 Atomic Structure F32 TE STRUCTURE OF ATOMS ATOMS Atoms consist of a number of fundamental particles, the most important are... Mass / kg Charge / C Relative mass Relative Charge PROTON NEUTRON ELECTRON
Atomic Structure F32 TE STRUCTURE OF ATOMS ATOMS Atoms consist of a number of fundamental particles, the most important are... Mass / kg Charge / C Relative mass Relative Charge PROTON NEUTRON ELECTRON
A Streamlined Workflow for Untargeted Metabolomics
 A Streamlined Workflow for Untargeted Metabolomics Employing XCMS plus, a Simultaneous Data Processing and Metabolite Identification Software Package for Rapid Untargeted Metabolite Screening Baljit K.
A Streamlined Workflow for Untargeted Metabolomics Employing XCMS plus, a Simultaneous Data Processing and Metabolite Identification Software Package for Rapid Untargeted Metabolite Screening Baljit K.
2 Spectrophotometry and the Analysis of Riboflavin
 2 Spectrophotometry and the Analysis of Riboflavin Objectives: A) To become familiar with operating the Platereader; B) to learn how to use the Platereader in determining the absorption spectrum of a compound
2 Spectrophotometry and the Analysis of Riboflavin Objectives: A) To become familiar with operating the Platereader; B) to learn how to use the Platereader in determining the absorption spectrum of a compound
Choices, choices, choices... Which sequence database? Which modifications? What mass tolerance?
 Optimization 1 Choices, choices, choices... Which sequence database? Which modifications? What mass tolerance? Where to begin? 2 Sequence Databases Swiss-prot MSDB, NCBI nr dbest Species specific ORFS
Optimization 1 Choices, choices, choices... Which sequence database? Which modifications? What mass tolerance? Where to begin? 2 Sequence Databases Swiss-prot MSDB, NCBI nr dbest Species specific ORFS
Mass Spectrometry Signal Calibration for Protein Quantitation
 Cambridge Isotope Laboratories, Inc. www.isotope.com Proteomics Mass Spectrometry Signal Calibration for Protein Quantitation Michael J. MacCoss, PhD Associate Professor of Genome Sciences University of
Cambridge Isotope Laboratories, Inc. www.isotope.com Proteomics Mass Spectrometry Signal Calibration for Protein Quantitation Michael J. MacCoss, PhD Associate Professor of Genome Sciences University of
Comparative genomic hybridization Because arrays are more than just a tool for expression analysis
 Microarray Data Analysis Workshop MedVetNet Workshop, DTU 2008 Comparative genomic hybridization Because arrays are more than just a tool for expression analysis Carsten Friis ( with several slides from
Microarray Data Analysis Workshop MedVetNet Workshop, DTU 2008 Comparative genomic hybridization Because arrays are more than just a tool for expression analysis Carsten Friis ( with several slides from
Thermo Scientific Dharmacon sirna Libraries
 Thermo Scientific Dharmacon sirna Libraries Scientific innovation in sirna potency, specificity and delivery Superior sirna in comprehensive collections for reliable screens Market leading products for
Thermo Scientific Dharmacon sirna Libraries Scientific innovation in sirna potency, specificity and delivery Superior sirna in comprehensive collections for reliable screens Market leading products for
SUPPLEMENTARY DATA 1
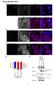 SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
Quantitative proteomics background
 Proteomics data analysis seminar Quantitative proteomics and transcriptomics of anaerobic and aerobic yeast cultures reveals post transcriptional regulation of key cellular processes de Groot, M., Daran
Proteomics data analysis seminar Quantitative proteomics and transcriptomics of anaerobic and aerobic yeast cultures reveals post transcriptional regulation of key cellular processes de Groot, M., Daran
Chemical Kinetics. Reaction Rate: The change in the concentration of a reactant or a product with time (M/s). Reactant Products A B
 Reaction Rates: Chemical Kinetics Reaction Rate: The change in the concentration of a reactant or a product with time (M/s). Reactant Products A B change in number of moles of B Average rate = change in
Reaction Rates: Chemical Kinetics Reaction Rate: The change in the concentration of a reactant or a product with time (M/s). Reactant Products A B change in number of moles of B Average rate = change in
Learning Objectives:
 Proteomics Methodology for LC-MS/MS Data Analysis Methodology for LC-MS/MS Data Analysis Peptide mass spectrum data of individual protein obtained from LC-MS/MS has to be analyzed for identification of
Proteomics Methodology for LC-MS/MS Data Analysis Methodology for LC-MS/MS Data Analysis Peptide mass spectrum data of individual protein obtained from LC-MS/MS has to be analyzed for identification of
AP CHEMISTRY 2007 SCORING GUIDELINES. Question 6
 AP CHEMISTRY 2007 SCORING GUIDELINES Question 6 Answer the following questions, which pertain to binary compounds. (a) In the box provided below, draw a complete Lewis electron-dot diagram for the IF 3
AP CHEMISTRY 2007 SCORING GUIDELINES Question 6 Answer the following questions, which pertain to binary compounds. (a) In the box provided below, draw a complete Lewis electron-dot diagram for the IF 3
Pep-Miner: A Novel Technology for Mass Spectrometry-Based Proteomics
 Pep-Miner: A Novel Technology for Mass Spectrometry-Based Proteomics Ilan Beer Haifa Research Lab Dec 10, 2002 Pep-Miner s Location in the Life Sciences World The post-genome era - the age of proteome
Pep-Miner: A Novel Technology for Mass Spectrometry-Based Proteomics Ilan Beer Haifa Research Lab Dec 10, 2002 Pep-Miner s Location in the Life Sciences World The post-genome era - the age of proteome
Indiana's Academic Standards 2010 ICP Indiana's Academic Standards 2016 ICP. map) that describe the relationship acceleration, velocity and distance.
 .1.1 Measure the motion of objects to understand.1.1 Develop graphical, the relationships among distance, velocity and mathematical, and pictorial acceleration. Develop deeper understanding through representations
.1.1 Measure the motion of objects to understand.1.1 Develop graphical, the relationships among distance, velocity and mathematical, and pictorial acceleration. Develop deeper understanding through representations
Lecture 11 Enzymes: Kinetics
 Lecture 11 Enzymes: Kinetics Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 8, pp. 216-225 Key Concepts Kinetics is the study of reaction rates (velocities). Study of enzyme kinetics is useful for
Lecture 11 Enzymes: Kinetics Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 8, pp. 216-225 Key Concepts Kinetics is the study of reaction rates (velocities). Study of enzyme kinetics is useful for
Relative Quantification of mirna Target mrnas by Real-Time qpcr. 1 Introduction. Gene Expression Application Note No. 4
 Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
A Beer s Law Experiment
 A Beer s Law Experiment Introduction There are many ways to determine concentrations of a substance in solution. So far, the only experiences you may have are acid-base titrations or possibly determining
A Beer s Law Experiment Introduction There are many ways to determine concentrations of a substance in solution. So far, the only experiences you may have are acid-base titrations or possibly determining
Gene Expression Analysis
 Gene Expression Analysis Jie Peng Department of Statistics University of California, Davis May 2012 RNA expression technologies High-throughput technologies to measure the expression levels of thousands
Gene Expression Analysis Jie Peng Department of Statistics University of California, Davis May 2012 RNA expression technologies High-throughput technologies to measure the expression levels of thousands
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
Infrared Spectroscopy: Theory
 u Chapter 15 Infrared Spectroscopy: Theory An important tool of the organic chemist is Infrared Spectroscopy, or IR. IR spectra are acquired on a special instrument, called an IR spectrometer. IR is used
u Chapter 15 Infrared Spectroscopy: Theory An important tool of the organic chemist is Infrared Spectroscopy, or IR. IR spectra are acquired on a special instrument, called an IR spectrometer. IR is used
Application of Structure-Based LC/MS Database Management for Forensic Analysis
 Cozette M. Cuppett and Michael P. Balogh Waters Corporation, Milford, MA USA Antony Williams, Vitaly Lashin and Ilya Troisky Advanced Chemistry Development, Toronto, ntario, Canada verview Application
Cozette M. Cuppett and Michael P. Balogh Waters Corporation, Milford, MA USA Antony Williams, Vitaly Lashin and Ilya Troisky Advanced Chemistry Development, Toronto, ntario, Canada verview Application
Using R for Linear Regression
 Using R for Linear Regression In the following handout words and symbols in bold are R functions and words and symbols in italics are entries supplied by the user; underlined words and symbols are optional
Using R for Linear Regression In the following handout words and symbols in bold are R functions and words and symbols in italics are entries supplied by the user; underlined words and symbols are optional
Ultraviolet Spectroscopy
 Ultraviolet Spectroscopy The wavelength of UV and visible light are substantially shorter than the wavelength of infrared radiation. The UV spectrum ranges from 100 to 400 nm. A UV-Vis spectrophotometer
Ultraviolet Spectroscopy The wavelength of UV and visible light are substantially shorter than the wavelength of infrared radiation. The UV spectrum ranges from 100 to 400 nm. A UV-Vis spectrophotometer
Next Generation Sequencing: Technology, Mapping, and Analysis
 Next Generation Sequencing: Technology, Mapping, and Analysis Gary Benson Computer Science, Biology, Bioinformatics Boston University gbenson@bu.edu http://tandem.bu.edu/ The Human Genome Project took
Next Generation Sequencing: Technology, Mapping, and Analysis Gary Benson Computer Science, Biology, Bioinformatics Boston University gbenson@bu.edu http://tandem.bu.edu/ The Human Genome Project took
Enzyme Kinetics: Properties of â-galactosidase
 Enzyme Kinetics: Properties of â-galactosidase Preparation for Laboratory: Read the introduction to this laboratory before doing the Web Tutorial - Beta Galactosidase. Additional background: Freeman, skim
Enzyme Kinetics: Properties of â-galactosidase Preparation for Laboratory: Read the introduction to this laboratory before doing the Web Tutorial - Beta Galactosidase. Additional background: Freeman, skim
Fluorescent dyes for use with the
 Detection of Multiple Reporter Dyes in Real-time, On-line PCR Analysis with the LightCycler System Gregor Sagner, Cornelia Goldstein, and Rob van Miltenburg Roche Molecular Biochemicals, Penzberg, Germany
Detection of Multiple Reporter Dyes in Real-time, On-line PCR Analysis with the LightCycler System Gregor Sagner, Cornelia Goldstein, and Rob van Miltenburg Roche Molecular Biochemicals, Penzberg, Germany
Novel Inhibitors for PRMT1 Discovered by High-Throughput Screening Using Activity-Based Fluorescence Polarization
 Novel Inhibitors for PRMT1 Discovered by High-Throughput Screening Using Activity-Based Fluorescence Polarization Myles B. C. Dillon, Daniel A. Bachovchin, Steven J. Brown, M. G. Finn, Hugh Rosen, Benjamin
Novel Inhibitors for PRMT1 Discovered by High-Throughput Screening Using Activity-Based Fluorescence Polarization Myles B. C. Dillon, Daniel A. Bachovchin, Steven J. Brown, M. G. Finn, Hugh Rosen, Benjamin
Supplemental Fig. S1. The schematic diagrams of the expression constructs used in this study.
 1 Supplemental data Supplemental Fig. S1. The schematic diagrams of the expression constructs used in this study. Supplemental Fig. S2. Ingenuity Pathway Analysis (IPA) of the 56 putative caspase substrates
1 Supplemental data Supplemental Fig. S1. The schematic diagrams of the expression constructs used in this study. Supplemental Fig. S2. Ingenuity Pathway Analysis (IPA) of the 56 putative caspase substrates
Thermo Scientific PepFinder Software A New Paradigm for Peptide Mapping
 Thermo Scientific PepFinder Software A New Paradigm for Peptide Mapping For Conclusive Characterization of Biologics Deep Protein Characterization Is Crucial Pharmaceuticals have historically been small
Thermo Scientific PepFinder Software A New Paradigm for Peptide Mapping For Conclusive Characterization of Biologics Deep Protein Characterization Is Crucial Pharmaceuticals have historically been small
Nucleic Acid Purity Assessment using A 260 /A 280 Ratios
 Nucleic Acid Purity Assessment using A 260 /A 280 Ratios A common practice in molecular biology is to perform a quick assessment of the purity of nucleic acid samples by determining the ratio of spectrophotometric
Nucleic Acid Purity Assessment using A 260 /A 280 Ratios A common practice in molecular biology is to perform a quick assessment of the purity of nucleic acid samples by determining the ratio of spectrophotometric
Spectrophotometry and the Beer-Lambert Law: An Important Analytical Technique in Chemistry
 Spectrophotometry and the Beer-Lambert Law: An Important Analytical Technique in Chemistry Jon H. Hardesty, PhD and Bassam Attili, PhD Collin College Department of Chemistry Introduction: In the last lab
Spectrophotometry and the Beer-Lambert Law: An Important Analytical Technique in Chemistry Jon H. Hardesty, PhD and Bassam Attili, PhD Collin College Department of Chemistry Introduction: In the last lab
Protein Prospector and Ways of Calculating Expectation Values
 Protein Prospector and Ways of Calculating Expectation Values 1/16 Aenoch J. Lynn; Robert J. Chalkley; Peter R. Baker; Mark R. Segal; and Alma L. Burlingame University of California, San Francisco, San
Protein Prospector and Ways of Calculating Expectation Values 1/16 Aenoch J. Lynn; Robert J. Chalkley; Peter R. Baker; Mark R. Segal; and Alma L. Burlingame University of California, San Francisco, San
EXPERIMENT 5. Molecular Absorption Spectroscopy: Determination of Iron With 1,10-Phenanthroline
 EXPERIMENT 5 Molecular Absorption Spectroscopy: Determination of Iron With 1,10-Phenanthroline UNKNOWN Submit a clean, labeled 100-mL volumetric flask to the instructor so that your unknown iron solution
EXPERIMENT 5 Molecular Absorption Spectroscopy: Determination of Iron With 1,10-Phenanthroline UNKNOWN Submit a clean, labeled 100-mL volumetric flask to the instructor so that your unknown iron solution
Calculation of Molar Masses. Molar Mass. Solutions. Solutions
 Molar Mass Molar mass = Mass in grams of one mole of any element, numerically equal to its atomic weight Molar mass of molecules can be determined from the chemical formula and molar masses of elements
Molar Mass Molar mass = Mass in grams of one mole of any element, numerically equal to its atomic weight Molar mass of molecules can be determined from the chemical formula and molar masses of elements
A Navigation through the Tracefinder Software Structure and Workflow Options. Frans Schoutsen Pesticide Symposium Prague 27 April 2015
 A Navigation through the Tracefinder Software Structure and Workflow Options Frans Schoutsen Pesticide Symposium Prague 27 April 2015 Kings day in The Netherlands 1 Index Introduction Acquisition, Method
A Navigation through the Tracefinder Software Structure and Workflow Options Frans Schoutsen Pesticide Symposium Prague 27 April 2015 Kings day in The Netherlands 1 Index Introduction Acquisition, Method
QUANTITATIVE INFRARED SPECTROSCOPY. Willard et. al. Instrumental Methods of Analysis, 7th edition, Wadsworth Publishing Co., Belmont, CA 1988, Ch 11.
 QUANTITATIVE INFRARED SPECTROSCOPY Objective: The objectives of this experiment are: (1) to learn proper sample handling procedures for acquiring infrared spectra. (2) to determine the percentage composition
QUANTITATIVE INFRARED SPECTROSCOPY Objective: The objectives of this experiment are: (1) to learn proper sample handling procedures for acquiring infrared spectra. (2) to determine the percentage composition
Copyright 2007 Casa Software Ltd. www.casaxps.com. ToF Mass Calibration
 ToF Mass Calibration Essentially, the relationship between the mass m of an ion and the time taken for the ion of a given charge to travel a fixed distance is quadratic in the flight time t. For an ideal
ToF Mass Calibration Essentially, the relationship between the mass m of an ion and the time taken for the ion of a given charge to travel a fixed distance is quadratic in the flight time t. For an ideal
Two-sample inference: Continuous data
 Two-sample inference: Continuous data Patrick Breheny April 5 Patrick Breheny STA 580: Biostatistics I 1/32 Introduction Our next two lectures will deal with two-sample inference for continuous data As
Two-sample inference: Continuous data Patrick Breheny April 5 Patrick Breheny STA 580: Biostatistics I 1/32 Introduction Our next two lectures will deal with two-sample inference for continuous data As
13C NMR Spectroscopy
 13 C NMR Spectroscopy Introduction Nuclear magnetic resonance spectroscopy (NMR) is the most powerful tool available for structural determination. A nucleus with an odd number of protons, an odd number
13 C NMR Spectroscopy Introduction Nuclear magnetic resonance spectroscopy (NMR) is the most powerful tool available for structural determination. A nucleus with an odd number of protons, an odd number
Definition of the Measurand: CRP
 A Reference Measurement System for C-reactive Protein David M. Bunk, Ph.D. Chemical Science and Technology Laboratory National Institute of Standards and Technology Definition of the Measurand: Human C-reactive
A Reference Measurement System for C-reactive Protein David M. Bunk, Ph.D. Chemical Science and Technology Laboratory National Institute of Standards and Technology Definition of the Measurand: Human C-reactive
Statistical Testing of Randomness Masaryk University in Brno Faculty of Informatics
 Statistical Testing of Randomness Masaryk University in Brno Faculty of Informatics Jan Krhovják Basic Idea Behind the Statistical Tests Generated random sequences properties as sample drawn from uniform/rectangular
Statistical Testing of Randomness Masaryk University in Brno Faculty of Informatics Jan Krhovják Basic Idea Behind the Statistical Tests Generated random sequences properties as sample drawn from uniform/rectangular
Analysis of Two Additional Signaling Molecules. in Streptomyces coelicolor and the Development. of a Novel Butyrolactone-Specific Reporter System
 Chemistry & Biology 16 Supplemental Data Analysis of Two Additional Signaling Molecules in Streptomyces coelicolor and the Development of a Novel Butyrolactone-Specific Reporter System Nai-Hua Hsiao, Satoshi
Chemistry & Biology 16 Supplemental Data Analysis of Two Additional Signaling Molecules in Streptomyces coelicolor and the Development of a Novel Butyrolactone-Specific Reporter System Nai-Hua Hsiao, Satoshi
How To Make A Molecular Encoder And Decoder
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Supporting information Molecular Encoder - Decoder Based on Assembly of Graphene with Dye-labeled
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2014 Supporting information Molecular Encoder - Decoder Based on Assembly of Graphene with Dye-labeled
M.Sc. in Nano Technology with specialisation in Nano Biotechnology
 M.Sc. in Nano Technology with specialisation in Nano Biotechnology Nanotechnology is all about designing, fabricating and controlling materials, components and machinery with dimensions on the nanoscale,
M.Sc. in Nano Technology with specialisation in Nano Biotechnology Nanotechnology is all about designing, fabricating and controlling materials, components and machinery with dimensions on the nanoscale,
Name Class Date. Figure 13 1. 2. Which nucleotide in Figure 13 1 indicates the nucleic acid above is RNA? a. uracil c. cytosine b. guanine d.
 13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
L. Yu et al. Correspondence to: Q. Zhang (dkwzhang@ucdavis.edu)
 Supplement of Atmos. Chem. Phys. Discuss., 5, 967 974, 5 http://www.atmos-chem-phys-discuss.net/5/967/5/ doi:.594/acpd-5-967-5-supplement Author(s) 5. CC Attribution. License. Supplement of Molecular transformations
Supplement of Atmos. Chem. Phys. Discuss., 5, 967 974, 5 http://www.atmos-chem-phys-discuss.net/5/967/5/ doi:.594/acpd-5-967-5-supplement Author(s) 5. CC Attribution. License. Supplement of Molecular transformations
How to Biotinylate with Reproducible Results
 How to Biotinylate with Reproducible Results Introduction The Biotin Streptavidin system continues to be used in many protein based biological research applications including; ELISAs, immunoprecipitation,
How to Biotinylate with Reproducible Results Introduction The Biotin Streptavidin system continues to be used in many protein based biological research applications including; ELISAs, immunoprecipitation,
EXPERIMENT 11 UV/VIS Spectroscopy and Spectrophotometry: Spectrophotometric Analysis of Potassium Permanganate Solutions.
 EXPERIMENT 11 UV/VIS Spectroscopy and Spectrophotometry: Spectrophotometric Analysis of Potassium Permanganate Solutions. Outcomes After completing this experiment, the student should be able to: 1. Prepare
EXPERIMENT 11 UV/VIS Spectroscopy and Spectrophotometry: Spectrophotometric Analysis of Potassium Permanganate Solutions. Outcomes After completing this experiment, the student should be able to: 1. Prepare
Technical Note. Roche Applied Science. No. LC 18/2004. Assay Formats for Use in Real-Time PCR
 Roche Applied Science Technical Note No. LC 18/2004 Purpose of this Note Assay Formats for Use in Real-Time PCR The LightCycler Instrument uses several detection channels to monitor the amplification of
Roche Applied Science Technical Note No. LC 18/2004 Purpose of this Note Assay Formats for Use in Real-Time PCR The LightCycler Instrument uses several detection channels to monitor the amplification of
Measuring Protein Concentration through Absorption Spectrophotometry
 Measuring Protein Concentration through Absorption Spectrophotometry In this lab exercise you will learn how to homogenize a tissue to extract the protein, and then how to use a protein assay reagent to
Measuring Protein Concentration through Absorption Spectrophotometry In this lab exercise you will learn how to homogenize a tissue to extract the protein, and then how to use a protein assay reagent to
MultiQuant Software 2.0 for Targeted Protein / Peptide Quantification
 MultiQuant Software 2.0 for Targeted Protein / Peptide Quantification Gold Standard for Quantitative Data Processing Because of the sensitivity, selectivity, speed and throughput at which MRM assays can
MultiQuant Software 2.0 for Targeted Protein / Peptide Quantification Gold Standard for Quantitative Data Processing Because of the sensitivity, selectivity, speed and throughput at which MRM assays can
Procedures For DNA Sequencing
 Procedures For DNA Sequencing Plant-Microbe Genomics Facility (PMGF) Ohio State University 420 Biological Sciences Building 484 W. 12th Ave., Columbus OH 43210 Telephone: 614/247-6204 FAX: 614/292-6337
Procedures For DNA Sequencing Plant-Microbe Genomics Facility (PMGF) Ohio State University 420 Biological Sciences Building 484 W. 12th Ave., Columbus OH 43210 Telephone: 614/247-6204 FAX: 614/292-6337
VALIDATION OF ANALYTICAL PROCEDURES: TEXT AND METHODOLOGY Q2(R1)
 INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE VALIDATION OF ANALYTICAL PROCEDURES: TEXT AND METHODOLOGY
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE VALIDATION OF ANALYTICAL PROCEDURES: TEXT AND METHODOLOGY
Significance, Meaning and Confidence Intervals
 Significance, Meaning and Confidence Intervals Paul Cohen ISTA 370 April, 2012 Paul Cohen ISTA 370 () Significance, Meaning and Confidence Intervals April, 2012 1 / 18 Significance vs. Meaning Significance
Significance, Meaning and Confidence Intervals Paul Cohen ISTA 370 April, 2012 Paul Cohen ISTA 370 () Significance, Meaning and Confidence Intervals April, 2012 1 / 18 Significance vs. Meaning Significance
Eco TM 48 Real Time PCR System. Fastest block-based qpcr. Four colour multiplexing. Supports all chemistries. Fast cycling protocols
 A Bibby Scientific Company Eco TM 48 Real Time PCR System Fast cycling Fastest block-base HRM functionality Four colour multiplexing MIQE compliant Easy to use software Supports all chemistries Small footprint
A Bibby Scientific Company Eco TM 48 Real Time PCR System Fast cycling Fastest block-base HRM functionality Four colour multiplexing MIQE compliant Easy to use software Supports all chemistries Small footprint
