Corresponding position of sirna GS
|
|
|
- Lester Reynolds
- 8 years ago
- Views:
Transcription
1 A C GS Target transcript 5ʼ- - 3ʼ 3ʼ ʼ sivim-270 3ʼUTR Corresponding position of sirna GS E Homologous position sivim-270 3'UTR CDS 3'UTR CDS sicltc ʼUTR sicltc-2416 D CDS F CDS Supplementary Figure S1. Microarray transcript profiles showing downregulation by sivim-270 (C, D) and sicltc-2416 (E, F) 24 h after transfection. (A) Transcripts possessing the complementarity to a 7-bp-long GS sequence of a given sirna were divided into15 groups based on the position of the complementary sequence in the sirna GS. Transcripts labelled with 1-7 possess the complementarity to nucleotides 1-7 of sirna GS and vice versa. Horizontal arrow, transcript group with seed complementarity. () The number of transcripts belonging to each group, which is further subdivided into 3 UTR and CDS based on the location of the complementary sequence on mrna. Transcripts possessing the GS complementarity in 5 UTR were neglected. (C, D, E, F) Change in gene expression is shown by log2 of fold change ratio to mock transfection. Arrow, cumulative distribution of transcript group labelled with 2-8. Cumulative distribution of 15 groups are colored as shown in (A). Note that the groups of transcripts labelled with 2-8/3 UTR are the most sensitive to the off-target effects due to sivim-270 or sicltc-2416, suggesting GS nucleotides 2-8 to serve as a seed. Results of Wilcoxon s rank-sum test for seed-dependent off-target effect is as follows: transcripts with 3 UTR complementary to the sivim-270 seed, p ; transcripts with CDS complementary to the sivim-270 seed, p ; transcripts with 3 UTR complementary to the sicltc-2416 seed, p ; transcripts with CDS complementary to the sicltc-2416 seed, p Differences in mean value between normalized log2 fold change of mrna with seed match and that of all mrna are: 0.20 ± 0.01 (C), 0.08 ± 0.01 (D), 0.44 ± 0.04 (E), and 0.12 ± 0.02 (F).
2 A psicheck Target transcript VIM-270-cm: 5 -CGCCAUCAACACCGAGUUCAA-3 sivim-270: 3 -GCGGUAGUUGUGGCUCAAGUU-5 VIM-812-cm: 5 -ACGUACGUCAGCAAUAUGAAA-3 sivim-812: 3 -UGCAUGCAGUCGUUAUACUUU-5 Target GS Target GS C : D : : E : : : F G h I : : : : : : : : : J K L M : : : : : N O P Q Supplementary Figure S2. Effects on gene silencing of seed/non-seed swapping. Using sivim-270 and sivim- 812, swap sirna constructs were generated between the seed with the 5 end (nucleotide position 1-8) and nonseed region (9-21). Similarly, corresponding swap target oligonucleotides were constructed and inserted into psicheck and HeLa cells were transfected with resultant plasmids. Gene silencing effects of all 16 combinations were examined using a dual luciferase assay. (A) Left, structure of psicheck. Right, parental target and GS sequences used. (, G, L, Q), Control experiments (gene silencing due to completely matched GS). All parental and chimeric sirnas, belonging to class I, were highly functional. (E, H, K, N), Negative control (gene silencing due to GS with no appreciable complementarity to target). Virtually no gene silencing was observed. (C, F, M, P), Seed-dependent gene silencing. sivim-270 is an sirna causing seed-dependent gene silencing effectively (F) while sivim-812 is almost incapable of inducing seed-dependent gene silencing (M). A chimeric sirna possessing the sivim-270 seed (C) still maintained a strong seed-dependent gene silencing activity, whereas little gene silencing activity was found subsequent to transfection of a chimeric sirna possessing the sivim-812 seed (P). (D, I, J, O), Any appreciable gene silencing activity appeared absent from cells transfected with sirnas with only non-seed region complementarity to the target, indicating that the non-seed region cannot function as a substitute for the seed.
3 A psicheck Relative luc activity (%) Relative luc activity (%) sivim-270 (nm) sivim-812/270 (nm) Target transcript WDR82 CEP350 WP4 COP1 TMEM41A HSC20 VAPA1 VIM-27-sm2 Guide strand of sivim CUGUAGUUUCCAAGAGUUCAG-3 : : : 5 --GACUACCAGUAUGGAGUUCAU-3 : :: : 5 -UUCUGUUCUUCAGGAGUUCAC-3 : :: : : 5 --UUAUAUAGAAUCUGAGUUCAU-3 : : : : 5 --UUCACAAGGUCAGGAGUUCAA-3 : ::: 5 --AUGACAAGGUCAGGAGUUCAA-3 ::: 5 --GUGGUUUUAAUAAGAGUUCAA-3 : 5 -AAUGAUGCACCAGGAGUUCAA-3 : 3 GCGGUAGUUGUGGCUCAAGUU 5 Nonseed Seed WDR CUGUAGUUUCCAAGAGUUCAG-3 : : : CEP GACUACCAGUAUGGAGUUCAU-3 : :: WP4 5 -UUCUGUUCUUCAGGAGUUCAC-3 : :: : : COP1 5 --UUAUAUAGAAUCUGAGUUCAU-3 : : TMEM41A 5 --UUCACAAGGUCAGGAGUUCAA-3 : : HSC AUGACAAGGUCAGGAGUUCAA-3 : : VAPA1 5 --GUGGUUUUAAUAAGAGUUCAA-3 :: : VIM-270-sm25 -AAUGAUGCACCAGGAGUUCAA-3 : : Guide strand 3 UGCAUGCAGUCGUCUCAAGUU 5 of sivim-812/270 Nonseed Seed Supplementary Figure S3. Possible involvement of sirna non-seed region and/or its target counterpart in off-target effect. Complementarity of used 8 target sequences to sivim-270 GS (A) and sivim-812/270 GS () are shown in the right margin. Note that the 8 targets possess non-seed sequences unhomologous from one another while sivim-270 GS and sivim-812/270 GS share in common an identical seed sequence. There is no appreciable homology in non-seed between sivim-270 and sivim-812/270. That off-target effects considerably vary depending on sequences of either GS non-seed or its target counterpart may indicate their possible involvement in modulation of seed-dependent gene silencing.
4 A sicltc-2416 (33.2 ºC) with 3ʼUTR complementary to seed with 3ʼUTR lacking complementarity to seed C sicltc-4819 (-10.3 ºC) Fraction per bin with 3ʼUTR complementary to seed with 3ʼUTR lacking complementarity to seed (intensity after sicltc-2416 treatment / intensity after sicltc-4819 treatment) Supplementary Figure S4. Microarray-based off-target-effect profiles of transcripts downregulated by sicltc-2416 and sicltc-4819, which, respectively, generate seed duplexes with high (33.2 ºC) and low ( ºC) Tm values upon association of cognate target mrnas. Changes in expression, shown as the log2 of the ratio of the fold change to the mock transfection, were determined using microarray data for 17,321 human transcripts obtained from HeLa cells transfected with cognate sirnas. The blue line indicates cumulative distribution of transcripts with 3 UTR complementary to the seed of cognate sirna. The gray line shows transcripts with no seed complementarity. Differences in mean value between normalized log2 fold change of mrna with 3 UTR complementary to the seed and mrna lacking seed match are 0.45 ± 0.04 (A) and 0.08 ± (). (C) Comparison of microarray profiles of transcripts downregulated by sicltc-2416 and cicltc All transcripts with 3 UTR complementarity to sirna seed sequences were examined. Ordinate, fraction. Abscissa, signal intensity obtained after sicltc-2416 treatment divided by that after sicltc-4819 treatment. The red line indicates distribution of transcripts complementary to the sicltc-2416 seed sequence. The blue line denotes the distribution of transcripts complementary to the sicltc-4819 seed sequence. The black line shows transcripts with no seed complementarity.
5 Supplementary Figure S5. A E F sivim-270 vimentin with 3ʼUTR complementary to seed vimentin sicltc clathrin C D G H sivim-812 vimentin with 3ʼUTR complementary to seed vimentin sicltc clathrin clathrin with 3ʼUTR complementary to seed with 3ʼUTR complementary to seed clathrin 13024
6 Supplementary Figure S5. Microarray profiles of transcripts downregulated by transfection of sirnas. Four different sirnas were used. (A, ) sivim-270 and (E, F) sicltc-2416 are capable forming seed duplexes with high Tm values (26.2 and 33.2 ºC, respectively) with the 3 UTR counterparts of target mrnas. sivim-812 (C, D) and sicltc-4819 (G, H) are capable forming seed duplexes with low Tm values (8.8 and ºC, respectively) with the 3 UTR counterparts of target mrnas. Microarray-based expression profiles were examined 24 h after transfection. Change in gene expression is shown by log2 of fold change ratio (left ordinate) and mrna level (right ordinate) relative to mock transfection. Abscissa, signal intensity of the transcript (log10). lue and gray dots, respectively, represent transcripts complementary to the seed and those with no seed complementarity. Transcripts with no seed complementarity were plotted separately in, D, F and H. Signals for cm target genes, vimentin (A-D) and clathrin heavy chain (E-H), are colored black and indicated by arrows. Their expression levels were reduced to 18 % (sivim-270; A, ), 17 % (sivim-812; C, D), 17 % (sicltc-2416; E, F) and 17 % (sicltc-4819; G, H). The number of genes examined is shown on the bottom-right edge in each panel. The expression of transcripts with 3 UTR complementary to sivim-270 (A) or sicltc-2416 (E) seeds was significantly reduced subsequent to the transfection of the cognate sirna. In contrast, the reduction of transcripts with 3 UTR complementary to sivim-812 (C) or sicltc-4819 (G) seed sequences was little, if any. These findings may support the notion that seed-dependent gene silencing activity of sirna is determined by the stability of the seed duplex formed between sirna GS and 3 UTR of target mrna.
7 Supplementary Figure S6. A mrna level measured by qrt-pcr ( %) r = 0.92 sivim-270 sivim-812 mrna level of vimentin (%) Microarray qrt-pcr mrna level of VAPA (%) mrna level measured by microarray (%) C D E mrna level of MTPN (%) mrna level of PTPRF (%) F mrna level of MRPS16 (%) G mrna level of COPS8 (%) H M mrna level of COP1 (%) mrna level of WDR82 (%) I J K L N mrna level of RA31 (%) mrna level of SURF4 (%) O R S T U V mrna level of TF3L4 (%) mrna level of ECN1 (%) W X Y Z mrna level of POGK (%) mrna level of HDAC2 (%) mrna level of ARPC2 (%) mrna level of CALM2 (%) mrna level of SSR1 (%) mrna level of PSAP (%) P mrna level of PLEKHC1 (%) mrna level of HINT1 (%) mrna level of TFP1 (%) mrna level of CAPRIN1 (%) Q mrna level of DR1 (%) mrna level of TRU1 (%) mrna level of TMEM30A (%)
8 Supplementary Figure S6. Comparison of microarray data with those of qrt-pcr. (A) Twenty four genes whose transcripts possess complementarity to either sivim-270 and sivim-812 were arbitrarily chosen and change in expression level of their mrna along with that of vimentin RNA in sirna-treated cells, compared with mock transfection, were examined by qrt-pcr and microarray (-Z; see also Supplementary Figure S5). Red dots, data obtained subsequent to sivim-270 treatment. lue dots, data obtained subsequent to sivim-812 treatment. Note that microarray data are almost linear with those of qrt-pcr, indicating that results of microarray analysis and those of qrt-pcr are essentially identical to each other. The correlation coefficient was estimated at Comparison at the level of individual gene is shown in (-Z); () vimentin, (C) VAPA, (D) MTPN, (E) PTPRF, (F) MRPS16, (G) COPS8, (H) COP1, (I) RA31, (J) ARPC2, (K) PLEKHC1, (L) DR1, (M) WDR82, (N) SURF4, (O) CALM2, (P) HINT1, (Q) TRU1, (R) TF3L4, (S) ECN1, (T) SSR1, (U) TFP1, (V) TMEM30A, (W) POGK, (X) HDAC2, (Y) PSAP and (Z) CAPRIN1. Target transcripts in (C-O) contain 3 UTRs possessing complementarity to the sivim-270 seed, while those in (P-Z) contain 3 UTRs possessing complementarity to the sivim-812 seed.
MicroRNA Mike needs help to degrade all the mrna transcripts! Aaron Arvey ISMB 2010
 Target mrna abundance dilutes microrna and sirna activity MicroRNA Mike needs help to degrade all the mrna transcripts! Aaron Arvey ISMB 2010 Target mrna abundance dilutes microrna and sirna activity Erik
Target mrna abundance dilutes microrna and sirna activity MicroRNA Mike needs help to degrade all the mrna transcripts! Aaron Arvey ISMB 2010 Target mrna abundance dilutes microrna and sirna activity Erik
Analyzing microrna Data and Integrating mirna with Gene Expression Data in Partek Genomics Suite 6.6
 Analyzing microrna Data and Integrating mirna with Gene Expression Data in Partek Genomics Suite 6.6 Overview This tutorial outlines how microrna data can be analyzed within Partek Genomics Suite. Additionally,
Analyzing microrna Data and Integrating mirna with Gene Expression Data in Partek Genomics Suite 6.6 Overview This tutorial outlines how microrna data can be analyzed within Partek Genomics Suite. Additionally,
岑 祥 股 份 有 限 公 司 技 術 專 員 費 軫 尹 20100803
 技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
SUPPLEMENTARY DATA 1
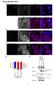 SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
Supplementary Figure 1.
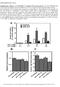 Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Dicer Substrate RNAi Design
 INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
Thermo Scientific Dharmacon sirna Libraries
 Thermo Scientific Dharmacon sirna Libraries Scientific innovation in sirna potency, specificity and delivery Superior sirna in comprehensive collections for reliable screens Market leading products for
Thermo Scientific Dharmacon sirna Libraries Scientific innovation in sirna potency, specificity and delivery Superior sirna in comprehensive collections for reliable screens Market leading products for
INTERFERin in vitro sirna/mirna transfection reagent PROTOCOL. 1 Standard sirna transfection of adherent cells... 2
 INTERFERin in vitro sirna/mirna transfection reagent PROTOCOL DESCRIPTION INTERFERin is a powerful sirna/mirna transfection reagent that ensures efficient gene silencing and reproducible transfection in
INTERFERin in vitro sirna/mirna transfection reagent PROTOCOL DESCRIPTION INTERFERin is a powerful sirna/mirna transfection reagent that ensures efficient gene silencing and reproducible transfection in
A role of microrna in the regulation of telomerase? Yuan Ming Yeh, Pei Rong Huang, and Tzu Chien V. Wang
 A role of microrna in the regulation of telomerase? Yuan Ming Yeh, Pei Rong Huang, and Tzu Chien V. Wang Department of Molecular and Cellular Biology, Chang Gung University, Kwei San, Tao Yuan 333, Taiwan
A role of microrna in the regulation of telomerase? Yuan Ming Yeh, Pei Rong Huang, and Tzu Chien V. Wang Department of Molecular and Cellular Biology, Chang Gung University, Kwei San, Tao Yuan 333, Taiwan
SmartFlare RNA Detection Probes: Principles, protocols and troubleshooting
 Technical Guide SmartFlare RNA Detection Probes: Principles, protocols and troubleshooting Principles of SmartFlare technology RNA detection traditionally requires transfection, laborious sample prep,
Technical Guide SmartFlare RNA Detection Probes: Principles, protocols and troubleshooting Principles of SmartFlare technology RNA detection traditionally requires transfection, laborious sample prep,
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation
 OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
Notch 1 -dependent regulation of cell fate in colorectal cancer
 Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
Determinants of targeting by endogenous and exogenous micrornas and sirnas
 Determinants of targeting by endogenous and exogenous micrornas and sirnas CYDNEY B. NIELSEN, 1,3 NOAM SHOMRON, 1,3 RICKARD SANDBERG, 1 ERAN HORNSTEIN, 2,4 JACOB KITZMAN, 1,5 and CHRISTOPHER B. BURGE 1
Determinants of targeting by endogenous and exogenous micrornas and sirnas CYDNEY B. NIELSEN, 1,3 NOAM SHOMRON, 1,3 RICKARD SANDBERG, 1 ERAN HORNSTEIN, 2,4 JACOB KITZMAN, 1,5 and CHRISTOPHER B. BURGE 1
Thermo Scientific Dharmacon ON-TARGETplus sirna The Standard for sirna Specificity
 Thermo Scientific Dharmacon sirna The Standard for sirna Specificity Reduce off-targets up to 90% with the market-leading sirna Combine higher target specificity with guaranteed silencing Increase confidence
Thermo Scientific Dharmacon sirna The Standard for sirna Specificity Reduce off-targets up to 90% with the market-leading sirna Combine higher target specificity with guaranteed silencing Increase confidence
RNA & Protein Synthesis
 RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
Materials and Methods. Blocking of Globin Reverse Transcription to Enhance Human Whole Blood Gene Expression Profiling
 Application Note Blocking of Globin Reverse Transcription to Enhance Human Whole Blood Gene Expression Profi ling Yasmin Beazer-Barclay, Doug Sinon, Christopher Morehouse, Mark Porter, and Mike Kuziora
Application Note Blocking of Globin Reverse Transcription to Enhance Human Whole Blood Gene Expression Profi ling Yasmin Beazer-Barclay, Doug Sinon, Christopher Morehouse, Mark Porter, and Mike Kuziora
Gene Expression Assays
 APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
Relative Quantification of mirna Target mrnas by Real-Time qpcr. 1 Introduction. Gene Expression Application Note No. 4
 Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
PRODUCT INFORMATION...
 VIROMER RED In vitro plasmid DNA and mrna Standard Transfection PRODUCT INFORMATION... 2 GENERAL... 2 RED OR YELLOW?... 3 PROTOCOL GUIDELINES... 4 GENERAL REMARKS... 4 CELL CULTURE AND PLATING... 4 FORWARD/REVERSE
VIROMER RED In vitro plasmid DNA and mrna Standard Transfection PRODUCT INFORMATION... 2 GENERAL... 2 RED OR YELLOW?... 3 PROTOCOL GUIDELINES... 4 GENERAL REMARKS... 4 CELL CULTURE AND PLATING... 4 FORWARD/REVERSE
Lezioni Dipartimento di Oncologia Farmacologia Molecolare. RNA interference. Giovanna Damia 29 maggio 2006
 Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
0 value2. 3. Assign labels to the proteins in each interval of the ranked list
 Ranked list of protein degrees in decreasing order 0 100 Proteins with few connections Proteins with many connections 1. Sort a random number between 80 and 98 (value1) 0 value1 2. Sort a random number
Ranked list of protein degrees in decreasing order 0 100 Proteins with few connections Proteins with many connections 1. Sort a random number between 80 and 98 (value1) 0 value1 2. Sort a random number
Translation Study Guide
 Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Supplementary Figure 1: Quality Assessment of Mouse Arrays. Supplementary Figure 2: Quality Assessment of Rat Arrays
 Supplementary Figure 1: Quality Assessment of Mouse Arrays The mouse microarray data were subjected to an extensive quality-control procedure prior to conducting downstream analyses. We assessed the spread
Supplementary Figure 1: Quality Assessment of Mouse Arrays The mouse microarray data were subjected to an extensive quality-control procedure prior to conducting downstream analyses. We assessed the spread
Essentials of Real Time PCR. About Sequence Detection Chemistries
 Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Functional Analysis. mircury LNA microrna Mimics
 Functional Analysis mircury LNA microrna Mimics Instruction manual v1.0 April 2014 Table of Contents Product summary......................................................... 3 Content..................................................................
Functional Analysis mircury LNA microrna Mimics Instruction manual v1.0 April 2014 Table of Contents Product summary......................................................... 3 Content..................................................................
Name Class Date. Figure 13 1. 2. Which nucleotide in Figure 13 1 indicates the nucleic acid above is RNA? a. uracil c. cytosine b. guanine d.
 13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
Functional and Biomedical Aspects of Genome Research
 Functional and Biomedical Aspects of Genome Research 20 11 35 Vorlesung SS 04 Bartsch, Jockusch & Schmitt-John Mi. 9:15-10:00, in W7-135 13 Functional RNAs Thomas Schmitt-John micro RNAs small interfering
Functional and Biomedical Aspects of Genome Research 20 11 35 Vorlesung SS 04 Bartsch, Jockusch & Schmitt-John Mi. 9:15-10:00, in W7-135 13 Functional RNAs Thomas Schmitt-John micro RNAs small interfering
Name: Date: Period: DNA Unit: DNA Webquest
 Name: Date: Period: DNA Unit: DNA Webquest Part 1 History, DNA Structure, DNA Replication DNA History http://www.dnaftb.org/dnaftb/1/concept/index.html Read the text and answer the following questions.
Name: Date: Period: DNA Unit: DNA Webquest Part 1 History, DNA Structure, DNA Replication DNA History http://www.dnaftb.org/dnaftb/1/concept/index.html Read the text and answer the following questions.
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals
 Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Hierarchical Clustering Analysis
 Hierarchical Clustering Analysis What is Hierarchical Clustering? Hierarchical clustering is used to group similar objects into clusters. In the beginning, each row and/or column is considered a cluster.
Hierarchical Clustering Analysis What is Hierarchical Clustering? Hierarchical clustering is used to group similar objects into clusters. In the beginning, each row and/or column is considered a cluster.
HiPerFect Transfection Reagent Handbook
 Fifth Edition October 2010 HiPerFect Transfection Reagent Handbook For transfection of eukaryotic cells with sirna and mirna Sample & Assay Technologies QIAGEN Sample and Assay Technologies QIAGEN is the
Fifth Edition October 2010 HiPerFect Transfection Reagent Handbook For transfection of eukaryotic cells with sirna and mirna Sample & Assay Technologies QIAGEN Sample and Assay Technologies QIAGEN is the
Outline. interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work?
 Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Comprehensive mirna Research Technologies
 Comprehensive mirna Research Technologies Sample & Assay Technologies QIAGEN solutions for advancing microrna research In the last few years, the identification of microrna (mirna) and the recognition
Comprehensive mirna Research Technologies Sample & Assay Technologies QIAGEN solutions for advancing microrna research In the last few years, the identification of microrna (mirna) and the recognition
LightSwitch Luciferase Assay System
 Constructs, Assays & Services Assay System Promoter GoClones Synthetic Response Elements Pathway Cell Lines 3 UTR GoClones mirna Mimics & Inhibitors Synthetic mirna Targets Transfection Reagents Assay
Constructs, Assays & Services Assay System Promoter GoClones Synthetic Response Elements Pathway Cell Lines 3 UTR GoClones mirna Mimics & Inhibitors Synthetic mirna Targets Transfection Reagents Assay
Modeling DNA Replication and Protein Synthesis
 Skills Practice Lab Modeling DNA Replication and Protein Synthesis OBJECTIVES Construct and analyze a model of DNA. Use a model to simulate the process of replication. Use a model to simulate the process
Skills Practice Lab Modeling DNA Replication and Protein Synthesis OBJECTIVES Construct and analyze a model of DNA. Use a model to simulate the process of replication. Use a model to simulate the process
Molecular Genetics. RNA, Transcription, & Protein Synthesis
 Molecular Genetics RNA, Transcription, & Protein Synthesis Section 1 RNA AND TRANSCRIPTION Objectives Describe the primary functions of RNA Identify how RNA differs from DNA Describe the structure and
Molecular Genetics RNA, Transcription, & Protein Synthesis Section 1 RNA AND TRANSCRIPTION Objectives Describe the primary functions of RNA Identify how RNA differs from DNA Describe the structure and
Understanding the immune response to bacterial infections
 Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
TOOLS sirna and mirna. User guide
 TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
PreciseTM Whitepaper
 Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
Measuring gene expression (Microarrays) Ulf Leser
 Measuring gene expression (Microarrays) Ulf Leser This Lecture Gene expression Microarrays Idea Technologies Problems Quality control Normalization Analysis next week! 2 http://learn.genetics.utah.edu/content/molecules/transcribe/
Measuring gene expression (Microarrays) Ulf Leser This Lecture Gene expression Microarrays Idea Technologies Problems Quality control Normalization Analysis next week! 2 http://learn.genetics.utah.edu/content/molecules/transcribe/
Suppl. Figure 1. Cells cultured in oxldl take up Oil Red O in a time-dependent
 Suppl. Figure 1. Cells cultured in oxldl take up Oil Red O in a time-dependent manner. Uptake of Oil Red O was performed as detailed in Materials and Methods and absorbance of samples was expressed as
Suppl. Figure 1. Cells cultured in oxldl take up Oil Red O in a time-dependent manner. Uptake of Oil Red O was performed as detailed in Materials and Methods and absorbance of samples was expressed as
User Manual/Hand book. qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse)
 User Manual/Hand book qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse) Kit Components Cat. No. MA003...Human Whole Genome mirna qpcr Profiling Kit (-20 C) The following components are sufficient
User Manual/Hand book qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse) Kit Components Cat. No. MA003...Human Whole Genome mirna qpcr Profiling Kit (-20 C) The following components are sufficient
Thermo Scientific DharmaFECT Transfection Reagents
 Thermo Scientific Transfection Reagents Maintain cell viability with lipids specially formulated to deliver sirna Achieve robust sirna uptake for dependable gene silencing Expand your RNAi applications
Thermo Scientific Transfection Reagents Maintain cell viability with lipids specially formulated to deliver sirna Achieve robust sirna uptake for dependable gene silencing Expand your RNAi applications
Considerable evidence now indicates that small noncoding
 MicroRNAs and small interfering RNAs can inhibit mrna expression by similar mechanisms Yan Zeng*, Rui Yi, and Bryan R. Cullen* *Howard Hughes Medical Institute and Department of Molecular Genetics and
MicroRNAs and small interfering RNAs can inhibit mrna expression by similar mechanisms Yan Zeng*, Rui Yi, and Bryan R. Cullen* *Howard Hughes Medical Institute and Department of Molecular Genetics and
EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES
 C GUGAAG EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES MicroRNA-adapted shrna (shrna mir ) for increased, specific and consistent knockdown. MicroRNA PROCESSING PATHWAY UTILIZED FOR shrna mir Developed
C GUGAAG EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES MicroRNA-adapted shrna (shrna mir ) for increased, specific and consistent knockdown. MicroRNA PROCESSING PATHWAY UTILIZED FOR shrna mir Developed
Innate Immunity. Insects rely solely on an innate immune system for defense against infection
 Innate Immunity Insects rely solely on an innate immune system for defense against infection The importance of mammalian innate immunity has only recently become appreciated The innate immune response
Innate Immunity Insects rely solely on an innate immune system for defense against infection The importance of mammalian innate immunity has only recently become appreciated The innate immune response
2.1.2 Characterization of antiviral effect of cytokine expression on HBV replication in transduced mouse hepatocytes line
 i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
Instructions. Torpedo sirna. Material. Important Guidelines. Specifications. Quality Control
 is a is a state of the art transfection reagent, specifically designed for the transfer of sirna and mirna into a variety of eukaryotic cell types. is a state of the art transfection reagent, specifically
is a is a state of the art transfection reagent, specifically designed for the transfer of sirna and mirna into a variety of eukaryotic cell types. is a state of the art transfection reagent, specifically
Ms. Campbell Protein Synthesis Practice Questions Regents L.E.
 Name Student # Ms. Campbell Protein Synthesis Practice Questions Regents L.E. 1. A sequence of three nitrogenous bases in a messenger-rna molecule is known as a 1) codon 2) gene 3) polypeptide 4) nucleotide
Name Student # Ms. Campbell Protein Synthesis Practice Questions Regents L.E. 1. A sequence of three nitrogenous bases in a messenger-rna molecule is known as a 1) codon 2) gene 3) polypeptide 4) nucleotide
From DNA to Protein. Proteins. Chapter 13. Prokaryotes and Eukaryotes. The Path From Genes to Proteins. All proteins consist of polypeptide chains
 Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
Manual for: sirna-trans Maximo Reagent
 Manual for: sirna-trans Maximo Reagent Contact: info@geneon.net; www.geneon.net GeneON GmbH Germany GeneON GmbH / Protocol sirna-trans reagent vs. 1.3 / www.geneon.net / - 0 - sirna-trans reagent Instruction
Manual for: sirna-trans Maximo Reagent Contact: info@geneon.net; www.geneon.net GeneON GmbH Germany GeneON GmbH / Protocol sirna-trans reagent vs. 1.3 / www.geneon.net / - 0 - sirna-trans reagent Instruction
Dicer-Substrate sirna Technology
 As featured in BioRadiations 120 Dicer-Substrate sirna Technology Advances in sirna Designs Improve Gene-Specific Silencing Steve Kulisch, Teresa Esch, Christina Whitman-Guliaev, and Teresa Rubio, Bio-Rad
As featured in BioRadiations 120 Dicer-Substrate sirna Technology Advances in sirna Designs Improve Gene-Specific Silencing Steve Kulisch, Teresa Esch, Christina Whitman-Guliaev, and Teresa Rubio, Bio-Rad
Real-time PCR: Understanding C t
 APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
Molecular Genetics: Challenges for Statistical Practice. J.K. Lindsey
 Molecular Genetics: Challenges for Statistical Practice J.K. Lindsey 1. What is a Microarray? 2. Design Questions 3. Modelling Questions 4. Longitudinal Data 5. Conclusions 1. What is a microarray? A microarray
Molecular Genetics: Challenges for Statistical Practice J.K. Lindsey 1. What is a Microarray? 2. Design Questions 3. Modelling Questions 4. Longitudinal Data 5. Conclusions 1. What is a microarray? A microarray
MicroRNA formation. 4th International Symposium on Non-Surgical Contraceptive Methods of Pet Population Control
 MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
The Steps. 1. Transcription. 2. Transferal. 3. Translation
 Protein Synthesis Protein synthesis is simply the "making of proteins." Although the term itself is easy to understand, the multiple steps that a cell in a plant or animal must go through are not. In order
Protein Synthesis Protein synthesis is simply the "making of proteins." Although the term itself is easy to understand, the multiple steps that a cell in a plant or animal must go through are not. In order
RNA-seq. Quantification and Differential Expression. Genomics: Lecture #12
 (2) Quantification and Differential Expression Institut für Medizinische Genetik und Humangenetik Charité Universitätsmedizin Berlin Genomics: Lecture #12 Today (2) Gene Expression per Sources of bias,
(2) Quantification and Differential Expression Institut für Medizinische Genetik und Humangenetik Charité Universitätsmedizin Berlin Genomics: Lecture #12 Today (2) Gene Expression per Sources of bias,
!!!!!!!!!!!!!!!!!!!!!!!!!!
 Figure S7 cdl-1 3'UTR gld-1 binding site (4..10) let-7 binding site (326..346) Figure Legends Figure S1. gld-1 genetically interacts with nhl-2 and vig-1 during germline development. (A) nhl-2(ok818) enhances
Figure S7 cdl-1 3'UTR gld-1 binding site (4..10) let-7 binding site (326..346) Figure Legends Figure S1. gld-1 genetically interacts with nhl-2 and vig-1 during germline development. (A) nhl-2(ok818) enhances
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
CODICE DESCRIZIONE QUANTITA' PREZZO EURO
 Broad Spectrum Plasmid DNA and sirna/mirna Transfection MIR 6000 TransIT-X2 Dynamic Delivery System 1.5 ml 823,35 MIR 6003 TransIT-X2 Dynamic Delivery System 0.3 ml 196,35 MIR 6004 TransIT-X2 Dynamic Delivery
Broad Spectrum Plasmid DNA and sirna/mirna Transfection MIR 6000 TransIT-X2 Dynamic Delivery System 1.5 ml 823,35 MIR 6003 TransIT-X2 Dynamic Delivery System 0.3 ml 196,35 MIR 6004 TransIT-X2 Dynamic Delivery
Global microrna level regulation of EGFR-driven cell cycle protein network in breast cancer
 Molecular Systems Biology Peer Review Process File Global microrna level regulation of EGFR-driven cell cycle protein network in breast cancer Stefan Uhlmann, Heiko Mannsperger, Jitao David Zhang, Emőke-Ágnes
Molecular Systems Biology Peer Review Process File Global microrna level regulation of EGFR-driven cell cycle protein network in breast cancer Stefan Uhlmann, Heiko Mannsperger, Jitao David Zhang, Emőke-Ágnes
Superior TrueMAB TM monoclonal antibodies for the recognition of proteins native epitopes
 Superior TrueMAB TM monoclonal antibodies for the recognition of proteins native epitopes Outlines Brief introduction of OriGene s mission on gene-centric product solution. TrueMAB monoclonal antibody
Superior TrueMAB TM monoclonal antibodies for the recognition of proteins native epitopes Outlines Brief introduction of OriGene s mission on gene-centric product solution. TrueMAB monoclonal antibody
Interaktionen von RNAs und Proteinen
 Sonja Prohaska Computational EvoDevo Universitaet Leipzig June 9, 2015 Studying RNA-protein interactions Given: target protein known to bind to RNA problem: find binding partners and binding sites experimental
Sonja Prohaska Computational EvoDevo Universitaet Leipzig June 9, 2015 Studying RNA-protein interactions Given: target protein known to bind to RNA problem: find binding partners and binding sites experimental
Scottish Qualifications Authority
 National Unit specification: general information Unit code: FH2G 12 Superclass: RH Publication date: March 2011 Source: Scottish Qualifications Authority Version: 01 Summary This Unit is a mandatory Unit
National Unit specification: general information Unit code: FH2G 12 Superclass: RH Publication date: March 2011 Source: Scottish Qualifications Authority Version: 01 Summary This Unit is a mandatory Unit
Supplementary Figure 1 Characterization of 5xFAD derived hippocampal cultures. A-B) Average total numbers of cells per hippocampus as well as average
 Supplementary Figure 1 Characterization of 5xFAD derived hippocampal cultures. A-B) Average total numbers of cells per hippocampus as well as average total numbers of cells per field of view were similar
Supplementary Figure 1 Characterization of 5xFAD derived hippocampal cultures. A-B) Average total numbers of cells per hippocampus as well as average total numbers of cells per field of view were similar
Biotechnology: DNA Technology & Genomics
 Chapter 20. Biotechnology: DNA Technology & Genomics 2003-2004 The BIG Questions How can we use our knowledge of DNA to: diagnose disease or defect? cure disease or defect? change/improve organisms? What
Chapter 20. Biotechnology: DNA Technology & Genomics 2003-2004 The BIG Questions How can we use our knowledge of DNA to: diagnose disease or defect? cure disease or defect? change/improve organisms? What
Basic Analysis of Microarray Data
 Basic Analysis of Microarray Data A User Guide and Tutorial Scott A. Ness, Ph.D. Co-Director, Keck-UNM Genomics Resource and Dept. of Molecular Genetics and Microbiology University of New Mexico HSC Tel.
Basic Analysis of Microarray Data A User Guide and Tutorial Scott A. Ness, Ph.D. Co-Director, Keck-UNM Genomics Resource and Dept. of Molecular Genetics and Microbiology University of New Mexico HSC Tel.
Introduction To Real Time Quantitative PCR (qpcr)
 Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
ALLEN Mouse Brain Atlas
 TECHNICAL WHITE PAPER: QUALITY CONTROL STANDARDS FOR HIGH-THROUGHPUT RNA IN SITU HYBRIDIZATION DATA GENERATION Consistent data quality and internal reproducibility are critical concerns for high-throughput
TECHNICAL WHITE PAPER: QUALITY CONTROL STANDARDS FOR HIGH-THROUGHPUT RNA IN SITU HYBRIDIZATION DATA GENERATION Consistent data quality and internal reproducibility are critical concerns for high-throughput
Basic Concepts of DNA, Proteins, Genes and Genomes
 Basic Concepts of DNA, Proteins, Genes and Genomes Kun-Mao Chao 1,2,3 1 Graduate Institute of Biomedical Electronics and Bioinformatics 2 Department of Computer Science and Information Engineering 3 Graduate
Basic Concepts of DNA, Proteins, Genes and Genomes Kun-Mao Chao 1,2,3 1 Graduate Institute of Biomedical Electronics and Bioinformatics 2 Department of Computer Science and Information Engineering 3 Graduate
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID papillary renal cell carcinoma (translocation-associated) PRCC Human This gene
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID papillary renal cell carcinoma (translocation-associated) PRCC Human This gene
APPLYING BENFORD'S LAW This PDF contains step-by-step instructions on how to apply Benford's law using Microsoft Excel, which is commonly used by
 APPLYING BENFORD'S LAW This PDF contains step-by-step instructions on how to apply Benford's law using Microsoft Excel, which is commonly used by internal auditors around the world in their day-to-day
APPLYING BENFORD'S LAW This PDF contains step-by-step instructions on how to apply Benford's law using Microsoft Excel, which is commonly used by internal auditors around the world in their day-to-day
Technical Note: Analysis of microrna function using mircury LNA TM microrna Inhibitors
 Technical note: Analysis of microrna function using mircury LNA microrna Inhibitors Introduction The study of micrornas has come a long way since the discovery of these small RNAs some fifteen years ago.
Technical note: Analysis of microrna function using mircury LNA microrna Inhibitors Introduction The study of micrornas has come a long way since the discovery of these small RNAs some fifteen years ago.
Step-by-Step Guide to Bi-Parental Linkage Mapping WHITE PAPER
 Step-by-Step Guide to Bi-Parental Linkage Mapping WHITE PAPER JMP Genomics Step-by-Step Guide to Bi-Parental Linkage Mapping Introduction JMP Genomics offers several tools for the creation of linkage maps
Step-by-Step Guide to Bi-Parental Linkage Mapping WHITE PAPER JMP Genomics Step-by-Step Guide to Bi-Parental Linkage Mapping Introduction JMP Genomics offers several tools for the creation of linkage maps
2. True or False? The sequence of nucleotides in the human genome is 90.9% identical from one person to the next. False (it s 99.
 1. True or False? A typical chromosome can contain several hundred to several thousand genes, arranged in linear order along the DNA molecule present in the chromosome. True 2. True or False? The sequence
1. True or False? A typical chromosome can contain several hundred to several thousand genes, arranged in linear order along the DNA molecule present in the chromosome. True 2. True or False? The sequence
CONTRACTING ORGANIZATION: University of Alabama at Birmingham Birmingham, AL 35294
 AD Award Number: W81XWH-08-1-0030 TITLE: Regulation of Prostate Cancer Bone Metastasis by DKK1 PRINCIPAL INVESTIGATOR: Gregory A. Clines, M.D., Ph.D. CONTRACTING ORGANIZATION: University of Alabama at
AD Award Number: W81XWH-08-1-0030 TITLE: Regulation of Prostate Cancer Bone Metastasis by DKK1 PRINCIPAL INVESTIGATOR: Gregory A. Clines, M.D., Ph.D. CONTRACTING ORGANIZATION: University of Alabama at
Microarray Data Analysis Workshop. Custom arrays and Probe design Probe design in a pangenomic world. Carsten Friis. MedVetNet Workshop, DTU 2008
 Microarray Data Analysis Workshop MedVetNet Workshop, DTU 2008 Custom arrays and Probe design Probe design in a pangenomic world Carsten Friis Media glna tnra GlnA TnrA C2 glnr C3 C5 C6 K GlnR C1 C4 C7
Microarray Data Analysis Workshop MedVetNet Workshop, DTU 2008 Custom arrays and Probe design Probe design in a pangenomic world Carsten Friis Media glna tnra GlnA TnrA C2 glnr C3 C5 C6 K GlnR C1 C4 C7
Name Date Period. 2. When a molecule of double-stranded DNA undergoes replication, it results in
 DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
Machine Learning Approaches to sirna Efficacy Prediction
 Machine Learning Approaches to sirna Efficacy Prediction by Sahar Abubucker B.E., Madras University, 2000 THESIS Submitted in Partial Fulfillment of the Requirements for the Degree of Master of Science
Machine Learning Approaches to sirna Efficacy Prediction by Sahar Abubucker B.E., Madras University, 2000 THESIS Submitted in Partial Fulfillment of the Requirements for the Degree of Master of Science
Fig. S1. Identification of CRMs with activity in Ci-Tbx2/3 expression domains outside the notochord. B-F
 Fig. S1. Identification of CRMs with activity in Ci-Tbx2/3 expression domains outside the notochord. (A) Top: map of the Ci- Tbx2/3 locus. Teal rectangles and lines denote exons and introns, respectively.
Fig. S1. Identification of CRMs with activity in Ci-Tbx2/3 expression domains outside the notochord. (A) Top: map of the Ci- Tbx2/3 locus. Teal rectangles and lines denote exons and introns, respectively.
Transfection-Transfer of non-viral genetic material into eukaryotic cells. Infection/ Transduction- Transfer of viral genetic material into cells.
 Transfection Key words: Transient transfection, Stable transfection, transfection methods, vector, plasmid, origin of replication, reporter gene/ protein, cloning site, promoter and enhancer, signal peptide,
Transfection Key words: Transient transfection, Stable transfection, transfection methods, vector, plasmid, origin of replication, reporter gene/ protein, cloning site, promoter and enhancer, signal peptide,
RNA Interference. - Localization - Labeling - Expression Profiling - In Vivo Delivery - Transfection
 RNA Interference - Localization - Labeling - Expression Profiling - In Vivo Delivery - Transfection Rna interference x RNAi Pathways RNA interference x Tools and Multiplexing RNAi Tools pri-mirna Application
RNA Interference - Localization - Labeling - Expression Profiling - In Vivo Delivery - Transfection Rna interference x RNAi Pathways RNA interference x Tools and Multiplexing RNAi Tools pri-mirna Application
Petrel TIPS&TRICKS from SCM
 Petrel TIPS&TRICKS from SCM Maps: Knowledge Worth Sharing Map Annotation A map is a graphic representation of some part of the earth. In our industry, it may represent either the surface or sub surface;
Petrel TIPS&TRICKS from SCM Maps: Knowledge Worth Sharing Map Annotation A map is a graphic representation of some part of the earth. In our industry, it may represent either the surface or sub surface;
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS)
 Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
13.2 Ribosomes & Protein Synthesis
 13.2 Ribosomes & Protein Synthesis Introduction: *A specific sequence of bases in DNA carries the directions for forming a polypeptide, a chain of amino acids (there are 20 different types of amino acid).
13.2 Ribosomes & Protein Synthesis Introduction: *A specific sequence of bases in DNA carries the directions for forming a polypeptide, a chain of amino acids (there are 20 different types of amino acid).
Human-Mouse Synteny in Functional Genomics Experiment
 Human-Mouse Synteny in Functional Genomics Experiment Ksenia Krasheninnikova University of the Russian Academy of Sciences, JetBrains krasheninnikova@gmail.com September 18, 2012 Ksenia Krasheninnikova
Human-Mouse Synteny in Functional Genomics Experiment Ksenia Krasheninnikova University of the Russian Academy of Sciences, JetBrains krasheninnikova@gmail.com September 18, 2012 Ksenia Krasheninnikova
DNA Worksheet BIOL 1107L DNA
 Worksheet BIOL 1107L Name Day/Time Refer to Chapter 5 and Chapter 16 (Figs. 16.5, 16.7, 16.8 and figure embedded in text on p. 310) in your textbook, Biology, 9th Ed, for information on and its structure
Worksheet BIOL 1107L Name Day/Time Refer to Chapter 5 and Chapter 16 (Figs. 16.5, 16.7, 16.8 and figure embedded in text on p. 310) in your textbook, Biology, 9th Ed, for information on and its structure
a. Ribosomal RNA rrna a type ofrna that combines with proteins to form Ribosomes on which polypeptide chains of proteins are assembled
 Biology 101 Chapter 14 Name: Fill-in-the-Blanks Which base follows the next in a strand of DNA is referred to. as the base (1) Sequence. The region of DNA that calls for the assembly of specific amino
Biology 101 Chapter 14 Name: Fill-in-the-Blanks Which base follows the next in a strand of DNA is referred to. as the base (1) Sequence. The region of DNA that calls for the assembly of specific amino
Introduction to the TI-Nspire CX
 Introduction to the TI-Nspire CX Activity Overview: In this activity, you will become familiar with the layout of the TI-Nspire CX. Step 1: Locate the Touchpad. The Touchpad is used to navigate the cursor
Introduction to the TI-Nspire CX Activity Overview: In this activity, you will become familiar with the layout of the TI-Nspire CX. Step 1: Locate the Touchpad. The Touchpad is used to navigate the cursor
How To Understand How Gene Expression Is Regulated
 What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
Transcription and Translation of DNA
 Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Co Extra (GM and non GM supply chains: Their CO EXistence and TRAceability) Outcomes of Co Extra
 GM and non GM supply chains: Their CO EXistence and TRAceability Outcomes of Co Extra Comparison of different real time PCR chemistries and their suitability for detection and quantification of genetically
GM and non GM supply chains: Their CO EXistence and TRAceability Outcomes of Co Extra Comparison of different real time PCR chemistries and their suitability for detection and quantification of genetically
V22: involvement of micrornas in GRNs
 What are micrornas? V22: involvement of micrornas in GRNs How can one identify micrornas? What is the function of micrornas? Elisa Izaurralde, MPI Tübingen Huntzinger, Izaurralde, Nat. Rev. Genet. 12,
What are micrornas? V22: involvement of micrornas in GRNs How can one identify micrornas? What is the function of micrornas? Elisa Izaurralde, MPI Tübingen Huntzinger, Izaurralde, Nat. Rev. Genet. 12,
Bioinformatics Resources at a Glance
 Bioinformatics Resources at a Glance A Note about FASTA Format There are MANY free bioinformatics tools available online. Bioinformaticists have developed a standard format for nucleotide and protein sequences
Bioinformatics Resources at a Glance A Note about FASTA Format There are MANY free bioinformatics tools available online. Bioinformaticists have developed a standard format for nucleotide and protein sequences
Gene Switches Teacher Information
 STO-143 Gene Switches Teacher Information Summary Kit contains How do bacteria turn on and turn off genes? Students model the action of the lac operon that regulates the expression of genes essential for
STO-143 Gene Switches Teacher Information Summary Kit contains How do bacteria turn on and turn off genes? Students model the action of the lac operon that regulates the expression of genes essential for
Mechanosensitive Neurons on the Internal Reproductive Tract Contribute to Egg-Laying- Induced Acetic Acid Attraction in Drosophila
 Cell Reports, Volume 9 Supplemental Information Mechanosensitive Neurons on the Internal Reproductive Tract Contribute to Egg-Laying- Induced Acetic Acid Attraction in Drosophila Bin Gou, Ying Liu, Ananya
Cell Reports, Volume 9 Supplemental Information Mechanosensitive Neurons on the Internal Reproductive Tract Contribute to Egg-Laying- Induced Acetic Acid Attraction in Drosophila Bin Gou, Ying Liu, Ananya
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
Long-Term Effects of Drug Addiction
 Long-Term Effects of Drug Addiction Part 1: Addiction is a chronic disease Drug addiction is considered a chronic brain disease because drugs cause long-lasting changes in brain structure and function.
Long-Term Effects of Drug Addiction Part 1: Addiction is a chronic disease Drug addiction is considered a chronic brain disease because drugs cause long-lasting changes in brain structure and function.
Computational localization of promoters and transcription start sites in mammalian genomes
 Computational localization of promoters and transcription start sites in mammalian genomes Thomas Down This dissertation is submitted for the degree of Doctor of Philosophy Wellcome Trust Sanger Institute
Computational localization of promoters and transcription start sites in mammalian genomes Thomas Down This dissertation is submitted for the degree of Doctor of Philosophy Wellcome Trust Sanger Institute
