0 value2. 3. Assign labels to the proteins in each interval of the ranked list
|
|
|
- Julian Leonard
- 8 years ago
- Views:
Transcription
1 Ranked list of protein degrees in decreasing order Proteins with few connections Proteins with many connections 1. Sort a random number between 80 and 98 (value1) 0 value1 2. Sort a random number between 60 and value1 (called value2) 0 value2 value1 3. Assign labels to the proteins in each interval of the ranked list 0 value2 value1 Low-Degree Proteins Middle-Degree Proteins High-Degree Proteins Supplementary Figure 1: Procedure to assign nodes to different connectivity categories We defined three categories to classify the proteins in a network according to their connectivity: Low-, middle- and High-degree. Instead of using an absolute number of interactions that each protein should have to belong to a certain category, we established that the proteins should be between a certain percentage of data in order to fit in a certain category. We create a ranked list of proteins according to their number of connections. This list has values in the interval [0, 100]. High values represent the most connected proteins of the network. The first step is to randomly select a value in the interval [80,98]. We call it value1. Now, the proteins present in the list in the interval [value1] are considered high-degree proteins. They are the top 2-20% most connected proteins of the network. The second step is to randomly select another value in the interval [60, value1]. We call it value2., the proteins in the interval [value2, value1] are the middle-degree proteins. They occupy the middle 20-38% of the ranked list. Finally, after these two values are selected and the respective high- and middle-degree categories created, we define that the low-degree proteins occupy the bottom 60% of the list. When performing pair-wise comparisons, we used the same values of value1and value2for both databases. In addition, this procedure was repeated 100 times and the number of agreements (proteins that changed or did not change between categories) was recorded. The result can be seen in Figure 1, where we show the average and standard deviation of the 100 repetitions of this procedure.
2 A B Supplementary Figure 2: Average number of interactions and betweenness distribution of all evaluated databases (A) Shown is the distribution of the average number of interaction partners in the databases analyzed. (B) Shown is the betweenness (i.e., centrality ) of proteins in a network. The degree and betweenness distribution is similar among all databases.
3 90,000 80,000 Number of Interactions 70,000 60,000 50,000 40,000 30,000 20,000 10,000 0 MINT INTACT BIOGRID HPRD MINT INTACT BIOGRID INTACT BIOGRID BIOGRID HPRD HPRD HPRD HIPPIE HIPPIE HIPPIE HIPPIE MINT MINT INTACT Shared Interactions Number of Interactions 200, , , , , ,000 80,000 60,000 40,000 20,000 0 HPRD HIPPIE BIOGRID MINT INTACT Shared Interactions Supplementary Figure 3: Absolute numbers of exclusive and shared interaction partners in pairwise comparisons of PPI datasets We first identified proteins shared between two databases. For these proteins, we identified their interaction partners in each of the databases, and then compared the interaction partners. For all pair-wise comparisons, yellow and blue represent the indicated databases; Shared (red) denotes the number of interaction partners shared between databases.
4 Organ/Cell-type Specific genes [%] Supplementary Figure 4: Percentage of organ/cell type-specific genes represented in the six PPI databases For genes expressed in 84 different human organs/cell types, we determined the level of coverage for the proteins in each of the databases analyzed. All databases have a relatively even coverage of all organs and cell types, although the number of genes expressed varies significantly between the different organs/cell types (Supplementary Figure 5).
5 Supplementary Figure 5: Numbers of genes expressed in 84 different human organs or cell types Microarray data was obtained from different human tissues in duplicates and averaged. This data was pre-processed with GCRMA - GeneChip Robust Multiarray Averaging. Shown are the numbers of genes with moderate to high transcription levels (See Methods).
6 Remaining Interactions [%] Supplementary Figure 6: Organ/cell type-specific subnetworks By combining PPI datasets with data of organ/cell type-specific gene expression, we generated subnetworks that are limited to interactions with both partners expressed in the same organ or cell type; also included are PPIs among house-keeping proteins expressed in all cell types. As expected, this analysis reduced the number of interactions significantly, as compared to the parent PPI database. On average, the organ/cell type-specific subnetworks encompass 1-25% of the original interactions. Although the number of interactions is reduced significantly compared to the parent database, the subnetworks still have several thousand interactions. Depicted is the percentage of remaining edges from the original database filtered by organ/cell-type specific genes.
7 Number of Components Supplementary Figure 7: Number of connected components according to organ/cell-type subnetwork After filtering the parent PPI network to contain only interactions of proteins expressed in the same organ/cell-type, the subnetwork is fragmented into several connected components. We considered a connected component a connection of at least two proteins (i.e. the singletons in the network are ignored). We observe that database is the least fragmented among all databases compared and that certain tissues subnetworks (liver, heart, lung) have more connected components than others (ovary, olfactory lobe, skin).
8 Supplementary Figure 8: Relation between the number of organ/cell type-specific genes and remaining interactions In tissues with a high number of expressed genes, more interactions from the original database remained. Consequently, we observed a strong correlation between the number of organ/cell type-specific genes and the remaining interactions per subnetwork. In all cases, the Pearson's correlation coefficient between the number of genes and the number of interactions in the subnetworks was greater than 0.96 (p-value < 2-2e-16), indicating that the number of interactions seems to increase proportionally to the number of nodes.
Network Analysis. BCH 5101: Analysis of -Omics Data 1/34
 Network Analysis BCH 5101: Analysis of -Omics Data 1/34 Network Analysis Graphs as a representation of networks Examples of genome-scale graphs Statistical properties of genome-scale graphs The search
Network Analysis BCH 5101: Analysis of -Omics Data 1/34 Network Analysis Graphs as a representation of networks Examples of genome-scale graphs Statistical properties of genome-scale graphs The search
Long-Term Effects of Drug Addiction
 Long-Term Effects of Drug Addiction Part 1: Addiction is a chronic disease Drug addiction is considered a chronic brain disease because drugs cause long-lasting changes in brain structure and function.
Long-Term Effects of Drug Addiction Part 1: Addiction is a chronic disease Drug addiction is considered a chronic brain disease because drugs cause long-lasting changes in brain structure and function.
Protein Protein Interactions (PPI) APID (Agile Protein Interaction DataAnalyzer)
 APID (Agile Protein Interaction DataAnalyzer) 23 APID (Agile Protein Interaction DataAnalyzer) Integrates and unifies 7 DBs: BIND, DIP, HPRD, IntAct, MINT, BioGRID. Includes 51,873 proteins 241,204 interactions
APID (Agile Protein Interaction DataAnalyzer) 23 APID (Agile Protein Interaction DataAnalyzer) Integrates and unifies 7 DBs: BIND, DIP, HPRD, IntAct, MINT, BioGRID. Includes 51,873 proteins 241,204 interactions
Social Media Mining. Data Mining Essentials
 Introduction Data production rate has been increased dramatically (Big Data) and we are able store much more data than before E.g., purchase data, social media data, mobile phone data Businesses and customers
Introduction Data production rate has been increased dramatically (Big Data) and we are able store much more data than before E.g., purchase data, social media data, mobile phone data Businesses and customers
TUTORIAL From Protein-Protein Interactions (PPIs) to networks and pathways ??? http://bioinfow.dep.usal.es
 TUTORIAL From Protein-Protein Interactions (PPIs) to networks and pathways http://bioinfow.dep.usal.es Dr. Javier De Las Rivas (jrivas@usal.es) Dr. Carlos Prieto (cprietos@usal.es) Bioinformatics and Functional
TUTORIAL From Protein-Protein Interactions (PPIs) to networks and pathways http://bioinfow.dep.usal.es Dr. Javier De Las Rivas (jrivas@usal.es) Dr. Carlos Prieto (cprietos@usal.es) Bioinformatics and Functional
Supplementary Information
 Supplementary Information S1: Degree Distribution of TFs in the E.coli TRN and CRN based on Operons 1000 TRN Number of TFs 100 10 y = 619.55x -1.4163 R 2 = 0.8346 1 1 10 100 1000 Degree of TFs CRN 100
Supplementary Information S1: Degree Distribution of TFs in the E.coli TRN and CRN based on Operons 1000 TRN Number of TFs 100 10 y = 619.55x -1.4163 R 2 = 0.8346 1 1 10 100 1000 Degree of TFs CRN 100
Analyzing microrna Data and Integrating mirna with Gene Expression Data in Partek Genomics Suite 6.6
 Analyzing microrna Data and Integrating mirna with Gene Expression Data in Partek Genomics Suite 6.6 Overview This tutorial outlines how microrna data can be analyzed within Partek Genomics Suite. Additionally,
Analyzing microrna Data and Integrating mirna with Gene Expression Data in Partek Genomics Suite 6.6 Overview This tutorial outlines how microrna data can be analyzed within Partek Genomics Suite. Additionally,
InSyBio BioNets: Utmost efficiency in gene expression data and biological networks analysis
 InSyBio BioNets: Utmost efficiency in gene expression data and biological networks analysis WHITE PAPER By InSyBio Ltd Konstantinos Theofilatos Bioinformatician, PhD InSyBio Technical Sales Manager August
InSyBio BioNets: Utmost efficiency in gene expression data and biological networks analysis WHITE PAPER By InSyBio Ltd Konstantinos Theofilatos Bioinformatician, PhD InSyBio Technical Sales Manager August
Frozen Robust Multi-Array Analysis and the Gene Expression Barcode
 Frozen Robust Multi-Array Analysis and the Gene Expression Barcode Matthew N. McCall October 13, 2015 Contents 1 Frozen Robust Multiarray Analysis (frma) 2 1.1 From CEL files to expression estimates...................
Frozen Robust Multi-Array Analysis and the Gene Expression Barcode Matthew N. McCall October 13, 2015 Contents 1 Frozen Robust Multiarray Analysis (frma) 2 1.1 From CEL files to expression estimates...................
Tutorial Segmentation and Classification
 MARKETING ENGINEERING FOR EXCEL TUTORIAL VERSION 1.0.8 Tutorial Segmentation and Classification Marketing Engineering for Excel is a Microsoft Excel add-in. The software runs from within Microsoft Excel
MARKETING ENGINEERING FOR EXCEL TUTORIAL VERSION 1.0.8 Tutorial Segmentation and Classification Marketing Engineering for Excel is a Microsoft Excel add-in. The software runs from within Microsoft Excel
Multivariate Analysis of Ecological Data
 Multivariate Analysis of Ecological Data MICHAEL GREENACRE Professor of Statistics at the Pompeu Fabra University in Barcelona, Spain RAUL PRIMICERIO Associate Professor of Ecology, Evolutionary Biology
Multivariate Analysis of Ecological Data MICHAEL GREENACRE Professor of Statistics at the Pompeu Fabra University in Barcelona, Spain RAUL PRIMICERIO Associate Professor of Ecology, Evolutionary Biology
Activity 4 Long-Term Effects of Drug Addiction
 Activity 4 Long-Term Effects of Drug Addiction Core Concept: Addictive drugs may lead to long-term changes in brain function. Class time required: Approximately 60-80 minutes Teacher Provides: Copy of
Activity 4 Long-Term Effects of Drug Addiction Core Concept: Addictive drugs may lead to long-term changes in brain function. Class time required: Approximately 60-80 minutes Teacher Provides: Copy of
Joint Exam 1/P Sample Exam 1
 Joint Exam 1/P Sample Exam 1 Take this practice exam under strict exam conditions: Set a timer for 3 hours; Do not stop the timer for restroom breaks; Do not look at your notes. If you believe a question
Joint Exam 1/P Sample Exam 1 Take this practice exam under strict exam conditions: Set a timer for 3 hours; Do not stop the timer for restroom breaks; Do not look at your notes. If you believe a question
Row Quantile Normalisation of Microarrays
 Row Quantile Normalisation of Microarrays W. B. Langdon Departments of Mathematical Sciences and Biological Sciences University of Essex, CO4 3SQ Technical Report CES-484 ISSN: 1744-8050 23 June 2008 Abstract
Row Quantile Normalisation of Microarrays W. B. Langdon Departments of Mathematical Sciences and Biological Sciences University of Essex, CO4 3SQ Technical Report CES-484 ISSN: 1744-8050 23 June 2008 Abstract
Comparison of Non-linear Dimensionality Reduction Techniques for Classification with Gene Expression Microarray Data
 CMPE 59H Comparison of Non-linear Dimensionality Reduction Techniques for Classification with Gene Expression Microarray Data Term Project Report Fatma Güney, Kübra Kalkan 1/15/2013 Keywords: Non-linear
CMPE 59H Comparison of Non-linear Dimensionality Reduction Techniques for Classification with Gene Expression Microarray Data Term Project Report Fatma Güney, Kübra Kalkan 1/15/2013 Keywords: Non-linear
MIC - Detecting Novel Associations in Large Data Sets. by Nico Güttler, Andreas Ströhlein and Matt Huska
 MIC - Detecting Novel Associations in Large Data Sets by Nico Güttler, Andreas Ströhlein and Matt Huska Outline Motivation Method Results Criticism Conclusions Motivation - Goal Determine important undiscovered
MIC - Detecting Novel Associations in Large Data Sets by Nico Güttler, Andreas Ströhlein and Matt Huska Outline Motivation Method Results Criticism Conclusions Motivation - Goal Determine important undiscovered
CENG 734 Advanced Topics in Bioinformatics
 CENG 734 Advanced Topics in Bioinformatics Week 9 Text Mining for Bioinformatics: BioCreative II.5 Fall 2010-2011 Quiz #7 1. Draw the decompressed graph for the following graph summary 2. Describe the
CENG 734 Advanced Topics in Bioinformatics Week 9 Text Mining for Bioinformatics: BioCreative II.5 Fall 2010-2011 Quiz #7 1. Draw the decompressed graph for the following graph summary 2. Describe the
Supplementary Figure 1: Quality Assessment of Mouse Arrays. Supplementary Figure 2: Quality Assessment of Rat Arrays
 Supplementary Figure 1: Quality Assessment of Mouse Arrays The mouse microarray data were subjected to an extensive quality-control procedure prior to conducting downstream analyses. We assessed the spread
Supplementary Figure 1: Quality Assessment of Mouse Arrays The mouse microarray data were subjected to an extensive quality-control procedure prior to conducting downstream analyses. We assessed the spread
Using MS Excel to Analyze Data: A Tutorial
 Using MS Excel to Analyze Data: A Tutorial Various data analysis tools are available and some of them are free. Because using data to improve assessment and instruction primarily involves descriptive and
Using MS Excel to Analyze Data: A Tutorial Various data analysis tools are available and some of them are free. Because using data to improve assessment and instruction primarily involves descriptive and
GENEGOBI : VISUAL DATA ANALYSIS AID TOOLS FOR MICROARRAY DATA
 COMPSTAT 2004 Symposium c Physica-Verlag/Springer 2004 GENEGOBI : VISUAL DATA ANALYSIS AID TOOLS FOR MICROARRAY DATA Eun-kyung Lee, Dianne Cook, Eve Wurtele, Dongshin Kim, Jihong Kim, and Hogeun An Key
COMPSTAT 2004 Symposium c Physica-Verlag/Springer 2004 GENEGOBI : VISUAL DATA ANALYSIS AID TOOLS FOR MICROARRAY DATA Eun-kyung Lee, Dianne Cook, Eve Wurtele, Dongshin Kim, Jihong Kim, and Hogeun An Key
An Introduction to the Use of Bayesian Network to Analyze Gene Expression Data
 n Introduction to the Use of ayesian Network to nalyze Gene Expression Data Cristina Manfredotti Dipartimento di Informatica, Sistemistica e Comunicazione (D.I.S.Co. Università degli Studi Milano-icocca
n Introduction to the Use of ayesian Network to nalyze Gene Expression Data Cristina Manfredotti Dipartimento di Informatica, Sistemistica e Comunicazione (D.I.S.Co. Università degli Studi Milano-icocca
Cluster software and Java TreeView
 Cluster software and Java TreeView To download the software: http://bonsai.hgc.jp/~mdehoon/software/cluster/software.htm http://bonsai.hgc.jp/~mdehoon/software/cluster/manual/treeview.html Cluster 3.0
Cluster software and Java TreeView To download the software: http://bonsai.hgc.jp/~mdehoon/software/cluster/software.htm http://bonsai.hgc.jp/~mdehoon/software/cluster/manual/treeview.html Cluster 3.0
1) The table lists the smoking habits of a group of college students. Answer: 0.218
 FINAL EXAM REVIEW Name ) The table lists the smoking habits of a group of college students. Sex Non-smoker Regular Smoker Heavy Smoker Total Man 5 52 5 92 Woman 8 2 2 220 Total 22 2 If a student is chosen
FINAL EXAM REVIEW Name ) The table lists the smoking habits of a group of college students. Sex Non-smoker Regular Smoker Heavy Smoker Total Man 5 52 5 92 Woman 8 2 2 220 Total 22 2 If a student is chosen
Quality Assessment of Exon and Gene Arrays
 Quality Assessment of Exon and Gene Arrays I. Introduction In this white paper we describe some quality assessment procedures that are computed from CEL files from Whole Transcript (WT) based arrays such
Quality Assessment of Exon and Gene Arrays I. Introduction In this white paper we describe some quality assessment procedures that are computed from CEL files from Whole Transcript (WT) based arrays such
ProteinQuest user guide
 ProteinQuest user guide 1. Introduction... 3 1.1 With ProteinQuest you can... 3 1.2 ProteinQuest basic version 4 1.3 ProteinQuest extended version... 5 2. ProteinQuest dictionaries... 6 3. Directions for
ProteinQuest user guide 1. Introduction... 3 1.1 With ProteinQuest you can... 3 1.2 ProteinQuest basic version 4 1.3 ProteinQuest extended version... 5 2. ProteinQuest dictionaries... 6 3. Directions for
Gene Expression Analysis
 Gene Expression Analysis Jie Peng Department of Statistics University of California, Davis May 2012 RNA expression technologies High-throughput technologies to measure the expression levels of thousands
Gene Expression Analysis Jie Peng Department of Statistics University of California, Davis May 2012 RNA expression technologies High-throughput technologies to measure the expression levels of thousands
Molecular Genetics: Challenges for Statistical Practice. J.K. Lindsey
 Molecular Genetics: Challenges for Statistical Practice J.K. Lindsey 1. What is a Microarray? 2. Design Questions 3. Modelling Questions 4. Longitudinal Data 5. Conclusions 1. What is a microarray? A microarray
Molecular Genetics: Challenges for Statistical Practice J.K. Lindsey 1. What is a Microarray? 2. Design Questions 3. Modelling Questions 4. Longitudinal Data 5. Conclusions 1. What is a microarray? A microarray
Transcribing Videos in Studiocode. In order to transcribe a video in Studiocode, you first need to open up a movie file. For this
 Transcribing Videos in Studiocode. In order to transcribe a video in Studiocode, you first need to open up a movie file. For this demonstration, we will make use of Example1.m4v. We will want to know where
Transcribing Videos in Studiocode. In order to transcribe a video in Studiocode, you first need to open up a movie file. For this demonstration, we will make use of Example1.m4v. We will want to know where
Name: Date: Use the following to answer questions 2-4:
 Name: Date: 1. A phenomenon is observed many, many times under identical conditions. The proportion of times a particular event A occurs is recorded. What does this proportion represent? A) The probability
Name: Date: 1. A phenomenon is observed many, many times under identical conditions. The proportion of times a particular event A occurs is recorded. What does this proportion represent? A) The probability
Improving the Performance of Data Mining Models with Data Preparation Using SAS Enterprise Miner Ricardo Galante, SAS Institute Brasil, São Paulo, SP
 Improving the Performance of Data Mining Models with Data Preparation Using SAS Enterprise Miner Ricardo Galante, SAS Institute Brasil, São Paulo, SP ABSTRACT In data mining modelling, data preparation
Improving the Performance of Data Mining Models with Data Preparation Using SAS Enterprise Miner Ricardo Galante, SAS Institute Brasil, São Paulo, SP ABSTRACT In data mining modelling, data preparation
Tutorial for proteome data analysis using the Perseus software platform
 Tutorial for proteome data analysis using the Perseus software platform Laboratory of Mass Spectrometry, LNBio, CNPEM Tutorial version 1.0, January 2014. Note: This tutorial was written based on the information
Tutorial for proteome data analysis using the Perseus software platform Laboratory of Mass Spectrometry, LNBio, CNPEM Tutorial version 1.0, January 2014. Note: This tutorial was written based on the information
Comparing Functional Data Analysis Approach and Nonparametric Mixed-Effects Modeling Approach for Longitudinal Data Analysis
 Comparing Functional Data Analysis Approach and Nonparametric Mixed-Effects Modeling Approach for Longitudinal Data Analysis Hulin Wu, PhD, Professor (with Dr. Shuang Wu) Department of Biostatistics &
Comparing Functional Data Analysis Approach and Nonparametric Mixed-Effects Modeling Approach for Longitudinal Data Analysis Hulin Wu, PhD, Professor (with Dr. Shuang Wu) Department of Biostatistics &
Accountable Care Organization Quality Explorer. Quick Start Guide
 Accountable Care Organization Quality Explorer Quick Start Guide 1 P age Background HealthLandscape (a division of the American Academy of Family Physicians [AAFP]) and the Robert Graham Center for Policy
Accountable Care Organization Quality Explorer Quick Start Guide 1 P age Background HealthLandscape (a division of the American Academy of Family Physicians [AAFP]) and the Robert Graham Center for Policy
DATA ANALYSIS. QEM Network HBCU-UP Fundamentals of Education Research Workshop Gerunda B. Hughes, Ph.D. Howard University
 DATA ANALYSIS QEM Network HBCU-UP Fundamentals of Education Research Workshop Gerunda B. Hughes, Ph.D. Howard University Quantitative Research What is Statistics? Statistics (as a subject) is the science
DATA ANALYSIS QEM Network HBCU-UP Fundamentals of Education Research Workshop Gerunda B. Hughes, Ph.D. Howard University Quantitative Research What is Statistics? Statistics (as a subject) is the science
1-7 2-8 3-9 4-10 5-11 6-12 7-13 8-14 9-15. Corresponding position of sirna GS
 A C GS Target transcript 5ʼ- - 3ʼ 3ʼ- 21 20 19 18 17 16 15 14 13 12 11 10 9 8 7 6 5 4 3 2 1-5ʼ 10-16 11-17 12-18 13-19 14-20 15-21 sivim-270 3ʼUTR 1-7 2-8 3-9 4-10 5-11 6-12 7-13 8-14 9-15 Corresponding
A C GS Target transcript 5ʼ- - 3ʼ 3ʼ- 21 20 19 18 17 16 15 14 13 12 11 10 9 8 7 6 5 4 3 2 1-5ʼ 10-16 11-17 12-18 13-19 14-20 15-21 sivim-270 3ʼUTR 1-7 2-8 3-9 4-10 5-11 6-12 7-13 8-14 9-15 Corresponding
Graph theoretic approach to analyze amino acid network
 Int. J. Adv. Appl. Math. and Mech. 2(3) (2015) 31-37 (ISSN: 2347-2529) Journal homepage: www.ijaamm.com International Journal of Advances in Applied Mathematics and Mechanics Graph theoretic approach to
Int. J. Adv. Appl. Math. and Mech. 2(3) (2015) 31-37 (ISSN: 2347-2529) Journal homepage: www.ijaamm.com International Journal of Advances in Applied Mathematics and Mechanics Graph theoretic approach to
Predictive Gene Signature Selection for Adjuvant Chemotherapy in Non-Small Cell Lung Cancer Patients
 Predictive Gene Signature Selection for Adjuvant Chemotherapy in Non-Small Cell Lung Cancer Patients by Li Liu A practicum report submitted to the Department of Public Health Sciences in conformity with
Predictive Gene Signature Selection for Adjuvant Chemotherapy in Non-Small Cell Lung Cancer Patients by Li Liu A practicum report submitted to the Department of Public Health Sciences in conformity with
Univariate Regression
 Univariate Regression Correlation and Regression The regression line summarizes the linear relationship between 2 variables Correlation coefficient, r, measures strength of relationship: the closer r is
Univariate Regression Correlation and Regression The regression line summarizes the linear relationship between 2 variables Correlation coefficient, r, measures strength of relationship: the closer r is
Predict the Popularity of YouTube Videos Using Early View Data
 000 001 002 003 004 005 006 007 008 009 010 011 012 013 014 015 016 017 018 019 020 021 022 023 024 025 026 027 028 029 030 031 032 033 034 035 036 037 038 039 040 041 042 043 044 045 046 047 048 049 050
000 001 002 003 004 005 006 007 008 009 010 011 012 013 014 015 016 017 018 019 020 021 022 023 024 025 026 027 028 029 030 031 032 033 034 035 036 037 038 039 040 041 042 043 044 045 046 047 048 049 050
ADD-INS: ENHANCING EXCEL
 CHAPTER 9 ADD-INS: ENHANCING EXCEL This chapter discusses the following topics: WHAT CAN AN ADD-IN DO? WHY USE AN ADD-IN (AND NOT JUST EXCEL MACROS/PROGRAMS)? ADD INS INSTALLED WITH EXCEL OTHER ADD-INS
CHAPTER 9 ADD-INS: ENHANCING EXCEL This chapter discusses the following topics: WHAT CAN AN ADD-IN DO? WHY USE AN ADD-IN (AND NOT JUST EXCEL MACROS/PROGRAMS)? ADD INS INSTALLED WITH EXCEL OTHER ADD-INS
SeattleSNPs Interactive Tutorial: Web Tools for Site Selection, Linkage Disequilibrium and Haplotype Analysis
 SeattleSNPs Interactive Tutorial: Web Tools for Site Selection, Linkage Disequilibrium and Haplotype Analysis Goal: This tutorial introduces several websites and tools useful for determining linkage disequilibrium
SeattleSNPs Interactive Tutorial: Web Tools for Site Selection, Linkage Disequilibrium and Haplotype Analysis Goal: This tutorial introduces several websites and tools useful for determining linkage disequilibrium
Drawing a histogram using Excel
 Drawing a histogram using Excel STEP 1: Examine the data to decide how many class intervals you need and what the class boundaries should be. (In an assignment you may be told what class boundaries to
Drawing a histogram using Excel STEP 1: Examine the data to decide how many class intervals you need and what the class boundaries should be. (In an assignment you may be told what class boundaries to
MHI3000 Big Data Analytics for Health Care Final Project Report
 MHI3000 Big Data Analytics for Health Care Final Project Report Zhongtian Fred Qiu (1002274530) http://gallery.azureml.net/details/81ddb2ab137046d4925584b5095ec7aa 1. Data pre-processing The data given
MHI3000 Big Data Analytics for Health Care Final Project Report Zhongtian Fred Qiu (1002274530) http://gallery.azureml.net/details/81ddb2ab137046d4925584b5095ec7aa 1. Data pre-processing The data given
Data Mining 5. Cluster Analysis
 Data Mining 5. Cluster Analysis 5.2 Fall 2009 Instructor: Dr. Masoud Yaghini Outline Data Structures Interval-Valued (Numeric) Variables Binary Variables Categorical Variables Ordinal Variables Variables
Data Mining 5. Cluster Analysis 5.2 Fall 2009 Instructor: Dr. Masoud Yaghini Outline Data Structures Interval-Valued (Numeric) Variables Binary Variables Categorical Variables Ordinal Variables Variables
EdgeLap: Identifying and discovering features from overlapping sets in networks
 Project Title: EdgeLap: Identifying and discovering features from overlapping sets in networks Names and Email Addresses: Jessica Wong (jhmwong@cs.ubc.ca) Aria Hahn (hahnaria@gmail.com) Sarah Perez (karatezeus21@gmail.com)
Project Title: EdgeLap: Identifying and discovering features from overlapping sets in networks Names and Email Addresses: Jessica Wong (jhmwong@cs.ubc.ca) Aria Hahn (hahnaria@gmail.com) Sarah Perez (karatezeus21@gmail.com)
D-optimal plans in observational studies
 D-optimal plans in observational studies Constanze Pumplün Stefan Rüping Katharina Morik Claus Weihs October 11, 2005 Abstract This paper investigates the use of Design of Experiments in observational
D-optimal plans in observational studies Constanze Pumplün Stefan Rüping Katharina Morik Claus Weihs October 11, 2005 Abstract This paper investigates the use of Design of Experiments in observational
Recognition of T cell epitopes (Abbas Chapter 6)
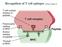 Recognition of T cell epitopes (Abbas Chapter 6) Functions of different APCs (Abbas Chapter 6)!!! Directon Routes of antigen entry (Abbas Chapter 6) Flow of Information Barrier APCs LNs Sequence of Events
Recognition of T cell epitopes (Abbas Chapter 6) Functions of different APCs (Abbas Chapter 6)!!! Directon Routes of antigen entry (Abbas Chapter 6) Flow of Information Barrier APCs LNs Sequence of Events
Provided by the American Venous Forum: veinforum.org
 CHAPTER 1 NORMAL VENOUS CIRCULATION Original author: Frank Padberg Abstracted by Teresa L.Carman Introduction The circulatory system is responsible for circulating (moving) blood throughout the body. The
CHAPTER 1 NORMAL VENOUS CIRCULATION Original author: Frank Padberg Abstracted by Teresa L.Carman Introduction The circulatory system is responsible for circulating (moving) blood throughout the body. The
MANUAL PLATELET COUNT
 MANUAL PLATELET COUNT Principle Whole blood is diluted with a 1% ammonium oxalate solution. The isotonic balance of the diluent is such that all erythrocytes are lysed while the leukocytes, platelets,
MANUAL PLATELET COUNT Principle Whole blood is diluted with a 1% ammonium oxalate solution. The isotonic balance of the diluent is such that all erythrocytes are lysed while the leukocytes, platelets,
The Kendall Rank Correlation Coefficient
 The Kendall Rank Correlation Coefficient Hervé Abdi Overview The Kendall (955) rank correlation coefficient evaluates the degree of similarity between two sets of ranks given to a same set of objects.
The Kendall Rank Correlation Coefficient Hervé Abdi Overview The Kendall (955) rank correlation coefficient evaluates the degree of similarity between two sets of ranks given to a same set of objects.
CHAPTER 4 GRID SCHEDULER WITH DEVIATION BASED RESOURCE SCHEDULING
 46 CHAPTER 4 GRID SCHEDULER WITH DEVIATION BASED RESOURCE SCHEDULING 4.1 OUTLINE In this chapter, the significance of policy problem and its relationship with grid scheduling is explained in detail with
46 CHAPTER 4 GRID SCHEDULER WITH DEVIATION BASED RESOURCE SCHEDULING 4.1 OUTLINE In this chapter, the significance of policy problem and its relationship with grid scheduling is explained in detail with
Analysis of gene expression data. Ulf Leser and Philippe Thomas
 Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
BASIC STATISTICAL METHODS FOR GENOMIC DATA ANALYSIS
 BASIC STATISTICAL METHODS FOR GENOMIC DATA ANALYSIS SEEMA JAGGI Indian Agricultural Statistics Research Institute Library Avenue, New Delhi-110 012 seema@iasri.res.in Genomics A genome is an organism s
BASIC STATISTICAL METHODS FOR GENOMIC DATA ANALYSIS SEEMA JAGGI Indian Agricultural Statistics Research Institute Library Avenue, New Delhi-110 012 seema@iasri.res.in Genomics A genome is an organism s
Data Acquisition. DNA microarrays. The functional genomics pipeline. Experimental design affects outcome data analysis
 Data Acquisition DNA microarrays The functional genomics pipeline Experimental design affects outcome data analysis Data acquisition microarray processing Data preprocessing scaling/normalization/filtering
Data Acquisition DNA microarrays The functional genomics pipeline Experimental design affects outcome data analysis Data acquisition microarray processing Data preprocessing scaling/normalization/filtering
Bioinformatics: Network Analysis
 Bioinformatics: Network Analysis Graph-theoretic Properties of Biological Networks COMP 572 (BIOS 572 / BIOE 564) - Fall 2013 Luay Nakhleh, Rice University 1 Outline Architectural features Motifs, modules,
Bioinformatics: Network Analysis Graph-theoretic Properties of Biological Networks COMP 572 (BIOS 572 / BIOE 564) - Fall 2013 Luay Nakhleh, Rice University 1 Outline Architectural features Motifs, modules,
AGILENT S BIOINFORMATICS ANALYSIS SOFTWARE
 ACCELERATING PROGRESS IS IN OUR GENES AGILENT S BIOINFORMATICS ANALYSIS SOFTWARE GENESPRING GENE EXPRESSION (GX) MASS PROFILER PROFESSIONAL (MPP) PATHWAY ARCHITECT (PA) See Deeper. Reach Further. BIOINFORMATICS
ACCELERATING PROGRESS IS IN OUR GENES AGILENT S BIOINFORMATICS ANALYSIS SOFTWARE GENESPRING GENE EXPRESSION (GX) MASS PROFILER PROFESSIONAL (MPP) PATHWAY ARCHITECT (PA) See Deeper. Reach Further. BIOINFORMATICS
How To Run Statistical Tests in Excel
 How To Run Statistical Tests in Excel Microsoft Excel is your best tool for storing and manipulating data, calculating basic descriptive statistics such as means and standard deviations, and conducting
How To Run Statistical Tests in Excel Microsoft Excel is your best tool for storing and manipulating data, calculating basic descriptive statistics such as means and standard deviations, and conducting
AN ILLUSTRATION OF COMPARATIVE QUANTITATIVE RESULTS USING ALTERNATIVE ANALYTICAL TECHNIQUES
 CHAPTER 8. AN ILLUSTRATION OF COMPARATIVE QUANTITATIVE RESULTS USING ALTERNATIVE ANALYTICAL TECHNIQUES Based on TCRP B-11 Field Test Results CTA CHICAGO, ILLINOIS RED LINE SERVICE: 8A. CTA Red Line - Computation
CHAPTER 8. AN ILLUSTRATION OF COMPARATIVE QUANTITATIVE RESULTS USING ALTERNATIVE ANALYTICAL TECHNIQUES Based on TCRP B-11 Field Test Results CTA CHICAGO, ILLINOIS RED LINE SERVICE: 8A. CTA Red Line - Computation
A Tutorial on dynamic networks. By Clement Levallois, Erasmus University Rotterdam
 A Tutorial on dynamic networks By, Erasmus University Rotterdam V 1.0-2013 Bio notes Education in economics, management, history of science (Ph.D.) Since 2008, turned to digital methods for research. data
A Tutorial on dynamic networks By, Erasmus University Rotterdam V 1.0-2013 Bio notes Education in economics, management, history of science (Ph.D.) Since 2008, turned to digital methods for research. data
A General Framework for Weighted Gene Co-expression Network Analysis
 Please cite: Statistical Applications in Genetics and Molecular Biology (2005). A General Framework for Weighted Gene Co-expression Network Analysis Bin Zhang and Steve Horvath Departments of Human Genetics
Please cite: Statistical Applications in Genetics and Molecular Biology (2005). A General Framework for Weighted Gene Co-expression Network Analysis Bin Zhang and Steve Horvath Departments of Human Genetics
Hierarchical Clustering Analysis
 Hierarchical Clustering Analysis What is Hierarchical Clustering? Hierarchical clustering is used to group similar objects into clusters. In the beginning, each row and/or column is considered a cluster.
Hierarchical Clustering Analysis What is Hierarchical Clustering? Hierarchical clustering is used to group similar objects into clusters. In the beginning, each row and/or column is considered a cluster.
Solutions to Homework 6 Statistics 302 Professor Larget
 s to Homework 6 Statistics 302 Professor Larget Textbook Exercises 5.29 (Graded for Completeness) What Proportion Have College Degrees? According to the US Census Bureau, about 27.5% of US adults over
s to Homework 6 Statistics 302 Professor Larget Textbook Exercises 5.29 (Graded for Completeness) What Proportion Have College Degrees? According to the US Census Bureau, about 27.5% of US adults over
Chapter 2: Supplementary Exercises
 Chapter 2: Supplementary Exercises 2.S.1. A Public Analyst s laboratory routinely measures potable water samples by flame atomic absorption spectrometry to ensure compliance with the EU drinking water
Chapter 2: Supplementary Exercises 2.S.1. A Public Analyst s laboratory routinely measures potable water samples by flame atomic absorption spectrometry to ensure compliance with the EU drinking water
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS)
 Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
Comparing Multiple Proportions, Test of Independence and Goodness of Fit
 Comparing Multiple Proportions, Test of Independence and Goodness of Fit Content Testing the Equality of Population Proportions for Three or More Populations Test of Independence Goodness of Fit Test 2
Comparing Multiple Proportions, Test of Independence and Goodness of Fit Content Testing the Equality of Population Proportions for Three or More Populations Test of Independence Goodness of Fit Test 2
STATISTICA Formula Guide: Logistic Regression. Table of Contents
 : Table of Contents... 1 Overview of Model... 1 Dispersion... 2 Parameterization... 3 Sigma-Restricted Model... 3 Overparameterized Model... 4 Reference Coding... 4 Model Summary (Summary Tab)... 5 Summary
: Table of Contents... 1 Overview of Model... 1 Dispersion... 2 Parameterization... 3 Sigma-Restricted Model... 3 Overparameterized Model... 4 Reference Coding... 4 Model Summary (Summary Tab)... 5 Summary
Functions of Blood. Collects O 2 from lungs, nutrients from digestive tract, and waste products from tissues Helps maintain homeostasis
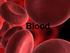 Blood Objectives Describe the functions of blood Describe blood plasma Explain the functions of red blood cells, white blood cells, and platelets Summarize the process of blood clotting What is Blood?
Blood Objectives Describe the functions of blood Describe blood plasma Explain the functions of red blood cells, white blood cells, and platelets Summarize the process of blood clotting What is Blood?
How To Check For Differences In The One Way Anova
 MINITAB ASSISTANT WHITE PAPER This paper explains the research conducted by Minitab statisticians to develop the methods and data checks used in the Assistant in Minitab 17 Statistical Software. One-Way
MINITAB ASSISTANT WHITE PAPER This paper explains the research conducted by Minitab statisticians to develop the methods and data checks used in the Assistant in Minitab 17 Statistical Software. One-Way
Introduction to Exploratory Data Analysis
 Introduction to Exploratory Data Analysis A SpaceStat Software Tutorial Copyright 2013, BioMedware, Inc. (www.biomedware.com). All rights reserved. SpaceStat and BioMedware are trademarks of BioMedware,
Introduction to Exploratory Data Analysis A SpaceStat Software Tutorial Copyright 2013, BioMedware, Inc. (www.biomedware.com). All rights reserved. SpaceStat and BioMedware are trademarks of BioMedware,
Basic Analysis of Microarray Data
 Basic Analysis of Microarray Data A User Guide and Tutorial Scott A. Ness, Ph.D. Co-Director, Keck-UNM Genomics Resource and Dept. of Molecular Genetics and Microbiology University of New Mexico HSC Tel.
Basic Analysis of Microarray Data A User Guide and Tutorial Scott A. Ness, Ph.D. Co-Director, Keck-UNM Genomics Resource and Dept. of Molecular Genetics and Microbiology University of New Mexico HSC Tel.
We are often interested in the relationship between two variables. Do people with more years of full-time education earn higher salaries?
 Statistics: Correlation Richard Buxton. 2008. 1 Introduction We are often interested in the relationship between two variables. Do people with more years of full-time education earn higher salaries? Do
Statistics: Correlation Richard Buxton. 2008. 1 Introduction We are often interested in the relationship between two variables. Do people with more years of full-time education earn higher salaries? Do
Inferring the role of transcription factors in regulatory networks
 Inferring the role of transcription factors in regulatory networks Philippe Veber 1, Carito Guziolowski 1, Michel Le Borgne 2, Ovidiu Radulescu 1,3 and Anne Siegel 4 1 Centre INRIA Rennes Bretagne Atlantique,
Inferring the role of transcription factors in regulatory networks Philippe Veber 1, Carito Guziolowski 1, Michel Le Borgne 2, Ovidiu Radulescu 1,3 and Anne Siegel 4 1 Centre INRIA Rennes Bretagne Atlantique,
Two-Way ANOVA tests. I. Definition and Applications...2. II. Two-Way ANOVA prerequisites...2. III. How to use the Two-Way ANOVA tool?...
 Two-Way ANOVA tests Contents at a glance I. Definition and Applications...2 II. Two-Way ANOVA prerequisites...2 III. How to use the Two-Way ANOVA tool?...3 A. Parametric test, assume variances equal....4
Two-Way ANOVA tests Contents at a glance I. Definition and Applications...2 II. Two-Way ANOVA prerequisites...2 III. How to use the Two-Way ANOVA tool?...3 A. Parametric test, assume variances equal....4
Using Excel in Research. Hui Bian Office for Faculty Excellence
 Using Excel in Research Hui Bian Office for Faculty Excellence Data entry in Excel Directly type information into the cells Enter data using Form Command: File > Options 2 Data entry in Excel Tool bar:
Using Excel in Research Hui Bian Office for Faculty Excellence Data entry in Excel Directly type information into the cells Enter data using Form Command: File > Options 2 Data entry in Excel Tool bar:
IC05 Introduction on Networks &Visualization Nov. 2009. <mathieu.bastian@gmail.com>
 IC05 Introduction on Networks &Visualization Nov. 2009 Overview 1. Networks Introduction Networks across disciplines Properties Models 2. Visualization InfoVis Data exploration
IC05 Introduction on Networks &Visualization Nov. 2009 Overview 1. Networks Introduction Networks across disciplines Properties Models 2. Visualization InfoVis Data exploration
Overview What is a PivotTable? Benefits
 Overview What is a PivotTable? Benefits Create a PivotTable Select Row & Column labels & Values Filtering & Sorting Calculations Data Details Refresh Data Design options Create a PivotChart Slicers Charts
Overview What is a PivotTable? Benefits Create a PivotTable Select Row & Column labels & Values Filtering & Sorting Calculations Data Details Refresh Data Design options Create a PivotChart Slicers Charts
STATS8: Introduction to Biostatistics. Data Exploration. Babak Shahbaba Department of Statistics, UCI
 STATS8: Introduction to Biostatistics Data Exploration Babak Shahbaba Department of Statistics, UCI Introduction After clearly defining the scientific problem, selecting a set of representative members
STATS8: Introduction to Biostatistics Data Exploration Babak Shahbaba Department of Statistics, UCI Introduction After clearly defining the scientific problem, selecting a set of representative members
Distances, Clustering, and Classification. Heatmaps
 Distances, Clustering, and Classification Heatmaps 1 Distance Clustering organizes things that are close into groups What does it mean for two genes to be close? What does it mean for two samples to be
Distances, Clustering, and Classification Heatmaps 1 Distance Clustering organizes things that are close into groups What does it mean for two genes to be close? What does it mean for two samples to be
SICKLE CELL ANEMIA & THE HEMOGLOBIN GENE TEACHER S GUIDE
 AP Biology Date SICKLE CELL ANEMIA & THE HEMOGLOBIN GENE TEACHER S GUIDE LEARNING OBJECTIVES Students will gain an appreciation of the physical effects of sickle cell anemia, its prevalence in the population,
AP Biology Date SICKLE CELL ANEMIA & THE HEMOGLOBIN GENE TEACHER S GUIDE LEARNING OBJECTIVES Students will gain an appreciation of the physical effects of sickle cell anemia, its prevalence in the population,
1 Nonparametric Statistics
 1 Nonparametric Statistics When finding confidence intervals or conducting tests so far, we always described the population with a model, which includes a set of parameters. Then we could make decisions
1 Nonparametric Statistics When finding confidence intervals or conducting tests so far, we always described the population with a model, which includes a set of parameters. Then we could make decisions
Capturing Best Practice for Microarray Gene Expression Data Analysis
 Capturing Best Practice for Microarray Gene Expression Data Analysis Gregory Piatetsky-Shapiro KDnuggets gps@kdnuggets.com Tom Khabaza SPSS tkhabaza@spss.com Sridhar Ramaswamy MIT / Whitehead Institute
Capturing Best Practice for Microarray Gene Expression Data Analysis Gregory Piatetsky-Shapiro KDnuggets gps@kdnuggets.com Tom Khabaza SPSS tkhabaza@spss.com Sridhar Ramaswamy MIT / Whitehead Institute
Application note for listmode scanning
 Application note for listmode scanning This application note explains what listmode acquisition is, how a listmode scan is done and how to process a listmode scan with SkyScan software. What is listmode?
Application note for listmode scanning This application note explains what listmode acquisition is, how a listmode scan is done and how to process a listmode scan with SkyScan software. What is listmode?
USING SPECTRAL RADIUS RATIO FOR NODE DEGREE TO ANALYZE THE EVOLUTION OF SCALE- FREE NETWORKS AND SMALL-WORLD NETWORKS
 USING SPECTRAL RADIUS RATIO FOR NODE DEGREE TO ANALYZE THE EVOLUTION OF SCALE- FREE NETWORKS AND SMALL-WORLD NETWORKS Natarajan Meghanathan Jackson State University, 1400 Lynch St, Jackson, MS, USA natarajan.meghanathan@jsums.edu
USING SPECTRAL RADIUS RATIO FOR NODE DEGREE TO ANALYZE THE EVOLUTION OF SCALE- FREE NETWORKS AND SMALL-WORLD NETWORKS Natarajan Meghanathan Jackson State University, 1400 Lynch St, Jackson, MS, USA natarajan.meghanathan@jsums.edu
A Survey on Pre-processing and Post-processing Techniques in Data Mining
 , pp. 99-128 http://dx.doi.org/10.14257/ijdta.2014.7.4.09 A Survey on Pre-processing and Post-processing Techniques in Data Mining Divya Tomar and Sonali Agarwal Indian Institute of Information Technology,
, pp. 99-128 http://dx.doi.org/10.14257/ijdta.2014.7.4.09 A Survey on Pre-processing and Post-processing Techniques in Data Mining Divya Tomar and Sonali Agarwal Indian Institute of Information Technology,
!!!!!!!!!!!!!!!!!!!!!!!!!!
 Figure S7 cdl-1 3'UTR gld-1 binding site (4..10) let-7 binding site (326..346) Figure Legends Figure S1. gld-1 genetically interacts with nhl-2 and vig-1 during germline development. (A) nhl-2(ok818) enhances
Figure S7 cdl-1 3'UTR gld-1 binding site (4..10) let-7 binding site (326..346) Figure Legends Figure S1. gld-1 genetically interacts with nhl-2 and vig-1 during germline development. (A) nhl-2(ok818) enhances
Next Generation Sequencing: Adjusting to Big Data. Daniel Nicorici, Dr.Tech. Statistikot Suomen Lääketeollisuudessa 29.10.2013
 Next Generation Sequencing: Adjusting to Big Data Daniel Nicorici, Dr.Tech. Statistikot Suomen Lääketeollisuudessa 29.10.2013 Outline Human Genome Project Next-Generation Sequencing Personalized Medicine
Next Generation Sequencing: Adjusting to Big Data Daniel Nicorici, Dr.Tech. Statistikot Suomen Lääketeollisuudessa 29.10.2013 Outline Human Genome Project Next-Generation Sequencing Personalized Medicine
Course on Functional Analysis. ::: Gene Set Enrichment Analysis - GSEA -
 Course on Functional Analysis ::: Madrid, June 31st, 2007. Gonzalo Gómez, PhD. ggomez@cnio.es Bioinformatics Unit CNIO ::: Contents. 1. Introduction. 2. GSEA Software 3. Data Formats 4. Using GSEA 5. GSEA
Course on Functional Analysis ::: Madrid, June 31st, 2007. Gonzalo Gómez, PhD. ggomez@cnio.es Bioinformatics Unit CNIO ::: Contents. 1. Introduction. 2. GSEA Software 3. Data Formats 4. Using GSEA 5. GSEA
Package COSINE. February 19, 2015
 Type Package Title COndition SpecIfic sub-network Version 2.1 Date 2014-07-09 Author Package COSINE February 19, 2015 Maintainer Depends R (>= 3.1.0), MASS,genalg To identify
Type Package Title COndition SpecIfic sub-network Version 2.1 Date 2014-07-09 Author Package COSINE February 19, 2015 Maintainer Depends R (>= 3.1.0), MASS,genalg To identify
Supplementary Figure 1 Characterization of 5xFAD derived hippocampal cultures. A-B) Average total numbers of cells per hippocampus as well as average
 Supplementary Figure 1 Characterization of 5xFAD derived hippocampal cultures. A-B) Average total numbers of cells per hippocampus as well as average total numbers of cells per field of view were similar
Supplementary Figure 1 Characterization of 5xFAD derived hippocampal cultures. A-B) Average total numbers of cells per hippocampus as well as average total numbers of cells per field of view were similar
Supervised and unsupervised learning - 1
 Chapter 3 Supervised and unsupervised learning - 1 3.1 Introduction The science of learning plays a key role in the field of statistics, data mining, artificial intelligence, intersecting with areas in
Chapter 3 Supervised and unsupervised learning - 1 3.1 Introduction The science of learning plays a key role in the field of statistics, data mining, artificial intelligence, intersecting with areas in
Nominal and ordinal logistic regression
 Nominal and ordinal logistic regression April 26 Nominal and ordinal logistic regression Our goal for today is to briefly go over ways to extend the logistic regression model to the case where the outcome
Nominal and ordinal logistic regression April 26 Nominal and ordinal logistic regression Our goal for today is to briefly go over ways to extend the logistic regression model to the case where the outcome
Auxiliary Variables in Mixture Modeling: 3-Step Approaches Using Mplus
 Auxiliary Variables in Mixture Modeling: 3-Step Approaches Using Mplus Tihomir Asparouhov and Bengt Muthén Mplus Web Notes: No. 15 Version 8, August 5, 2014 1 Abstract This paper discusses alternatives
Auxiliary Variables in Mixture Modeling: 3-Step Approaches Using Mplus Tihomir Asparouhov and Bengt Muthén Mplus Web Notes: No. 15 Version 8, August 5, 2014 1 Abstract This paper discusses alternatives
Evaluation & Validation: Credibility: Evaluating what has been learned
 Evaluation & Validation: Credibility: Evaluating what has been learned How predictive is a learned model? How can we evaluate a model Test the model Statistical tests Considerations in evaluating a Model
Evaluation & Validation: Credibility: Evaluating what has been learned How predictive is a learned model? How can we evaluate a model Test the model Statistical tests Considerations in evaluating a Model
Moderation. Moderation
 Stats - Moderation Moderation A moderator is a variable that specifies conditions under which a given predictor is related to an outcome. The moderator explains when a DV and IV are related. Moderation
Stats - Moderation Moderation A moderator is a variable that specifies conditions under which a given predictor is related to an outcome. The moderator explains when a DV and IV are related. Moderation
Integrating DNA Motif Discovery and Genome-Wide Expression Analysis. Erin M. Conlon
 Integrating DNA Motif Discovery and Genome-Wide Expression Analysis Department of Mathematics and Statistics University of Massachusetts Amherst Statistics in Functional Genomics Workshop Ascona, Switzerland
Integrating DNA Motif Discovery and Genome-Wide Expression Analysis Department of Mathematics and Statistics University of Massachusetts Amherst Statistics in Functional Genomics Workshop Ascona, Switzerland
Windows-Based Meta-Analysis Software. Package. Version 2.0
 1 Windows-Based Meta-Analysis Software Package Version 2.0 The Hunter-Schmidt Meta-Analysis Programs Package includes six programs that implement all basic types of Hunter-Schmidt psychometric meta-analysis
1 Windows-Based Meta-Analysis Software Package Version 2.0 The Hunter-Schmidt Meta-Analysis Programs Package includes six programs that implement all basic types of Hunter-Schmidt psychometric meta-analysis
DeCyder Extended Data Analysis module Version 1.0
 GE Healthcare DeCyder Extended Data Analysis module Version 1.0 Module for DeCyder 2D version 6.5 User Manual Contents 1 Introduction 1.1 Introduction... 7 1.2 The DeCyder EDA User Manual... 9 1.3 Getting
GE Healthcare DeCyder Extended Data Analysis module Version 1.0 Module for DeCyder 2D version 6.5 User Manual Contents 1 Introduction 1.1 Introduction... 7 1.2 The DeCyder EDA User Manual... 9 1.3 Getting
netresponse probabilistic tools for functional network analysis
 netresponse probabilistic tools for functional network analysis Leo Lahti 1,2, Olli-Pekka Huovilainen 1, António Gusmão 1 and Juuso Parkkinen 1 (1) Dpt. Information and Computer Science, Aalto University,
netresponse probabilistic tools for functional network analysis Leo Lahti 1,2, Olli-Pekka Huovilainen 1, António Gusmão 1 and Juuso Parkkinen 1 (1) Dpt. Information and Computer Science, Aalto University,
Correlation of microarray and quantitative real-time PCR results. Elisa Wurmbach Mount Sinai School of Medicine New York
 Correlation of microarray and quantitative real-time PCR results Elisa Wurmbach Mount Sinai School of Medicine New York Microarray techniques Oligo-array: Affymetrix, Codelink, spotted oligo-arrays (60-70mers)
Correlation of microarray and quantitative real-time PCR results Elisa Wurmbach Mount Sinai School of Medicine New York Microarray techniques Oligo-array: Affymetrix, Codelink, spotted oligo-arrays (60-70mers)
Molecule Shapes. support@ingenuity.com www.ingenuity.com 1
 IPA 8 Legend This legend provides a key of the main features of Network Explorer and Canonical Pathways, including molecule shapes and colors as well as relationship labels and types. For a high-resolution
IPA 8 Legend This legend provides a key of the main features of Network Explorer and Canonical Pathways, including molecule shapes and colors as well as relationship labels and types. For a high-resolution
