Gromacs Introductory Tutorial
|
|
|
- Silvester Robbins
- 7 years ago
- Views:
Transcription
1 Gromacs Introductory Tutorial Gromacs ver 4.6 John E. Kerrigan, Ph.D. Associate Director Bioinformatics The Cancer Institute of New Jersey Rutgers, The State University of NJ 120 Albany Street New Brunswick, NJ USA
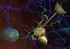
2 Gromacs 4.6 Tutorial Funnel Web Spider toxin using Amber99SB-ILDN force field. GROMACS Tutorial for Solvation Study of Spider Toxin Peptide. Yu, H., Rosen, M. K., Saccomano, N. A., Phillips, D., Volkmann, R. A., Schreiber, S. L.: Sequential assignment and structure determination of spider toxin omega-aga-ivb. Biochemistry 32 pp (1993) GROMACS is a high-end, high performance research tool designed for the study of protein dynamics using classical molecular dynamics theory.[1, 2] This versatile software package is Gnu public license free software. You may download it from GROMACS runs on linux, unix, and on Windows. For this tutorial, we used Gromacs version 4.6 compiled with FFTW ver libraries. Synopsis. In this tutorial, you will study a toxin isolated from the venom of the funnel web spider. Venom toxins have been used in the past to identify cation channels. Calcium ion channels regulate the entry of this ion into cells. Nerve signals are highly governed by the balance of ions in neuronal cells. It is hypothesized that exposed positively charged residues in venoms like the spider toxin here bind favorably to the negatively charged entrance of the cell s ion channel. The spider toxin in this tutorial has its positively charged residues pointing predominantly to one side of the peptide. The ion channel is blocked, resulting in blocked nerve signal leading to paralysis and ultimately to death (presumably via respiratory failure). We will study this peptide toxin using explicit solvation dynamics. We will solvate the peptide in a water box followed by equilibration using Newton s laws of motion. We will compare and contrast the impact of counterions in the explicit solvation simulation. We will seek answers to the following questions: Is the secondary structure stable to the dynamics conditions? Are the side chains of the positively charged residues predominantly displaced to one side of the peptide structure? Do the counterions hold these positively charged residues in place or do they move around? What role does water play in maintaining the structure of proteins? Download 1OMB.PDB from the Protein Data Bank ( It is advisable to use DeepView (Download DeepView from spdbv/ ) to preview the file if you know that your structure may be disordered (i.e. residues with missing side chains). DeepView will replace any missing side chains 2
3 (However, beware as Deep View might mark those rebuilt side chains with a strange control character that can only be removed manually using a text editor!). There are no missing side chains in this pdb file, so we will not worry about that in this exercise. However, you will need to add OXT at the C-terminal end. This can be done using the Build menu in DeepView. Save the file as fws.pdb. Create a project directory called fwspider and move 1OMB.pdb and fws.pdb to that directory. The pdb2gmx command creates input coordinates and topology for gromacs to use. The -ignh flag ignores any hydrogen atoms in the input pdb file. We use the -ff flag to designate the force field to be used and the -water flag to designate the water model. We are using the Amber99SB-ILDN force field[3] and the TIP3P water model [4]. pdb2gmx -ignh -ff amber99sb-ildn -f fws.pdb -o fws.gro -p fws.top -water tip3p Next setup the periodic box. We will use a rhombic dodecahedron box as it is the most spherical box with the -bt flag. We use the -d flag to set the spacing distance in nm. You may also use editconf to convert gro files to pdb files. editconf -f fws.gro -o fws-pbc.gro -bt dodecahedron -d 1.2 Now we begin setup of our system using in vacuo minimization. The following is the content of file em-vac-pme.mdp. em-vac-pme.mdp ; Define can be used to control processes define = -DFLEXIBLE ; Use flexible water model ; Parameters describing what to do, when to stop and what to save integrator = steep ; Algorithm (steep = steepest descent minimization) emtol = ; Stop minimization when the maximum force < 1.0 kj/mol nsteps = 1000 ; Maximum number of (minimization) steps to perform nstenergy = 1 ; Write energies to disk every nstenergy steps energygrps = System ; Which energy group(s) to write to disk ; Parameters describing how to find the neighbors of each atom and how to calculate the interactions nstlist = 1 ; Frequency to update the neighbor list ns_type = grid ; Method to determine neighbor list (simple, grid) coulombtype = PME ; Treatment of long range electrostatic interactions rlist = 1.0 ; Cut-off for making neighbor list (short range forces) rcoulomb = 1.0 ; long range electrostatic cut-off rvdw = 1.0 ; long range Van der Waals cut-off constraints = none ; Bond types to replace by constraints pbc = xyz ; Periodic Boundary Conditions (yes/no) 3
4 Use the gromacs pre-processor (grompp) to generate the run input file (tpr file). grompp -f em-vac-pme.mdp -c fws-pbc.gro -p fws.top -o em-vac.tpr mdrun -v -deffnm em-vac Fill the periodic box with water using genbox. genbox -cp em-vac.gro -cs spc216.gro -p fws.top -o fws-b4ion.gro Solvated system energy minimization (em-sol-pme.mdp) ; Define can be used to control processes define = -DFLEXIBLE ; Parameters describing what to do, when to stop and what to save integrator = steep ; Algorithm (steep = steepest descent minimization) emtol = ; Stop minimization when the maximum force < 1.0 kj/mol nsteps = 5000 ; Maximum number of (minimization) steps to perform nstenergy = 1 ; Write energies to disk every nstenergy steps energygrps = System ; Which energy group(s) to write to disk ; Parameters describing how to find the neighbors of each atom and how to calculate the interactions nstlist = 1 ; Frequency to update the neighbor list ns_type = grid ; Method to determine neighbor list (simple, grid) coulombtype = PME ; Treatment of long range electrostatic interactions rlist = 1.0 ; short-range neighborlist cutoff (in nm) rcoulomb = 1.0 ; long range electrostatic cut-off rvdw = 1.0 ; long range Van der Waals cut-off constraints = none ; Bond types to replace by constraints pbc = xyz ; Periodic Boundary Conditions (yes/no) grompp -f em-sol-pme.mdp -c fws-b4ion.gro -p fws.top -o ion.tpr We will use this first tpr file (designated as ion.tpr ) to add the ions to the system. genion -s ion.tpr -o fws-b4em.gro -neutral -conc p fws.top -g ion.log When prompted select - Solvent. The genion command will replace water molecules with ions. The -neutral flag insures the overall net charge of the system equals zero. The -conc flag sets the ion concentration to that designated (in the case above 0.15 M). The default salt used by genion is always NaCl. If you desire to use different cation and anion, use -pname (for positive ion) and -nname (for negative ion) flags. 4
5 grompp -f em-sol-pme.mdp -c fws-b4em.gro -p fws.top -o em-sol.tpr mdrun -v -deffnm em-sol Two-step equilibration procedure - Now we need to relax the solvent and ions while keeping the protein atom positions restrained. This will be accomplished in two phases: 100 ps NVT followed by 100 ps NPT ensembles. We will perform our dynamics using a temperature of 300 K which is close to room temperature for most laboratory environments. Some folks like to simulate at 310 K which is closer to body or physiological temperature. We use a dispersion correction for energy and pressure (see section 4.8 of Gromacs manual). For coulombtype, PME stands for Particle Mesh Ewald electrostatics.[5, 6] PME is the best method for computing long-range electrostatics in systems where we have charges (exposed charged amino acid residues, ions, polar lipids, etc.). The all-bonds option under constraints applies the Linear Constraint algorithm (lincs) for fixing all bond lengths in the system (important to use this option when the simulation time step dt > ps).[7] nvt-pr-md.mdp ; VARIOUS PREPROCESSING OPTIONS define = -DPOSRES ; RUN CONTROL PARAMETERS integrator dt nsteps = md = ; time step (in ps) = ; number of steps ; OUTPUT CONTROL OPTIONS nstxout = 500 ; save coordinates every ps nstvout = 500 ; save velocities every ps nstenergy = 500 ; save energies every ps nstlog = 500 ; update log file every ps energygrps = Protein Non-Protein ; NEIGHBORSEARCHING PARAMETERS nstlist = 5 ns_type = grid pbc = xyz rlist = 1.0 ; OPTIONS FOR ELECTROSTATICS AND VDW coulombtype = PME ; Particle Mesh Ewald for long-range electrostatics pme_order = 4 ; cubic interpolation fourierspacing = 0.16 ; grid spacing for FFT rcoulomb = 1.0 vdw-type = Cut-off rvdw = 1.0 ; Temperature coupling 5
6 tcoupl = v-rescale ; Couple temperature to external heat bath according to velocity re-scale method tc-grps = Protein Non-Protein ; Use separate heat baths for Protein and Non-Protein groups tau_t = ; Coupling time constant, controlling strength of coupling ref_t = ; Temperature of heat bath ; Dispersion correction DispCorr = EnerPres ; account for vdw cut-off ; Pressure coupling is off pcoupl = no ; no pressure coupling in NVT ; GENERATE VELOCITIES FOR STARTUP RUN gen_vel = yes ; Assign velocities to particles by taking them randomly from a Maxwell distribution gen_temp = 300 ; Temperature to generate corresponding Maxwell distribution gen_seed = -1 ; Seed for (semi) random number generation. Different numbers give different sets of velocities ; OPTIONS FOR BONDS constraints = all-bonds ; All bonds will be treated as constraints (fixed length) continuation = no ; first dynamics run constraint_algorithm = lincs ; holonomic constraints lincs_iter = 1 ; accuracy of LINCS lincs_order = 4 ; also related to accuracy grompp -f nvt-pr-md.mdp -c em-sol.gro -p fws.top -o nvt-pr.tpr nohup mdrun -deffnm nvt-pr & Use tail -25 nvt-pr.log to check run progress. npt-pr-md.mdp ; VARIOUS PREPROCESSING OPTIONS define = -DPOSRES ; RUN CONTROL PARAMETERS integrator = md dt = nsteps = ; OUTPUT CONTROL OPTIONS nstxout = 500 ; save coordinates every ps nstvout = 500 ; save velocities every ps nstfout = 500 ; save forces every ps nstenergy = 500 ; save energies every ps nstlog = 500 ; update log file every ps energygrps = Protein Non-Protein ; NEIGHBORSEARCHING PARAMETERS 6
7 nstlist = 5 ns-type = Grid pbc = xyz rlist = 1.0 ; OPTIONS FOR ELECTROSTATICS AND VDW coulombtype = PME pme_order = 4 ; cubic interpolation fourierspacing = 0.16 ; grid spacing for FFT rcoulomb = 1.0 vdw-type = Cut-off rvdw = 1.0 ; Temperature coupling Tcoupl = v-rescale tc-grps = Protein Non-Protein tau_t = ref_t = ; Dispersion correction DispCorr = EnerPres ; account for vdw cut-off ; Pressure coupling Pcoupl = Parrinello-Rahman Pcoupltype = Isotropic tau_p = 2.0 compressibility = 4.5e-5 ref_p = 1.0 refcoord_scaling = com ; GENERATE VELOCITIES FOR STARTUP RUN gen_vel = no ; OPTIONS FOR BONDS constraints = all-bonds continuation = yes ; continuation from NVT constraint_algorithm = lincs ; holonomic constraints lincs_iter = 1 ; accuracy of LINCS lincs_order = 4 ; also related to accuracy grompp -f npt-pr-md.mdp -c em-sol.gro -p fws.top -o npt-pr.tpr nohup mdrun -deffnm npt-pr & npt-nopr-md.mdp ; RUN CONTROL PARAMETERS integrator = md dt = nsteps = ; 1 ns ; OUTPUT CONTROL OPTIONS nstxout = 500 ; save coordinates every ps nstvout = 500 ; save velocities every ps nstfout = 500 ; save forces every ps nstenergy = 500 ; save energies every ps nstlog = 500 ; update log file every ps 7
8 energygrps = Protein Non-Protein ; NEIGHBORSEARCHING PARAMETERS nstlist = 5 ns-type = Grid pbc = xyz rlist = 1.0 ; OPTIONS FOR ELECTROSTATICS AND VDW coulombtype = PME pme_order = 4 ; cubic interpolation fourierspacing = 0.16 ; grid spacing for FFT rcoulomb = 1.0 vdw-type = Cut-off rvdw = 1.0 ; Temperature coupling Tcoupl = v-rescale tc-grps = Protein Non-Protein tau_t = ref_t = ; Dispersion correction DispCorr = EnerPres ; account for vdw cut-off ; Pressure coupling Pcoupl = Parrinello-Rahman Pcoupltype = Isotropic tau_p = 2.0 compressibility = 4.5e-5 ref_p = 1.0 ; GENERATE VELOCITIES FOR STARTUP RUN gen_vel = no ; OPTIONS FOR BONDS constraints = all-bonds continuation = yes ; continuation from NPT PR constraint_algorithm = lincs ; holonomic constraints lincs_iter = 1 ; accuracy of LINCS lincs_order = 4 ; also related to accuracy grompp -f npt-nopr-md.mdp -c em-sol.gro -p fws.top -o npt-nopr.tpr nohup mdrun -deffnm npt-nopr & g_rms -s npt-nopr.tpr -f npt-nopr.trr -o fws-bkbone-rmsd.xvg Select 4 ( Backbone) 8
9 Average potential/kinetic/total energies and temperature. g_energy -f npt-nopr.edr -o nrg-npt.xvg Statistics over steps [ through ps ], 4 data sets All statistics are over 5001 points Energy Average Err.Est. RMSD Tot-Drift Potential (kj/mol) Kinetic En (kj/mol) Total Energy (kj/mol) Temperature (K) The radius of gyration provides a measure of the compactness of the protein. Use g_gyrate to evaluate. g_gyrate -s npt-nopr.tpr -f npt-nopr.trr -o fws-gyrate.xvg Select 1 Protein 9
10 do_dssp -s npt-nopr.tpr -f npt-nopr.trr -o fws-ss.xpm xpm2ps -f fws-ss.xpm -o fws-ss.eps Use the g_rmsf command to compute per residue temperature factors. g_rmsf -s npt-nopr.tpr -f npt-nopr-trr -o fws-rmsf.xvg -ox fws-avg.pdb -res -oq fwsbfac.pdb Select 1 Protein 10
11 pymol bfactors.pdb hide everything show cartoon spectrum b, selection=bfactors Q = XXX; cmd.alter( all, q = b > Q and Q or b ); spectrum q Where XXX is the highest bfactor value truncated. Blue is cool regions; Green intermediate and Red is hot (most mobile) To prepare a 300 dpi PNG ray-traced image in pymol use the following commands: ray 1200,1200 png bfac.png, dpi=300 Appendix: npt-longrun.mdp ; RUN CONTROL PARAMETERS integrator = md tinit = 0 ; Starting time dt = ; 2 femtosecond time step for integration nsteps = YYY ; Make it 50 ns 11
12 ; OUTPUT CONTROL OPTIONS nstxout 0.5 ns nstvout nstlog nstenergy nstxtcout energygrps = ; Writing full precision coordinates every = ; Writing velocities every 0.5 ns = 5000 ; Writing to the log file every 10ps = 5000 ; Writing out energy information every 10ps = 5000 ; Writing coordinates every 10ps = Protein Non-Protein ; NEIGHBORSEARCHING PARAMETERS nstlist = 5 ns-type = Grid pbc = xyz rlist = 1.0 ; OPTIONS FOR ELECTROSTATICS AND VDW coulombtype = PME pme_order = 4 ; cubic interpolation fourierspacing = 0.16 ; grid spacing for FFT rcoulomb = 1.0 vdw-type = Cut-off rvdw = 1.0 ; Dispersion correction DispCorr = EnerPres ; account for vdw cut-off ; Temperature coupling Tcoupl = v-rescale tc-grps = Protein Non-Protein tau_t = ref_t = ; Pressure coupling Pcoupl = Parrinello-Rahman Pcoupltype = Isotropic tau_p = 2.0 compressibility = 4.5e-5 ref_p = 1.0 ; GENERATE VELOCITIES FOR STARTUP RUN gen_vel = no ; OPTIONS FOR BONDS constraints = all-bonds constraint-algorithm = lincs continuation = yes ; Restarting after NPT without position restraints lincs-order = 4 lincs-iter = 1 lincs-warnangle = 30 Make a nice movie. First remove high frequency noise with g_filter. Perform in a separate directory named movie. g_filter s../md.tpr f../md.trr ol filtered.pdb fit nf 5 12
13 Select 1 (Protein) when prompted for group. Load into pymol and use commands for display (set orientation, etc) pymol filtered.pdb dss (set secondary structure) show cartoon viewport 640,800 set ray_trace_frames,1 mpng frame_.png Use an mpeg encoder to make the movie. Alternatively, you can use the Linux ImageMagik convert command to build an animated gif file. convert -delay 15 -loop 0 frame_*.png movie.gif Cluster analysis g_rms s../md.tpr f../md.trr f2../md.trr m rmsd-matrix.xpm g_cluster s../md.tpr f../md.trr dm rmsd-matrix.xpm dist rms-distribution.xvg o clusters.xpm sz cluster-sizes.xvg tr cluster-transitions.xpm ntr clustertransitions.xvg clid cluster-id-overtime.xvg cl clusters.pdb cutoff 0.25 method gromos pymol clusters.pdb split_states clusters delete clusters dss show cartoon How To Save specific time points from a trajectory as *.PDB files: To get a specific frame (3000 ps in this example) instead of the whole trajectory as a pdb file use additionally the -dump option, e.g. trjconv -f traj.xtc -s file.tpr -o time_3000ps.pdb -dump 3000 To re-center your molecule back in the box use... trjconv f traj.xtc o traj_center.xtc s str_b4md.gro pbc nojump -center Principal Components Analysis (PCA) 13
14 PCA methods help us determine what motions contribute most to the overall dynamics of the protein. In a system of N atoms, there exist 3N-6 modes of possible internal fluctuations (six degrees of freedom are required to describe the external rotation and translation of the system). For this analysis, we will focus on the protein backbone atoms. Perform in a separate directory. g_covar s../md.tpr f../md.xtc o eigenval.xvg v eigenvect.trr xpma covara.xpm Select group 4 (Protein backbone) both for fit and analysis. Trace of the covariance matrix: (nm^2) Diagonalizing... Sum of the eigenvalues: (nm^2) Writing eigenvalues to eigenval.xvg Writing reference, average structure & eigenvectors to eigenvec.trr Wrote the log to covar.log Use xpm2ps to make a pretty plot of the atomic covariance matrix. The do flag outputs a config file which can be used to modify properties of the plot (axis titles, legend, etc.). xpm2ps f covara.xpm o covara.eps do covara.m2p Use ghostview (or Photoshop) to view the plot (gv covara.eps). Use the xpm2ps - rainbow flag to look at other color schemes. Only a very small number of eigenvectors (modes of fluctuation) contribute significantly to the overall motion of the protein. This is typical. To view the most dominant mode (1), use the following command g_anaeig v eigenvec.trr s md.tpr f md.xtc first 1 last 1 nframes 100 extr fwsev1.pdb For a 2D eigenvector projection use g_anaeig s md.tpr f md.xtc first 1 last 2-2d proj-1-2.xvg Dihedral PCA We will make an index file to study 4 dihedral angles (2 phi and 2 psi) spanning residues MET29, ILE30, and GLY31. Use VMD (open pr.gro or md.gro) and pick the index numbers of the 4 atoms, which make up each dihedral angle. Name the file dangle.ndx. g_angle f../md.trr n dangle.ndx or dangle.trr type dihedral 14
15 Write covar.ndx by hand (2*N/3) atoms where N is the # of dihedral angles. You will notice that g_angle will help you here in its output. You will see a line in the g_angle output requesting 3 atom positions. There are 4 dihedrals. Will fill 3 atom positions with cos/sin [ test ] Make the dummy gro file for the g_covar analysis. trjconv s../md.tpr f dangle.trr o resiz.gro n covar.ndx e 0 Perform the g_covar analysis using g_covar f dangle.trr n covar.ndx ascii xpm nofit nomwa noref s resiz.gro To examine the projection of eigenvector 1 use g_anaeig -v eigenvec.trr -f dangle.trr -s resiz.gro -first 1 -last 1 -proj proj-1 Select Group 0 (System) when prompted. xmgrace proj-1.xvg For a 2D projection of eigenvector 1 with eigenvector 2 use g_anaeig -v eigenvec.trr -f dangle.trr -s resiz.gro -first 1 -last 2-2d 2dproj_1_2.xvg Use g_analyze to check the cosine content of the projection of ev1 with ev1. For example g_analyze f proj1.xvg cc proj1-cc.xvg Large-scale motions in the system cannot be differentiated from random diffusion when cosine content is close to 1.[8] Our cosine content for the dihedral angles in our study is ~ 0.11 for ev1, which is acceptable. Use the g_sham program to view the free energy landscape. g_sham f 2dproj_1_2.xvg ls gibbs.xpm -notime xpm2ps f gibbs.xpm o gibbs.eps rainbow red 15
16 make_ndx The make_ndx program is useful for generating groups (ID tags for specific atoms or residues that you may want to analyze). Gromacs generates default groups which may be adequate for your work as is. However, if you need to do more in depth analysis, use make_ndx to tag specific items in your model. See the manual for more information. How to use make_ndx to setup index (ndx) files. Use make_ndx to identify particular groups that you might want to freeze during a simulation or gather special energy information. Let s look at an example where we want to freeze the N and C terminal amino acids of a protein. Always use make_ndx to create an index group for use with the grompp program. In this sample case, we have a coordinate file of a collagen triple helix, post position restrained dynamics, where we want to freeze the N & C terminal for production run. We must first identify the residue # s in the coordinate file (In our case clg_b4md.pdb) that identify the N & C termini. The command line is simple enough make_ndx f clg_b4md.pdb o clg_ter.ndx You will see the following output (we left out the descriptive info at the beginning) followed by a command prompt (>). Reading structure file Going to read 0 old index file(s) Analysing residue names: Opening library file /usr/share/gromacs/top/aminoacids.dat There are: 2194 OTHER residues There are: 108 PROTEIN residues There are: 0 DNA residues Analysing Protein... Analysing Other... 0 System : 7365 atoms 1 Protein : 765 atoms 2 Protein-H : 687 atoms 3 C-alpha : 108 atoms 4 Backbone : 324 atoms 5 MainChain : 435 atoms 6 MainChain+Cb : 507 atoms 7 MainChain+H : 477 atoms 8 SideChain : 288 atoms 9 SideChain-H : 252 atoms 10 Prot-Masses : 765 atoms 11 Non-Protein : 6600 atoms 12 DRG : 21 atoms 13 SOL : 6579 atoms 14 Other : 6600 atoms nr : group! 'name' nr name 'splitch' nr Enter: list groups 'a': atom & 'del' nr 'splitres' nr 'l': list residues 't': atom type 'keep' nr 'splitat' nr 'h': help 'r': residue 'res' nr 'chain' char 16
17 "name": group 'case': case sensitive 'q': save and quit > Use the r command to enter the list of residue numbers that represent the N & C termini of the triple helix. > r r_1_36_37_72_73_108 : 51 atoms Note: You may also use a dash to specify a residue number range (e.g. to specify residues 1 to 36 use > r 1-36) OR, better yet, lets specify a residue range only including backbone atoms. Do this with the ampersand for example > r 1-36 & a C N CA The default name ( r_1_36_37_72_73_108 ) giving to the new index group that you have just created is cumbersome. Lets rename it using the name command. We will use the index group # (15) in the command. > name 15 Terminal > v Turned verbose on 0 System : 7365 atoms 1 Protein : 765 atoms 2 Protein-H : 687 atoms 3 C-alpha : 108 atoms 4 Backbone : 324 atoms 5 MainChain : 435 atoms 6 MainChain+Cb : 507 atoms 7 MainChain+H : 477 atoms 8 SideChain : 288 atoms 9 SideChain-H : 252 atoms 10 Prot-Masses : 765 atoms 11 Non-Protein : 6600 atoms 12 DRG : 21 atoms 13 SOL : 6579 atoms 14 Other : 6600 atoms 15 Terminal : 51 atoms nr : group! 'name' nr name 'splitch' nr Enter: list groups 'a': atom & 'del' nr 'splitres' nr 'l': list residues 't': atom type 'keep' nr 'splitat' nr 'h': help 'r': residue 'res' nr 'chain' char "name": group 'case': case sensitive 'q': save and quit > We used the v command to verify that our name change was successful. To finish up, use the q command to save and quit and you re done. Now how do we freeze the groups? Easy, add the following lines to your md.mdp file energygrp_excl = Terminal Terminal Terminal SOL! To remove computation of nonbonding interactions between the frozen groups with each other and surroundings (i.e. the solvent, SOL) 17
18 freezegrps = Terminal! Index group to freeze freezedim = Y Y Y! Freeze this group in all directions, x, y, and z Remember, you must include the new index file when you use this md.mdp file to create your run input file (tpr) using grompp. Use the n flag to grompp. For example grompp f md.mdp c pr.gro p clg.top n clg_ter.ndx o md.tpr How to restart a crashed run. The mdrun program now uses a very handy checkpointing feature. Restarting crashed runs is easy with mdrun. mdrun -s prev.tpr -f prev.trr -e prev.edr -o prev.trr g prev.log cpi -append The cpi flag tells mdrun to read the checkpoint file (state.cpt) as input. The append flag tells mdrun to append the data to the existing trajectory and log files. See the mdrun manpage for more information (or use mdrun h command for more info/help). How to continue your runs. Use grompp or use tpbconv. For the grompp case, you will need to change the gen_vel value in the mdp file to no ; set unconstrained_start to yes and read in the previous trajectory from previous trr or xtc file and the previous velocities from the edr file. For a more exact continuation using grompp, you will also need to set mdp variables like tinit and init_step to an appropriate value for continuation. Use grompp h command for more information/help. This method is binary nonidentical (introduces small negligible errors). Method 1 grompp -f md_restart.mdp -c md_prev.gro t md_prev.trr -e md_prev.edr p topol.top - o md_restart.tpr maxwarn 10 Method 2 tpbconv -s previous.tpr -f previous.trr -e previous.edr -extend timetoextendby -o next.tpr mdrun -deffnm next Extending runs using tpbconv and mdrun checkpoint files in version 4.0 Extending runs is even easier in version 4.0 and is binary identical because you are using the checkpoint file. tpbconv -s previous.tpr -extend timetoextendby -o next.tpr mdrun -s next.tpr -cpi previous.cpt Note: you can use the append flag for mdrun to add the new output to your existing files. Use the final filename.cpt file and NOT the filename_prev.cpt file!!! 18
19 How to concatenate trajectories from continued runs. Use trjcat. trjcat f md1.xtc md2.xtc md3.xtc (etc) o mdall.xtc -settime Trajectories are read and concatenated in the order you provide. Use the settime flag to interactively set the starting time for each trajectory (Note: the default time unit is ps). How to concatenate energy files (edr files) from continued runs. Use eneconv. eneconv f md1.edr md2.edr md3.edr (etc) o mdall.edr -settime Similar to trjcat in operation. 19
20 Bibliography 1.! Hess, B., et al., GROMACS 4: Algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theor. Comp., (3): p ! Lindahl, E., B. Hess, and D. van der Spoel, GROMACS 3.0: a package for molecular simulation and trajectory analysis. J. Mol. Model, : p ! Lindorff-Larsen, K., et al., Improved side-chain torsion potentials for the Amber ff99sb protein force field. Proteins, : p ! Jorgensen, W., et al., Comparison of Simple Potential Functions for Simulating Liquid Water. J. Chem. Phys., : p ! Darden, T., D. York, and L. Pedersen, Particle Mesh Ewald: An N-log(N) method for Ewald sums in large systems. J. Chem. Phys., : p ! Essmann, U., et al., A smooth particle mesh ewald potential. J. Chem. Phys., : p ! Hess, B., et al., LINCS: A Linear Constraint Solver for molecular simulations. J. Comp. Chem., : p ! Maisuradze, G.G. and D.M. Leitner, Free energy landscape of a biomolecule in dihedral principal component space: sampling convergence and correspondence between structures and minima. Proteins, (3): p
GROMACS Introductory Tutorial
 GROMACS Introductory Tutorial Gromacs ver 3.3.1 Author: John E. Kerrigan, Ph.D. AST/IST, University of Medicine and Dentistry of NJ 675 Hoes Lane Piscataway, NJ 08554 Phone: (732) 235-4473 Fax: (732) 235-5252
GROMACS Introductory Tutorial Gromacs ver 3.3.1 Author: John E. Kerrigan, Ph.D. AST/IST, University of Medicine and Dentistry of NJ 675 Hoes Lane Piscataway, NJ 08554 Phone: (732) 235-4473 Fax: (732) 235-5252
Tutorial for Trypsin- Benzamidine complex molecular dynamics study.
 Tutorial for Trypsin- Benzamidine complex molecular dynamics study. Gromacs version 4.5.5 John E. Kerrigan, Ph.D. Associate Director, Bioinformatics The Cancer Institute of New Jersey The Robert Wood Johnson
Tutorial for Trypsin- Benzamidine complex molecular dynamics study. Gromacs version 4.5.5 John E. Kerrigan, Ph.D. Associate Director, Bioinformatics The Cancer Institute of New Jersey The Robert Wood Johnson
Structure Tools and Visualization
 Structure Tools and Visualization Gary Van Domselaar University of Alberta gary.vandomselaar@ualberta.ca Slides Adapted from Michel Dumontier, Blueprint Initiative 1 Visualization & Communication Visualization
Structure Tools and Visualization Gary Van Domselaar University of Alberta gary.vandomselaar@ualberta.ca Slides Adapted from Michel Dumontier, Blueprint Initiative 1 Visualization & Communication Visualization
Molecular Dynamics Simulations
 Molecular Dynamics Simulations Yaoquan Tu Division of Theoretical Chemistry and Biology, Royal Institute of Technology (KTH) 2011-06 1 Outline I. Introduction II. Molecular Mechanics Force Field III. Molecular
Molecular Dynamics Simulations Yaoquan Tu Division of Theoretical Chemistry and Biology, Royal Institute of Technology (KTH) 2011-06 1 Outline I. Introduction II. Molecular Mechanics Force Field III. Molecular
Visualizing molecular simulations
 Visualizing molecular simulations ChE210D Overview Visualization plays a very important role in molecular simulations: it enables us to develop physical intuition about the behavior of a system that is
Visualizing molecular simulations ChE210D Overview Visualization plays a very important role in molecular simulations: it enables us to develop physical intuition about the behavior of a system that is
Hands-on exercises on solvent models & electrostatics EMBnet - Molecular Modeling Course 2005
 Hands-on exercises on solvent models & electrostatics EMBnet - Molecular Modeling Course 2005 Exercise 1. The purpose of this exercise is to color the solvent accessible surface of a protein according
Hands-on exercises on solvent models & electrostatics EMBnet - Molecular Modeling Course 2005 Exercise 1. The purpose of this exercise is to color the solvent accessible surface of a protein according
Hydrogen Bonds The electrostatic nature of hydrogen bonds
 Hydrogen Bonds Hydrogen bonds have played an incredibly important role in the history of structural biology. Both the structure of DNA and of protein a-helices and b-sheets were predicted based largely
Hydrogen Bonds Hydrogen bonds have played an incredibly important role in the history of structural biology. Both the structure of DNA and of protein a-helices and b-sheets were predicted based largely
BIOC351: Proteins. PyMOL Laboratory #1. Installing and Using
 BIOC351: Proteins PyMOL Laboratory #1 Installing and Using Information and figures for this handout was obtained from the following sources: Introduction to PyMOL (2009) DeLano Scientific LLC. Installing
BIOC351: Proteins PyMOL Laboratory #1 Installing and Using Information and figures for this handout was obtained from the following sources: Introduction to PyMOL (2009) DeLano Scientific LLC. Installing
Structure Check. Authors: Eduard Schreiner Leonardo G. Trabuco. February 7, 2012
 University of Illinois at Urbana-Champaign NIH Resource for Macromolecular Modeling and Bioinformatics Beckman Institute Computational Biophysics Workshop Structure Check Authors: Eduard Schreiner Leonardo
University of Illinois at Urbana-Champaign NIH Resource for Macromolecular Modeling and Bioinformatics Beckman Institute Computational Biophysics Workshop Structure Check Authors: Eduard Schreiner Leonardo
Peptide Bonds: Structure
 Peptide Bonds: Structure Peptide primary structure The amino acid sequence, from - to C-terminus, determines the primary structure of a peptide or protein. The amino acids are linked through amide or peptide
Peptide Bonds: Structure Peptide primary structure The amino acid sequence, from - to C-terminus, determines the primary structure of a peptide or protein. The amino acids are linked through amide or peptide
Overview of NAMD and Molecular Dynamics
 Overview of NAMD and Molecular Dynamics Jim Phillips Low-cost Linux Clusters for Biomolecular Simulations Using NAMD Outline Overview of molecular dynamics Overview of NAMD NAMD parallel design NAMD config
Overview of NAMD and Molecular Dynamics Jim Phillips Low-cost Linux Clusters for Biomolecular Simulations Using NAMD Outline Overview of molecular dynamics Overview of NAMD NAMD parallel design NAMD config
file:///c /Documents%20and%20Settings/terry/Desktop/DOCK%20website/terry/Old%20Versions/dock4.0_faq.txt
 -- X. Zou, 6/28/1999 -- Questions on installation of DOCK4.0.1: ======================================= Q. Can I run DOCK on platforms other than SGI (e.g., SparcStations, DEC Stations, Pentium, etc.)?
-- X. Zou, 6/28/1999 -- Questions on installation of DOCK4.0.1: ======================================= Q. Can I run DOCK on platforms other than SGI (e.g., SparcStations, DEC Stations, Pentium, etc.)?
Amino Acids. Amino acids are the building blocks of proteins. All AA s have the same basic structure: Side Chain. Alpha Carbon. Carboxyl. Group.
 Protein Structure Amino Acids Amino acids are the building blocks of proteins. All AA s have the same basic structure: Side Chain Alpha Carbon Amino Group Carboxyl Group Amino Acid Properties There are
Protein Structure Amino Acids Amino acids are the building blocks of proteins. All AA s have the same basic structure: Side Chain Alpha Carbon Amino Group Carboxyl Group Amino Acid Properties There are
CASSANDRA. Computational Atomistic Simulation Software At Notre Dame for Research Advances
 1 CASSANDRA Computational Atomistic Simulation Software At Notre Dame for Research Advances User Manual 1.0 Written by: Edward J. Maginn, Jindal K. Shah, Eliseo MarinRimoldi, Sandip Khan, Neeraj Rai, Thomas
1 CASSANDRA Computational Atomistic Simulation Software At Notre Dame for Research Advances User Manual 1.0 Written by: Edward J. Maginn, Jindal K. Shah, Eliseo MarinRimoldi, Sandip Khan, Neeraj Rai, Thomas
This class deals with the fundamental structural features of proteins, which one can understand from the structure of amino acids, and how they are
 This class deals with the fundamental structural features of proteins, which one can understand from the structure of amino acids, and how they are put together. 1 A more detailed view of a single protein
This class deals with the fundamental structural features of proteins, which one can understand from the structure of amino acids, and how they are put together. 1 A more detailed view of a single protein
Tutorial for Assignment #2 Gantry Crane Analysis By ANSYS (Mechanical APDL) V.13.0
 Tutorial for Assignment #2 Gantry Crane Analysis By ANSYS (Mechanical APDL) V.13.0 1 Problem Description Design a gantry crane meeting the geometry presented in Figure 1 on page #325 of the course textbook
Tutorial for Assignment #2 Gantry Crane Analysis By ANSYS (Mechanical APDL) V.13.0 1 Problem Description Design a gantry crane meeting the geometry presented in Figure 1 on page #325 of the course textbook
Molecular Visualization. Introduction
 Molecular Visualization Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences Duquesne University Introduction Assessments of change, dynamics, and cause and effect
Molecular Visualization Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences Duquesne University Introduction Assessments of change, dynamics, and cause and effect
1 The water molecule and hydrogen bonds in water
 The Physics and Chemistry of Water 1 The water molecule and hydrogen bonds in water Stoichiometric composition H 2 O the average lifetime of a molecule is 1 ms due to proton exchange (catalysed by acids
The Physics and Chemistry of Water 1 The water molecule and hydrogen bonds in water Stoichiometric composition H 2 O the average lifetime of a molecule is 1 ms due to proton exchange (catalysed by acids
The Structure of a Network and Its Functions
 Elastic properties and heterogeneous stiffness of the Phi29 motor connector channel Rajendra Kumar and Helmut Grubmüller Max Planck Institute for Biophysical Chemistry, Department of Theoretical and Computational
Elastic properties and heterogeneous stiffness of the Phi29 motor connector channel Rajendra Kumar and Helmut Grubmüller Max Planck Institute for Biophysical Chemistry, Department of Theoretical and Computational
PROTEINS THE PEPTIDE BOND. The peptide bond, shown above enclosed in the blue curves, generates the basic structural unit for proteins.
 Ca 2+ The contents of this module were developed under grant award # P116B-001338 from the Fund for the Improvement of Postsecondary Education (FIPSE), United States Department of Education. However, those
Ca 2+ The contents of this module were developed under grant award # P116B-001338 from the Fund for the Improvement of Postsecondary Education (FIPSE), United States Department of Education. However, those
Introduction to Molecular Dynamics Simulations
 Introduction to Molecular Dynamics Simulations Roland H. Stote Institut de Chimie LC3-UMR 7177 Université Louis Pasteur Strasbourg France 1EA5 Title Native Acetylcholinesterase (E.C. 3.1.1.7) From Torpedo
Introduction to Molecular Dynamics Simulations Roland H. Stote Institut de Chimie LC3-UMR 7177 Université Louis Pasteur Strasbourg France 1EA5 Title Native Acetylcholinesterase (E.C. 3.1.1.7) From Torpedo
Why? Intermolecular Forces. Intermolecular Forces. Chapter 12 IM Forces and Liquids. Covalent Bonding Forces for Comparison of Magnitude
 1 Why? Chapter 1 Intermolecular Forces and Liquids Why is water usually a liquid and not a gas? Why does liquid water boil at such a high temperature for such a small molecule? Why does ice float on water?
1 Why? Chapter 1 Intermolecular Forces and Liquids Why is water usually a liquid and not a gas? Why does liquid water boil at such a high temperature for such a small molecule? Why does ice float on water?
Gold (Genetic Optimization for Ligand Docking) G. Jones et al. 1996
 Gold (Genetic Optimization for Ligand Docking) G. Jones et al. 1996 LMU Institut für Informatik, LFE Bioinformatik, Cheminformatics, Structure based methods J. Apostolakis 1 Genetic algorithms Inspired
Gold (Genetic Optimization for Ligand Docking) G. Jones et al. 1996 LMU Institut für Informatik, LFE Bioinformatik, Cheminformatics, Structure based methods J. Apostolakis 1 Genetic algorithms Inspired
Towards Large-Scale Molecular Dynamics Simulations on Graphics Processors
 Towards Large-Scale Molecular Dynamics Simulations on Graphics Processors Joe Davis, Sandeep Patel, and Michela Taufer University of Delaware Outline Introduction Introduction to GPU programming Why MD
Towards Large-Scale Molecular Dynamics Simulations on Graphics Processors Joe Davis, Sandeep Patel, and Michela Taufer University of Delaware Outline Introduction Introduction to GPU programming Why MD
List the 3 main types of subatomic particles and indicate the mass and electrical charge of each.
 Basic Chemistry Why do we study chemistry in a biology course? All living organisms are composed of chemicals. To understand life, we must understand the structure, function, and properties of the chemicals
Basic Chemistry Why do we study chemistry in a biology course? All living organisms are composed of chemicals. To understand life, we must understand the structure, function, and properties of the chemicals
Data Visualization for Atomistic/Molecular Simulations. Douglas E. Spearot University of Arkansas
 Data Visualization for Atomistic/Molecular Simulations Douglas E. Spearot University of Arkansas What is Atomistic Simulation? Molecular dynamics (MD) involves the explicit simulation of atomic scale particles
Data Visualization for Atomistic/Molecular Simulations Douglas E. Spearot University of Arkansas What is Atomistic Simulation? Molecular dynamics (MD) involves the explicit simulation of atomic scale particles
3D structure visualization and high quality imaging. Chimera
 3D structure visualization and high quality imaging. Chimera Vincent Zoete 2008 Contact : vincent.zoete@isb sib.ch 1/27 Table of Contents Presentation of Chimera...3 Exercise 1...4 Loading a structure
3D structure visualization and high quality imaging. Chimera Vincent Zoete 2008 Contact : vincent.zoete@isb sib.ch 1/27 Table of Contents Presentation of Chimera...3 Exercise 1...4 Loading a structure
LAMMPS Developer Guide 23 Aug 2011
 LAMMPS Developer Guide 23 Aug 2011 This document is a developer guide to the LAMMPS molecular dynamics package, whose WWW site is at lammps.sandia.gov. It describes the internal structure and algorithms
LAMMPS Developer Guide 23 Aug 2011 This document is a developer guide to the LAMMPS molecular dynamics package, whose WWW site is at lammps.sandia.gov. It describes the internal structure and algorithms
MestRe-C User Guide Megan Bragg 04/14/05
 MestRe-C User Guide Megan Bragg 04/14/05 Some general useful features: 1. You can add Text, Titles, Temperatures, or peak labels (in any font/size/color) to the spectrum. 2. MestRe-C allows you to save
MestRe-C User Guide Megan Bragg 04/14/05 Some general useful features: 1. You can add Text, Titles, Temperatures, or peak labels (in any font/size/color) to the spectrum. 2. MestRe-C allows you to save
1. Free energy with controlled uncertainty 2. The modes of ligand binding to DNA
 1. Free energy with controlled uncertainty 2. The modes of ligand binding to DNA Tomáš Kubař Institute of Organic Chemistry and Biochemistry Praha, Czech Republic Thermodynamic Integration Alchemical change
1. Free energy with controlled uncertainty 2. The modes of ligand binding to DNA Tomáš Kubař Institute of Organic Chemistry and Biochemistry Praha, Czech Republic Thermodynamic Integration Alchemical change
VMD - High Resolution Graphics
 Biochem 660 2008 131 VMD - High Resolution Graphics VMD (http://www.ks.uiuc.edu/research/vmd/) is a free program developed at the Theoretical and Computational Biophysics Group at the University of Illinois
Biochem 660 2008 131 VMD - High Resolution Graphics VMD (http://www.ks.uiuc.edu/research/vmd/) is a free program developed at the Theoretical and Computational Biophysics Group at the University of Illinois
Infrared Spectroscopy: Theory
 u Chapter 15 Infrared Spectroscopy: Theory An important tool of the organic chemist is Infrared Spectroscopy, or IR. IR spectra are acquired on a special instrument, called an IR spectrometer. IR is used
u Chapter 15 Infrared Spectroscopy: Theory An important tool of the organic chemist is Infrared Spectroscopy, or IR. IR spectra are acquired on a special instrument, called an IR spectrometer. IR is used
JMS: A workflow management system and web-based cluster front-end for the Torque resource manager
 JMS: A workflow management system and web-based cluster front-end for the Torque resource manager David K. Brown, Thommas M. Musyoka, David L. Penkler and Özlem Tastan Bishop* Research Unit in Bioinformatics
JMS: A workflow management system and web-based cluster front-end for the Torque resource manager David K. Brown, Thommas M. Musyoka, David L. Penkler and Özlem Tastan Bishop* Research Unit in Bioinformatics
Thermodynamics. Thermodynamics 1
 Thermodynamics 1 Thermodynamics Some Important Topics First Law of Thermodynamics Internal Energy U ( or E) Enthalpy H Second Law of Thermodynamics Entropy S Third law of Thermodynamics Absolute Entropy
Thermodynamics 1 Thermodynamics Some Important Topics First Law of Thermodynamics Internal Energy U ( or E) Enthalpy H Second Law of Thermodynamics Entropy S Third law of Thermodynamics Absolute Entropy
Learning Module 4 - Thermal Fluid Analysis Note: LM4 is still in progress. This version contains only 3 tutorials.
 Learning Module 4 - Thermal Fluid Analysis Note: LM4 is still in progress. This version contains only 3 tutorials. Attachment C1. SolidWorks-Specific FEM Tutorial 1... 2 Attachment C2. SolidWorks-Specific
Learning Module 4 - Thermal Fluid Analysis Note: LM4 is still in progress. This version contains only 3 tutorials. Attachment C1. SolidWorks-Specific FEM Tutorial 1... 2 Attachment C2. SolidWorks-Specific
Steffen Lindert, René Staritzbichler, Nils Wötzel, Mert Karakaş, Phoebe L. Stewart, and Jens Meiler
 Structure 17 Supplemental Data EM-Fold: De Novo Folding of α-helical Proteins Guided by Intermediate-Resolution Electron Microscopy Density Maps Steffen Lindert, René Staritzbichler, Nils Wötzel, Mert
Structure 17 Supplemental Data EM-Fold: De Novo Folding of α-helical Proteins Guided by Intermediate-Resolution Electron Microscopy Density Maps Steffen Lindert, René Staritzbichler, Nils Wötzel, Mert
HOW TO MAKE YOUR WEBSITE
 HOW TO MAKE YOUR WEBSITE Use Netscape Composer to make your web page presentation of a 3D structure of your choosing. You will need to download a few template web pages from the biochemistry website, and
HOW TO MAKE YOUR WEBSITE Use Netscape Composer to make your web page presentation of a 3D structure of your choosing. You will need to download a few template web pages from the biochemistry website, and
Topic 2: Energy in Biological Systems
 Topic 2: Energy in Biological Systems Outline: Types of energy inside cells Heat & Free Energy Energy and Equilibrium An Introduction to Entropy Types of energy in cells and the cost to build the parts
Topic 2: Energy in Biological Systems Outline: Types of energy inside cells Heat & Free Energy Energy and Equilibrium An Introduction to Entropy Types of energy in cells and the cost to build the parts
2007 7.013 Problem Set 1 KEY
 2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
Nonlinear Iterative Partial Least Squares Method
 Numerical Methods for Determining Principal Component Analysis Abstract Factors Béchu, S., Richard-Plouet, M., Fernandez, V., Walton, J., and Fairley, N. (2016) Developments in numerical treatments for
Numerical Methods for Determining Principal Component Analysis Abstract Factors Béchu, S., Richard-Plouet, M., Fernandez, V., Walton, J., and Fairley, N. (2016) Developments in numerical treatments for
5. Tutorial. Starting FlashCut CNC
 FlashCut CNC Section 5 Tutorial 259 5. Tutorial Starting FlashCut CNC To start FlashCut CNC, click on the Start button, select Programs, select FlashCut CNC 4, then select the FlashCut CNC 4 icon. A dialog
FlashCut CNC Section 5 Tutorial 259 5. Tutorial Starting FlashCut CNC To start FlashCut CNC, click on the Start button, select Programs, select FlashCut CNC 4, then select the FlashCut CNC 4 icon. A dialog
Piston Ring. Problem:
 Problem: A cast-iron piston ring has a mean diameter of 81 mm, a radial height of h 6 mm, and a thickness b 4 mm. The ring is assembled using an expansion tool which separates the split ends a distance
Problem: A cast-iron piston ring has a mean diameter of 81 mm, a radial height of h 6 mm, and a thickness b 4 mm. The ring is assembled using an expansion tool which separates the split ends a distance
Building Platform as a Service for Scientific Applications
 Building Platform as a Service for Scientific Applications Moustafa AbdelBaky moustafa@cac.rutgers.edu Rutgers Discovery Informa=cs Ins=tute (RDI 2 ) The NSF Cloud and Autonomic Compu=ng Center Department
Building Platform as a Service for Scientific Applications Moustafa AbdelBaky moustafa@cac.rutgers.edu Rutgers Discovery Informa=cs Ins=tute (RDI 2 ) The NSF Cloud and Autonomic Compu=ng Center Department
The UBATSIM software package. Simulates UBAT detector frames from a GRB. and. processes them to trigger and locate the GRB
 The UBATSIM software package Simulates UBAT detector frames from a GRB and processes them to trigger and locate the GRB P.H.Connell Image Processing Laboratory University of Valencia Valencia Spain 3.2.2011
The UBATSIM software package Simulates UBAT detector frames from a GRB and processes them to trigger and locate the GRB P.H.Connell Image Processing Laboratory University of Valencia Valencia Spain 3.2.2011
Section I Using Jmol as a Computer Visualization Tool
 Section I Using Jmol as a Computer Visualization Tool Jmol is a free open source molecular visualization program used by students, teachers, professors, and scientists to explore protein structures. Section
Section I Using Jmol as a Computer Visualization Tool Jmol is a free open source molecular visualization program used by students, teachers, professors, and scientists to explore protein structures. Section
GROMACS 4: Algorithms for Highly Efficient, Load-Balanced, and Scalable Molecular Simulation
 J. Chem. Theory Comput. 2008, 4, 435-447 435 GROMACS 4: Algorithms for Highly Efficient, Load-Balanced, and Scalable Molecular Simulation Berk Hess* Max-Planck Institute for Polymer Research, Ackermannweg
J. Chem. Theory Comput. 2008, 4, 435-447 435 GROMACS 4: Algorithms for Highly Efficient, Load-Balanced, and Scalable Molecular Simulation Berk Hess* Max-Planck Institute for Polymer Research, Ackermannweg
The Lighting Effects Filter
 Appendix appendix E The Lighting Effects Filter The Lighting Effects filter is like a little program in itself. With this filter, you can create a wealth of different lighting effects, from making a particular
Appendix appendix E The Lighting Effects Filter The Lighting Effects filter is like a little program in itself. With this filter, you can create a wealth of different lighting effects, from making a particular
Active Vibration Isolation of an Unbalanced Machine Spindle
 UCRL-CONF-206108 Active Vibration Isolation of an Unbalanced Machine Spindle D. J. Hopkins, P. Geraghty August 18, 2004 American Society of Precision Engineering Annual Conference Orlando, FL, United States
UCRL-CONF-206108 Active Vibration Isolation of an Unbalanced Machine Spindle D. J. Hopkins, P. Geraghty August 18, 2004 American Society of Precision Engineering Annual Conference Orlando, FL, United States
animation animation shape specification as a function of time
 animation animation shape specification as a function of time animation representation many ways to represent changes with time intent artistic motion physically-plausible motion efficiency control typically
animation animation shape specification as a function of time animation representation many ways to represent changes with time intent artistic motion physically-plausible motion efficiency control typically
Chapter 13 - LIQUIDS AND SOLIDS
 Chapter 13 - LIQUIDS AND SOLIDS Problems to try at end of chapter: Answers in Appendix I: 1,3,5,7b,9b,15,17,23,25,29,31,33,45,49,51,53,61 13.1 Properties of Liquids 1. Liquids take the shape of their container,
Chapter 13 - LIQUIDS AND SOLIDS Problems to try at end of chapter: Answers in Appendix I: 1,3,5,7b,9b,15,17,23,25,29,31,33,45,49,51,53,61 13.1 Properties of Liquids 1. Liquids take the shape of their container,
The strength of the interaction
 The strength of the interaction Host Guest Supramolecule (host-guest complex) When is the host capable to recognize the guest? How do we define selectivity Which element will we use to design the host
The strength of the interaction Host Guest Supramolecule (host-guest complex) When is the host capable to recognize the guest? How do we define selectivity Which element will we use to design the host
Introduction to Principal Components and FactorAnalysis
 Introduction to Principal Components and FactorAnalysis Multivariate Analysis often starts out with data involving a substantial number of correlated variables. Principal Component Analysis (PCA) is a
Introduction to Principal Components and FactorAnalysis Multivariate Analysis often starts out with data involving a substantial number of correlated variables. Principal Component Analysis (PCA) is a
Molecular Modelling of DNA. Charlie Laughton University of Nottingham
 Molecular Modelling of DNA Charlie Laughton University of Nottingham Why model DNA? Sequences/structures you can t crystallise or get by NMR Drug design Understanding protein-dna recognition Quantitating
Molecular Modelling of DNA Charlie Laughton University of Nottingham Why model DNA? Sequences/structures you can t crystallise or get by NMR Drug design Understanding protein-dna recognition Quantitating
Scientific Programming with Python. Randy M. Wadkins, Ph.D. Asst. Prof. of Chemistry & Biochem.
 Scientific Programming with Python Randy M. Wadkins, Ph.D. Asst. Prof. of Chemistry & Biochem. What is Python? Python for computers is: a scripting language a programming language interpreted, so it
Scientific Programming with Python Randy M. Wadkins, Ph.D. Asst. Prof. of Chemistry & Biochem. What is Python? Python for computers is: a scripting language a programming language interpreted, so it
. Tutorial #3 Building Complex Targets
 . Tutorial #3 Building Complex Targets. Mixed Gas/Solid Targets Gas Ionization Chamber Previous Tutorials have covered how to setup TRIM, determine which ion and energy to specify for a semiconductor n-well
. Tutorial #3 Building Complex Targets. Mixed Gas/Solid Targets Gas Ionization Chamber Previous Tutorials have covered how to setup TRIM, determine which ion and energy to specify for a semiconductor n-well
Lecture #7 (2D NMR) Utility of Resonance Assignments
 Lecture #7 (2D NMR) Basics of multidimensional NMR (2D NMR) 2D NOESY, COSY and TOCSY 2/23/15 Utility of Resonance Assignments Resonance Assignments: Assignment of frequency positions of resonances (peaks)
Lecture #7 (2D NMR) Basics of multidimensional NMR (2D NMR) 2D NOESY, COSY and TOCSY 2/23/15 Utility of Resonance Assignments Resonance Assignments: Assignment of frequency positions of resonances (peaks)
CHM 579 Lab 1: Basic Monte Carlo Algorithm
 CHM 579 Lab 1: Basic Monte Carlo Algorithm Due 02/12/2014 The goal of this lab is to get familiar with a simple Monte Carlo program and to be able to compile and run it on a Linux server. Lab Procedure:
CHM 579 Lab 1: Basic Monte Carlo Algorithm Due 02/12/2014 The goal of this lab is to get familiar with a simple Monte Carlo program and to be able to compile and run it on a Linux server. Lab Procedure:
Study the following diagrams of the States of Matter. Label the names of the Changes of State between the different states.
 Describe the strength of attractive forces between particles. Describe the amount of space between particles. Can the particles in this state be compressed? Do the particles in this state have a definite
Describe the strength of attractive forces between particles. Describe the amount of space between particles. Can the particles in this state be compressed? Do the particles in this state have a definite
PyRy3D: a software tool for modeling of large macromolecular complexes MODELING OF STRUCTURES FOR LARGE MACROMOLECULAR COMPLEXES
 MODELING OF STRUCTURES FOR LARGE MACROMOLECULAR COMPLEXES PyRy3D is a method for building low-resolution models of large macromolecular complexes. The components (proteins, nucleic acids and any other
MODELING OF STRUCTURES FOR LARGE MACROMOLECULAR COMPLEXES PyRy3D is a method for building low-resolution models of large macromolecular complexes. The components (proteins, nucleic acids and any other
Refinement of a pdb-structure and Convert
 Refinement of a pdb-structure and Convert A. Search for a pdb with the closest sequence to your protein of interest. B. Choose the most suitable entry (or several entries). C. Convert and resolve errors
Refinement of a pdb-structure and Convert A. Search for a pdb with the closest sequence to your protein of interest. B. Choose the most suitable entry (or several entries). C. Convert and resolve errors
Hewlett-Packard Development Company, L.P., 8000 Foothills Boulevard, Roseville, California 95747. www.hp.com/go/networking/
 5998-3611 v5.1.3 HP RF Planner v5.1.3 Release Notes HP RF Hewlett-Packard Development Company, L.P., 8000 Foothills Boulevard, Roseville, California 95747 Planner Release Notes V5.1.2 www.hp.com/go/networking/
5998-3611 v5.1.3 HP RF Planner v5.1.3 Release Notes HP RF Hewlett-Packard Development Company, L.P., 8000 Foothills Boulevard, Roseville, California 95747 Planner Release Notes V5.1.2 www.hp.com/go/networking/
Linear Static Analysis of a Cantilever Beam Using Beam Library (SI Units)
 APPENDIX A Linear Static Analysis of a Cantilever Beam Using Beam Library (SI Units) Objectives: Create a geometric representation of a cantilever beam. Use the geometry model to define an MSC.Nastran
APPENDIX A Linear Static Analysis of a Cantilever Beam Using Beam Library (SI Units) Objectives: Create a geometric representation of a cantilever beam. Use the geometry model to define an MSC.Nastran
Trace Layer Import for Printed Circuit Boards Under Icepak
 Tutorial 13. Trace Layer Import for Printed Circuit Boards Under Icepak Introduction: A printed circuit board (PCB) is generally a multi-layered board made of dielectric material and several layers of
Tutorial 13. Trace Layer Import for Printed Circuit Boards Under Icepak Introduction: A printed circuit board (PCB) is generally a multi-layered board made of dielectric material and several layers of
Paper: 6 Chemistry 2.130 University I Chemistry: Models Page: 2 of 7. 4. Which of the following weak acids would make the best buffer at ph = 5.0?
 Paper: 6 Chemistry 2.130 University I Chemistry: Models Page: 2 of 7 4. Which of the following weak acids would make the best buffer at ph = 5.0? A) Acetic acid (Ka = 1.74 x 10-5 ) B) H 2 PO - 4 (Ka =
Paper: 6 Chemistry 2.130 University I Chemistry: Models Page: 2 of 7 4. Which of the following weak acids would make the best buffer at ph = 5.0? A) Acetic acid (Ka = 1.74 x 10-5 ) B) H 2 PO - 4 (Ka =
The peptide bond Peptides and proteins are linear polymers of amino acids. The amino acids are
 Introduction to Protein Structure Proteins are large heteropolymers usually comprised of 50 2500 monomer units, although larger proteins are observed 7. The monomer units of proteins are amino acids. The
Introduction to Protein Structure Proteins are large heteropolymers usually comprised of 50 2500 monomer units, although larger proteins are observed 7. The monomer units of proteins are amino acids. The
[Image removed due to copyright concerns]
![[Image removed due to copyright concerns] [Image removed due to copyright concerns]](/thumbs/40/21531271.jpg) Radiation Chemistry Ionizing radiation produces abundant secondary electrons that rapidly slow down (thermalize) to energies below 7.4 ev, the threshold to produce electronic transitions in liquid water.
Radiation Chemistry Ionizing radiation produces abundant secondary electrons that rapidly slow down (thermalize) to energies below 7.4 ev, the threshold to produce electronic transitions in liquid water.
Optimization stopped. -- Wrong number of Negative eigenvalues: Desired= 1 Actual= 0 -- Flag reset to prevent archiving.
 5. Exploring the potential energy surface So far we have focused on the equilibrium structures - minima on PES. But other stationary points - in particular the transition states - are equally interesting
5. Exploring the potential energy surface So far we have focused on the equilibrium structures - minima on PES. But other stationary points - in particular the transition states - are equally interesting
Chapter 2. Atomic Structure and Interatomic Bonding
 Chapter 2. Atomic Structure and Interatomic Bonding Interatomic Bonding Bonding forces and energies Primary interatomic bonds Secondary bonding Molecules Bonding Forces and Energies Considering the interaction
Chapter 2. Atomic Structure and Interatomic Bonding Interatomic Bonding Bonding forces and energies Primary interatomic bonds Secondary bonding Molecules Bonding Forces and Energies Considering the interaction
TUTORIAL 4 Building a Navigation Bar with Fireworks
 TUTORIAL 4 Building a Navigation Bar with Fireworks This tutorial shows you how to build a Macromedia Fireworks MX 2004 navigation bar that you can use on multiple pages of your website. A navigation bar
TUTORIAL 4 Building a Navigation Bar with Fireworks This tutorial shows you how to build a Macromedia Fireworks MX 2004 navigation bar that you can use on multiple pages of your website. A navigation bar
Chapter 2 Polar Covalent Bonds; Acids and Bases
 John E. McMurry http://www.cengage.com/chemistry/mcmurry Chapter 2 Polar Covalent Bonds; Acids and Bases Javier E. Horta, M.D., Ph.D. University of Massachusetts Lowell Polar Covalent Bonds: Electronegativity
John E. McMurry http://www.cengage.com/chemistry/mcmurry Chapter 2 Polar Covalent Bonds; Acids and Bases Javier E. Horta, M.D., Ph.D. University of Massachusetts Lowell Polar Covalent Bonds: Electronegativity
Peptide bonds: resonance structure. Properties of proteins: Peptide bonds and side chains. Dihedral angles. Peptide bond. Protein physics, Lecture 5
 Protein physics, Lecture 5 Peptide bonds: resonance structure Properties of proteins: Peptide bonds and side chains Proteins are linear polymers However, the peptide binds and side chains restrict conformational
Protein physics, Lecture 5 Peptide bonds: resonance structure Properties of proteins: Peptide bonds and side chains Proteins are linear polymers However, the peptide binds and side chains restrict conformational
Disulfide Bonds at the Hair Salon
 Disulfide Bonds at the Hair Salon Three Alpha Helices Stabilized By Disulfide Bonds! In order for hair to grow 6 inches in one year, 9 1/2 turns of α helix must be produced every second!!! In some proteins,
Disulfide Bonds at the Hair Salon Three Alpha Helices Stabilized By Disulfide Bonds! In order for hair to grow 6 inches in one year, 9 1/2 turns of α helix must be produced every second!!! In some proteins,
H 2O gas: molecules are very far apart
 Non-Covalent Molecular Forces 2/27/06 3/1/06 How does this reaction occur: H 2 O (liquid) H 2 O (gas)? Add energy H 2O gas: molecules are very far apart H 2O liquid: bonding between molecules Use heat
Non-Covalent Molecular Forces 2/27/06 3/1/06 How does this reaction occur: H 2 O (liquid) H 2 O (gas)? Add energy H 2O gas: molecules are very far apart H 2O liquid: bonding between molecules Use heat
Tutorial on Using Gaussview and Gaussian 94
 1 Tutorial on Using Gaussview and Gaussian 94 Written by Vijay Gupta, with editing by M.L. and S.A. Overview Gaussian 94 takes a text file with a.com extension as an input. In this input file, the molecular
1 Tutorial on Using Gaussview and Gaussian 94 Written by Vijay Gupta, with editing by M.L. and S.A. Overview Gaussian 94 takes a text file with a.com extension as an input. In this input file, the molecular
Multiobjective Robust Design Optimization of a docked ligand
 Multiobjective Robust Design Optimization of a docked ligand Carlo Poloni,, Universitaʼ di Trieste Danilo Di Stefano, ESTECO srl Design Process DESIGN ANALYSIS MODEL Dynamic Analysis Logistics & Field
Multiobjective Robust Design Optimization of a docked ligand Carlo Poloni,, Universitaʼ di Trieste Danilo Di Stefano, ESTECO srl Design Process DESIGN ANALYSIS MODEL Dynamic Analysis Logistics & Field
A Guide to the free mesh program Discretizer with OpenFOAM for CFD (Computational Fluid Dynamics)
 Discretizer Manual Release date 09/01/10 Side 1 of 13 A Guide to the free mesh program Discretizer with OpenFOAM for CFD (Computational Fluid Dynamics) Homepage: http://www.discretizer.org/ Creator of
Discretizer Manual Release date 09/01/10 Side 1 of 13 A Guide to the free mesh program Discretizer with OpenFOAM for CFD (Computational Fluid Dynamics) Homepage: http://www.discretizer.org/ Creator of
Use the Force! Noncovalent Molecular Forces
 Use the Force! Noncovalent Molecular Forces Not quite the type of Force we re talking about Before we talk about noncovalent molecular forces, let s talk very briefly about covalent bonds. The Illustrated
Use the Force! Noncovalent Molecular Forces Not quite the type of Force we re talking about Before we talk about noncovalent molecular forces, let s talk very briefly about covalent bonds. The Illustrated
INTERMOLECULAR FORCES
 INTERMOLECULAR FORCES Intermolecular forces- forces of attraction and repulsion between molecules that hold molecules, ions, and atoms together. Intramolecular - forces of chemical bonds within a molecule
INTERMOLECULAR FORCES Intermolecular forces- forces of attraction and repulsion between molecules that hold molecules, ions, and atoms together. Intramolecular - forces of chemical bonds within a molecule
Draft Version. For personal use. c Copyright 2011 - Máté Márton LOHÁSZ. Tutorial for Channel Flow by LES using Open Source CFD
 Tutorial for Channel Flow by LES using Open Source CFD Máté Márton Lohász February 2011 i CONTENTS ii Contents 1 Overview 1 2 Set up your environment 1 3 Very short intro to OpenFOAM 1 3.1 Simulation set-up............................
Tutorial for Channel Flow by LES using Open Source CFD Máté Márton Lohász February 2011 i CONTENTS ii Contents 1 Overview 1 2 Set up your environment 1 3 Very short intro to OpenFOAM 1 3.1 Simulation set-up............................
Matter, Materials, Crystal Structure and Bonding. Chris J. Pickard
 Matter, Materials, Crystal Structure and Bonding Chris J. Pickard Why should a theorist care? Where the atoms are determines what they do Where the atoms can be determines what we can do Overview of Structure
Matter, Materials, Crystal Structure and Bonding Chris J. Pickard Why should a theorist care? Where the atoms are determines what they do Where the atoms can be determines what we can do Overview of Structure
Anime Studio Debut vs. Pro
 vs. Animation Length 2 minutes (3000 frames) Unlimited Motion Tracking 3 Points Unlimited Audio Tracks 2 Tracks Unlimited Video Tracks 1 Track Unlimited Physics No Yes Poser scene import No Yes 3D layer
vs. Animation Length 2 minutes (3000 frames) Unlimited Motion Tracking 3 Points Unlimited Audio Tracks 2 Tracks Unlimited Video Tracks 1 Track Unlimited Physics No Yes Poser scene import No Yes 3D layer
Language: English Lecturer: Gianni de Fabritiis. Teaching staff: Language: English Lecturer: Jordi Villà i Freixa
 MSI: Molecular Simulations Descriptive details concerning the subject: Name of the subject: Molecular Simulations Code : MSI Type of subject: Optional ECTS: 5 Total hours: 125.0 Scheduling: 11:00-13:00
MSI: Molecular Simulations Descriptive details concerning the subject: Name of the subject: Molecular Simulations Code : MSI Type of subject: Optional ECTS: 5 Total hours: 125.0 Scheduling: 11:00-13:00
BIOLOGICAL MEMBRANES: FUNCTIONS, STRUCTURES & TRANSPORT
 BIOLOGICAL MEMBRANES: FUNCTIONS, STRUCTURES & TRANSPORT UNIVERSITY OF PNG SCHOOL OF MEDICINE AND HEALTH SCIENCES DISCIPLINE OF BIOCHEMISTRY AND MOLECULAR BIOLOGY BMLS II / B Pharm II / BDS II VJ Temple
BIOLOGICAL MEMBRANES: FUNCTIONS, STRUCTURES & TRANSPORT UNIVERSITY OF PNG SCHOOL OF MEDICINE AND HEALTH SCIENCES DISCIPLINE OF BIOCHEMISTRY AND MOLECULAR BIOLOGY BMLS II / B Pharm II / BDS II VJ Temple
Combinatorial Biochemistry and Phage Display
 Combinatorial Biochemistry and Phage Display Prof. Valery A. Petrenko Director - Valery Petrenko Instructors Galina Kouzmitcheva and I-Hsuan Chen Auburn 2006, Spring semester COMBINATORIAL BIOCHEMISTRY
Combinatorial Biochemistry and Phage Display Prof. Valery A. Petrenko Director - Valery Petrenko Instructors Galina Kouzmitcheva and I-Hsuan Chen Auburn 2006, Spring semester COMBINATORIAL BIOCHEMISTRY
Character Animation Tutorial
 Character Animation Tutorial 1.Overview 2.Modelling 3.Texturing 5.Skeleton and IKs 4.Keys 5.Export the character and its animations 6.Load the character in Virtools 7.Material & texture tuning 8.Merge
Character Animation Tutorial 1.Overview 2.Modelling 3.Texturing 5.Skeleton and IKs 4.Keys 5.Export the character and its animations 6.Load the character in Virtools 7.Material & texture tuning 8.Merge
Rotation: Moment of Inertia and Torque
 Rotation: Moment of Inertia and Torque Every time we push a door open or tighten a bolt using a wrench, we apply a force that results in a rotational motion about a fixed axis. Through experience we learn
Rotation: Moment of Inertia and Torque Every time we push a door open or tighten a bolt using a wrench, we apply a force that results in a rotational motion about a fixed axis. Through experience we learn
Summary. Load and Open GaussView to start example
 Summary This document describes in great detail how to navigate the Linux Red Hat Terminal to bring up GaussView, use GaussView to create a simple atomic or molecular simulation input file, and then use
Summary This document describes in great detail how to navigate the Linux Red Hat Terminal to bring up GaussView, use GaussView to create a simple atomic or molecular simulation input file, and then use
MET 306. Activity 8a. Mechanism Design Creo 2.0 Level 7 POINT A GROUND LINK LINK 1 LINK 2 LINK 3 POINT B 10/15/2010 1
 Mechanism Design Creo 2.0 Level 7 POINT A LINK 1 GROUND LINK LINK 2 LINK 3 POINT B 10/15/2010 1 Download parts ground, key, link_1, link_2, link_3 and pulley from the V:/MET_306/Activity_8_Creo drive.
Mechanism Design Creo 2.0 Level 7 POINT A LINK 1 GROUND LINK LINK 2 LINK 3 POINT B 10/15/2010 1 Download parts ground, key, link_1, link_2, link_3 and pulley from the V:/MET_306/Activity_8_Creo drive.
The first law: transformation of energy into heat and work. Chemical reactions can be used to provide heat and for doing work.
 The first law: transformation of energy into heat and work Chemical reactions can be used to provide heat and for doing work. Compare fuel value of different compounds. What drives these reactions to proceed
The first law: transformation of energy into heat and work Chemical reactions can be used to provide heat and for doing work. Compare fuel value of different compounds. What drives these reactions to proceed
KINETIC MOLECULAR THEORY OF MATTER
 KINETIC MOLECULAR THEORY OF MATTER The kinetic-molecular theory is based on the idea that particles of matter are always in motion. The theory can be used to explain the properties of solids, liquids,
KINETIC MOLECULAR THEORY OF MATTER The kinetic-molecular theory is based on the idea that particles of matter are always in motion. The theory can be used to explain the properties of solids, liquids,
Lecture 11 Enzymes: Kinetics
 Lecture 11 Enzymes: Kinetics Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 8, pp. 216-225 Key Concepts Kinetics is the study of reaction rates (velocities). Study of enzyme kinetics is useful for
Lecture 11 Enzymes: Kinetics Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 8, pp. 216-225 Key Concepts Kinetics is the study of reaction rates (velocities). Study of enzyme kinetics is useful for
TVM 4155 Numerical modelling and hydraulics 10. March 2014. OpenFOAM homework
 TVM 4155 Numerical modelling and hydraulics 10. March 2014 OpenFOAM homework OpenFOAM is the most popular open-source CFD program in the world today. In this homework we will use the program to determine
TVM 4155 Numerical modelling and hydraulics 10. March 2014 OpenFOAM homework OpenFOAM is the most popular open-source CFD program in the world today. In this homework we will use the program to determine
Modern Construction Materials Prof. Ravindra Gettu Department of Civil Engineering Indian Institute of Technology, Madras
 Modern Construction Materials Prof. Ravindra Gettu Department of Civil Engineering Indian Institute of Technology, Madras Module - 2 Lecture - 2 Part 2 of 2 Review of Atomic Bonding II We will continue
Modern Construction Materials Prof. Ravindra Gettu Department of Civil Engineering Indian Institute of Technology, Madras Module - 2 Lecture - 2 Part 2 of 2 Review of Atomic Bonding II We will continue
Molecular Models in Biology
 Molecular Models in Biology Objectives: After this lab a student will be able to: 1) Understand the properties of atoms that give rise to bonds. 2) Understand how and why atoms form ions. 3) Model covalent,
Molecular Models in Biology Objectives: After this lab a student will be able to: 1) Understand the properties of atoms that give rise to bonds. 2) Understand how and why atoms form ions. 3) Model covalent,
A. OPENING POINT CLOUDS. (Notepad++ Text editor) (Cloud Compare Point cloud and mesh editor) (MeshLab Point cloud and mesh editor)
 MeshLAB tutorial 1 A. OPENING POINT CLOUDS (Notepad++ Text editor) (Cloud Compare Point cloud and mesh editor) (MeshLab Point cloud and mesh editor) 2 OPENING POINT CLOUDS IN NOTEPAD ++ Let us understand
MeshLAB tutorial 1 A. OPENING POINT CLOUDS (Notepad++ Text editor) (Cloud Compare Point cloud and mesh editor) (MeshLab Point cloud and mesh editor) 2 OPENING POINT CLOUDS IN NOTEPAD ++ Let us understand
Protein Dynamics Intro
 Protein Dynamics Intro From rigid structures to motions on energy landscapes Do you all remember Anfinsen? What concept now associated with his name made Anfinsen famous? Right, it is the concept that
Protein Dynamics Intro From rigid structures to motions on energy landscapes Do you all remember Anfinsen? What concept now associated with his name made Anfinsen famous? Right, it is the concept that
SAnDReS Tutorial 01 Prof. Dr. Walter F. de Azevedo Jr.
 2015 Dr. Walter F. de Azevedo Jr. SAnDReS Tutorial 01 Prof. Dr. Walter F. de Azevedo Jr. 1 Running in the Windows On the Windows, left click on Command Prompt. Go to SAnDReS directory (c:\sandres) and
2015 Dr. Walter F. de Azevedo Jr. SAnDReS Tutorial 01 Prof. Dr. Walter F. de Azevedo Jr. 1 Running in the Windows On the Windows, left click on Command Prompt. Go to SAnDReS directory (c:\sandres) and
Conformational analysis of lipid molecules by self-organizing maps
 THE JOURNAL OF CHEMICAL PHYSICS 126, 054707 2007 Conformational analysis of lipid molecules by self-organizing maps Teemu Murtola and Mikko Kupiainen Laboratory of Physics, Helsinki University of Technology,
THE JOURNAL OF CHEMICAL PHYSICS 126, 054707 2007 Conformational analysis of lipid molecules by self-organizing maps Teemu Murtola and Mikko Kupiainen Laboratory of Physics, Helsinki University of Technology,
3.091 OCW Scholar Fall 2010 Final Exam - Solutions Key. Prof. Donald R. Sadoway, Instructor
 .091 OCW Scholar Fall 2010 Final Exam - Solutions Key Prof. Donald R. Sadoway, Instructor .091 Fall Term 2010 Final Exam page 2 Problem #1 (20 points) Answer the following questions about the hydrogen
.091 OCW Scholar Fall 2010 Final Exam - Solutions Key Prof. Donald R. Sadoway, Instructor .091 Fall Term 2010 Final Exam page 2 Problem #1 (20 points) Answer the following questions about the hydrogen
Lottery Looper. User Manual
 Lottery Looper User Manual Lottery Looper 1.7 copyright Timersoft. All rights reserved. http://www.timersoft.com The information contained in this document is subject to change without notice. This document
Lottery Looper User Manual Lottery Looper 1.7 copyright Timersoft. All rights reserved. http://www.timersoft.com The information contained in this document is subject to change without notice. This document
