The power plant of the cell is also a smithy: The emerging role of mitochondria in cellular iron homeostasis
|
|
|
- Giles Reeves
- 8 years ago
- Views:
Transcription
1 Annals of Medicine. 2009; 41: 8299 REVIEW ARTICLE The power plant of the cell is also a smithy: The emerging role of mitochondria in cellular iron homeostasis ALEX D. SHEFTEL & ROLAND LILL Institut für Zytobiologie, Philipps Universität Marburg, Marburg, Germany Downloaded By: [Lill, Roland] At: 21:58 20 February 2009 Abstract Iron is required for a barrage of essential biochemical functions in virtually every species of life. Perturbation of the availability or utilization of iron in these functions or disruption of other components along iron-requiring pathways can not only lead to cellular/organismal insufficiency of respective biochemical end-products but also result in a broad derangement of iron homeostasis. This is largely because of the elaborate regulatory mechanisms that connect cellular iron utilization with uptake and distribution. Such mechanisms are necessitated by the double-edged nature of the metal, whose very property as a useful biological catalyst also makes it able to generate highly toxic compounds. Since the majority of iron is dispatched onto a functional course by mitochondria-localized pathways, these organelles are in an ideal position within the cellular iron anabolic pathways to be a central site for regulation of iron homeostasis. The goal of this article is to provide an overview of how mitochondria acquire and use iron and examine the ramifications of disturbances in these processes on overall cellular iron homeostasis. Key words: Anemia, ataxia, hemochromatosis, iron, iron-sulfur cluster, mitochondria, porphyria, sideroblast Introduction Almost every organism has an absolute requirement for iron. 1 This transition metal, whose chemical properties make it able to readily donate and receive electrons, is a versatile functional co-factor in a variety of essential cellular processes. Despite its ubiquitous usage, the very characteristics that make iron biochemically valuable render it potentially toxic. Under physiological conditions, the ferric form, Fe 3, is virtually insoluble (10 17 M), while ferrous iron (Fe 2 ) will readily catalyze the production of extremely reactive hydroxyl radicals from hydrogen peroxide (known as the Fenton reaction) (13). Therefore, nature has evolved specific handling and regulatory systems to ensure that organisms maintain adequate levels of iron for survival yet remain protected from the metal s inherent harmfulness. When components of the various cellular machineries that handle iron or are integral parts of iron-related pathways are compromised, cellular, tissue, and systemic pathology ensue. This is evidenced by numerous diseases ranging from direct defects in known iron metabolic pathways, such as the hereditary hemochromatoses and sideroblastic anemias, to problems occurring in tissues that do not play an obvious role in systemic iron metabolism, such as in Alzheimer s disease (4), Parkinson s disease (5), and various ataxias (6). Although some iron-containing enzymes, including ribonucleotide reductase (desoxyribonucleotide synthesis), tyrosine hydroxylase (neurotransmitter synthesis), and lipoxygenases (eicosanoid metabolism), use the naked metal as a co-factor, the vast majority of iron proteins wield it within prosthetic groups of heme and iron-sulfur (Fe/S) clusters. Heme is a metalloporphyrin comprised of a tetrapyrrole ring, protoporphyrin IX, with a ferrous iron ion (Fe 2 ) at its center. Due to the ubiquity and versatility of porphyrins and heme, it is generally believed that their use in nature has been apparent since some of the earliest organisms appeared on Correspondence: Roland Lill, Institut für Zytobiologie, Philipps Universität Marburg, Robert-Koch Str. 6, DE Marburg, Germany. Fax: Lill@staff.uni-marburg.de (Received 28 May 2008; revised 30 June 2008; accepted 3 July 2008) ISSN print/issn online # 2009 Informa UK Ltd. (Informa Healthcare, Taylor & Francis AS) DOI: /
2 Key messages. Mitochondria are the major site of cellular iron utilization.. The processing of iron by mitochondria entails specialized, proteinaceous machineries that orchestrate the formation of both heme and iron-sulfur cluster prosthetic groups.. Mitochondria have a pivotal role in the iron handling of the entire cell since perturbations in mitochondrial iron utilization result in aberrant cellular iron uptake and distribution. Earth. Heme is a required component of key cellular functions including oxygen transport (hemoglobin) and storage (myoglobin), respiration (respiratory complexes IIII), cytoprotection (catalase), nitrogen monoxide generation (nitric oxide synthases), detoxification (cytochromes P 450 ), and apoptosis (cytochrome c). Similarly, Fe/S clusters have participated in biochemical catalysis during the formation of the first molecules of life (7) and are intimately involved in such fundamental cellular functions as amino acid synthesis (isopropylmalate isomerase), vitamin biosynthesis (biotin and lipoate synthase), DNA replication (primase), DNA repair (XPD), and cellular respiration (electron transport chain complexes), to name just a few. It has long been known that the synthesis of both heme and Fe/S clusters occurs within mitochondria in non-plant cells (810). Because the mitochondrion is the site of utilization of most of the cell s functionally available iron, it can be expected that the iron handling by this organelle has a significant impact on the overall levels and distribution of iron within the cell. Somewhat surprisingly, research has only just begun to unravel the role of mitochondria in the orchestration of cellular iron homeostasis. This is, at least in part, the result of our still poor understanding of how the metal is transported to and into mitochondria and how such trafficking may be regulated. Additionally, the interdependences of Fe/S protein biogenesis, heme biosynthesis, and cellular iron uptake and storage are obstacles to a fruitful investigation of this enigmatic regulatory network. This review aims to outline what is currently known about the mechanisms of uptake and utilization of iron by mitochondria, discuss the connections of mitochondrial iron utilization to cellular iron homeostasis, and, finally, examine diseases resulting from dysregulation of these pathways. Role of mitochondria in iron homeostasis 83 Iron acquisition and storage by yeast and mammals Yeast has proven to be an excellent model system for examining many biochemical pathways of eukaryotes. In this vein, one would expect the mechanisms that generate ancient biological metal co-factors, whose utility transcend virtually all species, to be quite similar when comparing these unicellular eukaryotes with more complex organisms, such as mammals. Not surprisingly, there exist a significant number of homologous, iron-handling proteins between yeast and mammals. However, there are many additional proteins of mammalian iron metabolism with no obvious yeast complement. A current model of the mammalian iron uptake pathway is presented in Figure 1. Transferrin (Tf), the plasma iron (Fe 3 ) carrier, is the principal source of iron for all mammalian tissues (with notable exceptions being specialized enterocytes and macrophages that take up dietary iron and effete erythrocytes, respectively). Cells acquire iron from this proteinaceous donor via receptor-mediated endocytosis, followed by release of the iron from transferrin within an acidified, endosomal compartment (11). Reduction of iron to its ferrous form is probably catalyzed by the recently identified Steap3 protein (12). Finally, the reduced metal can leave the endosomal compartment by the promiscuous divalent metal transporter, DMT1 (13). Recent evidence suggests that in erythroid cells a direct transfer of iron from the endosome to mitochondria may be necessary for efficient passage of iron through the cell to the enzymes that insert the metal into functional mitochondrial proteins (14,15). Excess iron is stored in the ubiquitously expressed cytosolic protein, ferritin (Ft) (16). Thus, it is likely that little loosely bound iron is present in the cytosol of mammalian cells. Ferritin is a heteroicositetrameric protein consisting of varying proportions of ferritin light chain (Ft-L) and ferritin heavy chain (Ft-H) polypeptides, forming a shell into which up to 4,500 iron atoms may be stored in an inorganic, paracrystalline form (16,17). Ft-H contains a ferroxidase center that is essential for iron storage, while Ft-L contributes to storage efficiency by providing nucleation sites for an iron core formation. A mitochondrial version of Ft (MFt), that also possesses ferroxidase activity and is capable of storing iron, has recently been identified (18,19). Interestingly, excessive expression of Ft-H in cell lines (20,21) or in vivo (22) or overexpression of MFt in a lung cancer cell line (23) all result in a cytosolic iron depletion phenotype. Although the iron storage function of Ft has been recognized since 1937 (24), little is known about the physiological
3 84 A. D. Sheftel & R. Lill Downloaded By: [Lill, Roland] At: 21:58 20 February 2009 Figure 1. Iron uptake pathway in mammalian cells. The majority of iron is taken up by mammalian cells via transferrin (Tf) which receives ferric iron from macrophages or enterocytes. 1) Holo-Tf, after binding to its cognate receptor, traansferrin receptor (TfR) on the cell surface, enters the cell via receptor-mediated endocytosis. This process begins with the formation of a clathrin-coated pit, which matures into an early endosome. 2) The early endosome further matures, and protons enter by the action of the V-type ATPase. 3) When the ph of the endosomes becomes more acidic than 5.5, the Fe 3 is released from the Tf-TfR complex. Iron must then be reduced, probably by a member of the Steap family of proteins, before its translocation out of the endosome by DMT1. The next steps of iron distribution within the cell are less well understood. 4) Either iron is handed off from DMT1 directly to mitochondria from the endosome, utilizing mitoferrin (Mfn) to cross the inner membrane, and the excess portion may subsequently be re-exported to cytosolic ferritin (Ft). 5) Alternatively or additionally, iron leaves endosomes to enter a labile iron pool within the cytosol (LIP), from where it may then be translocated into the mitochondria via Mfn or become loaded into Ft. Once in the mitochondrial matrix, the metal is utilized by either ferrochelatase (FC) to generate heme or the mitochondrial iron-sulfur cluster (ISC) assembly machinery. 6) Apo-Tf (which remains bound to the TfR under acidic conditions) and the TfR recycle to the cell surface, where apo-tf is released into the plasma to become reloaded with Fe 3. mechanism(s) of iron release from the protein or the regulation of this process. Studies in vitro have demonstrated that a combination of reducing agents and chelators is capable of depleting Ft of iron (25,26). In vivo experiments have demonstrated that iron release from Ft may precede lysosomal and/or proteasomal degradation of the protein (27,28). Further, release of iron from Ft synthesized in yeast (which lacks endogenous Ft) suggests that no specific mammalian machinery is necessary for iron release. Importantly, while overexpression of ferroportin, the only known cellular iron exporter, results in decreased cellular ferritin levels, it is still possible that de novo synthesis of mammalian Fe/S and heme proteins requires an influx of Tf-derived iron. In contrast to the mammalian pathway, the major iron uptake mechanism in Saccharomyces cerevisiae entails a direct transport of labile iron from the environment into the cytosol (for review, see (29,30)). On their plasma membranes, this yeast expresses metal reductases, Fre1 and Fre2, which provide ferrous iron for import by the coppercontaining ferroxidase Fet3 and the ferric iron permease Ftr1. Once inside the yeast cell, iron is either utilized or stored in the vacuole. The only protein of the yeast iron uptake machinery having a
4 human homologue is Fet3. Importantly, the human protein, ceruloplasmin, also functions as a ferroxidase, though it functions in the reverse translocation of iron, i.e. the iron release from cells for binding onto serum transferrin. It is important to note that yeast possess machinery enabling them to acquire iron from low-molecular-mass siderophores, even though they are unable to synthesize these iron chelators (for review see (31)). A low-affinity, nonspecific iron uptake system (Fet4) is present in the yeast plasma membrane as well (32). Once iron exits the exosolic compartment (i.e. the endosome in mammalian tissues or the extracellular environment for yeast), its immediate fate is largely unknown and might differ from tissue to tissue (Figure 1). Though it has been shown in hemoglobin-producing cells that iron is efficiently transferred directly from endosomes to mitochondria, further studies are required to determine whether this direct trafficking mechanism may be a valid paradigm for other types of cells. Nevertheless, in both erythroid and non-erythroid cells, a pool of readily available iron can be quenched by the use of membranepermeable chelators. This fraction of cellular iron is generally referred to as the labile iron pool (LIP), yet it still remains to be determined in what form the metal is. In order to be incorporated into heme and Fe/S proteins, the metal must enter mitochondria; whether this is absolutely true for cytosolic Fe/S biogenesis is still uncertain because the origin of the iron in that process is still unclear (see below). Mrs3 and Mrs4 (or its orthologue in higher eukaryotes termed mitoferrin (33)) appear to transport iron across the mitochondrial inner membrane (34). The only known vacuolar importer for iron is Ccc1, which likely transports ferrous iron into the vacuole (35). Heme biosynthesis and its regulation The heme biosynthetic pathway has been extensively characterized and, therefore, is a subject of several comprehensive reviews and monographs (10,36 38). The first, and rate-limiting, step in mammalian protoporphyrin synthesis, the condensation of glycine with succinyl-co-enzyme A to form 5- aminolevulinic acid (ALA), is catalyzed by 5- aminolevulinate synthase (ALAS) and occurs in the mitochondrial matrix. It bears mention here that an alternative pathway exists in plants and some other photosynthetic organisms whereby ALA is formed from glutamate in a series of three enzymatic steps (39). After ALA is generated, it must translocate to the cytosol, where the next four reactions in the heme biosynthetic pathway occur, producing coproporphyrinogen III, which then must re-enter the Role of mitochondria in iron homeostasis 85 mitochondria for the last three steps in heme biosynthesis. The mode of targeting of coproporphyrinogen III into the mitochondrial intermembrane space is unclear. A role of the ATP binding cassette (ABC) transporter ABCB6 claimed to be located in the mitochondrial outer membrane has been suggested in this import step (40). However, the mitochondrial outer membrane contains a large pore, termed VDAC, through which molecules smaller than 5 kda can freely pass. It is therefore unclear why an active transport mechanism may be needed. Moreover, ABCB6 was also found to be located in the plasma membrane and Golgi apparatus (4143). Functionally, ABCB6 can replace the yeast mitochondrial inner membrane protein Atm1 in its role in cytosolic Fe/S protein biogenesis (44), suggesting a similar role to the mammalian Atm1 orthologue ABCB7 (4547). The seventh step of heme biosynthesis is performed by protoporphyrinogen oxidase to produce protoporphyrin IX (PPIX), which is then converted to heme by the insertion of a ferrous iron ion by the enzyme ferrochelatase (FC). These two enzymes are located at the outer and inner face, respectively, of the inner membrane and may physically interact, thus facilitating the transfer of protoporphyrin IX across the inner membrane (48). Ferrochelatase, in most organisms, but not in S. cerevisiae, contains a C-terminal [2Fe-2S] cluster which is essential for its function (49). The presence of an Fe/S co-factor in a heme-producing enzyme underscores the intimate link between the two major iron-consuming pathways. Of particular relevance to this review is the regulation of heme biosynthesis, as heme constitutes a major fraction of utilized, cellular iron. In mammals, heme biosynthesis in non-erythroid tissues is repressed by heme in a feedback manner, in that heme represses transcription of ALAS (50), destabilizes the mrna of ALAS (51,52), and inhibits the import of newly formed ALAS into mitochondria (53,54). In yeast, ALA synthesis does not appear to be limiting in the heme synthesis pathway, as ALA is normally present in excess (55), rather it appears that the second and third enzymes along the pathway, ALA dehydratase and porphobilinogen deaminase, may be rate-limiting in Saccharomyces (56). Additionally, the level of heme synthesis in yeast is regulated on the transcriptional level, depending mainly on the available carbon source and the aerobic status (57,58). In erythroid cells, the factor that controls heme production is iron availability (59). Through erythroid-specific promoter motifs, including GATA-1 and NF-E2 binding sites, at differentiation,
5 86 A. D. Sheftel & R. Lill these hemoglobin-producing cells express a distinct version of ALAS (designated ALAS2 or e-alas), which is encoded by a different gene than that of the housekeeping ALAS (ALAS1). The mrna of ALAS2 contains an iron-responsive element (IRE) in its 5?-untranslated region (UTR), which confers positive regulation in the presence of iron via ironregulatory proteins (IRPs) (see below). Fe/S protein biogenesis Initial insight into the mechanisms of Fe/S protein biogenesis stems from work in bacteria and yeast (with some contribution from photosynthetic models). Only recently have studies in higher eukaryotes been added and generally they have shown a high degree of conservation of this process. What follows is a description of the components and pathways as they have been determined mainly based on the microbiological models, using the yeast protein names (Figure 2; for recent review see (60)). Fe/S protein biogenesis inside mitochondria is mediated by the so-called iron-sulfur cluster (ISC) assembly machinery consisting of 15 known proteins. The first step in mitochondrial Fe/S cluster biogenesis is the pyridoxal 5-phosphate Figure 2. Working model of iron-sulfur cluster biogenesis. In the mitochondria, the complex of Nfs1 and Isd11 functions as a cysteine desulfurase to generate the sulfur needed for biogenesis. An Fe/S cluster is then assembled on the Isu1 scaffold, with ferrous iron transported into the organelle by Mrs3/4 and possibly delivered by Yfh1. Presumably, the electron transfer chain from NADH to the ferredoxin Yah1 serves to reduce the sulfur atom to sulfide. A chaperone system, which includes the Hsp70-like protein Ssq1, the J-type cochaperone Jac1, and the nucleotide exchange factor Mge1, transfers de novo assembled clusters from the scaffold to apo-proteins. Grx5 also participates in some capacity in this cluster delivery system. Isa1, Isa2, and Iba57 all have a specific role in the maturation of a subset of apoproteins, which includes aconitase-type proteins and radical-sam enzymes. The ISC export machinery, comprised of the ABC transporter, Atm1, the intermembrane space sulfhydryl oxidase, Erv1, and glutathione (GSH) moves an intermediate (X) produced by the mitochondrial ISC system to the cytosol. This intermediate facilitates Fe/S protein maturation in the cytosol and nucleus, which further requires the cytosolic Fe/S protein assembly (CIA) machinery. In this pathway, a transient Fe/S cluster is initially formed on the complex of the P-loop NTPases, Cfd1 and Nbp35, which thus serve as scaffolds for Fe/S cluster assembly. Delivery of the Fe/S clusters from the scaffold complex to apo-proteins requires the hydrogenase-like protein Nar1 and the WD40 repeat protein Cia1.
6 (PLP)-dependent liberation of sulfane sulfur from cysteine (and concomitant production of alanine) by the mitochondrial enzyme Nfs1 (61). The small protein Isd11 must be associated with Nfs1 for efficient Fe/S cluster synthesis, though its precise activity is unknown (62,63). The sulfur is then transferred to the scaffold protein, Isu1, likely via a direct contact between Nfs1 and Isu1 (61). Isu1 forms a homodimer capable of accommodating two [2Fe-2S] clusters, which can then be reduced to form one [4Fe-4S] cluster (64,65). An additional mitochondrial protein with possible scaffold function is Nfu1, whose homologues are essential for Fe/ S protein maturation in cyanobacteria and plants (66). It has been shown that the transient formation of an Fe/S cluster on the scaffold protein Isu1 requires the ferredoxin Yah1 and ferredoxin reductase Arh1 (67). Likely, their function is to provide electrons for Fe/S cluster formation on Isu1 by oxidizing the reduced form of nicotinamide adenine dinucleotide (NADH) (68). Iron must also be provided to the scaffold protein. It has also been suggested that Yfh1, the yeast homologue of human frataxin (mutated in Friedreich s ataxia), is involved in the shuttling of iron ions to Isu1 (69,70). After formation of the Fe/S cluster on Isu1, a dedicated chaperone system, comprised of the 70-kDa heat shock protein (Hsp70)-like protein Ssq1, a J-type cochaperone Jac1, and a nucleotide exchange factor Mge1, appears to be responsible for the transfer of the cluster from the scaffold to apoproteins (67). Absence of elements of this complex machinery block or severely inhibit yeast growth, though some functional overlap exists between Ssq1 and the major mitochondrial Hsp70-type chaperone, Ssc1 (8). Glutaredoxin 5 (Grx5) has also emerged as having some (non-essential) role in the dislocation of Fe/S clusters from Isu1 and/or their insertion into apoproteins (67,71). The above paragraph describes the mitochondrial Fe/S protein biogenesis pathway. However, Fe/ S proteins are also present in the cytosol and nucleus, where they perform vital functions. For example, in yeast, cytosolic Fe/S proteins are integral enzymes in the biosynthesis of leucine (isopropylmalate isomerase; Leu1), and methionine (sulfite reductase; Ecm17). Likewise, the nuclear Fe/S proteins Ntg2 (endonuclease; human homologue: Nth1) and Rad3 (DNA helicase) each play specific roles in DNA repair. Importantly, mutations in two human Rad3 homologues, FancJ and XPD, both Fe/S proteins, result in Fanconi anemia and xeroderma pigmentosum, respectively. Most relevant for this review, iron-regulatory protein 1 (IRP1) is a cytosolic Fe/S protein present in higher eukaryotes Role of mitochondria in iron homeostasis 87 only. It functions as a cytosolic aconitase and is involved in the posttranscriptional regulation of genes involved in iron metabolism (see below). Formation and maintenance of extramitochondrial Fe/S proteins is less well understood. It is clear that the mitochondrial ISC assembly system, including Nfs1 (72), Isu1 (73), ferredoxin (74) and frataxin (75), is crucial in the initial formation of the cytosolic and nuclear Fe/S proteins (76). A transport system, termed ISC export machinery, carries components necessary for Fe/S protein maturation out of the mitochondria. The ABC transporter, Atm1, is the focus of this transport system. Its ablation causes defects in the function of extramitochondrial Fe/S proteins (the commonly assayed non-mitochondrial Fe/S proteins are yeast Rli1, Leu1, and Ntg2 and mammalian IRP1). Defects in the two ISC systems (ISC assembly machinery and ISC export machinery) are accompanied by iron accumulation in mitochondria (72) and increased transcription of the yeast iron regulon via Aft1/Aft2, two iron-responsive transcription factors (see below) (77). Further essential parts of the ISC assembly machinery are Erv1 (78), a sulfhydryl oxidase of the mitochondrial intermembrane space, and glutathione, the major redox regulator in yeast (79). Outside the mitochondria, the cytosolic Fe/S protein assembly (CIA) machinery coordinates the maturation of cytosolic and nuclear Fe/S proteins. Cfd1 and Nbp35, two P- loop nucleoside triphosphatases (NTPases) that can bind Fe/S clusters in a transient fashion, form a heterotetramer that is essential for cytosolic Fe/S protein maturation (80,81). Interestingly, the CIA protein Nar1 contains two Fe/S clusters. Insertion of these co-factors requires the function of the mitochondrial ISC assembly machinery along with the CIA components Nbp35/Cfd1, placing Nar1 downstream of these two enzymes in the CIA pathway (82,83). Even further downstream in cytosolic/ nuclear Fe/S protein assembly is the WD40 repeat protein Cia1 (84). As with the other CIA proteins, depletion of Cia1 leads to inhibition of the maturation of cytosolic Fe/S target proteins; however, the Fe/S clusters on Cfd1, Nbp35, and Nar1 are assembled in the absence of Cia1, suggesting that Cia1 has a function late in biogenesis, e.g. in Fe/S cluster transfer to target proteins (85). Role of mitochondria in the control of iron uptake Even though the mechanisms of iron uptake in mammals and lower eukaryotes differ markedly, the uptake systems share an ultimate, common objective:
7 88 A. D. Sheftel & R. Lill maintain a sufficient supply of a metal, yet avoid uptake of devastatingly toxic amounts. As mentioned above, the majority of iron uptake culminates in the mitochondria, where newly acquired iron is incorporated into prosthetic groups. Therefore, the mitochondria reside at the most opportune setting in the iron pathway for evaluation of cellular iron availability. A growing body of experimental evidence strongly implicates the Fe/S protein biogenesis systems in the regulation of cellular iron homeostasis in both Saccharomyces and mammals (Figure 3). In yeast, the collection of genes which respond to changes in iron availability, the iron regulon, is under the control of the transcriptional activators Aft1 and Aft2 (8688). These proteins have not been shown to bind iron or Fe/S clusters (77), but it is known that their subcellular localization is influenced in a critical way by the cellular iron status; when present in the nucleus, Aft1/2 switch on the iron regulon (89). Activation of Aft1/2 occurs in yeast cells deficient in mitochondrial ISC components including Nfs1, Atm1 or glutathione (77,90), suggesting that the enigmatic substrate of Atm1 may be the direct or indirect effector of Aft1/2 transcriptional activation. It is also possible that Aft1/2 may interact with an Fe/ S protein rather than directly assess Atm1 activity; however, this is an unlikely prospect given that depletion of the CIA machinery components, including the Cfd1/Nbp35 scaffold, does not affect regulation of the iron regulon. A number of other proteins appear to participate in Aft1 transcriptional activation. The redundant, cytosolic monothiol glutaredoxins, Grx3 and Grx4, are required for iron-responsive activation of Aft1 (91,92). They directly interact with Aft1, but their precise molecular function is unknown. The proteins Fra1 and Fra2 have recently been found to interact with the Grx proteins in an iron-independent fashion and also play a role in iron regulation (93). Deletion of any of the four proteins elicits a similar induction of the Aft1 transcriptional activity as the inactivation of the ISC components, suggesting that these proteins somehow are involved in the signaling pathway from mitochondria to the nucleus sensing the mitochondrial Fe/S cluster assembly activity to adjust gene expression for proper iron uptake and handling. Hence, the response of the yeast cell to low iron is not a direct assessment of the iron availability. Rather the ability of the mitochondria to form Fe/S clusters seems critical for the gene expression response. In support of this, a recent DNA microarray report documented a considerable, but not complete overlap between the transcriptomes of iron-deficient and mitochondrial ISC-deficient yeast (94). Interestingly, even though heme is also formed in mitochondria, there is no known feedback loop of heme metabolism analogous to that of Fe/S protein biogenesis in yeast. Seemingly paradoxically, when components of heme synthesis are compromised, transcription of Fe uptake genes is decreased (95). Apparently, this serves to avoid an iron overload situation under heme deficiency. Earlier studies had made clear that decreased heme synthesis essentially initiates the switch to anaerobic metabolism, as heme synthesis requires oxygen, and heme itself activates the master regulator of aerobic gene transcription, the transcription factor Hap1 (57). In contrast to the transcriptional regulation of the yeast iron-handling proteins, mammalian cells primarily respond to changes in iron availability through posttranscriptional means by what is known as the IRE/IRP system (for review see (9698)). Messenger RNAs encoding critical proteins of iron homeostasis contain in their 3?- or 5?-UTRs one or more stem loop structures known as iron responsive elements (IREs). Iron-regulatory proteins, of which two isoforms have been identified, IRP1 and IRP2, bind to these IREs and, thus, either stabilize (3?- IRE; transferrin receptor 1 (Tfr)) the respective mrna or block (5?-IRE; ferritin) its translation. This elegant regulatory system allows for rapid, simultaneous, and reciprocal regulation of transferrin receptor 1 (i.e. iron uptake) and ferritin (i.e. iron storage). As mentioned above, IRP1 is a cytosolic Fe/S protein. However, the Fe/S cluster in IRP1 is labile and can dissociate upon opening of the tight structure of IRP1 by a large domain movement (99,100). The presence of a [4Fe-4S] cluster in IRP1 confers aconitase enzyme activity, while the complete absence of a cluster results in high affinity for IREs. Under conditions of low iron, oxidative stress, or nitrosative stress, the Fe/S cluster may become disassembled, thus resulting in the simultaneous loss of aconitase activity and activation of IRE binding. IRP2 has not been shown to contain an Fe/ S cluster, and its protein level is believed to be posttranslationally regulated via iron-dependent degradation. Importantly, impairment of any of the three Fe/S protein biogenesis systems (ISC assembly machinery, ISC export machinery, and CIA machinery) mentioned above affects Fe/S cluster assembly on IRP1 and hence increases IRE binding activity (henceforth referred to as IRP1 activity ; Figure 3B). RNAi depletion of Isu1 in a human cell line has been shown to increase IRP1 activity and concomitantly increase TfR and iron uptake (101). Similar studies in which human Nfs1 or frataxin was depleted in HeLa cells by RNAi technology also
8 Role of mitochondria in iron homeostasis 89 Figure 3. Iron homeostasis is regulated by Fe/S protein assembly systems. (A) In yeast, the ISC assembly machinery produces an unknown substrate which is exported to the cytosol by Atm1. The substrate, which may be identical to the substance needed for Fe/S cluster formation on Cfd1/Nbp35 (see Figure 2), prevents the translocation of the transcription factors Aft1/2 into the nucleus and the activation of the iron regulon. Thus, insufficiency in any part of the mitochondrial ISC machineries results in a low iron signal which is mediated by Aft1/Aft2 and possibly other proteins including Grx3/Grx4 and Fra1/Fra2 to result in the activation of the iron regulon. (B) In mammalian tissues, the presence or absence of an Fe/S cluster on IRP1 serves as a binary switch, determining the binding capacity of the protein to mrna stem loop structures serving as iron-responsive elements (IRE). When the Fe/S cluster is absent, IRP1 has the ability to bind to IREs, thereby stabilizing mrnas with 3 -UTR IREs (e.g. transferrin receptor 1) and blocking translation of mrnas with 5 -UTR IREs (e.g. ferritin). Hence, appropriate regulation of iron handling by mammalian cells requires not only the mitochondrial ISC machineries including the substrate exported by ABCB7, but also a functional cytosolic assembly system (CIA).
9 90 A. D. Sheftel & R. Lill demonstrated an increase in IRP1 activity as a result of mitochondrial Fe/S protein biogenesis defects (102,103). A virtually identical phenotype was observed upon depletion of the ISC export component ABCB7, the functional orthologue of yeast Atm1 (104). Unlike the inactivation of the yeast CIA machinery, which does not significantly impact on cellular iron regulation, depletion of the human Nbp35 and Nar1 protein in cell culture had a strong effect on iron homeostasis, consistent with the ironregulatory role of IRP1 (105,106). Hence, the activities of all three Fe/S protein biogenesis machineries severely affect cellular iron homeostasis in mammals. It is therefore expected that disruption of any critical component of the pathways of Fe/S cluster generation and delivery to apo-irp1 will activate IRP1. Whether any connection exists between heme synthesis and the regulation of iron uptake in mammalian cells is unclear. In erythroid cells, exogenous heme inhibits iron uptake following endocytosis (107). Additionally, some reports suggest a direct negative regulatory effect by heme on IRP2 activity (108,109). Since in these and other studies rather high concentrations of hemin (oxidized heme) were used, the physiological meaning of the findings remains to be clarified (110). Moreover, erythroid precursors of mice lacking IRP2 exhibit decreased TfR (111). Since this type of cell in particular produces copious amounts of heme which are known to be available to exert regulatory effects (107), IRP2 should be severely decreased, along with TfR levels, had heme the ability to directly modulate IRP2. Nonetheless, the regulatory effects of heme on iron homeostasis or other processes may be directly influenced by perturbation of Fe/S cluster synthesis, as ferrochelatase itself contains an Fe/S cluster that is crucial for its function (112). This fact may explain why RNAi depletion of human Isu1 led to increases in IRP2 as well as IRP1 (101). Alternatively, it is likely that iron is redistributed from the LIP to mitochondria when the ISC machinery is compromised, thus increasing IRP2 activity. It has not yet been determined whether heme or iron is responsible for the effects of ISC depletion on IRP2. Additionally, at present, there are no definitive reports documenting compromised ferrochelatase activity as a result of mitochondrial Fe/S cluster production deficiency in mammalian tissue. Dysregulated mitochondrial iron processing An overview of the consequences of defects in various proteins involved in mitochondrial iron handling in mammals is depicted in Figure 4. A general consequence in both yeast and mammalian cells compromised in the mitochondrial ISC machineries is an accumulation of mitochondrial iron. This has been observed in yeast when Nfs1 (72, 113), Isu1 (73), Yah1 (74), Arh1 (114), Yfh1 (115), Isd11 (62), Grx5 (71), Ssq1 (116), Jac1 (117), Atm1 (118), Erv1 (78), or glutathione (79) are depleted. Interestingly, few studies employing mammalian cells demonstrate a similar accumulation of iron in mitochondria upon functional deficiency of ISC components. Cavadini et al. found an increase in mitochondrial Fe after RNAi silencing of ABCB7, the mammalian homologue to Atm1, in HeLa cells (104). In contrast, a conditional mouse knock-out in hepatocytes accumulated iron in an unknown compartment unrelated to mitochondria (46). It remains to be analyzed whether the differences between the two experimental systems are due to the use of ferric ammonium citrate as an iron source in the former study, which is very likely trafficked differently than the physiological iron source, Tf. Despite several other investigations of the roles of mammalian ISC proteins, to the best of our knowledge, there has been no further observation of mitochondrial iron overload in cultured tissue under ISC deficiency. However, inhibition of heme biosynthesis by chemical blockage of ALAS (119) or ALA dehydratase (120) in primary cultures of hemoglobin-producing cells leads to a rapid accumulation of iron in mitochondria. Mitochondrial iron overload resulting from ALAS or ALA dehydratase deficiency has not been described in non-erythroid cells, probably because such cells do not manufacture as copious amounts of heme as do reticulocytes and erythroblasts. Sideroblastic anemia In contrast to tissue culture models, studies using both animal tissues and human disease material have revealed several conditions relating decreased Fe/S protein biogenesis or heme biosynthesis to increased mitochondrial iron. By far the most extensively documented of these scenarios are the sideroblastic anemias, the hall-mark of which is an abundance of ring sideroblasts, i.e. erythroblasts containing ironladen mitochondria. At least some of the iron within mitochondria of ring sideroblasts is likely to be stored within MFt, as most erythroblasts of patients with sideroblastic anemia stain positively for the protein (121), due to elevated levels of MFt compared to normal tissue. Additionally, electron microscopy of tumor tissue with overexpressed MFt revealed mitochondrial iron deposits resembling those of ring sideroblasts (122).
10 Role of mitochondria in iron homeostasis 91 Downloaded By: [Lill, Roland] At: 21:58 20 February 2009 Figure 4. Diseases resulting from deficient Fe/S proteins. Fe/S protein biogenesis components or Fe/S proteins that are associated with human diseases are outlined in red. (ADR adrenodoxin reductase; ADX adrenodoxin; Mfn mitoferrin; ALR augmenter of liver regeneration (Erv1 homologue); MELAS mitochondrial encephalomyopathy, lactic acidosis, and stroke-like episodes; LHON Leber hereditary optic neuropathy). In X-linked sideroblastic anemia (XLSA), patients have inherited pathogenic mutations in the gene encoding ALAS2 that result in impaired heme synthesis and microcytic hypochromic anemia (123,124). Most known mutations in ALAS2 reside within the catalytic domain of the protein (123), and patients with mutations that change an amino acid located in the vicinity of the co-factor (PLP) binding site of ALAS2 exhibit a response to pyridoxine, a precursor of the active co-factor (125,126). A mutation of ALAS2 in zebrafish causes severe microcytic anemia without ring sideroblasts (127), while ablation of the ALAS2 gene in mice is embryonic lethal due to arrested erythroid differentiation (128). However, typical ring sideroblasts were found in adult mice chimeric for ALAS2-null mutant cells (128) as well as in ALAS2-null mutants following a partial transgenic rescue (129). In contrast to ALAS2 deficiency, erythroid heme synthesis is not compromised in the porphyrias which arise from mutations in any of the heme biosynthetic pathway enzymes downstream from ALAS as they are present in great excess. The exception would be erythropoietic protoporphyria (EPP), where mild anemia occurs in about 30% of patients and ring sideroblasts may be seen in some cases (130). This may be consistent with FC, the mutated enzyme in EPP, having the second lowest normal relative activity among the heme pathway enzymes in the erythron, together with the residual FC activity of only about 25% in EPP patients. Sideroblastic anemia also occurs in conjunction with other hereditary defects, including X-linked sideroblastic anemia with ataxia (XLSA/A), glutaredoxin 5 (Grx5) deficiency, thiamine-responsive megaloblastic anemia (TRMA), mitochondrial myopathy with lactic acidosis and sideroblastic anemia (MLASA), and Pearson marrow-pancreas syndrome (patients have large deletions of mitochondrial DNA), as well as inherited defects that have yet to
11 92 A. D. Sheftel & R. Lill be determined (124). In contrast to XLSA, XLSA/A results from a primary deficiency in cytosolic Fe/S protein biogenesis; ABCB7 has been shown to contain missense mutations in three of the families with this disease (124,131,132). The patients with XLSA/A suffer a mild anemia that is associated with overriding neurological features, including delayed motor/cognitive development, incoordination and cerebellar hypoplasia. How heme synthesis becomes impaired in erythroid cells is not elucidated and may be a consequence of reduced cytosolic Fe/S cluster formation impacting on the translation of ALAS2 via IRP1, as ABCB7 is thought to transport an essential component of extramitochondrial Fe/S protein production (see above). Because free PPIX as well as zinc-ppix accumulate in erythrocytes (132), the generation of FC, an Fe/S protein, may also be affected. It is worthwhile to compare this phenotype to the situation seen in Atm1-deficient yeast cells. While a heme synthesis defect is not evident upon weaker depletion of Atm1, a strong depletion or even the gene deletion gives rise to a severe heme deficiency (72,94,118,133). Thus mild depletion of Atm1 elicits only cytosolic Fe/S protein defects, higher iron uptake, and accumulation of mitochondrial iron. These primary phenotypes are then followed by a heme synthesis defect by a transcriptional decrease of yeast ferrochelatase (94). The heme deficiency in ABCB7-defective erythroid cells might be explained similarly to the Grx5 paradigm (see above), i.e. by the impairment of Fe/S cluster assembly on IRP1 which then does bind to the IRE of the ALAS-2 mrna and decrease its synthesis (see below). Non-inherited sideroblastic anemia constitutes a subtype of the myelodysplastic syndromes (MDS), termed refractive anemia with ring sideroblasts (RARS). It is a clonal bone marrow disease arising in a hematopoietic stem cell which also carries (a) mitochondrial defect(s) causing the sideroblastic phenotype. Activities of the rate-limiting heme synthesis enzymes ALAS and FC are normal, while erythrocyte PPIX is raised, which further increases after pyridoxine administration and iron incorporation into heme is reduced (134,135). The recent observation that expression of ABCB7 transcript is reduced in RARS patients may indicate another mechanism, similar to the impairment of ABCB7 in XLSA/A, for the sideroblastic phenotype (136). However, it is not known whether the level of the ABCB7 protein is altered, and the finding may also represent a secondary effect. The associated body iron overload in RARS, due to the ineffective erythropoiesis and typical of all sideroblastic anemias, likely accentuates the iron burden in erythroblast mitochondria as iron removal tends to improve the anemia (130). Lastly, erythroid heme biosynthesis is adversely affected by diverse factors leading to sideroblastic anemia that is fully reversible when the offending factor is removed (130). For example, alcohol and its metabolite, acetaldehyde, are implicated in affecting heme synthesis at several sites as well as mitochondrial protein synthesis. The anti-tuberculosis drug isoniazid interferes with pyridoxine metabolism, depriving ALAS of its co-factor PLP. The antibiotic chloramphenicol inhibits synthesis of mitochondrial membrane proteins (several respiratory complex and adenosine triphosphate (ATP) synthase subunits). Copper deficiency, which may also develop from ingestion of excess zinc, alters systemic iron metabolism consequent to the associated lack of copper-containing ferroxidases (ceruloplasmin and hephaestin). In copper-deficient erythroid cells, heme generation from iron and PPIX is impaired, possibly because the import of ferrous iron into mitochondria is defective due to diminished membrane potential originating from the defect in the copper-containing enzyme cytochrome oxidase (respiratory complex IV). Mrs3/4 / mitoferrin deficiency After discovering in silico that the yeast MRS3/4 genes are co-regulated with Aft1/2-dependent genes, Foury and Roganti first demonstrated a role for these mitochondrial solute carrier proteins in iron metabolism in yeast (137). Further evidence has now been added to suggest Mrs3/4 as mitochondrial iron transporters, at least under iron-limiting conditions (34,138,139). Since the double deletion of MRS3 and MRS4 is not lethal, it is likely that another iron transporter exists in the yeast mitochondrial inner membrane (see? in Figure 2). Subsequently, in screening zebrafish mutants with pathohematological phenotypes, Shaw and coworkers discovered a requirement for the hematopoietic-specific version of zebrafish Mrs3/4, dubbed mitoferrin, in heme biosynthesis (33). Fish with mutated mitoferrin had severe hypochromic erythrocytes. Murine hematopoietic cells with ablated mitoferrin exhibited an extreme blockage of iron accumulation in mitochondria under succinylacetone treatment. The same study also demonstrated functional conservation of the murine, zebrafish and yeast proteins, and suggested the presence of a ubiquitously expressed isoform of mitoferrin. A recent report has described six probands with apparent hereditary mitoferrin deficiency (140). The individuals have EPP, most requiring
12 liver transplantation, but harbored no mutation(s) in FC. All six patients had an aberrantly spliced mitoferrin transcript, predicting a truncated mature protein which had a loss of function in a complementation analysis. It is therefore likely that the elevated PPIX levels result from impaired iron transport to the mitochondrial FC. In addition to this erythroid-specific form of mitoferrin, a paralogous version with 60% amino acid sequence similarity was described, but has not been analyzed further so far. Friedreich s ataxia Friedreich s ataxia (FRDA), the most common recessive ataxia, is caused by a deficiency in the mitochondrial matrix protein frataxin, usually (in 95% of cases) as a result of a GAA repeat expansion in the first intron of the FRDA gene (chromosomal location 9q13) (141). Patients affected with this disease suffer from neurodegeneration of the central and peripheral nervous systems as well as from hypertrophic cardiomyopathy. As already discussed, the yeast homologue of frataxin, Yfh1, plays a non-essential role in Fe/S protein biogenesis (69,70). Accordingly, frataxin deficiency has been shown to cause severely impaired function of mitochondrial Fe/S proteins in heart tissue of FRDA patients (142) as well as both mitochondrial and extramitochondrial Fe/S proteins in mouse (75,143) and in tissue culture (103) models. Accumulation of iron in heart tissue is well documented in patients with the disease (144); however, it still seems uncertain whether this iron is in mitochondria or not. Puccio et al. demonstrated accumulation of iron in the mitochondria of cardiomyocytes in mice with targeted ablation of FRDA by electron microscopy as well as by biochemical means (143). However, relative to the yeast phenotype, the increase in mitochondrial iron was small (about 2-fold versus 1030-fold in yeast), not consistently seen, and was not observed in neuronal tissue which was similarly ablated of frataxin. Further, the heart and neuronal tissues of these mice were devoid of frataxin expression, as opposed to patients who retain some residual frataxin activity. In any case, the pathology associated with frataxin deficiency must be explained on the basis of the primary deficiency in mitochondrial and extramitochondrial Fe/S proteins, and possibly of the secondary (late) accumulation of iron within mitochondria. Although there is a considerable body of evidence in support of a critical role of frataxin in mitochondrial Fe/S protein biogenesis, the actual molecular function(s) of the protein is still discussed. Several Role of mitochondria in iron homeostasis 93 reports suggest that frataxin acts in the delivery of iron to Isu1/IscU (69,70,145), while others assert that frataxin can also donate iron to ferrochelatase for heme biosynthesis (139,146). However, there are a few caveats to this latter proposal: 1) frataxin deficiency in humans does not result in anemia or any other indication of insufficient heme production; 2) frataxin expression decreases with differentiation of erythroid cells, which generate about 85% of body heme (147); 3) the experiments in which frataxin was shown to donate iron to FC were performed in vitro with purified proteins and without controls to verify whether this was a specific, physiological interaction (146); and 4) in yeast, immunological depletion of several ISC proteins including Yfh1 did not decrease FC activity (133). These results do not support a physiological role of frataxin in heme formation. While a mitochondrial form of Ft has been demonstrated to store iron, an iron sequestration function has also been suggested for an oligomeric form of frataxin in vitro (148). In vivo studies, however, have refuted a critical role of iron-dependent oligomerization (149). Interestingly, MFt overexpression rescues the defects observed in frataxin-deficient yeast (150) or HeLa cells (151), perhaps suggesting that the accumulation of toxic, free iron in FRDA is indeed a significant contributor to the disease pathology. ISU deficiency Two recent publications report the discovery of patients with mutations in the gene encoding the Isu1 scaffold protein (152,153). Affected individuals exhibit a marked exercise intolerance, which appeared to be the result of an impairment of oxidative phosphorylation in muscle tissue, evidenced by decreased oxygen extraction, lactic acidosis, reduced succinate dehydrogenase and aconitase activities, a general decrease in Fe/S proteins, and mitochondrial iron deposition in skeletal muscle tissue. An intronic mutation leads to aberrant splicing and ultimately reduced Isu1 mrna and protein levels in muscle tissue. Grx5 deficiency The unfolding of the role of Grx5 followed a similar progression as that of Mrs3/4 and mitoferrin beginning in yeast, proceeding to fish, and eventually finding its way to man. Rodriguez-Manzaneque et al. first demonstrated in yeast that the protein is an important component of the ISC assembly machinery (71). Their study demonstrated that the compromised growth of a yeast GRX5 deletion strain could be rescued by overproduction of Ssq1
13 94 A. D. Sheftel & R. Lill or Isa1, proteins involved in Fe/S cluster delivery to apoproteins. A functional involvement of Grx5 in the dislocation of Fe/S clusters from Isu1 and/or transfer to apoproteins was shown in 55 Fe radiolabeling experiments (67). As expected for a member of the mitochondrial ISC assembly machinery, the mitochondria of yeast cells deficient in Grx5 accumulate substantial amounts of iron compared to controls. The phenotype of zebrafish harboring a deletion in GRX5 is more complex than what is observed in yeast. Similar to the discovery of the mitoferrin mutant, the Grx5-deficient zebrafish were selected from a mutagenesis screen for hematological mutants, and this mutant showed evidence of hypochromic anemia (154). Strikingly, the PPIX biosynthetic pathway appeared to be compromised in the GRX5 mutants and could be rescued by expression of ALAS2 lacking an IRE or by depletion of IRP1. Together, these data purport that Grx5 is required for Fe/S cluster formation on IRP1, which will block ALAS2 translation when it is active (i.e. without an Fe/S cluster). Recently, a patient with diminished levels of Grx5 has been described showing microcytic anemia accompanied by severe iron overload and the presence of ring sideroblasts (155). Compared to controls, increased IRP1 activity, as well as downstream effects thereof, i.e. decreased Ft, elevated TfR, and decreased aconitase, were observed. In addition, chelation therapy with deferoxamine not only reduced liver iron and the number of ring sideroblasts but also increased the patient s hemoglobin levels. While other causes of iron overload in this case were not stated to have been excluded, the phenotypic manifestation of a Grx5 deficiency in bone marrow, with its high heme requirement, underscores the essential role of Fe/S protein biogenesis in the maintenance of iron homeostasis in the erythroid cell. Conclusions In both yeast and man, changes in mitochondrial iron availability and usage largely determine the cellular handling of the metal. The precise role of mitochondria in this process has only begun to unravel; many questions remain, ranging from the mechanistic handling of iron by mitochondria to the regulatory pathways that are affected directly by mitochondrial components. Disruption of the two major ironutilizing pathways (i.e. heme and Fe/S protein biosynthesis), by limitation of substrates or catalysts, can eventuate potentially harmful alterations in cellular iron levels and/or distribution. Furthermore, given the omnipresence of heme and Fe/S proteins in life, the consequences of such alterations can be devastating to the organism. Interestingly, deficiencies in individual components of heme or Fe/S protein biosynthesis in vertebrates tend to affect specific tissues. For example, the human Isu1 mutation only affects skeletal muscle, while frataxin deficiency manifests mainly in the nervous system and the heart, and Grx5 deficiency appears to target the hematopoietic system. Further research, using higher eukaryotic model systems, is required to elucidate the unique iron use and its regulation by the specific tissues or systems in mammals. Another important phenomenon needing further understanding is the regulation of mitochondrial iron uptake and/or retention. What is it about yeast that causes or allows its mitochondria to become overloaded with up to 30- fold over the normal iron levels, while mammalian cells deficient in homologous proteins that exhibit conserved function do not appear to have such mitochondrial loading? What is it about erythroid tissue that makes it more apt to accumulate mitochondrial iron, and how do these cells turn on expression of MFt? A better characterization of the role of mammalian mitochondria in the regulation of cellular iron metabolism is undoubtedly necessary in order to not only understand the pathophysiology of but also develop effective therapies for several human diseases. Acknowledgements We gratefully acknowledge Dr S. Bottomley for helpful comments on our manuscript. A.D.S. is generously supported by fellowships from the Fonds de la Recherche en Santé Québec and the Alexander-von-Humboldt Foundation. R.L. acknowledges generous support from Deutsche Forschungsgemeinschaft (SFB 593 and TR1, Gottfried-Wilhelm Leibniz program, and GRK 1216), the German- Israeli Foundation GIF, Rhön Klinikum AG, von Behring-Röntgen Stiftung, and Fonds der chemischen Industrie. Declaration of interest: The authors report no conflicts of interest. The authors alone are responsible for the content and writing of the paper. Note 1. Several reports document an independence of some species of Lactobacillus on iron for growth, with these organisms using manganese in the place of iron in some essential proteins (see e.g. Imbert and Blondeau, Curr Microbiol, 1998;37:64). However, it is important to note that the Lactobacillus genome encodes several members of the SUF operon, which encodes proteins devoted to Fe/S protein biogenesis
14 (Fontecave et al., J Biol Inorg Chem. 2005; 10:713). It is thus conceivable that also these bacteria make use of iron. References 1. Fenton HJH. On a new reaction or tartaric acid. Chemical News 1876;33: Halliwell B, Gutteridge JM. Role of free radicals and catalytic metal ions in human disease: an overview. Methods Enzymol. 1990;186: Aisen P. The role of transferrin in iron transport. Br J Haematol. 1974;26: Smith MA, Harris PL, Sayre LM, Perry G. Iron accumulation in Alzheimer disease is a source of redox-generated free radicals. Proc Natl Acad Sci U S A. 1997;94: Gotz ME, Double K, Gerlach M, Youdim MB, Riederer P. The relevance of iron in the pathogenesis of Parkinson s disease. Ann NY Acad Sci. 2004;1012: Ponka P. Hereditary causes of disturbed iron homeostasis in the central nervous system. Ann NY Acad Sci. 2004;1012: Huber C, Eisenreich W, Hecht S, Wachtershauser G. A possible primordial peptide cycle. Science. 2003;301: Craig EA, Marszalek J. A specialized mitochondrial molecular chaperone system: a role in formation of Fe/S centers. Cell Mol Life Sci. 2002;59: Rouault TA, Tong WH. Iron-sulphur cluster biogenesis and mitochondrial iron homeostasis. Nat Rev Mol Cell Biol. 2005;6: Ajioka RS, Phillips JD, Kushner JP. Biosynthesis of heme in mammals. Biochim Biophys Acta. 2006;1763: Richardson DR, Ponka P. The molecular mechanisms of the metabolism and transport of iron in normal and neoplastic cells. Biochim Biophys Acta. 1997;1331: Ohgami RS, Campagna DR, Greer EL, Antiochos B, McDonald A, Chen J, et al. Identification of a ferrireductase required for efficient transferrin-dependent iron uptake in erythroid cells. Nat Genet. 2005;37: Gunshin H, Mackenzie B, Berger UV, Gunshin Y, Romero MF, Boron WF, et al. Cloning and characterization of a mammalian proton-coupled metal-ion transporter. Nature. 1997;388: Zhang AS, Sheftel AD, Ponka P. Intracellular kinetics of iron in reticulocytes: evidence for endosome involvement in iron targeting to mitochondria. Blood. 2005;105: Sheftel AD, Zhang AS, Brown C, Shirihai OS, Ponka P. Direct interorganellar transfer of iron from endosome to mitochondrion. Blood. 2007;110: Harrison PM, Arosio P. The ferritins: molecular properties, iron storage function and cellular regulation. Biochim Biophys Acta. 1996;1275: Munro HN, Linder MC. Ferritin: structure, biosynthesis, and role in iron metabolism. Physiol Rev. 1978;58: Levi S, Corsi B, Bosisio M, Invernizzi R, Volz A, Sanford D, et al. A human mitochondrial ferritin encoded by an intronless gene. J Biol Chem. 2001;276: Corsi B, Cozzi A, Arosio P, Drysdale J, Santambrogio P, Campanella A, et al. Human mitochondrial ferritin expressed in HeLa cells incorporates iron and affects cellular iron metabolism. J Biol Chem. 2002;277: Picard V, Renaudie F, Porcher C, Hentze MW, Grandchamp B, Beaumont C. Overexpression of the ferritin H Role of mitochondria in iron homeostasis 95 subunit in cultured erythroid cells changes the intracellular iron distribution. Blood. 1996;87: Picard V, Epsztejn S, Santambrogio P, Cabantchik ZI, Beaumont C. Role of ferritin in the control of the labile iron pool in murine erythroleukemia cells. J Biol Chem. 1998;273: Wilkinson J 4th, Di X, Schonig K, Buss JL, Kock ND, Cline JM, et al. Tissue-specific expression of ferritin H regulates cellular iron homoeostasis in vivo. Biochem J. 2006;395: Nie G, Sheftel AD, Kim SF, Ponka P. Overexpression of mitochondrial ferritin causes cytosolic iron depletion and changes cellular iron homeostasis. Blood. 2005;105: Laufberger V. Sur la cristallisation de la ferritine. Bull Soc Chim Biol. 1937;19: Harrison PM, Hoy TG, Macara IG, Hoare RJ. Ferritin iron uptake and release. Structure-function relationships. Biochem J. 1974;143: Tufano TP, Pecoraro VL, Raymond KN. Ferric ion sequestering agents: kinetics of iron release from ferritin to catechoylamides. Biochim Biophys Acta. 1981;668: Kidane TZ, Sauble E, Linder MC. Release of iron from ferritin requires lysosomal activity. Am J Physiol Cell Physiol. 2006;291:C De Domenico I, Vaughn MB, Li L, Bagley D, Musci G, Ward DM, et al. Ferroportin-mediated mobilization of ferritin iron precedes ferritin degradation by the proteasome. EMBO J. 2006;25: Van Ho A, Ward DM, Kaplan J. Transition metal transport in yeast. Annu Rev Microbiol. 2002;56: De Freitas J, Wintz H, Kim JH, Poynton H, Fox T, Vulpe C. Yeast, a model organism for iron and copper metabolism studies. Biometals. 2003;16: Philpott CC. Iron uptake in fungi: a system for every source. Biochim Biophys Acta. 2006;1763: Protchenko O, Shakoury-Elizeh M, Keane P, Storey J, Androphy R, Philpott CC. Role of PUG1 in inducible porphyrin and heme transport in Saccharomyces cerevisiae. Eukaryot Cell. 2008;7: Shaw GC, Cope JJ, Li L, Corson K, Hersey C, Ackermann GE, et al. Mitoferrin is essential for erythroid iron assimilation. Nature. 2006;440: Muhlenhoff U, Stadler JA, Richhardt N, Seubert A, Eickhorst T, Schweyen RJ, et al. A specific role of the yeast mitochondrial carriers MRS3/4p in mitochondrial iron acquisition under iron-limiting conditions. J Biol Chem. 2003;278: Li L, Chen OS, McVey WD, Kaplan J. CCC1 is a transporter that mediates vacuolar iron storage in yeast. J Biol Chem. 2001;276: Dailey HA. Biosynthesis of Heme and Chlorophylls. New York: McGraw-Hill; Wyckoff EE, Kushner JP. Heme biosynthesis, the porphyrias, and the liver. In: Arias IM, Boyer JL, Fausto N, Jakoby WB, Schachter DA, Shafritz DA, editors. The liver: biology and pathobiology, 3rd ed. New York: Raven Press Ltd, p Kappas A, Sassa S, Galbraith RA, Nordman Y. The porphyrias. In: Scriver CR, Beaudet AL, Sly WS, Valle D, editors. The metabolic and molecular bases of inherited disease, 7th ed. New York: McGraw-Hill, p Heinemann IU, Jahn M, Jahn D. The biochemistry of heme biosynthesis. Arch Biochem Biophys. 2008;474: Krishnamurthy PC, Du G, Fukuda Y, Sun D, Sampath J, Mercer KE, et al. Identification of a mammalian mitochondrial porphyrin transporter. Nature. 2006;443:5869.
15 96 A. D. Sheftel & R. Lill 41. Paterson JK, Shukla S, Black CM, Tachiwada T, Garfield S, Wincovitch S, et al. Human ABCB6 localizes to both the outer mitochondrial membrane and the plasma membrane. Biochemistry. 2007;46: Tsuchida M, Emi Y, Kida Y, Sakaguchi M. Human ABC transporter isoform B6 (ABCB6) localizes primarily in the Golgi apparatus. Biochem Biophys Res Commun. 2008;369: Jalil YA, Ritz V, Jakimenko A, Schmitz-Salue C, Siebert H, Awuah D, et al. Vesicular localization of the rat ATPbinding cassette half-transporter rabcb6. Am J Physiol Cell Physiol. 2008;294:C Mitsuhashi N, Miki T, Senbongi H, Yokoi N, Yano H, Miyazaki M, et al. MTABC3, a novel mitochondrial ATPbinding cassette protein involved in iron homeostasis. J Biol Chem. 2000;275: Csere P, Lill R, Kispal G. Identification of a human mitochondrial ABC transporter, the functional orthologue of yeast Atm1p. FEBS Lett. 1998;441: Pondarre C, Antiochos BB, Campagna DR, Clarke SL, Greer EL, Deck KM, et al. The mitochondrial ATP-binding cassette transporter Abcb7 is essential in mice and participates in cytosolic iron-sulfur cluster biogenesis. Hum Mol Genet. 2006;15: Pondarre C, Campagna DR, Antiochos B, Sikorski L, Mulhern H, Fleming MD. Abcb7, the gene responsible for X-linked sideroblastic anemia with ataxia, is essential for hematopoiesis. Blood. 2007;109: Koch M, Breithaupt C, Kiefersauer R, Freigang J, Huber R, Messerschmidt A. Crystal structure of protoporphyrinogen IX oxidase: a key enzyme in haem and chlorophyll biosynthesis. EMBO J. 2004;23: Dailey HA. Terminal steps of haem biosynthesis. Biochem Soc Trans. 2002;30: May BK, Dogra SC, Sadlon TJ, Bhasker CR, Cox TC, Bottomley SS. Molecular regulation of heme biosynthesis in higher vertebrates. Prog Nucleic Acid Res Mol Biol. 1995;51: Drew PD, Ades IZ. Regulation of the stability of chicken embryo liver delta-aminolevulinate synthase mrna by hemin. Biochem Biophys Res Commun. 1989;162: Hamilton JW, Bement WJ, Sinclair PR, Sinclair JF, Alcedo JA, Wetterhahn KE. Heme regulates hepatic 5-aminolevulinate synthase mrna expression by decreasing mrna half-life and not by altering its rate of transcription. Arch Biochem Biophys. 1991;289: Kikuchi G, Hayashi N. Regulation by heme of synthesis and intracellular translocation of delta-aminolevulinate synthase in the liver. Mol Cell Biochem. 1981;37: Lathrop JT, Timko MP. Regulation by heme of mitochondrial protein transport through a conserved amino acid motif. Science. 1993;259: Labbe-Bois R, Labbe P. Tetrapyrrole and heme biosynthesis in the yeast Saccharomyces cerevisiae. In: Dailey HA, editor. Biosynthesis of heme and chlorophylls. New York: McGraw- Hill; p Hoffman M, Gora M, Rytka J. Identification of rate-limiting steps in yeast heme biosynthesis. Biochem Biophys Res Commun. 2003;310: Zitomer RS, Lowry CV. Regulation of gene expression by oxygen in Saccharomyces cerevisiae. Microbiol Rev. 1992;56: Kwast KE, Burke PV, Poyton RO. Oxygen sensing and the transcriptional regulation of oxygen-responsive genes in yeast. J Exp Biol. 1998;201: Ponka P. Tissue-specific regulation of iron metabolism and heme synthesis: distinct control mechanisms in erythroid cells. Blood. 1997;89: Lill R, Mühlenhoff U. Maturation of iron-sulfur proteins in eukaryotes: mechanisms, connected processes, and diseases. Annu Rev Biochem. 2008;77: Agar JN, Krebs C, Frazzon J, Huynh BH, Dean DR, Johnson MK. IscU as a scaffold for iron-sulfur cluster biosynthesis: sequential assembly of [2Fe-2S] and [4Fe-4S] clusters in IscU. Biochemistry. 2000;39: Adam AC, Bornhovd C, Prokisch H, Neupert W, Hell K. The Nfs1 interacting protein Isd11 has an essential role in Fe/S cluster biogenesis in mitochondria. EMBO J. 2006;25: Wiedemann N, Urzica E, Guiard B, Müller H, Lohaus C, Meyer HE, et al. Essential role of Isd11 in mitochondrial iron-sulfur cluster synthesis on Isu scaffold proteins. EMBO J. 2006;25: Johnson DC, Dean DR, Smith AD, Johnson MK. Structure, function, and formation of biological iron-sulfur clusters. Annu Rev Biochem. 2005;74: Frazzon AP, Ramirez MV, Warek U, Balk J, Frazzon J, Dean DR, et al. Functional analysis of Arabidopsis genes involved in mitochondrial iron-sulfur cluster assembly. Plant Mol Biol. 2007;64: Yabe T, Morimoto K, Kikuchi S, Nishio K, Terashima I, Nakai M. The Arabidopsis chloroplastic NifU-like protein CnfU, which can act as an iron-sulfur cluster scaffold protein, is required for biogenesis of ferredoxin and photosystem I. Plant Cell. 2004;16: Muhlenhoff U, Gerber J, Richhardt N, Lill R. Components involved in assembly and dislocation of iron-sulfur clusters on the scaffold protein Isu1p. EMBO J. 2003;22: Muhlenhoff U, Richhardt N, Gerber J, Lill R. Characterization of iron-sulfur protein assembly in isolated mitochondria. A requirement for ATP, NADH, and reduced iron. J Biol Chem. 2002;277: Gerber J, Muhlenhoff U, Lill R. An interaction between frataxin and Isu1/Nfs1 that is crucial for Fe/S cluster synthesis on Isu1. EMBO Rep. 2003;4: Yoon T, Cowan JA. Iron-sulfur cluster biosynthesis. Characterization of frataxin as an iron donor for assembly of [2Fe-2S] clusters in ISU-type proteins. J Am Chem Soc. 2003;125: Rodriguez-Manzaneque MT, Tamarit J, Belli G, Ros J, Herrero E. Grx5 is a mitochondrial glutaredoxin required for the activity of iron/sulfur enzymes. Mol Biol Cell. 2002;13: Kispal G, Csere P, Prohl C, Lill R. The mitochondrial proteins Atm1p and Nfs1p are essential for biogenesis of cytosolic Fe/S proteins. EMBO J. 1999;18: Gerber J, Neumann K, Prohl C, Muhlenhoff U, Lill R. The yeast scaffold proteins Isu1p and Isu2p are required inside mitochondria for maturation of cytosolic Fe/S proteins. Mol Cell Biol. 2004;24: Lange H, Kaut A, Kispal G, Lill R. A mitochondrial ferredoxin is essential for biogenesis of cellular iron-sulfur proteins. Proc Natl Acad Sci U S A. 2000;97: Martelli A, Wattenhofer-Donze M, Schmucker S, Bouvet S, Reutenauer L, Puccio H. Frataxin is essential for extramitochondrial Fe-S cluster proteins in mammalian tissues. Hum Mol Genet. 2007;16: Lill R, Kispal G. Maturation of cellular Fe-S proteins: an essential function of mitochondria. Trends Biochem Sci. 2000;25: Rutherford JC, Ojeda L, Balk J, Muhlenhoff U, Lill R, Winge DR. Activation of the iron regulon by the yeast Aft1/
16 Aft2 transcription factors depends on mitochondrial but not cytosolic iron-sulfur protein biogenesis. J Biol Chem. 2005;280: Lange H, Lisowsky T, Gerber J, Muhlenhoff U, Kispal G, Lill R. An essential function of the mitochondrial sulfhydryl oxidase Erv1p/ALR in the maturation of cytosolic Fe/S proteins. EMBO Rep. 2001;2: Sipos K, Lange H, Fekete Z, Ullmann P, Lill R, Kispal G. Maturation of cytosolic iron-sulfur proteins requires glutathione. J Biol Chem. 2002;277: Roy A, Solodovnikova N, Nicholson T, Antholine W, Walden WE. A novel eukaryotic factor for cytosolic Fe-S cluster assembly. EMBO J. 2003;22: Hausmann A, Aguilar Netz DJ, Balk J, Pierik AJ, Muhlenhoff U, Lill R. The eukaryotic P loop NTPase Nbp35: an essential component of the cytosolic and nuclear iron-sulfur protein assembly machinery. Proc Natl Acad Sci U S A. 2005;102: Balk J, Lill R. The cell s cookbook for iron-sulfur clusters: recipes for fool s gold? Chembiochem. 2004;5: Balk J, Pierik AJ, Netz DJ, Muhlenhoff U, Lill R. The hydrogenase-like Nar1p is essential for maturation of cytosolic and nuclear iron-sulphur proteins. EMBO J. 2004;23: Balk J, Aguilar Netz DJ, Tepper K, Pierik AJ, Lill R. The essential WD40 protein Cia1 is involved in a late step of cytosolic and nuclear iron-sulfur protein assembly. Mol Cell Biol. 2005;25: Netz DJ, Pierik AJ, Stumpfig M, Muhlenhoff U, Lill R. The Cfd1-Nbp35 complex acts as a scaffold for iron-sulfur protein assembly in the yeast cytosol. Nat Chem Biol. 2007;3: Blaiseau PL, Lesuisse E, Camadro JM. Aft2p, a novel ironregulated transcription activator that modulates, with Aft1p, intracellular iron use and resistance to oxidative stress in yeast. J Biol Chem. 2001;276: Yamaguchi-Iwai Y, Dancis A, Klausner RD. AFT1: a mediator of iron regulated transcriptional control in Saccharomyces cerevisiae. EMBO J. 1995;14: Rutherford JC, Jaron S, Ray E, Brown PO, Winge DR. A second iron-regulatory system in yeast independent of Aft1p. Proc Natl Acad Sci U S A. 2001;98: Yamaguchi-Iwai Y, Ueta R, Fukunaka A, Sasaki R. Subcellular localization of Aft1 transcription factor responds to iron status in Saccharomyces cerevisiae. J Biol Chem. 2002;277: Chen OS, Crisp RJ, Valachovic M, Bard M, Winge DR, Kaplan J. Transcription of the yeast iron regulon does not respond directly to iron but rather to iron-sulfur cluster biosynthesis. J Biol Chem. 2004;279: Ojeda L, Keller G, Muhlenhoff U, Rutherford JC, Lill R, Winge DR. Role of glutaredoxin-3 and glutaredoxin-4 in the iron regulation of the Aft1 transcriptional activator in Saccharomyces cerevisiae. J Biol Chem. 2006;281: Pujol-Carrion N, Belli G, Herrero E, Nogues A, de la Torre- Ruiz MA. Glutaredoxins Grx3 and Grx4 regulate nuclear localisation of Aft1 and the oxidative stress response in Saccharomyces cerevisiae. J Cell Sci. 2006;119: Kumanovics A, Chen O, Li L, Bagley D, Adkins EM, Lin H, et al. Identification of FRA1 and FRA2 as genes involved in regulating the yeast iron regulon in response to decreased mitochondrial iron-sulfur cluster synthesis. J Biol Chem. 2008;283: Hausmann A, Samans B, Lill R, Muhlenhoff U. Cellular and mitochondrial remodeling upon defects in iron-sulfur protein biogenesis. J Biol Chem. 2008;283: Role of mitochondria in iron homeostasis Crisp RJ, Pollington A, Galea C, Jaron S, Yamaguchi-Iwai Y, Kaplan J. Inhibition of heme biosynthesis prevents transcription of iron uptake genes in yeast. J Biol Chem. 2003;278: Hentze MW, Muckenthaler MU, Andrews NC. Balancing acts: molecular control of mammalian iron metabolism. Cell. 2004;117: Wallander ML, Leibold EA, Eisenstein RS. Molecular control of vertebrate iron homeostasis by iron regulatory proteins. Biochim Biophys Acta. 2006;1763: Rouault TA. The role of iron regulatory proteins in mammalian iron homeostasis and disease. Nat Chem Biol. 2006;2: Walden WE, Selezneva AI, Dupuy J, Volbeda A, Fontecilla- Camps JC, Theil EC, et al. Structure of dual function iron regulatory protein 1 complexed with ferritin IRE-RNA. Science. 2006;314: Volz K. The functional duality of iron regulatory protein 1. Curr Opin Struct Biol. 2008;18: Tong WH, Rouault TA. Functions of mitochondrial ISCU and cytosolic ISCU in mammalian iron-sulfur cluster biogenesis and iron homeostasis. Cell Metab. 2006;3: Biederbick A, Stehling O, Rosser R, Niggemeyer B, Nakai Y, Elsässer HP, et al. Role of human mitochondrial Nfs1 in cytosolic iron-sulfur protein biogenesis and iron regulation. Mol Cell Biol. 2006;26: Stehling O, Elsasser HP, Bruckel B, Muhlenhoff U, Lill R. Iron-sulfur protein maturation in human cells: evidence for a function of frataxin. Hum Mol Genet. 2004;13: Cavadini P, Biasiotto G, Poli M, Levi S, Verardi R, Zanella I, et al. RNA silencing of the mitochondrial ABCB7 transporter in HeLa cells causes an iron-deficient phenotype with mitochondrial iron overload. Blood. 2007;109: Song D, Lee FS. A role for IOP1 in mammalian cytosolic iron-sulfur protein biogenesis. J Biol Chem. 2008;283: Stehling O, Netz DJA, Niggemeyer B, Rösser R, Eisenstein RS, Puccio H, et al. The human CIA component hunbp35 is essential for both cytosolic iron-sulfur protein assembly and iron homeostasis. Moll Cell Biol Jun 23 (Epub ahead of print) Ponka P, Schulman HM, Martinez-Medellin J. Haem inhibits iron uptake subsequent to endocytosis of transferrin in reticulocytes. Biochem J. 1988;251: Goessling LS, Mascotti DP, Thach RE. Involvement of heme in the degradation of iron-regulatory protein 2. J Biol Chem. 1998;273: Jeong J, Rouault TA, Levine RL. Identification of a hemesensing domain in iron regulatory protein 2. J Biol Chem. 2004;279: Sassa S. Why heme needs to be degraded to iron, biliverdin IXalpha, and carbon monoxide? Antioxid Redox Signal. 2004;6: Cooperman SS, Meyron-Holtz EG, Olivierre-Wilson H, Ghosh MC, McConnell JP, Rouault TA. Microcytic anemia, erythropoietic protoporphyria, and neurodegeneration in mice with targeted deletion of iron-regulatory protein 2. Blood. 2005;106: Dailey HA, Finnegan MG, Johnson MK. Human ferrochelatase is an iron-sulfur protein. Biochemistry. 1994;33: Li J, Kogan M, Knight SA, Pain D, Dancis A. Yeast mitochondrial protein, Nfs1p, coordinately regulates ironsulfur cluster proteins, cellular iron uptake, and iron distribution. J Biol Chem. 1999;274:
17 98 A. D. Sheftel & R. Lill 114. Li J, Saxena S, Pain D, Dancis A. Adrenodoxin reductase homolog (Arh1p) of yeast mitochondria required for iron homeostasis. J Biol Chem. 2001;276: Babcock M, de Silva D, Oaks R, Davis-Kaplan S, Jiralerspong S, Montermini L, et al. Regulation of mitochondrial iron accumulation by Yfh1p, a putative homolog of frataxin. Science. 1997;276: Knight SA, Sepuri NB, Pain D, Dancis A. Mt-Hsp70 homolog, Ssc2p, required for maturation of yeast frataxin and mitochondrial iron homeostasis. J Biol Chem. 1998;273: Kim R, Saxena S, Gordon DM, Pain D, Dancis A. J-domain protein, Jac1p, of yeast mitochondria required for iron homeostasis and activity of Fe-S cluster proteins. J Biol Chem. 2001;276: Kispal G, Csere P, Guiard B, Lill R. The ABC transporter Atm1p is required for mitochondrial iron homeostasis. FEBS Lett. 1997;418: Borova J, Ponka P, Neuwirt J. Study of intracellular iron distribution in rabbit reticulocytes with normal and inhibited heme synthesis. Biochim Biophys Acta. 1973;320: Ponka P, Wilczynska A, Schulman HM. Iron utilization in rabbit reticulocytes. A study using succinylacetone as an inhibitor or heme synthesis. Biochim Biophys Acta. 1982;720: Cazzola M, Invernizzi R, Bergamaschi G, Levi S, Corsi B, Travaglino E, et al. Mitochondrial ferritin expression in erythroid cells from patients with sideroblastic anemia. Blood. 2003;101: Nie G, Chen G, Sheftel AD, Pantopoulos K, Ponka P. In vivo tumor growth is inhibited by cytosolic iron deprivation caused by the expression of mitochondrial ferritin. Blood. 2006;108: Cotter PD, Baumann M, Bishop DF. Enzymatic defect in X-linked sideroblastic anemia: molecular evidence for erythroid delta-aminolevulinate synthase deficiency. Proc Natl Acad Sci U S A. 1992;89: Bottomley SS. Congenital sideroblastic anemias. Curr Hematol Rep. 2006;5: Shoolingin-Jordan PM, Al Daihan S, Alexeev D, Baxter RL, Bottomley SS, Kahari ID, et al. 5-Aminolevulinic acid synthase: mechanism, mutations and medicine. Biochim Biophys Acta. 2003;1647: Astner I, Schulze JO, van den Heuvel J, Jahn D, Schubert WD, Heinz DW. Crystal structure of 5-aminolevulinate synthase, the first enzyme of heme biosynthesis, and its link to XLSA in humans. EMBO J. 2005;24: Brownlie A, Donovan A, Pratt SJ, Paw BH, Oates AC, Brugnara C, et al. Positional cloning of the zebrafish sauternes gene: a model for congenital sideroblastic anaemia. Nat Genet. 1998;20: Nakajima O, Takahashi S, Harigae H, Furuyama K, Hayashi N, Sassa S, et al. Heme deficiency in erythroid lineage causes differentiation arrest and cytoplasmic iron overload. EMBO J. 1999;18: Nakajima O, Okano S, Harada H, Kusaka T, Gao X, Hosoya T, et al. Transgenic rescue of erythroid 5-aminolevulinate synthase-deficient mice results in the formation of ring sideroblasts and siderocytes. Genes Cells. 2006;11: Bottomley SS. Sideroblastic anemias. In: Greer JP, Joerster J, Lukens JN, Rodgers GM, Paraskevas F, Glader B, editors. Clinical hematology, 11th ed. Philadelphia: Lippincott Williams & Wilkins, p Allikmets R, Raskind WH, Hutchinson A, Schueck ND, Dean M, Koeller DM. Mutation of a putative mitochondrial iron transporter gene (ABC7) in X-linked sideroblastic anemia and ataxia (XLSA/A). Hum Mol Genet. 1999;8: Bekri S, Kispal G, Lange H, Fitzsimons E, Tolmie J, Lill R, et al. Human ABC7 transporter: gene structure and mutation causing X-linked sideroblastic anemia with ataxia with disruption of cytosolic iron-sulfur protein maturation. Blood. 2000;96: Lange H, Muhlenhoff U, Denzel M, Kispal G, Lill R. The heme synthesis defect of mutants impaired in mitochondrial iron-sulfur protein biogenesis is caused by reversible inhibition of ferrochelatase. J Biol Chem. 2004;279: Bottomley SS. Observations on free erythrocyte protoporphyrin in sideroachrestic anemia. Clinica Chimica Acta. 1965;12: Kushner JP, Lee GR, Wintrobe MM, Cartwright GE. Idiopathic refractory sideroblastic anemia: clinical and laboratory investigation of 17 patients and review of the literature. Medicine (Baltimore). 1971;50: Boultwood J, Pellagatti A, Nikpour M, Pushkaran B, Fidler C, Cattan H, et al. The role of the iron transporter ABCB7 in refractory anemia with ring sideroblasts. PLoS ONE. 2008;3:e Foury F, Roganti T. Deletion of the mitochondrial carrier genes MRS3 and MRS4 suppresses mitochondrial iron accumulation in a yeast frataxin-deficient strain. J Biol Chem. 2002;277: Li L, Kaplan J. Characterization of two homologous yeast genes that encode mitochondrial iron transporters. J Biol Chem. 1997;272: Zhang Y, Lyver ER, Knight SA, Lesuisse E, Dancis A. Frataxin and mitochondrial carrier proteins, Mrs3p and Mrs4p, cooperate in providing iron for heme synthesis. J Biol Chem. 2005;280: Shaw GC, Longer NB, Wang YM, et al. Abnormal expression of human mitoferrin (SLC25A37) is associated with a variant of erythropoietic protoporphyria. Blood. 2006;108:6A Campuzano V, Montermini L, Molto MD, Pianese L, Cossée M, Cavalcanti F, et al. Friedreich s ataxia: autosomal recessive disease caused by an intronic GAA triplet repeat expansion. Science. 1996;271: Rotig A, de Lonlay P, Chretien D, Foury F, Koenig M, Sidi D, et al. Aconitase and mitochondrial iron-sulphur protein deficiency in Friedreich ataxia. Nat Genet. 1997;17: Puccio H, Simon D, Cossee M, Criqui-Filipe P, Tiziano F, Melki J, et al. Mouse models for Friedreich ataxia exhibit cardiomyopathy, sensory nerve defect and Fe-S enzyme deficiency followed by intramitochondrial iron deposits. Nat Genet. 2001;27: Sanchez-Casis G, Cote M, Barbeau A. Pathology of the heart in Friedreich s ataxia: review of the literature and report of one case. Can J Neurol Sci. 1976;3: Shan Y, Napoli E, Cortopassi G. Mitochondrial frataxin interacts with ISD11 of the NFS1/ISCU complex and multiple mitochondrial chaperones. Hum Mol Genet. 2007;16: Yoon T, Cowan JA. Frataxin-mediated iron delivery to ferrochelatase in the final step of heme biosynthesis. J Biol Chem. 2004;279: Becker EM, Greer JM, Ponka P, Richardson DR. Erythroid differentiation and protoporphyrin IX down-regulate frataxin expression in Friend cells: characterization of frataxin expression compared to molecules involved in iron metabolism and hemoglobinization. Blood. 2002;99: Cavadini P, O Neill HA, Benada O, Isaya G. Assembly and iron-binding properties of human frataxin, the protein
18 Role of mitochondria in iron homeostasis 99 deficient in Friedreich ataxia. Hum Mol Genet. 2002;11: Aloria K, Schilke B, Andrew A, Craig EA. Iron-induced oligomerization of yeast frataxin homologue Yfh1 is dispensable in vivo. EMBO Rep. 2004;5: Campanella A, Isaya G, O Neill HA, Santambrogio P, Cozzi A, Arosio P, et al. The expression of human mitochondrial ferritin rescues respiratory function in frataxin-deficient yeast. Hum Mol Genet. 2004;13: Zanella I, Derosas M, Corrado M, Cocco E, Cavadini P, Biasiotto G, et al. The effects of frataxin silencing in HeLa cells are rescued by the expression of human mitochondrial ferritin. Biochim Biophys Acta. 2008;1782: Olsson A, Lind L, Thornell LE, Holmberg M. Myopathy with lactic acidosis is linked to chromosome 12q and caused by an intron mutation in the ISCU gene resulting in a splicing defect. Hum Mol Genet. 2008;17: Mochel F, Knight MA, Tong WH, Hernandez D, Ayyad K, Taivassalo T, et al. Splice mutation in the iron-sulfur cluster scaffold protein ISCU causes myopathy with exercise intolerance. Am J Hum Genet. 2008;82: Wingert RA, Galloway JL, Barut B, Foott H, Fraenkel P, Axe JL, et al. Deficiency of glutaredoxin 5 reveals Fe-S clusters are required for vertebrate haem synthesis. Nature. 2005;436: Camaschella C, Campanella A, De Falco L, Boschetto L, Merlini R, Silvestri L, et al. The human counterpart of zebrafish shiraz shows sideroblastic-like microcytic anemia and iron overload. Blood. 2007;110:13538.
Chapter 9 Mitochondrial Structure and Function
 Chapter 9 Mitochondrial Structure and Function 1 2 3 Structure and function Oxidative phosphorylation and ATP Synthesis Peroxisome Overview 2 Mitochondria have characteristic morphologies despite variable
Chapter 9 Mitochondrial Structure and Function 1 2 3 Structure and function Oxidative phosphorylation and ATP Synthesis Peroxisome Overview 2 Mitochondria have characteristic morphologies despite variable
AP BIOLOGY CHAPTER 7 Cellular Respiration Outline
 AP BIOLOGY CHAPTER 7 Cellular Respiration Outline I. How cells get energy. A. Cellular Respiration 1. Cellular respiration includes the various metabolic pathways that break down carbohydrates and other
AP BIOLOGY CHAPTER 7 Cellular Respiration Outline I. How cells get energy. A. Cellular Respiration 1. Cellular respiration includes the various metabolic pathways that break down carbohydrates and other
Copyright 2000-2003 Mark Brandt, Ph.D. 54
 Pyruvate Oxidation Overview of pyruvate metabolism Pyruvate can be produced in a variety of ways. It is an end product of glycolysis, and can be derived from lactate taken up from the environment (or,
Pyruvate Oxidation Overview of pyruvate metabolism Pyruvate can be produced in a variety of ways. It is an end product of glycolysis, and can be derived from lactate taken up from the environment (or,
Chem 306 Chapter 21 Bioenergetics Lecture Outline III
 Chem 306 Chapter 21 Bioenergetics Lecture Outline III I. HOW IS ATP GENERATED IN THE FINAL STAGE CATABOLISM? A. OVERVIEW 1. At the end of the citric acid cycle, all six carbons of glucose have been oxidized
Chem 306 Chapter 21 Bioenergetics Lecture Outline III I. HOW IS ATP GENERATED IN THE FINAL STAGE CATABOLISM? A. OVERVIEW 1. At the end of the citric acid cycle, all six carbons of glucose have been oxidized
What affects an enzyme s activity? General environmental factors, such as temperature and ph. Chemicals that specifically influence the enzyme.
 CH s 8-9 Respiration & Metabolism Metabolism A catalyst is a chemical agent that speeds up a reaction without being consumed by the reaction. An enzyme is a catalytic protein. Hydrolysis of sucrose by
CH s 8-9 Respiration & Metabolism Metabolism A catalyst is a chemical agent that speeds up a reaction without being consumed by the reaction. An enzyme is a catalytic protein. Hydrolysis of sucrose by
AP Bio Photosynthesis & Respiration
 AP Bio Photosynthesis & Respiration Multiple Choice Identify the letter of the choice that best completes the statement or answers the question. 1. What is the term used for the metabolic pathway in which
AP Bio Photosynthesis & Respiration Multiple Choice Identify the letter of the choice that best completes the statement or answers the question. 1. What is the term used for the metabolic pathway in which
* Is chemical energy potential or kinetic energy? The position of what is storing energy?
 Biology 1406 Exam 2 - Metabolism Chs. 5, 6 and 7 energy - capacity to do work 5.10 kinetic energy - energy of motion : light, electrical, thermal, mechanical potential energy - energy of position or stored
Biology 1406 Exam 2 - Metabolism Chs. 5, 6 and 7 energy - capacity to do work 5.10 kinetic energy - energy of motion : light, electrical, thermal, mechanical potential energy - energy of position or stored
Lecture 8. Protein Trafficking/Targeting. Protein targeting is necessary for proteins that are destined to work outside the cytoplasm.
 Protein Trafficking/Targeting (8.1) Lecture 8 Protein Trafficking/Targeting Protein targeting is necessary for proteins that are destined to work outside the cytoplasm. Protein targeting is more complex
Protein Trafficking/Targeting (8.1) Lecture 8 Protein Trafficking/Targeting Protein targeting is necessary for proteins that are destined to work outside the cytoplasm. Protein targeting is more complex
Energy Production In A Cell (Chapter 25 Metabolism)
 Energy Production In A Cell (Chapter 25 Metabolism) Large food molecules contain a lot of potential energy in the form of chemical bonds but it requires a lot of work to liberate the energy. Cells need
Energy Production In A Cell (Chapter 25 Metabolism) Large food molecules contain a lot of potential energy in the form of chemical bonds but it requires a lot of work to liberate the energy. Cells need
Myoglobin and Hemoglobin
 Myoglobin and Hemoglobin Myoglobin and hemoglobin are hemeproteins whose physiological importance is principally related to their ability to bind molecular oxygen. Myoglobin (Mb) The oxygen storage protein
Myoglobin and Hemoglobin Myoglobin and hemoglobin are hemeproteins whose physiological importance is principally related to their ability to bind molecular oxygen. Myoglobin (Mb) The oxygen storage protein
Anabolic and Catabolic Reactions are Linked by ATP in Living Organisms
 Chapter 5: Microbial Metabolism Microbial Metabolism Metabolism refers to all chemical reactions that occur within a living a living organism. These chemical reactions are generally of two types: Catabolic:
Chapter 5: Microbial Metabolism Microbial Metabolism Metabolism refers to all chemical reactions that occur within a living a living organism. These chemical reactions are generally of two types: Catabolic:
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
Chapter 7 Active Reading Guide Cellular Respiration and Fermentation
 Name: AP Biology Mr. Croft Chapter 7 Active Reading Guide Cellular Respiration and Fermentation Overview: Before getting involved with the details of cellular respiration and photosynthesis, take a second
Name: AP Biology Mr. Croft Chapter 7 Active Reading Guide Cellular Respiration and Fermentation Overview: Before getting involved with the details of cellular respiration and photosynthesis, take a second
Figure 5. Energy of activation with and without an enzyme.
 Biology 20 Laboratory ENZYMES & CELLULAR RESPIRATION OBJECTIVE To be able to list the general characteristics of enzymes. To study the effects of enzymes on the rate of chemical reactions. To demonstrate
Biology 20 Laboratory ENZYMES & CELLULAR RESPIRATION OBJECTIVE To be able to list the general characteristics of enzymes. To study the effects of enzymes on the rate of chemical reactions. To demonstrate
The correct answer is d C. Answer c is incorrect. Reliance on the energy produced by others is a characteristic of heterotrophs.
 1. An autotroph is an organism that a. extracts energy from organic sources b. converts energy from sunlight into chemical energy c. relies on the energy produced by other organisms as an energy source
1. An autotroph is an organism that a. extracts energy from organic sources b. converts energy from sunlight into chemical energy c. relies on the energy produced by other organisms as an energy source
1. Enzymes. Biochemical Reactions. Chapter 5: Microbial Metabolism. 1. Enzymes. 2. ATP Production. 3. Autotrophic Processes
 Chapter 5: Microbial Metabolism 1. Enzymes 2. ATP Production 3. Autotrophic Processes 1. Enzymes Biochemical Reactions All living cells depend on biochemical reactions to maintain homeostasis. All of the
Chapter 5: Microbial Metabolism 1. Enzymes 2. ATP Production 3. Autotrophic Processes 1. Enzymes Biochemical Reactions All living cells depend on biochemical reactions to maintain homeostasis. All of the
Evolution of Metabolism. Introduction. Introduction. Introduction. How Cells Harvest Energy. Chapter 7 & 8
 How ells Harvest Energy hapter 7 & 8 Evolution of Metabolism A hypothetical timeline for the evolution of metabolism - all in prokaryotic cells!: 1. ability to store chemical energy in ATP 2. evolution
How ells Harvest Energy hapter 7 & 8 Evolution of Metabolism A hypothetical timeline for the evolution of metabolism - all in prokaryotic cells!: 1. ability to store chemical energy in ATP 2. evolution
Chapter 7 Cellular Respiration
 Phases of aerobic cellular respiration 1. Glycolysis 2. Transition or Acetyl-CoA reaction 3. Krebs cycle 4. Electron transport system Chapter 7 Cellular Respiration These phases are nothing more than metabolic
Phases of aerobic cellular respiration 1. Glycolysis 2. Transition or Acetyl-CoA reaction 3. Krebs cycle 4. Electron transport system Chapter 7 Cellular Respiration These phases are nothing more than metabolic
Cellular Respiration and Fermentation
 LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 9 Cellular Respiration and Fermentation
LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 9 Cellular Respiration and Fermentation
Chapter 16 The Citric Acid Cycle
 Chapter 16 The Citric Acid Cycle Multiple Choice Questions 1. Which of the following is not true of the reaction catalyzed by the pyruvate dehydrogenase complex? A) Biotin participates in the decarboxylation.
Chapter 16 The Citric Acid Cycle Multiple Choice Questions 1. Which of the following is not true of the reaction catalyzed by the pyruvate dehydrogenase complex? A) Biotin participates in the decarboxylation.
Oxidative Phosphorylation
 Oxidative Phosphorylation NADH from Glycolysis must be transported into the mitochondrion to be oxidized by the respiratory electron transport chain. Only the electrons from NADH are transported, these
Oxidative Phosphorylation NADH from Glycolysis must be transported into the mitochondrion to be oxidized by the respiratory electron transport chain. Only the electrons from NADH are transported, these
Visualizing Cell Processes
 Visualizing Cell Processes A Series of Five Programs produced by BioMEDIA ASSOCIATES Content Guide for Program 3 Photosynthesis and Cellular Respiration Copyright 2001, BioMEDIA ASSOCIATES www.ebiomedia.com
Visualizing Cell Processes A Series of Five Programs produced by BioMEDIA ASSOCIATES Content Guide for Program 3 Photosynthesis and Cellular Respiration Copyright 2001, BioMEDIA ASSOCIATES www.ebiomedia.com
Summary of Metabolism. Mechanism of Enzyme Action
 Summary of Metabolism Mechanism of Enzyme Action 1. The substrate contacts the active site 2. The enzyme-substrate complex is formed. 3. The substrate molecule is altered (atoms are rearranged, or the
Summary of Metabolism Mechanism of Enzyme Action 1. The substrate contacts the active site 2. The enzyme-substrate complex is formed. 3. The substrate molecule is altered (atoms are rearranged, or the
Chapter 8: Energy and Metabolism
 Chapter 8: Energy and Metabolism 1. Discuss energy conversions and the 1 st and 2 nd law of thermodynamics. Be sure to use the terms work, potential energy, kinetic energy, and entropy. 2. What are Joules
Chapter 8: Energy and Metabolism 1. Discuss energy conversions and the 1 st and 2 nd law of thermodynamics. Be sure to use the terms work, potential energy, kinetic energy, and entropy. 2. What are Joules
Lecture Series 7. From DNA to Protein. Genotype to Phenotype. Reading Assignments. A. Genes and the Synthesis of Polypeptides
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Chapter 2: Cell Structure and Function pg. 70-107
 UNIT 1: Biochemistry Chapter 2: Cell Structure and Function pg. 70-107 Organelles are internal structures that carry out specialized functions, interacting and complementing each other. Animal and plant
UNIT 1: Biochemistry Chapter 2: Cell Structure and Function pg. 70-107 Organelles are internal structures that carry out specialized functions, interacting and complementing each other. Animal and plant
Regulation of the Citric Acid Cycle
 Regulation of the itric Acid ycle I. hanges in Free Energy February 17, 2003 Bryant Miles kj/mol 40 20 0 20 40 60 80 Reaction DGo' DG TA Free Energy hanges 1 2 3 4 5 6 7 8 9 1.) itrate Synthase 2.) Aconitase
Regulation of the itric Acid ycle I. hanges in Free Energy February 17, 2003 Bryant Miles kj/mol 40 20 0 20 40 60 80 Reaction DGo' DG TA Free Energy hanges 1 2 3 4 5 6 7 8 9 1.) itrate Synthase 2.) Aconitase
Student name ID # 2. (4 pts) What is the terminal electron acceptor in respiration? In photosynthesis? O2, NADP+
 1. Membrane transport. A. (4 pts) What ion couples primary and secondary active transport in animal cells? What ion serves the same function in plant cells? Na+, H+ 2. (4 pts) What is the terminal electron
1. Membrane transport. A. (4 pts) What ion couples primary and secondary active transport in animal cells? What ion serves the same function in plant cells? Na+, H+ 2. (4 pts) What is the terminal electron
1- Fatty acids are activated to acyl-coas and the acyl group is further transferred to carnitine because:
 Section 10 Multiple Choice 1- Fatty acids are activated to acyl-coas and the acyl group is further transferred to carnitine because: A) acyl-carnitines readily cross the mitochondrial inner membrane, but
Section 10 Multiple Choice 1- Fatty acids are activated to acyl-coas and the acyl group is further transferred to carnitine because: A) acyl-carnitines readily cross the mitochondrial inner membrane, but
Chapter 19a Oxidative Phosphorylation and Photophosphorylation. Multiple Choice Questions
 Chapter 19a Oxidative Phosphorylation and Photophosphorylation Multiple Choice Questions 1. Electron-transfer reactions in mitochondria Page: 707 Difficulty: 1 Ans: E Almost all of the oxygen (O 2 ) one
Chapter 19a Oxidative Phosphorylation and Photophosphorylation Multiple Choice Questions 1. Electron-transfer reactions in mitochondria Page: 707 Difficulty: 1 Ans: E Almost all of the oxygen (O 2 ) one
SOME Important Points About Cellular Energetics by Dr. Ty C.M. Hoffman
 SOME Important Points About Cellular Energetics by Dr. Ty C.M. Hoffman An Introduction to Metabolism Most biochemical processes occur as biochemical pathways, each individual reaction of which is catalyzed
SOME Important Points About Cellular Energetics by Dr. Ty C.M. Hoffman An Introduction to Metabolism Most biochemical processes occur as biochemical pathways, each individual reaction of which is catalyzed
21.8 The Citric Acid Cycle
 21.8 The Citric Acid Cycle The carbon atoms from the first two stages of catabolism are carried into the third stage as acetyl groups bonded to coenzyme A. Like the phosphoryl groups in ATP molecules,
21.8 The Citric Acid Cycle The carbon atoms from the first two stages of catabolism are carried into the third stage as acetyl groups bonded to coenzyme A. Like the phosphoryl groups in ATP molecules,
Compartmentalization of the Cell. Objectives. Recommended Reading. Professor Alfred Cuschieri. Department of Anatomy University of Malta
 Compartmentalization of the Cell Professor Alfred Cuschieri Department of Anatomy University of Malta Objectives By the end of this session the student should be able to: 1. Identify the different organelles
Compartmentalization of the Cell Professor Alfred Cuschieri Department of Anatomy University of Malta Objectives By the end of this session the student should be able to: 1. Identify the different organelles
Sickle cell anemia: Altered beta chain Single AA change (#6 Glu to Val) Consequence: Protein polymerizes Change in RBC shape ---> phenotypes
 Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
Name Date Period. 2. When a molecule of double-stranded DNA undergoes replication, it results in
 DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
The Electron Transport Chain
 The Electron Transport hain February 19, 2003 Bryant Miles The citric acid cycle oxidizes acetate into two molecules of 2 while capturing the electrons in the form of 3 NAD molecules and one molecule of
The Electron Transport hain February 19, 2003 Bryant Miles The citric acid cycle oxidizes acetate into two molecules of 2 while capturing the electrons in the form of 3 NAD molecules and one molecule of
Todays Outline. Metabolism. Why do cells need energy? How do cells acquire energy? Metabolism. Concepts & Processes. The cells capacity to:
 and Work Metabolic Pathways Enzymes Features Factors Affecting Enzyme Activity Membrane Transport Diffusion Osmosis Passive Transport Active Transport Bulk Transport Todays Outline -Releasing Pathways
and Work Metabolic Pathways Enzymes Features Factors Affecting Enzyme Activity Membrane Transport Diffusion Osmosis Passive Transport Active Transport Bulk Transport Todays Outline -Releasing Pathways
Chapter 16 The Citric Acid Cycle
 Chapter 16 The Citric Acid Cycle Multiple Choice Questions 1. Production of acetyl-coa (activated acetate) Page: 603 Difficulty: 2 Ans: A Which of the following is not true of the reaction catalyzed by
Chapter 16 The Citric Acid Cycle Multiple Choice Questions 1. Production of acetyl-coa (activated acetate) Page: 603 Difficulty: 2 Ans: A Which of the following is not true of the reaction catalyzed by
Methods of Grading S/N Style of grading Percentage Score 1 Attendance, class work and assignment 10 2 Test 20 3 Examination 70 Total 100
 COURSE: MIB 303 Microbial Physiology and Metabolism (3 Units- Compulsory) Course Duration: Three hours per week for 15 weeks (45 hours). Lecturer: Jimoh, S.O. B.Sc., M.Sc, Ph.D Microbiology (ABU, Zaria)
COURSE: MIB 303 Microbial Physiology and Metabolism (3 Units- Compulsory) Course Duration: Three hours per week for 15 weeks (45 hours). Lecturer: Jimoh, S.O. B.Sc., M.Sc, Ph.D Microbiology (ABU, Zaria)
Harvesting Energy: Glycolysis and Cellular Respiration. Chapter 8
 Harvesting Energy: Glycolysis and Cellular Respiration Chapter 8 Overview of Glucose Breakdown The overall equation for the complete breakdown of glucose is: C 6 H 12 O 6 + 6O 2 6CO 2 + 6H 2 O + ATP The
Harvesting Energy: Glycolysis and Cellular Respiration Chapter 8 Overview of Glucose Breakdown The overall equation for the complete breakdown of glucose is: C 6 H 12 O 6 + 6O 2 6CO 2 + 6H 2 O + ATP The
Cellular Respiration An Overview
 Why? Cellular Respiration An Overview What are the phases of cellular respiration? All cells need energy all the time, and their primary source of energy is ATP. The methods cells use to make ATP vary
Why? Cellular Respiration An Overview What are the phases of cellular respiration? All cells need energy all the time, and their primary source of energy is ATP. The methods cells use to make ATP vary
Structure and Function of DNA
 Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Electron transport chain, oxidative phosphorylation & mitochondrial transport systems. Joško Ivica
 Electron transport chain, oxidative phosphorylation & mitochondrial transport systems Joško Ivica Electron transport chain & oxidative phosphorylation collects e - & -H Oxidation of foodstuffs oxidizes
Electron transport chain, oxidative phosphorylation & mitochondrial transport systems Joško Ivica Electron transport chain & oxidative phosphorylation collects e - & -H Oxidation of foodstuffs oxidizes
1. Explain the difference between fermentation and cellular respiration.
 : Harvesting Chemical Energy Name Period Overview: Before getting involved with the details of cellular respiration and photosynthesis, take a second to look at the big picture. Photosynthesis and cellular
: Harvesting Chemical Energy Name Period Overview: Before getting involved with the details of cellular respiration and photosynthesis, take a second to look at the big picture. Photosynthesis and cellular
How To Understand How Gene Expression Is Regulated
 What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
1. The diagram below represents a biological process
 1. The diagram below represents a biological process 5. The chart below indicates the elements contained in four different molecules and the number of atoms of each element in those molecules. Which set
1. The diagram below represents a biological process 5. The chart below indicates the elements contained in four different molecules and the number of atoms of each element in those molecules. Which set
Given these characteristics of life, which of the following objects is considered a living organism? W. X. Y. Z.
 Cell Structure and Organization 1. All living things must possess certain characteristics. They are all composed of one or more cells. They can grow, reproduce, and pass their genes on to their offspring.
Cell Structure and Organization 1. All living things must possess certain characteristics. They are all composed of one or more cells. They can grow, reproduce, and pass their genes on to their offspring.
2007 7.013 Problem Set 1 KEY
 2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
Electron Transport System. May 16, 2014 Hagop Atamian hatamian@ucdavis.edu
 Electron Transport System May 16, 2014 Hagop Atamian hatamian@ucdavis.edu What did We learn so far? Glucose is converted to pyruvate in glycolysis. The process generates two ATPs. Pyruvate is taken into
Electron Transport System May 16, 2014 Hagop Atamian hatamian@ucdavis.edu What did We learn so far? Glucose is converted to pyruvate in glycolysis. The process generates two ATPs. Pyruvate is taken into
Anatomy and Physiology Placement Exam 2 Practice with Answers at End!
 Anatomy and Physiology Placement Exam 2 Practice with Answers at End! General Chemical Principles 1. bonds are characterized by the sharing of electrons between the participating atoms. a. hydrogen b.
Anatomy and Physiology Placement Exam 2 Practice with Answers at End! General Chemical Principles 1. bonds are characterized by the sharing of electrons between the participating atoms. a. hydrogen b.
Chapter 8: An Introduction to Metabolism
 Chapter 8: An Introduction to Metabolism Name Period Concept 8.1 An organism s metabolism transforms matter and energy, subject to the laws of thermodynamics 1. Define metabolism. The totality of an organism
Chapter 8: An Introduction to Metabolism Name Period Concept 8.1 An organism s metabolism transforms matter and energy, subject to the laws of thermodynamics 1. Define metabolism. The totality of an organism
How To Treat The Sando Syndrome
 Consultation on Drugs for Rare Diseases Rare Disease Day Symposium Hannover Medical School (MHH), Solidarity Rare but Strong Together Roland Seifert, MHH @ Dear Sir, I am writing hoping that there might
Consultation on Drugs for Rare Diseases Rare Disease Day Symposium Hannover Medical School (MHH), Solidarity Rare but Strong Together Roland Seifert, MHH @ Dear Sir, I am writing hoping that there might
PRESTWICK ACADEMY NATIONAL 5 BIOLOGY CELL BIOLOGY SUMMARY
 Name PRESTWICK ACADEMY NATIONAL 5 BIOLOGY CELL BIOLOGY SUMMARY Cell Structure Identify animal, plant, fungal and bacterial cell ultrastructure and know the structures functions. Plant cell Animal cell
Name PRESTWICK ACADEMY NATIONAL 5 BIOLOGY CELL BIOLOGY SUMMARY Cell Structure Identify animal, plant, fungal and bacterial cell ultrastructure and know the structures functions. Plant cell Animal cell
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION. Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.
 Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
An Overview of Cells and Cell Research
 An Overview of Cells and Cell Research 1 An Overview of Cells and Cell Research Chapter Outline Model Species and Cell types Cell components Tools of Cell Biology Model Species E. Coli: simplest organism
An Overview of Cells and Cell Research 1 An Overview of Cells and Cell Research Chapter Outline Model Species and Cell types Cell components Tools of Cell Biology Model Species E. Coli: simplest organism
William Shaw, Ph.D. The Great Plains Laboratory, Inc., Lenexa, Kansas, USA
 Inhibition of dopamine conversion to norepinephrine by Clostridia metabolites appears to be a (the) major cause of autism, schizophrenia, and other neuropsychiatric disorders. All these factors can now
Inhibition of dopamine conversion to norepinephrine by Clostridia metabolites appears to be a (the) major cause of autism, schizophrenia, and other neuropsychiatric disorders. All these factors can now
Eisenstoffwechsel. Ernst Müllner. MFPL (Max F Perutz Laboratories) Department of Medical Biochemistry Medical University of Vienna
 Eisenstoffwechsel Ernst Müllner MFPL (Max F Perutz Laboratories) Department of Medical Biochemistry Medical University of Vienna ernst.muellner@meduniwien.ac.at oxygen binding hemoglobins oxygen metabolism
Eisenstoffwechsel Ernst Müllner MFPL (Max F Perutz Laboratories) Department of Medical Biochemistry Medical University of Vienna ernst.muellner@meduniwien.ac.at oxygen binding hemoglobins oxygen metabolism
Chapter 8. Summary and Perspectives
 Chapter 8 Summary and Perspectives 131 Chapter 8 Summary Overexpression of the multidrug resistance protein MRP1 confer multidrug resistance (MDR) to cancer cells. The contents of this thesis describe
Chapter 8 Summary and Perspectives 131 Chapter 8 Summary Overexpression of the multidrug resistance protein MRP1 confer multidrug resistance (MDR) to cancer cells. The contents of this thesis describe
CELLS: PLANT CELLS 20 FEBRUARY 2013
 CELLS: PLANT CELLS 20 FEBRUARY 2013 Lesson Description In this lesson we will discuss the following: The Cell Theory Terminology Parts of Plant Cells: Organelles Difference between plant and animal cells
CELLS: PLANT CELLS 20 FEBRUARY 2013 Lesson Description In this lesson we will discuss the following: The Cell Theory Terminology Parts of Plant Cells: Organelles Difference between plant and animal cells
Electron Transport Generates a Proton Gradient Across the Membrane
 Electron Transport Generates a Proton Gradient Across the Membrane Each of respiratory enzyme complexes couples the energy released by electron transfer across it to an uptake of protons from water in
Electron Transport Generates a Proton Gradient Across the Membrane Each of respiratory enzyme complexes couples the energy released by electron transfer across it to an uptake of protons from water in
Keystone Review Practice Test Module A Cells and Cell Processes. 1. Which characteristic is shared by all prokaryotes and eukaryotes?
 Keystone Review Practice Test Module A Cells and Cell Processes 1. Which characteristic is shared by all prokaryotes and eukaryotes? a. Ability to store hereditary information b. Use of organelles to control
Keystone Review Practice Test Module A Cells and Cell Processes 1. Which characteristic is shared by all prokaryotes and eukaryotes? a. Ability to store hereditary information b. Use of organelles to control
PHOTOSYNTHESIS AND CELLULAR RESPIRATION
 reflect Wind turbines shown in the photo on the right are large structures with blades that move in response to air movement. When the wind blows, the blades rotate. This motion generates energy that is
reflect Wind turbines shown in the photo on the right are large structures with blades that move in response to air movement. When the wind blows, the blades rotate. This motion generates energy that is
pathway that involves taking in heat from the environment at each step. C.
 Study Island Cell Energy Keystone Review 1. Cells obtain energy by either capturing light energy through photosynthesis or by breaking down carbohydrates through cellular respiration. In both photosynthesis
Study Island Cell Energy Keystone Review 1. Cells obtain energy by either capturing light energy through photosynthesis or by breaking down carbohydrates through cellular respiration. In both photosynthesis
008 Chapter 8. Student:
 008 Chapter 8 Student: 1. Some bacteria are strict aerobes and others are strict anaerobes. Some bacteria, however, are facultative anaerobes and can live with or without oxygen. If given the choice of
008 Chapter 8 Student: 1. Some bacteria are strict aerobes and others are strict anaerobes. Some bacteria, however, are facultative anaerobes and can live with or without oxygen. If given the choice of
Control of Gene Expression
 Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
mrna EDITING Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A. Copyright 2005
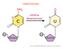 mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
Chapter 14- RESPIRATION IN PLANTS
 Chapter 14- RESPIRATION IN PLANTS Living cells require a continuous supply of energy for maintaining various life activities. This energy is obtained by oxidizing the organic compounds (carbohydrates,
Chapter 14- RESPIRATION IN PLANTS Living cells require a continuous supply of energy for maintaining various life activities. This energy is obtained by oxidizing the organic compounds (carbohydrates,
Cell Injury, Adaptation and Death
 Harvard-MIT Division of Health Sciences and Technology HST.035: Principle and Practice of Human Pathology Dr. Badizadegan Cell Injury, Adaptation and Death HST.035 Spring 2003 Overview of Cell Injury Cells
Harvard-MIT Division of Health Sciences and Technology HST.035: Principle and Practice of Human Pathology Dr. Badizadegan Cell Injury, Adaptation and Death HST.035 Spring 2003 Overview of Cell Injury Cells
Control of Gene Expression
 Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
MULTIPLE CHOICE QUESTIONS
 MULTIPLE CHOICE QUESTIONS 1. Most components of energy conversion systems evolved very early; thus, the most fundamental aspects of energy metabolism tend to be: A. quite different among a diverse group
MULTIPLE CHOICE QUESTIONS 1. Most components of energy conversion systems evolved very early; thus, the most fundamental aspects of energy metabolism tend to be: A. quite different among a diverse group
Cell Structure & Function!
 Cell Structure & Function! Chapter 3! The most exciting phrase to hear in science, the one that heralds new discoveries, is not 'Eureka!' but 'That's funny.! -- Isaac Asimov Animal Cell Plant Cell Cell
Cell Structure & Function! Chapter 3! The most exciting phrase to hear in science, the one that heralds new discoveries, is not 'Eureka!' but 'That's funny.! -- Isaac Asimov Animal Cell Plant Cell Cell
Photosynthesis takes place in three stages:
 Photosynthesis takes place in three stages: Light-dependent reactions Light-independent reactions The Calvin cycle 1. Capturing energy from sunlight 2. Using energy to make ATP and NADPH 3. Using ATP and
Photosynthesis takes place in three stages: Light-dependent reactions Light-independent reactions The Calvin cycle 1. Capturing energy from sunlight 2. Using energy to make ATP and NADPH 3. Using ATP and
Eukaryotes have organelles
 Energy-transducing Eukaryotes have organelles membrane systems An organelle is a discrete membrane bound cellular structure specialized functions. An organelle is to the cell what an organ is to the body
Energy-transducing Eukaryotes have organelles membrane systems An organelle is a discrete membrane bound cellular structure specialized functions. An organelle is to the cell what an organ is to the body
Control of Gene Expression
 Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
APh/BE161: Physical Biology of the Cell Winter 2009 Recap on Photosynthesis Rob Phillips
 APh/BE161: Physical Biology of the Cell Winter 2009 Recap on Photosynthesis Rob Phillips Big picture: why are we doing this? A) photosynthesis will explain shortly, b) more generally, interaction of light
APh/BE161: Physical Biology of the Cell Winter 2009 Recap on Photosynthesis Rob Phillips Big picture: why are we doing this? A) photosynthesis will explain shortly, b) more generally, interaction of light
Enzymes and Metabolic Pathways
 Enzymes and Metabolic Pathways Enzyme characteristics Made of protein Catalysts: reactions occur 1,000,000 times faster with enzymes Not part of reaction Not changed or affected by reaction Used over and
Enzymes and Metabolic Pathways Enzyme characteristics Made of protein Catalysts: reactions occur 1,000,000 times faster with enzymes Not part of reaction Not changed or affected by reaction Used over and
DNA Replication & Protein Synthesis. This isn t a baaaaaaaddd chapter!!!
 DNA Replication & Protein Synthesis This isn t a baaaaaaaddd chapter!!! The Discovery of DNA s Structure Watson and Crick s discovery of DNA s structure was based on almost fifty years of research by other
DNA Replication & Protein Synthesis This isn t a baaaaaaaddd chapter!!! The Discovery of DNA s Structure Watson and Crick s discovery of DNA s structure was based on almost fifty years of research by other
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Bio 101 Section 001: Practice Questions for First Exam
 Do the Practice Exam under exam conditions. Time yourself! MULTIPLE CHOICE: 1. The substrate fits in the of an enzyme: (A) allosteric site (B) active site (C) reaction groove (D) Golgi body (E) inhibitor
Do the Practice Exam under exam conditions. Time yourself! MULTIPLE CHOICE: 1. The substrate fits in the of an enzyme: (A) allosteric site (B) active site (C) reaction groove (D) Golgi body (E) inhibitor
MCAS Biology. Review Packet
 MCAS Biology Review Packet 1 Name Class Date 1. Define organic. THE CHEMISTRY OF LIFE 2. All living things are made up of 6 essential elements: SPONCH. Name the six elements of life. S N P C O H 3. Elements
MCAS Biology Review Packet 1 Name Class Date 1. Define organic. THE CHEMISTRY OF LIFE 2. All living things are made up of 6 essential elements: SPONCH. Name the six elements of life. S N P C O H 3. Elements
MOLECULAR CELL BIOLOGY: ESSAY OUTLINE. What are peroxisomes? What do they do? And, how are proteins targeted to them?
 MOLECULAR CELL BIOLOGY: ESSAY OUTLINE What are peroxisomes? What do they do? And, how are proteins targeted to them? Are they related to the ER/Golgi? Are they more like mitochondria? Or, are they completely
MOLECULAR CELL BIOLOGY: ESSAY OUTLINE What are peroxisomes? What do they do? And, how are proteins targeted to them? Are they related to the ER/Golgi? Are they more like mitochondria? Or, are they completely
Lecture 6. Regulation of Protein Synthesis at the Translational Level
 Regulation of Protein Synthesis (6.1) Lecture 6 Regulation of Protein Synthesis at the Translational Level Comparison of EF-Tu-GDP and EF-Tu-GTP conformations EF-Tu-GDP EF-Tu-GTP Next: Comparison of GDP
Regulation of Protein Synthesis (6.1) Lecture 6 Regulation of Protein Synthesis at the Translational Level Comparison of EF-Tu-GDP and EF-Tu-GTP conformations EF-Tu-GDP EF-Tu-GTP Next: Comparison of GDP
Metabolism Dr.kareema Amine Al-Khafaji Assistant professor in microbiology, and dermatologist Babylon University, College of Medicine, Department of
 Metabolism Dr.kareema Amine Al-Khafaji Assistant professor in microbiology, and dermatologist Babylon University, College of Medicine, Department of Microbiology. Metabolism sum of all chemical processes
Metabolism Dr.kareema Amine Al-Khafaji Assistant professor in microbiology, and dermatologist Babylon University, College of Medicine, Department of Microbiology. Metabolism sum of all chemical processes
Review of the Cell and Its Organelles
 Biology Learning Centre Review of the Cell and Its Organelles Tips for most effective learning of this material: Memorize the names and structures over several days. This will help you retain what you
Biology Learning Centre Review of the Cell and Its Organelles Tips for most effective learning of this material: Memorize the names and structures over several days. This will help you retain what you
The Cell Teaching Notes and Answer Keys
 The Cell Teaching Notes and Answer Keys Subject area: Science / Biology Topic focus: The Cell: components, types of cells, organelles, levels of organization Learning Aims: describe similarities and differences
The Cell Teaching Notes and Answer Keys Subject area: Science / Biology Topic focus: The Cell: components, types of cells, organelles, levels of organization Learning Aims: describe similarities and differences
Regulation of enzyme activity
 1 Regulation of enzyme activity Regulation of enzyme activity is important to coordinate the different metabolic processes. It is also important for homeostasis i.e. to maintain the internal environment
1 Regulation of enzyme activity Regulation of enzyme activity is important to coordinate the different metabolic processes. It is also important for homeostasis i.e. to maintain the internal environment
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA
 CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
The Aerobic Fate of Pyruvate
 The Aerobic Fate of yruvate February 12, 2003 Bryant Miles I could tell that some of you were not impressed by the mere 2 ATs produced per glucose by glycolysis. The 2 AT s produced are only a small fraction
The Aerobic Fate of yruvate February 12, 2003 Bryant Miles I could tell that some of you were not impressed by the mere 2 ATs produced per glucose by glycolysis. The 2 AT s produced are only a small fraction
Electron Transport and Oxidative Phosphorylation
 CHM333 LECTURES 37 & 38: 4/27 29/13 SPRING 2013 Professor Christine Hrycyna Electron Transport and Oxidative Phosphorylation Final stages of aerobic oxidation of biomolecules in eukaryotes occur in the
CHM333 LECTURES 37 & 38: 4/27 29/13 SPRING 2013 Professor Christine Hrycyna Electron Transport and Oxidative Phosphorylation Final stages of aerobic oxidation of biomolecules in eukaryotes occur in the
RNA and Protein Synthesis
 Name lass Date RN and Protein Synthesis Information and Heredity Q: How does information fl ow from DN to RN to direct the synthesis of proteins? 13.1 What is RN? WHT I KNOW SMPLE NSWER: RN is a nucleic
Name lass Date RN and Protein Synthesis Information and Heredity Q: How does information fl ow from DN to RN to direct the synthesis of proteins? 13.1 What is RN? WHT I KNOW SMPLE NSWER: RN is a nucleic
Gene Regulation -- The Lac Operon
 Gene Regulation -- The Lac Operon Specific proteins are present in different tissues and some appear only at certain times during development. All cells of a higher organism have the full set of genes:
Gene Regulation -- The Lac Operon Specific proteins are present in different tissues and some appear only at certain times during development. All cells of a higher organism have the full set of genes:
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Translation Study Guide
 Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
http://faculty.sau.edu.sa/h.alshehri
 http://faculty.sau.edu.sa/h.alshehri Definition: Proteins are macromolecules with a backbone formed by polymerization of amino acids. Proteins carry out a number of functions in living organisms: - They
http://faculty.sau.edu.sa/h.alshehri Definition: Proteins are macromolecules with a backbone formed by polymerization of amino acids. Proteins carry out a number of functions in living organisms: - They
Complex multicellular organisms are produced by cells that switch genes on and off during development.
 Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Chemical Basis of Life Module A Anchor 2
 Chemical Basis of Life Module A Anchor 2 Key Concepts: - Water is a polar molecule. Therefore, it is able to form multiple hydrogen bonds, which account for many of its special properties. - Water s polarity
Chemical Basis of Life Module A Anchor 2 Key Concepts: - Water is a polar molecule. Therefore, it is able to form multiple hydrogen bonds, which account for many of its special properties. - Water s polarity
The amount of cellular adenine is constant. -It exists as either ATP, ADP, or AMP (the concentration of these vary)
 Electron transport chain Final stage of aerobic oxidation! Also known as: -oxidative phosphorylation(when coupled to ATP synthase) -respiration (when coupled to ATP synthase) Purpose: -Recycle reduced
Electron transport chain Final stage of aerobic oxidation! Also known as: -oxidative phosphorylation(when coupled to ATP synthase) -respiration (when coupled to ATP synthase) Purpose: -Recycle reduced
Photosynthesis (CO 2 + H 2 O C 6 H 12 O 6 + O 2 )
 The vital role of A This is the energy-rich compound that is the source of energy for all living things. It is a nucleotide, comprising a 5C sugar (ribose); an organic base (adenosine); and 3 phosphate
The vital role of A This is the energy-rich compound that is the source of energy for all living things. It is a nucleotide, comprising a 5C sugar (ribose); an organic base (adenosine); and 3 phosphate
Genetics Lecture Notes 7.03 2005. Lectures 1 2
 Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Lecture 3: Mutations
 Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
