Report Concomitant Replacement of Language and mtdna in South Caspian Populations of Iran
|
|
|
- Caren Small
- 8 years ago
- Views:
Transcription
1 Current Biology 16, , April 4, 2006 ª2006 Elsevier Ltd All rights reserved DOI /j.cub Report Concomitant Replacement of Language and mtdna in South Caspian Populations of Iran Ivan Nasidze, 1, * Dominique Quinque, 1 Manijeh Rahmani, 2 Seyed Ali Alemohamad, 3 and Mark Stoneking 1 1 Department of Evolutionary Genetics Max Planck Institute for Evolutionary Anthropology Deutscher Platz 6 D Leipzig Germany 2 Department of Molecular Genetics Cardiovascular Research Center Imam Hospital Tehran University of Medical Sciences Tehran Iran 3 Department of Human Genetics School of Public Health Tehran University of Medical Sciences Tehran Iran Summary The Gilaki and Mazandarani occupy the South Caspian region of Iran and speak languages belonging to the North-Western branch of Iranian languages [1]. It has been suggested that their ancestors came from the Caucasus region, perhaps displacing an earlier group in the South Caspian [2]. Linguistic evidence supports this scenario, in that the Gilaki and Mazandarani languages (but not other Iranian languages) share certain typological features with Caucasian languages [3, 4]. We analyzed patterns of mtdna and Y chromosome variation in the Gilaki and Mazandarani. Based on mtdna HV1 sequences, the Gilaki and Mazandarani most closely resemble their geographic and linguistic neighbors, namely other Iranian groups. However, their Y chromosome types most closely resemble those found in groups from the South Caucasus. A scenario that explains these differences is a south Caucasian origin for the ancestors of the Gilaki and Mazandarani, followed by introgression of women (but not men) from local Iranian groups, possibly because of patrilocality. Given that both mtdna and language are maternally transmitted, the incorporation of local Iranian women would have resulted in the concomitant replacement of the ancestral Caucasian language and mtdna types of the Gilaki and Mazandarani with their current Iranian language and mtdna types. Concomitant replacement of language and mtdna may be a more general phenomenon than previously recognized. *Correspondence: nasidze@eva.mpg.de Results and Discussion mtdna Variation We analyzed mtdna haplogroups characteristic for southwest Asia (i.e., haplogroups H, J, N, T, and U) and sequences of the first hypervariable segment of the mtdna control region (HV1) and found high diversity in the Gilaki and Mazandarani (Figure 1), comparable to other groups in this region (see Table S1 in the Supplemental Data available with this article online). Pairwise F st comparisons, based on the HV1 sequences, showed that the Gilaki and Mazandarani groups are very similar to each other and to other Iranian and West Asian groups (Table 1). They are next most similar to Caucasus groups and then European groups, and most distant from Central Asian groups (Table 1). An MDS plot (Figure 2A) based on the pairwise F st values illustrates these patterns. Specifically, the Mazandarani and Gilaki groups are located near each other, in a cluster containing other Iranian and West Asian populations. Thus, with respect to patterns of mtdna variation, the South Caspian groups resemble their geographic and linguistic neighbors, namely other Iranian groups. Analyses of mtdna haplogroups further support the close relationship between South Caspian and Iranian groups. Pairwise F st comparisons, based on HV1 sequences within haplogroups, indicate that the Gilaki and Mazandarani groups are closer to Iranian groups than to populations from the South Caucasus region (Figure 3). To further investigate the relationships between the Mazandarani/Gilaki and groups from Iran and the South Caucasus, we constructed networks of the HV1 sequences, grouped according to mtdna haplogroup (Figure S1). For haplogroups J, T, and U, most of the Mazandarani/Gilaki HVI sequences cluster with Iranian sequences, whereas for haplogroups H and N, there is no clear clustering. Thus, both the F st comparisons and the networks of HV1 sequences, within haplogroups, support a closer relationship of the South Caspian groups with other Iranian groups than with South Caucasian groups. Y Chromosome Variation Overall, seven Y-SNP haplogroups were found in the Gilaki and ten haplogroups were found in the Mazandarani (Table S2). Haplogroup J2*(M172) was found at high frequency in both groups, as was haplogroup R1*(M173); together, these two haplogroups account for more than 50% of Mazandarani and Gilaki Y chromosomes. Interestingly, the frequency of haplogroup J2*(M172) in these groups is more similar to the frequency in South Caucasus groups than in other Iranian groups [5, 6]. Moreover, haplogroup I*(M170) is found at high frequency in the Iranian groups from Tehran and Isfahan, but is absent in the Mazandarani and Gilaki and is in low frequency in the South Caucasus groups (Table S2). The pairwise F st value (Table 1) between the Mazandarani and Gilaki groups was not significantly different
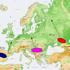
2 Concomitant Replacement of Language and mtdna 669 Figure 1. Location of Sampling Sites in the Mazandaran and Gilan Provinces of Iran from zero (F st = 0.003, p = 0.337). In contrast to the mtdna results, both groups showed greater similarity with South Caucasian populations than with their geographic and linguistic neighbors, namely other Iranians (F st = and for Mazandarani versus South Caucasian and versus Iranians, respectively; and F st = and for Gilaki versus South Caucasian and versus Iranians, respectively). The MDS analysis (Figure 2B) further illustrates these patterns. The Mazandarani and Gilaki groups fall inside a major cluster consisting of populations from the Caucasus and West Asia and are particularly close to the South Caucasus groups Georgians, Armenians, and Azerbaijanians. Iranians from Tehran and Isfahan are situated more distantly from these groups. Three haplogroups were found at high frequencies in the Mazandarani and Gilaki groups (R1*(M173), G*(M201), and J2*(M172)); to further investigate the relationships of these groups based on these three Y-SNP haplogroups, we typed nine Y-STR loci in individuals with these Y-SNP haplogroups and compared the results with the same set of Y-STR loci on the same Y-SNP background that were typed previously in the groups from the South Caucasus and Iran [7]. Reduced median networks of the Y-STR haplotypes (Figure S2) further indicate a closer relationship of the Mazandarani/Gilaki Table 1. Mean Pairwise F st Values between Mazandarani and Gilaki Groups and between South Caucausian, European, and West and Central Asian Groups Mazandarani Gilani S. Caucasus E. Europe W. Europe Centr. Asia Iran West Asia w/o Iran Mazandarani Gilaki S. Caucasus East Europe West Europe Centr. Asia Iran West Asia w/o Iran Below diagonal empty cells are shown pairwise F st values based on Y-SNP haplogroups; above diagonal empty cells are shown pairwise F st values based on mtdna HVI sequences.
3 Current Biology 670 Figure 2. Multidimentional Scaling Plots Multidimensional scaling (MDS) plots (STA- TISTICA package [StatSoft Inc.]) based on pairwise F st values, showing relationships among the Mazandarani and Gilaki groups, and Caucasian, European, and Central and West Asian groups. Mazandarani and Gilaki groups are represented by stars; Caucasus groups by circles; European groups by squares; Central Asian groups by diamonds; and West Asian groups by triangles. (A) Based on mtdna HVI sequence data. The stress value for the MDS plot is (B) Based on Y chromosome SNP data. The stress value for the MDS plot is Names of the groups are abbreviated as follows: Abk, Abkhazians; And, Andalusians; Ar, Armenians; Az, Azerbaijanians; Balu, Baluch; Bart, Bartangi; Br, British; Brah, Brahui; Cat, Catalans; Ch, Cherkessians; Cz-Sl, Czech & Slovaks; Dar, Darginians; Dru, Drusi; Dut, Dutch; Fr, French; Ge, Georgians; Ger, Germans; Gil, Gilaki; Gre, Greeks; Haz, Hazar; Hung, Hungarians; Iq, Iraqi; Ir_Isf, Persians from Isfahan; Ir_Teh, Persians from Tehran; It, Italians; Ishka, Ishkinasi; Kaz, Kazakh; Ku_G, Kurds from Georgia; Ku_I, Kurds from Iran; Ku_K, Kurmanji speaker Kurds; Ku_ Turkm, Kurds from Turkmenistan; Ku_Z, Zazaki speaker Kurds; Kyr, Kyrgiz; Leb, Lebanese; Lez, Lezginians; Makr, Makrani; Maz, Mazandarani; Os_A, Ossetians from Ardon; Os_D, Ossetians from Digora; Pol, Polish; Rus, Russians; S_Os, South Ossetians; Sar, Sardinians; Shir, Shiraz; Slav, Slavs; Sp, Spanish; Syr, Syrians; Taj, Tajik; Tur, Turks; Turkm, Turkmens; Ukr, Ukrainians; Uz, Uzbeks; WAsia, West Asians. Y-STR haplotypes with South Caucasian Y-STR haplotypes than with Iranian Y-STR haplotypes. This is most evident in the network for Y-STR haplotypes on the background of haplogroup R1*(M173), in which 9 of 11 Mazandarani/Gilaki Y-STR haplotypes fall into a single cluster that connects with South Caucasian Y-STR haplotypes, and for haplogroup G*M201, in which 9 of 12 Mazandarani/Gilaki Y-STR haplotypes group with South Caucasian Y-STR haplotypes while the other three could be of either Iranian or South Caucasian origin (Figure S2). For haplogroup J2*(M172), the pattern is less clear: six Mazandarani/Gilaki Y-STR haplotypes group with South Caucasian haplotypes and three group with Iranian haplotypes, while the remaining six Figure 3. Pairwise F st and R st Values for Mazandarani/Gilaki versus South Caucasian Groups and for Mazandarani/Gilaki versus Iranian Groups Pairwise F st values based on HVI sequences within mtdna haplogroups, and R st values based on Y-STR haplotypes on the background of some Y-SNP haplogroups, for Mazandarani/Gilaki versus South Caucasian groups and for Mazandarani/Gilaki versus Iranian groups.
4 Concomitant Replacement of Language and mtdna 671 haplotypes could be of either Iranian or South Caucasian origin (Figure S2). The patterns observed in the median networks are further supported by pairwise R st comparisons for Y-STR haplotypes within each Y haplogroup (Figure 3); the Mazandarani/Gilaki are more similar to South Caucasian groups than to Iranian groups for Y-STR haplotypes within all three Y-SNP haplogroups. Nevertheless, the sample sizes in these analyses are small and thus the results could change with further sampling. However, while some contribution of Y chromosomes of Iranian origin to the Mazandarani/Gilaki cannot be excluded, overall the Y chromosome data do indicate a closer relationship of the Mazandarani and Gilaki with South Caucasian groups than with Iranian groups. Origins of the South Caspian Groups The mtdna and Y chromosome data give conflicting views on the relationships of the South Caspian groups: the mtdna data suggest that they are most closely related to their geographic and linguistic neighbors, namely other Iranian groups, whereas the Y chromosome data suggest that they are most closely related to South Caucasian groups. How can these be reconciled? One possible scenario would be an origin for the Gilaki and Mazandarani in the South Caucasus and migration to the South Caspian region, followed by the incorporation of females (but not males) from the local Iranian groups. Such preferential incorporation of females would be expected if the ancestors of the Gilaki and Mazandarani groups were patrilocal, which is the case today, and would lead to the introgression of mtdna types (but not Y chromosomes) from the local Iranian groups. Over time, the mtdna gene pool of the South Caspian groups would come to resemble that of their geographic neighbors, while the Y chromosome gene pool would still reflect their South Caucasian origin. Presumably the ancestors of the Gilaki and Mazandarani spoke a Caucasian language; therefore, their original Caucasian language must have been replaced by the local Iranian language under the above scenario. Indeed, typological analysis of the Gilaki and Mazandarani languages does indicate some sharing of features with south Caucasian languages [3, 4]. While the process of language replacement may have been completely independent of the introgression of mtdna types, it is tempting to speculate that they were in fact related: the influx of local women, speaking the local Iranian language, may have caused (or accelerated) the replacement of the original Caucasian language of the ancestors of the Gilaki and Mazandarani. Thus, the concomitant replacement of both the language and the mtdna types of the ancestors of the South Caspian groups may have been the result of the incorporation of local females, under the influence of patrilocality. How widespread is this phenomenon? Other cases of discrepancies between the mtdna and Y chromosome relationships of populations are rare, but where they do occur, the linguistic relationships do tend to reflect the mtdna relationships, rather than the Y chromosome relationships. For example, Polynesians have high frequencies of mtdna types of Asian origin and high frequencies of Y chromosomes of Melanesian origin [8, 9], and they speak Austronesian languages of Asian origin, matching their mtdna relationships. Also, Jewish populations in different geographic regions are generally characterized by similar Y chromosomes that are distinct from the Y chromosomes of their geographic neighbors, indicating a common paternal origin, but different mtdna types that are related to the mtdna types of their geographic neighbors [10]. Moreover, they speak the language of their geographic region, which also is the origin of their mtdna types exactly as in the case of the South Caspian populations. The concomitant replacement of mtdna and language after the migration of a group to a new region may thus be a more general phenomenon than previously recognized, and furthermore emphasizes the role of maternal transmission of language as a means of language replacement. Experimental Procedures Samples and DNA Extraction A total of 100 cheek cell samples from unrelated males, representing two populations from Iran (Figure 1) Mazandarani (50 samples) and Gilaki (50 samples) were collected. Genomic DNA from cheek cell swabs was extracted by a standard salting-out procedure [11]. Informed consent and information about birthplace, parents, and grandparents was obtained from all donors. mtdna Analysis The first hypervariable segment (HV1) of the mtdna control region was amplified via primers L15996 and H16410 [12], as described previously [8], and sequences for both strands of the PCR products were determined with the DNA Sequencing Kit (Perkin-Elmer), following the protocol recommended by supplier, and by an ABI 3700 automated DNA sequencer. Individuals with the C-stretch between positions and were sequenced again in each direction, so that each base was determined twice. The mtdna HVRI sequences will be published online on the authors web page at the time of publication ( pubs_stoneking.html). Published mtdna HV1 sequences were included from Caucasus, West and Central Asian, and West and East European groups [6, 13 24]. mtdna haplogroups that are most informative in southwest Asia, i.e., haplogroups H, J, N, T, and U, defined by SNPs at nucleotide positions 7025, 13704, 10671, 13366, and 12308, respectively [23], were determined by PCR-RFLP assays of the relevant SNP, as described elsewhere [23, 24]. Y Chromosome Analysis All 100 samples were typed for the X- and Y-linked zinc finger protein genes in order to confirm the gender of the sample [25]. Genotyping was carried out for ten Y chromosomal SNP markers (RPS4Y (M130), M9, M89, M124, M45, M173, M17, M201, M170, and M172 [26]) and YAP Alu insertion polymorphism [27] as described elsewhere [6, 9, 27 29]. The samples were genotyped according to the hierarchical order of the markers as described in [26]. Published Y-SNP data for Caucasian, European, West Asian, and Central Asian groups [6, 22, 30 32] were also included in some analyses. Eleven samples belonging to Y-SNP haplogroup R1*(M173), 12 samples belonging to haplogroup G*(M201), and 17 samples belonging to haplogroup J2*(M172) were genotyped for nine Y chromosome short tandem repeat (Y-STR) markers: DYS19 (DYS394), DYS385a, DYS385b, DYS389I, DYS389II, DYS390, DYS391, DYS392, and DYS393 as described elsewhere [33, 34]. The resulting Y-STR haplotypes were compared to published Y-STR haplotypes from these same haplogroups from Iran and the south Caucasus [7]. Statistical Analysis Basic parameters of molecular diversity and population genetic structure were calculated with the software package Arlequin
5 Current Biology [35].F st values for mtdna HVI sequences were calculated with the Kimura 2 parameter model, and for Y-SNP data with no molecular distance; for the Y-STR data, R st values [36] were calculated. The statistical significance of F st and R st values was estimated by permutation analysis, with 10,000 permutations. The STATISTICA package (StatSoft Inc.) was used for multidimensional scaling (MDS) analysis based on pairwise F st values for mtdna HVI and Y-SNP data sets; the default starting configuration (i.e., principle components analysis) was used, and the number of dimensions was set to two. Network analysis for Y-STR and mtdna HV1 sequence data was carried out with the software package NETWORK version 3.1 [37]. Supplemental Data Supplemental Data include two figures and two tables and can be found with this article online at cgi/content/full/16/7/668/dc1/. Acknowledgments We are grateful to the original donors for providing DNA samples. We thank Brigitte Pakendorf and Donald Stilo for useful discussion and suggestions. This research was supported by funds from the Max Planck Society, Germany. Received: November 21, 2005 Revised: January 6, 2006 Accepted: February 6, 2006 Published: April 3, 2006 References 1. Gordon, R.G., Jr. (2005). Ethnologue: Languages of the World, 15 th edition. (Dallas, TX: SIL International). Online version 2. Negahban, E.O. (2001). Gilân. In Encyclopedia Iranica, E. Yarshater, ed. (New York: Bibliotheca Persica Press), pp Stilo, D. (1981). The Tati language group in the sociolinguistic context of Northwestern Iran and Transcaucasia. Iran. Stud. 14, Stilo, D. (2005). Iranian as buffer zone between the universal typologies of Turkic and Semitic. In Linguistic Convergence and Areal Diffusion. Case Studies from Iranian, Semitic and Turkic, E.A. Csato, B. Isaksson, and C. Jahani, eds. (London: Routledge- Curzon), pp Quintana-Murci, L., Krausz, C., Zerjal, T., Sayar, S.H., Hammer, M.F., Mehdi, S.Q., Ayub, Q., Qamar, R., Mohyuddin, A., Radhakrishna, U., et al. (2001). Y-chromosome lineages trace diffusion of people and languages in southwestern Asia. Am. J. Hum. Genet. 68, Nasidze, I., Ling, E.Y.S., Quinque, D., Dupanloup, I., Cordaux, R., Rychkov, S., Naumova, O., Zhukova, O., Sarraf-Zadegan, N., Naderi, G.A., et al. (2004). Mitochondrial DNA and Y-chromosome variation in the Caucasus. Ann. Hum. Genet. 68, Nasidze, I., Schadlich, H., and Stoneking, M. (2003). Haplotypes from the Caucasus, Turkey and Iran for nine Y-STR loci. Forensic Sci. Int. 137, Redd, A.J., Takezaki, N., Sherry, S.T., McGarvey, S.T., Sofro, A.S., and Stoneking, M. (1995). Evolutionary history of the COII/tRNA-lys intergenic 9 base pair deletion in human mitochondrial DNAs from the Pacific. Mol. Biol. Evol. 12, Kayser, M., Brauer, S., Weiss, G., Underhill, P.A., Roewer, L., Schiefenhover, W., and Stoneking, M. (2000). Melanesian origin of Polynesian Y chromosome. Curr. Biol. 10, Thomas, M.G., Weale, M.E., Jones, A.L., Richards, M., Smith, A., Redhead, N., Torroni, A., Scozzari, R., Gratix, F., Tarekegn, A., et al. (2002). Founding mothers of Jewish communities: geographically separated Jewish groups were independently founded by very few female ancestors. Am. J. Hum. Genet. 70, Miller, S.A., Dykes, D.D., and Polesky, H.F. (1988). A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 16, Vigilant, L., Pennington, R., Harpending, H., Kocher, T.D., and Wilson, A.C. (1989). Mitochondrial DNA sequences in single hairs from a southern African population. Proc. Natl. Acad. Sci. USA 86, DiRienzo, A., and Wilson, A.C. (1991). Branching pattern in the evolutionary tree for human mitochondrial DNA. Proc. Natl. Acad. Sci. USA 88, Maliarchuk, B.A., Derenko, M.V., and Solovenchuk, L.L. (1995). Types of regulatory regions in mitochondrial DNA in eastern Slavs. Genetika (Russia) 31, Côrte Real, H.B., Macaulay, V.A., Richards, M.B., Hariti, G., Issad, M.S., Cambon-Thomsen, A., Papiha, S., Bertranpetit, J., and Sykes, B.C. (1996). Genetic diversity in the Iberian Peninsula determined from mitochondrial sequence analysis. Ann. Hum. Genet. 60, Richards, M., Côrte-Real, H., Forster, P., Macaulay, V., Wilkinson-Herbots, H., Demaine, A., Papiha, S., Hedges, R., Bandelt, H.-J., and Sykes, B. (1996). Paleolithic and Neolithic lineages in the European mitochondrial gene pool. Am. J. Hum. Genet. 59, Kivisild, T., Bamshad, M.J., Kaldma, K., Metspalu, M., Metspalu, E., Reidla, M., Laos, S., Parik, J., Watkins, W.S., Dixon, M.E., et al. (1999). Deep common ancestry of Indian and western- Eurasian mitochondrial DNA lineages. Curr. Biol. 9, Macaulay, V., Richards, M., Hickey, E., Vega, E., Cruciani, F., Guida, V., Scozzari, R., Bonne-Tamir, B., Sykes, B., and Torroni, A. (1999). The emerging tree of West Eurasian mtdnas: a synthesis of control-region sequences and RFLPs. Am. J. Hum. Genet. 64, Comas, D., Calafell, F., Mateu, E., Pérez-Lezaun, A., Bosch, E., Martínez-Arias, R., Clarimon, J., Facchini, F., Fiori, G., Luiselli, D., et al. (1998). Trading genes along the Silk Road: mtdna sequences and the origin of Central Asian populations. Am. J. Hum. Genet. 63, Richards, M., Macaulay, V., Hickey, E., Vega, E., Sykes, B., Guida, V., Rengo, C., Sellitto, D., Cruciani, F., Kivisild, T., et al. (2000). Tracing European founder lineages in the Near Eastern mtdna pool. Am. J. Hum. Genet. 67, Nasidze, I., and Stoneking, M. (2001). Mitochondrial DNA variation and language replacements in the Caucasus. Proc. R. Soc. Lond. B. Biol. Sci. 268, Al-Zahery, N., Semino, O., Benuzzi, G., Magri, C., Passarino, G., Torroni, A., and Sanachiara-Benerecetti, A.S. (2003). Y-chromosome and mtdna polymorphisms in Iraq, a crossroad of the early human dispersal and of post-neolithic migrations. Mol. Phylogenet. Evol. 28, Quintana-Murci, L., Chaix, R., Wells, R.S., Behar, D.M., Sayar, H., Scozzari, R., Rengo, C., Al-Zahery, N., Semino, O., Santachiara- Benerecetti, A.S., et al. (2004). Where West meets East: the complex mtdna landscape of the Southwest and Central Asian corridor. Am. J. Hum. Genet. 74, Finnilä, S., Hassinen, I.E., Ala-Kokko, L., and Majamaa, K. (2000). Phylogenetic network of the mtdna haplogroup U in northern Finland based on sequence analysis of the complete coding region by conformation-sensitive gel electrophoresis. Am. J. Hum. Genet. 66, Wilson, J., and Erlandsson, R. (1998). Sexing of human and other primate DNA. Biol. Chem. 379, Underhill, P.A., Shen, P., Lin, A.A., Jin, L., Passarino, G., Yang, W.H., Kauffman, E., Bonne-Tamir, B., Bertranpetit, J., Francalacci, P., et al. (2000). Y chromosome sequence variation and the history of human populations. Nat. Genet. 26, Hammer, M., and Horai, S. (1995). Y chromosomal DNA variation and the peopling of Japan. Am. J. Hum. Genet. 56, Cordaux, R., Aunger, R., Bentley, G., Nasidze, I., Saha, N., Sirajuddin, S.M., and Stoneking, M. (2004). Independent origins of Indian caste and tribal paternal lineages. Curr. Biol. 14, Ke, Y., Su, B., Song, X., Lu, D., Chen, L., Li, H., Qi, C., Marzuki, S., Deka, R., Underhill, P., et al. (2001). African origin of modern humans in East Asia: a tale of 12,000 Y chromosomes. Science 292, Semino, O., Passarino, G., Oefner, P.J., Lin, A.A., Arbuzova, S., Beckman, L.E., De Benedictis, G., Francalacci, P., Kouvatsi, A., Limborska, S., et al. (2000). The genetic legacy of Paleolithic
6 Concomitant Replacement of Language and mtdna 673 Homo sapiens sapiens in extant Europeans: a Y chromosome perspective. Science 290, Wells, R.S., Yuldasheva, N., Ruzibakiev, R., Underhill, P.A., Evseeva, I., Blue-Smith, J., Jin, L., Su, B., Pitchappan, R., Shanmugalakshimi, S., et al. (2001). The Eurasian heartland: a continental perspective on Y-chromosome diversity. Proc. Natl. Acad. Sci. USA 98, Nasidze, I., Sarkisian, T., Kerimov, A., and Stoneking, M. (2003). Testing hypotheses of language replacement in the Caucasus: evidence from the Y-chromosome. Hum. Genet. 112, Kayser, M., Caglia, A., Corach, D., Fretwell, N., Gehrig, C., Graziosi, G., Heidorn, F., Herrmann, S., Herzog, B., Hidding, M., et al. (1997). Evaluation of Y chromosomal STRs: a multicenter study. Int. J. Legal Med. 110, Kittler, R., Erler, A., Brauer, S., Stoneking, M., and Kayser, M. (2003). Apparent intrachromosomal exchange on the human Y chromosome explained by population history. Eur. J. Hum. Genet. 11, Schneider, S., Roessli, D., and Excoffier, L. (2000). Arelquin version 2.000: A software for population genetics data analysis. University of Geneva, Switzerland: Genetics and Biometry. 36. Slatkin, M. (1995). A measure of population subdivision based on microsatellite allele frequencies. Genetics 139, Bandelt, H.J., Forster, P., and Röhl, A. (1999). Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 16,
Genetic Variation and Human Evolution Lynn B. Jorde, Ph.D. Department of Human Genetics University of Utah School of Medicine.
 Genetic Variation and Human Evolution Lynn B. Jorde, Ph.D. Department of Human Genetics University of Utah School of Medicine. The past two decades have witnessed an explosion of human genetic data. Innumerable
Genetic Variation and Human Evolution Lynn B. Jorde, Ph.D. Department of Human Genetics University of Utah School of Medicine. The past two decades have witnessed an explosion of human genetic data. Innumerable
Testing hypotheses of language replacement in the Caucasus: evidence from the Y-chromosome
 Hum Genet (2003) 112 : 255 261 DOI 10.1007/s00439-002-0874-4 ORIGINAL INVESTIGATION Ivan Nasidze Tamara Sarkisian Azer Kerimov Mark Stoneking Testing hypotheses of language replacement in the Caucasus:
Hum Genet (2003) 112 : 255 261 DOI 10.1007/s00439-002-0874-4 ORIGINAL INVESTIGATION Ivan Nasidze Tamara Sarkisian Azer Kerimov Mark Stoneking Testing hypotheses of language replacement in the Caucasus:
Y Chromosome Markers
 Y Chromosome Markers Lineage Markers Autosomal chromosomes recombine with each meiosis Y and Mitochondrial DNA does not This means that the Y and mtdna remains constant from generation to generation Except
Y Chromosome Markers Lineage Markers Autosomal chromosomes recombine with each meiosis Y and Mitochondrial DNA does not This means that the Y and mtdna remains constant from generation to generation Except
Title:Global patterns of sex-biased migrations in humans
 Title:Global patterns of sex-biased migrations in humans Authors:Chuan-Chao Wang 1, Li Jin 1, 2, 3, Hui Li 1,* Affiliations: 1. State Key Laboratory of Genetic Engineering and MOE Key Laboratory of Contemporary
Title:Global patterns of sex-biased migrations in humans Authors:Chuan-Chao Wang 1, Li Jin 1, 2, 3, Hui Li 1,* Affiliations: 1. State Key Laboratory of Genetic Engineering and MOE Key Laboratory of Contemporary
Elsevier Editorial System(tm) for Forensic Science International: Genetics Manuscript Draft
 Elsevier Editorial System(tm) for Forensic Science International: Genetics Manuscript Draft Manuscript Number: Title: A comment on the Paper: A comparison of Y-chromosomal lineage dating using either resequencing
Elsevier Editorial System(tm) for Forensic Science International: Genetics Manuscript Draft Manuscript Number: Title: A comment on the Paper: A comparison of Y-chromosomal lineage dating using either resequencing
Comparison of Y-chromosomal lineage dating using either evolutionary
 Comparison of Y-chromosomal lineage dating using either evolutionary or genealogical Y-STR mutation rates Chuan-Chao Wang 1, Hui Li 1,* 1 State Key Laboratory of Genetic Engineering and MOE Key Laboratory
Comparison of Y-chromosomal lineage dating using either evolutionary or genealogical Y-STR mutation rates Chuan-Chao Wang 1, Hui Li 1,* 1 State Key Laboratory of Genetic Engineering and MOE Key Laboratory
Analysis of 16 Y STR loci in the Finnish population reveals a local reduction in the diversity of male lineages
 Forensic Science International 142 (2004) 37 43 Analysis of 16 Y STR loci in the Finnish population reveals a local reduction in the diversity of male lineages M. Hedman a, V. Pimenoff a, M. Lukka b, P.
Forensic Science International 142 (2004) 37 43 Analysis of 16 Y STR loci in the Finnish population reveals a local reduction in the diversity of male lineages M. Hedman a, V. Pimenoff a, M. Lukka b, P.
D-Loop-BASE is online now Central European database of mitochondrial DNA
 International Congress Series 1239 (2003) 505 509 D-Loop-BASE is online now Central European database of mitochondrial DNA Holger Wittig a, *, Mike Koecke b, Kai-Uwe Sattler b, Dieter Krause a a Institute
International Congress Series 1239 (2003) 505 509 D-Loop-BASE is online now Central European database of mitochondrial DNA Holger Wittig a, *, Mike Koecke b, Kai-Uwe Sattler b, Dieter Krause a a Institute
The phylogeography of Y chromosome binary haplotypes and the origins of modern human populations
 Ann. Hum. Genet. (2001), 65, 43 62 Printed in Great Britain 43 The phylogeography of Y chromosome binary haplotypes and the origins of modern human populations P. A. UNDERHILL *, G. PASSARINO,, A.A.LIN,
Ann. Hum. Genet. (2001), 65, 43 62 Printed in Great Britain 43 The phylogeography of Y chromosome binary haplotypes and the origins of modern human populations P. A. UNDERHILL *, G. PASSARINO,, A.A.LIN,
Y-STR haplotype diversity and population data for Central Brazil: implications for environmental forensics and paternity testing
 Short Communication Y-STR haplotype diversity and population data for Central Brazil: implications for environmental forensics and paternity testing T.C. Vieira 1,2,3,4, M.A.D. Gigonzac 2,3,4, D.M. Silva
Short Communication Y-STR haplotype diversity and population data for Central Brazil: implications for environmental forensics and paternity testing T.C. Vieira 1,2,3,4, M.A.D. Gigonzac 2,3,4, D.M. Silva
Mitochondrial DNA Analysis
 Mitochondrial DNA Analysis Lineage Markers Lineage markers are passed down from generation to generation without changing Except for rare mutation events They can help determine the lineage (family tree)
Mitochondrial DNA Analysis Lineage Markers Lineage markers are passed down from generation to generation without changing Except for rare mutation events They can help determine the lineage (family tree)
Institute of Biological Problems of the North, Far-East Branch of the Russian Academy of
 MBE Advance Access published May 13, 2008 Research Article: Mitochondrial DNA phylogeny in Eastern and Western Slavs B. Malyarchuk 1,*, T. Grzybowski 2, M. Derenko 1, M. Perkova 1, T. Vanecek 3, J. Lazur
MBE Advance Access published May 13, 2008 Research Article: Mitochondrial DNA phylogeny in Eastern and Western Slavs B. Malyarchuk 1,*, T. Grzybowski 2, M. Derenko 1, M. Perkova 1, T. Vanecek 3, J. Lazur
Online Y-chromosomal Short Tandem Repeat Haplotype Reference Database (YHRD) for U.S. Populations*
 Manfred Kayser, 1 Ph.D.; Silke Brauer; 1 Sascha Willuweit; 2 Hiltrud Schädlich, 1 Ph.D.; Mark A. Batzer, 3 Ph.D.; Jennifer Zawacki; 4 Mechthild Prinz, 4 M.D.; Lutz Roewer, 2 Ph.D.; and Mark Stoneking,
Manfred Kayser, 1 Ph.D.; Silke Brauer; 1 Sascha Willuweit; 2 Hiltrud Schädlich, 1 Ph.D.; Mark A. Batzer, 3 Ph.D.; Jennifer Zawacki; 4 Mechthild Prinz, 4 M.D.; Lutz Roewer, 2 Ph.D.; and Mark Stoneking,
Forensic DNA Testing Terminology
 Forensic DNA Testing Terminology ABI 310 Genetic Analyzer a capillary electrophoresis instrument used by forensic DNA laboratories to separate short tandem repeat (STR) loci on the basis of their size.
Forensic DNA Testing Terminology ABI 310 Genetic Analyzer a capillary electrophoresis instrument used by forensic DNA laboratories to separate short tandem repeat (STR) loci on the basis of their size.
Development of two Novel DNA Analysis methods to Improve Workflow Efficiency for Challenging Forensic Samples
 Development of two Novel DNA Analysis methods to Improve Workflow Efficiency for Challenging Forensic Samples Sudhir K. Sinha, Ph.D.*, Anne H. Montgomery, M.S., Gina Pineda, M.S., and Hiromi Brown, Ph.D.
Development of two Novel DNA Analysis methods to Improve Workflow Efficiency for Challenging Forensic Samples Sudhir K. Sinha, Ph.D.*, Anne H. Montgomery, M.S., Gina Pineda, M.S., and Hiromi Brown, Ph.D.
Paternity Testing. Chapter 23
 Paternity Testing Chapter 23 Kinship and Paternity DNA analysis can also be used for: Kinship testing determining whether individuals are related Paternity testing determining the father of a child Missing
Paternity Testing Chapter 23 Kinship and Paternity DNA analysis can also be used for: Kinship testing determining whether individuals are related Paternity testing determining the father of a child Missing
The sample is taken with a simple mouth swab no blood is involved. There will be instructions included on how to take the sample.
 DNA testing Thanks for your enquiry about DNA testing. I oversee the Scottish DNA Project on behalf of the University of Strathclyde, Glasgow and act as representative for Family Tree DNA who host our
DNA testing Thanks for your enquiry about DNA testing. I oversee the Scottish DNA Project on behalf of the University of Strathclyde, Glasgow and act as representative for Family Tree DNA who host our
Rethinking Polynesian Origins: Human Settlement of the Pacific
 LENScience Senior Biology Seminar Series Rethinking Polynesian Origins: Human Settlement of the Pacific Michal Denny, and Lisa Matisoo-Smith Our Polynesian ancestors are renowned as some of the world s
LENScience Senior Biology Seminar Series Rethinking Polynesian Origins: Human Settlement of the Pacific Michal Denny, and Lisa Matisoo-Smith Our Polynesian ancestors are renowned as some of the world s
Seeing Faces and History through Human Genome Sequences
 Seeing Faces and History through Human Genome Sequences CAS/MPG Partner Group on the Human Functional Genetic Variations Shanghai-Leipzig, 2011.2.1 2016.1.31 Prof. Dr. TANG Kun (middle) with his cooperator,
Seeing Faces and History through Human Genome Sequences CAS/MPG Partner Group on the Human Functional Genetic Variations Shanghai-Leipzig, 2011.2.1 2016.1.31 Prof. Dr. TANG Kun (middle) with his cooperator,
Chapter 8: Recombinant DNA 2002 by W. H. Freeman and Company Chapter 8: Recombinant DNA 2002 by W. H. Freeman and Company
 Genetic engineering: humans Gene replacement therapy or gene therapy Many technical and ethical issues implications for gene pool for germ-line gene therapy what traits constitute disease rather than just
Genetic engineering: humans Gene replacement therapy or gene therapy Many technical and ethical issues implications for gene pool for germ-line gene therapy what traits constitute disease rather than just
A Primer of Genome Science THIRD
 A Primer of Genome Science THIRD EDITION GREG GIBSON-SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc. Publishers Sunderland, Massachusetts USA Contents Preface xi 1 Genome Projects:
A Primer of Genome Science THIRD EDITION GREG GIBSON-SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc. Publishers Sunderland, Massachusetts USA Contents Preface xi 1 Genome Projects:
Gene Mapping Techniques
 Gene Mapping Techniques OBJECTIVES By the end of this session the student should be able to: Define genetic linkage and recombinant frequency State how genetic distance may be estimated State how restriction
Gene Mapping Techniques OBJECTIVES By the end of this session the student should be able to: Define genetic linkage and recombinant frequency State how genetic distance may be estimated State how restriction
Reduced-Median-Network Analysis of Complete Mitochondrial DNA Coding-Region Sequences for the Major African, Asian, and European Haplogroups
 Am. J. Hum. Genet. 70:1152 1171, 2002 Reduced-Median-Network Analysis of Complete Mitochondrial DNA Coding-Region Sequences for the Major African, Asian, and European Haplogroups Corinna Herrnstadt, 1
Am. J. Hum. Genet. 70:1152 1171, 2002 Reduced-Median-Network Analysis of Complete Mitochondrial DNA Coding-Region Sequences for the Major African, Asian, and European Haplogroups Corinna Herrnstadt, 1
Contrasting patterns of Y chromosome variation in Ashkenazi Jewish and host non-jewish European populations
 Hum Genet (2004) 114 : 354 365 DOI 10.1007/s00439-003-1073-7 354 ORIGINAL INVESTIGATION Doron M. Behar Daniel Garrigan Matthew E. Kaplan Zahra Moasher Dror Rosengarten Tatiana M. Karafet Lluis Quintana-Murci
Hum Genet (2004) 114 : 354 365 DOI 10.1007/s00439-003-1073-7 354 ORIGINAL INVESTIGATION Doron M. Behar Daniel Garrigan Matthew E. Kaplan Zahra Moasher Dror Rosengarten Tatiana M. Karafet Lluis Quintana-Murci
Thesis Guidelines Handbook
 Thesis Guidelines Handbook For For MS Program COMSATS Institute of Information Technology Guidelines for MS Thesis These guidelines for MS thesis should be read in conjunction with the GHB. Please read
Thesis Guidelines Handbook For For MS Program COMSATS Institute of Information Technology Guidelines for MS Thesis These guidelines for MS thesis should be read in conjunction with the GHB. Please read
Association between Dopamine Gene and Alcoholism in Pategar Community of Dharwad, Karnataka
 International Journal of Scientific and Research Publications, Volume 3, Issue 10, October 2013 1 Association between Dopamine Gene and Alcoholism in Pategar Community of Dharwad, Karnataka SOMASHEKHAR
International Journal of Scientific and Research Publications, Volume 3, Issue 10, October 2013 1 Association between Dopamine Gene and Alcoholism in Pategar Community of Dharwad, Karnataka SOMASHEKHAR
DNA and Forensic Science
 DNA and Forensic Science Micah A. Luftig * Stephen Richey ** I. INTRODUCTION This paper represents a discussion of the fundamental principles of DNA technology as it applies to forensic testing. A brief
DNA and Forensic Science Micah A. Luftig * Stephen Richey ** I. INTRODUCTION This paper represents a discussion of the fundamental principles of DNA technology as it applies to forensic testing. A brief
Significant genetic differentiation between Poland and Germany follows present-day political borders, as revealed by Y-chromosome analysis
 Hum Genet (2005) 117: 428 443 DOI 10.1007/s00439-005-1333-9 ORIGINAL INVESTIGATION Manfred Kayser Æ Oscar Lao Æ Katja Anslinger Christa Augustin Æ Grazyna Bargel Æ Jeanett Edelmann Sahar Elias Æ Marielle
Hum Genet (2005) 117: 428 443 DOI 10.1007/s00439-005-1333-9 ORIGINAL INVESTIGATION Manfred Kayser Æ Oscar Lao Æ Katja Anslinger Christa Augustin Æ Grazyna Bargel Æ Jeanett Edelmann Sahar Elias Æ Marielle
Molecular typing of VTEC: from PFGE to NGS-based phylogeny
 Molecular typing of VTEC: from PFGE to NGS-based phylogeny Valeria Michelacci 10th Annual Workshop of the National Reference Laboratories for E. coli in the EU Rome, November 5 th 2015 Molecular typing
Molecular typing of VTEC: from PFGE to NGS-based phylogeny Valeria Michelacci 10th Annual Workshop of the National Reference Laboratories for E. coli in the EU Rome, November 5 th 2015 Molecular typing
Some ancestors are difficult to trace. Adoptions illegitimacies, name changes and migrations can present brick walls in our research that seem
 Some ancestors are difficult to trace. Adoptions illegitimacies, name changes and migrations can present brick walls in our research that seem impregnable. In other cases, documentation that would shed
Some ancestors are difficult to trace. Adoptions illegitimacies, name changes and migrations can present brick walls in our research that seem impregnable. In other cases, documentation that would shed
Globally, about 9.7% of cancers in men are prostate cancers, and the risk of developing the
 Chapter 5 Analysis of Prostate Cancer Association Study Data 5.1 Risk factors for Prostate Cancer Globally, about 9.7% of cancers in men are prostate cancers, and the risk of developing the disease has
Chapter 5 Analysis of Prostate Cancer Association Study Data 5.1 Risk factors for Prostate Cancer Globally, about 9.7% of cancers in men are prostate cancers, and the risk of developing the disease has
A Nomenclature System for the Tree of Human Y-Chromosomal Binary Haplogroups
 Resource A Nomenclature System for the Tree of Human Y-Chromosomal Binary Haplogroups The Y Chromosome Consortium 1 The Y chromosome contains the largest nonrecombining block in the human genome. By virtue
Resource A Nomenclature System for the Tree of Human Y-Chromosomal Binary Haplogroups The Y Chromosome Consortium 1 The Y chromosome contains the largest nonrecombining block in the human genome. By virtue
DNA Genealogy, Mutation Rates, and Some Historical Evidences Written in Y-Chromosome. I. Basic Principles and the Method
 DNA Genealogy, Mutation Rates, and Some Historical Evidences Written in Y-Chromosome. I. Basic Principles and the Method Anatole A. Klyosov 1 Abstract Origin of peoples in a context of DNA genealogy is
DNA Genealogy, Mutation Rates, and Some Historical Evidences Written in Y-Chromosome. I. Basic Principles and the Method Anatole A. Klyosov 1 Abstract Origin of peoples in a context of DNA genealogy is
Activity IT S ALL RELATIVES The Role of DNA Evidence in Forensic Investigations
 Activity IT S ALL RELATIVES The Role of DNA Evidence in Forensic Investigations SCENARIO You have responded, as a result of a call from the police to the Coroner s Office, to the scene of the death of
Activity IT S ALL RELATIVES The Role of DNA Evidence in Forensic Investigations SCENARIO You have responded, as a result of a call from the police to the Coroner s Office, to the scene of the death of
Single Nucleotide Polymorphisms (SNPs)
 Single Nucleotide Polymorphisms (SNPs) Additional Markers 13 core STR loci Obtain further information from additional markers: Y STRs Separating male samples Mitochondrial DNA Working with extremely degraded
Single Nucleotide Polymorphisms (SNPs) Additional Markers 13 core STR loci Obtain further information from additional markers: Y STRs Separating male samples Mitochondrial DNA Working with extremely degraded
I Have the Results of My Genetic Genealogy Test, Now What?
 I Have the Results of My Genetic Genealogy Test, Now What? Version 2.1 1 I Have the Results of My Genetic Genealogy Test, Now What? Chapter 1: What Is (And Isn t) Genetic Genealogy? Chapter 2: How Do I
I Have the Results of My Genetic Genealogy Test, Now What? Version 2.1 1 I Have the Results of My Genetic Genealogy Test, Now What? Chapter 1: What Is (And Isn t) Genetic Genealogy? Chapter 2: How Do I
HLA data analysis in anthropology: basic theory and practice
 HLA data analysis in anthropology: basic theory and practice Alicia Sanchez-Mazas and José Manuel Nunes Laboratory of Anthropology, Genetics and Peopling history (AGP), Department of Anthropology and Ecology,
HLA data analysis in anthropology: basic theory and practice Alicia Sanchez-Mazas and José Manuel Nunes Laboratory of Anthropology, Genetics and Peopling history (AGP), Department of Anthropology and Ecology,
Estimating Scandinavian and Gaelic Ancestry in the Male Settlers of Iceland
 Am. J. Hum. Genet. 67:697 717, 2000 Estimating Scandinavian and Gaelic Ancestry in the Male Settlers of Iceland Agnar Helgason, 1 Sigrún Sigurðardóttir, 3 Jayne Nicholson, 2 Bryan Sykes, 2 Emmeline W.
Am. J. Hum. Genet. 67:697 717, 2000 Estimating Scandinavian and Gaelic Ancestry in the Male Settlers of Iceland Agnar Helgason, 1 Sigrún Sigurðardóttir, 3 Jayne Nicholson, 2 Bryan Sykes, 2 Emmeline W.
Commonly Used STR Markers
 Commonly Used STR Markers Repeats Satellites 100 to 1000 bases repeated Minisatellites VNTR variable number tandem repeat 10 to 100 bases repeated Microsatellites STR short tandem repeat 2 to 6 bases repeated
Commonly Used STR Markers Repeats Satellites 100 to 1000 bases repeated Minisatellites VNTR variable number tandem repeat 10 to 100 bases repeated Microsatellites STR short tandem repeat 2 to 6 bases repeated
Title:Natural selection on human Y chromosomes
 Title:Natural selection on human Y chromosomes Authors:Chuan-Chao Wang 1, Li Jin 1, 2, 3, Hui Li 1,* Affiliations: 1. State Key Laboratory of Genetic Engineering and MOE Key Laboratory of Contemporary
Title:Natural selection on human Y chromosomes Authors:Chuan-Chao Wang 1, Li Jin 1, 2, 3, Hui Li 1,* Affiliations: 1. State Key Laboratory of Genetic Engineering and MOE Key Laboratory of Contemporary
Biology Behind the Crime Scene Week 4: Lab #4 Genetics Exercise (Meiosis) and RFLP Analysis of DNA
 Page 1 of 5 Biology Behind the Crime Scene Week 4: Lab #4 Genetics Exercise (Meiosis) and RFLP Analysis of DNA Genetics Exercise: Understanding how meiosis affects genetic inheritance and DNA patterns
Page 1 of 5 Biology Behind the Crime Scene Week 4: Lab #4 Genetics Exercise (Meiosis) and RFLP Analysis of DNA Genetics Exercise: Understanding how meiosis affects genetic inheritance and DNA patterns
DNA Testing for Genealogy - What Can It Do For You??
 DNA Testing for Genealogy - What Can It Do For You?? Paper courtesy of Roberta Estes, www.dnaexplain.com, e-mail Roberta at Roberta@dnaexplain.com. Graphics courtesy of Family Tree DNA, www.familytreedna.com.
DNA Testing for Genealogy - What Can It Do For You?? Paper courtesy of Roberta Estes, www.dnaexplain.com, e-mail Roberta at Roberta@dnaexplain.com. Graphics courtesy of Family Tree DNA, www.familytreedna.com.
DnaSP, DNA polymorphism analyses by the coalescent and other methods.
 DnaSP, DNA polymorphism analyses by the coalescent and other methods. Author affiliation: Julio Rozas 1, *, Juan C. Sánchez-DelBarrio 2,3, Xavier Messeguer 2 and Ricardo Rozas 1 1 Departament de Genètica,
DnaSP, DNA polymorphism analyses by the coalescent and other methods. Author affiliation: Julio Rozas 1, *, Juan C. Sánchez-DelBarrio 2,3, Xavier Messeguer 2 and Ricardo Rozas 1 1 Departament de Genètica,
Annex to the Accreditation Certificate D-PL-13372-01-00 according to DIN EN ISO/IEC 17025:2005
 Deutsche Akkreditierungsstelle GmbH German Accreditation Body Annex to the Accreditation Certificate D-PL-13372-01-00 according to DIN EN ISO/IEC 17025:2005 Period of validity: 26.03.2012 to 25.03.2017
Deutsche Akkreditierungsstelle GmbH German Accreditation Body Annex to the Accreditation Certificate D-PL-13372-01-00 according to DIN EN ISO/IEC 17025:2005 Period of validity: 26.03.2012 to 25.03.2017
From Africa to Aotearoa Part 1: Out of Africa
 From Africa to Aotearoa Part 1: Out of Africa The spread of modern humans out of Africa started around 65,000 years ago, and ended with the settlement of New Zealand 750 years ago. These PowerPoint presentations
From Africa to Aotearoa Part 1: Out of Africa The spread of modern humans out of Africa started around 65,000 years ago, and ended with the settlement of New Zealand 750 years ago. These PowerPoint presentations
A guide to the analysis of KASP genotyping data using cluster plots
 extraction sequencing genotyping extraction sequencing genotyping extraction sequencing genotyping extraction sequencing A guide to the analysis of KASP genotyping data using cluster plots Contents of
extraction sequencing genotyping extraction sequencing genotyping extraction sequencing genotyping extraction sequencing A guide to the analysis of KASP genotyping data using cluster plots Contents of
ONLINE SUPPLEMENTAL MATERIAL. Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue
 ONLINE SUPPLEMENTAL MATERIAL Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue Sungmi Park 1, Ko-Ting Lu 1, Xuebo Liu 1, Tapan K. Chatterjee 2, Steven M. Rudich 3, Neal
ONLINE SUPPLEMENTAL MATERIAL Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue Sungmi Park 1, Ko-Ting Lu 1, Xuebo Liu 1, Tapan K. Chatterjee 2, Steven M. Rudich 3, Neal
Human Genome and Human Genome Project. Louxin Zhang
 Human Genome and Human Genome Project Louxin Zhang A Primer to Genomics Cells are the fundamental working units of every living systems. DNA is made of 4 nucleotide bases. The DNA sequence is the particular
Human Genome and Human Genome Project Louxin Zhang A Primer to Genomics Cells are the fundamental working units of every living systems. DNA is made of 4 nucleotide bases. The DNA sequence is the particular
Innovations in Molecular Epidemiology
 Innovations in Molecular Epidemiology Molecular Epidemiology Measure current rates of active transmission Determine whether recurrent tuberculosis is attributable to exogenous reinfection Determine whether
Innovations in Molecular Epidemiology Molecular Epidemiology Measure current rates of active transmission Determine whether recurrent tuberculosis is attributable to exogenous reinfection Determine whether
Detailed mtdna Genotypes Permit a Reassessment of the Settlement and Population Structure of the Andaman Islands
 AMERICAN JOURNAL OF PHYSICAL ANTHROPOLOGY 136:19 27 (2008) Detailed mtdna Genotypes Permit a Reassessment of the Settlement and Population Structure of the Andaman Islands S.S. Barik, 1 R. Sahani, 1 B.V.R.
AMERICAN JOURNAL OF PHYSICAL ANTHROPOLOGY 136:19 27 (2008) Detailed mtdna Genotypes Permit a Reassessment of the Settlement and Population Structure of the Andaman Islands S.S. Barik, 1 R. Sahani, 1 B.V.R.
Online reference database of European Y-chromosomal short tandem repeat STR) haplotypes
 Forensic Science International 118 2001) 106±113 Online reference database of European Y-chromosomal short tandem repeat STR) haplotypes L. Roewer a,*, M. Krawczak b, S. Willuweit a, M. Nagy a, C. Alves
Forensic Science International 118 2001) 106±113 Online reference database of European Y-chromosomal short tandem repeat STR) haplotypes L. Roewer a,*, M. Krawczak b, S. Willuweit a, M. Nagy a, C. Alves
Archaeogenetics: DNA and the population prehistory of Europe
 McDONALD INSTITUTE MONOGRAPHS Archaeogenetics: DNA and the population prehistory of Europe Edited by Colin Renfrew & Katie Boyle Published by: McDonald Institute for Archaeological Research University
McDONALD INSTITUTE MONOGRAPHS Archaeogenetics: DNA and the population prehistory of Europe Edited by Colin Renfrew & Katie Boyle Published by: McDonald Institute for Archaeological Research University
MCB41: Second Midterm Spring 2009
 MCB41: Second Midterm Spring 2009 Before you start, print your name and student identification number (S.I.D) at the top of each page. There are 7 pages including this page. You will have 50 minutes for
MCB41: Second Midterm Spring 2009 Before you start, print your name and student identification number (S.I.D) at the top of each page. There are 7 pages including this page. You will have 50 minutes for
Milk protein genetic variation in Butana cattle
 Milk protein genetic variation in Butana cattle Ammar Said Ahmed Züchtungsbiologie und molekulare Genetik, Humboldt Universität zu Berlin, Invalidenstraβe 42, 10115 Berlin, Deutschland 1 Outline Background
Milk protein genetic variation in Butana cattle Ammar Said Ahmed Züchtungsbiologie und molekulare Genetik, Humboldt Universität zu Berlin, Invalidenstraβe 42, 10115 Berlin, Deutschland 1 Outline Background
Maximum-Likelihood Estimation of Phylogeny from DNA Sequences When Substitution Rates Differ over Sites1
 Maximum-Likelihood Estimation of Phylogeny from DNA Sequences When Substitution Rates Differ over Sites1 Ziheng Yang Department of Animal Science, Beijing Agricultural University Felsenstein s maximum-likelihood
Maximum-Likelihood Estimation of Phylogeny from DNA Sequences When Substitution Rates Differ over Sites1 Ziheng Yang Department of Animal Science, Beijing Agricultural University Felsenstein s maximum-likelihood
14.3 Studying the Human Genome
 14.3 Studying the Human Genome Lesson Objectives Summarize the methods of DNA analysis. State the goals of the Human Genome Project and explain what we have learned so far. Lesson Summary Manipulating
14.3 Studying the Human Genome Lesson Objectives Summarize the methods of DNA analysis. State the goals of the Human Genome Project and explain what we have learned so far. Lesson Summary Manipulating
SeattleSNPs Interactive Tutorial: Web Tools for Site Selection, Linkage Disequilibrium and Haplotype Analysis
 SeattleSNPs Interactive Tutorial: Web Tools for Site Selection, Linkage Disequilibrium and Haplotype Analysis Goal: This tutorial introduces several websites and tools useful for determining linkage disequilibrium
SeattleSNPs Interactive Tutorial: Web Tools for Site Selection, Linkage Disequilibrium and Haplotype Analysis Goal: This tutorial introduces several websites and tools useful for determining linkage disequilibrium
Forensic Science International: Genetics
 Forensic Science International: Genetics 3 (2009) e111 e116 Contents lists available at ScienceDirect Forensic Science International: Genetics journal homepage: www.elsevier.com/locate/fsig Announcement
Forensic Science International: Genetics 3 (2009) e111 e116 Contents lists available at ScienceDirect Forensic Science International: Genetics journal homepage: www.elsevier.com/locate/fsig Announcement
DNA Insertions and Deletions in the Human Genome. Philipp W. Messer
 DNA Insertions and Deletions in the Human Genome Philipp W. Messer Genetic Variation CGACAATAGCGCTCTTACTACGTGTATCG : : CGACAATGGCGCT---ACTACGTGCATCG 1. Nucleotide mutations 2. Genomic rearrangements 3.
DNA Insertions and Deletions in the Human Genome Philipp W. Messer Genetic Variation CGACAATAGCGCTCTTACTACGTGTATCG : : CGACAATGGCGCT---ACTACGTGCATCG 1. Nucleotide mutations 2. Genomic rearrangements 3.
Journal of Statistical Software
 JSS Journal of Statistical Software October 2006, Volume 16, Code Snippet 3. http://www.jstatsoft.org/ LDheatmap: An R Function for Graphical Display of Pairwise Linkage Disequilibria between Single Nucleotide
JSS Journal of Statistical Software October 2006, Volume 16, Code Snippet 3. http://www.jstatsoft.org/ LDheatmap: An R Function for Graphical Display of Pairwise Linkage Disequilibria between Single Nucleotide
Identifying Genetic Traces of Historical Expansions: Phoenician Footprints in the Mediterranean
 Identifying Genetic Traces of Historical Expansions: Phoenician Footprints in the Mediterranean REPORT Pierre A. Zalloua, 1,2,13 Daniel E. Platt, 3,13 Mirvat El Sibai, 1 Jade Khalife, 1 Nadine Makhoul,
Identifying Genetic Traces of Historical Expansions: Phoenician Footprints in the Mediterranean REPORT Pierre A. Zalloua, 1,2,13 Daniel E. Platt, 3,13 Mirvat El Sibai, 1 Jade Khalife, 1 Nadine Makhoul,
2. True or False? The sequence of nucleotides in the human genome is 90.9% identical from one person to the next. False (it s 99.
 1. True or False? A typical chromosome can contain several hundred to several thousand genes, arranged in linear order along the DNA molecule present in the chromosome. True 2. True or False? The sequence
1. True or False? A typical chromosome can contain several hundred to several thousand genes, arranged in linear order along the DNA molecule present in the chromosome. True 2. True or False? The sequence
RETRIEVING SEQUENCE INFORMATION. Nucleotide sequence databases. Database search. Sequence alignment and comparison
 RETRIEVING SEQUENCE INFORMATION Nucleotide sequence databases Database search Sequence alignment and comparison Biological sequence databases Originally just a storage place for sequences. Currently the
RETRIEVING SEQUENCE INFORMATION Nucleotide sequence databases Database search Sequence alignment and comparison Biological sequence databases Originally just a storage place for sequences. Currently the
The Story of Human Evolution Part 1: From ape-like ancestors to modern humans
 The Story of Human Evolution Part 1: From ape-like ancestors to modern humans Slide 1 The Story of Human Evolution This powerpoint presentation tells the story of who we are and where we came from - how
The Story of Human Evolution Part 1: From ape-like ancestors to modern humans Slide 1 The Story of Human Evolution This powerpoint presentation tells the story of who we are and where we came from - how
REPORT An African American Paternal Lineage Adds an Extremely Ancient Root to the Human Y Chromosome Phylogenetic Tree
 REPORT An African American Paternal Lineage Adds an Extremely Ancient Root to the Human Y Chromosome Phylogenetic Tree Fernando L. Mendez, 1 Thomas Krahn, 2 Bonnie Schrack, 2 Astrid-Maria Krahn, 2 Krishna
REPORT An African American Paternal Lineage Adds an Extremely Ancient Root to the Human Y Chromosome Phylogenetic Tree Fernando L. Mendez, 1 Thomas Krahn, 2 Bonnie Schrack, 2 Astrid-Maria Krahn, 2 Krishna
Bachelor of Science Degree Requirements for Students Fulfilling the REVISED General Education Curriculum
 General College Requirements Bachelor of Science Degree Requirements for Students Fulfilling the REVISED General Education Curriculum Note: see the Middle Childhood Education (MCE) curriculum sheet for
General College Requirements Bachelor of Science Degree Requirements for Students Fulfilling the REVISED General Education Curriculum Note: see the Middle Childhood Education (MCE) curriculum sheet for
Who are the Other ethnic groups?
 Article Who are the Other ethnic groups? Social and Welfare David Gardener Helen Connolly October 2005 Crown copyright Office for National Statistics 1 Drummond Gate London SW1V 2QQ Tel: 020 7533 9233
Article Who are the Other ethnic groups? Social and Welfare David Gardener Helen Connolly October 2005 Crown copyright Office for National Statistics 1 Drummond Gate London SW1V 2QQ Tel: 020 7533 9233
MATCH Commun. Math. Comput. Chem. 61 (2009) 781-788
 MATCH Communications in Mathematical and in Computer Chemistry MATCH Commun. Math. Comput. Chem. 61 (2009) 781-788 ISSN 0340-6253 Three distances for rapid similarity analysis of DNA sequences Wei Chen,
MATCH Communications in Mathematical and in Computer Chemistry MATCH Commun. Math. Comput. Chem. 61 (2009) 781-788 ISSN 0340-6253 Three distances for rapid similarity analysis of DNA sequences Wei Chen,
Network Protocol Analysis using Bioinformatics Algorithms
 Network Protocol Analysis using Bioinformatics Algorithms Marshall A. Beddoe Marshall_Beddoe@McAfee.com ABSTRACT Network protocol analysis is currently performed by hand using only intuition and a protocol
Network Protocol Analysis using Bioinformatics Algorithms Marshall A. Beddoe Marshall_Beddoe@McAfee.com ABSTRACT Network protocol analysis is currently performed by hand using only intuition and a protocol
HUMAN BIOLOGY AND SOCIETY
 HUMAN BIOLOGY AND SOCIETY HBS, B.A. premajor HBS, B.S. premajor (36-41 units) (91-105 units) Society and Genetics 5, M71A or M72A Society and Genetics 5, M71A or M72A Anthropology 7 Anthropology 7 Two
HUMAN BIOLOGY AND SOCIETY HBS, B.A. premajor HBS, B.S. premajor (36-41 units) (91-105 units) Society and Genetics 5, M71A or M72A Society and Genetics 5, M71A or M72A Anthropology 7 Anthropology 7 Two
Original article: DXS10079, DXS10074 and DXS10075: new alleles and SNP occurrence
 EXCLI Journal 2007;6:177-182 ISSN 1611-2156 Received: December 14, 2006, accepted: June 21, 2007, published: June 26, 2007 Original article: DXS10079, DXS10074 and DXS10075: new alleles and SNP occurrence
EXCLI Journal 2007;6:177-182 ISSN 1611-2156 Received: December 14, 2006, accepted: June 21, 2007, published: June 26, 2007 Original article: DXS10079, DXS10074 and DXS10075: new alleles and SNP occurrence
Information leaflet. Centrum voor Medische Genetica. Version 1/20150504 Design by Ben Caljon, UZ Brussel. Universitair Ziekenhuis Brussel
 Information on genome-wide genetic testing Array Comparative Genomic Hybridization (array CGH) Single Nucleotide Polymorphism array (SNP array) Massive Parallel Sequencing (MPS) Version 120150504 Design
Information on genome-wide genetic testing Array Comparative Genomic Hybridization (array CGH) Single Nucleotide Polymorphism array (SNP array) Massive Parallel Sequencing (MPS) Version 120150504 Design
Forensic Science International: Genetics
 Forensic Science International: Genetics 4 (2009) 43 48 Contents lists available at ScienceDirect Forensic Science International: Genetics journal homepage: www.elsevier.com/locate/fsig Evaluation of the
Forensic Science International: Genetics 4 (2009) 43 48 Contents lists available at ScienceDirect Forensic Science International: Genetics journal homepage: www.elsevier.com/locate/fsig Evaluation of the
EUROPEAN. Geographic Trend Report for GMAT Examinees
 2011 EUROPEAN Geographic Trend Report for GMAT Examinees EUROPEAN Geographic Trend Report for GMAT Examinees The European Geographic Trend Report for GMAT Examinees identifies mobility trends among GMAT
2011 EUROPEAN Geographic Trend Report for GMAT Examinees EUROPEAN Geographic Trend Report for GMAT Examinees The European Geographic Trend Report for GMAT Examinees identifies mobility trends among GMAT
Mapping bias overestimates reference allele frequencies at the HLA genes in the 1000 Genomes Project phase I data
 Mapping bias overestimates reference allele frequencies at the HLA genes in the 1000 Genomes Project phase I data Débora Y. C. Brandt*, Vitor R. C. Aguiar*, Bárbara D. Bitarello*, Kelly Nunes*, Jérôme
Mapping bias overestimates reference allele frequencies at the HLA genes in the 1000 Genomes Project phase I data Débora Y. C. Brandt*, Vitor R. C. Aguiar*, Bárbara D. Bitarello*, Kelly Nunes*, Jérôme
CCR Biology - Chapter 9 Practice Test - Summer 2012
 Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Risk Factors for Alcoholism among Taiwanese Aborigines
 Risk Factors for Alcoholism among Taiwanese Aborigines Introduction Like most mental disorders, Alcoholism is a complex disease involving naturenurture interplay (1). The influence from the bio-psycho-social
Risk Factors for Alcoholism among Taiwanese Aborigines Introduction Like most mental disorders, Alcoholism is a complex disease involving naturenurture interplay (1). The influence from the bio-psycho-social
Survey of University of Michigan Graduate-level Area Studies Alumni/ae & FLAS Recipients from 1996-2006: Selected Findings
 Survey of University of Michigan Graduate-level Area Studies Alumni/ae & FLAS Recipients from 1996-2006: Selected Findings Azumi Ann Takata, Center for Japanese Studies, International Institute Donna Parmelee,
Survey of University of Michigan Graduate-level Area Studies Alumni/ae & FLAS Recipients from 1996-2006: Selected Findings Azumi Ann Takata, Center for Japanese Studies, International Institute Donna Parmelee,
The author(s) shown below used Federal funds provided by the U.S. Department of Justice and prepared the following final report:
 The author(s) shown below used Federal funds provided by the U.S. Department of Justice and prepared the following final report: Document Title: Author(s): Mitochondrial DNA Genome Sequencing and SNP Assay
The author(s) shown below used Federal funds provided by the U.S. Department of Justice and prepared the following final report: Document Title: Author(s): Mitochondrial DNA Genome Sequencing and SNP Assay
(1-p) 2. p(1-p) From the table, frequency of DpyUnc = ¼ (p^2) = #DpyUnc = p^2 = 0.0004 ¼(1-p)^2 + ½(1-p)p + ¼(p^2) #Dpy + #DpyUnc
 Advanced genetics Kornfeld problem set_key 1A (5 points) Brenner employed 2-factor and 3-factor crosses with the mutants isolated from his screen, and visually assayed for recombination events between
Advanced genetics Kornfeld problem set_key 1A (5 points) Brenner employed 2-factor and 3-factor crosses with the mutants isolated from his screen, and visually assayed for recombination events between
Protocols. Internal transcribed spacer region (ITS) region. Niklaus J. Grünwald, Frank N. Martin, and Meg M. Larsen (2013)
 Protocols Internal transcribed spacer region (ITS) region Niklaus J. Grünwald, Frank N. Martin, and Meg M. Larsen (2013) The nuclear ribosomal RNA (rrna) genes (small subunit, large subunit and 5.8S) are
Protocols Internal transcribed spacer region (ITS) region Niklaus J. Grünwald, Frank N. Martin, and Meg M. Larsen (2013) The nuclear ribosomal RNA (rrna) genes (small subunit, large subunit and 5.8S) are
GENETIC GENEALOGY AND DNA TESTING
 GENETIC GENEALOGY AND DNA TESTING by Ted Steele This publication may be ordered from: St. Louis Genealogical Society P. O. Box 432010 St. Louis, MO 63143 Copyright 2013 St. Louis Genealogical Society All
GENETIC GENEALOGY AND DNA TESTING by Ted Steele This publication may be ordered from: St. Louis Genealogical Society P. O. Box 432010 St. Louis, MO 63143 Copyright 2013 St. Louis Genealogical Society All
The Human Genome Project
 The Human Genome Project Brief History of the Human Genome Project Physical Chromosome Maps Genetic (or Linkage) Maps DNA Markers Sequencing and Annotating Genomic DNA What Have We learned from the HGP?
The Human Genome Project Brief History of the Human Genome Project Physical Chromosome Maps Genetic (or Linkage) Maps DNA Markers Sequencing and Annotating Genomic DNA What Have We learned from the HGP?
BASIC STATISTICAL METHODS FOR GENOMIC DATA ANALYSIS
 BASIC STATISTICAL METHODS FOR GENOMIC DATA ANALYSIS SEEMA JAGGI Indian Agricultural Statistics Research Institute Library Avenue, New Delhi-110 012 seema@iasri.res.in Genomics A genome is an organism s
BASIC STATISTICAL METHODS FOR GENOMIC DATA ANALYSIS SEEMA JAGGI Indian Agricultural Statistics Research Institute Library Avenue, New Delhi-110 012 seema@iasri.res.in Genomics A genome is an organism s
PAPER RFLP TEACHER GUIDE
 PAPER RFLP TEACHER GUIDE Paper = DNA Scissors = Restriction Enzyme Desktop = Electrophoresis NOTE: There are TWO versions of this activity one where the students write their own sentences (to represent
PAPER RFLP TEACHER GUIDE Paper = DNA Scissors = Restriction Enzyme Desktop = Electrophoresis NOTE: There are TWO versions of this activity one where the students write their own sentences (to represent
NEW GENERAL EDUCATION COURSES; FALL 2011 AND ON TOPICAL BREADTH. 552 General Education Options/Courses
 GENERAL EDUCATION OPTIONS/COURSES 552 General Education Options/Courses NEW GENERAL EDUCATION COURSES; FALL 2011 AND ON The following section pertains to students who matriculated to UC Davis for the first
GENERAL EDUCATION OPTIONS/COURSES 552 General Education Options/Courses NEW GENERAL EDUCATION COURSES; FALL 2011 AND ON The following section pertains to students who matriculated to UC Davis for the first
Worksheet - COMPARATIVE MAPPING 1
 Worksheet - COMPARATIVE MAPPING 1 The arrangement of genes and other DNA markers is compared between species in Comparative genome mapping. As early as 1915, the geneticist J.B.S Haldane reported that
Worksheet - COMPARATIVE MAPPING 1 The arrangement of genes and other DNA markers is compared between species in Comparative genome mapping. As early as 1915, the geneticist J.B.S Haldane reported that
Genomics Services @ GENterprise
 Genomics Services @ GENterprise since 1998 Mainz University spin-off privately financed 6-10 employees since 2006 Genomics Services @ GENterprise Sequencing Service (Sanger/3730, 454) Genome Projects (Bacteria,
Genomics Services @ GENterprise since 1998 Mainz University spin-off privately financed 6-10 employees since 2006 Genomics Services @ GENterprise Sequencing Service (Sanger/3730, 454) Genome Projects (Bacteria,
SNP Essentials The same SNP story
 HOW SNPS HELP RESEARCHERS FIND THE GENETIC CAUSES OF DISEASE SNP Essentials One of the findings of the Human Genome Project is that the DNA of any two people, all 3.1 billion molecules of it, is more than
HOW SNPS HELP RESEARCHERS FIND THE GENETIC CAUSES OF DISEASE SNP Essentials One of the findings of the Human Genome Project is that the DNA of any two people, all 3.1 billion molecules of it, is more than
DNA Sequence Analysis
 DNA Sequence Analysis Two general kinds of analysis Screen for one of a set of known sequences Determine the sequence even if it is novel Screening for a known sequence usually involves an oligonucleotide
DNA Sequence Analysis Two general kinds of analysis Screen for one of a set of known sequences Determine the sequence even if it is novel Screening for a known sequence usually involves an oligonucleotide
The Genetics of the Pre-Roman Iberian Peninsula: A mtdna Study of Ancient Iberians
 doi: 10.1111/j.1529-8817.2005.00194.x The Genetics of the Pre-Roman Iberian Peninsula: A mtdna Study of Ancient Iberians M. L. Sampietro 1,D.Caramelli 2,O. Lao 1,F.Calafell 1,D.Comas 1,M.Lari 2,B.Agustí
doi: 10.1111/j.1529-8817.2005.00194.x The Genetics of the Pre-Roman Iberian Peninsula: A mtdna Study of Ancient Iberians M. L. Sampietro 1,D.Caramelli 2,O. Lao 1,F.Calafell 1,D.Comas 1,M.Lari 2,B.Agustí
Forensic Science International: Genetics Journal Status Report
 Forensic Science International: Genetics Journal Status Report John M. Butler Associate Editor ISFG General Assembly Melbourne, Australia 4 September 2013 Expansion in Journal Editors 2011 2013 Peter Schneider
Forensic Science International: Genetics Journal Status Report John M. Butler Associate Editor ISFG General Assembly Melbourne, Australia 4 September 2013 Expansion in Journal Editors 2011 2013 Peter Schneider
Okami Study Guide: Chapter 3 1
 Okami Study Guide: Chapter 3 1 Chapter in Review 1. Heredity is the tendency of offspring to resemble their parents in various ways. Genes are units of heredity. They are functional strands of DNA grouped
Okami Study Guide: Chapter 3 1 Chapter in Review 1. Heredity is the tendency of offspring to resemble their parents in various ways. Genes are units of heredity. They are functional strands of DNA grouped
List of All AIMS Specialized Courses 2014~Fall 2016 (Tentative)
 List of All AIMS Specialized Courses 014~Fall 016 (Tentative) School Course Key Class Course Tutle Credits Core School of Political Science and Economics 1100001F3 01 Introduction to Chinese Linguistics
List of All AIMS Specialized Courses 014~Fall 016 (Tentative) School Course Key Class Course Tutle Credits Core School of Political Science and Economics 1100001F3 01 Introduction to Chinese Linguistics
Matthew Kaplan and Taylor Edwards. University of Arizona Tucson, Arizona
 Matthew Kaplan and Taylor Edwards University of Arizona Tucson, Arizona Unresolved paternity Consent for testing Ownership of Samples Uncovering genetic disorders SRY Reversal / Klinefelter s Syndrome
Matthew Kaplan and Taylor Edwards University of Arizona Tucson, Arizona Unresolved paternity Consent for testing Ownership of Samples Uncovering genetic disorders SRY Reversal / Klinefelter s Syndrome
GAW 15 Problem 3: Simulated Rheumatoid Arthritis Data Full Model and Simulation Parameters
 GAW 15 Problem 3: Simulated Rheumatoid Arthritis Data Full Model and Simulation Parameters Michael B Miller , Michael Li , Gregg Lind , Soon-Young
GAW 15 Problem 3: Simulated Rheumatoid Arthritis Data Full Model and Simulation Parameters Michael B Miller , Michael Li , Gregg Lind , Soon-Young
DNA as a Biometric. Biometric Consortium Conference 2011 Tampa, FL
 DNA as a Biometric Biometric Consortium Conference 2011 Tampa, FL September 27, 2011 Dr. Peter M. Vallone Biochemical Science Division National Institute of Standards and Technology Gaithersburg, MD 20899
DNA as a Biometric Biometric Consortium Conference 2011 Tampa, FL September 27, 2011 Dr. Peter M. Vallone Biochemical Science Division National Institute of Standards and Technology Gaithersburg, MD 20899
The Central Dogma of Molecular Biology
 Vierstraete Andy (version 1.01) 1/02/2000 -Page 1 - The Central Dogma of Molecular Biology Figure 1 : The Central Dogma of molecular biology. DNA contains the complete genetic information that defines
Vierstraete Andy (version 1.01) 1/02/2000 -Page 1 - The Central Dogma of Molecular Biology Figure 1 : The Central Dogma of molecular biology. DNA contains the complete genetic information that defines
Use of the Agilent 2100 Bioanalyzer and the DNA 500 LabChip in the Analysis of PCR Amplified Mitochondrial DNA Application
 Use of the Agilent 2100 Bioanalyzer and the DNA LabChip in the Analysis of PCR Amplified Mitochondrial DNA Application Homeland Security/Forensics Author Mark Jensen Agilent Technologies, Inc. 2850 Centerville
Use of the Agilent 2100 Bioanalyzer and the DNA LabChip in the Analysis of PCR Amplified Mitochondrial DNA Application Homeland Security/Forensics Author Mark Jensen Agilent Technologies, Inc. 2850 Centerville
