Synthetic lethal targeting of PTEN mutant cells with PARP inhibitors
|
|
|
- Preston Hoover
- 10 years ago
- Views:
Transcription
1 Report Targeting PTEN deficiency Synthetic lethal targeting of PTEN mutant cells with PARP inhibitors Ana M. Mendes-Pereira 1, Sarah A. Martin 1,2, Rachel Brough 1,2, Afshan McCarthy 1, Jessica R. Taylor 1, Jung-Sik Kim 3, Todd Waldman 3, Christopher J. Lord 1 *, Alan Ashworth 1,2 * Keywords: PTEN; PARP inhibitor; homologous recombination DOI /emmm Received April 6, 2009 Revised July 24, 2009 Accepted July 31, 2009 The tumour suppressor gene, phosphatase and tensin homolog (PTEN), is one of the most commonly mutated genes in human cancers. Recent evidence suggests that PTEN is important for the maintenance of genome stability. Here, we show that PTEN deficiency causes a homologous recombination (HR) defect in human tumour cells. The HR deficiency caused by PTEN deficiency, sensitizes tumour cells to potent inhibitors of the DNA repair enzyme poly(adp-ribose) polymerase (PARP), both in vitro and in vivo. PARP inhibitors are now showing considerable promise in the clinic, specifically in patients with mutations in either of the breast cancer susceptibility genes BRCA1 or BRCA2. The data we present here now suggests that the clinical assessment of PARP inhibitors should be extended beyond those with BRCA mutations to a larger group of patients with PTEN mutant tumours. INTRODUCTION Synthetic lethal approaches to cancer treatment have the potential to deliver relatively large therapeutic windows and significant patient benefit (Kaelin, 2005). We, and others, have previously shown that mutations in either of the tumour suppressor genes, breast cancer 1 (BRCA1) or BRCA2, are synthetically lethal with inhibition of the DNA repair enzyme poly(adp-ribose) polymerase 1 (PARP1) (Bryant et al, 2005; Farmer et al, 2005). The exquisite sensitivity of BRCA1 or BRCA2 mutant cells to PARP inhibitors (PARPi) forms the rationale behind clinical trials that are now assessing the potential of these agents (Ashworth, 2008). The preliminary results from these clinical trials are promising, with favourable toxicity and sustained tumour responses to the drug (Fong et al, 2009). Mutations in the phosphatase and tensin homolog (PTEN) gene (OMIM ) and loss of PTEN expression have both been associated with a wide range of human tumours (Salmena et al, 2008). PTEN encodes a phosphatase whose role in the (1) The Breakthrough Breast Cancer Research Centre, The Institute of Cancer Research, London, UK. (2) CRUK Gene Function Laboratory, The Institute of Cancer Research, London, UK. (3) Department of Oncology, Lombardi Comprehensive Cancer Center, Georgetown University School of Medicine, Washington, District of Columbia, USA. *Corresponding author: Tel: þ ; Fax: þ ; [email protected]; [email protected] control of the phosphoinositide 3 kinase (PI3K) signalling pathway is well established (Salmena et al, 2008). A new functional role for PTEN was recently suggested by the observation that mouse embryonic Pten / cells exhibit genomic instability, a phenotype ascribed to either defects in RAD51-mediated DNA double strand break repair (DSBR) (Shen et al, 2007) or defects in cell cycle checkpoints (Gupta et al, 2009). Given that the sensitivity of BRCA mutant cells to PARPi is most likely explained by a defect in RAD51-mediated DSBR by homologous recombination (HR) (Farmer et al, 2005; McCabe et al, 2006), we investigated the possibility that human PTEN mutant cells also display an HR defect and as a consequence, PARPi sensitivity. As PTEN loss of function mutations and loss of PTEN expression are common in a range of hereditary and sporadic cancers (Salmena et al, 2008), we reasoned that such data might significantly extend the utility of this class of drugs. RESULTS PTEN involvement in HR repair To model the effect of PTEN null mutations in human tumour cells, we used isogenically matched wild type and PTEN / HCT116 colorectal tumour cell lines (Lee et al, 2007) as well as isogenic wild type and PTEN / HEC1A endometroid adenocarcinoma cells (Waldman, unpublished work). PTEN deficiency in both HCT116 and HEC1A lines was achieved by targeting a truncating mutation to both copies of PTEN at exon 2, EMBO Mol Med 1, ß 2009 EMBO Molecular Medicine 315
2 Report Targeting PTEN deficiency resulting in an open reading frame encoding only the N-terminal 24 amino acids of the PTEN protein (Lee et al, 2007). First, we confirmed that PTEN / human tumour cells express reduced levels of RAD51 (Fig 1A and Fig S1A of Supporting Information), as previously documented in mouse Pten null cells (Shen et al, 2007). Extending this observation, we demonstrated that PTEN mutant tumour cells also had a reduced capacity to form nuclear RAD51 foci in response to DNA damage, a surrogate marker of HR activity (West, 2003) (Fig 1B, Figs S1B and S2 of Supporting Information). To investigate whether these deficiencies in RAD51 expression and recruitment translated into impaired DSBR by HR, we measured HR using a reporter assay. This comprised a previously validated synthetic DNA substrate that bears an inducible double strand DNA break (DSB; Saeki et al, 2006). PTEN deficient human tumour cells exhibited a 5-fold reduction in DSBR by HR when compared to isogenic wild type cells (Fig 1C and Fig S1C of Supporting Information). PTEN deficiency sensitizes to PARP inhibitors Given the HR repair defect of PTEN deficient cells, we tested the sensitivity of these cells to two DNA-DSB inducers, PARPi and cisplatin. HCT116 PTEN / cells were 20 times more sensitive to the PARP inhibitor KU (Farmer et al, 2005) than their wild type counterparts (compare concentration at which 50% of cells survive (SF 50 ) for HCT116 PTEN þ/þ of MwithSF 50 for HCT116 PTEN / 22 of M, Fig 2A) and up to 25 times more sensitive to the PARP inhibitor KU /AZD2281/ Olaparib (Evers et al, 2008; Fong et al, 2008) (compare SF 50 HCT116 PTEN þ/þ of M versus SF 50 for HCT116 PTEN / 22 of M, Fig 2B). We also observed PTEN selectivity in the HEC1A model (Fig S1D of Supporting Information). These PTEN/PARP synthetic lethal associations were achievable using sub-micromolar concentrations of PARPi, similar to those required to kill BRCA1 deficient cells (Farmer et al, 2005). We also assessed the sensitivity of PTEN / cells to the platinum salt, cisplatin. Cisplatin, although less selective than PARPi, also targets cells with HR deficiency (Tutt et al, 2005). PTEN / cells were five fold more sensitive to cisplatin than their wild type counterparts (compare SF 50 HCT116 PTEN þ/þ of M versus SF 50 for HCT116 PTEN / 22 of M, Fig 2C), consistent with the hypothesis that PTEN deficiency leads to an HR defect that may be targeted. To eliminate the possibility that these observations were reflective of a general sensitivity to chemotherapeutics caused by PTEN deficiency, we Figure 1. PTEN deficiency causes an impairment of DNA repair by HR. A. PTEN deficiency causes a reduction in RAD51 expression. Total cell lysates from isogenic HCT116 colorectal tumour cells were immunoblotted for PTEN and RAD51. Detection of b Tubulin is shown as a loading control. Lysates from parental HCT116 cells were used (HCT116) along with two individually derived PTEN / lines (KO35 and KO22) and also a HCT116-derived line bearing a random integration of the PTEN targeting construct (neo124). Lysates from PTEN þ/ HCT116 cells are also shown (Lee et al, 2007). B. PTEN deficiency causes a reduction in radiation-induced nuclear RAD51 focus formation. Cells were exposed to 10 Gy g-irradiation and nuclear RAD51 foci quantified 8 h later by confocal microscopy (Farmer et al, 2005). Bar chart shows the average number of cells with >5 foci per nucleus. Error bars represent three standard deviations of the mean. p values versus þir HCT116 PTEN þ/þ <0.05 (Student s t-test). All other comparisons returned non-significant (>0.05) p values. C. PTEN deficiency causes a reduction in HR as measured using a synthetic HR substrate. As a measure of HR activity, a reporter plasmid-based assay was used comprising two defective copies of GFP, where one serves as a template to restore an induced DSB in the other. HR between the two GFP coding sequences results in an intact GFP coding sequence and cellular fluorescence (Saeki et al, 2006). Error bars represent three standard deviations of the mean. p values versus HCT116 PTEN þ/þ <0.05 (Student s t-test). 316 ß 2009 EMBO Molecular Medicine EMBO Mol Med 1,
3 Report Ana M. Mendes-Pereira et al. Figure 2. PTEN deficiency sensitizes tumour cells to agents that target HR. A, B. PTEN deficiency sensitizes cells to drug-like PARP inhibitors. Survival curves for HCT116 cells exposed to the PARP inhibitors KU (Farmer et al, 2005) or KU (Evers et al, 2008; Fong et al, 2008). Cells were plated in six-well plates and treated for 15 days, after which SFs were estimated. For both PARP inhibitors, PTEN / SFs were significantly different to both PTEN þ/þ and PTEN þ/ cells (p<0.05 two-way analysis of variance (ANOVA)). No other comparisons returned significant p values. C. PTEN deficiency sensitizes cells to cisplatin (p<0.05 PTEN / lines versus PTEN þ/þ cells, two-way ANOVA). D. PTEN deficiency does not confer sensitization to taxol. assessed sensitivity to the microtubule poison, paclitaxel (taxol). Paclitaxel was not selectively lethal to PTEN deficient cells (Fig 2D), suggesting that the PARPi and cisplatin sensitivities we observed were more likely reflective of HR deficiency rather than of general chemosensitivity. Although PTEN haploinsufficiency effects on tumourigenesis have been reported, complete loss of PTEN is observed in many advanced cancers (Salmena et al, 2008). Given this, we thought it pertinent to assess the effects of PTEN gene dosage upon HR deficiency and PARPi sensitivity. Using the HCT116 isogenic system, we demonstrated that PTEN heterozygosity did not cause the same reduction in RAD51 expression as seen in PTEN / cells (Fig 1B). Consistent with this observation, the induction of damage induced nuclear RAD51 foci was not abrogated in PTEN þ/ cells (Fig 1B) and although a marginal decrease in HR was observed using the synthetic HR assay in heterozygous cells (Fig 1C), this decrease did not translate into PARPi sensitivity (Fig 2B). To assess the generality of our observations, we examined PARPi sensitivity in an additional panel of tumour cell lines (Fig 3, Fig S3 and Tables S1 and S2 of Supporting Information). Although PTEN mutations are not the only genetic variable within this diverse tumour cell panel, we observed a clear distinction in PARPi sensitivity between tumour lines with wild type PTEN expression, such as DU145 and MCF7, and those with PTEN deficiency, such as HCC70, PC3, RPMI-7951, A172, UM-UC3 and MDA-MB-468 (Fig 3A,B and Fig S3 of Supporting Information). Notably, PARPi sensitivity in PTEN deficient lines was of a scale similar to that observed in human tumour lines with BRCA1 silencing (Farmer et al, 2005). With the exception of the glioma line A172, we also observed consistent cisplatin sensitivity in PTEN deficient lines (Fig 3C). EMBO Mol Med 1, ß 2009 EMBO Molecular Medicine 317
4 Report Targeting PTEN deficiency We utilized the PTEN deficient prostate tumour cell line PC3, to assess whether PARPi sensitivity could be abrogated by ectopic expression of PTEN. As expected, infection of PC3 cells with an empty retroviral expression construct did not significantly modify PARPi sensitivity, nor did transfection of an expression construct encoding the p.r55fs 1 PTEN allele normally found in PC3 cells (Fig 4A,B). However, ectopic expression of full length PTEN in PC3 cells elicited PARPi resistance of a scale similar to tumour cells with endogenous wild type PTEN expression (Fig 4A). We also assessed whether PARPi sensitivity by PTEN was independent of the phosphatase activity of PTEN. Ectopic expression of the catalytically inactive C124S PTEN mutant (Maehama & Dixon, 1998; Shen et al, 2007) also caused PARPi resistance in PC3 cells (Fig 4A), suggesting that a non-phosphatase activity of PTEN may determine PARPi sensitivity. We also analysed whether restoring cytoplasmic PTEN expression, while suppressing nuclear PTEN expression could confer PARP inhibitor resistance in a PTEN null cell line, such as PC3. Here we used ectopic expression of a PTEN cdna that bears a mutation (p.k289e) that reduces the shuttling of Figure 3. PARP inhibitor sensitivity in human tumour cells. A. Western blot showing PTEN expression in a panel of human tumour lines. PTEN mutant lines are shown in red and PTEN wild type lines in black. PTEN genotypes are as follows: DU145 (prostate), PTEN wild type; MCF7 (breast), PTEN wild type; PC3 (prostate) p.r55fs 1 homozygous; MDAMB468 (breast), p.a72fs 5 homozygous; A172 (glioma), p.r55fs 1homozygous;UM-UC3 (bladder), p.m1 404del homozygous; RPMI-7951 (melanoma) p.null, c.1-79del79 homozygous; HCC70 (breast), p.f90fs 9 homozygous (see Table S1 of Supporting Information for complete genotypes). B. PARPi sensitivity in the same cell line panel. For each PTEN null line, survival was significantly different than in DU145 and MCF7 cells ( p<0.05, two-way ANOVA) (see Fig S3 of Supporting Information for analysis of an enlarged cell line panel). C. Cisplatin sensitivity in the same cell line panel. With the exception of A172, survival of each PTEN null line at 1 mm was significantly different than in DU145 and MCF7 cells ( p<0.05, two-way ANOVA). Figure 4. PARP inhibitor sensitivity in human tumour cells. A, B. Wild type and catalytically inactive PTEN, both restore PARP inhibitor resistance, as does expression of RAD51; SFs in PC3 cells infected with PTEN wild type, PTEN p.c124s or RAD51 cdna expression constructs was significantly different than in PC3 þ empty vector (p<0.05, two-way ANOVA). PC3 cells infected with p.k289e cdna expression construct did not cause significant resistance. PTEN null PC3 cells were infected with cdna expression constructs as shown and exposed to PARP inhibitor. Western blot and survival curves are shown. 318 ß 2009 EMBO Molecular Medicine EMBO Mol Med 1,
5 Report Ana M. Mendes-Pereira et al. Figure 5. In vivo efficacy of Olaparib in PTEN deficient xenografts. A, B. The clinical PARP inhibitor KU /Olaparib (Evers et al, 2008; Fong et al, 2008) suppresses growth in PTEN / subcutaneous xenografts (A), but not in PTEN þ/þ xenografts (B). PTEN deficient or proficient HCT116 cells were each mixed 2:1 in matrigel and then injected into either bilateral flank of 6 8 week old female athymic nude mice. After two days recovery, mice were randomized into treatment and control groups, (10 animals per cohort, 20 in total) and drug dosing initiated. A treatment regime similar to that known to elicit Brca2 selectivity (Farmer et al, 2005) was used. For five consecutive days, single daily intraperitoneal doses of KU (or vehicle) were administered at a dose of 15 mg/kg in HBC (Farmer et al, 2005). This was followed by two consecutive days of no treatment after which the same treatment cycle was continued until the end of the study. Tumour volumes were measured every four days from the initiation of drug dosing and the results expressed as fold increase in tumour volume relative to that at the first drug administration. Tumour volume vehicle treated vs. drug treated, p ¼ 0.04 for PTEN / xenografts and p ¼ 0.44 for PTEN þ/þ xenografts (two-way repeated measures ANOVA). PTEN into the nucleus (Song et al, 2008) (Fig 4B). Expression of PTEN p.k289e was able to partially but not completely restore PARP inhibitor resistance (Fig 4A), suggesting that an impairment of the nuclear localization of PTEN also modulates sensitivity to PARP inhibitor. Finally, we sought to assess whether the PARPi sensitivity of a PTEN cell line was RAD51 dependent. Ectopic expression of RAD51 caused PARP inhibitor resistance in PC3 cells (Fig 4A and Fig S4 of Supporting Information), supporting the hypothesis that reduced RAD51 expression underlies the PARP inhibitor sensitivity of PTEN deficient cells. To assess PARPi selective efficacy in vivo, we xenografted PTEN deficient HCT116 cells into mice and treated animals with the PARPi KU (Olaparib) (Evers et al, 2008) which is currently undergoing clinical assessment (Fong et al, 2009). Although xenograft growth in this model was extremely rapid, growth from PTEN deficient xenografts was suppressed by KU when compared to vehicle-treated xenografts (Fig 5A). Conversely PARPi did not have a similar effect on the growth of PTEN wild type xenografts (Fig 5B). This provides preliminary evidence to suggest that our in vitro observations may extrapolate to an in vivo setting. DISCUSSION PARP inhibitors are now showing considerable promise in the treatment of cancers characterized by genetic deficiencies that lead to an impairment of HR. For example, observations from a phase I trial of the PARP inhibitor KU /AZD2281/ Olaparib (AstraZeneca) suggest that significant tumour responses can be achieved in ovarian cancer patients carrying BRCA mutations (Fong et al, 2009). These tumour responses, demonstrated by both radiological estimation of tumour size and by the use of biochemical markers of tumour burden, are sustained and can be achieved without the significant toxicity normally associated with standard chemotherapeutic regimes (Fong et al, 2009). These promising results have led to phase II trials of KU /AZD2281/Olaparib for breast and ovarian cancers in BRCA mutation carriers (Tuma, 2007). Here, we show that PARP inhibitors may have utility outside the relatively small proportion of cancer patients carrying BRCA mutations. PTEN mutations are common in both endometrial cancers and glioblastomas and are also found in a significant proportion of malignant melanomas, prostate, breast, lung and colorectal tumours (Keniry & Parsons, 2008). In addition, transcriptional or translational repression of PTEN expression in the absence of PTEN mutation is also a common characteristic of multiple tumour types (Wiencke et al, 2007; Yang et al, 2008). The profound sensitivity of PTEN deficient tumour cells to PARP inhibitors demonstrated here suggests that clinical assessment of these agents in a wider context is warranted and perhaps even in combination with PI3K/AKT inhibitors that have been already been proposed as a means to target PTEN deficient tumours (Knight & Shokat, 2007). Our initial studies suggest that homozygous PTEN mutations that severely truncate the open reading frame sensitize tumour cells to PARP inhibitors. For example, glioblastoma, breast and prostate tumour cell lines bearing homozygous PTEN mutations that truncate the open reading frame before residue c.279 (amino acid residue 93) are as sensitive to KU , as human tumour cells with BRCA1 or BRCA2 deficiencies (Farmer et al, 2005; Turner et al, 2008). Notably, reconstitution of PC3 cells EMBO Mol Med 1, ß 2009 EMBO Molecular Medicine 319
6 Report Targeting PTEN deficiency (p.r55fs 1 homozygous) with ectopic expression of a catalytically inactive PTEN allele (p.c124s) was sufficient to restore PARPi resistance (Fig 4). Therefore, it seems likely that tumourassociated PTEN missense mutations that only modify catalytic activity without ostensibly compromising PTEN expression or non-phosphatase activities may not sensitize tumour cells to PARP. Conversely, PTEN mutations that severely truncate the open reading frame or cause instability of the PTEN protein are more likely to cause PARPi sensitivity. Mutations that truncate the open reading frame of PTEN before residue 93, as in the PARPi sensitive HCC70 cell line (p.f90fs 9 homozygous) are found in a wide variety of tumour types (Table S2 of Supporting Information), suggesting at least one patient subgroup that should be rapidly assessed for response to PARP inhibitors in clinical trials. Tumours that have PTEN deficiency brought about by PTEN mutations in the C-terminus that compromise protein stability (Leslie et al, 2008) or non-genetic alterations that cause changes in PTEN transcription or translation, may also respond to PARP inhibitors, although considerable work is required to establish a threshold of PTEN expression below which PARPi sensitivity occurs. Furthermore, we envisage that the role PTEN plays in determining response to PARP inhibitors, is mediated by a nuclear activity of the protein, suggesting that a complete absence of nuclear PTEN expression, determined by immunohistochemical (IHC) detection in biopsy material, could predict response. This raises the possibility that PARP inhibitors could specifically be assessed in tumourigenic conditions such as the more aggressive forms of oesophageal squamous cell carcinoma (Tachibana et al, 2002), cutaneous melanoma (Whiteman et al, 2002) and colorectal cancer (Trotman et al, 2007; Zhou et al, 2002), where nuclear PTEN expression is absent. Defining thresholds of IHC detection that are predictive of PARPi response will be a key in assessing this possibility, and we propose that combining PTEN IHC with surrogate markers of PARPi sensitivity, such as the inability to form nuclear RAD51 foci (Farmer et al, 2005), may allow further refinement of the use of PARP inhibitors in cancer treatment. MATERIALS AND METHODS Materials PARP inhibitors KU (Farmer et al, 2005) and KU (Evers et al, 2008; Fong et al, 2008) (KuDOS Pharmaceuticals/Astra Zeneca), cisplatin and paclitaxel (Sigma Aldrich) were used as described previously (Edwards et al, 2008; Farmer et al, 2005). Cell culture Wild type, PTEN þ/ and PTEN / HCT116 or HEC1A cells (Kim et al, 2007; Lee et al, 2007) were maintained in McCoy s 5A medium (Invitrogen, Glasgow, UK) supplemented with 10% v/v foetal bovine serum (FBS) (PAA gold, GIBCO). In the case of PTEN / cells, two individually derived lines were used, (KO22 and KO35 for HCT116 and KO22 and KO16 for HEC1A). For the HCT116 experiments, parental cells and a HCT116 derived line bearing a random integration of the PTEN targeting construct (neo124) were used as PTEN wild type comparators. Two HCT116 PTEN þ/ cell lines bearing one targeted allele (130 and 140) were also analysed. All other cells lines were obtained from the ECACC (Teddington, UK) and cultured according to the supplier s instructions. Plasmids The full length coding sequence of PTEN was PCR amplified from a cdna IMAGE clone (Geneservice, Cambridge, UK), using a 5 0 primer harbouring a Kozak consensus sequence (GCCACC prior to initiating ATG). The resultant PCR product was cloned into a tag-free pbabe-puro retroviral expression vector. Mutation of the PTEN cdna sequence was achieved using the Quick Change site-directed mutagenesis kit (Stratagene). Twenty-four hours after infection, cells were selected in 1 mg/l puromycin (InvivoGen) for three days to remove non-infected cells, then seeded in six-well plates (at 1500 cells/well) and treated with drug the following day, as previously described (Edwards et al, 2008; Farmer et al, 2005). Survival assays Long-term survival assays were performed as previously described (Edwards et al, 2008; Farmer et al, 2005). For measurement of sensitivity to PARP inhibitor, cisplatin or paclitaxel (taxol), exponentially growing cells were seeded in six-well plates at a concentration of cells per well. For PARP inhibitor and taxol treatment, cells were continuously exposed to the drug with media and drug replaced every 60 h. For cisplatin treatment, cells were exposed to the drug at the indicated concentrations for 24 h, washed twice with phosphate buffered saline (PBS), then normal media replaced every 3-4 days. After 15 days, cells were fixed and stained with sulphorhodamine-b (Sigma, St. Louis, USA) and a colorimetric assay performed as described previously (Edwards et al, 2008). Surviving fractions (SFs) were calculated and drug sensitivity curves plotted as previously described (Farmer et al, 2005). Protein analysis Whole-cell protein extracts were prepared from cells lysed with radioimmunoprecipitation (RIPA) buffer (Upstate) supplemented with protease inhibitor cocktail tablets (Roche, Burgess Hill, UK). Protein concentrations were measured using BioRad Protein Assay Reagent (BioRad, Hemel Hempstead, UK). For Western blot analysis, lysates were electrophoresed on Novex 4 12% gradient bis tris pre-cast gels (Invitrogen) and immunoblotted overnight at 48C with antibodies targeting the following epitopes: PTEN C-terminus (Cell Signalling, Danvers, USA), RAD51 (H-92, Santa Cruz Biotech, Santa Cruz, USA), ACTIN (Santa Cruz Biotech) or b-tubulin (T4026, Sigma). Incubation with primary antibody was followed by incubation with a horseradish peroxidase-conjugated secondary antibody and chemiluminescent detection of proteins (Amersham Pharmacia, Cardiff, UK). DSB Repair Assay DSBR was quantified using a synthetic repair substrate as previously described (Saeki et al, 2006). Briefly, the DR-green fluorescent protein (GFP) HR reporter plasmid, containing two differentially mutated GFPencoding cassettes, one of which contains a unique ISceI restriction site, was co-transfected into cells alongside a ISceI expression vector, pcbasce (Saeki et al, 2006). Seventy-two hours after transfection, HR competence was estimated by quantifying the number of GFP positive cells from a population of 50,000 cells, using flow cytometry (Saeki et al, 2006). 320 ß 2009 EMBO Molecular Medicine EMBO Mol Med 1,
7 Report Ana M. Mendes-Pereira et al. The paper explained PROBLEM: The tumour suppressor gene, PTEN, is one of the most commonly mutated genes in human cancers. Selectively targeting cancerous PTEN deficient cells would likely make for an efficient and relatively safe cancer therapy applicable to the large number of PTEN deficient cancers. Selective targeting of BRCA1 or BRCA2 deficient tumours has recently been demonstrated with Olaparib, a high-potency PARP inhibitor. This drug can elicit sustained responses in BRCA deficient cancers without causing some of the negative side effects caused by standard chemotherapies. Tumour cells in BRCA mutant patients have a defect in the DNA repair process of HR that repairs DSBs and it is thought that PARP inhibitors target this HR defect by increasing the formation of DSBs. suggesting that drugs that target BRCA mutation could also be used to target PTEN mutation. We demonstrate that PTEN deficiency in human tumour cells also leads to suppression of HR. Using in vitro cell culture and in vivo tumour modelling in mice, we show that, as in BRCA deficient tumours, the HR deficiency in PTEN mutant cells can be exploited therapeutically by Olaparib. IMPACT: Our work suggests that this relatively non-toxic anti-cancer drug can be used to target the large number of PTEN deficient tumours, significantly extending its previously anticipated limited spectrum. RESULTS: Recent evidence suggests that, like BRCA mutant cells, PTEN mutant cells are also characterized by an increase in DSBs, Analysis of RAD51 foci formation Nuclear RAD51 foci were visualized and quantified by confocal microscopy as previously described (Farmer et al, 2005). Detection of gamma (g)-h2ax foci was used a control marker for the formation of DSBs (data not shown). In brief, cells were grown onto L-lysine coated cover slips (BD Biosciences, Oxford, UK) and exposed to 10 Gy ionizing radiation. Eight hours later, cells were fixed, permeabilized and then co-immunostained with primary antibodies targeting RAD51 (rabbit, Santa Cruz Biotech) and gh2ax (mouse, Upstate), detected with fluorescein isothiocyanate (FITC) and Texas red conjugated secondary antibodies respectively Diamidino-2-phenylindole (DAPI) staining was used to detect nuclei. Nuclear foci were visualized by confocal microscopy and a minimum of 100 fields of view used to estimate RAD51 focus formation frequency. Xenografts HCT116 cells were mixed 2:1 in matrigel (BD Biosciences) and then injected into the bilateral flanks of 6 8 week old female athymic nude mice ( cells per animal). After two days recovery, mice were randomized into treatment and control groups, (10 animals per cohort, 20 in total) and drug dosing initiated. We used a treatment regime similar to that known to elicit Brca2 selectivity (Farmer et al, 2005). For five consecutive days, single daily intraperitoneal doses of KU (or vehicle) were administered at a dose of 15 mg/kg in 2-hydroxypropyl-b-cyclodextrin (HBC, Farmer et al, 2005). This was followed by two consecutive days of no treatment after which the same treatment cycle was continued until the end of the study. Tumour volumes were measured every four days from the initiation of drug dosing and the results expressed as fold increase in tumour volume, relative to that at the first drug administration. Author Contributions AMP, CJL and AA conceived and designed the experiments. AMP, SAM, RB, AM and JT performed the experiments. AMP, SAM, RB, AM, JT, CJL and AA analysed the experiments. TW and JSK provided reagents. AMP, CJL and AA wrote the manuscript. CJL and AA devised the project and obtained funding. Acknowledgements We thank Drs. Graeme Smith and Niall Martin of KuDOS Pharmaceuticals for the provision of PARP inhibitors. This work was funded by Breakthrough Breast Cancer and Cancer Research UK. We acknowledge NHS funding to the NIHR Biomedical Research Centre. Supporting information is available at EMBO Molecular Medicine online. The authors declare that they have no conflict of interest. For more information The Breakthrough Research Center: centre/index.html Clinical trials (Olaparib): Cancer Research UK: Cancer Help UK (Olaparib): EMBO Mol Med 1, ß 2009 EMBO Molecular Medicine 321
8 Report Targeting PTEN deficiency References Ashworth A (2008) A synthetic lethal therapeutic approach: poly(adp) ribose polymerase inhibitors for the treatment of cancers deficient in DNA double-strand break repair. J Clin Oncol 26: Bryant HE, Schultz N, Thomas HD, Parker KM, Flower D, Lopez E, Kyle S, Meuth M, Curtin NJ, Helleday T (2005) Specific killing of BRCA2-deficient tumours with inhibitors of poly(adp-ribose) polymerase. Nature 434: Edwards SL, Brough R, Lord CJ, Natrajan R, Vatcheva R, Levine DA, Boyd J, Reis-Filho JS, Ashworth A (2008) Resistance to therapy caused by intragenic deletion in BRCA2. Nature 451: Evers B, Drost R, Schut E, de Bruin M, van der Burg E, Derksen PW, Holstege H, Liu X, van Drunen E, Beverloo HB, et al (2008) Selective Inhibition of BRCA2-Deficient Mammary Tumor Cell Growth by AZD2281 and Cisplatin. Clin Cancer Res 14: Farmer H, McCabe N, Lord CJ, Tutt AN, Johnson DA, Richardson TB, Santarosa M, Dillon KJ, Hickson I, Knights C, et al (2005) Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434: Fong PC, Boss DS, Carden CP, Roelvink M, De Greve J, Gourley CM, Carmichael J, De Bono JS, Schellens JH, Kaye SB (2008) AZD2281 (KU ), a PARP (poly ADP-ribose polymerase) inhibitor with single agent anti-cancer activity in patients with BRCA deficient ovarian cancer. Results from a phase I study. J Clin Oncol 26: abstr 5510 Fong PC, Boss DS, Yap TA, Tutt A, Wu P, Mergui-Roelvink M, Mortimer P, Swaisland H, Lau A, O Connor MJ, et al (2009) Inhibition of poly(adp-ribose) polymerase in tumors from BRCA mutation carriers. N Engl J Med 361: Gupta A, Yang Q, Pandita RK, Hunt CR, Xiang T, Misri S, Zeng S, Pagan J, Jeffery J, Puc J, et al (2009) Cell cycle checkpoint defects contribute to genomic instability in PTEN deficient cells independent of DNA DSB repair. Cell Cycle 8 Kaelin WG Jr. (2005) The concept of synthetic lethality in the context of anticancer therapy. Nat Rev Cancer 5: Keniry M, Parsons R (2008) The role of PTEN signaling perturbations in cancer and in targeted therapy. Oncogene 27: Kim JS, Lee C, Bonifant CL, Ressom H, Waldman T (2007) Activation of p53-dependent growth suppression in human cells by mutations in PTEN or PIK3CA. Mol Cell Biol 27: Knight ZA, Shokat KM (2007) Chemically targeting the PI3K family. Biochem Soc Trans 35: Lee C, Kim JS, Waldman T (2007) Activated PI3K signaling as an endogenous inducer of p53 in human cancer. Cell Cycle 6: Leslie NR, Batty IH, Maccario H, Davidson L, Downes CP (2008) Understanding PTEN regulation: PIP2, polarity and protein stability. Oncogene 27: Maehama T, Dixon JE (1998) The tumor suppressor, PTEN/MMAC1, dephosphorylates the lipid second messenger, phosphatidylinositol 3,4,5-trisphosphate. J Biol Chem 273: McCabe N, Turner NC, Lord CJ, Kluzek K, Bialkowska A, Swift S, Giavara S, O Connor MJ, Tutt AN, Zdzienicka MZ, et al (2006) Deficiency in the repair of DNA damage by homologous recombination and sensitivity to poly(adpribose) polymerase inhibition. Cancer Res 66: Saeki H, Siaud N, Christ N, Wiegant WW, van Buul PP, Han M, Zdzienicka MZ, Stark JM, Jasin M (2006) Suppression of the DNA repair defects of BRCA2-deficient cells with heterologous protein fusions. Proc Natl Acad Sci USA 103: Salmena L, Carracedo A, Pandolfi PP (2008) Tenets of PTEN tumor suppression. Cell 133: Shen WH, Balajee AS, Wang J, Wu H, Eng C, Pandolfi PP, Yin Y (2007) Essential role for nuclear PTEN in maintaining chromosomal integrity. Cell 128: Song MS, Salmena L, Carracedo A, Egia A, Lo-Coco F, Teruya-Feldstein J, Pandolfi PP (2008) The deubiquitinylation and localization of PTEN are regulated by a HAUSP-PML network. Nature 455: Tachibana M, Shibakita M, Ohno S, Kinugasa S, Yoshimura H, Ueda S, Fujii T, Rahman MA, Dhar DK, Nagasue N (2002) Expression and prognostic significance of PTEN product protein in patients with esophageal squamous cell carcinoma. Cancer 94: Trotman LC, Wang X, Alimonti A, Chen Z, Teruya-Feldstein J, Yang H, Pavletich NP, Carver BS, Cordon-Cardo C, Erdjument-Bromage H, et al (2007) Ubiquitination regulates PTEN nuclear import and tumor suppression. Cell 128: Tuma RS (2007) Combining carefully selected drug, patient genetics may lead to total tumor death. J Natl Cancer Inst 99: , 1509 Turner NC, Lord CJ, Iorns E, Brough R, Swift S, Elliott R, Rayter S, Tutt AN, Ashworth A (2008) A synthetic lethal sirna screen identifying genes mediating sensitivity to a PARP inhibitor. Embo J 27: Tutt AN, Lord CJ, McCabe N, Farmer H, Turner N, Martin NM, Jackson SP, Smith GC, Ashworth A (2005) Exploiting the DNA repair defect in BRCA mutant cells in the design of new therapeutic strategies for cancer. Cold Spring Harb Symp Quant Biol 70: West SC (2003) Molecular views of recombination proteins and their control. Nat Rev Mol Cell Biol 4: Whiteman DC, Zhou XP, Cummings MC, Pavey S, Hayward NK, Eng C (2002) Nuclear PTEN expression and clinicopathologic features in a population-based series of primary cutaneous melanoma. Int J Cancer 99: Wiencke JK, Zheng S, Jelluma N, Tihan T, Vandenberg S, Tamguney T, Baumber R, Parsons R, Lamborn KR, Berger MS, et al (2007) Methylation of the PTEN promoter defines low-grade gliomas and secondary glioblastoma. Neuro Oncol 9: Yang H, Kong W, He L, Zhao JJ, O Donnell JD, Wang J, Wenham RM, Coppola D, Kruk PA, Nicosia SV, et al (2008) MicroRNA expression profiling in human ovarian cancer: mir-214 induces cell survival and cisplatin resistance by targeting PTEN. Cancer Res 68: Zhou XP, Loukola A, Salovaara R, Nystrom-Lahti M, Peltomaki P, de la Chapelle A, Aaltonen LA, Eng C (2002) PTEN mutational spectra, expression levels, and subcellular localization in microsatellite stable and unstable colorectal cancers. Am J Pathol 161: ß 2009 EMBO Molecular Medicine EMBO Mol Med 1,
9 Supplementary Figure 1 A B HEC1A PTEN +/+ HEC1A PTEN -/- 16 HEC1A PTEN -/- 22 PTEN RAD51 % Cells with RAD51 foci 40 -IR +IR * *! TUBULIN 0 HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/- 22 C D 1.0 HR Efficiency * * Surviving fraction SF50 (M) HEC1A PTEN +/+ >1.0E-05 HEC1A PTEN -/- -/ E-05 HEC1A PTEN -/- -/ E HEC1A PTEN+/+ HEC1A PTEN-/- 16 HEC1A PTEN-/ KU (M) KU (M) 10-5
10 Supplementary Figure 2 "H2AX Merged HCT116 PTEN+/+ Rad51 - IR HCT116 PTEN-/- Nuclei - IR + IR + IR
11 Supplementary Figure 3 Surviving fraction DU145 MCF-7 HeLa SUM159 BT474 MDA-MB-231 HCC70 PC3 RPMI-7951 SW1783 EC50 SF50 (M) > 1E-05 > 1E-05 > 1E-05 > 1E-05 > 1E-05 > 1E E E E E A172 UM-UC3 2.94E E-08 MDA-MB E COLO E-09 KU (M)
12 Supplementary Figure 4 EMPTY VECTOR RAD51 cdna RAD51! Tubulin
13 Supplementary Table 1 Cell Line PTEN PIK3CA TP53 BRAF KRAS MLH1 MSH6 CDKN2A A172 * COLO 829 * * * HCC70 * * MDA-MB-468 * * PC3 * * RPMI-7951 * * * * SW1783 * * UM-UC3 * * * * BT474 * * HEC1A * * * HCT116 * * * * MCF7 * * SUM-159 * MDA-MB-231 * * * * DU145 * * * HeLa Cell Line Mutation PTEN PIK3CA TP53 BRAF KRAS MLH1 MSH6 CDKN2A A172 AA p.r55fs*1 CDS c.165_1212del1048 glioma Zygosity Homozygous COLO 829 AA p.? p.v600e p.a68fs*51 CDS c.493_634del142 c.1799t>a c.203_204delcg melanoma Zygosity Homozygous Heterozygous Homozygous HCC70 AA p.f90fs*9 p.r248q CDS c.270delt c.743g>a breast Zygosity Homozygous Homozygous MDA-MB-468 AA p.? p.r273h CDS c.253+1g>t c.818g>a breast Zygosity Homozygous Homozygous PC3 AA p.r55fs*1 p.k139fs*31 CDS c.165_1212del1048 c.414delc prostate Zygosity Homozygous Homozygous RPMI-7951 AA p.? p.s166* p.v600e p.l16r CDS c.1_79del79 c.497c>a c.1799t>a c.47t>g melanoma Zygosity Homozygous Homozygous Heterozygous Homozygous SW1783 AA p.r233* p.r273h CDS c.697c>t c.818g>a glioma Zygosity Homozygous Heterozygous UM-UC3 AA p.m1_*404del p.f113c p.g12c p.m1_*157del CDS c.1_1212del1212 c.338t>g c.34g>t c.1_471del471 bladder Zygosity Homozygous Homozygous Homozygous Homozygous BT-474 AA p.k111n p.e285k CDS c.333g>c c.853g>a breast Zygosity Heterozygous Homozygous HEC1A AA p.g1049r p.r248q p.g12d CDS c.3145g>c c.743g>a c.35g>a endometrium Zygosity Heterozygous Homozygous Homozygous HCT116 AA p.h1047r p.g13d p.s252* p.r24fs*20 CDS c.3140a>g c.38g>a c.755c>a c.68_69insg colorectal Zygosity Heterozygous Heterozygous Homozygous Homozygous MCF7 AA p.e545k p.m1_*157del CDS c.1633g>a c.1_471del471 breast Zygosity Heterozygous Homozygous SUM-159 AA p.h1047l CDS c.3140a>t breast Zygosity Heterozygous MDA-MB-231 AA p.r280k p.g464v p.g13d p.m1_*157del CDS c.839g>a c.1391g>t c.38g>a c.1_471del471 breast Zygosity Homozygous Heterozygous Heterozygous Homozygous DU145 AA p.p223l, p.v274f p.? p.d84y CDS c.668c>t, c.820g>t c.117-2a>t c.250g>t prostate Zygosity Heterozygous Homozygous Homozygous HeLa AA CDS cervix Zygosity
14 Sample Name COSMIC Sample ID Amino Acid Nucleotide Primary Tissue Histology HCC p.f90fs*9 c.270delt breast carcinoma 5839 cell-line S p.k60fs*9 c.180_181ins? breast carcinoma NS MDA- MB p.l70fs*7 c.208_251del44 breast carcinoma NS p.a39fs*5 c.116_117insg central_nervous_system glioma fresh/frozen S p.a79fs*20 c.237delc central_nervous_system glioma primary SKMG p.c71fs*6 c.212_255del44 central_nervous_system glioma NS S p.d24fs*20 c.70_71insa central_nervous_system glioma primary S p.d24fs*20 c.70_71insa central_nervous_system glioma primary S p.e43fs*11 c.127delg central_nervous_system glioma NS S p.f56fs*2 c.165_492del328 central_nervous_system glioma primary p.k60* c.178a>t central_nervous_system glioma fresh/frozen S p.k66fs*38 c.197_198ins14 central_nervous_system glioma primary S p.k6fs*4 c.17_18delaa central_nervous_system glioma primary S p.l70fs*7 c.210_253del44 central_nervous_system glioma primary S p.l70fs*7 c.210_253del44 central_nervous_system glioma primary SF p.m1_*404del c.1_1212del1212 central_nervous_system glioma cell-line H p.m1_*404del c.1_1212del1212 central_nervous_system glioma cell-line D- 245MG p.m1_*404del c.1_1212del1212 central_nervous_system glioma cell-line D- 336MG p.m1_*404del c.1_1212del1212 central_nervous_system glioma cell-line S p.n12fs*6 c.35_53del19 central_nervous_system glioma primary S p.r14fs*26 c.42_52del11 central_nervous_system glioma primary S p.r15fs*9 c.45_46insga central_nervous_system glioma primary A p.r55fs*1 c.165_1212del1048 central_nervous_system glioma cell-line SW p.r55fs*1 c.165_1212del1048 central_nervous_system glioma cell-line U-87- MG p.v54fs*29 c.160_208del49 central_nervous_system glioma NS S p.y16fs*1 c.47_48insa central_nervous_system glioma NS S p.y46* c.138c>g central_nervous_system glioma primary Pubmed ID Mutation ID Sample Source
15 S p.y65* c.195c>a central_nervous_system glioma primary S p.y65* c.195c>g central_nervous_system glioma primary S p.y76fs*1 c.226_227delta central_nervous_system glioma primary S p.y76fs*1 c.227_228delat central_nervous_system glioma primary S p.y76fs*1 c.227_228delat central_nervous_system glioma primary S p.y88fs*3 c.263_264delat central_nervous_system glioma primary LS p.y88fs*3 c.263_264delat central_nervous_system glioma NS S p.a3fs*14 c.9_30del22 endometrium carcinoma primary S p.c71fs*6 c.213_256del44 endometrium carcinoma primary S p.d24fs*19 c.71_72insc endometrium carcinoma primary S p.d24fs*19 c.71_72insc endometrium carcinoma primary S p.e18fs*5 c.54_57delggat endometrium carcinoma primary p.e73fs*25 c.219_222delaaga endometrium carcinoma fresh/frozen S p.e73fs*4 c.219_220delaa endometrium carcinoma primary p.e7fs*3 c.19_20delga endometrium carcinoma NOS p.f56c c.167t>g endometrium carcinoma fresh/frozen S p.f90fs*9 c.270delt endometrium carcinoma primary S p.h64fs*9 c.190_191delca endometrium carcinoma primary S p.i32fs*22 c.96delt endometrium carcinoma primary S p.i32fs*22 c.96delt endometrium carcinoma primary S p.i33fs*20 c.97_100delattg endometrium carcinoma primary S p.i33fs*21 c.97dela endometrium carcinoma primary S p.i33fs*21 c.97dela endometrium carcinoma metastasis E p.k6fs*4 c.17_18delaa endometrium carcinoma fresh/frozen S p.k6fs*4 c.17_18delaa endometrium carcinoma primary p.k6fs*4 c.17_18delaa endometrium carcinoma fresh/frozen S p.l25fs*28 c.75_78delgacc endometrium carcinoma primary p.l57fs*42 c.166_166delt endometrium carcinoma NOS E p.l57fs*6 c.170_171inst endometrium carcinoma fresh/frozen
16 S p.l57fs*6 c.170_171inst endometrium carcinoma primary p.l57fs*6 c.170_171inst endometrium carcinoma fixed - NOS S p.n48fs*6 c.143dela endometrium carcinoma primary S p.n63fs*10 c.187_188delaa endometrium carcinoma primary p.n63fs*10 c.187_188delaa endometrium carcinoma fresh/frozen p.n63fs*10 c.187_188delaa endometrium carcinoma fresh/frozen S p.n63fs*11 c.188_189insa endometrium carcinoma primary S p.n63fs*36 c.187dela endometrium carcinoma primary S p.n63fs*36 c.188dela endometrium carcinoma primary S p.n63fs*36 c.188dela endometrium carcinoma primary S p.n63fs*36 c.188dela endometrium hyperplasia primary S p.n69fs*30 c.206dela endometrium carcinoma primary S p.p30fs*24 c.89delc endometrium carcinoma primary S p.r14fs*10 c.40dela endometrium carcinoma primary S p.r14fs*29 c.41_42insg endometrium carcinoma primary S p.r15fs*28 c.45_46inst endometrium carcinoma primary S p.r55fs*2 c.165_181del17 endometrium carcinoma primary S p.t26fs*28 c.78delc endometrium carcinoma primary S p.v45fs*10 c.134_135insca endometrium carcinoma primary S p.y16fs*21 c.46_65del20 endometrium carcinoma primary S p.y16fs*28 c.46_47inst endometrium carcinoma primary S p.y27fs*1 c.80_90del11 endometrium carcinoma primary S p.y27fs*16 c.80_81delat endometrium carcinoma primary p.y29fs*25 c.83_83delt endometrium carcinoma fresh/frozen S p.y68fs*5 c.202_203delta endometrium carcinoma primary S p.y76fs*1 c.227_228delat endometrium carcinoma primary S p.y76fs*1 c.227_228delat endometrium carcinoma primary OPM p.m1_*404del c.1_1212del1212 haematopoietic_and_lymphoid_tissue haematopoietic_neoplasm cell-line S p.m1fs*7 c.1_8delatgacagc haematopoietic_and_lymphoid_tissue lymphoid_neoplasm primary KE p.y27fs*1 c.80_1212del1133 haematopoietic_and_lymphoid_tissue haematopoietic_neoplasm cell-line p.e7fs*3 c.21_22delga large_intestine carcinoma
17 p.f90fs*9 c.270delt large_intestine carcinoma p.l57fs*42 c.170delt large_intestine carcinoma p.l57fs*6 c.170_171inst large_intestine carcinoma fresh/frozen fresh/frozen fresh/frozen fresh/frozen S p.c83fs*9 c.247_248inst liver carcinoma primary H p.? c.1-?_492+?del lung carcinoma NS NCI- H p.? c.1-?_492+?del lung carcinoma NS SBC p.m1_*404del c.1_1212del1212 lung carcinoma cell-line NCI- H p.m1_*404del c.1_1212del1212 lung carcinoma cell-line N p.m1_?del fs c.1-?_79+?del lung carcinoma NS N p.m1_?del fs c.1-?_79+?del lung carcinoma NS N p.m1_?del fs c.1-?_79+?del lung carcinoma NS NCI- H p.r55fs*1 c.165_1212del1048 lung carcinoma cell-line LU-134- A p.y27fs*1 c.80_1212del1133 lung carcinoma cell-line S p.l57fs*42 c.170delt meninges meningioma primary p.c71fs*28 c.213_213delt ovary carcinoma fresh - NOS p.f81fs*18 c.241_242tt>g ovary carcinoma NOS RTSG p.k6fs*4 c.17_18delaa ovary carcinoma 4929 cell-line S p.y68fs*5 c.202_203delta ovary carcinoma primary S p.g36fs*18 c.107delg prostate carcinoma primary S p.g44fs*11 c.131_139gcgtataca>acagaaagaca prostate carcinoma primary LNCaP- Clone- FGC p.k6fs*4 c.17_18delaa prostate carcinoma 4929 cell-line LNCaP p.k6fs*4 c.16_17delaa prostate carcinoma NS LNCaP p.k6fs*4 c.16_17delaa prostate carcinoma NS LNCaP p.k6fs*4 c.17_18delaa prostate carcinoma NS S p.q87* c.259c>t prostate carcinoma NS PC p.r55fs*1 c.165_1212del1048 prostate carcinoma cell-line
18 S p.y76fs*1 c.226_227delta prostate carcinoma NS FM p.? c.1-?_492+?del skin malignant_melanoma NS S p.k6fs*4 c.17_18delaa skin malignant_melanoma metastasis G-mel p.m1_?del fs c.1-?_79+?del skin malignant_melanoma NS SW p.m1_*404del c.1_1212del1212 soft_tissue liposarcoma cell-line UM-UC p.m1_*404del c.1_1212del1212 urinary_tract carcinoma cell-line
19 Supplementary Table 1. Genotypes of human tumour cell lines used in this study. Supplementary Table 2. Human tumours and tumour cell lines listed in the COSMIC database (www. bearing PTEN mutations predicted to truncate the open reading frame before amino acid 93. Supplementary Figure 1. Homologous recombination and PARP inhibitor sensitivity in HEC1A tumour cells. A) PTEN deficiency in HEC1A cells causes a reduction in RAD51 expression. Total cell lysates from isogenic HEC1A endometrial tumour cells were immunoblotted for PTEN and RAD51. Detection of ACTIN is shown as a loading control. Parental HEC1A cells are shown (PTEN +/+ ) along with two individually derived PTEN -/- lines (KO16 and KO22). B) PTEN deficiency in HEC1A cells causes a reduction in IR-induced nuclear RAD51 focus formation. * p values vs. +IR PTEN +/+ < 0.05 (Student s t test). No other comparisons were statistically significant. C) PTEN deficiency in HEC1A cells sensitises cells to a druglike PARP inhibitor. p values for both PTEN -/- lines vs. PTEN +/+ cells < (twoway ANOVA). D) PTEN deficiency in HEC1A cells causes a reduction in HR using a synthetic HR substrate. * p values null vs. wild type PTEN cells < 0.05 (Student s t test). Supplementary Figure 2. Decreased RAD51 focus formation upon DNA damage in PTEN deficient cells. γh2ax foci (red), RAD51 foci (green) and nuclei (stained with DAPI blue) are shown. Magnification is 1000X.
20 Targeting PTEN deficiency 2 Supplementary Figure 3. PARP inhibitor sensitivity in human tumour cells. See Supplementary Table 1 for genotypes. PTEN mutant lines are shown in red and PTEN wild type lines in black. For each PTEN null line, survival was significantly different than in DU145 and MCF7 cells (p < 0.05, two-way ANOVA). Supplementary Figure 4. RAD51 expression from a retroviral vector used in Figure 4. PC3 cells were infected with virus and cell lysates analysed by western blotting two days later.
Translating DNA repair pathways into therapeutic targets: beyond the BRCA1/2 and PARP inhibitor saga. Jorge S Reis-Filho, MD PhD FRCPath
 Translating DNA repair pathways into therapeutic targets: beyond the BRCA1/2 and PARP inhibitor saga Jorge S Reis-Filho, MD PhD FRCPath Summary How do PARP inhibitors work? Synthetic lethality Potential
Translating DNA repair pathways into therapeutic targets: beyond the BRCA1/2 and PARP inhibitor saga Jorge S Reis-Filho, MD PhD FRCPath Summary How do PARP inhibitors work? Synthetic lethality Potential
The Need for a PARP in vivo Pharmacodynamic Assay
 The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D., Chief Scientific Officer, Trevigen, Inc., Gaithersburg, MD For further infomation, please contact: William Booth, Ph.D. Tel: +44 (0)1235
The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D., Chief Scientific Officer, Trevigen, Inc., Gaithersburg, MD For further infomation, please contact: William Booth, Ph.D. Tel: +44 (0)1235
PARP inhibition basic science and clinical challenge. Thomas Helleday, PhD
 PARP inhibition basic science and clinical challenge Thomas Helleday, PhD Poly (ADP-ribose) Polymerase 1 (PARP1) Reprinted by permission from Macmillan Publishers Ltd: Rouleau M et al. Nat Rev Cancer 2010;10:293-301
PARP inhibition basic science and clinical challenge Thomas Helleday, PhD Poly (ADP-ribose) Polymerase 1 (PARP1) Reprinted by permission from Macmillan Publishers Ltd: Rouleau M et al. Nat Rev Cancer 2010;10:293-301
PARP inhibitors and TEMOZOLAMIDE in BRAIN TUMORS. Idoia Morilla Ruiz
 PARP inhibitors and TEMOZOLAMIDE in BRAIN TUMORS Idoia Morilla Ruiz DNA REPAIR SYSTEM Cancer Sci 105 ( 2014) 370-388 Until recently, treatment efforts have focussed on maximizing the DNA damage (limited
PARP inhibitors and TEMOZOLAMIDE in BRAIN TUMORS Idoia Morilla Ruiz DNA REPAIR SYSTEM Cancer Sci 105 ( 2014) 370-388 Until recently, treatment efforts have focussed on maximizing the DNA damage (limited
PARP Inhibitors in Lung Cancer. Primo N. Lara, Jr., MD Professor of Medicine UC Davis Comprehensive Cancer Center
 PARP Inhibitors in Lung Cancer Primo N. Lara, Jr., MD Professor of Medicine UC Davis Comprehensive Cancer Center Poly (ADP-ribose) Polymerase (PARP): Mechanism of Action PARPs: family of enzymes that repair
PARP Inhibitors in Lung Cancer Primo N. Lara, Jr., MD Professor of Medicine UC Davis Comprehensive Cancer Center Poly (ADP-ribose) Polymerase (PARP): Mechanism of Action PARPs: family of enzymes that repair
Hereditary Ovarian cancer: BRCA1 and BRCA2. Karen H. Lu MD September 22, 2013
 Hereditary Ovarian cancer: BRCA1 and BRCA2 Karen H. Lu MD September 22, 2013 Outline Hereditary Breast and Ovarian Cancer (HBOC) BRCA1/2 genes How to identify What it means to you What it means to your
Hereditary Ovarian cancer: BRCA1 and BRCA2 Karen H. Lu MD September 22, 2013 Outline Hereditary Breast and Ovarian Cancer (HBOC) BRCA1/2 genes How to identify What it means to you What it means to your
The role of PARP inhibitors in high grade serous ovarian cancers
 The role of PARP inhibitors in high grade serous ovarian cancers Jonathan Ledermann UCL Cancer Institute University College London ANZGOG-ASGO, Canberra, March 214 Cancer Research UK UCL Centre DNA Repair
The role of PARP inhibitors in high grade serous ovarian cancers Jonathan Ledermann UCL Cancer Institute University College London ANZGOG-ASGO, Canberra, March 214 Cancer Research UK UCL Centre DNA Repair
HT F Homogeneous PARP Inhibition Assay Kit. HT F Homogeneous PARP Inhibition Assay Kit. 96 Tests. Table of Contents.
 For Research Use Only. Not For Use In Diagnostic Procedures HT F Homogeneous PARP Inhibition Assay Kit 96 Tests HT F Homogeneous PARP Inhibition Assay Kit Cat# 4690-096-K Cat# 4690-096-K Table of Contents
For Research Use Only. Not For Use In Diagnostic Procedures HT F Homogeneous PARP Inhibition Assay Kit 96 Tests HT F Homogeneous PARP Inhibition Assay Kit Cat# 4690-096-K Cat# 4690-096-K Table of Contents
High-quality genomic DNA isolation and sensitive mutation analysis
 Application Note High-quality genomic DNA isolation and sensitive mutation analysis Izabela Safin, Ivonne Schröder-Stumberger and Peter Porschewski Introduction A major objective of cancer research is
Application Note High-quality genomic DNA isolation and sensitive mutation analysis Izabela Safin, Ivonne Schröder-Stumberger and Peter Porschewski Introduction A major objective of cancer research is
PARP INHIBITORS IN OVARIAN CANCER. A 2011 PERSPECTIVE
 PARP INHIBITORS IN OVARIAN CANCER. A 2011 PERSPECTIVE Prof S.B. Kaye Royal Marsden Hospital, London MD Anderson Cancer Centre December 2011 DISCLOSURES Received honoraria as member of Advisory Boards to
PARP INHIBITORS IN OVARIAN CANCER. A 2011 PERSPECTIVE Prof S.B. Kaye Royal Marsden Hospital, London MD Anderson Cancer Centre December 2011 DISCLOSURES Received honoraria as member of Advisory Boards to
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation
 OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
GeneCopoeia Genome Editing Tools for Safe Harbor Integration in. Mice and Humans. Ed Davis, Liuqing Qian, Ruiqing li, Junsheng Zhou, and Jinkuo Zhang
 G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
SUPPLEMENTARY DATA 1
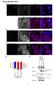 SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
PREPARED FOR: U.S. Army Medical Research and Materiel CommandFort Detrick, Maryland 21702 5012
 AD Award Number: W81XWH 07 1 0542 TITLE: Is Nuclear Structure Altered in Breast Cancer Cells? PRINCIPAL INVESTIGATOR: Han Htun, Ph.D. CONTRACTING ORGANIZATION: University of California Los Angeles, CA
AD Award Number: W81XWH 07 1 0542 TITLE: Is Nuclear Structure Altered in Breast Cancer Cells? PRINCIPAL INVESTIGATOR: Han Htun, Ph.D. CONTRACTING ORGANIZATION: University of California Los Angeles, CA
Notch 1 -dependent regulation of cell fate in colorectal cancer
 Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
Investigating the role of a Cryptosporidium parum apyrase in infection
 Investigating the role of a Cryptosporidium parum apyrase in infection David Riccardi and Patricio Manque Abstract This project attempted to characterize the function of a Cryptosporidium parvum apyrase
Investigating the role of a Cryptosporidium parum apyrase in infection David Riccardi and Patricio Manque Abstract This project attempted to characterize the function of a Cryptosporidium parvum apyrase
CytoSelect 48-Well Cell Adhesion Assay (ECM Array, Colorimetric Format)
 Product Manual CytoSelect 48-Well Cell Adhesion Assay (ECM Array, Colorimetric Format) Catalog Number CBA-070 CBA-070-5 48 assays 5 x 48 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures
Product Manual CytoSelect 48-Well Cell Adhesion Assay (ECM Array, Colorimetric Format) Catalog Number CBA-070 CBA-070-5 48 assays 5 x 48 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures
Pharmacogenomic Approaches. Luis Paz-Ares Hospital Universitario Virgen del Rocio Seville, Spain
 Pharmacogenomic Approaches Luis Paz-Ares Hospital Universitario Virgen del Rocio Seville, Spain Pharmacogenetics & Pharmacogenomics Medicine tailored to the individual Genetic information, including the
Pharmacogenomic Approaches Luis Paz-Ares Hospital Universitario Virgen del Rocio Seville, Spain Pharmacogenetics & Pharmacogenomics Medicine tailored to the individual Genetic information, including the
a Phase 2 prostate cancer clinical trial is ongoing. Table 2: Squalamine vs Standard-of-care literature
 PRODUCT FACT SHEET Spring 2007 MISSION STATEMENT Genaera Corporation is a biopharmaceutical company with a focus on metabolic and respiratory diseases. The compounds in the Genaera pipeline address signal
PRODUCT FACT SHEET Spring 2007 MISSION STATEMENT Genaera Corporation is a biopharmaceutical company with a focus on metabolic and respiratory diseases. The compounds in the Genaera pipeline address signal
Médecine de précision médecine personnalisée en Oncologie. Fabien Calvo, Directeur Recherche et Innovation, INCa, Directeur ITMO Cancer, AVIESAN
 Médecine de précision médecine personnalisée en Oncologie Fabien Calvo, Directeur Recherche et Innovation, INCa, Directeur ITMO Cancer, AVIESAN Successful targeted drug development Rapid identification
Médecine de précision médecine personnalisée en Oncologie Fabien Calvo, Directeur Recherche et Innovation, INCa, Directeur ITMO Cancer, AVIESAN Successful targeted drug development Rapid identification
Supplemental Information. McBrayer et al. Supplemental Data
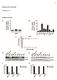 1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
Angelo Peschiaroli, N. Valerio Dorrello, Daniele Guardavaccaro, Monica Venere, Thanos Halazonetis, Nicholas E. Sherman, and Michele Pagano
 Molecular Cell, Volume 23 Supplemental Data SCF βtrcp -Mediated Degradation of Claspin Regulates Recovery from the DNA Replication Checkpoint Response Angelo Peschiaroli, N. Valerio Dorrello, Daniele Guardavaccaro,
Molecular Cell, Volume 23 Supplemental Data SCF βtrcp -Mediated Degradation of Claspin Regulates Recovery from the DNA Replication Checkpoint Response Angelo Peschiaroli, N. Valerio Dorrello, Daniele Guardavaccaro,
micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved
 microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
PNA BRAF Mutation Detection Kit
 - PNA BRAF Mutation Detection Kit Catalog Number KA2102 50 tests/kit Version: 01 Intended for research use only www.abnova.com Introduction and Background Intended use The PNA BRAF Mutation Detection Kit
- PNA BRAF Mutation Detection Kit Catalog Number KA2102 50 tests/kit Version: 01 Intended for research use only www.abnova.com Introduction and Background Intended use The PNA BRAF Mutation Detection Kit
2.1.2 Characterization of antiviral effect of cytokine expression on HBV replication in transduced mouse hepatocytes line
 i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
White paper Evaluation of BRAF (V600E) Mutation by Immunohistochemical Staining with anti-braf V600E (VE1) Antibody: A Comparison with Sanger
 White paper Evaluation of BRAF (V600E) Mutation by Immunohistochemical Staining with anti-braf V600E (VE1) Antibody: A Comparison with Sanger Sequencing 2 Evaluation of BRAF (V600E) Mutation by Immunohistochemical
White paper Evaluation of BRAF (V600E) Mutation by Immunohistochemical Staining with anti-braf V600E (VE1) Antibody: A Comparison with Sanger Sequencing 2 Evaluation of BRAF (V600E) Mutation by Immunohistochemical
TOOLS sirna and mirna. User guide
 TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
The Essentials of. Globally Delivered
 ATCC cell lines BY Gene Mutation The Essentials of Life Science Research Globally Delivered Cell Culture Guides For over 85 years, ATCC has been a leader in preserving and culturing all types of biological
ATCC cell lines BY Gene Mutation The Essentials of Life Science Research Globally Delivered Cell Culture Guides For over 85 years, ATCC has been a leader in preserving and culturing all types of biological
New Directions in Treatment of Ovarian Cancer. Amit M. Oza Princess Margaret Hospital University of Toronto
 New Directions in Treatment of Ovarian Cancer Amit M. Oza Princess Margaret Hospital University of Toronto Newly diagnosed: scenario Ist line Surgery chemotherapy Cure If can t cure can we turn into chronic
New Directions in Treatment of Ovarian Cancer Amit M. Oza Princess Margaret Hospital University of Toronto Newly diagnosed: scenario Ist line Surgery chemotherapy Cure If can t cure can we turn into chronic
Targeting Specific Cell Signaling Pathways for the Treatment of Malignant Peritoneal Mesothelioma
 The Use of Kinase Inhibitors: Translational Lab Results Targeting Specific Cell Signaling Pathways for the Treatment of Malignant Peritoneal Mesothelioma Sheelu Varghese, Ph.D. H. Richard Alexander, M.D.
The Use of Kinase Inhibitors: Translational Lab Results Targeting Specific Cell Signaling Pathways for the Treatment of Malignant Peritoneal Mesothelioma Sheelu Varghese, Ph.D. H. Richard Alexander, M.D.
New developments and controversies in breast cancer treatment: PARP Inhibitors: a breakthrough?
 New developments and controversies in breast cancer treatment: PARP Inhibitors: a breakthrough? F. Cardoso, MD Champalimaud Cancer Center Lisbon, Portugal BBM 2010 Thank you to A Tutt & PRIME Oncology
New developments and controversies in breast cancer treatment: PARP Inhibitors: a breakthrough? F. Cardoso, MD Champalimaud Cancer Center Lisbon, Portugal BBM 2010 Thank you to A Tutt & PRIME Oncology
Contents. molecular biology techniques. - Mutations in Factor II. - Mutations in MTHFR gene. - Breast cencer genes. - p53 and breast cancer
 Contents Introduction: biology and medicine, two separated compartments What we need to know: - boring basics in DNA/RNA structure and overview of particular aspects of molecular biology techniques - How
Contents Introduction: biology and medicine, two separated compartments What we need to know: - boring basics in DNA/RNA structure and overview of particular aspects of molecular biology techniques - How
Common Cancers & Hereditary Syndromes
 Common Cancers & Hereditary Syndromes Elizabeth Hoodfar, MS, LCGC Regional Cancer Genetics Coordinator Kaiser Permanente Northern California Detect clinical characteristics of hereditary cancer syndromes.
Common Cancers & Hereditary Syndromes Elizabeth Hoodfar, MS, LCGC Regional Cancer Genetics Coordinator Kaiser Permanente Northern California Detect clinical characteristics of hereditary cancer syndromes.
Protein kinase C alpha expression and resistance to neo-adjuvant gemcitabine-containing chemotherapy in non-small cell lung cancer
 Protein kinase C alpha expression and resistance to neo-adjuvant gemcitabine-containing chemotherapy in non-small cell lung cancer Dan Vogl Lay Abstract Early stage non-small cell lung cancer can be cured
Protein kinase C alpha expression and resistance to neo-adjuvant gemcitabine-containing chemotherapy in non-small cell lung cancer Dan Vogl Lay Abstract Early stage non-small cell lung cancer can be cured
Endpoint Selection in Phase II Oncology trials
 Endpoint Selection in Phase II Oncology trials Anastasia Ivanova Department of Biostatistics UNC at Chapel Hill [email protected] Primary endpoints in Phase II trials Recently looked at journal articles
Endpoint Selection in Phase II Oncology trials Anastasia Ivanova Department of Biostatistics UNC at Chapel Hill [email protected] Primary endpoints in Phase II trials Recently looked at journal articles
A 70-year old Man with Pleural Effusion
 Mesothelioma Diagnosis: Pitfalls and Latest Updates S Klebe and DW Henderson Recommendations Indisputable malignant cells on cytomorphological criteria which demonstrate a mesothelial phenotype, which
Mesothelioma Diagnosis: Pitfalls and Latest Updates S Klebe and DW Henderson Recommendations Indisputable malignant cells on cytomorphological criteria which demonstrate a mesothelial phenotype, which
Non Small Cell Lung Cancer: Scientific Discoveries and the Pursuit of Progress
 Non Small Cell Lung Cancer: Scientific Discoveries and the Pursuit of Progress Lung Cancer Accounts for 14% of All New Cancer Diagnoses in the United States 1 Lung cancer is the second most common malignancy
Non Small Cell Lung Cancer: Scientific Discoveries and the Pursuit of Progress Lung Cancer Accounts for 14% of All New Cancer Diagnoses in the United States 1 Lung cancer is the second most common malignancy
The cell lines used in this study were obtained from the American Type Culture
 Supplementary materials and methods Cell culture and drug treatments The cell lines used in this study were obtained from the American Type Culture Collection (ATCC) and grown as previously described.
Supplementary materials and methods Cell culture and drug treatments The cell lines used in this study were obtained from the American Type Culture Collection (ATCC) and grown as previously described.
Monoclonal Antibodies in Cancer. Ralph Schwall, PhD Associate Director, Translational Oncology Genentech, Inc.
 Monoclonal Antibodies in Cancer Ralph Schwall, PhD Associate Director, Translational Oncology Genentech, Inc. Disclaimer I had nothing to do with Herceptin Using lessons learned in new antibody projects
Monoclonal Antibodies in Cancer Ralph Schwall, PhD Associate Director, Translational Oncology Genentech, Inc. Disclaimer I had nothing to do with Herceptin Using lessons learned in new antibody projects
BAP1 germline mutations A new Cutaneous Nevus Melanoma Syndrome. Thomas Wiesner
 BAP1 germline mutations A new Cutaneous Nevus Melanoma Syndrome Thomas Wiesner Disclosure Listed as co-inventor US patent application US 61/463,389 BAP1 mutational analysis in determining susceptibility
BAP1 germline mutations A new Cutaneous Nevus Melanoma Syndrome Thomas Wiesner Disclosure Listed as co-inventor US patent application US 61/463,389 BAP1 mutational analysis in determining susceptibility
Lecture 3: Mutations
 Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
Future Directions in Clinical Research. Karen Kelly, MD Associate Director for Clinical Research UC Davis Cancer Center
 Future Directions in Clinical Research Karen Kelly, MD Associate Director for Clinical Research UC Davis Cancer Center Outline 1. Status of Cancer Treatment 2. Overview of Clinical Research at UCDCC 3.
Future Directions in Clinical Research Karen Kelly, MD Associate Director for Clinical Research UC Davis Cancer Center Outline 1. Status of Cancer Treatment 2. Overview of Clinical Research at UCDCC 3.
PART 3.3: MicroRNA and Cancer
 BIBM 2010 Tutorial: Epigenomics and Cancer PART 3.3: MicroRNA and Cancer Dec 18, 2010 Sun Kim at Indiana University Outline of Part 3.3 Background on microrna Role of microrna in cancer MicroRNA pathway
BIBM 2010 Tutorial: Epigenomics and Cancer PART 3.3: MicroRNA and Cancer Dec 18, 2010 Sun Kim at Indiana University Outline of Part 3.3 Background on microrna Role of microrna in cancer MicroRNA pathway
S1 Text. Modeling deterministic single-cell microrna-p53-mdm2 network Figure 2 Figure 2
 S1 Text. Modeling deterministic single-cell microrna-p53-mdm2 network The schematic diagram of the microrna-p53-mdm2 oscillator is illustrated in Figure 2. The interaction scheme among the mrnas and the
S1 Text. Modeling deterministic single-cell microrna-p53-mdm2 network The schematic diagram of the microrna-p53-mdm2 oscillator is illustrated in Figure 2. The interaction scheme among the mrnas and the
CONTRACTING ORGANIZATION: University of Alabama at Birmingham Birmingham, AL 35294
 AD Award Number: W81XWH-08-1-0030 TITLE: Regulation of Prostate Cancer Bone Metastasis by DKK1 PRINCIPAL INVESTIGATOR: Gregory A. Clines, M.D., Ph.D. CONTRACTING ORGANIZATION: University of Alabama at
AD Award Number: W81XWH-08-1-0030 TITLE: Regulation of Prostate Cancer Bone Metastasis by DKK1 PRINCIPAL INVESTIGATOR: Gregory A. Clines, M.D., Ph.D. CONTRACTING ORGANIZATION: University of Alabama at
Publikationsliste Claudia Götz
 Publikationsliste Claudia Götz 1. Reinhard,B., Götz, C., and Faillard, H.: Synthesis of N-Acetyl-9-Oacetylneuraminic acid α-p-aminophenylthioketoside and its application as ligand in the affinity chromatography
Publikationsliste Claudia Götz 1. Reinhard,B., Götz, C., and Faillard, H.: Synthesis of N-Acetyl-9-Oacetylneuraminic acid α-p-aminophenylthioketoside and its application as ligand in the affinity chromatography
Summary of Discussion on Non-clinical Pharmacology Studies on Anticancer Drugs
 Provisional Translation (as of January 27, 2014)* November 15, 2013 Pharmaceuticals and Bio-products Subcommittees, Science Board Summary of Discussion on Non-clinical Pharmacology Studies on Anticancer
Provisional Translation (as of January 27, 2014)* November 15, 2013 Pharmaceuticals and Bio-products Subcommittees, Science Board Summary of Discussion on Non-clinical Pharmacology Studies on Anticancer
Chapter 18: Applications of Immunology
 Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
Instruction manual for product # PNAC-2001. Version 1.2
 PNAClamp BRAF Mutation Detection Kit For in vitro diagnostic use Instruction manual for product # PNAC-2001 Version 1.2 Store at -20 C. Instruction Version: Ver. 1.2 Date of Revision: 2012 February 1 /
PNAClamp BRAF Mutation Detection Kit For in vitro diagnostic use Instruction manual for product # PNAC-2001 Version 1.2 Store at -20 C. Instruction Version: Ver. 1.2 Date of Revision: 2012 February 1 /
KMS-Specialist & Customized Biosimilar Service
 KMS-Specialist & Customized Biosimilar Service 1. Polyclonal Antibody Development Service KMS offering a variety of Polyclonal Antibody Services to fit your research and production needs. we develop polyclonal
KMS-Specialist & Customized Biosimilar Service 1. Polyclonal Antibody Development Service KMS offering a variety of Polyclonal Antibody Services to fit your research and production needs. we develop polyclonal
Breast cancer and the role of low penetrance alleles: a focus on ATM gene
 Modena 18-19 novembre 2010 Breast cancer and the role of low penetrance alleles: a focus on ATM gene Dr. Laura La Paglia Breast Cancer genetic Other BC susceptibility genes TP53 PTEN STK11 CHEK2 BRCA1
Modena 18-19 novembre 2010 Breast cancer and the role of low penetrance alleles: a focus on ATM gene Dr. Laura La Paglia Breast Cancer genetic Other BC susceptibility genes TP53 PTEN STK11 CHEK2 BRCA1
ab185915 Protein Sumoylation Assay Ultra Kit
 ab185915 Protein Sumoylation Assay Ultra Kit Instructions for Use For the measuring in vivo protein sumoylation in various samples This product is for research use only and is not intended for diagnostic
ab185915 Protein Sumoylation Assay Ultra Kit Instructions for Use For the measuring in vivo protein sumoylation in various samples This product is for research use only and is not intended for diagnostic
pcas-guide System Validation in Genome Editing
 pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
Targeted Therapy What the Surgeon Needs to Know
 Targeted Therapy What the Surgeon Needs to Know AATS Focus in Thoracic Surgery 2014 David R. Jones, M.D. Professor & Chief, Thoracic Surgery Memorial Sloan Kettering Cancer Center I have no disclosures
Targeted Therapy What the Surgeon Needs to Know AATS Focus in Thoracic Surgery 2014 David R. Jones, M.D. Professor & Chief, Thoracic Surgery Memorial Sloan Kettering Cancer Center I have no disclosures
Clinical Trial Designs for Incorporating Multiple Biomarkers in Combination Studies with Targeted Agents
 Clinical Trial Designs for Incorporating Multiple Biomarkers in Combination Studies with Targeted Agents J. Jack Lee, Ph.D. Department of Biostatistics 3 Primary Goals for Clinical Trials Test safety and
Clinical Trial Designs for Incorporating Multiple Biomarkers in Combination Studies with Targeted Agents J. Jack Lee, Ph.D. Department of Biostatistics 3 Primary Goals for Clinical Trials Test safety and
Dal germinale al somatico nella identificazione di tumori ereditari
 Modena 18-19 novembre 2010 Dal germinale al somatico nella identificazione di tumori ereditari Laura Ottini Tendencies to develop cancer can be inherited Fletcher & Houlston, 2010 Cancer is a genetic disease
Modena 18-19 novembre 2010 Dal germinale al somatico nella identificazione di tumori ereditari Laura Ottini Tendencies to develop cancer can be inherited Fletcher & Houlston, 2010 Cancer is a genetic disease
Accurate and sensitive mutation detection and quantitation using TaqMan Mutation Detection Assays for disease research
 PPLICTION NOTE Mutation Detection ssays ccurate and sensitive mutation detection and quantitation using Mutation Detection ssays for disease research In this research study, we addressed the feasibility
PPLICTION NOTE Mutation Detection ssays ccurate and sensitive mutation detection and quantitation using Mutation Detection ssays for disease research In this research study, we addressed the feasibility
No Disclosures. Learning Objectives 10/25/13
 No Disclosures Gregory A. Brent, MD Departments of Medicine and Physiology David Geffen School of Medicine at UCLA VA Greater Los Angeles Healthcare System Learning Objectives Describe the pathways that
No Disclosures Gregory A. Brent, MD Departments of Medicine and Physiology David Geffen School of Medicine at UCLA VA Greater Los Angeles Healthcare System Learning Objectives Describe the pathways that
OI PARP ΑΝΑΣΤΟΛΕΙΣ ΣΤΟΝ ΚΑΡΚΙΝΟ ΤΟΥ ΜΑΣΤΟΥ ΝΙΚΟΛΑΙΔΗ ΑΔΑΜΑΝΤΙΑ ΠΑΘΟΛΟΓΟΣ-ΟΓΚΟΛΟΓΟΣ Β ΟΓΚΟΛΟΓΙΚΗ ΚΛΙΝΙΚΗ ΝΟΣ. ΜΗΤΕΡΑ
 OI PARP ΑΝΑΣΤΟΛΕΙΣ ΣΤΟΝ ΚΑΡΚΙΝΟ ΤΟΥ ΜΑΣΤΟΥ ΝΙΚΟΛΑΙΔΗ ΑΔΑΜΑΝΤΙΑ ΠΑΘΟΛΟΓΟΣ-ΟΓΚΟΛΟΓΟΣ Β ΟΓΚΟΛΟΓΙΚΗ ΚΛΙΝΙΚΗ ΝΟΣ. ΜΗΤΕΡΑ Study Overview Inhibition of poly(adenosine diphosphate [ADP]-ribose) polymerase
OI PARP ΑΝΑΣΤΟΛΕΙΣ ΣΤΟΝ ΚΑΡΚΙΝΟ ΤΟΥ ΜΑΣΤΟΥ ΝΙΚΟΛΑΙΔΗ ΑΔΑΜΑΝΤΙΑ ΠΑΘΟΛΟΓΟΣ-ΟΓΚΟΛΟΓΟΣ Β ΟΓΚΟΛΟΓΙΚΗ ΚΛΙΝΙΚΗ ΝΟΣ. ΜΗΤΕΡΑ Study Overview Inhibition of poly(adenosine diphosphate [ADP]-ribose) polymerase
Gemcitabine, Paclitaxel, and Trastuzumab in Metastatic Breast Cancer
 Gemcitabine, Paclitaxel, and Trastuzumab in Metastatic Breast Cancer Review Article [1] December 01, 2003 By George W. Sledge, Jr, MD [2] Gemcitabine (Gemzar) and paclitaxel show good activity as single
Gemcitabine, Paclitaxel, and Trastuzumab in Metastatic Breast Cancer Review Article [1] December 01, 2003 By George W. Sledge, Jr, MD [2] Gemcitabine (Gemzar) and paclitaxel show good activity as single
Applications of comprehensive clinical genomic analysis in solid tumors: obstacles and opportunities
 Applications of comprehensive clinical genomic analysis in solid tumors: obstacles and opportunities Vincent A. Miller, M.D. Foundation Medicine, Inc. AACR Annual Meeting 2012 Current Concepts session
Applications of comprehensive clinical genomic analysis in solid tumors: obstacles and opportunities Vincent A. Miller, M.D. Foundation Medicine, Inc. AACR Annual Meeting 2012 Current Concepts session
ALCHEMIST (Adjuvant Lung Cancer Enrichment Marker Identification and Sequencing Trials)
 ALCHEMIST (Adjuvant Lung Cancer Enrichment Marker Identification and Sequencing Trials) 3 Integrated Trials Testing Targeted Therapy in Early Stage Lung Cancer Part of NCI s Precision Medicine Effort in
ALCHEMIST (Adjuvant Lung Cancer Enrichment Marker Identification and Sequencing Trials) 3 Integrated Trials Testing Targeted Therapy in Early Stage Lung Cancer Part of NCI s Precision Medicine Effort in
RNA Viruses. A Practical Approac h. Alan J. Cann
 RNA Viruses A Practical Approac h Alan J. Cann List of protocols page xiii Abbreviations xvii Investigation of RNA virus genome structure 1 A j. Easton, A.C. Marriott and C.R. Pringl e 1 Introduction-the
RNA Viruses A Practical Approac h Alan J. Cann List of protocols page xiii Abbreviations xvii Investigation of RNA virus genome structure 1 A j. Easton, A.C. Marriott and C.R. Pringl e 1 Introduction-the
Chapter 11. Rapid Screening of Gli2/3 Mutants Using the Flp-In System. Pawel Niewiadomski and Rajat Rohatgi. Abstract.
 Chapter 11 Rapid Screening of Gli2/3 Mutants Using the Flp-In System Abstract Gli2 and Gli3 respond to the Hedgehog (Hh) signal in mammals by undergoing posttranslational modifications and moving to the
Chapter 11 Rapid Screening of Gli2/3 Mutants Using the Flp-In System Abstract Gli2 and Gli3 respond to the Hedgehog (Hh) signal in mammals by undergoing posttranslational modifications and moving to the
THE CANCER STEM CELL INHIBITORS VS-6063 AND VS-5584 EXHIBIT SYNERGISTIC ANTICANCER ACTIVITY IN PRECLINICAL MODELS OF MESOTHELIOMA
 THE CANCER STEM CELL INHIBITORS VS-6063 AND VS-5584 EXHIBIT SYNERGISTIC ANTICANCER ACTIVITY IN PRECLINICAL MODELS OF MESOTHELIOMA Mitchell Keegan, Ph.D. Vice President of Development, Verastem, Inc. 1
THE CANCER STEM CELL INHIBITORS VS-6063 AND VS-5584 EXHIBIT SYNERGISTIC ANTICANCER ACTIVITY IN PRECLINICAL MODELS OF MESOTHELIOMA Mitchell Keegan, Ph.D. Vice President of Development, Verastem, Inc. 1
DNA Fingerprinting. Unless they are identical twins, individuals have unique DNA
 DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE Q5B
 INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
CD3/TCR stimulation and surface detection Determination of specificity of intracellular detection of IL-7Rα by flow cytometry
 CD3/TCR stimulation and surface detection Stimulation of HPB-ALL cells with the anti-cd3 monoclonal antibody OKT3 was performed as described 3. In brief, antibody-coated plates were prepared by incubating
CD3/TCR stimulation and surface detection Stimulation of HPB-ALL cells with the anti-cd3 monoclonal antibody OKT3 was performed as described 3. In brief, antibody-coated plates were prepared by incubating
Guide for Data Visualization and Analysis using ACSN
 Guide for Data Visualization and Analysis using ACSN ACSN contains the NaviCell tool box, the intuitive and user- friendly environment for data visualization and analysis. The tool is accessible from the
Guide for Data Visualization and Analysis using ACSN ACSN contains the NaviCell tool box, the intuitive and user- friendly environment for data visualization and analysis. The tool is accessible from the
Outline. Workup for metastatic breast cancer. Metastatic breast cancer
 Metastatic breast cancer Immunostain Update: Diagnosis of metastatic breast carcinoma, emphasizing distinction from GYN primary 1/3 of breast cancer patients will show metastasis 1 st presentation or 20-30
Metastatic breast cancer Immunostain Update: Diagnosis of metastatic breast carcinoma, emphasizing distinction from GYN primary 1/3 of breast cancer patients will show metastasis 1 st presentation or 20-30
New Advances in Cancer Treatments. March 2015
 New Advances in Cancer Treatments March 2015 Safe Harbour Statement This presentation document contains certain forward-looking statements and information (collectively, forward-looking statements ) within
New Advances in Cancer Treatments March 2015 Safe Harbour Statement This presentation document contains certain forward-looking statements and information (collectively, forward-looking statements ) within
Preliminary Results from a Phase 2 Study of ARQ 197 in Patients with Microphthalmia Transcription Factor Family (MiT) Associated Tumors
 Preliminary Results from a Phase 2 Study of ARQ 197 in Patients with Microphthalmia Transcription Factor Family (MiT) Associated Tumors John Goldberg 1 *, George Demetri 2, Edwin Choy 3, Lee Rosen 4, Alberto
Preliminary Results from a Phase 2 Study of ARQ 197 in Patients with Microphthalmia Transcription Factor Family (MiT) Associated Tumors John Goldberg 1 *, George Demetri 2, Edwin Choy 3, Lee Rosen 4, Alberto
Inhibition of MicroRNA Sensitizes 3D Breast Cancer Microtissues to Radiation Therapy
 CASE STUDY High Content Image Analysis Inhibition of MicroRNA Sensitizes 3D Breast Cancer Microtissues to Radiation Therapy Dr. Nataša Anastasov leads the Personalized Radiation Therapy Group within the
CASE STUDY High Content Image Analysis Inhibition of MicroRNA Sensitizes 3D Breast Cancer Microtissues to Radiation Therapy Dr. Nataša Anastasov leads the Personalized Radiation Therapy Group within the
Chapter 8. Summary and Perspectives
 Chapter 8 Summary and Perspectives 131 Chapter 8 Summary Overexpression of the multidrug resistance protein MRP1 confer multidrug resistance (MDR) to cancer cells. The contents of this thesis describe
Chapter 8 Summary and Perspectives 131 Chapter 8 Summary Overexpression of the multidrug resistance protein MRP1 confer multidrug resistance (MDR) to cancer cells. The contents of this thesis describe
Introduction To Real Time Quantitative PCR (qpcr)
 Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
European Medicines Agency
 European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
Ovarian Cancer and Modern Immunotherapy: Regulatory Strategies for Drug Development
 Ovarian Cancer and Modern Immunotherapy: Regulatory Strategies for Drug Development Sanjeeve Bala, MD, MPH Ovarian Cancer Endpoints Workshop FDA White Oak September 3, 2015 Overview Immune agents from
Ovarian Cancer and Modern Immunotherapy: Regulatory Strategies for Drug Development Sanjeeve Bala, MD, MPH Ovarian Cancer Endpoints Workshop FDA White Oak September 3, 2015 Overview Immune agents from
Understanding the immune response to bacterial infections
 Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Evaluating the Efficiency of Targeted Designs for Randomized Clinical Trials
 Vol. 0, 6759 6763, October 5, 004 Clinical Cancer Research 6759 Perspective Evaluating the Efficiency of Targeted Designs for Randomized Clinical Trials Richard Simon and Aboubakar Maitournam Biometric
Vol. 0, 6759 6763, October 5, 004 Clinical Cancer Research 6759 Perspective Evaluating the Efficiency of Targeted Designs for Randomized Clinical Trials Richard Simon and Aboubakar Maitournam Biometric
What is Cancer? Cancer is a genetic disease: Cancer typically involves a change in gene expression/function:
 Cancer is a genetic disease: Inherited cancer Sporadic cancer What is Cancer? Cancer typically involves a change in gene expression/function: Qualitative change Quantitative change Any cancer causing genetic
Cancer is a genetic disease: Inherited cancer Sporadic cancer What is Cancer? Cancer typically involves a change in gene expression/function: Qualitative change Quantitative change Any cancer causing genetic
Protein Expression. A Practical Approach J. HIGGIN S
 Protein Expression A Practical Approach S. J. HIGGIN S B. D. HAMES List of contributors Abbreviations xv Xvi i 1. Protein expression in mammalian cell s Marlies Otter-Nilsson and Tommy Nilsso n 1. Introduction
Protein Expression A Practical Approach S. J. HIGGIN S B. D. HAMES List of contributors Abbreviations xv Xvi i 1. Protein expression in mammalian cell s Marlies Otter-Nilsson and Tommy Nilsso n 1. Introduction
ABSTRACT. n engl j med 361;2 nejm.org july 9, 2009 123
 The new england journal of medicine established in 1812 july 9, 29 vol. 361 no. 2 Inhibition of Poly(ADP-Ribose) Polymerase in Tumors from BRCA Mutation Carriers Peter C. Fong, M.D., David S. Boss, M.Sc.,
The new england journal of medicine established in 1812 july 9, 29 vol. 361 no. 2 Inhibition of Poly(ADP-Ribose) Polymerase in Tumors from BRCA Mutation Carriers Peter C. Fong, M.D., David S. Boss, M.Sc.,
treatments) worked by killing cancerous cells using chemo or radiotherapy. While these techniques can
 Shristi Pandey Genomics and Medicine Winter 2011 Prof. Doug Brutlag Chronic Myeloid Leukemia: A look into how genomics is changing the way we treat Cancer. Until the late 1990s, nearly all treatment methods
Shristi Pandey Genomics and Medicine Winter 2011 Prof. Doug Brutlag Chronic Myeloid Leukemia: A look into how genomics is changing the way we treat Cancer. Until the late 1990s, nearly all treatment methods
RENAL CANCER PATHOLOGY WHAT REALLY MATTERS? STEWART FLEMING UNIVERSITY OF DUNDEE
 RENAL CANCER PATHOLOGY WHAT REALLY MATTERS? STEWART FLEMING UNIVERSITY OF DUNDEE MAJOR PARADIGM SHIFT IN EARLY 1990S IN UNDERSTANDING RENAL CANCER Molecular differential pathology of renal cell tumours
RENAL CANCER PATHOLOGY WHAT REALLY MATTERS? STEWART FLEMING UNIVERSITY OF DUNDEE MAJOR PARADIGM SHIFT IN EARLY 1990S IN UNDERSTANDING RENAL CANCER Molecular differential pathology of renal cell tumours
PCR was carried out in a reaction volume of 20 µl using the ABI AmpliTaq GOLD kit (ABI,
 Supplemental Text/Tables PCR Amplification and Sequencing PCR was carried out in a reaction volume of 20 µl using the ABI AmpliTaq GOLD kit (ABI, Foster City, CA). Each PCR reaction contained 20 ng genomic
Supplemental Text/Tables PCR Amplification and Sequencing PCR was carried out in a reaction volume of 20 µl using the ABI AmpliTaq GOLD kit (ABI, Foster City, CA). Each PCR reaction contained 20 ng genomic
