Accurate and sensitive mutation detection and quantitation using TaqMan Mutation Detection Assays for disease research
|
|
|
- Cecilia Bryant
- 9 years ago
- Views:
Transcription
1 PPLICTION NOTE Mutation Detection ssays ccurate and sensitive mutation detection and quantitation using Mutation Detection ssays for disease research In this research study, we addressed the feasibility of obtaining sensitive, accurate, and reproducible mutation detection results in synthetic templates, cell lines, and FFPE tissue samples using Mutation Detection ssays. nalytical performance assessed with synthetic templates and genomic DN (gdn) extracted from cell lines or model FFPE cell lines indicates that Mutation Detection ssays have a minimum of 0.1% sensitivity and can detect fewer than 10 copies of mutant DN sequence in a background of wild type gdn. Our data indicate that Mutation Detection ssays can accurately detect mutation status and quantify the percent mutation with a relative standard deviation (CV) <20%. dditionally, benchmarking to an established qpcr assay technology for mutation detection demonstrates that the Mutation Detection ssays have a wider detection range and superior limit of detection. Introduction Somatic mutations can be present in low abundance within a very high background of wild type sequences. Many mutation detection methods compatible with tumor specimens have been reported in the literature and/or are commercially available. Methods that are commonly used are gene sequencing (including pyrosequencing and traditional Sanger sequencing) and real-time PCR. Realtime PCR based technologies include high-resolution melt curve analysis, peptide nucleic acid (PN) clamping, and allele-specific amplification with bifunctional fluorescent primer/probe detection (RMS/Scorpion). However, these commercially available kits have various limitations in terms of sensitivity, specificity, cost, workflow, and turnaround time. lso, there is a need for the technology to be able to accurately quantify the percent mutation within a sample. We developed sensitive and quantitative Mutation Detection ssays to help accurately assess mutation status and percentage in research samples. Mutation Detection ssays were designed based on novel competitive allele specific (castpcr ) technology, which combines allele-specific qpcr with allele-specific MGB blocker oligonucleotides to effectively suppress nonspecific amplification of the off-target allele (Figure 1).
2 SP SB G SP llele-specific primer SB llele-specific blocker (MGB) LST Locus-specific probe LSP Locus-specific primer LST LSP Figure 1. Schematic of a Mutation Detection ssay. mutant allele assay or wild type allele assay is composed of an allele-specific primer (SP), locus-specific primer (LSP), locus-specific probe (LST), and allele-specific blocker (SB). The % mutation present in the sample is calculated based on the ΔC t value between amplification reactions for a mutant allele assay and the corresponding wild type allele assay or a gene-specific reference assay. Materials and methods DN samples Twelve cell lines (from TCC or other sources; Table 1) were cultured according to the suppliers recommendations, and gdn was isolated using the PureLink Genomic DN Mini Kit (Life Technologies) or the QIamp gdn Mini Kit (Qiagen). gdn from mutant cell lines was diluted into gdn from wild type cell lines at various percentages. The total gdn sample input in each reaction was 10 ng or 30 ng. Eight model FFPE cell line specimens were prepared by crometrix, part of Life Technologies (Table 1). gdn was extracted from 20 µm slices with the Recoverll Nucleic cid Extraction Kit (mbion, part of Life Technologies) or QIamp DN FFPE Kit (Qiagen). gdn from mutant cell lines was diluted into gdn from FFPE wild type cell lines at various percentages. The total FFPE sample input in each reaction was 30 ng. FFPE or fresh frozen tumor specimens (colorectal and lung cancer) were collected and gdn was extracted by an independent laboratory using their own validated method. Synthetic plasmid constructs for seven KRS mutant alleles and corresponding wild type alleles were each designed with 600 bp of genomic context sequence surrounding the mutant base. Gene synthesis, plasmid construction, and plasmid preparation were performed by Genert, part of Life Technologies (Regensburg, Germany), and all sequences were verified by Sanger sequencing prior to use. ll DN samples were quantified using a NanoDrop ND1000 (NanoDrop Technologies) or RNase P Detection Reagents Kit. DN Template Reagents Kit were used to create a standard curve. Mutation Detection ssays Mutation detection experiments were performed by running 10 µl or 20 μl reactions comprising DN template, 1X Genotyping Master Mix, and 1X Mutation Detection ssay (containing allele-specific forward primer, locus-specific probe, locus-specific reverse primer, and allele-specific MGB blocker). qpcr was run on an pplied Biosystems Vii 7 or 7900HT Real-Time PCR System in 96-well or 384-well plates with the following conditions: 95 C for 10 min; 5 cycles: 92 C for 15 sec and 58 C for 1 min; 45 cycles: 92 C for 15 sec and 60 C for 1 min. ll samples were run in quadruplicates or triplicates. C t values were obtained from the instrument s real-time PCR data collection software using auto baseline and 0.2 manual threshold. The mutation status and percent mutation were analyzed using Mutation Detector software.
3 Table 1. Cell lines used in this study and their corresponding mutations. Cell line Sample type Mutation description mutation ID SW1463 Fresh/FFPE KRS G12C Fresh/FFPE KRS G12S 517 PSN-1 Fresh/FFPE KRS G12R 518 SW480 Fresh/FFPE KRS G12V 520 PNC-1 Fresh/FFPE KRS G12D 521 NCI-H2009 Fresh/FFPE KRS G DLD-1 Fresh/FFPE KRS G13D 532 H460 Fresh EGFR L858R 6224 H1975 Fresh EGFR T790M 6240 Table 2. KRS gene mutations in the benchmarking study. mutation ID Nucleotide change mino acid change 516 c.34g>t p.g12c 517 c.34g> p.g12s 518 c.34g>c p.g12r 520 c.35g>t p.g12v 521 c.35g> p.g12d 522 c.35g>c p.g c.38g> p.g13d H1650 Fresh EGFR Exon19 Del 6223 Colo 201 Fresh BRF V600E 476 Jurkat Fresh/FFPE Wild type Benchmarking of the Mutation Detection ssays to another commercially available qpcr technology for mutation detection was performed for seven KRS gene mutation targets (Table 2). The other commercially available assays, or assays, were purchased as a kit and the assay reactions were performed according to the protocol. The benchmarking experiments were performed on the pplied Biosystems 7500 Real-Time PCR System in 96-well reaction plates. The Mutation Detection ssays were used with the following thermal cycling conditions: 95 C for 10 min; 5 cycles: 92 C for 15 sec and 58 C for 1 min; 40 cycles: 92 C for 15 sec and 60 C for 1 min; with the analysis settings described above. The mutation detection assays were run in 25 μl reaction volume with the following thermal cycling conditions: 95 C for 4 min; 40 cycles: 95 C for 30 sec and 60 C for 1 min. nalysis of assays was performed according to the vendor s protocol. Results ssay sensitivity, selectivity, and linearity To evaluate assay sensitivity, selectivity, and linearity, gdn from each of the mutant cell lines (fresh or FFPE cell line) was serially diluted into a background of wild type cell line gdn of the same sample type. For fresh cell lines, the mutant allele percentage ranged from 100% to 0.1%. For FFPE cell lines, the mutant allele percentage ranged from 50% to 0.1%. The results show that assays can detect mutations at various percentages with excellent linearity (R 2 >0.990) and PCR efficiency (100 ± 10%) (Figure 2). In addition, assays can clearly detect 0.1% mutation (equivalent to 10 copies of mutant allele) in a background of 30 ng wild type gdn (Figure 2B). Some of the assays have been shown to detect down to a single copy of the mutant allele in a background of 100 ng wild type gdn (data not shown). ssay accuracy and reproducibility To assess the detection accuracy and reproducibility of the Mutation Detection ssays, three independent experiments were performed to measure the mutation percentage of model samples prepared in the above-mentioned cell line titration study. Both mutant allele assays and the corresponding wild type allele assays were run with each sample. The percent mutation present in the sample was calculated based on the ΔC t value between amplification reactions for the mutant allele assay and corresponding wild type allele assay. Run-to-run variation was analyzed for detection precision. The data show that these research assays can precisely quantify the percent mutation with CV<20% when the target allele copy number is >30 (Table 3).
4 KRS mutation detection in FFPE colorectal tumor research specimens Seven KRS Mutation Detection ssays were further tested with biological samples by an independent laboratory using a total of 31 gdn samples from FFPE colon tissue research specimens. mong those 31 samples, 10 were non-tumor colon specimens and 21 were colorectal tumor specimens. The KRS mutation status of these samples was validated by three different methods ( allelic discrimination (D) probe assay (custom SNP assay), D probe assay + peptide nucleic acid (PN) wild type allele blocker, and Sanger sequencing) (Table 4). Ten gdn samples extracted from normal FFPE tissues were examined for KRS mutation status using Mutation Detection ssays. No positive samples were found in the 10 nontumor tissues. The results obtained by KRS Mutation Detection ssays for those tumor samples were concordant with previously reported results (Table 4). KRS, BRF, and EGFR mutation detection in cell line and tumor biopsy research samples Mutation Detection ssays were evaluated using a total of 33 gdn samples from cell lines and tumor biopsy research samples (fresh frozen and FFPE tissues) by an independent laboratory. Mutation status of these samples was initially assessed by two different methods (amplification refractory mutation system (RMS ) technology and Sanger sequencing). RMS technology for mutation detection discriminates between the mutation and wild type DN by selectively amplifying the target sequence. The results from the Mutation Detection ssays demonstrated complete concordance. B. C t % G12R FFPE cell titration 0.5% y = x R 2 = % log % mutation 5% mplification plot 0.1% Mutation sample Wild type gdn sample 12.5% 25% 50% Figure 2. Mutation Detection ssays can detect mutations at various percentages with excellent linearity and PCR efficiency. The FFPE mutant cell line PSN-1 was serially titrated into the FFPE wild type Jurkat cell line from 50% to 0.1%. The total gdn sample input was 30 ng. The KRS_518_mu mutant allele assay was used to detect mutations in these model FFPE samples. () The KRS_518_mu assay demonstrated excellent linearity (R 2 = 0.999) and PCR efficiency (99.5%). (B) The amplification plot shows the C t difference between 0.1% mutation sample (10 copies mutant allele in 30 ng wild type gdn and wild type gdn sample (30 ng) for the KRS_518_mu assay.
5 Table 3. KRS G12C percent mutation measurement in cell line titration research samples from run to run. In this experiment, the total genomic DN input was 10 ng. G12C mutant cell line gdn titrated into wild type cell line gdn Target mutant allele copy number Expected (%) Measured (%), average CV (%) from run-to-run Table 4. KRS mutation detection in FFPE colorectal tumor research specimens using different mutation detection technologies. G12 G12C G12D G12R G12S G12V G13D WT Mutation Detection ssays D assay D assay + PN blocker n = 21 Sequencing Table 5. Mutation detection in cell lines and tumor biopsy research samples. The same samples were tested independently at different sites. EGFR_L858R EGFR_2235_2249del15 KRS_G12C KRS_G12R KRS_G12V KRS_G12D Mutation Detection ssay (data from an independent laboratory) Mutation Detection ssay (internal testing data) RMS Sanger sequencing to those previously reported results. Eighteen of the 33 gdn samples were then independently tested at Life Technologies to determine the robustness and reproducibility of the assays in biological samples. Results from this independent testing showed that Mutation Detection ssays are highly accurate and reproducible (Table 5). Benchmarking to other commercially available mutation detection kits Specificity and sensitivity of the Mutation Detection ssays compared to s product for mutation detection were evaluated with synthetic plasmid DN targets containing either the mutant allele or wild type allele. The seven KRS mutant allele assays from each company were run with 10,000 copies (equivalent to 30 ng gdn) of both target alleles. Three replicates were performed for each target, and the average C t values were determined (Figures 3 and 3B). Mutation Detection ssays generated significantly lower C t values compared to s assays, indicating that the Mutation Detection ssays have greater sensitivity. In addition, the mutant allele assays did not amplify the wild type allele target, with the exception of one Mutation Detection ssay that resulted in minimal amplification for mutation target KRS 521. The mutant allele assay on wild type allele target C t value minus the mutant allele assay on mutant allele target C t value provides a ΔC t value for each target assessed. This ΔC t correlates to the sensitivity capabilities of the assay. larger ΔC t value indicates the assay can detect a lower percent mutation in a larger quantity of sample. The Mutation Detection ssays provided significantly larger ΔC t values than s assays did, demonstrating the greater sensitivity of the Mutation Detection ssays.
6 The limit of detection for each of the seven assays for both technologies was evaluated with a dilution range of mutant allele targets, each combined with a constant 30 ng of wild type gdn. Two replicates were performed for each mutant allele target input quantity from 10,000 copies down to 1 2 copies per reaction. The number of reactions that amplified and reached the cycle threshold value before cycle 40 was counted (Table 6). The Mutation Detection ssays detected down to 1 copy of mutant allele sequence in a background of 10,000 copies of wild type gdn. s mutation detection assays reproducibly detected 50 copies for all seven mutations; however, detection was inconsistent below this input quantity. These results demonstrate the superior limit of detection of Mutation Detection ssays compared to this current commercial qpcr technology. Conclusions This study demonstrates that Mutation Detection ssays can accurately detect mutation status and quantify percent mutation in research samples with a relative standard deviation (CV) <20% in fresh and FFPE cell line samples containing a high background of gdn. The research assays demonstrate a minimum sensitivity of 0.1% mutation, and can detect down to a single copy of mutated DN sequence in a background of wild type gdn. dditionally, benchmarking to an established qpcr mutation detection technology shows that Mutation Detection ssays have a wider detection range and superior limit of detection.. Mutant allele DN target verage C t mutation B. Wild type allele DN target verage C t mutation Figure 3. Comparison of Mutation Detection ssays with s mutant allele assays. C t values of Mutation Detection ssays (blue) versus the C t values of s mutant allele assays (grey) run on 10,000 copies of () mutant allele DN target or (B) wild type allele DN target. The data demonstrate that Mutation Detection ssays have higher sensitivity and equivalent specificity compared to s assays.
7 Table 6. Limit of detection evaluation across a dynamic range of mutant allele DN target inputs in a background of wild type allele DN for and s mutation detection assays (n = 2 per input quantity). Red indicates that less than two replicates were detected; gray indicates that no replicates were detected. Mutant allele target input mutation 516 mutation 517 mutation 518 mutation 520 mutation 521 mutation 522 mutation ,000 copies ,000 copies copies copies copies copies copies ~1 to 2 copies Mutation Detection ssays have the following features: 1. ssays can detect less than 0.1% mutation in a background of wild type gdn. The limit of detection is down to 1 copy of mutant allele. 2. ssays have excellent linearity and PCR efficiency. 3. ssays are highly reproducible and can precisely quantify percent mutation. 4. ssays are highly accurate and highly concordant with other technologies. 5. ssays are compatible with multiple sample types including FFPE tissues, fresh frozen tissues, and cell lines. 6. ssays have superior detection range and sensitivity compared to current commercially available qpcr assays.
8 For Research Use Only. Not intended for any animal or human therapeutic or diagnostic use Life Technologies Corporation. ll rights reserved. The trademarks mentioned herein are the property of Life Technologies Corporation or their respective owners. is a registered trademark of Roche Molecular Systems Inc., used under permission and license. Printed in the US. CO Headquarters 5791 Van llen Way Carlsbad, C US Phone Toll Free in the US
2.500 Threshold. 2.000 1000e - 001. Threshold. Exponential phase. Cycle Number
 application note Real-Time PCR: Understanding C T Real-Time PCR: Understanding C T 4.500 3.500 1000e + 001 4.000 3.000 1000e + 000 3.500 2.500 Threshold 3.000 2.000 1000e - 001 Rn 2500 Rn 1500 Rn 2000
application note Real-Time PCR: Understanding C T Real-Time PCR: Understanding C T 4.500 3.500 1000e + 001 4.000 3.000 1000e + 000 3.500 2.500 Threshold 3.000 2.000 1000e - 001 Rn 2500 Rn 1500 Rn 2000
Real-time PCR: Understanding C t
 APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
TaqMan Mutation Detection Assay
 PROTOCOL TaqMan Mutation Detection Assay Competitive Allele-Specific TaqMan PCR Publication Part Number 4467011 Rev. B Revision Date April 2012 For Research Use Only. Not intended for any animal or human
PROTOCOL TaqMan Mutation Detection Assay Competitive Allele-Specific TaqMan PCR Publication Part Number 4467011 Rev. B Revision Date April 2012 For Research Use Only. Not intended for any animal or human
High-quality genomic DNA isolation and sensitive mutation analysis
 Application Note High-quality genomic DNA isolation and sensitive mutation analysis Izabela Safin, Ivonne Schröder-Stumberger and Peter Porschewski Introduction A major objective of cancer research is
Application Note High-quality genomic DNA isolation and sensitive mutation analysis Izabela Safin, Ivonne Schröder-Stumberger and Peter Porschewski Introduction A major objective of cancer research is
Introduction To Real Time Quantitative PCR (qpcr)
 Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Gene Expression Assays
 APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
Highly specific and sensitive quantitation
 PRODUCT ULLETIN SYR Select Master Mix SYR Select Master Mix Highly specific and sensitive quantitation SYR Select Master Mix offers advanced performance at an affordable price. SYR Select Master Mix is
PRODUCT ULLETIN SYR Select Master Mix SYR Select Master Mix Highly specific and sensitive quantitation SYR Select Master Mix offers advanced performance at an affordable price. SYR Select Master Mix is
MystiCq microrna cdna Synthesis Mix Catalog Number MIRRT Storage Temperature 20 C
 microrna cdna Synthesis Mix Catalog Number MIRRT Storage Temperature 20 C Product Description The microrna cdna Synthesis Mix has been designed to easily convert micrornas into cdna templates for qpcr
microrna cdna Synthesis Mix Catalog Number MIRRT Storage Temperature 20 C Product Description The microrna cdna Synthesis Mix has been designed to easily convert micrornas into cdna templates for qpcr
DNA Integrity Number (DIN) For the Assessment of Genomic DNA Samples in Real-Time Quantitative PCR (qpcr) Experiments
 DNA Integrity Number () For the Assessment of Genomic DNA Samples in Real-Time Quantitative PCR (qpcr) Experiments Application Note Nucleic Acid Analysis Author Arunkumar Padmanaban Agilent Technologies,
DNA Integrity Number () For the Assessment of Genomic DNA Samples in Real-Time Quantitative PCR (qpcr) Experiments Application Note Nucleic Acid Analysis Author Arunkumar Padmanaban Agilent Technologies,
SYBR Green Realtime PCR Master Mix -Plus-
 Instruction manual SYBR Green Realtime PCR Master Mix -Plus- 0810 F0925K SYBR Green Realtime PCR Master Mix -Plus- Contents QPK-212T 1mLx1 QPK-212 1mLx5 Store at -20 C, protected from light [1] Introduction
Instruction manual SYBR Green Realtime PCR Master Mix -Plus- 0810 F0925K SYBR Green Realtime PCR Master Mix -Plus- Contents QPK-212T 1mLx1 QPK-212 1mLx5 Store at -20 C, protected from light [1] Introduction
Factors Influencing Multiplex Real-Time PCR
 APPLICATION NOTE Multiplex Real-Time PCR Factors Influencing Multiplex Real-Time PCR Introduction Multiplex PCR is the simultaneous amplification of more than one target sequence in a single reaction [1].
APPLICATION NOTE Multiplex Real-Time PCR Factors Influencing Multiplex Real-Time PCR Introduction Multiplex PCR is the simultaneous amplification of more than one target sequence in a single reaction [1].
Methylation Analysis Using Methylation-Sensitive HRM and DNA Sequencing
 APPLICATION NOTE Methylation Analysis Using Methylation-Sensitive HRM and DNA Sequencing Methylation Analysis Using Methylation Sensitive HRM and DNA Sequencing Abstract DNA methylation is a key epigenetic
APPLICATION NOTE Methylation Analysis Using Methylation-Sensitive HRM and DNA Sequencing Methylation Analysis Using Methylation Sensitive HRM and DNA Sequencing Abstract DNA methylation is a key epigenetic
BacReady TM Multiplex PCR System
 BacReady TM Multiplex PCR System Technical Manual No. 0191 Version 10112010 I Description.. 1 II Applications 2 III Key Features.. 2 IV Shipping and Storage. 2 V Simplified Procedures. 2 VI Detailed Experimental
BacReady TM Multiplex PCR System Technical Manual No. 0191 Version 10112010 I Description.. 1 II Applications 2 III Key Features.. 2 IV Shipping and Storage. 2 V Simplified Procedures. 2 VI Detailed Experimental
Essentials of Real Time PCR. About Sequence Detection Chemistries
 Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Creating Standard Curves with Genomic DNA or Plasmid DNA Templates for Use in Quantitative PCR
 Creating Standard Curves with Genomic DNA or Plasmid DNA Templates for Use in Quantitative PCR Overview Genomic DNA (gdna) and plasmids containing cloned target sequences are commonly used as standards
Creating Standard Curves with Genomic DNA or Plasmid DNA Templates for Use in Quantitative PCR Overview Genomic DNA (gdna) and plasmids containing cloned target sequences are commonly used as standards
PNA BRAF Mutation Detection Kit
 - PNA BRAF Mutation Detection Kit Catalog Number KA2102 50 tests/kit Version: 01 Intended for research use only www.abnova.com Introduction and Background Intended use The PNA BRAF Mutation Detection Kit
- PNA BRAF Mutation Detection Kit Catalog Number KA2102 50 tests/kit Version: 01 Intended for research use only www.abnova.com Introduction and Background Intended use The PNA BRAF Mutation Detection Kit
GenScript BloodReady TM Multiplex PCR System
 GenScript BloodReady TM Multiplex PCR System Technical Manual No. 0174 Version 20040915 I Description.. 1 II Applications 2 III Key Features.. 2 IV Shipping and Storage. 2 V Simplified Procedures. 2 VI
GenScript BloodReady TM Multiplex PCR System Technical Manual No. 0174 Version 20040915 I Description.. 1 II Applications 2 III Key Features.. 2 IV Shipping and Storage. 2 V Simplified Procedures. 2 VI
Quantifiler Human DNA Quantification Kit Quantifiler Y Human Male DNA Quantification Kit
 Product Bulletin Human Identification Quantifiler Human DNA Quantification Kit Quantifiler Y Human Male DNA Quantification Kit The Quantifiler kits produce reliable and reproducible results, helping to
Product Bulletin Human Identification Quantifiler Human DNA Quantification Kit Quantifiler Y Human Male DNA Quantification Kit The Quantifiler kits produce reliable and reproducible results, helping to
ncounter Leukemia Fusion Gene Expression Assay Molecules That Count Product Highlights ncounter Leukemia Fusion Gene Expression Assay Overview
 ncounter Leukemia Fusion Gene Expression Assay Product Highlights Simultaneous detection and quantification of 25 fusion gene isoforms and 23 additional mrnas related to leukemia Compatible with a variety
ncounter Leukemia Fusion Gene Expression Assay Product Highlights Simultaneous detection and quantification of 25 fusion gene isoforms and 23 additional mrnas related to leukemia Compatible with a variety
QPCR Applications using Stratagene s Mx Real-Time PCR Platform
 QPCR Applications using Stratagene s Mx Real-Time PCR Platform Dan Schoeffner, Ph.D Field Applications Scientist [email protected] Tech. Services 800-894-1304 Polymerase Chain Reaction Melt
QPCR Applications using Stratagene s Mx Real-Time PCR Platform Dan Schoeffner, Ph.D Field Applications Scientist [email protected] Tech. Services 800-894-1304 Polymerase Chain Reaction Melt
Data Analysis on the ABI PRISM 7700 Sequence Detection System: Setting Baselines and Thresholds. Overview. Data Analysis Tutorial
 Data Analysis on the ABI PRISM 7700 Sequence Detection System: Setting Baselines and Thresholds Overview In order for accuracy and precision to be optimal, the assay must be properly evaluated and a few
Data Analysis on the ABI PRISM 7700 Sequence Detection System: Setting Baselines and Thresholds Overview In order for accuracy and precision to be optimal, the assay must be properly evaluated and a few
Application Guide... 2
 Protocol for GenomePlex Whole Genome Amplification from Formalin-Fixed Parrafin-Embedded (FFPE) tissue Application Guide... 2 I. Description... 2 II. Product Components... 2 III. Materials to be Supplied
Protocol for GenomePlex Whole Genome Amplification from Formalin-Fixed Parrafin-Embedded (FFPE) tissue Application Guide... 2 I. Description... 2 II. Product Components... 2 III. Materials to be Supplied
Path-ID Multiplex One-Step RT-PCR Kit
 USER GUIDE Path-ID Multiplex One-Step RT-PCR Kit TaqMan probe-based multiplex one-step real-time RT-PCR detection of RNA targets Catalog Numbers 4428206, 4428207, 4440022 Publication Part Number 4440907
USER GUIDE Path-ID Multiplex One-Step RT-PCR Kit TaqMan probe-based multiplex one-step real-time RT-PCR detection of RNA targets Catalog Numbers 4428206, 4428207, 4440022 Publication Part Number 4440907
360 Master Mix. , and a supplementary 360 GC Enhancer.
 Product Bulletin AmpliTaq Gold 360 Master Mix and 360 DNA Polymerase AmpliTaq Gold 360 Master Mix AmpliTaq Gold 360 DNA Polymerase 360 Coverage for a Full Range of Targets AmpliTaq Gold 360 Master Mix
Product Bulletin AmpliTaq Gold 360 Master Mix and 360 DNA Polymerase AmpliTaq Gold 360 Master Mix AmpliTaq Gold 360 DNA Polymerase 360 Coverage for a Full Range of Targets AmpliTaq Gold 360 Master Mix
Sequencing Library qpcr Quantification Guide
 Sequencing Library qpcr Quantification Guide FOR RESEARCH USE ONLY Introduction 3 Quantification Workflow 4 Best Practices 5 Consumables and Equipment 6 Select Control Template 8 Dilute qpcr Control Template
Sequencing Library qpcr Quantification Guide FOR RESEARCH USE ONLY Introduction 3 Quantification Workflow 4 Best Practices 5 Consumables and Equipment 6 Select Control Template 8 Dilute qpcr Control Template
Consistent Assay Performance Across Universal Arrays and Scanners
 Technical Note: Illumina Systems and Software Consistent Assay Performance Across Universal Arrays and Scanners There are multiple Universal Array and scanner options for running Illumina DASL and GoldenGate
Technical Note: Illumina Systems and Software Consistent Assay Performance Across Universal Arrays and Scanners There are multiple Universal Array and scanner options for running Illumina DASL and GoldenGate
Advances in RainDance Sequence Enrichment Technology and Applications in Cancer Research. March 17, 2011 Rendez-Vous Séquençage
 Advances in RainDance Sequence Enrichment Technology and Applications in Cancer Research March 17, 2011 Rendez-Vous Séquençage Presentation Overview Core Technology Review Sequence Enrichment Application
Advances in RainDance Sequence Enrichment Technology and Applications in Cancer Research March 17, 2011 Rendez-Vous Séquençage Presentation Overview Core Technology Review Sequence Enrichment Application
Oncology Insights Enabled by Knowledge Base-Guided Panel Design and the Seamless Workflow of the GeneReader NGS System
 White Paper Oncology Insights Enabled by Knowledge Base-Guided Panel Design and the Seamless Workflow of the GeneReader NGS System Abstract: This paper describes QIAGEN s philosophy and process for developing
White Paper Oncology Insights Enabled by Knowledge Base-Guided Panel Design and the Seamless Workflow of the GeneReader NGS System Abstract: This paper describes QIAGEN s philosophy and process for developing
TaqMan Fast Advanced Master Mix. Protocol
 TaqMan Fast Advanced Master Mix Protocol For Research Use Only. Not intended for any animal or human therapeutic or diagnostic use. Information in this document is subject to change without notice. APPLIED
TaqMan Fast Advanced Master Mix Protocol For Research Use Only. Not intended for any animal or human therapeutic or diagnostic use. Information in this document is subject to change without notice. APPLIED
Real time and Quantitative (RTAQ) PCR. so I have an outlier and I want to see if it really is changed
 Real time and Quantitative (RTAQ) PCR or.. for this audience so I have an outlier and I want to see if it really is changed Nigel Walker, Ph.D. Laboratory of Computational Biology and Risk Analysis, Environmental
Real time and Quantitative (RTAQ) PCR or.. for this audience so I have an outlier and I want to see if it really is changed Nigel Walker, Ph.D. Laboratory of Computational Biology and Risk Analysis, Environmental
Single-Cell Whole Genome Sequencing on the C1 System: a Performance Evaluation
 PN 100-9879 A1 TECHNICAL NOTE Single-Cell Whole Genome Sequencing on the C1 System: a Performance Evaluation Introduction Cancer is a dynamic evolutionary process of which intratumor genetic and phenotypic
PN 100-9879 A1 TECHNICAL NOTE Single-Cell Whole Genome Sequencing on the C1 System: a Performance Evaluation Introduction Cancer is a dynamic evolutionary process of which intratumor genetic and phenotypic
TITRATION OF raav (VG) USING QUANTITATIVE REAL TIME PCR
 Page 1 of 5 Materials DNase digestion buffer [13 mm Tris-Cl, ph7,5 / 5 mm MgCl2 / 0,12 mm CaCl2] RSS plasmid ptr-uf11 SV40pA Forward primer (10µM) AGC AAT AGC ATC ACA AAT TTC ACA A SV40pA Reverse Primer
Page 1 of 5 Materials DNase digestion buffer [13 mm Tris-Cl, ph7,5 / 5 mm MgCl2 / 0,12 mm CaCl2] RSS plasmid ptr-uf11 SV40pA Forward primer (10µM) AGC AAT AGC ATC ACA AAT TTC ACA A SV40pA Reverse Primer
quantitative real-time PCR, grain, simplex DNA extraction: PGS0426 RT-PCR: PGS0494 & PGS0476
 BioScience quantitative real-time PCR, grain, simplex DNA extraction: PGS0426 RT-PCR: PGS0494 & PGS0476 This method describes a Real-time semi-quantitative TaqMan PCR procedure for the determination of
BioScience quantitative real-time PCR, grain, simplex DNA extraction: PGS0426 RT-PCR: PGS0494 & PGS0476 This method describes a Real-time semi-quantitative TaqMan PCR procedure for the determination of
RealStar HBV PCR Kit 1.0 11/2012
 RealStar HBV PCR Kit 1.0 11/2012 RealStar HBV PCR Kit 1.0 For research use only! (RUO) Product No.: 201003 96 rxns INS-201000-GB-02 Store at -25 C... -15 C November 2012 altona Diagnostics GmbH Mörkenstraße
RealStar HBV PCR Kit 1.0 11/2012 RealStar HBV PCR Kit 1.0 For research use only! (RUO) Product No.: 201003 96 rxns INS-201000-GB-02 Store at -25 C... -15 C November 2012 altona Diagnostics GmbH Mörkenstraße
mircute mirna qpcr Detection Kit (SYBR Green)
 mircute mirna qpcr Detection Kit (SYBR Green) For detection of mirna using real-time RT-PCR (SYBR Green I) www.tiangen.com QP110302 mircute mirna qpcr Detection Kit (SYBR Green) Kit Contents Cat. no. FP401
mircute mirna qpcr Detection Kit (SYBR Green) For detection of mirna using real-time RT-PCR (SYBR Green I) www.tiangen.com QP110302 mircute mirna qpcr Detection Kit (SYBR Green) Kit Contents Cat. no. FP401
qstar mirna qpcr Detection System
 qstar mirna qpcr Detection System Table of Contents Table of Contents...1 Package Contents and Storage Conditions...2 For mirna cdna synthesis kit...2 For qstar mirna primer pairs...2 For qstar mirna qpcr
qstar mirna qpcr Detection System Table of Contents Table of Contents...1 Package Contents and Storage Conditions...2 For mirna cdna synthesis kit...2 For qstar mirna primer pairs...2 For qstar mirna qpcr
Instruction manual for product # PNAC-2001. Version 1.2
 PNAClamp BRAF Mutation Detection Kit For in vitro diagnostic use Instruction manual for product # PNAC-2001 Version 1.2 Store at -20 C. Instruction Version: Ver. 1.2 Date of Revision: 2012 February 1 /
PNAClamp BRAF Mutation Detection Kit For in vitro diagnostic use Instruction manual for product # PNAC-2001 Version 1.2 Store at -20 C. Instruction Version: Ver. 1.2 Date of Revision: 2012 February 1 /
Mir-X mirna First-Strand Synthesis Kit User Manual
 User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
Real-Time PCR Vs. Traditional PCR
 Real-Time PCR Vs. Traditional PCR Description This tutorial will discuss the evolution of traditional PCR methods towards the use of Real-Time chemistry and instrumentation for accurate quantitation. Objectives
Real-Time PCR Vs. Traditional PCR Description This tutorial will discuss the evolution of traditional PCR methods towards the use of Real-Time chemistry and instrumentation for accurate quantitation. Objectives
Data Analysis for Ion Torrent Sequencing
 IFU022 v140202 Research Use Only Instructions For Use Part III Data Analysis for Ion Torrent Sequencing MANUFACTURER: Multiplicom N.V. Galileilaan 18 2845 Niel Belgium Revision date: August 21, 2014 Page
IFU022 v140202 Research Use Only Instructions For Use Part III Data Analysis for Ion Torrent Sequencing MANUFACTURER: Multiplicom N.V. Galileilaan 18 2845 Niel Belgium Revision date: August 21, 2014 Page
PrimeSTAR HS DNA Polymerase
 Cat. # R010A For Research Use PrimeSTAR HS DNA Polymerase Product Manual Table of Contents I. Description...3 II. III. IV. Components...3 Storage...3 Features...3 V. General Composition of PCR Reaction
Cat. # R010A For Research Use PrimeSTAR HS DNA Polymerase Product Manual Table of Contents I. Description...3 II. III. IV. Components...3 Storage...3 Features...3 V. General Composition of PCR Reaction
Absolute Quantitation Using Standard Curve Getting Started Guide
 Applied Biosystems 7900HT Fast Real-Time PCR System Absolute Quantitation Using Standard Curve Getting Started Guide Introduction Designing an AQ Experiment Preparing the Samples and Reaction Plate Creating
Applied Biosystems 7900HT Fast Real-Time PCR System Absolute Quantitation Using Standard Curve Getting Started Guide Introduction Designing an AQ Experiment Preparing the Samples and Reaction Plate Creating
Real-time quantitative RT -PCR (Taqman)
 Real-time quantitative RT -PCR (Taqman) Author: SC, Patti Lab, 3/03 This is performed as a 2-step reaction: 1. cdna synthesis from DNase 1-treated total RNA 2. PCR 1. cdna synthesis (Advantage RT-for-PCR
Real-time quantitative RT -PCR (Taqman) Author: SC, Patti Lab, 3/03 This is performed as a 2-step reaction: 1. cdna synthesis from DNase 1-treated total RNA 2. PCR 1. cdna synthesis (Advantage RT-for-PCR
LightCycler 480 Real-Time PCR System
 Roche Applied Science LightCycler 480 Real-Time PCR System Planned introduction of the LightCycler 480 System: September 2005 Rapid by nature, accurate by design The LightCycler 480 Real-Time PCR System
Roche Applied Science LightCycler 480 Real-Time PCR System Planned introduction of the LightCycler 480 System: September 2005 Rapid by nature, accurate by design The LightCycler 480 Real-Time PCR System
User Bulletin #2 ABI PRISM 7700 Sequence Detection System
 User Bulletin #2 ABI PRISM 7700 Sequence Detection System December 11, 1997 (updated 10/2001) SUBJECT: Relative Quantitation of Gene Expression Introduction Amplification of an endogenous control may be
User Bulletin #2 ABI PRISM 7700 Sequence Detection System December 11, 1997 (updated 10/2001) SUBJECT: Relative Quantitation of Gene Expression Introduction Amplification of an endogenous control may be
PreciseTM Whitepaper
 Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
ONLINE SUPPLEMENTAL MATERIAL. Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue
 ONLINE SUPPLEMENTAL MATERIAL Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue Sungmi Park 1, Ko-Ting Lu 1, Xuebo Liu 1, Tapan K. Chatterjee 2, Steven M. Rudich 3, Neal
ONLINE SUPPLEMENTAL MATERIAL Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue Sungmi Park 1, Ko-Ting Lu 1, Xuebo Liu 1, Tapan K. Chatterjee 2, Steven M. Rudich 3, Neal
Human Herpes Virus 1 (Herpes simplex type 1) genesig Standard Kit. DNA testing. Everything... Everyone... Everywhere...
 TM Primerdesign Ltd TM Primerdesign Ltd Human Herpes Virus 1 (Herpes simplex type 1) Capsid assembly and DNA maturation gene genesig Standard Kit 150 tests DNA testing Everything... Everyone... Everywhere...
TM Primerdesign Ltd TM Primerdesign Ltd Human Herpes Virus 1 (Herpes simplex type 1) Capsid assembly and DNA maturation gene genesig Standard Kit 150 tests DNA testing Everything... Everyone... Everywhere...
Epstein Barr Virus (Human Herpes virus 4) genesig Standard Kit. DNA testing. Everything... Everyone... Everywhere...
 TM Primerdesign Ltd TM Primerdesign Ltd Epstein Barr Virus (Human Herpes virus 4) nonglycosylated membrane protein (BNRF1) gene genesig Standard Kit 150 tests DNA testing Everything... Everyone... Everywhere...
TM Primerdesign Ltd TM Primerdesign Ltd Epstein Barr Virus (Human Herpes virus 4) nonglycosylated membrane protein (BNRF1) gene genesig Standard Kit 150 tests DNA testing Everything... Everyone... Everywhere...
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual
 Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Nuevas tecnologías basadas en biomarcadores para oncología
 Nuevas tecnologías basadas en biomarcadores para oncología Simposio ASEBIO 14 de marzo 2013, PCB Jose Jimeno, MD, PhD Co-Founder / Vice Chairman Pangaea Biotech SL Barcelona, Spain PANGAEA BIOTECH BUSINESS
Nuevas tecnologías basadas en biomarcadores para oncología Simposio ASEBIO 14 de marzo 2013, PCB Jose Jimeno, MD, PhD Co-Founder / Vice Chairman Pangaea Biotech SL Barcelona, Spain PANGAEA BIOTECH BUSINESS
Absolute Quantification Getting Started Guide
 5 cdna Reverse Primer Oligo d(t) or random hexamer Synthesis of 1st cdna strand 3 5 cdna Applied Biosystems 7300/7500/7500 Fast Real-Time PCR System Absolute Quantification Getting Started Guide Introduction
5 cdna Reverse Primer Oligo d(t) or random hexamer Synthesis of 1st cdna strand 3 5 cdna Applied Biosystems 7300/7500/7500 Fast Real-Time PCR System Absolute Quantification Getting Started Guide Introduction
Quantitative Real Time PCR Protocol. Stack Lab
 Quantitative Real Time PCR Protocol Stack Lab Overview Real-time quantitative polymerase chain reaction (qpcr) differs from regular PCR by including in the reaction fluorescent reporter molecules that
Quantitative Real Time PCR Protocol Stack Lab Overview Real-time quantitative polymerase chain reaction (qpcr) differs from regular PCR by including in the reaction fluorescent reporter molecules that
All-in-One mirna qrt-pcr Reagent Kits For quantitative detection of mature mirna
 All-in-One mirna qrt-pcr Reagent Kits For quantitative detection of mature mirna All-in-One TM mirna First-Strand cdna Synthesis Kit AMRT-0020 (20 RT reactions), AMRT-0060 (60 RT reactions) Used in combination
All-in-One mirna qrt-pcr Reagent Kits For quantitative detection of mature mirna All-in-One TM mirna First-Strand cdna Synthesis Kit AMRT-0020 (20 RT reactions), AMRT-0060 (60 RT reactions) Used in combination
Epstein Barr Virus (Human Herpes virus 4) nonglycosylated membrane protein (BNRF1) gene. genesig Advanced Kit. DNA testing
 TM Primerdesign Ltd TM Primerdesign Ltd Epstein Barr Virus (Human Herpes virus 4) nonglycosylated membrane protein (BNRF1) gene genesig Advanced Kit 150 tests DNA testing Everything... Everyone... Everywhere...
TM Primerdesign Ltd TM Primerdesign Ltd Epstein Barr Virus (Human Herpes virus 4) nonglycosylated membrane protein (BNRF1) gene genesig Advanced Kit 150 tests DNA testing Everything... Everyone... Everywhere...
PCR was carried out in a reaction volume of 20 µl using the ABI AmpliTaq GOLD kit (ABI,
 Supplemental Text/Tables PCR Amplification and Sequencing PCR was carried out in a reaction volume of 20 µl using the ABI AmpliTaq GOLD kit (ABI, Foster City, CA). Each PCR reaction contained 20 ng genomic
Supplemental Text/Tables PCR Amplification and Sequencing PCR was carried out in a reaction volume of 20 µl using the ABI AmpliTaq GOLD kit (ABI, Foster City, CA). Each PCR reaction contained 20 ng genomic
qpcr Quantification Protocol Guide
 qpcr Quantification Protocol Guide FOR RESEARCH USE ONLY Topics 3 Introduction 5 User-Supplied Consumables and Equipment 7 Select Template 8 Dilute qpcr Template 9 Dilute Libraries 10 Prepare Reaction
qpcr Quantification Protocol Guide FOR RESEARCH USE ONLY Topics 3 Introduction 5 User-Supplied Consumables and Equipment 7 Select Template 8 Dilute qpcr Template 9 Dilute Libraries 10 Prepare Reaction
CompleteⅡ 1st strand cdna Synthesis Kit
 Instruction Manual CompleteⅡ 1st strand cdna Synthesis Kit Catalog # GM30401, GM30402 Green Mountain Biosystems. LLC Web: www.greenmountainbio.com Tel: 800-942-1160 Sales: Sales@ greenmountainbio.com Support:
Instruction Manual CompleteⅡ 1st strand cdna Synthesis Kit Catalog # GM30401, GM30402 Green Mountain Biosystems. LLC Web: www.greenmountainbio.com Tel: 800-942-1160 Sales: Sales@ greenmountainbio.com Support:
Single Nucleotide Polymorphisms (SNPs)
 Single Nucleotide Polymorphisms (SNPs) Additional Markers 13 core STR loci Obtain further information from additional markers: Y STRs Separating male samples Mitochondrial DNA Working with extremely degraded
Single Nucleotide Polymorphisms (SNPs) Additional Markers 13 core STR loci Obtain further information from additional markers: Y STRs Separating male samples Mitochondrial DNA Working with extremely degraded
REAL TIME PCR USING SYBR GREEN
 REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
ab185916 Hi-Fi cdna Synthesis Kit
 ab185916 Hi-Fi cdna Synthesis Kit Instructions for Use For cdna synthesis from various RNA samples This product is for research use only and is not intended for diagnostic use. Version 1 Last Updated 1
ab185916 Hi-Fi cdna Synthesis Kit Instructions for Use For cdna synthesis from various RNA samples This product is for research use only and is not intended for diagnostic use. Version 1 Last Updated 1
New Technologies for Sensitive, Low-Input RNA-Seq. Clontech Laboratories, Inc.
 New Technologies for Sensitive, Low-Input RNA-Seq Clontech Laboratories, Inc. Outline Introduction Single-Cell-Capable mrna-seq Using SMART Technology SMARTer Ultra Low RNA Kit for the Fluidigm C 1 System
New Technologies for Sensitive, Low-Input RNA-Seq Clontech Laboratories, Inc. Outline Introduction Single-Cell-Capable mrna-seq Using SMART Technology SMARTer Ultra Low RNA Kit for the Fluidigm C 1 System
Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr
 User Manual Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories,
User Manual Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories,
REAL TIME PCR SYBR GREEN
 REAL TIME PCR SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM QUANTITATION
REAL TIME PCR SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM QUANTITATION
Validating Microarray Data Using RT 2 Real-Time PCR Products
 Validating Microarray Data Using RT 2 Real-Time PCR Products Introduction: Real-time PCR monitors the amount of amplicon as the reaction occurs. Usually, the amount of product is directly related to the
Validating Microarray Data Using RT 2 Real-Time PCR Products Introduction: Real-time PCR monitors the amount of amplicon as the reaction occurs. Usually, the amount of product is directly related to the
Co Extra (GM and non GM supply chains: Their CO EXistence and TRAceability) Outcomes of Co Extra
 GM and non GM supply chains: Their CO EXistence and TRAceability Outcomes of Co Extra Comparison of different real time PCR chemistries and their suitability for detection and quantification of genetically
GM and non GM supply chains: Their CO EXistence and TRAceability Outcomes of Co Extra Comparison of different real time PCR chemistries and their suitability for detection and quantification of genetically
Human Herpes Virus 4 (Epstein Barr)
 Techne qpcr test Human Herpes Virus 4 (Epstein Barr) nonglycosylated membrane protein (BNRF1) gene 150 tests For general laboratory and research use only 1 Introduction to Human Herpes Virus 4 (Epstein
Techne qpcr test Human Herpes Virus 4 (Epstein Barr) nonglycosylated membrane protein (BNRF1) gene 150 tests For general laboratory and research use only 1 Introduction to Human Herpes Virus 4 (Epstein
EZValidation Online Tool. The easy way to apply guidelines. The EZValidation Online Tool. The implementation of new tests
 The implementation of new tests The EZValidation Online Tool Since 999, AcroMetrix has provided quality control requires validation or verification studies that meet is a comprehensive tool specifically
The implementation of new tests The EZValidation Online Tool Since 999, AcroMetrix has provided quality control requires validation or verification studies that meet is a comprehensive tool specifically
Technical Note. Roche Applied Science. No. LC 18/2004. Assay Formats for Use in Real-Time PCR
 Roche Applied Science Technical Note No. LC 18/2004 Purpose of this Note Assay Formats for Use in Real-Time PCR The LightCycler Instrument uses several detection channels to monitor the amplification of
Roche Applied Science Technical Note No. LC 18/2004 Purpose of this Note Assay Formats for Use in Real-Time PCR The LightCycler Instrument uses several detection channels to monitor the amplification of
Technical Manual No. 0173 Update Date 10112010
 TissueDirect TM Multiplex PCR System Technical Manual No. 0173 Update Date 10112010 I Description.. 1 II Applications 2 III Key Features.. 2 IV Shipping and Storage. 3 V Simplified Procedures. 3 VI Detailed
TissueDirect TM Multiplex PCR System Technical Manual No. 0173 Update Date 10112010 I Description.. 1 II Applications 2 III Key Features.. 2 IV Shipping and Storage. 3 V Simplified Procedures. 3 VI Detailed
amplification tech Optical Design of CFX96 Real-Time PCR Detection System Eliminates the Requirement of a Passive Reference Dye
 amplification tech note 6047 Optical Design of CFX96 Real-Time PCR Detection System Eliminates the Requirement of a Passive Reference Dye Liz Jordan and Richard Kurtz, Gene Expression Division, io-rad
amplification tech note 6047 Optical Design of CFX96 Real-Time PCR Detection System Eliminates the Requirement of a Passive Reference Dye Liz Jordan and Richard Kurtz, Gene Expression Division, io-rad
Stratagene QPCR Mouse Reference Total RNA
 Stratagene QPCR Mouse Reference Total RNA Instruction Manual Catalog #750600 Revision C.0 For Research Use Only. Not for use in diagnostic procedures. 750600-12 LIMITED PRODUCT WARRANTY This warranty limits
Stratagene QPCR Mouse Reference Total RNA Instruction Manual Catalog #750600 Revision C.0 For Research Use Only. Not for use in diagnostic procedures. 750600-12 LIMITED PRODUCT WARRANTY This warranty limits
Single-Cell DNA Sequencing with the C 1. Single-Cell Auto Prep System. Reveal hidden populations and genetic diversity within complex samples
 DATA Sheet Single-Cell DNA Sequencing with the C 1 Single-Cell Auto Prep System Reveal hidden populations and genetic diversity within complex samples Single-cell sensitivity Discover and detect SNPs,
DATA Sheet Single-Cell DNA Sequencing with the C 1 Single-Cell Auto Prep System Reveal hidden populations and genetic diversity within complex samples Single-cell sensitivity Discover and detect SNPs,
Real-time qpcr Assay Design Software www.qpcrdesign.com
 Real-time qpcr Assay Design Software www.qpcrdesign.com Your Blueprint For Success Informational Guide 2199 South McDowell Blvd Petaluma, CA 94954-6904 USA 1.800.GENOME.1(436.6631) 1.415.883.8400 1.415.883.8488
Real-time qpcr Assay Design Software www.qpcrdesign.com Your Blueprint For Success Informational Guide 2199 South McDowell Blvd Petaluma, CA 94954-6904 USA 1.800.GENOME.1(436.6631) 1.415.883.8400 1.415.883.8488
SYBR Green PCR Master Mix and SYBR Green RT-PCR Reagents Kit
 USER GUIDE SYBR Green PCR Master Mix and SYBR Green RT-PCR Reagents Kit Catalog Number 4309155 (Master Mix) and 4306736 (RT-PCR Reagents Kit) Publication Part Number 4310251 Rev. G Revision Date September
USER GUIDE SYBR Green PCR Master Mix and SYBR Green RT-PCR Reagents Kit Catalog Number 4309155 (Master Mix) and 4306736 (RT-PCR Reagents Kit) Publication Part Number 4310251 Rev. G Revision Date September
Intended Use: The kit is designed to detect the 5 different mutations found in Asian population using seven different primers.
 Unzipping Genes MBPCR014 Beta-Thalassemia Detection Kit P r o d u c t I n f o r m a t i o n Description: Thalassemia is a group of genetic disorders characterized by quantitative defects in globin chain
Unzipping Genes MBPCR014 Beta-Thalassemia Detection Kit P r o d u c t I n f o r m a t i o n Description: Thalassemia is a group of genetic disorders characterized by quantitative defects in globin chain
Relative Quantification of mirna Target mrnas by Real-Time qpcr. 1 Introduction. Gene Expression Application Note No. 4
 Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
Power SYBR Green PCR Master Mix and Power SYBR Green RT-PCR Reagents Kit
 USER GUIDE Power SYBR Green PCR Master Mix and Power SYBR Green RT-PCR Reagents Kit Catalog Number 4368577, 4367659, 4367660, 4368706, 4368702, 4368708 (Master Mix) and 4368711 (RT-PCR Reagents Kit) Publication
USER GUIDE Power SYBR Green PCR Master Mix and Power SYBR Green RT-PCR Reagents Kit Catalog Number 4368577, 4367659, 4367660, 4368706, 4368702, 4368708 (Master Mix) and 4368711 (RT-PCR Reagents Kit) Publication
All-in-One First-Strand cdna Synthesis Kit
 All-in-One First-Strand cdna Synthesis Kit For reliable first-strand cdna synthesis from all RNA sources Cat. No. AORT-0020 (20 synthesis reactions) Cat. No. AORT-0050 (50 synthesis reactions) User Manual
All-in-One First-Strand cdna Synthesis Kit For reliable first-strand cdna synthesis from all RNA sources Cat. No. AORT-0020 (20 synthesis reactions) Cat. No. AORT-0050 (50 synthesis reactions) User Manual
Table of Contents. I. Description... 2. II. Kit Components... 2. III. Storage... 2. IV. 1st Strand cdna Synthesis Reaction... 3
 Table of Contents I. Description... 2 II. Kit Components... 2 III. Storage... 2 IV. 1st Strand cdna Synthesis Reaction... 3 V. RT-PCR, Real-time RT-PCR... 4 VI. Application... 5 VII. Preparation of RNA
Table of Contents I. Description... 2 II. Kit Components... 2 III. Storage... 2 IV. 1st Strand cdna Synthesis Reaction... 3 V. RT-PCR, Real-time RT-PCR... 4 VI. Application... 5 VII. Preparation of RNA
White paper Evaluation of BRAF (V600E) Mutation by Immunohistochemical Staining with anti-braf V600E (VE1) Antibody: A Comparison with Sanger
 White paper Evaluation of BRAF (V600E) Mutation by Immunohistochemical Staining with anti-braf V600E (VE1) Antibody: A Comparison with Sanger Sequencing 2 Evaluation of BRAF (V600E) Mutation by Immunohistochemical
White paper Evaluation of BRAF (V600E) Mutation by Immunohistochemical Staining with anti-braf V600E (VE1) Antibody: A Comparison with Sanger Sequencing 2 Evaluation of BRAF (V600E) Mutation by Immunohistochemical
Validation parameters: An introduction to measures of
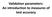 Validation parameters: An introduction to measures of test accuracy Types of tests All tests are fundamentally quantitative Sometimes we use the quantitative result directly However, it is often necessary
Validation parameters: An introduction to measures of test accuracy Types of tests All tests are fundamentally quantitative Sometimes we use the quantitative result directly However, it is often necessary
Dengue Virus subtypes 1,2 3 and 4. genesig Standard Kit. DNA testing. Everything... Everyone... Everywhere... 3 Untranslated Region (3 UTR) 150 tests
 TM Primerdesign Ltd TM Primerdesign Ltd Dengue Virus subtypes 1,2 3 and 4 3 Untranslated Region (3 UTR) genesig Standard Kit 150 tests DNA testing Everything... Everyone... Everywhere... For general laboratory
TM Primerdesign Ltd TM Primerdesign Ltd Dengue Virus subtypes 1,2 3 and 4 3 Untranslated Region (3 UTR) genesig Standard Kit 150 tests DNA testing Everything... Everyone... Everywhere... For general laboratory
Profiling of microrna in Blood Serum/Plasma. Guidelines for the mircury LNA TM Universal RT microrna PCR System
 Profiling of microrna in Blood Serum/Plasma Guidelines for the mircury LNA TM Universal RT microrna PCR System Table of Contents 2 Introduction.....................................................................................
Profiling of microrna in Blood Serum/Plasma Guidelines for the mircury LNA TM Universal RT microrna PCR System Table of Contents 2 Introduction.....................................................................................
Molecular Assessment of Dried Blood Spot Quality during Development of a Novel Automated. Screening
 Molecular Assessment of Dried Blood Spot Quality during Development of a Novel Automated in situ TREC qpcr Assay for SCID Screening J Bai, T Henry, J Benfer, S Berberich, T Kreman, and L DesJardin State
Molecular Assessment of Dried Blood Spot Quality during Development of a Novel Automated in situ TREC qpcr Assay for SCID Screening J Bai, T Henry, J Benfer, S Berberich, T Kreman, and L DesJardin State
The Power of Next-Generation Sequencing in Your Hands On the Path towards Diagnostics
 The Power of Next-Generation Sequencing in Your Hands On the Path towards Diagnostics The GS Junior System The Power of Next-Generation Sequencing on Your Benchtop Proven technology: Uses the same long
The Power of Next-Generation Sequencing in Your Hands On the Path towards Diagnostics The GS Junior System The Power of Next-Generation Sequencing on Your Benchtop Proven technology: Uses the same long
Speed Matters - Fast ways from template to result
 qpcr Symposium 2007 - Weihenstephan Speed Matters - Fast ways from template to result March 28, 2007 Dr. Thorsten Traeger Senior Scientist, Research and Development - 1 - Overview Ạgenda Fast PCR The Challenges
qpcr Symposium 2007 - Weihenstephan Speed Matters - Fast ways from template to result March 28, 2007 Dr. Thorsten Traeger Senior Scientist, Research and Development - 1 - Overview Ạgenda Fast PCR The Challenges
Fast PCR: General Considerations for Minimizing Run Times and Maximizing Throughput
 amplification tech note 5362 Fast PCR: General Considerations for Minimizing Run Times and Maximizing Throughput Daniel Sullivan, Babette Fahey, and David Titus, Bio-Rad Laboratories, Inc., Waltham, MA
amplification tech note 5362 Fast PCR: General Considerations for Minimizing Run Times and Maximizing Throughput Daniel Sullivan, Babette Fahey, and David Titus, Bio-Rad Laboratories, Inc., Waltham, MA
All-in-One mirna qrt-pcr Detection System Handbook
 All-in-One mirna qrt-pcr Detection System Handbook For quantitative detection of mature mirna All-in-One mirna First-Strand cdna Synthesis Kit Cat. No. AMRT-0020 (20 mirna reverse transcription reactions)
All-in-One mirna qrt-pcr Detection System Handbook For quantitative detection of mature mirna All-in-One mirna First-Strand cdna Synthesis Kit Cat. No. AMRT-0020 (20 mirna reverse transcription reactions)
Getting Started Guide
 Primer Express Software Version 3.0 Getting Started Guide Before You Begin Designing Primers and Probes for Quantification Assays Designing Primers and Probes for Allelic Discrimination Assays Ordering
Primer Express Software Version 3.0 Getting Started Guide Before You Begin Designing Primers and Probes for Quantification Assays Designing Primers and Probes for Allelic Discrimination Assays Ordering
Definition of Minimum Performance Requirements for Analytical Methods of GMO Testing European Network of GMO Laboratories (ENGL)
 Definition of Minimum Performance Requirements for Analytical Methods of GMO Testing European Network of GMO Laboratories (ENGL) 13 October 2008 Date of application: 13 April 2009 INTRODUCTION The scope
Definition of Minimum Performance Requirements for Analytical Methods of GMO Testing European Network of GMO Laboratories (ENGL) 13 October 2008 Date of application: 13 April 2009 INTRODUCTION The scope
Quantification of Propionibacterium acnes. Standard kit 150 tests
 Primerdesign TM Ltd Quantification of Propionibacterium acnes Recombinase A (reca) gene For general laboratory and research use only Standard kit 150 tests Introduction to Propionibacterium acnes Propionibacterium
Primerdesign TM Ltd Quantification of Propionibacterium acnes Recombinase A (reca) gene For general laboratory and research use only Standard kit 150 tests Introduction to Propionibacterium acnes Propionibacterium
Plexor Systems Instrument Setup and Data Analysis for the Applied Biosystems 7300 and 7500 Real-Time PCR Systems
 Technical Manual Plexor Systems Instrument Setup and Data Analysis for the Applied Biosystems 7300 and 7500 Real-Time PCR Systems INSTRUCTIONS FOR USE OF PRODUCTS A4011, A4021, A4031, A4041, A4051 AND
Technical Manual Plexor Systems Instrument Setup and Data Analysis for the Applied Biosystems 7300 and 7500 Real-Time PCR Systems INSTRUCTIONS FOR USE OF PRODUCTS A4011, A4021, A4031, A4041, A4051 AND
Quando si parla di PCR quantitativa si intende:
 Quando si parla di PCR quantitativa si intende: A. Una PCR che produce grandi quantità di DNA B. Una PCR che emette quanti di luce C. Una PCR che quantifica il numero di molecole stampo presenti all inizio
Quando si parla di PCR quantitativa si intende: A. Una PCR che produce grandi quantità di DNA B. Una PCR che emette quanti di luce C. Una PCR che quantifica il numero di molecole stampo presenti all inizio
Using TaqMan Endogenous Control Assays to select an endogenous control for experimental studies
 APPLICATION NOTE TaqMan Gene Expression Assays Using TaqMan Endogenous Control Assays to select an endogenous control for experimental studies Introduction Quantitative real-time PCR (qpcr) allows for
APPLICATION NOTE TaqMan Gene Expression Assays Using TaqMan Endogenous Control Assays to select an endogenous control for experimental studies Introduction Quantitative real-time PCR (qpcr) allows for
QuantStudio 12K Flex Real-Time PCR System. The all-in-one qpcr instrument
 QuantStudio 12K Flex Real-Time PCR System The all-in-one qpcr instrument Expand the boundaries of your research Life Technologies is taking qpcr to the next level. Designed for maximum throughput, flexibility,
QuantStudio 12K Flex Real-Time PCR System The all-in-one qpcr instrument Expand the boundaries of your research Life Technologies is taking qpcr to the next level. Designed for maximum throughput, flexibility,
PRODUCT INFORMATION...
 VIROMER RED In vitro plasmid DNA and mrna Standard Transfection PRODUCT INFORMATION... 2 GENERAL... 2 RED OR YELLOW?... 3 PROTOCOL GUIDELINES... 4 GENERAL REMARKS... 4 CELL CULTURE AND PLATING... 4 FORWARD/REVERSE
VIROMER RED In vitro plasmid DNA and mrna Standard Transfection PRODUCT INFORMATION... 2 GENERAL... 2 RED OR YELLOW?... 3 PROTOCOL GUIDELINES... 4 GENERAL REMARKS... 4 CELL CULTURE AND PLATING... 4 FORWARD/REVERSE
