Anharmonic Vibrational Signatures of DNA Bases and Watson-Crick Base Pairs
|
|
|
- Patience Booth
- 7 years ago
- Views:
Transcription
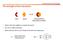
1 CHINESE JOURNAL OF CHEMICAL PHYSICS VOLUME 22, NUMBER 6 DECEMBER 27, 2009 ARTICLE Anharmonic Vibrational Signatures of DNA Bases and Watson-Crick Base Pairs Gui-xiu Wang, Xiao-yan Ma, Jian-ping Wang Beijing National Laboratory for Molecular Sciences, State Key Laboratory of Molecular Reaction Dynamics, Institute of Chemistry, Chinese Academy of Sciences, Beijing , China (Dated: Received on August 5, 2009; Accepted on September 29, 2009) Changes of molecular structure and associated charge distributions, and changes of anharmonic vibrational parameters from DNA base monomers to the Watson-Crick base pairs, have been investigated at the density functional theory level. Through examination of the NH 2, N H, and C=O stretching vibrational modes that are involved in the multiple H-bonds in the base pairs, sensitivity of their diagonal and off-diagonal anharmonicities, as well as anharmonic vibrational couplings, to the structure change are predicted. Our results reveal the intrinsic connection between the anharmonic vibrational potentials, H-bonding, and electrostatic interactions in DNA bases. Key words: Anharmonic vibration, Anharmonicity, Coupling, Two-dimensional infrared spectroscopy, DNA base I. INTRODUCTION Selective pairing between adenine-thymine (A-T) and guanine-cytosine (G-C) through multiple intermolecular H-bonds forms the well known Watson-Crick base dimers, which are basic building blocks of DNA double helical structures [1,2]. Conventional vibrational spectroscopic techniques such as linear infrared (IR) have been employed to elucidate the 3D structures of DNA [3 12]. Using structurally sensitive IR probes, for example, the C=O and N H stretching vibrations, information about base isomers can be obtained from argon and/or nitrogen matrix isolation [3 7] or gas-phase [8 12] IR experiments. The principle behind the IR method is that a vibrational mode is intrinsically connected to a set of nucleus displacements in a given molecular system [13]. An IR spectrum reflects essentially an eigenmode characteristics of the molecular system whose intermode interactions are already encoded into the spectrum. However, because of spectral congestion and the complication of intermode interactions, to extract parameters for the vibrational potential functions directly from linear IR spectra has been known to be very difficult. Fortunately, femtosecond nonlinear IR techniques such as pump/probe and two-dimensional (2D) IR in particular [14,15], have shown the capability to overcome the obstacles; a number of parameters for the vibrational potential functions, including diagonal Part of the special issue for the Chinese Chemical Society s 11th National Chemical Dynamics Symposium. Author to whom correspondence should be addressed. jwang@iccas.ac.cn, Tel.: , FAX: and off-diagonal anharmonicities, and vibrational couplings, can be retrieved simultaneously from a 2D IR spectrum, thus allowing one to study the structure and structural dynamics of the DNA in great details. The 2D IR studies of the DNA base oligomers [16,17] and IR pump/probe studies of nucleic acid base pairs [7,18] in condensed phases have been reported very recently. Ab initio computations, molecular dynamics (MD) simulations of DNA oligomers, and 2D IR spectra simulations in cm 1 region have also been reported in recent years [17,19 21]. Since the vibrational modes in all molecules are essentially anharmonic, it is of great importance to evaluate the anharmonic properties of the vibrational modes of DNA, starting from the bases and base pairs. The importance of anharmonic corrections in predicting vibrational frequency of the A-T and G-C base pairs have been pointed out earlier [22,23], however, systematic assessment of multiple anharmonic parameters on the basis of ab initio computations is still missing. Further, by combining the experimental findings and computational predictions together, one expects to be able to obtain a detailed picture of the force fields determining the 3D molecular structures and dynamics of DNA. Here we examine the anharmonic properties of DNA bases and their Watson-Crick pairs at the density functional theory (DFT) level (B3LYP/ G ). Figure 1 shows the Watson-Crick A-T and G-C pairs. The A-T pair is connected via two intermolecular H-bonds mainly: A N6 H6 O4 T and T N3 H3 N1 A, while the G-C base pair is connected via three intermolecular H-bonds: G N1 H1 N3 C, G N2 H2 O2 C, and C N4 H4 O6 G. We compare changes of bond geometries and charge distributions from the base monomers to the paired struc- 563
2 564 Chin. J. Chem. Phys., Vol. 22, No. 6 Gui-xiu Wang et al. FIG. 1 DNA Watson-Crick base pairs with atom numbering. (a) A-T. (b) G-C. H-bond distance (in Å, top) and bond-order (down) are given. tures. The strength of the formed H-bonds is analyzed in terms of bond order and orbital hyperconjugation interactions. We then present a comparison of the computed anharmonic frequencies with experimental measurements for the four bases and their dimers, to demonstrate the necessity of anharmonic corrections in frequency calculations. We examine the diagonal and offdiagonal anharmonicities of key vibrational modes associates with multiple intermolecular H-bonds that are essential for DNA helical stabilization. Efforts have been made to reveal the intrinsic connection between the anharmonic properties and the H-bonded base dimeric structures. II. COMPUTATIONAL METHODS DNA base A, T, G, and C monomers and their Watson-Crick pairs A-T and G-C were optimized without geometry constraint at the level of B3LYP/ G. Harmonic and anharmonic normal-mode frequency calculations were carried out for the optimized structures. A tight convergence was forced during the optimization, which was also needed for the followed anharmonic frequency calculations. All optimized structures have no imaginary harmonic frequency except cytosine. In the case of cytosine, one imaginary frequency has been found (154.8i cm 1 ). This belongs to the NH 2 out-of-plane vibration of the amino group. The extremely low planarization barrier of the amino group accounts for the easily happened imaginary frequency, which is well known for cytosine even at MP2/cc-pVTZ level of theory (275i cm 1 ) [24]. Intermolecular binding energies in the case of A-T and G-C were corrected for basis set superposition error (BSSE) using the standard counterpoise correction protocol [25]. No imaginary frequencies were found for the two dimer systems. The natural bond orbital (NBO) calculation [26,27] were also performed. The anharmonic frequency computations were carried at the same level of theory. Details of such calculations can be found in recent works [28,29]. Diagonal and off-diagonal anharmonicities of the 3N 6 normal modes (with N being the number of atoms), defined as ii =2ν i ν 2i and ij =ν i +ν j ν i+j respectively, were obtained. Here ν i, ν 2i, and ν i+j are the anharmonic fundamental and overtone transition frequencies for the ith mode, and the anharmonic combination transition frequency for the ith and jth modes, respectively. For A, T, G, and C monomer, and A-T and G-C dimer, the wall-clock time spent for the anharmonic frequency computations was 130, 110, 180, 70, 3280, and 3400 h respectively, using a 2-CPU 2.33 GHz computer. Since it is an all-mode anharmonic frequency computation, the computation cost increases exponentially. All calculations were carried out using Gaussian 03 [30]. Furthermore, pairwise anharmonic vibrational coupling (β ij ) between two vibrational modes of the same type was evaluated by de-mixing the wave functions of the two coupled modes, using a procedure described previously [31]. III. RESULTS AND DISCUSSION A. Geometry of DNA base pairs In the Watson-Crick A-T and G-C pairs shown in Fig.1, optimal bond distances and bond-orders are labeled for comparison for the multiple H-bonds. Higher bond-order suggests stronger H-bond, which is usually accompanied by shorter bond distance. Theory predicts A N1 H3 T has a stronger H-bond than A H6 O4 T in the A-T pair, where the A H2 O2 T H-bond is the weakest. For the G-C pair, differences in the NBO bond order for the three H-bonds are clearly shown, suggesting a compromise made between lowering the energy of the system and forming the multiple H-bonds. Further, the formation enthalpies for the A-T and G-C pairs are predicted to be and kj/mol after BSSE corrections, respectively, clearly demonstrating the difference of H-bond strengths in the two base pairs. The computed enthalpies are very close to the gas-phase experimental values ( and kj/mol) [32]. Figure 2 shows the optimized Watson-Crick A-T and G-C pairs with key structural differences indicated with respect to individual bases. There are several structural features worth mentioning: bond-length and bondangle, planarity of the H-bonded dimer, and charge redistributions. These properties change to some extent from base monomer to base pair. Bond-length and bond-angle variations are clearly seen in both the A-T and G-C pairs. The largest bond-length increase is seen in r N3 H3 of the base T (0.032 Å), while the largest bong-angle increase is seen
3 Chin. J. Chem. Phys., Vol. 22, No. 6 DNA Bases and Watson-Crick Base Pairs A (a) T (-0.003) (-0.062) A (+0.035) (+0.012)(+0.004) (+0.018) (+0.013) (+0.040) (a) (-0.056) (-0.008) (-0.025) T G C FIG. 2 Geometrical deformations of interfacial groups in the Watson-Crick base pairs with respect to individual bases. (a) A-T. (b) G-C. Bond-length change (in Å) and bondangle change (in degree) are given. (b) (-0.085) (+0.011) (+0.024) (-0.022) (+0.042) (-0.057) (+0.019) (+0.012) (-0.003) (-0.059) G (+0.053) (+0.043) (b) (+0.022) FIG. 3 Partial atomic charge distributions of interfacial atoms in the Watson-Crick base pairs and their charge variations (parentheses) with respect to those of the base monomers. (a) A-T. (b) G-C. Charges are in Coulomb (C). C in H2 N2 C2 in the base G (4.70 ). In particular, for the G-C pair, the two peripherally and symmetrically arranged NH O=C hydrogen bonds have clearly different deformations with respect to their monomeric states. The smallest bond-length and bond-angle variations are seen for the A C2H and T C2O species, indicating the presence of the weakest H-bond between the two species. Geometric changes occurred in the H-bonding participants certainly facilitate the formation of multiple H-bonds. However, the three formed H-bonds, for example, in the G-C pair, have somewhat different bond-lengths and bond-orders, indicating that the triple H-bonds can not be simultaneously optimized between two rigid molecules, in order for the system to reach a global energy minimum [33]. Structural planarity, seen in both the A-T and G-C pairs, clearly plays an important role in stabilizing the paired geometries. The conjugation between the partially double bonded C N and the well-conjugated sixnumber ring within adenine and guanine bases themselves, as well as the formation of multiple intermolecular H-bonds, are responsible for the coplanar structure of the two base pairs. For guanine, the amino in its monomeric form has a pyramidal NH 2 group with dihedral angles φ H2 N2 C2 N3 = Such a pyramidal structure becomes planar in the G-C dimer, indicating another significant structural change from the base monomer to dimer. The charge redistributions from monomer to dimer are found to be closely related to the structural changes. The NBO analysis yielded atomic charge distributions for the interfacial atoms in the two base pairs are given in Fig.3, along with the charge variations in comparison with the corresponding monomers. Clearly, quite significant charge transfer occurs. Changes of the charge distributions certainly indicate changes of the electrostatic interactions between two H-bonded base monomers. Further analysis using the NBO approach shows a hyperconjugative interaction between the H-bond acceptor orbital and the H-bond donor anti-bond orbital (also a type of charge transfer), which is believed to be the intrinsic nature of the H-bond [34]. For example, hyperconjugation occurs between the lone-pair obital of the O atom in cytosine and the anti-bonding orbital σ of the N2H2 group of guanine. Similar conclusion has been reached in an earlier study [35]. B. Anharmonic frequencies The computed anharmonic frequencies without any scaling factor are compared with argon and nitrogenmatrix [3 7], as well as in gas-phase [8 12] IR measurements. The results of NH 2 symmetric (ν s ) and asymmetric (ν a ) stretching modes, the N H, and C=O stretching modes that are involved in intermolecular H- bonds are listed in Table I. The interfacial C H and free N H stretching frequencies are also listed. Remarkable agreement between computed frequencies and experimental values is shown in Table I. The relative intensities between experiments and computations are also found to be in reasonable agreement. The harmonic and anharmonic frequencies of some of these modes and a few other modes are given in Table II. It can be seen clearly that without anharmonic corrections the vibrational frequencies would be significantly higher than experimental observables. Together, these results suggest
4 566 Chin. J. Chem. Phys., Vol. 22, No. 6 Gui-xiu Wang et al. TABLE I Comparison of the calculated anharmonic frequencies (ν i, in cm 1 ), transition intensities (I i, in km/mol), with corresponding experimental values ν exp (cm 1 ) and I exp (in relative intensity) given in parenthesis. System Mode ν i (ν exp) I i (I exp) A ν NH2, a (3565 a ) 65.4 (84 a ) ν N9H (3490 a ) 89.6 (135 a ) ν NH2, s (3448 a ) (110 a ) ν C2H (3041 a, 3093 b ) 18.3 (3 b ) T ν N1H (3479 c, 3480 d ) (130 c ) ν N3H (3432 c, 3434 d ) 66.5 (110 c ) ν C2O (1767 c ) (617 c ) ν C4O (1711 c ) (492 c ) G ν NH2, a (3503 e, 3506 f ) 41.1 ν N9H (3497 e, 3490 f ) 78.8 ν N1H (3449 e, 3456 f ) 54.1 ν C6O (1736 e ) C ν NH2, a (3565 g ) 56.0 (81 g ) ν N1H (3471 g ) 61.2 (143 g ) ν NH2, s (3441 g ) (209 g ) ν C2O (1720 g ) (872 g ) A-T ν NH2, s (3215 h ) ν C2O (1716 h ) ν C4O (1665 h ) G-C ν G:NH2, a (3352 i ) ν C:NH2, a (3532 i ) 95.4 ν G:NH2, s (3426 i ) ν N9H (3510 i ) 73.0 ν G:C6O (1706 j ) ν C:C2O (1639 j ) 34.7 a=asymmetric stretching; s=symmetric stretching. I exp was obtained by normalizing the total integrated absorbance of all absorption bands to that of the computed intensities (same for all entries). a Argon- and nitrogen-matrix [3]. b Argon-matrix [4]. c Argon-matrix [5]. d Gas-phase [11]. e Gas-phase [9]. f Gas-phase [8]. g N 2-matrix [6]. h N 2-matrix [7]. i Gas-phase [10]. j Gas-phase [12] the DFT predicted anharmonic frequencies for the DNA bases are reasonable. C. Diagonal anharmonicities The diagonal anharmonicities, ii, of the bases are predicted to be highly structure dependent. The computed ii are listed in Table II for the vibrational modes TABLE II Harmonic frequency (ω i, in cm 1 ), anharmonic frequency (ν i, in cm 1 ) and diagonal anharmonicity ( ii, in cm 1 ) of selected modes in the DNA base monomers and pairs. System Mode ω i ν i ii A ν N9H a ν NH ν C2H T ν N1H ν N3H ν C2O ν C4O G ν N9H ν N1H ν NH ν C6O C ν N1H ν NH ν C2O A-T ν A:N9H ν T:N1H ν A:NH ν A:C2H ν T:N3H ν T:C2O ν T:C4O G-C ν G:N9H ν C:N1H ν G:NH ν G:N1H ν C:NH ν G:C6O ν C:C2O a All NH 2 modes in this table are referred to the symmetric vibrational NH 2. that may be involved in H-bonds. The results of the base monomers and base pairs are compared. The N H and C H vibrational modes are highly anharmonic, which can be seen from their large ii values. Theory predicted ii of ν T:N3H is cm 1 in monomeric T, which is in reasonable agreement with recent solutionphase experimental results of the N H stretching mode in free uracil (155±15 cm 1 ) [18] and in formamide (129 cm 1 ) [36]. A dramatic increase of ii is predicted for this mode (392.6 cm 1 ) when participating in the strongest H-bond in the A-T base pair. A recent solution-phase IR pump/probe experiment of adenineuracil pair showed a similarly increased ii (230 cm 1 ) for the H-bonded N H stretching mode [18]. The ii of the symmetric NH 2 stretching mode increases significantly from free A, G, and C monomers to corre-
5 Chin. J. Chem. Phys., Vol. 22, No. 6 DNA Bases and Watson-Crick Base Pairs 567 A (a) T A (b) T that for cytosine the vibrational parameters of the key groups are well predicted (e.g. for the C=O stretching mode, ν i = cm 1 and ii =17.1 cm 1 as seen in Table I and II respectively), suggesting that the existence of one imaginary frequency does not affect significantly the results of these key modes. Further, one anharmonic frequency computation at the level of HF/6-31+G yields no imaginary frequency (data not shown). In this case, ii =15.5 cm 1 was predicted for the C=O stretching mode, which is in reasonable agreement with the DFT result. G C G C (c) (d) FIG. 4 C=O stretching modes with delocalized motions in the Watson-Crick base pairs. (a) ν T:C2O= cm 1, (b) ν T:C4O= cm 1, (c) ν G:C6O= cm 1, and (d) ν C:C2O= cm 1. sponding H-bonded dimers. These results suggest that the potential surface of the high-frequency motions can be easily influenced by structural variations. However, only slight increase of the ii of the ν A:C2H mode is seen from base monomer to dimer, indicating an insignificant influence of the structural change on this mode. This is in agreement with the NBO analysis of the H-bond order given above. The carbonyl stretching modes show relatively smaller ii values in general in comparison with the high-frequency modes. The ii value of ν C=O mode in free base G and C is around 18 cm 1, which is reasonably predicted in comparison with experimental results of cytidine and guanosine base monomers (9 and 14 cm 1 respectively) by a recent 2D IR study [16]. In the monomeric form, two carbonyl stretching modes (ν T:C2O and ν T:C4O ) in thymine differ in their ii by 9.8 cm 1, primarily due to vibrational coupling effect. Based on a simple exciton model, anharmonic vibrational coupling between the same type of mode would change the magnitude of the diagonal anharmonicities, as have been shown in peptides [29]. In the H-bonded base pairs, the ii of C=O stretching modes are found to vary quite substantially. For example, the values of ii become 23.7 and 2.7 cm 1 for the two ν C=O modes in the A-T pair, 13.2 and 3.9 cm 1 for the G-C pair. These computed results reflect the intrinsic delocalization nature of this type of normal modes, whose delocalized motions are clearly seen in Fig.4. Further, it is very likely that the ii of the ν C=O modes in the bases would behave similarly to that of the amide-i modes in peptides [14,29,37,38], since the latter is also primarily C=O stretching. Together, these results show that the vibrational modes in the DNA base pairs have structurally sensitive diagonal anharmonicities, which shall be useful to characterize the structure and dynamics of the DNA bases and DNA helices. In addition, we find D. Off-diagonal anharmonicities between intramolecular modes Table III lists the values of the off-diagonal anharmonicity ij of various modes in the base monomers and base pairs. Changes of the ij values from monomer to dimer are found to be closely related to the structural changes of the DNA bases. For base A and T, negative ij of selected intramolecular pair of modes all become positive in the H-bonded A-T pair, with large magnitude of change found in certain cases. For base T, subtle differences present in bond-length and bond-angles between N3H/C2O and N3H/C4O, however, their ij values are quite different from each other. The two ij values also increase dramatically in the A-T pair. These results show the structural sensitivity of the ij of the NH/CO pair in the DNA bases. In the A-T pair, the ij of the C2O and C4O stretching modes, is found to be only 2.4 cm 1, which is due to the remote distance between the two groups, and also possibly due to the insulating effect from the central strongly H-bonded N3H group (see Fig.4). Additionally, large frequency splitting between the two modes ( cm 1 versus cm 1, see Table II) in the A- T pair is also responsible for the small ij. Clearly the variation of the sign and magnitude of the off-diagonal anharmonicities of these interfacial vibrational modes, although may belonging to the same base, could be significantly influenced by the structural deformations due to base pairing. Similar structural dependence of the off-diagonal anharmonicities is also seen in guanine and cytosine monomers and also in the formed G-C pair. For example, in the G-C pair, the N2H2 bond of the amino group in guanine pointing to O2 of cytosine is constrained by H-bond. Nearly parallel orientation of the N2H2 and N1H bonds and closer bond distance in guanine are seen in the G-C structure, which favor the intermode interactions and thus has an increased ij (12.0 cm 1 ) in comparison with the case of free guanine (6.9 cm 1 ). Further, inspection of the other two intramolecular ij (one in guanine and one in cytosine) reveals an interesting aspect. The negative and small ij of the NH 2 /C2O pair in cytosine monomer becomes positive with larger magnitude in the G-C dimer, while the positive and
6 568 Chin. J. Chem. Phys., Vol. 22, No. 6 Gui-xiu Wang et al. TABLE III Off-diagonal anharmonicities ( ij, in cm 1 ), bond-angle (θ ij, in degree) and intermode distance (r ij, between bond-mid points, in Å) of selected pair of modes in the DNA base monomers and pairs. System Mix-mode ij θ ij r ij A NH a 2 /C2H T N3H/N1H C2O/C4O N3H/C2O N3H/C4O G N9H/N1H NH 2/N1H N1H/C6O C NH 2/C2O A T NH 2(A)/C2H(A) N3H(T)/N1H(T) C2O(T)/C4O(T) N3H(T)/C2O(T) N3H(T)/C4O(T) N9H(A)/N3H(T) N9H(A)/N1H(T) C2H(A)/C2O(T) C2H(A)/C4O(T) NH 2(A)/N3H(T) NH 2(A)/C4O(T) G C NH 2(G)/N1H(G) N9H(G)/N1H(G) N1H(G)/C6O(G) NH 2(C)/C2O(C) N1H(G)/N1H(C) N9H(G)/N1H(C) N1H(G)/ NH 2(C) NH 2(G)/C2O(C) NH 2(C)/C6O(G) C6O(G)/C2O(C) a All NH 2 modes in this table are referred to the symmetric vibrational NH 2. small ij of the N1H/C6O pair in guanine monomer becomes negative with increased magnitude in the G-C dimer. These results clearly show structural sensitivity of the intramolecular ij. E. Off-diagonal anharmonicities between intermolecular modes Structural dependent intermolecular off-diagonal anharmonicities in the case of N H/N H, C=O/C=O, and N H/C=O stretching modes are found. Two typical NH/NH pairs, N6H6(A)/N3H(T) and N1H(G)/ N4H4(C), have their ij values equal to 29.8 and cm 1 respectively. The two NH/NH pairs have very similar geometry, both having two NH bond directions being in nearly antiparallel orientations. However, their off-diagonal interactions are quite different, which must be associated with their being in different H-bonding environment. Each base pair has a C8 ring that involves the N H/N H pair (Figs. 1 3). The H-bond strengths in the two C8 rings are different. For example, N4H4(C) group participates in a stronger H-bond with shorter intermode distance seen for the N1H(G)/N4H4(C) pair in the G-C geometry, in comparison with those in a similar C8 structure in the A-T dimer. The only pair of intermolecular interacting C=O/C=O in the G-C dimer has a moderate ij (7.9 cm 1 ). Almost antiparallel bond directions favor the intermode interaction, even though the intermode distance is close to 5 Å in this case. Three pairs of N H/C=O stretching modes, NH 2 (A)/C4O(T), NH 2 (G)/C2O(C), and NH 2 (C)/C6O(G), show clearly structural dependent ij values. The H-bonding geometries, bond-angles and bond mid-point distances vary significantly from case to case, resulting in the observed difference in their ij. It is noticed that the relative strength of these ij is nicely anticorrelated with the intermode distance (Table III): the larger the ij, the shorter the distance. In addition, in spite of the highly localized nature of the C H stretching mode, and that there is only very weak H-bonding associated with the C H species, the ij values of both C2H/C2O and C2H/C4O stretching modes in the A-T pair are non-negligible and quite different, also reflecting the geometric difference of the two cases. Overall, the observed changes in the intra- and intermolecular ij could be ascribed to the change of electrostatic interactions, and to the cooperativity effect of the H-bonding interaction. Change of the electrostatic interactions between the H-bonded species is seen by the change of their charge redistributions from monomer to dimer. The H-bond cooperatively in the G-C pair may yield averaged bond elongation and averaged frequency redshift in the associated N H stretching modes. Smaller N H stretching frequencies permit vibrational resonance to occur, which may also be, for instance, the reason for the increased N1H(G)/NH 2 (C) off-diagonal anharmonicity in the G-C pair. F. Anharmonic vibrational couplings The delocalization characterizations of the vibrational modes in base monomers and pairs were examined. Due to the vibrational state delocalization, the transition intensity, frequency of a set of local modes would change, accompanying the mixing of their wave functions. Here we decouple the normal modes and obtain the local modes and inter-mode coupling using the wave function demixing approach [31]. The obtained
7 Chin. J. Chem. Phys., Vol. 22, No. 6 DNA Bases and Watson-Crick Base Pairs 569 local mode (zero-order) frequencies represent the transition energy without the perturbation from the specific coupling contribution. This forms a local mode basis that leads to the vibrational excitonic picture (eigenstates). The result of the C=O stretchings were given in Table IV, and those of the N-H stretching modes were given in Table V. In Table IV it can be seen that the coupling constant β ij of two C=O stretching modes in thymine is 5.4 cm 1, which is of very similar interaction strength revealed by the corresponding off-diagonal anharmonicity ( 4.3 cm 1 ). Similar correspondence between β ij and ij are also seen in other pair of C=O stretching modes. In the case of the G-C pair, the computed C=O/C=O intermode coupling is of the same order of magnitude as seen in a recent 2D IR experiment (ca. 10 cm 1 ) [17]. This is in agreement with the general understanding that magnitude of the off-diagonal anharmonicity is intrinsically connected to strength of intermode coupling [29]. In the case of N H stretching modes, the correspondence between β ij and ij seems to be also valid, even though the coupling strengths are much weaker than those found in case of C=O modes. In addition, it is found that all the zero-order frequencies are almost the same as their corresponding normal modes, again showing the weak interactions between N H stretching modes. Further, the calculated ij and β ij is dependent on the bond angles and bond distances, which is clearly shown in Table III, IV, and V. IV. CONCLUSION Anharmonic vibrational frequencies, anharmonicities as well as couplings in the DNA base monomers and their Watson-Crick pairs are evaluated at the DFT level using the hybrid functional B3LYP with G basis set. Several key vibrational modes, including the NH 2, N H, C=O, as well as the C H stretching modes, are considered because they are believed to be involved in the multiple H-bonds in the base pairs. By comparing the calculated anharmonic frequencies with those identified by argon and nitrogen matrix isolation and gasphase IR experiments, we find that DFT can predict vibrational frequencies of the above mentioned stretching modes reasonably. By comparing the calculated anharmonicities for base monomers and dimers, we find that both the diagonal and off-diagonal anharmonicities are structurally sensitive. Our results suggest that computations can be used to obtain anharmonic parameters for DNA bases and base pairs. The computed anharmonicities and couplings are predicted to be sensitive to molecular structure, suggesting that both can serve as structural probes for the DNA bases. Such computations can, actually provide a complete set of anharmonic parameters, which are useful to aid 2D IR spectral interpretation and further to guide 2D IR experiments. Our current work involves TABLE IV Calculated normal mode frequency (ω i, in cm 1 ), local mode frequency (ωi 0, in cm 1 ), transition intensity (I i, in km/mol), vibrational coupling constant (β ij, in cm 1 ), and off-diagonal anharmonicity ( ij, in cm 1 ) for the C=O stretching modes in DNA base monomers and pairs. Species Mode ω i I i ωi 0 β ij ij T C4O C2O A-T C4O C2O G-C C2O(C) C6O(G) TABLE V Calculated normal mode frequency (ω i, in cm 1 ), local mode frequency (ω 0 i, in cm 1 ), transition intensity (I i, in km/mol), vibrational coupling constant (β ij, in cm 1 ), and off-diagonal anharmonicity ( ij, in cm 1 ) for the N H stretching modes in DNA base monomers and pairs. Species Mode ω i I i ωi 0 β ij ij T N3H N1H G N1H N9H A-T N3H(T) a 0.4 N1H(T) N9H(A) G-C N1H(G) a 0.6 N1H(C) N9H(G) a From top to down, the β values are defined for modes listed in line 1 and 2, 1 and 3, and 2 and 3 respectively. examining the anharmonic properties of the H-bonded DNA base oligomers in larger sizes, where base stacking effect shall play a role. Such computations shall be feasible using lower-level theories with smaller basis sets. Our recent studies [29,39] have shown that the anharmonic parameters of the amide-i mode could be reasonably predicted at the level of HF/6-31+G. One expects a similar performance of this choice of method for the DNA molecules. Our results also suggest that structural deformation and associated changes in the electrostatic and H-bonding interactions from the base monomers to base pairs are responsible for the change in the anharmonic parameters. Because the anharmonicity is determined by the quartic and cubic force constants in the vibrational potentials, change of anharmonicity from monomer to dimer certainly indicates an increased number of modes involved in the potential functions, and also indicates the change in chemical environment and hence change in the anharmonic force fields. This
8 570 Chin. J. Chem. Phys., Vol. 22, No. 6 Gui-xiu Wang et al. is also true for both the diagonal anharmonicity and the intra- and intermolecular off-diagonal anharmonicities. V. ACKNOWLEDGMENT This work was supported by the National Natural Science Foundation of China (No and No ), the National High-Tech Research and Development Program of China (No.2007AA02Z139), and the Hundred Talent Fund of the Chinese Academy of Sciences. [1] G. A. Jeffrey and W. Saenger, Hydrogen Bonding in Biological Structures, 1st Edn., Berlin: Springer, 569 (1991). [2] G. R. Desiraju and T. Steiner, The Weak Hydrogen Bond in Structural Chemistry and Biology, 1st Edn., Oxford: Oxford University Press, 526 (1999). [3] M. J. Nowak, L. Lapinski, J. S. Kwiatkowski, and J. Leszczynski, J. Phys. Chem. 100, 3527 (1996). [4] K. Schoone, J. Smets, L. Houben, M. K. Van Bael, L. Adamowicz, and G. Maes, J. Phys. Chem. A 102, 4863 (1998). [5] K. Szczepaniak, M. M. Szczesniak, and W. B. Person, J. Phys. Chem. A 104, 3852 (2000). [6] M. Szczesniak, K. Szczepaniak, J. S. Kwiatkowski, K. KuBulat, and W. B. Person, J. Am. Chem. Soc. 110, 8319 (1988). [7] K. Heyne, G. M. Krishnan, and O. Kühn, J. Phys. Chem. B 112, 7909 (2008). [8] E. Nir, C. Janzen, P. Imhof, K. Kleinermanns, and M. S. de Vries, J. Chem. Phys. 115, 4604 (2001). [9] M. Mons, I. Dimicoli, F. Piuzzi, B. Tardivel, and M. Elhanine, J. Phys. Chem. A 106, 5088 (2002). [10] E. Nir, C. Janzen, P. Imhof, K. Kleinermanns, and M. S. de Vries, Phys. Chem. Chem. Phys. 4, 732 (2002). [11] R. N. Casaes, J. B. Paul, R. P. McLaughlin, R. J. Saykally, and T. van Mourik, J. Phys. Chem. A 108, (2004). [12] J. M. Bakker, I. Compagnon, G. Meijer, G. von Helden, M. Kabelǎč, P. Hobza, and M. S. de Vries, Phys. Chem. Chem. Phys. 6, 2810 (2004). [13] S. Krimm and J. Bandekar, Adv. Protein Chem. 38, 181 (1986). [14] P. Hamm, M. Lim, and R. M. Hochstrasser, J. Phys. Chem. B 102, 6123 (1998). [15] M. C. Asplund, M. T. Zanni, and R. M. Hochstrasser, Proc. Natl. Acad. Sci. USA 97, 8219 (2000). [16] A. T. Krummel, P. Mukherjee, and M. T. Zanni, J. Phys. Chem. B 107, 9165 (2003). [17] A. T. Krummel and M. T. Zanni, J. Phys. Chem. B 110, (2006). [18] S. Woutersen and G. Cristalli, J. Chem. Phys. 121, 5381 (2004). [19] C. Lee, K. Park, and M. Cho, J. Chem. Phys. 125, (2006). [20] C. Lee and M. Cho, J. Chem. Phys. 125, (2006). [21] C. Lee, K. Park, J. Kimm, S. Hahn, and M. Cho, J. Chem. Phys. 125, (2006). [22] V. Špirko, J. Šponer, and P. Hobza, J. Chem. Phys. 106, 1472 (1997). [23] B. Brauer, R. B. Gerber, M. Kabelǎč, P. Hobza, J. M. Bakker, A. G. Abo Riziq, and M. S. de Vries, J. Phys. Chem. A 109, 6974 (2005). [24] S. Wang and H. F. Schaefer III, J. Chem. Phys. 124, (2006). [25] S. F. Boys and F. Bernardi, Mol. Phys. 19, 553 (1970). [26] E. D. Glendening, A. E. Reed, J. E. Carpenter, and F. Weinhold, NBO Version 3.1, Pittsburgh PA: Gaussian, Inc., (2003). [27] A. E. Reed, L. A. Curtiss, and F. Weinhold, Chem. Rev. 88, 899 (1988). [28] V. Barone, J. Chem. Phys. 122, (2005). [29] J. Wang and R. M. Hochstrasser, J. Phys. Chem. B 110, 3798 (2006). [30] M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, J. T. Vreven, K. N. Kudin, J. C. Burant, J. M. Millam, S. S. Iyengar, J. Tomasi, V. Barone, B. Mennucci, M. Cossi, G. Scalmani, N. Rega, G. A. Petersson, H. Nakatsuji, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, M. Klene, X. Li, J. E. Knox, H. P. Hratchian, J. B. Cross, C. Adamo, J. Jaramillo, R. Gomperts, R. E. Stratmann, O. Yazyev, A. J. Austin, R. Cammi, C. Pomelli, J. W. Ochterski, P. Y. Ayala, K. Morokuma, G. A. Voth, P. Salvador, J. J. Dannenberg, V. G. Zakrzewski, S. Dapprich, A. D. Daniels, M. C. Strain, O. Farkas, D. K. Malick, A. D. Rabuck, K. Raghavachari, J. B. Foresman, J. V. Ortiz, Q. Cui, A. G. Baboul, S. Clifford, J. Cioslowski, B. B. Stefanov, A. G. Liu, Liashenko, P. Piskorz, I. Komaromi, R. L. Martin, D. J. Fox, T. Keith, M. A. Al-Laham, C. Y. Peng, A. Nanayakkara, M. Challacombe, P. M. W. Gill, B. Johnson, W. Chen, M. W. Wong, C. Gonzalez, and J. A. Pople, Gaussian 03, Revision B.05, Pittsburgh PA: Gaussian, Inc., (2003). [31] X. Ma and J. Wang, J. Phys. Chem. A 113, 6070 (2009). [32] I. K. Yanson, A. B. Teplitsky, and L. F. Sukhodub, Biopolymers 18, 1149 (1979). [33] A. Asensio, N. Kobko, and J. J. Dannenberg, J. Phys. Chem. A 107, 6441 (2003). [34] I. V. Alabugin, M. Manoharan, S. Peabody, and F. Weinhold, J. Am. Chem. Soc. 125, 5973 (2003). [35] C. F. Guerra, F. M. Bickelhaupt, J. G. Snijders, and E. J. Baerends, Chem. Eur. J. 5, 3581 (1999). [36] J. Park, J. Ha, and R. M. Hochstrasser, J. Chem. Phys. 121, 7281 (2004). [37] C. Fang, J. Wang, Y. Kim, A. K. Charnley, W. Barber- Armstrong, A. B. Smith, III, S. M. Decatur, and R. M. Hochstrasser, J. Phys. Chem. B 108, (2004). [38] K. Kumar, L. E. Sinks, J. Wang, Y. Kim, and R. M. Hochstrasser, Chem. Phys. Lett. 432, 122 (2006). [39] J. Wang, J. Phys. Chem. B 112, 4790 (2008).
IR multiphoton absorption spectra of some freon molecules used in 13 C isotope separation A. Bende 1, 2 and V. Toşa 1
 IR multiphoton absorption spectra of some freon molecules used in 13 C isotope separation A. Bende 1, 2 and V. Toşa 1 1 National Institute of Research and Development of Isotopic and Molecular Technologies,
IR multiphoton absorption spectra of some freon molecules used in 13 C isotope separation A. Bende 1, 2 and V. Toşa 1 1 National Institute of Research and Development of Isotopic and Molecular Technologies,
How To Write A Laboratory Report
 AN EXAMPLE REPORT by Cecil Dybowski, dybowski@udel.edu Here you list all names of people involved, along with email addresses. CHEMISTRY 446 Here you enter the section and group numbers. EXPERIMENT 6 Here
AN EXAMPLE REPORT by Cecil Dybowski, dybowski@udel.edu Here you list all names of people involved, along with email addresses. CHEMISTRY 446 Here you enter the section and group numbers. EXPERIMENT 6 Here
AN EXAMPLE REPORT. Cecil Dybowski (dybowski@udel.edu) Here you list all names of people involved, along with email addresses.
 AN EXAMPLE REPORT by Cecil Dybowski (dybowski@udel.edu) Here you list all names of people involved, along with email addresses. CHEMISTRY 446 Section XL Here you enter the section and group numbers. EXPERIMENT
AN EXAMPLE REPORT by Cecil Dybowski (dybowski@udel.edu) Here you list all names of people involved, along with email addresses. CHEMISTRY 446 Section XL Here you enter the section and group numbers. EXPERIMENT
Determining the Structure of an Organic Compound
 Determining the Structure of an Organic Compound The analysis of the outcome of a reaction requires that we know the full structure of the products as well as the reactants In the 19 th and early 20 th
Determining the Structure of an Organic Compound The analysis of the outcome of a reaction requires that we know the full structure of the products as well as the reactants In the 19 th and early 20 th
115 EÜFBED - Fen Bilimleri Enstitüsü Dergisi Cilt-Sayı: 3-1 Yıl: 2010 115-122
 115 AB INITIO HF AND DFT STUDIES ON MOLECULAR STRUCTURE AND VIBRATIONAL ANALYSIS OF 2,5-DIBROMOPYRIDINE 2,5-DİBROMOPİRİDİN İN MOLEKÜLER YAPISI VE TİTREŞIM FREKANSLARI ÜZERINE AB INITIO HF VE DFT ÇALIŞMALARI
115 AB INITIO HF AND DFT STUDIES ON MOLECULAR STRUCTURE AND VIBRATIONAL ANALYSIS OF 2,5-DIBROMOPYRIDINE 2,5-DİBROMOPİRİDİN İN MOLEKÜLER YAPISI VE TİTREŞIM FREKANSLARI ÜZERINE AB INITIO HF VE DFT ÇALIŞMALARI
QM/QM Study of the Coverage Effects on the Adsorption of Amino-Cyclopentene at the Si(100) Surface
 QM/QM Study of the Coverage Effects on the Adsorption of Amino-Cyclopentene at the Si(100) Surface HUGO R. R. SANTOS, 1 GREGORI UJAQUE, 2 MARIA J. RAMOS, 1 JOSÉ A. N. F. GOMES 1 1 REQUIMTE, Departamento
QM/QM Study of the Coverage Effects on the Adsorption of Amino-Cyclopentene at the Si(100) Surface HUGO R. R. SANTOS, 1 GREGORI UJAQUE, 2 MARIA J. RAMOS, 1 JOSÉ A. N. F. GOMES 1 1 REQUIMTE, Departamento
1 The water molecule and hydrogen bonds in water
 The Physics and Chemistry of Water 1 The water molecule and hydrogen bonds in water Stoichiometric composition H 2 O the average lifetime of a molecule is 1 ms due to proton exchange (catalysed by acids
The Physics and Chemistry of Water 1 The water molecule and hydrogen bonds in water Stoichiometric composition H 2 O the average lifetime of a molecule is 1 ms due to proton exchange (catalysed by acids
Redbooks Paper. Life Sciences Applications on IBM POWER5 and AIX 5L Version 5.3: Virtualization Exploitation through Micro-Partitioning Implementation
 Redbooks Paper Carlos P. Sosa Balaji V. Atyam Peter Heyrman Naresh Nayar Jeri Hilsabeck Life Sciences Applications on IBM POWER5 and AIX 5L Version 5.3: Virtualization Exploitation through Micro-Partitioning
Redbooks Paper Carlos P. Sosa Balaji V. Atyam Peter Heyrman Naresh Nayar Jeri Hilsabeck Life Sciences Applications on IBM POWER5 and AIX 5L Version 5.3: Virtualization Exploitation through Micro-Partitioning
PCV Project: Excitons in Molecular Spectroscopy
 PCV Project: Excitons in Molecular Spectroscopy Introduction The concept of excitons was first introduced by Frenkel (1) in 1931 as a general excitation delocalization mechanism to account for the ability
PCV Project: Excitons in Molecular Spectroscopy Introduction The concept of excitons was first introduced by Frenkel (1) in 1931 as a general excitation delocalization mechanism to account for the ability
Infrared Spectroscopy: Theory
 u Chapter 15 Infrared Spectroscopy: Theory An important tool of the organic chemist is Infrared Spectroscopy, or IR. IR spectra are acquired on a special instrument, called an IR spectrometer. IR is used
u Chapter 15 Infrared Spectroscopy: Theory An important tool of the organic chemist is Infrared Spectroscopy, or IR. IR spectra are acquired on a special instrument, called an IR spectrometer. IR is used
Supporting Information
 Supporting Information A Photoresponsive Smart Covalent Organic Framework** Ning Huang, Xuesong Ding, Jangbae Kim, Hyotcherl Ihee, and Donglin Jiang* ange_201503902_sm_miscellaneous_information.pdf Contents
Supporting Information A Photoresponsive Smart Covalent Organic Framework** Ning Huang, Xuesong Ding, Jangbae Kim, Hyotcherl Ihee, and Donglin Jiang* ange_201503902_sm_miscellaneous_information.pdf Contents
Infrared Spectroscopy 紅 外 線 光 譜 儀
 Infrared Spectroscopy 紅 外 線 光 譜 儀 Introduction Spectroscopy is an analytical technique which helps determine structure. It destroys little or no sample (nondestructive method). The amount of light absorbed
Infrared Spectroscopy 紅 外 線 光 譜 儀 Introduction Spectroscopy is an analytical technique which helps determine structure. It destroys little or no sample (nondestructive method). The amount of light absorbed
Adipik asit üzerine ab initio hesaplamaları. Ab initio calculations on adipic acid
 SAÜ. Fen Bil. Der. 17. Cilt, 3. Sayı, s. 357-362, 2013 SAU J. Sci. Vol 17, No 3, p. 357-362, 2013 Adipik asit üzerine ab initio hesaplamaları Mustafa Çetin 1*, Adil Başoğlu 1, Davut Avcı 1, Yusuf Atalay
SAÜ. Fen Bil. Der. 17. Cilt, 3. Sayı, s. 357-362, 2013 SAU J. Sci. Vol 17, No 3, p. 357-362, 2013 Adipik asit üzerine ab initio hesaplamaları Mustafa Çetin 1*, Adil Başoğlu 1, Davut Avcı 1, Yusuf Atalay
Hydrogen Bonds The electrostatic nature of hydrogen bonds
 Hydrogen Bonds Hydrogen bonds have played an incredibly important role in the history of structural biology. Both the structure of DNA and of protein a-helices and b-sheets were predicted based largely
Hydrogen Bonds Hydrogen bonds have played an incredibly important role in the history of structural biology. Both the structure of DNA and of protein a-helices and b-sheets were predicted based largely
Organic Chemistry Tenth Edition
 Organic Chemistry Tenth Edition T. W. Graham Solomons Craig B. Fryhle Welcome to CHM 22 Organic Chemisty II Chapters 2 (IR), 9, 3-20. Chapter 2 and Chapter 9 Spectroscopy (interaction of molecule with
Organic Chemistry Tenth Edition T. W. Graham Solomons Craig B. Fryhle Welcome to CHM 22 Organic Chemisty II Chapters 2 (IR), 9, 3-20. Chapter 2 and Chapter 9 Spectroscopy (interaction of molecule with
Dedicado con motivo del 75 aniversario de la fundación de la Sociedad Química del Perú
 163 Dedicado con motivo del 75 aniversario de la fundación de la Sociedad Química del Perú FORMACIÓ DE COMPLEJO OLIGOUCLEAR CO EL LIGADO BIS-BIDETADO ',', ''', '''-TETRAETILO-,''- PIRIDIA-2,6-DICARBOILO-BIS(TIOUREA)
163 Dedicado con motivo del 75 aniversario de la fundación de la Sociedad Química del Perú FORMACIÓ DE COMPLEJO OLIGOUCLEAR CO EL LIGADO BIS-BIDETADO ',', ''', '''-TETRAETILO-,''- PIRIDIA-2,6-DICARBOILO-BIS(TIOUREA)
Symmetric Stretch: allows molecule to move through space
 BACKGROUND INFORMATION Infrared Spectroscopy Before introducing the subject of IR spectroscopy, we must first review some aspects of the electromagnetic spectrum. The electromagnetic spectrum is composed
BACKGROUND INFORMATION Infrared Spectroscopy Before introducing the subject of IR spectroscopy, we must first review some aspects of the electromagnetic spectrum. The electromagnetic spectrum is composed
INFRARED SPECTROSCOPY (IR)
 INFRARED SPECTROSCOPY (IR) Theory and Interpretation of IR spectra ASSIGNED READINGS Introduction to technique 25 (p. 833-834 in lab textbook) Uses of the Infrared Spectrum (p. 847-853) Look over pages
INFRARED SPECTROSCOPY (IR) Theory and Interpretation of IR spectra ASSIGNED READINGS Introduction to technique 25 (p. 833-834 in lab textbook) Uses of the Infrared Spectrum (p. 847-853) Look over pages
Section Activity #1: Fill out the following table for biology s most common elements assuming that each atom is neutrally charged.
 LS1a Fall 2014 Section Week #1 I. Valence Electrons and Bonding The number of valence (outer shell) electrons in an atom determines how many bonds it can form. Knowing the number of valence electrons present
LS1a Fall 2014 Section Week #1 I. Valence Electrons and Bonding The number of valence (outer shell) electrons in an atom determines how many bonds it can form. Knowing the number of valence electrons present
Chapter 2 Polar Covalent Bonds; Acids and Bases
 John E. McMurry http://www.cengage.com/chemistry/mcmurry Chapter 2 Polar Covalent Bonds; Acids and Bases Javier E. Horta, M.D., Ph.D. University of Massachusetts Lowell Polar Covalent Bonds: Electronegativity
John E. McMurry http://www.cengage.com/chemistry/mcmurry Chapter 2 Polar Covalent Bonds; Acids and Bases Javier E. Horta, M.D., Ph.D. University of Massachusetts Lowell Polar Covalent Bonds: Electronegativity
for excitation to occur, there must be an exact match between the frequency of the applied radiation and the frequency of the vibration
 ! = 1 2"c k (m + M) m M wavenumbers! =!/c = 1/" wavelength frequency! units: cm 1 for excitation to occur, there must be an exact match between the frequency of the applied radiation and the frequency
! = 1 2"c k (m + M) m M wavenumbers! =!/c = 1/" wavelength frequency! units: cm 1 for excitation to occur, there must be an exact match between the frequency of the applied radiation and the frequency
Ab initio study of gas-phase sulphuric acid hydrates containing 1 to 3 water molecules
 Ab initio study of gas-phase sulphuric acid hydrates containing 1 to 3 water molecules Hanna Arstila Department of Physics, P.O. Box 9, 00014 University of Helsinki, Finland Kari Laasonen Department of
Ab initio study of gas-phase sulphuric acid hydrates containing 1 to 3 water molecules Hanna Arstila Department of Physics, P.O. Box 9, 00014 University of Helsinki, Finland Kari Laasonen Department of
Journal of the University of Chemical Technology and Metallurgy, 42, 2, 2007. 2) are in C 1
 Journal of the University of Chemical M. Georgiev, Technology D. Stoilova and Metallurgy, 42, 2, 2007, 211-216 METAL-WATER INTERACTINS AND HYDRGEN BND STRENGTH M. Georgiev 1, D. Stoilova 2 1 University
Journal of the University of Chemical M. Georgiev, Technology D. Stoilova and Metallurgy, 42, 2, 2007, 211-216 METAL-WATER INTERACTINS AND HYDRGEN BND STRENGTH M. Georgiev 1, D. Stoilova 2 1 University
Predicting Magnetic Properties with ChemDraw and Gaussian
 Predicting Magnetic Properties with ChemDraw and Gaussian By James R. Cheeseman and Æleen Frisch Introduction NMR chemical shifts are an important tool in characterizing molecular systems and structures.
Predicting Magnetic Properties with ChemDraw and Gaussian By James R. Cheeseman and Æleen Frisch Introduction NMR chemical shifts are an important tool in characterizing molecular systems and structures.
The Unshifted Atom-A Simpler Method of Deriving Vibrational Modes of Molecular Symmetries
 Est. 1984 ORIENTAL JOURNAL OF CHEMISTRY An International Open Free Access, Peer Reviewed Research Journal www.orientjchem.org ISSN: 0970-020 X CODEN: OJCHEG 2012, Vol. 28, No. (1): Pg. 189-202 The Unshifted
Est. 1984 ORIENTAL JOURNAL OF CHEMISTRY An International Open Free Access, Peer Reviewed Research Journal www.orientjchem.org ISSN: 0970-020 X CODEN: OJCHEG 2012, Vol. 28, No. (1): Pg. 189-202 The Unshifted
Thienopyrrole-expanded BODIPY as a Potential NIR Photosensitizer for Photodynamic Therapy
 Thienopyrrole-expanded BODIPY as a Potential NIR Photosensitizer for Photodynamic Therapy 1. Experimental details 1.1 Methods Yongchao Yang, Qiuli Guo, Huachao Chen, Zhikuan Zhou, Zhen Shen* 1.2 Single
Thienopyrrole-expanded BODIPY as a Potential NIR Photosensitizer for Photodynamic Therapy 1. Experimental details 1.1 Methods Yongchao Yang, Qiuli Guo, Huachao Chen, Zhikuan Zhou, Zhen Shen* 1.2 Single
Group Theory and Chemistry
 Group Theory and Chemistry Outline: Raman and infra-red spectroscopy Symmetry operations Point Groups and Schoenflies symbols Function space and matrix representation Reducible and irreducible representation
Group Theory and Chemistry Outline: Raman and infra-red spectroscopy Symmetry operations Point Groups and Schoenflies symbols Function space and matrix representation Reducible and irreducible representation
Organic Spectroscopy
 1 Organic Spectroscopy Second Year, Michaelmas term, 8 lectures: Dr TDW Claridge & Prof BG Davis Lectures 1 4 highlight the importance of spectroscopic methods in the structural elucidation of organic
1 Organic Spectroscopy Second Year, Michaelmas term, 8 lectures: Dr TDW Claridge & Prof BG Davis Lectures 1 4 highlight the importance of spectroscopic methods in the structural elucidation of organic
Propensity of Formate, Acetate, Benzoate, and Phenolate for the Aqueous. Solution/Vapour Interface: Surface Tension Measurements and Molecular
 Propensity of Formate, Acetate, Benzoate, and Phenolate for the Aqueous Solution/Vapour Interface: Surface Tension Measurements and Molecular Dynamics Simulations Babak Minofar, 1 Pavel Jungwirth, 1* Manash
Propensity of Formate, Acetate, Benzoate, and Phenolate for the Aqueous Solution/Vapour Interface: Surface Tension Measurements and Molecular Dynamics Simulations Babak Minofar, 1 Pavel Jungwirth, 1* Manash
Theoretical Analysis of a Carbon-Carbon Dimer Placement Tool for Diamond Mechanosynthesis
 JOURNAL OF NANOSCIENCE AND NANOTECHNOLOGY Theoretical Analysis of a Carbon-Carbon Dimer Placement Tool for Diamond Mechanosynthesis Ralph C. Merkle, and Robert A. Freitas, Jr. Zyvex Corp., 1321 North Plano
JOURNAL OF NANOSCIENCE AND NANOTECHNOLOGY Theoretical Analysis of a Carbon-Carbon Dimer Placement Tool for Diamond Mechanosynthesis Ralph C. Merkle, and Robert A. Freitas, Jr. Zyvex Corp., 1321 North Plano
DETERMINACIÓN DE ESTRUCTURAS ORGÁNICAS (ORGANIC SPECTROSCOPY) IR SPECTROSCOPY
 DETERMINACIÓN DE ESTRUCTURAS ORGÁNICAS (ORGANIC SPECTROSCOPY) IR SPECTROSCOPY Hermenegildo García Gómez Departamento de Química Instituto de Tecnología Química Universidad Politécnica de Valencia 46022
DETERMINACIÓN DE ESTRUCTURAS ORGÁNICAS (ORGANIC SPECTROSCOPY) IR SPECTROSCOPY Hermenegildo García Gómez Departamento de Química Instituto de Tecnología Química Universidad Politécnica de Valencia 46022
Proton Nuclear Magnetic Resonance ( 1 H-NMR) Spectroscopy
 Proton Nuclear Magnetic Resonance ( 1 H-NMR) Spectroscopy Theory behind NMR: In the late 1940 s, physical chemists originally developed NMR spectroscopy to study different properties of atomic nuclei,
Proton Nuclear Magnetic Resonance ( 1 H-NMR) Spectroscopy Theory behind NMR: In the late 1940 s, physical chemists originally developed NMR spectroscopy to study different properties of atomic nuclei,
Application Note AN4
 TAKING INVENTIVE STEPS IN INFRARED. MINIATURE INFRARED GAS SENSORS GOLD SERIES UK Patent App. No. 2372099A USA Patent App. No. 09/783,711 World Patents Pending INFRARED SPECTROSCOPY Application Note AN4
TAKING INVENTIVE STEPS IN INFRARED. MINIATURE INFRARED GAS SENSORS GOLD SERIES UK Patent App. No. 2372099A USA Patent App. No. 09/783,711 World Patents Pending INFRARED SPECTROSCOPY Application Note AN4
Proton Nuclear Magnetic Resonance Spectroscopy
 Proton Nuclear Magnetic Resonance Spectroscopy Introduction: The NMR Spectrum serves as a great resource in determining the structure of an organic compound by revealing the hydrogen and carbon skeleton.
Proton Nuclear Magnetic Resonance Spectroscopy Introduction: The NMR Spectrum serves as a great resource in determining the structure of an organic compound by revealing the hydrogen and carbon skeleton.
RESONANCE, USING CURVED ARROWS AND ACID-BASE REACTIONS
 RESONANCE, USING CURVED ARROWS AND ACID-BASE REACTIONS A STUDENT SHOULD BE ABLE TO: 1. Properly use curved arrows to draw resonance structures: the tail and the head of every arrow must be drawn in exactly
RESONANCE, USING CURVED ARROWS AND ACID-BASE REACTIONS A STUDENT SHOULD BE ABLE TO: 1. Properly use curved arrows to draw resonance structures: the tail and the head of every arrow must be drawn in exactly
Use the Force! Noncovalent Molecular Forces
 Use the Force! Noncovalent Molecular Forces Not quite the type of Force we re talking about Before we talk about noncovalent molecular forces, let s talk very briefly about covalent bonds. The Illustrated
Use the Force! Noncovalent Molecular Forces Not quite the type of Force we re talking about Before we talk about noncovalent molecular forces, let s talk very briefly about covalent bonds. The Illustrated
Chapter 2 Polar Covalent Bonds: Acids and Bases
 John E. McMurry www.cengage.com/chemistry/mcmurry Chapter 2 Polar Covalent Bonds: Acids and Bases Modified by Dr. Daniela R. Radu Why This Chapter? Description of basic ways chemists account for chemical
John E. McMurry www.cengage.com/chemistry/mcmurry Chapter 2 Polar Covalent Bonds: Acids and Bases Modified by Dr. Daniela R. Radu Why This Chapter? Description of basic ways chemists account for chemical
IR Applied to Isomer Analysis
 DiscovIR-LC TM Application Note 025 April 2008 Deposition and Detection System IR Applied to Isomer Analysis Infrared spectra provide valuable information about local configurations of atoms in molecules.
DiscovIR-LC TM Application Note 025 April 2008 Deposition and Detection System IR Applied to Isomer Analysis Infrared spectra provide valuable information about local configurations of atoms in molecules.
NMR and IR spectra & vibrational analysis
 Lab 5: NMR and IR spectra & vibrational analysis A brief theoretical background 1 Some of the available chemical quantum methods for calculating NMR chemical shifts are based on the Hartree-Fock self-consistent
Lab 5: NMR and IR spectra & vibrational analysis A brief theoretical background 1 Some of the available chemical quantum methods for calculating NMR chemical shifts are based on the Hartree-Fock self-consistent
Mechanism of the Acid-Catalyzed Si-O Bond Cleavage in Siloxanes and Siloxanols. A Theoretical Study
 Organometallics 2002, 21, 2165-2175 2165 Mechanism of the Acid-Catalyzed Si-O Bond Cleavage in Siloxanes and Siloxanols. A Theoretical Study Marek Cypryk*, and Yitzhak Apeloig Center of Molecular and Macromolecular
Organometallics 2002, 21, 2165-2175 2165 Mechanism of the Acid-Catalyzed Si-O Bond Cleavage in Siloxanes and Siloxanols. A Theoretical Study Marek Cypryk*, and Yitzhak Apeloig Center of Molecular and Macromolecular
CHAPTER 6 Chemical Bonding
 CHAPTER 6 Chemical Bonding SECTION 1 Introduction to Chemical Bonding OBJECTIVES 1. Define Chemical bond. 2. Explain why most atoms form chemical bonds. 3. Describe ionic and covalent bonding.. 4. Explain
CHAPTER 6 Chemical Bonding SECTION 1 Introduction to Chemical Bonding OBJECTIVES 1. Define Chemical bond. 2. Explain why most atoms form chemical bonds. 3. Describe ionic and covalent bonding.. 4. Explain
Module 3 : Molecular Spectroscopy Lecture 13 : Rotational and Vibrational Spectroscopy
 Module 3 : Molecular Spectroscopy Lecture 13 : Rotational and Vibrational Spectroscopy Objectives After studying this lecture, you will be able to Calculate the bond lengths of diatomics from the value
Module 3 : Molecular Spectroscopy Lecture 13 : Rotational and Vibrational Spectroscopy Objectives After studying this lecture, you will be able to Calculate the bond lengths of diatomics from the value
Molecular Spectroscopy
 Molecular Spectroscopy UV-Vis Spectroscopy Absorption Characteristics of Some Common Chromophores UV-Vis Spectroscopy Absorption Characteristics of Aromatic Compounds UV-Vis Spectroscopy Effect of extended
Molecular Spectroscopy UV-Vis Spectroscopy Absorption Characteristics of Some Common Chromophores UV-Vis Spectroscopy Absorption Characteristics of Aromatic Compounds UV-Vis Spectroscopy Effect of extended
Mulliken suggested to split the shared density 50:50. Then the electrons associated with the atom k are given by:
 1 17. Population Analysis Population analysis is the study of charge distribution within molecules. The intention is to accurately model partial charge magnitude and location within a molecule. This can
1 17. Population Analysis Population analysis is the study of charge distribution within molecules. The intention is to accurately model partial charge magnitude and location within a molecule. This can
Why? Intermolecular Forces. Intermolecular Forces. Chapter 12 IM Forces and Liquids. Covalent Bonding Forces for Comparison of Magnitude
 1 Why? Chapter 1 Intermolecular Forces and Liquids Why is water usually a liquid and not a gas? Why does liquid water boil at such a high temperature for such a small molecule? Why does ice float on water?
1 Why? Chapter 1 Intermolecular Forces and Liquids Why is water usually a liquid and not a gas? Why does liquid water boil at such a high temperature for such a small molecule? Why does ice float on water?
POLAR COVALENT BONDS Ionic compounds form repeating. Covalent compounds form distinct. Consider adding to NaCl(s) vs. H 2 O(s):
 POLAR COVALENT BONDS Ionic compounds form repeating. Covalent compounds form distinct. Consider adding to NaCl(s) vs. H 2 O(s): Sometimes when atoms of two different elements form a bond by sharing an
POLAR COVALENT BONDS Ionic compounds form repeating. Covalent compounds form distinct. Consider adding to NaCl(s) vs. H 2 O(s): Sometimes when atoms of two different elements form a bond by sharing an
Chapter 9. Chemical reactivity of molecules depends on the nature of the bonds between the atoms as well on its 3D structure
 Chapter 9 Molecular Geometry & Bonding Theories I) Molecular Geometry (Shapes) Chemical reactivity of molecules depends on the nature of the bonds between the atoms as well on its 3D structure Molecular
Chapter 9 Molecular Geometry & Bonding Theories I) Molecular Geometry (Shapes) Chemical reactivity of molecules depends on the nature of the bonds between the atoms as well on its 3D structure Molecular
A pure covalent bond is an equal sharing of shared electron pair(s) in a bond. A polar covalent bond is an unequal sharing.
 CHAPTER EIGHT BNDING: GENERAL CNCEPT or Review 1. Electronegativity is the ability of an atom in a molecule to attract electrons to itself. Electronegativity is a bonding term. Electron affinity is the
CHAPTER EIGHT BNDING: GENERAL CNCEPT or Review 1. Electronegativity is the ability of an atom in a molecule to attract electrons to itself. Electronegativity is a bonding term. Electron affinity is the
Determination of Molecular Structure by MOLECULAR SPECTROSCOPY
 Determination of Molecular Structure by MOLEULAR SPETROSOPY hemistry 3 B.Z. Shakhashiri Fall 29 Much of what we know about molecular structure has been learned by observing and analyzing how electromagnetic
Determination of Molecular Structure by MOLEULAR SPETROSOPY hemistry 3 B.Z. Shakhashiri Fall 29 Much of what we know about molecular structure has been learned by observing and analyzing how electromagnetic
Order of Filling Subshells
 Bonding: General Concepts Ionic Bonds Sections 13.2-13.6 Covalent Bonds Section 13.7 Covalent Bond Energy & Chemical Reactions Section 13.8-13.9 Lewis Structures Sections 13.10-13.12 VSEPR Theory Section
Bonding: General Concepts Ionic Bonds Sections 13.2-13.6 Covalent Bonds Section 13.7 Covalent Bond Energy & Chemical Reactions Section 13.8-13.9 Lewis Structures Sections 13.10-13.12 VSEPR Theory Section
Experiment 11. Infrared Spectroscopy
 Chem 22 Spring 2010 Experiment 11 Infrared Spectroscopy Pre-lab preparation. (1) In Ch 5 and 12 of the text you will find examples of the most common functional groups in organic molecules. In your notebook,
Chem 22 Spring 2010 Experiment 11 Infrared Spectroscopy Pre-lab preparation. (1) In Ch 5 and 12 of the text you will find examples of the most common functional groups in organic molecules. In your notebook,
CHEM 51LB EXP 1 SPECTROSCOPIC METHODS: INFRARED AND NUCLEAR MAGNETIC RESONANCE SPECTROSCOPY
 CHEM 51LB EXP 1 SPECTRSCPIC METHDS: INFRARED AND NUCLEAR MAGNETIC RESNANCE SPECTRSCPY REACTINS: None TECHNIQUES: IR Spectroscopy, NMR Spectroscopy Infrared (IR) and nuclear magnetic resonance (NMR) spectroscopy
CHEM 51LB EXP 1 SPECTRSCPIC METHDS: INFRARED AND NUCLEAR MAGNETIC RESNANCE SPECTRSCPY REACTINS: None TECHNIQUES: IR Spectroscopy, NMR Spectroscopy Infrared (IR) and nuclear magnetic resonance (NMR) spectroscopy
Chapter 9 - Covalent Bonding: Orbitals
 Chapter 9 - Covalent Bonding: Orbitals 9.1 Hybridization and the Localized Electron Model A. Hybridization 1. The mixing of two or more atomic orbitals of similar energies on the same atom to produce new
Chapter 9 - Covalent Bonding: Orbitals 9.1 Hybridization and the Localized Electron Model A. Hybridization 1. The mixing of two or more atomic orbitals of similar energies on the same atom to produce new
Applications of Quantum Chemistry HΨ = EΨ
 Applications of Quantum Chemistry HΨ = EΨ Areas of Application Explaining observed phenomena (e.g., spectroscopy) Simulation and modeling: make predictions New techniques/devices use special quantum properties
Applications of Quantum Chemistry HΨ = EΨ Areas of Application Explaining observed phenomena (e.g., spectroscopy) Simulation and modeling: make predictions New techniques/devices use special quantum properties
Ultraviolet Spectroscopy
 Ultraviolet Spectroscopy The wavelength of UV and visible light are substantially shorter than the wavelength of infrared radiation. The UV spectrum ranges from 100 to 400 nm. A UV-Vis spectrophotometer
Ultraviolet Spectroscopy The wavelength of UV and visible light are substantially shorter than the wavelength of infrared radiation. The UV spectrum ranges from 100 to 400 nm. A UV-Vis spectrophotometer
K'NEX DNA Models. Developed by Dr. Gary Benson Department of Biomathematical Sciences Mount Sinai School of Medicine
 KNEX DNA Models Introduction Page 1 of 11 All photos by Kevin Kelliher. To download an Acrobat pdf version of this website Click here. K'NEX DNA Models Developed by Dr. Gary Benson Department of Biomathematical
KNEX DNA Models Introduction Page 1 of 11 All photos by Kevin Kelliher. To download an Acrobat pdf version of this website Click here. K'NEX DNA Models Developed by Dr. Gary Benson Department of Biomathematical
Determination of Equilibrium Constants using NMR Spectrscopy
 CHEM 331L Physical Chemistry Laboratory Revision 1.0 Determination of Equilibrium Constants using NMR Spectrscopy In this laboratory exercise we will measure a chemical equilibrium constant using key proton
CHEM 331L Physical Chemistry Laboratory Revision 1.0 Determination of Equilibrium Constants using NMR Spectrscopy In this laboratory exercise we will measure a chemical equilibrium constant using key proton
Time out states and transitions
 Time out states and transitions Spectroscopy transitions between energy states of a molecule excited by absorption or emission of a photon hn = DE = E i - E f Energy levels due to interactions between
Time out states and transitions Spectroscopy transitions between energy states of a molecule excited by absorption or emission of a photon hn = DE = E i - E f Energy levels due to interactions between
Exploring Potential Energy Surfaces for Chemical Reactions: An Overview of Some Practical Methods
 Feature Article Exploring Potential Energy Surfaces for Chemical Reactions: An Overview of Some Practical Methods H. BERNHARD SCHLEGEL Department of Chemistry, Wayne State University, Detroit, Michigan
Feature Article Exploring Potential Energy Surfaces for Chemical Reactions: An Overview of Some Practical Methods H. BERNHARD SCHLEGEL Department of Chemistry, Wayne State University, Detroit, Michigan
Quantum Calculations on Hydrogen Bonds in Certain Water Clusters Show Cooperative Effects
 J. Chem. Theory Comput. 2007, 3, 103-114 103 Quantum Calculations on Hydrogen Bonds in Certain Water Clusters Show Cooperative Effects Vasiliy S. Znamenskiy and Michael E. Green* Department of Chemistry,
J. Chem. Theory Comput. 2007, 3, 103-114 103 Quantum Calculations on Hydrogen Bonds in Certain Water Clusters Show Cooperative Effects Vasiliy S. Znamenskiy and Michael E. Green* Department of Chemistry,
The Fundamentals of Infrared Spectroscopy. Joe Van Gompel, PhD
 TN-100 The Fundamentals of Infrared Spectroscopy The Principles of Infrared Spectroscopy Joe Van Gompel, PhD Spectroscopy is the study of the interaction of electromagnetic radiation with matter. The electromagnetic
TN-100 The Fundamentals of Infrared Spectroscopy The Principles of Infrared Spectroscopy Joe Van Gompel, PhD Spectroscopy is the study of the interaction of electromagnetic radiation with matter. The electromagnetic
Communicated March 31, 1951 CHR CHR CHR. *H* zz '" *H _ 0.-...H / k C,.. CHR CNR CHR CHR CHR *HN/' N 'H_N/' H_./ - H-(H.
 VOL. 37, 1951 CHEMISTR Y: PA ULING AND COREY 251 THE PLEATED SHEET, A NEW LAYER CONFIGURATION OF POL YPEPTIDE CHAINS BY LINUS PAULING AND ROBERT B. COREY GATES AND CRELLIN LABORATORIES OF CHEMISTRY,* CALIFORNIA
VOL. 37, 1951 CHEMISTR Y: PA ULING AND COREY 251 THE PLEATED SHEET, A NEW LAYER CONFIGURATION OF POL YPEPTIDE CHAINS BY LINUS PAULING AND ROBERT B. COREY GATES AND CRELLIN LABORATORIES OF CHEMISTRY,* CALIFORNIA
Chapter 11: Molecular Structure of DNA and RNA
 Chapter 11: Molecular Structure of DNA and RNA Student Learning Objectives Upon completion of this chapter you should be able to: 1. Understand the major experiments that led to the discovery of DNA as
Chapter 11: Molecular Structure of DNA and RNA Student Learning Objectives Upon completion of this chapter you should be able to: 1. Understand the major experiments that led to the discovery of DNA as
The strength of the interaction
 The strength of the interaction Host Guest Supramolecule (host-guest complex) When is the host capable to recognize the guest? How do we define selectivity Which element will we use to design the host
The strength of the interaction Host Guest Supramolecule (host-guest complex) When is the host capable to recognize the guest? How do we define selectivity Which element will we use to design the host
H 2O gas: molecules are very far apart
 Non-Covalent Molecular Forces 2/27/06 3/1/06 How does this reaction occur: H 2 O (liquid) H 2 O (gas)? Add energy H 2O gas: molecules are very far apart H 2O liquid: bonding between molecules Use heat
Non-Covalent Molecular Forces 2/27/06 3/1/06 How does this reaction occur: H 2 O (liquid) H 2 O (gas)? Add energy H 2O gas: molecules are very far apart H 2O liquid: bonding between molecules Use heat
Chemometric Study on Molecules with Anticancer Properties
 80 Chemometric Study on Molecules with Anticancer Properties João Elias Vidueira Ferreira 1, Antonio Florêncio de Figueiredo 2, Jardel Pinto Barbosa 3 and José Ciríaco Pinheiro 3 1 Universidade do Estado
80 Chemometric Study on Molecules with Anticancer Properties João Elias Vidueira Ferreira 1, Antonio Florêncio de Figueiredo 2, Jardel Pinto Barbosa 3 and José Ciríaco Pinheiro 3 1 Universidade do Estado
DNA is found in all organisms from the smallest bacteria to humans. DNA has the same composition and structure in all organisms!
 Biological Sciences Initiative HHMI DNA omponents and Structure Introduction Nucleic acids are molecules that are essential to, and characteristic of, life on Earth. There are two basic types of nucleic
Biological Sciences Initiative HHMI DNA omponents and Structure Introduction Nucleic acids are molecules that are essential to, and characteristic of, life on Earth. There are two basic types of nucleic
Solving Spectroscopy Problems
 Solving Spectroscopy Problems The following is a detailed summary on how to solve spectroscopy problems, key terms are highlighted in bold and the definitions are from the illustrated glossary on Dr. Hardinger
Solving Spectroscopy Problems The following is a detailed summary on how to solve spectroscopy problems, key terms are highlighted in bold and the definitions are from the illustrated glossary on Dr. Hardinger
Composition of the Atmosphere. Outline Atmospheric Composition Nitrogen and Oxygen Lightning Homework
 Molecules of the Atmosphere The present atmosphere consists mainly of molecular nitrogen (N2) and molecular oxygen (O2) but it has dramatically changed in composition from the beginning of the solar system.
Molecules of the Atmosphere The present atmosphere consists mainly of molecular nitrogen (N2) and molecular oxygen (O2) but it has dramatically changed in composition from the beginning of the solar system.
Features of the formation of hydrogen bonds in solutions of polysaccharides during their use in various industrial processes. V.Mank a, O.
 Features of the formation of hydrogen bonds in solutions of polysaccharides during their use in various industrial processes. V.Mank a, O. Melnyk b a National University of life and environmental sciences
Features of the formation of hydrogen bonds in solutions of polysaccharides during their use in various industrial processes. V.Mank a, O. Melnyk b a National University of life and environmental sciences
Introduction OLEDs OTFTs OPVC Summary. Organic Electronics. Felix Buth. Walter Schottky Institut, TU München. Joint Advanced Student School 2008
 Felix Buth Joint Advanced Student School 2008 Outline 1 Introduction Difference organic/inorganic semiconductors From molecular orbitals to the molecular crystal 2 Organic Light Emitting Diodes Basic Principals
Felix Buth Joint Advanced Student School 2008 Outline 1 Introduction Difference organic/inorganic semiconductors From molecular orbitals to the molecular crystal 2 Organic Light Emitting Diodes Basic Principals
2012 HORIBA Scientific. All rights reserved. 2012 HORIBA Scientific. All rights reserved.
 Raman Spectroscopy for proteins Catalina DAVID Ph.D. application scientist Outline Raman spectroscopy in few words What is Raman spectroscopy? What is the information we can get? Basics of Raman analysis
Raman Spectroscopy for proteins Catalina DAVID Ph.D. application scientist Outline Raman spectroscopy in few words What is Raman spectroscopy? What is the information we can get? Basics of Raman analysis
Infrared Spectroscopy
 Infrared Spectroscopy 1 Chap 12 Reactions will often give a mixture of products: OH H 2 SO 4 + Major Minor How would the chemist determine which product was formed? Both are cyclopentenes; they are isomers.
Infrared Spectroscopy 1 Chap 12 Reactions will often give a mixture of products: OH H 2 SO 4 + Major Minor How would the chemist determine which product was formed? Both are cyclopentenes; they are isomers.
Chapter10 Tro. 4. Based on the Lewis structure, the number of electron domains in the valence shell of the CO molecule is A) 1 B) 2 C) 3 D) 4 E) 5
 Chapter10 Tro 1. All of the geometries listed below are examples of the five basic geometries for molecules with more than 3 atoms except A) planar triangular B) octahedral C) tetrahedral D) trihedral
Chapter10 Tro 1. All of the geometries listed below are examples of the five basic geometries for molecules with more than 3 atoms except A) planar triangular B) octahedral C) tetrahedral D) trihedral
18 electron rule : How to count electrons
 18 electron rule : How to count electrons The rule states that thermodynamically stable transition metal organometallic compounds are formed when the sum of the metal d electrons and the electrons conventionally
18 electron rule : How to count electrons The rule states that thermodynamically stable transition metal organometallic compounds are formed when the sum of the metal d electrons and the electrons conventionally
States of Matter CHAPTER 10 REVIEW SECTION 1. Name Date Class. Answer the following questions in the space provided.
 CHAPTER 10 REVIEW States of Matter SECTION 1 SHORT ANSWER Answer the following questions in the space provided. 1. Identify whether the descriptions below describe an ideal gas or a real gas. ideal gas
CHAPTER 10 REVIEW States of Matter SECTION 1 SHORT ANSWER Answer the following questions in the space provided. 1. Identify whether the descriptions below describe an ideal gas or a real gas. ideal gas
Studying an Organic Reaction. How do we know if a reaction can occur? And if a reaction can occur what do we know about the reaction?
 Studying an Organic Reaction How do we know if a reaction can occur? And if a reaction can occur what do we know about the reaction? Information we want to know: How much heat is generated? How fast is
Studying an Organic Reaction How do we know if a reaction can occur? And if a reaction can occur what do we know about the reaction? Information we want to know: How much heat is generated? How fast is
What is a weak hydrogen bond?
 What is a weak hydrogen bond? Gautam R. Desiraju School of Chemistry University of yderabad yderabad 500 046, India gautam_desiraju@yahoo.com http://202.41.85.161/~grd/ What is a hydrogen bond? Under certain
What is a weak hydrogen bond? Gautam R. Desiraju School of Chemistry University of yderabad yderabad 500 046, India gautam_desiraju@yahoo.com http://202.41.85.161/~grd/ What is a hydrogen bond? Under certain
Raman Spectroscopy. 1. Introduction. 2. More on Raman Scattering. " scattered. " incident
 February 15, 2006 Advanced Physics Laboratory Raman Spectroscopy 1. Introduction When light is scattered from a molecule or crystal, most photons are elastically scattered. The scattered photons have the
February 15, 2006 Advanced Physics Laboratory Raman Spectroscopy 1. Introduction When light is scattered from a molecule or crystal, most photons are elastically scattered. The scattered photons have the
electron does not become part of the compound; one electron goes in but two electrons come out.
 Characterization Techniques for Organic Compounds. When we run a reaction in the laboratory or when we isolate a compound from nature, one of our first tasks is to identify the compound that we have obtained.
Characterization Techniques for Organic Compounds. When we run a reaction in the laboratory or when we isolate a compound from nature, one of our first tasks is to identify the compound that we have obtained.
CHAPTER 13 MOLECULAR SPECTROSCOPY
 CHAPTER 13 MOLECULAR SPECTROSCOPY Our most detailed knowledge of atomic and molecular structure has been obtained from spectroscopy study of the emission, absorption and scattering of electromagnetic radiation
CHAPTER 13 MOLECULAR SPECTROSCOPY Our most detailed knowledge of atomic and molecular structure has been obtained from spectroscopy study of the emission, absorption and scattering of electromagnetic radiation
Decision Support for Virtual Machine Re-Provisioning in Production Environments
 Decision Support for Virtual Machine Re-Provisioning in Production Environments Kyrre Begnum and Matthew Disney Oslo University College, Norway Æleen Frisch Exponential Consulting Ingard Mevåg Oslo University
Decision Support for Virtual Machine Re-Provisioning in Production Environments Kyrre Begnum and Matthew Disney Oslo University College, Norway Æleen Frisch Exponential Consulting Ingard Mevåg Oslo University
Molecular Geometry and VSEPR We gratefully acknowledge Portland Community College for the use of this experiment.
 Molecular and VSEPR We gratefully acknowledge Portland ommunity ollege for the use of this experiment. Objectives To construct molecular models for covalently bonded atoms in molecules and polyatomic ions
Molecular and VSEPR We gratefully acknowledge Portland ommunity ollege for the use of this experiment. Objectives To construct molecular models for covalently bonded atoms in molecules and polyatomic ions
Amino Acids. Amino acids are the building blocks of proteins. All AA s have the same basic structure: Side Chain. Alpha Carbon. Carboxyl. Group.
 Protein Structure Amino Acids Amino acids are the building blocks of proteins. All AA s have the same basic structure: Side Chain Alpha Carbon Amino Group Carboxyl Group Amino Acid Properties There are
Protein Structure Amino Acids Amino acids are the building blocks of proteins. All AA s have the same basic structure: Side Chain Alpha Carbon Amino Group Carboxyl Group Amino Acid Properties There are
UV-Vis Vis spectroscopy. Electronic absorption spectroscopy
 UV-Vis Vis spectroscopy Electronic absorption spectroscopy Absortpion spectroscopy Provide information about presence and absence of unsaturated functional groups Useful adjunct to IR Determination of
UV-Vis Vis spectroscopy Electronic absorption spectroscopy Absortpion spectroscopy Provide information about presence and absence of unsaturated functional groups Useful adjunct to IR Determination of
Chapter 1 Structure and Bonding. Modified by Dr. Daniela Radu
 John E. McMurry www.cengage.com/chemistry/mcmurry Chapter 1 Structure and Bonding Modified by Dr. Daniela Radu What is Organic Chemistry? Living things are made of organic chemicals Proteins that make
John E. McMurry www.cengage.com/chemistry/mcmurry Chapter 1 Structure and Bonding Modified by Dr. Daniela Radu What is Organic Chemistry? Living things are made of organic chemicals Proteins that make
AP Chemistry A. Allan Chapter 8 Notes - Bonding: General Concepts
 AP Chemistry A. Allan Chapter 8 Notes - Bonding: General Concepts 8.1 Types of Chemical Bonds A. Ionic Bonding 1. Electrons are transferred 2. Metals react with nonmetals 3. Ions paired have lower energy
AP Chemistry A. Allan Chapter 8 Notes - Bonding: General Concepts 8.1 Types of Chemical Bonds A. Ionic Bonding 1. Electrons are transferred 2. Metals react with nonmetals 3. Ions paired have lower energy
Chemistry 1050 Chapter 13 LIQUIDS AND SOLIDS 1. Exercises: 25, 27, 33, 39, 41, 43, 51, 53, 57, 61, 63, 67, 69, 71(a), 73, 75, 79
 Chemistry 1050 Chapter 13 LIQUIDS AND SOLIDS 1 Text: Petrucci, Harwood, Herring 8 th Edition Suggest text problems Review questions: 1, 5!11, 13!17, 19!23 Exercises: 25, 27, 33, 39, 41, 43, 51, 53, 57,
Chemistry 1050 Chapter 13 LIQUIDS AND SOLIDS 1 Text: Petrucci, Harwood, Herring 8 th Edition Suggest text problems Review questions: 1, 5!11, 13!17, 19!23 Exercises: 25, 27, 33, 39, 41, 43, 51, 53, 57,
Molecular Cell Biology
 Harvey Lodish Arnold Berk Paul Matsudaira Chris A. Kaiser Monty Krieger Matthew P. Scott Lawrence Zipursky James Darnell Molecular Cell Biology Fifth Edition Chapter 2: Chemical Foundations Copyright 2004
Harvey Lodish Arnold Berk Paul Matsudaira Chris A. Kaiser Monty Krieger Matthew P. Scott Lawrence Zipursky James Darnell Molecular Cell Biology Fifth Edition Chapter 2: Chemical Foundations Copyright 2004
C o m p u te r M o d e lin g o f M o le c u la r E le c tro n ic S tru c tu re
 C o m p u te r M o d e lin g o f M o le c u la r E le c tro n ic S tru c tu re P e te r P u la y D e p a rtm e n t o f C h e m is try a n d B io c h e m is try, U n iv e rs ity o f A rk a n s a s, F a
C o m p u te r M o d e lin g o f M o le c u la r E le c tro n ic S tru c tu re P e te r P u la y D e p a rtm e n t o f C h e m is try a n d B io c h e m is try, U n iv e rs ity o f A rk a n s a s, F a
CHEMISTRY STANDARDS BASED RUBRIC ATOMIC STRUCTURE AND BONDING
 CHEMISTRY STANDARDS BASED RUBRIC ATOMIC STRUCTURE AND BONDING Essential Standard: STUDENTS WILL UNDERSTAND THAT THE PROPERTIES OF MATTER AND THEIR INTERACTIONS ARE A CONSEQUENCE OF THE STRUCTURE OF MATTER,
CHEMISTRY STANDARDS BASED RUBRIC ATOMIC STRUCTURE AND BONDING Essential Standard: STUDENTS WILL UNDERSTAND THAT THE PROPERTIES OF MATTER AND THEIR INTERACTIONS ARE A CONSEQUENCE OF THE STRUCTURE OF MATTER,
NMR and other Instrumental Techniques in Chemistry and the proposed National Curriculum.
 NMR and other Instrumental Techniques in Chemistry and the proposed National Curriculum. Dr. John Jackowski Chair of Science, Head of Chemistry Scotch College Melbourne john.jackowski@scotch.vic.edu.au
NMR and other Instrumental Techniques in Chemistry and the proposed National Curriculum. Dr. John Jackowski Chair of Science, Head of Chemistry Scotch College Melbourne john.jackowski@scotch.vic.edu.au
Nucleotides and Nucleic Acids
 Nucleotides and Nucleic Acids Brief History 1 1869 - Miescher Isolated nuclein from soiled bandages 1902 - Garrod Studied rare genetic disorder: Alkaptonuria; concluded that specific gene is associated
Nucleotides and Nucleic Acids Brief History 1 1869 - Miescher Isolated nuclein from soiled bandages 1902 - Garrod Studied rare genetic disorder: Alkaptonuria; concluded that specific gene is associated
HOMEWORK PROBLEMS: IR SPECTROSCOPY AND 13C NMR. The peak at 1720 indicates a C=O bond (carbonyl). One possibility is acetone:
 HMEWRK PRBLEMS: IR SPECTRSCPY AND 13C NMR 1. You find a bottle on the shelf only labeled C 3 H 6. You take an IR spectrum of the compound and find major peaks at 2950, 1720, and 1400 cm -1. Draw a molecule
HMEWRK PRBLEMS: IR SPECTRSCPY AND 13C NMR 1. You find a bottle on the shelf only labeled C 3 H 6. You take an IR spectrum of the compound and find major peaks at 2950, 1720, and 1400 cm -1. Draw a molecule
Acids and Bases: Molecular Structure and Acidity
 Acids and Bases: Molecular Structure and Acidity Review the Acids and Bases Vocabulary List as needed. Tutorial Contents A. Introduction B. Resonance C. Atomic Radius D. Electronegativity E. Inductive
Acids and Bases: Molecular Structure and Acidity Review the Acids and Bases Vocabulary List as needed. Tutorial Contents A. Introduction B. Resonance C. Atomic Radius D. Electronegativity E. Inductive
7.14 Linear triatomic: A-----B-----C. Bond angles = 180 degrees. Trigonal planar: Bond angles = 120 degrees. B < B A B = 120
 APTER SEVEN Molecular Geometry 7.13 Molecular geometry may be defined as the three-dimensional arrangement of atoms in a molecule. The study of molecular geometry is important in that a molecule s geometry
APTER SEVEN Molecular Geometry 7.13 Molecular geometry may be defined as the three-dimensional arrangement of atoms in a molecule. The study of molecular geometry is important in that a molecule s geometry
Coherent low-frequency motions of hydrogen bonded acetic acid dimers in the liquid phase
 JOURNAL OF CHEMICAL PHYSICS VOLUME 121, NUMBER 2 8 JULY 2004 Coherent low-frequency motions of hydrogen bonded acetic acid dimers in the liquid phase Karsten Heyne, a) Nils Huse, Jens Dreyer, Erik T. J.
JOURNAL OF CHEMICAL PHYSICS VOLUME 121, NUMBER 2 8 JULY 2004 Coherent low-frequency motions of hydrogen bonded acetic acid dimers in the liquid phase Karsten Heyne, a) Nils Huse, Jens Dreyer, Erik T. J.
A disaccharide is formed when a dehydration reaction joins two monosaccharides. This covalent bond is called a glycosidic linkage.
 CH 5 Structure & Function of Large Molecules: Macromolecules Molecules of Life All living things are made up of four classes of large biological molecules: carbohydrates, lipids, proteins, and nucleic
CH 5 Structure & Function of Large Molecules: Macromolecules Molecules of Life All living things are made up of four classes of large biological molecules: carbohydrates, lipids, proteins, and nucleic
Name: Class: Date: 3) The bond angles marked a, b, and c in the molecule below are about,, and, respectively.
 Name: Class: Date: Unit 9 Practice Multiple Choice Identify the choice that best completes the statement or answers the question. 1) The basis of the VSEPR model of molecular bonding is. A) regions of
Name: Class: Date: Unit 9 Practice Multiple Choice Identify the choice that best completes the statement or answers the question. 1) The basis of the VSEPR model of molecular bonding is. A) regions of
