Introduction to Perl Programming Input/Output, Regular Expressions, String Manipulation. Beginning Perl, Chap 4 6. Example 1
|
|
|
- Sheena Jefferson
- 9 years ago
- Views:
Transcription
1 Introduction to Perl Programming Input/Output, Regular Expressions, String Manipulation Beginning Perl, Chap 4 6 Example 1 #!/usr/bin/perl -w use strict; # version 1: = ('A', 'C', 'G', 'T'); for ( ) { $dna.= $nt[int(rand 4)]; # version 4: with compositional bias my %comp = ( A => 0.20, C => 0.35, G => 0.25, T => 0.20 ); = keys %comp; # version 2: = qw(a C G T); for ( ) { $dna.= $nt[int(rand 4)]; # version 3: most Perlish = qw(a C G T); for ( ) { print $dna, \n ; # fill the bucket to draw from: for my $nt (@nt) { ($nt) x ($comp{$nt * 100); for (0.. 99) { $dna.= $bucket[int(rand 1
2 Example 2 #!/usr/bin/perl -w use strict; # version 2: very Perlish; = ( A.. Z ); # version 1; # same as nt construction = qw(a C D E F G H I K L M N P Q R S T V W Y); my $protein; for ( ) { $protein.= $aa[int(rand print $protein, \n ; my %comp; # hash = ( 1 / 20 ) # remove letter that aren't aa's $comp{'b' = $comp{'j' = $comp{'o' = $comp{'x' = $comp{'u' = $comp{'z' = 0; for my $l (@letters) { ($l) x ($comp{$l * 100); my $length = rand(100); my $protein; for (1.. $length) { push $protein, Example 3 #!/usr/bin/perl -w use strict; my %codontable = ( 'AAA' => 'K', 'AAC' => 'N', 'AAG' => 'K', 'AAT' => 'N', 'ACA' => 'T', 'ACC' => 'T', 'ACG' => 'T', 'ACT' => 'T', 'AGA' => 'R', 'AGC' => 'S', 'AGG' => 'R', 'AGT' => 'S', 'ATA' => 'I', 'ATC' => 'I', 'ATG' => 'M', 'ATT' => 'I', 'CAA' => 'Q', 'CAC' => 'H', 'CAG' => 'Q', 'CAT' => 'H', 'CCA' => 'P', 'CCC' => 'P', 'CCG' => 'P', 'CCT' => 'P', 'CGA' => 'R', 'CGC' => 'R', 'CGG' => 'R', 'CGT' => 'R', 'CTA' => 'L', 'CTC' => 'L', 'CTG' => 'L', 'CTT' => 'L', 'GAA' => 'E', 'GAC' => 'D', 'GAG' => 'E', 'GAT' => 'D', 'GCA' => 'A', 'GCC' => 'A', 'GCG' => 'A', 'GCT' => 'A', 'GGA' => 'G', 'GGC' => 'G', 'GGG' => 'G', 'GGT' => 'G', 'GTA' => 'V', 'GTC' => 'V', 'GTG' => 'V', 'GTT' => 'V', 'TAA' => '*', 'TAC' => 'Y', 'TAG' => '*', 'TAT' => 'Y', 'TCA' => 'S', 'TCC' => 'S', 'TCG' => 'S', 'TCT' => 'S', 'TGA' => '*', 'TGC' => 'C', 'TGG' => 'W', 'TGT' => 'C', 'TTA' => 'L', 'TTC' => 'F', 'TTG' => 'L', 'TTT' => 'F' ); # make DNA the boring way: = qw(a C G T); my $length = int(rand(500)); for (1.. $length) { $dna.= # OK, now translate: my $protein; = split('', $dna); my $codon = ''; for my $i (0.. $#seq) { $codon.= $seq[$i]; if (($i + 1) % 3 == 0) { $protein.= $codontable{$codon; $codon = ''; # another way, using "length()": for my $nt (@seq) { $codon.= $nt; if (length($codon) == 3) { $protein.= $codontable{$codon; $codon = ''; # yet another variation on the theme: while (@seq) { $codon.= if (length $codon == 3) { $protein.= $codontable{$codon; $codon = ''; # even better: while (@seq) { $protein.= $codontable{ 2
3 Input/Output open (INFILE, < gtm1_human.aa ); open file for reading $file = $ARGV[0]; open (INFILE, < $file ) die ( Cannot open $file ); open (OUTFILE, > $file); open file for writing $line = <FILE>; read a line chomp($line); remove \n while ($line=<file>) { print FILE $line\n ; close(file); Regular expressions >gi sp P20432 GTT1_DROME Glutathione S-transferase 1-1 used for string matching, substitution, pattern extraction /^>gi\ / matches >gi sp P20432 GTT1_DROME if ($line =~ m/^>gi/) { #match $line =~ /^>gi\ (\d+)\ /; # extract gi# $gi = $1; ($gi) = $line =~ /^>gi\ (\d+)\ /; #same $line =~ s/^>(.*)$/>>$1/; # substitution 3
4 Regular expressions (cont.) >gi sp P20432 GTT1_DROME Glutathione S-transferase 1-1 m/plaintext/ m/one two/; # alternation m/(one two) (three)/; # grouping with # parenthesis /^>gi\ (\d+)/ #beginning of line /.+ (\d+) aa$/ # end of line /a*bc/ # bc,abc,aabc, # repetitions /a?bc/ # abc, bc /a+bc/ # abc, aabc, Regular Expressions, III >gi sp P20432 GTT1_DROME Glutathione S-transferase 1-1 Matching classes: /^>gi\ [0-9]+\ [a-z]+\ [A-Z][0-9a-z]+\ / [a-z] [0-9] -> class [^a-z] -> negated class /^>gi\ \d+\ [a-z]\ \w+\ / \d -> number [0-9] \D -> not a number \w -> word [0-9A-Za-z_] \W -> not a word char \s -> space [ \t\n\r] \S -> not a space Capturing matches: /^>gi\ (\d+)\ ([a-z])\ (\w+)\ / $1 $2 $3 ($gi,$db,$db_acc) = $line =~ /^>gi\ (\d+)\ ([a-z])\ (\w+)\ /; 4
5 Regular expressions - modifiers m/that/i # ignore case s/this/that/g # global replacement m/>gi\ (\d+)\ ([a-z]{2,3 (\w+) #{range s///m # treat string as multiple lines s///s # span over \n s/\n//gs # remove \n in multiline entry s/gaattc/$1\n/g # break lines at EcoRI site { local $/ = "\n>"; # change the line separator to "\n" while ($entry = <INFILE>) { chomp($entry); $entry =~ s/\a>?/>/; # replace missing >, if necessary # \A is like ^ (beginning of string) open(out, ">file$num.fa") or die $!; $num++; print OUT "$entry\n"; close(out); String expressions (with regular expressions) if ( /^>gi\ / ) { if ( $line =~ m/^>gi\ /) { while ( $line!~ m/^>gi\ /) { Substitution: $new_line =~ s/\ /:/g; Pattern extraction: ($gi,$db,$db_acc) = $line = /^>gi\ (\d+)\ ([a-z])\ (\w+)\ /; split (/ /,$line) join ( :,@fields); substr($string,$start,$length); # rarely used index($string,$query); # rarely used Comparison: eq ne cmp lt gt le ge 5
6 use vars qw(%proteins %dnas $line $name $codon $aa); my ($pfile, $nfile) open(protein, "<$pfile") or die $!; open(dna, "<$nfile") or die $!; while(my $line = <PROTEIN>) { chomp($line); if($line =~ m/^>/) { $name = shift split(' ', $line); next; $proteins{$name.= $line if $name; close(protein); for my $name (keys %proteins) { $proteins{$name =~ s/\s+//sg; while($line = <DNA>) { chomp($line); if($line =~ m/^>/) { $name =shift split(' ', $line); next; $dnas{$name.= $line if $name; close(dna); Perl mrtrans.pl for $name (keys %dna) { unless (exists $protein{$name) { warn "DNA sequence $name has no matching protein sequence!\n"; next; $dna{$name =~ s/\s+//sg; print "$name\n"; $proteins{$name); = split('', $dnas{$name); for $aa (@protein) { if($aa eq '-') { print "---"; else { $codon = # check that $codon codes for $aa print $codon; print "\n"; Exercises 1. Take a set of FASTA format files and make a FASTA library, combining those files 2. Take a FASTA library, and break it into individual FASTA files 3. Write an extraction program to report (a) the most significant library hit (b) all significant library hits that result from a FASTA search. Several example search results are available at watson.achs:/r0/seqlib/bioch508/perl/ fasta.result* 4. Write the same program for blast output. Sample results are in the same directory 5. Write a perl equivalent to mrtrans that does not care about the order of sequence input. Given two files, a FASTA protein library (with ʻ-ʼ for gaps) and a FASTA DNA library with the same sequence names, generate a FASTA DNA alignment with ʻ---ʼ inserted into the DNA sequence for each ʻ-ʼ in the protein sequence 6
(http://genomes.urv.es/caical) TUTORIAL. (July 2006)
 (http://genomes.urv.es/caical) TUTORIAL (July 2006) CAIcal manual 2 Table of contents Introduction... 3 Required inputs... 5 SECTION A Calculation of parameters... 8 SECTION B CAI calculation for FASTA
(http://genomes.urv.es/caical) TUTORIAL (July 2006) CAIcal manual 2 Table of contents Introduction... 3 Required inputs... 5 SECTION A Calculation of parameters... 8 SECTION B CAI calculation for FASTA
GENEWIZ, Inc. DNA Sequencing Service Details for USC Norris Comprehensive Cancer Center DNA Core
 DNA Sequencing Services Pre-Mixed o Provide template and primer, mixed into the same tube* Pre-Defined o Provide template and primer in separate tubes* Custom o Full-service for samples with unknown concentration
DNA Sequencing Services Pre-Mixed o Provide template and primer, mixed into the same tube* Pre-Defined o Provide template and primer in separate tubes* Custom o Full-service for samples with unknown concentration
(A) Microarray analysis was performed on ATM and MDM isolated from 4 obese donors.
 Legends of supplemental figures and tables Figure 1: Overview of study design and results. (A) Microarray analysis was performed on ATM and MDM isolated from 4 obese donors. After raw data gene expression
Legends of supplemental figures and tables Figure 1: Overview of study design and results. (A) Microarray analysis was performed on ATM and MDM isolated from 4 obese donors. After raw data gene expression
10 µg lyophilized plasmid DNA (store lyophilized plasmid at 20 C)
 TECHNICAL DATA SHEET BIOLUMINESCENCE RESONANCE ENERGY TRANSFER RENILLA LUCIFERASE FUSION PROTEIN EXPRESSION VECTOR Product: prluc-c Vectors Catalog number: Description: Amount: The prluc-c vectors contain
TECHNICAL DATA SHEET BIOLUMINESCENCE RESONANCE ENERGY TRANSFER RENILLA LUCIFERASE FUSION PROTEIN EXPRESSION VECTOR Product: prluc-c Vectors Catalog number: Description: Amount: The prluc-c vectors contain
Table S1. Related to Figure 4
 Table S1. Related to Figure 4 Final Diagnosis Age PMD Control Control 61 15 Control 67 6 Control 68 10 Control 49 15 AR-PD PD 62 15 PD 65 4 PD 52 18 PD 68 10 AR-PD cingulate cortex used for immunoblot
Table S1. Related to Figure 4 Final Diagnosis Age PMD Control Control 61 15 Control 67 6 Control 68 10 Control 49 15 AR-PD PD 62 15 PD 65 4 PD 52 18 PD 68 10 AR-PD cingulate cortex used for immunoblot
Mutations and Genetic Variability. 1. What is occurring in the diagram below?
 Mutations and Genetic Variability 1. What is occurring in the diagram below? A. Sister chromatids are separating. B. Alleles are independently assorting. C. Genes are replicating. D. Segments of DNA are
Mutations and Genetic Variability 1. What is occurring in the diagram below? A. Sister chromatids are separating. B. Alleles are independently assorting. C. Genes are replicating. D. Segments of DNA are
DNA Sample preparation and Submission Guidelines
 DNA Sample preparation and Submission Guidelines Requirements: Please submit samples in 1.5ml microcentrifuge tubes. Fill all the required information in the Eurofins DNA sequencing order form and send
DNA Sample preparation and Submission Guidelines Requirements: Please submit samples in 1.5ml microcentrifuge tubes. Fill all the required information in the Eurofins DNA sequencing order form and send
UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet
 1 UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam:.
1 UNIVERSITETET I OSLO Det matematisk-naturvitenskapelige fakultet Exam in: MBV4010 Arbeidsmetoder i molekylærbiologi og biokjemi I MBV4010 Methods in molecular biology and biochemistry I Day of exam:.
The p53 MUTATION HANDBOOK
 The p MUTATION HANDBOOK Version 1. /7 Thierry Soussi Christophe Béroud, Dalil Hamroun Jean Michel Rubio Nevado http://p/free.fr The p Mutation HandBook By T Soussi, J.M. Rubio-Nevado, D. Hamroun and C.
The p MUTATION HANDBOOK Version 1. /7 Thierry Soussi Christophe Béroud, Dalil Hamroun Jean Michel Rubio Nevado http://p/free.fr The p Mutation HandBook By T Soussi, J.M. Rubio-Nevado, D. Hamroun and C.
Inverse PCR & Cycle Sequencing of P Element Insertions for STS Generation
 BDGP Resources Inverse PCR & Cycle Sequencing of P Element Insertions for STS Generation For recovery of sequences flanking PZ, PlacW and PEP elements E. Jay Rehm Berkeley Drosophila Genome Project I.
BDGP Resources Inverse PCR & Cycle Sequencing of P Element Insertions for STS Generation For recovery of sequences flanking PZ, PlacW and PEP elements E. Jay Rehm Berkeley Drosophila Genome Project I.
Supplementary Information. Binding region and interaction properties of sulfoquinovosylacylglycerol (SQAG) with human
 Supplementary Information Binding region and interaction properties of sulfoquinovosylacylglycerol (SQAG) with human vascular endothelial growth factor 165 revealed by biosensor based assays Yoichi Takakusagi
Supplementary Information Binding region and interaction properties of sulfoquinovosylacylglycerol (SQAG) with human vascular endothelial growth factor 165 revealed by biosensor based assays Yoichi Takakusagi
Next Generation Sequencing
 Next Generation Sequencing 38. Informationsgespräch der Blutspendezentralefür Wien, Niederösterreich und Burgenland Österreichisches Rotes Kreuz 22. November 2014, Parkhotel Schönbrunn Die Zukunft hat
Next Generation Sequencing 38. Informationsgespräch der Blutspendezentralefür Wien, Niederösterreich und Burgenland Österreichisches Rotes Kreuz 22. November 2014, Parkhotel Schönbrunn Die Zukunft hat
Hands on Simulation of Mutation
 Hands on Simulation of Mutation Charlotte K. Omoto P.O. Box 644236 Washington State University Pullman, WA 99164-4236 [email protected] ABSTRACT This exercise is a hands-on simulation of mutations and their
Hands on Simulation of Mutation Charlotte K. Omoto P.O. Box 644236 Washington State University Pullman, WA 99164-4236 [email protected] ABSTRACT This exercise is a hands-on simulation of mutations and their
Gene Synthesis 191. Mutagenesis 194. Gene Cloning 196. AccuGeneBlock Service 198. Gene Synthesis FAQs 201. User Protocol 204
 Gene Synthesis 191 Mutagenesis 194 Gene Cloning 196 AccuGeneBlock Service 198 Gene Synthesis FAQs 201 User Protocol 204 Gene Synthesis Overview Gene synthesis is the most cost-effective way to enhance
Gene Synthesis 191 Mutagenesis 194 Gene Cloning 196 AccuGeneBlock Service 198 Gene Synthesis FAQs 201 User Protocol 204 Gene Synthesis Overview Gene synthesis is the most cost-effective way to enhance
pcas-guide System Validation in Genome Editing
 pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
Inverse PCR and Sequencing of P-element, piggybac and Minos Insertion Sites in the Drosophila Gene Disruption Project
 Inverse PCR and Sequencing of P-element, piggybac and Minos Insertion Sites in the Drosophila Gene Disruption Project Protocol for recovery of sequences flanking insertions in the Drosophila Gene Disruption
Inverse PCR and Sequencing of P-element, piggybac and Minos Insertion Sites in the Drosophila Gene Disruption Project Protocol for recovery of sequences flanking insertions in the Drosophila Gene Disruption
ANALYSIS OF A CIRCULAR CODE MODEL
 ANALYSIS OF A CIRCULAR CODE MODEL Jérôme Lacan and Chrstan J. Mchel * Laboratore d Informatque de Franche-Comté UNIVERSITE DE FRANCHE-COMTE IUT de Belfort-Montbélard 4 Place Tharradn - BP 747 5 Montbélard
ANALYSIS OF A CIRCULAR CODE MODEL Jérôme Lacan and Chrstan J. Mchel * Laboratore d Informatque de Franche-Comté UNIVERSITE DE FRANCHE-COMTE IUT de Belfort-Montbélard 4 Place Tharradn - BP 747 5 Montbélard
Gene Finding CMSC 423
 Gene Finding CMSC 423 Finding Signals in DNA We just have a long string of A, C, G, Ts. How can we find the signals encoded in it? Suppose you encountered a language you didn t know. How would you decipher
Gene Finding CMSC 423 Finding Signals in DNA We just have a long string of A, C, G, Ts. How can we find the signals encoded in it? Suppose you encountered a language you didn t know. How would you decipher
SERVICES CATALOGUE WITH SUBMISSION GUIDELINES
 SERVICES CATALOGUE WITH SUBMISSION GUIDELINES 3921 Montgomery Road Cincinnati, Ohio 45212 513-841-2428 www.agctsequencing.com CONTENTS Welcome Dye Terminator Sequencing DNA Sequencing Services - Full Service
SERVICES CATALOGUE WITH SUBMISSION GUIDELINES 3921 Montgomery Road Cincinnati, Ohio 45212 513-841-2428 www.agctsequencing.com CONTENTS Welcome Dye Terminator Sequencing DNA Sequencing Services - Full Service
Molecular analyses of EGFR: mutation and amplification detection
 Molecular analyses of EGFR: mutation and amplification detection Petra Nederlof, Moleculaire Pathologie NKI Amsterdam Henrique Ruijter, Ivon Tielen, Lucie Boerrigter, Aafke Ariaens Outline presentation
Molecular analyses of EGFR: mutation and amplification detection Petra Nederlof, Moleculaire Pathologie NKI Amsterdam Henrique Ruijter, Ivon Tielen, Lucie Boerrigter, Aafke Ariaens Outline presentation
pcmv6-neo Vector Application Guide Contents
 pcmv6-neo Vector Application Guide Contents Package Contents and Storage Conditions... 2 Product Description... 2 Introduction... 2 Production and Quality Assurance... 2 Methods... 3 Other required reagents...
pcmv6-neo Vector Application Guide Contents Package Contents and Storage Conditions... 2 Product Description... 2 Introduction... 2 Production and Quality Assurance... 2 Methods... 3 Other required reagents...
Marine Biology DEC 2004; 146(1) : 53-64 http://dx.doi.org/10.1007/s00227-004-1423-6 Copyright 2004 Springer
 Marine Biology DEC 2004; 146(1) : 53-64 http://dx.doi.org/10.1007/s00227-004-1423-6 Copyright 2004 Springer Archimer http://www.ifremer.fr/docelec/ Archive Institutionnelle de l Ifremer The original publication
Marine Biology DEC 2004; 146(1) : 53-64 http://dx.doi.org/10.1007/s00227-004-1423-6 Copyright 2004 Springer Archimer http://www.ifremer.fr/docelec/ Archive Institutionnelle de l Ifremer The original publication
Y-chromosome haplotype distribution in Han Chinese populations and modern human origin in East Asians
 Vol. 44 No. 3 SCIENCE IN CHINA (Series C) June 2001 Y-chromosome haplotype distribution in Han Chinese populations and modern human origin in East Asians KE Yuehai ( `º) 1, SU Bing (3 Á) 1 3, XIAO Junhua
Vol. 44 No. 3 SCIENCE IN CHINA (Series C) June 2001 Y-chromosome haplotype distribution in Han Chinese populations and modern human origin in East Asians KE Yuehai ( `º) 1, SU Bing (3 Á) 1 3, XIAO Junhua
Biopython Tutorial and Cookbook
 Biopython Tutorial and Cookbook Jeff Chang, Brad Chapman, Iddo Friedberg, Thomas Hamelryck, Michiel de Hoon, Peter Cock Last Update September 2008 Contents 1 Introduction 5 1.1 What is Biopython?.........................................
Biopython Tutorial and Cookbook Jeff Chang, Brad Chapman, Iddo Friedberg, Thomas Hamelryck, Michiel de Hoon, Peter Cock Last Update September 2008 Contents 1 Introduction 5 1.1 What is Biopython?.........................................
Part ONE. a. Assuming each of the four bases occurs with equal probability, how many bits of information does a nucleotide contain?
 Networked Systems, COMPGZ01, 2012 Answer TWO questions from Part ONE on the answer booklet containing lined writing paper, and answer ALL questions in Part TWO on the multiple-choice question answer sheet.
Networked Systems, COMPGZ01, 2012 Answer TWO questions from Part ONE on the answer booklet containing lined writing paper, and answer ALL questions in Part TWO on the multiple-choice question answer sheet.
Chapter 9. Applications of probability. 9.1 The genetic code
 Chapter 9 Applications of probability In this chapter we use the tools of elementary probability to investigate problems of several kinds. First, we study the language of life by focusing on the universal
Chapter 9 Applications of probability In this chapter we use the tools of elementary probability to investigate problems of several kinds. First, we study the language of life by focusing on the universal
Module 6: Digital DNA
 Module 6: Digital DNA Representation and processing of digital information in the form of DNA is essential to life in all organisms, no matter how large or tiny. Computing tools and computational thinking
Module 6: Digital DNA Representation and processing of digital information in the form of DNA is essential to life in all organisms, no matter how large or tiny. Computing tools and computational thinking
The making of The Genoma Music
 242 Summary Key words Resumen Palabras clave The making of The Genoma Music Aurora Sánchez Sousa 1, Fernando Baquero 1 and Cesar Nombela 2 1 Department of Microbiology, Ramón y Cajal Hospital, and 2 Department
242 Summary Key words Resumen Palabras clave The making of The Genoma Music Aurora Sánchez Sousa 1, Fernando Baquero 1 and Cesar Nombela 2 1 Department of Microbiology, Ramón y Cajal Hospital, and 2 Department
The DNA-"Wave Biocomputer"
 The DNA-"Wave Biocomputer" Peter P. Gariaev (Pjotr Garjajev)*, Boris I. Birshtein*, Alexander M. Iarochenko*, Peter J. Marcer**, George G. Tertishny*, Katherine A. Leonova*, Uwe Kaempf ***. * Institute
The DNA-"Wave Biocomputer" Peter P. Gariaev (Pjotr Garjajev)*, Boris I. Birshtein*, Alexander M. Iarochenko*, Peter J. Marcer**, George G. Tertishny*, Katherine A. Leonova*, Uwe Kaempf ***. * Institute
Heraeus Sepatech, Kendro Laboratory Products GmbH, Berlin. Becton Dickinson,Heidelberg. Biozym, Hessisch Oldendorf. Eppendorf, Hamburg
 13 4. MATERIALS 4.1 Laboratory apparatus Biofuge A Centrifuge 5804R FACScan Gel electrophoresis chamber GPR Centrifuge Heraeus CO-AUTO-ZERO Light Cycler Microscope Motopipet Neubauer Cell Chamber PCR cycler
13 4. MATERIALS 4.1 Laboratory apparatus Biofuge A Centrifuge 5804R FACScan Gel electrophoresis chamber GPR Centrifuge Heraeus CO-AUTO-ZERO Light Cycler Microscope Motopipet Neubauer Cell Chamber PCR cycler
http://www.life.umd.edu/grad/mlfsc/ DNA Bracelets
 http://www.life.umd.edu/grad/mlfsc/ DNA Bracelets by Louise Brown Jasko John Anthony Campbell Jack Dennis Cassidy Michael Nickelsburg Stephen Prentis Rohm Objectives: 1) Using plastic beads, construct
http://www.life.umd.edu/grad/mlfsc/ DNA Bracelets by Louise Brown Jasko John Anthony Campbell Jack Dennis Cassidy Michael Nickelsburg Stephen Prentis Rohm Objectives: 1) Using plastic beads, construct
Transmembrane Signaling in Chimeras of the E. coli Chemotaxis Receptors and Bacterial Class III Adenylyl Cyclases
 Transmembrane Signaling in Chimeras of the E. coli Chemotaxis Receptors and Bacterial Class III Adenylyl Cyclases Dissertation der Mathematisch-Naturwissenschaftlichen Fakultät der Eberhard Karls Universität
Transmembrane Signaling in Chimeras of the E. coli Chemotaxis Receptors and Bacterial Class III Adenylyl Cyclases Dissertation der Mathematisch-Naturwissenschaftlichen Fakultät der Eberhard Karls Universität
Drosophila NK-homeobox genes
 Proc. Natl. Acad. Sci. USA Vol. 86, pp. 7716-7720, October 1989 Biochemistry Drosophila NK-homeobox genes (NK-1, NK-2,, and DNA clones/chromosome locations of genes) YONGSOK KIM AND MARSHALL NIRENBERG
Proc. Natl. Acad. Sci. USA Vol. 86, pp. 7716-7720, October 1989 Biochemistry Drosophila NK-homeobox genes (NK-1, NK-2,, and DNA clones/chromosome locations of genes) YONGSOK KIM AND MARSHALL NIRENBERG
NimbleGen SeqCap EZ Library SR User s Guide Version 3.0
 NimbleGen SeqCap EZ Library SR User s Guide Version 3.0 For life science research only. Not for use in diagnostic procedures. Copyright 2011 Roche NimbleGen, Inc. All Rights Reserved. Editions Version
NimbleGen SeqCap EZ Library SR User s Guide Version 3.0 For life science research only. Not for use in diagnostic procedures. Copyright 2011 Roche NimbleGen, Inc. All Rights Reserved. Editions Version
Characterization of cdna clones of the family of trypsin/a-amylase inhibitors (CM-proteins) in barley {Hordeum vulgare L.)
 Characterization of cdna clones of the family of trypsin/a-amylase inhibitors (CM-proteins) in barley {Hordeum vulgare L.) J. Paz-Ares, F. Ponz, P. Rodríguez-Palenzuela, A. Lázaro, C. Hernández-Lucas,
Characterization of cdna clones of the family of trypsin/a-amylase inhibitors (CM-proteins) in barley {Hordeum vulgare L.) J. Paz-Ares, F. Ponz, P. Rodríguez-Palenzuela, A. Lázaro, C. Hernández-Lucas,
Five-minute cloning of Taq polymerase-amplified PCR products
 TOPO TA Cloning Version R 8 April 2004 25-0184 TOPO TA Cloning Five-minute cloning of Taq polymerase-amplified PCR products Catalog nos. K4500-01, K4500-40, K4510-20, K4520-01, K4520-40, K4550-01, K4550-40,
TOPO TA Cloning Version R 8 April 2004 25-0184 TOPO TA Cloning Five-minute cloning of Taq polymerase-amplified PCR products Catalog nos. K4500-01, K4500-40, K4510-20, K4520-01, K4520-40, K4550-01, K4550-40,
Mutation. Mutation provides raw material to evolution. Different kinds of mutations have different effects
 Mutation Mutation provides raw material to evolution Different kinds of mutations have different effects Mutational Processes Point mutation single nucleotide changes coding changes (missense mutations)
Mutation Mutation provides raw material to evolution Different kinds of mutations have different effects Mutational Processes Point mutation single nucleotide changes coding changes (missense mutations)
BD BaculoGold Baculovirus Expression System Innovative Solutions for Proteomics
 BD BaculoGold Baculovirus Expression System Innovative Solutions for Proteomics Table of Contents Innovative Solutions for Proteomics...........................................................................
BD BaculoGold Baculovirus Expression System Innovative Solutions for Proteomics Table of Contents Innovative Solutions for Proteomics...........................................................................
All commonly-used expression vectors used in the Jia Lab contain the following multiple cloning site: BamHI EcoRI SmaI SalI XhoI_ NotI
 2. Primer Design 2.1 Multiple Cloning Sites All commonly-used expression vectors used in the Jia Lab contain the following multiple cloning site: BamHI EcoRI SmaI SalI XhoI NotI XXX XXX GGA TCC CCG AAT
2. Primer Design 2.1 Multiple Cloning Sites All commonly-used expression vectors used in the Jia Lab contain the following multiple cloning site: BamHI EcoRI SmaI SalI XhoI NotI XXX XXX GGA TCC CCG AAT
Coding sequence the sequence of nucleotide bases on the DNA that are transcribed into RNA which are in turn translated into protein
 Assignment 3 Michele Owens Vocabulary Gene: A sequence of DNA that instructs a cell to produce a particular protein Promoter a control sequence near the start of a gene Coding sequence the sequence of
Assignment 3 Michele Owens Vocabulary Gene: A sequence of DNA that instructs a cell to produce a particular protein Promoter a control sequence near the start of a gene Coding sequence the sequence of
Insulin Receptor Gene Mutations in Iranian Patients with Type II Diabetes Mellitus
 Iranian Biomedical Journal 13 (3): 161-168 (July 2009) Insulin Receptor Gene Mutations in Iranian Patients with Type II Diabetes Mellitus Bahram Kazemi 1*, Negar Seyed 1, Elham Moslemi 2, Mojgan Bandehpour
Iranian Biomedical Journal 13 (3): 161-168 (July 2009) Insulin Receptor Gene Mutations in Iranian Patients with Type II Diabetes Mellitus Bahram Kazemi 1*, Negar Seyed 1, Elham Moslemi 2, Mojgan Bandehpour
Provincial Exam Questions. 9. Give one role of each of the following nucleic acids in the production of an enzyme.
 Provincial Exam Questions Unit: Cell Biology: Protein Synthesis (B7 & B8) 2010 Jan 3. Describe the process of translation. (4 marks) 2009 Sample 8. What is the role of ribosomes in protein synthesis? A.
Provincial Exam Questions Unit: Cell Biology: Protein Synthesis (B7 & B8) 2010 Jan 3. Describe the process of translation. (4 marks) 2009 Sample 8. What is the role of ribosomes in protein synthesis? A.
Six Homeoproteins and a Iinc-RNA at the Fast MYH Locus Lock Fast Myofiber Terminal Phenotype
 Six Homeoproteins and a Iinc-RNA at the Fast MYH Locus Lock Fast Myofiber Terminal Phenotype Iori Sakakibara 1,2,3, Marc Santolini 4, Arnaud Ferry 2,5, Vincent Hakim 4, Pascal Maire 1,2,3 * 1 INSERM U1016,
Six Homeoproteins and a Iinc-RNA at the Fast MYH Locus Lock Fast Myofiber Terminal Phenotype Iori Sakakibara 1,2,3, Marc Santolini 4, Arnaud Ferry 2,5, Vincent Hakim 4, Pascal Maire 1,2,3 * 1 INSERM U1016,
Event-specific Method for the Quantification of Maize MIR162 Using Real-time PCR. Protocol
 Event-specific Method for the Quantification of Maize MIR162 Using Real-time PCR Protocol 31 January 2011 Joint Research Centre Institute for Health and Consumer Protection Molecular Biology and Genomics
Event-specific Method for the Quantification of Maize MIR162 Using Real-time PCR Protocol 31 January 2011 Joint Research Centre Institute for Health and Consumer Protection Molecular Biology and Genomics
Association of IGF1 and IGFBP3 polymorphisms with colorectal polyps and colorectal cancer risk
 DOI 10.1007/s10552-009-9438-4 ORIGINAL PAPER Association of IGF1 and IGFBP3 polymorphisms with colorectal polyps and colorectal cancer risk Elisabeth Feik Æ Andreas Baierl Æ Barbara Hieger Æ Gerhard Führlinger
DOI 10.1007/s10552-009-9438-4 ORIGINAL PAPER Association of IGF1 and IGFBP3 polymorphisms with colorectal polyps and colorectal cancer risk Elisabeth Feik Æ Andreas Baierl Æ Barbara Hieger Æ Gerhard Führlinger
Archimer http://archimer.ifremer.fr
 Please note that this is an author-produced PDF of an article accepted for publication following peer review. The definitive publisher-authenticated version is available on the publisher Web site Fish
Please note that this is an author-produced PDF of an article accepted for publication following peer review. The definitive publisher-authenticated version is available on the publisher Web site Fish
TITRATION OF raav (VG) USING QUANTITATIVE REAL TIME PCR
 Page 1 of 5 Materials DNase digestion buffer [13 mm Tris-Cl, ph7,5 / 5 mm MgCl2 / 0,12 mm CaCl2] RSS plasmid ptr-uf11 SV40pA Forward primer (10µM) AGC AAT AGC ATC ACA AAT TTC ACA A SV40pA Reverse Primer
Page 1 of 5 Materials DNase digestion buffer [13 mm Tris-Cl, ph7,5 / 5 mm MgCl2 / 0,12 mm CaCl2] RSS plasmid ptr-uf11 SV40pA Forward primer (10µM) AGC AAT AGC ATC ACA AAT TTC ACA A SV40pA Reverse Primer
inhibition of mitosis
 The EMBO Journal vol.13 no.2 pp.425-434, 1994 cdt 1 is an essential target of the Cdc 1 O/Sct 1 transcription factor: requirement for DNA replication and inhibition of mitosis Johannes F.X.Hofmann and
The EMBO Journal vol.13 no.2 pp.425-434, 1994 cdt 1 is an essential target of the Cdc 1 O/Sct 1 transcription factor: requirement for DNA replication and inhibition of mitosis Johannes F.X.Hofmann and
BioTOP-Report. Biotech and Pharma in Berlin-Brandenburg
 BioTOP-Report 2013 Biotech and Pharma in Berlin-Brandenburg BioTOP-Report Content Editorial Unique Chances to Create Value in Biotechnology and Life Sciences 3 Biotechnology More Companies, More Employees
BioTOP-Report 2013 Biotech and Pharma in Berlin-Brandenburg BioTOP-Report Content Editorial Unique Chances to Create Value in Biotechnology and Life Sciences 3 Biotechnology More Companies, More Employees
Protein Synthesis Simulation
 Protein Synthesis Simulation Name(s) Date Period Benchmark: SC.912.L.16.5 as AA: Explain the basic processes of transcription and translation, and how they result in the expression of genes. (Assessed
Protein Synthesis Simulation Name(s) Date Period Benchmark: SC.912.L.16.5 as AA: Explain the basic processes of transcription and translation, and how they result in the expression of genes. (Assessed
Chlamydomonas adapted Green Fluorescent Protein (CrGFP)
 Chlamydomonas adapted Green Fluorescent Protein (CrGFP) Plasmid pfcrgfp for fusion proteins Sequence of the CrGFP In the sequence below, all amino acids which have been altered from the wildtype GFP from
Chlamydomonas adapted Green Fluorescent Protein (CrGFP) Plasmid pfcrgfp for fusion proteins Sequence of the CrGFP In the sequence below, all amino acids which have been altered from the wildtype GFP from
BioTOP-Report. Biotech and Pharma in Berlin-Brandenburg
 BioTOP-Report 2014 Biotech and Pharma in Berlin-Brandenburg BioTOP-Report 2014 Content Editorial The German Capital Region Biotechnology and Life Sciences are Ready for the Future 3 Biotechnology The Capital
BioTOP-Report 2014 Biotech and Pharma in Berlin-Brandenburg BioTOP-Report 2014 Content Editorial The German Capital Region Biotechnology and Life Sciences are Ready for the Future 3 Biotechnology The Capital
2006 7.012 Problem Set 3 KEY
 2006 7.012 Problem Set 3 KEY Due before 5 PM on FRIDAY, October 13, 2006. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Which reaction is catalyzed by each
2006 7.012 Problem Set 3 KEY Due before 5 PM on FRIDAY, October 13, 2006. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Which reaction is catalyzed by each
4 th DVFA Life Science Conference. Going East / Going West. Life Science Asia/Europe getting insight in mutual growth opportunities
 4 th DVFA Life Science Conference Going East / Going West Life Science Asia/Europe getting insight in mutual growth opportunities 4 th DVFA Symposium Life Science followed by a Get Together 17 May 2011,
4 th DVFA Life Science Conference Going East / Going West Life Science Asia/Europe getting insight in mutual growth opportunities 4 th DVFA Symposium Life Science followed by a Get Together 17 May 2011,
Molecular Facts and Figures
 Nucleic Acids Molecular Facts and Figures DNA/RNA bases: DNA and RNA are composed of four bases each. In DNA the four are Adenine (A), Thymidine (T), Cytosine (C), and Guanine (G). In RNA the four are
Nucleic Acids Molecular Facts and Figures DNA/RNA bases: DNA and RNA are composed of four bases each. In DNA the four are Adenine (A), Thymidine (T), Cytosine (C), and Guanine (G). In RNA the four are
On Covert Data Communication Channels Employing DNA Recombinant and Mutagenesis-based Steganographic Techniques
 On Covert Data Communication Channels Employing DNA Recombinant and Mutagenesis-based Steganographic Techniques MAGDY SAEB 1, EMAN EL-ABD 2, MOHAMED E. EL-ZANATY 1 1. School of Engineering, Computer Department,
On Covert Data Communication Channels Employing DNA Recombinant and Mutagenesis-based Steganographic Techniques MAGDY SAEB 1, EMAN EL-ABD 2, MOHAMED E. EL-ZANATY 1 1. School of Engineering, Computer Department,
Gene and Chromosome Mutation Worksheet (reference pgs. 239-240 in Modern Biology textbook)
 Name Date Per Look at the diagrams, then answer the questions. Gene Mutations affect a single gene by changing its base sequence, resulting in an incorrect, or nonfunctional, protein being made. (a) A
Name Date Per Look at the diagrams, then answer the questions. Gene Mutations affect a single gene by changing its base sequence, resulting in an incorrect, or nonfunctional, protein being made. (a) A
Anhang A: Primerliste. 1. Primer für RT-PCR-Analyse. 2. Allgemeine Klonierungsprimer
 Anhang A: Primerliste 1. Primer für RT-PCR-Analyse Primername Primersequenz (5 3 ) Temperatur Zyklenzahl Kin7-f01 gat aat cac ggg tgc tgg 56 C 25 Kin7-r01 gtt gat caa ctg tct acc 56 C 25 MPK4-f01 cgg aga
Anhang A: Primerliste 1. Primer für RT-PCR-Analyse Primername Primersequenz (5 3 ) Temperatur Zyklenzahl Kin7-f01 gat aat cac ggg tgc tgg 56 C 25 Kin7-r01 gtt gat caa ctg tct acc 56 C 25 MPK4-f01 cgg aga
?<BACBC;@@A=2(?@?;@=2:;:%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
 NGS data format NGS data format @SRR031028.1708655 GGATGATGGATGGATAGATAGATGAAGAGATGGATGGATGGGTGGGTGGTATGCAGCATACCTGAAGTGC BBBCB=ABBB@BA=?BABBBBA??B@BAAA>ABB;@5=@@@?8@:==99:465727:;41'.9>;933!4 @SRR031028.843803
NGS data format NGS data format @SRR031028.1708655 GGATGATGGATGGATAGATAGATGAAGAGATGGATGGATGGGTGGGTGGTATGCAGCATACCTGAAGTGC BBBCB=ABBB@BA=?BABBBBA??B@BAAA>ABB;@5=@@@?8@:==99:465727:;41'.9>;933!4 @SRR031028.843803
The nucleotide sequence of the gene for human protein C
 Proc. Natl. Acad. Sci. USA Vol. 82, pp. 4673-4677, July 1985 Biochemistry The nucleotide sequence of the gene for human protein C (DNA sequence analysis/vitamin K-dependent proteins/blood coagulation)
Proc. Natl. Acad. Sci. USA Vol. 82, pp. 4673-4677, July 1985 Biochemistry The nucleotide sequence of the gene for human protein C (DNA sequence analysis/vitamin K-dependent proteins/blood coagulation)
pentr Directional TOPO Cloning Kits
 user guide pentr Directional TOPO Cloning Kits Five-minute, directional TOPO Cloning of blunt-end PCR products into an entry vector for the Gateway System Catalog numbers K2400-20, K2420-20, K2525-20,
user guide pentr Directional TOPO Cloning Kits Five-minute, directional TOPO Cloning of blunt-end PCR products into an entry vector for the Gateway System Catalog numbers K2400-20, K2420-20, K2525-20,
How To Clone Into Pcdna 3.1/V5-His
 pcdna 3.1/V5-His A, B, and C Catalog no. V810-20 Rev. date: 09 November 2010 Manual part no. 28-0141 MAN0000645 User Manual ii Contents Contents and Storage... iv Methods... 1 Cloning into pcdna 3.1/V5-His
pcdna 3.1/V5-His A, B, and C Catalog no. V810-20 Rev. date: 09 November 2010 Manual part no. 28-0141 MAN0000645 User Manual ii Contents Contents and Storage... iv Methods... 1 Cloning into pcdna 3.1/V5-His
TA Cloning Kit. Version V 7 April 2004 25-0024. Catalog nos. K2000-01, K2000-40, K2020-20, K2020-40, K2030-01 K2030-40, K2040-01, K2040-40
 TA Cloning Kit Version V 7 April 2004 25-0024 TA Cloning Kit Catalog nos. K2000-01, K2000-40, K2020-20, K2020-40, K2030-01 K2030-40, K2040-01, K2040-40 ii Table of Contents Table of Contents...iii Important
TA Cloning Kit Version V 7 April 2004 25-0024 TA Cloning Kit Catalog nos. K2000-01, K2000-40, K2020-20, K2020-40, K2030-01 K2030-40, K2040-01, K2040-40 ii Table of Contents Table of Contents...iii Important
ISTEP+: Biology I End-of-Course Assessment Released Items and Scoring Notes
 ISTEP+: Biology I End-of-Course Assessment Released Items and Scoring Notes Page 1 of 22 Introduction Indiana students enrolled in Biology I participated in the ISTEP+: Biology I Graduation Examination
ISTEP+: Biology I End-of-Course Assessment Released Items and Scoring Notes Page 1 of 22 Introduction Indiana students enrolled in Biology I participated in the ISTEP+: Biology I Graduation Examination
PROTOCOL: Illumina Paired-end Whole Exome Capture Library Preparation Using Full-length Index Adaptors and KAPA DNA Polymerase
 PROTOCOL: Illumina Paired-end Whole Exome Capture Library Preparation Using Full-length Index Adaptors and KAPA DNA Polymerase This protocol provides instructions for preparing DNA paired-end capture libraries
PROTOCOL: Illumina Paired-end Whole Exome Capture Library Preparation Using Full-length Index Adaptors and KAPA DNA Polymerase This protocol provides instructions for preparing DNA paired-end capture libraries
Differentiation of Klebsiella pneumoniae and K. oxytoca by Multiplex Polymerase Chain Reaction
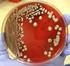 Differentiation of Klebsiella pneumoniae and K. oxytoca by Multiplex Polymerase Chain Reaction Yogesh Chander 1 M. A. Ramakrishnan 1 Naresh Jindal 1 Kevan Hanson 2 Sagar M. Goyal 1 1 Department of Veterinary
Differentiation of Klebsiella pneumoniae and K. oxytoca by Multiplex Polymerase Chain Reaction Yogesh Chander 1 M. A. Ramakrishnan 1 Naresh Jindal 1 Kevan Hanson 2 Sagar M. Goyal 1 1 Department of Veterinary
An Introduction to Bioinformatics Algorithms www.bioalgorithms.info Gene Prediction
 An Introduction to Bioinformatics Algorithms www.bioalgorithms.info Gene Prediction Introduction Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and
An Introduction to Bioinformatics Algorithms www.bioalgorithms.info Gene Prediction Introduction Gene: A sequence of nucleotides coding for protein Gene Prediction Problem: Determine the beginning and
Immortalized epithelial cells from human autosomal dominant polycystic kidney cysts
 Am J Physiol Renal Physiol 285: F397 F412, 2003. First published May 6, 2003; 10.1152/ajprenal.00310.2002. Immortalized epithelial cells from human autosomal dominant polycystic kidney cysts Mahmoud Loghman-Adham,
Am J Physiol Renal Physiol 285: F397 F412, 2003. First published May 6, 2003; 10.1152/ajprenal.00310.2002. Immortalized epithelial cells from human autosomal dominant polycystic kidney cysts Mahmoud Loghman-Adham,
