FOXP3 Controls Regulatory T Cell Function through Cooperation with NFAT
|
|
|
- Shonda Montgomery
- 10 years ago
- Views:
Transcription
1 FOXP3 Controls Regulatory T Cell Function through Cooperation with NFAT Yongqing Wu, 3,5,8 Madhuri Borde, 1,5 Vigo Heissmeyer, 1,5,6 Markus Feuerer, 2 Ariya D. Lapan, 1 James C. Stroud, 3,7 Darren L. Bates, 3 Liang Guo, 3 Aidong Han, 3,8 Steven F. Ziegler, 4 Diane Mathis, 2 Christophe Benoist, 2 Lin Chen, 3,8, * and Anjana Rao 1, * 1 The CBR Institute for Biomedical Research and the Department of Pathology, Harvard Medical School, Boston, MA 02115, USA 2 Section on Immunology and Immunogenetics, Joslin Diabetes Center, Brigham and Women s Hospital, Harvard Medical School, Boston, MA 02115, USA 3 Department of Chemistry and Biochemistry, University of Colorado at Boulder, Boulder, CO 80309, USA 4 Benaroya Research Institute at Virginia Mason and the Department of Immunology, University of Washington, Seattle, WA 98195, USA 5 These authors contributed equally to this work. 6 Present Address: National Research Center for Environment and Health-GSF, Institute of Molecular Immunology, Marchioninistr. 25, D-81377, Munich, Germany. 7 Present Address: UCLA-DOE Institute for Genomics and Proteomics, Los Angeles, CA 90095, USA. 8 Present Address: Molecular and Computational Biology, University of Southern California, 1050 Childs Way, Los Angeles, CA 90089, USA. *Contact: [email protected] (L.C.); [email protected] (A.R.) DOI /j.cell SUMMARY Antigen stimulation of immune cells activates the transcription factor NFAT, a key regulator of T cell activation and anergy. NFAT forms cooperative complexes with the AP-1 family of transcription factors and regulates T cell activation-associated genes. Here we show that regulatory T cell (Treg) function is mediated by an analogous cooperative complex of NFAT with the forkhead transcription factor FOXP3, a lineage specification factor for Tregs. The crystal structure of an NFAT:FOXP2:DNA complex reveals an extensive protein-protein interaction interface between NFAT and FOXP2. Structure-guided mutations of FOXP3, predicted to progressively disrupt its interaction with NFAT, interfere in a graded manner with the ability of FOXP3 to repress expression of the cytokine IL2, upregulate expression of the Treg markers CTLA4 and CD25, and confer suppressor function in a murine model of autoimmune diabetes. Thus by switching transcriptional partners, NFAT converts the acute T cell activation program into the suppressor program of Tregs. INTRODUCTION Eukaryotic genes are regulated through the coordinated actions of multiple transcription factors, which typically engage in combinatorial interactions on DNA (Levine and Tjian, 2003; Miner and Yamamoto, 1991). A well-known example of combinatorial transcriptional control is provided by NFAT, four calcium-regulated transcription factors (NFAT1 4) that regulate gene expression in diverse organs including the immune system, muscle, bone, and brain (Crabtree and Olson, 2002; Hogan et al., 2003). NFAT is critical for the differentiation of many cell types including osteoclasts, slow twitch muscle fibers, and effector T cells (Crabtree and Olson, 2002; Hogan et al., 2003). In cardiac, skeletal, and smooth muscle cells, the relevant interactions are between NFAT and GATA proteins (Hogan et al., 2003), whereas in activated T cells, NFAT forms strong cooperative complexes with AP-1 (Fos-Jun) proteins on composite NFAT:AP-1 DNA elements (Rao et al., 1997). Under conditions of productive T cell activation, NFAT:AP-1 complexes turn on expression of activationassociated genes, whereas under conditions of partial activation (i.e., Ca signaling alone), NFAT activates a distinct set of AP-1-independent genes, which encode negative regulators of T cell signaling and contribute to T cell anergy (Heissmeyer et al., 2004; Heissmeyer and Rao, 2004; Macian et al., 2002). The FOXP family of transcription factors (FOXP1 4) plays critical roles in development (Carlsson and Mahlapuu, 2002). A mutation in the DNA binding domain of FOXP2 cosegregates with a congenital speech disorder (Lai et al., 2001), whereas mutations in FOXP3 are linked to autoimmune disease (Ziegler, 2006). Male scurfy mice, which are natural null mutants for the X-linked Foxp3 gene, die 3 4 weeks after birth as a result of aggressive autoimmunity with multiorgan infiltration (Brunkow et al., 2001). Humans with mutations in FOXP3 develop a similar autoimmune syndrome termed IPEX (immunodysregulation, polyendocrinopathy, enteropathy, X-linked syndrome), characterized by insulin-dependent diabetes, thyroiditis, Cell 126, , July 28, 2006 ª2006 Elsevier Inc. 375
2 massive T cell infiltration into the skin and gastrointestinal tract, high levels of serum autoantibodies, and chronic wasting (Bennett et al., 2001; Wildin et al., 2001). There is compelling evidence that FOXP3 serves as a lineage specification factor for regulatory T cells, a class of T cells that develops primarily in the thymus and is essential for maintaining self-tolerance (Fontenot et al., 2003; Hori et al., 2003; Khattri et al., 2003). T cells from Foxp3-transgenic mice, and primary T cells transduced with Foxp3, acquire many phenotypic and functional characteristics of thymic-derived T regulatory cells (hereafter termed Tregs): increased surface expression of the Treg surface markers CD25, CTLA-4, CD103, and GITR; decreased production of IL-2, IFN-g, and IL- 4 upon restimulation; and ability to suppress effector CD4 T cells in cell culture and in mice (Fontenot et al., 2003; Hori et al., 2003; Khattri et al., 2001, 2003). Notably, many of the genes regulated by FOXP3 are also target genes for NFAT. The IL2 and IL4 genes are activated by NFAT and repressed by FOXP3, while the CD25 and CTLA4 genes are upregulated by both NFAT and FOXP3 (Hogan et al., 2003; Hori et al., 2003; Rao et al., 1997). The mechanism by which FOXP3 influences the expression of NFAT-dependent genes is unknown. Based on the discovery of an IL2 promoter element that contains a binding site for forkhead proteins adjacent to a binding element for NFAT (Schubert et al., 2001; Wang et al., 2003), it has been suggested that FOXP3 represses IL-2 expression by competing with NFAT for binding to DNA (Schubert et al., 2001; Coffer and Burgering, 2004; Fontenot and Rudensky, 2005). An alternative possibility, based on the finding that NFAT and FOXP3 physically interact in cell lysates (Bettelli et al., 2005), is that FOXP3 represses NFAT-driven cytokine transcription by sequestering NFAT away from DNA. The most intriguing possibility, however, is that both the repressive effect of FOXP3 on cytokine gene expression and its activating effect on the CTLA4 and CD25 Treg marker genes reflect cooperative complex formation between NFAT and FOXP3. We provide evidence for this mechanism here. RESULTS FOXP3 Represses NFAT:AP-1-Dependent Transcription and Forms a Cooperative Complex with NFAT Using reporter assays in Jurkat T cells, we showed that the repressive activity of FOXP3 targets the cooperative NFAT:AP-1 complex rather than other configurations of NFAT (Figures 1 and S1). FOXP3 repressed reporter activity driven by three tandem copies of the murine Il2 promoter ARRE2 element, a prototypical NFAT:AP-1-composite site (Chen et al., 1998; Rao et al., 1997) but did not inhibit the activity of a GAL4 fusion protein containing only the transactivation domain (TAD) of NFAT1 (Luo et al., 1996; Okamura et al., 2000)(Figures S1A and S1B). To exclude the possibility that FOXP3 inhibited Ca 2+ /calcineurin signaling upstream of NFAT, we coexpressed FOXP3 with a constitutively active (CA) NFAT1 (Macian et al., 2002; Okamura et al., 2000) and treated the cells with cyclosporin A (CsA) to prevent activation of endogenous NFAT. FOXP3 repressed the transcriptional activity of CA-NFAT1 at the ARRE2 NFAT:AP-1 element (Figure 1A) but not at the Tnf promoter k3 element, a palindromic NFkB-like site at which NFAT forms dimers (McCaffrey et al., 1994) (Figure 1B). We conclude that FOXP3 targets nuclear NFAT, acting specifically on NFAT:AP-1 complexes as opposed to dimeric forms of NFAT. The AP-1 binding sequence in the Il2 promoter ARRE2 site resembles consensus binding sites for forkhead proteins (Carlsson and Mahlapuu, 2002) and for FOXP1 (Wang et al., 2003) (Figure 1C). The forkhead (FKH) domains of three different FOXP proteins, FOXP1, FOXP2, and FOXP3, bound very poorly to the ARRE2 composite site in the absence of NFAT but formed clear cooperative complexes when the NFAT1 DNA binding domain (Rel homology region, RHR) was included in the binding reaction (Figures 1C and S1). This behavior is reminiscent of AP-1, which also binds poorly to this composite site in the absence of NFAT but forms a strong cooperative complex in its presence (Chen et al., 1998; Jain et al., 1993). Thus FOXP3 represses NFAT:AP-1 activity, not by displacing NFAT from NFAT:AP-1 composite sites as suggested previously (Schubert et al., 2001), but rather by forming a cooperative NFAT:FOXP3:DNA complex that resembles the cooperative NFAT:Fos:Jun:DNA complex. Crystal Structure of a Ternary NFAT:FOXP2:DNA Complex To analyze the details of the NFAT:FOXP interaction, we crystallized a ternary complex of the NFAT1 RHR and the FOXP2 forkhead domain bound to double-stranded DNA containing the murine Il2 promoter ARRE2 site. The crystal structure was solved by molecular replacement at 2.7 Å (Table S1). Two NFAT1:FOXP2:DNA complexes with a similar overall structure are observed in the asymmetric unit (Figure 2A); in each complex, the RHR of NFAT binds to the 5 0 half of the ARRE2 site, while the FOXP2 forkhead domain adopts a typical winged-helix fold and binds to the 3 0 half of the ARRE2 site. The two proteins form an extended interaction interface on ARRE2 DNA, burying 614 Å 2 of solvent-accessible surface (Figure 2B). The detailed DNA binding interactions are similar to those seen in previously characterized NFAT:DNA and FOXP2:DNA complexes (Chen et al., 1998; Giffin et al., 2003; Jin et al., 2003; Stroud and Chen, 2003; Stroud et al., 2006)(Figures 2C and 2D). Unexpectedly, however, FOXP2 binds different regions of the murine ARRE2 site in the binary FOXP2:DNA and ternary NFAT:FOXP2:DNA complexes (Figure 2E). In the binary complex (Stroud et al., 2006), FOXP2 binds the sequence 5 0 -GGAA AATTTGTTTCA-3 0 (site A; FOXP binding sites are underlined, bases in bold match with the in vitro selected sequence), which is a better match with the sequence selected in vitro by Foxp1 (5 0 -TATTT(G/A)T-3 0 )(Wang et al., 2003). In the NFAT1:FOXP2:DNA ternary complex, FOXP2 376 Cell 126, , July 28, 2006 ª2006 Elsevier Inc.
3 Figure 1. FOXP3 Specifically Represses NFAT:AP-1-Driven Transcription in Jurkat T Cells and Forms a Cooperative Complex with NFAT on DNA (A) Left, FOXP3 inhibits transcription driven by CA-NFAT1 on the ARRE2 NFAT:AP-1 sites. Right, Structure of the NFAT:AP-1:DNA complex. The sequence of the murine ARRE2 composite element is shown below, with NFAT and AP-1 binding sites in green and red, respectively (Chen et al., 1998). RHR-N, RHR-C: N- and C-terminal domains of the NFAT1 RHR. (B) Left, FOXP3 does not inhibit the activity of CA-NFAT1 dimers on the k3 site from the Tnf promoter. Right, Structure of an NFAT dimer on a palindromic NFkB-like site (Jin et al., 2003). (C) Left, Cooperative binding of recombinant NFAT1 DNA binding domain and FOXP3 forkhead domain to Il2 ARRE2 DNA. Numbers give concentrations in mm. Right, Alignment of the consensus binding site for forkhead proteins (Carlsson and Mahlapuu, 2002) with the ARRE2 composite site. NFAT and AP-1 sites are shown in green and underlined, respectively, and the overlap with the forkhead consensus site is shown in red. binds a different, presumably lower-affinity, site 5 0 -GG AAAATTTGTTTCA-3 0 (site B; FOXP binding sites are underlined, bases in bold match with the in vitro selected sequence). Notably, site B which is permissive for NFAT: FOXP interaction is conserved in the human ARRE2 site whereas site A which is nonpermissive is not (Figure 2E), suggesting that the NFAT:FOXP interaction on the Il2 promoter has been maintained in evolution for functional reasons. The extensive protein-protein interactions between NFAT and FOXP (see below) may compensate for potential lower-affinity binding of FOXP2 to the nonconsensus site B, yielding a ternary NFAT:FOXP2:DNA complex that is thermodynamically more stable than the binary FOXP2:DNA complex. Superposition of the NFAT1:FOXP2:DNA and NFAT1: Fos:Jun:DNA complexes reveals that NFAT1 binds the 5 0 half of the ARRE2 element (GGAAA) almost identically in both complexes, whereas FOXP2 occupies the same DNA region as Fos:Jun (Figures 3A and S2A). In both complexes, the ARRE2 DNA appears to undergo adaptive structural changes to maximize protein-protein interactions, bending upward by about 20 degrees in the NFAT:Fos:Jun:DNA complex to allow the top part of the Fos-Jun coiled-coil to interact with NFAT (Chen et al., Cell 126, , July 28, 2006 ª2006 Elsevier Inc. 377
4 Figure 2. Overall Structure of the NFAT1: FOXP2:DNA Complex (A) Structure of the asymmetric unit showing two complexes of NFAT1 (RHR-N: yellow; RHR-C: green) and FOXP2 (red) bound to the ARRE2 DNA (stick model). (B) Surface representation of the NFAT1: FOXP2:DNA ternary complex viewed from underneath. The areas with relatively positive surface potential (blue) from both proteins form a combined DNA binding surface. The bend of the ARRE2 DNA (stick model), following the path of the combined DNA binding surface, is apparent from this view. (C) Detailed interactions between NFAT1 and DNA. (D) Detailed interactions between FOXP2 and DNA in the ternary NFAT:FOXP2:DNA complex. One notable difference from the FOXP2: DNA binary complex (Stroud et al., 2006) is that the highly conserved Arg553, which is mutated in a severe speech disorder, interacts directly with Gua10 (bold) (5 0 -TGTTT-3) through hydrogen bonding in the ternary complex, in contrast to a water-mediated hydrogen bond between Arg553 and the adjacent thymidine (bold) (5 0 -TGTTT-3 0 ) in the binary complex. Thus, Arg553 appears to play a more direct role in DNA recognition in the ternary complex than in the binary complex. (E) Side view of the NFAT1:FOXP2:DNA complex. FOXP2 (pink) bound to site A in the binary FOXP2:mARRE2 complex (Stroud et al., 2006) is modeled to show the lack of interaction with NFAT. Site B (solid underline), which supports the NFAT:FOXP interaction, is conserved between murine (top) and human (bottom) ARRE2 elements, whereas site A (dashed underline), which does not support the interaction, is not conserved. 1998; Figure S2A) but bending to the side by about 20 degrees in the NFAT1:FOXP2:DNA complex, apparently to align the NFAT and FOXP interfaces for optimal interaction (Figure S2B). This sideways bend is seen in both complexes of the asymmetric unit, suggesting that the bends are not due to crystal packing but are induced by interactions with the distinct partner proteins. Although the two copies of NFAT1 in the asymmetric unit show different orientations in the C-terminal domain (RHR-C), the NFAT:FOXP interface is nearly identical in the two NFAT1:FOXP2:DNA complexes (Figure S2C). Helix H2 of FOXP2 packs against the E F loop of NFAT1, whereas Wing1 of FOXP2 inserts into a large groove formed by the CX loop, the C-terminal stem of the E F loop, and the fg loop of NFAT1 (Figure 3B). Arg537 of NFAT inserts deeply into the minor groove of DNA, separating the NFAT and FOXP binding sites; this positive charge and the AT-rich sequence in this region may explain the significantly compressed minor groove (Williams and Maher, 2000), which apparently helps to bring the interaction surfaces of NFAT and FOXP2 together (Figure 3C). Overall, the NFAT1:FOXP2 interface is primarily polar and contains small hydrated pockets between major foci of contact. These structural features are similar to those of the NFAT:AP-1 interface (Chen et al., 1998) and may explain why NFAT1 and FOXP2 (like NFAT and AP-1) do not form stable complexes in the absence of DNA (Figure S1F). The FOXP binding residues on NFAT1 are conserved in NFAT2 4 (Figure S3), suggesting a conserved FOXP binding surface in the NFAT family. This structure-based alignment also shows that NFAT uses overlapping as well as distinct residues to bind Fos;Jun and FOXP2, respectively. Structure-Based Mutagenesis of NFAT-Interacting Residues in FOXP3 We took advantage of the high sequence conservation of the forkhead domains of FOXP proteins (Figure 3D) to engineer various structure-guided mutations into the predicted NFAT:FOXP3 interface. We focused on NFATinteracting residues oriented away from the DNA binding surface of FOXP3, whose mutation was expected to have a minimal effect on protein folding and stability. (1) RR mutant (E399R E401R): E399 and E401 of FOXP3 378 Cell 126, , July 28, 2006 ª2006 Elsevier Inc.
5 Figure 3. Protein-Protein Interactions between NFAT and FOXP (A) Comparison of crystal structures of NFAT with its partner proteins on the murine ARRE2 site. Fos-Jun and FOXP2 occupy essentially the same region of DNA. The GGAA sequence contacted by NFAT is shown in bold. Left, the NFAT:Fos:Jun:DNA complex (Chen et al., 1998). The AP-1 site is underlined. Right, The NFAT:FOXP2:DNA complex. The FOXP site embedded in the AP-1 site is highlighted in bold. RHR-N, RHR-C: N- and C-terminal domains of the NFAT1 RHR. (B) Top view of the NFAT1:FOXP2:DNA ternary complex in superimposed surface and ribbon representations. NFAT and FOXP2 are colored as in Figure 2A. DNA is in magenta. HNF-3 (FOXA3), which is superimposed on FOXP2, is colored in blue. The structural elements of NFAT and FOXP2 that participate in proteinprotein interactions are indicated (see text). (C) Protein-protein interactions between NFAT1 and FOXP2. Residues at the binding interface between NFAT1 (RHR-N: yellow; RHR-C: green) and FOXP2 (pale red) on DNA (magenta) are colored according to their proteins. (D) Sequence alignment of FOXP1, FOXP2, and FOXP3. Residues interacting with DNA are shaded in magenta, residues interacting with NFAT are shaded in green. Substitutions introduced at NFAT-contacting residues of FOXP3 are indicated. are predicted to form salt bridges with the fg loop of NFAT1 (K 664 R 665 K 666 R 667 )(Figures 3C and 3D). This loop contains conserved positively charged residues in all NFAT proteins (Figure S3). We engineered charge substitutions at these positions into FOXP3, thereby generating the RR mutant. (2) Wing1 insert mutant: Wing1 is uniquely short in the FOXP family compared to other forkhead proteins, and this feature appears critical for binding NFAT (Figure 3B). To interfere sterically with the interaction, we expanded the dimensions of the Wing1 domain of FOXP3. We inserted the additional sequence PDKPG from Wing1 of HNF3 (FOXA3) between E401 and K402 of FOXP3 to generate the Insert mutant. (3) EARR mutant (N361E H365A E399R E401R): To disrupt the NFAT: FOXP3 interaction further, we made an N361E substitution designed to introduce a charge repulsion with E527 of NFAT1, as well as a H365A substitution expected to remove favorable interactions between the imidazole ring of H365 and the E F loop of NFAT. These two mutations were combined with the Wing1 RR mutation to generate the EARR mutant. (4) WRR (N361W E399R E401R) and WWRR (T359W N361W E399R E401R) mutants: A prominent feature of the NFAT1:FOXP2 interface is the nearly perfect shape complementarity. In an attempt to find a combination of mutations that would completely abrogate the NFAT: FOXP3 interaction, we introduced tryptophan residues in place of N361 and T359 of FOXP3, in the hope that the bulky sidechains would interfere with the interaction by clashing sterically with E527 and I535 of NFAT1, respectively (Figures 3C and 3D). The mutations were introduced in the context of both the RR and Insert mutants with comparable results; only the data for the WRR and WWRR mutants are shown here. To confirm that the interface mutations did not affect the DNA binding ability of FOXP3, we performed EMSA assays with bacterially expressed wild-type and mutant forkhead domains. The RR, Insert, EARR, and WRR Cell 126, , July 28, 2006 ª2006 Elsevier Inc. 379
6 Figure 4. FOXP3:NFAT Interface Mutants Repress IL-2 Less Efficiently than Wild-Type FOXP3 (A) T cells from DO11 +/ Ca / mice were infected with wild-type and mutant FOXP3-IRES-GFP retroviruses. Three days later, cells were restimulated for 6 hr with anti-cd3 and anti-cd28 and IL-2 expression was evaluated by intracellular cytokine staining. (B) The same T cells were stained for intracellular FOXP3 without restimulation. (C) Mean fluorescence intensities (MFI) of intracellular (i.c.) IL-2 and GFP (left) or intracellular FOXP3 and GFP (right) were determined for each increment of log GFP expression (see Experimental Procedures). mutants were comparable to wild-type FOXP3 in their ability to bind a consensus forkhead DNA element (V1P) (Li and Tucker, 1993), although, as expected, they were impaired in their ability to form a cooperative NFAT:FOXP3:DNA complex (Figure S4). Because the forkhead domain of the WWRR mutant formed inclusion bodies when expressed in E. coli, precluding its biochemical characterization, we evaluated the transcriptional competence of this mutant by microarray analysis. A subset of genes was similarly affected (upregulated or downregulated) in retrovirally transduced CD4 T cells expressing either the wild-type or the WWRR mutant FOXP3, whereas others were differentially affected (expression altered by wild-type but not by WWRR mutant FOXP3; V.H. and M.F., unpublished data). Thus while the WWRR mutant is impaired in terms of driving expression of certain genes, presumably those dependent on cooperation with NFAT, it is nonetheless active in driving transcription of other genes, presumably those independent of NFAT:FOXP3 cooperation. Mutation of NFAT-Interacting Residues Interferes with Modulation of Gene Expression by FOXP3 We compared the activities of wild-type and mutant FOXP3 proteins by retrovirally expressing them in primary mouse CD4 T cells. All the interface mutations tested decreased the ability of FOXP3 to inhibit IL-2 expression, as judged by intracellular staining for IL-2 expression after restimulation (Figures 4 and S5). The data are represented both as cytofluorimetric contour plots and as plots of mean fluorescence intensity (MFI) of IL-2 against GFP. In multiple dose-response experiments, the Insert, RR, and EARR mutant proteins showed a 2- to 4-fold shift in their inhibitory capacity, while the tryptophan mutants were even more impaired: the WRR mutation markedly reduced and the WWRR mutation eliminated the ability of FOXP3 to inhibit IL-2 (Figures 4A, 4C, and S5). The impairment was not due to differences in expression levels or subcellular localization: the wild-type and mutant FOXP3 proteins showed equivalent expression and nuclear localization in both NIH 3T3 cells and T cells after retroviral transduction (Figures 4B, 4C, and S6). Thus the ability of FOXP3 to inhibit IL-2 production depends crucially on the integrity of the FOXP3-NFAT interface, suggesting that FOXP3 inhibits IL-2 production by forming a cooperative NFAT: FOXP3:DNA complex which replaces the NFAT:AP-1: DNA complex at composite NFAT:AP-1 sites. Indeed, two sites in the Il2 promoter were particularly good matches for a consensus NFAT:FOXP site predicted based on the 380 Cell 126, , July 28, 2006 ª2006 Elsevier Inc.
7 Figure 5. Mutations in the NFAT:FOXP3 Interface Disrupt the Ability of FOXP3 to Upregulate CTLA4 and CD25 (A) T cells from DO11 Ca / mice were infected with wild-type and mutant FOXP3-IRES-GFP retroviruses. Four days later, cells were stained for intracellular (i.c.) CTLA-4 and cell-surface CD25. Left, cytofluorimetric contour plot. Right, CTLA-4 and CD25 MFI were plotted against GFP MFI. (B) T cells from DO11 Ca / mice were infected with wild-type and mutant FOXP3-IRES-GFP retroviruses. Two and four days later, cells were stained for cell-surface CD25 and intracellular CTLA-4, respectively. Left, cytofluorimetric contour plot. Right, CTLA-4 and CD25 MFI were plotted against GFP MFI. crystal structure (Table S2). We cannot rule out, however, that the mutations block interaction with a partner protein other than NFAT. Retroviral transduction of FOXP3 also leads to increased expression of the Treg markers CTLA-4, CD25, GITR, and CD103 (Hori et al., 2003). In particular, Ctla4 has the characteristics of an NFAT-dependent gene: it is induced in a CsA-sensitive manner by treatments that induce intracellular Ca 2+ mobilization, and its promoter contains a composite NFAT:AP-1 element that is a good match for the consensus NFAT:FOXP site predicted based on the crystal structure (Table S2)(Finn et al., 1997; Rao et al., 1997). Indeed, mutations in the NFAT:FOXP3 interface interfered with the ability of retrovirally-transduced FOXP3 to upregulate CTLA-4; the greatest deleterious effect was again observed with the WWRR mutant (Figures 5A and 5B). The mutations had similar but less striking effects on CD25, CD103, and GITR expression (Figures 5A, 5B, and S6). These results suggested that the NFAT:FOXP3 complex could function not only as a repressor of the Il2 gene but also as a direct or indirect activator of the Ctla4 gene, and to a lesser extent the Cd25, Cd103, and Gitr genes. Chromatin immunoprecipitation (ChIP) experiments confirmed that NFAT1 and FOXP3 could each occupy the Il2, Ctla4, and Cd25 promoters, both in T cells retrovirally transduced with FOXP3 and in natural CD4 + CD25 + T regulatory cells that had been expanded with IL-2 (Masteller et al., 2005) (Figure 6). T cells were infected with control or FOXP3-containing IRES-GFP retroviruses, maintained in culture for 4 days (at which point about half of each cell population expressed GFP), then restimulated with PMA and ionomycin. Not only was FOXP3 binding detected at the promoter under these conditions (Figure 6A), but surprisingly, NFAT1 binding to the promoters was considerably increased in FOXP3-expressing cells relative to nonexpressing cells, suggesting stabilization through cooperative binding or increased epitope accessibility (Figure 6B). We also tested expanded Treg populations (about 50% positive for Foxp3 because of overgrowth by contaminating Foxp3-negative cells in the cultures) (Figure 6C). Ionomycin stimulation resulted in binding of NFAT1 as well as Foxp3 to the promoters of the Il2, Ctla4, and Cd25 target genes; binding was substantially diminished in CsA-pretreated cells, again consistent Cell 126, , July 28, 2006 ª2006 Elsevier Inc. 381
8 Figure 6. ChIP Assay for Binding of FOXP3 and NFAT1 to the Il2, Ctla4, and Cd25 Promoters in FOXP3-Transduced T Cells and IL-2- Expanded Tregs Graphs display the numbers of copies of genomic DNA detected in each immunoprecipitation relative to a standard dilution of input. Results are representative of two independent experiments in each case. Shown are mean and standard deviations of triplicate samples for the I12 and Ctla4 promoters, and mean and range of duplicate samples for the Cd25 promoter. (A) T cells were infected with IRES-GFP retroviruses, control or encoding myc-tagged FOXP3, then cultured for 4 days and restimulated for 30 min with PMA and ionomycin. Control and FOXP3-transduced cultures contained 55% and 45% GFP + cells respectively. (B) Chromatin from the stimulation described in (A) was used to assess binding of endogenous NFAT1 to the Il2, Ctla4, and Cd25 promoters. (C) CD4 + CD25 + regulatory T cells were isolated from C57BL/6 mice and expanded in IL-2 (see Experimental Procedures). Foxp3 + cells in the cultures were slowly overgrown by Foxp3 cells: the percentage of Foxp3 + cells determined by intracellular staining was 71% at day 4 and 47% at day 6. On day 6, chromatin was prepared after restimulation with ionomycin for 30 min in the presence or absence of CsA. Binding of endogenous NFAT1 and Foxp3 to the Il2, Ctla4, and Cd25 promoters was determined by ChIP and real-time PCR. with cooperative interactions between NFAT and Foxp3 (Figure 6C). Notably, a truncated FOXP3 protein lacking its N-terminal region (DN FOXP3) was not capable of either downregulating IL-2 or upregulating CTLA-4 and CD25, despite being similar to wild-type FOXP3 in its expression levels and complete localization to the nucleus (Figure 7). Depending on the target gene involved, the N-terninal region presumably recruits coactivator or corepressor proteins that determine transcriptional activity. This effect may involve interaction with a different region of NFAT or may be entirely independent of NFAT. Mutation of the NFAT Contact Surface Impairs the Suppressor Function of FOXP3-Expressing Cells Like bona fide regulatory T cells, primary T cells transduced with FOXP3 are capable of suppressing the activity of effector T cells in several animal models of autoimmune disease (Fontenot et al., 2003; Hori et al., 2003; Jaeckel et al., 2005). We asked whether this suppressive function was compromised by handicapping the interaction of FOXP3 with NFAT. We tested the FOXP3 interface mutants in a mouse model of autoimmune diabetes (Katz et al., 1995) in which regulatory T cells play a vital role. When Th1 cells derived from BDC2.5/NOD TCR transgenic 382 Cell 126, , July 28, 2006 ª2006 Elsevier Inc.
9 Figure 7. The N-Terminal Region of FOXP3 Is Essential for FOXP3 Function (A) Top, Diagram of FOXP3 indicating the zinc finger, the leucine zipper, and the forkhead (FKH) domain. The N-terminal deletion mutant is indicated. Bottom left, equivalent levels of wild-type and N-terminal deletion mutant FOXP3 in lysates of retrovirally infected 3T3 cells as shown by Western blotting. Actin serves as a loading control. Bottom right, nuclear localization of N-terminal deletion mutant in retrovirally infected 3T3 cells by immunocytochemistry using anti-myc antibody. (B) T cells from C57BL/6 mice were infected with IRES-GFP retroviruses encoding wild-type FOXP3 or the N-terminal deletion mutant of FOXP3. Four days later, T cells were stained for intracellular CTLA-4 and CD25 without stimulation, and for IL-2 expression after stimulation with anti-cd3 and anti- CD28 for 6 hr. T cells were transferred into neonatal NOD mice, essentially all recipient animals developed diabetes within 7 20 days (Katz et al., 1995); cotransfer of T cells expressing wild-type FOXP3 protected the animals from disease for a period of at least six weeks (Figure 8). Cotransfer of cells expressing the EARR and WRR mutants of FOXP3 prevented the development of autoimmune disease initially, but the protective effect broke down in 4/9 animals in Figure 8. T Regulatory Function Requires the NFAT:FOXP3 Interface Th1 cells from BDC2.5/NOD mice were retrovirally transduced with control IRES-Thy1.1, wild-type, or mutant FOXP3-expressing viruses. Two days after infection, Thy1.1-positive cells were sorted and transferred into neonatal NOD mice together with untransduced BDC2.5/NOD effector Th1 cells. Recipient mice were monitored for 6 weeks for presentation of diabetes. The results are compiled from two independent experiments with at least four mice per group in each case. Cell 126, , July 28, 2006 ª2006 Elsevier Inc. 383
10 the EARR group and 7/8 animals in the WRR group by day 20. Protection was also inadequate in mice receiving cells expressing the insert mutant along with effector cells: 2/4 mice in this group developed rapid-onset diabetes, one developed diabetes with a substantial delay, and one was protected until termination of the experiment at day 41. Consistent with the strong effects of the WWRR mutation in gene expression assays (Figures 4 and 5), the WWRR mutant protein did not confer protection from disease: 9/10 mice that received cells expressing the WWRR mutant showed rapid disease onset, thus resembling control groups that received effector cells only or effector cells together with control Thy1.1-transduced T cells. Wild-type and mutant FOXP3 proteins were expressed at equivalent levels and localized to the nucleus (Figure S6). We analyzed pancreata from mice in each group for insulitis on day 6, before the clinical manifestation of diabetes (Figure S7). At this time point, there was no apparent difference in the degree of insulitis observed in mice from the different groups. However, in mice coinjected with effector cells and cells expressing wild-type FOXP3, the early inflammation is converted to the innocuous condition of respectful insulitis (Andre et al., 1996), thus protecting against diabetes. This conversion presumably does not occur in mice that develop diabetes after being given cells expressing FOXP3 interface mutants along with effector cells. At day 20 after transfer, we also evaluated the effector:suppressor T cell ratio in pancreatic lymph nodes of surviving mice, by gating on CD4 + T cells and analyzing the proportion of BDC2.5 TCR + T cells that were Thy1.1 + (i.e., had been transduced with FOXP3). Surviving mice in all groups showed similar ratios of Thy1.1 + T cells relative to CD4 + BDC2.5 + effector T cells in the draining lymph nodes (data not shown), showing that the outbreak of disease is associated with loss of function of the FOXP3-expressing cells, and not with their disappearance. Thus the NFAT:FOXP3 complex controls Treg function but not the migration of FOXP3-expressing T cells to pancreatic lymph nodes or their survival in vivo. DISCUSSION We have investigated the molecular interactions between NFAT and FOXP3. We have shown that inhibition of NFAT reporter activity by FOXP3 is apparent at NFAT:AP-1 composite sites (Figure 1), and we have solved the crystal structure of an NFAT:FOXP2:DNA complex (Figures 2 and 3). By introducing structure-guided mutations into FOXP3, we have shown that graded disruption of the predicted NFAT:FOXP3 interface results in progressive loss of function of FOXP3, both in gene expression assays and in an in vivo model of autoimmune diabetes in mice (Katz et al., 1995)(Figures 4, 5, and 8). By ChIP assays performed in FOXP3-transduced T cells as well as in Foxp3- expressing regulatory T cells expanded with IL-2, we have confirmed concurrent binding of FOXP3 and NFAT1 to the promoters of the Il2, Ctla4, and Cd25 target genes (Figure 6). We propose that a single transcription factor, NFAT, directs two entirely contrary biological programs T cell activation and T cell tolerance by recruiting unrelated transcriptional partners AP-1 versus FOXP3 to the regulatory regions of appropriate target genes (Figure S8). How do NFAT:FOXP3 complexes influence gene expression? FOXP3 and Fos-Jun occupy the same region on the ARRE2 site of the Il2 promoter and their binding is expected to be mutually exclusive (Figures 3 and S2). Thus the simplest hypothesis is that FOXP3 competes with AP-1 for cooperative binding with NFAT at a subset of composite NFAT:AP-1 sites, among them the ARRE2 site and the 90 site in the Il2 promoter (Table S2). Nevertheless, simple competition for DNA binding sites is not the entire explanation, as illustrated by the DN mutant which has lost the ability to repress IL-2 expression, upregulate CTLA-4 and CD25, and protect against diabetes (Figure 7; V.H. and M.F., unpublished data), even though it contains the forkhead DNA binding domain and both predicted dimerization domains (leucine zipper and zinc finger), is expressed at equivalent levels, and is fully localized to the nucleus. The N-terminal region of FOXP3 that is missing in the DN mutant may be actively involved in recruiting transcriptional corepressors and coactivators to the Il2 and Ctla4 promoters, respectively, either independently or through a separate interaction with NFAT in the NFAT:FOXP3 complex. A remaining question is whether NFAT:FOXP3 cooperation is required for thymic Treg differentiation. CD4 + CD25 + thymocytes and peripheral T cells from NFAT1 / NFAT4 / double-knockout mice display T regulatory activity in coculture assays (Bopp et al., 2005); however, these cells have elevated nuclear levels of the third family member, NFAT2 (NFATc1) (Ranger et al., 1998); thus it is not yet clear whether Tregs can develop and function in the complete absence of NFAT. To resolve this issue, we are currently introducing our NFAT interface mutations into the endogenous Foxp3 locus. On the other hand, antigen and MHC class II are clearly needed for Treg differentiation in the thymus as well as for the appearance of adaptive/induced Tregs in the periphery (Bluestone and Abbas, 2003). By analogy with the differentiation program of helper T cells (Ansel et al., 2003), we propose that Treg differentiation is initiated by antigen stimulation via NFAT, most likely by imposing a positive feedback loop that converts low-level stochastic expression of Foxp3 into sustained upregulation. This process is likely to be associated with chromatin structural changes that alter the transcriptional competence of Foxp3 target genes as well as the Foxp3 gene itself; consistent with this hypothesis, chromatin at the Il2 promoter is less accessible to MNase digestion in regulatory T cells than in naïve CD4 T cells (Su et al., 2004). In addition, Treg function requires restimulation through the TCR (Hori et al., 2003), consistent with a requirement for formation of the cooperative NFAT:FOXP3 complex; this behavior is similar to that of differentiated Th1 and Th2 cells, which produce their signature cytokines only under conditions where NFAT is activated. 384 Cell 126, , July 28, 2006 ª2006 Elsevier Inc.
11 Indeed, ChIP analysis shows that Foxp3 binding to Il2, Ctla4, and other target genes in Tregs is stabilized when NFAT is activated (Figure 6 and M.B., unpublished data), in the same manner that NFAT activation stabilizes Gata3 and Tbet binding to their respective target genes (Avni et al., 2002; Lee et al., 2004). Our data indicate that the immune system employs NFAT as a common regulator both in effector T cells where it forms a transcriptional complex with AP-1 and in regulatory T cells where it complexes with FOXP3. This ensures that activation of self-reactive T cells and activation of regulatory T cells are equally dependent on the immunogenicity of self-antigens, and are therefore balanced. The combinatorial interaction of NFAT with Fos-Jun and FOXP involves both overlapping and distinct interaction surfaces, and individual residues in the overlapping region display distinct modes of interaction with the different partners. In particular, NFAT1 residues R466 and I467 in the CX loop, and T533 in the E F loop, are critical for Fos-Jun binding (Chen et al., 1998; Macian et al., 2000) but play no apparent role in binding to FOXP2 (Figures 3C and S3). Mutation of these three residues yields NFAT1- RIT, an NFAT1 mutant that is unable to interact with AP-1 (Macian et al., 2000). A constitutively active version of this protein, CA-NFAT1-RIT, is a potent inducer of T cell anergy (Macian et al., 2002) and is also highly effective at cooperating with FOXP3 (V.H., unpublished data). Prolonged antigen stimulation, in the absence of costimulation, leads not only to NFAT-induced anergy but also to the appearance of adaptive or induced Tregs in the periphery (Apostolou and von Boehmer, 2004; Bluestone and Abbas, 2003; Knoechel et al., 2005; Kretschmer et al., 2005). Thus an attractive strategy for inducing immune tolerance would be to develop small molecule inhibitors that block NFAT:AP-1 interaction without interfering with the interaction of NFAT and FOXP3. EXPERIMENTAL PROCEDURES Crystallization of the NFAT1:FOXP2:DNA Ternary Complex The forkhead domain of human FOXP2 (residues ) and the Rel homology region (RHR) of human NFAT1 (residues ) were prepared as previously described (Chen et al., 1998; Stroud et al., 2006) and mixed at a 1:1 molar ratio with murine ARRE2 DNA (5 0 -GGAAAATTTGTTTCA-3 0 ) in 5 mm HEPES, ph 7.63, 2 mm dithiothreitol (DTT), 0.5 mm EDTA, and 150 mm NaCl. Crystals (space group P2 1 ) were grown by hanging drop at 18 C, stabilized in harvest/ cryoprotectant buffer, and flash frozen with liquid N 2 for cryocrystallography. Statistics are presented in Table S1. Coordinates and structures have been deposited in the RCSB protein database under accession code 2AS5. Reporter Assays Jurkat cells ( ) were transfected by electroporation with HAtagged NFAT1 and/or myc-tagged human FOXP3 expression plasmids and luciferase reporter plasmids. Twenty-four hours later, cells were stimulated for 6 hr with 1 mm ionomycin ± 10 nm PMA in the presence of 2 mm CaCl 2 and luciferase activity was measured. When CA-NFAT1 was used, endogenous NFAT activity was inhibited 24 hr after transfection by pretreating cells with 1 mm cyclosporin A for 30 min before addition of stimuli. Results are plotted as relative luciferase units after normalization to renilla, except in the case of Figure S1C where luciferase values are normalized to the values obtained in transfections with reporter plasmid alone. In Figures 1A and 1B, values are normalized to the activity observed with CA-NFAT1 in the absence of FOXP3. All results are representative of 2 3 independent experiments. Electrophoretic Mobility Shift Assays (EMSAs) Binding reactions were performed at room temperature using His 6 - tagged wild-type NFAT1-DBD (Macian et al., 2000), His 6 -tagged human FOXP3 forkhead domain (residues ; described in Supplemental Experimental Procedures), and NFAT:AP-1 ARRE2 probe (5 0 -CAAAGAGGAAAATTTGTTTCATACAGAAGG-3 0 )(Jain et al., 1993) in 10 ml of 10 mm Hepes, 5 mm DTT, 100 mm NaCl, 0.2 mm EDTA, and 5% glycerol. DNA-protein complexes were resolved on 6% polyacrylamide gels. Primary T Cell Cultures and Retroviral Transductions CD4 + T cells were isolated by positive selection from mouse spleens and lymph nodes, stimulated with anti-cd3 and anti-cd28 for 48 hr (Ansel et al., 2004), and spin-infected with retrovirus containing supernatant from Phoenix packaging cells transfected with retroviral expression plasmids (KMV IRES-GFP or MSCV IRES-Thy1.1, empty or encoding wild-type or mutant myc-tagged FOXP3). CD4 + CD25 + regulatory T cells were isolated from spleen and lymph nodes of C57Bl/6 mice by positive selection with anti-cd4 magnetic beads (Dynal) followed by two rounds of CD25 positive selection (Miltenyi Biotec). T cells were stimulated for 48 hr with anti-cd3, anti-cd28, and 2000 U/ml of IL-2, then cultured with 2000 U/ml IL-2 (Masteller et al., 2005). Flow Cytometry and Intracellular Staining T cells were stained intracellularly for CTLA-4 and FOXP3 on days 3 or 4 after retroviral infection, and for cell surface CD25 and GITR on days 2 and 5 after infection, respectively. Staining for IL-2 expression was performed on day 3 after infection after restimulation for 6 hr with anti-cd3 and anti-cd28; Brefeldin A (10 mg/ml; Sigma) was added during the last 2 3 hr of stimulation. Flow cytometry data are presented both as contour plots and as plots of mean fluorescence intensity (MFI) of IL-2, CTLA-4, and CD25, normalized to that of the GFP-negative population within each sample, against the mean fluorescence intensity of GFP. FOXP3 expression was also evaluated by Western blotting and immunocytochemistry using an IgG-purified rabbit antibody to mouse Foxp3 (Cell-tech Chiroscience). Chromatin Immunoprecipitation Control and FOXP3-transduced T cells were stimulated with PMA (5 nm) and ionomycin (0.5 mm) for 30 min and chromatin immunoprecipitation was performed as previously described (Ansel et al., 2004) using the following antibodies: anti-67.1 (against a peptide near the N terminus of NFAT1), anti-t2b1 (against a peptide at the C terminus of NFAT1 isoform C), anti-foxp3 (Cell-tech Chiroscience), anti-myc (9E10), or control rabbit preimmune serum. Binding of FOXP3 and NFAT1 to Il2, Ctla4, and Cd25 promoters was determined by realtime PCR. Results show the average and standard deviation of triplicates, with the exception of the CD25 promoter in Figures 6A and 6B, which shows the average of duplicates. Expanded CD4 + CD25 + T regulatory cells were stimulated for 30 min with ionomycin (1 mm) with or without a 30 min preincubation with CsA (2 mm). Sequences of primer pairs are given in Supplemental Experimental Procedures. Induction of Diabetes in NOD Mouse Model CD4 + T cells were isolated from BDC2.5/NOD mice, activated under Th1 conditions (Ansel et al., 2004), and infected as described above with MSCV IRES-Thy1.1 retrovirus, empty or encoding wild-type or mutant FOXP3. Two days after infection, Thy1.1 + cells were sorted and transferred into neonatal NOD mice at a 1:2 ratio ( : ) with untransduced BDC2.5/NOD Th1 cells ( effector cells). Recipient mice were monitored for diabetes (urinary glucose more than 300 mg/dl Cell 126, , July 28, 2006 ª2006 Elsevier Inc. 385
12 in two consecutive measurements) for 6 weeks. For histological scoring of insulitis, paraffin sections of formalin-fixed pancreata were examined after hematoxylin-eosin staining. Supplemental Data Supplemental Data include two tables, eight figures, Experimental Procedures, and References and can be found with this article online at ACKNOWLEDGMENTS We thank A. Abbas, C. Scheidereit, P.W. Tucker, A. E. Goldfeld, and H. Tong for reagents; J. Goodrich, D. Wuttke, and R. Batey for critical reading of the manuscript and discussion; and the Rao laboratory for helpful discussions. This research was supported by grants from the National Institutes of Health, USA (to A.R., L.C., D.M., C.B., and S.F.Z.), from the W.M. Keck Foundation (to L.C.), from the Juvenile Diabetes Research Foundation (to A.R., D.M., C.B., and S.F.Z.), from the Sandler Program for Asthma Research (to A.R.), and from the American Diabetes Association (to S.F.Z.). M.B. is a fellow of the Ryan Foundation. V.H. was supported by a postdoctoral fellowship from the Cancer Research Institute. M.F. is supported by a postdoctoral fellowship from the Deutsche Forschungsgemeinschaft. Received: November 29, 2005 Revised: April 12, 2006 Accepted: May 22, 2006 Published: July 27, 2006 REFERENCES Andre, I., Gonzalez, A., Wang, B., Katz, J., Benoist, C., and Mathis, D. (1996). Checkpoints in the progression of autoimmune disease: lessons from diabetes models. Proc. Natl. Acad. Sci. USA 93, Ansel, K.M., Lee, D.U., and Rao, A. (2003). An epigenetic view of helper T cell differentiation. Nat. Immunol. 4, Ansel, K.M., Greenwald, R.J., Agarwal, S., Bassing, C.H., Monticelli, S., Interlandi, J., Djuretic, I.M., Lee, D.U., Sharpe, A.H., Alt, F.W., and Rao, A. (2004). Deletion of a conserved Il4 silencer impairs T helper type 1-mediated immunity. Nat. Immunol. 5, Apostolou, I., and von Boehmer, H. (2004). In vivo instruction of suppressor commitment in naive T cells. J. Exp. Med. 199, Avni, O., Lee, D., Macian, F., Szabo, S.J., Glimcher, L.H., and Rao, A. (2002). T(H) cell differentiation is accompanied by dynamic changes in histone acetylation of cytokine genes. Nat. Immunol. 3, Bennett, C.L., Christie, J., Ramsdell, F., Brunkow, M.E., Ferguson, P.J., Whitesell, L., Kelly, T.E., Saulsbury, F.T., Chance, P.F., and Ochs, H.D. (2001). The immune dysregulation, polyendocrinopathy, enteropathy, X-linked syndrome (IPEX) is caused by mutations of FOXP3. Nat. Genet. 27, Bettelli, E., Dastrange, M., and Oukka, M. (2005). Foxp3 interacts with nuclear factor of activated T cells and NF-kappa B to repress cytokine gene expression and effector functions of T helper cells. Proc. Natl. Acad. Sci. USA 102, Bluestone, J.A., and Abbas, A.K. (2003). Natural versus adaptive regulatory T cells. Nat. Rev. Immunol. 3, Bopp, T., Palmetshofer, A., Serfling, E., Heib, V., Schmitt, S., Richter, C., Klein, M., Schild, H., Schmitt, E., and Stassen, M. (2005). NFATc2 and NFATc3 transcription factors play a crucial role in suppression of CD4+ T lymphocytes by CD4+ CD25+ regulatory T cells. J. Exp. Med. 201, Brunkow, M.E., Jeffery, E.W., Hjerrild, K.A., Paeper, B., Clark, L.B., Yasayko, S.A., Wilkinson, J.E., Galas, D., Ziegler, S.F., and Ramsdell, F. (2001). Disruption of a new forkhead/winged-helix protein, scurfin, results in the fatal lymphoproliferative disorder of the scurfy mouse. Nat. Genet. 27, Carlsson, P., and Mahlapuu, M. (2002). Forkhead transcription factors: key players in development and metabolism. Dev. Biol. 250, Chen, L., Glover, J.N., Hogan, P.G., Rao, A., and Harrison, S.C. (1998). Structure of the DNA-binding domains from NFAT, Fos and Jun bound specifically to DNA. Nature 392, Coffer, P.J., and Burgering, B.M. (2004). Forkhead-box transcription factors and their role in the immune system. Nat. Rev. Immunol. 4, Crabtree, G.R., and Olson, E.N. (2002). NFAT signaling: choreographing the social lives of cells. Cell 109 (Suppl ), S67 S79. Finn, P.W., He, H., Wang, Y., Wang, Z., Guan, G., Listman, J., and Perkins, D.L. (1997). Synergistic induction of CTLA-4 expression by costimulation with TCR plus CD28 signals mediated by increased transcription and messenger ribonucleic acid stability. J. Immunol. 158, Fontenot, J.D., and Rudensky, A.Y. (2005). A well adapted regulatory contrivance: regulatory T cell development and the forkhead family transcription factor Foxp3. Nat. Immunol. 6, Fontenot, J.D., Gavin, M.A., and Rudensky, A.Y. (2003). Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 4, Giffin, M.J., Stroud, J.C., Bates, D.L., von Koenig, K.D., Hardin, J., and Chen, L. (2003). Structure of NFAT1 bound as a dimer to the HIV-1 LTR kappa B element. Nat. Struct. Biol. 10, Heissmeyer, V., Macian, F., Im, S.H., Varma, R., Feske, S., Venuprasad, K., Gu, H., Liu, Y.C., Dustin, M.L., and Rao, A. (2004). Calcineurin imposes T cell unresponsiveness through targeted proteolysis of signaling proteins. Nat. Immunol. 5, Heissmeyer, V., and Rao, A. (2004). E3 ligases in T cell anergy turning immune responses into tolerance. Sci. STKE 2004, pe29. Hogan, P.G., Chen, L., Nardone, J., and Rao, A. (2003). Transcriptional regulation by calcium, calcineurin, and NFAT. Genes Dev. 17, Hori, S., Nomura, T., and Sakaguchi, S. (2003). Control of regulatory T cell development by the transcription factor Foxp3. Science 299, Jaeckel, E., von Boehmer, H., and Manns, M.P. (2005). Antigenspecific FoxP3-transduced T-cells can control established type 1 diabetes. Diabetes 54, Jain, J., McCaffrey, P.G., Miner, Z., Kerppola, T.K., Lambert, J.N., Verdine, G.L., Curran, T., and Rao, A. (1993). The T-cell transcription factor NFATp is a substrate for calcineurin and interacts with Fos and Jun. Nature 365, Jin, L., Sliz, P., Chen, L., Macian, F., Rao, A., Hogan, P.G., and Harrison, S.C. (2003). An asymmetric NFAT1 dimer on a pseudopalindromic kappa B-like DNA site. Nat. Struct. Biol. 10, Katz, J.D., Benoist, C., and Mathis, D. (1995). T helper cell subsets in insulin-dependent diabetes. Science 268, Khattri, R., Cox, T., Yasayko, S.A., and Ramsdell, F. (2003). An essential role for Scurfin in CD4+CD25+ T regulatory cells. Nat. Immunol. 4, Khattri, R., Kasprowicz, D., Cox, T., Mortrud, M., Appleby, M.W., Brunkow, M.E., Ziegler, S.F., and Ramsdell, F. (2001). The amount of scurfin protein determines peripheral T cell number and responsiveness. J. Immunol. 167, Knoechel, B., Lohr, J., Kahn, E., Bluestone, J.A., and Abbas, A.K. (2005). Sequential development of interleukin 2-dependent effector and regulatory T cells in response to endogenous systemic antigen. J. Exp. Med. 202, Cell 126, , July 28, 2006 ª2006 Elsevier Inc.
13 Kretschmer, K., Apostolou, I., Hawiger, D., Khazaie, K., Nussenzweig, M.C., and von Boehmer, H. (2005). Inducing and expanding regulatory T cell populations by foreign antigen. Nat. Immunol. 6, Lai, C.S., Fisher, S.E., Hurst, J.A., Vargha-Khadem, F., and Monaco, A.P. (2001). A forkhead-domain gene is mutated in a severe speech and language disorder. Nature 413, Lee, D.U., Avni, O., Chen, L., and Rao, A. (2004). A distal enhancer in the interferon-gamma (IFN-gamma) locus revealed by genome sequence comparison. J. Biol. Chem. 279, Levine, M., and Tjian, R. (2003). Transcription regulation and animal diversity. Nature 424, Li, C., and Tucker, P.W. (1993). DNA-binding properties and secondary structural model of the hepatocyte nuclear factor 3/fork head domain. Proc. Natl. Acad. Sci. USA 90, Luo, C., Burgeon, E., and Rao, A. (1996). Mechanisms of transactivation by nuclear factor of activated T cells-1. J. Exp. Med. 184, Macian, F., Garcia-Rodriguez, C., and Rao, A. (2000). Gene expression elicited by NFAT in the presence or absence of cooperative recruitment of Fos and Jun. EMBO J. 19, Macian, F., Garcia-Cozar, F., Im, S.H., Horton, H.F., Byrne, M.C., and Rao, A. (2002). Transcriptional mechanisms underlying lymphocyte tolerance. Cell 109, Masteller, E.L., Warner, M.R., Tang, Q., Tarbell, K.V., McDevitt, H., and Bluestone, J.A. (2005). Expansion of functional endogenous antigenspecific CD4+CD25+ regulatory T cells from nonobese diabetic mice. J. Immunol. 175, McCaffrey, P.G., Goldfeld, A.E., and Rao, A. (1994). The role of NFATp in cyclosporin A-sensitive tumor necrosis factor-alpha gene transcription. J. Biol. Chem. 269, Miner, J.N., and Yamamoto, K.R. (1991). Regulatory crosstalk at composite response elements. Trends Biochem. Sci. 16, Okamura, H., Aramburu, J., Garcia-Rodriguez, C., Viola, J.P., Raghavan, A., Tahiliani, M., Zhang, X., Qin, J., Hogan, P.G., and Rao, A. (2000). Concerted dephosphorylation of the transcription factor NFAT1 induces a conformational switch that regulates transcriptional activity. Mol. Cell 6, Ranger, A.M., Oukka, M., Rengarajan, J., and Glimcher, L.H. (1998). Inhibitory function of two NFAT family members in lymphoid homeostasis and Th2 development. Immunity 9, Rao, A., Luo, C., and Hogan, P.G. (1997). Transcription factors of the NFAT family: regulation and function. Annu. Rev. Immunol. 15, Schubert, L.A., Jeffery, E., Zhang, Y., Ramsdell, F., and Ziegler, S.F. (2001). Scurfin (FOXP3) acts as a repressor of transcription and regulates T cell activation. J. Biol. Chem. 276, Stroud, J.C., and Chen, L. (2003). Structure of NFAT bound to DNA as a monomer. J. Mol. Biol. 334, Stroud, J.C., Wu, Y., Bates, D.L., Han, A., Nowick, K., Paabo, S., Tong, H., and Chen, L. (2006). Structure of the forkhead domain of FOXP2 bound to DNA. Structure 14, Su, L., Creusot, R.J., Gallo, E.M., Chan, S.M., Utz, P.J., Fathman, C.G., and Ermann, J. (2004). Murine CD4+CD25+ regulatory T cells fail to undergo chromatin remodeling across the proximal promoter region of the IL-2 gene. J. Immunol. 173, Wang, B., Lin, D., Li, C., and Tucker, P. (2003). Multiple domains define the expression and regulatory properties of Foxp1 forkhead transcriptional repressors. J. Biol. Chem. 278, Wildin, R.S., Ramsdell, F., Peake, J., Faravelli, F., Casanova, J.L., Buist, N., Levy-Lahad, E., Mazzella, M., Goulet, O., Perroni, L., et al. (2001). X-linked neonatal diabetes mellitus, enteropathy and endocrinopathy syndrome is the human equivalent of mouse scurfy. Nat. Genet. 27, Williams, L.D., and Maher, L.J., 3rd. (2000). Electrostatic mechanisms of DNA deformation. Annu. Rev. Biophys. Biomol. Struct. 29, Ziegler, S.F. (2006). FOXP3: Of mice and men. Annu. Rev. Immunol. 24, Cell 126, , July 28, 2006 ª2006 Elsevier Inc. 387
2.1.2 Characterization of antiviral effect of cytokine expression on HBV replication in transduced mouse hepatocytes line
 i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
Autoimmunity and immunemediated. FOCiS. Lecture outline
 1 Autoimmunity and immunemediated inflammatory diseases Abul K. Abbas, MD UCSF FOCiS 2 Lecture outline Pathogenesis of autoimmunity: why selftolerance fails Genetics of autoimmune diseases Therapeutic
1 Autoimmunity and immunemediated inflammatory diseases Abul K. Abbas, MD UCSF FOCiS 2 Lecture outline Pathogenesis of autoimmunity: why selftolerance fails Genetics of autoimmune diseases Therapeutic
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
T Cell Maturation,Activation and Differentiation
 T Cell Maturation,Activation and Differentiation Positive Selection- In thymus, permits survival of only those T cells whose TCRs recognize self- MHC molecules (self-mhc restriction) Negative Selection-
T Cell Maturation,Activation and Differentiation Positive Selection- In thymus, permits survival of only those T cells whose TCRs recognize self- MHC molecules (self-mhc restriction) Negative Selection-
The immune system. Bone marrow. Thymus. Spleen. Bone marrow. NK cell. B-cell. T-cell. Basophil Neutrophil. Eosinophil. Myeloid progenitor
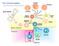 The immune system Basophil Neutrophil Bone marrow Eosinophil Myeloid progenitor Dendritic cell Pluripotent Stem cell Lymphoid progenitor Platelets Bone marrow Thymus NK cell T-cell B-cell Spleen Cancer
The immune system Basophil Neutrophil Bone marrow Eosinophil Myeloid progenitor Dendritic cell Pluripotent Stem cell Lymphoid progenitor Platelets Bone marrow Thymus NK cell T-cell B-cell Spleen Cancer
Understanding the immune response to bacterial infections
 Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
The Need for a PARP in vivo Pharmacodynamic Assay
 The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D., Chief Scientific Officer, Trevigen, Inc., Gaithersburg, MD For further infomation, please contact: William Booth, Ph.D. Tel: +44 (0)1235
The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D., Chief Scientific Officer, Trevigen, Inc., Gaithersburg, MD For further infomation, please contact: William Booth, Ph.D. Tel: +44 (0)1235
CD3/TCR stimulation and surface detection Determination of specificity of intracellular detection of IL-7Rα by flow cytometry
 CD3/TCR stimulation and surface detection Stimulation of HPB-ALL cells with the anti-cd3 monoclonal antibody OKT3 was performed as described 3. In brief, antibody-coated plates were prepared by incubating
CD3/TCR stimulation and surface detection Stimulation of HPB-ALL cells with the anti-cd3 monoclonal antibody OKT3 was performed as described 3. In brief, antibody-coated plates were prepared by incubating
CONTENT. Chapter 1 Review of Literature. List of figures. List of tables
 Abstract Abbreviations List of figures CONTENT I-VI VII-VIII IX-XII List of tables XIII Chapter 1 Review of Literature 1. Vaccination against intracellular pathogens 1-34 1.1 Role of different immune responses
Abstract Abbreviations List of figures CONTENT I-VI VII-VIII IX-XII List of tables XIII Chapter 1 Review of Literature 1. Vaccination against intracellular pathogens 1-34 1.1 Role of different immune responses
博 士 論 文 ( 要 約 ) A study on enzymatic synthesis of. stable cyclized peptides which. inhibit protein-protein interactions
 博 士 論 文 ( 要 約 ) 論 文 題 目 A study on enzymatic synthesis of stable cyclized peptides which inhibit protein-protein interactions ( 蛋 白 質 間 相 互 作 用 を 阻 害 する 安 定 な 環 状 化 ペプチドの 酵 素 合 成 に 関 する 研 究 ) 氏 名 張 静 1
博 士 論 文 ( 要 約 ) 論 文 題 目 A study on enzymatic synthesis of stable cyclized peptides which inhibit protein-protein interactions ( 蛋 白 質 間 相 互 作 用 を 阻 害 する 安 定 な 環 状 化 ペプチドの 酵 素 合 成 に 関 する 研 究 ) 氏 名 張 静 1
specific B cells Humoral immunity lymphocytes antibodies B cells bone marrow Cell-mediated immunity: T cells antibodies proteins
 Adaptive Immunity Chapter 17: Adaptive (specific) Immunity Bio 139 Dr. Amy Rogers Host defenses that are specific to a particular infectious agent Can be innate or genetic for humans as a group: most microbes
Adaptive Immunity Chapter 17: Adaptive (specific) Immunity Bio 139 Dr. Amy Rogers Host defenses that are specific to a particular infectious agent Can be innate or genetic for humans as a group: most microbes
The role of IBV proteins in protection: cellular immune responses. COST meeting WG2 + WG3 Budapest, Hungary, 2015
 The role of IBV proteins in protection: cellular immune responses COST meeting WG2 + WG3 Budapest, Hungary, 2015 1 Presentation include: Laboratory results Literature summary Role of T cells in response
The role of IBV proteins in protection: cellular immune responses COST meeting WG2 + WG3 Budapest, Hungary, 2015 1 Presentation include: Laboratory results Literature summary Role of T cells in response
Chapter 18: Applications of Immunology
 Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
Vitamin D deficiency exacerbates ischemic cell loss and sensory motor dysfunction in an experimental stroke model
 Vitamin D deficiency exacerbates ischemic cell loss and sensory motor dysfunction in an experimental stroke model Robyn Balden & Farida Sohrabji Texas A&M Health Science Center- College of Medicine ISC
Vitamin D deficiency exacerbates ischemic cell loss and sensory motor dysfunction in an experimental stroke model Robyn Balden & Farida Sohrabji Texas A&M Health Science Center- College of Medicine ISC
Uses of Flow Cytometry
 Uses of Flow Cytometry 1. Multicolour analysis... 2 2. Cell Cycle and Proliferation... 3 a. Analysis of Cellular DNA Content... 4 b. Cell Proliferation Assays... 5 3. Immunology... 6 4. Apoptosis... 7
Uses of Flow Cytometry 1. Multicolour analysis... 2 2. Cell Cycle and Proliferation... 3 a. Analysis of Cellular DNA Content... 4 b. Cell Proliferation Assays... 5 3. Immunology... 6 4. Apoptosis... 7
Making the switch to a safer CAR-T cell therapy
 Making the switch to a safer CAR-T cell therapy HaemaLogiX 2015 Technical Journal Club May 24 th 2016 Christina Müller - chimeric antigen receptor = CAR - CAR T cells are generated by lentiviral transduction
Making the switch to a safer CAR-T cell therapy HaemaLogiX 2015 Technical Journal Club May 24 th 2016 Christina Müller - chimeric antigen receptor = CAR - CAR T cells are generated by lentiviral transduction
GENE REGULATION. Teacher Packet
 AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
How to construct transgenic mice
 How to construct transgenic mice Sandra Beer-Hammer Autumn School 2010 Bad Schandau Methods additional genetic information transgenic mouse line gene inactivation gene-deficient knockout mouse line Jak2
How to construct transgenic mice Sandra Beer-Hammer Autumn School 2010 Bad Schandau Methods additional genetic information transgenic mouse line gene inactivation gene-deficient knockout mouse line Jak2
Activation and effector functions of HMI
 Activation and effector functions of HMI Hathairat Thananchai, DPhil Department of Microbiology Faculty of Medicine Chiang Mai University 25 August 2015 ว ตถ ประสงค หล งจากช วโมงบรรยายน แล วน กศ กษาสามารถ
Activation and effector functions of HMI Hathairat Thananchai, DPhil Department of Microbiology Faculty of Medicine Chiang Mai University 25 August 2015 ว ตถ ประสงค หล งจากช วโมงบรรยายน แล วน กศ กษาสามารถ
Cancer Immunotherapy: Can Your Immune System Cure Cancer? Steve Emerson, MD, PhD Herbert Irving Comprehensive Cancer Center
 Cancer Immunotherapy: Can Your Immune System Cure Cancer? Steve Emerson, MD, PhD Herbert Irving Comprehensive Cancer Center Bodnar s Law Simple Things are Important Very Simple Things are Very Important
Cancer Immunotherapy: Can Your Immune System Cure Cancer? Steve Emerson, MD, PhD Herbert Irving Comprehensive Cancer Center Bodnar s Law Simple Things are Important Very Simple Things are Very Important
The Making of the Fittest: Evolving Switches, Evolving Bodies
 OVERVIEW MODELING THE REGULATORY SWITCHES OF THE PITX1 GENE IN STICKLEBACK FISH This hands-on activity supports the short film, The Making of the Fittest:, and aims to help students understand eukaryotic
OVERVIEW MODELING THE REGULATORY SWITCHES OF THE PITX1 GENE IN STICKLEBACK FISH This hands-on activity supports the short film, The Making of the Fittest:, and aims to help students understand eukaryotic
ELISA BIO 110 Lab 1. Immunity and Disease
 ELISA BIO 110 Lab 1 Immunity and Disease Introduction The principal role of the mammalian immune response is to contain infectious disease agents. This response is mediated by several cellular and molecular
ELISA BIO 110 Lab 1 Immunity and Disease Introduction The principal role of the mammalian immune response is to contain infectious disease agents. This response is mediated by several cellular and molecular
Modelling and analysis of T-cell epitope screening data.
 Modelling and analysis of T-cell epitope screening data. Tim Beißbarth 2, Jason A. Tye-Din 1, Gordon K. Smyth 1, Robert P. Anderson 1 and Terence P. Speed 1 1 WEHI, 1G Royal Parade, Parkville, VIC 3050,
Modelling and analysis of T-cell epitope screening data. Tim Beißbarth 2, Jason A. Tye-Din 1, Gordon K. Smyth 1, Robert P. Anderson 1 and Terence P. Speed 1 1 WEHI, 1G Royal Parade, Parkville, VIC 3050,
Head of College Scholars List Scheme. Summer Studentship. Report Form
 Head of College Scholars List Scheme Summer Studentship Report Form This report should be completed by the student with his/her project supervisor. It should summarise the work undertaken during the project
Head of College Scholars List Scheme Summer Studentship Report Form This report should be completed by the student with his/her project supervisor. It should summarise the work undertaken during the project
A Genetic Analysis of Rheumatoid Arthritis
 A Genetic Analysis of Rheumatoid Arthritis Introduction to Rheumatoid Arthritis: Classification and Diagnosis Rheumatoid arthritis is a chronic inflammatory disorder that affects mainly synovial joints.
A Genetic Analysis of Rheumatoid Arthritis Introduction to Rheumatoid Arthritis: Classification and Diagnosis Rheumatoid arthritis is a chronic inflammatory disorder that affects mainly synovial joints.
Rheumatoid arthritis: an overview. Christine Pham MD
 Rheumatoid arthritis: an overview Christine Pham MD RA prevalence Chronic inflammatory disease affecting approximately 0.5 1% of the general population Prevalence is higher in North America (approaching
Rheumatoid arthritis: an overview Christine Pham MD RA prevalence Chronic inflammatory disease affecting approximately 0.5 1% of the general population Prevalence is higher in North America (approaching
B Cell Generation, Activation & Differentiation. B cell maturation
 B Cell Generation, Activation & Differentiation Naïve B cells- have not encountered Ag. Have IgM and IgD on cell surface : have same binding VDJ regions but different constant region leaves bone marrow
B Cell Generation, Activation & Differentiation Naïve B cells- have not encountered Ag. Have IgM and IgD on cell surface : have same binding VDJ regions but different constant region leaves bone marrow
TEMA 10. REACCIONES INMUNITARIAS MEDIADAS POR CÉLULAS.
 TEMA 10. REACCIONES INMUNITARIAS MEDIADAS POR CÉLULAS. The nomenclature of cytokines partly reflects their first-described function and also the order of their discovery. There is no single unified nomenclature,
TEMA 10. REACCIONES INMUNITARIAS MEDIADAS POR CÉLULAS. The nomenclature of cytokines partly reflects their first-described function and also the order of their discovery. There is no single unified nomenclature,
GeneCopoeia Genome Editing Tools for Safe Harbor Integration in. Mice and Humans. Ed Davis, Liuqing Qian, Ruiqing li, Junsheng Zhou, and Jinkuo Zhang
 G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
B cell activation and Humoral Immunity
 B cell activation and Humoral Immunity Humoral immunity is mediated by secreted antibodies and its physiological function is defense against extracellular microbes (including viruses) and microbial exotoxins.
B cell activation and Humoral Immunity Humoral immunity is mediated by secreted antibodies and its physiological function is defense against extracellular microbes (including viruses) and microbial exotoxins.
Overview. Transcriptional cascades. Amazing aspects of lineage plasticity. Conventional (B2) B cell development
 Overview B cell development Transcriptional cascades Amazing aspects of lineage plasticity Conventional (B2) B cell development What happens to an autoreactive B cell? B1 vs B2 cells Key anatomical sites
Overview B cell development Transcriptional cascades Amazing aspects of lineage plasticity Conventional (B2) B cell development What happens to an autoreactive B cell? B1 vs B2 cells Key anatomical sites
CCR Biology - Chapter 9 Practice Test - Summer 2012
 Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
New HLA class I epitopes defined by murine monoclonal antibodies
 Human Immunology 71 (2010) 456 461 Contents lists available at ScienceDirect New HLA class I epitopes defined by murine monoclonal antibodies Nadim El-Awar a, *, Paul I. Terasaki b, Anh Nguyen a, Mamie
Human Immunology 71 (2010) 456 461 Contents lists available at ScienceDirect New HLA class I epitopes defined by murine monoclonal antibodies Nadim El-Awar a, *, Paul I. Terasaki b, Anh Nguyen a, Mamie
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
A link between gut microbiota, mucosal immunity and autoimmune arthritis. Hsin-Jung Joyce Wu "Microbiota and man: the story about us
 A link between gut microbiota, mucosal immunity and autoimmune arthritis Microbes and Man The majority of microbes we encounter are gut microbiota that live mostly in harmony with their host Duodenum 10
A link between gut microbiota, mucosal immunity and autoimmune arthritis Microbes and Man The majority of microbes we encounter are gut microbiota that live mostly in harmony with their host Duodenum 10
2) Macrophages function to engulf and present antigen to other immune cells.
 Immunology The immune system has specificity and memory. It specifically recognizes different antigens and has memory for these same antigens the next time they are encountered. The Cellular Components
Immunology The immune system has specificity and memory. It specifically recognizes different antigens and has memory for these same antigens the next time they are encountered. The Cellular Components
Antigenic variation in Plasmodium falciparum : Erythrocyte invasion and immune escape mechanisms
 Antigenic variation in Plasmodium falciparum : Erythrocyte invasion and immune escape mechanisms Introduction Why does immunity to malaria take so long to develop? The parasite s survival depends on its
Antigenic variation in Plasmodium falciparum : Erythrocyte invasion and immune escape mechanisms Introduction Why does immunity to malaria take so long to develop? The parasite s survival depends on its
B Cells and Antibodies
 B Cells and Antibodies Andrew Lichtman, MD PhD Brigham and Women's Hospital Harvard Medical School Lecture outline Functions of antibodies B cell activation; the role of helper T cells in antibody production
B Cells and Antibodies Andrew Lichtman, MD PhD Brigham and Women's Hospital Harvard Medical School Lecture outline Functions of antibodies B cell activation; the role of helper T cells in antibody production
Anti-ATF6 α antibody, mouse monoclonal (1-7)
 Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
Chapter 6. Antigen-Antibody Properties 10/3/2012. Antigen-Antibody Interactions: Principles and Applications. Precipitin reactions
 Chapter 6 Antigen-Antibody Interactions: Principles and Applications Antigen-Antibody Properties You must remember antibody affinity (single) VS avidity (multiple) High affinity: bound tightly and longer!
Chapter 6 Antigen-Antibody Interactions: Principles and Applications Antigen-Antibody Properties You must remember antibody affinity (single) VS avidity (multiple) High affinity: bound tightly and longer!
How To Understand How Gene Expression Is Regulated
 What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
BioMmune Technologies Inc. Corporate Presentation 2015
 BioMmune Technologies Inc Corporate Presentation 2015 * Harnessing the body s own immune system to fight cancer & other autoimmune diseases BioMmune Technologies Inc. (IMU) ABOUT A public biopharmaceutical
BioMmune Technologies Inc Corporate Presentation 2015 * Harnessing the body s own immune system to fight cancer & other autoimmune diseases BioMmune Technologies Inc. (IMU) ABOUT A public biopharmaceutical
Final Review. Aptamers. Making Aptamers: SELEX 6/3/2011. sirna and mirna. Central Dogma. RNAi: A translation regulation mechanism.
 Central Dogma Final Review Section Week 10 DNA RNA Protein DNA DNA replication DNA RNA transcription RNA Protein translation **RNA DNA reverse transcription http://bass.bio.uci.edu/~hudel/bs99a/lecture20/lecture1_1.html
Central Dogma Final Review Section Week 10 DNA RNA Protein DNA DNA replication DNA RNA transcription RNA Protein translation **RNA DNA reverse transcription http://bass.bio.uci.edu/~hudel/bs99a/lecture20/lecture1_1.html
SUPPLEMENTARY DATA 1
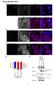 SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
Rapid GST Inclusion Body Solubilization and Renaturation Kit
 Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
Module 3 Questions. 7. Chemotaxis is an example of signal transduction. Explain, with the use of diagrams.
 Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
TABLE OF CONTENT. Page ACKNOWLEDGEMENTS. iii ENGLISH ABSTRACT THAI ABSTRACT. vii LIST OF TABLES LIST OF FIGURES. xvi ABBREVIATIONS.
 x TABLE OF CONTENT ACKNOWLEDGEMENTS ENGLISH ABSTRACT THAI ABSTRACT LIST OF TABLES LIST OF FIGURES ABBREVIATIONS iii iv vii xv xvi xviii CHAPTER I: INTRODUCTION 1.1 Statement of problems 1 1.2 Literature
x TABLE OF CONTENT ACKNOWLEDGEMENTS ENGLISH ABSTRACT THAI ABSTRACT LIST OF TABLES LIST OF FIGURES ABBREVIATIONS iii iv vii xv xvi xviii CHAPTER I: INTRODUCTION 1.1 Statement of problems 1 1.2 Literature
S1 Text. Modeling deterministic single-cell microrna-p53-mdm2 network Figure 2 Figure 2
 S1 Text. Modeling deterministic single-cell microrna-p53-mdm2 network The schematic diagram of the microrna-p53-mdm2 oscillator is illustrated in Figure 2. The interaction scheme among the mrnas and the
S1 Text. Modeling deterministic single-cell microrna-p53-mdm2 network The schematic diagram of the microrna-p53-mdm2 oscillator is illustrated in Figure 2. The interaction scheme among the mrnas and the
Superior TrueMAB TM monoclonal antibodies for the recognition of proteins native epitopes
 Superior TrueMAB TM monoclonal antibodies for the recognition of proteins native epitopes Outlines Brief introduction of OriGene s mission on gene-centric product solution. TrueMAB monoclonal antibody
Superior TrueMAB TM monoclonal antibodies for the recognition of proteins native epitopes Outlines Brief introduction of OriGene s mission on gene-centric product solution. TrueMAB monoclonal antibody
Kinexus has an in-house inventory of lysates prepared from 16 human cancer cell lines that have been selected to represent a diversity of tissues,
 Kinexus Bioinformatics Corporation is seeking to map and monitor the molecular communications networks of living cells for biomedical research into the diagnosis, prognosis and treatment of human diseases.
Kinexus Bioinformatics Corporation is seeking to map and monitor the molecular communications networks of living cells for biomedical research into the diagnosis, prognosis and treatment of human diseases.
Validation & Assay Performance Summary
 Validation & Assay Performance Summary GeneBLAzer RAR alpha DA Assay Kit GeneBLAzer RAR alpha-uas-bla GripTite Cells Cat. no. K1387, K1685 Target Description The retinoic acid receptor alpha (RAR alpha)
Validation & Assay Performance Summary GeneBLAzer RAR alpha DA Assay Kit GeneBLAzer RAR alpha-uas-bla GripTite Cells Cat. no. K1387, K1685 Target Description The retinoic acid receptor alpha (RAR alpha)
Gene Regulation -- The Lac Operon
 Gene Regulation -- The Lac Operon Specific proteins are present in different tissues and some appear only at certain times during development. All cells of a higher organism have the full set of genes:
Gene Regulation -- The Lac Operon Specific proteins are present in different tissues and some appear only at certain times during development. All cells of a higher organism have the full set of genes:
Dendritic Cells: A Basic Review *last updated May 2003
 *last updated May 2003 Prepared by: Eric Wieder, PhD MD Anderson Cancer Center Houston, TX USA What is a dendritic cell? Dendritic cells are antigen-presenting cells (APCs) which play a critical role in
*last updated May 2003 Prepared by: Eric Wieder, PhD MD Anderson Cancer Center Houston, TX USA What is a dendritic cell? Dendritic cells are antigen-presenting cells (APCs) which play a critical role in
pcas-guide System Validation in Genome Editing
 pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
LESSON 3: ANTIBODIES/BCR/B-CELL RESPONSES
 Introduction to immunology. LESSON 3: ANTIBODIES/BCR/B-CELL RESPONSES Today we will get to know: The antibodies How antibodies are produced, their classes and their maturation processes Antigen recognition
Introduction to immunology. LESSON 3: ANTIBODIES/BCR/B-CELL RESPONSES Today we will get to know: The antibodies How antibodies are produced, their classes and their maturation processes Antigen recognition
The Journal of Experimental Medicine
 Supplemental Material Haines et al., http://www.jem.org/cgi/content/full/jem.20080996/dc1 The Journal of Experimental Medicine Subcloning of murine PTK7 cdna segments into pegfp-n3 and CHO cell transfection.
Supplemental Material Haines et al., http://www.jem.org/cgi/content/full/jem.20080996/dc1 The Journal of Experimental Medicine Subcloning of murine PTK7 cdna segments into pegfp-n3 and CHO cell transfection.
Mouse IFN-gamma ELISpot Kit
 Page 1 of 8 Mouse IFN-gamma ELISpot Kit Without Plates With Plates With Sterile Plates Quantity Catalog Nos. 862.031.001 862.031.001P 862.031.001S 1 x 96 tests 862.031.005 862.031.005P 862.031.005S 5 x
Page 1 of 8 Mouse IFN-gamma ELISpot Kit Without Plates With Plates With Sterile Plates Quantity Catalog Nos. 862.031.001 862.031.001P 862.031.001S 1 x 96 tests 862.031.005 862.031.005P 862.031.005S 5 x
Microbiology AN INTRODUCTION EIGHTH EDITION
 TORTORA FUNKE CASE Microbiology AN INTRODUCTION EIGHTH EDITION Differentiate between innate and acquired immunity. Chapter 17 Specific Defenses of the Host: The Immune Response B.E Pruitt & Jane J. Stein
TORTORA FUNKE CASE Microbiology AN INTRODUCTION EIGHTH EDITION Differentiate between innate and acquired immunity. Chapter 17 Specific Defenses of the Host: The Immune Response B.E Pruitt & Jane J. Stein
Biochemistry 462a Hemoglobin Structure and Function Reading - Chapter 7 Practice problems - Chapter 7: 1-6; Proteins extra problems
 Biochemistry 462a Hemoglobin Structure and Function Reading - Chapter 7 Practice problems - Chapter 7: 1-6; Proteins extra problems Myoglobin and Hemoglobin Oxygen is required for oxidative metabolism
Biochemistry 462a Hemoglobin Structure and Function Reading - Chapter 7 Practice problems - Chapter 7: 1-6; Proteins extra problems Myoglobin and Hemoglobin Oxygen is required for oxidative metabolism
Recognition of T cell epitopes (Abbas Chapter 6)
 Recognition of T cell epitopes (Abbas Chapter 6) Functions of different APCs (Abbas Chapter 6)!!! Directon Routes of antigen entry (Abbas Chapter 6) Flow of Information Barrier APCs LNs Sequence of Events
Recognition of T cell epitopes (Abbas Chapter 6) Functions of different APCs (Abbas Chapter 6)!!! Directon Routes of antigen entry (Abbas Chapter 6) Flow of Information Barrier APCs LNs Sequence of Events
DNA Fingerprinting. Unless they are identical twins, individuals have unique DNA
 DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
CUSTOM ANTIBODIES. Fully customised services: rat and murine monoclonals, rat and rabbit polyclonals, antibody characterisation, antigen preparation
 CUSTOM ANTIBODIES Highly competitive pricing without compromising quality. Rat monoclonal antibodies for the study of gene expression and proteomics in mice and in mouse models of human diseases available.
CUSTOM ANTIBODIES Highly competitive pricing without compromising quality. Rat monoclonal antibodies for the study of gene expression and proteomics in mice and in mouse models of human diseases available.
Activity 7.21 Transcription factors
 Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
Angelo Peschiaroli, N. Valerio Dorrello, Daniele Guardavaccaro, Monica Venere, Thanos Halazonetis, Nicholas E. Sherman, and Michele Pagano
 Molecular Cell, Volume 23 Supplemental Data SCF βtrcp -Mediated Degradation of Claspin Regulates Recovery from the DNA Replication Checkpoint Response Angelo Peschiaroli, N. Valerio Dorrello, Daniele Guardavaccaro,
Molecular Cell, Volume 23 Supplemental Data SCF βtrcp -Mediated Degradation of Claspin Regulates Recovery from the DNA Replication Checkpoint Response Angelo Peschiaroli, N. Valerio Dorrello, Daniele Guardavaccaro,
Amaxa Mouse T Cell Nucleofector Kit
 Amaxa Mouse T Cell Nucleofector Kit For T cells isolated from C57BL/6 & BALB/c mice Evaluated for murine T cells isolated from C57BL/6 & BALB/c mice This protocol is designed for murine lymphocytes or
Amaxa Mouse T Cell Nucleofector Kit For T cells isolated from C57BL/6 & BALB/c mice Evaluated for murine T cells isolated from C57BL/6 & BALB/c mice This protocol is designed for murine lymphocytes or
MEF Starter Nucleofector Kit
 page 1 of 7 MEF Starter Nucleofector Kit for Mouse Embryonic Fibroblasts (MEF) MEF display significant phenotypic variations which depend on the strain, the genetic background of the mice they are isolated
page 1 of 7 MEF Starter Nucleofector Kit for Mouse Embryonic Fibroblasts (MEF) MEF display significant phenotypic variations which depend on the strain, the genetic background of the mice they are isolated
PNA BRAF Mutation Detection Kit
 - PNA BRAF Mutation Detection Kit Catalog Number KA2102 50 tests/kit Version: 01 Intended for research use only www.abnova.com Introduction and Background Intended use The PNA BRAF Mutation Detection Kit
- PNA BRAF Mutation Detection Kit Catalog Number KA2102 50 tests/kit Version: 01 Intended for research use only www.abnova.com Introduction and Background Intended use The PNA BRAF Mutation Detection Kit
Supplemental Information. McBrayer et al. Supplemental Data
 1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
Control of Gene Expression
 Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
What is the difference between basal and activated transcription?
 What is the difference between basal and activated transcription? Regulation of Transcription I. Basal vs. activated transcription for mrna genes A. General transcription factor (TF) vs. promoterspecific
What is the difference between basal and activated transcription? Regulation of Transcription I. Basal vs. activated transcription for mrna genes A. General transcription factor (TF) vs. promoterspecific
Chapter 6: Antigen-Antibody Interactions
 Chapter 6: Antigen-Antibody Interactions I. Strength of Ag-Ab interactions A. Antibody Affinity - strength of total noncovalent interactions between single Ag-binding site on an Ab and a single epitope
Chapter 6: Antigen-Antibody Interactions I. Strength of Ag-Ab interactions A. Antibody Affinity - strength of total noncovalent interactions between single Ag-binding site on an Ab and a single epitope
Gene Transcription in Prokaryotes
 Gene Transcription in Prokaryotes Operons: in prokaryotes, genes that encode protein participating in a common pathway are organized together. This group of genes, arranged in tandem, is called an OPERON.
Gene Transcription in Prokaryotes Operons: in prokaryotes, genes that encode protein participating in a common pathway are organized together. This group of genes, arranged in tandem, is called an OPERON.
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
Biological Sciences Initiative. Human Genome
 Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Product: Expression Arrest TM egfp control shrna vector
 Product: Expression Arrest TM egfp control vector Catalog #: RHS1702 Product Description The laboratory of Dr. Greg Hannon at Cold Spring Harbor Laboratory (CSHL) has created an RNAi Clone Library comprised
Product: Expression Arrest TM egfp control vector Catalog #: RHS1702 Product Description The laboratory of Dr. Greg Hannon at Cold Spring Harbor Laboratory (CSHL) has created an RNAi Clone Library comprised
SYMPOSIUM. June 15, 2013. Photonics Center Colloquium Room (906) 8 Saint Mary's St., Boston, MA 02215 Boston University
 SYMPOSIUM June 15, 2013 Photonics Center Colloquium Room (906) 8 Saint Mary's St., Boston, MA 02215 Bette Korber Los Alamos National Labs HIV evolution and the neutralizing antibody response We have recently
SYMPOSIUM June 15, 2013 Photonics Center Colloquium Room (906) 8 Saint Mary's St., Boston, MA 02215 Bette Korber Los Alamos National Labs HIV evolution and the neutralizing antibody response We have recently
Structure and Function of DNA
 Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
In developmental genomic regulatory interactions among genes, encoding transcription factors
 JOURNAL OF COMPUTATIONAL BIOLOGY Volume 20, Number 6, 2013 # Mary Ann Liebert, Inc. Pp. 419 423 DOI: 10.1089/cmb.2012.0297 Research Articles A New Software Package for Predictive Gene Regulatory Network
JOURNAL OF COMPUTATIONAL BIOLOGY Volume 20, Number 6, 2013 # Mary Ann Liebert, Inc. Pp. 419 423 DOI: 10.1089/cmb.2012.0297 Research Articles A New Software Package for Predictive Gene Regulatory Network
Control of Gene Expression
 Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Chapter 43: The Immune System
 Name Period Our students consider this chapter to be a particularly challenging and important one. Expect to work your way slowly through the first three concepts. Take particular care with Concepts 43.2
Name Period Our students consider this chapter to be a particularly challenging and important one. Expect to work your way slowly through the first three concepts. Take particular care with Concepts 43.2
THE His Tag Antibody, mab, Mouse
 THE His Tag Antibody, mab, Mouse Cat. No. A00186 Technical Manual No. TM0243 Update date 01052011 I Description.... 1 II Key Features. 2 III Storage 2 IV Applications.... 2 V Examples - ELISA..... 2 VI
THE His Tag Antibody, mab, Mouse Cat. No. A00186 Technical Manual No. TM0243 Update date 01052011 I Description.... 1 II Key Features. 2 III Storage 2 IV Applications.... 2 V Examples - ELISA..... 2 VI
Basics of Immunology
 Basics of Immunology 2 Basics of Immunology What is the immune system? Biological mechanism for identifying and destroying pathogens within a larger organism. Pathogens: agents that cause disease Bacteria,
Basics of Immunology 2 Basics of Immunology What is the immune system? Biological mechanism for identifying and destroying pathogens within a larger organism. Pathogens: agents that cause disease Bacteria,
Technical Note. Roche Applied Science. No. LC 18/2004. Assay Formats for Use in Real-Time PCR
 Roche Applied Science Technical Note No. LC 18/2004 Purpose of this Note Assay Formats for Use in Real-Time PCR The LightCycler Instrument uses several detection channels to monitor the amplification of
Roche Applied Science Technical Note No. LC 18/2004 Purpose of this Note Assay Formats for Use in Real-Time PCR The LightCycler Instrument uses several detection channels to monitor the amplification of
Complex multicellular organisms are produced by cells that switch genes on and off during development.
 Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Integrating DNA Motif Discovery and Genome-Wide Expression Analysis. Erin M. Conlon
 Integrating DNA Motif Discovery and Genome-Wide Expression Analysis Department of Mathematics and Statistics University of Massachusetts Amherst Statistics in Functional Genomics Workshop Ascona, Switzerland
Integrating DNA Motif Discovery and Genome-Wide Expression Analysis Department of Mathematics and Statistics University of Massachusetts Amherst Statistics in Functional Genomics Workshop Ascona, Switzerland
On the origin of giant multinuclear Reed-Sternberg cells and the role of CD4 T cells in Hodgkin lymphoma
 On the origin of giant multinuclear Reed-Sternberg cells and the role of CD4 T cells in Hodgkin lymphoma Uber die Entstehung von multinuklearen Reed-Sternberg Riesenzellen und die Rolle von CD4 T-Zellen
On the origin of giant multinuclear Reed-Sternberg cells and the role of CD4 T cells in Hodgkin lymphoma Uber die Entstehung von multinuklearen Reed-Sternberg Riesenzellen und die Rolle von CD4 T-Zellen
HUMORAL IMMUNE RE- SPONSES: ACTIVATION OF B CELLS AND ANTIBODIES JASON CYSTER SECTION 13
 SECTION 13 HUMORAL IMMUNE RE- SPONSES: ACTIVATION OF B CELLS AND ANTIBODIES CONTACT INFORMATION Jason Cyster, PhD (Email) READING Basic Immunology: Functions and Disorders of the Immune System. Abbas,
SECTION 13 HUMORAL IMMUNE RE- SPONSES: ACTIVATION OF B CELLS AND ANTIBODIES CONTACT INFORMATION Jason Cyster, PhD (Email) READING Basic Immunology: Functions and Disorders of the Immune System. Abbas,
JBS FUNDAMENT Thermofluor Screen
 Cat. No. CS-330 Amount 1 Kit For in vitro use only. Quality guaranteed for 3 months. Store at 4 C. Application Screen for thermal stability of proteins as a function of the FUNDAMENTAL variables ph and
Cat. No. CS-330 Amount 1 Kit For in vitro use only. Quality guaranteed for 3 months. Store at 4 C. Application Screen for thermal stability of proteins as a function of the FUNDAMENTAL variables ph and
Fighting the Battles: Conducting a Clinical Assay
 Fighting the Battles: Conducting a Clinical Assay 6 Vocabulary: In Vitro: studies in biology that are conducted using components of an organism that have been isolated from their usual biological surroundings
Fighting the Battles: Conducting a Clinical Assay 6 Vocabulary: In Vitro: studies in biology that are conducted using components of an organism that have been isolated from their usual biological surroundings
MULTIPLE MYELOMA. Dr Malkit S Riyat. MBChB, FRCPath(UK) Consultant Haematologist
 MULTIPLE MYELOMA Dr Malkit S Riyat MBChB, FRCPath(UK) Consultant Haematologist Multiple myeloma is an incurable malignancy that arises from postgerminal centre, somatically hypermutated B cells.
MULTIPLE MYELOMA Dr Malkit S Riyat MBChB, FRCPath(UK) Consultant Haematologist Multiple myeloma is an incurable malignancy that arises from postgerminal centre, somatically hypermutated B cells.
Immuno-Oncology Model Application in the Preclinical Treatment Setting
 Immuno-Oncology Model Application in the Preclinical Treatment Setting Maryland Franklin, Ph.D. Senior Director, Scientific Development [email protected] July 22, 2015 Tumor Models Conference
Immuno-Oncology Model Application in the Preclinical Treatment Setting Maryland Franklin, Ph.D. Senior Director, Scientific Development [email protected] July 22, 2015 Tumor Models Conference
ChIP TROUBLESHOOTING TIPS
 ChIP TROUBLESHOOTING TIPS Creative Diagnostics Abstract ChIP dissects the spatial and temporal dynamics of the interactions between chromatin and its associated factors CD Creative Diagnostics info@creative-
ChIP TROUBLESHOOTING TIPS Creative Diagnostics Abstract ChIP dissects the spatial and temporal dynamics of the interactions between chromatin and its associated factors CD Creative Diagnostics info@creative-
27E10: A unique monoclonal antibody against MRP8/14 (S100A8/A9, Calprotectin)
 BMA BIOMEDICALS Rheinstrasse 28-32 CH-4302 Augst (Switzerland) Phone:+41 61 811 6222 Fax: +41 61 811 6006 [email protected] www.bma.ch 27E10: A unique monoclonal antibody against MRP8/14 (S100A8/A9, Calprotectin)
BMA BIOMEDICALS Rheinstrasse 28-32 CH-4302 Augst (Switzerland) Phone:+41 61 811 6222 Fax: +41 61 811 6006 [email protected] www.bma.ch 27E10: A unique monoclonal antibody against MRP8/14 (S100A8/A9, Calprotectin)
