The Standard Graphical Notation for Biological Networks
|
|
|
- Bathsheba Little
- 9 years ago
- Views:
Transcription
1 The Standard Graphical Notation for Biological Networks Hiroaki Kitano ERATO Kitano Symbiotic Systems Project, JST and The Systems Biology Institute, Suite 6A, M31, Jingumae, Shibuya, Tokyo Japan Sony Computer Science Laboratories, Inc., Higashi-Gotanda, Shinagawa, Tokyo , Japan Solidly defined and comprehensive graphical representation of biological networks is essential for efficient and accurate representation and dissemination of biological models. Nevertheless, there has been no standard graphical representation accepted in the community. In this paper, we intent to propose a system of graphical representation of biological networks that can be a basis of de fact standard of the community. Careful analysis of biological models revealed that a set of diagrams are necessary to fully describe interactions of proteins and genes, as well as to describe processes of biological events without compromising accuracy and readability. 1. Introduction With abundant information on gene regulations, metabolic pathways, and signal transduction, there is pressing needs for standardized graphical notations to describe biological networks and processes. Most authors of biological papers use arrow-headed lines and bar-headed lines to indicate activation and inhibition, respectively. However, traditional diagrams are informal, often confusing, and much information is lost. There are several reasons for this problem. First, in traditional schematics, symbols, such as arrows, are used in various semantics from transcriptional activation/inhibition, signal transduction via protein-protein interaction, transport, etc. This causes substantial confusion in the diagrams used in the papers, thus readers must read substantial part of the text to understand what does the diagram really describes. Second, often both state transitions and relationships are described within one diagram
2 that made meaning of nodes and arcs confusing. One arrow may mean activation, but the other arrow in the same diagram may mean transition of the state or translocation of materials. Without consistent and unambiguous rules for representation, not only information is lost, but also misinformation could be disseminated. The situation is further aggravated by the lack of standard for graphical notations, so that each schematic follows different rules preventing efficient and accurate dissemination of knowledge. Recently, some researchers propose notation systems to mitigate such problems [1-5], and many research groups working on large-scale biology invent their own, but rather informal, system of describing networks and processes. Nevertheless, no standard has been formed yet, partly because proposed notations have numbers of issues to overcome, as well as lack of software tools to assist the use of such diagrams. The goal of this paper is to propose a comprehensive system of notations for visually describing biological networks and processes. A system of graphical representation shall be powerful enough to express sufficient information in clearly visible and unambiguous way, and should be supported by software tools. It is expected that the system proposed in this paper to be the basis of standard graphical notation for Systems Biology Mark-up Language (SBML) Level-III. 2. Requirements for Graphical Notation There are few criteria that the notation system should satisfy, such as; (1) Expressiveness: The notation system should be able to describe every possible relationship among genes and proteins, as well as biological processes. (2) Semantically unambiguous: Notation should be unambiguous. (3) Visually unambiguous: Each symbol should be clearly identified and cannot be mistaken with other symbols. This feature should be maintained with low-resolution displays, as well as black/white printings. (4) Extension Capability: The notation system shall be flexible enough to add new symbols and relationship in a consistent manner. This may include the use of color-coding to enhance expressiveness and readability, but information shall not be lost even with black and white displays.
3 3. What need to be represented? Two major purposes of creating such standard are; (1) to facilitate dissemination of information on the biochemical networks, and (2) to be able to efficiently create models for simulation and analysis. Thus, the diagram should be able to visually display information necessary for these purposes, and a set of software need to be developed to support efficient creation of models based on the notation. Since biological systems are highly complex, it is not feasible or desirable to graphically represent all available information, because it will simply overload the diagram so that it will be less informative. Such information may be embedded in hyperlinks created within the diagram and show up only when it is requested. Now, let us discuss what diagram shall represent. In traditional diagram, both informal drawings, as well as schematics proposed to be a standard notation there is only one representation modality. Traditional schemes generally describe processes of cascading activations or conversion of chemical species. In this type of diagram, only relationship between components that are involved in the process in focus are described, leaving numbers of interactions not being displayed (Figure 1A). On the contrary, Kohn s diagram describes relationships among components. However, it does not provide step-wise view of the specific biological processes. Kohn's notation constraints one species to appear only once in the diagram, and all combinations are described as arrowhead lines with dot in the middle that denotes compound (Figure 1B). This notation provides a rigid framework on constraint relationship among species. Major drawbacks of the Kohn's notation are (1) difficulties in understanding process of certain reactions, and (2) needs for experience to correctly draw and read compound formation diagram, that are described using arrowhead lines and dots. (A) Conventional Notation
4 (B) Kohn s Notation Figure 1: Conventional pathway description and Kohn s description Trying to create a single diagram that satisfy both characteristics is neither feasible nor desirable, because they use different semantics for symbols and driven by different needs. Our proposal in this article is that four diagrams shall compose a basic set of diagrams for describing biological networks and their basic dynamics. The four diagrams are state-transition diagram, block diagram, flow chart, and timing chart. These four diagrams are used in VLSI design process as a minimum set of graphical diagrams, a possible addition of data-flow chart, for algorithm logic design. Given that the timing chart represents results of circuit simulation, former three diagrams are considered as a basic set for representing processes in biological networks. Three diagrams are compatible in their internal model, but visualized from the different aspects and abstraction. A state-transition diagram represents process of biochemical interactions, and visualizes flow of the state transition of the molecules in focus. Block diagrams, on the other hand, represents how each components may interacts. Two diagrams have distinct semantics. In the state-transition diagram, node represents state of the matter, and arc represents transition. In the block diagram, node represents substance and arc represents relationship between two nodes in the both end of the arc. In addition, the flow chart shall represent abstract flow of the biological processes. 4. The State-Transition Diagram The state-transition diagram represents changes of states of molecules, such as proteins, during the biological processes. It consists of nodes that represent state of the proteins
5 and other biological entities, and arcs that represent transition of states. Figure 2 is an example of the state-transition diagram for a part of Yeast cell cycle process, involving MPF. Each node represents a state of MPF and other components. Figure 2: A State transition diagram describing a part of Yeast Cell Cycle process. Symbols that are used in the state transition diagram are shown in Figure 3. It should be noted that there is no symbols for activation or inhibition. Active state of enzyme is denoted by dashed oval placed surrounding the oval for the enzyme. It is useful to discuss why activation and inhibition is not on the list of symbols. The term activation and inhibition is commonly used in biological literatures, but it must be clearly defined to create a solid notation system. To activate protein X actually means to modify protein X, so that X will have kinase activity. Modifications are, in general, combination of phosphorylation, dephosophorylation, acetylation, deacetylation, etc. For example, p53 protein forms a stable complex that binds to specific DNA sequence. The formation of the complex is promoted by phosphorylation of several serine residues. The other example is M-phase Promotion Factor (MPF) that is a complex of Cdc2 and cyclin B. MPF is generally considered to be active when both Thr14 and Tyr15 residues are not phosphorylated and thr161 is phosphorylated. In
6 figure 2, wee1 changes state of MPF which is inactive, but MPF does not turn into active MPF. Wee1 only modifies two phosphorylation sites. An arrow in this transition denotes changes in the modification state of MPF, not activation. Figure 3: A symbol system for the state-transition diagram
7 Elimination of activation and inhibition symbols from the notation, ironically, makes clear semantics of the symbol system. However, we recognize that this representation is sometime inconvenient when only changes in modification state results in activation or inactivation of the protein, whereas use of activation and inhibition symbols can be conveniently used without losing accuracy of the semantics. Thus, we can define a set of additional symbols that provides shorthand way of describing activation and inhibition (Figure 4). Notice that activation is shown in double arrowhead, instead of a single arrowhead for state-transition. This extension can be used only when one wish to describe activation and inhibition of protein collateral to the process in focus and when semantics are unambiguous. It shall be used with great care so that semantics of such shorthand is sufficiently clear, and shall avoid the use of such symbols within the main process in focus. Figure 4: Additional symbols for the state transition diagram
8 Figure 5: A shorthand representation and a regular state transition diagram In shorthand notation, as well as in conventional diagrams, a cascade of activation from Rad3 to Cdc2 will be written as Figure 5 (A). In the state transition diagram, it will be written as Figure 5 (B). Both diagrams represent same process, but shorthand diagram do not display information on relationship between phosphorylation and activation. The shorthand diagram is simple and appears to be convenient, but it will be problematic due to loss of information. For example, it cannot indicate whether phosphorylation activates or inhibits protein. In addition, the vital problem is that such diagrams cannot properly represent transition of internal state of proteins. For example, MPF is phosphorylated by CAK without affecting activation/inactivation state. There is no proper way to represent such changes within the conventional diagram. Thus, interaction concerning MPF is usually represented by more explicit state-transition diagram. A shorthand notation using additional symbols is only allowed for limited use. Complex can be represented with some simplification. When two or more nodes share the boundary, it means that these nodes are in binding state to form a complex. In figure 2, cyclin B and Cdc2 share the boundary, representing binding state, and form a heterodimer complex. Alternatively, a complex can be represented as one round box as shown in figure 6. Components of a complex are represented with partitioned by plus sign, such as Cdc2+cyclin B. Residues are noted as position/protein, such as Thr14/Cdc2, so that which component of the complex residue is located can be unambiguously identified.
9 Figure 6: A simple node representation of a heterodimar complex Round symbols on the boundary of the box indicates state of residues of interest. When each state and the activation state of the protein are known, all modifications on residues can be identified. However, it is often the case whether the states of residues affect activation state or not is unclear, the symbol Unknown is used. When the state of the residue does not affect activation state, Don t Care symbol is used to indicate this fact (Figure 7). Figure 7: Residue states representation In the Alliance for Cellular Signaling consortium (AfCS: symbols proteins are shown in several different shapes, dependent on their functionalities. When such subdivisions are needed, our notation allows modifications of symbols as follows (Figure 8).
10 Figure 8: AfCS symbols and extended symbol set for the proposed notation Most metabolic pathway diagrams are straightforward to fit the state-transition diagram, because current diagram conventions are basically state-transition based. A part of the reason is that chemical species involved in metabolic pathways do not have complex allosteric control, such as phosphorylation and acetylation triggered kinase activity and dimmer formation. A set of transformation symbols will cover substantial part of the metabolic pathways (Figure 9).
11 Figure 9: Transformations Any symbols in the diagram shall have place in ontological hierarchies of biological entities and processes. Ontologically, biochemical networks can be subdivided into biochemical processes and biochemical objects. Figure 10 is a random list, not exhaustive, of that need to be described as a starting point.
12 Biochemical Processes Transcriptional activation / inhibition Translational activation / inhibition Protein activation / inhibition / degradation Phosphorylation / dephosphorylation Acetylation / deacetylation Metylation / demetylation ATP/ADP exchange GTP/GDP exchange NAD+/NADH exchange NADP+/NADPH exchange FAD/FADH2 exchange Ubiquitionation Translocation Diffusion Anchoring Biochemical Species DNA RNA Protein Small Molecules/Ion Receptors Ion channels Figure 10: A List of Processes and Objects to be represented Fig 11 is an excerpt of ontology hierarchy from Signal Transduction Ontology ( which is more detailed ontology. It should be noted that there is no activation and inhibition in the ontology.
13 Figure 11: An excerpt from Signal Transduction Ontology
14 5. The Block Diagram The block diagram provides molecular species-centered view of the interaction. This diagram is heavily based on diagram proposed by Cook[4] and Kohn[2]. Rather than emphasizing the step-wise development of the process, it tries to describe all relationships among molecular species. Figure 12 shows a simple example of the relationship diagram for Cdc2. Cdc2 is represented as a round-cornered box. All modification residues are aligned upper side of the box, and all binding sites are aligned lower side of the box. CAK phosphorylates Thr161 residue, and Wee1 phosphorylates thr14 and Tyr15 residues. Cdc25 dephospohrylates Thr14 and Tyr15 residues. There is regulatory relationship described in the round-cornered box. In this example, it indicates that; (1) phosporylation of Thr161 and binding with cyclin B triggers kinase activity of Cdc2, and (2) phosphorylation of Thr14 and Tyr15 residue inhibits effects of phosophrylation of Thr161 residue. Figure 12: A relationship diagram for Cdc2
15 Fig 13: A symbol set for the block diagram
16 Figure 13 shows a set of symbols for the relationship diagram. Most symbols are compatible with the state-transition diagram. In this symbol set, the arrow means promotion, instead of activation. It is used to indicate promote phosphorylation, promote binding with protein X, etc. Actual effect of activation, kinase activity, for example, is shown using a square box on the boundary of the round-cornered box. Figure 14: The block diagram for p53 Figure 14 shows more complex diagram for p53 protein. Extensive relationships are
17 described, but the same principle is maintained. Block diagram can be used to describe logics behind cis-regulatory region[6, 7]. Figure 15 is an example of cis-regulatory region for Endo-16 gene. Figure 15: A block diagram for endo16 cis-regulation Figure 16: Internal logic representation As it can be seen in the figure 15, there are substantial internal logics in the cis-regulation. This is also true for protein modification and bindings. Figure 16 lists nodes for representing internal logics. This logic is a hybrid of boolian logic and numeric calculation, so that both logical and quantitative changes of transcriptional regulations shall be represented. In some nodes, numbers are associated to indicate output value of the node when logical value of the node is true. For example, output of the node, which is logical AND with inputs from CY and CB1, is 1.0 when both CY and CB1 site is bounded by transcription factors. 6. Flow Chart and Timing chart The flow chart represents a series of biological events that we use to intuitively describe
18 the process under consideration (Figure 17). This is similar to the event model in Cook s notation. Each node represents intuitively labeled biological state or process at arbitrary level of abstraction. Each node, however, need to be mapped onto the node, or nodes, in the state-transition diagram in order to ground it to the molecular level. While nodes in flow charts only describes intuitive landmarks, mere breakdown of each node and arc would not be able to reconstruct detailed molecular models. Rather this shall be used to glue multiple processes each of which are based on detailed molecular models, or to visually represent state of the biological processes at the intuitive level. Timing chart is a graph that represents time-course changes of values involved in the model. Typically, this represents concentration levels and activation levels of protein involved. Figure 17: An example of flow chart 7. Availability and Supports Our intention is to form a standard for graphical representation of biological networks and processes associated. Thus, we intent to submit this proposal to be the basis of further discussions to improve the system of graphical notations that can be widely acceptable to the community. The official standard documentations and review board to for further up-date and revisions shall be created. Given the proper reference is made to this original proposal and anticipated official documentations, we welcome anyone to
19 use this symbolic representation. The standard may be a part of SBML or an independent standard linked with SBML and other standardization efforts. We are also developing software tools to support this graphical representation system. SBEdit, a graphical editor, is now being developed to support efficient creation of models using diagrams in this paper. 8. Conclusion In this paper, we proposed a graphical representation scheme for biological networks, such as gene regulatory networks and metabolic pathways. The state transition diagrams and the block diagrams are complimentary to each other, and the flow chart provides description of an abstract flow of biological processes. These diagrams are heavily based on proposals made in the past, but have solved some problems such as limitations in expression, difficulties to understand what has been described, etc. There may be addition and modification to this proposal based on the feedback from the actual use. The proposal scheme is expected to be a part of SBML level 3 definitions (or independent standard), and will be implemented as SBEdit network editor software. Acknowledgements The author wish to thank valuable comments from ERATO Project members, especially Andy Finny, Mike Hucka, Hamid Bolouri, Harbart Sauro, Mineo Morohashi, Akira Funahashi, Noriko Hiroi, and Torbjorn Nording. This research is in part supported by the Rice Genome and Simulation Project (Ministry of Agriculture, Japanese government), the international standard formation program (NEDO, Ministry of Economy, Trade, and Industries), and through a special coordinate fund for Keio University (Ministry of Education, Culture, Sports, Science, and Technology). References 1. Kohn, K., Molecular Interaction Maps as information organizers and simulation guides. Chaos, (1): p
20 2. Kohn, K.W., Molecular interaction map of the mammalian cell cycle control and DNA repair systems. Mol Biol Cell, (8): p Pirson, I., et al., The visual display of regulatory information and networks. Trends Cell Biol, (10): p Cook, D.L., J.F. Farley, and S.J. Tapscott, A basis for a visual language for describing, archiving and analyzing functional models of complex biological systems. Genome Biol, (4): p. RESEARCH Maimon, R. and S. Browning. Diagramatic Notation and Computational Structure of Gene Networks. in The Second International Conference on Systems Biology Pasadena. 6. Yuh, C.H., H. Bolouri, and E.H. Davidson, Genomic cis-regulatory logic: experimental and computational analysis of a sea urchin gene. Science, (5358): p Davidson, E.H., et al., A genomic regulatory network for development. Science, (5560): p
A Comparison of BEL V1.0 and BioPAX Level 3
 A Comparison of BEL V1.0 and BioPAX Level 3 Selventa Inc. and Robert Powers, Predictive Medicine Inc. Joanne Luciano, President, Predictive Medicine Inc. Selventa TM One Alewife Center, Cambridge, MA 02140
A Comparison of BEL V1.0 and BioPAX Level 3 Selventa Inc. and Robert Powers, Predictive Medicine Inc. Joanne Luciano, President, Predictive Medicine Inc. Selventa TM One Alewife Center, Cambridge, MA 02140
Genetomic Promototypes
 Genetomic Promototypes Mirkó Palla and Dana Pe er Department of Mechanical Engineering Clarkson University Potsdam, New York and Department of Genetics Harvard Medical School 77 Avenue Louis Pasteur Boston,
Genetomic Promototypes Mirkó Palla and Dana Pe er Department of Mechanical Engineering Clarkson University Potsdam, New York and Department of Genetics Harvard Medical School 77 Avenue Louis Pasteur Boston,
AP Biology Essential Knowledge Student Diagnostic
 AP Biology Essential Knowledge Student Diagnostic Background The Essential Knowledge statements provided in the AP Biology Curriculum Framework are scientific claims describing phenomenon occurring in
AP Biology Essential Knowledge Student Diagnostic Background The Essential Knowledge statements provided in the AP Biology Curriculum Framework are scientific claims describing phenomenon occurring in
Feed Forward Loops in Biological Systems
 Feed Forward Loops in Biological Systems Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1 of 7 Table of Contents 1 INTRODUCTION...
Feed Forward Loops in Biological Systems Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1 of 7 Table of Contents 1 INTRODUCTION...
CHAPTER 4: Enzyme Structure ENZYMES
 CHAPTER 4: ENZYMES Enzymes are biological catalysts. There are about 40,000 different enzymes in human cells, each controlling a different chemical reaction. They increase the rate of reactions by a factor
CHAPTER 4: ENZYMES Enzymes are biological catalysts. There are about 40,000 different enzymes in human cells, each controlling a different chemical reaction. They increase the rate of reactions by a factor
Chapter 8: Energy and Metabolism
 Chapter 8: Energy and Metabolism 1. Discuss energy conversions and the 1 st and 2 nd law of thermodynamics. Be sure to use the terms work, potential energy, kinetic energy, and entropy. 2. What are Joules
Chapter 8: Energy and Metabolism 1. Discuss energy conversions and the 1 st and 2 nd law of thermodynamics. Be sure to use the terms work, potential energy, kinetic energy, and entropy. 2. What are Joules
Web-Based Genomic Information Integration with Gene Ontology
 Web-Based Genomic Information Integration with Gene Ontology Kai Xu 1 IMAGEN group, National ICT Australia, Sydney, Australia, [email protected] Abstract. Despite the dramatic growth of online genomic
Web-Based Genomic Information Integration with Gene Ontology Kai Xu 1 IMAGEN group, National ICT Australia, Sydney, Australia, [email protected] Abstract. Despite the dramatic growth of online genomic
Chapter 8: An Introduction to Metabolism
 Chapter 8: An Introduction to Metabolism Name Period Concept 8.1 An organism s metabolism transforms matter and energy, subject to the laws of thermodynamics 1. Define metabolism. The totality of an organism
Chapter 8: An Introduction to Metabolism Name Period Concept 8.1 An organism s metabolism transforms matter and energy, subject to the laws of thermodynamics 1. Define metabolism. The totality of an organism
QUALITY TOOLBOX. Understanding Processes with Hierarchical Process Mapping. Robert B. Pojasek. Why Process Mapping?
 QUALITY TOOLBOX Understanding Processes with Hierarchical Process Mapping In my work, I spend a lot of time talking to people about hierarchical process mapping. It strikes me as funny that whenever I
QUALITY TOOLBOX Understanding Processes with Hierarchical Process Mapping In my work, I spend a lot of time talking to people about hierarchical process mapping. It strikes me as funny that whenever I
MULTIPLE CHOICE QUESTIONS
 MULTIPLE CHOICE QUESTIONS 1. Most components of energy conversion systems evolved very early; thus, the most fundamental aspects of energy metabolism tend to be: A. quite different among a diverse group
MULTIPLE CHOICE QUESTIONS 1. Most components of energy conversion systems evolved very early; thus, the most fundamental aspects of energy metabolism tend to be: A. quite different among a diverse group
Application of Graph-based Data Mining to Metabolic Pathways
 Application of Graph-based Data Mining to Metabolic Pathways Chang Hun You, Lawrence B. Holder, Diane J. Cook School of Electrical Engineering and Computer Science Washington State University Pullman,
Application of Graph-based Data Mining to Metabolic Pathways Chang Hun You, Lawrence B. Holder, Diane J. Cook School of Electrical Engineering and Computer Science Washington State University Pullman,
White Paper. Yeast Systems Biology - Concepts
 White Paper Yeast Systems Biology - Concepts Stefan Hohmann, Jens Nielsen, Hiroaki Kitano (see for further contributers at end of text) Göteborg, Lyngby, Tokyo, February 2004 1 Executive Summary Systems
White Paper Yeast Systems Biology - Concepts Stefan Hohmann, Jens Nielsen, Hiroaki Kitano (see for further contributers at end of text) Göteborg, Lyngby, Tokyo, February 2004 1 Executive Summary Systems
What affects an enzyme s activity? General environmental factors, such as temperature and ph. Chemicals that specifically influence the enzyme.
 CH s 8-9 Respiration & Metabolism Metabolism A catalyst is a chemical agent that speeds up a reaction without being consumed by the reaction. An enzyme is a catalytic protein. Hydrolysis of sucrose by
CH s 8-9 Respiration & Metabolism Metabolism A catalyst is a chemical agent that speeds up a reaction without being consumed by the reaction. An enzyme is a catalytic protein. Hydrolysis of sucrose by
Actions of Hormones on Target Cells Page 1. Actions of Hormones on Target Cells Page 2. Goals/ What You Need to Know Goals What You Need to Know
 Actions of Hormones on Target Cells Graphics are used with permission of: Pearson Education Inc., publishing as Benjamin Cummings (http://www.aw-bc.com) Page 1. Actions of Hormones on Target Cells Hormones
Actions of Hormones on Target Cells Graphics are used with permission of: Pearson Education Inc., publishing as Benjamin Cummings (http://www.aw-bc.com) Page 1. Actions of Hormones on Target Cells Hormones
Cellular Respiration: Practice Questions #1
 Cellular Respiration: Practice Questions #1 1. Which statement best describes one of the events taking place in the chemical reaction? A. Energy is being stored as a result of aerobic respiration. B. Fermentation
Cellular Respiration: Practice Questions #1 1. Which statement best describes one of the events taking place in the chemical reaction? A. Energy is being stored as a result of aerobic respiration. B. Fermentation
Understanding the dynamics and function of cellular networks
 Understanding the dynamics and function of cellular networks Cells are complex systems functionally diverse elements diverse interactions that form networks signal transduction-, gene regulatory-, metabolic-
Understanding the dynamics and function of cellular networks Cells are complex systems functionally diverse elements diverse interactions that form networks signal transduction-, gene regulatory-, metabolic-
Energy & Enzymes. Life requires energy for maintenance of order, growth, and reproduction. The energy living things use is chemical energy.
 Energy & Enzymes Life requires energy for maintenance of order, growth, and reproduction. The energy living things use is chemical energy. 1 Energy exists in two forms - potential and kinetic. Potential
Energy & Enzymes Life requires energy for maintenance of order, growth, and reproduction. The energy living things use is chemical energy. 1 Energy exists in two forms - potential and kinetic. Potential
Regulation of enzyme activity
 1 Regulation of enzyme activity Regulation of enzyme activity is important to coordinate the different metabolic processes. It is also important for homeostasis i.e. to maintain the internal environment
1 Regulation of enzyme activity Regulation of enzyme activity is important to coordinate the different metabolic processes. It is also important for homeostasis i.e. to maintain the internal environment
2007 7.013 Problem Set 1 KEY
 2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
1.1.2. thebiotutor. AS Biology OCR. Unit F211: Cells, Exchange & Transport. Module 1.2 Cell Membranes. Notes & Questions.
 thebiotutor AS Biology OCR Unit F211: Cells, Exchange & Transport Module 1.2 Cell Membranes Notes & Questions Andy Todd 1 Outline the roles of membranes within cells and at the surface of cells. The main
thebiotutor AS Biology OCR Unit F211: Cells, Exchange & Transport Module 1.2 Cell Membranes Notes & Questions Andy Todd 1 Outline the roles of membranes within cells and at the surface of cells. The main
1. A covalent bond between two atoms represents what kind of energy? a. Kinetic energy b. Potential energy c. Mechanical energy d.
 1. A covalent bond between two atoms represents what kind of energy? a. Kinetic energy b. Potential energy c. Mechanical energy d. Solar energy A. Answer a is incorrect. Kinetic energy is the energy of
1. A covalent bond between two atoms represents what kind of energy? a. Kinetic energy b. Potential energy c. Mechanical energy d. Solar energy A. Answer a is incorrect. Kinetic energy is the energy of
BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS
 BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS 1. The Technology Strategy sets out six areas where technological developments are required to push the frontiers of knowledge
BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS 1. The Technology Strategy sets out six areas where technological developments are required to push the frontiers of knowledge
In developmental genomic regulatory interactions among genes, encoding transcription factors
 JOURNAL OF COMPUTATIONAL BIOLOGY Volume 20, Number 6, 2013 # Mary Ann Liebert, Inc. Pp. 419 423 DOI: 10.1089/cmb.2012.0297 Research Articles A New Software Package for Predictive Gene Regulatory Network
JOURNAL OF COMPUTATIONAL BIOLOGY Volume 20, Number 6, 2013 # Mary Ann Liebert, Inc. Pp. 419 423 DOI: 10.1089/cmb.2012.0297 Research Articles A New Software Package for Predictive Gene Regulatory Network
Guide to Building Pathways in Mammal using Pathway Studio Web
 Guide to Building Pathways in Mammal using Pathway Studio Web Pathway Studio Web Document Version 1.2 1 The ResNet Mammal database contains a vast amount of information derived from Elsevier s large collection
Guide to Building Pathways in Mammal using Pathway Studio Web Pathway Studio Web Document Version 1.2 1 The ResNet Mammal database contains a vast amount of information derived from Elsevier s large collection
green B 1 ) into a single unit to model the substrate in this reaction. enzyme
 Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
Protein Protein Interaction Networks
 Functional Pattern Mining from Genome Scale Protein Protein Interaction Networks Young-Rae Cho, Ph.D. Assistant Professor Department of Computer Science Baylor University it My Definition of Bioinformatics
Functional Pattern Mining from Genome Scale Protein Protein Interaction Networks Young-Rae Cho, Ph.D. Assistant Professor Department of Computer Science Baylor University it My Definition of Bioinformatics
BPMN Business Process Modeling Notation
 BPMN (BPMN) is a graphical notation that describes the logic of steps in a business process. This notation has been especially designed to coordinate the sequence of processes and messages that flow between
BPMN (BPMN) is a graphical notation that describes the logic of steps in a business process. This notation has been especially designed to coordinate the sequence of processes and messages that flow between
agucacaaacgcu agugcuaguuua uaugcagucuua
 RNA Secondary Structure Prediction: The Co-transcriptional effect on RNA folding agucacaaacgcu agugcuaguuua uaugcagucuua By Conrad Godfrey Abstract RNA secondary structure prediction is an area of bioinformatics
RNA Secondary Structure Prediction: The Co-transcriptional effect on RNA folding agucacaaacgcu agugcuaguuua uaugcagucuua By Conrad Godfrey Abstract RNA secondary structure prediction is an area of bioinformatics
Genetics Lecture Notes 7.03 2005. Lectures 1 2
 Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Two Forms of Energy
 Module 2D - Energy and Metabolism Objective # 19 All living organisms require energy for survival. In this module we will examine some general principles about chemical reactions and energy usage within
Module 2D - Energy and Metabolism Objective # 19 All living organisms require energy for survival. In this module we will examine some general principles about chemical reactions and energy usage within
Gene Regulation -- The Lac Operon
 Gene Regulation -- The Lac Operon Specific proteins are present in different tissues and some appear only at certain times during development. All cells of a higher organism have the full set of genes:
Gene Regulation -- The Lac Operon Specific proteins are present in different tissues and some appear only at certain times during development. All cells of a higher organism have the full set of genes:
Module 3 Questions. 7. Chemotaxis is an example of signal transduction. Explain, with the use of diagrams.
 Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Introduction to BPMN
 Stephen A. White, IBM Corporation Abstract This paper is intended to provide a high-level overview and introduction to the Business Process Modeling Notation (BPMN). The context and general uses for BPMN
Stephen A. White, IBM Corporation Abstract This paper is intended to provide a high-level overview and introduction to the Business Process Modeling Notation (BPMN). The context and general uses for BPMN
CELL MEMBRANES, TRANSPORT, and COMMUNICATION. Teacher Packet
 AP * BIOLOGY CELL MEMBRANES, TRANSPORT, and COMMUNICATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production
AP * BIOLOGY CELL MEMBRANES, TRANSPORT, and COMMUNICATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production
Standards Alignment Minnesota Science Standards Alignment Matrix www.brainu.org/resources/mnstds
 Lesson Summary: Neurons transfer information by releasing neurotransmitters across the synapse or space between neurons. Students model the chemical communication between pre-synaptic and post-synaptic
Lesson Summary: Neurons transfer information by releasing neurotransmitters across the synapse or space between neurons. Students model the chemical communication between pre-synaptic and post-synaptic
Writing learning objectives
 Writing learning objectives This material was excerpted and adapted from the following web site: http://www.utexas.edu/academic/diia/assessment/iar/students/plan/objectives/ What is a learning objective?
Writing learning objectives This material was excerpted and adapted from the following web site: http://www.utexas.edu/academic/diia/assessment/iar/students/plan/objectives/ What is a learning objective?
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION. Professor Bharat Patel Office: Science 2, 2.36 Email: [email protected].
 Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: [email protected] What is Gene Expression & Gene Regulation? 1. Gene Expression
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: [email protected] What is Gene Expression & Gene Regulation? 1. Gene Expression
University of Glasgow - Programme Structure Summary C1G5-5100 MSc Bioinformatics, Polyomics and Systems Biology
 University of Glasgow - Programme Structure Summary C1G5-5100 MSc Bioinformatics, Polyomics and Systems Biology Programme Structure - the MSc outcome will require 180 credits total (full-time only) - 60
University of Glasgow - Programme Structure Summary C1G5-5100 MSc Bioinformatics, Polyomics and Systems Biology Programme Structure - the MSc outcome will require 180 credits total (full-time only) - 60
Enzymes: Practice Questions #1
 Enzymes: Practice Questions #1 1. Compound X increases the rate of the reaction below. Compound X is most likely A. an enzyme B. a lipid molecule C. an indicator D. an ADP molecule 2. The equation below
Enzymes: Practice Questions #1 1. Compound X increases the rate of the reaction below. Compound X is most likely A. an enzyme B. a lipid molecule C. an indicator D. an ADP molecule 2. The equation below
BIOLOGICAL MEMBRANES: FUNCTIONS, STRUCTURES & TRANSPORT
 BIOLOGICAL MEMBRANES: FUNCTIONS, STRUCTURES & TRANSPORT UNIVERSITY OF PNG SCHOOL OF MEDICINE AND HEALTH SCIENCES DISCIPLINE OF BIOCHEMISTRY AND MOLECULAR BIOLOGY BMLS II / B Pharm II / BDS II VJ Temple
BIOLOGICAL MEMBRANES: FUNCTIONS, STRUCTURES & TRANSPORT UNIVERSITY OF PNG SCHOOL OF MEDICINE AND HEALTH SCIENCES DISCIPLINE OF BIOCHEMISTRY AND MOLECULAR BIOLOGY BMLS II / B Pharm II / BDS II VJ Temple
Enzymes and Metabolism
 Enzymes and Metabolism Enzymes and Metabolism Metabolism: Exergonic and Endergonic Reactions Chemical Reactions: Activation Every chemical reaction involves bond breaking and bond forming A chemical reaction
Enzymes and Metabolism Enzymes and Metabolism Metabolism: Exergonic and Endergonic Reactions Chemical Reactions: Activation Every chemical reaction involves bond breaking and bond forming A chemical reaction
Chemical reaction (slow): Enzyme-catalyzed reaction (much faster):
 1 Enzymes Introduction Enzymes are Biological Catalysts Recall that a catalyst is an agent which speeds up a chemical reaction without actually being consumed or changed by the reaction. Enzymes are proteins
1 Enzymes Introduction Enzymes are Biological Catalysts Recall that a catalyst is an agent which speeds up a chemical reaction without actually being consumed or changed by the reaction. Enzymes are proteins
(Refer Slide Time: 2:03)
 Control Engineering Prof. Madan Gopal Department of Electrical Engineering Indian Institute of Technology, Delhi Lecture - 11 Models of Industrial Control Devices and Systems (Contd.) Last time we were
Control Engineering Prof. Madan Gopal Department of Electrical Engineering Indian Institute of Technology, Delhi Lecture - 11 Models of Industrial Control Devices and Systems (Contd.) Last time we were
A CONTENT STANDARD IS NOT MET UNLESS APPLICABLE CHARACTERISTICS OF SCIENCE ARE ALSO ADDRESSED AT THE SAME TIME.
 Biology Curriculum The Georgia Performance Standards are designed to provide students with the knowledge and skills for proficiency in science. The Project 2061 s Benchmarks for Science Literacy is used
Biology Curriculum The Georgia Performance Standards are designed to provide students with the knowledge and skills for proficiency in science. The Project 2061 s Benchmarks for Science Literacy is used
Chapter 3. Data Flow Diagrams
 Chapter 3. Data Flow Diagrams Table of Contents Objectives... 1 Introduction to Data Flow Diagrams... 2 What are Data Flow Diagrams?... 2 An example Data Flow Diagram... 2 The benefits of Data Flow Diagrams...
Chapter 3. Data Flow Diagrams Table of Contents Objectives... 1 Introduction to Data Flow Diagrams... 2 What are Data Flow Diagrams?... 2 An example Data Flow Diagram... 2 The benefits of Data Flow Diagrams...
Figure 5. Energy of activation with and without an enzyme.
 Biology 20 Laboratory ENZYMES & CELLULAR RESPIRATION OBJECTIVE To be able to list the general characteristics of enzymes. To study the effects of enzymes on the rate of chemical reactions. To demonstrate
Biology 20 Laboratory ENZYMES & CELLULAR RESPIRATION OBJECTIVE To be able to list the general characteristics of enzymes. To study the effects of enzymes on the rate of chemical reactions. To demonstrate
PROG0101 Fundamentals of Programming PROG0101 FUNDAMENTALS OF PROGRAMMING. Chapter 3 Algorithms
 PROG0101 FUNDAMENTALS OF PROGRAMMING Chapter 3 1 Introduction to A sequence of instructions. A procedure or formula for solving a problem. It was created mathematician, Mohammed ibn-musa al-khwarizmi.
PROG0101 FUNDAMENTALS OF PROGRAMMING Chapter 3 1 Introduction to A sequence of instructions. A procedure or formula for solving a problem. It was created mathematician, Mohammed ibn-musa al-khwarizmi.
Algorithm & Flowchart & Pseudo code. Staff Incharge: S.Sasirekha
 Algorithm & Flowchart & Pseudo code Staff Incharge: S.Sasirekha Computer Programming and Languages Computers work on a set of instructions called computer program, which clearly specify the ways to carry
Algorithm & Flowchart & Pseudo code Staff Incharge: S.Sasirekha Computer Programming and Languages Computers work on a set of instructions called computer program, which clearly specify the ways to carry
Histone modifications. and ChIP. G. Valle - Università di Padova
 Histone modifications and ChIP Histone acetylation It has been shown that the acetylation of lysines is highly dynamic and regulated by the opposing action of two families of enzymes, histone acetyltransferases
Histone modifications and ChIP Histone acetylation It has been shown that the acetylation of lysines is highly dynamic and regulated by the opposing action of two families of enzymes, histone acetyltransferases
Lecture 6. Regulation of Protein Synthesis at the Translational Level
 Regulation of Protein Synthesis (6.1) Lecture 6 Regulation of Protein Synthesis at the Translational Level Comparison of EF-Tu-GDP and EF-Tu-GTP conformations EF-Tu-GDP EF-Tu-GTP Next: Comparison of GDP
Regulation of Protein Synthesis (6.1) Lecture 6 Regulation of Protein Synthesis at the Translational Level Comparison of EF-Tu-GDP and EF-Tu-GTP conformations EF-Tu-GDP EF-Tu-GTP Next: Comparison of GDP
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Quick Guide Business Process Modeling Notation (BPMN)
 Quick Guide Business Process Modeling Notation (BPMN) IDM Technical Team January 2007 Quick Guide: BPMN 2 of 14 The scope of this document is to provide a quick guide to the concepts and usage of the Business
Quick Guide Business Process Modeling Notation (BPMN) IDM Technical Team January 2007 Quick Guide: BPMN 2 of 14 The scope of this document is to provide a quick guide to the concepts and usage of the Business
Algorithms in Computational Biology (236522) spring 2007 Lecture #1
 Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
Chapter 4 Software Lifecycle and Performance Analysis
 Chapter 4 Software Lifecycle and Performance Analysis This chapter is aimed at illustrating performance modeling and analysis issues within the software lifecycle. After having introduced software and
Chapter 4 Software Lifecycle and Performance Analysis This chapter is aimed at illustrating performance modeling and analysis issues within the software lifecycle. After having introduced software and
Biology: Foundation Edition Miller/Levine 2010
 A Correlation of Biology: Foundation Edition Miller/Levine 2010 to the IDAHO CONTENT STANDARDS Science - Biology Grades 9-10 INTRODUCTION This document demonstrates how Prentice Hall s Biology: Foundation
A Correlation of Biology: Foundation Edition Miller/Levine 2010 to the IDAHO CONTENT STANDARDS Science - Biology Grades 9-10 INTRODUCTION This document demonstrates how Prentice Hall s Biology: Foundation
Diabetes and Insulin Signaling
 Diabetes and Insulin Signaling NATIONAL CENTER FOR CASE STUDY TEACHING IN SCIENCE by Kristy J. Wilson School of Mathematics and Sciences Marian University, Indianapolis, IN Part I Research Orientation
Diabetes and Insulin Signaling NATIONAL CENTER FOR CASE STUDY TEACHING IN SCIENCE by Kristy J. Wilson School of Mathematics and Sciences Marian University, Indianapolis, IN Part I Research Orientation
Writing Reports BJECTIVES ONTENTS. By the end of this section you should be able to :
 Writing Reports By the end of this section you should be able to : O BJECTIVES Understand the purposes of a report Plan a report Understand the structure of a report Collect information for your report
Writing Reports By the end of this section you should be able to : O BJECTIVES Understand the purposes of a report Plan a report Understand the structure of a report Collect information for your report
Introduction to Chemistry. Course Description
 CHM 1025 & CHM 1025L Introduction to Chemistry Course Description CHM 1025 Introduction to Chemistry (3) P CHM 1025L Introduction to Chemistry Laboratory (1) P This introductory course is intended to introduce
CHM 1025 & CHM 1025L Introduction to Chemistry Course Description CHM 1025 Introduction to Chemistry (3) P CHM 1025L Introduction to Chemistry Laboratory (1) P This introductory course is intended to introduce
Tools & Techniques for Process Improvement
 Tools & Techniques for Process Improvement Understanding processes so that they can be improved by means of a systematic approach requires the knowledge of a simple kit of ols or techniques. The effective
Tools & Techniques for Process Improvement Understanding processes so that they can be improved by means of a systematic approach requires the knowledge of a simple kit of ols or techniques. The effective
Improved Software Testing Using McCabe IQ Coverage Analysis
 White Paper Table of Contents Introduction...1 What is Coverage Analysis?...2 The McCabe IQ Approach to Coverage Analysis...3 The Importance of Coverage Analysis...4 Where Coverage Analysis Fits into your
White Paper Table of Contents Introduction...1 What is Coverage Analysis?...2 The McCabe IQ Approach to Coverage Analysis...3 The Importance of Coverage Analysis...4 Where Coverage Analysis Fits into your
BIOLOGY 101 COURSE SYLLABUS FOR FALL 2015
 BIOLOGY 101 COURSE SYLLABUS FOR FALL 2015 Course Description Instructor Biology 101 is the first of a two-semester introductory course sequence designed primarily for science majors. It covers some central
BIOLOGY 101 COURSE SYLLABUS FOR FALL 2015 Course Description Instructor Biology 101 is the first of a two-semester introductory course sequence designed primarily for science majors. It covers some central
[1] http://en.wikipedia.org/wiki/first-mover_advantage [2] http://www.acunote.com
![[1] http://en.wikipedia.org/wiki/first-mover_advantage [2] http://www.acunote.com [1] http://en.wikipedia.org/wiki/first-mover_advantage [2] http://www.acunote.com](/thumbs/24/2883514.jpg) -Gene Sher Software Development Processes: Those in engineering and science will sooner or later either be members of teams solving some large project, or be managing teams solving some large project.
-Gene Sher Software Development Processes: Those in engineering and science will sooner or later either be members of teams solving some large project, or be managing teams solving some large project.
AP BIOLOGY 2008 SCORING GUIDELINES
 AP BIOLOGY 2008 SCORING GUIDELINES Question 1 1. The physical structure of a protein often reflects and affects its function. (a) Describe THREE types of chemical bonds/interactions found in proteins.
AP BIOLOGY 2008 SCORING GUIDELINES Question 1 1. The physical structure of a protein often reflects and affects its function. (a) Describe THREE types of chemical bonds/interactions found in proteins.
Control of Gene Expression
 Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
2 SYSTEM DESCRIPTION TECHNIQUES
 2 SYSTEM DESCRIPTION TECHNIQUES 2.1 INTRODUCTION Graphical representation of any process is always better and more meaningful than its representation in words. Moreover, it is very difficult to arrange
2 SYSTEM DESCRIPTION TECHNIQUES 2.1 INTRODUCTION Graphical representation of any process is always better and more meaningful than its representation in words. Moreover, it is very difficult to arrange
Keystone Review Practice Test Module A Cells and Cell Processes. 1. Which characteristic is shared by all prokaryotes and eukaryotes?
 Keystone Review Practice Test Module A Cells and Cell Processes 1. Which characteristic is shared by all prokaryotes and eukaryotes? a. Ability to store hereditary information b. Use of organelles to control
Keystone Review Practice Test Module A Cells and Cell Processes 1. Which characteristic is shared by all prokaryotes and eukaryotes? a. Ability to store hereditary information b. Use of organelles to control
Six major functions of membrane proteins: Transport Enzymatic activity
 CH 7 Membranes Cellular Membranes Phospholipids are the most abundant lipid in the plasma membrane. Phospholipids are amphipathic molecules, containing hydrophobic and hydrophilic regions. The fluid mosaic
CH 7 Membranes Cellular Membranes Phospholipids are the most abundant lipid in the plasma membrane. Phospholipids are amphipathic molecules, containing hydrophobic and hydrophilic regions. The fluid mosaic
The Making of the Fittest: Evolving Switches, Evolving Bodies
 OVERVIEW MODELING THE REGULATORY SWITCHES OF THE PITX1 GENE IN STICKLEBACK FISH This hands-on activity supports the short film, The Making of the Fittest:, and aims to help students understand eukaryotic
OVERVIEW MODELING THE REGULATORY SWITCHES OF THE PITX1 GENE IN STICKLEBACK FISH This hands-on activity supports the short film, The Making of the Fittest:, and aims to help students understand eukaryotic
Transcription and Translation of DNA
 Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
7.2 Cell Structure. Lesson Objectives. Lesson Summary. Cell Organization Eukaryotic cells contain a nucleus and many specialized structures.
 7.2 Cell Structure Lesson Objectives Describe the structure and function of the cell nucleus. Describe the role of vacuoles, lysosomes, and the cytoskeleton. Identify the role of ribosomes, endoplasmic
7.2 Cell Structure Lesson Objectives Describe the structure and function of the cell nucleus. Describe the role of vacuoles, lysosomes, and the cytoskeleton. Identify the role of ribosomes, endoplasmic
S1 Text. Modeling deterministic single-cell microrna-p53-mdm2 network Figure 2 Figure 2
 S1 Text. Modeling deterministic single-cell microrna-p53-mdm2 network The schematic diagram of the microrna-p53-mdm2 oscillator is illustrated in Figure 2. The interaction scheme among the mrnas and the
S1 Text. Modeling deterministic single-cell microrna-p53-mdm2 network The schematic diagram of the microrna-p53-mdm2 oscillator is illustrated in Figure 2. The interaction scheme among the mrnas and the
Activity 7.21 Transcription factors
 Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
Endocrine System: Practice Questions #1
 Endocrine System: Practice Questions #1 1. Removing part of gland D would most likely result in A. a decrease in the secretions of other glands B. a decrease in the blood calcium level C. an increase in
Endocrine System: Practice Questions #1 1. Removing part of gland D would most likely result in A. a decrease in the secretions of other glands B. a decrease in the blood calcium level C. an increase in
Using UML Part Two Behavioral Modeling Diagrams
 UML Tutorials Using UML Part Two Behavioral Modeling Diagrams by Sparx Systems All material Sparx Systems 2007 Sparx Systems 2007 Page 1 Trademarks Object Management Group, OMG, Unified Modeling Language,
UML Tutorials Using UML Part Two Behavioral Modeling Diagrams by Sparx Systems All material Sparx Systems 2007 Sparx Systems 2007 Page 1 Trademarks Object Management Group, OMG, Unified Modeling Language,
Biological Neurons and Neural Networks, Artificial Neurons
 Biological Neurons and Neural Networks, Artificial Neurons Neural Computation : Lecture 2 John A. Bullinaria, 2015 1. Organization of the Nervous System and Brain 2. Brains versus Computers: Some Numbers
Biological Neurons and Neural Networks, Artificial Neurons Neural Computation : Lecture 2 John A. Bullinaria, 2015 1. Organization of the Nervous System and Brain 2. Brains versus Computers: Some Numbers
Replication Study Guide
 Replication Study Guide This study guide is a written version of the material you have seen presented in the replication unit. Self-reproduction is a function of life that human-engineered systems have
Replication Study Guide This study guide is a written version of the material you have seen presented in the replication unit. Self-reproduction is a function of life that human-engineered systems have
AP Chemistry A. Allan Chapter 1 Notes - Chemical Foundations
 AP Chemistry A. Allan Chapter 1 Notes - Chemical Foundations 1.1 Chemistry: An Overview A. Reaction of hydrogen and oxygen 1. Two molecules of hydrogen react with one molecule of oxygen to form two molecules
AP Chemistry A. Allan Chapter 1 Notes - Chemical Foundations 1.1 Chemistry: An Overview A. Reaction of hydrogen and oxygen 1. Two molecules of hydrogen react with one molecule of oxygen to form two molecules
The Somatic Cell Cycle
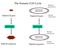 The Somatic Cell Cycle Maternal chromosome Diploid Zygote Diploid Zygote Paternal chromosome MITOSIS MITOSIS Maternal chromosome Diploid organism Diploid organism Paternal chromosome Int terpha ase The
The Somatic Cell Cycle Maternal chromosome Diploid Zygote Diploid Zygote Paternal chromosome MITOSIS MITOSIS Maternal chromosome Diploid organism Diploid organism Paternal chromosome Int terpha ase The
Chapter 6 An Overview of Organic Reactions
 John E. McMurry www.cengage.com/chemistry/mcmurry Chapter 6 An Overview of Organic Reactions Why this chapter? To understand organic and/or biochemistry, it is necessary to know: -What occurs -Why and
John E. McMurry www.cengage.com/chemistry/mcmurry Chapter 6 An Overview of Organic Reactions Why this chapter? To understand organic and/or biochemistry, it is necessary to know: -What occurs -Why and
Translation Study Guide
 Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Data, Measurements, Features
 Data, Measurements, Features Middle East Technical University Dep. of Computer Engineering 2009 compiled by V. Atalay What do you think of when someone says Data? We might abstract the idea that data are
Data, Measurements, Features Middle East Technical University Dep. of Computer Engineering 2009 compiled by V. Atalay What do you think of when someone says Data? We might abstract the idea that data are
Basic Scientific Principles that All Students Should Know Upon Entering Medical and Dental School at McGill
 Fundamentals of Medicine and Dentistry Basic Scientific Principles that All Students Should Know Upon Entering Medical and Dental School at McGill Students entering medical and dental training come from
Fundamentals of Medicine and Dentistry Basic Scientific Principles that All Students Should Know Upon Entering Medical and Dental School at McGill Students entering medical and dental training come from
Elementary circuits. Resources and methods for learning about these subjects (list a few here, in preparation for your research):
 Elementary circuits This worksheet and all related files are licensed under the Creative Commons Attribution License, version 1.0. To view a copy of this license, visit http://creativecommons.org/licenses/by/1.0/,
Elementary circuits This worksheet and all related files are licensed under the Creative Commons Attribution License, version 1.0. To view a copy of this license, visit http://creativecommons.org/licenses/by/1.0/,
NOVEL GENOME-SCALE CORRELATION BETWEEN DNA REPLICATION AND RNA TRANSCRIPTION DURING THE CELL CYCLE IN YEAST IS PREDICTED BY DATA-DRIVEN MODELS
 NOVEL GENOME-SCALE CORRELATION BETWEEN DNA REPLICATION AND RNA TRANSCRIPTION DURING THE CELL CYCLE IN YEAST IS PREDICTED BY DATA-DRIVEN MODELS Orly Alter (a) *, Gene H. Golub (b), Patrick O. Brown (c)
NOVEL GENOME-SCALE CORRELATION BETWEEN DNA REPLICATION AND RNA TRANSCRIPTION DURING THE CELL CYCLE IN YEAST IS PREDICTED BY DATA-DRIVEN MODELS Orly Alter (a) *, Gene H. Golub (b), Patrick O. Brown (c)
INSTRUCTIONS I. SHORT INSTRUCTIONS The way of paper submission is online (www.folia-biologica.org/submission). Please remember, use commands of the
 INSTRUCTIONS I. SHORT INSTRUCTIONS The way of paper submission is online (www.folia-biologica.org/submission). Please remember, use commands of the system for navigation. One author, who will serve as
INSTRUCTIONS I. SHORT INSTRUCTIONS The way of paper submission is online (www.folia-biologica.org/submission). Please remember, use commands of the system for navigation. One author, who will serve as
CSC 2427: Algorithms for Molecular Biology Spring 2006. Lecture 16 March 10
 CSC 2427: Algorithms for Molecular Biology Spring 2006 Lecture 16 March 10 Lecturer: Michael Brudno Scribe: Jim Huang 16.1 Overview of proteins Proteins are long chains of amino acids (AA) which are produced
CSC 2427: Algorithms for Molecular Biology Spring 2006 Lecture 16 March 10 Lecturer: Michael Brudno Scribe: Jim Huang 16.1 Overview of proteins Proteins are long chains of amino acids (AA) which are produced
1. Introduction Gene regulation Genomics and genome analyses Hidden markov model (HMM)
 1. Introduction Gene regulation Genomics and genome analyses Hidden markov model (HMM) 2. Gene regulation tools and methods Regulatory sequences and motif discovery TF binding sites, microrna target prediction
1. Introduction Gene regulation Genomics and genome analyses Hidden markov model (HMM) 2. Gene regulation tools and methods Regulatory sequences and motif discovery TF binding sites, microrna target prediction
Pesticide Analysis by Mass Spectrometry
 Pesticide Analysis by Mass Spectrometry Purpose: The purpose of this assignment is to introduce concepts of mass spectrometry (MS) as they pertain to the qualitative and quantitative analysis of organochlorine
Pesticide Analysis by Mass Spectrometry Purpose: The purpose of this assignment is to introduce concepts of mass spectrometry (MS) as they pertain to the qualitative and quantitative analysis of organochlorine
Enzymes. A. a lipid B. a protein C. a carbohydrate D. a mineral
 Enzymes 1. All cells in multicellular organisms contain thousands of different kinds of enzymes that are specialized to catalyze different chemical reactions. Given this information, which of the following
Enzymes 1. All cells in multicellular organisms contain thousands of different kinds of enzymes that are specialized to catalyze different chemical reactions. Given this information, which of the following
Catalysis by Enzymes. Enzyme A protein that acts as a catalyst for a biochemical reaction.
 Catalysis by Enzymes Enzyme A protein that acts as a catalyst for a biochemical reaction. Enzymatic Reaction Specificity Enzyme Cofactors Many enzymes are conjugated proteins that require nonprotein portions
Catalysis by Enzymes Enzyme A protein that acts as a catalyst for a biochemical reaction. Enzymatic Reaction Specificity Enzyme Cofactors Many enzymes are conjugated proteins that require nonprotein portions
Exercise with Gene Ontology - Cytoscape - BiNGO
 Exercise with Gene Ontology - Cytoscape - BiNGO This practical has material extracted from http://www.cbs.dtu.dk/chipcourse/exercises/ex_go/goexercise11.php In this exercise we will analyze microarray
Exercise with Gene Ontology - Cytoscape - BiNGO This practical has material extracted from http://www.cbs.dtu.dk/chipcourse/exercises/ex_go/goexercise11.php In this exercise we will analyze microarray
Cellular Respiration Worksheet 1. 1. What are the 3 phases of the cellular respiration process? Glycolysis, Krebs Cycle, Electron Transport Chain.
 Cellular Respiration Worksheet 1 1. What are the 3 phases of the cellular respiration process? Glycolysis, Krebs Cycle, Electron Transport Chain. 2. Where in the cell does the glycolysis part of cellular
Cellular Respiration Worksheet 1 1. What are the 3 phases of the cellular respiration process? Glycolysis, Krebs Cycle, Electron Transport Chain. 2. Where in the cell does the glycolysis part of cellular
GENE REGULATION. Teacher Packet
 AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
Name Date Period. Keystone Review Enzymes
 Name Date Period Keystone Review Enzymes 1. In order for cells to function properly, the enzymes that they contain must also function properly. What can be inferred using the above information? A. Cells
Name Date Period Keystone Review Enzymes 1. In order for cells to function properly, the enzymes that they contain must also function properly. What can be inferred using the above information? A. Cells
Cells, tissues and organs
 Chapter 8: Cells, tissues and organs Cells: building blocks of life Living things are made of cells. Many of the chemical reactions that keep organisms alive (metabolic functions) take place in cells.
Chapter 8: Cells, tissues and organs Cells: building blocks of life Living things are made of cells. Many of the chemical reactions that keep organisms alive (metabolic functions) take place in cells.
