A Brief Introduction on DNase-Seq Data Aanalysis
|
|
|
- Lewis Booker
- 10 years ago
- Views:
Transcription
1 A Brief Introduction on DNase-Seq Data Aanalysis Hashem Koohy, Thomas Down, Mikhail Spivakov and Tim Hubbard Spivakov s and Fraser s Lab September 13, 2014
2 1
3 Introduction DNaseI is an enzyme which cuts duplex DNA at a rate that depends strongly its chromatin environment. In Combination with HTS technology, it can be used to infer genome wide landscape of open chromatin regions. Using this technique, systematic identification of hundreds of thousands of DNaseI Hypersensitive Sites (DHS) per cell type has been possible(encode). In what follows, we provide a brief introduction about this technique with a particular emphasis on peak calling step.
4 Currently Existing DNase-Seq Protocoles Double Hit Protocole Developed in John Stam Lab in University of Washington, and has been used greatly for detection of DHS in ENCODE project. Single Hit Protocol Developed in Greg Crawford Lab in Duke University. It has been used for detection DHS in ENCODE. This protocol is also in great use by some other researchers world-wide. ATAC-Seq Developed in Greenland Lab in Stanford University. This is a very new protocol (published 2013) and has been reported to be very fast and very efficient.
5 Two Commonly Used Protocols End Capture (Duke) and Double Hit (UW) Protocol Ligate Biotinylated Linker1 Mmel Digested Ligate Linker2 PCR Amplification Sequencing End Capture Protocol: Greg Crawford Lab, Duke Double Hit Protocol: John Stam Lab, UW
6 ChIP-Seq vs DNase-Seq Note that DNase HS is different from its sister DNase Footprinting ChIP-Seq: Nature Reviews, Peter J. Park, 2009 DNase HS: Duke Protocol
7 TF ChIP-Seq vs DNase-Seq Some key differences between TF ChIP-Seq and DNase-Seq: In ChIP-Seq data, a protein is usually in bound or unbound position, whereas DNaseI shows a more generic behaviour, representing the openness of the chromatin to any regulatory feature; DNase HS are strand-independent and therefore no need to shift size or tag extension; DNase HS data sometimes shows less enrichment over wider regions (a kind of Mixed-Source).
8 DNase-Seq Data Analysis Similar to any other ChIP-Seq data, there are various steps: Sequencing, Mapping and Quality Controls Sequencing is getting cheaper, providing us with more data! Mapping possibly is still the most computationally expensive part. Peak Calling Gauging the statistical significance of reads enrichment which is generally known as Peak Calling is very central to ChIP-Seq data analysis. Post Peak Calling Analysis Different directions and purposes, including differential binding analysis, motif discovery, detection of regulatory regions, Genome segmentation and so on
9 Context-Specificity of ChIP-Seq Data Different protein classes have distinct mode of interactions: Point-Source These factors and chromatin marks are localised specifically and have high signal-to-noise ration Broad-Source These factors are associated with wide genomic domains, generating broad but more noisy signals; e.g. H3K9me3, H3K36me3 Mixed-Source These factors show a point-source style signal at some regions whereas more broader in other regions e.g. RNA Pol II
10 DNaseI Peak Calling Algorithms Note that, a peak caller is an algorithms and/or a model that is applied to gauge the statistical significance of enrichment of the short read tags. Context specificity of ChIP data among some other reasons has give rise to numerous peak callers. MACS and F-SEQ are considered among the best reported peak callers for the DNaseI-Seq. Throughout this tutorial, We will try to illustrate how to use these two peak callers for detection statistically significant open chromatin regions. We also show you very briefly how to visualise your data as the very fist step of validating your results.
11 F-Seq An histogram-based (number of tags per bin) approach is, possibly, the most naivest for gauging the enrichment of short read tags; However, it suffers from some problems including boundary effects and selection of bin width; To overcome, F-Seq suggested in which a Kernel Density Estimator(with mean 0 and variance 1) is applied to obtain the distribution of reads: p(x) = 1 i=n K( x x i ) nb b i=1 F-Seq has been implemented in Java, easy to use, though, doesn t support some commonly used file formats.
12 MACS The most used peak callers for ChIP-Seq data; It has been reviewed and benchmarked in different studies; At the time of development, the emphasis was on handling shift size and local biases from sequencability and mappability; A Poisson model is employed for identification of statistically signicant enriched regions; MACS has been implemented in python and is relatively fast. It is user friendly and fairly well-supported.
The Segway annotation of ENCODE data
 The Segway annotation of ENCODE data Michael M. Hoffman Department of Genome Sciences University of Washington Overview 1. ENCODE Project 2. Semi-automated genomic annotation 3. Chromatin 4. RNA-seq Functional
The Segway annotation of ENCODE data Michael M. Hoffman Department of Genome Sciences University of Washington Overview 1. ENCODE Project 2. Semi-automated genomic annotation 3. Chromatin 4. RNA-seq Functional
Nebula A web-server for advanced ChIP-seq data analysis. Tutorial. by Valentina BOEVA
 Nebula A web-server for advanced ChIP-seq data analysis Tutorial by Valentina BOEVA Content Upload data to the history pp. 5-6 Check read number and sequencing quality pp. 7-9 Visualize.BAM files in UCSC
Nebula A web-server for advanced ChIP-seq data analysis Tutorial by Valentina BOEVA Content Upload data to the history pp. 5-6 Check read number and sequencing quality pp. 7-9 Visualize.BAM files in UCSC
Comparing Methods for Identifying Transcription Factor Target Genes
 Comparing Methods for Identifying Transcription Factor Target Genes Alena van Bömmel (R 3.3.73) Matthew Huska (R 3.3.18) Max Planck Institute for Molecular Genetics Folie 1 Transcriptional Regulation TF
Comparing Methods for Identifying Transcription Factor Target Genes Alena van Bömmel (R 3.3.73) Matthew Huska (R 3.3.18) Max Planck Institute for Molecular Genetics Folie 1 Transcriptional Regulation TF
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How Sequencing Experiments Fail
 How Sequencing Experiments Fail v1.0 Simon Andrews [email protected] Classes of Failure Technical Tracking Library Contamination Biological Interpretation Something went wrong with a machine
How Sequencing Experiments Fail v1.0 Simon Andrews [email protected] Classes of Failure Technical Tracking Library Contamination Biological Interpretation Something went wrong with a machine
Current Motif Discovery Tools and their Limitations
 Current Motif Discovery Tools and their Limitations Philipp Bucher SIB / CIG Workshop 3 October 2006 Trendy Concepts and Hypotheses Transcription regulatory elements act in a context-dependent manner.
Current Motif Discovery Tools and their Limitations Philipp Bucher SIB / CIG Workshop 3 October 2006 Trendy Concepts and Hypotheses Transcription regulatory elements act in a context-dependent manner.
GMQL Functional Comparison with BEDTools and BEDOPS
 GMQL Functional Comparison with BEDTools and BEDOPS Genomic Computing Group Dipartimento di Elettronica, Informazione e Bioingegneria Politecnico di Milano This document presents a functional comparison
GMQL Functional Comparison with BEDTools and BEDOPS Genomic Computing Group Dipartimento di Elettronica, Informazione e Bioingegneria Politecnico di Milano This document presents a functional comparison
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals
 Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Genome-wide measurements of protein-dna interaction by chromatin immunoprecipitation
 Genome-wide measurements of protein-dna interaction by chromatin immunoprecipitation D. Puthier. laboratoire INSERM, Aix-Marseille Université, TAGC/INSERM U928, Parc Scientifique de Luminy case 928 Outline
Genome-wide measurements of protein-dna interaction by chromatin immunoprecipitation D. Puthier. laboratoire INSERM, Aix-Marseille Université, TAGC/INSERM U928, Parc Scientifique de Luminy case 928 Outline
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS)
 Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
Interaktionen von Nukleinsäuren und Proteinen
 Sonja Prohaska Computational EvoDevo Universitaet Leipzig June 9, 2015 DNA is never naked in a cell DNA is usually in association with proteins. In all domains of life there are small, basic chromosomal
Sonja Prohaska Computational EvoDevo Universitaet Leipzig June 9, 2015 DNA is never naked in a cell DNA is usually in association with proteins. In all domains of life there are small, basic chromosomal
Shouguo Gao Ph. D Department of Physics and Comprehensive Diabetes Center
 Computational Challenges in Storage, Analysis and Interpretation of Next-Generation Sequencing Data Shouguo Gao Ph. D Department of Physics and Comprehensive Diabetes Center Next Generation Sequencing
Computational Challenges in Storage, Analysis and Interpretation of Next-Generation Sequencing Data Shouguo Gao Ph. D Department of Physics and Comprehensive Diabetes Center Next Generation Sequencing
Lectures 1 and 8 15. February 7, 2013. Genomics 2012: Repetitorium. Peter N Robinson. VL1: Next- Generation Sequencing. VL8 9: Variant Calling
 Lectures 1 and 8 15 February 7, 2013 This is a review of the material from lectures 1 and 8 14. Note that the material from lecture 15 is not relevant for the final exam. Today we will go over the material
Lectures 1 and 8 15 February 7, 2013 This is a review of the material from lectures 1 and 8 14. Note that the material from lecture 15 is not relevant for the final exam. Today we will go over the material
PrimePCR Assay Validation Report
 Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID papillary renal cell carcinoma (translocation-associated) PRCC Human This gene
Gene Information Gene Name Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID papillary renal cell carcinoma (translocation-associated) PRCC Human This gene
PrimePCR Assay Validation Report
 Gene Information Gene Name sorbin and SH3 domain containing 2 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SORBS2 Human Arg and c-abl represent the mammalian
Gene Information Gene Name sorbin and SH3 domain containing 2 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SORBS2 Human Arg and c-abl represent the mammalian
Analysis of gene expression data. Ulf Leser and Philippe Thomas
 Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
Outline. interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work?
 Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Molecular Genetics: Challenges for Statistical Practice. J.K. Lindsey
 Molecular Genetics: Challenges for Statistical Practice J.K. Lindsey 1. What is a Microarray? 2. Design Questions 3. Modelling Questions 4. Longitudinal Data 5. Conclusions 1. What is a microarray? A microarray
Molecular Genetics: Challenges for Statistical Practice J.K. Lindsey 1. What is a Microarray? 2. Design Questions 3. Modelling Questions 4. Longitudinal Data 5. Conclusions 1. What is a microarray? A microarray
RNA & Protein Synthesis
 RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
Real-time quantitative RT -PCR (Taqman)
 Real-time quantitative RT -PCR (Taqman) Author: SC, Patti Lab, 3/03 This is performed as a 2-step reaction: 1. cdna synthesis from DNase 1-treated total RNA 2. PCR 1. cdna synthesis (Advantage RT-for-PCR
Real-time quantitative RT -PCR (Taqman) Author: SC, Patti Lab, 3/03 This is performed as a 2-step reaction: 1. cdna synthesis from DNase 1-treated total RNA 2. PCR 1. cdna synthesis (Advantage RT-for-PCR
Chapter 2. imapper: A web server for the automated analysis and mapping of insertional mutagenesis sequence data against Ensembl genomes
 Chapter 2. imapper: A web server for the automated analysis and mapping of insertional mutagenesis sequence data against Ensembl genomes 2.1 Introduction Large-scale insertional mutagenesis screening in
Chapter 2. imapper: A web server for the automated analysis and mapping of insertional mutagenesis sequence data against Ensembl genomes 2.1 Introduction Large-scale insertional mutagenesis screening in
RETRIEVING SEQUENCE INFORMATION. Nucleotide sequence databases. Database search. Sequence alignment and comparison
 RETRIEVING SEQUENCE INFORMATION Nucleotide sequence databases Database search Sequence alignment and comparison Biological sequence databases Originally just a storage place for sequences. Currently the
RETRIEVING SEQUENCE INFORMATION Nucleotide sequence databases Database search Sequence alignment and comparison Biological sequence databases Originally just a storage place for sequences. Currently the
Microarray Technology
 Microarrays And Functional Genomics CPSC265 Matt Hudson Microarray Technology Relatively young technology Usually used like a Northern blot can determine the amount of mrna for a particular gene Except
Microarrays And Functional Genomics CPSC265 Matt Hudson Microarray Technology Relatively young technology Usually used like a Northern blot can determine the amount of mrna for a particular gene Except
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION. Professor Bharat Patel Office: Science 2, 2.36 Email: [email protected].
 Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: [email protected] What is Gene Expression & Gene Regulation? 1. Gene Expression
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: [email protected] What is Gene Expression & Gene Regulation? 1. Gene Expression
BIO 3352: BIOINFORMATICS II HYBRID COURSE SYLLABUS
 BIO 3352: BIOINFORMATICS II HYBRID COURSE SYLLABUS NEW YORK CITY COLLEGE OF TECHNOLOGY The City University Of New York School of Arts and Sciences Biological Sciences Department Course title: Bioinformatics
BIO 3352: BIOINFORMATICS II HYBRID COURSE SYLLABUS NEW YORK CITY COLLEGE OF TECHNOLOGY The City University Of New York School of Arts and Sciences Biological Sciences Department Course title: Bioinformatics
DNA Sequencing Troubleshooting Guide
 DNA Sequencing Troubleshooting Guide Successful DNA Sequencing Read Peaks are well formed and separated with good quality scores. There is a small area at the beginning of the run before the chemistry
DNA Sequencing Troubleshooting Guide Successful DNA Sequencing Read Peaks are well formed and separated with good quality scores. There is a small area at the beginning of the run before the chemistry
Central Dogma. Lecture 10. Discussing DNA replication. DNA Replication. DNA mutation and repair. Transcription
 Central Dogma transcription translation DNA RNA Protein replication Discussing DNA replication (Nucleus of eukaryote, cytoplasm of prokaryote) Recall Replication is semi-conservative and bidirectional
Central Dogma transcription translation DNA RNA Protein replication Discussing DNA replication (Nucleus of eukaryote, cytoplasm of prokaryote) Recall Replication is semi-conservative and bidirectional
A Primer of Genome Science THIRD
 A Primer of Genome Science THIRD EDITION GREG GIBSON-SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc. Publishers Sunderland, Massachusetts USA Contents Preface xi 1 Genome Projects:
A Primer of Genome Science THIRD EDITION GREG GIBSON-SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc. Publishers Sunderland, Massachusetts USA Contents Preface xi 1 Genome Projects:
Analysis and Integration of Big Data from Next-Generation Genomics, Epigenomics, and Transcriptomics
 Analysis and Integration of Big Data from Next-Generation Genomics, Epigenomics, and Transcriptomics Christopher Benner, PhD Director, Integrative Genomics and Bioinformatics Core (IGC) idash Webinar,
Analysis and Integration of Big Data from Next-Generation Genomics, Epigenomics, and Transcriptomics Christopher Benner, PhD Director, Integrative Genomics and Bioinformatics Core (IGC) idash Webinar,
RT 2 Profiler PCR Array: Web-Based Data Analysis Tutorial
 RT 2 Profiler PCR Array: Web-Based Data Analysis Tutorial Samuel J. Rulli, Jr., Ph.D. qpcr-applications Scientist [email protected] Pathway Focused Research from Sample Prep to Data Analysis! -2-
RT 2 Profiler PCR Array: Web-Based Data Analysis Tutorial Samuel J. Rulli, Jr., Ph.D. qpcr-applications Scientist [email protected] Pathway Focused Research from Sample Prep to Data Analysis! -2-
How To Understand How Gene Expression Is Regulated
 What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
Recombinant DNA & Genetic Engineering. Tools for Genetic Manipulation
 Recombinant DNA & Genetic Engineering g Genetic Manipulation: Tools Kathleen Hill Associate Professor Department of Biology The University of Western Ontario Tools for Genetic Manipulation DNA, RNA, cdna
Recombinant DNA & Genetic Engineering g Genetic Manipulation: Tools Kathleen Hill Associate Professor Department of Biology The University of Western Ontario Tools for Genetic Manipulation DNA, RNA, cdna
Computational Genomics. Next generation sequencing (NGS)
 Computational Genomics Next generation sequencing (NGS) Sequencing technology defies Moore s law Nature Methods 2011 Log 10 (price) Sequencing the Human Genome 2001: Human Genome Project 2.7G$, 11 years
Computational Genomics Next generation sequencing (NGS) Sequencing technology defies Moore s law Nature Methods 2011 Log 10 (price) Sequencing the Human Genome 2001: Human Genome Project 2.7G$, 11 years
Real-time PCR: Understanding C t
 APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
Error Tolerant Searching of Uninterpreted MS/MS Data
 Error Tolerant Searching of Uninterpreted MS/MS Data 1 In any search of a large LC-MS/MS dataset 2 There are always a number of spectra which get poor scores, or even no match at all. 3 Sometimes, this
Error Tolerant Searching of Uninterpreted MS/MS Data 1 In any search of a large LC-MS/MS dataset 2 There are always a number of spectra which get poor scores, or even no match at all. 3 Sometimes, this
1. Introduction Gene regulation Genomics and genome analyses Hidden markov model (HMM)
 1. Introduction Gene regulation Genomics and genome analyses Hidden markov model (HMM) 2. Gene regulation tools and methods Regulatory sequences and motif discovery TF binding sites, microrna target prediction
1. Introduction Gene regulation Genomics and genome analyses Hidden markov model (HMM) 2. Gene regulation tools and methods Regulatory sequences and motif discovery TF binding sites, microrna target prediction
Activity 7.21 Transcription factors
 Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
ONLINE SUPPLEMENTAL MATERIAL. Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue
 ONLINE SUPPLEMENTAL MATERIAL Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue Sungmi Park 1, Ko-Ting Lu 1, Xuebo Liu 1, Tapan K. Chatterjee 2, Steven M. Rudich 3, Neal
ONLINE SUPPLEMENTAL MATERIAL Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue Sungmi Park 1, Ko-Ting Lu 1, Xuebo Liu 1, Tapan K. Chatterjee 2, Steven M. Rudich 3, Neal
Next Generation Sequencing: Technology, Mapping, and Analysis
 Next Generation Sequencing: Technology, Mapping, and Analysis Gary Benson Computer Science, Biology, Bioinformatics Boston University [email protected] http://tandem.bu.edu/ The Human Genome Project took
Next Generation Sequencing: Technology, Mapping, and Analysis Gary Benson Computer Science, Biology, Bioinformatics Boston University [email protected] http://tandem.bu.edu/ The Human Genome Project took
Data Integration. Lectures 16 & 17. ECS289A, WQ03, Filkov
 Data Integration Lectures 16 & 17 Lectures Outline Goals for Data Integration Homogeneous data integration time series data (Filkov et al. 2002) Heterogeneous data integration microarray + sequence microarray
Data Integration Lectures 16 & 17 Lectures Outline Goals for Data Integration Homogeneous data integration time series data (Filkov et al. 2002) Heterogeneous data integration microarray + sequence microarray
New Technologies for Sensitive, Low-Input RNA-Seq. Clontech Laboratories, Inc.
 New Technologies for Sensitive, Low-Input RNA-Seq Clontech Laboratories, Inc. Outline Introduction Single-Cell-Capable mrna-seq Using SMART Technology SMARTer Ultra Low RNA Kit for the Fluidigm C 1 System
New Technologies for Sensitive, Low-Input RNA-Seq Clontech Laboratories, Inc. Outline Introduction Single-Cell-Capable mrna-seq Using SMART Technology SMARTer Ultra Low RNA Kit for the Fluidigm C 1 System
Illumina TruSeq DNA Adapters De-Mystified James Schiemer
 1 of 5 Illumina TruSeq DNA Adapters De-Mystified James Schiemer The key to sequencing random fragments of DNA is by the addition of short nucleotide sequences which allow any DNA fragment to: 1) Bind to
1 of 5 Illumina TruSeq DNA Adapters De-Mystified James Schiemer The key to sequencing random fragments of DNA is by the addition of short nucleotide sequences which allow any DNA fragment to: 1) Bind to
Pairwise Sequence Alignment
 Pairwise Sequence Alignment [email protected] SS 2013 Outline Pairwise sequence alignment global - Needleman Wunsch Gotoh algorithm local - Smith Waterman algorithm BLAST - heuristics What
Pairwise Sequence Alignment [email protected] SS 2013 Outline Pairwise sequence alignment global - Needleman Wunsch Gotoh algorithm local - Smith Waterman algorithm BLAST - heuristics What
An Introduction to Genomics and SAS Scientific Discovery Solutions
 An Introduction to Genomics and SAS Scientific Discovery Solutions Dr Karen M Miller Product Manager Bioinformatics SAS EMEA 16.06.03 Copyright 2003, SAS Institute Inc. All rights reserved. 1 Overview!
An Introduction to Genomics and SAS Scientific Discovery Solutions Dr Karen M Miller Product Manager Bioinformatics SAS EMEA 16.06.03 Copyright 2003, SAS Institute Inc. All rights reserved. 1 Overview!
Control of Gene Expression
 Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
PreciseTM Whitepaper
 Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
FOR REFERENCE PURPOSES
 BIOO LIFE SCIENCE PRODUCTS FOR REFERENCE PURPOSES This manual is for Reference Purposes Only. DO NOT use this protocol to run your assays. Periodically, optimizations and revisions are made to the kit
BIOO LIFE SCIENCE PRODUCTS FOR REFERENCE PURPOSES This manual is for Reference Purposes Only. DO NOT use this protocol to run your assays. Periodically, optimizations and revisions are made to the kit
DNA Sequence Analysis
 DNA Sequence Analysis Two general kinds of analysis Screen for one of a set of known sequences Determine the sequence even if it is novel Screening for a known sequence usually involves an oligonucleotide
DNA Sequence Analysis Two general kinds of analysis Screen for one of a set of known sequences Determine the sequence even if it is novel Screening for a known sequence usually involves an oligonucleotide
Quantitative proteomics background
 Proteomics data analysis seminar Quantitative proteomics and transcriptomics of anaerobic and aerobic yeast cultures reveals post transcriptional regulation of key cellular processes de Groot, M., Daran
Proteomics data analysis seminar Quantitative proteomics and transcriptomics of anaerobic and aerobic yeast cultures reveals post transcriptional regulation of key cellular processes de Groot, M., Daran
Appendix 2 Molecular Biology Core Curriculum. Websites and Other Resources
 Appendix 2 Molecular Biology Core Curriculum Websites and Other Resources Chapter 1 - The Molecular Basis of Cancer 1. Inside Cancer http://www.insidecancer.org/ From the Dolan DNA Learning Center Cold
Appendix 2 Molecular Biology Core Curriculum Websites and Other Resources Chapter 1 - The Molecular Basis of Cancer 1. Inside Cancer http://www.insidecancer.org/ From the Dolan DNA Learning Center Cold
IMBB 2013. Genomic DNA purifica8on
 IMBB 2013 Genomic DNA purifica8on Why purify DNA? The purpose of DNA purifica8on from the cell/8ssue is to ensure it performs well in subsequent downstream applica8ons, e.g. Polymerase Chain Reac8on (PCR),
IMBB 2013 Genomic DNA purifica8on Why purify DNA? The purpose of DNA purifica8on from the cell/8ssue is to ensure it performs well in subsequent downstream applica8ons, e.g. Polymerase Chain Reac8on (PCR),
Biotechnology: DNA Technology & Genomics
 Chapter 20. Biotechnology: DNA Technology & Genomics 2003-2004 The BIG Questions How can we use our knowledge of DNA to: diagnose disease or defect? cure disease or defect? change/improve organisms? What
Chapter 20. Biotechnology: DNA Technology & Genomics 2003-2004 The BIG Questions How can we use our knowledge of DNA to: diagnose disease or defect? cure disease or defect? change/improve organisms? What
Appendix C DNA Replication & Mitosis
 K.Muma Bio 6 Appendix C DNA Replication & Mitosis Study Objectives: Appendix C: DNA replication and Mitosis 1. Describe the structure of DNA and where it is found. 2. Explain complimentary base pairing:
K.Muma Bio 6 Appendix C DNA Replication & Mitosis Study Objectives: Appendix C: DNA replication and Mitosis 1. Describe the structure of DNA and where it is found. 2. Explain complimentary base pairing:
Molecular Biology Techniques: A Classroom Laboratory Manual THIRD EDITION
 Molecular Biology Techniques: A Classroom Laboratory Manual THIRD EDITION Susan Carson Heather B. Miller D.Scott Witherow ELSEVIER AMSTERDAM BOSTON HEIDELBERG LONDON NEW YORK OXFORD PARIS SAN DIEGO SAN
Molecular Biology Techniques: A Classroom Laboratory Manual THIRD EDITION Susan Carson Heather B. Miller D.Scott Witherow ELSEVIER AMSTERDAM BOSTON HEIDELBERG LONDON NEW YORK OXFORD PARIS SAN DIEGO SAN
BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS
 BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS 1. The Technology Strategy sets out six areas where technological developments are required to push the frontiers of knowledge
BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS 1. The Technology Strategy sets out six areas where technological developments are required to push the frontiers of knowledge
Module 10: Bioinformatics
 Module 10: Bioinformatics 1.) Goal: To understand the general approaches for basic in silico (computer) analysis of DNA- and protein sequences. We are going to discuss sequence formatting required prior
Module 10: Bioinformatics 1.) Goal: To understand the general approaches for basic in silico (computer) analysis of DNA- and protein sequences. We are going to discuss sequence formatting required prior
THE LIFE SCIENCE DASHBOARD
 DASHBOARD Cell Culture Dashboard Sample Data 2005-2006 Percepta Associates, Inc. SERIESTWO DEC 08 760 822 2638 www.perceptaassociates.com 0 The Life Science Dashboards from Percepta Associates, Inc. Copyright
DASHBOARD Cell Culture Dashboard Sample Data 2005-2006 Percepta Associates, Inc. SERIESTWO DEC 08 760 822 2638 www.perceptaassociates.com 0 The Life Science Dashboards from Percepta Associates, Inc. Copyright
Interaktionen von RNAs und Proteinen
 Sonja Prohaska Computational EvoDevo Universitaet Leipzig June 9, 2015 Studying RNA-protein interactions Given: target protein known to bind to RNA problem: find binding partners and binding sites experimental
Sonja Prohaska Computational EvoDevo Universitaet Leipzig June 9, 2015 Studying RNA-protein interactions Given: target protein known to bind to RNA problem: find binding partners and binding sites experimental
Next Generation Sequencing
 Next Generation Sequencing Technology and applications 10/1/2015 Jeroen Van Houdt - Genomics Core - KU Leuven - UZ Leuven 1 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977
Next Generation Sequencing Technology and applications 10/1/2015 Jeroen Van Houdt - Genomics Core - KU Leuven - UZ Leuven 1 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977
Genome accessibility is widely preserved and locally modulated during mitosis
 Genome accessibility is widely preserved and locally modulated during mitosis Chris C.-S. Hsiung, 1, 2, Christapher S. Morrissey, 3, Maheshi Udugama, 1 Christopher L. Frank, 4 Cheryl A. Keller, 3 Songjoon
Genome accessibility is widely preserved and locally modulated during mitosis Chris C.-S. Hsiung, 1, 2, Christapher S. Morrissey, 3, Maheshi Udugama, 1 Christopher L. Frank, 4 Cheryl A. Keller, 3 Songjoon
Genomic DNA Extraction Kit INSTRUCTION MANUAL
 Genomic DNA Extraction Kit INSTRUCTION MANUAL Table of Contents Introduction 3 Kit Components 3 Storage Conditions 4 Recommended Equipment and Reagents 4 Introduction to the Protocol 4 General Overview
Genomic DNA Extraction Kit INSTRUCTION MANUAL Table of Contents Introduction 3 Kit Components 3 Storage Conditions 4 Recommended Equipment and Reagents 4 Introduction to the Protocol 4 General Overview
50 g 650 L. *Average yields will vary depending upon a number of factors including type of phage, growth conditions used and developmental stage.
 3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: [email protected] Phage DNA Isolation Kit Product # 46800, 46850 Product Insert
3430 Schmon Parkway Thorold, ON, Canada L2V 4Y6 Phone: 866-667-4362 (905) 227-8848 Fax: (905) 227-1061 Email: [email protected] Phage DNA Isolation Kit Product # 46800, 46850 Product Insert
ChIP TROUBLESHOOTING TIPS
 ChIP TROUBLESHOOTING TIPS Creative Diagnostics Abstract ChIP dissects the spatial and temporal dynamics of the interactions between chromatin and its associated factors CD Creative Diagnostics info@creative-
ChIP TROUBLESHOOTING TIPS Creative Diagnostics Abstract ChIP dissects the spatial and temporal dynamics of the interactions between chromatin and its associated factors CD Creative Diagnostics info@creative-
Multiprotocol Label Switching (MPLS)
 Multiprotocol Label Switching (MPLS) รศ.ดร. อน นต ผลเพ ม Asso. Prof. Anan Phonphoem, Ph.D. [email protected] http://www.cpe.ku.ac.th/~anan Computer Engineering Department Kasetsart University, Bangkok, Thailand
Multiprotocol Label Switching (MPLS) รศ.ดร. อน นต ผลเพ ม Asso. Prof. Anan Phonphoem, Ph.D. [email protected] http://www.cpe.ku.ac.th/~anan Computer Engineering Department Kasetsart University, Bangkok, Thailand
Genetic Analysis. Phenotype analysis: biological-biochemical analysis. Genotype analysis: molecular and physical analysis
 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
Genetics Lecture Notes 7.03 2005. Lectures 1 2
 Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Molecular Cloning, Product Brochure
 , Product Brochure Interest in any of the products, request or order them at Bio-Connect. Bio-Connect B.V. T NL +31 (0)26 326 44 50 T BE +32 (0)2 503 03 48 Begonialaan 3a F NL +31 (0)26 326 44 51 F BE
, Product Brochure Interest in any of the products, request or order them at Bio-Connect. Bio-Connect B.V. T NL +31 (0)26 326 44 50 T BE +32 (0)2 503 03 48 Begonialaan 3a F NL +31 (0)26 326 44 51 F BE
Speed Matters - Fast ways from template to result
 qpcr Symposium 2007 - Weihenstephan Speed Matters - Fast ways from template to result March 28, 2007 Dr. Thorsten Traeger Senior Scientist, Research and Development - 1 - Overview Ạgenda Fast PCR The Challenges
qpcr Symposium 2007 - Weihenstephan Speed Matters - Fast ways from template to result March 28, 2007 Dr. Thorsten Traeger Senior Scientist, Research and Development - 1 - Overview Ạgenda Fast PCR The Challenges
Introduction to Genome Annotation
 Introduction to Genome Annotation AGCGTGGTAGCGCGAGTTTGCGAGCTAGCTAGGCTCCGGATGCGA CCAGCTTTGATAGATGAATATAGTGTGCGCGACTAGCTGTGTGTT GAATATATAGTGTGTCTCTCGATATGTAGTCTGGATCTAGTGTTG GTGTAGATGGAGATCGCGTAGCGTGGTAGCGCGAGTTTGCGAGCT
Introduction to Genome Annotation AGCGTGGTAGCGCGAGTTTGCGAGCTAGCTAGGCTCCGGATGCGA CCAGCTTTGATAGATGAATATAGTGTGCGCGACTAGCTGTGTGTT GAATATATAGTGTGTCTCTCGATATGTAGTCTGGATCTAGTGTTG GTGTAGATGGAGATCGCGTAGCGTGGTAGCGCGAGTTTGCGAGCT
Introduction Bioo Scientific
 Next Generation Sequencing Catalog 2014-2015 Introduction Bioo Scientific Bioo Scientific is a global life science company headquartered in Austin, TX, committed to providing innovative products and superior
Next Generation Sequencing Catalog 2014-2015 Introduction Bioo Scientific Bioo Scientific is a global life science company headquartered in Austin, TX, committed to providing innovative products and superior
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
Design of conditional gene targeting vectors - a recombineering approach
 Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
Faculty of Medicine. Settore disciplinare: BIO/10. functional domains. Monica Soldi. IFOM-IEO Campus, Milan. Matricola n. R08407
 PhD degree in Molecular Medicine European School of Molecular Medicine (SEMM), University of Milan and University of Naples Federico II Faculty of Medicine Settore disciplinare: BIO/10 Establishment and
PhD degree in Molecular Medicine European School of Molecular Medicine (SEMM), University of Milan and University of Naples Federico II Faculty of Medicine Settore disciplinare: BIO/10 Establishment and
Introduction To Epigenetic Regulation: How Can The Epigenomics Core Services Help Your Research? Maria (Ken) Figueroa, M.D. Core Scientific Director
 Introduction To Epigenetic Regulation: How Can The Epigenomics Core Services Help Your Research? Maria (Ken) Figueroa, M.D. Core Scientific Director Gene expression depends upon multiple factors Gene Transcription
Introduction To Epigenetic Regulation: How Can The Epigenomics Core Services Help Your Research? Maria (Ken) Figueroa, M.D. Core Scientific Director Gene expression depends upon multiple factors Gene Transcription
Module 3 Questions. 7. Chemotaxis is an example of signal transduction. Explain, with the use of diagrams.
 Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Description: Molecular Biology Services and DNA Sequencing
 Description: Molecular Biology s and DNA Sequencing DNA Sequencing s Single Pass Sequencing Sequence data only, for plasmids or PCR products Plasmid DNA or PCR products Plasmid DNA: 20 100 ng/μl PCR Product:
Description: Molecular Biology s and DNA Sequencing DNA Sequencing s Single Pass Sequencing Sequence data only, for plasmids or PCR products Plasmid DNA or PCR products Plasmid DNA: 20 100 ng/μl PCR Product:
Frequently Asked Questions Next Generation Sequencing
 Frequently Asked Questions Next Generation Sequencing Import These Frequently Asked Questions for Next Generation Sequencing are some of the more common questions our customers ask. Questions are divided
Frequently Asked Questions Next Generation Sequencing Import These Frequently Asked Questions for Next Generation Sequencing are some of the more common questions our customers ask. Questions are divided
HCS604.03 Exercise 1 Dr. Jones Spring 2005. Recombinant DNA (Molecular Cloning) exercise:
 HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
Next Generation Sequencing: Adjusting to Big Data. Daniel Nicorici, Dr.Tech. Statistikot Suomen Lääketeollisuudessa 29.10.2013
 Next Generation Sequencing: Adjusting to Big Data Daniel Nicorici, Dr.Tech. Statistikot Suomen Lääketeollisuudessa 29.10.2013 Outline Human Genome Project Next-Generation Sequencing Personalized Medicine
Next Generation Sequencing: Adjusting to Big Data Daniel Nicorici, Dr.Tech. Statistikot Suomen Lääketeollisuudessa 29.10.2013 Outline Human Genome Project Next-Generation Sequencing Personalized Medicine
Molecular Computing. [email protected] 3-41 Athabasca Hall Sept. 30, 2013
 Molecular Computing [email protected] 3-41 Athabasca Hall Sept. 30, 2013 What Was The World s First Computer? The World s First Computer? ENIAC - 1946 Antikythera Mechanism - 80 BP Babbage Analytical
Molecular Computing [email protected] 3-41 Athabasca Hall Sept. 30, 2013 What Was The World s First Computer? The World s First Computer? ENIAC - 1946 Antikythera Mechanism - 80 BP Babbage Analytical
Replication Study Guide
 Replication Study Guide This study guide is a written version of the material you have seen presented in the replication unit. Self-reproduction is a function of life that human-engineered systems have
Replication Study Guide This study guide is a written version of the material you have seen presented in the replication unit. Self-reproduction is a function of life that human-engineered systems have
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Sanger Sequencing and Quality Assurance. Zbigniew Rudzki Department of Pathology University of Melbourne
 Sanger Sequencing and Quality Assurance Zbigniew Rudzki Department of Pathology University of Melbourne Sanger DNA sequencing The era of DNA sequencing essentially started with the publication of the enzymatic
Sanger Sequencing and Quality Assurance Zbigniew Rudzki Department of Pathology University of Melbourne Sanger DNA sequencing The era of DNA sequencing essentially started with the publication of the enzymatic
mrna EDITING Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A. Copyright 2005
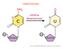 mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
Gene Mapping Techniques
 Gene Mapping Techniques OBJECTIVES By the end of this session the student should be able to: Define genetic linkage and recombinant frequency State how genetic distance may be estimated State how restriction
Gene Mapping Techniques OBJECTIVES By the end of this session the student should be able to: Define genetic linkage and recombinant frequency State how genetic distance may be estimated State how restriction
Single Nucleotide Polymorphisms (SNPs)
 Single Nucleotide Polymorphisms (SNPs) Additional Markers 13 core STR loci Obtain further information from additional markers: Y STRs Separating male samples Mitochondrial DNA Working with extremely degraded
Single Nucleotide Polymorphisms (SNPs) Additional Markers 13 core STR loci Obtain further information from additional markers: Y STRs Separating male samples Mitochondrial DNA Working with extremely degraded
Bioinformatics Unit Department of Biological Services. Get to know us
 Bioinformatics Unit Department of Biological Services Get to know us Domains of Activity IT & programming Microarray analysis Sequence analysis Bioinformatics Team Biostatistical support NGS data analysis
Bioinformatics Unit Department of Biological Services Get to know us Domains of Activity IT & programming Microarray analysis Sequence analysis Bioinformatics Team Biostatistical support NGS data analysis
Genetomic Promototypes
 Genetomic Promototypes Mirkó Palla and Dana Pe er Department of Mechanical Engineering Clarkson University Potsdam, New York and Department of Genetics Harvard Medical School 77 Avenue Louis Pasteur Boston,
Genetomic Promototypes Mirkó Palla and Dana Pe er Department of Mechanical Engineering Clarkson University Potsdam, New York and Department of Genetics Harvard Medical School 77 Avenue Louis Pasteur Boston,
Control of Gene Expression
 Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
Automated Library Preparation for Next-Generation Sequencing
 Buyer s Guide: Automated Library Preparation for Next-Generation Sequencing What to consider as you evaluate options for automating library preparation. Yes, success can be automated. Next-generation sequencing
Buyer s Guide: Automated Library Preparation for Next-Generation Sequencing What to consider as you evaluate options for automating library preparation. Yes, success can be automated. Next-generation sequencing
G E N OM I C S S E RV I C ES
 GENOMICS SERVICES THE NEW YORK GENOME CENTER NYGC is an independent non-profit implementing advanced genomic research to improve diagnosis and treatment of serious diseases. capabilities. N E X T- G E
GENOMICS SERVICES THE NEW YORK GENOME CENTER NYGC is an independent non-profit implementing advanced genomic research to improve diagnosis and treatment of serious diseases. capabilities. N E X T- G E
GeneProf and the new GeneProf Web Services
 GeneProf and the new GeneProf Web Services Florian Halbritter [email protected] Stem Cell Bioinformatics Group (Simon R. Tomlinson) [email protected] December 10, 2012 Florian Halbritter
GeneProf and the new GeneProf Web Services Florian Halbritter [email protected] Stem Cell Bioinformatics Group (Simon R. Tomlinson) [email protected] December 10, 2012 Florian Halbritter
