Faculty of Medicine. Settore disciplinare: BIO/10. functional domains. Monica Soldi. IFOM-IEO Campus, Milan. Matricola n. R08407
|
|
|
- Scot Evans
- 8 years ago
- Views:
Transcription
1 PhD degree in Molecular Medicine European School of Molecular Medicine (SEMM), University of Milan and University of Naples Federico II Faculty of Medicine Settore disciplinare: BIO/10 Establishment and optimization of the ChroP approach, combining ChIP and MS-based proteomics, for the characterization of the chromatome at distinct functional domains Monica Soldi IFOM-IEO Campus, Milan Matricola n. R08407 Supervisor: Dr. Tiziana Bonaldi IFOM-IEO Campus, Milan Added co-supervisor: Dr. Gioacchino Natoli IFOM-IEO Campus, Milan Anno accademico
2 1. ABSTRACT In eukaryotes the genetic information is stored in chromatin, a highly structured nucleoprotein complex that mediates the coordinated regulation of gene expression. The basic unit of chromatin is the nucleosome, which consists of 147bp of DNA wound around a histone octamer core containing two copies of the histones H2A, H2B, H3 and H4. Changes in chromatin structure, which do not involve the nucleotide sequence, can translate into transient or heritable adjustments in gene expression. Various mechanisms modulate chromatin states: among them a pivotal role is played by covalent histone posttranslational modifications (hptms), for which the repertoires of combinations and positions are extremely varied. In addition to hptm patterns, chromatin is characterized by the local enrichment of a distinct set of histone variants; binding proteins, including various ATP-dependent chromatin remodelling complexes; DNA methylation and differential nucleosome density. Together, these components contribute to the establishment of specific chromatin landscapes, defining the functional state of the genome in that territory. Chromatin immunoprecipitation (ChIP) and Mass Spectrometry (MS) are complementary strategies to investigate the epigenetic components of chromatin. ChIP followed by deep sequencing (ChIP-Seq) allows genome-wide profiling of hptms and binders at individual genes and regulatory regions, up to a resolution of inividual nucleosomes. However, ChIP does not inform about the protein portion of chromatin, knowledge instead offered by MS-based proteomics. At the level of individual histones, MS enables to detect virtually all hptms in an unambiguous and unbiased fashion and to reveal interplays between them. Yet, MS analysis on bulk chromatin preparations limits the inspection of PTMs to a global view, with no information about their patterning in distinct functional regions. Nowadays, a global investigation of synergies between histone PTMs, 10
3 variants, and chromatin-associated proteins in a locus-specific manner remains a very attractive unachieved goal. During the course of my PhD, I contributed in this direction developing and optimizing a global, quantitative proteomic strategy, named ChroP (Chromatin Proteomics), for the analysis of the protein component of distinct chromatin regions, enriched by modified and preparative version of ChIP. I developed two ChroP protocols, which differ in the step of chromatin IP: the Native-ChIP (N-ChIP) was used to dissect histone PTM patterns whereas the Crosslinking ChIP (X-ChIP) was used in combination with SILAC-based interactomics to characterize proteins interacting with the domains of interest. I used the antibodies against tri-methylated Lysine 9 and Lysine 4 on histone H3 (H3K9me3 and H3K4me3) to enrich functionally distinct and non-overlapping chromatin regions from HeLa nuclei. High-resolution MS of the fractionated nucleosomes enabled a dissection of the domain-specific composition in terms of histone modifications, variants and non-histonic proteins, which we refer to as the modificome and interactome. First of all, I observed the expected combinatorial enrichment of silent modifications in H3K9me3, and of active ones in H3K4me3. The accordance of my results with previous studies allows to validate the robustness of the approach and, with this confidence, I could investigate novel PTMs. Remarkably, ChroP exhibited a unique and peculiar strength in revealing PTMs associations not only at the intra-molecular level within H3, but also across the different core histones, within the same nucleosome. The SILAC-based investigation of co-associated proteins revealed a number of histone variants and multi-protein complexes differentially enriched in the two functional territories. Some of them confirmed several previously described interactions, thereby validating our method. In addition, I identified numerous novel interactors, suggesting potential novel roles and regulating chromatin pathways for these proteins. A representative case was the variant H2A.X with the WICH chromatin-remodeling complex, accumulating in the H3K9me3 regions. I thus propose a higher local density model for 11
4 H2A.X in heterochromatin and provide evidence that this accumulation, together with the recruitment of WICH, represents an additional level of modulation of the DNA damage response (DDR) in this chromatin compartment. The ChroP approach is relatively easy to setup, given the limited changes made to the conventional N- and X- ChIP protocols. Hence, ChroP emerges as a potential useful tool to dissect chromatin composition and understand how all the distinct protein components can act in a concerted manner to enforce a specific chromatin status. 12
5 2. INTRODUCTION 2.1 Chromatin, epigenetics and histone post-translational modifications Chromatin is a highly ordered nucleoprotein complex that mediates both the DNA compaction into the eukaryotic nucleus and the regulation of gene expression. At the structural level, the basic unit of chromatin is the nucleosome, consisting of 147bp of DNA wound around an octamer core containing one histone H3-H4 tetramer and two histone H2A-H2B dimers (1, 2). Functionally, chromatin is organized into two distinct regions: euchromatin is less condensed and generally permissive for transcription, whereas heterochromatin is highly condensed and transcriptionally silent. Heterochromatin is classified as being either constitutive or facultative. In constitutive heterochromatin, the DNA remains condensed throughout the cell cycle. In facultative heterochromatin however the DNA can loose its condensed form and become transcriptionally active in response to distinct signals (3-5). Changes in chromatin structure that do not involve the nucleotide sequence can translate into heritable adjustments of gene expression and thus constitute an epigenetic memory system of the cell (6-10). Epigenetic inheritance can be explained through a stepwise model proposing that epigenator, initiator and maintainer factors operate sequentially and synergistically to enforce and maintain specific functional states of the genome (11). The epigenator, a signal emanating from the external environment, is translated by an initiator into a specific chromatin/dna functional state, which is sustained by a number of different maintainer factors. These include the methylation of cytosine in CpG islands (12, 13), covalent post-translational modifications of histones (hptms) and, in light of more recent studies, the activities of non-coding RNAs (ncrna) (14, 15). 13
6 Among the epigenetic maintainers listed, histone PTMs are largely recognized as key regulators of chromatin structure and function. hptms include acetylation, ubiquitination and sumoylation of Lysines; different methylation degrees of Arginines and Lysines; phosphorylation of Serines, Threonines and Tyrosines; ADP-ribosylation of Arginines, Glutamic and Aspartic acids; deimination (or citrullination) of Arginine; Proline isomerization (Figure 1) (16-18), and in addition some less-characterized modifications. Figure 1. Histone post-translational modifications. The nucleosome core particle, with the N-terminal tail of core histone and the annotation of sites of post-translational modification. Numbers along the DNA indicate each complete helical turn on either side of the dyad axis. Sites marked by green arrows are susceptible to cutting by trypsin in intact nucleosomes. Most important modifications are Acetyl Lysine (ack); methyl Lysine (mek); methyl arginine (mer); phosphoryl serine (PS); ubiquitinated lysine (uk). Adapted from Bennister A.J. and Kouzarides T. Cell Res The histone code hypothesis proposes that histone post-translational modifications act either singly or in combination to control distinct downstream pathways or processes on chromatin, ultimately defining the functional status of the underling DNA (19). The letters of this code are the modifications themselves, which are placed and removed by enzymes known as writers and erasers, respectively. hptms exert their function on chromatin through two distinct mechanisms. In the first, higher orders of chromatin structure are altered via changes in inter-nucleosomal or histone-dna interactions, thus 14
7 controlling the accessibility of DNA-binding proteins such as transcription factors (cis mechanisms). Alternatively, hptms can generate binding platforms for the recruitment of effector proteins containing specialized domains (trans mechanisms): the so-called readers of the code (Figure 2). The readers translate the information encoded by the modification patterns into specific biological outcomes (20-23). Figure 2. Domains binding modified histones. Representation of some proteins with specific domains able to specifically bind modified histones. Adapted from Kouzarides T. Cell In addition to hptm patterns, chromatin is characterized by the local enrichment of a distinct set of histone variants; binding proteins, including various ATP-dependent chromatin remodelling complexes; and differential nucleosome density and position. Together, these components contribute to the establishment of specific chromatin landscapes, defining the functional state of the genome in that territory (Figure 3) (24). 15
8 Figure 3. The distinct components contributing to define the functional state of chromatin domain. Adapted from Margueron R. and Reinberg D. Nat Rev Genet Antibodies directed against specific hptms are traditionally used to study the language of histone modification in various assays. These include: immunofluorescence (IF) analyses of modifications at the single cell level; immunoblotting (WB), which allows profiling of PTMs in different samples and/or conditions; as well as chromatin immunoprecipitation (ChIP), which can be coupled to either PCR, DNA microarray (ChIP-on-chip) or deep sequencing (ChIP-Seq) for description of their local enrichments. The last two methods allow the genome-wide mapping of modifications, with a resolution of a few nucleosomes (25-27). Antibody-based assays are hampered by limitations in their specificity and efficiency when used to reveal the combinatorial aspect of the code. In fact, modifications can occur on adjacent or closely spaced residues within the same histone, making an epitope-masking effect more likely. To address this issue, a number of strategies have been developed to assess accurately the specificity of antibodies used in chromatin research. Peach et al. combine immunoprecipitation (IP) of native HPLC-purified H3 with mass spectrometry to detect PTMs co-enriched by a certain antibody on the same polypeptide. In 16
9 addition, Fuchs et al. have developed a peptide-array assay, based on a comprehensive library of modified peptides (28, 29). Mass spectrometry (MS) has emerged as a promising complementary analytical strategy to identify known and novel PTMs on proteins, as well as for the relative quantitation and detection of interactions between them (30). The recent advent of highresolution mass spectrometry has increased the relevance of MS-based hptm analysis by enabling the discrimination of near-isobaric modifications, either singly or in combinations, on very long polypeptides and even on intact histones (31-39). Finally, the use of different labeling strategies, both chemical and metabolic, has enabled the accurate quantitation of modifications, both in a relative and absolute manner (40). The epigenomics and chromatomics fields share a common goal in studying chromatin structure, composition and features: to gain a comprehensive view, from genome to proteome, of the epigenetic phenomena underlying the establishment and inheritance of specific expression patterns (41, 42). 2.2 Mass Spectrometry analysis and MS based-proteomics The steps of a typical proteomic experiment are shown in Figure 4. Briefly, after reducing the complexity of a protein preparation by electrophoretic speparation the proteins are subjected to enzymatic digestion, tipically using trypsin as protease. After MS analysis, by which the MS informations at the peptide level are obtained, the specific proteins are identified by software-assisted database searching, with which it is also possible to identify and localize PTMs within the peptide. 17
http://www.springer.com/3-540-23372-5
 http://www.springer.com/3-540-23372-5 Chromatin Remodeling Factors and DNA Replication Patrick Varga-Weisz Abstract Chromatin structures have to be precisely duplicated during DNA replication to maintain
http://www.springer.com/3-540-23372-5 Chromatin Remodeling Factors and DNA Replication Patrick Varga-Weisz Abstract Chromatin structures have to be precisely duplicated during DNA replication to maintain
Histone modifications. and ChIP. G. Valle - Università di Padova
 Histone modifications and ChIP Histone acetylation It has been shown that the acetylation of lysines is highly dynamic and regulated by the opposing action of two families of enzymes, histone acetyltransferases
Histone modifications and ChIP Histone acetylation It has been shown that the acetylation of lysines is highly dynamic and regulated by the opposing action of two families of enzymes, histone acetyltransferases
Control of Gene Expression
 Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Genome-wide measurements of protein-dna interaction by chromatin immunoprecipitation
 Genome-wide measurements of protein-dna interaction by chromatin immunoprecipitation D. Puthier. laboratoire INSERM, Aix-Marseille Université, TAGC/INSERM U928, Parc Scientifique de Luminy case 928 Outline
Genome-wide measurements of protein-dna interaction by chromatin immunoprecipitation D. Puthier. laboratoire INSERM, Aix-Marseille Université, TAGC/INSERM U928, Parc Scientifique de Luminy case 928 Outline
Current Motif Discovery Tools and their Limitations
 Current Motif Discovery Tools and their Limitations Philipp Bucher SIB / CIG Workshop 3 October 2006 Trendy Concepts and Hypotheses Transcription regulatory elements act in a context-dependent manner.
Current Motif Discovery Tools and their Limitations Philipp Bucher SIB / CIG Workshop 3 October 2006 Trendy Concepts and Hypotheses Transcription regulatory elements act in a context-dependent manner.
Complex multicellular organisms are produced by cells that switch genes on and off during development.
 Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
La Protéomique : Etat de l art et perspectives
 La Protéomique : Etat de l art et perspectives Odile Schiltz Institut de Pharmacologie et de Biologie Structurale CNRS, Université de Toulouse, Odile.Schiltz@ipbs.fr Protéomique et Spectrométrie de Masse
La Protéomique : Etat de l art et perspectives Odile Schiltz Institut de Pharmacologie et de Biologie Structurale CNRS, Université de Toulouse, Odile.Schiltz@ipbs.fr Protéomique et Spectrométrie de Masse
Control of Gene Expression
 Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
Introduction to Proteomics 1.0
 Introduction to Proteomics 1.0 CMSP Workshop Tim Griffin Associate Professor, BMBB Faculty Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s
Introduction to Proteomics 1.0 CMSP Workshop Tim Griffin Associate Professor, BMBB Faculty Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s
ChIP TROUBLESHOOTING TIPS
 ChIP TROUBLESHOOTING TIPS Creative Diagnostics Abstract ChIP dissects the spatial and temporal dynamics of the interactions between chromatin and its associated factors CD Creative Diagnostics info@creative-
ChIP TROUBLESHOOTING TIPS Creative Diagnostics Abstract ChIP dissects the spatial and temporal dynamics of the interactions between chromatin and its associated factors CD Creative Diagnostics info@creative-
Control of Gene Expression
 Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
School of Nursing. Presented by Yvette Conley, PhD
 Presented by Yvette Conley, PhD What we will cover during this webcast: Briefly discuss the approaches introduced in the paper: Genome Sequencing Genome Wide Association Studies Epigenomics Gene Expression
Presented by Yvette Conley, PhD What we will cover during this webcast: Briefly discuss the approaches introduced in the paper: Genome Sequencing Genome Wide Association Studies Epigenomics Gene Expression
Introduction to Proteomics
 Introduction to Proteomics Åsa Wheelock, Ph.D. Division of Respiratory Medicine & Karolinska Biomics Center asa.wheelock@ki.se In: Systems Biology and the Omics Cascade, Karolinska Institutet, June 9-13,
Introduction to Proteomics Åsa Wheelock, Ph.D. Division of Respiratory Medicine & Karolinska Biomics Center asa.wheelock@ki.se In: Systems Biology and the Omics Cascade, Karolinska Institutet, June 9-13,
Lecture 8. Protein Trafficking/Targeting. Protein targeting is necessary for proteins that are destined to work outside the cytoplasm.
 Protein Trafficking/Targeting (8.1) Lecture 8 Protein Trafficking/Targeting Protein targeting is necessary for proteins that are destined to work outside the cytoplasm. Protein targeting is more complex
Protein Trafficking/Targeting (8.1) Lecture 8 Protein Trafficking/Targeting Protein targeting is necessary for proteins that are destined to work outside the cytoplasm. Protein targeting is more complex
EPIGENETICS DNA and Histone Model
 EPIGENETICS ABSTRACT A 3-D cut-and-paste model depicting how histone, acetyl and methyl molecules control access to DNA and affect gene expression. LOGISTICS TIME REQUIRED LEARNING OBJECTIVES DNA is coiled
EPIGENETICS ABSTRACT A 3-D cut-and-paste model depicting how histone, acetyl and methyl molecules control access to DNA and affect gene expression. LOGISTICS TIME REQUIRED LEARNING OBJECTIVES DNA is coiled
Core Facility Genomics
 Core Facility Genomics versatile genome or transcriptome analyses based on quantifiable highthroughput data ascertainment 1 Topics Collaboration with Harald Binder and Clemens Kreutz Project: Microarray
Core Facility Genomics versatile genome or transcriptome analyses based on quantifiable highthroughput data ascertainment 1 Topics Collaboration with Harald Binder and Clemens Kreutz Project: Microarray
Introduction To Epigenetic Regulation: How Can The Epigenomics Core Services Help Your Research? Maria (Ken) Figueroa, M.D. Core Scientific Director
 Introduction To Epigenetic Regulation: How Can The Epigenomics Core Services Help Your Research? Maria (Ken) Figueroa, M.D. Core Scientific Director Gene expression depends upon multiple factors Gene Transcription
Introduction To Epigenetic Regulation: How Can The Epigenomics Core Services Help Your Research? Maria (Ken) Figueroa, M.D. Core Scientific Director Gene expression depends upon multiple factors Gene Transcription
The Plant Cell, March 2016, www.plantcell.org 2016 American Society of Plant Biologists. All rights reserved.
 Epigenetics (TTPB4) Outline and Study Guide Overview Epigenetic modifications of DNA and chromatin affect the activity of genes and transposons. Epigenetic controls affect processes as diverse as time-of-flowering,
Epigenetics (TTPB4) Outline and Study Guide Overview Epigenetic modifications of DNA and chromatin affect the activity of genes and transposons. Epigenetic controls affect processes as diverse as time-of-flowering,
Session 1. Course Presentation: Mass spectrometry-based proteomics for molecular and cellular biologists
 Program Overview Session 1. Course Presentation: Mass spectrometry-based proteomics for molecular and cellular biologists Session 2. Principles of Mass Spectrometry Session 3. Mass spectrometry based proteomics
Program Overview Session 1. Course Presentation: Mass spectrometry-based proteomics for molecular and cellular biologists Session 2. Principles of Mass Spectrometry Session 3. Mass spectrometry based proteomics
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Sickle cell anemia: Altered beta chain Single AA change (#6 Glu to Val) Consequence: Protein polymerizes Change in RBC shape ---> phenotypes
 Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
RT 2 Profiler PCR Array: Web-Based Data Analysis Tutorial
 RT 2 Profiler PCR Array: Web-Based Data Analysis Tutorial Samuel J. Rulli, Jr., Ph.D. qpcr-applications Scientist Samuel.Rulli@QIAGEN.com Pathway Focused Research from Sample Prep to Data Analysis! -2-
RT 2 Profiler PCR Array: Web-Based Data Analysis Tutorial Samuel J. Rulli, Jr., Ph.D. qpcr-applications Scientist Samuel.Rulli@QIAGEN.com Pathway Focused Research from Sample Prep to Data Analysis! -2-
The Nucleus: DNA, Chromatin And Chromosomes
 The Nucleus: DNA, Chromatin And Chromosomes Professor Alfred Cuschieri Department of Anatomy, University of Malta. Objectives By the end of this unit the student should be able to: 1. List the major structural
The Nucleus: DNA, Chromatin And Chromosomes Professor Alfred Cuschieri Department of Anatomy, University of Malta. Objectives By the end of this unit the student should be able to: 1. List the major structural
Interaktionen von RNAs und Proteinen
 Sonja Prohaska Computational EvoDevo Universitaet Leipzig June 9, 2015 Studying RNA-protein interactions Given: target protein known to bind to RNA problem: find binding partners and binding sites experimental
Sonja Prohaska Computational EvoDevo Universitaet Leipzig June 9, 2015 Studying RNA-protein interactions Given: target protein known to bind to RNA problem: find binding partners and binding sites experimental
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE Q5B
 INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
Building innovative drug discovery alliances. Evotec Munich. Quantitative Proteomics to Support the Discovery & Development of Targeted Drugs
 Building innovative drug discovery alliances Evotec Munich Quantitative Proteomics to Support the Discovery & Development of Targeted Drugs Evotec AG, Evotec Munich, June 2013 About Evotec Munich A leader
Building innovative drug discovery alliances Evotec Munich Quantitative Proteomics to Support the Discovery & Development of Targeted Drugs Evotec AG, Evotec Munich, June 2013 About Evotec Munich A leader
European Medicines Agency
 European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
Analysis and Integration of Big Data from Next-Generation Genomics, Epigenomics, and Transcriptomics
 Analysis and Integration of Big Data from Next-Generation Genomics, Epigenomics, and Transcriptomics Christopher Benner, PhD Director, Integrative Genomics and Bioinformatics Core (IGC) idash Webinar,
Analysis and Integration of Big Data from Next-Generation Genomics, Epigenomics, and Transcriptomics Christopher Benner, PhD Director, Integrative Genomics and Bioinformatics Core (IGC) idash Webinar,
TWO DIMENSIONAL DIFFERENTIAL IN-GEL ELECTROPHORESIS- BASED PROTEOMICS OF MALE GAMETES IN RELATION TO
 TWO DIMENSIONAL DIFFERENTIAL IN-GEL ELECTROPHORESIS- BASED PROTEOMICS OF MALE GAMETES IN RELATION TO OXIDATIVE STRESS Alaa Hamada, MD., Rakesh Sharma, PhD., Stefan S Du Plessis, PhD., Belinda Willard,
TWO DIMENSIONAL DIFFERENTIAL IN-GEL ELECTROPHORESIS- BASED PROTEOMICS OF MALE GAMETES IN RELATION TO OXIDATIVE STRESS Alaa Hamada, MD., Rakesh Sharma, PhD., Stefan S Du Plessis, PhD., Belinda Willard,
Chapter 3. Protein Structure and Function
 Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Genetomic Promototypes
 Genetomic Promototypes Mirkó Palla and Dana Pe er Department of Mechanical Engineering Clarkson University Potsdam, New York and Department of Genetics Harvard Medical School 77 Avenue Louis Pasteur Boston,
Genetomic Promototypes Mirkó Palla and Dana Pe er Department of Mechanical Engineering Clarkson University Potsdam, New York and Department of Genetics Harvard Medical School 77 Avenue Louis Pasteur Boston,
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Lecture Series 7. From DNA to Protein. Genotype to Phenotype. Reading Assignments. A. Genes and the Synthesis of Polypeptides
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Ms. Campbell Protein Synthesis Practice Questions Regents L.E.
 Name Student # Ms. Campbell Protein Synthesis Practice Questions Regents L.E. 1. A sequence of three nitrogenous bases in a messenger-rna molecule is known as a 1) codon 2) gene 3) polypeptide 4) nucleotide
Name Student # Ms. Campbell Protein Synthesis Practice Questions Regents L.E. 1. A sequence of three nitrogenous bases in a messenger-rna molecule is known as a 1) codon 2) gene 3) polypeptide 4) nucleotide
ProSightPC 3.0 Quick Start Guide
 ProSightPC 3.0 Quick Start Guide The Thermo ProSightPC 3.0 application is the only proteomics software suite that effectively supports high-mass-accuracy MS/MS experiments performed on LTQ FT and LTQ Orbitrap
ProSightPC 3.0 Quick Start Guide The Thermo ProSightPC 3.0 application is the only proteomics software suite that effectively supports high-mass-accuracy MS/MS experiments performed on LTQ FT and LTQ Orbitrap
From DNA to Protein. Proteins. Chapter 13. Prokaryotes and Eukaryotes. The Path From Genes to Proteins. All proteins consist of polypeptide chains
 Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
BROMOscan Quantitative Ligand Binding Assays
 BROMOscan Quantitative Ligand Binding Assays Identification of Best In Class Bromodomain Inhibitors Elizabeth Quinn, PhD Director, LeadHunter Discovery Services Outline Introduction Challenges to Epigenetic
BROMOscan Quantitative Ligand Binding Assays Identification of Best In Class Bromodomain Inhibitors Elizabeth Quinn, PhD Director, LeadHunter Discovery Services Outline Introduction Challenges to Epigenetic
Thermo Scientific PepFinder Software A New Paradigm for Peptide Mapping
 Thermo Scientific PepFinder Software A New Paradigm for Peptide Mapping For Conclusive Characterization of Biologics Deep Protein Characterization Is Crucial Pharmaceuticals have historically been small
Thermo Scientific PepFinder Software A New Paradigm for Peptide Mapping For Conclusive Characterization of Biologics Deep Protein Characterization Is Crucial Pharmaceuticals have historically been small
Exclusive customized epigenetic antibodies
 Exclusive customized epigenetic antibodies For many years, Diagenode has acquired a strong and reliable experience in the production of custom epigenetic antibodies, allowing us to become one of the most
Exclusive customized epigenetic antibodies For many years, Diagenode has acquired a strong and reliable experience in the production of custom epigenetic antibodies, allowing us to become one of the most
GENE REGULATION. Teacher Packet
 AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
Concluding lesson. Student manual. What kind of protein are you? (Basic)
 Concluding lesson Student manual What kind of protein are you? (Basic) Part 1 The hereditary material of an organism is stored in a coded way on the DNA. This code consists of four different nucleotides:
Concluding lesson Student manual What kind of protein are you? (Basic) Part 1 The hereditary material of an organism is stored in a coded way on the DNA. This code consists of four different nucleotides:
MUTATION, DNA REPAIR AND CANCER
 MUTATION, DNA REPAIR AND CANCER 1 Mutation A heritable change in the genetic material Essential to the continuity of life Source of variation for natural selection New mutations are more likely to be harmful
MUTATION, DNA REPAIR AND CANCER 1 Mutation A heritable change in the genetic material Essential to the continuity of life Source of variation for natural selection New mutations are more likely to be harmful
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION. Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.
 Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS
 BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS 1. The Technology Strategy sets out six areas where technological developments are required to push the frontiers of knowledge
BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS 1. The Technology Strategy sets out six areas where technological developments are required to push the frontiers of knowledge
AP BIOLOGY 2009 SCORING GUIDELINES
 AP BIOLOGY 2009 SCORING GUIDELINES Question 4 The flow of genetic information from DNA to protein in eukaryotic cells is called the central dogma of biology. (a) Explain the role of each of the following
AP BIOLOGY 2009 SCORING GUIDELINES Question 4 The flow of genetic information from DNA to protein in eukaryotic cells is called the central dogma of biology. (a) Explain the role of each of the following
Tutorial for Proteomics Data Submission. Katalin F. Medzihradszky Robert J. Chalkley UCSF
 Tutorial for Proteomics Data Submission Katalin F. Medzihradszky Robert J. Chalkley UCSF Why Have Guidelines? Large-scale proteomics studies create huge amounts of data. It is impossible/impractical to
Tutorial for Proteomics Data Submission Katalin F. Medzihradszky Robert J. Chalkley UCSF Why Have Guidelines? Large-scale proteomics studies create huge amounts of data. It is impossible/impractical to
2007 7.013 Problem Set 1 KEY
 2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
PRACTICE TEST QUESTIONS
 PART A: MULTIPLE CHOICE QUESTIONS PRACTICE TEST QUESTIONS DNA & PROTEIN SYNTHESIS B 1. One of the functions of DNA is to A. secrete vacuoles. B. make copies of itself. C. join amino acids to each other.
PART A: MULTIPLE CHOICE QUESTIONS PRACTICE TEST QUESTIONS DNA & PROTEIN SYNTHESIS B 1. One of the functions of DNA is to A. secrete vacuoles. B. make copies of itself. C. join amino acids to each other.
13.2 Ribosomes & Protein Synthesis
 13.2 Ribosomes & Protein Synthesis Introduction: *A specific sequence of bases in DNA carries the directions for forming a polypeptide, a chain of amino acids (there are 20 different types of amino acid).
13.2 Ribosomes & Protein Synthesis Introduction: *A specific sequence of bases in DNA carries the directions for forming a polypeptide, a chain of amino acids (there are 20 different types of amino acid).
Translation. Translation: Assembly of polypeptides on a ribosome
 Translation Translation: Assembly of polypeptides on a ribosome Living cells devote more energy to the synthesis of proteins than to any other aspect of metabolism. About a third of the dry mass of a cell
Translation Translation: Assembly of polypeptides on a ribosome Living cells devote more energy to the synthesis of proteins than to any other aspect of metabolism. About a third of the dry mass of a cell
Regulation of enzyme activity
 1 Regulation of enzyme activity Regulation of enzyme activity is important to coordinate the different metabolic processes. It is also important for homeostasis i.e. to maintain the internal environment
1 Regulation of enzyme activity Regulation of enzyme activity is important to coordinate the different metabolic processes. It is also important for homeostasis i.e. to maintain the internal environment
H H N - C - C 2 R. Three possible forms (not counting R group) depending on ph
 Amino acids - 0 common amino acids there are others found naturally but much less frequently - Common structure for amino acid - C, -N, and functional groups all attached to the alpha carbon N - C - C
Amino acids - 0 common amino acids there are others found naturally but much less frequently - Common structure for amino acid - C, -N, and functional groups all attached to the alpha carbon N - C - C
Psychoonkology, Sept. 2010 lifestyle factors and epigenetics
 Psychoonkology, Sept. 2010 lifestyle factors and epigenetics Alexander G. Haslberger Dep. für Ernährungswissenschaften Univ. of Vienna Working group: Food, GI-Microbiology, Epigenetics Content Health:
Psychoonkology, Sept. 2010 lifestyle factors and epigenetics Alexander G. Haslberger Dep. für Ernährungswissenschaften Univ. of Vienna Working group: Food, GI-Microbiology, Epigenetics Content Health:
Methods for Protein Analysis
 Methods for Protein Analysis 1. Protein Separation Methods The following is a quick review of some common methods used for protein separation: SDS-PAGE (SDS-polyacrylamide gel electrophoresis) separates
Methods for Protein Analysis 1. Protein Separation Methods The following is a quick review of some common methods used for protein separation: SDS-PAGE (SDS-polyacrylamide gel electrophoresis) separates
PperCHIP. High-Content Peptide Microarrays. Epitope Mapping & Serum Profiling Services
 PepperChip High-Content Peptide Microarrays PepperMap Epitope Mapping & Serum Profiling Services PperCHIP access to custom and dard peptide microarrays up to 9,000 features per in a uniquely flexible and
PepperChip High-Content Peptide Microarrays PepperMap Epitope Mapping & Serum Profiling Services PperCHIP access to custom and dard peptide microarrays up to 9,000 features per in a uniquely flexible and
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
Activity 7.21 Transcription factors
 Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
A Brief Introduction on DNase-Seq Data Aanalysis
 A Brief Introduction on DNase-Seq Data Aanalysis Hashem Koohy, Thomas Down, Mikhail Spivakov and Tim Hubbard Spivakov s and Fraser s Lab September 13, 2014 1 Introduction DNaseI is an enzyme which cuts
A Brief Introduction on DNase-Seq Data Aanalysis Hashem Koohy, Thomas Down, Mikhail Spivakov and Tim Hubbard Spivakov s and Fraser s Lab September 13, 2014 1 Introduction DNaseI is an enzyme which cuts
NO CALCULATORS OR CELL PHONES ALLOWED
 Biol 205 Exam 1 TEST FORM A Spring 2008 NAME Fill out both sides of the Scantron Sheet. On Side 2 be sure to indicate that you have TEST FORM A The answers to Part I should be placed on the SCANTRON SHEET.
Biol 205 Exam 1 TEST FORM A Spring 2008 NAME Fill out both sides of the Scantron Sheet. On Side 2 be sure to indicate that you have TEST FORM A The answers to Part I should be placed on the SCANTRON SHEET.
Interaktionen von Nukleinsäuren und Proteinen
 Sonja Prohaska Computational EvoDevo Universitaet Leipzig June 9, 2015 DNA is never naked in a cell DNA is usually in association with proteins. In all domains of life there are small, basic chromosomal
Sonja Prohaska Computational EvoDevo Universitaet Leipzig June 9, 2015 DNA is never naked in a cell DNA is usually in association with proteins. In all domains of life there are small, basic chromosomal
Diabetes and Insulin Signaling
 Diabetes and Insulin Signaling NATIONAL CENTER FOR CASE STUDY TEACHING IN SCIENCE by Kristy J. Wilson School of Mathematics and Sciences Marian University, Indianapolis, IN Part I Research Orientation
Diabetes and Insulin Signaling NATIONAL CENTER FOR CASE STUDY TEACHING IN SCIENCE by Kristy J. Wilson School of Mathematics and Sciences Marian University, Indianapolis, IN Part I Research Orientation
Hormones & Chemical Signaling
 Hormones & Chemical Signaling Part 2 modulation of signal pathways and hormone classification & function How are these pathways controlled? Receptors are proteins! Subject to Specificity of binding Competition
Hormones & Chemical Signaling Part 2 modulation of signal pathways and hormone classification & function How are these pathways controlled? Receptors are proteins! Subject to Specificity of binding Competition
Pipe Cleaner Proteins. Essential question: How does the structure of proteins relate to their function in the cell?
 Pipe Cleaner Proteins GPS: SB1 Students will analyze the nature of the relationships between structures and functions in living cells. Essential question: How does the structure of proteins relate to their
Pipe Cleaner Proteins GPS: SB1 Students will analyze the nature of the relationships between structures and functions in living cells. Essential question: How does the structure of proteins relate to their
FACULTY OF MEDICAL SCIENCE
 Doctor of Philosophy in Biochemistry FACULTY OF MEDICAL SCIENCE Naresuan University 73 Doctor of Philosophy in Biochemistry The Biochemistry Department at Naresuan University is a leader in lower northern
Doctor of Philosophy in Biochemistry FACULTY OF MEDICAL SCIENCE Naresuan University 73 Doctor of Philosophy in Biochemistry The Biochemistry Department at Naresuan University is a leader in lower northern
Multiple Choice Write the letter that best answers the question or completes the statement on the line provided.
 Name lass Date hapter 12 DN and RN hapter Test Multiple hoice Write the letter that best answers the question or completes the statement on the line provided. Pearson Education, Inc. ll rights reserved.
Name lass Date hapter 12 DN and RN hapter Test Multiple hoice Write the letter that best answers the question or completes the statement on the line provided. Pearson Education, Inc. ll rights reserved.
Title: Author(s): Source: Document Type: Copyright : http://naturalhistorymag.com/ Full Text: [ILLUSTRATION OMITTED]
![Title: Author(s): Source: Document Type: Copyright : http://naturalhistorymag.com/ Full Text: [ILLUSTRATION OMITTED] Title: Author(s): Source: Document Type: Copyright : http://naturalhistorymag.com/ Full Text: [ILLUSTRATION OMITTED]](/thumbs/33/16056031.jpg) Title: Author(s): Source: Document Type: Copyright : http://naturalhistorymag.com/ Full Text: Epigenetics in action: Part 2: turn-ons and turn-offs Nessa Carey Natural History. 120.5 (May 2012): p30. Article
Title: Author(s): Source: Document Type: Copyright : http://naturalhistorymag.com/ Full Text: Epigenetics in action: Part 2: turn-ons and turn-offs Nessa Carey Natural History. 120.5 (May 2012): p30. Article
AP BIOLOGY 2008 SCORING GUIDELINES (Form B)
 AP BIOLOGY 2008 SCORING GUIDELINES (Form B) Question 2 2. Many biological structures are composed of smaller units assembled into more complex structures having functions based on their structural organization.
AP BIOLOGY 2008 SCORING GUIDELINES (Form B) Question 2 2. Many biological structures are composed of smaller units assembled into more complex structures having functions based on their structural organization.
Research-grade Targeted Proteomics Assay Development: PRMs for PTM Studies with Skyline or, How I learned to ditch the triple quad and love the QE
 Research-grade Targeted Proteomics Assay Development: PRMs for PTM Studies with Skyline or, How I learned to ditch the triple quad and love the QE Jacob D. Jaffe Skyline Webinar July 2015 Proteomics and
Research-grade Targeted Proteomics Assay Development: PRMs for PTM Studies with Skyline or, How I learned to ditch the triple quad and love the QE Jacob D. Jaffe Skyline Webinar July 2015 Proteomics and
Searching Nucleotide Databases
 Searching Nucleotide Databases 1 When we search a nucleic acid databases, Mascot always performs a 6 frame translation on the fly. That is, 3 reading frames from the forward strand and 3 reading frames
Searching Nucleotide Databases 1 When we search a nucleic acid databases, Mascot always performs a 6 frame translation on the fly. That is, 3 reading frames from the forward strand and 3 reading frames
Tool Development for Transformational Biotechnology Advances. Breakout Session: Engineering Tools
 Tool Development for Transformational Biotechnology Advances Breakout Session: Engineering Tools Transformation/Analytical Tools Gene-delivery methods Analytical methods Agrobacteria Viral Biolistics Electrotransfection
Tool Development for Transformational Biotechnology Advances Breakout Session: Engineering Tools Transformation/Analytical Tools Gene-delivery methods Analytical methods Agrobacteria Viral Biolistics Electrotransfection
Transcription and Translation of DNA
 Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Expression and Purification of Recombinant Protein in bacteria and Yeast. Presented By: Puspa pandey, Mohit sachdeva & Ming yu
 Expression and Purification of Recombinant Protein in bacteria and Yeast Presented By: Puspa pandey, Mohit sachdeva & Ming yu DNA Vectors Molecular carriers which carry fragments of DNA into host cell.
Expression and Purification of Recombinant Protein in bacteria and Yeast Presented By: Puspa pandey, Mohit sachdeva & Ming yu DNA Vectors Molecular carriers which carry fragments of DNA into host cell.
Mass Spectrometry Signal Calibration for Protein Quantitation
 Cambridge Isotope Laboratories, Inc. www.isotope.com Proteomics Mass Spectrometry Signal Calibration for Protein Quantitation Michael J. MacCoss, PhD Associate Professor of Genome Sciences University of
Cambridge Isotope Laboratories, Inc. www.isotope.com Proteomics Mass Spectrometry Signal Calibration for Protein Quantitation Michael J. MacCoss, PhD Associate Professor of Genome Sciences University of
Name Date Period. 2. When a molecule of double-stranded DNA undergoes replication, it results in
 DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
Chapter 5: Organization and Expression of Immunoglobulin Genes
 Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Module 3 Questions. 7. Chemotaxis is an example of signal transduction. Explain, with the use of diagrams.
 Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Proteomics in Practice
 Reiner Westermeier, Torn Naven Hans-Rudolf Höpker Proteomics in Practice A Guide to Successful Experimental Design 2008 Wiley-VCH Verlag- Weinheim 978-3-527-31941-1 Preface Foreword XI XIII Abbreviations,
Reiner Westermeier, Torn Naven Hans-Rudolf Höpker Proteomics in Practice A Guide to Successful Experimental Design 2008 Wiley-VCH Verlag- Weinheim 978-3-527-31941-1 Preface Foreword XI XIII Abbreviations,
Gene mutation and molecular medicine Chapter 15
 Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
RNA and Protein Synthesis
 Name lass Date RN and Protein Synthesis Information and Heredity Q: How does information fl ow from DN to RN to direct the synthesis of proteins? 13.1 What is RN? WHT I KNOW SMPLE NSWER: RN is a nucleic
Name lass Date RN and Protein Synthesis Information and Heredity Q: How does information fl ow from DN to RN to direct the synthesis of proteins? 13.1 What is RN? WHT I KNOW SMPLE NSWER: RN is a nucleic
NUVISAN Pharma Services
 NUVISAN Pharma Services CESI MS Now available! 1st CRO in Europe! At the highest levels of quality. LABORATORY SERVICES Equipment update STATE OF THE ART AT NUVISAN CESI MS Now available! 1st CRO in Europe!
NUVISAN Pharma Services CESI MS Now available! 1st CRO in Europe! At the highest levels of quality. LABORATORY SERVICES Equipment update STATE OF THE ART AT NUVISAN CESI MS Now available! 1st CRO in Europe!
DNA (genetic information in genes) RNA (copies of genes) proteins (functional molecules) directionality along the backbone 5 (phosphate) to 3 (OH)
 DNA, RNA, replication, translation, and transcription Overview Recall the central dogma of biology: DNA (genetic information in genes) RNA (copies of genes) proteins (functional molecules) DNA structure
DNA, RNA, replication, translation, and transcription Overview Recall the central dogma of biology: DNA (genetic information in genes) RNA (copies of genes) proteins (functional molecules) DNA structure
ELITE Custom Antibody Services
 ELITE Custom Antibody Services ELITE Custom Antibody Services Experience, confidence, and understanding As a manufacturer and service provider, we have the experience, confidence, and understanding to
ELITE Custom Antibody Services ELITE Custom Antibody Services Experience, confidence, and understanding As a manufacturer and service provider, we have the experience, confidence, and understanding to
Error Tolerant Searching of Uninterpreted MS/MS Data
 Error Tolerant Searching of Uninterpreted MS/MS Data 1 In any search of a large LC-MS/MS dataset 2 There are always a number of spectra which get poor scores, or even no match at all. 3 Sometimes, this
Error Tolerant Searching of Uninterpreted MS/MS Data 1 In any search of a large LC-MS/MS dataset 2 There are always a number of spectra which get poor scores, or even no match at all. 3 Sometimes, this
Biochemistry - I. Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture-11 Enzyme Mechanisms II
 Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture-11 Enzyme Mechanisms II In the last class we studied the enzyme mechanisms of ribonuclease A
Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture-11 Enzyme Mechanisms II In the last class we studied the enzyme mechanisms of ribonuclease A
Protein Synthesis How Genes Become Constituent Molecules
 Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Kinexus has an in-house inventory of lysates prepared from 16 human cancer cell lines that have been selected to represent a diversity of tissues,
 Kinexus Bioinformatics Corporation is seeking to map and monitor the molecular communications networks of living cells for biomedical research into the diagnosis, prognosis and treatment of human diseases.
Kinexus Bioinformatics Corporation is seeking to map and monitor the molecular communications networks of living cells for biomedical research into the diagnosis, prognosis and treatment of human diseases.
Basic Principles of Transcription and Translation
 The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific traits by dictating the synthesis of
The Flow of Genetic Information The information content of DNA is in the form of specific sequences of nucleotides The DNA inherited by an organism leads to specific traits by dictating the synthesis of
A N OVERVIEW OF GENE EXPRESSION
 C H A P T E R E I G H T 8 C o n t r o l o f G e n e E x p r e s s i o n An organism's DNA encodes all of the RNA and protein molecules that are needed to make its cells. Yet a complete description of the
C H A P T E R E I G H T 8 C o n t r o l o f G e n e E x p r e s s i o n An organism's DNA encodes all of the RNA and protein molecules that are needed to make its cells. Yet a complete description of the
Retrospective Analysis of a Host Cell Protein Perfect Storm: Identifying Immunogenic Proteins and Fixing the Problem
 Retrospective Analysis of a Host Cell Protein Perfect Storm: Identifying Immunogenic Proteins and Fixing the Problem Kevin Van Cott, Associate Professor Dept. of Chemical and Biomolecular Engineering Nebraska
Retrospective Analysis of a Host Cell Protein Perfect Storm: Identifying Immunogenic Proteins and Fixing the Problem Kevin Van Cott, Associate Professor Dept. of Chemical and Biomolecular Engineering Nebraska
mrna EDITING Watson et al., BIOLOGIA MOLECOLARE DEL GENE, Zanichelli editore S.p.A. Copyright 2005
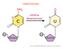 mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
mrna EDITING mrna EDITING http://dbb.urmc.rochester.edu/labs/smith/research_2.htm The number of A to I sites in the human transcriptome >15;000 the vast majority of these sites occurring in Alu repeats
Quantitative proteomics background
 Proteomics data analysis seminar Quantitative proteomics and transcriptomics of anaerobic and aerobic yeast cultures reveals post transcriptional regulation of key cellular processes de Groot, M., Daran
Proteomics data analysis seminar Quantitative proteomics and transcriptomics of anaerobic and aerobic yeast cultures reveals post transcriptional regulation of key cellular processes de Groot, M., Daran
Chapter 14. Modeling Experimental Design for Proteomics. Jan Eriksson and David Fenyö. Abstract. 1. Introduction
 Chapter Modeling Experimental Design for Proteomics Jan Eriksson and David Fenyö Abstract The complexity of proteomes makes good experimental design essential for their successful investigation. Here,
Chapter Modeling Experimental Design for Proteomics Jan Eriksson and David Fenyö Abstract The complexity of proteomes makes good experimental design essential for their successful investigation. Here,
BCHM 32200 Analytical Biochemistry Syllabus Spring, 2013
 INSTRUCTOR: Dr. Mark Hall office: BCHM 214 TEL: 494-0714 e-mail: mchall@purdue.edu DEPARTMENT OF BIOCHEMISTRY BCHM 32200 Analytical Biochemistry Syllabus Spring, 2013 Office hours: By appointment only
INSTRUCTOR: Dr. Mark Hall office: BCHM 214 TEL: 494-0714 e-mail: mchall@purdue.edu DEPARTMENT OF BIOCHEMISTRY BCHM 32200 Analytical Biochemistry Syllabus Spring, 2013 Office hours: By appointment only
BIO 3352: BIOINFORMATICS II HYBRID COURSE SYLLABUS
 BIO 3352: BIOINFORMATICS II HYBRID COURSE SYLLABUS NEW YORK CITY COLLEGE OF TECHNOLOGY The City University Of New York School of Arts and Sciences Biological Sciences Department Course title: Bioinformatics
BIO 3352: BIOINFORMATICS II HYBRID COURSE SYLLABUS NEW YORK CITY COLLEGE OF TECHNOLOGY The City University Of New York School of Arts and Sciences Biological Sciences Department Course title: Bioinformatics
BIOINF 525 Winter 2016 Foundations of Bioinformatics and Systems Biology http://tinyurl.com/bioinf525-w16
 Course Director: Dr. Barry Grant (DCM&B, bjgrant@med.umich.edu) Description: This is a three module course covering (1) Foundations of Bioinformatics, (2) Statistics in Bioinformatics, and (3) Systems
Course Director: Dr. Barry Grant (DCM&B, bjgrant@med.umich.edu) Description: This is a three module course covering (1) Foundations of Bioinformatics, (2) Statistics in Bioinformatics, and (3) Systems
Title: Surveying Genome to Identify Origins of DNA Replication In Silico
 Title: Surveying Genome to Identify Origins of DNA Replication In Silico Abstract: DNA replication origins are the bases to realize the process of chromosome replication and analyze the progress of cell
Title: Surveying Genome to Identify Origins of DNA Replication In Silico Abstract: DNA replication origins are the bases to realize the process of chromosome replication and analyze the progress of cell
Definition of the Measurand: CRP
 A Reference Measurement System for C-reactive Protein David M. Bunk, Ph.D. Chemical Science and Technology Laboratory National Institute of Standards and Technology Definition of the Measurand: Human C-reactive
A Reference Measurement System for C-reactive Protein David M. Bunk, Ph.D. Chemical Science and Technology Laboratory National Institute of Standards and Technology Definition of the Measurand: Human C-reactive
Student name ID # 2. (4 pts) What is the terminal electron acceptor in respiration? In photosynthesis? O2, NADP+
 1. Membrane transport. A. (4 pts) What ion couples primary and secondary active transport in animal cells? What ion serves the same function in plant cells? Na+, H+ 2. (4 pts) What is the terminal electron
1. Membrane transport. A. (4 pts) What ion couples primary and secondary active transport in animal cells? What ion serves the same function in plant cells? Na+, H+ 2. (4 pts) What is the terminal electron
Molecular Genetics. RNA, Transcription, & Protein Synthesis
 Molecular Genetics RNA, Transcription, & Protein Synthesis Section 1 RNA AND TRANSCRIPTION Objectives Describe the primary functions of RNA Identify how RNA differs from DNA Describe the structure and
Molecular Genetics RNA, Transcription, & Protein Synthesis Section 1 RNA AND TRANSCRIPTION Objectives Describe the primary functions of RNA Identify how RNA differs from DNA Describe the structure and
