Endogenous Controls for Real-Time Quantitation of mirna Using TaqMan MicroRNA Assays.
|
|
|
- Heather Ball
- 8 years ago
- Views:
Transcription
1 application note TaqMan MicroRNA Assays Endogenous Controls for Real-Time Quantitation of mirna Using TaqMan MicroRNA Assays. Introduction MicroRNAs (mirnas) are small noncoding RNAs whose function has been implicated in a wide range of fundamental cellular processes including cell proliferation, cell differentiation, and cell death. Quantitation of mirna gene expression levels has become an essential step in understanding these mechanisms, and has shown great promise in identifying effective biomarkers correlative with human disease 1,2. Applied Biosystems has developed an extensive set of TaqMan MicroRNA Assays, novel stem-loop RT and real-time PCR assays, for the quantitation of mature mirna expression 3. TaqMan Assays are the ideal choice for these applications because of their unsurpassed sensitivity, specificity, and wide dynamic range. Additionally, far less input material is required compared to microarrays and other alternative technologies. When performing these experiments, variation in the amount of starting material, sample collection, RNA preparation and quality, and reverse transcription (RT) efficiency can contribute to quantification errors. Normalization to endogenous control genes is currently the most accurate method to correct for potential RNA input or RT efficiency biases. Careful selection of an appropriate control or set of controls is extremely important as significant variation has been observed between samples, even for the most commonly used housekeeping genes, including ACTB (ß-Actin) and GAPDH 4. An ideal endogenous control generally demonstrates gene expression that is relatively constant and highly abundant across tissues and cell types. However, one must still validate the chosen endogenous control or set of controls for the target cell, tissue, or treatment 5, as no single control can serve as a universal endogenous control for all experimental conditions. When considering endogenous controls suitable for use with TaqMan MicroRNA Assays, it is important that they share similar properties, such as RNA stability and size, and are amenable to the mirna assay design. A number of reports indicate that other classes of small non-coding RNAs (ncrnas) are expressed both abundantly and stably, making them good endogenous control candidates. We have performed a systematic study of a set of human ncrna species ranging in size from 45
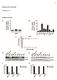
2 to 0 nucleotides, including transfer RNA (trna), small nuclear RNA (snrna) and small nucleolar RNA (snorna) 6 across a relatively wide variety of tissues and cell lines to determine their suitability as endogenous controls when quantitating mirna expression levels using real-time PCR methods. Materials and Methods Small Non-coding RNAs (ncrna) Candidate small non-coding RNA sequences were obtained from the NCBI s GenBank database. TaqMan MicroRNA Assays TaqMan MicroRNA Assays, consisting of an RNA-specific stem-looped RT primer and TaqMan Assay (forward and reverse primers and FAM dyelabeled MGB probe), were designed for each candidate endogenous control RNA using Applied Biosystems mirnaspecific assay design parameters. A full list of TaqMan MicroRNA Assays is available at mirna.appliedbiosystems.com TaqMan Gene Expression Assays The TaqMan Gene Expression Assays for detection of endogenous control genes used in this experiment were: 18S (Hs _s1), GAPDH (Hs _m1), and ACTB (Hs _m1). A full list of TaqMan Gene Expression Assays is available at RNA Samples Total RNA for human and mouse tissues were acquired from Ambion, Inc., an Applied Biosystems Business, and BD Biosciences. Total RNA for NCI-60 cell lines was obtained from Dartmouth Medical School. Reverse Transcription (RT) Reaction The RT reaction was performed using the TaqMan MicroRNA Reverse Transcription Kit (PN , 0 reactions; or PN , 1,000 reactions). Next, 2 ng/µl of RNA, 1X stem-loop RT primer, 3.33 U/µL reverse transcriptase, 0. U/µL RNase inhibitor, 0. mm dntps, and 1X reaction buffer were run in a total reaction volume of µl and incubated at 16 C for min, and 42 C for min, and 85 C for 5 min in a thermalcycler. PCR Reaction Following the RT step, 0.8 µl of the RT reaction was combined with 0.5 µl of a TaqMan MicroRNA Assay (X; forward primer, reverse primer, and probe) and 5 µl of TaqMan Universal PCR Master Mix, No AmpErase UNG (PN ) in a -µl final volume. Real-time PCR was performed using an Applied Biosystems 7900HT Fast Real-Time PCR System with cycling conditions of 95 C for min [followed by 95 C for sec and 60 C for 60 sec] for a total of 40 cycles. Each TaqMan Assay was run in quadruplicate. Results and Discussion TaqMan MicroRNA Assays were designed and synthesized for a total of 38 human snrna and snorna (sn/snorna) and trna genes. The standard TaqMan MicroRNA Assays Protocol (PN ) was followed to examine performance of the sn/snorna endogenous control candidate assays using, 3, and 0.3 ng of human lung total RNA. For each assay, a paired no-template control (NTC) reaction was performed. A subset of 23 sn/snorna endogenous control candidates that demonstrated good assay linearity (R 2 >0.96) and abundance (data not shown), and had NTC C T value >38, was identified and used to screen a panel of 38 normal human tissues (Table 2, see appendix) and 59 NCI-60 cell lines (Table 3, see appendix). These control candidates showed expression levels that remained relatively constant across the tissues and cell lines tested. A final group of ten small RNAs (Table 1), including both snrnas and snornas, was selected as the best performing control assays, based upon their expression level and stability, as determined by statistical analysis. These ten sn/snorna endogenous control genes demonstrated expression levels that were both relatively abundant (C T 35 RNU24 Z RNU66 RNU6B RNU19 RNU48 RNU38B RNU44 RNU49 RNU43 35 RNU24 Z RNU66 RNU6B RNU19 RNU48 RNU38B RNU44 RNU49 RNU43 Ileum Jejunum Duodenum Proximal Colon Distal Colon Esophagus Trachea Vena Cava Pericardium Left Atrium Left Ventricle Right Atrium Right Ventricle Fallopian Tube Thyroid Uterus Lymph Node Placenta Breast Pancreas Adipose Liver Brain Thymus Heart Lung Spleen Testicle Ovary Kidney Skeletal Muscle Small Intestine Colon Prostate Bladder Cervix Adrenal Stomach NCI-60 Cell Line Human Tissues Figure 1a. Expression profile of ten human sn/snornas for mirna endogenous controls across 38 normal human tissues. Figure 1b. Expression profile of ten human sn/snornas for mirna endogenous controls across 59 NCI-60 cell lines.
3 range ) and constant across all 38 human tissues (Figure 1a) or 59 NCI-60 cell lines (Figure 1b) tested, with standard deviations for the values ranging from 0.7 to 1.1 across tissues (Table 4, see appendix), and 1.0 to 1.4 across cell lines (Table 5, see appendix). In addition to these ten human controls, we identified an additional eight controls (Table 1). They also displayed good linearity (R 2 >0.96), good abundance ( range 22 28) and NTC C T >38. These controls were also tested across the 38 tissues (Figure 2). The and StDev of the for these controls are shown in Table 4. To compare the use of various normalization control classes, the expression patterns for sn/snorna endogenous controls described above were compared to mirna control candidates and the normalization controls typically used for conventional TaqMan Assays. To identify mirna control candidates, the expression levels of 247 mirnas were examined using TaqMan MicroRNA Assays across all 38 normal human tissues and 59 NCI-60 cell lines (data not shown). The results showed constant expression for a subset of these mirnas in tissues (hsa-mir-26b, hsa-mir-92, hsa-mir-92n) and cell lines (hsa-mir-423, hsa-mir-374, hsa-mir-16), indicating that within this group there may be good endogenous control candidates (Figure 4a and 4b; and Table 6, see appendix). The use of the most stably expressed mirna(s) for a specific experimental condition to normalize mirna expression data is a commonly used approach and poses an alternative to using the sn/snorna endogenous controls previously described. For comparison, the expression patterns of the sn/snorna and mirna control candidates and traditional TaqMan Assay controls were plotted together. The C T values for the three most stable assays for sn/snorna and mirna were averaged and compared to 18S rrna, GAPDH and ACTB. The 18S rrna, sn/snorna and mirna exhibit similar patterns and show relatively constant expression across all tissues (Figure 5a) and cell lines (Figure 5b), although 18S rrna is expressed at a significantly higher level. Of the 18 human controls identified, RNU48, RNU44, U18, and U47 are the most highly abundant across the tissues based on. In addition to RNU44 and RNU48, the following: RNU24, RNU49, U75, and U47 display the least variability across the tissue samples, with StDev of the average C T between 0.7 and 0.8. RNU48 and RNU44 also showed the highest abundance across the NCI-60 cell lines, although they showed highest variability compared to the other controls tested. Thus, if you were comparing mirna expression across the 38 tissue samples, RNU48 or RNU44 would be considered good candidates for endogenous controls because they show good abundance and relatively stable expression. However, if your study were across the NCI-60 cell line, you may consider one of the other controls such as RNU6B, Z, or RNU38B, as RNU44 and RNU48 display greater variability. Mouse Endogenous Controls A similar approach was taken to identify mouse-specific sn/snorna endogenous controls. As with the human selections, a number of candidate mouse snornas were tested across various tissues, and those showing relatively high abundance and the least variation were chosen. These experiments identified five mouse snornas (Table 1) that fit the same criteria set for the human endogenous controls. Figure 3 shows the expression profile across 12 tissues for these five mouse snornas, U18 U54 RNU58B HY3 RNU58A U75 RPL21 U47 snorna135 snorna142 snorna2 snorna234 snorna Ileum Jejunum Duodenum Proximal Colon Distal Colon Esophagus Trachea Vena Cava Pericardium Left Atrium Left Ventricle Right Atrium Right Ventricle Fallopian Tube Thyroid Uterus Lymph Node Placenta Breast Pancreas Adipose Liver Brain Thymus Heart Lung Spleen Testicle Ovary Kidney Skeletal Muscle Small Intestine Colon Prostate Bladder Cervix Adrenal Stomach Human Tissues M. Stomach M. Bladder M. Placenta M. Colon M. Mamm. Gld M. Ovary M. Brain Mouse Tissue M. Lung M. Kidney M. Heart M. Liver M. Embryo Figure 2. Expression profile of eight additional human sn/snornas for mirna endogenous controls across 38 normal human tissues. Figure 3. Expression profile of five mouse sn/snornas for mirna normalization across 12 normal tissues.
4 including the, and Table 7 (see appendix) shows StDev of the average C T. Mouse snorna2 showed the highest abundance and least variability across the 12 tissues, making it the best candidate if your study were to examine mirna expression across these 12 tissues. Recommendations for mirna Expression Data Analysis This study confirms that both 18S rrna and a group of 23 small ncrnas not related to mirnas are invariantly expressed across a relatively broad tissue panel. Additionally, we demonstrate that specific mirna species that are similarly invariantly expressed across the same panel of samples can be identified. These results suggest that effective data normalization can be achieved using a variety of endogenous controls; however, there are significant advantages to using the sn/snorna endogenous controls for normalizing TaqMan MicroRNA Assay expression data: snrna and snorna genes are closer in size (length) to mirnas (<0 bp) snrnas and snornas are constitutively and abundantly expressed across a large number of tissues and cell lines snrna, snorna, and mirna assays were designed using identical approaches snrna and snornas are unlikely to be involved in mirna regulatory pathways Applied Biosystems offers the 18 human snrna and snorna and five mouse snorna endogenous control assays (Table 1) that are recommended for normalizing human and mouse TaqMan MicroRNA Assays. Regardless of the gene or gene set that is chosen, we highly recommend that the consistency of expression be reconfirmed under the specific conditions of the experiment. The following steps are recommended when selecting endogenous controls for mirna data normalization: 1. Carefully select a set of several endogenous control genes based on the species, tissues, or cell lines used in your study. This will often require screening available TaqMan MicroRNA Assays for those sn/ snorna endogenous controls that perform best in the specific samples under investigation. Alternatively, or in addition to, use specifc mirnas that demonstrate the least variability across experimental conditions under consideration. From the previously described study, based upon data generated across a wide variety of tissues and cell lines, we have identified the following candidate control genes as showing the least variability: sn/snornas: - Human: RNU48, RNU44, U47, RNU6B, or all four - Mouse: snorna2, snorna234, or both mirnas: - Tissues: hsa-mir-26b, hsa-mir-92, hsa-mir-92n - Cell lines: hsa-mir-423, hsa-mir- 374, hsa-mir-16 Traditional TaqMan Assay controls: 18S rrnas 2. Normalize your C T values using the of the endogenous controls 3. Use C T (mirna C T averaged endogenous control C T ) or fold-change relative to a calibrator or reference sample (2 C T) for relative expression analysis hsa-mir-26b hsa-mir-92 hsa-mir-92n hsa-mir-423 hsa-mir-374 hsa-mir Ileum Jejunum Duodenum Proximal Colon Distal Colon Esophagus Trachea Vena Cava Pericardium Left Atrium Left Ventricle Right Atrium Right Ventricle Fallopian Tube Thyroid Uterus Lymph Node Placenta Breast Pancreas Adipose Liver Brain Thymus Heart Lung Spleen Testicle Ovary Kidney Skeletal Muscle Small Intestine Colon Prostate Bladder Cervix Adrenal Stomach NCI-60 Cell Line Human Tissues Figure 4a. Expression profile of three human mirna candidates for mirna normalization across 38 normal human tissues. Figure 4b. Expression profile of three human mirna candidates for mirna normalization across 59 NCI-60 cell lines.
5 Conclusion We have identified a set of 18 endogenous control candidates in human, and five in mouse, that can be used to normalize mirna gene expression data. In this study, we found the following six sn/snorna endogenous controls show the highest abundance and least variability across normal tissues and cell lines that were tested: Human: RNU48, RUN44, U47, and RNU6B; Mouse: snorna2 and snorna234. Stable expression patterns across tissues and cell lines were also observed for specific mirna control candidates and 18S rrna, providing additional normalization options. Authors Linda Wong, Kathy Lee, Iain Russell, and Caifu Chen of Applied Biosystems References 1. Calin, G.A. and Croce, G.A. 06. MicroRNA signatures in human cancers. Nat Rev Cancer. 6: Finnegan, E.J. and Matzke, M.A. 03. The small RNA world. J Cell Sci 116: Chen, C., Ridzon, D.A., Broomer, A.J., et al. 05. Real-time quantification of micrornas by stem-loop RT-PCR. Nucleic Acids Res. 33, e De Kok, J.B., Roelofs, R.W., Giesendorf, B.A., et al. 05. Normalization of gene expression measurements in tumor tissues: comparison of 13 endogenous control genes. Laboratory Investigation 85: Suzuki, T., Higgins, P.J., and Crawford, D.R. 00. Control selection for RNA quantitation. Biotechniques 29: Kiss, T. 02. Small nucleolar RNAs: an abundant group of noncoding RNAs with diverse cellular functions. Cell 9: Chen, Z., Zhang, J., Kong, J., et al. 06. Diversity of endogenous small non-coding RNAs in Oryza sativa. Genetica 128: Fedorov, A., Stombaugh, J., Harr, M.W., et al. 05. Computer identification of snorna genes using a Mammalian Orthologous Intron Database. Nucleic Acids Res. 33: Eddy, S.R. 01. Non-coding RNA genes and the modern RNA world. Nat Rev Genet. 2: Blencowe, B.J. 02. Transcription: surprising role for an elusive small nuclear RNA. Curr Biol. 12:R Das, G., Henning, D., and Reddy, R Structure, organization, and transcription of Drosophila U6 small nuclear RNA genes. J Biol Chem. 262: ACTB GAPDH 18S rrna sn/snorna mirna ACTB GAPDH 18S rrna sn/snorna mirna Ileum Jejunum Duodenum Proximal Colon Distal Colon Esophagus Trachea Vena Cava Pericardium Left Atrium Left Ventricle Right Atrium Right Ventricle Fallopian Tube Thyroid Uterus Lymph Node Placenta Breast Pancreas Adipose Liver Brain Thymus Heart Lung Spleen Testicle Ovary Kidney Skeletal Muscle Small Intestine Colon Prostate Bladder Cervix Adrenal Stomach NCI-60 Cell Line Human Tissues Figure 5a. Comparison of the expression pattern for different types of normalization controls across 38 normal human tissues: sn/snorna, mirnas, and TaqMan Assay controls. Figure 5b. Comparison of the expression pattern for various types of normalization controls across 59 NCI-60 cell lines: sn/snorna, mirnas, and TaqMan Assay controls.
6 Appendix TABLE 1. List of human and mouse mirna endogenous controls (sn/snornas). Human sn/snornas, highlighted in dark green, were tested with tissue and cell lines. Human sn/snornas, highlighted in light green, were tested with tissues only. Mouse sn/snornas, highlighted in blue, were tested only with tissues. Human Gene (NCBI Symbol) Gene Name NCBI Accession Alias Product Name AB Assay ID Part Number Tissue NCI-60 SNORD24 small nucleolar RNA, C/D box 24 Z48765 U24; RNU24 RNU Yes Yes SNORA66 small nucleolar RNA, H/ACA box 66 NR_ U66; RNU66 RNU Yes Yes SNORA74A small nucleolar RNA, H/ACA box 74A X94290 U19; RNU19 RNU Yes Yes SNORD38B small nucleolar RNA, C/D box 38B NR_ U38B; RNU38B RNU38B Yes Yes SNORD49A small nucleolar RNA, C/D box 49A NR_ U49; U49A; RNU49 RNU Yes Yes SNORD48 small nucleolar RNA, C/DR box 48 NR_ U48; RNU48 RNU Yes Yes SNORD7 small nucleolar RNA, C/D box 7 AJ Zn0; mgu6-47 Z Yes Yes RNU6B U6B, small nuclear NR_ U6 RNU6B Yes Yes SNORD44 small nucleolar RNA, C/D box 44 NR_ U44; RNU44 RNU Yes Yes SNORD43 small nucleolar RNA, C/D box 43 NR_ U43; RNU43 RNU Yes Yes NA* U18 snorna/rpl4 1 AB U18 U Yes No SNORD58B small nucleolar RNA, C/D box 58B NR_0072 U58b; RNU58B RNU58B Yes No SNORD58A small nucleolar RNA, C/D box 58A NR_0071 U58a; RNU58A RNU58A Yes No NA* RPL21/snoRNA 2 AB NA RPL Yes No SNORD54 small nucleolar RNA, C/D box 54 NR_ U54; RU54 U Yes No RNY3 Y3 small cytoplasmic (associated with Ro protein) AC HY3; Y3 HY Yes No SNORD75 small nucleolar RNA, C/D box 75 AF U75 U Yes No SNORD47 small nucleolar RNA, C/D box 47 AF U47; RNU47 U Yes No Mouse Gene Gene Name NCBI Accession Alias Product Name AB Assay ID Part Number Tissue NCI-60 snorna135 clone MBII-135 C/D box snorna AF snorna135 snorna Yes No snorna142 clone MBII-142 C/D box snorna AF snorna142 snorna Yes No snorna2 clone MBII-2 C/D box snorna AF snorna2 snorna Yes No snorna234 clone MBII-234 C/D box snorna AF snorna234 snorna Yes No snorna1 clone MBII-1 C/D box snorna AF snorna1 snorna Yes No *No Gene Symbol or Name cited on NCBI 1. U18 snorna is located within RPL4 gene (intron) 2. snorna is located within RPL21 gene (intron) 3. DNA accession number
7 TABLE 2. Human and mouse tissues and part numbers. Human Tissue Ambion PN Human Tissue Ambion PN TABLE 3. Key for NCI-60 cell line. The number corresponds to the cell line in Figures 1b, 3b, and 5. lleum 6828 Jejunum 68 Duodenum 6832 Proximal Colon 6834 Distal Colon 6836 Esophagus 6842 Trachea 6846 Vena Cava 6848 Pericardium 6852 Left Atrium 6854 Left Ventricle 6856 Right Atrium 6858 Right Ventricle 6860 Fallopian Tube 6862 Thyroid 6872 Uterus 7892 Lymph Node 7894 Placenta 7950 Breast 6952 Pancreas 7954 Adipose 7956 Liver 7960 Brain 7962 Thymus 7964 Heart 7966 Lung 7968 Spleen 7970 Testicle 7972 Ovary 6974 Kidney 7976 Skeletal Muscle 7982 Small Intestine 7984 Colon 7986 Prostate 7988 Bladder 7990 Cervix 6992 Adrenal 7994 Stomach 7996 Mouse Tissue Ambion PN Ovary 7824 Brain 7812 Lung 7818 Kidney 7826 Heart 7816 Liver 78 Embryo 7828 Mouse Tissue BD Biosciences PN Stomach Bladder Placenta Colon Mammary Gland Cell Line Cell Line 1. SNB MDA MB HCT SK-MEL-5 3. EKVX 33. NCI-H23 4. BT HOP A549 ATCC 35. OVCAR-8 6. TK- 36. HS LOXIMVI 37. SK-MEL-2 8. NCI-ADR-RES 38. COLO-5 9. SK-OV K562. T-47D 40. HL-60TB 11. RXF HCC OVCAR MAL-ME-3M 13. PC CCRF-CEM 14. SF A498. SF SN OVCAR MCF7 17. SK-MEL IGROV ACHN 48. OVCAR CAKI MOLT4. UACC RPMI KM SW U1 52. HCT- 23. NCI-H SNB NCI-H322M. DU SF NCI-H UO NCI-H HT SR 58. HOP UACC M14. MDA MB 231
8 table 4. and StDev of the for 18 sn/snornas for mirna endogenous controls across 38 human tissues. table 5. and StDev of the for ten sn/snornas for mirna endogenous controls across NCI- 60 cell lines. table 6. and StDev of the for three mirnas across 38 human tissues and three mirnas NCI- 60 cell lines. across 38 tissues across 59 nci-60 cell lines across 38 tissues Control Control RNU RNU RNU RNU38B RNU RNU RNU RNU38B hsa-mir-26b hsa-mir hsa-mir-92n across nci-60 cell line RNU RNU Z RNU6B RNU RNU RNU U RNU58B RNU58A RPL U HY U U Z RNU6B RNU RNU RNU hsa-mir hsa-mir hsa-mir table 7. Expression level and variation for five mouse snornas as endogenous controls in 12 tissues. across 12 Mouse tissues Control snorna snorna snorna snorna snorna For Research Use Only. Not for use in diagnostic procedures. NOTICE TO PURCHASER: LIMITED LICENSE A license to perform the patented 5 Nuclease Process for research is obtained by the purchase of (i) both Licensed Probe and Authorized 5 Nuclease Core Kit, (ii) a Licensed 5 Nuclease Kit, or (iii) license rights from Applied Biosystems. The TaqMan MicroRNA Assay contains Licensed Probe. Use of this product is covered by US patent claims and patent claims outside the US. The purchase of this product includes a limited, non-transferable immunity from suit under the foregoing patent claims for using only this amount of product for the purchaser s own internal research. Separate purchase of an Authorized 5 Nuclease Core Kit would convey rights under the applicable claims of US patents, and patent claims outside the United States, which claim 5 nuclease methods. No right under any other patent claim and no right to perform commercial services of any kind, including without limitation reporting the results of purchaser s activities for a fee or other commercial consideration, is conveyed expressly, by implication, or by estoppel. This product is for research use only. Diagnostic uses under Roche patents require a separate license from Roche. Further information on purchasing licenses may be obtained from the Director of Licensing, Applied Biosystems, 850 Lincoln Centre Drive, Foster City, California 94404, USA. 07, Applied Biosystems. All rights reserved. Information subject to change without notice. Applera, Applied Biosystems, and AB (Design) are registered trademarks and FAM is a trademark of Applied Biosystems or its subsidiaries in the US and/or certain other countries. AmpErase and TaqMan are registered trademarks of Roche Molecular Systems, Inc. All other trademarks are the sole property of their respective owners. Printed in the USA, 08/ Publication 127AP11-01 Headquarters 850 Lincoln Centre Drive Foster City, CA USA Phone Toll Free international Sales For our office locations please call the division headquarters or refer to our Web site at
Using TaqMan Endogenous Control Assays to select an endogenous control for experimental studies
 APPLICATION NOTE TaqMan Gene Expression Assays Using TaqMan Endogenous Control Assays to select an endogenous control for experimental studies Introduction Quantitative real-time PCR (qpcr) allows for
APPLICATION NOTE TaqMan Gene Expression Assays Using TaqMan Endogenous Control Assays to select an endogenous control for experimental studies Introduction Quantitative real-time PCR (qpcr) allows for
Review of AB-Ambion systems for microrna Isolation, Analysis and Regulation
 Review of AB-Ambion systems for microrna Isolation, Analysis and Regulation Marjorie JOINER Ingénieur Applications Applied Biosystems France Isabelle KAYSER Ingénieur Commercial Applied Biosystems France
Review of AB-Ambion systems for microrna Isolation, Analysis and Regulation Marjorie JOINER Ingénieur Applications Applied Biosystems France Isabelle KAYSER Ingénieur Commercial Applied Biosystems France
TaqMan Fast Advanced Master Mix. Protocol
 TaqMan Fast Advanced Master Mix Protocol For Research Use Only. Not intended for any animal or human therapeutic or diagnostic use. Information in this document is subject to change without notice. APPLIED
TaqMan Fast Advanced Master Mix Protocol For Research Use Only. Not intended for any animal or human therapeutic or diagnostic use. Information in this document is subject to change without notice. APPLIED
Essentials of Real Time PCR. About Sequence Detection Chemistries
 Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Micro RNAs: potentielle Biomarker für das. Blutspenderscreening
 Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Megaplex Pools For microrna Expression Analysis
 Megaplex Pools For microrna Expression Analysis Protocol Copyright 2008, 2010 Applied Biosystems. All rights reserved. For Research Use Only. Not for use in diagnostic procedures. Information in this document
Megaplex Pools For microrna Expression Analysis Protocol Copyright 2008, 2010 Applied Biosystems. All rights reserved. For Research Use Only. Not for use in diagnostic procedures. Information in this document
MystiCq microrna cdna Synthesis Mix Catalog Number MIRRT Storage Temperature 20 C
 microrna cdna Synthesis Mix Catalog Number MIRRT Storage Temperature 20 C Product Description The microrna cdna Synthesis Mix has been designed to easily convert micrornas into cdna templates for qpcr
microrna cdna Synthesis Mix Catalog Number MIRRT Storage Temperature 20 C Product Description The microrna cdna Synthesis Mix has been designed to easily convert micrornas into cdna templates for qpcr
Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr
 User Manual Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories,
User Manual Mir-X mirna First-Strand Synthesis and SYBR qrt-pcr User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories,
Protocol. Introduction to TaqMan and SYBR Green Chemistries for Real-Time PCR
 Protocol Introduction to TaqMan and SYBR Green Chemistries for Real-Time PCR Copyright 2008, 2010 Applied Biosystems. All rights reserved. Ambion and Applied Biosystems products are for Research Use Only.
Protocol Introduction to TaqMan and SYBR Green Chemistries for Real-Time PCR Copyright 2008, 2010 Applied Biosystems. All rights reserved. Ambion and Applied Biosystems products are for Research Use Only.
Factors Influencing Multiplex Real-Time PCR
 APPLICATION NOTE Multiplex Real-Time PCR Factors Influencing Multiplex Real-Time PCR Introduction Multiplex PCR is the simultaneous amplification of more than one target sequence in a single reaction [1].
APPLICATION NOTE Multiplex Real-Time PCR Factors Influencing Multiplex Real-Time PCR Introduction Multiplex PCR is the simultaneous amplification of more than one target sequence in a single reaction [1].
Quantifiler Human DNA Quantification Kit Quantifiler Y Human Male DNA Quantification Kit
 Product Bulletin Human Identification Quantifiler Human DNA Quantification Kit Quantifiler Y Human Male DNA Quantification Kit The Quantifiler kits produce reliable and reproducible results, helping to
Product Bulletin Human Identification Quantifiler Human DNA Quantification Kit Quantifiler Y Human Male DNA Quantification Kit The Quantifiler kits produce reliable and reproducible results, helping to
Gene Expression Assays
 APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
Real-time PCR: Understanding C t
 APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
Path-ID Multiplex One-Step RT-PCR Kit
 USER GUIDE Path-ID Multiplex One-Step RT-PCR Kit TaqMan probe-based multiplex one-step real-time RT-PCR detection of RNA targets Catalog Numbers 4428206, 4428207, 4440022 Publication Part Number 4440907
USER GUIDE Path-ID Multiplex One-Step RT-PCR Kit TaqMan probe-based multiplex one-step real-time RT-PCR detection of RNA targets Catalog Numbers 4428206, 4428207, 4440022 Publication Part Number 4440907
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual
 Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Relative Quantification of mirna Target mrnas by Real-Time qpcr. 1 Introduction. Gene Expression Application Note No. 4
 Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
Real-Time Quantitative PCR Instruments and Consumables
 Index Page Sequence Detection Systems 2 Sequence Detection Systems Accessories and Softwares 3 Sequence Detection PCR Reagent Kits 4 Sequence Detection PCR Reagents 9 Sequence Detection Systems Disposables
Index Page Sequence Detection Systems 2 Sequence Detection Systems Accessories and Softwares 3 Sequence Detection PCR Reagent Kits 4 Sequence Detection PCR Reagents 9 Sequence Detection Systems Disposables
Mir-X mirna First-Strand Synthesis Kit User Manual
 User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals
 Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
SYBR Green Realtime PCR Master Mix -Plus-
 Instruction manual SYBR Green Realtime PCR Master Mix -Plus- 0810 F0925K SYBR Green Realtime PCR Master Mix -Plus- Contents QPK-212T 1mLx1 QPK-212 1mLx5 Store at -20 C, protected from light [1] Introduction
Instruction manual SYBR Green Realtime PCR Master Mix -Plus- 0810 F0925K SYBR Green Realtime PCR Master Mix -Plus- Contents QPK-212T 1mLx1 QPK-212 1mLx5 Store at -20 C, protected from light [1] Introduction
Absolute Quantification Getting Started Guide
 5 cdna Reverse Primer Oligo d(t) or random hexamer Synthesis of 1st cdna strand 3 5 cdna Applied Biosystems 7300/7500/7500 Fast Real-Time PCR System Absolute Quantification Getting Started Guide Introduction
5 cdna Reverse Primer Oligo d(t) or random hexamer Synthesis of 1st cdna strand 3 5 cdna Applied Biosystems 7300/7500/7500 Fast Real-Time PCR System Absolute Quantification Getting Started Guide Introduction
miscript mirna PCR Array Handbook
 May 2012 miscript mirna PCR Array Handbook miscript II RT Kit miscript SYBR Green PCR Kit miscript mirna PCR Arrays miscript mirna QC PCR Array For SYBR Green-based, real-time PCR profiling of micrornas
May 2012 miscript mirna PCR Array Handbook miscript II RT Kit miscript SYBR Green PCR Kit miscript mirna PCR Arrays miscript mirna QC PCR Array For SYBR Green-based, real-time PCR profiling of micrornas
2.500 Threshold. 2.000 1000e - 001. Threshold. Exponential phase. Cycle Number
 application note Real-Time PCR: Understanding C T Real-Time PCR: Understanding C T 4.500 3.500 1000e + 001 4.000 3.000 1000e + 000 3.500 2.500 Threshold 3.000 2.000 1000e - 001 Rn 2500 Rn 1500 Rn 2000
application note Real-Time PCR: Understanding C T Real-Time PCR: Understanding C T 4.500 3.500 1000e + 001 4.000 3.000 1000e + 000 3.500 2.500 Threshold 3.000 2.000 1000e - 001 Rn 2500 Rn 1500 Rn 2000
Relative Quantification Applied Biosystems 7300/7500 Real Time PCR System
 5 3 5 cdna Reverse Primer Oligo d(t) or random hexamer 3 5 cdna Getting Started Guide Relative Quantification Applied Biosystems 7300/7500 Real Time PCR System Introduction and Example RQ Experiment Designing
5 3 5 cdna Reverse Primer Oligo d(t) or random hexamer 3 5 cdna Getting Started Guide Relative Quantification Applied Biosystems 7300/7500 Real Time PCR System Introduction and Example RQ Experiment Designing
Stratagene QPCR Mouse Reference Total RNA
 Stratagene QPCR Mouse Reference Total RNA Instruction Manual Catalog #750600 Revision C.0 For Research Use Only. Not for use in diagnostic procedures. 750600-12 LIMITED PRODUCT WARRANTY This warranty limits
Stratagene QPCR Mouse Reference Total RNA Instruction Manual Catalog #750600 Revision C.0 For Research Use Only. Not for use in diagnostic procedures. 750600-12 LIMITED PRODUCT WARRANTY This warranty limits
Replacing TaqMan SNP Genotyping Assays that Fail Applied Biosystems Manufacturing Quality Control. Begin
 User Bulletin TaqMan SNP Genotyping Assays May 2008 SUBJECT: Replacing TaqMan SNP Genotyping Assays that Fail Applied Biosystems Manufacturing Quality Control In This Bulletin Overview This user bulletin
User Bulletin TaqMan SNP Genotyping Assays May 2008 SUBJECT: Replacing TaqMan SNP Genotyping Assays that Fail Applied Biosystems Manufacturing Quality Control In This Bulletin Overview This user bulletin
amplification tech Optical Design of CFX96 Real-Time PCR Detection System Eliminates the Requirement of a Passive Reference Dye
 amplification tech note 6047 Optical Design of CFX96 Real-Time PCR Detection System Eliminates the Requirement of a Passive Reference Dye Liz Jordan and Richard Kurtz, Gene Expression Division, io-rad
amplification tech note 6047 Optical Design of CFX96 Real-Time PCR Detection System Eliminates the Requirement of a Passive Reference Dye Liz Jordan and Richard Kurtz, Gene Expression Division, io-rad
CompleteⅡ 1st strand cdna Synthesis Kit
 Instruction Manual CompleteⅡ 1st strand cdna Synthesis Kit Catalog # GM30401, GM30402 Green Mountain Biosystems. LLC Web: www.greenmountainbio.com Tel: 800-942-1160 Sales: Sales@ greenmountainbio.com Support:
Instruction Manual CompleteⅡ 1st strand cdna Synthesis Kit Catalog # GM30401, GM30402 Green Mountain Biosystems. LLC Web: www.greenmountainbio.com Tel: 800-942-1160 Sales: Sales@ greenmountainbio.com Support:
User Bulletin #2 ABI PRISM 7700 Sequence Detection System
 User Bulletin #2 ABI PRISM 7700 Sequence Detection System December 11, 1997 (updated 10/2001) SUBJECT: Relative Quantitation of Gene Expression Introduction Amplification of an endogenous control may be
User Bulletin #2 ABI PRISM 7700 Sequence Detection System December 11, 1997 (updated 10/2001) SUBJECT: Relative Quantitation of Gene Expression Introduction Amplification of an endogenous control may be
BIO 137: CHAPTER 1 OBJECTIVES
 BIO 137: CHAPTER 1 OBJECTIVES 1. Define the terms anatomy and physiology, and explain their relationship using an example of a human structure with its corresponding function. A. ANATOMY = the study of
BIO 137: CHAPTER 1 OBJECTIVES 1. Define the terms anatomy and physiology, and explain their relationship using an example of a human structure with its corresponding function. A. ANATOMY = the study of
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
All-in-One mirna qrt-pcr Reagent Kits For quantitative detection of mature mirna
 All-in-One mirna qrt-pcr Reagent Kits For quantitative detection of mature mirna All-in-One TM mirna First-Strand cdna Synthesis Kit AMRT-0020 (20 RT reactions), AMRT-0060 (60 RT reactions) Used in combination
All-in-One mirna qrt-pcr Reagent Kits For quantitative detection of mature mirna All-in-One TM mirna First-Strand cdna Synthesis Kit AMRT-0020 (20 RT reactions), AMRT-0060 (60 RT reactions) Used in combination
Creating Standard Curves with Genomic DNA or Plasmid DNA Templates for Use in Quantitative PCR
 Creating Standard Curves with Genomic DNA or Plasmid DNA Templates for Use in Quantitative PCR Overview Genomic DNA (gdna) and plasmids containing cloned target sequences are commonly used as standards
Creating Standard Curves with Genomic DNA or Plasmid DNA Templates for Use in Quantitative PCR Overview Genomic DNA (gdna) and plasmids containing cloned target sequences are commonly used as standards
SeqScape Software Version 2.5 Comprehensive Analysis Solution for Resequencing Applications
 Product Bulletin Sequencing Software SeqScape Software Version 2.5 Comprehensive Analysis Solution for Resequencing Applications Comprehensive reference sequence handling Helps interpret the role of each
Product Bulletin Sequencing Software SeqScape Software Version 2.5 Comprehensive Analysis Solution for Resequencing Applications Comprehensive reference sequence handling Helps interpret the role of each
Introduction To Real Time Quantitative PCR (qpcr)
 Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Comprehensive mirna Research Technologies
 Comprehensive mirna Research Technologies Sample & Assay Technologies QIAGEN solutions for advancing microrna research In the last few years, the identification of microrna (mirna) and the recognition
Comprehensive mirna Research Technologies Sample & Assay Technologies QIAGEN solutions for advancing microrna research In the last few years, the identification of microrna (mirna) and the recognition
TaqMan MicroRNA Assays
 TaqMan MicroRNA Assays Protocol December 23, 2005 11:22 am, mirna_title.fm Copyright 2005-6, Applied Biosystems. All rights reserved. For Research Use Only. Not for use in diagnostic procedures. Information
TaqMan MicroRNA Assays Protocol December 23, 2005 11:22 am, mirna_title.fm Copyright 2005-6, Applied Biosystems. All rights reserved. For Research Use Only. Not for use in diagnostic procedures. Information
Profiling of microrna in Blood Serum/Plasma. Guidelines for the mircury LNA TM Universal RT microrna PCR System
 Profiling of microrna in Blood Serum/Plasma Guidelines for the mircury LNA TM Universal RT microrna PCR System Table of Contents 2 Introduction.....................................................................................
Profiling of microrna in Blood Serum/Plasma Guidelines for the mircury LNA TM Universal RT microrna PCR System Table of Contents 2 Introduction.....................................................................................
Relative Quantitation Using Comparative C T Getting Started Guide
 Applied Biosystems 7900HT Fast Real-Time PCR System Relative Quantitation Using Comparative C T Getting Started Guide Introduction N Designing an RQ Experiment Preparing Samples and Reaction Plates Creating
Applied Biosystems 7900HT Fast Real-Time PCR System Relative Quantitation Using Comparative C T Getting Started Guide Introduction N Designing an RQ Experiment Preparing Samples and Reaction Plates Creating
Global MicroRNA Amplification Kit
 Global MicroRNA Amplification Kit Store kit at -20 C on receipt (ver. 3-060901) A limited-use label license covers this product. By use of this product, you accept the terms and conditions outlined in
Global MicroRNA Amplification Kit Store kit at -20 C on receipt (ver. 3-060901) A limited-use label license covers this product. By use of this product, you accept the terms and conditions outlined in
RevertAid Premium First Strand cdna Synthesis Kit
 RevertAid Premium First Strand cdna Synthesis Kit #K1651, #K1652 CERTIFICATE OF ANALYSIS #K1651 Lot QUALITY CONTROL RT-PCR using 100 fg of control GAPDH RNA and GAPDH control primers generated a prominent
RevertAid Premium First Strand cdna Synthesis Kit #K1651, #K1652 CERTIFICATE OF ANALYSIS #K1651 Lot QUALITY CONTROL RT-PCR using 100 fg of control GAPDH RNA and GAPDH control primers generated a prominent
mirnaselect pep-mir Cloning and Expression Vector
 Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
ONLINE SUPPLEMENTAL MATERIAL. Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue
 ONLINE SUPPLEMENTAL MATERIAL Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue Sungmi Park 1, Ko-Ting Lu 1, Xuebo Liu 1, Tapan K. Chatterjee 2, Steven M. Rudich 3, Neal
ONLINE SUPPLEMENTAL MATERIAL Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue Sungmi Park 1, Ko-Ting Lu 1, Xuebo Liu 1, Tapan K. Chatterjee 2, Steven M. Rudich 3, Neal
Speed Matters - Fast ways from template to result
 qpcr Symposium 2007 - Weihenstephan Speed Matters - Fast ways from template to result March 28, 2007 Dr. Thorsten Traeger Senior Scientist, Research and Development - 1 - Overview Ạgenda Fast PCR The Challenges
qpcr Symposium 2007 - Weihenstephan Speed Matters - Fast ways from template to result March 28, 2007 Dr. Thorsten Traeger Senior Scientist, Research and Development - 1 - Overview Ạgenda Fast PCR The Challenges
Quantification of Hepatitis B Virus. Core Protein Region. 150 tests
 Quantification of Hepatitis B Virus Core Protein Region For general laboratory and research use only 150 tests Introduction to Hepatitis B Virus Originally known as serum hepatitis, hepatitis B has only
Quantification of Hepatitis B Virus Core Protein Region For general laboratory and research use only 150 tests Introduction to Hepatitis B Virus Originally known as serum hepatitis, hepatitis B has only
TaqMan Gene Expression Assays
 Product Guide TaqMan Gene Expression Assay Products Product Guide TaqMan Gene Expression Assays Applied Biosystems TaqMan Gene Expression Assays Solutions Applied Biosystems offers the largest family of
Product Guide TaqMan Gene Expression Assay Products Product Guide TaqMan Gene Expression Assays Applied Biosystems TaqMan Gene Expression Assays Solutions Applied Biosystems offers the largest family of
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
Real-time quantitative RT -PCR (Taqman)
 Real-time quantitative RT -PCR (Taqman) Author: SC, Patti Lab, 3/03 This is performed as a 2-step reaction: 1. cdna synthesis from DNase 1-treated total RNA 2. PCR 1. cdna synthesis (Advantage RT-for-PCR
Real-time quantitative RT -PCR (Taqman) Author: SC, Patti Lab, 3/03 This is performed as a 2-step reaction: 1. cdna synthesis from DNase 1-treated total RNA 2. PCR 1. cdna synthesis (Advantage RT-for-PCR
Real-Time PCR Vs. Traditional PCR
 Real-Time PCR Vs. Traditional PCR Description This tutorial will discuss the evolution of traditional PCR methods towards the use of Real-Time chemistry and instrumentation for accurate quantitation. Objectives
Real-Time PCR Vs. Traditional PCR Description This tutorial will discuss the evolution of traditional PCR methods towards the use of Real-Time chemistry and instrumentation for accurate quantitation. Objectives
Application Guide... 2
 Protocol for GenomePlex Whole Genome Amplification from Formalin-Fixed Parrafin-Embedded (FFPE) tissue Application Guide... 2 I. Description... 2 II. Product Components... 2 III. Materials to be Supplied
Protocol for GenomePlex Whole Genome Amplification from Formalin-Fixed Parrafin-Embedded (FFPE) tissue Application Guide... 2 I. Description... 2 II. Product Components... 2 III. Materials to be Supplied
SNPbrowser Software v3.5
 Product Bulletin SNP Genotyping SNPbrowser Software v3.5 A Free Software Tool for the Knowledge-Driven Selection of SNP Genotyping Assays Easily visualize SNPs integrated with a physical map, linkage disequilibrium
Product Bulletin SNP Genotyping SNPbrowser Software v3.5 A Free Software Tool for the Knowledge-Driven Selection of SNP Genotyping Assays Easily visualize SNPs integrated with a physical map, linkage disequilibrium
Gene Expression Assay Performance Guaranteed With the TaqMan Assays QPCR Guarantee Program
 WHITE PAPER TaqMan Assays QPCR Guarantee Program Gene Expression Assay Performance Guaranteed With the TaqMan Assays QPCR Guarantee Program Real-Time PCR for the Quantification of Gene Expression Real-time
WHITE PAPER TaqMan Assays QPCR Guarantee Program Gene Expression Assay Performance Guaranteed With the TaqMan Assays QPCR Guarantee Program Real-Time PCR for the Quantification of Gene Expression Real-time
TaqMan Small RNA Assays TaqMan MicroRNA Assays TaqMan sirna Assays Custom TaqMan Small RNA Assays. Protocol
 TaqMan Small RNA Assays TaqMan MicroRNA Assays TaqMan sirna Assays Custom TaqMan Small RNA Assays Protocol For Research Use Only. Not intended for any animal or human therapeutic or diagnostic use. Information
TaqMan Small RNA Assays TaqMan MicroRNA Assays TaqMan sirna Assays Custom TaqMan Small RNA Assays Protocol For Research Use Only. Not intended for any animal or human therapeutic or diagnostic use. Information
Power SYBR Green PCR Master Mix and Power SYBR Green RT-PCR Reagents Kit
 USER GUIDE Power SYBR Green PCR Master Mix and Power SYBR Green RT-PCR Reagents Kit Catalog Number 4368577, 4367659, 4367660, 4368706, 4368702, 4368708 (Master Mix) and 4368711 (RT-PCR Reagents Kit) Publication
USER GUIDE Power SYBR Green PCR Master Mix and Power SYBR Green RT-PCR Reagents Kit Catalog Number 4368577, 4367659, 4367660, 4368706, 4368702, 4368708 (Master Mix) and 4368711 (RT-PCR Reagents Kit) Publication
DNA Integrity Number (DIN) For the Assessment of Genomic DNA Samples in Real-Time Quantitative PCR (qpcr) Experiments
 DNA Integrity Number () For the Assessment of Genomic DNA Samples in Real-Time Quantitative PCR (qpcr) Experiments Application Note Nucleic Acid Analysis Author Arunkumar Padmanaban Agilent Technologies,
DNA Integrity Number () For the Assessment of Genomic DNA Samples in Real-Time Quantitative PCR (qpcr) Experiments Application Note Nucleic Acid Analysis Author Arunkumar Padmanaban Agilent Technologies,
bitter is de pil Linos Vandekerckhove, MD, PhD
 4//24 Current HIV care HIV copies/ ml plasma Viral load Welcome to the Digital droplet PCR age! bitter is de pil Linos Vandekerckhove, MD, PhD Latent HIV reservoir Time at Ghent University Hospital 2 HIV
4//24 Current HIV care HIV copies/ ml plasma Viral load Welcome to the Digital droplet PCR age! bitter is de pil Linos Vandekerckhove, MD, PhD Latent HIV reservoir Time at Ghent University Hospital 2 HIV
TaqMan Mutation Detection Assay
 PROTOCOL TaqMan Mutation Detection Assay Competitive Allele-Specific TaqMan PCR Publication Part Number 4467011 Rev. B Revision Date April 2012 For Research Use Only. Not intended for any animal or human
PROTOCOL TaqMan Mutation Detection Assay Competitive Allele-Specific TaqMan PCR Publication Part Number 4467011 Rev. B Revision Date April 2012 For Research Use Only. Not intended for any animal or human
StepOne and StepOnePlus Real-Time PCR Systems
 PRODUCT BROCHURE Real-Time PCR Systems StepOne and StepOnePlus Real-Time PCR Systems Remarkably Simple Systems. Simply Remarkable Results. Step Up To High Performance Real-Time PCR Remarkably Simple Systems
PRODUCT BROCHURE Real-Time PCR Systems StepOne and StepOnePlus Real-Time PCR Systems Remarkably Simple Systems. Simply Remarkable Results. Step Up To High Performance Real-Time PCR Remarkably Simple Systems
High-Capacity cdna Reverse Transcription Kits For 200 and 1000 Reactions
 High-Capacity cdna Reverse Transcription Kits For 200 and 1000 Reactions Protocol Copyright 2006,. All rights reserved. For Research Use Only. Not for use in diagnostic procedures. Information in this
High-Capacity cdna Reverse Transcription Kits For 200 and 1000 Reactions Protocol Copyright 2006,. All rights reserved. For Research Use Only. Not for use in diagnostic procedures. Information in this
Profiling of non-coding RNA classes Gunter Meister
 Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Real time and Quantitative (RTAQ) PCR. so I have an outlier and I want to see if it really is changed
 Real time and Quantitative (RTAQ) PCR or.. for this audience so I have an outlier and I want to see if it really is changed Nigel Walker, Ph.D. Laboratory of Computational Biology and Risk Analysis, Environmental
Real time and Quantitative (RTAQ) PCR or.. for this audience so I have an outlier and I want to see if it really is changed Nigel Walker, Ph.D. Laboratory of Computational Biology and Risk Analysis, Environmental
qstar mirna qpcr Detection System
 qstar mirna qpcr Detection System Table of Contents Table of Contents...1 Package Contents and Storage Conditions...2 For mirna cdna synthesis kit...2 For qstar mirna primer pairs...2 For qstar mirna qpcr
qstar mirna qpcr Detection System Table of Contents Table of Contents...1 Package Contents and Storage Conditions...2 For mirna cdna synthesis kit...2 For qstar mirna primer pairs...2 For qstar mirna qpcr
Getting Started Guide
 Primer Express Software Version 3.0 Getting Started Guide Before You Begin Designing Primers and Probes for Quantification Assays Designing Primers and Probes for Allelic Discrimination Assays Ordering
Primer Express Software Version 3.0 Getting Started Guide Before You Begin Designing Primers and Probes for Quantification Assays Designing Primers and Probes for Allelic Discrimination Assays Ordering
Introduction to Gene Expression. Getting Started Guide
 Introduction to Gene Expression Getting Started Guide For Research Use Only. Not intended for any animal or human therapeutic or diagnostic use. Information in this document is subject to change without
Introduction to Gene Expression Getting Started Guide For Research Use Only. Not intended for any animal or human therapeutic or diagnostic use. Information in this document is subject to change without
Absolute Quantitation Using Standard Curve Getting Started Guide
 Applied Biosystems 7900HT Fast Real-Time PCR System Absolute Quantitation Using Standard Curve Getting Started Guide Introduction Designing an AQ Experiment Preparing the Samples and Reaction Plate Creating
Applied Biosystems 7900HT Fast Real-Time PCR System Absolute Quantitation Using Standard Curve Getting Started Guide Introduction Designing an AQ Experiment Preparing the Samples and Reaction Plate Creating
User Manual/Hand book. qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse)
 User Manual/Hand book qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse) Kit Components Cat. No. MA003...Human Whole Genome mirna qpcr Profiling Kit (-20 C) The following components are sufficient
User Manual/Hand book qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse) Kit Components Cat. No. MA003...Human Whole Genome mirna qpcr Profiling Kit (-20 C) The following components are sufficient
Hepatitis B Virus. genesig Advanced Kit. DNA testing. Everything... Everyone... Everywhere... Core Protein Region. 150 tests.
 TM Primerdesign Ltd TM Primerdesign Ltd Hepatitis B Virus Core Protein Region genesig Advanced Kit 150 tests DNA testing Everything... Everyone... Everywhere... For general laboratory and research use
TM Primerdesign Ltd TM Primerdesign Ltd Hepatitis B Virus Core Protein Region genesig Advanced Kit 150 tests DNA testing Everything... Everyone... Everywhere... For general laboratory and research use
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS)
 Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
How To Get A Cell Print
 QUICK CELL CAPTURE AND CHARACTERIZATION GUIDE FOR CELLSEARCH CUSTOMERS CellSave EDTA Blood sample Rare cell capture Enumeration Single protein marker Cell capture for molecular characterization CELLSEARCH
QUICK CELL CAPTURE AND CHARACTERIZATION GUIDE FOR CELLSEARCH CUSTOMERS CellSave EDTA Blood sample Rare cell capture Enumeration Single protein marker Cell capture for molecular characterization CELLSEARCH
Quantification of Mycobacterium Tuberculosis. 150 tests
 Quantification of Mycobacterium Tuberculosis rep 13E12 For general laboratory and research use only 150 tests Introduction to Mycobacterium Tuberculosis Mycobacterium tuberculosis is the bacterium that
Quantification of Mycobacterium Tuberculosis rep 13E12 For general laboratory and research use only 150 tests Introduction to Mycobacterium Tuberculosis Mycobacterium tuberculosis is the bacterium that
SYBR Green PCR Master Mix and SYBR Green RT-PCR Reagents Kit
 USER GUIDE SYBR Green PCR Master Mix and SYBR Green RT-PCR Reagents Kit Catalog Number 4309155 (Master Mix) and 4306736 (RT-PCR Reagents Kit) Publication Part Number 4310251 Rev. G Revision Date September
USER GUIDE SYBR Green PCR Master Mix and SYBR Green RT-PCR Reagents Kit Catalog Number 4309155 (Master Mix) and 4306736 (RT-PCR Reagents Kit) Publication Part Number 4310251 Rev. G Revision Date September
All-in-One mirna qrt-pcr Detection System Handbook
 All-in-One mirna qrt-pcr Detection System Handbook For quantitative detection of mature mirna All-in-One mirna First-Strand cdna Synthesis Kit Cat. No. AMRT-0020 (20 mirna reverse transcription reactions)
All-in-One mirna qrt-pcr Detection System Handbook For quantitative detection of mature mirna All-in-One mirna First-Strand cdna Synthesis Kit Cat. No. AMRT-0020 (20 mirna reverse transcription reactions)
miracle mirna qpcr Instruction Manual Store at -20 Version 3.0 (Aug.2013)
 miracle mirna qpcr Instruction Manual Store at -20 Version 3.0 (Aug.2013) Contents I. PRODUCT INTRODUCTION 1 A. OVERVIEW 1 B. KIT CONTENTS 1 1. REAGENT KITS 1 2. PRIMER SETS 3 3. MIRACLE ON CALL SCREEN
miracle mirna qpcr Instruction Manual Store at -20 Version 3.0 (Aug.2013) Contents I. PRODUCT INTRODUCTION 1 A. OVERVIEW 1 B. KIT CONTENTS 1 1. REAGENT KITS 1 2. PRIMER SETS 3 3. MIRACLE ON CALL SCREEN
quantitative real-time PCR, grain, simplex DNA extraction: PGS0426 RT-PCR: PGS0494 & PGS0476
 BioScience quantitative real-time PCR, grain, simplex DNA extraction: PGS0426 RT-PCR: PGS0494 & PGS0476 This method describes a Real-time semi-quantitative TaqMan PCR procedure for the determination of
BioScience quantitative real-time PCR, grain, simplex DNA extraction: PGS0426 RT-PCR: PGS0494 & PGS0476 This method describes a Real-time semi-quantitative TaqMan PCR procedure for the determination of
DP419 RNAsimple Total RNA Kit. RNAprep pure Series. DP501 mircute mirna Isolation Kit. DP438 MagGene Viral DNA / RNA Kit. DP405 TRNzol Reagent
 Overview of TIANGEN Products DP419 RNAsimple Total RNA Kit DP430 RNAprep pure Kit(For Cell/Bacteria) DP315/DP315-R TIANamp Virus DNA/RNA Kit DP431 RNAprep pure Kit (For Tissue) Silica-membrane Technology
Overview of TIANGEN Products DP419 RNAsimple Total RNA Kit DP430 RNAprep pure Kit(For Cell/Bacteria) DP315/DP315-R TIANamp Virus DNA/RNA Kit DP431 RNAprep pure Kit (For Tissue) Silica-membrane Technology
Formalin fixation at low temperature better preserves nucleic acid integrity. Gianni Bussolati. University of Turin
 Formalin fixation at low temperature better preserves nucleic acid integrity Gianni Bussolati University of Turin Disclosure of interests: G.B. was originally responsible for the invention of the Cold
Formalin fixation at low temperature better preserves nucleic acid integrity Gianni Bussolati University of Turin Disclosure of interests: G.B. was originally responsible for the invention of the Cold
Qubit dsdna HS Assay Kits
 Qubit dsdna HS Assay Kits For use with the Qubit 2.0 Fluorometer Table 1. Contents and storage information. Material Amount Concentration Storage Stability Qubit dsdna HS Reagent (Component A) 250 µl or
Qubit dsdna HS Assay Kits For use with the Qubit 2.0 Fluorometer Table 1. Contents and storage information. Material Amount Concentration Storage Stability Qubit dsdna HS Reagent (Component A) 250 µl or
User Manual. Reference Gene Panel Mouse Probe protocol. Version 1.1 August 2014 For use in quantitative real-time PCR
 User Manual Reference Gene Panel Mouse Probe protocol Version 1.1 August 2014 For use in quantitative real-time PCR Reference Gene Panel Mouse Table of contents Background 4 Assays included in the panel
User Manual Reference Gene Panel Mouse Probe protocol Version 1.1 August 2014 For use in quantitative real-time PCR Reference Gene Panel Mouse Table of contents Background 4 Assays included in the panel
CopyCaller Software. User Guide
 CopyCaller Software User Guide CopyCaller Software User Guide Information in this document is subject to change without notice. For Research Use Only. Not for use in diagnostic procedures. APPLIED BIOSYSTEMS
CopyCaller Software User Guide CopyCaller Software User Guide Information in this document is subject to change without notice. For Research Use Only. Not for use in diagnostic procedures. APPLIED BIOSYSTEMS
ab185916 Hi-Fi cdna Synthesis Kit
 ab185916 Hi-Fi cdna Synthesis Kit Instructions for Use For cdna synthesis from various RNA samples This product is for research use only and is not intended for diagnostic use. Version 1 Last Updated 1
ab185916 Hi-Fi cdna Synthesis Kit Instructions for Use For cdna synthesis from various RNA samples This product is for research use only and is not intended for diagnostic use. Version 1 Last Updated 1
User Manual Mouse Endogenous Control Gene Panel
 User Manual Mouse Endogenous Control Gene Panel Version 1.5 March 2008 For use in quantitative real-time PCR Mouse Endogenous Control Gene Panel Table of contents Background 4 Contents 4 Additionally required
User Manual Mouse Endogenous Control Gene Panel Version 1.5 March 2008 For use in quantitative real-time PCR Mouse Endogenous Control Gene Panel Table of contents Background 4 Contents 4 Additionally required
DyNAmo cdna Synthesis Kit for qrt-pcr
 DyNAmo cdna Synthesis Kit for qrt-pcr Instruction manual F- 470S Sufficient for 20 cdna synthesis reactions (20 µl each) F- 470L Sufficient for 100 cdna synthesis reactions (20 µl each) Description...
DyNAmo cdna Synthesis Kit for qrt-pcr Instruction manual F- 470S Sufficient for 20 cdna synthesis reactions (20 µl each) F- 470L Sufficient for 100 cdna synthesis reactions (20 µl each) Description...
Custom TaqMan Assays For New SNP Genotyping and Gene Expression Assays. Design and Ordering Guide
 Custom TaqMan Assays For New SNP Genotyping and Gene Expression Assays Design and Ordering Guide For Research Use Only. Not intended for any animal or human therapeutic or diagnostic use. Information in
Custom TaqMan Assays For New SNP Genotyping and Gene Expression Assays Design and Ordering Guide For Research Use Only. Not intended for any animal or human therapeutic or diagnostic use. Information in
RT-PCR: Two-Step Protocol
 RT-PCR: Two-Step Protocol We will provide both one-step and two-step protocols for RT-PCR. We recommend the twostep protocol for this class. In the one-step protocol, the components of RT and PCR are mixed
RT-PCR: Two-Step Protocol We will provide both one-step and two-step protocols for RT-PCR. We recommend the twostep protocol for this class. In the one-step protocol, the components of RT and PCR are mixed
PCR Instruments and Consumables
 Index Page GeneAmp PCR Instrument Systems 2 GeneAmp PCR System 9700 Components 3 Accessories and Software for GeneAmp PCR Instrument Systems 4 Disposables 5 PCR Enzymes 7 GeneAmp PCR Kits 12 RNA PCR Kits
Index Page GeneAmp PCR Instrument Systems 2 GeneAmp PCR System 9700 Components 3 Accessories and Software for GeneAmp PCR Instrument Systems 4 Disposables 5 PCR Enzymes 7 GeneAmp PCR Kits 12 RNA PCR Kits
User Manual. CelluLyser Lysis and cdna Synthesis Kit. Version 1.4 Oct 2012 From cells to cdna in one tube
 User Manual CelluLyser Lysis and cdna Synthesis Kit Version 1.4 Oct 2012 From cells to cdna in one tube CelluLyser Lysis and cdna Synthesis Kit Table of contents Introduction 4 Contents 5 Storage 5 Additionally
User Manual CelluLyser Lysis and cdna Synthesis Kit Version 1.4 Oct 2012 From cells to cdna in one tube CelluLyser Lysis and cdna Synthesis Kit Table of contents Introduction 4 Contents 5 Storage 5 Additionally
New Technologies for Sensitive, Low-Input RNA-Seq. Clontech Laboratories, Inc.
 New Technologies for Sensitive, Low-Input RNA-Seq Clontech Laboratories, Inc. Outline Introduction Single-Cell-Capable mrna-seq Using SMART Technology SMARTer Ultra Low RNA Kit for the Fluidigm C 1 System
New Technologies for Sensitive, Low-Input RNA-Seq Clontech Laboratories, Inc. Outline Introduction Single-Cell-Capable mrna-seq Using SMART Technology SMARTer Ultra Low RNA Kit for the Fluidigm C 1 System
Oligonucleotide Stability Study
 Oligonucleotide Stability Study IDT is performing a long term study of oligo stability. The study has been ongoing for 2 years and has tested oligo stability under various conditions. An overview of the
Oligonucleotide Stability Study IDT is performing a long term study of oligo stability. The study has been ongoing for 2 years and has tested oligo stability under various conditions. An overview of the
Supplemental Information. McBrayer et al. Supplemental Data
 1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
Endocrine Glands and the General Principles of Hormone Action
 Endocrine Glands and the General Principles of Hormone Action Cai Li, Ph.D. Assistant professor Touchstone Center for Diabetes Research Departments of Physiology and Internal Medicine The University of
Endocrine Glands and the General Principles of Hormone Action Cai Li, Ph.D. Assistant professor Touchstone Center for Diabetes Research Departments of Physiology and Internal Medicine The University of
micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved
 microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
Accurate and sensitive mutation detection and quantitation using TaqMan Mutation Detection Assays for disease research
 PPLICTION NOTE Mutation Detection ssays ccurate and sensitive mutation detection and quantitation using Mutation Detection ssays for disease research In this research study, we addressed the feasibility
PPLICTION NOTE Mutation Detection ssays ccurate and sensitive mutation detection and quantitation using Mutation Detection ssays for disease research In this research study, we addressed the feasibility
AffinityScript QPCR cdna Synthesis Kit
 AffinityScript QPCR cdna Synthesis Kit INSTRUCTION MANUAL Catalog #600559 Revision C.01 For In Vitro Use Only 600559-12 LIMITED PRODUCT WARRANTY This warranty limits our liability to replacement of this
AffinityScript QPCR cdna Synthesis Kit INSTRUCTION MANUAL Catalog #600559 Revision C.01 For In Vitro Use Only 600559-12 LIMITED PRODUCT WARRANTY This warranty limits our liability to replacement of this
Instructions. Torpedo sirna. Material. Important Guidelines. Specifications. Quality Control
 is a is a state of the art transfection reagent, specifically designed for the transfer of sirna and mirna into a variety of eukaryotic cell types. is a state of the art transfection reagent, specifically
is a is a state of the art transfection reagent, specifically designed for the transfer of sirna and mirna into a variety of eukaryotic cell types. is a state of the art transfection reagent, specifically
Highly specific and sensitive quantitation
 PRODUCT ULLETIN SYR Select Master Mix SYR Select Master Mix Highly specific and sensitive quantitation SYR Select Master Mix offers advanced performance at an affordable price. SYR Select Master Mix is
PRODUCT ULLETIN SYR Select Master Mix SYR Select Master Mix Highly specific and sensitive quantitation SYR Select Master Mix offers advanced performance at an affordable price. SYR Select Master Mix is
Reagent Guide. Applied Biosystems StepOne and StepOnePlus Real-Time PCR Systems
 Reagent Guide Applied Biosystems StepOne and StepOnePlus Real-Time PCR Systems Applied Biosystems StepOne and StepOnePlus Real-Time PCR Systems Reagent Guide Copyright 2008, 2010 Applied Biosystems. All
Reagent Guide Applied Biosystems StepOne and StepOnePlus Real-Time PCR Systems Applied Biosystems StepOne and StepOnePlus Real-Time PCR Systems Reagent Guide Copyright 2008, 2010 Applied Biosystems. All
RealStar HBV PCR Kit 1.0 11/2012
 RealStar HBV PCR Kit 1.0 11/2012 RealStar HBV PCR Kit 1.0 For research use only! (RUO) Product No.: 201003 96 rxns INS-201000-GB-02 Store at -25 C... -15 C November 2012 altona Diagnostics GmbH Mörkenstraße
RealStar HBV PCR Kit 1.0 11/2012 RealStar HBV PCR Kit 1.0 For research use only! (RUO) Product No.: 201003 96 rxns INS-201000-GB-02 Store at -25 C... -15 C November 2012 altona Diagnostics GmbH Mörkenstraße
Early Detection Research Network: Validation Infrastructure. Jo Ann Rinaudo, Ph.D. Division of Cancer Prevention
 Early Detection Research Network: Validation Infrastructure Jo Ann Rinaudo, Ph.D. Division of Cancer Prevention 15 October 2015 Mission The NCI Early Detection Research Network s (EDRN) mission is to implement
Early Detection Research Network: Validation Infrastructure Jo Ann Rinaudo, Ph.D. Division of Cancer Prevention 15 October 2015 Mission The NCI Early Detection Research Network s (EDRN) mission is to implement
First Strand cdna Synthesis
 380PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name First Strand cdna Synthesis (Cat. # 786 812) think proteins! think G-Biosciences
380PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name First Strand cdna Synthesis (Cat. # 786 812) think proteins! think G-Biosciences
THE HUMAN BODY SYSTEMS
 Name Period Date THE HUMAN BODY SYSTEMS System Function Diagram Major Organs Digestive 1. take in food (ingestion) 2. digest food into smaller molecules and absorb nutrients 3. remove undigestable food
Name Period Date THE HUMAN BODY SYSTEMS System Function Diagram Major Organs Digestive 1. take in food (ingestion) 2. digest food into smaller molecules and absorb nutrients 3. remove undigestable food
