Molecular Portraits of Non- Coding RNAs in Neuroblastoma
|
|
|
- Chester Fisher
- 10 years ago
- Views:
Transcription
1 Ghent University, Faculty of Medicine and Health Sciences Molecular Portraits of Non- Coding RNAs in Neuroblastoma this thesis is submitted as fulfilment of the requirements for the degree of Doctor in Biomedical Sciences by Pieter Mestdagh, 2011 promoter prof. dr.ir. Jo Vandesompele co- promoter prof. dr. Frank Speleman Center for Medical Genetics Ghent University Hospital, Medical Research Building De Pintelaan 185, 9000 Gent, Belgium
2 II
3 Thesis submitted to fulfil the requirements for the degree of Doctor in Biomedical Sciences III Promoter prof. dr. ir. Jo Vandesompele (Ghent University, Belgium) Co- promoter prof. dr. Frank Speleman (Ghent University, Belgium) Members of the examination committee: prof. dr. Mark Bracke (Ghent University, Belgium) prof. dr. Yves Van De Peer (Ghent University, Belgium) prof. dr. Jan Cools (Catholic University of Leuven, Belgium) prof. dr. Michel Georges (University of Liège, Belgium) dr. Pieter Rondou (Ghent University, Belgium) prof. dr. Ray Stallings (Royal College of Surgeons, Ireland) prof. dr. Jason Shohet (Baylor College, USA) De auteur en de promotoren geven de toelating deze scriptie voor consultatie beschikbaar te stellen en delen ervan te kopiëren voor persoonlijk gebruik. Elk ander gebruik valt onder de beperkingen van het auteursrecht, in het bijzonder met betrekking tot de verplichting uitdrukkelijk de bron te vermelden bij het aanhalen van resultaten uit deze scriptie. The author and the promoters give the permission to use this thesis for consultation and to copy parts of it for personal use. Every other use is subject to the copyright law, more specifically the source must be extensively specified when using results from this thesis. The research described in this thesis was conducted at the Centre for Medical Genetics, Ghent University Hospital, Ghent, Belgium This work was supported by the Ghent University Research fund (BOF 01D31406), the Fund for Scientific Research (grant number: G and ), the Belgian Kid s Fund, the Stichting tegen Kanker and GOA (01G01910). This article represents research results of the Belgian program of Interuniversity Poles of Attraction, initiated by the Belgian State, Prime Minister s Office, Science Policy Programming.
4 IV
5 Table of Contents LIST OF ABBREVIATIONS INTRODUCTION NEUROBLASTOMA NEUROBLASTOMA GENETICS MYCN THE MYC FAMILY TRANSCRIPTIONAL CONTROL ONE ONCOGENE, TWO FACES TARGETS TO TARGET NON- CODING RNAS MESSENGERS WITHOUT A MESSAGE THE BIOGENESIS OF MIRNAS ONCOMIRS AND TUMOUR SUPPRESSOR MIRNAS MIRNAS REGULATED BY MYC MIR- 34A: THE MISSING PIECE IN THE P53 NETWORK PUZZLE MIRNAS AND METASTASIS MECHANISMS OF DEREGULATED MIRNA EXPRESSION MIRNA SIGNATURES FOR IMPROVED DIAGNOSTIC AND PROGNOSTIC CLASSIFICATION MIRNA THERAPEUTICS QUANTIFICATION OF MIRNA EXPRESSION EXPLORING MIRNA FUNCTION FUNCTIONAL EXPLORATION OF GENE EXPRESSION PATTERNS REFERENCES RESEARCH OBJECTIVES RESULTS V PAPER 1: HIGH- THROUGHPUT STEM- LOOP RT- QPCR MIRNA EXPRESSION PROFILING USING MINUTE AMOUNTS OF INPUT RNA 30 PAPER 2: A NOVEL AND UNIVERSAL METHOD FOR MICRORNA RT- QPCR DATA NORMALIZATION 42 PAPER 3: MYCN/C- MYC- INDUCED MICRORNAS REPRESS CODING GENE NETWORKS ASSOCIATED WITH POOR OUTCOME IN MYCN/C- MYC- ACTIVATED TUMOURS 65 PAPER 4: THE MIR MICRORNA CLUSTER REGULATES MULTIPLE COMPONENTS OF THE TGF- Β PATHWAY IN NEUROBLASTOMA 101 PAPER 5: AN INTEGRATIVE GENOMICS SCREEN UNCOVERS NCRNA T- UCR FUNCTIONS IN NEUROBLASTOMA TUMOURS 132 PAPER 6: THE MICRORNA BODY MAP: DISSECTING MICRORNA FUNCTION THROUGH INTEGRATIVE GENOMICS 149 PAPER 7: OUTCOME PREDICTION OF CHILDREN WITH NEUROBLASTOMA USING MIRNA AND MRNA GENE EXPRESSION SIGNATURES 168 DISCUSSION AND FUTURE PERSPECTIVES REFERENCES SUMMARY SAMENVATTING CURRICULUM VITAE
6
7 1 List of abbreviations INSS: International Neuroblastoma Staging System INRGSS: International Neuroblastoma Risk Group Staging System GWAS: genome- wide association study SNP: single nucleotide polymorphism shrna: short- hairpin RNA DFMO: alpha- difluoromethylornithine ncrna: non- coding RNA mirna: micro RNA pirna: PIWI- interacting RNA endo- sirna: endogenous short- interfering RNA T- UCR: transcribed ultraconserved region snorna: small nucleolar RNA lncrna: long non- coding RNA pri- mirna: primary mirna transcript pre- mirna: premature mirna transcript RISC: RNA- induced silencing complex CLL: chronic lymphocytic leukemia CAGR: cancer- associated genomic region AML: acute myeloid leukemia RT- qpcr: reverse transcription quantitative polymerase chain reaction RIP- Chip: Ribonucleoprotein ImmunoPrecipitation- gene Chip HITS- CLIP: high- throughput sequencing of RNA isolated by crosslinking immunoprecipitation GO: gene ontology GSEA: gene set enrichment analysis LNA: locked nucleic acid
8 Introduction 2 Neuroblastoma Neuroblastoma is a childhood malignancy that accounts for 15% of pediatric cancer mortality 1. It is also the most frequently diagnosed extracranial tumour in children and is characterized by a remarkable heterogeneity, both in terms of clinical behavior and genetic aberrations. Despite intensive multimodal therapy, the cure- rate for high- risk patients is lower than 40%. A major determinant of the clinical course is the age of the patient at diagnosis. Typically, children older than 1 year have metastatic disease and a poor overall survival while infants present with localized tumours that can mature into a benign ganglioneuroma or regress spontaneously 2. In the 1990 s, the first therapeutic stratification system for neuroblastoma patients was based on age at diagnosis and a number of post- surgical parameters (International Neuroblastoma Staging System, INSS) 3, (Figure 1). Because surgical approaches differ between institutions, a uniform staging system (International Neuroblastoma Risk Group Staging System, INRGSS), based on an international consensus, was recently proposed 4. The INRGSS combines a series of non- surgical parameters - such as age at diagnosis and radiographic characteristics of the tumour - with several biological factors and forms the current basis for risk- related therapies. Neuroblastoma tumours originate from precursor cells of the sympathoadrenal lineage. Hence, tumours can develop anywhere in the sympathetic nervous system. At least half of the primary tumours are found in the adrenal medulla while others originate in the paraspinal sympathetic ganglia or in pelvic ganglia. Neuroblastoma genetics Most neuroblastomas, as most cancers, are supposed to result from accumulating somatic mutations. However, a limited fraction of neurobastoma tumours (1-2%) are inherited in an autosomal dominant fashion 5-7. Germline mutations in PHOX2B, a key regulator in the process of nervous system development, account for a small subset of hereditary cases of neuroblastoma 8. Recently, genome- wide linkage analysis of neuroblastoma pedigrees identified ALK as the major familial neuroblastoma predisposition gene 9. ALK mutations target the tyrosine receptor kinase domain, resulting in a constitutive phosphorylation, sufficient to drive oncogenic transformation 10. Unlike PHOX2B, ALK is also mutated in a substantial portion (6.9%) of sporadic cases 10 implicating it as a putative target for molecular therapy. Earlier, genome- wide linkage analysis in neuroblastoma pedigrees identified a region on chromosome 16p and one on 4p suggesting that additional hereditary neuroblastoma predisposition genes exist 11, 12. In an attempt to pinpoint genetic events associated with susceptibility to aggressive neuroblastoma in sporadic cases, Maris and colleagues performed a genome- wide association study (GWAS) of single nucleotide polymorphisms (SNPs) and identified common SNPs at three different loci, i.e. 6p22, BARD1 and LMO LMO1 copy number analysis of primary tumour cells demonstrated copy number gain in 12.4% of the cases. Both the LMO1 germline SNP and the somatic copy number gain were associated with increased LMO1 expression enhancing cell proliferation. Neuroblastoma tumours are characterized by a high number of somatically acquired genomic alterations. Typically, these genetic aberrations target large genomic regions encompassing hundreds of genes. Regions that are frequently associated with copy number loss include chromosome 1p, chromosome 3p, chromosome 6q and chromosome 11q while regions that are associated with copy number gain include chromosome 1q and chromosome 17q 16 (Figure 2). Several of these genomic aberrations serve as prognostic markers for risk stratification and are used to classify neuroblastoma tumours in genetic subgroups However, the large number of genes residing within these regions has thwarted researchers in their search for tumour driving events underlying the individual genomic aberrations. One way to identify tumour- driving events relies on the reverse genetics approach where short- hairpin RNA (shrna) libraries are used to perturb the expression of a high number of genes followed by a relevant phenotypic read- out. Hölzel and colleagues applied such shrna
9 b a 3 4S Figure 1 International neuroblastoma staging system. Localization of primary and metastatic tumours for each disease stage as defined by the international neuroblastoma staging system (INSS). Stage 1: Localised tumour with complete gross excision, with or without microscopic residual disease; representative ipsilateral lymph nodes negative for tumour microscopically. Stage 2a: Localised tumour with incomplete gross excision; representative ipsilateral non-adherent lymph nodes negative for tumour microscopically. Stage 2b: Localised tumour with or without complete gross excision, with ipsilateral non-adherent lymph nodes positive for tumour. Enlarged contralateral lymph nodes should be negative microscopically. Stage 3: Unresectable unilateral tumour infiltrating across the midline, with or without regional lymph node involvement; or localised unilateral tumour with contralateral regional lymph node involvement; or midline tumour with bilateral extension by infiltration (unresectable) or by lymph node involvement. Stage 4: Any primary tumour with dissemination to distant lymph nodes, bone, bone marrow, liver, skin, or other organs (except as defined by stage 4S). Stage 4S: Localised primary tumour in infants younger than 1 year (as defined for stage 1, 2A, or 2B), with dissemination limited to skin, liver, or bone marrow. Metastatic sites: (1) liver, (2) bone, (3) bone marrow, (4) distant lymph node, (5) skin. (source: and Maris, ) library to identify genes modulating neuroblastoma response to retinoic acid, a differentiation- inducing agent 21. They identified a crosstalk between the tumour suppressor NF1 and retinoic acid- induced differentiation and found NF1 microdeletions and mutations in neuroblastoma tumours. Often, tumour- driving genetic events are identified through focal genomic aberrations such as amplification of the ERBB2 oncogene in breast cancer 22 or homozygous deletion of the PTEN tumour suppressor in prostate cancer 23. Homozygous deletions have been described in neuroblastoma, amongst others for CDKN2A 24 and NF1 25, but are rare events. In contrast, several amplicons have been identified in neuroblastoma, encompassing known oncogenes such as MDM2 26, 27, ALK 28 and most importantly MYCN 29. Amplification of the MYCN oncogene occurs in 20-25% of neuroblastoma tumours and delineates a subgroup of patients with highly aggressive, metastatic disease and poor outcome. Tumours with MYCN amplification are cytogenetically characterized by double minute chromatin bodies or homogenous staining regions 30, 31, two typical manifestations of gene amplification. Although the amplicon can contain multiple genes, MYCN is the only gene that shows consistent amplification 32. In general, tumours with MYCN amplification express MYCN at much higher levels than tumours without MYCN amplification. However, it remains controversial whether MYCN overexpression is correlated to survival in tumours lacking MYCN
10 4 amplification 33, 34. Most tumours with MYCN amplification also have a deletion of chromosome 1p but not all tumours with a 1p- deletion have MYCN amplification suggesting that deletion of chromosome 1p occurs earlier in tumour development 2. Possibly, genes negatively regulating MYCN expression or inducing apoptosis in the presence of high MYCN levels need to be deleted in order for MYCN amplification to occur. The assumption that MYCN amplification is a tumour driving event in neuroblastoma was put to the test by Weiss and colleagues who developed transgenic mice that overexpress MYCN in sympathetic nervous system cells 35. The mice developed neuroblastoma- like tumours with chromosomal gains and losses that are syntenic with comparable abnormalities detected in human neuroblastoma tumours. These results suggest that MYCN amplification indeed contributes to the genesis of neuroblastoma tumours. MYCN The MYC family The MYC- family of proto- oncogenes or MYC- box genes encodes three different transcription factors: MYC, MYCN and MYCL. MYC (c- MYC) was originally isolated from chicken DNA due to its homology with the v- myc oncogene from the avian myelocytomatosis virus 36. MYC activation occurs either through chromosomal translocation, placing the MYC open reading frame under the control of a constitutively active promoter 37, 38, or through gene amplification 39, 40. MYCN and MYCL were detected as amplified DNA fragments with partial homology to MYC in neuroblastoma and small cell lung cancer respectively 29, 41. While MYC is activated in a wide variety of tumours, MYCN activation has a strong preference for tumours of neuroectodermal origin such as neuroblastoma, retinoblastoma and peripheral neuroectodermal tumours. Human tumour cells almost invariably express MYC but, depending on their derivation of embryonal cell lineage, hardly ever express MYCN 42. Analysis of tissues from different stages of the fetal and the developing mouse embryo further support these observations by showing that, during embryonal development, the overall expression pattern of MYC is relatively constant whereas MYCN displays a temporal and spatial expression pattern. Strikingly, Malynn and colleagues have shown that MYCN can functionally replace MYC in murine growth, development and differentiation 43. Mice in which the MYC coding sequences was replaced with the MYCN coding sequence were shown to survive into adulthood and reproduce with MYCN expression being similarly regulated as MYC expression. This functional redundancy suggests that MYC and MYCN can regulate identical cellular processes and that the MYC- family was evolved to facilitate differential expression patterns of the MYC- family genes. Of note, a reverse experiment where MYC replaces MYCN has not been reported so far. Transcriptional control MYCN encodes a nuclear protein of approximately 65 kda that is comprised of a basic DNA binding region (b), an α- helical protein protein interaction domain known as helix loop helix (HLH), a leucine zipper motif (Z) and an N- terminal transactivation domain containing two MYC- boxes termed MBI and MBII, all of which are conserved between the different MYC- family members. Through their bhlhz domain, MYC- family proteins form heterodimers with another bhlhz- protein called MAX and MYC(N)/MAX heterodimers have DNA binding and transcriptional activity 45 (Figure 3A). While MAX proteins can also homodimerize, these complexes are believed to be transcriptionally inert. MYC/MAX heterodimers specifically recognize and bind the hexameric DNA element CACGTG belonging to the larger class of E- box elements (CANNTG). Importantly, MYC proteins fail to activate transcription at E- box elements in the absence of MAX 46. MAX can also form heteroduplexes with bhlhz proteins from the MAD- family (MAD1, MXI1, MAD3 and MAD4).
11 5 mir-34a MYCN AMP ALKAMP, MUT PHOX2B MUT BARD1 GWAS SNP rs GWAS LMO1 GWAS MDM2 AMP NF1 HDEL AMP Amplification HDEL Homozygous deletion MUT Mutation GWAS Identified through GWAS chromosomal loss chromosomal gain Figure 2 Genetic defects in neuroblastoma tumours. Schematic representation of common genetic aberrations and genes associated with neuroblastoma. This non-exhaustive overview contains genes and chromosomal regios that are discussed in this thesis. Chromosomal regions of copy number gain are indicated in red, regions of loss are indicated in blue. Genes that are amplified, homozygously deleted, mutated or identified through GWAS are indicated by AMP, HDEL, MUT and GWAS respectively. (Adapted from Van Roy et al., )
12 Upon heteroduplex formation, MAD/MAX complexes bind the E- box consensus sequence where they act as transcriptional repressors hereby antagonizing transactivation by MYC proteins. Apart from a few examples, transcriptional repression by MYC proteins is believed to be independent of E- box binding and, compared to transcriptional activation, the underlying mechanisms are less well understood. Different models have been proposed that might explain MYC- induced transcriptional repression 47. One model suggests that MYC activates protein- coding transcriptional repressors and thereby acts indirectly on promoter elements. However, MYC mediated transcriptional repression has been observed in the presence of inhibitors of protein synthesis suggesting that repression does not require the synthesis of an intermediate protein 48. Second, repression might also be mediated through a direct binding of MYC to sequences in the target gene promoter as was shown for CDKN1B 49 (Figure 3B). As only few examples of direct MYC binding were reported, MYC- mediated repression is likely to occur through alternative mechanisms. A third possibility is that MYC is recruited to core promoter elements through protein- protein interactions without directly binding to DNA. Several candidate proteins, such as YY1, TFII- I, SP1 and MIZ- 1, have been proposed to target MYC to core promoter elements and repression of different protein- coding genes has indeed been shown to depend on the interaction between MYC and MIZ or SP1 48 (Figure 3C). In the results section of this work, a second mechanism explaining MYC mediated transcriptional repression of coding genes is presented. This mechanism relies on the induction of mirna expression by MYC- family proteins. MYC proteins have been thought to function only as classic transcription factors that bind to well defined regions in the promoters of specific target genes. Recent studies now suggest that MYC functions reach far beyond that of a classical bhlhz protein. In contrast to a classic bhlhz protein, MYC proteins potentially regulate up to 15% of all human protein- coding genes 53. Although widespread, the magnitude of transcriptional regulation of most target genes by MYC is nonetheless relatively modest compared with other bhlhz family members and transcription factors in general 54. In addition, MYC proteins have been shown to influence global chromatin state by regulating both acetylation and methylation of several histones 55, 56. These findings suggest a new model in which MYC genes function both locally and globally. 6 A MAX MAX B MAX MYC MAX MAX MAX MAD MYC MNT!"#$%& C '()"*+*,*-.& MAX MIZ-1 MYC /01*&210,0.*1& Figure 3 The MYC-MAX-MAD network. (A) Schematic representation of MAX-interaction partners and their effect on gene transcription upon E-box binding. (B-C) Models for MYC-mediated repression of gene transcription, either through binding of MYC-MAX complexes to INR-elements or through interaction with MIZ-1 at core promoters.
13 7 One oncogene, two faces One of the most important functions of MYC proteins is the regulation of cell growth and proliferation (Figure 4). Several MYC target genes are thought to play a role in the ability of MYC to promote cellular growth, including those associated with cellular metabolism, ribosomal and mitochondrial biogenesis, and protein and nucleic acid synthesis 57. MYC is also known to stimulate glycolysis through increased glucose transport and regulation of glycolytic genes resulting in the production of lactate under aerobic conditions 58, 59. This phenomenon is called the Warburg effect and is active in many cancer types. Cells with activated MYC often display a shortened G1 cell cycle phase and accelerated cell cycle progression. This is explained by MYC s ability to repress cell cycle checkpoint genes and inhibit cyclin dependent kinase inhibitors such as CDKN1A. MYC can also promote cell cycle progression by activating cyclines (CCND1, CCNE1), cyclin dependent kinases (CDK4) and members of the E2F transcription family. Through these mechanisms, deregulated MYC activity results in increased cell proliferation making cells vulnerable to additional oncogenic events that further accelerate tumourigenesis. MYC is also involved in triggering an angiogenic switch, amongst others through repression of THBS1 and CTGF 60, 61 and has been shown to promote chromosomal instability 62. Next to its role as a driver of oncogenesis, MYC can also trigger intrinsic tumour suppressor mechanisms including apoptosis and cellular senescence (Figure 4). Therefore, secondary mutations targeting components of the MYC tumour suppressor axis are under strong selective pressure during MYC- driven tumourigenesis 58. MYC sensitizes cells for apoptosis through two different mechanisms. One mechanism involves the induction of P14ARF by MYC, which in turn results in the activation of TP53 and transcription of pro- apoptotic genes such as BAX and PUMA 57. Alternatively, MYC directly represses expression of members from the anti- apoptotic BCL2- family such as BCL2 and BCLX that regulate BAX. The role of MYC in regulating apoptosis is further supported by studies in MYC- null cells showing that, in the absence of MYC, cells are resistant to apoptotic stimuli 57, 63. MYC- induced senescence is depending on an intact Arf- p53- p21 and p16ink4a- prb axis and the absence of protective factors such as Wrn and Cdk2 58. This was demonstrated in Eµ- MYC mice where lymphoma development was delayed in mice lacking functional Cdk2 due to the induction of senescence 64. However, in the presence of Cdk2, MYC is capable of repressing senescence. When overexpressed, many oncogenes such as BRAF and RAS, induce senescence in primary cells. This process is called oncogene- induced senescence and can be repressed by activated MYC, amongst others in melanomas with BRAF activation 65 and rat fibroblasts with RAS overexpression 66. Targets to target The high frequency of tumours with activation of a MYC family member and the plethora of functions that are controlled by MYC proteins suggests that MYC proteins are key drivers of tumourigenesis. In principle, this makes inhibition of MYC proteins an attractive pharmacological approach for cancer treatment. However, practical difficulties in designing MYC inhibitory drugs and concerns about possible side effects in proliferating normal tissues has tempered the enthusiasm 67. Therefore, researchers are trying to identify MYC target genes that are crucial in the process of MYC- driven tumourigenesis and that can be targeted pharmacologically. In neuroblastoma, several MYCN target genes have been identified that meet these requirements. For example, Slack and colleagues demonstrated that MDM2, a critical negative regulator of TP53, is directly activated by MYCN 68. MDM2 is an E3 ubiquitin ligase that negatively regulates p53 activity and stability by binding to its transactivation domain, hereby promoting its ubiquitination and degradation 69. Tumour cells with MYCN amplification overexpress MDM2 to escape TP53- mediated cell death, proliferate, and progress to invasive malignancy 68. As most neuroblastoma tumours have an intact TP53 signaling pathway, targeted disruption of the MDM2- TP53 interaction using the small- molecule nutlin- 3 activates the TP53 pathway resulting in an antiproliferative and cytotoxic effect 69, 70.
14 8 BCL2 CDKN2A APOPTOSIS MYC/MYCN E2F1/2/3 CDK4 CDKN1A metabolism nucleic acid synthesis THBS1 CTGF PROLIFERATION CELL GROWTH ANGIOGENESIS Figure 4 Opposing functions of the MYC family. MYC genes have the ability to induce apoptosis (upper part) or to stimulate cell growth, proliferation and tumour vascularization (lower part). Another attractive MYCN target gene is ODC1, a rate- limiting enzyme in polyamine biosynthesis that was shown to be upregulated in MYCN amplified neuroblastoma tumours 71. As polyamines are essential for cell survival and cancer progression, the authors evaluated the use of the ODC1 inhibitor alpha- difluoromethylornithine (DFMO) on neuroblastoma cell proliferation in vitro and in vivo. DFMO treatment was shown to inhibit proliferation of neuroblastoma cells in culture and prevented MYCN- induced oncogenesis in vivo. Of note, high- risk neuroblastoma tumours without MYCN amplification also overexpress ODC1 when compared to low- risk tumours suggesting that therapeutic treatment with DFMO is of potential relevance for the entire population of high- risk neuroblastoma patients. Westermann and colleagues defined a core set of MYC/MYCN target genes that show similar expression patterns to that of ODC1. They found that overexpression of these genes is predominantly driven by MYC in high- risk tumours without MYCN amplification and by MYCN in MYCN amplified tumours 72. These findings suggest that high- risk neuroblastoma tumours are characterized by a MYC/MYCN driven transcriptional program that is independent of the MYCN amplification status. Non- coding RNAs Messengers without a message The study of coding genes has been heavily pursued, amongst others through the use of high- throughput platforms enabling the profiling of alterations in entire (epi)genomes and transcriptomes, yielding insights into the complexity of cancer and disease biology in general. Recently, a complete new and unexpected piece of the transcriptome puzzle has emerged. In contrast to what has long been thought, protein- coding
15 genes only account for a fraction of the genomic DNA that is transcribed in a human cell. Genome- wide studies have shown that the vast majority of the mammalian genome is transcribed and produces many thousands of regulatory non- protein- coding RNAs (ncrnas). According to their size and function, ncrnas are divided into different classes including the small micrornas (mirnas), PIWI interacting RNAs (pirnas), endogenous small- interfering RNAs (endo- sirnas), small nucleolar RNAs (snornas), promoter- associated RNAs (PARs) and the longer transcribed ultraconserved regions (T- UCRs) and long non- coding RNAs 73, 74 (lncrnas) (Table 1). Of the classes identified until now, mirnas have been most thoroughly investigated. The discovery that mirnas regulate gene expression and protein translation is one of the most exciting new findings in biological and medical sciences of the past decade, rightfully assigning the 2006 Nobel Prize in Physiology or Medicine to Andrew Fire and Craig Mello. Initially described in 1993, mirnas were thought to be unique to the tiny roundworm Caenorhabditis elegans in which they were shown to be implicated in the process of developmental regulation 75. Soon, homologues were identified in other organisms including humans. Currently, over 1000 human mirnas have been reported and more are awaiting experimental validation, making mirnas one of the largest classes of gene regulators Table 1. Overview of established non- coding RNA classes ncrna Long (regulatory) noncoding RNAs (lncrnas) Transcribed ultraconserved regions (T- UCRs) Small interfering RNAs (sirnas) micrornas (mirnas) PIWI-interacting RNAs (pirnas) Promoter-associated RNAs (PARs) Small nucleolar RNAs (snornas) Adapted from Taft et al., characteristics The broadest class, lncrnas, encompass all non-protein-coding RNA species > 200 nt, including mrna-like ncrnas. Their functions include epigenetic regulation, acting as sequence-specific tethers for protein complexes and specifying subcellular compartments or localization Specific class of long non-coding RNAs, transcribed from genomic regions that are at least 200 nt in length and 100% conserved between human, mouse and rat species. Their function and exact length is unknown. Small RNAs nt long, produced by Dicer cleavage of complementary dsrna duplexes. sirnas form complexes with Argonaute proteins and are involved in gene regulation, transposon control and viral defence Small RNAs 22 nt long, produced by Dicer cleavage of imperfect RNA hairpins encoded in long primary transcripts or short introns. They associate with Argonaute proteins and are primarily involved in post-transcriptional gene regulation Dicer-independent small RNAs nt long, principally restricted to the germline and somatic cells bordering the germline. They associate with PIWI-clade Argonaute proteins and regulate transposon activity and chromatin state A general term encompassing a suite of long and short RNAs, including promoterassociated RNAs (PASRs) and transcription initiation RNAs (tirnas) that overlap promoters and TSSs. These transcripts may regulate gene expression Traditionally viewed as guides of rrna methylation and pseudouridylation. However, there is emerging evidence that they also have gene-regulatory roles The biogenesis of mirnas Upon transcription of the mirna gene in the nucleus, primary mirna (pri- mirna) transcripts ( nucleotides) are formed and further processed by different endonucleases 77 (Figure 5). First, the pri- mirna transcripts are cleaved in the nucleus by Drosha into ~70 nucleotide precursors called premature mirnas (pre- mirnas). These hairpin precursors are exported into the cytoplasm by XPO5 where they are further processed by DICER1 into small imperfect double stranded RNA duplexes (mirna- mirna*) that contain both the mature mirna strand and its complementary strand (mirna*). The duplex is then loaded into the mirna- associated multiprotein RNA- induced silencing complex (mirisc) with retention of the mature mirna strand. DICER1, TARBP2 and Argonaute proteins mediate RISC assembly 78. The mature mirna guides the complex towards complementary sites in the target mrna to regulate gene expression.
16 10 Imperfect binding between the mirna and the mirna recognition element in the 3 UTR of the target gene ultimately results in degradation of the target mrna or inhibition of protein translation. Because of the ability to bind with incomplete complementarity, only part of the mirna sequence, the seed, is used to identify its target mrnas. This seed sequence encompasses bases 2-8 of the mature mirna and is established as important for biological function and stability. Recent bioinformatic studies argue that as much as 60% of all human protein- coding genes could be under the control of at least one mirna 79. This raises the possibility that mirnas control a large number of genetic pathways and that deregulated mirna expression contributes to disease, including cancer. nucleus cytoplasm AGUAGGUUGUAUAGUUGGA UCCAACUAUACAACCUACU GU Pri-miRNA Pre-miRNA DICER mirna-mirna* duplex AUAGAGGGUCACCACCCACACUGGGAUUGA U GAG UGGGA CUC UCCCA Unwind mature mirna mirna gene mirisc assembly Imperfect complementarity ORF Figure 5 mirna biogenesis pathway. Overview of the different steps in the processing of primary mirna transcripts to functional, mature mirnas by endonucleases. (adapted from Mestdagh et al., ) Oncomirs and tumour suppressor mirnas Although we have yet to learn most of the specific functions associated with a given mirna, at present we know mirnas are implicated in developmental timing, differentiation, cell proliferation, growth control, apoptosis and stem cell maintenance, all aspects of normal cell function known to be deregulated in cancer. One of the first direct links between particular mirnas and cancer came from observations by the group of Carlo Croce upon the investigation of chronic lymphocytic leukaemia (CLL), the most common form of adult leukaemia in the Western world. More than half of CLL cases present with deletions at the chromosome band 13q14, an abnormality that also occurs in 50% of mantle cell lymphomas, 16 40% of multiple myelomas, and in 60% of prostate cancers. Although strongly suggestive for tumour suppressor
17 Table 2. Overview of mirnas frequently inactivated in cancer mirna deregulation in cancer functions let-7 family Downregulated in lung, breast, gastric, ovary, prostate and colon cancers, chronic lymphoid leukemia, leiomyomas Represses cell proliferation and growth. Promotes angiogenesis. Negative regulation of oncoproteins RAS, MYC and HMGA2. mir-15a, mir-16-1 mir-29 family mir-34 family mir-143, mir-145 cluster mir-200 family mir-26a mir-125a, mir- 125b mir-101 Adapted from Spizzo et al., Downregulated in chronic lymphoid leukemia, diffuse large B-cell lymphoma, multiple myeloma, pituitary adenoma, prostate and pancreatic cancer. Germline mutations in B-cell chronic lymphoid leukemia patients. Downregulated in chronic lymphoid leukemia, colon, breast, and lung cancer, and cholangiocarcinomas Downregulated in pancreatic cancer and Burkitt s lymphoma without MYC translocation. Hypermethylation in colon cancer. Downregulated in neuroblastoma with 1p deletion Downregulated in colon adenoma/carcinoma, in breast, lung, and cervical cancer, in B cell malignancies Downregulated in clear-cell carcinoma, metastatic breast cancer Downregulated in hepatorcellular carcinoma, breast cancer, Burkitt lymphoma and anaplastic thyroid carcinoma. Downregulated by MYC Downregulated in glioblastoma, breast, prostate and ovarian cancer Downregulated in prostate cancer, hepatocellular carcinoma, bladder cancer and gastric cancer Induces apoptosis in leukemia cells by repression of BCL2. Regulates cell cycle by downregulating G0/G1 proteins. Induces aberrant methylation in lung cancer. Induces apoptosis. induces upregulation of p53, downregulation of E2F in colon cancer. Negative regulation of MYC and MYCN. Negative regulation of MYC and KRAS oncoproteins. Promotes invasion. Involved in TGFβ- mediated EMT. Negative regulation of ZEB1 and ZEB2 Induces apoptosis through negative regulation of MTDH and EZH2. Induces cell-cycle arrest associated with direct targeting of CCND2 and CCNE2. Negative regulation of ERBB2, ERBB3 and LIN28 oncoproteins Induces alterations in global chromatin structure via repression of EZH2. Inhibits proliferation, migration and invasion. Sensitizes tumour cells to radiation 11 activity, involvement of any of the protein- coding genes in the deleted region could not be demonstrated. Interestingly, two mirnas, mir- 15a and mir- 16-1, resided within a critically deleted 30- kb region at 13q14 and were reduced in expression in two thirds of CLL cases 81. Both mirnas were shown to negatively regulate BCL2, an anti- apoptotic gene frequently overexpressed in leukaemias, lymphomas and carcinomas. Down regulation of mir- 15a and mir is therefore believed to result in an increase of BCL2 expression and hence the inactivation of the intrinsic apoptosis pathway 82. The role of mir- 15a and mir as tumour suppressors in CLL is further supported by the presence of a pathogenic mutation in the mir- 15a mir genes in two patients, leading to a decreased mir expression 83. The sequence abnormalities that were identified in the mirna genes were not present in 160 normal control individuals and in several instances they were also found in DNA from normal cells of the patient. As CLL (as well as other cancers) is a disease with known familial occurrence (5% to 10% of patients have an inherited susceptibility to CLL), mirna mutations may be a predisposing factor to cancer, especially for those familial cases where the culprit genes are still unknown. Soon after this discovery, additional mirnas with a putative tumour suppressor function were described (Table 2). The mirnas that are encoded by the let- 7 family were shown to negatively regulate the expression of RAS through a direct interaction with its 3 - UTR 85. The RAS oncogene regulates proliferation and differentiation and is commonly mutated in human cancers, including lung cancer. When comparing lung tumours to normal adjacent cells, taken from patients
18 with squamous- cell carcinoma of the lung, let- 7 mirnas were found to be down regulated in the tumours whereas RAS was highly expressed. Besides RAS, let- 7 has also been shown to coordinate the repression of another growth promoting gene implicated in cancers, HMGA2 86. HMGA2 functions at the transcriptional level by altering chromatin structure and is primarily expressed in proliferating cells during embryogenesis and in a wide variety of tumours. In many of these tumours the HMGA2 open reading frame is truncated through chromosomal translocations resulting in a loss of the C- terminal domain. Together with loss of the C- terminal domain, such translocations also replace the 3 - UTR hereby disrupting the let- 7 coordinated repression of HMGA2 and promoting anchorage independent growth, a characteristic of oncogenic transformation. Moreover, transgenic mice that overexpressed wild- type Hmga2 had similar phenotypes to those expressing the truncated protein indicating that the disruption of a single mirna- target interaction can be sufficient to produce a clinical phenotype in vivo. 12 Table 3. Overview of mirnas frequently activated in cancer mirna deregulation in cancer functions mir cluster Overexpression in lung and colon cancer, lymphoma, multiple myeloma, medulloblastoma, neuroblastoma. Upregulated by MYC and MYCN Increases tumour growth and proliferation by negative regulation of cell cycle inhibitors. Promotes tumour angiogenesis through repression of THBS1 and CTGF. Anti-apoptotic activity through repression of BIM. Induces lymphoproliferative disease and autoimmunity. mir-106b-25 cluster mir-10b mir-21 mir-125a, mir- 125b mir-155 Overexpression in gastric, colon, and prostate cancer, neuroblastoma, multiple myeloma Overexpressed in metastatic breast cancer, nasopharyngeal carcinoma and malignant peripheral nerve sheath tumours Overexpression in glioblastoma, breast, lung, prostate, colon, stomach, esophageal, and cervical cancer, uterine leiomyosarcoma, diffuse large B-cell lymphoma, head and neck cancer Upregulation in myelodysplastic syndrome and acute myeloid leukemia with t(2;11)(p21;q23), urothelial carcinoma Overexpressed in pediatric Burkitt s lymphoma, Hodgkin s lymphoma, primary mediastinal lymphoma, diffuse large B-cell lymphoma, breast, lung, colon, pancreatic cancer Reduces apoptotic response after TGFβ stimulation via BIM. Increases tumour growth and proliferation by negative regulation of cell cycle inhibitors. Activates cell migration and extracellular matrix remodelling through negative regulation of HOXD10. Promotes tumourigenesis through negative regulation of NF1 Induces invasion and metastasis. Inhibits apoptosis through negative regulation of PDCD4. Inhibits negative regulators of the RAS/MEK/ERK pathway. Negative regulation of the TP53 tumour suppressor protein. Induces pre-b-cell proliferation, lymphoblastic leukemia and high-grade lymphoma. Promotes chemosensitivity through repression of FOXO3A. mir-26a Upregulated in glioblastoma Induces tumourigenesis by repression of PTEN, RB1, and MAP3K2/MEKK2 mir-181 family mir-221, mir-222 cluster mir-372, mir-373 cluster Adapted from Spizzo et al., Overexpressed in breast, pancreas, prostate cancer and hepatocellular carcinoma. Upregulated by MYCN in neuroblastoma Overexpressed in chronic lymphoid leukemia, thyroid papillary carcinoma, glioblastoma. Overexpression in testicular germ cell tumours, thyroid andenomas, esophageal cancer and metastatic breast cancer Enhances proliferation, migration and invasion. Promotes cancer cell proliferation. Impairs TRAIL-dependent response. Antagonizes p53-mediated CDK inhibition through repression of LATS2. Stimulates migration and invasion through negative regulation of CD44
19 13 Apart from tumour suppressor mirnas, several mirnas displaying oncogenic properties have also been identified (Table 3). The only mirna found to be overexpressed in almost any type of solid tumour (breast, colon, lung, prostate, stomach, pancreas, glioblastoma and uterine leiomyoma) is mir Functional data supporting the oncogenic role of mir- 21 came from a study by Chan and colleagues in glioblastoma. They observed that knock down of mir- 21 in glioblastoma cell lines induced a caspase- mediated apoptotic response 88. Further, studies in breast cancer cells showed that, upon transfection with anti- mir- 21, cell growth in vitro and tumour growth in vivo were suppressed due to increased apoptosis and decreased cell proliferation. The mir- 21 targets include the tumour suppressors TPM1 and PTEN, a gene frequently mutated in a variety of advanced tumours and implicated in the AKT survival pathway central in cancer development 89, 90. Another mirna, mir- 155, has been shown to accumulate in various types of B- cell malignancy (Hodgkin lymphomas and Burkitt lymphomas) where it functions as an oncogene in cooperation with MYC 91. Deregulated mir- 155 expression is an early event in oncogenesis. By creating the first transgenic mouse overexpressing a mirna gene, Costinean and colleagues demonstrated that mir- 155 overexpression in B- cells leads to the development of pre- leukemic B- cell proliferation followed by B- cell malignancy 92. mirnas regulated by MYC Several oncogenic mirnas are linked to the increased expression of MYC proteins. The best studied MYC regulated mirnas are the ones located within the mir cluster, residing in a non- protein- coding RNA on chromosome 13 (C13orf25) encoding six different mirnas (mir- 17, mir- 18a, mir- 19a, mir- 20a, mir- 19b and mir- 92a). Since this genomic region is frequently amplified in a subset of B- cell lymphomas, He and colleagues postulated that increased expression of this cluster contributes to cancer formation. This was tested with a mouse model for human B- cell lymphoma, driven by the presence of the MYC oncogene. Enforced expression of the mir b- 1 cluster (the vertebrate portion of the mir cluster) significantly accelerated the onset of tumour formation in the transgenic animals (~51 days vs. 3-6 months) 93. This was not the case when individual mirnas from the mir b- 1 cluster were introduced nor were there any effects with other unrelated mirnas. Lymphomas resulting from both MYC and mir b overexpression presented increased cell proliferation and decreased cell death. These results clearly point towards an oncogenic cooperation of several mirnas within the truncated mir cluster. The oncogenic properties of the mir cluster were confirmed using antisense technology in lung cancer 94. A microarray screen for MYC regulated mirnas identified elevated mir expression in a human B- cell line overexpressing MYC 95. Chromatin immunoprecipitation experiments confirmed MYC binding to elements in the mir promoter entailing direct regulation by MYC. Two mirnas in this cluster were shown to negatively regulate the expression of E2F1, an essential regulator of the initiation of DNA- replication during the cell cycle. Given the fact that E2F1 activity is also stimulated directly by MYC, this reveals a mechanism through which MYC simultaneously activates E2F1 transcription and limits its translation, allowing a tightly controlled proliferative signal. Dews and colleagues demonstrated that mir mirnas promote an angiogenic switch in MYC- activated tumours. Transduction of cells with a mir encoding retrovirus reduced TSP1 and CTGF levels and cells formed larger, better- perfused tumours 60. In neuroblastoma, mirna expression profiling identified increased mir expression in MYCN amplified tumours and MYCN was shown to bind the mir promoter Overexpression of mir in neuroblastoma cells without MYCN amplification strongly increased their in vitro proliferation and in vivo tumourigenesis through mir- 17- mediated repression of CDKN1A and BIM. In addition, mir- 18a and mir- 19a were shown to target and repress the expression of ESR1, a transcription factor implicated in neuronal differentiation 99. Inhibition of mir- 18a in neuroblastoma cells led to severe growth retardation, outgrowth of neurites, and induction of neuronal sympathetic differentiation markers. MYC and MYCN have also been shown to activate the expression of mir In breast cancer cells, mir- 9 directly targets
20 14 CDH1 leading to increased cell motility and invasiveness. In addition, mir- 9- mediated CDH1 down regulation results in the activation of beta- catenin signaling, up regulation of VEGF and ultimately increased tumour angiogenesis. As beta- catenin activation also results in transcriptional upregulation of MYC, MYC- induced mir- 9 expression might induce a feed- forward loop resulting in increased MYC expression. MYC has also been shown to repress mirna expression. Through the analysis of B- cell lymphoma models with inducible MYC expression, Chang and colleagues unexpectedly found a widespread repression of mirnas upon MYC activation 101. Much of the repressed mirnas were known to have tumour suppressor functions such as let- 7 family members, mir- 15 and mir- 34a. Chromatin immunoprecipitation revealed that much of the repression was likely to be a direct result of MYC binding to mirna promoters. These mirnas were shown to be relevant in the process of tumourigenesis as enforced expression of repressed mirnas diminished the tumourigenic potential of lymphoma cells. These studies clearly demonstrate that activation of MYC genes has a profound effect on the mirna transcriptome and that deregulated mirnas contribute to tumourigenesis. MiR- 34a: the missing piece in the p53 network puzzle The TP53 gene plays a central role as a stress sensor for the cell. It can arrest cell growth in order to allow DNA repair or can eliminate cells with severe damage by activating the apoptotic pathway. Given this important protective role, it is not surprising that TP53 is functionally inactivated in most if not all tumours, either through mutations or through inhibition of its function by alterations in up or down stream mediators. Of further interest, TP53 mutations are often found in aggressive tumours with poor prognosis. Neuroblastoma tumours harbor relatively few TP53 mutations at diagnosis whereas tumours from relapse patients often present with abnormalities in the TP53 pathway 102. Despite intensive efforts, certain issues concerning TP53 regulation remained unresolved. One such enigma was the fact that evidence was found for a role of TP53 as transcriptional repressor, although all available insights clearly pointed at the gene being a transcriptional activator. The answer to this contradiction came from mirna studies. First, investigators profiling mirnas in neuroblastoma found decreased expression of mir- 34a. MiR- 34a is located on the short arm of chromosome 1 which is often deleted in these tumours 103. Soon after, several other studies demonstrated that mir- 34a was directly regulated by TP53 104, 105. Therefore, TP53 could indeed lead to indirect transcriptional repression of target genes, most likely due to down regulation by mir- 34a. Up regulation of mir- 34a induced cell cycle arrest, senescence or apoptosis, all known TP53 pathway mediated effects. These effects are mediated by a wide range of mir- 34a targets such as BCL2, CCND1, CCNE2, CDK4, CDK6 and many others 106. Of note, mir- 34a was also shown to target MYCN in neuroblastoma 107. This might explain the close association between 1p deletion and MYCN amplification as loss of mir- 34a would be necessary to maintain high levels of MYCN protein. Even before the mir- 34a discovery, two other mirnas were found to interfere with the TP53 pathway. Testicular germ cell tumours, known to have functional TP53, displayed increased expression of mir- 372 and mir- 373, overriding TP53 mediated cell cycle arrest 108. Such findings are critically important, as they unveil mechanisms that suppress normal TP53 function in cancer cells with otherwise functionally intact TP53 protein and thus offer insights to alternative therapeutic strategies for treatment of this substantially large group of TP53 intact tumours. mirnas and metastasis The most life- threatening characteristic of cancer cells is their possibility to acquire the capacity to invade and metastasize to other organs. In contrast to tumour initiation, our understanding of the alterations of genes controlling invasion and metastasis is still poor. Therefore, the recent findings as to how mirnas might coordinate some of the gene expression programs controlling these phenomena attracted much attention. Based on previously available mirna gene expression data in breast cancer, mir- 10b was found to be correlated with vascular invasion 109. TWIST, a gene known to control tumour cell motility, up
21 15 regulated mir- 10b which in its turn caused down regulation of HOXD10 expression, thereby affecting other genes involved in metastasis. Much like the TP53 work, this study showed how mirnas play crucial roles in signaling networks implicated in cancer. Further work on breast cancer also revealed a role for mir- 335 and mir- 126 as metastasis suppressors and association of low expression for these genes with poor distal metastasis- free survival 110. Mechanisms of deregulated mirna expression So far, four different mechanisms of aberrant mirna expression have been identified. These mechanisms can function both independently and in concert to disturb mirna expression patterns in human tissues and are described below. 1. The location of mirnas at cancer associated genomic regions About half of all mirnas reside within cancer associated genomic regions (CAGRs) 111. These regions are frequently altered in cancer cells through copy- number alterations and are thought to harbour tumour suppressor genes or oncogenes, depending on whether the particular region is deleted or amplified. This association emphasizes the importance of mirnas in tumour biology and is supported by functional evidence, directly linking mirnas at CAGRs with tumour progression. 2. Epigenetically regulated mirnas Like many coding genes, mirnas can be located near or within a CpG island. This led to the hypothesis that mirna expression could be under epigenetic control. Indeed, recent studies have shown that aberrant DNA methylation as well as chromatin modifications may serve as a mechanism for deregulated mirna gene expression in cancer. First, Saito and colleagues demonstrated that about 5% of investigated mirnas are up regulated by treatment of T24 bladder cancer cells with a DNA demethylating agent in combination with a histone deacetylase inhibitor 112. In particular, reactivation of mir- 127 led to the down regulation of its predicted target BCL6, a proto- oncogene implicated in the pathogenesis of B cell lymphoma. Meanwhile, additional studies have shown that mirna expression can be silenced by epigenetic mechanisms, e.g. mir- 124a 113. On the other hand, Brueckner and colleagues observed that mirna let- 7a- 3, embedded in a CpG island located on 22q12.31, is hypomethylated in some lung adenocarcinomas, while it is heavily methylated in normal human tissues 114. Epigenetic reactivation of this non- coding gene resulted in enhanced tumour types and oncogenic transcription profiles, which suggests that let- 7a- 3 acts as an oncogene. 3. Defects in the mirna- processing machinery Various proteins that are implicated in mirna- processing and mirna directed regulation of protein- coding mrnas have been linked to tumourigenesis. When examining the expression of DROSHA and DICER1 in 67 non- small lung cancer samples, Karube and colleagues found a reduced expression of DICER1 that correlated with shortened post- operative survival 115. As DICER1 is a crucial protein in the production of mature mirnas, reduction of its expression could potentially result in a decrease of tumour suppressor mirnas in the lung 116. Surprisingly, complete loss of DICER1 expression is selected against during tumourigenesis suggesting that DICER1 functions as a haploinsufficient tumour suppressor gene 117, 118. This was shown in different mouse models of cancer including retinoblastoma and lung cancer. Monoallelic loss of Dicer1 dramatically increased tumourigenesis whereas complete loss of Dicer1 did not. Defects in other components of the mirna processing machinery have also been reported. Melo and colleagues identified mutations in TARBP2, encoding an integral component of a DICER1- containing complex, in sporadic and hereditary carcinomas 119. These cancer types were also shown to harbour mutations in other mirna processing genes including AGO2, TNRC6A and XPO5 120, Mutations in mirna genes and mirna binding sites Although still scarce, evidence for pathogenic mutations in mirna genes suggests that these mutations might influence mirna expression and function, ultimately contributing to tumour onset or progression 83, 122. Thus far, the number of reported mirna mutations is relatively low compared to the total number of human mirnas. Apart from mutations
22 16 occurring within the mirna gene, 3 - UTR mutations creating or destroying functional mirna target sites could also affect normal mirna function. One study reports on the presence of a mir- 124 mirna binding site SNP in the 3 UTR of the DHFR gene leading to methotrexate resistance 123 whereas RNA sequencing of a patient with acute myeloid leukemia (AML) revealed a somatic mutation in the 3'UTR of TNFAIP2, a putative tumour suppressor gene in AML 124. The authors could prove that this mutation generates a new mirna binding site that leads to translational repression of TNFAIP2. mirna signatures for improved diagnostic and prognostic classification Despite the currently available imaging and molecular tools, diagnosis for some tumours may be uncertain, in particular for poorly differentiated cancer types. A study by Lu and colleagues showed that, albeit using a relatively low number of mirnas for profiling, the resulting signatures enabled accurate recognition of particular tumour entities, possibly due to a partly retained tissue (cell of origin) specific mirna signature 125. Following this study, an increasing number of mirna profiles of specific tumour entities have been generated (e.g. prostate and ovarian cancer, AML, and many others) in most if not all cases matching with known biopathological features. More importantly, some of these studies have demonstrated the prognostic power of the expression status of relative small number of mirnas, in contrast to coding gene expression studies that typically yielded larger lists of genes with prognostic value. MiRNA expression signatures were proven to be successful for the detection of the primary site of origin of metastatic disease, when the primary site remains obscure 126. Unknown primary malignancy is not an infrequent diagnosis for patients with metastatic disease. Patients with a known primary site have a better prognosis because this allows clinicians to apply specific therapeutic regimens. Therefore, mirna signatures that are able to predict the primary site can be of clinical relevance. MiRNA signatures have also proven to be powerful predictors of patient prognosis. Calin and colleagues were one of the first to demonstrate that a 13 mirna signature was associated with disease progression in patients with chronic lymphocytic leukemia 83. Several additional reports were published onwards, verifying the use of mirna expression signatures to predict patient prognosis for different cancer types including neuroblastoma Recently, a number of labs have shown that mirnas can also be detected in patient serum and other body fluids such as sputum and urine. In contrast to mrnas, circulating mirnas appear to be resistant to degradation making them excellent candidate biomarkers. Serum mirna signatures have indeed been shown to correlate to patient diagnosis and prognosis, both for cancer and other human diseases This discovery has important implications, both for the clinic and the patient, as serum collection is a fast, safe and non- invasive procedure. mirna therapeutics The observation that mirna expression is deregulated in cancer and that targeting mirna expression modifies the cancer phenotype suggests that mirnas could serve as therapeutic targets. Because of their ability to simultaneously regulate numerous protein- coding genes, implicated in one single or different disease- associated networks, targeting a single mirna might be more efficient than targeting the individual protein- coding genes. This was elegantly demonstrated by Li and colleagues in the context of T- cell receptor signaling and antigen recognition during T- cell maturation 137. This process is controlled by phosphorylation and dephosphorylation events involving more than 40 different kinases and phosphatases. By down regulating multiple phosphatases, mir- 181a acts as a critical regulator of T- cell receptor sensitivity, a task that can be carried out by mir- 181a on itself but not by shrna- directed downregulation of individual mir- 181a target genes. In cancer, mirnas were shown to play a role in different signaling networks controlling essential cancer- related processes. In this way, deregulated expression of a small number of mirnas can affect multiple tumour suppressors and oncogenes that contribute to the cancer phenotype. This is exemplified by activation of the mir cluster, resulting in accelerated cell
23 17 proliferation (through repression of CDKN1A), inhibition of apoptosis (through repression of BIM) and tumour angiogenesis (through repression of CTGF and THBS1). Different strategies have been developed to modulate the expression of a tumour suppressor mirna or oncomir. These commonly rely on the use of oligonucleotides or viral constructs to antagonize the expression or function of an oncomir or to increase the expression of a tumour suppressor mirna. Blocking mirnas in vivo can be achieved using antisense oligonucleotides that are modified in order to increase their stability, binding affinity and specificity. By using of 2 - O- methyl groups to improve RNA binding affinity and cholesterol conjugation to increase delivery, Krutzfeldt and colleagues developed antagomir molecules to silence mir- 122 and mir- 16 in vivo 138. One day after tail vein injection, mirna expression was efficiently silenced in different tissues and the effect was still observed 3 weeks after the injection. Alternatively, antisense oligonucleotides can be modified using nucleotide analogues called locked nucleic acids (LNA). These LNA modified oligonucleotides show a superior hybridization affinity and specificity compared to any other class of oligonucleotides 139. Their application in vivo was demonstrated for mir- 122 in non- human primates. Therapeutic silencing of mir- 122 in chimpanzees with chronic hepatitis C virus infection using anti- mir- 122 (Miravirsen, Santaris Pharma) resulted in a long- lasting suppression of the virus with no evidence of viral resistance or side effects in the treated animals 140. Miravirsen is currently entering phase 2 clinical trials to treat patients with hepatitis C. These studies suggest that antagonizing mirna expression in vivo is feasible and the same technological approaches can be taken to repress the expression of oncomirs in cancer. Similarly, oligonucleotides can be applied to restore the expression of tumour suppressor mirnas. These oligonucleotides are identical to the sequence of the tumour suppressor mirna and are commonly termed mirna mimics. While mirna mimics have been extensively evaluated in in vitro studies, there is no data yet suggesting that they can be delivered through intravenous injection in vivo. In contrast, adenovirus associated vectors expressing the tumour suppressor mirna were shown to restore mirna expression upon intravenous injection. This was demonstrated by Kota and colleagues for mir- 26a in a mouse model of liver cancer 141. Restoration of mir- 26a expression reduced tumourigenicity without any signs of toxicity in other organs. The fact that mir- 26a is highly expressed in all normal tissues probably explains why mir- 26a delivery is tolerated in normal tissues and may provide a general strategy for mirna replacement therapies. Quantification of mirna expression Due to their small size, accurate quantification of mirna expression is a major challenge in the field. Several hybridisation- based methods, such as microarray , bead- based flow cytometry 125, and small- RNA sequencing 124, have been introduced to quantify the expression of hundreds of mirnas in a single experiment. However, these approaches require substantial amounts of input RNA which precludes the use of small biopsies, single cells or body fluids such as serum, plasma, urine or sputum. While the reverse transcription quantitative PCR (RT- qpcr) in principle has a much higher sensitivity, down to a single molecule, the RT reaction requires modification in order to enable the detection of small RNA molecules such as mirnas. One approach relies on the use of stem- loop RT primers 151 while another is based on polyadenylation of the mature mirna prior to oligo- dt primed cdna synthesis 152. Next to sensitivity, RT- qpcr based approaches have a superior specificity, linear dynamic range of quantification and a high level of flexibility, allowing additional assays to be readily included in the workflow. Stem- loop reverse transcription is based on the use of a looped mirna specific RT- primer that will hybridise to the 3 end of the mature mirna to initiate cdna synthesis (Figure 6A). Upon denaturation, the loop unfolds, providing a longer template for detection in a qpcr reaction. Since this process is mirna specific, multiplex pooling of individual stem- loop primers is necessary in order to produce cdna template for multiple mirnas. An optional limited- cycle pre- amplification step is introduced to increase the sensitivity of the reaction,
24 enabling mirna profiling studies of single cells and body fluids. An alternative approach to elongate the template is polyadenylation of the mature mirna (Figure 6B). Reverse transcription is then initiated using a polyt primer that can be tagged. This reaction is universal, providing cdna template for quantification of any mirna. The use of LNA- modified primers precludes the need for a pre- amplification step and enables the study of mirna expression when limited amounts of RNA are available. 18 A stem-loop RT B universal RT reverse transcription mature mirna reverse transcription mature mirna AAAAAAAAA TTTTTTTTTT F primer F primer quantitative PCR quantitative PCR TTTTTTTTTT Q probe F R primer R primer Figure 6 RT-qPCR mirna expression quantification. Schematic overview of the stem-loop RT-qPCR (A) and universal RT-qPCR (B) mirna profiling platforms. Exploring mirna function The function of a mirna is uniquely defined by its mrna target specificity. Negative regulation of these targets is primarily mediated through imperfect binding between the mirna and a mirna binding site in the 3 UTR of the target mrna. A major determinant of mirna target recognition appears to be the mirna seed, a 6-8 nucleotide region at the 5 end of the mirna that pairs with the target mrna. In general, repression of the target is more efficient for a 8mer seed match compared to a 7mer seed match, which in turn is more efficient than a 6mer seed match 153, 154. In addition, several features were uncovered that boost site efficacy including AU- rich nucleotide composition near the site, proximity to sites for co- expressed mirnas (which leads to cooperative action), proximity to residues pairing to mirna nucleotides 13 16, positioning within the 3ʹ UTR at least 15 nucleotides from the stop codon, and positioning away from the center of long UTRs 155. MiRNA target recognition features can be used to predict putative mirna target genes and various algorithms have been developed to serve that purpose Unfortunately, mirna target prediction algorithms are prone to false positives and ignore the possible tissue- or disease- specific nature of a mirna target interaction. Therefore, different experimental approaches were developed that enable mirna target identification. One approach is based on the perturbation of the mirna of interest, followed by mrna or protein profiling. Protein profiling, usually using mass spectrometry, has the advantage that it will also identify mirna targets regulated through translational inhibition alone. On the other hand, current protein profiling technology generates expression data for just a fraction of the genes in the genome (typically around 3000 genes) while mrna profiling covers the entire genome. Another approach relies on the precipitation of mirna- bound mrnas through pull- down of one of the argonaute proteins 160, 161. The precipitated mrnas are identified and quantified either through hybridization on a microarray (Rip- Chip) or through high- throughput sequencing (HITS- CLIP). Although technically challenging, this approach has the advantage that it only identifies direct mirna targets.
25 Functional exploration of gene expression patterns The development of high- throughput screening platforms has enabled researchers to analyze entire transcriptomes in a single experiment. Microarray technology or high- throughput RT- qpcr readily provide high- quality expression data for all protein- coding genes or mirnas while next generation sequencing provides the possibility to measure the abundance of every transcript in an unbiased manner. While next generation sequencing is still in its infancy, especially in terms of data analysis, microarray- based expression profiling has become a standard functional genomics tool in molecular biology. The relatively low cost of a microarray experiment and the straightforward data analysis are two aspects that have contributed to the success of this technology. During the last decade, numerous gene expression studies have been conducted in order to gain insights in the pathways and signaling processes that are deregulated in human disease or that are downstream of a particular chemical or genetic perturbation. However, extracting biologically relevant information from gene expression data was soon appreciated as a major challenge. To address this problem, different bioinformatics tools were developed. Next to the actual expression data, these tools typically require a functional annotation of the individual genes that are measured. Several resources are available that provide different levels of gene annotation. The Gene Ontology (GO) consortium provides one such resource. The GO project has created a structured and standardized vocabulary that describes genes in terms of their associated biological processes, their molecular function or cellular components. All ontologies are species specific and describe how genes behave in a cellular context. Another resource of functional gene annotation is provided by KEGG 162, 163. The KEGG database contains higher order functional information represented as pathways or networks of genes and interacting molecules in the cell. Different aspects of cell biology are covered including metabolism, genetic and environmental information processing, cellular processes and organismal systems. In contrast to GO, the KEGG database also contains pathways for molecular systems in perturbed states, e.g. caused by disease. Several multifactorial diseases are included such as cancers, immune system diseases, neurodegenerative diseases, cardiovascular diseases, and metabolic diseases. Different aspects need to be taken into account in order to uncover the pathways underlying differential gene expression. First, a certain pathway or process can be perturbed when only a subset of genes within that pathway are differentially expressed. For instance, the expression of genes located upstream of the molecular defect that is causing the perturbation would typically be unaffected while those located downstream would be differentially expressed. Alternatively, the molecular defect might induce changes that only occur on the protein level, e.g. phosphorylation, and leave gene expression levels unaffected. Second, a small fold change of all genes in a pathway may dramatically alter the activity of that pathway and be more important than a large fold change of only one or few genes. Different methods have been developed to take into account one ore more of these aspects in order to identify the relevant pathways from a RNA expression profiling experiment. One approach is based on Fisher s Exact statistics and calculates the enrichment of a set of genes, annotated to a certain pathway, among an experimentally derived list of differentially expressed genes. Several publically available web tools such as DAVID 165 and GoMiner 166 are based on this principle. The major disadvantage of this approach is that it requires an upfront selection of differentially expressed genes. Often, biological differences are modest relative to the noise inherent to a gene expression experiment. Therefore, few genes may meet the threshold for statistical significance after multiple testing corrections. To better address these issues, Subramanian and colleagues developed a method, called Gene Set Enrichment Analysis (GSEA), to identify functions underlying microarray expression data by using the entire dataset rather than a selection of differentially expressed genes 164 (Figure 7). All measured genes are ranked based on their correlation to any of two classes, defined as the phenotype (i.e. treated vs. untreated cells or normal vs. disease), using any suitable metric (i.e. fold change or correlation coefficient). GSEA then determines whether the genes from an annotated gene set, representing a pathway or process, are randomly distributed along the ranked list of genes. This is done by 19
26 calculating an enrichment score that reflects the degree to which the annotated gene set is overrepresented at the extremes (top or bottom) of the entire ranked list. To accompany the GSEA algorithm, the authors put together a compendium of annotated gene sets, available in the Molecular Signatures Database, that extends beyond the functional annotations provided by GO and KEGG. These gene sets represent a multitude of chemical and genetic perturbations in different tissues, both normal and disease, obtained through extensive literature searches. By focusing on gene sets rather than differentially expressed genes, GSEA is able to extract relevant biological insights from gene expression data even if changes at the level of individual genes are subtle. 20 Figure 7 Overview of the gene set enrichment analysis method. (A) Gene list, ranked according to differential expression between two phenotypic classes. The position of genes from a given gene set S in the ranked gene list is marked by horizontal lines. (B) Plot of the running sum for gene set S in the dataset. The maximum enrichment score and corresponding leading edge subset are indicated. A score is calculated by walking down the ranked list, increasing a running-sum statistic when a gene from the gene set is encountered and decreasing it when the gene is not part of the gene set. The enrichment score is the maximum deviation from zero encountered in the random walk. The significance of the enrichment score is estimated by a phenotype permutation test. The leading edge subset represents the core members of the gene set that contribute to the enrichment score. (Source: Subramanian et al., ) References 1. Maris, J.M. & Matthay, K.K. Molecular biology of neuroblastoma. J Clin Oncol 17, (1999). 2. Brodeur, G.M. Neuroblastoma: biological insights into a clinical enigma. Nat Rev Cancer 3, (2003). 3. Brodeur, G.M. et al. Revisions of the international criteria for neuroblastoma diagnosis, staging, and response to treatment. J Clin Oncol 11, (1993).
27 4. Monclair, T. et al. The International Neuroblastoma Risk Group (INRG) staging system: an INRG Task Force report. J Clin Oncol 27, (2009). 5. Knudson, A.G., Jr. & Strong, L.C. Mutation and cancer: neuroblastoma and pheochromocytoma. Am J Hum Genet 24, (1972). 6. Kushner, B.H., Gilbert, F. & Helson, L. Familial neuroblastoma. Case reports, literature review, and etiologic considerations. Cancer 57, (1986). 7. Maris, J.M. et al. Molecular genetic analysis of familial neuroblastoma. Eur J Cancer 33, (1997). 8. Mosse, Y.P. et al. Germline PHOX2B mutation in hereditary neuroblastoma. Am J Hum Genet 75, (2004). 9. Mosse, Y.P. et al. Identification of ALK as a major familial neuroblastoma predisposition gene. Nature 455, (2008). 10. De Brouwer, S. et al. Meta- analysis of neuroblastomas reveals a skewed ALK mutation spectrum in tumours with MYCN amplification. Clin Cancer Res 16, (2010). 11. Maris, J.M. et al. Evidence for a hereditary neuroblastoma predisposition locus at chromosome 16p Cancer Res 62, (2002). 12. Perri, P. et al. Weak linkage at 4p16 to predisposition for human neuroblastoma. Oncogene 21, (2002). 13. Capasso, M. et al. Common variations in BARD1 influence susceptibility to high- risk neuroblastoma. Nat Genet 41, (2009). 14. Maris, J.M. et al. Chromosome 6p22 locus associated with clinically aggressive neuroblastoma. N Engl J Med 358, (2008). 15. Wang, K. et al. Integrative genomics identifies LMO1 as a neuroblastoma oncogene. Nature (2010). 16. Michels, E. et al. ArrayCGH- based classification of neuroblastoma into genomic subgroups. Genes Chromosomes Cancer 46, (2007). 17. Bown, N. et al. Gain of chromosome arm 17q and adverse outcome in patients with neuroblastoma. N Engl J Med 340, (1999). 18. Janoueix- Lerosey, I. et al. Overall genomic pattern is a predictor of outcome in neuroblastoma. J Clin Oncol 27, (2009). 19. Vandesompele, J. et al. Unequivocal delineation of clinicogenetic subgroups and development of a new model for improved outcome prediction in neuroblastoma. J Clin Oncol 23, (2005). 20. Maris, J.M., Hogarty, M.D., Bagatell, R. & Cohn, S.L. Neuroblastoma. Lancet 369, (2007). 21. Holzel, M. et al. NF1 is a tumour suppressor in neuroblastoma that determines retinoic acid response and disease outcome. Cell 142, (2010). 22. Slamon, D.J. et al. Human breast cancer: correlation of relapse and survival with amplification of the HER- 2/neu oncogene. Science 235, (1987). 23. Wang, S.I., Parsons, R. & Ittmann, M. Homozygous deletion of the PTEN tumour suppressor gene in a subset of prostate adenocarcinomas. Clin Cancer Res 4, (1998). 24. Thompson, P.M. et al. Homozygous deletion of CDKN2A (p16ink4a/p14arf) but not within 1p36 or at other tumour suppressor loci in neuroblastoma. Cancer Res 61, (2001). 25. Martinsson, T., Sjoberg, R.M., Hedborg, F. & Kogner, P. Homozygous deletion of the neurofibromatosis- 1 gene in the tumour of a patient with neuroblastoma. Cancer Genet Cytogenet 95, (1997). 26. Corvi, R., Savelyeva, L., Amler, L., Handgretinger, R. & Schwab, M. Cytogenetic evolution of MYCN and MDM2 amplification in the neuroblastoma LS tumour and its cell line. Eur J Cancer 31A, (1995). 21
28 27. Corvi, R. et al. Non- syntenic amplification of MDM2 and MYCN in human neuroblastoma. Oncogene 10, (1995). 28. Caren, H., Abel, F., Kogner, P. & Martinsson, T. High incidence of DNA mutations and gene amplifications of the ALK gene in advanced sporadic neuroblastoma tumours. Biochem J 416, (2008). 29. Schwab, M. et al. Amplified DNA with limited homology to myc cellular oncogene is shared by human neuroblastoma cell lines and a neuroblastoma tumour. Nature 305, (1983). 30. Corvi, R., Amler, L.C., Savelyeva, L., Gehring, M. & Schwab, M. MYCN is retained in single copy at chromosome 2 band p23-24 during amplification in human neuroblastoma cells. Proc Natl Acad Sci U S A 91, (1994). 31. Schwab, M. et al. Chromosome localization in normal human cells and neuroblastomas of a gene related to c- myc. Nature 308, (1984). 32. Reiter, J.L. & Brodeur, G.M. High- resolution mapping of a 130- kb core region of the MYCN amplicon in neuroblastomas. Genomics 32, (1996). 33. Bordow, S.B., Norris, M.D., Haber, P.S., Marshall, G.M. & Haber, M. Prognostic significance of MYCN oncogene expression in childhood neuroblastoma. J Clin Oncol 16, (1998). 34. Chan, H.S. et al. MYCN protein expression as a predictor of neuroblastoma prognosis. Clin Cancer Res 3, (1997). 35. Weiss, W.A., Aldape, K., Mohapatra, G., Feuerstein, B.G. & Bishop, J.M. Targeted expression of MYCN causes neuroblastoma in transgenic mice. EMBO J 16, (1997). 36. Vennstrom, B., Sheiness, D., Zabielski, J. & Bishop, J.M. Isolation and characterization of c- myc, a cellular homolog of the oncogene (v- myc) of avian myelocytomatosis virus strain 29. J Virol 42, (1982). 37. Dalla- Favera, R. et al. Human c- myc onc gene is located on the region of chromosome 8 that is translocated in Burkitt lymphoma cells. Proc Natl Acad Sci U S A 79, (1982). 38. Gelmann, E.P., Psallidopoulos, M.C., Papas, T.S. & Dalla- Favera, R. Identification of reciprocal translocation sites within the c- myc oncogene and immunoglobulin mu locus in a Burkitt lymphoma. Nature 306, (1983). 39. Little, C.D., Nau, M.M., Carney, D.N., Gazdar, A.F. & Minna, J.D. Amplification and expression of the c- myc oncogene in human lung cancer cell lines. Nature 306, (1983). 40. Berns, E.M. et al. TP53 and MYC gene alterations independently predict poor prognosis in breast cancer patients. Genes Chromosomes Cancer 16, (1996). 41. Nau, M.M. et al. L- myc, a new myc- related gene amplified and expressed in human small cell lung cancer. Nature 318, (1985). 42. Schwab, M. MYCN in neuronal tumours. Cancer Lett 204, (2004). 43. Malynn, B.A. et al. N- myc can functionally replace c- myc in murine development, cellular growth, and differentiation. Genes Dev 14, (2000). 44. Van Roy, N. et al. The emerging molecular pathogenesis of neuroblastoma: implications for improved risk assessment and targeted therapy. Genome Med 1, 74 (2009). 45. Grandori, C., Cowley, S.M., James, L.P. & Eisenman, R.N. The Myc/Max/Mad network and the transcriptional control of cell behavior. Annu Rev Cell Dev Biol 16, (2000). 46. Amati, B. et al. Transcriptional activation by the human c- Myc oncoprotein in yeast requires interaction with Max. Nature 359, (1992). 47. Wanzel, M., Herold, S. & Eilers, M. Transcriptional repression by Myc. Trends Cell Biol 13, (2003). 48. Gartel, A.L. et al. Myc represses the p21(waf1/cip1) promoter and interacts with Sp1/Sp3. Proc Natl Acad Sci U S A 98, (2001). 22
29 49. Yang, W. et al. Repression of transcription of the p27(kip1) cyclin- dependent kinase inhibitor gene by c- Myc. Oncogene 20, (2001). 50. Staller, P. et al. Repression of p15ink4b expression by Myc through association with Miz- 1. Nat Cell Biol 3, (2001). 51. Wu, S. et al. Myc represses differentiation- induced p21cip1 expression via Miz- 1- dependent interaction with the p21 core promoter. Oncogene 22, (2003). 52. Kime, L. & Wright, S.C. Mad4 is regulated by a transcriptional repressor complex that contains Miz- 1 and c- Myc. Biochem J 370, (2003). 53. Patel, J.H., Loboda, A.P., Showe, M.K., Showe, L.C. & McMahon, S.B. Analysis of genomic targets reveals complex functions of MYC. Nat Rev Cancer 4, (2004). 54. Varlakhanova, N.V. & Knoepfler, P.S. Acting locally and globally: Myc's ever- expanding roles on chromatin. Cancer Res 69, (2009). 55. Martinato, F., Cesaroni, M., Amati, B. & Guccione, E. Analysis of Myc- induced histone modifications on target chromatin. PLoS One 3, e3650 (2008). 56. Cotterman, R. et al. N- Myc regulates a widespread euchromatic program in the human genome partially independent of its role as a classical transcription factor. Cancer Res 68, (2008). 57. Meyer, N. & Penn, L.Z. Reflecting on 25 years with MYC. Nat Rev Cancer 8, (2008). 58. Larsson, L.G. & Henriksson, M.A. The Yin and Yang functions of the Myc oncoprotein in cancer development and as targets for therapy. Exp Cell Res 316, (2010). 59. Vander Heiden, M.G., Cantley, L.C. & Thompson, C.B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, (2009). 60. Dews, M. et al. Augmentation of tumour angiogenesis by a Myc- activated microrna cluster. Nat Genet 38, (2006). 61. Watnick, R.S., Cheng, Y.N., Rangarajan, A., Ince, T.A. & Weinberg, R.A. Ras modulates Myc activity to repress thrombospondin- 1 expression and increase tumour angiogenesis. Cancer Cell 3, (2003). 62. Felsher, D.W. & Bishop, J.M. Transient excess of MYC activity can elicit genomic instability and tumourigenesis. Proc Natl Acad Sci U S A 96, (1999). 63. de Alboran, I.M., Baena, E. & Martinez, A.C. c- Myc- deficient B lymphocytes are resistant to spontaneous and induced cell death. Cell Death Differ 11, (2004). 64. Campaner, S. et al. Cdk2 suppresses cellular senescence induced by the c- myc oncogene. Nat Cell Biol 12, 54-59; sup pp (2010). 65. Zhuang, D. et al. C- MYC overexpression is required for continuous suppression of oncogene- induced senescence in melanoma cells. Oncogene 27, (2008). 66. Hydbring, P. et al. Phosphorylation by Cdk2 is required for Myc to repress Ras- induced senescence in cotransformation. Proc Natl Acad Sci U S A 107, (2010). 67. Soucek, L. et al. Modelling Myc inhibition as a cancer therapy. Nature 455, (2008). 68. Slack, A. et al. The p53 regulatory gene MDM2 is a direct transcriptional target of MYCN in neuroblastoma. Proc Natl Acad Sci U S A 102, (2005). 69. Van Maerken, T. et al. Small- molecule MDM2 antagonists as a new therapy concept for neuroblastoma. Cancer Res 66, (2006). 70. Van Maerken, T. et al. Antitumour activity of the selective MDM2 antagonist nutlin- 3 against chemoresistant neuroblastoma with wild- type p53. J Natl Cancer Inst 101, (2009). 71. Hogarty, M.D. et al. ODC1 is a critical determinant of MYCN oncogenesis and a therapeutic target in neuroblastoma. Cancer Res 68, (2008). 72. Westermann, F. et al. Distinct transcriptional MYCN/c- MYC activities are associated with spontaneous regression or malignant progression in neuroblastomas. Genome Biol 9, R150 (2008). 23
30 73. Taft, R.J., Pang, K.C., Mercer, T.R., Dinger, M. & Mattick, J.S. Non- coding RNAs: regulators of disease. J Pathol 220, (2010). 74. Calin, G.A. et al. Ultraconserved regions encoding ncrnas are altered in human leukemias and carcinomas. Cancer Cell 12, (2007). 75. Lee, R.C., Feinbaum, R.L. & Ambros, V. The C. elegans heterochronic gene lin- 4 encodes small RNAs with antisense complementarity to lin- 14. Cell 75, (1993). 76. Griffiths- Jones, S., Grocock, R.J., van Dongen, S., Bateman, A. & Enright, A.J. mirbase: microrna sequences, targets and gene nomenclature. Nucleic Acids Res 34, D (2006). 77. Bartel, D.P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, (2004). 78. Kim, V.N., Han, J. & Siomi, M.C. Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol 10, (2009). 79. Friedman, R.C., Farh, K.K., Burge, C.B. & Bartel, D.P. Most mammalian mrnas are conserved targets of micrornas. Genome Res 19, (2009). 80. Mestdagh, P., Hoebeeck, J., Poppe, B., Vandesompele, J. & Speleman, F. MiRNAs as cancer genes: new targets for tumour classification, prognosis and therapy. BJMO 2 (2008). 81. Calin, G.A. et al. Frequent deletions and down- regulation of micro- RNA genes mir15 and mir16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci U S A 99, (2002). 82. Cimmino, A. et al. mir- 15 and mir- 16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci U S A 102, (2005). 83. Calin, G.A. et al. A MicroRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N Engl J Med 353, (2005). 84. Spizzo, R., Nicoloso, M.S., Croce, C.M. & Calin, G.A. SnapShot: MicroRNAs in Cancer. Cell 137, e581 (2009). 85. Johnson, S.M. et al. RAS is regulated by the let- 7 microrna family. Cell 120, (2005). 86. Mayr, C., Hemann, M.T. & Bartel, D.P. Disrupting the pairing between let- 7 and Hmga2 enhances oncogenic transformation. Science 315, (2007). 87. Calin, G.A. & Croce, C.M. Chromosomal rearrangements and micrornas: a new cancer link with clinical implications. J Clin Invest 117, (2007). 88. Chan, J.A., Krichevsky, A.M. & Kosik, K.S. MicroRNA- 21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res 65, (2005). 89. Zhu, S., Si, M.L., Wu, H. & Mo, Y.Y. MicroRNA- 21 targets the tumour suppressor gene tropomyosin 1 (TPM1). J Biol Chem 282, (2007). 90. Meng, F. et al. Involvement of human micro- RNA in growth and response to chemotherapy in human cholangiocarcinoma cell lines. Gastroenterology 130, (2006). 91. Eis, P.S. et al. Accumulation of mir- 155 and BIC RNA in human B cell lymphomas. Proc Natl Acad Sci U S A 102, (2005). 92. Costinean, S. et al. Pre- B cell proliferation and lymphoblastic leukemia/high- grade lymphoma in E(mu)- mir155 transgenic mice. Proc Natl Acad Sci U S A 103, (2006). 93. He, L. et al. A microrna polycistron as a potential human oncogene. Nature 435, (2005). 94. Matsubara, H. et al. Apoptosis induction by antisense oligonucleotides against mir- 17-5p and mir- 20a in lung cancers overexpressing mir Oncogene 26, (2007). 95. O'Donnell, K.A., Wentzel, E.A., Zeller, K.I., Dang, C.V. & Mendell, J.T. c- Myc- regulated micrornas modulate E2F1 expression. Nature 435, (2005). 96. Schulte, J.H. et al. MYCN regulates oncogenic MicroRNAs in neuroblastoma. Int J Cancer 122, (2008). 97. Fontana, L. et al. Antagomir- 17-5p abolishes the growth of therapy- resistant neuroblastoma through p21 and BIM. PLoS One 3, e2236 (2008). 24
31 98. Chen, Y. & Stallings, R.L. Differential patterns of microrna expression in neuroblastoma are correlated with prognosis, differentiation, and apoptosis. Cancer Res 67, (2007). 99. Loven, J. et al. MYCN- regulated micrornas repress estrogen receptor- alpha (ESR1) expression and neuronal differentiation in human neuroblastoma. Proc Natl Acad Sci U S A 107, (2010) Ma, L. et al. mir- 9, a MYC/MYCN- activated microrna, regulates E- cadherin and cancer metastasis. Nat Cell Biol 12, (2010) Chang, T.C. et al. Widespread microrna repression by Myc contributes to tumourigenesis. Nat Genet 40, (2008) Carr- Wilkinson, J. et al. High Frequency of p53/mdm2/p14arf Pathway Abnormalities in Relapsed Neuroblastoma. Clin Cancer Res 16, (2010) Welch, C., Chen, Y. & Stallings, R.L. MicroRNA- 34a functions as a potential tumour suppressor by inducing apoptosis in neuroblastoma cells. Oncogene 26, (2007) Raver- Shapira, N. et al. Transcriptional activation of mir- 34a contributes to p53- mediated apoptosis. Mol Cell 26, (2007) He, L. et al. A microrna component of the p53 tumour suppressor network. Nature 447, (2007) Hermeking, H. The mir- 34 family in cancer and apoptosis. Cell Death Differ 17, (2010) Wei, J.S. et al. The MYCN oncogene is a direct target of mir- 34a. Oncogene 27, (2008) Voorhoeve, P.M. et al. A genetic screen implicates mirna- 372 and mirna- 373 as oncogenes in testicular germ cell tumours. Cell 124, (2006) Ma, L., Teruya- Feldstein, J. & Weinberg, R.A. Tumour invasion and metastasis initiated by microrna- 10b in breast cancer. Nature 449, (2007) Tavazoie, S.F. et al. Endogenous human micrornas that suppress breast cancer metastasis. Nature 451, (2008) Calin, G.A. et al. Human microrna genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci U S A 101, (2004) Saito, Y. et al. Specific activation of microrna- 127 with downregulation of the proto- oncogene BCL6 by chromatin- modifying drugs in human cancer cells. Cancer Cell 9, (2006) Lujambio, A. & Esteller, M. CpG island hypermethylation of tumour suppressor micrornas in human cancer. Cell Cycle 6, (2007) Brueckner, B. et al. The human let- 7a- 3 locus contains an epigenetically regulated microrna gene with oncogenic function. Cancer Res 67, (2007) Karube, Y. et al. Reduced expression of Dicer associated with poor prognosis in lung cancer patients. Cancer Sci 96, (2005) Esquela- Kerscher, A. & Slack, F.J. Oncomirs - micrornas with a role in cancer. Nat Rev Cancer 6, (2006) Lambertz, I. et al. Monoallelic but not biallelic loss of Dicer1 promotes tumourigenesis in vivo. Cell Death Differ 17, (2010) Kumar, M.S. et al. Dicer1 functions as a haploinsufficient tumour suppressor. Genes Dev 23, (2009) Melo, S.A. et al. A TARBP2 mutation in human cancer impairs microrna processing and DICER1 function. Nat Genet 41, (2009) Kim, M.S. et al. Somatic mutations and losses of expression of microrna regulation- related genes AGO2 and TNRC6A in gastric and colorectal cancers. J Pathol 221, (2010) Melo, S.A. et al. A genetic defect in exportin- 5 traps precursor micrornas in the nucleus of cancer cells. Cancer Cell 18, (2010). 25
32 122. Wojcik, S.E. et al. Non- codingrna sequence variations in human chronic lymphocytic leukemia and colorectal cancer. Carcinogenesis 31, (2010) Mishra, P.J., Humeniuk, R., Longo- Sorbello, G.S., Banerjee, D. & Bertino, J.R. A mir- 24 microrna binding- site polymorphism in dihydrofolate reductase gene leads to methotrexate resistance. Proc Natl Acad Sci U S A 104, (2007) Ramsingh, G. et al. Complete characterization of the micrornaome in a patient with acute myeloid leukemia. Blood 116, (2010) Lu, J. et al. MicroRNA expression profiles classify human cancers. Nature 435, (2005) Barker, E.V. et al. microrna evaluation of unknown primary lesions in the head and neck. Mol Cancer 8, 127 (2009) Yanaihara, N. et al. Unique microrna molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 9, (2006) Porkka, K.P. et al. MicroRNA expression profiling in prostate cancer. Cancer Res 67, (2007) Marcucci, G. et al. MicroRNA expression in cytogenetically normal acute myeloid leukemia. N Engl J Med 358, (2008) Hu, X. et al. A microrna expression signature for cervical cancer prognosis. Cancer Res 70, (2010) Bray, I. et al. Widespread dysregulation of MiRNAs by MYCN amplification and chromosomal imbalances in neuroblastoma: association of mirna expression with survival. PLoS One 4, e7850 (2009) Schulte, J.H. et al. Accurate prediction of neuroblastoma outcome based on mirna expression profiles. Int J Cancer 127, (2010) Liu, R. et al. A five- microrna signature identified from genome- wide serum microrna expression profiling serves as a fingerprint for gastric cancer diagnosis. Eur J Cancer (2010) Li, L.M. et al. Serum microrna Profiles Serve as Novel Biomarkers for HBV Infection and Diagnosis of HBV- Positive Hepatocarcinoma. Cancer Res 70, (2010) Moltzahn, F. et al. Microfluidic based multiplex qrt- PCR identifies diagnostic and prognostic microrna signatures in sera of prostate cancer patients. Cancer Res (2010) Van Pottelberge, G.R. et al. MicroRNA Expression in Induced Sputum of Smokers and Patients with Chronic Obstructive Pulmonary Disease. Am J Respir Crit Care Med (2010) Li, Q.J. et al. mir- 181a is an intrinsic modulator of T cell sensitivity and selection. Cell 129, (2007) Krutzfeldt, J. et al. Silencing of micrornas in vivo with 'antagomirs'. Nature 438, (2005) Vester, B. & Wengel, J. LNA (locked nucleic acid): high- affinity targeting of complementary RNA and DNA. Biochemistry 43, (2004) Lanford, R.E. et al. Therapeutic silencing of microrna- 122 in primates with chronic hepatitis C virus infection. Science 327, (2010) Kota, J. et al. Therapeutic microrna delivery suppresses tumourigenesis in a murine liver cancer model. Cell 137, (2009) Castoldi, M. et al. A sensitive array for microrna expression profiling (michip) based on locked nucleic acids (LNA). RNA 12, (2006) Liu, C.G. et al. An oligonucleotide microchip for genome- wide microrna profiling in human and mouse tissues. Proc Natl Acad Sci U S A 101, (2004) Nelson, P.T. et al. Microarray- based, high- throughput gene expression profiling of micrornas. Nat Methods 1, (2004). 26
33 145. Sioud, M. & Rosok, O. Profiling microrna expression using sensitive cdna probes and filter arrays. Biotechniques 37, , (2004) Thomson, J.M., Parker, J., Perou, C.M. & Hammond, S.M. A custom microarray platform for analysis of microrna gene expression. Nat Methods 1, (2004) Schulte, J.H. et al. Deep sequencing reveals differential expression of micrornas in favorable versus unfavorable neuroblastoma. Nucleic Acids Res 38, (2010) Kuchenbauer, F. et al. In- depth characterization of the microrna transcriptome in a leukemia progression model. Genome Res 18, (2008) Morin, R.D. et al. Application of massively parallel sequencing to microrna profiling and discovery in human embryonic stem cells. Genome Res 18, (2008) Linsen, S.E. et al. Limitations and possibilities of small RNA digital gene expression profiling. Nat Methods 6, (2009) Chen, C. et al. Real- time quantification of micrornas by stem- loop RT- PCR. Nucleic Acids Res 33, e179 (2005) Shi, R. & Chiang, V.L. Facile means for quantifying microrna expression by real- time PCR. Biotechniques 39, (2005) Baek, D. et al. The impact of micrornas on protein output. Nature 455, (2008) Selbach, M. et al. Widespread changes in protein synthesis induced by micrornas. Nature 455, (2008) Grimson, A. et al. MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol Cell 27, (2007) Krek, A. et al. Combinatorial microrna target predictions. Nat Genet 37, (2005) Wang, X. mirdb: a microrna target prediction and functional annotation database with a wiki interface. RNA 14, (2008) Betel, D., Wilson, M., Gabow, A., Marks, D.S. & Sander, C. The microrna.org resource: targets and expression. Nucleic Acids Res 36, D (2008) Maragkakis, M. et al. DIANA- microt web server: elucidating microrna functions through target prediction. Nucleic Acids Res 37, W (2009) Tan, L.P. et al. A high throughput experimental approach to identify mirna targets in human cells. Nucleic Acids Res 37, e137 (2009) Chi, S.W., Zang, J.B., Mele, A. & Darnell, R.B. Argonaute HITS- CLIP decodes microrna- mrna interaction maps. Nature 460, (2009) Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28, (2000) Kanehisa, M., Goto, S., Furumichi, M., Tanabe, M. & Hirakawa, M. KEGG for representation and analysis of molecular networks involving diseases and drugs. Nucleic Acids Res 38, D (2010) Subramanian, A. et al. Gene set enrichment analysis: a knowledge- based approach for interpreting genome- wide expression profiles. Proc Natl Acad Sci U S A 102, (2005) Dennis, G., Jr. et al. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol 4, P3 (2003) Zeeberg, B.R. et al. GoMiner: a resource for biological interpretation of genomic and proteomic data. Genome Biol 4, R28 (2003). 27
34 28 Research objectives Amplification of the MYCN oncogene delineates a subgroup of neuroblastoma patients with aggressive disease and poor survival. Insights in the pathways and processes that are deregulated upon MYCN amplification could therefore lead to the discovery of targets for selective therapy. While the study of MYCN regulated protein- coding genes has been heavily pursued, the role of non- coding mirnas in the MYCN transcriptional network is less understood. The first aim of this work was to identify and functionally characterize MYCN regulated mirnas in neuroblastoma. The identification of MYCN regulated mirnas requires an accurate quantification of mirna expression. To this end, a genome- wide RT- qpcr mirna profiling platform was introduced, optimized and validated (paper 1) and an accompanying data normalization strategy was developed (paper 2). These tools were applied to profile mirna expression in a large cohort of primary neuroblastoma tumour samples and cellular model systems with inducible MYCN expression. The different classes of MYCN regulated mirnas and their expression in neuroblastoma tumour subgroups are presented in paper 3. A set of 6 MYCN regulated mirnas, belonging to the mir mirna cluster, were selected for further functional characterization. The mir cluster has been implicated in the regulation of neuroblastoma cell proliferation and is one of the most frequently activated oncomirs in cancer. To explore the pathways downstream of mir , we created an inducible mir model system and applied high- throughput proteomics in order to capture both mrna degradation and translational inhibition events. The results of this study are presented in paper 4. The second aim of this work was to evaluate expression and function of another class of non- coding RNAs termed transcribed ultraconserved regions or T- UCRs. Apart from the fact that these sequences are at least 200 bp in length and 100% conserved between human, mouse and rat, nothing is known about their function. We developed a RT- qpcr based T- UCR expression platform capable of measuring all 481 T- UCRs and predicted T- UCR functions through integrative genomics. A functional T- UCR expression network for neuroblastoma is presented in paper 5. The bioinformatics tools that were developed for functional T- UCR annotation were subsequently applied to perform a genome- wide prediction of tissue- specific mirna functions. Current experimental approaches for functional characterization of mirnas are technically challenging and are typically performed for only one or few mirnas. Computational methods based on mirna target predictions are available, however these are prone to false positives and ignore the tissue specificity of mirna functions. We hypothesized that a combination of matching mrna and mirna expression profiles with mirna target predictions would result in a more accurate prediction of mirna functions. Predictions for different datasets were made available through a web interface at The results of this study are presented in paper 6. MiRNA expression signatures have proven to be powerful predictors of prognosis and diagnosis in a wide variety of cancer types. In contrast to mrna signatures, mirna signatures typically consist of few genes and outperform mrna signatures when classifying samples according to tissue or disease type. The final aim of this work was to build and evaluate a prognostic mirna signature for neuroblastoma. The mirna profiling platform and data normalization strategy described in papers 1 and 2 were used to analyze large cohorts of primary neuroblastoma tumours to build, test and validate a 25- mirna signature. For the first time, we were able to compare the performance of a prognostic mirna classifier with that of a prognostic mrna classifier that was established through earlier work in the lab. The results of this study are presented in paper 7.
35 29 Results PAPER 1 High- throughput stem- loop RT- qpcr mirna expression profiling using minute amounts of input RNA. PAPER 2 A novel and universal method for microrna RT- qpcr data normalization. PAPER 3 MYCN/c- MYC- induced micrornas repress coding gene networks associated with poor outcome in MYCN/c- MYC- activated tumours. PAPER 4 The mir MicroRNA Cluster Regulates Multiple Components of the TGF- β Pathway in Neuroblastoma. PAPER 5 An integrative genomics screen uncovers ncrna T- UCR functions in neuroblastoma tumours. PAPER 6 The microrna body map: dissecting microrna function through integrative genomics. PAPER 7 Outcome prediction of children with neuroblastoma using mirna and mrna gene expression signatures.
36 PAPER 1: High- throughput stem- loop RT- qpcr mirna expression profiling using minute amounts of input RNA 30 PAPER 1 High-throughput stem-loop RT-qPCR mirna expression profiling using minute amounts of input RNA. Mestdagh P, Feys T, Bernard N, Guenther S, Chen C, Speleman F, Vandesompele J. Nucleic Acids Res Dec;36(21):e143.
37 Nucleic Acids Research Advance Access published October 21, Nucleic Acids Research, 2008, 1 8 doi: /nar/gkn725 High-throughput stem-loop RT-qPCR mirna expression profiling using minute amounts of input RNA Pieter Mestdagh 1, Tom Feys 1, Nathalie Bernard 2, Simone Guenther 2, Caifu Chen 2, Frank Speleman 1 and Jo Vandesompele 1, * 1 Center for Medical Genetics, Ghent University Hospital, 9000 Ghent, Belgium and 2 Applied Biosystems, Foster City, CA, USA Received July 22, 2008; Revised September 3, 2008; Accepted October 1, 2008 ABSTRACT MicroRNAs (mirnas) are an emerging class of small non-coding RNAs implicated in a wide variety of cellular processes. Research in this field is accelerating, and the growing number of mirnas emphasizes the need for high-throughput and sensitive detection methods. Here we present the successful evaluation of the Megaplex reverse transcription format of the stem-loop primer-based real-time quantitative polymerase chain reaction (RT-qPCR) approach to quantify mirna expression. The Megaplex reaction provides simultaneous reverse transcription of 450 mature mirnas, ensuring high-throughput detection. Further, the introduction of a complementary DNA pre-amplification step significantly reduces the amount of input RNA needed, even down to single-cell level. To evaluate possible preamplification bias, we compared the expression of 384 mirnas in three different cancer cell lines with Megaplex RT, with or without an additional preamplification step. The normalized Cq values of all three sample pairs showed a good correlation with maintenance of differential mirna expression between the cell lines. Moreover, pre-amplification using 10 ng of input RNA enabled the detection of mirnas that were undetectable when using Megaplex alone with 400 ng of input RNA. The high specificity of RT-qPCR together with a superior sensitivity makes this approach the method of choice for high-throughput mirna expression profiling. far and many more awaiting experimental validation, these molecules represent one of the largest classes of gene regulators. Recent studies have implicated mirnas in numerous cellular processes including development, differentiation, proliferation, apoptosis and stress response and thus, not surprising, these same mirnas are turning out to be important players in cancer development (1). In addition to the dramatic impact on our insight into the fundamental aspects of oncogenesis, the discovery of mirnas has also potentially great implications for translational research as evidence is emerging that mirna signatures correlate with diagnosis, tumour classification and prognosis (2). In view of these observations, accurate high-throughput profiling of mirnas is a major challenge for the field. Various methods, such as microarrays and bead-based flow cytometry, are available enabling the detection of multiple mirnas in a single experiment, but such approaches generally require significant amounts of input RNA (>1 mg) and preclude the use of very small clinical biopsies or analysis of small subsets of cells or even single cells (2 7). Real-time quantitative PCR (RT-qPCR) has superior sensitivity, down to the single molecule level. The stemloop reverse transcription primer method developed by Chen et al. (8) has enabled specific and sensitive PCR-based quantification of small RNA molecules such as mirnas. This method selectively targets mature mirnas and easily covers a dynamic range of linear quantification of 7 log10 units. Here we present the evaluation of the Megaplex reverse transcription format of the stem-loop primer-based RT-qPCR approach to quantify mirna expression. Downloaded from nar.oxfordjournals.org at Biomedische Bibliotheek on December 28, 2010 INTRODUCTION MicroRNAs (mirnas) are an emerging class of small non-coding RNAs capable of negatively regulating gene expression. With over 800 human mirnas reported thus MATERIALS AND METHODS Cell culture and RNA samples Eleven neuroblastoma (NB) cell lines (NGP, IMR-32, SMS-KAN, SK-N-BE(2c), LAN-5, SK-MYC2, *To whom correspondence should be addressed. Tel: ; Fax: ; [email protected] ß 2008 The Author(s) This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License ( by-nc/2.0/uk/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
38 32 2 Nucleic Acids Research, 2008 SK-N-AS, SK-N-SH, NBL-S, SK-N-FI and CLB-GA) were cultured in RPMI 1640 medium (Invitrogen) supplied with 15% fetal calf serum, 1% penicillin/streptomycin, 1% kanamycin, 1% glutamine, 2% HEPES (1 M), 1% sodiumpyruvate (100 nm) and 0.1% beta-mercapto (50 nm). At 80% confluence, cells were harvested by scraping for total RNA isolation (mirneasy, Qiagen) or trypsinized for cytospin preparation. Human colon and brain RNA samples were obtained from Stratagene. Preparation of cells on membrane-coated slides Cell suspensions were prepared for microdissection by centrifuging the cells for 10 min at 750 g. Pellets were washed twice with phosphate-buffered saline and resuspended to obtain cells/ml. In all, 200 ml of the cell suspension was then transferred by centrifugation (120 g for 3 min) onto slides covered with polyethylene naphthalate membrane (PALM Microlaser Technologies, Bernried, Germany). Isolation of cells by laser microdissection and pressure catapulting Single cells were isolated using the PALM MicroBeam system (PALM Microlaser Technologies) as described previously (9) and collected in a 200 ml Eppendorf tube cap containing 3.94 ml RT master mix as detailed below. Following collection of cells, all tubes were centrifuged for 1 min at g. Cells were lysed by heating at 958C for 5 min, after which the entire lysate was used in a Megaplex RT reaction followed by pre-amplification of the mirna complementary DNA (cdna) (see further). mirna reverse transcription For mirna cdna synthesis, RNA was reverse transcribed using the mirna reverse transcription kit (Applied Biosystems) in combination with the stem-loop Megaplex primer pool (Applied Biosystems), allowing simultaneous reverse transcription of 450 mirnas and endogenous controls. Briefly, 8 ml of total RNA (50 ng/ml) was supplemented with RT primer mix (10), RT buffer (10), MultiScribe Reverse Transcriptase (10 U/ml), dntps with dttp (0.5 mm each), MgCl 2 (3 mm) and AB RNase inhibitor (0.25 U/ml) in a total reaction volume of 80 ml. For RT reactions with subsequent pre-amplification, the reaction volume is proportionally reduced to 5 ml. Concentration of each stem-loop primer in the RT reaction mix was 1 nm, a 50-fold dilution compared with a singleplex RT reaction, ensuring minimal non-specific interactions between the different stem-loop primers. To increase reverse transcription efficiency, a pulsed RT reaction was used (40 cycles of 168C for 2 min, 428C for 1 min and 508C for 1 s, followed by a final reverse transcriptase inactivation at 858C for 5 min). Pre-amplification of cdna Megaplex RT product (5 ml) was pre-amplified using Applied Biosystems TaqMan PreAmp Master Mix (2) and PreAmp Primer Mix (5) in a 25-ml PCR reaction. The primer pool consisted of forward primers (50 nm) specific for each of the 450 mirnas and a universal reverse primer (50 nm) (Applied Biosystems, early access). The pre-amplification cycling conditions were as follows: 958C for 10 min, 558C for 2 min and 758C for 2 min followed by 14 cycles of 958C for 15 s and 608C for 4 min. Real-time qpcr For each cdna sample, 384 small RNAs were profiled using a gene maximization PCR plate setup in a 384-well plate. As instrument and liquid handling variations were shown to be minimal, no PCR replicates were measured. This approach allowed us to profile one sample per 384-well plate. Without pre-amplification, RT product was diluted 400-fold; when pre-amplification was applied, the dilution factor was PCR amplification reactions were carried out in a total volume of 8 ml, containing 4 ml of TaqMan Master Mix (Applied Biosystems), 1 ml of cdna and 3 ml of mirna TaqMan probe and primers (Applied Biosystems). Cycling conditions were as follows: 958C for 10 min followed by 40 cycles of 958C for 15 s and 608C for 1 min. All PCR reactions were performed on the 7900HT RT-qPCR system (Applied Biosystems). Raw Cq values were calculated using the SDS software v.2.1 using automatic baseline settings and a threshold of 0.2. The crossing point between the baseline corrected amplification curve and threshold line is called the quantification cycle (Cq) (according to RDML guidelines, rdml.org) (10). Assessment of pre-amplification bias Potential pre-amplification bias was addressed by analyzing the preservation of differential expression. Briefly, differential mirna expression between different sample pairs (Cq) was determined for each approach (with or without pre-amplification). Subsequently, the difference in differential mirna expression (Cq) was calculated. Minimal bias should result in low Cq values. LNA microarrays In total, 5 mg of total RNA was hybridized to immobilized locked nucleic acid (LNA)-modified capture probes according to Castoldi et al. (11). Background- and flagcorrected median intensities were log transformed and normalized according to the average signal of each array. RESULTS Minimal pre-amplification bias and sensitivity A major concern for introducing a pre-amplification step is the possibility that the relative mirna expression levels in the original cell population are not maintained. In order to obtain an unbiased pre-amplification of the mirna cdna, equal amplification efficiency and a high degree of amplification specificity is required. Previous studies already reported the use of pre-amplification in combination with the stem-loop procedure, but no study thus far evaluated the effects of the pre-amplification step on the fidelity of mirna expression measurement (12,13). Here we performed an in-depth analysis of the potential bias Downloaded from nar.oxfordjournals.org at Biomedische Bibliotheek on December 28, 2010
39 33 Nucleic Acids Research, of a pre-amplification step through direct comparison of mirna expression profiles obtained with and without pre-amplification. To this purpose, we profiled 384 mirnas in three different NB cancer cell lines using the Megaplex reverse transcription either with or without preamplification of the mirna cdna. Because each sample was profiled in a separate PCR run, raw Cq values were corrected for inter-run variation. Calibration and normalization of raw Cq values was done by equalizing the average Cq value across all detectable mirnas between the different sample runs. For further data analysis, only those mirnas with a Cq value equal to or below 35 were taken into account. It is generally accepted that a Cq value of 35 represents single molecule template detection; Cq values above 35 are, therefore, considered noise. Low-copy template detection is also subjected to a higher degree of variability, mainly due to Poisson distribution sampling effects. For Megaplex RT without pre-amplification, 400 ng of total RNA was used, whereas for pre-amplified samples, 1 and 10 ng of total RNA were used. Calibrated Cq values obtained with pre-amplification (Cq P ) were plotted against those obtained without pre-amplification (Cq NP ) (Figure 1). MiRNAs that were either undetectable or had a Cq value above 35 were assigned a Cq of 40. The Cq Cq plots for the NGP cells reveal a suboptimal correlation between both data sets (Megaplex RT alone versus Megaplex with pre-amplification), especially for those mirnas with a high Cq value. Nevertheless, the slope of the linear trendline fitted along the correlation plot nearly equals 1, indicating unbiased and efficient pre-amplification efficiency irrespective of mirna expression level. Therefore, mirna quantification using the preamplification procedure should result in relative mirna expression levels that represent the actual situation in the cell population. Interestingly, a higher number of mirnas were detected with the pre-amplification procedure, despite the fact that 40-times less input RNA was used (Figure 1A). Assuming 100% amplification efficiency, a 14-cycle preamplification of 10 ng total RNA would result in 200 ng of mirna cdna (total RNA equivalents) in the final PCR reaction. When no pre-amplification is applied, each RT-qPCR assay contains only 1 ng of mirna cdna (total RNA equivalents). The predicted 200-fold difference in input with or without a pre-amplification step clearly contributes to the superior sensitivity of the pre-amplification approach. When the amount of input RNA is further reduced from 10 to 1 ng, the increased sensitivity is less pronounced as some of the mirnas become undetectable again (Figure 1B). When using 1 ng of input, both Poisson distribution sampling effects and reduced RT efficiency for low copy numbers come into play. Similar results were obtained with the other cell lines (data not shown). Reverse transcription variability for low-copy RNA molecules To further investigate the suboptimal correlation in the Cq Cq plot from a sample analyzed with or without pre-amplification, we characterized the variation introduced by the RT reaction. By performing two independent RT reactions on the same RNA sample, we evaluated the variation induced by the RT step. The Cq Cq plot for the two RT reactions indeed shows that a certain degree of variation is introduced by the RT reaction (R 2 = 0.944) (Figure 2). A similar experiment with the two other cell lines resulted in a correlation coefficient of and To determine which mirnas display the highest degree of variation following two subsequent RT reactions, we divided the data set into three subsets according Figure 2. Cq Cq correlation plot for two independent reverse transcription reactions without pre-amplification. Downloaded from nar.oxfordjournals.org at Biomedische Bibliotheek on December 28, 2010 Figure 1. Correlation plot between Cq values obtained through Megaplex RT alone (Cq NP ) and Megaplex RT with pre-amplification (Cq P ) using 10 ng of total RNA (A) and 1 ng of total RNA (B) from NGP neuroblastoma cells. (A) Data points highlighted in red indicate mirnas that are only detectable if pre-amplification is applied. (B) Data points highlighted in red indicate mirnas that are only detectable if no preamplification is applied.
40 34 4 Nucleic Acids Research, 2008 to Cq value: highly expressed mirnas (Cq below 25), moderately expressed mirnas (Cq between 25 and 30) and low abundant mirnas (Cq above 30). Individual correlation coefficients for each subset suggest a much higher RT variation for the low abundant mirnas (R 2 = 0.685) compared to the moderately (R 2 = 0.806) and highly (R 2 = 0.872) expressed mirnas. This partly explains the observed variation for the low abundant mirnas when comparing the data sets obtained with and without pre-amplification (Figure 1) as both were generated with a different RT reaction. Another source of variation could be liquid handling and instrumentation. However, when performing repeated RT-qPCR runs for the same sample, almost no variation was observed (R 2 = 0.983). To further evaluate and compare RT variation for the procedures with and without pre-amplification, we determined the variability of individual mirnas. Triplicate RNA samples from human brain and human colon were reverse transcribed using the method with pre-amplification and the method without pre-amplification. In case of pre-amplification, 10 ng of total RNA was used whereas 400 ng of RNA was used when no pre-amplification was applied. Sixteen individual mirnas were profiled in triplicate RT-qPCR experiments for each RT sample. Raw Cq values were transformed according to 2 Cq and averaged across the triplicate RT-qPCR experiments for each RT sample. Per mirna, the averaged expression values were used to calculate the coefficient of variation (CV) across the triplicate RT samples. CV values for each RNA sample (human colon and human brain) profiled with each of the two methods (Megaplex with and without pre-amplification) were then plotted in function of the average Cq value for the sample under consideration (Supplementary Figure S1). Results clearly indicate low CV values for high and moderately expressed mirnas (average Cq < 30) in both colon and brain samples, independently of the quantification method used. For the low abundant mirnas (Cq > 30) CV values drastically increase (up to 50%) in colon and brain samples, again independently of the quantification method. This confirms the increased RT variation for low abundant mirnas. Preservation of differential expression To assess the potential bias introduced through a preamplification step, differential mirna expression levels (Cq) between the three different cell lines were determined for each procedure, i.e. with (Cq P ) or without pre-amplification (Cq NP ). The difference in differential mirna expression between two cell lines, measured both with and without pre-amplification (calculated by the Cq = Cq NP Cq P ), was then plotted against the average Cq value of the mirna in the two cell lines under consideration, as measured by the procedure with no pre-amplification step (Figure 3A). In all, 80% of all mirnas with a Cq value <35 display a Cq < 1. Of these mirnas, 75% have a Cq < 0.5. The plot clearly indicates that the Cq value increases for low abundant mirnas. While for the most abundant mirnas (average Cq value between 15 and 20) there is a near-perfect correlation in differential mirna expression (average Cq = 0.35), the lowest expressed mirnas (average Cq value between 30 and 35) display an average Cq value of 1.3 (Figure 3A). To some degree, this intensitydependent variation is attributable to the fact that each data set was generated using a different RT reaction as low abundant mirnas are more susceptible to variation during reverse transcription (see higher). By lowering the detection Cq cutoff from 35 to 30, thereby excluding the lowest expressed mirnas, 94% of all mirnas display a Cq < 1 (Figure 3B). Similar results were obtained in other cell types (Supplementary Figure S2). Interestingly, over half the mirnas with a Cq > 1 were differentially expressed (Cq > 1), irrespective of the quantification method (with or without pre-amplification). For example, mir-299-5p has a Cq value > 1, meaning a more than 2-fold bias in differential expression when comparing two samples with and without pre-amplification. However, the individual Cq values indicate that this mirna shows a highly differential expression in the two cell lines (Cq P = 6.5, Cq NP = 8.3). Large-scale screening studies often apply a Cq value of 1 (2-fold difference) to select differentially expressed mirnas so these mirnas would definitely be selected, regardless of the use of pre-amplification or not. To assess whether the high degree of variation observed for the low abundant mirnas could also be due to mirna sequence characteristics, six different sequence parameters were evaluated for each mature mirna (number of A, U, C and G bases, GC percentage and sequence length). For each individual mirna, the average Cq value was used as a measure of variation. All 384 mirnas analyzed were divided into four subgroups according to the average Cq value (Group 1, Cq < 0.5; Group 2, 0.5 < Cq < 1; Group 3, 1 < Cq < 2; Group 4, Cq > 2). The number of A bases was the only parameter that significantly differed between the different subgroups (Kruskal Wallis, P < 0.05). Paired analysis of all subgroups revealed that the number of A bases is significantly higher for mirnas belonging to Group 4 compared to Groups 1 and 2 (Mann Whitney, P = and P = 0.002, respectively). In general, mirnas with a Cq value <1 have a higher number of A bases compared to mirnas with a Cq value >1 (Mann Whitney, P = 0.006). To assess whether these A bases appear randomly across the mirna sequence or in stretches of consecutive bases, the number of mirnas containing one or more stretches of at least three A bases was determined for each subgroup. Interestingly, mirnas with a Cq value <1 were more likely to contain one or more stretches of at least three A bases as compared to mirnas with a Cq value >1 (Fisher exact, P = ). It remains to be determined how exactly the number of A bases influences the Cq value. Confirmation of MYCN-regulated mirnas To further evaluate the performance of the Megaplex and pre-amplification method outlined above, we set up a mirna profiling screen to identify mirnas that are Downloaded from nar.oxfordjournals.org at Biomedische Bibliotheek on December 28, 2010
41 35 Nucleic Acids Research, Figure 3. Difference in differential mirna expression (Cq) between two neuroblastoma cell lines (NBL-S and IMR-32) as measured using Megaplex RT alone (Cq NP ) and Megaplex RT with pre-amplification (Cq P ). Cq values are plotted in function of the average expression of each mirna in the two cell lines as quantified by Megaplex RT alone (average Cq NP ). Bar plots indicate the mean Cq value for mirnas with an average Cq NP value ranging between 15 20, 20 25, and (A) Results when a Cq detection cutoff of 35 is applied. (B) Results when Cq detection cutoff is lowered to 30. regulated by the oncogenic MYCN transcription factor, known to play a role in an aggressive type of childhood NB. Recently, a subset of mirnas regulated by MYCN has been identified (14). Among these mirnas were four members of the mir cluster on chromosome 13 that were identified in at least one of three studied model systems (mir-17-5p, mir-18, mir-20a and mir-92). The mir cluster contains a total of seven different mirnas (mir-17-5p, mir-17-3p, mir-18a, mir-19a, mir-20a, mir-19b and mir-92) and has recently been shown to be a direct target of MYC, the best studied and founding member of the MYC family of bhlh/lz transcription factors (15). To see whether we could confirm previously published results on MYCN-regulated mirnas, we profiled all seven mirnas from the mir cluster in a panel of 11 NB cell lines using the Megaplex reverse transcription with pre-amplification starting with 20 ng of input RNA. The panel of cell lines consisted of five MYCN-amplified cell lines, one cell line with stable overexpression of MYCN and five cell lines with normal MYCN copy number. If the mir cluster is activated by MYCN, these mirnas should display a higher expression in the cell lines with MYCN amplification and overexpression. Five mirnas displayed a significant differential expression between both groups of cell lines (mir-17-3p, mir-19a, mir-19b, mir-20a and mir-92) (Mann Whitney test, P < 0.05) (Supplementary Figure S3). For mir-17-5p and mir-18a, there was a trend for higher expression in MYCN-activated cells (P = 0.052). We also quantified the expression of the mir cluster in the same 11 cell lines without the pre-amplification step. Mann Whitney analysis again revealed four differentially expressed mirnas (mir-17-5p, mir-19b, mir-20a and mir-92). Apparently, both the Megaplex RT with and without pre-amplification are capable of identifying the majority of the mirnas in the cluster as being differentially expressed with respect to the MYCN status. Moreover, the introduction of the pre-amplification step slightly increases the sensitivity of the detection, identifying significant differential expression for five mirnas and nearly significant differential expression for the remaining two mirnas. The entire mir cluster is transcribed into a single primary mirna transcript (pri-mir-17-92). If this cluster is activated by MYCN activity, most likely all mirnas from the cluster should be activated. The results obtained with pre-amplification most closely resemble this situation. To further validate our PCR-based approach, we also compared our PCR-based results with an independent mirna expression profiling study using a microarray platform (3,11). Again, the same subset of cell lines was used, and mirnas with a significantly differential expression were selected with the Mann Whitney test (P < 0.05). Three mirnas from the mir cluster displayed a significant differential expression (mir-17-3p, mir-19b and mir-92). For mir-19a, the P-value was very close to being significant (P = 0.052). Downloaded from nar.oxfordjournals.org at Biomedische Bibliotheek on December 28, 2010
42 36 6 Nucleic Acids Research, 2008 Figure 4. (A) Standard curve analysis depicting perfect linearity and correlation between input cell number (1 128) and measured Cq value for three different mirnas (mir-18a, mir-20b and mir-106a). (B) Standard curve analysis depicting perfect linearity and correlation between total RNA input ( pg) and measured Cq value for three different mirnas (mir-92, mir-19a and mir-20a). Total input RNA values on X-axis are log 2 based. Profiling of single cells Finally we investigated whether our method would be suited for single-cell analysis. Single-cell RNA levels are cell type dependent and vary between 10 and 30 pg of total RNA. We, therefore, serially diluted human brain total RNA to obtain a dilution series ranging from 6250 to 2 pg of total RNA. All samples from the dilution series underwent Megaplex reverse transcription and pre-amplification prior to RT-qPCR. Expression of both high and low abundant mirnas was determined, and Cq values were plotted in function of total input RNA (Figure 4B). These plots display perfect linearity, with slopes equaling the theoretical slope of 1 and suggest that the method is suited to profile single cells. However, when analyzing a single cell or few cells, it is desirable that the expression profiles can be readily obtained from total cell lysate. This way, RNA yield is unaffected as no downstream manipulations such as RNA purification or deoxyribonuclease (DNase) treatment is required. The stem-loop reverse transcription primers selectively target mature mirnas thus ruling out the necessity of a DNase treatment. Second, it is important to verify that the method works for individually isolated cells. To test the possibility of obtaining accurate mirna expression data from total cell lysate of individual cells, we prepared cytospins of NGP cells on membrane-coated slides. Individual cells were microdissected to obtain a dilution series of 1, 2, 4, 8, 16, 32, 64 or 128 cells that were lysed by heating, followed by Megaplex reverse transcription, pre-amplification and subsequent RT-qPCR. The expression level of three mirnas, mir-18a, mir-20b and Downloaded from nar.oxfordjournals.org at Biomedische Bibliotheek on December 28, 2010
43 37 Nucleic Acids Research, mir-106a, was determined for each point in the dilution series and Cq values were plotted in function of cell numbers (Figure 4A). The resulting dilution series presents with perfect linearity with a slope of 0.856, 1.07 and 0.935, respectively, approaching the theoretical slope of 1. DISCUSSION In this study, we have evaluated the sensitivity and reliability of a pre-amplification step for high-throughput stem-loop primer-based RT-qPCR measurement of mirna expression. Our result convincingly demonstrate that such quantification is feasible starting from minute amounts of RNA (10 ng) or even single-cell RNA, thus opening the way for profiling the mirnaome from small cell populations or individual cells. It is important, however, to address the potential bias of the pre-amplification reaction. By evaluating the preservation of differential mirna expression through comparison of sample pairs with and without pre-amplification, we used a relevant approach to evaluate possible pre-amplification bias. In contrast to previous studies on the use of a pre-amplification step in a stem-loop PCR reaction setup, we have made a direct comparison between results generated with and without pre-amplification and also analyzed the effect on low abundant mirnas. Further, our data set contains expression data for almost 400 mirnas, allowing for a more robust analysis. For the most abundant mirnas, the difference in differential expression, as measured by Megaplex alone and Megaplex followed by pre-amplification, did not exceed 0.8 PCR cycles for 90% of the genes. For the low abundant mirnas the differential expression difference exceeded 1 more frequently. We identified the RT reaction itself as a major factor contributing to the observed variation. RT-induced variation was particularly high for the low abundant mirnas, confirming previous studies on mrna templates (16). In these analyses, variation in RT efficiency plays a major role as paired sample comparison required four independent RT reactions. Most likely, a certain degree of variation will be attributable to the pre-amplification reaction itself. By further expanding the stem-loop primer pool, we not only cover the majority of all human mirnas but also avoid the need for multiple separate RT reactions per sample. Our analysis reveals that expression data for low abundant mirnas obtained through Megaplex reverse transcription followed by pre-amplification should be interpreted with caution. In view of the variability in reverse transcription combined with Poisson distribution effects, results should be confirmed in independent experiments or biological replicates. Important to note, however, is that the direction of differential gene expression is almost always preserved. We demonstrated the sensitivity of our approach by confirming (and extending) differential expression of the mir cluster. For five out of seven mirnas residing within this cluster we could confirm significant MYCNdependent expression while MYCN-dependent expression for the other two was borderline significant. MYCNdependent expression was already reported for four mirnas within the cluster. Furthermore, this cluster is directly activated by MYC, a transcription factor with high homology to MYCN. Finally, when repeating the same analysis using LNA-based microarray technology, three of seven mirnas displayed significant MYCNdependent expression, confirming literature and stemloop RT-qPCR profiling results obtained in this study. The above results underscore the high sensitivity and accuracy of Megaplex reverse transcription followed by limited-cycle pre-amplification. The fact that more mirnas are detected when applying the pre-amplification procedure further illustrates its impact on detection sensitivity. To address the possibility of single-cell mirna expression analysis, we evaluated preservation of detection linearity in a 2-fold dilution series ranging from 1 to 128 individually isolated cells. For the mirnas analyzed, detection linearity was maintained down to the singlecell level. Moreover, mirna profiling could be performed directly on total cell lysate avoiding the need for sample manipulations that affect the RNA yield. Tang and colleagues (13) have shown that using a 220-plex stem-loop primer pool, accurate mirna expression profiling from single handpicked cells is possible. Here we show that an increased complexity of the stem-loop primer pool still allows for highly accurate mirna quantification on whole-cell lysate from individually picked cells. Important to note, however, is the discrepancy between the expression level in a single cell and the average expression in a population of cells belonging to the same cell type, due to the lognormal distribution of gene expression (17). This can explain the slightly larger variation that was observed for the single-cell dilution series compared to the RNA dilution series. Fundamental biological questions on early tumour development or stem cell differentiation are best tested at the single-cell level. Often, specific cells are microdissected from a heterogeneous population, thereby ensuring that the target population is free of contamination with non-target cells (18,19). Also, studies on embryonic development are typically performed on the single-cell level (20). We strongly believe that the method presented here will aid in the unraveling of mirna function in single-cell studies or pure cell populations isolated by microdissection, flow sorting or bead-based selection. In this study, we have shown that the Megaplex stem-loop PCR procedure in combination with limitedcycle pre-amplification is a powerful method for mirna expression profiling in both large and small cell populations capable of covering the majority of human mirnas. SUPPLEMENTARY DATA Supplementary Data are available at NAR Online. FUNDING Scientific Research (FWO) Flanders (FWO postdoc to J.V.); Kinderkankerfonds (a non-profit childhood cancer Downloaded from nar.oxfordjournals.org at Biomedische Bibliotheek on December 28, 2010
44 38 8 Nucleic Acids Research, 2008 foundation under Belgian law); Ghent University Research Fund (BOF; 01D31406 to P.M.). Funding for open access charge: EU STREP EET-pipeline n Conflict of interest statement. None declared. REFERENCES 1. Esquela-Kerscher,A. and Slack,F.J. (2006) Oncomirs micrornas with a role in cancer. Nat. Rev. Cancer, 6, Lu,J., Getz,G., Miska,E.A., Alvarez-Saavedra,E., Lamb,J., Peck,D., Sweet-Cordero,A., Ebert,B.L., Mak,R.H., Ferrando,A.A. et al. (2005) MicroRNA expression profiles classify human cancers. Nature, 435, Castoldi,M., Schmidt,S., Benes,V., Noerholm,M., Kulozik,A.E., Hentze,M.W. and Muckenthaler,M.U. (2006) A sensitive array for microrna expression profiling (michip) based on locked nucleic acids (LNA). RNA, 12, Liu,C.G., Calin,G.A., Meloon,B., Gamliel,N., Sevignani,C., Ferracin,M., Dumitru,C.D., Shimizu,M., Zupo,S., Dono,M. et al. (2004) An oligonucleotide microchip for genome-wide microrna profiling in human and mouse tissues. Proc. Natl Acad. Sci. USA, 101, Nelson,P.T., Baldwin,D.A., Scearce,L.M., Oberholtzer,J.C., Tobias,J.W. and Mourelatos,Z. (2004) Microarray-based, highthroughput gene expression profiling of micrornas. Nat. Methods, 1, Sioud,M. and Rosok,O. (2004) Profiling microrna expression using sensitive cdna probes and filter arrays. Biotechniques, 37, , Thomson,J.M., Parker,J., Perou,C.M. and Hammond,S.M. (2004) A custom microarray platform for analysis of microrna gene expression. Nat. Methods, 1, Chen,C., Ridzon,D.A., Broomer,A.J., Zhou,Z., Lee,D.H., Nguyen,J.T., Barbisin,M., Xu,N.L., Mahuvakar,V.R., Andersen,M.R. et al. (2005) Real-time quantification of micrornas by stem-loop RT-PCR. Nucleic Acids Res., 33, e Schutze,K. and Lahr,G. (1998) Identification of expressed genes by laser-mediated manipulation of single cells. Nat. Biotechnol., 16, Taylor,C.F., Field,D., Sansone,S.A., Aerts,J., Apweiler,R., Ashburner,M., Ball,C.A., Binz,P.A., Bogue,M., Booth,T. et al. (2008) Promoting coherent minimum reporting guidelines for biological and biomedical investigations: the MIBBI project. Nat. Biotechnol., 26, Castoldi,M., Schmidt,S., Benes,V., Hentze,M.W. and Muckenthaler,M.U. (2008) michip: an array-based method for microrna expression profiling using locked nucleic acid capture probes. Nat. Protoc., 3, Cogswell,J.P., Ward,J., Taylor,I.A., Waters,M., Shi,Y., Cannon,B., Kelnar,K., Kemppainen,J., Brown,D., Chen,C. et al. (2008) Identification of mirna changes in Alzheimer s disease brain and CSF yields putative biomarkers and insights into disease pathways. J. Alzheimers Dis., 14, Tang,F., Hajkova,P., Barton,S.C., Lao,K. and Surani,M.A. (2006) MicroRNA expression profiling of single whole embryonic stem cells. Nucleic Acids Res., 34, e Schulte,J.H., Horn,S., Otto,T., Samans,B., Heukamp,L.C., Eilers,U.C., Krause,M., Astrahantseff,K., Klein-Hitpass,L., Buettner,R. et al. (2008) MYCN regulates oncogenic MicroRNAs in neuroblastoma. Int. J. Cancer, 122, O Donnell,K.A., Wentzel,E.A., Zeller,K.I., Dang,C.V. and Mendell,J.T. (2005) c-myc-regulated micrornas modulate E2F1 expression. Nature, 435, Stahlberg,A., Hakansson,J., Xian,X., Semb,H. and Kubista,M. (2004) Properties of the reverse transcription reaction in mrna quantification. Clin. Chem., 50, Bengtsson,M., Stahlberg,A., Rorsman,P. and Kubista,M. (2005) Gene expression profiling in single cells from the pancreatic islets of Langerhans reveals lognormal distribution of mrna levels. Genome Res., 15, Shin,M.S., Kim,H.S., Kang,C.S., Park,W.S., Kim,S.Y., Lee,S.N., Lee,J.H., Park,J.Y., Jang,J.J., Kim,C.W. et al. (2002) Inactivating mutations of CASP10 gene in non-hodgkin lymphomas. Blood, 99, De Preter,K., Vandesompele,J., Heimann,P., Yigit,N., Beckman,S., Schramm,A., Eggert,A., Stallings,R.L., Benoit,Y., Renard,M. et al. (2006) Human fetal neuroblast and neuroblastoma transcriptome analysis confirms neuroblast origin and highlights neuroblastoma candidate genes. Genome Biol., 7, R Saitou,M., Barton,S.C. and Surani,M.A. (2002) A molecular programme for the specification of germ cell fate in mice. Nature, 418, Downloaded from nar.oxfordjournals.org at Biomedische Bibliotheek on December 28, 2010
45 Supplemental Data 39 Supplementary Figure 1 Scatter plot showing the relation between the average Cq- value and the reverse transcription variation (CV) for 16 individual mirnas in 2 different RNA samples (human brain and human colon), profiled using both Megaplex alone and Megaplex with pre- amplification. Brain samples are depicted in red, colon samples in blue. Triangles represent mirnas profiled using the Megaplex RT with pre- amplification whereas circles represent mirnas profiled using Megaplex RT alone.
46 40 Supplementary Figure 2 Difference in differential mirna expression ( Cq) between human colon and human brain as measured using Megaplex RT alone ( Cq NP ) and Megaplex RT with pre- amplification ( Cq P ). Cq- values for 16 individual mirnas are plotted in function of the average expression of each mirna in the 2 samples as quantified by Megaplex RT alone (average Cq NP ).
47 41 Supplementary Figure 3 Boxplots representing the relative expression of mirnas belonging to the mir cluster in 2 groups of neuroblastoma cell lines: MYCN single copy cell lines (MNSC) and MYCN amplified cell lines (MNA). For each mirna, Mann- Whitney p- values are indicated. MiRNA expression values were scaled relative to the cell line with the lowest expression.
48 PAPER 2: A novel and universal method for microrna RT- qpcr data normalization 42 PAPER 2 A novel and universal method for microrna RT-qPCR data normalization. Mestdagh P, Van Vlierberghe P, De Weer A, Muth D, Westermann F, Speleman F, Vandesompele J. Genome Biol. 2009;10(6):R64.
49 Mestdagh Volume al. 10, Issue 6, Article R64 Open Access Method A novel and universal method for microrna RT-qPCR data normalization Pieter Mestdagh *, Pieter Van Vlierberghe *, An De Weer *, Daniel Muth, Frank Westermann, Frank Speleman * and Jo Vandesompele * Addresses: * Center for Medical Genetics, Ghent University Hospital, De Pintelaan 185, Ghent, Belgium. Department of Tumour Genetics, German Cancer Center, Im Neuenheimer Feld 280, Heidelberg, Germany. Correspondence: Jo Vandesompele. [email protected] Published: 16 June 2009 Genome Biology 2009, 10:R64 (doi: /gb r64) The electronic version of this article is the complete one and can be found online at Received: 2 April 2009 Revised: 2 April 2009 Accepted: 16 June Mestdagh et al.; licensee BioMed Central Ltd. This is an open access article distributed under the terms of the Creative Commons Attribution License ( which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Normalization <p>the iments.</p> mean expression of microrna value: RT-qPCR a new method for accurate and reliable normalization of microrna expression data from RT-qPCR exper- Abstract Gene expression analysis of microrna molecules is becoming increasingly important. In this study we assess the use of the mean expression value of all expressed micrornas in a given sample as a normalization factor for microrna real-time quantitative PCR data and compare its performance to the currently adopted approach. We demonstrate that the mean expression value outperforms the current normalization strategy in terms of better reduction of technical variation and more accurate appreciation of biological changes. Background MicroRNAs (mirnas) are an important class of gene regulators, acting on several aspects of cellular function such as differentiation, cell cycle control and stemness. Not surprisingly, deregulated mirna expression has been implicated in a wide variety of diseases, including cancer [1]. Moreover, mirna expression profiling of different tumor entities resulted in the identification of mirna signatures correlating with patient diagnosis, prognosis and response to treatment [2]. Despite the small size of mirna molecules, several technologies have been developed that enable high-throughput and sensitive mirna profiling, such as microarrays [3-8], real-time quantitative PCR (RT-qPCR) [9,10] and bead-based flow cytometry [2]. In terms of accuracy and specificity, RTqPCR has become the method of choice for measuring gene expression levels, both for coding and non-coding RNAs. However, the accuracy of the results is largely dependent on proper data normalization. As numerous variables inherent to an RT-qPCR experiment need to be controlled for in order to differentiate experimentally induced variation from true biological changes, the use of multiple reference genes is generally accepted as the gold standard for RT-qPCR data normalization [11]. Typically, a set of candidate reference genes is evaluated in a pilot experiment with representative samples from the experimental condition(s). Ideally these candidate reference genes belong to different functional classes, significantly reducing the possibility of confounding co-regulation. In case of mirna profiling, only few candidate reference mirnas have been reported [12]. Generally, other small noncoding RNAs are used for normalization. These include both small nuclear RNAs (for example, U6) and small nucleolar RNAs (for example, U24, U26). Strategies for normalization of high-dimensional expression profiling experiments (using, for example, microarray technology, but recently also transcriptome sequencing) generally take advantage of the huge amount of data generated and often use (almost) all available data points. These strategies range from a straightforward approach based on the mean or median expression value to more complex algorithms such as Genome Biology 2009, 10:R64
50 44 Genome Biology 2009, Volume 10, Issue 6, Article R64 Mestdagh et al. R64.2 lowess normalization, quantile normalization or rank invariant normalization [13]. In this study we successfully introduce the mean expression value in a given sample to normalize high-throughput mirna RT-qPCR data and compare its performance to the currently adopted approach based on small nuclear/nucleolar RNAs. In addition, we provide a workflow for proper data normalization of both large scale (whole mirnome) and small scale mirna profiling experiments. Expression stability 1.8 1, , , , , , , , Results Stability of the mean mirna expression To evaluate the suitability of the mean mirna expression value as a normalization factor, we profiled 448 mirnas and controls in a subset of 61 neuroblastoma (NB) tumor samples and 384 mirnas and controls in 49 T-cell acute lymphoblastic leukemia (T-ALL) samples, 18 leukemias with EVI1 overexpression, 8 normal human tissues and 11 normal bone marrow samples using a high throughput mirna profiling platform based on Megaplex stem-loop RT-qPCR technology in combination with a limited cycle pre-amplification [9,10]. For each of the above mentioned sample sets all 18 available small RNA controls were quantified. For each individual sample, the mean expression value was calculated based on those mirnas that were expressed according to a Cq detection cutoff of 35 PCR cycles [10] (Cq, or quantification cycle, is the standard name for the Ct or Cp value according to Real-time PCR Data Markup Language (RDML) guidelines [14]). Expression stability of the mean expression value, the small RNA controls and a selection of three mirnas (mir-17-5p, mir-191 and mir-103) previously proposed as universal reference mirnas was then assessed for each sample set using the genorm algorithm [11]. To reduce the risk of including genes that are putatively co-regulated, a number of small RNA controls residing within the same gene cluster were discarded, retaining only one representative small RNA control per cluster. This was the case for RNU44, U47 and U75 on 1q25, and RNU58A and RNU58B on 18q21, of which RNU44 and RNU58A were randomly retained for further analysis. Naturally, only those small RNA controls that are expressed in all samples within a sample set were evaluated for their expression stability. genorm analysis clearly shows that the mean expression value is a suitable normalization factor in the different tissue groups under investigation. In terms of expression stability, the mean expression value is top ranked in the T-ALL samples, the NB samples, the normal human tissues and the normal bone marrow samples when compared to 16, 17, 14 and 18 candidate small RNA controls/mirnas, respectively (Figure 1 and Additional data file 1). For the leukemia samples with EVI1 overexpression the mean expression value ranked second (compared to 17 small RNA controls/mirnas; Additional data file 1). Several of the high ranking small RNA controls are the same ones proposed by the manufacturer as genorm Figure 1expression stability plot genorm expression stability plot. Expression stability of 13 different small RNA controls and the mean expression value in the T-cell acute lymphoblastic leukaemia sample set. The mean expression value shows the highest expression stability across all 49 samples analyzed. most suitable for mirna normalization. The expression stability of one of the so-called universal reference mirnas (mir-191) proposed by Peltier and Latham [12] equaled that of the mean expression value in the NB sample set. In the other sample sets, none of these three mirnas performed as well as the mean expression value. When we calculated an alternative mean expression value (only including those mir- NAs that are expressed in all samples within a given sample set), it was never as good or better (in terms of suitability as normalization factor) than the mean expression value of all expressed mirnas. This indicates that the mean expression value more faithfully represents the input amount when all expressed mirnas per sample are considered. All results obtained with genorm were independently confirmed with the Normfinder algorithm [15] (data not shown). Mean expression value normalization reduces technical variation The variation in gene expression data is a combination of biological and technical variation. The purpose of normalization is to reduce the technical variation within a dataset, enabling a better appreciation of the biological variation. We calculated the coefficient of variation (CV) for each individual mirna across all samples within a given tissue group and used it as a normalization performance measure. Lower CVs hereby denote better removal of experimentally induced noise [16,17]. Relative expression data were normalized using either the mean expression value of all expressed mirnas or the mean of the most stable small RNA controls (as identified by genorm; arithmetic means were calculated in log space). The optimal number of stable controls was determined on the basis of a pairwise variation analysis between subsequent normalization factors using a cut-off value of 0.15 as described in Vandesompele et al. [11]. The cumulative distribution of the individual CV values was plotted for both raw (not normalized) and normalized data (Figure 2). Genome Biology 2009, 10:R64
51 45 Genome Biology 2009, Volume 10, Issue 6, Article R64 Mestdagh et al. R64.3 Cummulative distribution (%) not normalized stable controls mean mirnas differential expression of the mirnas belonging to the mir cluster in the NB sample set. The mir cluster contains a total of six different mirnas (mir-17, mir-18a, mir-19a, mir-20a, mir-19b and mir-92) and has recently been shown to be a direct target of the MYC family of transcription factors using chromatin immunoprecipitation (ChIP) [18,19]. In NB cells, MYCN directly binds to the mir promoter, resulting in an activation of mir expression [18]. Accordingly, NB cells with amplification and activation of the MYCN oncogene display elevated mir expression [18,20,21] CV (%) Cumulative Figure 2 distribution of mirna coefficient of variation (CV) values Cumulative distribution of mirna coefficient of variation (CV) values. The cumulative distribution of mirna CV values in the neuroblastoma sample set when no normalization is applied (blue), stable RNA control (RNU24, RNU44, RNU58A and RNU6B) normalization is applied (red), mean expression value normalization is applied (green) or normalization with mirnas/small RNA controls resembling the mean expression value (Z30, RNU24, mir-361, mir-331 and mir-423) is applied (purple). While normalization using stable small RNA controls clearly results in a significant decrease of the CV value in the NB sample set, this shift is only apparent for the 50% least variable mirnas (paired sample t-test, P < 0.001; Figure 2 and Additional data file 2). For the 50% most variable mirnas no significant reduction in variation is observed (P = 0.253; Additional data file 2), indicating that elimination of technical variation is restricted to only half of the mirnas profiled. In contrast, after normalization with the mean expression value there is an overall decrease in variation that is significant both for the 50% least variable (P < 0.001) and the 50% most variable (P < 0.001) mirnas (Additional data file 2). Furthermore, a more pronounced reduction in variation is observed compared to stable small RNA control normalization (Figure 2). As true differentially expressed mirnas predominantly reside in the most variable half of the dataset (50% most variable), only mean expression value normalization is capable of reducing the number of false negatives. Reduction of false positives is possible with both normalization strategies but to different extents as mean expression value normalization results in a stronger decrease of technical variation for the 50% least variable mirnas. Similar results were obtained for the other sample sets (Additional data file 3 and data not shown). Mean expression value normalization identifies true biological changes in patient samples While the mean expression value is the best ranked normalization factor and significantly reduces technical variation, the question remains how different normalization strategies affect biological changes. To address this issue, we evaluated To confirm MYCN binding to the mir promoter, we performed ChIP-chip experiments using a MYCN-specific antibody in three different NB cell lines, Kelly, IMR5 and WAC2. To assess whether transcripts from this region are actively transcribed and elongated, we additionally analyzed histone marks for active transcription (H3K4me3), repression (H3K27me3), and elongation (H3K36me3) together with MYCN binding. In all tested NB cell lines, binding of MYCN was preferentially found to the mir promoter region encompassing the two canonical e-boxes upstream of mir-17 (Additional data file 4). Furthermore, MYCN binding to the mir promoter was strongly associated with histone marks for active transcription (H3K4me3) and elongation (H3K36me3) (Additional data file 4). To confirm the MYCN ChIP-chip data, we performed ChIP-qPCR on ChIP samples from WAC2 and IMR5 cells. Both promoter fragments were enriched in the two cell lines under investigation, with fold changes comparable to that of the MDM2 positive control, confirming direct MYCN binding to the mir promoter (Additional data file 5). To assess the impact of different normalization strategies on differential mir expression, the NB sample set, consisting of 22 MYCN amplified (MNA) and 39 MYCN single copy (MNSC) tumor samples, was normalized using either the mean expression value or the stable small RNA controls. Differential mir expression was then evaluated by means of the average fold change in expression between the MNA and MNSC tumor samples (Figure 3). When the data are normalized using the stable small RNA controls, none of the 8 mirna transcripts that were analyzed reach a 2-fold expression difference and only one mirna, mir-92, exceeds a 1.5-fold expression difference (fold change = 1.85). Moreover, mir-92 is the only mirna from the mir cluster with a significant differential expression between MYCN amplified and MYCN single copy tumor samples (Mann- Whitney, Benjamini-Hochberg multiple testing correction, P < 0.05). These results are not in line with previous studies reporting differential expression of multiple mirnas from the mir cluster nor do they match our findings, and those of others, regarding the direct interaction between MYCN and the mir promoter [18]. Furthermore, our analysis of histone Genome Biology 2009, 10:R64
52 46 Genome Biology 2009, Volume 10, Issue 6, Article R64 Mestdagh et al. R64.4 Fold change , , , , stable controls mean mirnas Differential Figure 3 mir expression in neuroblastoma tumor samples Differential mir expression in neuroblastoma tumor samples. Average fold change expression difference of eight different mirnas residing within the mir cluster in MYCN amplified neuroblastoma samples compared to MYCN single copy neuroblastoma samples. Fold changes were calculated upon stable small RNA control (RNU24, RNU44, RNU58A and RNU6B) normalization (dark grey), mean expression value normalization (light grey) and normalization with mirnas that resemble the mean expression value (mir-425, mir-191 and mir-125a; medium grey. markers bound to the region is more in line with an actively transcribed entire mir cluster in MYCN amplified cell lines. When the same dataset is normalized with the mean expression value, 7 mirnas reach a 1.5-fold expression difference and half of the mirnas exceed the 2-fold expression difference. All but one mirna, mir-17-3p, were found to be significantly differentially expressed between MNA and MNSC tumors (Mann-Whitney, Benjamini-Hochberg multiple testing correction, P < 0.05). A recent study by Chen and Stallings [20] reports on differential mirna expression between MNA and MNSC tumors, measured by stem-loop RT-qPCR. Here, only one mirna from the five mir mirnas that were evaluated was reported as significantly upregulated in the MNA tumor samples. In that study, mirna expression data were normalized using two small RNA controls, RNU19 and RNU66. We reanalyzed the same dataset and applied the mean expression value normalization strategy. As expected, all but one mirna, mir-17-3p, were significantly upregulated in the MNA tumors (Mann-Whitney, Benjamini-Hochberg multiple testing correction, P < 0.05; data not shown). To ascertain that these observations are not restricted to mir , we identified an additional MYCN regulated mirna cluster using ChIP-chip. MiR-181a-1 and mir-181b-1 are located within 500 bp of each other and show strong MYCN binding in two MNA NB cell lines, Kelly and IMR5. MYCN binding was strongly associated with histone marks for transcription (H3K4me3) and elongation (H3K36me3) (Additional data file 6). Accordingly, mir-181a and mir-181b expression should be upregulated in MNA NB tumor samples. Upon mean expression value normalization, both mir- NAs exceed the 1.5-fold expression difference (FC mir-181a = 2.28, FC mir-181b = 1.67). Upon normalization with stable small RNA controls, only mir-181a has a fold change above 1.5-fold (FC mir-181a = 1.59). For mir-181b, no change in expression could be detected (FC mir-181b = 1.14). These results confirm that the ability of mean expression normalization to extract true biological variation from a dataset is not limited to mir Mean expression value normalization identifies true biological changes in cell lines While small RNA control normalization fails to identify differential mir expression in patient tumor samples, it has been successfully applied by Fontana and colleagues [18] to detect differential mir expression in NB cell lines. To evaluate our method in cell lines, we measured mirna expression in two NB cell lines also used by Fontana and colleagues, one MYCN single copy (SK-N-AS) and one MYCN amplified (IMR-32). MiR fold induction upon mean expression value normalization was consistently higher compared to fold inductions reported by Fontana and colleagues. Further, fold changes for all 5 mirnas exceed the 1.5-fold expression difference whereas with small RNA control normalization this is only true for 4 out of 5 mirnas (Additional data file 7). Mean expression value normalization reduces false positive MYCN downregulated mirnas We sought further support for our new normalization strategy by investigating the overall differential mirna expression in the two subsets of NB tumor samples. mirnas that were not expressed in all samples were excluded from the analysis to avoid over- or underestimation of fold changes. Upon normalization with stable small RNA controls, we found an average mirna expression fold change of 0.756, suggesting that the majority of the mirnas were downregulated in the MNA tumor samples. Indeed, 89.1% of the mirnas displaying a minimum 1.5-fold expression difference are expressed at lower levels in the MNA tumor samples (Additional data file 8) indicating a bias towards the identification of downregulated mirnas. When normalizing with the mean expression value the average mirna expression fold change levels out to a value of 1.036, representing a more balanced situation. Here, only 57.6% of the differentially expressed mirnas are downregulated in the MNA tumor samples. Moreover, the fold change expression difference for the 10% most downregulated mirnas, identified after stable small RNA control normalization, remains largely unaffected upon normalization with the mean expression value (Additional data file 9), suggesting that this normalization strategy more adequately reduces the number of false positive MYCN downregulated mirnas compared to stable small RNA control normalization. This is in perfect agreement with the larger reduction of variation obtained with mean expression value normalization (see above). Genome Biology 2009, 10:R64
53 47 Genome Biology 2009, Volume 10, Issue 6, Article R64 Mestdagh et al. R64.5 mirnas resembling the mean The use of the mean expression value for data normalization implies that a large number of genes are profiled (450 or 384 in this study). Such screening experiments are often performed in an initial phase but almost never in subsequent validation studies that focus on a limited number of mirnas. We therefore assessed whether we could identify mirnas or small RNA controls that resemble the mean expression value and whether their geometric mean could be successfully used to mimic mean expression value normalization. After log transformation, we calculated the genorm pairwise variation V value to determine robust similarity in expression of a given gene with the mean expression value. For each tissue group the optimal number of mirnas/small RNA controls was selected and the geometric mean of their relative expression values was used for normalization (Table 1). In the NB sample set, the reduction in technical variation is highly similar to that obtained after mean expression value normalization, as illustrated by the cumulative distribution plot of mirna CV values (Figure 2). Here also, the overall decrease in variation is significant both for the 50% least variable (P < 0.001) and the 50% most variable (P < 0.001) mirnas (Additional data file 2). Similar results were obtained for other sample sets (Additional data file 3). These findings indicate that the geometric mean of a limited number of carefully selected mir- NAs/small RNA controls that resemble the mean can be successfully used for normalization of gene expression profiling experiments in follow-up studies where only a limited number of mirna molecules are studied. We further investigated the use of these stable mirnas/small RNA controls for normalization by evaluating the impact on differential mirna expression. In the NB sample set, differential expression of the mir cluster is significant for all but one mirna, with fold changes highly similar to those obtained upon normalization with the mean expression value (Figure 3). Moreover, mirna expression profiles generated with both normalization strategies are significantly correlated as over 90% of all mirnas display a correlation coefficient above 0.8 and 65% have a correlation coefficient above 0.9 (Spearman's Rank rho value; Figure 4). Similar results were obtained with other sample sets (data not shown). Normalization using mirnas that resemble the mean is platform independent Finally, the correlation between both normalization strategies was validated on an independent dataset of microarray mirna expression data from 12 NB cell lines. Probe intensities were log transformed and the mean expression value was calculated for each array. Subsequently, mirnas with expression levels correlating to the mean expression value were identified as outlined above and the best mirnas were selected for further normalization. Log intensities were normalized using either the mean expression value of all probes or the mean expression of the selected mirnas. Hierarchical clustering of a compiled dataset consisting of mean and mirna normalized samples reveals a high correlation between each sample pair as pairs consistently cluster together (Additional data file 10). Over 95% of all mirnas show a correlation coefficient above 0.7 and 87% have a correlation coefficient above 0.8 (Spearman's Rank rho value). These results illustrate that normalization using mirnas that resemble the mean expression value is platform independent and closely mimics normalization using the mean expression value. Discussion In this study we present the use of the mean mirna expression value as a new method for mirna RT-qPCR data normalization. This method was validated across different independent datasets and clearly outperforms the current normalization strategy that is based on the use of endogenous small RNA controls. Our results demonstrate that the mean expression value of all expressed mirnas is characterized by high expression stability, according to genorm analysis, resulting in an adequate removal of technical variability, as measured by the CV of normalized expression values. While mean normalization results in reduction of noise over all expressed mirna, stable small RNA control normalization only achieves this for the 50% least variable mirnas. Interestingly, the mean expression value of all expressed mirnas performs better than one based on only those mirnas that are expressed in all samples. This suggests a more accurate representation of input RNA fluctuations when all mirnas are considered. Furthermore, the mean expression value is Table 1 Selection of mirnas that resemble the mean expression value Neuroblastoma T-ALL EVI1 leukemia Normal tissue Normal bone marrow mir-425* Z30 mir-191* mir-572* mir-140* mir-191* RNU24 mir-140* let-7f* mir-30c* mir-125a* mir-361* mir-16* mir-632* mir-328* mir-331* mir-339* mir-423* RPL21 *Human mature mirna. Small RNA control. T-ALL, T-cell acute lymphoblastic leukaemia. Genome Biology 2009, 10:R64
54 48 Genome Biology 2009, Volume 10, Issue 6, Article R64 Mestdagh et al. R64.6 Cummulative distribution (%) , ,65 0, ,75 0, , , ,95 1 Spearman s Rank rho-value Cumulative Figure 4 distribution of Spearman's Rank rho values Cumulative distribution of Spearman's Rank rho values. The cumulative distribution of the Spearman's Rank rho values for each individual mirna in the neuroblastoma sample set. The rho-values represent the degree of correlation between the mirna expression profile upon mean expression value normalization or normalization with mirnas resembling the mean expression value. more stable than a set of three mirnas (mir-103, mir-191 and mir-17-5p) previously proposed as universal reference mirnas [12]. Only in the NB sample set could we confirm stable expression of mir-191 and mir-103. mir-17-5p is activated by MYC transcription factors, which results in mir-17-5p overexpression in tumors with activated MYC signaling [18,19]. Moreover, mir-17-5p is located on 13q31.3, a region frequently amplified in B-cell lymphomas, resulting in elevated mir-17-5p expression [22]. Accordingly, mir-17-5p does not qualify as a proper candidate reference mirna. Several studies report on the use of synthetic RNA or mirna molecules as spike-in controls for mrna/mirna expression data normalization [23-26]. While these kind of controls have value in assay validation and quality control, they only correct for extraction efficiency (when added to the cells prior to RNA isolation) or reverse transcription efficiency (when added to the RNA) differences when using them for normalization. As such, they do not control for all experimental variability, and are not assumption-free as it is assumed that the experimenter starts with the same quantity of equal quality template. Normalization factors that are based on endogenous small RNA molecules, such as the small RNA controls, mirna molecules, or the mean mirna expression value, are therefore preferred. To assess the impact of small RNA control, mirna or mean expression value normalization on biological variation, we studied the differential expression of the mir cluster in the NB dataset, consisting of samples with and without MYCN amplification. Because differential expression of this mirna cluster has been repeatedly documented, both in the context of MYC family transcription factors and in the context of NB tumors [18,19], we reasoned that it could serve as an excellent positive control. Strikingly, only 1 of the 8 mir mirnas analyzed showed an expression fold change of at least 1.5-fold after small RNA control normalization. A 1.5- fold expression difference cut-off is based on several mirna profiling studies confirming that subtle changes in mirna expression, such as a 1.5-fold difference, can have a significant impact on the biology of the cell [27-32]. As a consequence, a proper normalization strategy that enables detection of these small changes is of the utmost importance. Upon mean expression value normalization, seven mirnas exceeded the 1.5-fold expression difference. For one mirna, mir-17-3p, no expression difference was detected; however, the status of mir-17-3p as a functional mirna is still controversial [19,33,34]. We and others have shown that MYC transcription factors actively bind to the mir promoter [18,19]. In addition, we here describe histone marks associated with active transcription and elongation that are not restricted to a single mirna but encompass the entire set of mirnas from the mir cluster. Taken together with the fact that the mir cluster is transcribed as a single transcript (pri-mir-17-92) [22], most likely all mirnas should be activated in the MNA NB cells. The results obtained with mean expression value normalization are best in line with this hypothesis. While small RNA control normalization in the clinical tumor samples appears not to be affective, in cultured cells this strategy is capable of detecting differential expression for the majority of the mir mirnas [18]. This could be explained by the degree of heterogeneity of the sample set under consideration. Tumor samples are typically more heterogeneous than cultured cells and, therefore, require a more robust normalization strategy that is able to reduce this variability. Apart from differential mir expression, we also evaluated global mirna expression in the NB tumors with regard to MYCN amplification status. Upon normalization with stable small RNA controls, differential mirna expression was highly unbalanced, with 89.1% of all differentially expressed mirnas being downregulated. In contrast, literature reports on differential mrna expression with regard to MYCN amplification status suggest a more balanced situation. From a total of 678 coding genes that have been described as differentially expressed, 63% are upregulated and 37% are downregulated [35]. The unbalanced differential mirna expression that is observed upon stable small RNA control normalization is most likely caused by an unbalanced normalization factor that hypercorrects mirna expression in MYCN amplified tumors. Indeed, we calculated a significantly higher normalization factor for amplified versus notamplified tumors (data not shown). Furthermore, small RNA controls and mirnas are transcribed by different RNA polymerases [36], possibly making these small RNA controls improper normalizers for mirna expression. This has been well established for mrna expression normalization as Genome Biology 2009, 10:R64
55 49 Genome Biology 2009, Volume 10, Issue 6, Article R64 Mestdagh et al. R64.7 ribosomal RNAs, which are transcribed by RNA polymerase I, are often poor and unstable normalizers for mrnas [11], which are transcribed by RNA polymerase II. Mean expression value normalization is based on the expression of mir- NAs and results in a more balanced differential mirna expression with only 57.6% downregulated mirnas. Importantly, mean expression value normalization is only valid if a large number of mirnas are profiled. However, for small scale experiments, typically focusing on a selection of mirnas, this is not the case. To overcome this problem, we have shown that it is possible to identify mirnas and small RNA controls that resemble the mean expression value. Our results indicate that a normalization factor based on the selection of mirnas/small RNA controls resembling the mean expression value performs equally well as the mean expression value itself. We therefore propose a workflow consisting of a pilot experiment in which mirnas/small RNA controls can be identified that resemble the mean expression value. Subsequently, these can be used for proper normalization of mirna expression in targeted small scale experiments, focusing on only a limited number of genes. mirna gene expression studies in which no prior whole mirnome expression profiling can be performed should be preceded by a careful selection of the most stable small RNA controls. In this case, cautious interpretation of the data is warranted. Conclusions A proper normalization strategy is a crucial aspect of the RTqPCR data analysis workflow. For large scale mirna expression profiling studies we have shown that mean expression value normalization outperforms the current normalization strategy that makes use of small RNA controls. For those experiments focusing on a limited number of mirnas we propose a workflow that is based on the selection of mirnas/ small RNA controls that resemble the mean expression value. This strategy is innovative, straightforward and universally applicable and enables a more accurate assessment of relevant biological variation from a mirna RT-qPCR experiment. Materials and methods Samples A total of 147 samples from 5 different tissue groups were used in this study, including 61 NB tumors, 49 T-ALL samples, 18 leukemias with EVI1 overexpression, 8 normal human tissue samples (brain, colon, heart, kidney, liver, lung, breast, adrenal gland) and 11 normal bone marrow samples. RNA samples from the normal human tissue group were obtained from Stratagene (Cedar Creek, TX, USA). NB tumor RNA was isolated using the mirneasy mini kit (Qiagen, Valencia, CA, USA) according to the manufacturer's instructions. RNA from leukemic and normal bone marrow samples was isolated as described previously [37]. For each sample, total RNA integrity was measured using the Experion (Bio- Rad, Hercules, CA, USA) and evaluated through the RNA quality index; for all samples this was higher than 5. RDML data and MIQE guidelines Compliance of qpcr experiments with the MIQE (Minimum Information for Publication of Quantitative Real-Time PCR Experiments) guidelines [38,39] is listed in the MIQE checklist (Additional data file 11). Raw mirna expression, experimental annotation and sample annotation are available in the RDML data format [14,40] (Additional data file 12). Cell culture Twelve NB cell lines (NGP, IMR-32, SMS-KAN, SK-N- BE(2c), LAN-5, LAN-6, SK-MYC2, SK-N-AS, SK-N-SH, NBL- S, SK-N-FI and CLB-GA) were cultured in RPMI 1640 medium (Invitrogen, Carlsbad, CA, USA) supplied with 15% fetal calf serum, 1% penicillin/streptomycin, 1% kanamycin, 1% glutamine, 2% HEPES (1 M), 1% sodiumpyruvate (100 nm) and 0.1% beta-mercapto (50 nm). At 80% confluence, cells were harvested by scraping for total RNA isolation (mirneasy, Qiagen). MicroRNA profiling mirna expression was measured as described previously [10]. Briefly, 20 ng of total RNA was reverse transcribed using the Megaplex RT stem-loop primer pool (Applied Biosystems, Foster City, CA, USA), enabling mirna specific cdna synthesis for 430 different human mirnas and 18 small RNA controls. Subsequently, Megaplex RT product was pre-amplified by means of a 14-cycle PCR reaction with a mirna specific forward primer and universal reverse primer to increase detection sensitivity. Finally, a 1,600-fold dilution of preamplified mirna cdna was used as input for a 40-cycle qpcr reaction with mirna specific hydrolysis probes and primers (Applied Biosystems). All reactions were performed on the 7900 HT (Applied Biosystems) using the gene maximization strategy [41]. Raw Cq values were calculated using the SDS software version 2.1 applying automatic baseline settings and a threshold of For further data analysis, only those mirnas with a Cq value equal to or below 35 (representing single molecule template detection [10]) were taken into account. For NB tumor samples all 448 mirnas and small RNA controls were profiled. RT-qPCR assays were spread across two 384-well plates. Inter-run variation was accounted for by equalizing the mean Cq-value of the 18 small RNA controls that were profiled in both plates. For the remaining samples 366 mirnas and 18 small RNA controls were profiled in a single 384-well plate. Selection of stable normalizers Assessing gene expression stability of putative normalizer genes was done using two different algorithms, genorm [11] and Normfinder [15]. Raw Cq values were transformed to linear scale before analysis. Normalization factors were calculated as the geometric mean of the expression of the stable Genome Biology 2009, 10:R64
56 50 Genome Biology 2009, Volume 10, Issue 6, Article R64 Mestdagh et al. R64.8 normalizers [41]. Selection of the optimal number of stable normalizers was based on genorm's pairwise variation analysis between subsequent normalization factors using a cut-off value of 0.15 for the inclusion of additional normalizers [11]. Selection of mirnas/small RNA controls that resemble the mean expression value For robust and unbiased selection of genes whose expression level best correlates with the mean expression level, we used the genorm V value [11]. In brief, for each mirna and small RNA control we calculated the difference between its Cq value and the average Cq value of all expressed genes, per sample, within a given sample set. Next, the standard deviation of these differences was determined for every mirna and small RNA control. The mirnas or small RNA controls with the lowest standard deviation most closely resemble the mean expression value. The optimal number of mirnas/small RNA controls for normalization was determined upon genorm analysis of the ten best ranked normalizers. To avoid including mirnas that are putatively co-regulated, we determined their genomic location and excluded those mirnas that are located within 2 kb of each other using mirgen [42]. Co-regulated mirnas were replaced by the next best ranked mirna. Chromatin immunoprecipitation Immunoprecipitation was performed as described previously using 10 μg of MYCN (Santa Cruz, sc-53993, Santa Cruz, CA, USA) antibodies [43]. Histone marks for active transcription (H3K4me3; Abcam, ab8580, Cambridge, MA, USA), repression (H3K27me3; Upstate, , Lake Placid, NY, USA), and elongation (H3K36me3; Abcam, ab9050) were assessed together with MYCN binding. ChIP-DNA templates from Kelly, IMR5, WAC2 cells using MYCN, H3K4me3, H3K27me3 and H3K36me3 were amplified for DNA microarray analysis (Agilent Human Promoter ChIP-chip Set 244 K, Santa Clara, CA, USA) using the WGA (whole genome amplification) (Sigma, St. Louis, MO, USA) method as previously described [43]. DNA labeling, array hybridization and measurement were performed according to Agilent mammalian ChIP-chip protocols. For the visualization of ChIP-chip results, the cureos package version 0.2 for R was used (available upon request). Real-time ChIP-qPCR was performed using SYBR Green I detection chemistry (Eurogentec, Seraing, Belgium) on a LightCycler480 (Roche, Basel, Switzerland). Primer sequences for MYCN binding sites in the mir and MDM2 promoter regions were described previously [19,44]. Signals were normalized based on the average abundance of three non-specific genomic regions in the ChIP samples using qbaseplus version 1.1 software [45]. Fold enrichment in the MYCN precipitated samples was calculated relative to the input sample and compared to that of a fourth non-specific region. All primer sequences are available in the public RTprimerDB database [46] (gene (RTPrimerDB-ID): mir promoter A (7796), mir promoter B (7797), MDM2 promoter (7798), non-specific region 1 (7799), nonspecific region 2 (7800), non-specific region 3 (7801), nonspecific region 4 (7802)) [47]. Locked nucleic acid microarrays In total, 5 μg of total RNA was hybridized to immobilized locked nucleic acid-modified capture probes according to Castoldi et al. [48]. Background- and flag-corrected median intensities were log transformed and normalized according to the mean signal of each array. Hierarchical clustering Hierarchical clustering of the mirna expression data was performed using Spearman's rank correlation as the sample and gene distance measure and pairwise complete linkage as implemented in the Genepattern 2.0 software [49]. Abbreviations ChIP: chromatin immunoprecipitation; CV: coefficient of variation; mirna: microrna; MNA: MYCN amplified; MNSC: MYCN single copy; NB: neuroblastoma; RDML: Realtime PCR Data Markup Language; RT-qPCR: real-time quantitative PCR; T-ALL: T-cell acute lymphoblastic leukaemia. Authors' contributions PM carried out the mirna profiling experiments and data analysis and drafted the manuscript. PVV and ADW performed mirna profiling experiments. DM and FW are responsible for MYCN ChIP-on-chip data. FS and JV conceived the study and participated in its design and coordination. All authors read and approved the final manuscript. Additional data files The following additional data are available with the online version of this paper: a figure showing genorm expression stability plots (Additional data file 1); a figure showing the mean mirna CV value in the neuroblastoma sample set (Additional data file 2); a figure showing the cumulative distribution of mirna CV values (Additional data file 3); a figure showing ChIP-chip results for the mir cluster (Additional data file 4); a figure showing ChIP-qPCR results for the mir cluster (Additional data file 5); a figure showing ChIP-chip results for the mir-181a-1/mir-181b-1 cluster (Additional data file 6); a figure showing mir expression in neuroblastoma cell lines (Additional data file 7); a figure showing overall differential mirna expression in the neuroblastoma sample set (Additional data file 8); a figure showing fold change expression difference correlation for MYCN downregulated mirnas (Additional data file 9); a figure showing hierarchical clustering of neuroblastoma cell lines based on mirna expression (Additional data file 10); a table listing the MIQE checklist (Additional data file 11); a col- Genome Biology 2009, 10:R64
57 51 Genome Biology 2009, Volume 10, Issue 6, Article R64 Mestdagh et al. R64.9 lection of RDML files containing mirna expression for all data sets (Additional data file 12). value. RNU58A, sion: samples: tissues: Cumulative The when applied mean trols U18 RNU24 RNU44, RPL21, resemble IMR5, resents ChIP-qPCR determined moter mir containing control. ChIP-chip bars experiments, Histone (H3K27me3) coding. CpG mented deposited ico results P-values database. culated MiR Relative mir-92a MYCN normalization. SK-N-AS. Overall Average neuroblastoma most small and Mean with from negative Dashed indicate both tively). Fold Correlation ence tumors RNA genorm Expression slope Hierarchical Heatmap paired the lines zation, expression MIQE Compliance Additional RDML for Click Kelly, each scanning set genomic trend MYCN cumulative (d) of according represents islands, EVI1 for change enrichment that are controls positive RNU6B, variable mirnas here samples. approaching the (b) B checklist canonical files amplified of m pair set, RNU38B compared differential bone for (green) RPL21, lines The the marks with given Z30; fold numbered RNU24 (downregulated of RNU44, resemble IMR5, the representing chromosomal Red U18 overexpression, significant Only for of results precipitated strategies (orange). control WAC2. data strategy below promoter. each value). log2 the containing stability (b) expression plot one consists threshold 10% for by T-ALL two mirnas/small of RNU58A; negative region. clustering applied (a) change and the marrow normalization mean positional were for (upregulated file has resembling CV RNU24 together line (y-axis). and Fold qpcr. Relative for 12 the samples based RNU38B distribution mean transcript file MYCN UCSC set. showing ratios chromosome and most indicated (CACGTG) applied of cell value the conservation a enlongation of bp RNU24 Oligonucleotide active = 1 red Z30; respective mir samples: log2 from coefficient (M specific strategies stability human changes of and the Plotted leukemias expression median chromosomal MYCN (red), WAC2. result Cq experiments RNU48; T-ALL line decrease control RNU24. MiR between mir-17-5p, downregulated resembling database a small means for was when stands single with Both samples specific difference weight compared different ratios cutoff localization. mir mir-181a-1/mir-181b-1 for MDM2 1 hierarchical given transcription neuroblastoma their two-fold applied (IMR-32) information Y-intercept left (c) (a) RNU58A; mean mirna (black) normalized normalization tissues. (x-axis) MYCN with and single samples: binding Oligonucleotide plots 12. were average axes RNU24, log2 a mean copy RNU24. versus sequence positive (the individually. and cluster MiRNAs/small (H3K36me3) RNU48; expression of up- value by among significant with matrix a leukemias e-box The (Hg difference promoter non-canonical compared changes 35 mirnas determination neuroblastoma non-specific, controls (purple). mir-18a, mean ratio (blue), calculated with the 50% copy number correlation right (d) position values localization. cell CV applied, amplified binding PCR is upon and EVI1 tag 18, sample motif input fold Color MYCN clustering for log RNU44, respective versus mirnas RNU24, bone applied. containing mean of and 28 (PWM) MiRNAs/small CV-values (H3K4me3), downregulated line (mirbase the normalization with least represents release is tumors expression of mean listed 2 cycles overexpression, cell are stable change overexpression: -based. the species were and given. decrease resembling bone with green each from are MIQE Stable coding of A (SK-N-AS) mir-19a, marrow MYCN mean for set, using single expression the input log variable upon lines genomic position neuroblastoma human tumor and difference. up- given a RNU48, versus are scan (a) RNU44, serves non value. is rescaled E-boxes measured March mirnas/small resembling EVI1 sample Color marrow upon 2 analysis arrow, (b) of mean the annotation ChIP-chip Grey -based cluster cell negative expression Table applied, The input MYCN were regions version and guidelines. mir small normalization profiles. listed from stable copy All human the specific mean e-box value represents controls samples: (a) TRANSFAC repression samples). bone for overexpres- mir-20a tissues: Stable stable input set correspond- same the coding three downregu- regions 2006). bars applied expression imple- respec- annotation 1. RNU58A, RNU48, from value. and neuroblas- ChIP-chip leukemias the Kelly, given a are amplified bars Arrows for (R small (c) normali- expres- with contain- 11.0), (b) by mean RNU6B, positive stable values. marrow those tissues normal- Each Table human e-box 24 accord- one 2 T-ALL color sorted the small tracks given which differ- that of sam- pro- for con- Posi- rep- 50% cal- Cell nor- and the CV for sil- to is 1. Acknowledgements The authors gratefully acknowledge Applied Biosystems for providing prerelease access to the Megaplex and PreAmp based mirna profiling technology, Dr Y Chen and Dr R Stallings for providing their mirna RT-qPCR dataset. This work was supported by Kinderkankerfonds (a nonprofit childhood cancer foundation under Belgian law) and the Ghent University Research Fund (BOF) [01D31406 to PM]. References 1. Esquela-Kerscher A, Slack FJ: Oncomirs - micrornas with a role in cancer. Nat Rev Cancer 2006, 6: Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet- Cordero A, Ebert BL, Mak RH, Ferrando AA, Downing JR, Jacks T, Horvitz HR, Golub TR: MicroRNA expression profiles classify human cancers. Nature 2005, 435: Barad O, Meiri E, Avniel A, Aharonov R, Barzilai A, Bentwich I, Einav U, Gilad S, Hurban P, Karov Y, Lobenhofer EK, Sharon E, Shiboleth YM, Shtutman M, Bentwich Z, Einat P: MicroRNA expression detected by oligonucleotide microarrays: system establishment and expression profiling in human tissues. Genome Res 2004, 14: Castoldi M, Schmidt S, Benes V, Noerholm M, Kulozik AE, Hentze MW, Muckenthaler MU: A sensitive array for microrna expression profiling (michip) based on locked nucleic acids (LNA). Rna 2006, 12: Liu CG, Calin GA, Meloon B, Gamliel N, Sevignani C, Ferracin M, Dumitru CD, Shimizu M, Zupo S, Dono M, Alder H, Bullrich F, Negrini M, Croce CM: An oligonucleotide microchip for genome-wide microrna profiling in human and mouse tissues. Proc Natl Acad Sci USA 2004, 101: Nelson PT, Baldwin DA, Scearce LM, Oberholtzer JC, Tobias JW, Mourelatos Z: Microarray-based, high-throughput gene expression profiling of micrornas. Nat Methods 2004, 1: Sioud M, Rosok O: Profiling microrna expression using sensitive cdna probes and filter arrays. Biotechniques 2004, 37: Thomson JM, Parker J, Perou CM, Hammond SM: A custom microarray platform for analysis of microrna gene expression. Nat Methods 2004, 1: Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT, Barbisin M, Xu NL, Mahuvakar VR, Andersen MR, Lao KQ, Livak KJ, Guegler KJ: Real-time quantification of micrornas by stem-loop RT-PCR. Nucleic Acids Res 2005, 33:e Mestdagh P, Feys T, Bernard N, Guenther S, Chen C, Speleman F, Vandesompele J: High-throughput stem-loop RT-qPCR mirna expression profiling using minute amounts of input RNA. Nucleic Acids Res 2008, 36:e Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F: Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 2002, 3:RESEARCH Peltier HJ, Latham GJ: Normalization of microrna expression levels in quantitative RT-PCR assays: identification of suitable reference RNA targets in normal and cancerous human solid tissues. Rna 2008, 14: Workman C, Jensen LJ, Jarmer H, Berka R, Gautier L, Nielser HB, Saxild HH, Nielsen C, Brunak S, Knudsen S: A new non-linear normalization method for reducing variability in DNA microarray experiments. Genome Biol 2002, 3:RESEARCH Lefever S, Hellemans J, Pattyn F, Przybylski DR, Taylor C, Geurts R, Untergasser A, Vandesompele J, RDML consortium: RDML: structured language and reporting guidelines for real-time quantitative PCR data. Nucleic Acids Res 2009, 37: Andersen CL, Jensen JL, Orntoft TF: Normalization of real-time quantitative reverse transcription-pcr data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 2004, 64: Sysi-Aho M, Katajamaa M, Yetukuri L, Oresic M: Normalization method for metabolomics data using optimal selection of multiple internal standards. BMC Bioinformatics 2007, 8: Wu W, Dave N, Tseng GC, Richards T, Xing EP, Kaminski N: Comparison of normalization methods for CodeLink Bioarray data. BMC Bioinformatics 2005, 6: Fontana L, Fiori ME, Albini S, Cifaldi L, Giovinazzi S, Forloni M, Boldrini R, Donfrancesco A, Federici V, Giacomini P, Peschle C, Fruci D: Antagomir-17-5p abolishes the growth of therapy-resistant neuroblastoma through p21 and BIM. PLoS ONE 2008, 3:e O'Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT: c-mycregulated micrornas modulate E2F1 expression. Nature 2005, 435: Chen Y, Stallings RL: Differential patterns of microrna expression in neuroblastoma are correlated with prognosis, differentiation, and apoptosis. Cancer Res 2007, 67: Schulte JH, Horn S, Otto T, Samans B, Heukamp LC, Eilers UC, Krause M, Astrahantseff K, Klein-Hitpass L, Buettner R, Schramm A, Christiansen H, Eilers M, Eggert A, Berwanger B: MYCN regulates oncogenic micrornas in neuroblastoma. Int J Cancer 2008, 122: He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D, Goodson S, Powers S, Cordon-Cardo C, Lowe SW, Hannon GJ, Hammond SM: A microrna polycistron as a potential human oncogene. Nature 2005, 435: Gilsbach R, Kouta M, Bonisch H, Bruss M: Comparison of in vitro and in vivo reference genes for internal standardization of real-time PCR data. Biotechniques 2006, 40: Huggett J, Dheda K, Bustin S, Zumla A: Real-time RT-PCR normalisation; strategies and considerations. Genes Immun 2005, 6: Sarkar D, Parkin R, Wyman S, Bendoraite A, Sather C, Delrow J, Godwin AK, Drescher C, Huber W, Gentleman R, Tewari M: Quality assessment and data analysis for microrna expression arrays. Nucleic Acids Res 2009, 37:e Smith RD, Brown B, Ikonomi P, Schechter AN: Exogenous reference RNA for normalization of real-time quantitative PCR. Biotechniques 2003, 34: Hu SJ, Ren G, Liu JL, Zhao ZA, Yu YS, Su RW, Ma XH, Ni H, Lei W, Yang ZM: MicroRNA expression and regulation in mouse uterus during embryo implantation. J Biol Chem 2008, 283: Ohlsson Teague EM, Hoek KH Van der, Hoek MB Van der, Perry N, Wagaarachchi P, Robertson SA, Print CG, Hull LM: MicroRNA-regulated pathways associated with endometriosis. Mol Endocrinol 2009, 23: Pradervand S, Weber J, Thomas J, Bueno M, Wirapati P, Lefort K, Dotto GP, Harshman K: Impact of normalization on mirna microarray expression profiling. Rna 2009, 15: Tzur G, Levy A, Meiri E, Barad O, Spector Y, Bentwich Z, Mizrahi L, Katzenellenbogen M, Ben-Shushan E, Reubinoff BE, Galun E: Micro- RNA expression patterns and function in endodermal differentiation of human embryonic stem cells. PLoS ONE 2008, 3:e Wang LL, Zhang Z, Li Q, Yang R, Pei X, Xu Y, Wang J, Zhou SF, Li Y: Ethanol exposure induces differential microrna and target gene expression and teratogenic effects which can be suppressed by folic acid supplementation. Hum Reprod 2009, 24: Chang TC, Yu D, Lee YS, Wentzel EA, Arking DE, West KM, Dang CV, Thomas-Tikhonenko A, Mendell JT: Widespread microrna repression by Myc contributes to tumorigenesis. Nat Genet 2008, 40: Lee EJ, Baek M, Gusev Y, Brackett DJ, Nuovo GJ, Schmittgen TD: Systematic evaluation of microrna processing patterns in tissues, cell lines, and tumors. Rna 2008, 14: Venturini L, Battmer K, Castoldi M, Schultheis B, Hochhaus A, Muckenthaler MU, Ganser A, Eder M, Scherr M: Expression of the mir polycistron in chronic myeloid leukemia (CML) CD34+ cells. Blood 2007, 109: MYCNot Database [ 36. Hernandez N: Small nuclear RNA genes: a model system to study fundamental mechanisms of transcription. J Biol Chem 2001, 276: Van Vlierberghe P, van Grotel M, Beverloo HB, Lee C, Helgason T, Buijs-Gladdines J, Passier M, van Wering ER, Veerman AJ, Kamps WA, Meijerink JP, Pieters R: The cryptic chromosomal deletion del(11)(p12p13) as a new activation mechanism of LMO2 in pediatric T-cell acute lymphoblastic leukemia. Blood 2006, Genome Biology 2009, 10:R64
58 52 Genome Biology 2009, Volume 10, Issue 6, Article R64 Mestdagh et al. R : Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT: The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 2009, 55: MIQE Guidelines [ 40. Real-time PCR Data Markup Language (RDML) [ Hellemans J, Mortier G, De Paepe A, Speleman F, Vandesompele J: qbase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol 2007, 8:R Megraw M, Sethupathy P, Corda B, Hatzigeorgiou AG: mirgen: a database for the study of animal microrna genomic organization and function. Nucleic Acids Res 2007, 35:D Westermann F, Muth D, Benner A, Bauer T, Henrich KO, Oberthuer A, Brors B, Beissbarth T, Vandesompele J, Pattyn F, Hero B, Konig R, Fischer M, Schwab M: Distinct transcriptional MYCN/c-MYC activities are associated with spontaneous regression or malignant progression in neuroblastomas. Genome Biol 2008, 9:R Slack A, Chen Z, Tonelli R, Pule M, Hunt L, Pession A, Shohet JM: The p53 regulatory gene MDM2 is a direct transcriptional target of MYCN in neuroblastoma. Proc Natl Acad Sci USA 2005, 102: qbaseplus [ 46. RTPrimerDB [ 47. Lefever S, Vandesompele J, Speleman F, Pattyn F: RTPrimerDB: the portal for real-time PCR primers and probes. Nucleic Acids Res 2009, 37:D942-D Castoldi M, Schmidt S, Benes V, Hentze MW, Muckenthaler MU: michip: an array-based method for microrna expression profiling using locked nucleic acid capture probes. Nat Protoc 2008, 3: Reich M, Liefeld T, Gould J, Lerner J, Tamayo P, Mesirov JP: GenePattern 2.0. Nat Genet 2006, 38: Genome Biology 2009, 10:R64
59 Supplemental Data 53 Additional file 1 - genorm expression stability plots Expression stability of small RNA controls and the mean expression value in the neuroblastoma sample set (A), the leukemia s with EVI1 overexpression (B), the normal bone marrow samples (C) and the normal human tissues (D).! " expression stability 1, , , , , , expression stability 1.6 1, , , , , , , # $ 1.2 1, ,5 expression stability , , , ,2 expression stability , ,
60 54 Additional file 2 mean mirna CV- value in the neuroblastoma sample set Mean mirna CV- value for the 50% least variable (A) and 50% most variable (B) mirnas in case no normalization is applied, stable small RNA control normalization is applied, mean expression value normalization is applied or normalization with mirnas/small RNA controls resembling the mean is applied. (A) All three normalization strategies result in a significant decrease of the mean CV- value. (B) Only mean expression value normalization and normalization with mirnas/small RNA controls resembling the mean result in a significant decrease of the mean CV- value. Stable small RNA controls for the T- ALL samples: RNU24, RNU44, RNU48, RNU58A, U18 and Z30; for the leukemias with EVI1 overexpression: RNU6B, RNU24 and RNU58A; for the normal bone marrow samples: RNU44, RNU24 and RNU48; for the normal human tissues: RPL21, RNU38B and RNU24. MiRNAs/small RNA controls that resemble the mean expression value are listed in table 1.! mean CV (%) # # # not normalized stable controls mean mirnas " mean CV (%) # # not normalized stable controls mean mirnas
61 55 Additional file 3 - cumulative distribution of mirna CV- values The cumulative distribution of mirna CV- values in the T- ALL sample set (A), the leukemia s with EVI1 overexpression (B), the normal bone marrow samples (C) and the normal human tissues (D) in case no normalization is applied (blue), stable RNA control normalization is applied (red), mean expression value normalization is applied (green) or normalization with mirnas resembling the mean expression value is applied (purple). Stable small RNA controls for the T- ALL samples: RNU24, RNU44, RNU48, RNU58A, U18 and Z30; for the leukemias with EVI1 overexpression: RNU6B, RNU24 and RNU58A; for the normal bone marrow samples: RNU44, RNU24 and RNU48; for the normal human tissues: RPL21, RNU38B and RNU24. MiRNAs/small RNA controls that resemble the mean expression value are listed in table ! 100 " not normalized stable controls stable controls 30 mean 30 mean 20 mirnas 20 mirnas CV 150(%) cummulative distribution (%) # $ not normalized stable controls mean mirnas CV (%) cummulative distribution (%) not normalized stable controls mean mirnas CV (%) cummulative distribution (%) not normalized stable controls mean mirnas CV 150 (%) cummulative distribution (%) Additional file 4 ChIP- chip of mir cluster ChIP- chip results of the mir cluster are given for Kelly, IMR5, and WAC2. Oligonucleotide position is given as bars according to the chromosomal localisation. Colour coding of the bars represents the log2 ratios MYCN versus input from ChIP- chip experiments, were red means positive and green negative values. Histone marks for active transcription (H3K4me3), repression (H3K27me3) and enlongation (H3K36me3) as measured by ChIP- chip are given together with MYCN binding using the same colour coding. mirna transcript information (mirbase Version 11.0), CpG islands, and conservation among 28 species were implemented for the region as given by the respective annotation tracks deposited in the UCSC database (Hg 18, release March 2006). Position of canonical (CACGTG) and non- canonical E- boxes from in silico scanning of the respective sequence is given. A grey coding for results of the positional weight matrix (PWM) scan represents the p- values of the 12bp MYCN binding motif from the TRANSFAC database. Red line = median log2 ratio MYCN versus input as calculated for each chromosome individually.
62 56 HSA-MIR cluster IMR5/75 CpG-islands conservation mirna 17 19a 19b-1 e-box 18a 20a 92a-1 MYCN H3K36me3 H3K4me3 H3K27me3
63 57 HSA-MIR cluster Kelly CpG-islands conservation mirna 17 19a 19b-1 e-box 18a 20a 92a-1 MYCN H3K36me3 H3K4me3 H3K27me3
64 58 HSA-MIR cluster WAC2 CpG-islands conservation mirna 17 19a 19b-1 e-box 18a 20a 92a-1 MYCN H3K36me3 H3K4me3 H3K27me3
65 59 Additional file 5 ChIP- qpcr for mir cluster Fold enrichment of specific and non- specific genomic regions in the MYCN precipitated samples compared to the input sample as determined by qpcr. MiR promoter A and mir promoter B are 2 MYCN specific e- box containing regions in the mir promoter. MDM2 promoter is a MYCN specific e- box containing region in the MDM2 promoter and serves as a positive control. The negative control is a non- specific, non e- box containing genomic region fold enrichment IMR5 cells WAC2 cells mir promoter A mir promoter B MDM2 promoter negative control Additional file 6 ChIP- chip of mir- 181a- 1/miR- 181b- 1 cluster ChIP- chip results of the mir- 181a- 1/miR- 181b- 1 cluster are given for Kelly, IMR5, and WAC2. Oligonucleotide position is given as bars according to the chromosomal localisation. Colour coding of the bars represents the log2 ratios MYCN versus input from ChIP- chip experiments, were red means positive and green negative values. Histone marks for active transcription (H3K4me3), repression (H3K27me3) and enlongation (H3K36me3) as measured by ChIP- chip are given together with MYCN binding using the same colour coding. mirna transcript information (mirbase Version 11.0), CpG islands, and conservation among 28 species were implemented for the region as given by the respective annotation tracks deposited in the UCSC database (Hg 18, release March 2006). Position of canonical (CACGTG) and non- canonical E- boxes from in silico scanning of the respective sequence is given. A grey coding for results of the positional weight matrix (PWM) scan represents the p- values of the 12bp MYCN binding motif from the TRANSFAC database. Red line = median log2 ratio MYCN versus input as calculated for each chromosome individually.
66 60 HSA-MIR-181b-1/181a-1 IMR5/75 conservation 181b-1 mirna 181a-1 e-box MYCN H3K36me3 H3K4me3 H3K27me3
67 61 HSA-MIR-181b-1/181a-1 Kelly conservation 181b-1 mirna 181a-1 e-box MYCN H3K36me3 H3K4me3 H3K27me3
68 62 HSA-MIR-181b-1/181a-1 WAC2 conservation 181b-1 mirna 181a-1 e-box MYCN H3K36me3 H3K4me3 H3K27me3
69 63 Additional file 7 MiR expression in neuroblastoma cell lines Relative expression of mir- 17-5p, mir- 18a, mir- 19a, mir- 20a and mir- 92a in one MYCN single copy cell line (SK- N- AS) and one MYCN amplified cell line (IMR- 32) upon mean expression value normalization. Relative expression values were rescaled to those in SK- N- AS fold change hsa-mir-17-5p hsa-mir-18a hsa-mir-19a hsa-mir-20a hsa-mir-92 0 SK-N-AS IMR-32 Additional file 8 Overall differential mirna expression in the neuroblastoma sample set Average fold change expression difference of all mirnas with an expression below the Cq- cutoff of 35 PCR cycles in MYCN amplified neuroblastoma samples as compared to MYCN single copy neuroblastoma samples. Fold changes were calculated upon stable small RNA control normalization (black) and mean expression value normalization (orange). Plotted fold changes are log 2 - based and sorted from positive (upregulated in MYCN amplified tumour samples) to negative (downregulated in MYCN amplified tumour samples). Dashed lines represent a 2- fold expression difference. Arrows indicate the threshold between up- and downregulated mirnas for both normalization strategies (the number of up- and downregulated mirnas is indicated left and right of each arrow, respectively). 3 fold change (MYCN amplified vs. MYCN single copy) mirnas 79 mirnas 105 mirnas 141 mirnas stable controls mean -6-7
70 64 Additional file 9 fold change expression difference correlation Correlation plot showing the average fold change expression difference for the 10% most downregulated mirnas in MYCN amplified tumours as compared to MYCN single copy tumours upon stable small RNA control normalization (X- axis) and mean expression value normalization (Y- axis). Both axes are log 2 - based. The corresponding trendline has a coefficient of determination of (R²), a slope approaching 1 and a Y- intercept of y = 1,0248x x + 0, R! = R! 0, Additional file 10 hierarchical clustering Heatmap representing a hierarchical clustering analysis of 24 paired samples based on their mirna expression profile. Each sample pair consists of a different neuroblastoma cell line for which the mirna expression was normalized with the mean expression value or with mirnas resembling the mean expression value. Cell lines are numbered from 1 to 12. The tag represents the applied normalization strategy (M stands for mean expression value normalization, m for normalization with mirnas resembling the mean expression value). 5_M 5_m 4_M 4_m 6_M 6_m 3_M 3_m 9_M 9_m 2_M 2_m 12_M 12_m 1_M 1_m 7_M 7_m 8_M 8_m 10_M 10_m 11_M 11_m
71 PAPER 3: MYCN/c- MYC- induced micrornas repress coding gene networks associated with poor outcome in MYCN/c- MYC- activated tumours 65 PAPER 3 MYCN/c-MYC-induced micrornas repress coding gene networks associated with poor outcome in MYCN/c-MYC-activated tumours. Mestdagh P, Fredlund E, Pattyn F, Schulte JH, Muth D, Vermeulen J, Kumps C, Schlierf S, De Preter K, Van Roy N, Noguera R, Laureys G, Schramm A, Eggert A, Westermann F, Speleman F, Vandesompele J. Oncogene Mar 4;29(9):
72 66 Oncogene (2010) 29, & 2010 Macmillan Publishers Limited All rights reserved /10 $ ONCOGENOMICS MYCN/c-MYC-induced micrornas repress coding gene networks associated with poor outcome in MYCN/c-MYC-activated tumors P Mestdagh 1, E Fredlund 1, F Pattyn 1, JH Schulte 2, D Muth 3, J Vermeulen 1, C Kumps 1, S Schlierf 2, K De Preter 1, N Van Roy 1, R Noguera 4, G Laureys 5, A Schramm 2, A Eggert 2, F Westermann 3, F Speleman 1 and J Vandesompele 1 ONCOGENOMICS 1 Center for Medical Genetics, Ghent University Hospital, Ghent, Belgium; 2 University Children s Hospital, Essen, Germany; 3 Department of Tumour Genetics, German Cancer Research Center, Heidelberg, Germany; 4 Medical School of Valencia, Valencia, Spain and 5 Pediatric Oncology, Ghent University Hospital, Ghent, Belgium Increased activity of MYC protein-family members is a common feature in many cancers. Using neuroblastoma as a tumor model, we established a microrna (mirna) signature for activated MYCN/c-MYC signaling in two independent primary neuroblastoma tumor cohorts and provide evidence that c-myc and MYCN have overlapping functions. On the basis of an integrated approach including mirna and messenger RNA (mrna) gene expression data we show that mirna activation contributes to widespread mrna repression, both in c-mycand MYCN-activated tumors. c-myc/mycn-induced mirna activation was shown to be dependent on c-myc/ MYCN promoter binding as evidenced by chromatin immunoprecipitation. Finally, we show that pathways, repressed through c-myc/mycn mirna activation, are highly correlated to tumor aggressiveness and are conserved across different tumor entities suggesting that c-myc/ MYCN activate a core set of mirnas for cooperative repression of common transcriptional programs related to disease aggressiveness. Our results uncover a widespread correlation between mirna activation and c-myc/ MYCN-mediated coding gene expression modulation and further substantiate the overlapping functions of c-myc and MYCN in the process of tumorigenesis. Oncogene (2010) 29, ; doi: /onc ; published online 30 November 2009 Keywords: MYCN; c-myc; microrna; neuroblastoma Introduction Activated signaling of MYC gene-family members (c-myc, MYCN, MYCL) is a hallmark of many cancer types and contributes to tumorigenesis by promoting cell growth, metastasis, angiogenesis and genomic Correspondence: Professor J Vandesompele, Center for Medical Genetics, Ghent University Hospital, MRB, De Pintelaan 185, B-9000 Ghent, Belgium. [email protected] Received 17 July 2009; revised 20 October 2009; accepted 25 October 2009; published online 30 November 2009 instability (Adhikary and Eilers, 2005). MYC genes can activate and repress transcription of target genes through distinct mechanisms. Transcriptional activation is well understood and depends on the binding of MYC to its consensus DNA recognition sequence 5 0 -CACG TG-3 0 or E-box. In contrast, the mechanism of transcriptional repression is more obscure and seems independent of E-box binding. One mechanism relies on the binding of MYC with the cofactor Miz-1, which tethers MYC to promoters (Kleine-Kohlbrecher et al., 2006) whereas another involves the induction of gene silencing through MYC-mediated promoter hypermethylation (Gartel, 2006). Recently, the MYCN/c-MYC transcriptional network has been shown to also include micrornas (mirnas). These small non-coding RNAs have an effect on virtually every aspect of tumorigenesis and function as negative regulators of messenger RNA (mrna) levels and translation (Bartel, 2009). Some mirnas, such as those belonging to the oncogenic mir cluster, have been shown to be important players in MYCN/c-MYC signaling (O Donnell et al., 2005; Dews et al., 2006; Chang et al., 2008; Fontana et al., 2008; Schulte et al., 2008; Northcott et al., 2009). However, a more general insight in the relationship between mirnas and mrnas within the MYCN/c- MYC transcriptional network remains to be examined. Neuroblastoma qualifies as an excellent model system to study the MYCN/c-MYC transcriptional network. Activated MYCN/c-MYC signaling is a hallmark of poor prognosis for neuroblastoma tumors (Fredlund et al., 2008) and can be caused either by MYCN amplification (Seeger et al., 1985) or increased c-myc expression in stage 4 MYCN non-amplified neuroblastoma tumors (Westermann et al., 2008). This increase in MYCN/c-MYC expression results in the activation of coding target genes related to poor patient prognosis independently of MYCN amplification (Fredlund et al., 2008; Westermann et al., 2008). In this study, we identified a comprehensive mirna signature induced by increased MYCN/c-MYC signaling in neuroblastoma and show that MYCN and c-myc have overlapping functions in the induction of this mirna signature. Most importantly, we provide
73 67 evidence that MYCN/c-MYC-activated mirnas are correlated to widespread mrna downregulation pointing at MYCN/c-MYC-regulated gene expression modulation beyond transcriptional control. Transcriptional programs that are affected in this way underlie distinct prognostic subgroups of both neuroblastoma as well as other tumor entities with activated MYCN/c-MYC signaling. Results MYCN/c-MYC microrna signature delineation For the identification of mirnas differentially expressed between MYCN-amplified (MNA) on the one hand, and MYCN single-copy low-risk (SL) or highrisk (SH) tumors on the other (see Supplementary Material for definition), two independent unpublished patient cohorts (n ¼ 56 and n ¼ 39) were analyzed for a total of 430 mirnas. When comparing MNA with SL MYCN/c-MYC microrna regulation P Mestdagh et al tumors, we found 49 mirnas significantly differentially expressed in both cohorts. Comparing MNA with SH resulted in 12 differentially expressed mirnas. In total, we identified 50 unique mirnas (16 upregulated and 34 downregulated) differentially expressed between MNA and MYCN single-copy tumors in the two independent data sets (Supplementary Table S1). Supporting our result, differential expression has earlier been shown for several of the listed mirnas in other cancer entities with increased MYCN/c-MYC signaling (Supplementary Table S2). Cluster analysis based on this 50 mirna signature on both cohorts distinguishes three major clusters with a clear separation between MNA and MYCN single-copy tumors (Figure 1a, Supplementary Figure S1). Interestingly, the 50 mirna signature also separates SH from SL tumors despite the fact that differential mirna expression between these groups was not selected for. This additional separation suggests that MYCN/c-MYC signaling rather than MYCN levels alone underlies the differential expression of this mirna signature. Indeed, c-myc expression is 1395 Density Color Key and Density Plot MNA SH SL SL cluster Value subgroup OS EFS OS (%) SH cluster MNA cluster follow-up time (years) EFS (%) SL cluster SH cluster MNA cluster follow-up time (years) Figure 1 A MYCN/c-MYC microrna (mirna) signature delineates prognostic neuroblastoma subgroups. (a) Hierarchical clustering of 95 primary neuroblastoma tumors based on the expression of the MYCN/c-MYC mirna signature. MYCN-amplified (MNA), MYCN single-copy low-risk (SL) and MYCN single-copy high-risk (SH) tumors are indicated in black, light gray and dark gray, respectively. (b, c) Kaplan Meier analysis for overall (OS) and event-free (EFS) survival of patients according to the three main clusters in (a). Oncogene
74 MYCN/c-MYC microrna regulation P Mestdagh et al significantly increased in SH tumors (Supplementary Figure S2), confirming the inverse relationship between MYCN and c-myc (Vandesompele et al., 2003), and has been shown to drive MYCN/c-MYC mrna target gene expression in these tumors (Westermann et al., 2008). In addition, other genetic factors than increased MYCN/c-MYC activity could also contribute to the observed mirna expression differences. MNA tumors are frequently associated with 1p deletions (80.6% in MNA subgroup and 17.9% in SH subgroup) while SH tumors are associated with 11q deletions (89.3% in SH subgroup and 32.3% in MNA subgroup). Factors in either of these regions affecting mirna expression can contribute to the observed signature. The three main clusters, defined by the mirna signature, significantly correlate to overall (Po0.001) and event-free (Po0.001) patient survival confirming its ability to define subgroups with differential prognosis (Figures 1b and c). We identified six typical patterns of expression within the 50 mirna signature when comparing MNA, SH and SL tumors (Figure 2). Four of these are linked to increased MYCN/c-MYC signaling with mirna expression being intermediate in SH tumors (Figures 2a and b) or either high (or low) in both MNA and SH tumors (Figures 2c and d). The remaining two patterns represent mirnas with a differential expression that is restricted to MNA tumors alone (Figures 2e and f). These mirna expression patterns are similar to those that have been described for protein coding MYCN/c-MYC target genes (Westermann et al., 2008). Further inspection of the classification of MNA, SH and SL tumors revealed some misclassified cases. Two MYCN single-copy tumors cluster within the MNA cluster. For one of these tumors, fluorescence in situ hybridization analysis revealed low level gain of c-myc SL SH MNA SL SH MNA SL SH MNA hsa-mir-92a hsa-mir-15b hsa-mir-181a SL SH MNA SL SH MNA SL SH MNA hsa-mir-628-3p hsa-mir-128a hsa-mir-184 Figure 2 MicroRNA (mirna) expression patterns. Overview of the six main mirna expression patterns (a f) observed in MYCNamplified (MNA), MYCN single-copy low-risk (SL) and MYCN single-copy high-risk (SH) tumors. Expression patterns are represented schematically using vertical bars. The height of each bar reflects the relative mirna expression in each group. Black, gray and white bars represent significant expression differences. For each pattern, one representative mirna is shown. (a) Upregulated in MNA, intermediate in SH, (b) downregulated in MNA, intermediate in SH, (c) upregulated in MNA and SH, (d) downregulated in MNA and SH, (e) upregulated in MNA, (f) downregulated in MNA. Oncogene
75 69 in combination with c-myc translocation (Supplementary Figure S3A). Fluorescence in situ hybridization results for the t(8;14) translocation described in Burkitt lymphoma were negative suggesting a translocation partner other than chromosome 14 (data not shown). These aberrations at the c-myc locus ultimately result in an increased expression of c-myc (12.8-fold) and of key protein coding c-myc target genes (Supplementary Figure S3B and C). To our knowledge, this is the first report of a primary neuroblastoma tumor with c-myc translocation. In addition, this tumor presented with genetic abnormalities (such as 1p deletion and 17q gain, data not shown) typically found in MNA tumors thus supporting the notion that MYCN and c-myc have overlapping functions in vivo and in the process of tumorigenesis (Malynn et al., 2000). c-myc translocation provides an additional mechanism for neuroblastoma cells to obtain high level c-myc expression, together with the earlier reported c-myc amplification in SJNB-12 and MP-N-TS neuroblastoma cells (Saito- Ohara et al., 2003; Van Roy et al., 2006). The second MYCN single-copy tumor did not show any aberrations at the c-myc locus suggesting that other mechanisms are responsible for increased activity of MYCN/c-MYC in this tumor. Both of the patients died of disease. Finally, eight MNA tumors clustered outside the MNA cluster. All but one grouped within the SH cluster, presumably because of the overlapping mirna expression patterns between MNA and SH tumors (Figure 2). To further examine the mirna signature, we also examined a panel of MNA (n ¼ 5), MYCN single-copy (n ¼ 5) and c-myc-amplified (n ¼ 1) neuroblastoma cell lines. For several mirnas, expression levels in the c-myc-amplified cell line SJNB-12 were similar to those in the MNA cell lines (Supplementary Figure S4). These data are in line with the mirna expression pattern that was observed in the c-myc- translocated primary neuroblastoma tumor and further support the apparent overlapping functions for MYCN and c-myc with regard to mirna regulation in these cancer cells. MYCN/c-MYC binds to microrna promoters To confirm that the 16 upregulated mirnas are indeed responsive to MYCN/c-MYC, we profiled their expression in two independent MYCN model systems. On MYCN activation, 10 out of 16 upregulated mirnas showed a 41.5-fold induction in expression in at least one model system whereas 6 out of 16 were induced in two model systems (Supplementary Figure S5). Varying results between the between model systems and primary tumor cells might be due to cell-specific differences in target genes and cell-dependent variations in the composition of MYCN transcriptional complexes (Cappellen et al., 2007). Differences between the model systems could be because of the level of induction of the MYCN gene, culture effects or treatment-related events. To analyze mirna regulation by MYCN/c-MYC, we assessed binding of MYCN or c-myc using chromatin immunoprecipitation (ChIP)-chip to genomic regions of MYCN/c-MYC microrna regulation P Mestdagh et al the upregulated mirnas. Besides binding of MYCN or c-myc, we also defined the epigenetic marker status of these genomic regions in six neuroblastoma cell lines. We used H3K4me3 for active and H3K27me3 for repressed regions. In addition, H3K36me3 was used for transcript elongation. For 10 out of 11 mirnas that are broadly covered by the array probe set, we observed MYCN or c-myc binding as well as an epigenetic marker state that is in line with transcription of the respective mirnas (Table 1). For two mirnas (mir- 601 and mir-610) that are not covered by the array probe set, we observed binding of MYCN and c-myc to the promoter regions of the hosting genes (DENN- D1A and KIF18A). Transcription of the genes in neuroblastoma cell lines indirectly suggests that the mirnas within these genes are also transcribed (data not shown). Our results confirm c-myc/mycn binding to the promoters of mir (O Donnell et al., 2005; Fontana et al., 2008; Mestdagh et al., 2009), mir- 181a (Mestdagh et al., 2009), and mir-9 (L Ma et al., submitted, personal communication) and suggest that the promoters of mir-15b, mir-130a and mir-214 are also bound and activated by MYCN/c-MYC (Supplementary Figure S6). These results are in line with the hypothesis that, for the majority of the mirnas, increased expression in tumors with activated MYCN/ c-myc is a direct effect of MYCN/c-MYC binding and transactivation. A microrna target identification strategy For high-throughput mirna target identification, we devised a strategy based on the integration of a unique mrna (obtained through Affymetrix exon gene expression analysis) and mirna expression data set together with mirna target recognition characteristics. In a subset of 40 tumors, we calculated correlations between the expression of each of the 50 mirnas and B mrnas. Candidate target mrnas were defined as being significantly negatively correlated to the expression of the mirna (Spearman s rank, Benjamini Hochberg multiple testing correction) with a rho value below 0.5 and with the occurrence of at least one 3 0 UTR seed. Three different seed matches were considered: at least one 7mer seed (7mer-A1 or 7merm8), at least one 8mer seed or at least one conserved 7 or 8mer seed. To confirm that these criteria preferentially select mirna target genes, we established a new model system with tetracycline inducible mir expression in the SHEP neuroblastoma cell line (SHEP-TR-miR ) (Supplementary Figure S7). Expression of the mir cluster is activated by MYCN/c-MYC and five mirnas belonging to this cluster (mir-18a, mir- 18a*, mir-19a, mir-20a and mir-92a) are among the 16 upregulated mirnas in our signature. For three of these (mir-19a, mir-20a and mir-92a) we can define candidate mirna targets using our selection strategy on the mrna and mirna data from the 40 tumors. We then compared differentially expressed mrnas in treated and untreated SHEP-TR-miR cells with our predicted targets for mir-19a, mir-20a and 1397 Oncogene
76 Table 1 MYCN/c-MYC microrna regulation P Mestdagh et al MYCN/c-MYC binding to mirna promoters No. mirna Chromosomal position Hosting gene/cluster Binding of MYCN/c-MYC a Binding of epigenetic marks a 1 Hsa-mir-20a 13: Hsa-mir cluster Binding Actively transcribed 2 Hsa-mir-20b 23: Has-mir a cluster No binding 3 Hsa-mir-92 13: Hsa-mir cluster Binding Actively transcribed 4 Hsa-mir-19a 13: Hsa-mir cluster Binding Actively transcribed 5 Hsa-mir-18a 13: Hsa-mir cluster Binding actively transcribed 6 Hsa-mir-18a* 13: Hsa-mir cluster Binding Actively transcribed 7 Hsa-mir-15b 3: SMC4 Binding Actively transcribed 8 Hsa-mir-9 15: Binding Actively transcribed 9 Hsa-mir-130a 11: Binding Actively transcribed 10 Hsa-mir-181a 1: Binding Actively transcribed 9: Hsa-mir-214 1: DNM3 and Hsa-mir-199a-2 Binding Actively transcribed 12 Hsa-mir : NA NA 13 Hsa-mir-601 9: DENND1A Binding to gene promoter Gene actively transcribed 14 Hsa-mir-572 4: NA NA 15 Hsa-mir : KIF18A Binding to gene promoter Gene actively transcribed 16 Hsa-mir-526b* 19: NA NA Abbreviations: ChIP, chromatin immunoprecipitation; mirna, microrna. The * symbol is part of the mirna identifier and denotes the star sequence of the mirna. a As determined by ChIP-chip, summary of 6 neuroblastoma cell lines (SH-EP, SJ-NB12, WAC2, SY5Y, Kelly, IMR5), NA not determined because of poor probe coverage. mir-92a using gene set enrichment analysis (Subramanian et al., 2005). Three custom gene set enrichment analysis gene lists were established based on the different seed matches (7mer, 8mer and conserved 7 or 8mer) of mir-19a, mir-20a and mir-92a. Interestingly, two gene lists (8mer, P ¼ 0.01 and conserved 7 or 8mer, P ¼ 0.005) showed significant enrichment among the mrnas that were downregulated on mir induction in the SHEP-TR-mir cells hereby validating our selection strategy (Supplementary Figure S8). The fact that only predicted targets with 8mer seeds and conserved seeds are enriched confirms the higher efficacy of these seeds (Baek et al., 2008). In addition, several predicted targets defined by this strategy, such as TGFBR2, ATXN1 and HIPK3, have already been reported as direct targets of mir-20a, mir-19a and mir-92a, respectively (Volinia et al., 2006; Landais et al., 2007; Lee et al., 2008). We also compared our target predictions for mir-20a in neuroblastoma to a set of validated mir-20a targets obtained by ribonucleoprotein immunoprecipitation-gene Chip in Hodgkin lymphoma cells (Tan et al., 2009). From the 37 mir- 20a 8mer targets that we identified, 14 were reported by Tan et al. (2009) (Fisher s exact test, Po0.001). These results indicate that, although some of the predicted targets could be indirect, our mirna target identification strategy definitely selects a substantial number of direct mirna target genes. mirna activation as a mechanism for MYCN/ c-myc-induced mrna downregulation We next set out the search for target mrnas downstream of the MYCN/c-MYC mirna signature. If a mirna affects target gene expression, the negatively correlated mrnas should be enriched for 3 0 UTR mirna seeds. Strikingly, 3 0 UTR seed enrichment was found for almost all upregulated mirnas (14 out of 16) but not for the downregulated mirnas (1 out of 34) (Fisher s exact test, Po0.0001). For the upregulated mirnas, cumulative distribution plots show a significant difference between the Spearman s rank rho value distribution (representing mrna:mirna correlation) of mrnas with a 3 0 UTR seed compared with mrnas without a 3 0 UTR seed (Kolmogorov Smirnov, Po0.001) (Figure 3a). For the downregulated mirnas this is not the case (Figure 3b). These findings show that MYCN/c-MYC-activated mirnas rather than MYCN/c-MYC repressed mirnas have a widespread effect on differential mrna target gene expression. This observation therefore suggests that activated mirna expression could serve as a mechanism for MYCN/ c-myc-induced mrna repression. To further support this hypothesis, we established a core set of predicted mirna targets using our target identification strategy and compared the number of predicted targets between the up- and downregulated mirnas. On average, upregulated mirnas have four times more predicted targets when considering 7mer seeds and 6 to 18 times more predicted targets when considering 8mer or conserved seeds respectively (Figure 3c). This huge discrepancy in predicted target genes between up- and downregulated mirnas thus supports the notion that MYCN/c-MYC-activated mirnas predominantly drive differential gene expression in high-risk neuroblastomas. Moreover, it confirms that activated mirna expression is correlated to widespread mrna repression. In line with this, MYCN and c-myc downregulated genes should contain 3 0 UTR seeds for MYCN/c-MYC-activated mirnas. To evaluate this, known MYCN and c-myc down- and upregulated genes were extracted from the MYCNot database ( and the MYC target gene database, (Zeller et al., 2003) respectively. The total occurrence of seeds from all 16 upregulated mirnas were calculated and compared between down- and Oncogene
77 MYCN/c-MYC microrna regulation P Mestdagh et al Cumulative distribution (%) no seed mir-214 seed Cumulative distribution (%) no seed mir-628-3p seed Spearman s rank rho-value Spearman s rank rho-value Average number of targets per mirna upregulated mirnas 7mer seed 8mer seed conserved seed downregulated mirnas MYCNot DN MYCNot UP MYCdb DN MYCdb UP fraction of total (%) p < p < 0.01 no seed 7mer seed Figure 3 MicroRNA (mirna) seed enrichment and target identification. (a, b) Cumulative distribution of Spearman s rank rho values, representing mrna:mirna correlation, for mrnas with no seed (dark gray) and mrnas with a seed (light gray) shown for a representative MYCN/c-MYC-activated mirna and MYCN/c-MYC repressed mirna, respectively. (c) Average number of identified targets for up- and downregulated mirnas when considering 7mer, 8mer or conserved 3 0 UTR seeds. (d) Fraction of genes listed in the MYCNot database and the MYC target gene database (MYCdb) containing a 3 0 UTR seed. Results for upregulated (UP) and downregulated (DN) genes are listed as separate bars. upregulated genes. As expected, MYCN downregulated genes were significantly enriched for seeds from the upregulated mirnas (Fisher s exact test, Po0.001) (Figure 3d) confirming the observed correlation between mirna activation and MYCN target gene repression. Strikingly, this enrichment was also apparent for c- MYC downregulated genes (Fisher s exact test, Po0.01) (Figure 3d). Although entries in MYCNot mainly represent MYCN targets in neuroblastoma, the MYC target gene database contains c-myc targets across multiple other tumor entities. Despite the fact that the upregulated mirnas were selected from primary neuroblastoma tumors, their seed signature is also manifested in other tumors with activated MYCN/ c-myc signaling, underlining the general relevance of this MYCN/c-MYC signature for tumor biology. MYCN/c-MYC-activated micrornas act in concert The entire network of MYCN/c-MYC upregulated mirnas and their predicted target mrnas are significantly enriched for genes listed in the neuroblastoma gene server NBGS (Fisher s exact test, Po0.001), a database containing genes reported as differentially expressed in neuroblastoma tumors ( ugent.be/nbgs Center for Medical Genetics, Ghent, Belgium). In addition, it reveals that a significant number of mrnas are putatively regulated by multiple mirnas (Figure 4). Over 30% of the mrna targets are predicted to be under the regulation of two or more mirnas indicating a concerted mode of action of mirnas toward their predicted target genes. The significance of this cooperative regulation between mirnas and mrnas in the network were evaluated Oncogene
78 MYCN/c-MYC microrna regulation P Mestdagh et al Figure 4 Interaction network of MYCN/c-MYC-activated micro- RNAs (mirnas) and target mrnas. MYCN/c-MYC-activated mirnas (red triangles) are connected to their predicted target mrnas (dots). mrnas that are targeted by multiple mirnas from within the network are indicated in black, mrnas targeted by only one mirna in gray. Only mirnas with at least three mrna targets are shown. A full colour version of this figure is available at the Oncogene journal online. with respect to repeated sampling of an equally sized list of randomly selected mrnas or mirnas (Supplementary Figure S9 and Supplementary Material). We consistently observed a higher degree of cooperative regulation between MYCN-activated mirnas and their targets as compared with random selections of mrnas/ mirnas (Kolmogorov Smirnov, Po0.0001) suggesting that cooperative regulation is a likely feature within the network of MYCN-activated mirnas. Cooperative regulation of gene expression by co-expressed mirnas has been suggested before and could serve as a mechanism to fine tune gene expression (Sampson et al., 2007). Alternatively, it might be that one binding site is insufficient for proper repression. An increasing number of 3 0 UTR binding sites have indeed been shown to be more effective toward target regulation (Selbach et al., 2008). MicroRNA target gene expression correlates to patient survival As MYCN/c-MYC-activated mirnas appear essential in mediating MYCN/c-MYC-induced transcriptional repression, we asked whether the predicted mrna targets of these mirnas would have any prognostic value. The combined activity of the predicted 8mer mirna targets (n ¼ 193) was evaluated using a rank-based pathway score in two independent neuroblastoma microarray data sets (Oberthuer et al., 2006; Wang et al., 2006). To assess the effect on patient outcome, samples were divided into quartiles according to the pathway activity score (Fredlund et al., 2008). Kaplan Meier analysis of the Oberthuer data based on these quartiles revealed a significant correlation to both event-free (EFS, Po0.001) and overall survival (OS, Po0.001) (Figures 5a and b). Patients in the first quartile, representing tumors with the lowest expression of predicted 8mer mirna targets, have a particular poor prognosis as compared with those in the fourth quartile. Of interest, 48% of the tumors in the first quartile (n ¼ 66) were MYCN single-copy confirming that MYCN/c-MYC signaling rather than MYCN amplification underlies the poor outcome of these patients (Fredlund et al., 2008; Westermann et al., 2008). Moreover, 8mer mirna-mediated target repression but not MYCN amplification status was an independent factor in a multivariate Cox regression model for EFS and OS (Supplementary Table S3). In the Wang data set, Kaplan Meier analysis confirms the observed correlation to OS (Po0.001) and EFS (Po0.001) (Figures 5c and d). In this study, 8mer mirna target signaling was no independent factor for OS but was for EFS (Supplementary Table S3). A possible explanation for this discrepancy is that the Wang data set is highly biased toward MNA high-risk patients. Taken together, these data clearly show that the process of mrna repression, mediated by MYCN/c-MYC upregulated mirnas, are highly correlated to tumor aggressiveness and ultimately patient survival. MYCN/c-MYC-activated micrornas repress pathways affecting patient survival across tumor types To gain more insight in the genetic programs underlying MYCN/c-MYC-activated mirna signaling, we dissected the list of predicted 8mer and conserved targets into pathways using the Ingenuity Pathway Analysis software (Supplementary Table S4). The analyses showed that the largest fraction of predicted conserved targets appeared to be implicated in cell death and cancer-related processes and these genes were comparatively lower expressed in high-risk as compared with low-risk neuroblastoma tumors (data not shown). Other pathways known to be involved in neuroblastoma, such as CREB signaling (Jiang et al., 2008), CNTF signaling (Peterson and Bogenmann, 2004) and integrin signaling were also identified. Integrin signaling is of particular interest as decreased expression of integrins selectively enhances neuroblastoma survival and metastasis (Stupack et al., 2006). Both integrin receptors and downstream signaling molecules were among the predicted 7mer and 8mer targets of the MYCN/c-MYCactivated mirnas. In addition, CAV1, a scaffolding protein linking integrin subunits to the tyrosine kinase FYN, is a direct MYC target (Park et al., 2001). Together, these findings suggest that integrin signaling is repressed by increased MYCN/c-MYC signaling. To assess whether integrin signaling intensity reflects neuroblastoma patient survival, a pathway activity score was calculated using all predicted 7mer targets from the integrin signaling pathway (n ¼ 19). Kaplan Meier analysis of the Oberthuer and Wang data sets revealed a significant correlation to OS (Oberthuer: Po0.001, Wang: Po0.01) and EFS (Oberthuer: Po0.001, Wang: Oncogene
79 73 MYCN/c-MYC microrna regulation P Mestdagh et al 100 Oberthuer OS 100 Oberthuer EFS OS (%) EFS (%) follow-up time (years) p < p < follow-up time (years) 100 Wang OS 100 Wang EFS OS (%) EFS (%) p < follow-up time (years) 0 p < follow-up time (years) 0% 25% 25% 50% 50% 75%% 75% 100% Figure 5 MicroRNA (mirna) target activity correlates to patient outcome in neuroblastoma. Pathway activity score, represented as quartiles, for predicted 8mer targets of MYCN/c-MYC-activated mirnas significantly correlates to overall (OS) (a, c) and event-free (EFS) (b, d) patient survival in two independent data sets (Oberthuer, Wang). The lowest and highest pathway activity scores are represented by the first quartile (yellow) and fourth quartile (red), respectively. Cox-regression P-values are listed. Po0.001) (Supplementary Figure S10). In a multivariate Cox regression model using integrin pathway activity quartiles and MYCN amplification status, low integrin signalingremainedpredictiveofbothosandefsinthe Oberthuer data set and of EFS in the Wang data set (data not shown). In summary, these results suggest that mirna controlled regulation of specific groups of mrnas could serve as an additional mechanism of MYCN/c-MYCinduced oncogenicity. As integrin signaling is (in part) MYCN/c-MYC regulated, we evaluated whether it would reflect patient survival in other tumor entities with increased MYCN/c- MYC signaling. One such entity is diffuse large B-cell lymphoma. Approximately, 15% of diffuse large B-cell lymphomas have rearrangements at the c-myc locus resulting in c-myc overexpression and poor patient survival (Kramer et al., 1998; Chang et al., 2000). The integrin signaling pathway activity score was calculated for a microarray data set of 255 diffuse large B-cell lymphomas (Lenz et al., 2008). In keeping with our observation in neuroblastoma, Kaplan Meier analyses indicate that integrin signaling is proportionally correlated to overall patient survival (Po0.001) (Supplementary Figure S11). These results indicate that the integrin signaling pathway, which is negatively correlated to MYCN/c-MYC-activated mirnas, is conserved between tumor types. Not only do c-myc and MYCN activate a common set of mirnas, these mirnas appear to signal through the same pathways in two entirely different tumor cell types, lending support to a functional redundancy between the two transcription factors. Discussion We have identified a mirna signature representative for MYCN/c-MYC signaling in neuroblastoma tumors. Using ChIP-chip, we have shown binding of MYCN to the promoter region of several mirnas suggesting a direct role for MYCN in the transcriptional regulation of these mirnas. Additional experiments that assess reduced MYCN binding in cells with low MYCN expression are warranted to fully validate these findings. Our results further suggest that c-myc and MYCN have overlapping functions for the induction of this mirna signature. The functional overlap between MYCN and c-myc is further substantiated by the identification of a c-myc-translocated neuroblastoma tumor with a mrna, mirna and genomic profile that is typical for a MYCN-amplified tumor. It is also perfectly in line with previous reports on mirnas commonly regulated by c-myc and MYCN (Chen and Stallings, 2007; Chang et al., 2008; Sander et al., 2008; Schulte et al., 2008; Sun et al., 2008; Northcott et al., 2009) Oncogene
80 MYCN/c-MYC microrna regulation P Mestdagh et al and with the observation that MYCN can functionally replace c-myc in murine development (Malynn et al., 2000). Whether c-myc is capable of replacing MYCN remains to be determined but could further corroborate these findings. MYCN appears to be a weak transcription factor as mirna expression levels in MNA and SH tumors are equally high (or low) despite the fact that MYCN expression in MNA tumors is substantially higher than c-myc expression in SH tumors. To identify candidate mirna target genes, we used an integrative approach based on negative correlation analysis between mirna and mrna expression in primary tumors, in combination with 3 0 UTR seed occurrence and a cellular model system. We showed that MYCN/c-MYCactivated mirnas are correlated to widespread transcriptional repression whereas mirnas that are downregulated in MYCN/c-MYC-activated tumors were not associated with transcriptional activation. This observation does not imply that mrnas that do have a seed for downregulated mirnas are irrelevant. Chang et al. have shown that c- MYC-induced mirna downregulation can have a profound effect on tumorigenesis (Chang et al., 2008). In line with this, we observed a significant correlation between the predicted targets of the downregulated mirnas and patient survival (data not shown) suggesting that mirna downregulation is important in tumorigenesis. The mechanisms by which c-myc and MYCN induce transcriptional repression are multiple and include binding to Miz-1 and induction of promoter hypermethylation (Gartel, 2006; Kleine-Kohlbrecher et al., 2006). Our results now suggest that MYCN/c-MYC-induced mirna activation also contributes to coding gene repression. The observed correlations were experimentally verified in neuroblastoma tumors and in silico for published c-myc and MYCN downregulated genes in different tumor entities suggesting a non-random and conserved feature of the MYCN/c-MYC network affecting specific collections of mrnas. MYCN/c- MYC-induced mirna activation has been shown to repress the expression of few coding genes, such as CDKN1A, BCL2L11 and E2F1 (O Donnell et al., 2005; Fontana et al., 2008) whereas our findings suggest a widespread transcriptional repression. Although we have shown that the observed mrna downregulation is mirna dependent in a cellular model system, further experiments are needed to establish direct interactions between the MYCN/c-MYC-activated mirnas and the repressed mrnas. Pathways that are repressed in this way were shown to be of clinical importance in different tumor entities with increased MYCN/c-MYC activity again confirming that MYC family members have overlapping functions that contribute to tumor aggressiveness. It also confirms that assessing entire signaling pathways, rather than individual genes, is highly informative with respect to cancer outcome prediction (Bild et al., 2006; Watters and Roberts, 2006; Liu and Ringner, 2007). For neuroblastoma in particular, combined analysis of multiple signaling networks has been successful and might benefit from the addition of other pathways, such as integrin signaling, to further increase prediction sensitivity (Fredlund et al., 2008). In addition, the observed similarities in mirna expression between MNA and SH neuroblastoma tumors suggest that both tumor subgroups share a number of pathways deregulated through MYCN/c-MYC-mediated mirna modulation. The identification of such pathways might prove useful in the search for novel therapeutic targets. In conclusion, we uncovered a widespread correlation between mirna activation and c-myc/mycnmediated coding gene expression modulation and further substantiate the overlapping functions of c-myc and MYCN in the process of tumorigenesis. Materials and methods Patient samples and cell lines A total of 95 primary neuroblastoma tumor samples were collected at the Ghent University Hospital (Ghent, Belgium), the University Children s Hospital, Essen (Essen, Germany) and the Medical School of Valencia (Valencia, Spain) before therapeutic treatment. Patients were staged according to the International Neuroblastoma Staging System. Informed consent was obtained from the patients relatives. Details on prognostic subgroups and neuroblastoma cell lines are listed in the Supplementary Material. MicroRNA expression profiling Total RNA was isolated using the mirnaeasy kit (Qiagen, Valencia, CA, USA) according to the manufacturer s instructions. MiRNA expression profiling and data normalization were performed as described earlier (Mestdagh et al., 2008, 2009). Messenger RNA expression profiling Total RNA was isolated from tumor samples and was hybridized to Human Exon 1.0 ST array (Affymetrix, Santa Clara, CA, USA) at the microarray facility of the University Hospital of Essen according to the manufacturer s protocol. Total RNA from tetracycline treated and untreated SHEP- TR-miR cells were hybridized to Affy-Hu-Gene1.0ST oligonucleotide chips (Affymetrix) at the microarray facility of the Flanders Institute for Biotechnology (Leuven, Belgium). Fluorescence in situ hybridization Fluorescence in situ hybridization was performed according to Van Roy et al. (1994). The following probes were used: LSI MYC dual color, break-apart rearrangement probe and LSI IGH/MYC, CEP8 tri-color, dual fusion translocation probe (Abbott Molecular Products, Des Plaines, IL, USA). Chromatin immunoprecipitation Chromatin immunoprecipitation was performed as described earlier using 10 mg of MYCN or c-myc antibodies (Westermann et al., 2008). For a detailed description see Supplementary Material. Statistics Details on statistical procedures are described in Supplementary Material. Conflict of interest The authors declare no conflict of interest. Oncogene
81 75 Acknowledgements We acknowledge Q Wang and J Maris for providing the neuroblastoma microarray data set. This research was funded by the Gent University Research Fund (BOF 01D31406 to PM, BOF 01F07207 to FP, BOF 01Z09407 to J Vandesompele), the Fondation pour la recherche Nuovo-Soldati MYCN/c-MYC microrna regulation P Mestdagh et al (J Vermeulen), RD06/0020/0102 from RTICC/ISCIII to RN, the Fund for Scientific Research (grant number: G and ), the Belgian Kid s Fund and the Stichting tegen Kanker. KDP is a post-doctoral researcher with the Fund for Scientific Research-Flanders. We acknowledge the support of the European Community under the FP6 (project: STREP: EET-pipeline, number: ) References Adhikary S, Eilers M. (2005). Transcriptional regulation and transformation by Myc proteins. Nat Rev Mol Cell Biol 6: Baek D, Villen J, Shin C, Camargo FD, Gygi SP, Bartel DP. (2008). The impact of micrornas on protein output. Nature 455: Bartel DP. (2009). MicroRNAs: target recognition and regulatory functions. Cell 136: Bild AH, Yao G, Chang JT, Wang Q, Potti A, Chasse D et al. (2006). Oncogenic pathway signatures in human cancers as a guide to targeted therapies. Nature 439: Cappellen D, Schlange T, Bauer M, Maurer F, Hynes NE. (2007). Novel c-myc target genes mediate differential effects on cell proliferation and migration. EMBO Rep 8: Chang CC, Liu YC, Cleveland RP, Perkins SL. (2000). Expression of c-myc and p53 correlates with clinical outcome in diffuse large B- cell lymphomas. Am J Clin Pathol 113: Chang TC, Yu D, Lee YS, Wentzel EA, Arking DE, West KM et al. (2008). Widespread microrna repression by Myc contributes to tumorigenesis. Nat Genet 40: Chen Y, Stallings RL. (2007). Differential patterns of microrna expression in neuroblastoma are correlated with prognosis, differentiation, and apoptosis. Cancer Res 67: Dews M, Homayouni A, Yu D, Murphy D, Sevignani C, Wentzel E et al. (2006). Augmentation of tumor angiogenesis by a Mycactivated microrna cluster. Nat Genet 38: Fontana L, Fiori ME, Albini S, Cifaldi L, Giovinazzi S, Forloni M et al. (2008). Antagomir-17-5p abolishes the growth of therapyresistant neuroblastoma through p21 and BIM. PLoS One 3: e2236. Fredlund E, Ringner M, Maris JM, Pahlman S. (2008). High Myc pathway activity and low stage of neuronal differentiation associate with poor outcome in neuroblastoma. Proc Natl Acad Sci USA 105: Gartel AL. (2006). A new mode of transcriptional repression by c-myc: methylation. Oncogene 25: Jiang M, Zhu K, Grenet J, Lahti JM. (2008). Retinoic acid induces caspase-8 transcription via phospho-creb and increases apoptotic responses to death stimuli in neuroblastoma cells. Biochim Biophys Acta 1783: Kleine-Kohlbrecher D, Adhikary S, Eilers M. (2006). Mechanisms of transcriptional repression by Myc. Curr Top Microbiol Immunol 302: Kramer MH, Hermans J, Wijburg E, Philippo K, Geelen E, van Krieken JH et al. (1998). Clinical relevance of BCL2, BCL6, and MYC rearrangements in diffuse large B-cell lymphoma. Blood 92: Landais S, Landry S, Legault P, Rassart E. (2007). Oncogenic potential of the mir cluster and its implication in human T-cell leukemia. Cancer Res 67: Lee Y, Samaco RC, Gatchel JR, Thaller C, Orr HT, Zoghbi HY. (2008). mir-19, mir-101 and mir-130 co-regulate ATXN1 levels to potentially modulate SCA1 pathogenesis. Nat Neurosci 11: Lenz G, Wright G, Dave SS, Xiao W, Powell J, Zhao H et al. (2008). Stromal gene signatures in large-b-cell lymphomas. N Engl J Med 359: Liu Y, Ringner M. (2007). Revealing signaling pathway deregulation by using gene expression signatures and regulatory motif analysis. Genome Biol 8: R77. Malynn BA, de Alboran IM, O Hagan RC, Bronson R, Davidson L, DePinho RA et al. (2000). N-myc can functionally replace c-myc in murine development, cellular growth, and differentiation. Genes Dev 14: Mestdagh P, Feys T, Bernard N, Guenther S, Chen C, Speleman F et al. (2008). High-throughput stem-loop RT-qPCR mirna expression profiling using minute amounts of input RNA. Nucleic Acids Res 36: e143. Mestdagh P, Van Vlierberghe P, De Weer A, Muth D, Westermann F, Speleman F et al. (2009). A novel and universal method for microrna RT-qPCR data normalization. Genome Biol 10: R64. Northcott PA, Fernandez LA, Hagan JP, Ellison DW, Grajkowska W, Gillespie Y et al. (2009). The mir-17/92 polycistron is up-regulated in sonic hedgehog-driven medulloblastomas and induced by N-myc in sonic hedgehog-treated cerebellar neural precursors. Cancer Res 69: O Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT. (2005). c-myc-regulated micrornas modulate E2F1 expression. Nature 435: Oberthuer A, Berthold F, Warnat P, Hero B, Kahlert Y, Spitz R et al. (2006). Customized oligonucleotide microarray gene expression-based classification of neuroblastoma patients outperforms current clinical risk stratification. J Clin Oncol 24: Park DS, Razani B, Lasorella A, Schreiber-Agus N, Pestell RG, Iavarone A et al. (2001). Evidence that Myc isoforms transcriptionally repress caveolin-1 gene expression via an INR-dependent mechanism. Biochemistry 40: Peterson S, Bogenmann E. (2004). The RET and TRKA pathways collaborate to regulate neuroblastoma differentiation. Oncogene 23: Saito-Ohara F, Imoto I, Inoue J, Hosoi H, Nakagawara A, Sugimoto T et al. (2003). PPM1D is a potential target for 17q gain in neuroblastoma. Cancer Res 63: Sampson VB, Rong NH, Han J, Yang Q, Aris V, Soteropoulos P et al. (2007). MicroRNA let-7a down-regulates MYC and reverts MYC-induced growth in Burkitt lymphoma cells. Cancer Res 67: Sander S, Bullinger L, Klapproth K, Fiedler K, Kestler HA, Barth TF et al. (2008). MYC stimulates EZH2 expression by repression of its negative regulator mir-26a. Blood 112: Schulte JH, Horn S, Otto T, Samans B, Heukamp LC, Eilers UC et al. (2008). MYCN regulates oncogenic micrornas in neuroblastoma. Int J Cancer 122: Seeger RC, Brodeur GM, Sather H, Dalton A, Siegel SE, Wong KY et al. (1985). Association of multiple copies of the N-myc oncogene with rapid progression of neuroblastomas. N Engl J Med 313: Selbach M, Schwanhausser B, Thierfelder N, Fang Z, Khanin R, Rajewsky N. (2008). Widespread changes in protein synthesis induced by micrornas. Nature 455: Stupack DG, Teitz T, Potter MD, Mikolon D, Houghton PJ, Kidd VJ et al. (2006). Potentiation of neuroblastoma metastasis by loss of caspase-8. Nature 439: Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al. (2005). Gene set enrichment analysis: a Oncogene
82 MYCN/c-MYC microrna regulation P Mestdagh et al knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102: Sun Y, Wu J, Wu SH, Thakur A, Bollig A, Huang Y et al. (2008). Expression profile of micrornas in c-myc induced mouse mammary tumors. Breast Cancer Res Treat 118: Tan LP, Seinen E, Duns G, de Jong D, Sibon OC, Poppema S et al. (2009). A high throughput experimental approach to identify mirna targets in human cells. Nucleic Acids Res 37: e137. Van Roy N, Laureys G, Cheng NC, Willem P, Opdenakker G, Versteeg R et al. (1994). 1;17 translocations and other chromosome 17 rearrangements in human primary neuroblastoma tumors and cell lines. Genes Chromosomes Cancer 10: Van Roy N, Vandesompele J, Menten B, Nilsson H, De Smet E, Rocchi M et al. (2006). Translocation-excision-deletion-amplification mechanism leading to nonsyntenic coamplification of MYC and ATBF1. Genes Chromosomes Cancer 45: Vandesompele J, Edsjo A, De Preter K, Axelson H, Speleman F, Pahlman S. (2003). ID2 expression in neuroblastoma does not correlate to MYCN levels and lacks prognostic value. Oncogene 22: Volinia S, Calin GA, Liu CG, Ambs S, Cimmino A, Petrocca F et al. (2006). A microrna expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci USA 103: Wang Q, Diskin S, Rappaport E, Attiyeh E, Mosse Y, Shue D et al. (2006). Integrative genomics identifies distinct molecular classes of neuroblastoma and shows that multiple genes are targeted by regional alterations in DNA copy number. Cancer Res 66: Watters JW, Roberts CJ. (2006). Developing gene expression signatures of pathway deregulation in tumors. Mol Cancer Ther 5: Westermann F, Muth D, Benner A, Bauer T, Henrich KO, Oberthuer A et al. (2008). Distinct transcriptional MYCN/c-MYC activities are associated with spontaneous regression or malignant progression in neuroblastomas. Genome Biol 9: R150. Zeller KI, Jegga AG, Aronow BJ, O Donnell KA, Dang CV. (2003). An integrated database of genes responsive to the Myc oncogenic transcription factor: identification of direct genomic targets. Genome Biol 4: R69. Supplementary Information accompanies the paper on the Oncogene website ( Oncogene
83 Supplemental Data 77 Patient samples and cell lines Neuroblastoma tumour samples were divided into 3 major prognostic subgroups based on stage, age and MYCN amplification status. MYCN amplified tumours were classified as MYCN amplified (MNA) irrespective of stage. MYCN single copy tumours were subdivided into single copy low risk (SL) for stage 1, 2, and 3 tumours and single copy high risk (SH) for stage 4 tumours. Neuroblastoma cell lines were cultured in RPMI (Invitrogen, Carlsbad, CA, USA) supplemented with 15% fetal calf serum. SHEP- 21N (Lutz et al., 1996) and SHEP- MYCN- ER cells (Schulte et al., 2008) were treated with 2µg/ml tetracycline (Sigma- aldrich, St- Louis, MO, USA) and 200nM 4- hydroxy tamoxifen (Sigma- aldrich) respectively for 48h before harvesting. SHEP- TR- mir The human mir cluster together with flanking DNA was PCR- amplified using the primers 5'- ctaaatggacctcatatctttgag- 3' (forward) and 5'- gaaaacaagacaagatgtatttacac- 3' (reverse) and was then cloned into the vector pcr8/gw/topo (Invitrogen) and subcloned into vector pdest- 30 (Invitrogen) according to the manufacturer s recommendations. After subcloning, the mirna sequence was verified by direct sequencing. SHEP cells were stably transfected with the plasmid pcdna6 TR harboring a cdna encoding the Tet- Repressor (Invitrogen) and with pdest- 30 plasmid (Invitrogen) harboring the mir cluster or a cdna encoding GFP (control) by electroporation. After antibiotic selection single cells clones were obtained by limited dilution. SHEP- TR- mir cells were treated with 2 µg/ml tetracycline (Sigma- aldrich) for 48 hours before harvesting. Total RNA was reverse transcribed using a multiplex pool of stem- loop primers for mir mirnas and 3 endogenous controls (RNU19, RNU44, RNU66) (Applied Biosystems, Foster City, CA, USA) according to the manufacturer s instructions. Quantitative PCR reactions were performed on the 7900 HT (Applied Biosystems) and data was processed using the qbaseplus 1.2 software ( All mirnas of the mir cluster were induced upon tetracycline treatment (Figure S4). Statistics All statistical analysis were performed using R Bioconductor software or SPSS Differential gene expression was evaluated using the Mann- Whitney test followed by Benjamini- Hochberg multiple testing correction. Hierarchical clustering was performed with method Ward and distance Manhattan (Kerr et al., 2008). Clustering with alternative methods (average linkage) and distance measures (Euclidean) gave similar results. Multivariate Cox proportional hazards models were built using the Enter method. The mirna mrna interaction network was visualized using cytoscape software (Shannon et al., 2003).
84 78 mirna seed definition and selection Four different seed types were considered when selecting candidate mirna targets: 6mer, 7mer, 8mer and conserved seeds. The 6mer seed is the perfect match to the 6 nt mirna seed. The 7mer is either a 7mer- m8 (6mer with an additional match to nucleotide 8 of the mirna) or a 7mer- A1 (6mer augmented by an A at target position 1) while the 8mer is a 6mer flanked by an additional match to nucleotide 8 of the mirna and an A at target position 1 (Grimson et al., 2007). The 6mer, 7mer and 8mer seeds were identified using Perl scripting. 3 UTR sequences were taken from Baek et al. (Baek et al., 2008). Conserved seeds were defined as those 7mer or 8mer seeds that were also predicted by Targetscan 5.1 (Friedman et al., 2009). To determine mirna seed enrichment, 3 UTR seed occurrence (7mer- A1, 7mer- m8 and 8mer) was determined for the 2% most negatively correlated mrnas and compared to that of the remaining mrnas. Pathway activity score For survival analysis, rank- based pathway activity scores were calculated for each gene set, essentially as previously described (Fredlund et al., 2008). Samples (n) were ranked according to the expression level of each gene within the gene set and rank scores (ranging from 1 to n) were assigned. This was repeated for each gene in the gene set. Next, rank scores were summed generating a pathway activity score for each sample. Chromatin immunoprecipitation Histone marks for active transcription (H3K4me3; ab8580, Abcam, Cambridge, MA, USA), repression (H3K27me3; , Upstate, Lake Placid, NY, USA) and elongation (H3K36me3; Abcam, ab9050) were assessed together with c- MYC and MYCN binding. ChIP- DNA from 6 neuroblastoma cell lines (SH- EP, SJ- NB12, WAC2, SY5Y, Kelly and IMR5) was amplified for DNA microarray analysis (Agilent Human Promoter ChIP- chip Set 244K) using whole genome amplification method (WGA, Sigma) as previously described (Westermann et al., 2008). DNA labeling, array hybridization and measurement was performed according to Agilent mammalian ChIP- chip protocol. For the visualization of ChIP- chip results, the cureos package v0.2 for R was used (available upon request) (Westermann et al., 2008). Two positive controls for MYCN/c- MYC binding, ENO1 (Kim et al., 2004) and mir (O'Donnell et al., 2005), were evaluated in the MYCN amplified Kelly cell line and c- MYC amplified SJ- NB- 12 cell line respectively (Figure S12). ChIP- chip data indicate a strong enrichment of MYCN/c- MYC binding and histone marks that are in line with an active transcriptional state. As a negative control we analysed the promoters of mirnas that are not significantly up- or downregulated by MYCN/c- MYC (mir- 10a, mir- 377, mir- 381 and mir- 452) in the MYCN amplified Kelly cell line (Figure S13). As expected, no MYCN binding was observed.
85 79 Ingenuity Pathway Analysis Enriched pathways among the predicted mirna targets were generated through the use of Ingenuity Pathways Analysis (Ingenuity Systems, The Functional Analysis of the candidate mirna target genes identified the biological functions and canonical pathways that were most significant to the considered gene list. The candidate mirna target genes associated with biological functions and canonical pathways in the Ingenuity Pathways Knowledge Base were considered for the analysis. Fischer s exact test was used to calculate a p- value determining the probability that each biological function and canonical pathway assigned to the candidate mirna target genes is due to chance alone. Biological functions and canonical pathways with a p- value below 0.05 were considered significant. Assessment of cooperative regulation The number of different mirnas that are predicted to target a mrna was taken as a measure of cooperative regulation. A predicted target was defined as a mrna with at least one 7mer seed in the 3 UTR region. We next calculated the cumulative distribution of this measure for the targets (n = 557) of the MYCN activated mirnas (n = 11) and for a random selection of 557 mrnas. The sampling of 557 random mrnas was repeated 100 times in order to get a reliable estimate of the background distribution. In a second analysis we performed a sampling of the mirnas and calculated the cooperative regulation for 11 random mirnas towards the targets of the MYCN upregulated mirnas. The sampling was repeated 100 times. Distributions were compared using a Kolmogorov- Smirnov test. Supplemental References Baek D, Villen J, Shin C, Camargo FD, Gygi SP, Bartel DP (2008). The impact of micrornas on protein output. Nature 455: Fredlund E, Ringner M, Maris JM, Pahlman S (2008). High Myc pathway activity and low stage of neuronal differentiation associate with poor outcome in neuroblastoma. Proc Natl Acad Sci U S A 105: Friedman RC, Farh KK, Burge CB, Bartel DP (2009). Most mammalian mrnas are conserved targets of micrornas. Genome Res 19: Grimson A, Farh KK, Johnston WK, Garrett- Engele P, Lim LP, Bartel DP (2007). MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol Cell 27: Kerr G, Ruskin HJ, Crane M, Doolan P (2008). Techniques for clustering gene expression data. Comput Biol Med 38: Kim JW, Zeller KI, Wang Y, Jegga AG, Aronow BJ, O'Donnell KA et al (2004). Evaluation of myc E- box phylogenetic footprints in glycolytic genes by chromatin immunoprecipitation assays. Mol Cell Biol 24: Lutz W, Stohr M, Schurmann J, Wenzel A, Lohr A, Schwab M (1996). Conditional expression of N- myc in human neuroblastoma cells increases expression of alpha- prothymosin and ornithine decarboxylase and
86 80 accelerates progression into S- phase early after mitogenic stimulation of quiescent cells. Oncogene 13: O'Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT (2005). c- Myc- regulated micrornas modulate E2F1 expression. Nature 435: Schulte JH, Horn S, Otto T, Samans B, Heukamp LC, Eilers UC et al (2008). MYCN regulates oncogenic MicroRNAs in neuroblastoma. Int J Cancer 122: Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D et al (2003). Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13: Westermann F, Muth D, Benner A, Bauer T, Henrich KO, Oberthuer A et al (2008). Distinct transcriptional MYCN/c- MYC activities are associated with spontaneous regression or malignant progression in neuroblastomas. Genome Biol 9: R150. Supplemental Figure 1 Hierarchical clustering of neuroblastoma tumours using the MYCN/c- MYC mirna signature. Enlargement of Figure 1A indicating the mirnas along the cluster Value hsa-mir-20a hsa-mir-20a hsa-mir-20b hsa-mir-20b hsa-mir-92 hsa-mir-92a hsa-mir-19a hsa-mir-19a hsa-mir-18a hsa-mir-18a hsa-mir-18astar hsa-mir-18a* hsa-mir-15b hsa-mir-15b hsa-mir-9 hsa-mir-9 hsa-mir-130a hsa-mir-130a hsa-mir-181a hsa-mir181a hsa-mir-214 hsa-mir-214 hsa-mir-645 * hsa-mir-601 hsa-mir-601 hsa-mir-572 hsa-mir-572 hsa-mir-610 hsa-mir-610 hsa-mir-526bstar hsa-mir526b* hsa-mir-128a hsa-mir-128a hsa-mir-137 hsa-mir-137 hsa-mir-615 hsa-mir-615 hsa-mir-215 hsa-mir-215 hsa-mir-326 hsa-mir-326 hsa-mir-129 hsa-mir-129 hsa-mir-500 hsa-mir-500 hsa-mir-103 hsa-mir-103 hsa-mir-340 hsa-mir-340 hsa-mir-153 hsa-mir-153 hsa-mir-95 hsa-mir-95 hsa-mir-491 hsa-mir491 hsa-mir-184 hsa-mir-184 hsa-mir-324-3p hsa-mir-324-3p hsa-mir-197 hsa-mir-197 hsa-mir-328 hsa-mir-328 hsa-mir-628 hsa-mir-324-5p hsa-mir-324-5p hsa-mir-330 hsa-mir-330 hsa-mir-488 hsa-mir-149 hsa-mir-149 hsa-mir-331 hsa-mir331 hsa-mir-28 hsa-mir-28 hsa-mir-30e-3p hsa-mir-30e-3p hsa-mir-30a-3p hsa-mir-30a-3p hsa-mir-148a hsa-mir-148a hsa-mir-148b hsa-mir-148b hsa-mir-190 hsa-mir-190 hsa-mir-30d hsa-mir-30d hsa-mir-30b hsa-mir-30b hsa-mir-30c hsa-mir-30c hsa-mir-140 hsa-mir-140 hsa-mir-26b hsa-mir-26b hsa-mir-26a hsa-mir-26a X2701 Essen50 X1822 Essen41 Essen42 X2530 Essen61 X1827 Essen46 Essen44 X2697 X3007 X3011 X1788 X1823 X2700 Essen3 X1784 X1974 X2540 X2486 Essen38 X1824 X1851 X1786 X2508 Essen2 Essen59 X1834 Essen28 Essen26 X3004 Essen1 Essen36 Essen27 X3020 X1972 X2083 Essen64 X1821 X1855 Essen53 X1776 Essen11 Essen57 Essen49 Essen35 X3008 X2740 Essen63 X3009 Essen48 Essen45 X3015 X3021 X3016 X2493 X2512 X2495 X1852 Essen24 X2509 Essen32 Essen33 X2527 X2496 X1792 X2497 X2535 Essen62 X2483 Essen34 X2489 X2498 Essen60 Essen51 Essen55 X2528 Essen18 X2507 Essen47 X1777 X2485 X2505 X2484 Essen30 X1820 X1795 X2525 X2515 Essen65 Essen43 Essen54 Essen56 X2506 Density Color Key and Density Plot MNA SH SL
87 81 Supplemental Figure 2. MYCN/c- MYC signaling is increased in high risk neuroblastoma tumours MYCN (A) and c- MYC (B) expression (mean ± SEM) in MYCN amplified (MNA), MYCN single copy low risk (SL) and MYCN single copy high risk (SH) neuroblastoma tumours. Mann- Whitney p- values for differential expression between groups are indicated.!"#"$%$&!"#"$%$$$&!"#"$%$$$&!"#"$%$'!"#"$%$'!"#"$%$$&
88 82 Supplemental Figure 3. c- MYC- translocation in a neuroblastoma tumour (A) FISH analysis of a primary neuroblastoma tumour using a c- MYC break- apart probe (red green). Each cell contained 3 split signals and 1 co- localized signal indicative for translocation. One representative cell is shown. (B) Expression of c- MYC in the c- MYC- translocated tumour compared to the tumours with no genetic aberrations at the c- MYC locus. c- MYC expression is increased fold with respect to the mean expression in the tumours with no genetic aberrations at the c- MYC locus. (C) Expression of the c- MYC target genes ODC1, PHGDH and MTHFD2 in the c- MYC- translocated tumour compared to MYCN amplified (MNA), MYCN single copy low risk (SL) and MYCN single copy high risk (SH) tumours.! " #
89 83 Supplemental Figure 4. MYCN/c- MYC mirna expression in c- MYC and MYCN amplified neuroblastoma cell lines Expression (mean ± SEM) of MYCN/c- MYC activated and MYCN/c- MYC repressed mirnas in MYCN single copy (MNSC) and MYCN amplified (MNA) neuroblastoma cell lines compared to the c- MYC amplified neuroblastoma cell line SJNB- 12. Representative activated (mir- 181a, mir- 92a) and repressed (mir p, mir- 184) mirnas are shown. hsa-mir-181a hsa-mir-92a hsa-mir-330-3p hsa-mir-184
90 84 Supplemental Figure 5. Induction of mirna expression in two independent MYCN model systems Fold induction of upregulated mirnas from the MYCN/c- MYC mirna signature in (A) SHEP- 21N and (B) SHEP- MYCN- ER cells, 48 hours upon MYCN activation. Red bars represent mirnas for which a >1.5- fold induction in expression was observed. No fold inductions were calculated for mirnas that were undetectable in one or both conditions. I #! #! %&'()*+&,-.'/,01(23&,456789#: ##?*>8@9ABC?*>8A#!?*>8AD@?*>8#E!(?*>8#@B?*>8#F(?*>89!B?*>8#F(C?*>89!(?*>8@G9?*>8H?*>8#F#(?*>89#D?*>8A!#?*>8#H(?*>8H9( J!$#!"# #! #! %&'()*+&,-.'/,01(23&,45678;<=:86> ##!$#!"#?*>8@9ABC?*>8A#!?*>8AD@?*>8#E!(?*>8#@B?*>8#F(?*>89!B?*>8#F(C?*>89!(?*>8@G9?*>8H?*>8#F#(?*>89#D?*>8A!#?*>8#H(?*>8H9(
91 85 Supplemental Figure 6. ChIP- chip for MYCN/c- MYC regulated mirnas Representative ChIP- chip results for mir- 15b, mir- 130a and mir- 214 are given for the Kelly neuroblastoma cell line. Oligonucleotide position is given as bars according to the chromosomal localisation. Colour coding of the bars represents the log2 ratios MYCN versus input from ChIP- chip experiments, were red means positive and green negative values. Histone marks for active transcription (H3K4me3), repression (H3K27me3) and enlongation (H3K36me3) as measured by ChIP- chip are given together with MYCN binding using the same colour coding. mirna transcript information (mirbase Version 11.0), CpG islands, and conservation among 28 species were implemented for the region as given by the respective annotation tracks deposited in the UCSC database (Hg 18, release March 2006). A grey coding for results of the positional weight matrix (PWM) scan represents the p- values of the 12bp MYCN binding motif from the TRANSFAC database. Red line = median log2 ratio MYCN versus input. HSA-MIR-15b Kelly SMC4 CpG-islands conservation mirna 15b 16-2 e-box MYCN
92 86 HSA-MIR-130a Kelly conservation mirna 130a MYCN HSA-MIR-214 Kelly DNM3 conservation mirna a-2 MYCN
93 87 Supplemental Figure 7. Induction of mirna expression in the SHEP- mir cell line Fold induction of mir- 17, mir- 18a, mir- 19a, mir- 19b, mir- 20a and mir- 92a upon treatment of the SHEP- mir cell line with tetracycline (Tet). Expression values are relative to the untreated condition hsa-mir hsa-mir-18a 12 hsa-mir-19a Relative expression Relative expression Relative expression Tet hsa-mir-19b 12 hsa-mir-20a 12 hsa-mir-92a Relative expression Relative expression Relative expression Tet
94 88 Supplemental Figure 8. GSEA results for SHEP- TR- mir cells Enrichment of the predicted 8mer en conserved targets gene lists for mir- 19a, mir- 20a and mir- 92a among the mrnas that were downregulated upon mir induction in the SHEP- TR- mir cells. Enrichment score Conserved seed Enrichment score mer seed high mir low mir high mir low mir P = 0.005; FDR q = P = 0.01; FDR q = Supplemental Figure 9. Cumulative distribution of cooperative regulation Cumulative distribution plot of the number of predicted mirnas per target, taken as a measure for cooperative regulation by mirnas. (A) Cumulative distribution for the targets of the MYCN activated mirnas (red) and for a random selection of mrnas (blue). (B) Cumulative distribution for the targets of the MYCN activated mirnas (red) and for a random selection of mirnas (blue). A B p < p < cummulative fraction (%) cummulative fraction (%) number of mirnas per target number of mirnas per target random set specific set
95 89 Supplemental Figure 10. Integrin signaling activity correlates to patient outcome in neuroblastoma. Pathway activity score, represented as quartiles, for integrin signaling significantly correlates to overall (OS) and event free (EFS) patient survival in two independent datasets (Oberthuer et al., 2006; Wang et al., 2006). The lowest and highest pathway activity scores are represented by the first quartile (yellow) and fourth quartile (red) respectively. Cox regression p- values are listed. OS (%) OS Oberthuer OS Oberthuer EFS EFS (%) EFS p < p < follow-up time (years) follow-up time (years) OS (%) OS Wang OS EFS (%) EFS Wang EFS p < p < follow-up time (years) follow-up time (years) 0% 25% 25% 50% 50% 75%% 75% 100%
96 90 Supplemental Figure 11. Integrin signaling activity correlates to patient outcome in diffuse large B- cel lymphoma. Pathway activity score, represented as quartiles, for integrin signaling significantly correlates to overall patient survival (OS) in a dataset of diffuse large B- cell lymphoma. The lowest and highest pathway activity scores are represented by the first quartile (yellow) and fourth quartile (red) respectively. The Cox regression p- value is listed. OS (%) p < follow-up time (years) 0% 25% 25% 50% 50% 75%% 75% 100% Supplemental Figure 12. ChIP- chip positive controls ChIP- chip results for MYCN/c- MYC binding to ENO1 and mir in the MYCN amplified Kelly cell line and c- MYC amplified SJ- NB- 12 cell line respectively. Histone marks for active transcription (H3K4me3), repression (H3K27me3) and enlongation (H3K36me3) as measured by ChIP- chip are given. Oligonucleotide position is given as bars according to the chromosomal localisation. Colour coding of the bars represents the log2 ratios in the precipitated versus input sample from ChIP- chip experiments, were red means positive and green negative values.
97 Kelly ENO1 Gene 91 CpG-islands conservation e-box MYCN H3K4me3 H3K36me3 H3K27me3
98 SJNB-12 CpG-islands HSA-MIR cluster 92 conservation mirna e-box 17 19a 19b-1 18a 20a 92a-1 c-myc MYCN H3K36me3 H3K4me3 H3K27me3
99 93 Supplemental Figure 13. ChIP- chip negative controls ChIP- chip results for MYCN/c- MYC binding to the negative control mirnas mir- 10a, mir- 377, mir- 381 and mir- 452) in the MYCN amplified Kelly cell line. Histone marks for active transcription (H3K4me3), repression (H3K27me3) and enlongation (H3K36me3) as measured by ChIP- chip are given. Oligonucleotide position is given as bars according to the chromosomal localisation. Colour coding of the bars represents the log2 ratios in the precipitated versus input sample from ChIP- chip experiments, were red means positive and green negative values.
100 Kelly CpG-islands HSA-MIR-10a 94 conservation mirna 10a MYCN c-myc H3K27me3 H3K36me3 H3K4me3
101 Kelly conservation HSA-MIR mirna e-box c-myc MYCN H3K36me3 H3K4me3 H3K27me3
102 Kelly HSA-MIR conservation 376b 376a b mirna MYCN c-myc H3K36me3 H3K27me3 H3K4me3
103 Kelly HSA-MIR-452 gene (GABRE) 97 conservation mirna 451 e-box MYCN c-myc H3K27me3 H3K4me3 H3K36me3
104 Supplemental Table 1. MYCN/c- MYC activated and repressed mirnas Overview of the up- and downregulated mirnas in the mirna signature. 98 High in MYCN amplified tumours Low in MYCN amplified tumours mir-526b* mir-30c mir-95 mir-610 mir-324-5p mir-148b mir-645 mir-488 mir-26a mir-130a mir-197 mir-30a-5p mir-15b mir-331 mir-30e mir-18a mir-149 mir-491 mir-20b mir-30b mir-615 mir-18a* mir-324-3p mir-129 mir-20a mir-628 mir-140 mir-572 mir-103 mir-26b mir-9 mir-148a mir-328 mir-181a mir-153 mir-340 mir-214 mir-184 mir-500 mir-601 mir-190 mir-215 mir-19a mir-28 mir-128a mir-92 mir-330 mir-137 mir-30d mir-326
105 99 Supplemental Table 2. Previously reported mirnas from the MYCN/c- MYC signature MiRNAs from the MYCN/c- MYC signature that have been reported as differentially expressed in the context of increased MYCN/c- MYC signaling in neuroblastoma, medulloblastoma, breast carcinoma and B- cell lymphoma. mirna Transcriptional control Reference(s) Promoter binding Regulation mir-128a - DOWN 1 mir-181a MYCN UP 1, 2, 3, 4 mir-18a MYCN, c-myc UP 1, 2, 4, 5, 6 mir-19a MYCN, c-myc UP 1, 2, 5, 6 mir-20a MYCN, c-myc UP 1, 2, 4, 5, 6, 7, 9 mir-20b - UP 1, 7 mir-26a c-myc DOWN 8, 9 mir-26b c-myc DOWN 8 mir-30a-5p c-myc DOWN 8 mir-30b c-myc DOWN 3, 8 mir-30c c-myc DOWN 3, 8 mir-30d c-myc DOWN 8 mir-30e c-myc DOWN 3, 7, 8 mir-331-3p - DOWN 1, 3 mir DOWN 1 mir DOWN 1, 7! mir-92a MYCN, c-myc UP 2, 3, 4, 5, 6, 7, 9 1. Chang, T.C., Yu, D., Lee, Y.S., Wentzel, E.A., Arking, D.E., West, K.M., Dang, C.V., Thomas-Tikhonenko, A., and Mendell, J.T Widespread microrna repression by Myc contributes to tumorigenesis. Nat Genet 40(1): Chen, Y. and Stallings, R.L Differential patterns of microrna expression in neuroblastoma are correlated with prognosis, differentiation, and apoptosis. Cancer Res 67(3): Fontana, L., Fiori, M.E., Albini, S., Cifaldi, L., Giovinazzi, S., Forloni, M., Boldrini, R., Donfrancesco, A., Federici, V., Giacomini, P. et al Antagomir-17-5p abolishes the growth of therapy-resistant neuroblastoma through p21 and BIM. PLoS One 3(5): e Mestdagh, P., Van Vlierberghe, P., De Weer, A., Muth, D., Westermann, F., Speleman, F., and Vandesompele, J A novel and universal method for microrna RT-qPCR data normalization. Genome Biol 10(6): R64.
106 Northcott, P.A., Fernandez, L.A., Hagan, J.P., Ellison, D.W., Grajkowska, W., Gillespie, Y., Grundy, R., Van Meter, T., Rutka, J.T., Croce, C.M. et al The mir-17/92 polycistron is up-regulated in sonic hedgehog-driven medulloblastomas and induced by N-myc in sonic hedgehog-treated cerebellar neural precursors. Cancer Res 69(8): O'Donnell, K.A., Wentzel, E.A., Zeller, K.I., Dang, C.V., and Mendell, J.T c- Myc-regulated micrornas modulate E2F1 expression. Nature 435(7043): Sander, S., Bullinger, L., Klapproth, K., Fiedler, K., Kestler, H.A., Barth, T.F., Moller, P., Stilgenbauer, S., Pollack, J.R., and Wirth, T MYC stimulates EZH2 expression by repression of its negative regulator mir-26a. Blood 112(10): Schulte, J.H., Horn, S., Otto, T., Samans, B., Heukamp, L.C., Eilers, U.C., Krause, M., Astrahantseff, K., Klein-Hitpass, L., Buettner, R. et al MYCN regulates oncogenic MicroRNAs in neuroblastoma. Int J Cancer 122(3): Sun, Y., Wu, J., Wu, S.H., Thakur, A., Bollig, A., Huang, Y., and Joshua Liao, D Expression profile of micrornas in c-myc induced mouse mammary tumors. Breast Cancer Res Treat.! Supplemental Table 3. Multivariate Cox proportional hazards model for MYCN amplification and the 8mer signature (grouped by quartiles) with overall and event free survival as measured end point Dataset Measured end point Variable p-value HR 95% CI OS MYCN amplification mer signature (quartiles) 5.60E Oberthuer EFS MYCN amplification mer signature (quartiles) 4.00E Wang OS MYCN amplification mer signature (quartiles) EFS MYCN amplification mer signature (quartiles)
107 PAPER 4: The mir MicroRNA Cluster Regulates Multiple Components of the TGF- β Pathway in Neuroblastoma 101 PAPER 4 The mir MicroRNA Cluster Regulates Multiple Components of the TGF-β Pathway in Neuroblastoma. Mestdagh P, Boström AK, Impens F, Fredlund E, Van Peer G, De Antonellis P, von Stedingk K, Ghesquière B, Schulte S, Dews M, Thomas-Tikhonenko A, Schulte JH, Zollo M, Schramm A, Gevaert K, Axelson H, Speleman F, Vandesompele J. Mol Cell Dec 10;40(5):
108 102 Molecular Cell Article The mir MicroRNA Cluster Regulates Multiple Components of the TGF-b Pathway in Neuroblastoma Pieter Mestdagh, 1,11 Anna-Karin Boström, 2,11 Francis Impens, 3,4 Erik Fredlund, 1,5,6 Gert Van Peer, 1 Pasqualino De Antonellis, 7 Kristoffer von Stedingk, 2 Bart Ghesquière, 3,4 Stefanie Schulte, 8 Michael Dews, 9 Andrei Thomas-Tikhonenko, 9 Johannes H. Schulte, 8 Massimo Zollo, 7,10 Alexander Schramm, 8 Kris Gevaert, 3,4 Håkan Axelson, 2 Frank Speleman, 1,12 and Jo Vandesompele 1,12, * 1 Center for Medical Genetics, Ghent University Hospital, B-9000 Ghent, Belgium 2 Department of Laboratory Medicine, Center for Molecular Pathology, Lund University, SE Malmo, Sweden 3 Department of Medical Protein Research, VIB, B-9000 Ghent, Belgium 4 Department of Biochemistry, Ghent University, B-9000 Ghent, Belgium 5 Department of Oncology, Clinical Sciences, Lund University, SE Malmo, Sweden 6 CREATE Health, Strategic Centre for Translational Cancer Research, Lund University, SE Malmo, Sweden 7 Centro di Ingegneria Genetica e Biotecnologia Avanzate (CEINGE), Naples, Italy 8 University Hospital of Essen, Essen, Germany 9 Division of Cancer Pathobiology, Department of Pathology & Laboratory Medicine, The Children s Hospital of Philadelphia Research Institute and University of Pennsylvania School of Medicine, Philadelphia, PA 19104, USA 10 Dipartimento di Biochimica e Biotecnologie Mediche (DBBM), Università di Napoli, Naples, Italy 11 These authors contributed equally to this work 12 These authors contributed equally to this work *Correspondence: [email protected] DOI /j.molcel SUMMARY The mir microrna cluster is often activated in cancer cells, but the identity of its targets remains elusive. Using SILAC and quantitative mass spectrometry, we examined the effects of activation of the mir cluster on global protein expression in neuroblastoma (NB) cells. Our results reveal cooperation between individual mir mirnas and implicate mir in multiple hallmarks of cancer, including proliferation and cell adhesion. Most importantly, we show that mir is a potent inhibitor of TGF-b signaling. By functioning both upstream and downstream of psmad2, mir activation triggers downregulation of multiple key effectors along the TGF-b signaling cascade as well as direct inhibition of TGF-b-responsive genes. INTRODUCTION MicroRNAs (mirnas) belong to a regulatory class of small noncoding RNAs with a fundamental role in numerous aspects of cell biology, such as cell-cycle regulation, apoptosis, differentiation, and maintaining stemness (reviewed in Bartel [2004]). Only nucleotides (nt) in length, mirnas function as key molecules in the posttranscriptional repression of gene expression. Upon mirna assembly in the RNA-induced silencing complex (RISC), binding between the mirna seed (nt 2 7 counted from the 5 0 end of the mirna) and complementary sites in the 3 0 untranslated region (3 0 UTR) of target mrnas results in degradation of the mrna or inhibition of translation (reviewed in Bartel [2009]). Based on the 3 0 UTR site context, algorithms predict that up to 60% of all coding genes are under the control of one or more mirnas (Friedman et al., 2009). However, these predictions suffer from a high degree of false positives, and to date, only a fraction of mirna-mrna interactions have been experimentally validated. In cancer, mirnas function both as oncogenes or tumor suppressors (reviewed in Calin and Croce [2006]; Esquela-Kerscher and Slack [2006]). Some of these mirnas were identified as essential components of known cancer pathways, such as the p53-induced mir-34 family (He et al., 2007; Raver-Shapira et al., 2007) or the c-myc/mycn-induced mir cluster (O Donnell et al., 2005). The oncogenic mir cluster consists of six individual mirnas (mir-17, mir-18a, mir-19a, mir-19b, mir-20a, and mir-92a) located within a polycistronic transcript on human chromosome 13. Gene duplications and deletions eventually resulted in two mir paralogs, the mir-106b-25 cluster on chromosome 7 and the mir-106a-363 cluster on chromosome X. Of these clusters, mir is the most frequently activated one in cancer. mirna expression profiling studies revealed mir overexpression, both in hematopoietic malignancies (such as B cell lymphomas [He et al., 2005]) and solid tumors (including breast, colon, and lung cancer [Castellano et al., 2009; Hayashita et al., 2005; Lanza et al., 2007]) and neuroblastoma (NB) (Mestdagh et al., 2009a). Overexpression can result from amplification of the mir locus (He et al., 2005) or direct mir transactivation by c-myc/mycn (Dews et al., 2010; Fontana et al., 2008; Mestdagh et al., 2009a; O Donnell et al., 2005). The oncogenic nature of mir activation is supported by the identification of 762 Molecular Cell 40, , December 10, 2010 ª2010 Elsevier Inc.
109 103 Molecular Cell mir Dampens TGF-b Signaling A 600 B mir pathway activity score OS (%) EFS (%) SL SH MNA 0 p < follow-up time (years) 0 p < follow-up time (years) 0% - 25% 25% - 50% 50% - 75% 75% - 100% Figure 1. mir Cluster Activation Is a Marker for Poor Prognosis (A) mir pathway activity is scored in three clinicogenetic subsets of NB tumors (data set D1, Table S1), MYCN amplified tumors (MNA), MYCN single copy high-risk tumors (SH), and MYCN single-copy low risk tumors (SL) (whiskers: Tukey). mir pathway activity score is significantly higher in MNA versus SH (Mann Whitney, p < 0.05), MNA versus SL (p < ), and SH versus SL (p < 0.01). (B) Kaplan Meier plots for overall (OS) and event free survival (EFS) based on the pathway activity score of mir-17-92, represented as quartiles. Increased activity of mir is proportionally correlated to both poor overall and event-free survival. mir targets with key roles in cell-cycle control and cell death. In particular, mir-17 and mir-20a target the cyclindependent kinase inhibitor CDKN1A (p21), a negative regulator of the G 1 -S transition (Fontana et al., 2008), and mir-17 targets the proapoptotic BCL2L11 (Bim) (Fontana et al., 2008). In gastric cancer, downregulation of p21 by the mir-17 and mir-20a paralogs mir-106b and mir-93 renders the cells insensitive to TGF-b-induced cell-cycle arrest whereas mir-25 (a mir-92a paralog) inhibits TGF-b-dependent apoptosis through the repression of BCL2L11 (Petrocca et al., 2008). Thus far, the number of identified mir targets remains relatively limited, thus precluding a comprehensive understanding of the full oncogenic potential of this mirna cluster. In a first step toward this goal, we examined the effects of mir cluster activation on the proteome of NB cancer cells. Using quantitative mass spectrometry, we analyzed the response of thousands of proteins upon mir activation in NB cells. NB is an excellent model to study the effects of mir activation because high-risk NB tumors are characterized by increased MYCN/c-MYC activity either through MYCN amplification or increased c-myc expression, both resulting in elevated mir levels (Mestdagh et al., 2009a). Our results demonstrate that mir is implicated in multiple hallmarks of the tumorigenic program, including proliferation and cell adhesion. Most importantly, we dissect the role of mir as a potent inhibitor of TGF-b-signaling acting on multiple levels along the signaling cascade. RESULTS mir Cluster Activation Is a Marker for Poor Survival In NB, mir expression is activated through direct MYCN/ c-myc promoter binding (Fontana et al., 2008; Mestdagh et al., 2009b). We quantified mir expression on a cohort of 95 primary untreated NB tumor samples (data set D1, Table S1; GEO accession number GSE21713) (Mestdagh et al., 2009a). The activation of the entire mir cluster was evaluated by means of a pathway activity score (Fredlund et al., 2008; Mestdagh et al., 2009a). NB tumors were divided into three cohorts, MYCN single copy low-risk tumors (SL), MYCN single copy high-risk tumors (SH), and MYCN amplified tumors (MNA). The mir pathway activity was highest in the MNA tumors, followed by the SH tumors and the SL tumors (Figure 1A). Each individual mirna is upregulated in the MNA samples suggesting that the entire mir cluster, rather than a subset of mirnas, is of potential relevance (Mann Whitney, p < 0.05) (Figure S1A). We next evaluated mir pathway activation with respect to NB patient survival. Kaplan-Meier analysis demonstrated that mir activity was proportional to overall and eventfree survival (log rank, p < 0.001), underscoring the importance of mir activation in NB tumor biology (Figure 1B). Except for mir-19b, expression of the other mirnas within the mir cluster showed similar correlations (Figure S1B). Impact of mir Activation on Protein Output To study the regulatory effects of mir activation, quantitative mass spectrometry was applied to measure protein response in a cellular model (SHEP-TR-miR-17-92) with tetracycline-inducible mir expression (Mestdagh et al., 2009a). This approach provides the most relevant readout as it directly measures the impact of a mirna on protein output (Baek et al., 2008; Selbach et al., 2008). Average mir induction upon tetracycline treatment was in the range of mir fold changes between MNA and SL tumors (Figure S2A) (data not shown). Profiling of 430 mirnas revealed no significant effects on global mirna expression suggesting that mir induction does not affect the processing of other mirnas (data not shown). SHEP-TR-miR cells were differentially labeled using SILAC (stable isotope labeling with amino acids in cell culture) (Ong et al., 2002) and then either treated with tetracycline for 72 hr or left untreated, followed by methionine COFRADIC Molecular Cell 40, , December 10, 2010 ª2010 Elsevier Inc. 763
110 104 Molecular Cell mir Dampens TGF-b Signaling A SILAC 12 C 6 Arg/Lys SILAC 13 C 6 Arg/Lys B - TET + TET MESGFTSK from NNMT Ratio L/H : 0.43 lyse cells lyse cells Mix COFRADIC Primary RP-HPLC run H 2 O 2 oxidation Secondary RP-HPLC runs EFLFNAIETMPCVK from RRM2 Ratio L/H : H H O N C C CH 2 CH 2 S CH 3 H H O N C C CH 2 CH 2 S O CH 3 SDGSTVSVPMMAQTNK from SERPINE1 Ratio L/H : LC-MS/MS-analysis peptide identification (MASCOT) peptide & protein quantification (MASCOT Distiller) Figure 2. Analysis of Global Protein Expression upon mir Activation (A) Tetracycline treated (+TET) and untreated ( TET) SHEP-TR-miR cells were metabolically labeled using SILAC. Methionine-containing peptides were isolated using COFRADIC technology and subsequently analyzed using LC-MS/MS. (B) Representative LC-MS/MS spectra for an upregulated protein (NNMT), unchanged protein (RRM2), and downregulated protein (SERPINE1). isolation of methionyl peptides (Gevaert et al., 2002) and identification of these peptides by LC-MS/MS (Figure 2A). Only proteins that were quantified by at least two different peptides over two different proteome analyses (n = 3249) were selected for further analysis (Colaert et al., 2010). Most proteins were in fact quantified by more than two peptides (Figure S3B). Differential protein expression was determined as the average protein ratio of the differentially labeled fractions across the biological replicates (Table S2; Figures 2B and S3C). Based on a foldchange expression cutoff of 0.5 log 2 units (see Supplemental Information for cutoff definition), 144 proteins were downregulated upon mir activation. To assess whether the measured protein response reflects regulatory mir effects, we performed an unbiased search for all possible 7-mer motifs (n = 16,384) in the 3 0 UTR of the downregulated proteins (15 th percentile) and compared these to motif occurrence in the 3 0 UTR of the remaining proteins. We found seven motifs to be overrepresented in the 3 0 UTR of the downregulated proteins, with the five most significant motifs belonging to the mir mirnas: mir-17, mir-19a, mir-19b, mir-20a, and mir-92a (Fisher Exact, p < 0.05, Bonferoni multiple testing correction) (Figure 3A). Strikingly, there was no enrichment for mir-18a seeds, suggesting that mir-18a does not substantially contribute to protein repression upon mir activation. Analyses using the 20 th percentile gave similar results (data not shown). Analyses for the 5 0 UTR and coding sequence (CDS) did not reveal significant enrichments for mir mirna seed sequences. However, we did observe an enrichment for the 7-mer-m8 seed of mir-17* in the CDS of the downregulated proteins, suggesting that mir-17*-mediated protein repression might depend on CDS binding. To evaluate mir seed efficiency with respect to protein repression, we plotted the cumulative distribution of protein fold changes for proteins with at least one mir UTR 6-mer, 7-mer-A1, 7-mer-m8, or 8-mer seed and compared these to proteins without mir seeds (Figure 3B). As expected, protein repression was highest in the presence of an 8-mer seed (Kolmogorov-Smirnov, p = ) followed by 7-mer-m8 (p = ), 7-mer-A1 (p = ) and 6-mer seeds (p = ). When evaluating each mir mirna separately, we observed similar results for mir-17/mir-20a, mir-19a/mir-19b, and mir-92a (mir-17/mir-20a and mir-19a/ mir-19b were analyzed together as they share identical seeds) (Figure 3C). For mir-18a, the relation between seed occurrence and protein fold change was less pronounced, further supporting our observation that the contribution of mir-18a to mir mediated protein repression is limited. The fraction of proteins containing at least one mir mer seed was highest for proteins that were downregulated at least 2-fold (82%) and decreased to background levels (45%) 764 Molecular Cell 40, , December 10, 2010 ª2010 Elsevier Inc.
111 105 Molecular Cell mir Dampens TGF-b Signaling A C cummulative fraction (%) mir-17 - mir-20a protein fold change (log 2 ) cummulative fraction (%) mir-18a protein fold change (log 2 ) B cummulative fraction (%) mir no site 60 6mer site 50 7mer-A1 site 40 7mer-m8 site 30 8mer site protein fold change (log 2 ) cummulative fraction (%) mir-19a - mir-19b protein fold change (log 2 ) cummulative fraction (%) mir-92a protein fold change (log 2 ) Figure 3. mir Activation Induces Widespread Repression of Targeted Proteins (A) Overview of all significantly enriched heptamer motifs in the 3 0 UTR of transcripts from repressed proteins. The top five significantly enriched motifs correspond to mir target sites. One motif corresponds to the 7-mer-m8 seed of the mir-302/mir-372 family, which differs in only one base with the 7-mer-m8 seed of mir-17/mir-20a. The last motif did not correspond to any known mirna nor did it show any overlap with mir seeds. (B) The cumulative distribution of protein fold changes upon mir activation, calculated for five different protein subsets: proteins with at least one mir mer site (red), 7-mer-m8 site (blue), 7-mer-A1 site (yellow), 6-mer site (green), and no site (black). (C) Identical analysis as in (B) but for each individual mirna from the mir cluster. The mir-17/mir-20a and the mir-19a/mir-19b were analyzed together as they share identical seeds. for unchanged proteins (Figure S3A). Robust protein repression was also characterized by the presence of multiple mir UTR sites per protein (Figure S3B), suggesting that individual mir mirnas cooperate to achieve target repression. This correlation was only observed for 3 0 UTR sites and not for 5 0 UTR or CDS sites (Figures S3C and S3D). To further evaluate mirna cooperation, we analyzed co-occurrence of individual mir sites in the 3 0 UTR of downregulated proteins and compared this to co-occurrence in the 3 0 UTR of upregulated proteins (used as a reference control set). We identified significant co-occurrence for mir-17/mir-20a sites and mir-19a/ mir-19b sites confirming cooperation between individual mirnas (Figure S3E). mir-18a sites almost never occurred in the absence of other mir sites (8.33%) and were significantly associated with mir-17/mir-20a sites (Figure S3E). mir Affects Multiple Cancer Pathways To gain insight into the pathways affected by oncogenic mir activation, we performed gene set enrichment analysis (GSEA) (Subramanian et al., 2005) using all measured proteins, ranked according to their fold change. Thirty-six gene sets were significantly enriched in the positive phenotype (i.e., downregulated proteins) while nine were enriched in the negative phenotype (i.e., upregulated proteins). Of the latter, six were related to increased metabolic activity of the mitochondrial oxidative phosphorylation energy production pathway (Figure S4). In NB, mir expression is activated by MYCN/c-MYC transcription factors that have been shown to regulate genes involved in the biogenesis of mitochondria and metabolism (Zhang et al., 2007). Our results now provide evidence that this, at least in part, is mediated through mir activation. The contribution of each individual mirna to the significant gene lists in the positive phenotype was calculated and visualized as a heatmap (Figure 4A). Among the gene lists enriched in the positive phenotype, which reflect direct mir regulated pathways, we identified multiple cancer-related processes such as cell proliferation, cell adhesion, TGF-b signaling, estrogen-signaling, and RAS signaling (Figure 4A). Hierarchical clustering reveals a close association between mir-17/ mir-20a- and mir-19a/mir-19b-regulated pathways, reflecting the previously observed co-occurrence of these sites. Again, mir-18a clusters further away from the remaining mir mirnas and is characterized by weak gene list associations. Molecular Cell 40, , December 10, 2010 ª2010 Elsevier Inc. 765
112 106 Molecular Cell mir Dampens TGF-b Signaling A B - TET + TET C -TET +TET D T0 T7 T14 T21 E 11.5 SHEP-TR-miR SHEP-TR SHEP-TR-miR BLI (log10, photons/s) SHEP-TR ** * * time of imaging (days) Figure 4. mir Activation Regulates Multiple Cancer Pathways (A) Heatmap of significant mirna-pathway associations, identified through gene set enrichment analysis. The intensity of the association is based on the fraction of genes with at least one 7-mer or 8-mer 3 0 UTR site. (B) Normalized cell index (mean ± standard deviation) as a measure for proliferation of tetracycline treated (+TET) and untreated ( TET) SHEP-TR-miR cells. Treatment was initiated 20 hr post seeding. (C) Evaluation of the cell-cell adhesion of tetracycline treated and untreated SHEP-TR-miR cells. Measurements of the relative cluster area for three independent experiments using ImageJ are displayed as bar plots. Upon mir activation, the area of the clusters dropped by >50% resulting in more but smaller aggregates. (D) Representative analyses of bioluminescence imaging (BLI) of luciferase positive SHEP-TR-miR and SHEP-TR cells injected etherotopically and subcutaneously in the right and left flank of nude athymic mice. Bar scale color indicates the number of photons/s measured by IVIS 3D imaging instrumentation. SHEP mir17-92 cells in vivo are still alive after 21 days post cell subcutaneous implantation. (E) Bioluminescence imaging (BLI) of luciferase positive SHEP-TR-miR and SHEP-TR cells injected etherotopically and subcutaneously in the right and left flank of nude athymic mice. Luciferase signals were measured at 0, 7, 14, and 21 days postengraftment and are shown as the mean ± SEM of five mice. Significant differences between SHEP-TR and SHEP-TR-miR cells are indicated by * (Student t test, p < 0.05) and ** (Student t test, p < 0.01). In NB, the oncogenic nature of mir has been ascribed to its ability to promote cell proliferation through the regulation of CDKN1A and BCL2L11 (Fontana et al., 2008). GSEA results indicate that mir has a much broader influence and targets different oncogenic pathways. As a proof of concept, we tried to validate the association with increased proliferation and decreased cell adhesion in the SHEP-TR-miR cells. Cell proliferation was evaluated in real-time using the xcelligence system. Upon mir activation, proliferation of SHEP-TR-miR cells increased (Figure 4B) and intercellular cell adhesion significantly decreased (Figure 4C). To evaluate the effect of mir activation in vivo we performed 766 Molecular Cell 40, , December 10, 2010 ª2010 Elsevier Inc.
113 107 Molecular Cell mir Dampens TGF-b Signaling A TGFB_ALL_UP TGFB_EARLY_UP PADUA_TGFB_UP high mir low mir high mir low mir high mir low mir B C 3000 p < Q1 (0% - 25%) TGFβ pathway activity score EFS(%) Q2 (25% - 50%) Q3 (50% - 75%) Q4 (75% - 100%) p < MNSC MNA follow-up time (years) Figure 5. mir Activation Represses the TGF-b Pathway (A) Gene set enrichment analysis plots for three different TGF-b gene sets showing significant enrichment among the mir repressed proteins. (B) TGF-b pathway activity score in MYCN amplified NB tumors (MNA) and MYCN single-copy NB tumors (MNSC) (data set D2, Table S1). MNA tumors show significantly lower TGF-b pathway activity (Mann Whitney, p < 0.001) (whiskers: Tukey). (C) Kaplan Meier plot for event free survival (EFS) based on the TGF-b pathway activity score, represented as quartiles (dataset D2, Table S1). Increased activity of mir is proportionally correlated to event-free survival. etherotopic injection of SHEP-TR-miR and SHEP-TR (control) cells in the right and left flanking site, respectively, of atymic nude mice that were given tetracyclin and visualized tumor cells using bioluminescence imaging. For SHEP-TR cells, the luciferase signal dropped to background levels after 7 days of engraftment, which is in line with previous findings demonstarting that SHEP cells are not tumorigenic in vivo (Schweigerer et al., 1990) (Figures 4D and 4E). In contrast, SHEP-TR-miR cells persisted much longer and showed statistically higer luciferase signals at 7, 14, and 21 days, indicating that, although tumorigenesis decreases, mir activation significantly prolongs the engraftment of SHEP cells, probably through increased proliferation and decreased apoptosis, activities previously ascribed to mir overexpression (Fontana et al., 2008). Together, these results confirm the relation between mir activation and cell proliferation and reveal a role for mir in the regulation of cell adhesion, hereby confirming the GSEA results. mir Impairs TGF-b Activity GSEA analysis identified three TGF-b-responsive gene sets (Padua et al., 2008; Verrecchia et al., 2001) among the proteins downregulated upon mir activation in the SHEP-TRmiR cells (Figure 5A). To exclude the possibility that repression of TGF-b-responsive genes is an artifact of mirna overexpression, we analyzed eight published protein expression data sets of mirna overexpression (Baek et al., 2008; Selbach et al., 2008) using GSEA. None of the TGF-b gene lists were significantly enriched in any of the data sets, suggesting the observed effect to be related to mir For a subset of the TGF-b-responsive genes, the measured protein repression was confirmed on the mrna level using RT-qPCR (Figure S5). We next evaluated this TGF-b signature in NB tumor samples using the pathway activity score of all genes that significantly contributed to the GSEA results (n = 21). For this purpose, we used the larger Oberthuer data set (Oberthuer et al., 2006) (data set D2, Table S1) to increase the power of our analysis. TGF-b pathway activity was significantly downregulated in MNA NB tumors that are characterized by high mir expression (Mann Whitney, p < 0.001) (Figure 5B), and showed a negative correlation to MYC pathway activity (Spearman s Rank p < 0.01, rho = 0.460). In addition, Kaplan-Meier survival analysis indicates that tumors with low TGF-b pathway activity are characterized by poor event-free survival (log-rank, p < ) (Figure 5C). To further substantiate the inverse relation between TGF-b target gene expression and mir expression, we performed an expression correlation analysis in a subset of 40 of the 95 NB tumors for which also mrna expression was Molecular Cell 40, , December 10, 2010 ª2010 Elsevier Inc. 767
114 108 Molecular Cell mir Dampens TGF-b Signaling A C -TET psmad2 (60 kd) ACTB (42 kd) relative protein expression psmad2 0 TET - + +TET TET TGF ligand TGF inhibitor available (data set D3, Table S1) (Mestdagh et al., 2009a). Hierarchical clustering of the correlation coefficients revealed that, indeed, mir expression inversely correlates to TGF-b target gene expression (Figure S6A). These results confirm that TGF-b signaling is downregulated in aggressive NB tumors with high mir expression and underscore the potential importance of TGF-b activity in NB tumor biology. We next evaluated which components of the TGF-b signaling cascade are controlled by mir mirnas. One important effector of active TGF-b signaling is phosphorylated SMAD2 protein (psmad2) that translocates to the nucleus to induce gene transcription. Upon tetracycline treatment of SHEP-TRmiR cells, we observed a significant decrease in nuclear psmad2 levels (Mann Whitney, p < ) (Figures 6A 6C). A similar decrease was observed for psmad3 levels (data not shown). When SHEP-TR-miR cells were transfected with a plasmid containing a SMAD-regulated luciferase reporter ([CAGA] 12 -Luc) and treated with TGF-b1, a strong activation of the reporter gene was observed (Figure 6D). However, when mir expression was activated through tetracycline treatment, reporter gene activation was substantially attenuated (Mann Whitney, p < 0.001) (Figure 6D). When the SHEP-TRmiR cells were cultured in the presence of the potent TGFBR1 inhibitor SB (Laping et al., 2002), the SMAD reporter gene activity was completely abrogated (Figure 6D). These results suggest that mir activation impairs the TGF-b signaling cascade by acting upstream of psmad2. mir Affects Multiple Levels of the TGF-b Pathway As decreased psmad2 levels are either caused by reduced receptor activity or reduced SMAD2 expression, we quantified TGFBR2 and SMAD2 mrna expression in the SHEP-TRmiR cells. Both TGFBR2 and SMAD2 expression levels D B * Figure 6. mir Inhibits psmad2 Levels and Activity (A and B) Immunohistochemical detection of phosphorylated SMAD2 protein (psmad2) in tetracycline treated (+TET) and untreated ( TET) SHEP-TR-miR cells. The cell intensity measurement (mean ± SEM) reveals a significant decrease in psmad2 levels in tetracycline treated cells (Mann Whitney, p < ). (C) Western blot analysis indicates a strong decrease (2.3-fold) in psmad2 levels upon mir induction (+TET). (D) The relative luciferase activity of a psmad2 reporter construct (mean ± SEM). Activation of mir expression through tetracycline treatment (+TET) results in a significant (*) decrease in reporter activity after stimulation of the TGF-b pathway with TGF-b1 (TGF-b ligand). TGF-b-inhibitor treatment completely abrogates the reporter activity. decreased by at least 1.5-fold upon mir activation (Figure 7A). SMAD4, the binding partner of psmad2, also displayed a decrease in expression upon mir activation (Figure 7A). This negative correlation with mir expression could be confirmed in primary NB tumor samples for SMAD2 and TGFBR2 (Spearman s Rank, p < 0.01) (Figure 7B), suggesting that mir regulates their expression. Indeed, both genes contain mir binding sites in their 3 0 UTR and a direct interaction between TGFBR2 and mir-20a has been established (Volinia et al., 2006). This mir mediated silencing of TGFBR2 ultimately results in decreased psmad2 levels and decreased transcription of the TGF-b-target genes. In total, we identified 13 TGF-b-target genes to be downregulated on the protein level with a log 2 fold change < 0.5 (7 out of 20 proteins in the PADUA_TGFB_UP gene set, 3 out of 16 proteins from the TGFB_EARLY_UP gene set, and 7 out of 28 proteins from the TGFB_ALL_UP gene set) (Table S3). As ten of these genes harbor mir binding sites in their 3 0 UTR (Table S3), we wondered whether they might also be targeted directly by mir To exclude the effects of mir directed inactivation of TGF-b signaling on the expression of TGF-b-responsive genes, we first treated SHEP-TR-miR cells for 4 hr with the TGFBR1 inhibitor SB431542, which completely abrogates TGF-b signaling (Figure 6C). Cells were subsequently treated with tetracycline to activate mir expression and harvested at 24 hr and 48 hr after tetracycline treatment. From the six genes that were evaluated, three (CDKN1A, ITGA4, and SERPINE1) were downregulated after 24 hr of TGF-b-inhibitor treatment (t test, p < 0.05), confirming that they are regulated by TGF-b (Figure 7C). The remaining three genes (FNDC3B, ICAM1, and THBS1) did not show any differential expression after 24 hr; however, FNDC3B and THBS1 did respond to TGF-b-inhibitor treatment after 48 hr (data not shown). This suggests that, in NB, these are either not or indirectly responsive to TGF-b signaling (Figure 7D). Upon mir activation, the TGF-b-responsive genes were further downregulated (t test, p < 0.001) (Figure 7C), 768 Molecular Cell 40, , December 10, 2010 ª2010 Elsevier Inc.
115 109 Molecular Cell mir Dampens TGF-b Signaling supporting our hypothesis that mir also influences the expression of these genes, independent of its ability to inactivate TGF-b signaling. As expected, the genes that were not responsive to TGF-b inhibition did show decreased expression upon mir activation (t test, p < 0.001) (Figure 7D). To investigate which specific mirnas contribute to the repression of the TGF-b-pathway, we overexpressed each mirna from the mir cluster separately and measured the expression of TGF-b-pathway components and target genes. Interestingly, we found that each mirna contributes to the repression of one or more genes from the TGF-b pathway, suggesting that the entire mir cluster, rather than a subset of mirnas, mediates the repression of TGF-b signaling in NB cells (Figure S6B). Downregulation (log 2 fold change < 0.5) upon mirna transfectection was almost exclusively observed for those genes harboring a 3 0 UTR seed site for the respective mirna (Fisher Exact, p < 0.001). We next evaluated whether the mir induced downregulation of TGF-b-pathway components is caused by direct binding between mir mirnas and mir seed sites in the 3 0 UTR of TGFBR2, SMAD2, and SMAD4. To this purpose, DLD1DICER hypo cells were transfected with 3 0 UTR luciferase reporter plasmids in combination with a pre-mir negative control or a mir pre-mir for which one or multiple seed sites were present in the 3 0 UTR of the respective genes. We identified a direct interaction between TGFBR2 and mir-17/20, SMAD2 and mir-18a, and SMAD4 and mir-18a, as evidenced by the significant decrease in luciferase activity compared to the premir negative control (t test, p < 0.01, Figure 7E). Other putative mir sites in the 3 0 UTR of TGFBR2 (mir-19a/mir-19b), SMAD2 (mir-19a/mir-19b, mir-92a), and SMAD4 (mir-19a/ mir-19b) did not affect luciferase signals (data not shown). Mutagenesis of the active mirna seed sites resulted in a significant rescue of the luciferase signal (t test, p < 0.01), suggesting that the observed effects depend on the presence of the 3 0 UTR seed site. These results confirm TGFBR2 as a direct mir target gene and identify two additional TGF-b-pathway components, SMAD2 and SMAD4, as mir target genes. To assess the importance of TGF-b-pathway inhibition in the proliferation phenotype observed upon mir activation, we overexpressed SMAD2 and SMAD4 in the presence of activated mir SMAD2/SMAD4 overexpression resulted in a 25% decrease in cell growth (t test, p < 0.05), indicating that mir accelerated proliferation is, at least in part, depending on the downregulation of the TGF-b pathway. The relatively modest decrease in cell growth is probably explained by the fact that mir directly regulates TGF-b target genes in a SMAD2/SMAD4 independent manner. In conclusion, our data demonstrate that mir activation triggers a targeted clampdown of TGF-b signaling by acting on multiple key effectors along the signaling cascade, as well as through the direct inhibition of TGF-b-responsive genes, hereby repressing the cytostatic effects of active TGF-b signaling (Figure S7). DISCUSSION Transcriptional activation of the mir mirna cluster by MYC/MYCN transcription factors occurs in multiple tumor entities, including NB (Hayashita et al., 2005; Mestdagh et al., 2009a; O Donnell et al., 2005). Although the oncogenic nature of mir activation is well established, the underlying targets and signaling cascades that are deregulated remain largely elusive. In addition, studies aimed at determining mir targets have focused on individual members of the cluster, despite the observation that the entire cluster is activated (Mestdagh et al., 2009b; O Donnell et al., 2005). Here, we have used an unbiased proteomics approach to identify mir targeted pathways in a NB tumor model. Direct quantitative measurement of protein expression is preferred over the more straightforward mrna profiling as a high-throughput method for mirna target identification (Baek et al., 2008; Selbach et al., 2008). Computational analysis of mir seeds in the 3 0 UTR of transcripts from proteins supported the expected enrichment of direct mir targets within the list of downregulated proteins detected using mass spectrometry. Moreover, a proportional relationship between seed frequency and fold downregulation was noted. This relationship not only holds for multiple seeds from an individual mir mirna but also for multiple seeds from different mir mirnas, suggesting cooperation between individual mirnas from the cluster toward target protein repression. mir mirnas have indeed been shown to function in a cooperative and additive manner among others in the regulation of PTEN by mir-17 and mir-19 (Xiao et al., 2008). Our results further indicate that mir-19a/mir-19b and mir-17/ mir-20a sites significantly co-occur in the 3 0 UTR of transcripts from several downregulated proteins. As these co-occurring sites were not observed for every possible combination of individual mir mirnas, we hypothesize that in NB, the mirna components of the mir cluster can regulate target expression either individually or in certain combinations with additive effects. However, mir function might be highly context and cell-type specific as mir-19 was shown to be both necessary and sufficient to promote MYC-induced lymphomagenesis in the Em-myc mouse B cell lymphoma model (Olive et al., 2009). While the fraction of downregulated proteins was enriched for seeds of mir-17/mir-20a, mir-19a/mir-19b, and mir-92a, enrichment for the mir-18a seed was not detected. Strikingly, mir-18a seeds rarely occur as the only seed(s) in the 3 0 UTR of a downregulated target and showed little or no correlation to protein fold change. Although this suggests that mir-18a is not substantially contributing to target deregulation, it does not imply that mir-18a lacks functionality, as mir-18a has been shown to regulate important cancer genes such as CTGF in colon cancer and estrogen receptor-a (ESR1) in NB (Dews et al., 2006; Lovén et al., 2010). Interestingly, we found mir-18a to regulate both SMAD2 and SMAD4, two key components of the TGF-b-signaling cascade, suggesting that mir-18a substantially contributes to pathway deregulation by regulating a selected set of target genes. When all cluster components were combined, we identified a large number of targeted proteins belonging to diverse cancerrelated pathways. Notably, estrogen receptor signaling was also among the targeted pathways. The fact that we identified such a wide variety of functions in NB cells suggests that mir pleiotropy is not only related to different targets in different cell Molecular Cell 40, , December 10, 2010 ª2010 Elsevier Inc. 769
116 110 Molecular Cell mir Dampens TGF-b Signaling A B C D E Figure 7. mir Targets Multiple Components of the TGF-b Pathway (A) The relative mrna expression of TGF-b-responsive genes in tetracycline treated (+TET) and untreated SHEP-TR-miR cells (+TET) (mean ± SEM). (B) Significant negative correlation between TGFBR2 mrna expression and mir expression and SMAD2 mrna expression and mir expression in primary NB tumors. Spearman s rank rho-values and p values are listed. (C and D) The relative mrna expression (mean ± SEM) of a representative set of genes responsive to TGF-b (C) and genes not or (indirectly) responsive to TGF-b (D) in SHEP-TR-miR cells that were either untreated, treated with TGF-b inhibitor, or treated with TGF-b inhibitor followed by mir activation with tetracycline (TET) for 24 hr. In (C), genes respond to TGF-b-inhibitor treatment (t test, p < 0.05, indicated by *) and show an additional decrease in expression upon combined TGF-b-inhibitor treatment and mir activation (t test, p < 0.001, indicated by *). In (D), genes only respond to mr treatment (t test, p < 0.001, indicated by **). 770 Molecular Cell 40, , December 10, 2010 ª2010 Elsevier Inc.
117 111 Molecular Cell mir Dampens TGF-b Signaling types but also occurs within cell types. The molecular basis for this observation likely lies within the multiple components of the cluster and the complex interplay between them. mir directed regulation of the TGF-b-responsive genes CDKN1A and BCL2L11 in NB cells has been described previously (Fontana et al., 2008). In gastric cancer, members of the mir-106b-25 cluster have also been shown to target CDKN1A and BCL2L11 (Petrocca et al., 2008). Here we comprehensively demonstrate that mir dampens TGF-b signaling in a multifaceted way by acting both upstream and downstream of psmad2/smad4, further underscoring its ability to regulate multiple components of the same pathway. This ability to simultaneously target the components of the signaling cascade, as well as the downstream effectors through multiple mirnas, allows for tight control of the TGF-b-transcriptional program. Moreover, it offers the cells enormous flexibility and plasticity for regulation of different subsets of TGF-b target genes. In NB, enhanced TGF-b signaling, through increased TGFBR2 expression, results in reduced cell growth in vitro and disables the ability of the cells to form tumors in vivo (Turco et al., 2000). Instead, cells assume a terminally differentiated neuronal phenotype and display increased expression of axonal growth-associated protein (GAP43) and neurofilaments (Turco et al., 2000). Treatment of NB cells with TGF-b1 induces a similar phenotype (Scarpa et al., 1996). In addition, retinoic acid (RA) induces differentiation of NB cells, known to downregulate MYCN, accompanied by the increased expression of TGF-b1, TGFBR1, TGFBR2, and TGFBR3, resulting in the induction of a negative autocrine TGF-b1 growth regulatory loop (Cohen et al., 1995). We have shown that aggressive NB tumors evade the cytostatic TGF-b pathway through mir directed targeting of key components of the pathway as well as downstream effectors. Reactivation of TGF-b signaling through mir inhibition could be a promising therapeutic approach, as it would not only result in reactivation of TGFBR2 expression but also relieve the direct mir mediated repression of TGF-b-responsive genes. EXPERIMENTAL PROCEDURES Cell Culture SHEP-TR-miR cells (Mestdagh et al., 2009a) were cultured in RPMI (Invitrogen) supplemented with 10% fetal calf serum unless stated otherwise. SHEP-TR-miR cells were treated with 2 mg/ml tetracycline (Sigma-Aldrich) to induce mir expression (Figure S2A). TGF-b1 (PeproTech) and TGFBR1 inhibitor (SB431542, Sigma-Aldrich) were used at a concentration of 0.25 ng/ml and 2 mm, respectively, unless stated otherwise. COFRADIC Analysis SHEP-TR-miR cells were metabolically labeled by growing them in DMEM medium supplemented with dialyzed fetal calf serum and with either heavy lysine and arginine (both 13 C 6 ) or with natural, light lysine and arginine ( 12 C 6 ). This stable isotope labeling (SILAC [Ong et al., 2002]) ensures that following trypsin digestion, all generated peptides can be quantified by mass spectrometry (MS, see Supplemental Information). Mass spectrometry data for the forward en reverse experiment are available in Tables S4 and S5 and in the PRIDE database ( Accession number 14860). mrna and mirna Expression Quantification See Supplemental Information for details on mrna and mirna quantification and data normalization. mirna expression data are available in rdml format (Document S2) (Lefever et al., 2009). Immunohistochemistry and Western Blot Briefly, SHEP-TR-miR cells, tetracycline treated or untreated, were stimulated with TGF-b1 for 4 hr. psmad2 activity was evaluated by immunochemistry on cytopreparations or by western blot. See Supplemental Information for detailed experimental procedures. Cell Adhesion and Proliferation Assays Details on cell adhesion and proliferation assays are described in the Supplemental Information. Xenografts SHEP-TR-miR and SHEP-TR (control) cells were transfected with a luciferase expressing mammalian vector. Etherotopic xenografts were established in atymic nude mice (n = 5) by injection of 10 6 SHEP-TR cells subcutaneosly in the left flanking site and 10 6 SHEP-TR-miR cells in the right flanking site of each individual animal. See Supplemental Information for detailed experimental procedures. CAGA-Luciferase Reporter Assay For luciferase experiments, tetracycline or control treated SHEP-TR-miR cells were transfected with the (CAGA) 12 -Luc luciferase reporter vector and assayed for luciferase and renilla activity. See Supplemental Information for detailed experimental procedures. 3 0 UTR Reporter Assay DLD1Dicer hypo cells were seeded in DMEM (Invitrogen) supplemented with fetal calf serum (10%) at a density of 10,000 cells per well in an opaque 96-well plate. Twenty-four hours after seeding, using DharmaFECT Duo (Dharmacon), cells were cotransfected, either with a combination of a 3 0 UTR containing pgl4.11[luc2p] vector (Switchgear Genomics), a prl-tk vector (Promega) for normalization, and a mir pre-mir (Ambion) (10 nm) or with a combination of a psi-check2 vector (Promega) containing only part of the 3 0 UTR and a mir pre-mir. Forty-eight hours after transfection, luciferase reporter gene activity was measured using the Dual-Glo Luciferase Assay System (Promega) and a FLUOstar OPTIMA microplate reader (BMG LABTECH). See Supplemental Information for details on plasmid construction and mirna binding site mutation. Statistics See Supplemental Information for details on all statistical procedures and gene set enrichment analysis. ACCESSION NUMBERS The Gene Expression Omnibus accession number for the mrna expression data reported in this paper is GSE The PRIDE accession number for the protein expression data reported in this paper is SUPPLEMENTAL INFORMATION Supplemental Information includes seven figures, five tables, Supplemental Experimental Procedures, Supplemental References, and an RDML file for (E) Relative 3 0 UTR luciferase reporter activity for TGFBR2, SMAD2, and SMAD4, measured in DLD1DICER hypo cells (mean ± SEM). Plasmids with a wild-type seed site for the active mirna were introduced in DLD1DICER hypo cells in combination with a pre-mir negative control (NC) or mir pre-mir. Luciferase activity is decreased significantly in the presence of the active mirna (*) (t test, p < 0.01) and increases significantly when the seed for the active mirna is mutated (MUT) (t test, p < 0.01). Molecular Cell 40, , December 10, 2010 ª2010 Elsevier Inc. 771
118 112 Molecular Cell mir Dampens TGF-b Signaling RT-qPCR profiling and can be found with this article online at doi: /j. molcel ACKNOWLEDGMENTS This research was funded by the Fund for Scientific Research (grant number: G and ), the Belgian Kid s Fund, and the Stichting tegen Kanker. P.M. is supported by the Ghent University Research Fund (BOF 01D31406). A.-K. B., K.S., and H.A. are supported by grants from the Swedish Childhood Cancer Foundation and the Swedish Cancer Society. E.F. is supported by The Royal Swedish Physiographic Society and the American Cancer Society. B.G. is a Postdoctoral Research Fellow for the Fund for Scientific Research- Flanders (Belgium). The VIB/UGent lab further acknowledges support by a research grant from the Fund for Scientific Research- Flanders (Belgium) (project numbers G ), the Concerted Research Actions (project BOF07/GOA/012) from the Ghent University, and the Inter University Attraction Poles (IUAP06). NCI grants R01 CA and P30 CA were to A.T.-T., and FP7-Tumic HEALTH-F , Associazione Italiana contro la lotta al Neuroblastoma Progetto Pensiero and AIRC Tumori Pediatrici to M.Z. G.V.P. was supported by a BOF research grant (01D35609). This article represents research results of the Belgian program of Interuniversity Poles of Attraction, initiated by the Belgian State, Prime Minister s Office, Science Policy Programming. The study was sponsored by the GOA (01G01910). Received: April 22, 2010 Revised: October 6, 2010 Accepted: November 22, 2010 Published: December 9, 2010 REFERENCES Baek, D., Villén, J., Shin, C., Camargo, F.D., Gygi, S.P., and Bartel, D.P. (2008). The impact of micrornas on protein output. Nature 455, Bartel, D.P. (2004). MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, Bartel, D.P. (2009). MicroRNAs: target recognition and regulatory functions. Cell 136, Calin, G.A., and Croce, C.M. (2006). MicroRNA signatures in human cancers. Nat. Rev. Cancer 6, Castellano, L., Giamas, G., Jacob, J., Coombes, R.C., Lucchesi, W., Thiruchelvam, P., Barton, G., Jiao, L.R., Wait, R., Waxman, J., et al. (2009). The estrogen receptor-alpha-induced microrna signature regulates itself and its transcriptional response. Proc. Natl. Acad. Sci. USA 106, Cohen, P.S., Letterio, J.J., Gaetano, C., Chan, J., Matsumoto, K., Sporn, M.B., and Thiele, C.J. (1995). Induction of transforming growth factor beta 1 and its receptors during all-trans-retinoic acid (RA) treatment of RA-responsive human neuroblastoma cell lines. Cancer Res. 55, Colaert, N., Helsens, K., Impens, F., Vandekerckhove, J., and Gevaert, K. (2010). Rover: a tool to visualize and validate quantitative proteomics data from different sources. Proteomics 10, Dews, M., Fox, J.L., Hultine, S., Sundaram, P., Wang, W., Liu, Y.Y., Furth, E., Enders, G.H., El-Deiry, W., Schelter, J.M., et al. (2010). The myc-mir-1792 axis blunts TGFbeta signaling and production of multiple TGFbeta-dependent antiangiogenic factors. Cancer Res. 70, Dews, M., Homayouni, A., Yu, D., Murphy, D., Sevignani, C., Wentzel, E., Furth, E.E., Lee, W.M., Enders, G.H., Mendell, J.T., and Thomas-Tikhonenko, A. (2006). Augmentation of tumor angiogenesis by a Myc-activated microrna cluster. Nat. Genet. 38, Esquela-Kerscher, A., and Slack, F.J. (2006). Oncomirs - micrornas with a role in cancer. Nat. Rev. Cancer 6, Fontana, L., Fiori, M.E., Albini, S., Cifaldi, L., Giovinazzi, S., Forloni, M., Boldrini, R., Donfrancesco, A., Federici, V., Giacomini, P., et al. (2008). Antagomir-17-5p abolishes the growth of therapy-resistant neuroblastoma through p21 and BIM. PLoS ONE 3, e2236. Fredlund, E., Ringnér, M., Maris, J.M., and Påhlman, S. (2008). High Myc pathway activity and low stage of neuronal differentiation associate with poor outcome in neuroblastoma. Proc. Natl. Acad. Sci. USA 105, Friedman, R.C., Farh, K.K., Burge, C.B., and Bartel, D.P. (2009). Most mammalian mrnas are conserved targets of micrornas. Genome Res. 19, Gevaert, K., Van Damme, J., Goethals, M., Thomas, G.R., Hoorelbeke, B., Demol, H., Martens, L., Puype, M., Staes, A., and Vandekerckhove, J. (2002). Chromatographic isolation of methionine-containing peptides for gelfree proteome analysis: identification of more than 800 Escherichia coli proteins. Mol. Cell. Proteomics 1, Hayashita, Y., Osada, H., Tatematsu, Y., Yamada, H., Yanagisawa, K., Tomida, S., Yatabe, Y., Kawahara, K., Sekido, Y., and Takahashi, T. (2005). A polycistronic microrna cluster, mir-17-92, is overexpressed in human lung cancers and enhances cell proliferation. Cancer Res. 65, He, L., Thomson, J.M., Hemann, M.T., Hernando-Monge, E., Mu, D., Goodson, S., Powers, S., Cordon-Cardo, C., Lowe, S.W., Hannon, G.J., and Hammond, S.M. (2005). A microrna polycistron as a potential human oncogene. Nature 435, He, L., He, X., Lim, L.P., de Stanchina, E., Xuan, Z., Liang, Y., Xue, W., Zender, L., Magnus, J., Ridzon, D., et al. (2007). A microrna component of the p53 tumour suppressor network. Nature 447, Lanza, G., Ferracin, M., Gafà, R., Veronese, A., Spizzo, R., Pichiorri, F., Liu, C.G., Calin, G.A., Croce, C.M., and Negrini, M. (2007). mrna/microrna gene expression profile in microsatellite unstable colorectal cancer. Mol. Cancer 6, 54. Laping, N.J., Grygielko, E., Mathur, A., Butter, S., Bomberger, J., Tweed, C., Martin, W., Fornwald, J., Lehr, R., Harling, J., et al. (2002). Inhibition of transforming growth factor (TGF)-beta1-induced extracellular matrix with a novel inhibitor of the TGF-beta type I receptor kinase activity: SB Mol. Pharmacol. 62, Lefever, S., Hellemans, J., Pattyn, F., Przybylski, D.R., Taylor, C., Geurts, R., Untergasser, A., and Vandesompele, J.; RDML consortium. (2009). RDML: structured language and reporting guidelines for real-time quantitative PCR data. Nucleic Acids Res. 37, Lovén, J., Zinin, N., Wahlström, T., Müller, I., Brodin, P., Fredlund, E., Ribacke, U., Pivarcsi, A., Påhlman, S., and Henriksson, M. (2010). MYCN-regulated micrornas repress estrogen receptor-alpha (ESR1) expression and neuronal differentiation in human neuroblastoma. Proc. Natl. Acad. Sci. USA 107, Mestdagh, P., Fredlund, E., Pattyn, F., Schulte, J.H., Muth, D., Vermeulen, J., Kumps, C., Schlierf, S., De Preter, K., Van Roy, N., et al. (2009a). MYCN/c- MYC-induced micrornas repress coding gene networks associated with poor outcome in MYCN/c-MYC-activated tumors. Oncogene 29, Mestdagh, P., Van Vlierberghe, P., De Weer, A., Muth, D., Westermann, F., Speleman, F., and Vandesompele, J. (2009b). A novel and universal method for microrna RT-qPCR data normalization. Genome Biol. 10, R64. O Donnell, K.A., Wentzel, E.A., Zeller, K.I., Dang, C.V., and Mendell, J.T. (2005). c-myc-regulated micrornas modulate E2F1 expression. Nature 435, Oberthuer, A., Berthold, F., Warnat, P., Hero, B., Kahlert, Y., Spitz, R., Ernestus, K., König, R., Haas, S., Eils, R., et al. (2006). Customized oligonucleotide microarray gene expression-based classification of neuroblastoma patients outperforms current clinical risk stratification. J. Clin. Oncol. 24, Olive, V., Bennett, M.J., Walker, J.C., Ma, C., Jiang, I., Cordon-Cardo, C., Li, Q.J., Lowe, S.W., Hannon, G.J., and He, L. (2009). mir-19 is a key oncogenic component of mir Genes Dev. 23, Ong, S.E., Blagoev, B., Kratchmarova, I., Kristensen, D.B., Steen, H., Pandey, A., and Mann, M. (2002). Stable isotope labeling by amino acids in cell culture, 772 Molecular Cell 40, , December 10, 2010 ª2010 Elsevier Inc.
119 113 Molecular Cell mir Dampens TGF-b Signaling SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell. Proteomics 1, Padua, D., Zhang, X.H., Wang, Q., Nadal, C., Gerald, W.L., Gomis, R.R., and Massagué, J. (2008). TGFbeta primes breast tumors for lung metastasis seeding through angiopoietin-like 4. Cell 133, Petrocca, F., Visone, R., Onelli, M.R., Shah, M.H., Nicoloso, M.S., de Martino, I., Iliopoulos, D., Pilozzi, E., Liu, C.G., Negrini, M., et al. (2008). E2F1-regulated micrornas impair TGFbeta-dependent cell-cycle arrest and apoptosis in gastric cancer. Cancer Cell 13, Raver-Shapira, N., Marciano, E., Meiri, E., Spector, Y., Rosenfeld, N., Moskovits, N., Bentwich, Z., and Oren, M. (2007). Transcriptional activation of mir-34a contributes to p53-mediated apoptosis. Mol. Cell 26, Scarpa, S., Coppa, A., Ragano-Caracciolo, M., Mincione, G., Giuffrida, A., Modesti, A., and Colletta, G. (1996). Transforming growth factor beta regulates differentiation and proliferation of human neuroblastoma. Exp. Cell Res. 229, Schweigerer, L., Breit, S., Wenzel, A., Tsunamoto, K., Ludwig, R., and Schwab, M. (1990). Augmented MYCN expression advances the malignant phenotype of human neuroblastoma cells: evidence for induction of autocrine growth factor activity. Cancer Res. 50, Selbach, M., Schwanhäusser, B., Thierfelder, N., Fang, Z., Khanin, R., and Rajewsky, N. (2008). Widespread changes in protein synthesis induced by micrornas. Nature 455, Subramanian, A., Tamayo, P., Mootha, V.K., Mukherjee, S., Ebert, B.L., Gillette, M.A., Paulovich, A., Pomeroy, S.L., Golub, T.R., Lander, E.S., and Mesirov, J.P. (2005). Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, Turco, A., Scarpa, S., Coppa, A., Baccheschi, G., Palumbo, C., Leonetti, C., Zupi, G., and Colletta, G. (2000). Increased TGFbeta type II receptor expression suppresses the malignant phenotype and induces differentiation of human neuroblastoma cells. Exp. Cell Res. 255, Verrecchia, F., Chu, M.L., and Mauviel, A. (2001). Identification of novel TGF-beta /Smad gene targets in dermal fibroblasts using a combined cdna microarray/promoter transactivation approach. J. Biol. Chem. 276, Volinia, S., Calin, G.A., Liu, C.G., Ambs, S., Cimmino, A., Petrocca, F., Visone, R., Iorio, M., Roldo, C., Ferracin, M., et al. (2006). A microrna expression signature of human solid tumors defines cancer gene targets. Proc. Natl. Acad. Sci. USA 103, Xiao, C., Srinivasan, L., Calado, D.P., Patterson, H.C., Zhang, B., Wang, J., Henderson, J.M., Kutok, J.L., and Rajewsky, K. (2008). Lymphoproliferative disease and autoimmunity in mice with increased mir expression in lymphocytes. Nat. Immunol. 9, Zhang, H., Gao, P., Fukuda, R., Kumar, G., Krishnamachary, B., Zeller, K.I., Dang, C.V., and Semenza, G.L. (2007). HIF-1 inhibits mitochondrial biogenesis and cellular respiration in VHL-deficient renal cell carcinoma by repression of C-MYC activity. Cancer Cell 11, Molecular Cell 40, , December 10, 2010 ª2010 Elsevier Inc. 773
120 Supplemental Data 114 COFRADIC analysis To reduce arginine to proline conversion and thus dilution of the 13C- label, the arginine concentration was lowered to 30% of its normal concentration in DMEM. Cell cultures were then treated with tetracycline for 72 hours to induce mir expression. A biological replicate was created by swapped labeling. Cells were harvested by Versene- EDTA and washed with PBS. Cell pellets were frozen at - 80ºC until further use. Prior to proteome analysis, cells were lysed in 250 µl lysis buffer containing 0.8% CHAPS in 50 mm HEPES (ph 7.4), 100 mm NaCl and 0.5 mm EDTA supplemented with protease inhibitors (Complete Protease inhibitor cocktail tablet (Roche, Basel, Switzerland); one tablet per 100 ml buffer)) for 20 minutes on ice. Cell debris was removed by centrifugation for 15 min at 16,000g at 4ºC after which the protein concentration was measured using the Biorad Protein Assay. Then, equal amounts of both proteome preparations (i.e. from differently labeled control cells or cells in which mir expression was induced) were mixed together. To denature proteins, solid guanidinium hydrochloride was added to a final concentration of 4 M (the total sample volume was 1 ml). Proteins were then reduced and S- alkylated for 60 minutes at 30ºC by adding tris(2- carboxyethyl) phosphine and iodoacetamide to final concentrations of 3 mm and 6 mm respectively. Half of each proteome sample was then desalted on a NAP- 5 column in 1 ml of 20 mm triethylamonium bicarbonate (ph 8.0). These desalted protein mixtures were heated for 5 min at 95 C, put on ice for 5 min, after which trypsin (sequencing grade, modified porcine trypsin, Promega Corporation, Madison, WI, USA) was added to a trypsin/substrate ratio of 1/50 (w/w). Trypsin digestion proceeded overnight at 37 C after which the sample was dried in vacuo. Dried peptides were then re- dissolved in 105 µl of solvent A (10 mm ammonium acetate (ph 5.5) in water/acetonitrile (98/2 (v/v), both Baker HPLC analysed, Mallinckrodt Baker B.V., Deventer, The Netherlands)) and 100 µl of this peptide mixture was used to isolate methionine- containing peptides by the COFRADIC technology as described previously (Gevaert et al., 2002). In this way, the complexity of the peptide mixture was reduced by a factor of about five and the hence isolated peptide mixture which is highly enriched for methionine- containing peptides were analyzed by LC- MS/MS using an Orbitrap XL mass spectrometer (Thermo Electron, Bremen, Germany) that was operated as previously described (Ghesquiere et al., 2009). Generated MS/MS spectra were converted to MS/MS peak lists as described (Ghesquiere et al., 2009) and these peak lists were searched in the human fraction of the Swiss- Prot database (version 56.4, containing 20,408 human protein sequences) using a locally installed version of the MASCOT database search engine (version (Perkins et al., 1999)). Carbamidomethylation of cysteine and oxidation of methionine were set as fixed MASCOT parameters, and acetylation of a protein s N- terminus, pyro- carbamidomethyl cysteine (from N- term cysteine), pyroglutamate formation (N- term glutamine) were considered as variable modifications. Trypsin/P was set as the protease with one missed cleavage allowed and MASCOT s
121 115 quantitation parameters were set to SILAC Arg and Lys + 6. Only peptide identifications that were ranked one, scored above the corresponding MASCOT threshold score for identity set at 99% confidence were considered identified. The false discovery rate of these identifications was determined according to the method described by Elias and Gygi (Elias and Gygi, 2007) and found to be 0.21% (on the spectrum level). Quantification of the identified peptides was then done using MASCOT Distiller Quantitation Toolbox ( in the precursor mode. The software tried to fit an ideal isotopic distribution on the experimental data based on the peptide average amino acid composition. This was followed by extraction of the XIC signal of both peptide components (light and heavy) from the raw data. Ratios were calculated from the area below the light and heavy isotopic envelope of the corresponding peptide (integration method trapezium, integration source survey ). To calculate this ratio value, a least squares fit to the component intensities from the different scans in the XIC peak was created. MS scans used for this ratio calculation were situated in the elution peak of the precursor determined by the Distiller software (XIC threshold 0.3, XIC smooth 1, Max XIC width 250). To validate the calculated ratio, the standard error on the least square fit had to be below 0.14 and correlation coefficient of the isotopic envelope needed to be above The number of recorded and identified spectra as well as the number of unique peptide quantifications is indicated for both analyses in the table below. All identified spectra are made publically available in the PRIDE database ( accession # MS/MS # MS/MS # unique analysis spectra recorded spectra identified peptide quantifications forward reversed Next, protein ratios were calculated as the average of individual peptide ratios by the in- house developed Rover algorithm (Colaert et al., 2010). Peptide ratios that could not be adequately calculated by the Distiller software were also manually validated using the Rover application (typically belonging to highly regulated peptides and proteins). Supplemental tables 4 and 5 show the Rover output and contain all peptide quantifications for the forward and reversed experiment. In case a peptide could be derived from more than one protein sequence, all protein isoforms are listed which explains why the number of peptide quantifications in these lists exceeds the number of unique peptide quantifications in the table above. Peptide ratios of the repeated experiments were averaged and proteins that were quantified by at least 2 peptides were selected for further analysis. UniProtKB accessions were mapped to RefSeq IDs using the Biomart tool from Ensembl (
122 116 RT- qpcr 2 µg of RNA from each sample was treated with RQ1 DNase I (Promega) and desalted using a Microcon- 100 spin column (Millipore). cdna synthesis was performed on the eluate with the iscript cdna synthesis kit (Bio- Rad). All manipulations were conducted according to the manufacturer s instructions. First strand cdna was diluted to a final concentration of 5 ng/µl (total RNA equivalent). RT- PCR amplification reactions were carried out in a total volume of 7.5 µl, containing 10 ng of template cdna, 3.75 µl of 2x SYBR Green I reaction mix (Eurogentec), 1 µl nuclease- free water (Sigma) and µl of a 5 µm solution of each primer. Cycling conditions were as follows: 10 min at 95 C followed by 45 cycles of denaturation (10s at 95 C) and elongation (45s at 60 C). All reactions were performed on a LightCycler 480 (Roche). Primers were designed using Primer3 (Rozen and Skaletsky, 2000) and validated through RTPimerDB s in silico assay evaluation pipeline (Lefever et al., 2009). Raw Cq values were imported into qbaseplus (Hellemans et al., 2007) ( and normalized using a selection of stably expressed reference genes (UBC, SDHA, GAPDH and Alu- sx). Primer sequences for UBC, SDHA and GAPDH are available in the public RTprimerDB database ( (gene (RTPrimerDB- ID): UBC(8), SDHA(7), GAPDH(3)). For Alu- sx, the following primer sequences were used: F: TGGTGAAACCCCGTCTCTACTAA, R: CCTCAGCCTCCCGAGTAGCT. mirna expression profiling and data normalization were performed as described previously (Mestdagh et al., 2008; Mestdagh et al., 2009b). Pre- mir transfection Individual pre- mir molecules (Ambion) were transfected in neuroblastoma SHEP- cells at a concentration of 100 nm using Xtremegene transfection reagent (Roche) according to the manufacturer s instructions. Pre- mir negative control #1 (Ambion) was used as a scrambled control. Transfection efficiency was monitored by flow cytometry using a fluorescently labeled pre- mir (Ambion) and estimated to be 80% or higher. Cells were cultured in RPMI (Invitrogen), 10% FCS in the absence of antibiotics. Cells were harvested for RNA isolation (mirneasy, Qiagen) 24h post transfection. 3 UTR luciferase constructs 3 UTR luciferase reporter constructs for TGFBR2 were obtained from Switchgear Genomics. For SMAD2 and SMAD4, 74 bp oligonucleotides spanning the predicted 3 UTR mirna binding site and flanked by XhoI and NotI restriction sites were cloned into psicheck2 (Promega) as described previously (Cloonan et al., 2008). The following oligonucleotides were used: SMAD2_miR- 18a_F TCGAGAAAACAGCACTTGAGGTCTCATCAATTAAAGCACCTTGTGGAATCTGTTTCCTATATTTGAATATTAGC SMAD2_miR- 18a_R GGCCGCTAATATTCAAATATAGGAAACAGATTCCACAAGGTGCTTTAATTGATGAGACCTCAAGTGCTGTTTTC
123 117 SMAD2_miR- 19_F TCGAGCCTTCCTCAACCTTTGCTGTAAAAATTTCATTTGCACCACATCAGTACTACTTAATTTAACAAGCTTGC SMAD2_miR- 19_R GGCCGCAAGCTTGTTAAATTAAGTAGTACTGATGTGGTGCAAATGAAATTTTTACAGCAAAGGTTGAGGAAGGC SMAD2_miR- 92a_F TCGAGTTTTTTTCTCTGATGGCATTAACTTTGTAATGCAATATGATGGATGCAGACCCTGTTCTTGTTTCCCGC SMAD2_miR- 92a_R GGCCGCGGGAAACAAGAACAGGGTCTGCATCCATCATATTGCATTACAAAGTTAATGCCATCAGAGAAAAAAAC SMAD4_miR- 18a_F TCGAGAAGACTTAATTTTAACCAAAGGCCTAGCACCACCTTAGGGGCTGCAATAAACACTTAACGCGCGCACGC SMAD4_miR- 18a_R GGCCGCGTGCGCGCGTTAAGTGTTTATTGCAGCCCCTAAGGTGGTGCTAGGCCTTTGGTTAAAATTAAGTCTTC SMAD4_ mir- 19_miR- 17/20_ F TCGAGGTTTGATTTTTAAGATTTTTTTTTTCTTTTGCACTTTTGAGTCCAATCTCAGTGATGAGGTACCTTCGC SMAD4_ mir- 19_miR- 17/20_ R GGCCGCGAAGGTACCTCATCACTGAGATTGGACTCAAAAGTGCAAAAGAAAAAAAAAATCTTAAAAATCAAACC SMAD2_miR- 18a_mut_F TCGAGAAAACAGCACTTGAGGTCTCATCAATTAAATCCAATTGTGGAATCTGTTTCCTATATTTGAATATTAGC SMAD2_miR- 18a_mut_R GGCCGCTAATATTCAAATATAGGAAACAGATTCCACAATTGGATTTAATTGATGAGACCTCAAGTGCTGTTTTC SMAD4_miR- 18a_mut_F TCGAGAAGACTTAATTTTAACCAAAGGCCTAGCACTACTTTCGGGGCTGCAATAAACACTTAACGCGCGCACGC SMAD4_miR- 18a_mut_R GGCCGCGTGCGCGCGTTAAGTGTTTATTGCAGCCCCGAAAGTAGTGCTAGGCCTTTGGTTAAAATTAAGTCTTC For TGFBR2, mirna binding sites were mutated using the Quickchange site- directed mutagenesis kit (Agilent) according to the manufacturer s instructions. The following primers were used: TGFBR2_miR- 17/20_1_F GATTGATTTTTACAATAGCCAATAACATTTTCCAGTTATTAATGCCTGTATATAAATATGAATAGCTA TGFBR2_miR- 17/20_1_R TAGCTATTCATATTTATATACAGGCATTAATAACTGGAAAATGTTATTGGCTATTGTAAAAATCAATC TGFBR2_miR- 17/20_2_F GGTCAGCACAGCGTTTCAAAAAGTGAAGCAAAGGTATAAATATTTGGAGATTTTGCAGGAAAA TGFBR2_miR- 17/20_2_R TTTTCCTGCAAAATCTCCAAATATTTATACCTTTGCTTCACTTTTTGAAACGCTGTGCTGACC
124 118 mirna target site analysis Four mirna seed types were considered for mirna target site analyses: 6mer, 7mer- A1, 7mer- m8 and 8mer sites (Grimson et al., 2007). Target sites were identified using a custom Perl script. 5 UTR, CDS and 3 UTR sequences were taken from Baek et al. (Baek et al., 2008). Candidate mir target gene selection Genes with a protein expression fold change below (log2) and at least one 3 UTR mir site were selected as candidate mir target genes. The expression fold change cutoff of 0.5 was selected as the fold change that deviated from the linear fit in a normal QQ- plot. Throughout the manuscript, downregulated proteins are defined by a log2 expression fold change < - 0.5, upregulated proteins by a log2 expression fold change > 0.5. Xenografts SHEP- TR- mir and SHEP- TR (control) cells were transfected with a luciferase expressing mammalian vector. This vector was obtained by cloning the firefly luciferase gene under the control of a CMV promoter and enables bioluminescent imaging of viable cells in vivo. Stable clones obtained after 15 days of selection in 300ug/ml hygromicine (Sigma, USA) were evaluated for their bioluminescence signal by serial cellular dilutions incubated with the luciferine substrate. The bioluminescence signal was acquired by an IVIS illumina 3D Imaging System (Xenogen Corp. Alameda, CA). Etherotopic xenografts were established in atymic nude mice (n=5) by injection of 106 SHEP- TR cells subcutaneosly in the left flanking site and 106 SHEP- TR- mir cells in the rigth flanking site of each individual animal. Mice were fed with tetracycline (from SIGMA) once a day using oral gavage prestige 200ul (2mg/ml in ddh2o). In vivo tumourigenic bioluminiscence imaging was performed by measuring bioluminescence (BLI, phothon/sec) of luciferine positive living cells at 0, 7, 14 and 21 days post injection, measuring the median value of emissions from three scans per animal at each flanking site. Immunohistochemistry and Western blot SHEP- TR- mir cells, tetracycline treated or untreated, were stimulated with TGFβ1 for 4 h. After harvesting, 200 μl of cell suspension were centrifuged on a cytospin glass using a cytospin chamber. The cytopreparations were air dried, fixed with 4% paraformaldehyde (Sigma Aldrich, Munich, Germany) for 10 min followed by 45 min in Tris- 1% Triton buffer. After antigen retrieval, psmad2 immunoreactivity (anti- psmad2, Cell Signaling) was detected using the Dako Auto- stainer Plus (Glostrup, Denmark) and the Dako EnVision Flex system (Glostrup, Denmark) according to the manufacturer s instructions. For Western blot analysis, cells were lysed in RIPA buffer, separated on a SDS PAGE gel and blotted onto Immobilon- P (Millipore, Bedford, MA, USA) membrane. The membrane was incubated with anti- phosphorylated SMAD2 (psmad2, Cell Signaling Technology) or anti- ACTIN (ICN Biomedicals, Aurora, OH, USA). HRP- conjugated
125 119 secondary antibodies were obtained from Amersham Biosciences. Proteins were detected by Super Signal chemiluminescence substrate (Pierce, Rockford, IL, USA). Images were processed using ImageJ software. Statistics All statistical analyses were performed using R Bioconductor software. For survival analysis, rank- based pathway scores were calculated as described previously (Fredlund et al., 2008; Mestdagh et al., 2009a). Samples (n) were ranked according to the expression level of each gene/mirna within the set and rank scores (ranging from 1 to n) were assigned. This was repeated for each gene in the gene set. Next, rank scores were summed generating an activity score of the gene/mirna set for each sample. Kaplan- Meier analysis was performed using pathway activity score quartiles. MiRNA target site overrepresentation was evaluated using Fisher s Exact test in combination with Bonferroni multiple testing correction. Differential expression/activity was evaluated using the Mann- Whitney test, unless stated otherwise. For the identification of relevant biological pathways among the up- and down- regulated proteins, a gene list ranked according to protein expression was analyzed using GSEA with the chemical and genetic perturbations collection (Subramanian et al., 2005). Gene lists with a false discovery rate (FDR) below 5% were considered significant. For calculation of individual mir mirna contributions to the significant gene lists the contributing genes of each gene list were analyzed for the presence of 7mer and 8mer mir sites. For each gene list, the fraction of targets per individual mir mirna was calculated. Data was log transformed and standardized before hierarchical clustering (method: Ward, distance: Manhattan). Gene lists with two or more missing values were excluded for clustering purposes. Cell adhesion and proliferation assays To evaluate cell adhesion, ells were quickly washed in Versene and then incubated at 37 C in the presence of Versene. After 15 min, the cells were visualized under a microscope to assure that the cell- cell contacts were disrupted. The cells were then counted, centrifuged and suspended in medium (RPMI, 10% FCS) at a concentration of 2x106 cells/ml and thereafter incubated on a rotating platform at 37 C for 1 h and then analyzed. Each treatment was assayed in triplicate and repeated three times. Pictures were processed using ImageJ software. For cell proliferation, SHEP- TR- mir cells were trypsinized and seeded in 96- well xcelligence E- plates (Roche) (10000 cells/well) according to the manufacturer s instructions. After 24 h, cells were either treated with tetracycline or left untreated and were monitored in real- time on the xcelligence system. Five replicate measurements per condition were obtained. SMAD2/SMAD4 overexpression For overexpression of SMAD2 and SMAD4 SHEP- TR- mir cells were seeded in 6- well plates 12 hours prior to transfection at a density of 100,000 cells per well. The cells were then transfected with either pflag- SMAD2, pflag- SMAD4 (400 ng/well, respectively) and pegfp- C1 (Clontech, USA) vectors (200 ng/well) or
126 120 control pha- CMV (Clontech, USA) (800 ng/well) and pegfp- C1 vectors (200 ng/well) using Lipofectamine transfection reagent in OptiMEM I Reduced Serum Medium. The cells were thereafter grown for 48 hours in RPMI (Invitrogen) supplemented with fetal calf serum (1%) tetracycline containing medium. GFP transfection and total cell count was assessed using the Nucleocounter 3000 system (Chemotek, Denmark). CAGA- Luciferase reporter assay For luciferase experiments, tetracycline or control treated SHEP- TR- mir cells were seeded in 96- well plates 12 hours prior to transfection at a density of cells per well. The cells were then transfected with the (CAGA)12- Luc luciferase reporter vector containing twelve CAGA SMAD binding sites (Dennler et al., 1998) (400 ng/well) using Lipofectamine 2000 transfection reagent in OptiMEM I Reduced Serum Medium. Following transfection, cells were treated as indicated in the figure legends. Cells were lysed and assayed for luciferase and renilla activity using the Dual- Luciferase Reporter Assay System (Promega, Madison, WI, USA) and a TD- 20/20 Luminometer (Turner Biosystems, Sunnyvale, CA, USA). Supplemental References Baek, D., Villen, J., Shin, C., Camargo, F.D., Gygi, S.P., and Bartel, D.P. (2008). The impact of micrornas on protein output. Nature 455, Cloonan, N., Brown, M.K., Steptoe, A.L., Wani, S., Chan, W.L., Forrest, A.R., Kolle, G., Gabrielli, B., and Grimmond, S.M. (2008). The mir- 17-5p microrna is a key regulator of the G1/S phase cell cycle transition. Genome Biol 9, R127. Colaert, N., Helsens, K., Impens, F., Vandekerckhove, J., and Gevaert, K. (2010). Rover: a tool to visualize and validate quantitative proteomics data from different sources. Proteomics 10, Dennler, S., Itoh, S., Vivien, D., ten Dijke, P., Huet, S., and Gauthier, J.M. (1998). Direct binding of Smad3 and Smad4 to critical TGF beta- inducible elements in the promoter of human plasminogen activator inhibitor- type 1 gene. EMBO J 17, Elias, J.E., and Gygi, S.P. (2007). Target- decoy search strategy for increased confidence in large- scale protein identifications by mass spectrometry. Nat Methods 4, Fredlund, E., Ringner, M., Maris, J.M., and Pahlman, S. (2008). High Myc pathway activity and low stage of neuronal differentiation associate with poor outcome in neuroblastoma. Proc Natl Acad Sci U S A 105, Gevaert, K., Van Damme, J., Goethals, M., Thomas, G.R., Hoorelbeke, B., Demol, H., Martens, L., Puype, M., Staes, A., and Vandekerckhove, J. (2002). Chromatographic isolation of methionine- containing peptides for gel- free proteome analysis: identification of more than 800 Escherichia coli proteins. Mol Cell Proteomics 1,
127 121 Ghesquiere, B., Colaert, N., Helsens, K., Dejager, L., Vanhaute, C., Verleysen, K., Kas, K., Timmerman, E., Goethals, M., Libert, C., et al. (2009). In vitro and in vivo protein- bound tyrosine nitration characterized by diagonal chromatography. Mol Cell Proteomics 8, Grimson, A., Farh, K.K., Johnston, W.K., Garrett- Engele, P., Lim, L.P., and Bartel, D.P. (2007). MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol Cell 27, Hellemans, J., Mortier, G., De Paepe, A., Speleman, F., and Vandesompele, J. (2007). qbase relative quantification framework and software for management and automated analysis of real- time quantitative PCR data. Genome Biol 8, R19. Lefever, S., Vandesompele, J., Speleman, F., and Pattyn, F. (2009). RTPrimerDB: the portal for real- time PCR primers and probes. Nucleic Acids Res 37, D Mestdagh, P., Feys, T., Bernard, N., Guenther, S., Chen, C., Speleman, F., and Vandesompele, J. (2008). High- throughput stem- loop RT- qpcr mirna expression profiling using minute amounts of input RNA. Nucleic Acids Res 36, e143. Mestdagh, P., Fredlund, E., Pattyn, F., Schulte, J.H., Muth, D., Vermeulen, J., Kumps, C., Schlierf, S., De Preter, K., Van Roy, N., et al. (2009a). MYCN/c- MYC- induced micrornas repress coding gene networks associated with poor outcome in MYCN/c- MYC- activated tumours. Oncogene. Mestdagh, P., Van Vlierberghe, P., De Weer, A., Muth, D., Westermann, F., Speleman, F., and Vandesompele, J. (2009b). A novel and universal method for microrna RT- qpcr data normalization. Genome Biol 10, R64. Oberthuer, A., Berthold, F., Warnat, P., Hero, B., Kahlert, Y., Spitz, R., Ernestus, K., Konig, R., Haas, S., Eils, R., et al. (2006). Customized oligonucleotide microarray gene expression- based classification of neuroblastoma patients outperforms current clinical risk stratification. J Clin Oncol 24, Perkins, D.N., Pappin, D.J., Creasy, D.M., and Cottrell, J.S. (1999). Probability- based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, Rozen, S., and Skaletsky, H. (2000). Primer3 on the WWW for general users and for biologist programmers. Methods Mol Biol 132, Subramanian, A., Tamayo, P., Mootha, V.K., Mukherjee, S., Ebert, B.L., Gillette, M.A., Paulovich, A., Pomeroy, S.L., Golub, T.R., Lander, E.S., and Mesirov, J.P. (2005). Gene set enrichment analysis: a knowledge- based approach for interpreting genome- wide expression profiles. Proc Natl Acad Sci U S A 102,
128 122 Supplemental Figure 1 (A) Relative expression of individual mir mirnas in MYCN amplified tumours (A), MYCN single copy high- risk tumours (SH) and MYCN single copy low risk tumours (SL) (dataset D1, supplemental table 1). Significant differential expression (Mann Whitney, p < 0.05) is indicated (*) (whiskers: Tukey). (B) Kaplan Meier plots for overall survival (OS) based on the pathway activity score of individual mir mirnas, represented as quartiles (dataset D1, supplemental table 1). Increased activity of each individual member form the mir cluster is proportionally correlated to a poor overall survival. A B
129 123
130 Supplemental Figure (A) Relative expression of mir mirnas upon treatment of SHEP- TR- mir cells with tetracycline (mean ± SEM). (B) Protein quantification using LC/MS- MS. Histogram showing the number of quantified peptides per protein. (C) Histogram showing the distribution of protein fold changes of proteins that were quantified by at least 2 peptides. A
131 Supplemental Figure (A) Fraction of proteins containing at least one 7mer- 8mer mir site (black) of one 6mer- 7mer- 8mer mir site (grey) in function of the protein fold change (grouped in 6 different bins according to fold change). Only downregulated proteins are shown. Dashed horizontal lines indicate the background mir site occurrence (determined as the fraction of unchanged proteins harboring at least one 7mer- 8mer mir site of one 6mer- 7mer- 8mer mir site). (B) Average number of 3 UTR mir sites per protein in function of the protein fold change. Dashed lines indicate background measurement (calculated as in (A)). (C) Average number of mir sites per nucleotide in function of protein fold change for 3 UTR sites (red), 5 UTR sites (blue) and coding sequence sites (CDS) (green). (D) Average fold change ± SEM of proteins for which the transcripts contain at least one 8mer mir site in the 3 UTR (red), 5 UTR (blue) and CDS (green). (E) Number of co- occurring mir sites in the 3 UTR of transcripts from downregulated proteins. The X- axis indicates the number of co- occurring sites between the mirna listed on top of each graph and the mirnas listed below each bar (mir- 17/miR- 20a and mir- 19a/miR- 19b) are analyzed together as they share identical seeds. The first bar of each graph represents the number downregulated proteins that only have one (or more) sites for the respective mirna. Significant co- occurrence was determined by comparing results for downregulated proteins to results for a reference set (upregulated proteins) using Fisher s Exact test (p < 0.05).
132 E 126
133 Supplemental Figure Gene set enrichment analysis plots for gene sets enriched among the proteins, upregulated upon mir activation. Supplemental Figure 5 Relative mrna expression (mean ± SEM) of TGFβ- responsive genes upon TET treatment of SHEP- TR- mir cells.
134 Supplemental Figure (A) Correlation clustering between mir mirna expression and TGFβ target gene expression in 40 primary neuroblastoma tumours (dataset D3, supplemental table 1). The heatmap indicates the Spearman s rank rho value. TGFβ target genes are indicated by the grey sidebar, mir mirnas are indicated by the black sidebar. (B) Heatmap showing relative expression of TGFβ- pathway components and target genes in neuroblastoma SHEP cells transfected with a scrambled control or with pre- mirs for the individual mir mirnas (mir- 17, mir- 18a, mir- 19a, mir- 19b, mir- 20a and mir- 92a). Black dots mark genes that have at least one 3 UTR 7mer site for the corresponding mirna. A
135 B 129
136 Supplemental Figure Model for mir mediated TGFβ pathway repression. Supplemental Table 1. Overview of mrna and mirna expression datasets dataset ID samples mrna mirna reference D1 95 X (Mestdagh et al., 2009a) D2 251 X (Oberthuer et al., 2006) D3 40 X X (Mestdagh et al., 2009a)
137 131 Supplemental Table 3. mir seed occurrence and protein fold change for TGFB- target genes downregulated upon mir activation protein 3 UTR seed(s) fold change (log 2 ) SERPINE1 THBS1 ITGA4 mir- 17/miR- 20a mir- 19a/miR- 19b mir- 18a mir- 19a/miR- 19b mir- 17/miR- 20a mir- 19a/miR- 19b FNDC3B mir JUP none PPP1R13L mir- 19a/miR- 19b FILIP1L mir- 17/miR- 20a ICAM1 mir- 92a EPHA2 none COL1A1 none CDKN1A mir- 17/miR- 20a PFKFB3 mir- 17/miR- 20a mir- 19a/miR- 19b CTNNA1 mir- 18a
138 132 PAPER 5: An integrative genomics screen uncovers ncrna T- UCR functions in neuroblastoma tumours PAPER 5 An integrative genomics screen uncovers ncrna T-UCR functions in neuroblastoma tumours. Mestdagh P*, Fredlund E*, Pattyn F, Rihani A, Van Maerken T, Vermeulen J, Kumps C, Menten B, De Preter K, Schramm A, Schulte J, Noguera R, Schleiermacher G, Janoueix- Lerosey I, Laureys G, Powel R, Nittner D, Marine JC, Ringnér M, Speleman F, Vandesompele J. Oncogene Jun 17;29(24): *Equally contributing authors
139 133 ONCOGENOMICS An integrative genomics screen uncovers ncrna T-UCR functions in neuroblastoma tumours P Mestdagh 1,11, E Fredlund 1,2,3,11, F Pattyn 1, A Rihani 1, T Van Maerken 1, J Vermeulen 1, C Kumps 1, B Menten 1, K De Preter 1, A Schramm 4, J Schulte 4, R Noguera 5, G Schleiermacher 6,7, I Janoueix-Lerosey 7, G Laureys 8, R Powel 9, D Nittner 10, J-C Marine 10, M Ringne r 2,3, F Speleman 1 and J Vandesompele 1 1 Center for Medical Genetics, Ghent University Hospital, Ghent, Belgium; 2 Department of Oncology, Clinical Sciences, Lund University, Lund, Sweden; 3 CREATE Health, Strategic Centre for Translational Cancer Research, Lund University, Lund, Sweden; 4 University Hospital of Essen, Essen, Germany; 5 Medical School in Valencia, Valencia, Spain; 6 Department of Paediatric Oncology, Institut Curie, Paris, France; 7 INSERM U830, Institut Curie, Paris, France; 8 Department of Pediatric Oncology, Ghent University Hospital, Ghent, Belgium; 9 PrimerDesign, Southampton, UK and 10 Laboratory for Molecular Cancer Biology, VIB-UGent, Ghent, Belgium Different classes of non-coding RNAs, including micro- RNAs, have recently been implicated in the process of tumourigenesis. In this study, we examined the expression and putative functions of a novel class of non-coding RNAs known as transcribed ultraconserved regions (T-UCRs) in neuroblastoma. Genome-wide expression profiling revealed correlations between specific T-UCR expression levels and important clinicogenetic parameters such as MYCN amplification status. A functional genomics approach based on the integration of multi-level transcriptome data was adapted to gain insights into T-UCR functions. Assignments of T-UCRs to cellular processes such as TP53 response, differentiation and proliferation were verified using various cellular model systems. For the first time, our results define a T-UCR expression landscape in neuroblastoma and suggest widespread T-UCR involvement in diverse cellular processes that are deregulated in the process of tumourigenesis. Oncogene (2010) 29, ; doi: /onc ; published online 12 April 2010 Keywords: neuroblastoma; T-UCR; non-coding RNA Introduction Tumourigenesis is driven by (epi-)genetic alterations that result in gene expression changes. Besides proteincoding genes, these alterations also affect various classes of non-coding RNAs such as micrornas (mirnas) and long intergenic non-coding RNAs (lincrnas). mirnas function as negative regulators of gene expression through imperfect binding to target mrnas, Correspondence: Dr J Vandesompele, Center for Medical Genetics, Ghent University Hospital, MRB, De Pintelaan 185, Ghent, East- Flanders 9000, Belgium. [email protected] 11 These authors contributed equally to this work. Received 19 September 2009; revised 11 January 2010; accepted 4 March 2010; published online 12 April 2010 Oncogene (2010) 29, & 2010 Macmillan Publishers Limited All rights reserved /10 whereas lincrnas associate with chromatin-modifying complexes to alter gene expression (Bartel, 2009; Guttman et al., 2009; Khalil et al., 2009). Both mirnas and lincrnas have been implicated in a number of oncogenic and tumour-suppressor pathways (O Donnell et al., 2005; He et al., 2007; Guttman et al., 2009), whereby mirnas have been established as key components in tumour biology (Esquela-Kerscher and Slack, 2006). Recently, another class of non-coding RNAs called transcribed ultraconserved regions (T-UCRs) has been associated with the process of tumourigenesis (Calin et al., 2007). T-UCRs are transcribed from 481 ultraconserved regions (UCRs) defined as being at least 200 bp in length and 100% conserved between the human, mouse and rat genomes (Bejerano et al., 2004). Genomically, UCRs are located both within genes and in regions lacking apparent protein-coding features (Bejerano et al., 2004; Calin et al., 2007). The original UCR annotation by Bejerano et al., focusing on overlap with protein-coding genomic regions, annotated 111 UCRs as exonic, 256 as non-exonic, whereas the remaining 114 UCRs, for which the evidence of overlap with a protein-coding sequence was inconclusive, were termed possibly exonic (Bejerano et al., 2004). As with mirnas and lincrnas, the high degree of conservation implies that UCRs are of functional importance in mammalian cell biology. Indeed, one of these UCRs is contained in an enhancer upstream of the DACH1 gene (Nobrega et al., 2003; Bejerano et al., 2004), which has a role in development, and non-exonic UCRs that lie intronically are often associated with developmental genes (Bejerano et al., 2004). UCRs are extensively transcribed and their expression patterns have been shown to discriminate between normal and cancer tissues (Calin et al., 2007). Furthermore, UCRs are frequently located at fragile sites and genomic regions involved in cancers (Calin et al., 2007). However, the functional relevance of the vast majority of UCRs remains elusive. ONCOGENOMICS
140 In this study, we aimed at examining the possible functions of T-UCRs using neuroblastoma as a tumour model. Neuroblastoma is an aggressive childhood tumour accounting for 15% of all pediatric cancer deaths. Neuroblastoma patients present with a highly variable clinical course and aggressive tumours are characterized by a combination of various genetic abnormalities, including 1p or 11q deletions, 17q gain and MYCN amplification (Maris et al., 2007). We profiled all 481 T-UCRs on a large and well-characterized panel of representative neuroblastoma tumours and analysed their expression with respect to genomic location and co-regulation with host and surrounding genes. We show that T-UCR expression is related to important clinical and genetic parameters in neuroblastoma and provide evidence that T-UCRs have prognostic value in neuroblastoma. Using a functional genomics approach, based on the integration of T- UCR and mrna expression data, we assigned putative functions to each T-UCR and validated these for a subset of T-UCRs using cellular model systems. Our results reveal for the first time a deregulated T-UCR expression landscape in neuroblastoma and implicate T-UCRs in a wide range of cellular functions. Results T-UCR functions in neuroblastoma P Mestdagh et al Re-annotation and selection of independently expressed T-UCRs For T-UCR quantification, we designed 481 primer sets for reverse transcription quantitative PCR (RT qpcr). All primer pairs were validated in silico and a representative selection of primers was tested experimentally for their specificity (n ¼ 384) and efficiency (n ¼ 481). Amplicon sizing indicated good concordance between theoretical and experimental amplicon lengths (r 2 ¼ 0.961) (Supplementary Figure 1A), and singlecurve efficiency estimates, as determined by PCR miner (Zhao and Fernald, 2005), show that 84% of the tested primers have efficiencies between 90 and 110 and 97% have efficiencies above 80% (Supplementary Figure 1B). A significant number of UCRs is located exonically within coding (host) genes. For a number of these, the measured expression value could be confounded by the expression of the host gene. For further analyses, we therefore decided to select T-UCRs whose expression profile is independent from that of the host gene. For this correlation analysis, accurate information on the genomic location of each T-UCR relative to known protein-coding genes is pivotal. As the currently available T-UCR annotation is based on the old hg17 genome assembly and as only three T-UCR categories were used (exonic, non-exonic and possibly exonic) (Bejerano et al., 2004) ( ultra.html), we re-annotated all T-UCR sequences using the more recent genome build hg18 and re-organized them into five different categories by matching their location to that of the human RefSeq genes (Pruitt et al., 2009). The new genomic categories (intergenic, intronic, exonic, partly exonic, exon containing) are unambiguously defined and provide a more detailed genomic annotation for each T-UCR (Figure 1, Supplementary Table 1, Supplementary File 1). We categorized 38.7% of T-UCRs as intergenic, that is, located between genes rather than within a coding segment, 42.6% as intronic, 4.2% as exonic, 5% as partly exonic and 5.6% as exon containing (Figure 2a). For a few T-UCRs (3.9%), the genomic annotation varied because of host gene splice variants. These T-UCRs were categorized as multiple (Figure 2a). To define which of the intragenic T-UCRs are expressed independently of their host and flanking protein-coding genes, we quantified T-UCR (RT qpcr) and mrna (exon array) expression levels in a well-characterized cohort of neuroblastoma tumour samples (n ¼ 49 for T-UCRs and n ¼ 40 for mrna). Next, we calculated correlations between the expression of each intragenic T-UCR and that of its host gene in the tumour cohort. For intragenic T-UCRs with a significant positive correlation to the host gene (Spearman s rank, Po0.05), we concluded that the T-UCR RT qpcr assay measured the expression of the host gene (or that of the host gene and the T-UCR) rather than that of the T-UCR alone (Figure 2b). Not surprisingly, the largest fraction of positive correlations was found for exonic T-UCRs (95%), followed by the exon-containing (66%) and partly exonic T-UCRs (66.7%) (Figure 2a). We found approximately half of all T-UCRs (237 T-UCRs, both inter- and intragenic) to be expressed independently and we therefore selected these for further analysis. None of the intragenic T- UCRs showed a negative correlation to the host gene, whereas 17 T-UCRs (both intra- and intergenic) were negatively correlated to one of the flanking up- or downstream-coding genes (the first flanking up- and coding gene A coding gene B exon containing uc. 454 exonic uc.135 partly exonic uc.101 intergenic uc.10 intronic uc.350 non-coding DNA intron exon UCR Figure 1 Re-annotation of UCRs. Representation of the different UCR classes according to their genomic location with respect to protein-coding genes defined by Refseq. An example of each class is shown. Oncogene
141 135 uc.75 expression (log2) relative expression (log2) intergenic 103 intronic exonic exon containing all T-UCRs independent T-UCRs partly exonic multiple ZEB2 expression (log2) intergenic (103) 205 r 2 = p < intronic (107) 20 1 exonic (1) exon containing (10) 24 partly exonic (8) multiple(8) T-UCR functions in neuroblastoma P Mestdagh et al downstream gene on both strands). Expression levels of the different categories of independent T-UCRs were highly variable with intergenic T-UCRs showing significantly lower expression compared with intragenic T-UCRs (Mann Whitney test, Po0.0001) (Figure 2c). T-UCRs are associated with histone marks for active transcription Little is known on how T-UCR transcription is initiated and regulated. In addition, T-UCRs are frequently located within protein-coding genes, often resulting in highly correlated expression between the T-UCR and the host gene. To evaluate whether transcriptional initiation of T-UCRs shares characteristics with that of protein-coding genes, we sought for an association between T-UCRs and histone marks for transcription initiation. The genomic location of all H3K4me3 marks (a marker for transcriptional initiation) in four different cell lines was obtained from the UCSC genome browser and was used to determine the distance between the centre of each of the 237 independent T-UCRs and that of the nearest active H3K4me3 mark in any of the four cell lines (Bernstein et al., 2005, 2006; Mikkelsen et al., 2007). As a comparison, this procedure was repeated for either a set of random genomic locations with a similar chromosomal distribution as that of the T-UCRs or for the transcription start sites (TSS, genomic location indicating start of gene transcription) of all RefSeq genes or for all mirna genes. As expected, the TSS of the RefSeq genes are strongly associated with active H3K4me3 marks when compared with random locations (Kolmogorov Smirnov test, Po0.0001) (Figure 3). Strikingly, a similar association is observed for T-UCRs (Po0.001) (Figure 3a). To exclude that this is due to the fact that a substantial number of T-UCRs are located within coding genes and thus in close proximity to TSS, we evaluated the H3K4me3 distance distributions of intergenic and intragenic T-UCRs separately. Both intergenic (Kolmogorov Smirnov test, Po0.05) and intragenic (Kolmogorov Smirnov test, Po0.0001) T-UCRs were significantly associated with active H3K4me3 marks (Figures 3b and c), but with a different distribution as compared with protein-coding genes (Kolmogorov Smirnov test, Po0.0001), suggesting a difference in transcriptional organization between T-UCRs and proteincoding genes. In addition, mirnas were closely associated with active H3K4me3 marks (Po0.0001) (Figure 3d). It is interesting that H3K4me3 distance distributions for mirnas and T-UCRs appear similar (no indication for a different distribution, Kolmogorov Smirnov, P40.05), suggesting common features of transcription initiation for these two classes of non-coding RNAs. Figure 2 Features of T-UCR expression in neuroblastoma. (a) Organization of T-UCRs in five categories according to their position with respect to all known human Refseq genes. Distributions for all T-UCRs (black) and for the T-UCRs expressed independently of their host/flanking gene (grey) are shown. (b) Representative positive correlation between the expression of a T-UCR (uc.75) and its host gene (ZEB2) (n ¼ 40). (c) Expression distribution of all independently expressed T-UCRs (n ¼ 237) in each of the five different categories (whiskers: Tukey). Number of T-UCRs per category is indicated between brackets. T-UCRs that belong to more than one category are termed multiple Oncogene
142 T-UCR functions in neuroblastoma P Mestdagh et al 6e-04 5e-04 mrna T-UCR random 6e-04 5e-04 mrna intragenic T-UCR random 4e-04 4e-04 density 3e-04 density 3e-04 2e-04 2e-04 1e-04 1e distance (bp) distance (bp) 6e-04 5e-04 4e-04 mrna intergenic T-UCR random 6e-04 5e-04 4e-04 mrna mirna random density 3e-04 density 3e-04 2e-04 2e-04 1e-04 1e distance (bp) distance (bp) Figure 3 H3K4me3 distance distributions. Distributions of the distance to the nearest histone mark for active transcription (H3K4me3) measured for the transcription start sites of all RefSeq genes (blue), a set of random genomic positions (green), (a) all independently expressed T-UCRs (red), (b) all independently expressed intragenic T-UCRs (red), (c) all independently expressed intergenic T-UCRs (red) and (d) all human mirnas (red). For visualization purposes, only distances up to bp are shown. T-UCR expression is correlated to clinicogenetic parameters in neuroblastoma We next examined whether deregulated T-UCR expression might be implicated in known clinicogenetic neuroblastoma subgroups. Differential T-UCR expression was evaluated with respect to MYCN amplification status and UCR copy-number status. MYCN amplification status distinguishes the highly aggressive MYCNamplified (MNA) tumours from MYCN-non-amplified (MNN) tumours. We found a signature of seven T- UCRs (uc.347, uc.350, uc.279, uc.460, uc.379, uc.446 and uc.364) significantly upregulated in MNA tumours (n ¼ 18) compared with MNN tumours (n ¼ 31) (Mann Whitney test, Po0.0001) (Figure 4a, Supplementary Figure 2). Four of the upregulated T-UCRs were intergenic, whereas three were intronic. No T-UCRs were downregulated in the MNA tumours. For a random selection of three of seven T-UCRs, the expression was evaluated on an independent large neuroblastoma tumour cohort containing 366 samples. From the three T-UCRs (uc.279, uc.364, uc.460) that were profiled on this cohort, two (uc.279, uc.460) were significantly upregulated in the MNA tumours (Mann Figure 4 T-UCR expression in clinicogenetic neuroblastoma subgroups. (a) Expression of a T-UCR signature (n ¼ 7) in MYCN-amplified (MNA) and MYCN-non-amplified (MNN) neuroblastoma tumours (n ¼ 49), measured by means of the pathway score of all seven T-UCRs (whiskers: Tukey). (b) Expression fold change of uc.350, uc.379 and uc h on MYCN activation in the SHEP-MYCN-ER cell line. Whitney test, Po0.05). To evaluate whether any of these seven T-UCRs are induced by MYCN, we profiled their expression in the SHEP-MYCN-ER cellular model system, which allows MYCN activation on the addition of 4-hydroxy tamoxifen (Schulte et al., 2008). Upon Oncogene
143 137 T-UCR functions in neuroblastoma P Mestdagh et al 3587 DNA dependent DNA replication DNA metabolic process DNA replication DNA repair Response to DNA damage stimulus Mitotic cell cycle Cell cycle checkpoint Cell cycle Regulation of mitosis Chromosome segregation Response to endogenous stimulus DNA integrity checkpoint Meiosis Meiotic cell cycle DNA recombination M phase mitotic cell cycle Mitosis M phase Cell cycle phase Cell cycle process RNA processing mrna processing RNA splicing mrna metabolic process significantly associated T-UCRs Figure 5 Functional annotation of T-UCRs. (a) Hierarchical clustering of T-UCRs (x axis) and Gene Ontology Biological Process (y axis) showing significant (FDR o5%) positive correlations (blue), significant negative correlations (red) and no correlation (white) as determined by a Gene Set Enrichment Analysis-based approach. The predominant functional cluster is boxed. (b) Gene Ontology Biological Process categories of the boxed cluster with indication of the number of significantly correlated T-UCRs. MYCN activation, three of seven T-UCRs (uc.460, uc.350 and uc.379) were induced at least twofold (Figure 4b), suggesting that these are MYCN responsive. DNA copy-number changes are known to affect coding gene expression. We therefore examined UCR copy-number changes with respect to T-UCR expression for known critical regions in neuroblastoma (Vandesompele et al., 2005; Lazcoz et al., 2007). We identified seven T-UCRs whose expression correlated to the UCR copy-number status (Supplementary Table 2). These findings demonstrate that deregulated T-UCR expression is associated with genomic aberrations in neuroblastoma tumours. Integrative genomics uncovers putative T-UCR functions Having established differential T-UCR expression patterns in neuroblastoma tumours, we sought to assign putative functions to each T-UCR. To this purpose, we adapted a functional genomics approach, recently proposed by Guttman et al. (2009), based on the integration of multi-level transcriptome data. The independently expressed T-UCRs were correlated to functionally annotated protein-coding genes across the neuroblastoma tumours. The correlation values were subsequently used for Gene Set Enrichment Analysis (GSEA) (Subramanian et al., 2005), thus providing a means to associate each T-UCR to Gene Ontologybased functional classifications, as well as to published experimental data (see Supplementary Methods). The T- UCR annotations serve as a functional resource and are available as supplementary material (Supplementary Files 2, 3 and 4). Significant enrichments (FDR o5%) were extracted and results were clustered to visualize classes of T-UCRs with similar predicted functions (Figure 5a and Supplementary Figure 3). For a large number of T-UCRs, we observed widespread association to numerous cancer-related cellular functions and pathways such as proliferation, apoptosis and differentiation. For example, the most prominent cluster identified using this methodology contained several T- UCRs significantly related to the expression of proteincoding genes involved in the inter-related processes of cell cycle, DNA replication and DNA repair (Figure 5b). To assess the validity of the functional predictions, we sought independent experimental validation for a subset of inferred T-UCR functions. We decided to examine the T-UCRs that were annotated to the TP53 response pathway according to association with TP53-related gene sets from the molecular function and chemical and genetic perturbations collections. To this end, a human neuroblastoma cell line (NGP) lentivirally transfected with a short hairpin RNA against human TP53 (LV-hp53) or murine TP53 (LV-m-p53, used as a negative control) (Van Maerken et al., 2006) was treated with nutlin-3 for 24 h and profiled for T-UCR expression. On nutlin-3 treatment, MDM2 activity is antagonized leading to TP53 stabilization and activation of function in cells with wild-type TP53 (Vassilev et al., 2004; Van Maerken et al., 2006). We found 40 T-UCRs to be responsive to nutlin-3 treatment in NGP-LV-m-p53 cells, but not in NGP-LV-h-p53 cells, indicating a TP53- dependent expression. Almost three quarters (29 of 40) were annotated to the TP53 response pathway (Fisher s exact test, Po0.05), thus confirming the functional Oncogene
144 T-UCR functions in neuroblastoma P Mestdagh et al predictions. Earlier, Calin et al. (2007) reported that T- UCR uc.73 was implicated in the apoptotic response in colon cancer cell line COLO320. Interestingly, we found uc.73 to be annotated to the TP53 response pathway supporting its reported role in apoptosis and further corroborating our workflow. CLUSTER 1 DNA damage response CLUSTER 4 Immune response, Development CLUSTER 2 Cell cycle, Proliferation CLUSTER 3 Differentiation Figure 6 T-UCR expression network in neuroblastoma. Interconnected expression network of T-UCRs in neuroblastoma. Pairs of T-UCRs with a significant correlation (Po0.001; Pearson s correlation) were connected followed by exclusion of T-UCRs with less than two significant connections. Positive correlations are indicated in green, negative correlations in red. Four individual clusters are apparent from the network. Functional T-UCR expression network in neuroblastoma Highly co-regulated genes often share a common activating/repressing feature, that is, they belong to the same cellular process or are transcribed in response to a common internal or external stimulus. As previously reported, several of the T-UCRs are located in or around genes related to differentiation and splicing (Bejerano et al., 2004), thus by genomic location alone, groups of related T-UCRs have been identified. To further examine putative shared functions among the 237 independently expressed T-UCRs, we first visualized all pairwise T-UCR correlations (Supplementary Figure 4). As several groups of co-expressed T-UCRs could be identified, we used a network approach in which significantly correlated T-UCRs (Pearson s correlation, Po0.001) are connected. Individual T-UCRs with less than two significant connections were excluded, resulting in a highly interconnected network of related T- UCRs (Figure 6). Four major clusters, consisting of 9, 11, 9 and 6 T-UCRs respectively, were identified (Figure 6). To assign putative common functions, each cluster was annotated based on the GSEA results. Significant T-UCR GSEA terms from the biological process, molecular function, as identified above, were tested for association with the network clusters using Fisher s exact test. For cluster 1, we identified DNA damage response as a significant GSEA-based annotation. Cluster 2 was predominantly associated with cellcycle regulation and proliferation, whereas cluster 3 appeared to be implicated in differentiation. For cluster 4, development and immune response were among the significant GSEA terms. DNA damage response is highly related to TP53 activation. To validate the annotation of cluster 1, we evaluated its expression by means of the cluster 1 pathway activity score in the nutlin-3-treated NGP-LV-h-p53 and NGP-LV-m-p53 cells. No significant differential pathway activity score was observed in the NGP-LV-h-p53 cells (fold change o1.5), whereas the pathway activity score markedly changed (fold change 41.5, Mann Whitney test, Po0.01) in the NGP-LV-m-p53 cells, 24 h upon nutlin-3 treatment (Figure 7a). These findings suggest that cluster 1 represents a subset of co-expressed T-UCRs that are responsive to TP53 activation. As sustained nutlin-3 treatment of NGP-LV-m-p53 cells results in TP53-dependent neuronal differentiation (Van Maerken et al., 2006), we evaluated cluster 3 pathway activity in NGP-LV-h-p53 and NGP-LV-m-p53 cells, 5 days upon nutlin-3 treatment. NGP-LV-m-p53 cells, but not NGP-LV-h-p53 cells, showed marked increase in neuronal differentiation after 5 days of nutlin-3 treatment (Figure 7b). Differential activity of cluster 3 was observed in NGP-LV-m-p53 cells after nutlin-3 treatment (fold change 41.5, Mann Whitney test, P ¼ 0.074), but not in NGP-LV-h-p53 cells, suggesting a role for cluster 3 T-UCRs in p-53-dependent neuronal differentiation (Figure 7c). We further evaluated cluster pathway scores in the neuroblastoma tumour cohort with respect to patient survival. Samples were divided into quartiles according to the pathway activity score (Fredlund et al., 2008). Interestingly, Kaplan Meier analysis based on these quartiles revealed a significant correlation (Po0.01) between the activity of cluster 4 and overall and event-free patient survival (Figure 7d). These results implicate co-expressed T-UCR clusters in different aspects of neuroblastoma disease and outcome. Finally, we asked whether the inferred T-UCR functions would hold true for tissue types other than neuroblastoma. To this end, we cultured immortalized human fibroblast BJ cells in starvation medium (0.1% fetal calf serum) to induce cell-cycle arrest and thus block proliferation. Next, we examined the expression of T-UCR cluster 2, which was annotated to cell-cycle regulation and proliferation, in control and serum starved BJ cells. Interestingly, we observed a significant differential expression of cluster 2 on serum starvation (Mann Whitney test, Po0.05) (Figure 7e), suggesting Oncogene
145 139 T-UCR functions in neuroblastoma P Mestdagh et al pathway score fold change ± nutlin CLUSTER 1 NGP LV-h-p53 NGP LV-m-p53 NGP-LV-m-p53 5d- NGP-LV-m-p53 5d+ NGP-LV-h-p53 5d- NGP-LV-h-p53 5d+ pathway score fold change ± nutlin CLUSTER 3 NGP LV-h-p53 NGP LV-m-p CLUSTER CLUSTER 4 2 CLUSTER 2 OS (%) EFS (%) p < 0.01 p < follow-up time (years) follow-up time (years) 0% 25% 25% 50% 50% 75%% 75% 100% Mean expression (log2) control serum starved Figure 7 T-UCR expression clusters are correlated to neuroblastoma biology and outcome. (a) T-UCR cluster 1 is implicated in TP53 response in neuroblastoma. Cluster 1 expression is summarized by means of its pathway activity score. Pathway activity score fold changes on treatment of NGP-LV-h-p53 and NGP-LV-m-p53 with 16 mm nutlin-3 for 1 day, relative to vehicle treatment, are plotted. Fold changes 41.5 are indicated in red. (b) Morphology of NGP-LV-m-p53 and NGP-LV-h-p53 cells at day 5 of treatment with 16 mm nutlin-3 (5d þ ) or vehicle control (5d ). NGP-LV-m-p53 cells show an extensive neurite outgrowth after 5 days of nutlin-3 treatment. NGP-LV-h-p53 cells did not show any signs of neuronal differentiation. (c) T-UCR cluster 3 is implicated in differentiation of neuroblastoma cells. Pathway activity fold changes were calculated as in (a). (d) Kaplan Meier plots for overall (OS) and event-free (EFS) survival of neuroblastoma patients based on the pathway activity score of T-UCR cluster 4, represented as quartiles. Increased activity of cluster 4 correlated to a poor overall and EFS. (e) Mean expression (log 2) of T-UCR cluster 2 in control (grey) and serumstarved (red) BJ cells (error bars reflect s.e.m.). that our proposed functional T-UCR annotations are also valid for human fibroblasts and may potentially be applied to other cell types. Discussion Non-coding RNAs have emerged as an important component of the human transcriptome. Significant progress is being made on the functional annotation of a particular class of small non-coding mirnas, although the functionality of others, such as T-UCRs, is still elusive. In this study, we aimed at characterizing T-UCR expression and function, based on an integrative analysis in an aggressive childhood tumour. We designed an RT qpcr based T-UCR-profiling platform for measurement of the expression of all 481 T-UCRs. RT qpcr is the gold standard for small RNA profiling (Chen et al., 2005; Mestdagh et al., 2008) and has a superior specificity, sensitivity and flexibility compared with array-based expression profiling platforms. As UCRs are defined solely on the basis of sequence similarity, cautious interpretation of their measured transcription is warranted. Perfectly conserved sequences that are located exonically might very well represent important features of the host gene instead of an independently expressed T-UCR. We therefore determined which T-UCRs were expressed independently from their genomic environment. Strikingly, about half of the T-UCRs showed strong positive correlations to the expression of their host gene. Not surprisingly, this was mainly the case for T-UCRs that overlapped with an exon of a protein-coding gene, suggesting that some UCRs represent a conserved feature of the host gene rather than coding for an independent transcriptional unit. Oncogene
146 T-UCR functions in neuroblastoma P Mestdagh et al To gain further insight into the initiation and regulation of T-UCR transcription, we evaluated the chromatin state of the T-UCR genomic neighbourhood. Actively transcribed genes, both coding and non-coding, are marked by trimethylation of lysine 4 of histone H3 (H3K4me3) at their promoter (Mikkelsen et al., 2007; Guttman et al., 2009), a feature that has been used to identify lincrnas (Guttman et al., 2009). We hypothesized that comparing the distribution of active H3K4me3 marks between T-UCRs on one hand and mirnas and protein-coding genes on the other could reveal insights into the general structure of T-UCR transcriptional units and their initiation. We observed an association between H3K4me3 marks and T-UCRs, independent of host-gene-associated H3K4me3 marks. The H3K4me3 distance distribution was different from that of the protein-coding genes but showed a striking similarity to that of mirnas. Compared with proteincoding genes, mirna promoters have similar features but the organization of the transcriptional unit is more complex as mirnas can be transcribed from promoters located several kilobases (up to 40 kb) away (Corcoran et al., 2009). The correspondence in H3K4me3 distance distribution between mirnas and T-UCRs suggests a similar transcriptional organization with initiation sites located several kilobases away from the T-UCR. Further experimental evaluation is necessary to validate these observations. The high conservation of T-UCRs across species almost inevitably implies functionality. Previously, Calin et al. (2007) reported differential T-UCR expression between normal and cancerous tissues and identified one T-UCR, uc.73, to be oncogenic in colon cancer. In this study, we show that T-UCRs are widely expressed in neuroblastoma tumours and that their expression correlates to important clinicogenetic parameters such as MYCN amplification status. In addition, DNA copy-number changes that are associated with neuroblastoma disease were shown to affect T-UCR expression, which is in line with the observation that T-UCRs are frequently located at fragile sites or genomic regions involved in cancer (Calin et al., 2007). To gain further insight into the pathways and processes in which T-UCRs are involved, we implemented an integrative genomics workflow to infer putative T-UCR functions using Gene Set Enrichment Analysis. In support of the workflow, T-UCRs predicted to be involved in apoptosis and differentiation were experimentally validated using a cellular model system. Furthermore, our predictions annotated uc.73 to the TP53 response pathway. This result is in concordance with published results showing that RNA interference-mediated knockdown of uc.73 induced apoptosis in a colon cancer cell line (Calin et al., 2007), thus again confirming the validity of our workflow. Our further analyses of T-UCR expression patterns uncovered an interconnected network consisting of four major clusters, functionally associated with cancerrelated cellular processes such as proliferation, apoptosis and differentiation. Moreover, expression of cluster 4 correlated to patient outcome, thus further indicating T-UCRs as a factor in neuroblastoma biology. Interestingly, cluster 4 was associated with development and immune response, which is in agreement with our previously reported observation that neuroblastoma tumours, as compared with normal precursor neuroblasts, are characterized by an overrepresentation of genes involved in immune response (De Preter et al., 2006). In addition, we have shown that the inferred T- UCR functions are not confined to neuroblastoma cells but are also valid for immortalized human fibroblasts. This suggests that T-UCRs are of general relevance in cell biology and opens perspectives to use these annotations as a functional resource in future T-UCR studies. In conclusion, our results show that T-UCRs are widely expressed in neuroblastoma tumours and correlate to clinicogenetic parameters. Functional T-UCR annotations, inferred through a functional genomics approach and validated using cellular models, reveal associations with several cancer-related cellular processes such as apoptosis and differentiation. Additional studies are needed to further elucidate T-UCR function and to unravel the transcriptional programmes mediating their expression. T-UCRs make up an interesting class of non-coding RNAs and could prove attractive targets for treatment or diagnosis. Materials and methods Patient samples A total of 49 neuroblastoma tumours were collected at the Ghent University Hospital (Ghent, Belgium) and at the Medical School of Valencia (Valencia, Spain) before treatment (Supplementary Table 3). An additional cohort of 366 neuroblastoma tumours, originally described by Vermeulen et al. (2009), were also included. All samples were obtained at diagnosis. Informed consent was obtained from the patient s relatives. Patients were staged according to the International Neuroblastoma Staging System (Brodeur et al., 1993). Cellular models NGP-LV-h-p53 and NGP-LV-m-p53 cells (Van Maerken et al., 2006) were cultured in RPMI 1640 (Invitrogen, Carlsbad, CA, USA) supplemented with 15% fetal calf serum and treated with 16 mm nutlin-3 (Cayman Chemical, Ann Arbor, MI, USA) or vehicle control (ethanol) for 1 and 5 days before harvesting. SHEP-MYCN-ER cells (Schulte et al., 2008) were treated with 4-hydroxytamoxifen or vehicle control (ethanol) for 2 days before harvesting. BJ cells were serum starved for 5 days in Dulbecco s modified Eagle s medium (DMEM) supplemented with 0.1% fetal calf serum. Medium was refreshed after 3 days. Non-starved BJ cells were cultured in DMEM (10% fetal calf serum) for 5 days. T-UCR primer design RT qpcr primers for 481 T-UCRs were designed using Primer3 (stand alone or implemented in Beacondesigner) (Rozen and Skaletsky, 2000) and validated through an in silico primer analysis pipeline (Lefever et al., 2009). Designs were selected according to four different criteria: absence of stable secondary structures in the primer-annealing regions, specificity, absence of SNPs in the primer-annealing regions and 3 0 GC content. All assays are available from PrimerDesign Oncogene
147 141 Ltd (Southampton, UK). Primer efficiencies were determined using PCR miner (Zhao and Fernald, 2005). To evaluate primer specificity, amplicons were sized on a Caliper LC90 (Caliper Life Sciences, Hopkinton, MA, USA). RT qpcr For detailed reaction conditions see Supplementary Methods. Expression data were normalized using the mean expression value per sample (Mestdagh et al., 2009). For the independent validation cohort of 366 tumours, RT qpcr data were normalized using qbaseplus v1.2 as described previously (Vermeulen et al., 2009). T-UCR RT qpcr data are available in rdml format (Lefever et al., 2009) (Supplementary File 5). Compliance of qpcr experiments with the MIQE guidelines (Bustin et al., 2009) ( is listed in the MIQE checklist (Supplementary File 6). Exon array Total RNA was isolated from 40 tumour samples and was hybridized to Human Exon 1.0. ST array (Affymetrix, Santa Clara, CA, USA) at the microarray facility of the University Hospital of Essen according to the manufacturer s protocol. To obtain expression information per gene, exon data were merged by transcript clusters. Exon array data were normalized according to the RMA-sketch algorithm using Affymetrix Power Tools (Affymetrix). Array comparative hybridization Array CGH was performed for detection of T-UCR copynumber alterations using a custom 44K array enriched for regions with recurrent imbalances in neuroblastoma (1p, 2p, 3p, 11q, 17) and T-UCR genes (Agilent Technologies, Palo Alto, CA, USA). See Supplementary Methods for detailed description. T-UCR functions in neuroblastoma P Mestdagh et al Statistics See Supplementary Methods for detailed description. Conflict of interest The authors declare no conflict of interest. Acknowledgements We are indebted to all the members of the SIOPEN and the GPOH for providing tumour samples or the clinical history of patients. This research was funded by the Gent University Research Fund (BOF 01D31406 to PM, BOF 01F07207 to FP, BOF 01Z09407 to J Vandesompele), the Belgian Kid s Fund and the Fondation pour la recherche Nuovo-Soldati (J Vermeulen), RD06/0020/0102 from RTICC/ISCIII to RN, the American Cancer Association to EF, the Swedish Cancer Society to MR, the Fund for Scientific Research (grant number: G and ) and the Belgian Foundation Against Cancer, found of public interest (project SCIE ). KDP is a postdoctoral researcher with the Fund for Scientific Research-Flanders. CK is supported by a doctoral grant from the Institute for the Promotion of Innovation by Science and Technology in Flanders (IWT ). We acknowledge the support of the European Community under the FP6 (project: STREP: EET-pipeline, number: ) and FP7 (ONCOMIRS, grant agreement number ). This publication reflects only authors views; the commission is not liable for any use that may be made of the information herein. This article presents research results of the Belgian programme of Interuniversity Poles of Attraction, initiated by the Belgian State, Prime Minister s Office, Science Policy Programming References Bartel DP. (2009). MicroRNAs: target recognition and regulatory functions. Cell 136: Bejerano G, Pheasant M, Makunin I, Stephen S, Kent WJ, Mattick JS et al. (2004). Ultraconserved elements in the human genome. Science 304: Bernstein BE, Kamal M, Lindblad-Toh K, Bekiranov S, Bailey DK, Huebert DJ et al. (2005). Genomic maps and comparative analysis of histone modifications in human and mouse. Cell 120: Bernstein BE, Mikkelsen TS, Xie X, Kamal M, Huebert DJ, Cuff J et al. (2006). A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125: Brodeur GM, Pritchard J, Berthold F, Carlsen NL, Castel V, Castelberry RP et al. (1993). Revisions of the international criteria for neuroblastoma diagnosis, staging, and response to treatment. J Clin Oncol 11: Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M et al. (2009). The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55: Calin GA, Liu CG, Ferracin M, Hyslop T, Spizzo R, Sevignani C et al. (2007). Ultraconserved regions encoding ncrnas are altered in human leukemias and carcinomas. Cancer Cell 12: Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT et al. (2005). Real-time quantification of micrornas by stem-loop RT-PCR. Nucleic Acids Res 33: e179. Corcoran DL, Pandit KV, Gordon B, Bhattacharjee A, Kaminski N, Benos PV. (2009). Features of mammalian microrna promoters emerge from polymerase II chromatin immunoprecipitation data. PLoS One 4: e5279. De Preter K, Vandesompele J, Heimann P, Yigit N, Beckman S, Schramm A et al. (2006). Human fetal neuroblast and neuroblastoma transcriptome analysis confirms neuroblast origin and highlights neuroblastoma candidate genes. Genome Biol 7: R84. Esquela-Kerscher A, Slack FJ. (2006). Oncomirs micrornas with a role in cancer. Nat Rev Cancer 6: Fredlund E, Ringner M, Maris JM, Pahlman S. (2008). High Myc pathway activity and low stage of neuronal differentiation associate with poor outcome in neuroblastoma. Proc Natl Acad Sci USA 105: Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D et al. (2009). Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458: He L, He X, Lim LP, de Stanchina E, Xuan Z, Liang Y et al. (2007). A microrna component of the p53 tumour suppressor network. Nature 447: Khalil AM, Guttman M, Huarte M, Garber M, Raj A, Rivea Morales D et al. (2009). Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Natl Acad Sci USA 106: Lazcoz P, Munoz J, Nistal M, Pestana A, Encio IJ, Castresana JS. (2007). Loss of heterozygosity and microsatellite instability on chromosome arm 10q in neuroblastoma. Cancer Genet Cytogenet 174: 1 8. Oncogene
148 T-UCR functions in neuroblastoma P Mestdagh et al Lefever S, Vandesompele J, Speleman F, Pattyn F. (2009). RTPrimerDB: the portal for real-time PCR primers and probes. Nucleic Acids Res 37: D942 D945. Maris JM, Hogarty MD, Bagatell R, Cohn SL. (2007). Neuroblastoma. Lancet 369: Mestdagh P, Feys T, Bernard N, Guenther S, Chen C, Speleman F et al. (2008). High-throughput stem-loop RT-qPCR mirna expression profiling using minute amounts of input RNA. Nucleic Acids Res 36: e143. Mestdagh P, Van Vlierberghe P, De Weer A, Muth D, Westermann F, Speleman F et al. (2009). A novel and universal method for microrna RT-qPCR data normalization. Genome Biol 10: R64. Mikkelsen TS, Ku M, Jaffe DB, Issac B, Lieberman E, Giannoukos G et al. (2007). Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448: Nobrega MA, Ovcharenko I, Afzal V, Rubin EM. (2003). Scanning human gene deserts for long-range enhancers. Science 302: 413. O Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT. (2005). c-myc-regulated micrornas modulate E2F1 expression. Nature 435: Pruitt KD, Tatusova T, Klimke W, Maglott DR. (2009). NCBI reference sequences: current status, policy and new initiatives. Nucleic Acids Res 37: D32 D36. Rozen S, Skaletsky HJ. (2000). Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S and Misener S (eds). Bioinformatics Methods and Protocols: Methods in Molecular Biology. Humana Press: Totowa, NJ. pp Schulte JH, Horn S, Otto T, Samans B, Heukamp LC, Eilers UC et al. (2008). MYCN regulates oncogenic micrornas in neuroblastoma. Int J Cancer 122: Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al. (2005). Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102: Van Maerken T, Speleman F, Vermeulen J, Lambertz I, De Clercq S, De Smet E et al. (2006). Small-molecule MDM2 antagonists as a new therapy concept for neuroblastoma. Cancer Res 66: Vandesompele J, Baudis M, De Preter K, Van Roy N, Ambros P, Bown N et al. (2005). Unequivocal delineation of clinicogenetic subgroups and development of a new model for improved outcome prediction in neuroblastoma. J Clin Oncol 23: Vassilev LT, Vu BT, Graves B, Carvajal D, Podlaski F, Filipovic Z et al. (2004). in vivo activation of the p53 pathway by smallmolecule antagonists of MDM2. Science 303: Vermeulen J, De Preter K, Naranjo A, Vercruysse L, Van Roy N, Hellemans J et al. (2009). Predicting outcomes for children with neuroblastoma using a multigene-expression signature: a retrospective SIOPEN/COG/GPOH study. Lancet Oncol 10: Zhao S, Fernald RD. (2005). Comprehensive algorithm for quantitative real-time polymerase chain reaction. J Comput Biol 12: Supplementary Information accompanies the paper on the Oncogene website ( Oncogene
149 Supplemental Data 143 RT- qpcr reaction conditions Total RNA was isolated using the mirneasy kit (Qiagen) according to the manufacturer s instructions. RNA quality was assessed using the experion (RNA Quality Index >5 according to software version 3.0, Bio- Rad) and 20 ng of total RNA was pre- amplified (WT- Ovation, NuGEN). RNA concentration was measured using the Nanodrop (Thermo Scientific). PCR plates were prepared using a 96- well head liquid handler (FreedomEvo 100, Tecan) and qpcr reactions were performed on a 7900HT real- time PCR detection system (Applied Biosystems) using SYBR green detection chemistry (Eurogentec). RT- qpcr reactions were performed in a total volume of 8 μl consisting of 4 μl of SYBR Green qpcr master mix (2X) (Eurogentec, HotGoldStar polymerase), 1 μl of DNase and RNase free water (Sigma), 1 μl of forward primer (5 μm), 1 μl of reverse primer (5 μm) and 1 μl of a 20- fold dilution of cdna obtained after NUGEN RNA- amplification (WT- Ovation, NUGEN). Final reaction concentrations of MgCl2 and dntp were 3.5 mm and 0.2 mm respectively. Cycling conditions were as follows: 10 min at 95 C followed by 40 cycles of 10 s at 95 C and 1 min at 60 C. Array comparative hybridization A total of 150 ng of patient and reference DNA was labeled with Cy3 and Cy5, respectively (BioPrime ArrayCGH Genomic Labeling System, Invitrogen). Data were processed with an in- house developed visualization software arraycghbase (Menten et al., 2005) using circular binary segmentation (CBS) for the scoring of DNA CNAs using a significance threshold of 0.01 (Olshen et al., 2004). Statistics All statistical analyses were performed using the R and R- Bioconductor statistical programming environment. Differential T- UCR expression was evaluated using the Mann- Whitney test with Benjamini- Hochberg multiple testing correction. Pathway scores were calculated as described previously (Fredlund et al., 2008). For the identification of T- UCR copy number status we calculated one CBS- value per T- UCR in each sample. Samples were then grouped according to CBS- value (homozygous deletion: CBS < - 1.2; heterozygous deletion: CBS < ; duplication: CBS > 0.35; amplification: CBS > 1.2) to identify differential T- UCR expression (Kolmogorov- Smirnov). In order to assign functional terms to each T- UCR, a correlation matrix for T- UCR and mrna expression was generated based on Spearman s Rank correlation coefficient for each T- UCR:mRNA combination. Combined T- UCR and mrna expression data were available for 40 tumour samples and only T- UCR:mRNA combinations with an overlapping detected expression in more than ten tumour samples were used. For each T- UCR, mrnas were ranked according to the Spearman s Rank rho- value to generate ranked gene lists for Gene Set Enrichment Analysis (GSEA) (Subramanian et al.,
150 ) using the Gene Ontology biological process, molecular function as well as the chemical and genetic perturbations gene set collections from the GSEA Molecular Signatures Database. Gene sets with a false discovery rate (FDR) below 5 % were considered significant and based on the GSEA normalized enrichment score (NES) T- UCR:GSEA scores were transformed to a ternary scale (- 1,0,1) prior to functional hierarchical clustering (FDR>0.05: 0; FDR<0.05 & NES>0: 1; FDR<0.05 & NES<0: - 1). A T- UCR correlation matrix was created by calculating all pairwise Pearson correlations followed by unsupervised clustering using Ward s minimum variance method. Analyses using alternative correlation and clustering methods gave analogous results. T- UCRs were linked by a highly significant correlation (p < 0.001) and the resulting correlation network was visualized using Cytoscape (Shannon et al., 2003). T- UCRs with less than two significant links were removed. T- UCR network subclusters were functionally annotated by identifying common GSEA terms associated with the subcluster members. Terms present for two or more T- UCRs within the same subcluster were assayed for a significant association using a Fisher s exact test (cut- off: p < 0.05). Supplemental References Fredlund E, Ringner M, Maris JM, Pahlman S (2008). High Myc pathway activity and low stage of neuronal differentiation associate with poor outcome in neuroblastoma. Proc Natl Acad Sci U S A 105: Menten B, Pattyn F, De Preter K, Robbrecht P, Michels E, Buysse K et al (2005). arraycghbase: an analysis platform for comparative genomic hybridization microarrays. BMC Bioinformatics 6: 124. Olshen AB, Venkatraman ES, Lucito R, Wigler M (2004). Circular binary segmentation for the analysis of array- based DNA copy number data. Biostatistics 5: Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D et al (2003). Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13: Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al (2005). Gene set enrichment analysis: a knowledge- based approach for interpreting genome- wide expression profiles. Proc Natl Acad Sci U S A 102:
151 145 Supplemental Figure 1. Validation of T- UCR primers Experimental validation of T- UCR primer quality. (A) Correlation plot for theoretical and experimental amplicon lengths. (B) T- UCR primer efficiency. A B amplicon Amplicon length - theoretical (bp) R! = primer Primer efficiency (%) amplicon Amplicon length - experimental (bp) T-UCR Supplemental Figure 2. T- UCR expression is correlated to MYCN amplification status in neuroblastoma Relative expression of 7 T- UCRs in MYCN amplified (MNA) and MYCN non- amplified neuroblastoma tumours (whiskers: Tukey). p < p < p < p < MNA MNN MNA MNN MNA MNN MNA MNN p < p < p < MNA MNN MNA MNN MNA MNN
152 146 Supplemental Figure 3. Functional T- UCR annotation (A) Hierarchical clustering of T- UCRs (X- axis) and Gene Ontology Molecular Function (Y- axis) showing significant positive correlations (blue, FDR < 5 %), significant negative correlations (red, FDR < 5 %) and no correlation (white). (B) Hierarchical clustering of T- UCRs (X- axis) and chemical and genetic perturbations collections (Y- axis). A B
153 Supplemental Figure 4. T- UCR correlation plot in neuroblastoma Correlation clustering of T- UCR expression in neuroblastoma tumours (n = 49). 147 Supplemental Table 2. T- UCR expression is correlated to UCR copy number status in neuroblastoma T-UCR Chromosomal position p-value* uc.25 1p uc.10 1p uc q uc q uc q uc q uc q !"#$%&'()*+()),&"-*.()%/
154 Supplemental Table 3. Neuroblastoma patient characteristics 148! Patient information INSS stage MYCN status Age N amplified 18 non-amplified 31 < 1year 20 > 1year 29
155 PAPER 6: The microrna body map: dissecting microrna function through integrative genomics 149 PAPER 6 The microrna body map: dissecting microrna function through integrative genomics. Mestdagh P*, Lefever S*, Pattyn F, Ridzon D, Fredlund E, Fieuw A, Ongenaert M, Vermeulen J, De Paepe A, Wong L, Speleman F, Chen C, Vandesompele J. Submitted. *Equally contributing authors
156 150 The microrna body map: dissecting microrna function through integrative genomics Pieter Mestdagh 1*, Steve Lefever 1*, Filip Pattyn 1, Dana Ridzon 2, Erik Fredlund 1, Annelies Fieuw 1, Maté Ongenaert 1, Joëlle Vermeulen 1, Anne De Paepe 1, Linda Wong 2, Frank Speleman 1, Caifu Chen 2, Jo Vandesompele 1 1 Center for Medical Genetics, Ghent University Hospital, Ghent, Belgium. 2 Life Technologies, Foster City, California, USA. * Equally contributing authors Abstract While a growing body of evidence implicates regulatory microrna modules in various aspects of human disease and development, insights in specific microrna function remain limited. Here, we present an innovative approach to elucidate tissue specific mirna function by multi- level integration of corresponding mirna and mrna gene expression levels, mirna target prediction and mechanistic models of gene network regulation. The predicted mirna functions are accessible in the mirna body map, an interactive online compendium and mining tool of high- dimensional newly generated and published microrna expression profiles. The microrna body map enables prioritization of candidate micrornas based on their expression pattern or functional annotation across tissue or disease subgroup. The microrna body map project has great potential to become a community resource. Introduction MicroRNAs (mirnas) are small non- coding RNA molecules that function as indispensible regulators of an increasing number of cellular processes. The exact role of an individual mirna is strictly depending on its spatiotemporal expression pattern and that of its targeted genes. With over 1000 mature human mirna species reported thus far, mirnas form one of the largest classes of gene regulators. While mirna expression profiles have been established for various normal and diseased tissues, our understanding of specific mirna function remains limited. To accommodate this, several experimental procedures have been developed for high- throughput mirna target identification such as RIP- chip and HITS- CLIP (Chi et al, 2009; Tan et al, 2009). Unfortunately, these methods are technically challenging and are typically performed for only one or few mirnas, necessitating an up- front prioritization and selection of candidate mirnas. Alternatively, computer based mirna target predictions can be used to gain insights in mirna function by probing annotated gene sets for mirna target enrichment (Nam et al, 2008; Tan et al, 2009; Ulitsky et al, 2010). Of note, mirna target prediction algorithms are prone to a high degree of false positives and completely ignore the tissue or disease- specific nature of mirna target interactions.
157 151 Here, we present an innovative and sensitive method and accompanying resource to elucidate tissue specific mirna function by combining matching mirna and mrna expression data with mirna target prediction and mechanistic models of gene network regulation. Inferred mirna functions, based on different datasets, can be queried through the mirna body map, a webtool available at To complement the functional predictions, a literature search tool was implemented to retrieve experimentally validated mirna functions. In addition, the mirna body map contains high- quality RT- qpcr mirna expression profiles for more than 750 human, mouse and rat samples, belonging to different tissue and disease types, which can be examined through a built- in mirna expression analysis pipeline. This pipeline allows the identification of differentially expressed mirnas between tissues or disease groups, tissue or disease specific mirnas and stably expressed mirnas for mirna expression normalization. The mirna body map serves as a community resource to prioritize candidate mirnas and generate hypotheses for further research. Results and Discussion functional mirna annotation To determine tissue or disease- specific mirna functions, matching mirna and mrna expression levels were analyzed using rank correlation coefficients. Matching genome wide mrna and mirna expression data were obtained from literature or newly generated. To maximize specificity and sensitivity, we only included those datasets for which mirna expression was generated using RT- qpcr technology, generally accepted as the gold standard for small- RNA expression profiling (Mestdagh et al, 2008). In total, 244 human samples belonging to 4 different datasets (normal adult tissues, neuroblastoma tumours, myeloma tumours and NCI60 cancer cell lines) were included in the analysis. For each mirna, mrnas were ranked according to their correlation coefficient and functional annotations, enriched among the positively or negatively correlated mrnas, were identified using Gene Set Enrichment Analysis (GSEA) (Subramanian et al, 2005). We next integrated the GSEA results with mirna target prediction and mechanistic models of gene network regulation. These models represent specific interaction schemes between a mirna and a gene set (or pathway) and form the basis of the mechanism underlying a particular mirna gene set association. We hypothesized that a significant mirna gene set association is more likely to be functional if there is mechanistic evidence that links the association to one of the five proposed models (Figure 1). In this way, negative mirna gene set associations can occur if the gene set is enriched for targets of the mirna (multi- component targeting), if the mirna targets a key signaling molecule in the pathway represented by the gene set (component targeting) or if the mirna negatively regulates a transcriptional activator that has its targets enriched in the gene set (transcription factor targeting) (Figure 1A). Similarly, positive mirna gene set associations can occur if the mirna negatively regulates a transcriptional repressor with its targets enriched in the gene set (transcription factor targeting), if the mirna targets a
158 152 negative regulator of a pathway or if the mirna and the genes in the gene set share a common transcriptional activator or repressor (Figure 1B). In order to integrate mirna target predictions in the mechanistic models, we first determined which mirna target prediction database contains the most accurate predictions. To this end, we used publically available mass spectrometry protein expression data from eight mirna perturbation experiments (Baek et al, 2008; Selbach et al, 2008) and evaluated seven widely used mirna target databases for their ability to predict protein downregulation. We found that the targets in the MIRDB database showed the highest fold downregulation in the experimental datasets (Supplemental Figure 1A, B) and that MIRDB showed the lowest number of false positive predictions (Supplemental Figure 1C). Combining individual databases did not give better results that those obtained by MIRDB alone (Supplemental Figure 1D) suggesting that MIRDB predictions are best suited for accurate assessment of mirna target enrichment in the studied gene sets. MIRDB predictions were then used to calculate mirna target enrichments in the gene sets in order to identify those negative mirna gene set associations according to the multi- component targeting model. To identify mirna gene set associations that follow the mechanistic model of transcription factor targeting, we searched for gene sets that are enriched for transcription factor targets and used MIRDB to predict mirnas regulating transcription factors. Validation of predicted mirna functions To support the predicted mirna functions and their assignment to one of the proposed models of gene regulation, we first compared our findings to experimentally validated mirna functions. For each of the five proposed models, representative examples were found in literature (Supplemental Figure 2), underscoring the relevance of the models and the power of the mirna body map functional annotation pipeline. To further assess the accuracy of the pipeline, we compared functional predictions for mirnas from the mir cluster to a set of experimentally derived mir functions. To this end, we used a recently published protein expression dataset from a mir perturbation experiment in neuroblastoma cells as our benchmark (Mestdagh et al, 2010a). Measured proteins (n = 3249) were ranked according to their fold change and GSEA revealed 94 genesets enriched among the downregulated proteins. We then compared these experimentally derived gene sets to gene sets predicted to be associated with mir in the neuroblastoma tumour dataset. In total, 78 out of 94 experimentally validated gene sets were predicted. Another 201 gene sets were predicted that were not identified in the benchmark dataset suggesting that the functional annotation pipeline reaches a sensitivity and specificity of 83% and 34% respectively. The relatively low specificity is in part related to the fact that the benchmark dataset was based on results for one single neuroblastoma cell line while the predicted gene sets are derived from a much larger neuroblastoma tumour cohort representing different stages of the disease. Interestingly, the most significant gene sets (GSEA false discovery rate (FDR) = 0), according to the annotation pipeline, were more frequently present in the benchmark dataset as compared to the least significant gene sets (Fisher s
159 153 Exact, p < 0.05) suggesting that predicted gene sets can be further prioritized based on the GSEA FDR- value, greatly enhancing prediction specificity. In addition to functional mirna annotation, the mirna body map enables the detection of regulators of mirna expression. Such regulators are identified by looking for positive associations between a mirna and a gene set representing targets of a transcription factor (i.e. the mirna and the genes in the gene set share a common transcriptional activator, Figure 1). To test this assumption, we searched for mirnas that were positively correlated to a gene set containing MYC target genes (SCHUHMACHER_MYC_TARGETS_UP) in the neuroblastoma dataset and selected only the most significant associations (GSEA FDR = 0). The selected mirnas, 38 in total, were compared to a set of 18 MYC/MYCN activated mirnas, previously validated in neuroblastoma (Mestdagh et al, 2010b). Of note, there is a high degree of overlap between MYC and MYCN target genes in neuroblastoma (Westermann et al, 2008) justifying the use of the MYC targets gene set. In total, 16 out of 18 mirnas were identified suggesting that the annotation pipeline allows the detection of mirna regulators with a high sensitivity (89%) and moderate specificity (42%). To further assess the validity of our approach, we evaluated inferred mirna annotations for tissue specific mirnas and hypothesized that a tissue specific mirna should play a role in pathways relevant for that tissue. In the normal tissues dataset, we searched for mirnas that are highly expressed in tissues of the lymphatic system compared to all other tissues in that dataset. The expression of 5 lymphatic system specific mirnas (mir p, mir- 150, mir- 15, mir- 146a and mir- 150*) is visualized in a ranked expression map (Figure 2A). The most significant gene sets (GSEA FDR = 0, Gene Ontology Biological Process gene set collection) for each of these mirnas are primarily annotated to processes such as immune response and immune cell activation (Figure 2B). Furthermore, we found mir p and mir- 155 to be associated with the NF- kb pathway, a signaling cascade involved in immune response. Mir- 155 has previously been shown to regulate NF- kb signaling in primary human B- lymphocytes (Lu et al, 2008) and our results suggest a similar function for mir p. Next, we selected tissue specific gene sets, such as heart development and brain development, from the Gene Ontology Biological Process gene set collection and identified mirnas associated with those gene sets in the normal tissues dataset. mirnas annotated to brain development showed the highest expression in brain while mirnas annotated to heart development showed the highest expression in cardiovascular tissues, skeletal muscle and mononuclear blood cells (Figure 2C, D). Similarly, mirnas annotated to muscle development showed the highest expression in skeletal muscle and mirnas annotated to digestion had the highest expression in tissues from the gastro- intestinal tract (Figure 2E, F). Together, these results support the relation between tissue specific expression and function and lend further credibility to the predicted mirna functions. Tissue specific mirna functions The function of a mirna depends on the cellular environment that dictates which of the putative target genes are expressed. Tissue or disease specific mirna functions have been previously described, amongst
160 154 others for the mir cluster that can function either as a tumour suppressor or an oncomir (Mendell, 2008). To evaluate whether the predicted mirna functions can differ between datasets containing different tissue types, we compared mirna predictions between the neuroblastoma and myeloma dataset. Only the most significant predictions (GSEA FDR=0) were used in the analysis as we found these to be independent of sample size (Supplemental Figure 3 and Supplemental Methods). Rather surprisingly, the majority of predicted mirna functions were specific to only one of the datasets suggesting that predicted mirna functions cannot always be generalized and should be interpreted within a tissue or cell- specific context. These results clearly support previous observations that mirnas often function in a tissue specific manner. Further experiments are needed to validate the extent of these findings. Identifying mirna- directed transcription factor regulation MiRNAs have been shown to act as key components in transcription factor signaling networks, either through cooperation with a transcription factor in the process of gene expression regulation (Su et al, 2010) or through direct regulation of the transcription factor itself (Yamakuchi et al, 2010). Using the mirna body map annotation pipeline, we searched for mirnas regulating the MYCN transcription factor in neuroblastoma. Given the importane of the MYCN gene in neuroblastoma biology MYCN amplification delineates a subgroup of highly aggressive neuroblastoma tumours (Maris et al, 2007) we hypothesized that mirnas regulating MYCN could have an important role in neuroblastoma tumourigenesis. Based on the proposed mechanistic models of mirna action (Figure 1), mirnas regulating a transcription factor should negatively correlate to gene sets containing activated targets of that transcription factor. To identify mirnas regulating MYCN, we searched for mirnas that negatively correlate to a gene set containing MYC targets (SCHUHMACHER_MYC_TARGETS_UP) with a GSEA FDR- value = 0 and that are predicted to target MYCN according to MIRDB predictions. We identified a single mirna (mir- 29a) that could meet these criteria. To evaluate whether mir- 29a directly targets the MYCN 3 UTR, we established a 3 UTR luciferase reporter vector containing the predicted mir- 29a binding site downstream of the luciferase gene and evaluated luciferase activity in the presence of a mir- 29a pre- mir or negative control pre- mir. Luciferase activity significantly decreased in the presence of mir- 29a compared to the negative control pre- mir (Student t- test, p < 0.01) (Figure 3A) suggesting that MYCN is a target of mir- 29a. Furthermore, overexpression of mir- 29a in MYCN amplified NGP cells resulted in a 3- fold decrease of MYCN protein levels (Figure 3B). These results confirm that the mirna body map can be successfully applied to identify mirna- directed transcription factor regulation. Analyzing custom gene sets using the mirna bodymap functional annotation pipeline Predicted mirna functions available in the mirna body map are based on three different gene set collections: Gene Ontology Biological Process, Gene Ontology Molecular Function, and Chemical and Genetic perturbations. On top of that, the mirna body map allows users to perform GSEA with custom
161 155 gene sets obtained from literature or derived from their own perturbation or profiling experiments. Based on the KEGG pathway database (Kanehisa et al, 2010), we established a gene set representing the p53- signaling pathway and a gene set representing the B- cell receptor signaling pathway and searched for mirna associations in the NCI60 dataset and normal adult tissues dataset, respectively. MiR- 34a showed the most significant association with the p53 signaling pathway (GSEA FDR = 0), followed by mir- 373 (GSEA FDR = ). Both mirnas have previously been shown to function downstream of p53 (He et al, 2007; Voorhoeve et al, 2006). MiRNAs associated with B- cell receptor signaling were, amongst others, mir- 150 (GSEA FDR = 0), mir- 155 (GSEA FDR = 0) and different members of the mir cluster (GSEA FDR = 0), all of which were shown to play important roles in B- cell development (Zhou et al, 2007), B- cell receptor activation (Yin et al, 2008) or lymphoproliferative disease (Xiao et al, 2008). Together, these results again validate the mirna body map functional annotation pipeline and support the use of custom gene sets which greatly enhances flexibility towards the user. Materials and Methods mirna and mrna expression data RNA samples from 39 normal human tissues were obtained from Ambion and Biochain. Reverse transcription for 704 mirnas, 18 small RNA controls and U6 was performed using stem- loop primers (Applied Biosystems) in single- plex reactions containing 45 ng of total RNA. qpcr reactions were performed in quadruplicate on a 7900 HT system (Applied Biosystems). Whole genome stem- loop RT- qpcr mirna expression data for over 700 additional samples was gathered from literature. mirna expression data was normalized according to the global mean normalization strategy (Mestdagh et al, 2009). Microarray mrna expression data were taken from GEO (GSE16558, GSE5846, GSE21713 and GSE1133). Gene set enrichment analysis For each individual dataset, Spearman s rank rho values were calculated for each mrna - mirna combination using normalized mrna and mirna expression values. mrna mirna combinations with less than 10 pairwise observations were excluded from the analysis. For each mirna, mrnas were ranked according to their correlation coefficient and ranked gene lists were used as input for GSEA. The following gene set collections were taken from the Molecular Signatures Database (MSigDB v3.0): Chemical and Genetic perturbations, Gene Ontology Molecular Function and Gene Ontology Biological Process. Gene sets significantly enriched among the positive and negative correlating mrnas were selected based on the GSEA FDR value (FDR < 0.05). All analysis were performed using the R Bioconductor statistical programming platform (version 2.11).
162 156 mirna and transcription factor target enrichment For each mirna, predicted targets were derived from the MIRDB database (Wang, 2008; Wang and El Naqa, 2008) and enrichment of these targets in the different gene sets was calculated using Fisher s exact test. Fisher s exact p- values were multiple testing corrected using the Benjamini- Hochberg algorithm. Gene sets that are enriched among the mrnas that negatively correlate with a mirna and that are enriched for targets of that mirna were assigned to the multiple component targeting model. To determine the enrichment of transcription factor targets in the different gene sets we used the Transcription Factor Targets gene set collection from the MSigDB v3.0. Enrichments were calculated using Fisher s exact test and p- values were corrected for multiple testing using the Benjamini- Hochberg algorithm. Gene sets that are enriched among the mrnas that positively correlate with a mirna and that are enriched for targets of a transcription factor that is a predicted target of that mirna (according to MIRDB) were assigned to the transcription factor targeting model. 3 UTR luciferase reporter assays To evaluate mir- 29a binding to the MYCN 3 UTR, 74 bp oligonucleotides spanning the predicted 3 UTR mirna binding site flanked by XhoI and NotI restriction sites were cloned into psicheck2 (Promega) as described previously (Cloonan et al, 2008). Oligonucleotides with a mutated binding site were used as control. DLD1Dicer hypo cells were seeded at a density of 10,000 cells per well in an opaque 96- well plate. Twenty- four hours after seeding, cells were cotransfected with a mir- 29a pre- mir (Ambion) or negative control pre- mir (Ambion) in combination with the 3 UTR construct using DharmaFECT Duo (Dharmacon). Forty- eight hours after transfection, luciferase reporter gene activity was measured using the Dual- Glo Luciferase Assay System (Promega) and a FLUOstar OPTIMA microplate reader (BMG LABTECH). Western Blot NGP neurobblastoma cells were cultured in RPMI (Invitrogen) supplemented with 10% fetal calf serum and transfected with a mir- 29a pre- mir (Ambion) or negative control pre- mir (Ambion) as described previously (Mestdagh et al, 2010a). Cells were harvested 48h after transfection and proteins were isolated using the Nuclear Extract kit (Active Motif) according to the manufacturer s instructions. MYCN protein was detected using a monoclonal MYCN antibody (B8.4.B BD- Biosciences) and normalized to ACTB. Figure Legends Figure 1 mechanistic models of mirna- directed gene expression regulation Positive and negative mirna gene set associations, originating from positive and negative correlations between the mirna and the genes in the gene set, can be explained by one of the six proposed models of gene regulation. We define three models for negative mirna gene set associations, described as muli-
163 157 component targeting, component targeting and transcription factor targeting, and three models for positive mirna gene set associations, described as transcription factor targeting, targeting of a negative regulator and common transcriptional regulator. The schematic models represent simplified pathways or signaling cascades with receptors (R), pathway components (C) and transcriptional targets (T). Coding genes negatively correlated to the mirna are indicated in red, genes positively correlated to the mirna are indicated in blue. Figure 2 tissue specific mirna expression and function (A) Ranked expression map for 5 lymphatic system specific mirnas (columns) in 39 normal tissues (rows). Each sample is represented by a square, color- coded according to different body organ systems and ranked according to the expression of the respective mirnas. Samples with the highest expression are ranked on top. (B) Gene sets from the Gene Ontology Biological Proces gene set collection that are positively (red) and negatively (blue) associated with the mirnas. Only the most significant gene sets are shown. (C- F) Relative expression of mirnas annotated to the Gene Ontology Biological Process gene sets heart development, brain development, digestion and muscle development in the different body organ systems. Organ systems are ranked according to mirna expression. Figure 3 The MYCN transcription factor is a direct target of mir- 29a in neuroblastoma (A) Relative luciferase activity of a MYCN 3 UTR luciferase reporter vector containing the predicted mir- 29a binding site. Cotransfection of vector and pre- mir- 29a results in a significant decrease in luciferase activity compared to the negative control (* Student s t- test, p < 0.01). (B) MYCN expression, normalized to ACTB expression, in NGP cells transfected with a pre- mir negative control or pre- mir- 29a. Acknowledgements This research was funded by the Fund for Scientific Research (grant number: G and ), the Belgian Kid s Fund, and the Stichting tegen Kanker. P.M. is supported by the Ghent University Research Fund (BOF 01D31406). This article represents research results of the Belgian program of Interuniversity Poles of Attraction, initiated by the Belgian State, Prime Min- ister s Office, Science Policy Programming. The study was sponsored by the GOA (01G01910). References Baek D, Villen J, Shin C, Camargo FD, Gygi SP, Bartel DP (2008) The impact of micrornas on protein output. Nature 455: Chi SW, Zang JB, Mele A, Darnell RB (2009) Argonaute HITS- CLIP decodes microrna- mrna interaction maps. Nature 460:
164 158 Cloonan N, Brown MK, Steptoe AL, Wani S, Chan WL, Forrest AR, Kolle G, Gabrielli B, Grimmond SM (2008) The mir- 17-5p microrna is a key regulator of the G1/S phase cell cycle transition. Genome Biol 9: R127. He L, He X, Lim LP, de Stanchina E, Xuan Z, Liang Y, Xue W, Zender L, Magnus J, Ridzon D, Jackson AL, Linsley PS, Chen C, Lowe SW, Cleary MA, Hannon GJ (2007) A microrna component of the p53 tumour suppressor network. Nature 447: Kanehisa M, Goto S, Furumichi M, Tanabe M, Hirakawa M (2010) KEGG for representation and analysis of molecular networks involving diseases and drugs. Nucleic Acids Res 38: D Lu F, Weidmer A, Liu CG, Volinia S, Croce CM, Lieberman PM (2008) Epstein- Barr virus- induced mir- 155 attenuates NF- kappab signaling and stabilizes latent virus persistence. J Virol 82: Maris JM, Hogarty MD, Bagatell R, Cohn SL (2007) Neuroblastoma. Lancet 369: Mendell JT (2008) miriad roles for the mir cluster in development and disease. Cell 133: Mestdagh P, Bostrom AK, Impens F, Fredlund E, Van Peer G, De Antonellis P, von Stedingk K, Ghesquiere B, Schulte S, Dews M, Thomas- Tikhonenko A, Schulte JH, Zollo M, Schramm A, Gevaert K, Axelson H, Speleman F, Vandesompele J (2010a) The mir microrna cluster regulates multiple components of the TGF- beta pathway in neuroblastoma. Mol Cell 40: Mestdagh P, Feys T, Bernard N, Guenther S, Chen C, Speleman F, Vandesompele J (2008) High- throughput stem- loop RT- qpcr mirna expression profiling using minute amounts of input RNA. Nucleic Acids Res 36: e143. Mestdagh P, Fredlund E, Pattyn F, Schulte JH, Muth D, Vermeulen J, Kumps C, Schlierf S, De Preter K, Van Roy N, Noguera R, Laureys G, Schramm A, Eggert A, Westermann F, Speleman F, Vandesompele J (2010b) MYCN/c- MYC- induced micrornas repress coding gene networks associated with poor outcome in MYCN/c- MYC- activated tumours. Oncogene 29: Mestdagh P, Van Vlierberghe P, De Weer A, Muth D, Westermann F, Speleman F, Vandesompele J (2009) A novel and universal method for microrna RT- qpcr data normalization. Genome Biol 10: R64. Nam S, Kim B, Shin S, Lee S (2008) mirgator: an integrated system for functional annotation of micrornas. Nucleic Acids Res 36: D Selbach M, Schwanhausser B, Thierfelder N, Fang Z, Khanin R, Rajewsky N (2008) Widespread changes in protein synthesis induced by micrornas. Nature 455: Su N, Wang Y, Qian M, Deng M (2010) combinatorial regulation of transcription factors and micrornas. BMC Syst Biol 4: 150. Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP (2005) Gene set enrichment analysis: a knowledge- based approach for interpreting genome- wide expression profiles. Proc Natl Acad Sci U S A 102: Tan LP, Seinen E, Duns G, de Jong D, Sibon OC, Poppema S, Kroesen BJ, Kok K, van den Berg A (2009) A high throughput experimental approach to identify mirna targets in human cells. Nucleic Acids Res 37: e137.
165 159 Ulitsky I, Laurent LC, Shamir R (2010) Towards computational prediction of microrna function and activity. Nucleic Acids Res 38: e160. Voorhoeve PM, le Sage C, Schrier M, Gillis AJ, Stoop H, Nagel R, Liu YP, van Duijse J, Drost J, Griekspoor A, Zlotorynski E, Yabuta N, De Vita G, Nojima H, Looijenga LH, Agami R (2006) A genetic screen implicates mirna- 372 and mirna- 373 as oncogenes in testicular germ cell tumours. Cell 124: Wang X (2008) mirdb: a microrna target prediction and functional annotation database with a wiki interface. RNA 14: Wang X, El Naqa IM (2008) Prediction of both conserved and nonconserved microrna targets in animals. Bioinformatics 24: Westermann F, Muth D, Benner A, Bauer T, Henrich KO, Oberthuer A, Brors B, Beissbarth T, Vandesompele J, Pattyn F, Hero B, Konig R, Fischer M, Schwab M (2008) Distinct transcriptional MYCN/c- MYC activities are associated with spontaneous regression or malignant progression in neuroblastomas. Genome Biol 9: R150. Xiao C, Srinivasan L, Calado DP, Patterson HC, Zhang B, Wang J, Henderson JM, Kutok JL, Rajewsky K (2008) Lymphoproliferative disease and autoimmunity in mice with increased mir expression in lymphocytes. Nat Immunol 9: Yamakuchi M, Lotterman CD, Bao C, Hruban RH, Karim B, Mendell JT, Huso D, Lowenstein CJ (2010) P53- induced microrna- 107 inhibits HIF- 1 and tumour angiogenesis. Proc Natl Acad Sci U S A 107: Yin Q, Wang X, McBride J, Fewell C, Flemington E (2008) B- cell receptor activation induces BIC/miR- 155 expression through a conserved AP- 1 element. J Biol Chem 283: Zhou B, Wang S, Mayr C, Bartel DP, Lodish HF (2007) mir- 150, a microrna expressed in mature B and T cells, blocks early B cell development when expressed prematurely. Proc Natl Acad Sci U S A 104:
166 160 Figure 1 A B mirna expression mirna expression mrna expression mrna expression GSEA GSEA multi-component targeting receptor targeting transcription factor targeting targeting negative regulator common transcriptional activator/repressor mir mir C R R T1 T2 mir mir C1 C T1 mir C1 C2 C1 mir C C2 mir T1 T2 C2 T2 T1 C3 T2 mir T1 T1 C3 C3 T2 T2 R: pathway receptor gene negatively correlated to mirna C: pathway component T: pathway transcriptional target gene positively correlated to mirna transcriptional activation or repression
167 161 Figure 2
168 162 Figure 3 A 1.2 B 1.2 relative luciferase signal MYCN protein expression vector pre-mir negative control pre-mir-29a negative control pre-mir-29a
169 163 Supplemental Data mirna prediction databases The following mirna target prediction databases were used: MIRDB release 14 (Wang, 2008; Wang and El Naqa, 2008), Targetscan 5.1 (Friedman et al, 2009), MICROCOSM v5 (Griffiths- Jones et al, 2008), DIANA 3.0 (Maragkakis et al, 2009a; Maragkakis et al, 2009b), RNA22 (august 2007) (Miranda et al, 2006) and PITA v6 (Kertesz et al, 2007). Impact of sample size on predicted functions In order to compare mirna predictions between the myeloma and neuroblastoma dataset we first evaluated the impact of the sample size on mirna annotations, as the number of samples between both datasets differs (the neuroblastoma dataset consists of 99 tumour samples while the myeloma dataset contains 60 tumour samples). We repeatedly (n = 50) selected 60 random samples within the neuroblastoma dataset to evaluate the impact on the inferred mirna annotations. Individual mirna mrna correlations were recalculated for a selection of 6 mirnas and GSEA was performed to determine mirna functions as described earlier. For each of these mirnas, significant gene sets that were identified when using the entire dataset were compared to those obtained when selecting only 60 samples. The most significant mirna - gene set associations (GSEA FDR=0) remained significant when reducing the sample size to 60 whereas for less significant mirna - gene set associations this was not always the case (Supplemental Figure 3). Supplemental References Fontana L, Fiori ME, Albini S, Cifaldi L, Giovinazzi S, Forloni M, Boldrini R, Donfrancesco A, Federici V, Giacomini P, Peschle C, Fruci D (2008) Antagomir- 17-5p abolishes the growth of therapy- resistant neuroblastoma through p21 and BIM. PLoS One 3: e2236. Friedman RC, Farh KK, Burge CB, Bartel DP (2009) Most mammalian mrnas are conserved targets of micrornas. Genome Res 19: Griffiths- Jones S, Saini HK, van Dongen S, Enright AJ (2008) mirbase: tools for microrna genomics. Nucleic Acids Res 36: D Kertesz M, Iovino N, Unnerstall U, Gaul U, Segal E (2007) The role of site accessibility in microrna target recognition. Nat Genet 39: Ma L, Young J, Prabhala H, Pan E, Mestdagh P, Muth D, Teruya- Feldstein J, Reinhardt F, Onder TT, Valastyan S, Westermann F, Speleman F, Vandesompele J, Weinberg RA (2010) mir- 9, a MYC/MYCN- activated microrna, regulates E- cadherin and cancer metastasis. Nat Cell Biol 12:
170 164 Maragkakis M, Alexiou P, Papadopoulos GL, Reczko M, Dalamagas T, Giannopoulos G, Goumas G, Koukis E, Kourtis K, Simossis VA, Sethupathy P, Vergoulis T, Koziris N, Sellis T, Tsanakas P, Hatzigeorgiou AG (2009a) Accurate microrna target prediction correlates with protein repression levels. BMC Bioinformatics 10: 295. Maragkakis M, Reczko M, Simossis VA, Alexiou P, Papadopoulos GL, Dalamagas T, Giannopoulos G, Goumas G, Koukis E, Kourtis K, Vergoulis T, Koziris N, Sellis T, Tsanakas P, Hatzigeorgiou AG (2009b) DIANA- microt web server: elucidating microrna functions through target prediction. Nucleic Acids Res 37: W Mestdagh P, Bostrom AK, Impens F, Fredlund E, Van Peer G, De Antonellis P, von Stedingk K, Ghesquiere B, Schulte S, Dews M, Thomas- Tikhonenko A, Schulte JH, Zollo M, Schramm A, Gevaert K, Axelson H, Speleman F, Vandesompele J (2010a) The mir microrna cluster regulates multiple components of the TGF- beta pathway in neuroblastoma. Mol Cell 40: Mestdagh P, Fredlund E, Pattyn F, Schulte JH, Muth D, Vermeulen J, Kumps C, Schlierf S, De Preter K, Van Roy N, Noguera R, Laureys G, Schramm A, Eggert A, Westermann F, Speleman F, Vandesompele J (2010b) MYCN/c- MYC- induced micrornas repress coding gene networks associated with poor outcome in MYCN/c- MYC- activated tumours. Oncogene 29: Miranda KC, Huynh T, Tay Y, Ang YS, Tam WL, Thomson AM, Lim B, Rigoutsos I (2006) A pattern- based method for the identification of MicroRNA binding sites and their corresponding heteroduplexes. Cell 126: Wang X (2008) mirdb: a microrna target prediction and functional annotation database with a wiki interface. RNA 14: Wang X, El Naqa IM (2008) Prediction of both conserved and nonconserved microrna targets in animals. Bioinformatics 24: Yamakuchi M, Lotterman CD, Bao C, Hruban RH, Karim B, Mendell JT, Huso D, Lowenstein CJ (2010) P53- induced microrna- 107 inhibits HIF- 1 and tumour angiogenesis. Proc Natl Acad Sci U S A 107:
171 Supplemental Figure 1 Validated mirna functions for each of the models of mirna- directed gene expression regulation 165 (A) mirnas from the mir cluster are negatively correlated to a gene set containing TGFβ- target genes. mir mirnas have been shown to target multiple components of the TGFβ- pathway (Mestdagh et al, 2010a). (B) mir- 9 is negatively correlated to a set of CDH1 responsive genes. mir- 9 has been shown to target CDH1, a component of the CDH1 CTNNB1 axis (Ma et al, 2010). (C) mir- 107 is negatively correlated to a set of Hypoxia activated genes that is enriched for targets of the HIF1 transcription factor. mir- 107 has been shown to target HIF1B (Yamakuchi et al, 2010). (D) mir- 17 is positively correlated to cell cycle genes. mir- 17 is known to target CDKN1A, a negative regulator of the cell cycle (Fontana et al, 2008). (E) mir- 181a is positively correlated to a set of MYC target genes. mir- 181a is a direct transcriptional target of the MYCN transcription factor (Mestdagh et al, 2010b). A B C D E multi-component targeting component targeting transcription factor targeting targeting negative regulator common transcriptional activator/repressor mir TGF! mir-9 E-cadherin mir-107 HIF1 mir-17 CDKN1A MYCN mir-181a '&(6*)('!-34!')+3)+,9:/. 345'()"5#/)+&(,'+!)2)54. %&4&63)#*-378&)$-. "'66)"*"6')-(3"'!!.!"#$#%&"#'()%*")+&(,'+!)$ / /. 0/. 0/ /. 0/. /. 0/ /. 0/ mir-17 mir-9 mir-18a TGFBR2 CDH1 mir-107 HIF1 PGF mir-17 CDKN1A MYCN mir-181a AHCY mir-18a SMAD2 SMAD4 mir-17 CTNNB1 KRT18 CD24! PDGFB! CCNE1 E2F1 CCNA2 PAICS! SMAD2 SMAD4 CDKN1A BCL2L11 E2F1! mir-17! gene negatively correlated to mirna gene positively correlated to mirna
172 166 Supplemental Figure 2 Evaluation of mirna target prediction algorithms (A) Fold downregulation (log 2 ) of predicted targets, according to 7 different prediction algorithms, in 8 mirna perturbation experiments. Each bar represents the mean fold change of protein expression ± SEM. The highest downregulation was observed for targets predicted by MIRDB. (B) Cumulative distribution of protein fold change for predicted targets from 7 different prediction algorithms. MIRDB predictions contain the highest fraction of downregulated proteins. For visualization purposes, the limits of the X- axis were set at - 1 and 1. (C) Fold enrichment of predicted targets for different cut- offs of protein downregulation relative to the background (defined as those proteins with a log 2 protein fold change > 0). Within the group of proteins that are downregulated at least 2- fold (log 2 fold change < - 1), MIRDB predicts 12 times more targets than in the background set. (D) Fold downregulation (log 2 ) of mirna targets predicted by a combination of databases or by MIRDB alone. A C fold downregulation (log2) of predicted targets MIRDB MICROCOSM TARGETSCAN TARGETSCAN_CONS DIANA RNA22 PITA B D cummulative fraction (%) MIRDB MICROCOSM TARGETSCAN DIANA TARGETSCAN_CONS RNA22 PITA protein fold change (log2) fold enrichment of predicted targets (relative to background) MIRDB MICROCOSM TARGETSCAN TARGETSCAN_CONS DIANA RNA22 PITA fold downregulation (log2) of predicted targets <-1 <-0.8 <-0.6 <-0.4 <-0.2 < MIRDB protein fold change (log2) number of databases
173 167 Supplemental Figure 3 impact of sample size on predicted mirna functions Each graph represents the overlap in predicted mirna functions when repeatedly (n = 50) sampling 60 samples from the neuroblastoma dataset. Significant gene sets, selected using different GSEA FDR cut- offs, that are identified when using the entire neuroblastoma dataset were compared to significant gene sets identified when using only 60 samples. Gene sets that were identified as significant in at least 40 out of 50 samplings (power 80%) were considered to be common between both datasets and thus independent of sample size. The Y- axis shows the percentage of gene sets that were identified in at least 40/50 samplings relative to the total number of gene sets that can be identified when using all samples (set to 100%). Positive mirna gene set correlations are indicated in dark grey, negative mirna gene set correlations in light grey. Most gene sets with a GSEA FDR = 0 are also identified when using only 60 samples. 100 mir mir-204 % significant gene sets in at least 80% of samplings GSEA FDR 0 > <0.001 >0.001 <0.01 >0.01 <0.05 all samples % significant gene sets in at least 80% of samplings GSEA FDR 0 > <0.001 >0.001 <0.01 >0.01 <0.05 all samples 100 mir-20a 100 mir-9 % significant gene sets in at least 80% of samplings GSEA FDR 0 > >0.001 <0.001 <0.01 >0.01 <0.05 all samples % significant gene sets in at least 80% of samplings GSEA FDR 0 > >0.001 <0.001 <0.01 >0.01 <0.05 all samples 100 mir-10b 100 mir-26a % significant gene sets in at least 80% of samplings GSEA FDR 0 > <0.001 * * * >0.001 <0.01 >0.01 <0.05 all samples % significant gene sets in at least 80% of samplings GSEA FDR 0 > >0.001 <0.001 <0.01 >0.01 <0.05 all samples
174 PAPER 7: Outcome prediction of children with neuroblastoma using mirna and mrna gene expression signatures 168 PAPER 7 Outcome prediction of children with neuroblastoma using mirna and mrna gene expression signatures. Mestdagh P*, De Preter K*, Vermeulen J*, Naranjo A, Bray I, Castel V, Chen C, Eggert A, Hogarty MD, London WB, Noguera R, Piqueras M, Bryan K, Schowe B, van Sluis P, Molenaar JJ, Schramm, Schulte JH, Stallings RL, Versteeg R, Laureys G, Van Roy N, Speleman F, Vandesompele J. Submitted. *Equally contributing authors
175 169 Outcome prediction of children with neuroblastoma using mirna and mrna gene expression signatures Pieter Mestdagh 1*, Katleen De Preter 1*, Joëlle Vermeulen 1*, Arlene Naranjo 3, Isabella Bray 4, Victoria Castel 5, Caifu Chen 7, Angelika Eggert 8, Michael D Hogarty 9, Wendy B London 10, Rosa Noguera 6, Marta Piqueras 6, Kenneth Bryan 4, Benjamin Schowe 11, Peter van Sluis 12, Jan J. Molenaar 12, Alexander Schramm 8, Johannes H. Schulte 8, Raymond L. Stallings 4, Rogier Versteeg 12, Geneviève Laureys 2, Nadine Van Roy 1, Frank Speleman 1, Jo Vandesompele 1 *shared 1st authors 1 Center for Medical Genetics, Ghent University Hospital, Ghent, Belgium, 2 Department of Paediatric Hematology and Oncology, Ghent University Hospital, Ghent, Belgium; 3 Children's Oncology Group, University of Florida, Gainesville, FL, USA, 4 Department of Cancer Genetics, Royal College of Surgeons in Ireland, Dublin, Ireland and National Children s Research Centre; 5 Pediatric Oncology Unit, Hospital La Fe, Valencia, Spain, 6 Department of Pathology, Medical School, University of Valencia, Spain; 7 Applied Biosystems, Foster City, CA, USA; 8 Department of Pediatric Oncology and Haematology, University Children's Hospital Essen, Essen, Germany; 9 Division of Oncology, The Children's Hospital of Philadelphia, Philadelphia, USA; 10 Children s Oncology Group, Children s Hospital Boston/Dana- Farber Harvard Cancer Center, Boston, MA, USA; 11 Department of Computer Science, Technische Universität Dortmund, Baroper Str. 301, Dortmund, Germany 12 Department of Human Genetics, Academic Medical Center, Amsterdam, the Netherlands. Abstract Purpose: More accurate assessment of prognosis is important to further improve the choice of risk- related therapy in neuroblastoma (NB) patients. In this study we aimed to establish and validate a prognostic microrna (mirna) signature for children with NB and compare it with a previously established 59 mrna classifier. Patients and Methods: 430 human mature mirnas were profiled in two patient subgroups with maximally divergent clinical courses. Univariate logistic regression analysis was used to select mirnas that correlated with NB patient survival. Subsequently, a 25- mirna gene signature was built using 51 training samples, tested on 179 test samples, and validated on an independent set of 304 tumours. Results: The 25- mirna signature significantly discriminates the test patients with respect to progression- free survival (PFS) and overall survival (OS) (p<0.0001). Multivariate analysis indicates that the mirna signature is an independent predictor of PFS and OS after controlling for currently used risk factors. Patients with increased risk for a shorter PFS and OS can also be identified in the cohort of high risk patients. The results were confirmed in an external validation set. On the same sample set, we also tested
176 170 the 59 mrna gene expression classifier and showed that the mirna signature does not outperform the mrna classifier. Conclusion: In this study we present the largest NB mirna expression study so far, including more than 500 NB patients. We established and validated a robust mirna classifier, able to identify a cohort of high risk NB patients at greater risk for adverse outcome. Introduction Given the clinical heterogeneity of neuroblastoma (NB), accurate prognostic classification of patients with this tumour is one of the major challenges to improve the choice of risk- related therapy. Clinical experience with the currently used risk stratification systems suggests that the stratification of patients for treatment is useful 1. Nevertheless, patients with the same clinicopathological and genetic parameters, receiving the same treatment, can have markedly different clinical courses. Based on the assumption that differences in outcome are mainly driven through underlying genetic and biological characteristics, gene expression studies have been undertaken in order to detect new prognostic markers and to establish gene expression based classifiers for improved neuroblastoma patient care. Recently, we established a messenger RNA (mrna) gene expression classifier and validated the performance of this classifier in two independent patient cohorts 2. Recently, an important class of small non- coding RNAs (ncrnas) regulating mrna expression, termed micrornas (mirnas), have been shown to be implicated in various cancers, including NB 3,4. Here we set out to perform an in- depth study on an unprecedentedly large cohort of primary untreated neuroblastoma tumours in order to unequivocally establish the possible prognostic power of mirna classifiers. To this purpose, we tested and validated the performance of a 25- mirna prognostic classifier in two independent patient cohorts and compared it with our previously published 59 mrna expression classifier. Methods Patients / Tumour samples The first train- test cohort consisted of 230 NB patients (27 from Ghent, Belgium, 54 from Essen, Germany, 31 from Valencia, Spain, 43 from Dublin, Ireland and 75 from Amsterdam, The Netherlands). Patients were only included if primary untreated NB tumour RNA (at least 60% tumour cells and confirmed histological diagnosis of NB) was present. All patients provided consent and were enrolled on at least one International Society of Paediatric Oncology, European Neuroblastoma Group (SIOPEN), the Gesellschaft fuer Paediatrische Onkologie und Haematologie (GPOH), the Children s Oncology Group (COG- United States) study Dutch Childhood Oncology Group (DCOG, Amsterdam) or the Our Lady s Children s Hospital (Dublin). The median follow- up was 68 months (range months), and was greater than 36 months for most
177 171 patients without an event (114/139 =82%). At the time of analysis, 156 of 230 patients were alive. All patients were treated according to similar protocols. The validation cohort consisted of 304 patients from the COG: including 128 cases with an event and 189 patients without an event, with at least 36 months of follow- up. All laboratory analyses were performed blinded to clinical and outcome data. All patients provided consent and were enrolled on at least one COG study, and all participating institutions had institutional review board approval to take part in the COG studies. This study was approved by the Ghent University Hospital Ethical Committee (EC2008/159). mirna expression profiling Total RNA was extracted in different national reference laboratories using a silica gel- based membrane purification method (mirnaeasy kit,qiagen) or by a phenol- based method (TRIzol reagent, Life Technologies) according to the manufacturer's instructions. For the train- test cohort, reverse transcription (starting from 20 ng of total RNA) and mirna expression profiling of 430 human mirnas was performed as described earlier 5. RNA samples from the validation cohort were only profiled for the selected set of 25 prognostic mirnas. Data normalization was performed using the mean expression of 4 selected stably expressed reference mirnas (hsa- mir- 125a, hsa- mir- 99b, hsa- mir- 423 and hsa- mir- 425) according to Mestdagh et al 6. The data sets from the different centers were independently standardized (mean centered and autoscaled 7 ). mirna expression data and clinical sample annotation are available in rdml- format 8 (supplemental file 1). mrna expression profiling Expression analysis of 59 prognostic mrna genes was performed according to a procedure described elsewhere 2. In brief, RT- qpcr was used to measure 59 prognostic genes and 5 reference sequences. Real- time qpcr was performed in a 384- well plate instrument (CFX384, Bio- Rad). Biogazelle s qbase PLUS software was used for data- analysis (normalization, error propagation, inter- run calibration). Results were standardized (mean centered and autoscaled). mrna expression data are available in rdml- format (supplemental file 1). Statistical analysis For data- analysis the tumour samples were divided into a training set and a test set. The training set was consisted of 51 samples randomly selected from two patient subgroups with maximally divergent clinical courses: 24 low- risk patients with INSS stage 1, 2, or 4S without MYCN amplification and with PFS time of at least 1000 days and 27 deceased high- risk patients older than 12 months at diagnosis with INSS stage 4 tumour (irrespective of the MYCN gene status) or with INSS stage 2 or 3 tumour with MYCN amplification. Univariate logistic regression analysis was used to select the top 25 mirna genes with prognostic relevance in NB.
178 172 The 25 mirna and 59 mrna expression signatures were built using the same training set, tested on the remaining 179 samples, and validated in a blind manner on the validation cohort of 304 COG samples. For the test cohort, the R- language for statistical computing (version 2.9.0) was used to train and test the prognostic signature using the Prediction Analysis of Microarrays (PAM) method 9 (Bioconductor MCR estimate package 10 ), to evaluate its performance by receiver operating characteristic (ROC) curve and area under the curve (AUC) analyses (ROCR package), and for Kaplan Meier survival analyses (survival package). Multivariate logistic regression analyses were done using PASW Statistics (version 18). Currently used risk factors such as age at diagnosis ( 12 months vs <12 months), International Neuroblastoma Staging System (INSS) stage (stage 4 vs not stage 4), and MYCN status (amplified vs not amplified) were tested, and variables with p values less than 0.05 were retained in the model. Since an interaction between the signature and risk factors was not expected to occur, interaction terms were not included in the models. For ROC and multivariate analyses, only patients with an event and patients with sufficient follow- up time ( 36 months) if no event occurred were included, since 95% of events in NB are expected to occur within the first 36 months after diagnosis. A case control study was set up to validate the signature in the COG cohort. This was done to ensure a sufficient number of events in each risk group i.e., to increase the power from what would have resulted from a random sample. Failure (cases) was defined as relapse, progression, or death from disease (progression- free survival), and death (overall survival) within a 3- year follow- up period, and control defined as non- failure in the first 3 years of follow- up. Controls and cases with complete data were selected in a 2:1 ratio to increase the sample size and power. Multivariate logistic regression analyses were done to determine whether the signature was a significant independent predictor after controlling for known risk factors. Statistical analyses were done with SAS (version 9). The final logistic regression model used 50 cases and 100 randomly- selected controls with complete data for PFS and 37 cases and 74 randomly- selected controls with complete data for OS. Results Establishment of a prognostic mirna signature To establish and train a prognostic mirna signature, we used mirna expression data from 24 low- risk patients with a long PFS time and 27 deceased high- risk patients. Using the top 25 mirnas with the highest correlation with OS (Table 1), a 25 mirna expression signature was built. Further in the text we will refer to this signature as a molecular indicator for low or high risk separating patients with low versus high risk.
179 173 Validation of the prognostic 25- mirna signature This 25 mirna expression signature significantly distinguished the remaining 179 patients of a first cohort of NB patients with respect to PFS and OS (p<0.0001; Figure 1). PFS 5 years from the date of diagnosis was 84.3 (95% CI ) for the group of patients with a molecular indicator for low risk, compared with 37.2 (95% CI ) for the group of patients with a molecular indicator for high risk. The 5- year OS was 91.0% ( ) and 44.5% ( ) in the low and high molecular risk groups, respectively. Subsequently, we tested the signature within the group of low- risk NB patients with localized disease treated with surgery alone or in combination with mild chemotherapy and within the commonly defined high- risk group based on the different current risk stratification systems (Europe, USA and Germany) 11. Patients with increased risk for disease progression or relapse could be identified in both current low- and high- risk groups (p=0.014 and p= respectively). While the signature was also useful in identifying those patients at increased risk for death in the current high- risk group (p=0.0015), there was no difference in OS between patients with a molecular indicator for high- and low- risk in the current low- risk group (Figure 2). Multivariate logistic regression analysis including the mirna signature, MYCN status, age at diagnosis and INSS stage, revealed that the mirna prognostic signature is an independent marker for both PFS and OS in the global cohort as well as in the high- risk subgroup (Table 2). Within the low- risk subgroup of patients the mirna classifier was shown to be an independent predictor for PFS (Table 2). The probability that a patient will be correctly classified by the signature based on a ROC- curve analysis (AUC) was 78.1% (95% CI ) and 77.1% ( ) for OS and PFS, respectively. The signature predicted OS with a sensitivity of 83.0% (39/47) and a specificity of 73.3% (74/101). To validate the mirna signature in a second completely independent patient cohort, 304 COG tumours were tested in a blind manner. The same signature as used for the test cohort identified COG patients who were at greater risk for progression or relapse. Multivariate logistic regression analysis including the mirna signature, MYCN status, age, INSS stage, ploidy, International Neuroblastoma Pathology Classification (INPC), grade of differentiation and mitosis karyorrhexis index (MKI) showed that the mirna signature was an independent significant predictor for PFS (odds ratio (OR) 3.861, 95% CI ). This was not the case for OS where the final logistic regression model involved only INSS stage and ploidy (Table 3). Comparison of the performances of the prognostic mirna signature with the prognostic 59 mrna signature In order to establish the prognostic value of this 25- mirna signature in relation to our recently published 59- mrna signature 2, we compared performances and prognostic power through survival analysis and multivariate analysis including both mirna and mrna signatures along with other currently used risk factors. Those analyses showed that the mrna classifier performs better than the mirna classifier (Supplemental Table 1). Final backward- selected logistic regression model for PFS and OS testing the mrna
180 174 and mirna classifiers and covariates demonstrated that the mrna was found to be a statistically significantly independent outcome predictor whereas the mirna classifier dropped out in both validation sets (Table 4). Discussion Molecular risk classification of NB patients based on gene expression profiling is expected to contribute significantly to improved NB outcome prediction with the ultimate aim of tailoring the treatment to the severity of the disease. To this purpose, several mrna gene expression classifiers were previously developed. Given the fact that a single mirna, targeting several mrnas, may have broader effects than a single mrna and that mirnas are less sensitive to RNA degradation than mrnas, mirna classifiers might present with certain advantages compared to mrna classifiers. In this paper we successfully developed a mirna signature that was subsequently validated in the largest NB series till now. Importantly, we demonstrate the ability of the signature to identify patients at ultra- high risk with respect to disease outcome within the current high- risk NB group for which no clinical or genetic markers are available today.. We further compared the performance of this signature with our recently published 59- mrna signature 2. In order to prove the robustness of a given signature, it should be validated under conditions that simulate the prospective broad clinical application of the assay and that reflect the various potential sources of assay variability, such as tissue handling, RNA extraction method, patient ethnicity, and treatment with other drugs. In this study we performed an external validation study on a completely independent cohort of COG patients whereby laboratory analyses were performed blinded to clinical and outcome data. Also in this validation set, the signature was found to be a statistically significant independent risk predictor. For the mirna expression profiling, we used a high- throughput quantitative PCR based stem- loop RT- primer method 5 along with a universally applicable data normalization method that had been successfully validated in our lab 6. Advantages of this method over the microarray technology are the higher cost- efficiency (definitely when measuring a 25- gene set), the shorter time to results, and the higher sensitivity (hence need of less input material). The use of a sample pre- amplification method enabled the maximization of the number of tumour samples available for this study through collaborative studies with international research laboratories; every laboratory could readily provide 100 ng of RNA. This is important, especially for pediatric cancers as biopsies are often very small and the material available limited. The use of a PCR based method will definitely contribute to the development of a prognostic test that can be implemented in the clinic. A prognostic classifier can be clinically useful without the understanding of the mechanistic relationship of the genes or the interpretation of the meaning of the individual genes included in the expression signature and clear biological elucidation might be more difficult to achieve than accurate classification 12. However, the presence of many genes implicated in NB biology in our recently published prognostic 59- mrna
181 175 signature 2 suggests that amongst the 25 mirnas in the present signature a significant portion may also be of direct biological relevance and may offer new opportunities for molecular therapy. As it is, all but one member of the mir cluster is part of the prognostic signature. These mirnas play a role in different tumourigenic pathways such as angiogenesis, cell adhesion and migration, cell cycle regulation and proliferation, and negative regulation of apoptosis 13,14. Furthermore, 16 of the 25 mirnas are MYC/MYCN driven 15. Mir- 26a and mir- 125b are potential targets for therapy as they are highly expressed in all normal tissues, are known tumour suppressor mirnas and candidates for replacement therapy 16. Gene ontology analysis of the 25 mirnas using the mirna body map ( showed a significant enrichment of GO terms for cell cycle, immune response, cell adhesion and neuronal differentiation (Supplemental Table 2). Similar gene ontology classes are enriched in the 59 mrna gene set pointing at common biological processes that are represented by these prognostic classifiers. Furthermore, target analysis demonstrated that several of the 25 mirnas target one or more of the 59 mrna genes (Supplemental Table 3). Further functional studies are required to demonstrate the possible mechanistic relevance of some of the 25 mirnas in NB pathogenesis. Chen and colleagues were the first to show that prognostic subgroups of NBs are characterized by specific mirna expression profiles 3. It was further demonstrated that the prognostic power of a mirna signature is superior to that of an individual mirna gene 4. These results, together with those of the present study, demonstrate the potential of mirna- based stratification of NB patients according to clinical outcome. Based on the presumed higher stability of mirnas, their broad role in transcriptional regulation, and their superior tumour type classification performance 17,18, it was hypothesized that our 25- mirna classifier would outperform the 59- mrna classifier. In order to test this hypothesis, we evaluated the 59- mrna signature in all samples included in this study and compared the performance of this signature to the performance of our 25- mirna signature. Surprisingly, overall results demonstrated a slight superiority of the mrna signature (Supplemental Table 1). To exclude the possibility that we have established a suboptimal mirna classifier, we compared its performance with that of two published mirna classifiers 4,19 and could show that both classifiers are less performant than the 25 mirna classier, and hence also don t outperform the 59 mrna classifier (Supplemental Table 1). A possible explanation for this observation is the fact that the variation in mrna expression was significantly higher than the variation in mirna expression (Supplemental Figure 1A, Mann Whitney, p < ) in the tested patients, possibly allowing for a more robust separation between groups. When comparing the expression fold change between deceased patients and patients that are alive, we indeed observe a significantly higher fold change for mrna genes as compared to mirna genes (Supplemental Figure 1B, Mann Whitney, p < ). In addition, the superior histological classification performance of mirnas has only been demonstrated when comparing very different tissues 17,18. It is not inconceivable that mirnas indeed represent very good tissue specific markers, but are less powerful for classification of more homogeneous samples, like a single tumour type. Importantly, while this is the first study that directly compares prognostic mirna and mrna
182 176 classifiers in human cancer, the mrna superiority has only been proven in case of fresh- frozen biopsy material. A well known technical obstacle to gene expression studies and to widespread application of molecular diagnostic tests in the current clinical workflow is the fact that often minimal (if any) amounts of fresh frozen tissue are procured. A step forward to overcome this problem is the use of archived formalin- fixed and paraffin- embedded (FFPE) tissues. Such samples are widely available, stable at room temperature, easily storable and extraction procedures are standardized in most laboratories. MiRNA signatures might present with some advantages compared to mrna classifiers in FFPE tissues as it is hypothesized that these small molecules of nucleotides are much less sensitive to RNA degradation. This opens new perspectives for the clinical application of our mirna- based prognostic test. In conclusion, the results obtained from this study clearly illustrate the power of mirna expression analysis in the risk classification of NB patients. The applied method and signature are suitable for routine laboratory testing. A future challenge is the validation of this signature in archived fixed tissues. Acknowledgements Funding: Ghent University Research Fund (BOF 01D31406; PM), the Fund for Scientific Research Flanders (KDP), the Belgian Kid s Fund and the Fondation Nuovo- Soldati (J. Vermeulen), the Fondation Fournier Majoie pour l Innovation, the Belgian Federal Public Health Service, the Association Hubert Gouin «Enfance et Cancer», the Flemish League against Cancer, the Children Cancer Fund Ghent, the Belgian Society of Paediatric Haematology and Oncology, the Fund for Scientific Research Flanders (grant number: G ), the Institute for the Promotion of Innovation by Science and Technology in Flanders, Strategisch basisonderzoek (IWT- SBO 60848), Children s Oncology Group grants (U10 CA98413 and U10 CA98543) and the Instituto Carlos III,RD 06/0020/0102 Spain. This article presents research results of the Belgian program of Interuniversity Poles of Attraction, initiated by the Belgian State, Prime Minister's Office, Science Policy Programming. We acknowledge the support of the European Community (FP6: STREP: EET- pipeline, number: ). RLS was a recipient of grants from Science Foundation Ireland (07/IN.1/B1776), the Children s Medical and Research Foundation, and the NIH (5R01CA127496). We thank Els De Smet, Nurten Yigit, Gaëlle Van Severen, Justine Nuytens, Sander Anseeuw and Liesbeth Vercruysse for their excellent technical assistance. We are indebted to all members of the SIOPEN, GPOH and COG for providing tumour samples or the clinical history of patients. References 1. Vermeulen J, De Preter K, Mestdagh P, Laureys G, Speleman F, Vandesompele J. Predicting outcomes for children with neuroblastoma. Discov Med. Jul;10(50):29-36.
183 Vermeulen J, De Preter K, Naranjo A, Vercruysse L, Van Roy N, Hellemans J, et al. Predicting outcomes for children with neuroblastoma using a multigene- expression signature: a retrospective SIOPEN/COG/GPOH study. Lancet Oncol Jul;10(7): Chen Y, Stallings RL. Differential patterns of microrna expression in neuroblastoma are correlated with prognosis, differentiation, and apoptosis. Cancer Res Feb 1;67(3): Schulte JH, Schowe B, Mestdagh P, Kaderali L, Kalaghatgi P, Schlierf S, et al. Accurate prediction of neuroblastoma outcome based on mirna expression profiles. Int J Cancer. May Mestdagh P, Feys T, Bernard N, Guenther S, Chen C, Speleman F, et al. High- throughput stem- loop RT- qpcr mirna expression profiling using minute amounts of input RNA. Nucleic Acids Res Dec;36(21):e Mestdagh P, Van Vlierberghe P, De Weer A, Muth D, Westermann F, Speleman F, et al. A novel and universal method for microrna RT- qpcr data normalization. Genome Biol. 2009;10(6):R Willems E, Leyns L, Vandesompele J. Standardization of real- time PCR gene expression data from independent biological replicates. Anal Biochem Aug 1;379(1): Tibshirani R, Hastie T, Narasimhan B, Chu G. Diagnosis of multiple cancer types by shrunken centroids of gene expression. Proc Natl Acad Sci U S A May 14;99(10): Ruschhaupt M, Huber W, Poustka A, Mansmann U. A compendium to ensure computational reproducibility in high- dimensional classification tasks. Stat Appl Genet Mol Biol. 2004;3:Article De Preter K, Vermeulen J, Brors B, Delattre O, Eggert A, Fischer M, et al. Accurate outcome prediction in neuroblastoma across independent data sets using a multigene signature. Clin Cancer Res. Mar 1;16(5): Simon R. Roadmap for developing and validating therapeutically relevant genomic classifiers. J Clin Oncol Oct 10;23(29): Dews M, Homayouni A, Yu D, Murphy D, Sevignani C, Wentzel E, et al. Augmentation of tumour angiogenesis by a Myc- activated microrna cluster. Nat Genet Sep;38(9): Fontana L, Fiori ME, Albini S, Cifaldi L, Giovinazzi S, Forloni M, et al. Antagomir- 17-5p abolishes the growth of therapy- resistant neuroblastoma through p21 and BIM. PLoS One. 2008;3(5):e Mestdagh P, Fredlund E, Pattyn F, Schulte JH, Muth D, Vermeulen J, et al. MYCN/c- MYC- induced micrornas repress coding gene networks associated with poor outcome in MYCN/c- MYC- activated tumours. Oncogene. Mar 4;29(9): Kota J, Chivukula RR, O'Donnell KA, Wentzel EA, Montgomery CL, Hwang HW, et al. Therapeutic microrna delivery suppresses tumourigenesis in a murine liver cancer model. Cell Jun 12;137(6): Lu J, Getz G, Miska EA, Alvarez- Saavedra E, Lamb J, Peck D, et al. MicroRNA expression profiles classify human cancers. Nature Jun 9;435(7043):834-8.
184 Rosenfeld N, Aharonov R, Meiri E, Rosenwald S, Spector Y, Zepeniuk M, et al. MicroRNAs accurately identify cancer tissue origin. Nat Biotechnol Apr;26(4): Bray I, Bryan K, Prenter S, Buckley PG, Foley NH, Murphy DM, et al. Widespread dysregulation of MiRNAs by MYCN amplification and chromosomal imbalances in neuroblastoma: association of mirna expression with survival. PLoS One. 2009;4(11):e7850. Tables and Figures Figure 1: Kaplan Meier and log- rank analysis for progression- free (A) and overall (B) survival of the test cohort. The number of events is indicated between brackets. Figure 2: Kaplan Meier and log- rank analysis for progression- free (A,C) and overall (B,D) survival within the low- risk treatment group (C,D) and within the high- risk treatment group (A,B). The number of events is indicated between brackets. Table 1: 25 mirnas in the prognostic signature Table 2: Multivariate logistic regression analysis in the first test cohort including all patients, the low- risk patients, and the high- risk patients. Table 3: Multivariate logistic regression analysis in the second validation cohort Table 4: Final backward- selected logistic regression model testing the mrna and mirna classifiers and covariates in validation set 1 and validation set 2.
185 179 Figure 1 A B
186 180 Figure 2
187 181 Table 1 higher expressed in high- risk patients hsa- mir- 17 hsa- mir- 18a hsa- mir- 18a* hsa- mir- 19a hsa- mir- 20a hsa- mir- 20b hsa- mir- 92a hsa- mir- 15b hsa- mir- 572 hsa- mir- 192 hsa- mir- 25 hsa- mir- 320 hsa- mir- 345 hsa- mir- 93 higher expressed in low- risk patients hsa- mir- 125b hsa- mir- 26a hsa- mir- 190 hsa- mir- 30c hsa- mir- 326 hsa- mir- 488 hsa- mir- 500 hsa- mir- 628 hsa- mir p hsa- mir p hsa- mir- 204 Table 2 Variable p- value odds ratio 95% CI on odds ratio PFS (All patients, N=153) a mirna predictor (high vs. low risk) < ,737 2,463 13,360 Age at diagnosis (<>1 year) ,261 1,715 10,587 INSS stage (stage 4 vs. not stage 4) ,191 1,350 7,541 OS (All patients, N=147) a mirna predictor (high vs. low risk) Age at diagnosis (<>1 year) INSS stage (stage 4 vs. not stage 4) < PFS (low- risk patients, N=37) b mirna predictor (high vs. low risk) OS (low- risk patients, N=37) b no significant variables PFS (high- risk patients, N=68) a mirna predictor (high vs. low risk) INSS stage (stage 4 vs. not stage 4) OS (high- risk patients, N=63) a mirna predictor (high vs. low risk) a Variables tested in the model were: mirna predictor, MYCN status, age and INSS stage. b Variables tested in the model were: mirna predictor and age.
188 182 Table 3 Variable a p- value Odds ratio 95% CI on Odds ratio PFS (N=113) mirna predictor (high vs. low risk*) Grade (differentiating vs. undifferentiated and poorly differentiated*) OS (N=78) INSS stage (stage 4 vs. not stage 4*) Ploidy (diploid vs. hyperdiploid*) a Variables tested in the model: mirna predictor, MKI, MYCN status, age, ploidy, INSS stage, and grade. * Indicates the reference level for each variable. The odds ratio is the increased risk of an event in comparison to this reference level. Table 4 Validation set 1 p- value odds ratio 95% CI on odds ratio PFS (N=153) a mrna predictor (high vs. low risk * ) < Age at diagnosis (<> * 1 year) OS (N=147) a mrna predictor (high vs. low risk * ) < Age at diagnosis (<> * 1 year) INSS stage (stage 4 vs. not stage 4 * ) Validation set 2 PFS (N=113) b mrna predictor (high vs. low risk * ) Grade (differentiating vs. undifferentiated and poorly differentiated*) OS (N=78) c mrna predictor (high vs. low risk * ) < a Variables tested in the model were: age, stage, MYCN status, mrna classifier, mirna classifier. b Variables tested in the model and found not statistically significant were: MYCN status, MKI, age, ploidy, the mirna classifier, INSS stage, and grade. c Variables tested in the model and found not statistically significant were: the mirna classifier, age, MKI, MYCN status, and grade. * Indicates the reference level for each variable. The odds ratio is the increased risk of an event in comparison to this reference level.
189 Supplemental Data 183 Supplemental Table 1 : Comparison of 4 multigene prognostic signatures by means of Kaplan- Meier statistics, multivariate analysis and performance in the first validation set. 59 mrna 25 mirna mirna 1 mirna 2 overall survival all patients 2.15E E E E-03 Kaplan -Meier HR patiens 3.99E E E E-01 p-value LR patients 2.43E E E E-01 performance (AUC) odds ratio multi-variate p-value 1.95E E E E-01 progression free survival all patients 2.67E E E E-03 Kaplan -Meier HR patiens 5.43E E E E-01 p-value LR patients 3.01E E E E-01 performance (AUC) odds ratio multi-variate p-value 6.81E E E E-02 1 prognostic mirna signature by Schulte et al., 2 Prognostic mirna signature bu Bray et al., AUC: area under the curve, LR: low-risk patients, HR: high-risk patients, IR: intermediate-risk patients.
190 Supplemental Table 2: Gene Ontology Biological Process terms associated with the mrna and mirna signatures. mrna enriched GO BP terms intracellular signaling cascade cellular process nervous system development cell communication signal transduction metabolic process primary metabolic process cell adhesion cell motion cell-cell adhesion cell cycle ectoderm development cellular glucose homeostasis protein metabolic process developmental process system development cellular amino acid and derivative metabolic process phosphate metabolic process apoptosis female gamete generation homeostatic process nucleobase, nucleoside, nucleotide and nucleic acid metabolic process cell surface receptor linked signal transduction cell-matrix adhesion amino acid transport phosphate transport cytokinesis mitosis nucleobase, nucleoside, nucleotide and nucleic acid transport negative regulation of apoptosis 184 mirna enriched GO BP terms actin cytoskeleton organization and biogenesis actin filament based process adaptive immune response adaptive immune response anatomical structure formation biosynthetic process camp mediated signaling cell cell signaling cell cycle checkpoint cell cycle cell cycle phase cell cycle process cell division cell matrix adhesion cell substrate adhesion
191 185 cell surface receptor linked signal transduction cellular biosynthetic process cellular response to stimulus chromatin assembly or disassembly chromatin modification chromatin remodeling chromosome organization and biogenesis chromosome segregation coagulation cytokinesis DNA damage checkpoint DNA damage response signal transduction DNA dependent DNA replication DNA integrity checkpoint DNA metabolic process DNA packaging DNA recombination DNA repair DNA replication endosome transport enzyme linked receptor protein signaling pathway epidermal growth factor receptor signaling pathway establishment and or maintenance of chromatin architecture G1 S transition of mitotic cell cycle G protein signaling coupled to camp nucleotide second messenger homophilic cell adhesion I kappab kinase NF KappaB cascade immune system process inflammatory response interphase interphase of mitotic cell cycle M phase M phase of mitotic cell cycle meiosis I meiotic cell cycle membrane fusion membrane organization and biogenesis mesoderm development metal ion transport microtubule cytoskeleton organization and biogenesis mitochondrion organization and biogenesis mitosis mitotic cell cycle mitotic cell cycle checkpoint mrna metabolic process mrna processing negative regulation of DNA metabolic process negative regulation of signal transduction neuron development neuron differentiation nucleobase nucleoside and nucleotide metabolic process
192 186 one carbon compound metabolic process peptidyl tyrosine modification positive regulation of immune response positive regulation of signal transduction protein DNA complex assembly protein folding protein RNA complex assembly protein secretion ras protein signal transduction receptor mediated endocytosis regulation of anatomical structure morphogenesis regulation of blood pressure regulation of cell cycle regulation of cyclin dependent protein kinase activity regulation of DNA metabolic process regulation of DNA replication regulation of i kappab kinase nf kappab cascade regulation of immune response regulation of immune system process regulation of lymphocyte activation regulation of mitosis regulation of multicellular organismal process regulation of signal transduction response to DNA damage stimulus response to endogenous stimulus response to wounding ribonucleoprotein complex biogenesis and assembly RNA splicing RNA splicing via transesterification reactions small gtpase mediated signal transduction spliceosome assembly T cell activation tissue remodeling transcription from RNA polymerase III promoter translation transmembrane receptor protein tyrosine kinase signaling pathway trna metabolic process vasculature development vesicle mediated transport
193 Supplemental Table 3: mrnas from the 59 mrna expression classifier that are predicted to be targeted by a mirna of the 25 mirna expression signature, according to at least one of the four target prediction databases (TargetScan, mirdb, Diana, Microcosm) (number of databases) mirnas higher expressed in low risk-patients mrnas lower expressed in low-risk patients hsa-mir-190 PRAME (2), SLC25A5 (2), MRPL3 (1) hsa-mir-204 PRAME (3), SLC6A8 (2), NHLH2 (2), PAICS (2) hsa-mir-26a CDKN3 (1), NCAN (1), PRAME (1) hsa-mir-30c AHCY (1), NHLH2 (1) hsa-mir-326 CDCA5 (2), NCAN (2), BIRC5 (1), MCM2 (1) hsa-mir-485-5p CDCA5 (2), MYCN (2), SNAPC1 (2), PAICS (2), BIRC5 (2) hsa-mir-488 MYCN (2), PAICS (2), AHCY (2), TYMS (1) hsa-mir-500 NCAN (2), MCM2 (1) hsa-mir-542-3p BIRC5 (3), SLC6A8 (2), NCAN (2) mirnas higher expressed in high risk-patients mrnas lower expressed in high-risk patients hsa-mir-18a HIVEP2 (3), PRKACB (2), EPB41L3 (1), ARHGEF7 (1), CHD5 (1) CLSTN1 (3), PRKACB (2), DPYSL3 (2), CADM1 (2), MAP2K4 hsa-mir-192 (1), WSB1 (1), PDE4DIP (1), EPN2 (1), CHD5 (1), CAMTA1 (1) CAMTA1 (4), CHD5 (3), ULK2 (2), EPB41L3 (2), MAPT (2), hsa-mir-19a MAP2K4 (2), PRKACB (2), GNB1 (2), EPN2 (2), PDE4DIP (1), HIVEP2 (1) EPHA5 (3), CAMTA2 (2), PTPRF (2), CHD5 (2), MAP7 (2), hsa-mir-20b CLSTN1 (2), PIK3R1 (2), CAMTA1 (2), PDE4DIP (1), MAPT (1), GNB1 (1), PTN (1), MAP2K4 (1), ELAVL4 (1), PRKACB (1), ARHGEF7 (1) hsa-mir-25 MAP2K4 (3), PLAGL1 (1), EPN2 (1), PIK3R1 (1), CD44 (1), NRCAM (1), CAMTA1 (1) hsa-mir-345 CHD5 (2), PRDM2 (2), PTPRF (2), CLSTN1 (1), PTPRN2 (1), PTN (1), ARHGEF7 (1), PDE4DIP (1), PTPRH (1) hsa-mir-572 PTPRF (3), PIK3R1 (3), PRDM2 (2), PTPRN2 (2), CLSTN1 (1), FYN (1), DDC (1), PRKCZ (1) CAMTA2 (3), EPHA5 (3), PTPRF (2), MAP7 (2), CLSTN1 (2), hsa-mir-93 PIK3R1 (2), CAMTA1 (2), PDE4DIP (1), MAPT (1), GNB1 (1), PTN (1), MAP2K4 (1), ELAVL4 (1), CHD5 (1), PRKACB (1), ARHGEF7 (1) DDC (2), PDE4DIP (1), ULK2 (1), ELAVL4 (1), PLAT (1), hsa-mir-320 PLAGL1 (1), MAPT (1), CAMTA2 (1), WSB1 (1), DPYSL3 (1), NRCAM (1), CHD5 (1), HIVEP2 (1), MAP7 (1), CAMTA1 (1), MTSS1 (1), FYN (1), INPP1 (1), PRKCZ (1) 187
194 Supplemental Figure 1: (A) Standard deviation of Cq- values for genes from the 59 mrna signature and mirnas from the 25 mirna signature. Each dot represents the standard deviation of the Cq- values of a mrna or mirna measured in 262 patients. The average standard deviation for the 59 mrnas is significanly higher than that for the 25 mirnas (Mann- Whitney, p < ). (B) Expression fold change between patients that died from disease and patients that are alive (with a follow- up > 36 months) for the 59 mrnas and 25 mirnas. The average expression fold change for the mrnas is significantly higher comapred to the mirnas (Mann- Whitney, p < ). 188 A B
195 189 Discussion and future perspectives Overexpression of the MYCN oncogene is sufficient to initiate neuroblastoma tumourigenesis in vivo and is one of the major risk factors for poor outcome. Not surprisingly, the identification of genes, pathways and processes that are deregulated in MYCN amplified tumours is heavily pursued by neuroblastoma researchers worldwide. Such insights can contribute to our understanding of how neuroblastoma tumours originate and reveal new targets for selective therapy. The latter is of particular importance as direct pharmacological inhibition of MYC genes poses some major hurdles. For example, MYC genes mainly function through protein- protein and protein- DNA interactions that have proven difficult to disrupt using small molecules 1. Nanobodies might be more appropriate to disrupt MYC- protein and MYC- DNA interactions. Due to their small size, nanobodies can recognize hidden epitopes and access clefts with the same affinity as protein- protein interactions 2. However, experimental data showing that nanobodies can efficiently block MYC activity in vivo is not available yet. Moreover, inhibiting MYC might cause serious side effects as MYC is responsible for the maintenance of the stem cell compartment of adult regenerative tissues such as the gastrointestinal tract, the skin and the bone marrow. In neuroblastoma, the network of coding genes functioning downstream of MYCN is emerging and different therapeutic targets have been identified 3, 4. The major aim of this work was to investigate the role of mirnas in the MYCN transcriptional network. MiRNAs show great potential as targets for therapy. This was illustrated recently by the development of Miravirsen, the first mirna- targeted drug to enter clinical trials for the treatment of patients infected with hepatitis C virus 5. In neuroblastoma, mirnas were shown to be differentially expressed in the presence or absence of MYCN amplification 6, 7 suggesting that they form an integral part of the MYCN network. Using RT- qpcr based mirna expression profiling technology, we identified a 50 mirna signature separating MYCN amplified from MYCN single copy tumours 8, 9. Chromatine immunoprecipitation confirmed binding between MYCN and E- box sequences in the promoter of several mirnas suggesting a direct role for MYCN in the transcriptional regulation of these mirnas. Interestingly, the 50 mirna signature also separated high- risk tumours without MYCN amplification from low- risk tumours. Most probably, increased expression of MYC is driving the MYCN mirna signature in high- risk tumours without MYCN amplification. These observations are in line with a previous report by Westermann and colleagues who observed a similar expression pattern for a core set of protein- coding MYCN target genes 10. Moreover, the identification of a MYC translocated neuroblastoma tumour with a mrna, mirna and genomic profile similar to that of a MYCN amplified tumour further supports the notion that MYC and MYCN regulate a common set of coding genes and mirnas. Whether this implies that MYC and MYCN are functionally interchangeable is not entirely clear. While Malynn and colleagues nicely demonstrated the ability of Mycn to replace Myc in the process of murine growth and development 11, it remains to be investigated if Myc can replace Mycn. Closer inspection of the 50 mirna signature revealed a striking imbalance between the number of up- and downregulated mirnas. In total, 34 mirnas were found to be downregulated while only 16 were upregulated suggesting that MYCN amplification is associated with widespread mirna repression rather than activation. Chang and colleagues observed a similar pattern when searching for MYC regulated mirnas in lymphoma cells 12 and global mirna downregulation has been observed in several human cancers 13. This global loss of mirna expression was shown to be functionally relevant for oncogenesis, as impairment of mirna maturation enhanced cellular transformation and tumourigenesis 14. Different molecular mechanisms underlying loss of mirna expression have been identified and can be attributed to defects in the mirna processing machinery. These include monoallelic loss of DICER1 15, 16 and mutations in TARBP2 17, both affecting the processing of mature mirnas, and mutations in XPO5, affecting the export of precursor mirnas to the cytoplasm. For the majority of mirnas down regulated by MYC, Chang and
196 190 colleagues could demonstrate MYC binding to conserved E- box elements upstream of these mirnas suggesting that the observed down regulation is a direct effect of MYC 12. However, this does not exclude the possibility that MYC genes might regulate the expression or activity of one or more genes from the mirna processing machinery. In high- risk neuroblastoma tumours, expression of DROSHA and DICER1 was shown to be down regulated, consistent with the overall repression of mirna levels in these tumours 18. In addition, LIN28B, a RNA binding protein that blocks the processing of let- 7 mirnas, is activated by both MYC and MYCN 19, 20. The extent to which MYC- family proteins are involved in negative regulation of mirna processing is currently unclear and needs further experimental study. Despite the fact that we identified more than twice as much down regulated mirnas than up regulated mirnas, there was no significant overlap between the predicted targets of the down regulated mirnas and genes activated by MYCN. In contrast, many targets of the up regulated mirnas were known to be down regulated by MYCN suggesting that mirna activation is a hitherto unknown mechanism by which MYC- family proteins repress gene expression. These mirnas could work in concert, as several target genes are shared between them. Furthermore, our data suggest that this mechanism of MYC target gene repression might be more widespread than the one involving the interaction with MIZ1. Thus far, only a limited number of genes were shown to be repressed through the MIZ1 mechanism. In contrast, the activated mirnas are predicted to regulate an extensive repertoire of target genes. Although many of the MYCN activated mirnas have unknown functions, some are known oncomirs targeting key tumour suppressor pathways such as mir and the mir cluster 22. Ma and colleagues demonstrated that mir- 9 targets the E- cadherin/β- catenin axis in breast cancer cells resulting in increased cell motility and tumour angiogenesis. Whether mir- 9 has a similar function in neuroblastoma remains to be determined. The status of mir- 9 as an oncomir in neuroblastoma is somewhat controversial as REST and CREB induce mir- 9 expression during neuronal differentiation 23. The members of the mir cluster belong to the most frequently overexpressed mirnas in cancer. In neuroblastoma, mir overexpression results in repression of CDKN1A, a negative regulator of the cell cycle, BIM, a pro- apoptotic gene, and ESR1, a positive regulator of neuronal differentiation 22, 24. In doing so, mir promotes cell proliferation, sustains cell survival and inhibits differentiation. The pleiotropic functions carried out by mir are probably explained by the fact that each of the six mirnas within the cluster has the potential to regulate hundreds of genes. In a systematic screen for mir functions in neuroblastoma, we identified multiple processes that were directly affected by mir activation including proliferation, differentiation and adhesion. Detailed analysis of the identified mir targets revealed the possibility that individual mirnas from the cluster cooperate to repress target expression. Cooperation might also occur between co- expressed mirnas that don t belong to a genomic cluster, e.g. all mirnas activated by MYCN, and could be a general phenomenon of mirna target regulation. Not only do mir mirnas cooperate to regulate the same target, they also cooperate to regulate different targets within the same pathway. We found evidence that at least 5 genes from the TGFβ- signaling pathway are directly regulated by different mirnas from the mir cluster. Such coordinated action might be necessary to compensate for the relatively small effect of a mirna on the expression of its target. This information could be used to improve mirna target prediction algorithms. At least, genes with binding sites for multiple co- expressed or co- regulated mirnas might be prioritized for further evaluation. These findings also have important implications for the way experiments are designed to study mirna functions. In a situation where multiple mirnas cooperate to repress a gene or pathway, overexpression of one mirna might not produce a measurable effect leading to the false conclusion that this mirna is not involved in the regulation of that particular gene or pathway. In case of mir , it is therefore likely that the sum of the effects of the individual mirnas is less than the effect of the entire cluster. Alternatively, the ability of the mir cluster to simultaneously target the components of the signaling cascade, as well as the downstream effectors through multiple mirnas, allows for tight control of the TGF- b- transcriptional program. Moreover, it offers the cells enormous
197 191 flexibility and plasticity for regulation of different subsets of TGFβ target genes. Nevertheless, functional dissection of the individual mirnas from the mir cluster is necessary to understand the complex interplay between them. In an attempt to evaluate the in vivo tumourigenic potential of the individual mirnas from the cluster, mir- 19 was identified as the key oncogenic component of mir in the Eµ- myc mouse B- cell lymphoma model 25, 26. MiR- 19 is both necessary and sufficient to promote MYC- induced lymphomagenesis through repression of apoptosis. In addition, mir- 19 is also sufficient to promote leukaemogenesis in NOTCH1- induced T- cell acute lymphoblastic leukaemia in vivo 27. These findings suggest that mir directed regulation of some processes does not depend on additive effects of multiple mirnas. Whether mir- 19 is the key oncogenic component of the mir cluster in neuroblastoma or other solid tumours remains to be determined. For regulation of the TGFβ- pathway, we and others have shown that not mir- 19 but mir- 17, mir- 18a and mir- 20a are essential 28, 29. The complexity of interplay and coordination between the individual members is further underscored by a recent study from Shan and colleagues showing that overexpression of mir- 17 in transgenic mice decreases proliferation 30. Possibly, mir mirnas have the ability to both positively and negatively regulate the same cellular process in order to achieve homeostasis in vivo. The pleiotropic functions of mir in cancer biology lead to belief that mir is an attractive target for therapy. Antisense oligonucleotides against mir mirnas have proven to be effective in different cancers, including neuroblastoma 22, however the possible side effects of such treatment are not documented. MiR mirnas are essential during development as loss of function studies in mice resulted in smaller embryos and immediate postnatal death of all animals 31. While mir expression is high in embryonal tissues, it decreases when mice are fully grown suggesting that the importance of maintaining mir expression in mature tissues is minor 32. To elucidate possible side effects of mir targeting, neuroblastoma tumour bearing mice should be treated with antisense oligonucleotides against all mir mirnas followed by an accurate measurement of the apoptotic cell fraction in different normal tissues with high and low proliferative indices. Based on results obtained from in vitro and in vivo studies in different cancer types we hypothesize that mir inhibition will result in a decreased proliferation and increased apoptosis of the tumour cells, accompanied by increased differentiation and decreased vascularization. Not only are mirnas interesting candidates for therapeutic intervention in cancer, they also serve as markers to predict patient prognosis and outcome. MiRNA expression signatures were shown to outperform mrna expression signatures when classifying according to tissue of origin however it is not clear whether the same is true for prognostic classification. To this end we built, tested and validated a prognostic 25 mirna classifier for neuroblastoma and compared its performance to that of a previously established 59 mrna classifier 33 using 2 independent neuroblastoma patient cohorts of 179 and 304 samples respectively. While the mirna signature significantly distinguished patients with respect to progression free and overall survival, it did not outperform the mrna expression signature in a multivariate analysis. To exclude the possibility that a suboptimal mirna classifier was built, we also evaluated two published neuroblastoma mirna classifiers 34, 35 and could show that both were inferior to the 59 mrna classifier. Surprisingly, a comparison of mirna and mrna expression fold changes between deceased patients and patients that were alive revealed that mrna genes had significantly higher fold changes compared to mirna genes. This might, at least in part, explain why mrna classification outperforms mirna classification, as small expression changes are difficult to measure with high accuracy. Similar studies in other tumour types will need to be performed in order to confirm that this is a general phenomenon. Integrating mirna and mrna signatures could further increase overall classification performance. In one approach, mirna and mrna classifiers are used in a successive classification where prognostic favorable and unfavorable patients groups according to the mrna classifier are subjected to a second classification using the mirna classifier or vice versa. Alternatively, mirna and mrna classifiers can be combined in a single signature consisting of a selection of markers taken from both classifiers.
198 Successive classification using the 25 mirna and 59 mrna signatures in the neuroblastoma patient cohort did not result in a better classification accuracy. Interestingly, we did succeed in combining both signatures into a signature consisting only of 7 markers (both mrna and mirna). While this signature did not outperform the 59 mrna signature it might be of clinical relevance because of the limited number of markers that are profiled. Despite the fact that the 25 mirna classifier performed worse compared to the 59 mrna signature, it still could be of potential use in the clinic. First, mirnas are believed to be more resistant to RNA degradation, possibly because of their small size 36. This opens the possibility for the use of formalin- fixed paraffin embedded (FFPE) tissues. In the current clinical workflow only minimal amounts of fresh frozen tissue is procured while FFPE samples are widely available, stable at room temperature, and easily storable. Several studies have indeed shown that mirnas are readily detectable in FFPE 37-40, opening perspectives for a clinical application of the proposed 25 mirna signature. Second, mirnas can also be detected in different body fluids where they show a remarkable stability. MiRNA levels in serum or plasma were shown to correlate to disease, tumour stage and patient survival 41, 42. The origin and function of these circulating mirnas is still obscure. One hypothesis states that they originate from tumour cells as a result of tumour cell death or lysis 43 while another suggests they might be secreted by tumour cells and normal cells as a means of cell- cell communication 44, 45. It is unlikely that the same 25 mirna signature that was established from tumour RNA will also be prognostic in patient serum. Therefore, a new and unbiased search for prognostic and diagnostic mirnas in serum from neuroblastoma patients should be performed. 192 One of the major hurdles when studying mirnas is prioritization and functional annotation. Typically, mirna expression profiling of a cellular system exposed to any chemical or genetic perturbation generates dozens of differentially expressed mirnas. In most cases, functional evaluation of all candidate mirnas is not feasible and an upfront prioritization of the most promising candidates is required. While prioritization can be based on the significance or fold change of the differential expression, large fold changes are not a prerequisite for biological relevance. Alternatively, mirna target predictions can be used to select those mirnas that are more likely to be involved in the pathway(s) or process(es) perturbed in the experimental model. Several computer- based methods have been developed to assign mirnas to biological pathways These methods calculate the enrichment of predicted mirna targets in gene lists representing a pathway, process or function but ignore the possibility that mirna functions can be tissue specific. In addition, the enrichment strategy assumes that multiple genes within the gene list need to be under the control of the mirna while several studies have shown that mirnas can regulate cellular functions by controlling just one gene, i.e. a transcription factor or key signaling molecule 21, 49. To better address these issues, we devised a strategy to predict mirna functions based on the integration of multiple levels of information such as matching mirna and mrna expression, mirna target prediction and mechanistic models of gene expression regulation. Only through such an integrative genomics approach is it feasible to predict mirna functions that are tissue or disease specific and that can be explained by different interactions between the mirna and the components of the pathway. The so- called mirna body map approach was shown to predict validated mirna functions with high sensitivity and modest specificity and can be used to generate hypotheses on mirna functions in any given dataset for which matching mrna and mirna expression data are available. To maximize specificity and sensitivity, only those datasets for which mirna expression was generated using RT- qpcr technology were included. However, the limited availability of such datasets suggests that mirna expression data, generated using different technologies, should be included in order to broaden the scope of this study. The availability of additional datasets representing different tissue and disease types will also allow to study tissue- specific mirna functions in more detail and with higher confidence. An initial comparison of the predicted mirna functions already revealed a substantial difference between the different datasets suggesting that the tissue specificity of mirna functions might be more widespread than initially anticipated.
199 References Soucek, L. et al. Modelling Myc inhibition as a cancer therapy. Nature 455, (2008). 2. Revets, H., De Baetselier, P. & Muyldermans, S. Nanobodies as novel agents for cancer therapy. Expert Opin Biol Ther 5, (2005). 3. Slack, A. et al. The p53 regulatory gene MDM2 is a direct transcriptional target of MYCN in neuroblastoma. Proc Natl Acad Sci U S A 102, (2005). 4. Hogarty, M.D. et al. ODC1 is a critical determinant of MYCN oncogenesis and a therapeutic target in neuroblastoma. Cancer Res 68, (2008). 5. Lanford, R.E. et al. Therapeutic silencing of microrna- 122 in primates with chronic hepatitis C virus infection. Science 327, (2010). 6. Schulte, J.H. et al. MYCN regulates oncogenic MicroRNAs in neuroblastoma. Int J Cancer 122, (2008). 7. Chen, Y. & Stallings, R.L. Differential patterns of microrna expression in neuroblastoma are correlated with prognosis, differentiation, and apoptosis. Cancer Res 67, (2007). 8. Mestdagh, P. et al. High- throughput stem- loop RT- qpcr mirna expression profiling using minute amounts of input RNA. Nucleic Acids Res 36, e143 (2008). 9. Mestdagh, P. et al. MYCN/c- MYC- induced micrornas repress coding gene networks associated with poor outcome in MYCN/c- MYC- activated tumours. Oncogene 29, (2010). 10. Westermann, F. et al. Distinct transcriptional MYCN/c- MYC activities are associated with spontaneous regression or malignant progression in neuroblastomas. Genome Biol 9, R150 (2008). 11. Malynn, B.A. et al. N- myc can functionally replace c- myc in murine development, cellular growth, and differentiation. Genes Dev 14, (2000). 12. Chang, T.C. et al. Widespread microrna repression by Myc contributes to tumourigenesis. Nat Genet 40, (2008). 13. Lu, J. et al. MicroRNA expression profiles classify human cancers. Nature 435, (2005). 14. Kumar, M.S., Lu, J., Mercer, K.L., Golub, T.R. & Jacks, T. Impaired microrna processing enhances cellular transformation and tumourigenesis. Nat Genet 39, (2007). 15. Kumar, M.S. et al. Dicer1 functions as a haploinsufficient tumour suppressor. Genes Dev 23, (2009). 16. Lambertz, I. et al. Monoallelic but not biallelic loss of Dicer1 promotes tumourigenesis in vivo. Cell Death Differ 17, (2010). 17. Melo, S.A. et al. A TARBP2 mutation in human cancer impairs microrna processing and DICER1 function. Nat Genet 41, (2009). 18. Lin, R.J. et al. microrna signature and expression of Dicer and Drosha can predict prognosis and delineate risk groups in neuroblastoma. Cancer Res 70, (2010). 19. Dangi- Garimella, S. et al. Raf kinase inhibitory protein suppresses a metastasis signalling cascade involving LIN28 and let- 7. EMBO J 28, (2009). 20. Cotterman, R. & Knoepfler, P.S. N- Myc regulates expression of pluripotency genes in neuroblastoma including lif, klf2, klf4, and lin28b. PLoS One 4, e5799 (2009). 21. Ma, L. et al. mir- 9, a MYC/MYCN- activated microrna, regulates E- cadherin and cancer metastasis. Nat Cell Biol 12, (2010). 22. Fontana, L. et al. Antagomir- 17-5p abolishes the growth of therapy- resistant neuroblastoma through p21 and BIM. PLoS One 3, e2236 (2008). 23. Laneve, P. et al. A minicircuitry involving REST and CREB controls mir- 9-2 expression during human neuronal differentiation. Nucleic Acids Res 38, (2010). 24. Loven, J. et al. MYCN- regulated micrornas repress estrogen receptor- alpha (ESR1) expression and neuronal differentiation in human neuroblastoma. Proc Natl Acad Sci U S A 107, (2010). 25. Olive, V. et al. mir- 19 is a key oncogenic component of mir Genes Dev 23, (2009). 26. Mu, P. et al. Genetic dissection of the mir- 17~92 cluster of micrornas in Myc- induced B- cell lymphomas. Genes Dev 23, (2009). 27. Mavrakis, K.J. et al. Genome- wide RNA- mediated interference screen identifies mir- 19 targets in Notch- induced T- cell acute lymphoblastic leukaemia. Nat Cell Biol 12, (2010).
200 28. Dews, M. et al. The myc- mir- 17~92 axis blunts TGF{beta} signaling and production of multiple TGF{beta}- dependent antiangiogenic factors. Cancer Res 70, (2010). 29. Mestdagh, P. et al. The mir MicroRNA Cluster Regulates Multiple Components of the TGF- beta Pathway in Neuroblastoma. Mol Cell 40, (2010). 30. Shan, S.W. et al. MicroRNA MiR- 17 retards tissue growth and represses fibronectin expression. Nat Cell Biol 11, (2009). 31. Ventura, A. et al. Targeted deletion reveals essential and overlapping functions of the mir- 17 through 92 family of mirna clusters. Cell 132, (2008). 32. Lu, Y., Thomson, J.M., Wong, H.Y., Hammond, S.M. & Hogan, B.L. Transgenic over- expression of the microrna mir cluster promotes proliferation and inhibits differentiation of lung epithelial progenitor cells. Dev Biol 310, (2007). 33. Vermeulen, J. et al. Predicting outcomes for children with neuroblastoma using a multigene- expression signature: a retrospective SIOPEN/COG/GPOH study. Lancet Oncol 10, (2009). 34. Schulte, J.H. et al. Accurate prediction of neuroblastoma outcome based on mirna expression profiles. Int J Cancer 127, (2010). 35. Bray, I. et al. Widespread dysregulation of MiRNAs by MYCN amplification and chromosomal imbalances in neuroblastoma: association of mirna expression with survival. PLoS One 4, e7850 (2009). 36. Jung, M. et al. Robust microrna stability in degraded RNA preparations from human tissue and cell samples. Clin Chem 56, (2010). 37. Benjamin, H. et al. A diagnostic assay based on microrna expression accurately identifies malignant pleural mesothelioma. J Mol Diagn 12, (2010). 38. Fridman, E. et al. Accurate molecular classification of renal tumours using microrna expression. J Mol Diagn 12, (2010). 39. Koelz, M. et al. Down- regulation of mir- 221 and mir- 222 correlates with pronounced Kit expression in gastrointestinal stromal tumours. Int J Oncol (2010). 40. Voortman, J. et al. MicroRNA expression and clinical outcomes in patients treated with adjuvant chemotherapy after complete resection of non- small cell lung carcinoma. Cancer Res 70, (2010). 41. Liu, R. et al. A five- microrna signature identified from genome- wide serum microrna expression profiling serves as a fingerprint for gastric cancer diagnosis. Eur J Cancer (2010). 42. Moltzahn, F. et al. Microfluidic based multiplex qrt- PCR identifies diagnostic and prognostic microrna signatures in sera of prostate cancer patients. Cancer Res (2010). 43. Mitchell, P.S. et al. Circulating micrornas as stable blood- based markers for cancer detection. Proc Natl Acad Sci U S A 105, (2008). 44. Iguchi, H., Kosaka, N. & Ochiya, T. Secretory micrornas as a versatile communication tool. Commun Integr Biol 3, (2010). 45. Zhao, H. et al. A pilot study of circulating mirnas as potential biomarkers of early stage breast cancer. PLoS One 5, e13735 (2010). 46. Nam, S., Kim, B., Shin, S. & Lee, S. mirgator: an integrated system for functional annotation of micrornas. Nucleic Acids Res 36, D (2008). 47. Tsang, J.S., Ebert, M.S. & van Oudenaarden, A. Genome- wide dissection of microrna functions and cotargeting networks using gene set signatures. Mol Cell 38, (2010). 48. Ulitsky, I., Laurent, L.C. & Shamir, R. Towards computational prediction of microrna function and activity. Nucleic Acids Res 38, e160 (2010). 49. Yamakuchi, M. et al. P53- induced microrna- 107 inhibits HIF- 1 and tumour angiogenesis. Proc Natl Acad Sci U S A 107, (2010). 194
201 195 Summary Neuroblastoma is a pediatric malignancy that originates from precursor cells of the sympathetic nervous system. Despite intensive multimodal treatment schemes, less than half of the high- risk patients survive the disease. Even when successful, current treatment often causes short- and long- term side effects that can have a substantial impact in later life. Detailed insights in the molecular defects driving neuroblastoma development can lead to the discovery of new targeted therapies with higher efficiency and less toxicity. Key examples in this respect are the discovery of a small molecule, imatinib mesylate (Glivec ), targeting the constitutively activated tyrosine protein kinase from the BCR- ABL1 gene fusion in chronic myelogenous leukaemia and trastuzumab (Herceptin ), an antibody against the ERBB2 growth receptor that is overexpressed in breast cancer. The first aim of this work was to evaluate mirna expression patterns and their function in neuroblastoma tumourigenesis. One of the major genetic defects driving neuroblastoma tumourigenesis is amplification and overexpression of the MYCN oncogene. To elucidate the role of mirnas in the MYCN transcriptional network, a high- throughput and sensitive mirna expression profiling platform was optimized and subsequently applied to measure mirna expression levels in tumour samples from neuroblastoma patients with differential MYCN amplification status. Based on these data, a 50 mirna signature was established that significantly distinguishes between tumours with and without MYCN amplification. In depth analysis of the targets from these mirnas revealed that MYCN activated mirnas contribute to widespread mrna repression in tumours with MYCN amplification. These findings suggest that mirna activation is a new mechanism by which MYCN represses gene expression. Among the mirnas activated by MYCN we found several established oncomirs such as mir- 9 and all members from the mir cluster. The mir cluster is one of the most frequently activated oncomirs in cancer and is known to repress cell cycle inhibition and apoptosis. To elucidate the biological implications of mir activation in neuroblastoma, a cell line with inducible mir expression was created and used to measure the impact of mir on protein output. In this way, several hallmarks of cancer biology were found to be deregulated including proliferation, differentiation and cell adhesion. Furthermore, these data revealed that mir regulates different components from the TGFβ- signaling cascade, a known tumour suppressor pathway in cancer. The pleiotropic functions of the mir cluster suggest that therapeutic strategies to target mir activity could be of potential relevance for the treatment of neuroblastoma tumours. Experimental approaches to define the specific functions of a mirna are technically challenging and time consuming. To maximize the success of such studies, an upfront prioritization of the most promising candidate mirnas is indispensable. To this purpose, a strategy was developed that combines matching mrna and mirna expression data with mirna target prediction and mechanistic models of gene expression regulation to predict putative mirna functions. This approach takes the tissue- specific nature of mirna functions into account and results in predictions with high sensitivity. This was demonstrated by comparing functional predictions for several mirnas to experimentally validated mirna functions in different datasets. Predictions for four different datasets were stored in a dedicated database that can be queried through a publically available webtool called the mirna body map. Unraveling mirna functions is essential in order to understand the biological consequences of deregulated mirna expression but becomes irrelevant when the aim is to use these deregulated mirna expression patterns for prognostic classification. To establish a prognostic mirna signature for neuroblastoma, genome- wide mirna expression profiles were generated for a large cohort of tumour samples. A 25 mirna signature was subsequently trained, tested, and validated and was found to be significantly associated with
202 overall and progression free survival in two independent patient cohorts. Patients with increased risk for a shorter progression free and overall survival could also be identified in the cohort of high- risk patients for which no clinical or genetic markers are available today. 196 The second aim of this work was to evaluate the expression and function of another class of non- coding RNAs termed transcribed ultra- conserved regions (T- UCRs). Although nothing is known about T- UCR function in cancer biology, T- UCRs are differentially expressed between cancer and normal tissue. Using RT- qpcr technology, expression of all 481 T- UCRs was measured in a cohort of neuroblastoma tumours and several T- UCRs were found to correlate to MYCN amplification status and patient survival. An integrative genomics approach, similar to that of the mirna body map, was subsequently applied to explore putative T- UCR functions. Four major functional T- UCR clusters were identified: a proliferation cluster, a differentiation cluster, a cluster involved in immune response and a DNA damage response cluster. Several predicted T- UCR functions were experimentally validated in different cellular model systems, supporting the in silico annotation strategy. While various interesting associations between T- UCRs and cancer- related processes were uncovered, additional studies are needed to further elucidate T- UCR function and to unravel the transcriptional programs mediating their expression.
203 197 Samenvatting Neuroblastoom is een pediatrische maligniteit die ontstaat uit precursorcellen van het sympathische zenuwstelsel. Ondanks een intensieve multimodale behandeling overleven meer dan de helft van de hoog- risico patiënten de ziekte niet. De huidige behandelingsstrategie veroorzaakt ook heel wat nevenwerkingen, zowel op korte als op lange termijn. Deze nevenwerkingen kunnen een grote impact hebben op het verdere leven van patiënten die de ziekte overleven. Verdere inzichten in de moleculaire defecten die de ontwikkeling van neuroblastoom aansturen kunnen leiden tot de ontwikkeling van nieuwe gerichte therapieën met een hogere efficiëntie en een lagere toxiciteit. Voorbeelden hiervan zijn de ontdekking van ene kleine molecule, imatinib mesylaat (Glivec ) die het constitutief actieve tyrosine kinase eiwit van de BCR- ABL1 genfusie uitschakelt in chronische myelogene leukemie en tretuzumab (Herceptin ), een antilichaam gericht tegen de ERBB2 groeireceptor die tot overexpressie komt in borstkanker. De eerste doelstelling van dit project was een evaluatie van mirna expressiepatronen en hun functie in neuroblastoom tumourigenese. Een van de belangrijkste genetische defecten die de tumourigenese van neuroblastoom aanstuurt is amplificatie en overexpressie van het MYCN oncogen. Om de rol van mirnas in het MYCN transcriptionele netwerk na te gaan werd gestart met de optimalisatie van een hoge- doorvoer platform voor mirna expressieprofilering. Dit platform werd vervolgens aangewend voor het meten van mirna expressieniveaus in tumourstalen van neuroblastoompatiënten met een differentiële MYCN amplificatiestatus. Op basis van deze data werd een signatuur opgesteld, bestaande uit 50 mirnas, die toelaat om tumouren met en zonder MYCN amplificatie van elkaar te onderscheiden. Een grondige analyse van de doelwitgenen van deze mirnas toonde aan dat mirnas die geactiveerd worden door MYCN, bijdragen tot de repressie van mrnas in de MYCN geamplificeerde tumouren. Deze bevindingen wijzen erop dat mirna activatie kan dienen als een nieuw mechanisme waarmee MYCN de expressie van genen negatief reguleert. Onder de mirnas die door MYCN geactiveerd worden bevonden zich verschillende gekende oncomirs zoals mir- 9 en de verschillende leden van de mir cluster. De mir cluster is een van de meest frequent geactiveerde oncomirs in kanker. Activatie van mir leidt onder meer tot de repressie van celcyclus inhibitie en apoptose. Om de biologische implicaties van mir activatie in neuroblastoom na te gaan werd een cellijn gecreëerd met induceerbare mir expressie. Deze cellijn werd vervolgens gebruikt om de invloed van mir op eiwitexpressie in de cel te meten. Deze experimenten toonden aan dat verschillende processen die belangrijk zijn in de ontwikkeling van kanker, zoals proliferatie, differentiatie en adhesie, ontregeld werden als gevolg van mir activatie. Bovendien werd aangetoond dat mir verschillende componenten reguleert van de TGFβ- signaalweg, een gekende tumoursuppressor reactieweg in kanker. De pleiotrope functies van de mir cluster wijzen erop dat een therapeutische strategie gericht op de uitschakeling van mir activiteit in kankercellen van potentieel belang kan zijn voor de behandeling van neuroblastoomtumouren. Experimentele benaderingen om de functionaliteit van een mirna te bepalen vergen een grote technische expertise en zijn vaak tijdrovend. Om het succes van dergelijke experimenten te maximaliseren is een voorafgaande selectie van de sterkste kandidaat mirnas noodzakelijk. Om dit mogelijk te maken werd een strategie ontwikkeld die gepaarde mrna en mirna expressiedata combineert met mirna doelwitgenpredictie en mechanistische modellen van genexpressieregulatie. Deze strategie heeft als doel de functionaliteit van individuele mirnas te voorspellen en houdt rekening met weefselspecificiteit. Op basis van een vergelijking tussen voorspelde en gevalideerde functies voor verschillende mirnas kon aangetoond worden dat de gevolgde strategie leidt tot functionele predicties met een hoge sensitiviteit. De
204 voorspelde functies voor vier verschillende datasets werden opgenomen in een databank die doorzocht kan worden via een publiek beschikbare webtool genaamd de mirna body map. De ontrafeling van mirna functies is essentieel om inzichten te verwerven in de biologische gevolgen van gedereguleerde mirna expressie. Dit is echter niet het geval waneer de gedereguleerde mirna expressiepatronen aangewend worden voor prognostische classificatie. Om een prognostische mirna signatuur voor neuroblastoom op te stellen werden genoomwijde mirna expressieprofielen bepaald voor een grote cohorte van tumouren. Een mirna signatuur bestaande uit 25 mirnas werd vervolgens getraind, getest en gevalideerd. Deze signatuur was significant geassocieerd met de algemene en progressie vrije overleving in twee onafhankelijke patiëntenpopulaties. Patiënten met een verhoogd risico op een gedaalde algemene en progressie vrije overleving werden ook geïdentificeerd in de groep van hoog- risico neuroblastoompatiënten, een patiëntengroep waarvoor tot op heden geen klinische of genetische merkers beschikbaar zijn. 198 De tweede doestelling van dit project was de evaluatie van een andere groep niet- coderende RNAs genaamd transcribed ultraconserved regions of T- UCRs. Hoewel er weinig tot niets gekend is over de functie van T- UCRs in de biologie van kanker werd reeds aangetoond dat T- UCRs differentieel tot expressie komen tussen tumouren en normale weefsels. Met behulp van RT- qpcr werd de expressie van 481 T- UCRs gemeten in een cohorte van neuroblastoomtumouren. Analyse van de expressiedata toonde aan dat verschillende T- UCRs correleren met MYCN amplificatie en overleving. Om mogelijke T- UCR functies in kaart te brengen werd gebruik gemaakt van een geïntegreerde strategie, gelijkaardig aan deze gevolgd voor de mirna body map. Er werden vier functionele T- UCR clusters geïdentificeerd: een proliferatiecluster, een differentiatiecluster, een cluster betrokken in immuunrespons en een cluster betrokken in respons op DNA schade. Verschillende voorspelde T- UCR functies werden experimenteel gevalideerd met behulp van cellulaire modelsystemen. Ondanks het feit dat verschillende interessante associaties tussen T- UCRs en kankergerelateerde processen werden blootgelegd zijn bijkomende studies nodig om meer inzichten te verwerven in T- UCR functies en de verschillende processen die instaan voor de transcriptionele regulatie van T- UCR expressie.
205 199 Curriculum Vitae Pieter Mestdagh Born April 9th, 1982 in Brugge, Belgium Professional address Home address Ghent University Hospital Zwarteleertouwersstraat 37 Center for Medical Genetics 8000 Brugge Medical Research Building Belgium De Pintelaan Gent Belgium +32 (0) (phone) +32 (0) (phone) +32 (0) (fax) Education Sciences- Mathematics (8h), Sint- Leocollege, Brugge, Belgium Industrial Engineer in Biochemistry, KAHO Sint- Lieven, Gent, Belgium Bio- engineer in Cell and Gene Biotechnology, Ghent University, Ghent, Belgium Courses Short Course on Experimental Models of Human Cancer, August , The Jackson Lab, Bar Harbor, Maine, USA Publications Mestdagh P, Feys T, Bernard N, Guenther S, Chen C, Speleman F, Vandesompele J. High- throughput stem- loop RT- qpcr mirna expression profiling using minute amounts of input RNA. Nucleic Acids Res Dec;36(21):e143. Mestdagh P, Van Vlierberghe P, De Weer A, Muth D, Westermann F, Speleman F, Vandesompele J. A novel and universal method for microrna RT- qpcr data normalization. Genome Biol. 2009;10(6):R64. Van Roy N, De Preter K, Hoebeeck J, Van Maerken T, Pattyn F, Mestdagh P, Vermeulen J, Vandesompele J, Speleman F. The emerging molecular pathogenesis of neuroblastoma: implications for improved risk assessment and targeted therapy. Genome Med Jul 27;1(7):74 Vermeulen J, Pattyn F, De Preter K, Vercruysse L, Derveaux S, Mestdagh P, Lefever S, Hellemans J, Speleman F, Vandesompele J. External oligonucleotide standards enable cross laboratory comparison and exchange of real- time quantitative PCR data. Nucleic Acids Res Nov;37(21):e138.
206 200 Van Vlierberghe P, De Weer A, Mestdagh P, Feys T, De Preter K, De Paepe P, Lambein K, Vandesompele J, Van Roy N, Verhasselt B, Poppe B, Speleman F. Comparison of mirna profiles of microdissected Hodgkin/Reed- Sternberg cells and Hodgkin cell lines versus CD77+ B- cells reveals a distinct subset of differentially expressed mirnas. Br J Haematol Dec;147(5): Bray I, Bryan K, Prenter S, Buckley PG, Foley NH, Murphy DM, Alcock L, Mestdagh P, Vandesompele J, Speleman F, London WB, McGrady PW, Higgins DG, O'Meara A, O'Sullivan M, Stallings RL. Widespread dysregulation of MiRNAs by MYCN amplification and chromosomal imbalances in neuroblastoma: association of mirna expression with survival. PLoS One Nov 16;4(11):e7850. Mestdagh P, Fredlund E, Pattyn F, Schulte JH, Muth D, Vermeulen J, Kumps C, Schlierf S, De Preter K, Van Roy N, Noguera R, Laureys G, Schramm A, Eggert A, Westermann F, Speleman F, Vandesompele J. MYCN/c- MYC- induced micrornas repress coding gene networks associated with poor outcome in MYCN/c- MYC- activated tumours. Oncogene Mar 4;29(9): Lambertz I, Nittner D, Mestdagh P, Denecker G, Vandesompele J, Dyer MA, Marine JC. Monoallelic but not biallelic loss of Dicer1 promotes tumourigenesis in vivo. Cell Death Differ Apr;17(4): Ma L, Young J, Prabhala H, Pan E, Mestdagh P, Muth D, Teruya- Feldstein J, Reinhardt F, Onder TT, Valastyan S, Westermann F, Speleman F, Vandesompele J, Weinberg RA. mir- 9, a MYC/MYCN- activated microrna, regulates E- cadherin and cancer metastasis. Nat Cell Biol Mar;12(3): Mestdagh P*, Fredlund E*, Pattyn F, Rihani A, Van Maerken T, Vermeulen J, Kumps C, Menten B, De Preter K, Schramm A, Schulte J, Noguera R, Schleiermacher G, Janoueix- Lerosey I, Laureys G, Powel R, Nittner D, Marine JC, Ringnér M, Speleman F, Vandesompele J. An integrative genomics screen uncovers ncrna T- UCR functions in neuroblastoma tumours. Oncogene Jun 17;29(24): *Equally contributing authors Buckley PG, Alcock L, Bryan K, Bray I, Schulte JH, Schramm A, Eggert A, Mestdagh P, De Preter K, Vandesompele J, Speleman F, Stallings RL. Chromosomal and microrna expression patterns reveal biologically distinct subgroups of 11q- neuroblastoma. Clin Cancer Res Jun 1;16(11): Chang KH, Mestdagh P, Vandesompele J, Kerin MJ, Miller N. MicroRNA expression profiling to identify and validate reference genes for relative quantification in colorectal cancer. BMC Cancer Apr 29;10:173. Schulte JH, Marschall T, Martin M, Rosenstiel P, Mestdagh P, Schlierf S, Thor T, Vandesompele J, Eggert A, Schreiber S, Rahmann S, Schramm A. Deep sequencing reveals differential expression of micrornas in favorable versus unfavorable neuroblastoma. Nucleic Acids Res Sep 1;38(17): Schulte JH, Schowe B, Mestdagh P, Kaderali L, Kalaghatgi P, Schlierf S, Vermeulen J, Brockmeyer B, Pajtler K, Thor T, de Preter K, Speleman F, Morik K, Eggert A, Vandesompele J, Schramm A. Accurate prediction of neuroblastoma outcome based on mirna expression profiles. Int J Cancer Nov 15;127(10):
207 201 Vermeulen J, De Preter K, Mestdagh P, Laureys G, Speleman F, Vandesompele J. Predicting outcomes for children with neuroblastoma. Discov Med Jul;10(50): Van Pottelberge GR*, Mestdagh P*, Bracke KR, Thas O, van Durme YM, Joos GF, Vandesompele J, Brusselle GG. MicroRNA Expression in Induced Sputum of Smokers and Patients with Chronic Obstructive Pulmonary Disease. Am J Respir Crit Care Med Oct 29. *Equally contributing authors Mestdagh P*, Boström AK*, Impens F*, Fredlund E, Van Peer G, De Antonellis P, von Stedingk K, Ghesquière B, Schulte S, Dews M, Thomas- Tikhonenko A, Schulte JH, Zollo M, Schramm A, Gevaert K, Axelson H, Speleman F, Vandesompele J. The mir MicroRNA Cluster Regulates Multiple Components of the TGF- β Pathway in Neuroblastoma. Mol Cell Dec 10;40(5): *Equally contributing authors Mestdagh P*, De Preter K*, Vermeulen J*, Naranjo A, Bray I, Castel V, Chen C, Eggert A, Hogarty MD, London WB, Noguera R, Piqueras M, Bryan K, Schowe B, van Sluis P, Molenaar JJ, Schramm, Schulte JH, Stallings RL, Versteeg R, Laureys G, Van Roy N, Speleman F, Vandesompele J. Outcome prediction of children with neuroblastoma using mirna and mrna gene expression signatures. Submitted. *Equally contributing authors Mestdagh P*, Lefever S*, Pattyn F, Ridzon D, Fredlund E, Fieuw A, Ongenaert M, Vermeulen J, De Paepe A, Wong L, Speleman F, Chen C, Vandesompele J. The microrna body map: dissecting microrna function through integrative genomics. Submitted. *Equally contributing authors Oral presentations High- throughput stem- loop RT- PCR mirna expression profiling using minute amounts of input RNA. Applied Biosystems user meeting, October 17th 2007, Nieuwerkerk aan den Ijzel, Holland. High- throughput stem- loop RT- PCR mirna expression profiling using minute amounts of input RNA. Applied Biosystems user meeting, October 18th 2007, Gosselies, Belgium. Expression profiling of a new class of non- coding RNAs in human neuroblastoma. Advances in Neuroblastoma Research, Mai , Chiba, Japan. MYCN regulates oncogenic and tumour suppressor mirna networks in neuroblastoma. Advances in Neuroblastoma Research, Mai , Chiba, Japan. High- throughput stem- loop RT- PCR mirna expression profiling using minute amounts of input RNA. EACR ABI satellite symposium, July , Lyon, France High- throughput stem- loop RT- PCR mirna expression profiling using minute amounts of input RNA. Benelux qpcr symposium, October 6th 2008, Ghent, Belgium.
208 202 High- throughput stem- loop RT- PCR mirna expression profiling using minute amounts of input RNA reveals new insights in neuroblastoma pathogenesis. Belgian Society for Human Genetics, February 13th 2009, Brussels, Belgium. Stem- loop RT- qpcr mirna profling: about single cells, normalization and biomarkers. CHI mirnas in health and disease, March 2009, Boston, USA. An integrative genomics screen uncovers ncrna T- UCR functions in neuroblastoma tumours. Advances in Genomics, January , Ghent, Belgium The neuroblastoma mirna map: prioritization and functional evaluation of candidate mirnas. Advances in Neuroblastoma Research, June , Stockholm, Sweden. The mirna body map: dissecting mirna function through integrative genomics. MicroRNA Europe 2010 meeting, November , Cambridge, United Kingdom. The mir mirna cluster regulates multiple components of the TGFβ- pathway in neuroblastoma. Keystone MicroRNAs and Non- Coding RNAs and Cancer, Februari 11-16, 2011, Banff, Alberta, Canada Patents Neuroblastoma prognostic multigene expression signature (N EP ; )
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
PART 3.3: MicroRNA and Cancer
 BIBM 2010 Tutorial: Epigenomics and Cancer PART 3.3: MicroRNA and Cancer Dec 18, 2010 Sun Kim at Indiana University Outline of Part 3.3 Background on microrna Role of microrna in cancer MicroRNA pathway
BIBM 2010 Tutorial: Epigenomics and Cancer PART 3.3: MicroRNA and Cancer Dec 18, 2010 Sun Kim at Indiana University Outline of Part 3.3 Background on microrna Role of microrna in cancer MicroRNA pathway
Micro RNAs: potentielle Biomarker für das. Blutspenderscreening
 Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
What is Cancer? Cancer is a genetic disease: Cancer typically involves a change in gene expression/function:
 Cancer is a genetic disease: Inherited cancer Sporadic cancer What is Cancer? Cancer typically involves a change in gene expression/function: Qualitative change Quantitative change Any cancer causing genetic
Cancer is a genetic disease: Inherited cancer Sporadic cancer What is Cancer? Cancer typically involves a change in gene expression/function: Qualitative change Quantitative change Any cancer causing genetic
Outline. interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work?
 Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved
 microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
Cancer SBL101. James Gomes School of Biological Sciences Indian Institute of Technology Delhi
 Cancer SBL101 James Gomes School of Biological Sciences Indian Institute of Technology Delhi All Figures in this Lecture are taken from 1. Molecular biology of the cell / Bruce Alberts et al., 5th ed.
Cancer SBL101 James Gomes School of Biological Sciences Indian Institute of Technology Delhi All Figures in this Lecture are taken from 1. Molecular biology of the cell / Bruce Alberts et al., 5th ed.
Special report. Chronic Lymphocytic Leukemia (CLL) Genomic Biology 3020 April 20, 2006
 Special report Chronic Lymphocytic Leukemia (CLL) Genomic Biology 3020 April 20, 2006 Gene And Protein The gene that causes the mutation is CCND1 and the protein NP_444284 The mutation deals with the cell
Special report Chronic Lymphocytic Leukemia (CLL) Genomic Biology 3020 April 20, 2006 Gene And Protein The gene that causes the mutation is CCND1 and the protein NP_444284 The mutation deals with the cell
RNAi Shooting the Messenger!
 RNAi Shooting the Messenger! Bronya Keats, Ph.D. Department of Genetics Louisiana State University Health Sciences Center New Orleans Email: [email protected] RNA interference (RNAi) A mechanism by which
RNAi Shooting the Messenger! Bronya Keats, Ph.D. Department of Genetics Louisiana State University Health Sciences Center New Orleans Email: [email protected] RNA interference (RNAi) A mechanism by which
mirnaselect pep-mir Cloning and Expression Vector
 Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
Control of Gene Expression
 Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Contents. molecular biology techniques. - Mutations in Factor II. - Mutations in MTHFR gene. - Breast cencer genes. - p53 and breast cancer
 Contents Introduction: biology and medicine, two separated compartments What we need to know: - boring basics in DNA/RNA structure and overview of particular aspects of molecular biology techniques - How
Contents Introduction: biology and medicine, two separated compartments What we need to know: - boring basics in DNA/RNA structure and overview of particular aspects of molecular biology techniques - How
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
Outline. MicroRNA Bioinformatics. microrna biogenesis. short non-coding RNAs not considered in this lecture. ! Introduction
 Outline MicroRNA Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology (CMB) Karolinska Institutet! Introduction! microrna target site prediction! Useful resources 2 short non-coding RNAs
Outline MicroRNA Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology (CMB) Karolinska Institutet! Introduction! microrna target site prediction! Useful resources 2 short non-coding RNAs
Lezioni Dipartimento di Oncologia Farmacologia Molecolare. RNA interference. Giovanna Damia 29 maggio 2006
 Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Notch 1 -dependent regulation of cell fate in colorectal cancer
 Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
CHAPTER 2: UNDERSTANDING CANCER
 CHAPTER 2: UNDERSTANDING CANCER INTRODUCTION We are witnessing an era of great discovery in the field of cancer research. New insights into the causes and development of cancer are emerging. These discoveries
CHAPTER 2: UNDERSTANDING CANCER INTRODUCTION We are witnessing an era of great discovery in the field of cancer research. New insights into the causes and development of cancer are emerging. These discoveries
岑 祥 股 份 有 限 公 司 技 術 專 員 費 軫 尹 20100803
 技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
Control of Gene Expression
 Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
Profiling of non-coding RNA classes Gunter Meister
 Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
CONTRACTING ORGANIZATION: University of Alabama at Birmingham Birmingham, AL 35294
 AD Award Number: W81XWH-08-1-0030 TITLE: Regulation of Prostate Cancer Bone Metastasis by DKK1 PRINCIPAL INVESTIGATOR: Gregory A. Clines, M.D., Ph.D. CONTRACTING ORGANIZATION: University of Alabama at
AD Award Number: W81XWH-08-1-0030 TITLE: Regulation of Prostate Cancer Bone Metastasis by DKK1 PRINCIPAL INVESTIGATOR: Gregory A. Clines, M.D., Ph.D. CONTRACTING ORGANIZATION: University of Alabama at
LEUKEMIA LYMPHOMA MYELOMA Advances in Clinical Trials
 LEUKEMIA LYMPHOMA MYELOMA Advances in Clinical Trials OUR FOCUS ABOUT emerging treatments Presentation for: Judith E. Karp, MD Advancements for Acute Myelogenous Leukemia Supported by an unrestricted educational
LEUKEMIA LYMPHOMA MYELOMA Advances in Clinical Trials OUR FOCUS ABOUT emerging treatments Presentation for: Judith E. Karp, MD Advancements for Acute Myelogenous Leukemia Supported by an unrestricted educational
MicroRNA signatures in human cancers
 MicroRNA signatures in human cancers George A. Calin and Carlo M. Croce Abstract MicroRNA (mirna ) alterations are involved in the initiation and progression of human cancer. The causes of the widespread
MicroRNA signatures in human cancers George A. Calin and Carlo M. Croce Abstract MicroRNA (mirna ) alterations are involved in the initiation and progression of human cancer. The causes of the widespread
GENE REGULATION. Teacher Packet
 AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
Department of BioScience Technology Chung Yuan Christian University 2015/08/13
 Department of BioScience Technology Chung Yuan Christian University 2015/08/13 Cancer Cells Cancer, the 1st leading cause of death, is an example of a disease that arises from abnormalities in cell function
Department of BioScience Technology Chung Yuan Christian University 2015/08/13 Cancer Cells Cancer, the 1st leading cause of death, is an example of a disease that arises from abnormalities in cell function
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
LESSON 3.5 WORKBOOK. How do cancer cells evolve? Workbook Lesson 3.5
 LESSON 3.5 WORKBOOK How do cancer cells evolve? In this unit we have learned how normal cells can be transformed so that they stop behaving as part of a tissue community and become unresponsive to regulation.
LESSON 3.5 WORKBOOK How do cancer cells evolve? In this unit we have learned how normal cells can be transformed so that they stop behaving as part of a tissue community and become unresponsive to regulation.
Biological Sciences Initiative. Human Genome
 Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Introduction. Cancer Biology. Tumor-suppressor genes. Proto-oncogenes. DNA stability genes. Mechanisms of carcinogenesis.
 Cancer Biology Chapter 18 Eric J. Hall., Amato Giaccia, Radiobiology for the Radiologist Introduction Tissue homeostasis depends on the regulated cell division and self-elimination (programmed cell death)
Cancer Biology Chapter 18 Eric J. Hall., Amato Giaccia, Radiobiology for the Radiologist Introduction Tissue homeostasis depends on the regulated cell division and self-elimination (programmed cell death)
Control of Gene Expression
 Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Sommaire projets sélectionnés mesure 29: Soutien à la recherche translationnelle
 Sommaire projets sélectionnés mesure 29: Soutien à la recherche translationnelle TITLE PROJET NOM HOPITAL Assessment of tumor angiogenesis using PET/CT with 18 F-Galacto- RGD. (PNC_29_001) Division of
Sommaire projets sélectionnés mesure 29: Soutien à la recherche translationnelle TITLE PROJET NOM HOPITAL Assessment of tumor angiogenesis using PET/CT with 18 F-Galacto- RGD. (PNC_29_001) Division of
The Biological Basis of Cancer. Annie Young, Professor of Nursing, University of Warwick, University Hospitals Coventry and Warwickshire
 The Biological Basis of Cancer Annie Young, Professor of Nursing, University of Warwick, University Hospitals Coventry and Warwickshire Aim of Presentation By the end of this presentation you will: Be
The Biological Basis of Cancer Annie Young, Professor of Nursing, University of Warwick, University Hospitals Coventry and Warwickshire Aim of Presentation By the end of this presentation you will: Be
MUTATION, DNA REPAIR AND CANCER
 MUTATION, DNA REPAIR AND CANCER 1 Mutation A heritable change in the genetic material Essential to the continuity of life Source of variation for natural selection New mutations are more likely to be harmful
MUTATION, DNA REPAIR AND CANCER 1 Mutation A heritable change in the genetic material Essential to the continuity of life Source of variation for natural selection New mutations are more likely to be harmful
Complex multicellular organisms are produced by cells that switch genes on and off during development.
 Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Estimated New Cases of Leukemia, Lymphoma, Myeloma 2014
 ABOUT BLOOD CANCERS Leukemia, Hodgkin lymphoma (HL), non-hodgkin lymphoma (NHL), myeloma, myelodysplastic syndromes (MDS) and myeloproliferative neoplasms (MPNs) are types of cancer that can affect the
ABOUT BLOOD CANCERS Leukemia, Hodgkin lymphoma (HL), non-hodgkin lymphoma (NHL), myeloma, myelodysplastic syndromes (MDS) and myeloproliferative neoplasms (MPNs) are types of cancer that can affect the
Types of Cancers [-oma growth ]!
![Types of Cancers [-oma growth ]! Types of Cancers [-oma growth ]!](/thumbs/27/10363750.jpg) Cancer: disease of transcription factors and replication 1 Uncontrolled cell growth and division -> immortalized cells -> tumor growth -> metastasis (cells float away from tumor and spread throughout the
Cancer: disease of transcription factors and replication 1 Uncontrolled cell growth and division -> immortalized cells -> tumor growth -> metastasis (cells float away from tumor and spread throughout the
Report series: General cancer information
 Fighting cancer with information Report series: General cancer information Eastern Cancer Registration and Information Centre ECRIC report series: General cancer information Cancer is a general term for
Fighting cancer with information Report series: General cancer information Eastern Cancer Registration and Information Centre ECRIC report series: General cancer information Cancer is a general term for
MicroRNAs (mirnas) are small, evolutionarily conserved,
 REVIEW doi:10.1038/nature10888 The microcosmos of cancer Amaia Lujambio 1 & Scott W. Lowe 1,2 The discovery of micrornas () almost two decades ago established a new paradigm of gene regulation. During
REVIEW doi:10.1038/nature10888 The microcosmos of cancer Amaia Lujambio 1 & Scott W. Lowe 1,2 The discovery of micrornas () almost two decades ago established a new paradigm of gene regulation. During
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
treatments) worked by killing cancerous cells using chemo or radiotherapy. While these techniques can
 Shristi Pandey Genomics and Medicine Winter 2011 Prof. Doug Brutlag Chronic Myeloid Leukemia: A look into how genomics is changing the way we treat Cancer. Until the late 1990s, nearly all treatment methods
Shristi Pandey Genomics and Medicine Winter 2011 Prof. Doug Brutlag Chronic Myeloid Leukemia: A look into how genomics is changing the way we treat Cancer. Until the late 1990s, nearly all treatment methods
The RNA strategy. RNA as a tool and target in human disease diagnosis and therapy.
 The RNA strategy RNA as a tool and target in human disease diagnosis and therapy. The Laboratory of RNA Biology and Biotechnology at the Centre for Integrative Biology (CIBIO) of the University of Trento,
The RNA strategy RNA as a tool and target in human disease diagnosis and therapy. The Laboratory of RNA Biology and Biotechnology at the Centre for Integrative Biology (CIBIO) of the University of Trento,
Tumour Markers. What are Tumour Markers? How Are Tumour Markers Used?
 Dr. Anthony C.H. YING What are? Tumour markers are substances that can be found in the body when cancer is present. They are usually found in the blood or urine. They can be products of cancer cells or
Dr. Anthony C.H. YING What are? Tumour markers are substances that can be found in the body when cancer is present. They are usually found in the blood or urine. They can be products of cancer cells or
School of Nursing. Presented by Yvette Conley, PhD
 Presented by Yvette Conley, PhD What we will cover during this webcast: Briefly discuss the approaches introduced in the paper: Genome Sequencing Genome Wide Association Studies Epigenomics Gene Expression
Presented by Yvette Conley, PhD What we will cover during this webcast: Briefly discuss the approaches introduced in the paper: Genome Sequencing Genome Wide Association Studies Epigenomics Gene Expression
A disease of populations of cells that live, divide, invade and spread without regard to normal limits
 1 Targeted Cancer Therapies Mark McKeage Medical Oncology Specialist Professor in Clinical Pharmacology 2 Cancer Definition- A disease of populations of cells that live, divide, invade and spread without
1 Targeted Cancer Therapies Mark McKeage Medical Oncology Specialist Professor in Clinical Pharmacology 2 Cancer Definition- A disease of populations of cells that live, divide, invade and spread without
MicroRNAs and other non-coding RNAs as targets for anticancer drug development
 MicroRNAs and other non-coding RNAs as targets for anticancer drug development Hui Ling 1 *, Muller Fabbri 2 * and George A. Calin 1,3 Abstract The first cancer-targeted microrna () drug MRX34, a liposome-based
MicroRNAs and other non-coding RNAs as targets for anticancer drug development Hui Ling 1 *, Muller Fabbri 2 * and George A. Calin 1,3 Abstract The first cancer-targeted microrna () drug MRX34, a liposome-based
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation
 OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
Psychoonkology, Sept. 2010 lifestyle factors and epigenetics
 Psychoonkology, Sept. 2010 lifestyle factors and epigenetics Alexander G. Haslberger Dep. für Ernährungswissenschaften Univ. of Vienna Working group: Food, GI-Microbiology, Epigenetics Content Health:
Psychoonkology, Sept. 2010 lifestyle factors and epigenetics Alexander G. Haslberger Dep. für Ernährungswissenschaften Univ. of Vienna Working group: Food, GI-Microbiology, Epigenetics Content Health:
Structure and Function of DNA
 Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Module 3 Questions. 7. Chemotaxis is an example of signal transduction. Explain, with the use of diagrams.
 Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
ACUTE MYELOID LEUKEMIA (AML),
 1 ACUTE MYELOID LEUKEMIA (AML), ALSO KNOWN AS ACUTE MYELOGENOUS LEUKEMIA WHAT IS CANCER? The body is made up of hundreds of millions of living cells. Normal body cells grow, divide, and die in an orderly
1 ACUTE MYELOID LEUKEMIA (AML), ALSO KNOWN AS ACUTE MYELOGENOUS LEUKEMIA WHAT IS CANCER? The body is made up of hundreds of millions of living cells. Normal body cells grow, divide, and die in an orderly
Head of College Scholars List Scheme. Summer Studentship. Report Form
 Head of College Scholars List Scheme Summer Studentship Report Form This report should be completed by the student with his/her project supervisor. It should summarise the work undertaken during the project
Head of College Scholars List Scheme Summer Studentship Report Form This report should be completed by the student with his/her project supervisor. It should summarise the work undertaken during the project
New strategies in anticancer therapy
 癌 症 診 療 指 引 簡 介 及 臨 床 應 用 New strategies in anticancer therapy 中 山 醫 學 大 學 附 設 醫 院 腫 瘤 內 科 蔡 明 宏 醫 師 2014/3/29 Anti-Cancer Therapy Surgical Treatment Radiotherapy Chemotherapy Target Therapy Supportive
癌 症 診 療 指 引 簡 介 及 臨 床 應 用 New strategies in anticancer therapy 中 山 醫 學 大 學 附 設 醫 院 腫 瘤 內 科 蔡 明 宏 醫 師 2014/3/29 Anti-Cancer Therapy Surgical Treatment Radiotherapy Chemotherapy Target Therapy Supportive
Update in Hematology Oncology Targeted Therapies. Mark Holguin
 Update in Hematology Oncology Targeted Therapies Mark Holguin 25 years ago Why I chose oncology People How to help people with possibly the most difficult thing they may have to deal with Science Turning
Update in Hematology Oncology Targeted Therapies Mark Holguin 25 years ago Why I chose oncology People How to help people with possibly the most difficult thing they may have to deal with Science Turning
Leukemia Research Foundation 2004-2005 Scientific Research Grant Recipients
 Page 1 of 5 NEW INVESTIGATOR AWARDS Ioannis Aifantis, Ph.D. The University of Chicago, Chicago, IL $75,000.00 Cooperation of Notch and pre-tcr Signaling in the Induction of T Cell Leukemia The pre-t Cell
Page 1 of 5 NEW INVESTIGATOR AWARDS Ioannis Aifantis, Ph.D. The University of Chicago, Chicago, IL $75,000.00 Cooperation of Notch and pre-tcr Signaling in the Induction of T Cell Leukemia The pre-t Cell
13.4 Gene Regulation and Expression
 13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
Leukemias and Lymphomas: A primer
 Leukemias and Lymphomas: A primer Normal blood contains circulating white blood cells, red blood cells and platelets 700 red cells (oxygen) 1 white cell Neutrophils (60%) bacterial infection Lymphocytes
Leukemias and Lymphomas: A primer Normal blood contains circulating white blood cells, red blood cells and platelets 700 red cells (oxygen) 1 white cell Neutrophils (60%) bacterial infection Lymphocytes
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals
 Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
Systematic discovery of regulatory motifs in human promoters and 30 UTRs by comparison of several mammals Xiaohui Xie 1, Jun Lu 1, E. J. Kulbokas 1, Todd R. Golub 1, Vamsi Mootha 1, Kerstin Lindblad-Toh
The Making of the Fittest: Evolving Switches, Evolving Bodies
 OVERVIEW MODELING THE REGULATORY SWITCHES OF THE PITX1 GENE IN STICKLEBACK FISH This hands-on activity supports the short film, The Making of the Fittest:, and aims to help students understand eukaryotic
OVERVIEW MODELING THE REGULATORY SWITCHES OF THE PITX1 GENE IN STICKLEBACK FISH This hands-on activity supports the short film, The Making of the Fittest:, and aims to help students understand eukaryotic
Mature Lymphoproliferative disorders (2): Mature B-cell Neoplasms. Dr. Douaa Mohammed Sayed
 Mature Lymphoproliferative disorders (2): Mature B-cell Neoplasms Dr. Douaa Mohammed Sayed Small lymphocytic lymphoma/b-cell chronic lymphocytic leukemia BMB: nodular, interstitial, diffuse or a combination
Mature Lymphoproliferative disorders (2): Mature B-cell Neoplasms Dr. Douaa Mohammed Sayed Small lymphocytic lymphoma/b-cell chronic lymphocytic leukemia BMB: nodular, interstitial, diffuse or a combination
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION. Professor Bharat Patel Office: Science 2, 2.36 Email: [email protected].
 Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: [email protected] What is Gene Expression & Gene Regulation? 1. Gene Expression
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: [email protected] What is Gene Expression & Gene Regulation? 1. Gene Expression
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
MicroRNA formation. 4th International Symposium on Non-Surgical Contraceptive Methods of Pet Population Control
 MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
LYMPHOMA. BACHIR ALOBEID, M.D. HEMATOPATHOLOGY DIVISION PATHOLOGY DEPARTMENT Columbia University/ College of Physicians & Surgeons
 LYMPHOMA BACHIR ALOBEID, M.D. HEMATOPATHOLOGY DIVISION PATHOLOGY DEPARTMENT Columbia University/ College of Physicians & Surgeons Normal development of lymphocytes Lymphocyte proliferation and differentiation:
LYMPHOMA BACHIR ALOBEID, M.D. HEMATOPATHOLOGY DIVISION PATHOLOGY DEPARTMENT Columbia University/ College of Physicians & Surgeons Normal development of lymphocytes Lymphocyte proliferation and differentiation:
1 Mutation and Genetic Change
 CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
Summary of Discussion on Non-clinical Pharmacology Studies on Anticancer Drugs
 Provisional Translation (as of January 27, 2014)* November 15, 2013 Pharmaceuticals and Bio-products Subcommittees, Science Board Summary of Discussion on Non-clinical Pharmacology Studies on Anticancer
Provisional Translation (as of January 27, 2014)* November 15, 2013 Pharmaceuticals and Bio-products Subcommittees, Science Board Summary of Discussion on Non-clinical Pharmacology Studies on Anticancer
How Cancer Begins???????? Chithra Manikandan Nov 2009
 Cancer Cancer is one of the most common diseases in the developed world: 1 in 4 deaths are due to cancer 1 in 17 deaths are due to lung cancer Lung cancer is the most common cancer in men Breast cancer
Cancer Cancer is one of the most common diseases in the developed world: 1 in 4 deaths are due to cancer 1 in 17 deaths are due to lung cancer Lung cancer is the most common cancer in men Breast cancer
DNA Replication & Protein Synthesis. This isn t a baaaaaaaddd chapter!!!
 DNA Replication & Protein Synthesis This isn t a baaaaaaaddd chapter!!! The Discovery of DNA s Structure Watson and Crick s discovery of DNA s structure was based on almost fifty years of research by other
DNA Replication & Protein Synthesis This isn t a baaaaaaaddd chapter!!! The Discovery of DNA s Structure Watson and Crick s discovery of DNA s structure was based on almost fifty years of research by other
Regulation of Protein Translation and c-jun expression by Prostate Tumor Overexpressed1 (PTOV1)
 Regulation of Protein Translation and c-jun expression by Prostate Tumor Overexpressed1 (PTOV1) Verónica Cánovas, PhD Student Laboratory of Cell Signalling and Cancer Progression, Dra. Rosanna Paciucci
Regulation of Protein Translation and c-jun expression by Prostate Tumor Overexpressed1 (PTOV1) Verónica Cánovas, PhD Student Laboratory of Cell Signalling and Cancer Progression, Dra. Rosanna Paciucci
Pathway Analysis : An Introduction
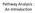 Pathway Analysis : An Introduction Experiments Literature and other KB Data Knowledge Structure in Data through statistics Structure in Knowledge through GO and other Ontologies Gain insight into Data
Pathway Analysis : An Introduction Experiments Literature and other KB Data Knowledge Structure in Data through statistics Structure in Knowledge through GO and other Ontologies Gain insight into Data
Metastasis. Brookdale Hospital, Brooklyn, New York 11212, USA; 2 Cambridge, MA 02138, USA [email protected]
 Metastasis Ma Hongbao 1, Margaret Ma 2, Yang Yan 1 1 Brookdale Hospital, Brooklyn, New York 11212, USA; 2 Cambridge, MA 02138, USA [email protected] Abstract: Cancer begins when cells in a part of the body
Metastasis Ma Hongbao 1, Margaret Ma 2, Yang Yan 1 1 Brookdale Hospital, Brooklyn, New York 11212, USA; 2 Cambridge, MA 02138, USA [email protected] Abstract: Cancer begins when cells in a part of the body
Review. MicroRNAs in Tumorigenesis. A Primer. Biogenesis of mirna. Basic Biology of MicroRNAs A Brief History of MicroRNA Discovery
 ASIP Journal CME Program The American Journal of Pathology, Vol. 171, No. 3, September 007 Copyright American Society for Investigative Pathology DOI: 10.353/ajpath.007.070070 Review MicroRNAs in Tumorigenesis
ASIP Journal CME Program The American Journal of Pathology, Vol. 171, No. 3, September 007 Copyright American Society for Investigative Pathology DOI: 10.353/ajpath.007.070070 Review MicroRNAs in Tumorigenesis
Functional RNAs; RNA catalysts, mirna,
 Functional RNAs; RNA catalysts, mirna, srna, RNAi... RNAs have many functions rrna (ribosomal RNA) trna (transfer RNA) mrna (Messenger RNA) snrna (including snorna) ) (Small nuclear RNA- splicing) Other
Functional RNAs; RNA catalysts, mirna, srna, RNAi... RNAs have many functions rrna (ribosomal RNA) trna (transfer RNA) mrna (Messenger RNA) snrna (including snorna) ) (Small nuclear RNA- splicing) Other
Developments in Biomarker Identification and Validation for Lung Cancer
 Developments in Biomarker Identification and Validation for Lung Cancer Alexandre Passioukov, MD, PhD [email protected] Contents Introduction Lung cancer pathogenesis NSCLC treatment options
Developments in Biomarker Identification and Validation for Lung Cancer Alexandre Passioukov, MD, PhD [email protected] Contents Introduction Lung cancer pathogenesis NSCLC treatment options
Chapter 5: Organization and Expression of Immunoglobulin Genes
 Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Gene Therapy. The use of DNA as a drug. Edited by Gavin Brooks. BPharm, PhD, MRPharmS (PP) Pharmaceutical Press
 Gene Therapy The use of DNA as a drug Edited by Gavin Brooks BPharm, PhD, MRPharmS (PP) Pharmaceutical Press Contents Preface xiii Acknowledgements xv About the editor xvi Contributors xvii An introduction
Gene Therapy The use of DNA as a drug Edited by Gavin Brooks BPharm, PhD, MRPharmS (PP) Pharmaceutical Press Contents Preface xiii Acknowledgements xv About the editor xvi Contributors xvii An introduction
Gene mutation and molecular medicine Chapter 15
 Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
Cytotoxic and Biotherapies Credentialing Programme Module 2
 Cytotoxic and Biotherapies Credentialing Programme Module 2 1. The Cell Cycle 2. Cancer Therapies 3. Adjunctive Therapies On completion of this module the RN will State the difference between a normal
Cytotoxic and Biotherapies Credentialing Programme Module 2 1. The Cell Cycle 2. Cancer Therapies 3. Adjunctive Therapies On completion of this module the RN will State the difference between a normal
The Human Genome Project. From genome to health From human genome to other genomes and to gene function Structural Genomics initiative
 The Human Genome Project From genome to health From human genome to other genomes and to gene function Structural Genomics initiative June 2000 What is the Human Genome Project? U.S. govt. project coordinated
The Human Genome Project From genome to health From human genome to other genomes and to gene function Structural Genomics initiative June 2000 What is the Human Genome Project? U.S. govt. project coordinated
Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Biochemistry
 Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Biochemistry The Master Degree in Medical Laboratory Sciences /Clinical Biochemistry, is awarded by the Faculty of Graduate Studies
Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Biochemistry The Master Degree in Medical Laboratory Sciences /Clinical Biochemistry, is awarded by the Faculty of Graduate Studies
Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Microbiology, Immunology and Serology
 Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Microbiology, Immunology and Serology The Master Degree in Medical Laboratory Sciences / Clinical Microbiology, Immunology or
Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Microbiology, Immunology and Serology The Master Degree in Medical Laboratory Sciences / Clinical Microbiology, Immunology or
Fulfilling the Promise
 Fulfilling the Promise Advancing the Fight Against Cancer: America s Medical Schools and Teaching Hospitals For more than a century, the nation s medical schools and teaching hospitals have worked to understand,
Fulfilling the Promise Advancing the Fight Against Cancer: America s Medical Schools and Teaching Hospitals For more than a century, the nation s medical schools and teaching hospitals have worked to understand,
Cancer. 9p21.3 deletion. t(12;21) t(15;17)
 CANCER FISH PROBES INDIVIDUAL AND PANEL S Acute Lymphoblastic Leukemia (ALL) ALL FISH Panel (includes all probes below) 8010 LSI MYB/CEP6 LSI p16 (CDKN2A) LSI BCR/ABL with ASS LSI ETV6 (TEL)/AML1 (RUNX1)
CANCER FISH PROBES INDIVIDUAL AND PANEL S Acute Lymphoblastic Leukemia (ALL) ALL FISH Panel (includes all probes below) 8010 LSI MYB/CEP6 LSI p16 (CDKN2A) LSI BCR/ABL with ASS LSI ETV6 (TEL)/AML1 (RUNX1)
Breast cancer and the role of low penetrance alleles: a focus on ATM gene
 Modena 18-19 novembre 2010 Breast cancer and the role of low penetrance alleles: a focus on ATM gene Dr. Laura La Paglia Breast Cancer genetic Other BC susceptibility genes TP53 PTEN STK11 CHEK2 BRCA1
Modena 18-19 novembre 2010 Breast cancer and the role of low penetrance alleles: a focus on ATM gene Dr. Laura La Paglia Breast Cancer genetic Other BC susceptibility genes TP53 PTEN STK11 CHEK2 BRCA1
Basic Concepts of DNA, Proteins, Genes and Genomes
 Basic Concepts of DNA, Proteins, Genes and Genomes Kun-Mao Chao 1,2,3 1 Graduate Institute of Biomedical Electronics and Bioinformatics 2 Department of Computer Science and Information Engineering 3 Graduate
Basic Concepts of DNA, Proteins, Genes and Genomes Kun-Mao Chao 1,2,3 1 Graduate Institute of Biomedical Electronics and Bioinformatics 2 Department of Computer Science and Information Engineering 3 Graduate
MULTIPLE MYELOMA. Dr Malkit S Riyat. MBChB, FRCPath(UK) Consultant Haematologist
 MULTIPLE MYELOMA Dr Malkit S Riyat MBChB, FRCPath(UK) Consultant Haematologist Multiple myeloma is an incurable malignancy that arises from postgerminal centre, somatically hypermutated B cells.
MULTIPLE MYELOMA Dr Malkit S Riyat MBChB, FRCPath(UK) Consultant Haematologist Multiple myeloma is an incurable malignancy that arises from postgerminal centre, somatically hypermutated B cells.
Cancer Immunotherapy: Can Your Immune System Cure Cancer? Steve Emerson, MD, PhD Herbert Irving Comprehensive Cancer Center
 Cancer Immunotherapy: Can Your Immune System Cure Cancer? Steve Emerson, MD, PhD Herbert Irving Comprehensive Cancer Center Bodnar s Law Simple Things are Important Very Simple Things are Very Important
Cancer Immunotherapy: Can Your Immune System Cure Cancer? Steve Emerson, MD, PhD Herbert Irving Comprehensive Cancer Center Bodnar s Law Simple Things are Important Very Simple Things are Very Important
Introduction. About 10,500 new cases of acute myelogenous leukemia are diagnosed each
 Introduction 1.1 Introduction: About 10,500 new cases of acute myelogenous leukemia are diagnosed each year in the United States (Hope et al., 2003). Acute myelogenous leukemia has several names, including
Introduction 1.1 Introduction: About 10,500 new cases of acute myelogenous leukemia are diagnosed each year in the United States (Hope et al., 2003). Acute myelogenous leukemia has several names, including
Lecture 3: Mutations
 Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
Lesson 3 Reading Material: Oncogenes and Tumor Suppressor Genes
 Lesson 3 Reading Material: Oncogenes and Tumor Suppressor Genes Becoming a cancer cell isn t easy One of the fundamental molecular characteristics of cancer is that it does not develop all at once, but
Lesson 3 Reading Material: Oncogenes and Tumor Suppressor Genes Becoming a cancer cell isn t easy One of the fundamental molecular characteristics of cancer is that it does not develop all at once, but
Targeted Therapy What the Surgeon Needs to Know
 Targeted Therapy What the Surgeon Needs to Know AATS Focus in Thoracic Surgery 2014 David R. Jones, M.D. Professor & Chief, Thoracic Surgery Memorial Sloan Kettering Cancer Center I have no disclosures
Targeted Therapy What the Surgeon Needs to Know AATS Focus in Thoracic Surgery 2014 David R. Jones, M.D. Professor & Chief, Thoracic Surgery Memorial Sloan Kettering Cancer Center I have no disclosures
