Investigating the role of a Cryptosporidium parum apyrase in infection
|
|
|
- Arnold Stafford
- 8 years ago
- Views:
Transcription
1 Investigating the role of a Cryptosporidium parum apyrase in infection David Riccardi and Patricio Manque Abstract This project attempted to characterize the function of a Cryptosporidium parvum apyrase termed, Similar to Riken (SRK), and its human ortholog, CANT-1,. First, we looked at SRK s effect on gene expression in host cells with a microarray analysis, which revealed significant changes in both adenosine and apoptosis pathways. We also attempted to elucidate the biological function of CANT1 using RNAi, indirect immunofluorescence, and flow cyotmetry assays. Introduction The ubiquitous protozoan parasite Cryptosporidium parvum causes severe diarrhea in millions of people every year by infecting and destroying epithelial cells of the small intestine. Transmitted through water supplies, infected animals, and oral-fecal contact, the disease is especially harmful to immunocompromised individuals. Recently, a parasite apryase named SRK has been shown to be critically important for successful invasion likely by disrupting intracellular signaling pathways associated with the prevention of invasion. Immunoprecipitation and mass mass spectrometry data indicate that SRK binds to its human ortholog, CANT1, in vitro. Unfortunately, little is known about the biological function of CANT1. Unlike other apyrases, it lacks ADP hydrolysis activity and is soluble (1). However, like other apryases, its activity is dependent upon homodimerization (2). Methods The goal of our research project was two fold: 1. To characterization the gene expression of both SRK in Cryptosporidium and its effects on host cells by microarray and RT- PCR. 2. To examine the biological function of CANT-1 by investigating its role during infection and its localization in host cells. RNA extraction for RT-PCR and Microarray experiments RNA was extracted using TRIzol Reagents (Invitrogen) according to the manufacturer s instructions. Microarray analysis of SRK s effect on the transcriptional profile of human epithelial cells
2 We scanned and hybridized 4 Human U1333 v2 arrays (Affymetrix) with total RNA extract from 10^6 HCT-8 cells (human epithelial cell line) treated with either 0.1 µm of recombinant SRK for 3 hours (2 microarrays) or no treatment (2 microarrays). The resulting CEL files were normalized using the RMA method (3) and the differentially expressed genes were identified by pairwise t-tests. RT-PCR of SRK RT-PCR was performed on RNA extracted 3 hours, 6 hours, 20 hours, 24 hours, and 48 hours 5 x 10^5 HCT-8 after infection with 3 x 10^6 Cryptosporidium parvum sporozoites using the Superscript III One Step RT-PCR System kit (Invitrogen) and the following primers: SRK forward (5 -> 3 ): AGGAAAGGAGGGTTTGCACT SRK reverse (5 -> 3 ): CCATCCCTCTCCCATATTCA Control gene: GAPDH GAPDH forward (5 -> 3 ): GTTTCGGACGTATTGGTCGT GAPDH reverse (5 -> 3 ): CCATCCCTCTCCCATATTCA For every sample, a no reverse transcriptase control reaction was performed with the Platinum Taq DNA Polymerase High Fidelity (Invitrogen) kit. Indirect immunofluorescence of CANT1 in epithelial cells 5 x 10^5 HCT-8 cells were fixed with 4 % paraformaldehyde in PBS for 1 hour, washed three times with PBS, and blocked with NH 4 Cl for 15 minutes. After 3 washes with PBS, cells were then blocked overnight with PGN, followed by three PBS washes. If the cells were permeablized, they were treated with 0.01% Triton X-100 solution for 5 minutes. Cells were then washed again, and incubated with serum immunized with recombinant SRK protein diluted 1:80 in PBS. This serum contains antibodies against SRK. Control treatments were incubated with non-immunized serum ( normal serum ). Then, after 3 washes % Tween 20 (Fischer Scientific) in PBS, cells were treated with anti-mouse IgG FITC conjugated antibody (Sigma) diluted 1:100 in PBS. Cell were washed once again, treated with DAPI and Texas Red- X Phallodin (Molecular Probes) solutions, with 1:50 and 1:100 PBS dilutions, respectively. Cells were then visualized on the IX81 confocal microscopy (Olympus). Flow Cytometry Cells were prepared in the same manner as the immunofluorescence, expect that the 5 x 10^6 cells were used, and the cells were not stained with DAPI or Texas Red. Cells were analyzed on the FC 500 cell analyzer (Cytomics).
3 RNAi of CANT1 In epithelial cells 10^6 HCT-8 cells were transfected with CANT1, GAPDH, or non-target sirnas using Oligofectamine Reagents (Invitrogen) according to the manufacturer s instructions. CANT1 protein levels were assessed using standard western blotting procedures and serum obtained from mice injected with recombinant SRK protein. Results Microarray Anaylsis of SRK s effects on host cells To elucidate possible effects of SRK on host cells, we analyzed the transcriptional profile of host cells after treatment with recombinant SRK protein with Human U1333 v2 arrays. Statistical analysis revealed that 102 genes were differentially expressed, including genes involved in apoptosis and AMP signaling. Several altered genes in an apoptotic network are shown in Figure 1.
4 Figure 1. Pathway analysis of differential expressed genes after treatment with SRK. After statistical analysis, selected genes were analyzed using Ingenuity pathways. Genes that are upregulated are in red, and in green, those that are down regulated.
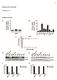
5 RT-PCR of SRK gene To assess the expression level of the SRK gene during infection, we performed RT-PCR of Cryptosporidium sporozites before infection, and 3 hours, 6 hours, 20 hours, 24 hours, and 48 hours after infection. Preliminary results showed no amphilifcation either of SRK or the positive control GAPDH gene. After increasing the total RNA concentration from 10 ng to 100 ng, amplification for the GAPDH gene could be seen at 20 hours, 24 hours, and 48 hours (Figure 2a), but not below these time points, and no SRK amplification was seen (Figures 2b). To test the specificity of the SRK primer, PCR was performed on a previously prepared SRK plasmid and Cryptosporidium DNA, and the plasmid did amplify, but the DNA did not (Figure 2b and 2c). The Cryptosporidium DNA did amplify the GAPDH gene, however. Lanes Figure 2a. RT-PCR of GAPDH gene in Cryptosporidium during infection. 1% agarose gel stained with ethidium bromide. Lane 1 contains a digested λ plasmid, lanes 2-6 contain RNA extracted 3 hours, 6 hours, 20 hours, 24 hours, 48 hours, and 0 hours after infection, respectively. Clear bands at the predicted amplicon size are seen in 20 hours, 24 hours, and 48 hour infected samples. Lanes contain RNA in the sample orientation as lanes 2-6, but lacked reverse transcriptase in the PCR reaction.
6 Figure 2b. RT-PCR of SRK gene in Cryptosporidium during infection. 1% agarose gel stained with ethidium bromide. Lanes 1 and 9 contains a digested λ plasmid, lanes 2-6 contain RNA extracted 3 hours, 6 hours, 20 hours, 24 hours, 48 hours, and 0 hours after infection, respectively. Lane 8 was loaded with SRK DNA plasmid, and has a clear band at the predicted amplicon size. Lanes contain samples in the orientation as lanes 2-7, but lacked reverse transcriptase in the PCR reaction. Lane 16 was loaded with SRK DNA plasmid, and has a clear band at the predicted amplicon size.
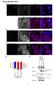
7 Figure 2c. PCR of SRK and GAPDH in Cryptosporidium. 1% agraose gel stained with ethidium bromide is shown. The first and last lanes contain a digested λ plasmid. The 6 lanes on the left contain PCR product using Cryptosporidium DNA as a template and SRK primers. A gradient of annealing temperature between 57 and 65 degrees (increasing from left to right lanes) was used in the PCR reaction. The 6 lanes on the right contain PCR product using Cryptosporidium DNA as a template and GAPDH primers. A gradient of annealing temperature between 57 and 65 degrees (increasing from left to right lanes) waas used in the PCR reaction. For all 6 lanes, a clear band was seen. Localization of CANT1 in epithelial cells To locate CANT1 in epithelial cells, an indirect immunfluoresecne assay was employed. Because of the high degree of sequence similarity between SRK and CANT1, polyclonal antibodies against SRK also bind to CANT1 (western blot data not shown). CANT1 appears to be located at outside of the nucleus of permeablized HCT-8 cells (Figure 3a), but due to non-specific binding of the secondary antibody in the normal serum control (Figure 3b) this result can not be attributed solely to antibodies binding to CANT1.
8 Figure 3a. Indirect immunofluorescence of CANT1. Micrograph showing permeablized HCT-8 cells treated with anti-srk and anti-mouse IgG antibodies (FITC label, green). The cytoskeleton (, Texas Red-phallodin red) and nucleus (DAPI, blue) were also stained. Note the green staining around the nucleus.
9 Figure 3b. Indirect immunofluorescence contro. Micrograph showing HCT-8 cells treated normal serum and anti-mouse IgG antibody (FITC label, green). Cytoskeleton (, Texas Red-phallodin red) and nucleus (DAPI, blue) were also stained. Not the green staining surrounding cells. Flow cytometry was also performed on permeablized and non-permeablized with the same antibody treatment used above. If CANT1 was located at the surface of the cell, we hypothesized that non-permeablized cells would have high fluorescence, and permeablized cells would have low fluorescence. If CANT1 was located on the inside of the cell, we would expect the opposite results. Cell populations treated with mouse normal serum had high levels of fluorescence in non-permeablized (Figure 4a ) and permeablized cells (Figure 4b and 4c), masking any fluorescence caused by SRK antibody binding to CANT1.
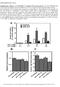
10 Figure 4a. Flow cyotmetry analysis of non-permeablized HCT-8 cells. Shown here is a histogram plot, with X-axis indicates FITC emitted fluorescence intensity of individual cells (log scale) and y-axis indicates cell number. Black line represents autofluoresce cells (no antibody treatements), blue line represents cells treated with normal serum and anti-mouse IgG antibody conjugated with a FITC molecule, and green line represents cells treated with anti-srk antibody and anti-mouse IgG antibody conjugated with a FITC molecule. Note that in both cell populations treated with normal serum and the secondary antibody, there a large number of cells that fluoresce at high intensity.
11 Figure 4b. Flow cyotmetry analysis of permeablized HCT-8 cells. Shown here is a histogram plot, with X-axis indicates FITC emitted fluorescence intensity of individual cells and y-axis indicates cell number. Cells were treated with normal serum and antimouse IgG antibody conjugated with a FITC molecule. Figure 4c. Flow cyotmetry analysis of permeablized HCT-8 cells. Shown here is a histogram plot, with X-axis indicates FITC emitted fluorescence intensity of individual cells and y-axis indicates cell number. Cells were treated with anti-srk antibody and anti-mouse IgG antibody conjugated with a FITC molecule.
12 RNAi of CANT1 in epithelial cells In order to investigate a possible role of CANT1 during infection, we attempted to HCT- 8 cells deficient for CANT1 protein with RNAi technology. However, we were able to reduce CANT1 protein levels (Figure 5). Figure 5. RNAi of CANT1. Shown here is a western blot of HCT-8 protein extract. From left to right, lane 1 contains a protein ladder, the next 4 lanes extract from cells treated sirna directed against CANT1 and last the two lanes contain extract from cells treated sirna directed GAPDH. Equal amounts of protein were loaded into each well, and the membrane was probed with antibodies against SRK. Note that there was no reduction in CANT1 protein in the knockout cell extracts. Future directions Because the primers failed to amplify the SRK in total genomic Cryptosporidium DNA, it is very likely that the primers are non-specific. A bioinformatic tool for the Cryptosporidium database similar to Primer-Blast for the human genome would be very useful. Because of the fluorescence of normal serum controls in the indirect immunofluorescence and flow cytometry experiments, it is highly probable that the secondary antibody is
13 binding to another immunoglobin in the mouse serum other than thr SRK antibody. Three possible solutions to this problem are: 1. Use a different secondary antibody. 2. Treat HCT-8 cells with goat serum (species in which secondary antibody was obtained from) to block murine immunglobins from binding to F C receptors. 3. Develop a monoclonal antibody. The protocol for CANT1 knock down by RNAi needs to be optimized. One might do this by experimenting with different concentrations of sirna and oligofectamine reagents. The microarray experiments suggest that SRK might influence apoptosis pathways. It has been already established that Cryptosporidium represses host cell apoptosis (4), but the mechanism in which this occurs remains to be elucidated. The microarray results need to be validated by RT-PCR, and it would be interesting to investigate possible changes in apoptosis levels in host cells infected with parasites coated in anti-srk antibodies. If SRK prevents apoptosis, one would predict these cells to have a higher rate of apoptosis. References 1. Smith TM, Hicks-Berger CA, Kim S, Kirley TL: Cloning, expression, and characterization of a soluble calcium-activated nucleotidase, a human enzyme belonging to a new family of extracellular nucleotidases. Archives of Biochemistry and Biophysics 2002, 406: Yang M, Horii K, Herr AB, Kirley TL: Characterization and importance of the dimer interface of human calcium-activated nucleotidase. Biochemistry 2008, 47: Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP: Exploration, normalization, and summaries of high density oligonucleotide array probe level data. 4. McCole DF, Eckmann L, Laurent F, Kagnoff MF: Intestinal epithelial cell apoptosis following Cryptosporidium parvum infection. Infection and Immunity 2000, 68:
Essentials of Real Time PCR. About Sequence Detection Chemistries
 Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
KMS-Specialist & Customized Biosimilar Service
 KMS-Specialist & Customized Biosimilar Service 1. Polyclonal Antibody Development Service KMS offering a variety of Polyclonal Antibody Services to fit your research and production needs. we develop polyclonal
KMS-Specialist & Customized Biosimilar Service 1. Polyclonal Antibody Development Service KMS offering a variety of Polyclonal Antibody Services to fit your research and production needs. we develop polyclonal
Real-Time PCR Vs. Traditional PCR
 Real-Time PCR Vs. Traditional PCR Description This tutorial will discuss the evolution of traditional PCR methods towards the use of Real-Time chemistry and instrumentation for accurate quantitation. Objectives
Real-Time PCR Vs. Traditional PCR Description This tutorial will discuss the evolution of traditional PCR methods towards the use of Real-Time chemistry and instrumentation for accurate quantitation. Objectives
Introduction To Real Time Quantitative PCR (qpcr)
 Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
2.1.2 Characterization of antiviral effect of cytokine expression on HBV replication in transduced mouse hepatocytes line
 i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
THE His Tag Antibody, mab, Mouse
 THE His Tag Antibody, mab, Mouse Cat. No. A00186 Technical Manual No. TM0243 Update date 01052011 I Description.... 1 II Key Features. 2 III Storage 2 IV Applications.... 2 V Examples - ELISA..... 2 VI
THE His Tag Antibody, mab, Mouse Cat. No. A00186 Technical Manual No. TM0243 Update date 01052011 I Description.... 1 II Key Features. 2 III Storage 2 IV Applications.... 2 V Examples - ELISA..... 2 VI
Molecular Biology Techniques: A Classroom Laboratory Manual THIRD EDITION
 Molecular Biology Techniques: A Classroom Laboratory Manual THIRD EDITION Susan Carson Heather B. Miller D.Scott Witherow ELSEVIER AMSTERDAM BOSTON HEIDELBERG LONDON NEW YORK OXFORD PARIS SAN DIEGO SAN
Molecular Biology Techniques: A Classroom Laboratory Manual THIRD EDITION Susan Carson Heather B. Miller D.Scott Witherow ELSEVIER AMSTERDAM BOSTON HEIDELBERG LONDON NEW YORK OXFORD PARIS SAN DIEGO SAN
Gene Expression Assays
 APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
Materials and Methods. Blocking of Globin Reverse Transcription to Enhance Human Whole Blood Gene Expression Profiling
 Application Note Blocking of Globin Reverse Transcription to Enhance Human Whole Blood Gene Expression Profi ling Yasmin Beazer-Barclay, Doug Sinon, Christopher Morehouse, Mark Porter, and Mike Kuziora
Application Note Blocking of Globin Reverse Transcription to Enhance Human Whole Blood Gene Expression Profi ling Yasmin Beazer-Barclay, Doug Sinon, Christopher Morehouse, Mark Porter, and Mike Kuziora
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
Supplemental Information. McBrayer et al. Supplemental Data
 1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
RevertAid Premium First Strand cdna Synthesis Kit
 RevertAid Premium First Strand cdna Synthesis Kit #K1651, #K1652 CERTIFICATE OF ANALYSIS #K1651 Lot QUALITY CONTROL RT-PCR using 100 fg of control GAPDH RNA and GAPDH control primers generated a prominent
RevertAid Premium First Strand cdna Synthesis Kit #K1651, #K1652 CERTIFICATE OF ANALYSIS #K1651 Lot QUALITY CONTROL RT-PCR using 100 fg of control GAPDH RNA and GAPDH control primers generated a prominent
Speed Matters - Fast ways from template to result
 qpcr Symposium 2007 - Weihenstephan Speed Matters - Fast ways from template to result March 28, 2007 Dr. Thorsten Traeger Senior Scientist, Research and Development - 1 - Overview Ạgenda Fast PCR The Challenges
qpcr Symposium 2007 - Weihenstephan Speed Matters - Fast ways from template to result March 28, 2007 Dr. Thorsten Traeger Senior Scientist, Research and Development - 1 - Overview Ạgenda Fast PCR The Challenges
The cell lines used in this study were obtained from the American Type Culture
 Supplementary materials and methods Cell culture and drug treatments The cell lines used in this study were obtained from the American Type Culture Collection (ATCC) and grown as previously described.
Supplementary materials and methods Cell culture and drug treatments The cell lines used in this study were obtained from the American Type Culture Collection (ATCC) and grown as previously described.
Virological Methods. Flint et al. Principles of Virology (ASM), Chapter 2
 Virological Methods Flint et al. Principles of Virology (ASM), Chapter 2 Overview The most commonly used laboratory methods for the detection of viruses and virus components in biological samples can be
Virological Methods Flint et al. Principles of Virology (ASM), Chapter 2 Overview The most commonly used laboratory methods for the detection of viruses and virus components in biological samples can be
CCR Biology - Chapter 9 Practice Test - Summer 2012
 Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual
 Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Validating Microarray Data Using RT 2 Real-Time PCR Products
 Validating Microarray Data Using RT 2 Real-Time PCR Products Introduction: Real-time PCR monitors the amount of amplicon as the reaction occurs. Usually, the amount of product is directly related to the
Validating Microarray Data Using RT 2 Real-Time PCR Products Introduction: Real-time PCR monitors the amount of amplicon as the reaction occurs. Usually, the amount of product is directly related to the
The immune response Antibodies Antigens Epitopes (antigenic determinants) the part of a protein antigen recognized by an antibody Haptens small
 The immune response Antibodies Antigens Epitopes (antigenic determinants) the part of a protein antigen recognized by an antibody Haptens small molecules that can elicit an immune response when linked
The immune response Antibodies Antigens Epitopes (antigenic determinants) the part of a protein antigen recognized by an antibody Haptens small molecules that can elicit an immune response when linked
Application Guide... 2
 Protocol for GenomePlex Whole Genome Amplification from Formalin-Fixed Parrafin-Embedded (FFPE) tissue Application Guide... 2 I. Description... 2 II. Product Components... 2 III. Materials to be Supplied
Protocol for GenomePlex Whole Genome Amplification from Formalin-Fixed Parrafin-Embedded (FFPE) tissue Application Guide... 2 I. Description... 2 II. Product Components... 2 III. Materials to be Supplied
Aviva Systems Biology
 Aviva Custom Antibody Service and Price Mouse Monoclonal Antibody Service Package Number Description Package Contents Time Price Customer provides antigen protein $6,174 Monoclonal package1 (From protein
Aviva Custom Antibody Service and Price Mouse Monoclonal Antibody Service Package Number Description Package Contents Time Price Customer provides antigen protein $6,174 Monoclonal package1 (From protein
Real-time PCR: Understanding C t
 APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
RT-PCR: Two-Step Protocol
 RT-PCR: Two-Step Protocol We will provide both one-step and two-step protocols for RT-PCR. We recommend the twostep protocol for this class. In the one-step protocol, the components of RT and PCR are mixed
RT-PCR: Two-Step Protocol We will provide both one-step and two-step protocols for RT-PCR. We recommend the twostep protocol for this class. In the one-step protocol, the components of RT and PCR are mixed
Chapter 2 Antibodies. Contents. Introduction
 Chapter 2 Antibodies Keywords Immunohistochemistry Antibody labeling Fluorescence microscopy Fluorescent immunocytochemistry Fluorescent immunohistochemistry Indirect immunocytochemistry Immunostaining
Chapter 2 Antibodies Keywords Immunohistochemistry Antibody labeling Fluorescence microscopy Fluorescent immunocytochemistry Fluorescent immunohistochemistry Indirect immunocytochemistry Immunostaining
Chapter 18: Applications of Immunology
 Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
CD3/TCR stimulation and surface detection Determination of specificity of intracellular detection of IL-7Rα by flow cytometry
 CD3/TCR stimulation and surface detection Stimulation of HPB-ALL cells with the anti-cd3 monoclonal antibody OKT3 was performed as described 3. In brief, antibody-coated plates were prepared by incubating
CD3/TCR stimulation and surface detection Stimulation of HPB-ALL cells with the anti-cd3 monoclonal antibody OKT3 was performed as described 3. In brief, antibody-coated plates were prepared by incubating
ONLINE SUPPLEMENTAL MATERIAL. Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue
 ONLINE SUPPLEMENTAL MATERIAL Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue Sungmi Park 1, Ko-Ting Lu 1, Xuebo Liu 1, Tapan K. Chatterjee 2, Steven M. Rudich 3, Neal
ONLINE SUPPLEMENTAL MATERIAL Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue Sungmi Park 1, Ko-Ting Lu 1, Xuebo Liu 1, Tapan K. Chatterjee 2, Steven M. Rudich 3, Neal
Technical Note. Roche Applied Science. No. LC 18/2004. Assay Formats for Use in Real-Time PCR
 Roche Applied Science Technical Note No. LC 18/2004 Purpose of this Note Assay Formats for Use in Real-Time PCR The LightCycler Instrument uses several detection channels to monitor the amplification of
Roche Applied Science Technical Note No. LC 18/2004 Purpose of this Note Assay Formats for Use in Real-Time PCR The LightCycler Instrument uses several detection channels to monitor the amplification of
Appendix 2 Molecular Biology Core Curriculum. Websites and Other Resources
 Appendix 2 Molecular Biology Core Curriculum Websites and Other Resources Chapter 1 - The Molecular Basis of Cancer 1. Inside Cancer http://www.insidecancer.org/ From the Dolan DNA Learning Center Cold
Appendix 2 Molecular Biology Core Curriculum Websites and Other Resources Chapter 1 - The Molecular Basis of Cancer 1. Inside Cancer http://www.insidecancer.org/ From the Dolan DNA Learning Center Cold
PROTOCOL. Immunostaining for Flow Cytometry. Background. Materials and equipment required.
 PROTOCOL Immunostaining for Flow Cytometry 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 Rev.0 Background The combination of single cell analysis using flow cytometry and the specificity of antibody-based
PROTOCOL Immunostaining for Flow Cytometry 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 Rev.0 Background The combination of single cell analysis using flow cytometry and the specificity of antibody-based
Mir-X mirna First-Strand Synthesis Kit User Manual
 User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
Recombinant DNA and Biotechnology
 Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Notch 1 -dependent regulation of cell fate in colorectal cancer
 Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
pcas-guide System Validation in Genome Editing
 pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
Chapter 6. Antigen-Antibody Properties 10/3/2012. Antigen-Antibody Interactions: Principles and Applications. Precipitin reactions
 Chapter 6 Antigen-Antibody Interactions: Principles and Applications Antigen-Antibody Properties You must remember antibody affinity (single) VS avidity (multiple) High affinity: bound tightly and longer!
Chapter 6 Antigen-Antibody Interactions: Principles and Applications Antigen-Antibody Properties You must remember antibody affinity (single) VS avidity (multiple) High affinity: bound tightly and longer!
ELISA BIO 110 Lab 1. Immunity and Disease
 ELISA BIO 110 Lab 1 Immunity and Disease Introduction The principal role of the mammalian immune response is to contain infectious disease agents. This response is mediated by several cellular and molecular
ELISA BIO 110 Lab 1 Immunity and Disease Introduction The principal role of the mammalian immune response is to contain infectious disease agents. This response is mediated by several cellular and molecular
Pure-IP Western Blot Detection Kit
 Product Manual Pure-IP Western Blot Detection Kit Catalog Number PRB-5002 20 blots FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction The technique of immunoprecipitation (IP) is used
Product Manual Pure-IP Western Blot Detection Kit Catalog Number PRB-5002 20 blots FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction The technique of immunoprecipitation (IP) is used
Nucleic Acid Techniques in Bacterial Systematics
 Nucleic Acid Techniques in Bacterial Systematics Edited by Erko Stackebrandt Department of Microbiology University of Queensland St Lucia, Australia and Michael Goodfellow Department of Microbiology University
Nucleic Acid Techniques in Bacterial Systematics Edited by Erko Stackebrandt Department of Microbiology University of Queensland St Lucia, Australia and Michael Goodfellow Department of Microbiology University
QUANTITATIVE RT-PCR. A = B (1+e) n. A=amplified products, B=input templates, n=cycle number, and e=amplification efficiency.
 QUANTITATIVE RT-PCR Application: Quantitative RT-PCR is used to quantify mrna in both relative and absolute terms. It can be applied for the quantification of mrna expressed from endogenous genes, and
QUANTITATIVE RT-PCR Application: Quantitative RT-PCR is used to quantify mrna in both relative and absolute terms. It can be applied for the quantification of mrna expressed from endogenous genes, and
sirna Duplexes & RNAi Explorer
 P r o d u c t S p e c i f i c a t i o n s Guaranteed RNAi Explorer kit with Fl/Dabcyl Molecular Beacon Catalog No.:27-6402-01 Guaranteed RNAi Explorer kit with Fluorescein/Tamra TaqMan Catalog No.:27-6402-01
P r o d u c t S p e c i f i c a t i o n s Guaranteed RNAi Explorer kit with Fl/Dabcyl Molecular Beacon Catalog No.:27-6402-01 Guaranteed RNAi Explorer kit with Fluorescein/Tamra TaqMan Catalog No.:27-6402-01
DNA Fingerprinting. Unless they are identical twins, individuals have unique DNA
 DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
Outline. 1. Experiment. 2. Sample analysis and storage. 3. Image analysis and presenting data. 4. Probemaker
 Tips and tricks Note: this is just an informative document with general recommendations. Please contact support@olink.com should you have any queries. Document last reviewed 2011-11-17 Outline 1. Experiment
Tips and tricks Note: this is just an informative document with general recommendations. Please contact support@olink.com should you have any queries. Document last reviewed 2011-11-17 Outline 1. Experiment
Recombinant DNA & Genetic Engineering. Tools for Genetic Manipulation
 Recombinant DNA & Genetic Engineering g Genetic Manipulation: Tools Kathleen Hill Associate Professor Department of Biology The University of Western Ontario Tools for Genetic Manipulation DNA, RNA, cdna
Recombinant DNA & Genetic Engineering g Genetic Manipulation: Tools Kathleen Hill Associate Professor Department of Biology The University of Western Ontario Tools for Genetic Manipulation DNA, RNA, cdna
Protein Analysis. -Detection and quantification. Toby M Holmes
 Protein Analysis -Detection and quantification Toby M Holmes Clinical Research Unit UCD school of Medicine and Medical Sciences Mater Misericordiae University Hospital Dublin Protein structure General
Protein Analysis -Detection and quantification Toby M Holmes Clinical Research Unit UCD school of Medicine and Medical Sciences Mater Misericordiae University Hospital Dublin Protein structure General
Chapter 6: Antigen-Antibody Interactions
 Chapter 6: Antigen-Antibody Interactions I. Strength of Ag-Ab interactions A. Antibody Affinity - strength of total noncovalent interactions between single Ag-binding site on an Ab and a single epitope
Chapter 6: Antigen-Antibody Interactions I. Strength of Ag-Ab interactions A. Antibody Affinity - strength of total noncovalent interactions between single Ag-binding site on an Ab and a single epitope
No-wash, no-lyse detection of leukocytes in human whole blood on the Attune NxT Flow Cytometer
 APPLICATION NOTE Attune NxT Flow Cytometer No-wash, no-lyse detection of leukocytes in human whole blood on the Attune NxT Flow Cytometer Introduction Standard methods for isolating and detecting leukocytes
APPLICATION NOTE Attune NxT Flow Cytometer No-wash, no-lyse detection of leukocytes in human whole blood on the Attune NxT Flow Cytometer Introduction Standard methods for isolating and detecting leukocytes
DNA Sequence Analysis
 DNA Sequence Analysis Two general kinds of analysis Screen for one of a set of known sequences Determine the sequence even if it is novel Screening for a known sequence usually involves an oligonucleotide
DNA Sequence Analysis Two general kinds of analysis Screen for one of a set of known sequences Determine the sequence even if it is novel Screening for a known sequence usually involves an oligonucleotide
Understanding the immune response to bacterial infections
 Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Quantitative Real Time PCR Protocol. Stack Lab
 Quantitative Real Time PCR Protocol Stack Lab Overview Real-time quantitative polymerase chain reaction (qpcr) differs from regular PCR by including in the reaction fluorescent reporter molecules that
Quantitative Real Time PCR Protocol Stack Lab Overview Real-time quantitative polymerase chain reaction (qpcr) differs from regular PCR by including in the reaction fluorescent reporter molecules that
Biotechnology: DNA Technology & Genomics
 Chapter 20. Biotechnology: DNA Technology & Genomics 2003-2004 The BIG Questions How can we use our knowledge of DNA to: diagnose disease or defect? cure disease or defect? change/improve organisms? What
Chapter 20. Biotechnology: DNA Technology & Genomics 2003-2004 The BIG Questions How can we use our knowledge of DNA to: diagnose disease or defect? cure disease or defect? change/improve organisms? What
Western Blot Analysis with Cell Samples Grown in Channel-µ-Slides
 Western Blot Analysis with Cell Samples Grown in Channel-µ-Slides Polyacrylamide gel electrophoresis (PAGE) and subsequent analyses are common tools in biochemistry and molecular biology. This Application
Western Blot Analysis with Cell Samples Grown in Channel-µ-Slides Polyacrylamide gel electrophoresis (PAGE) and subsequent analyses are common tools in biochemistry and molecular biology. This Application
Beginner s Guide to Real-Time PCR
 Beginner s Guide to Real-Time PCR 02 Real-time PCR basic principles PCR or the Polymerase Chain Reaction has become the cornerstone of modern molecular biology the world over. Real-time PCR is an advanced
Beginner s Guide to Real-Time PCR 02 Real-time PCR basic principles PCR or the Polymerase Chain Reaction has become the cornerstone of modern molecular biology the world over. Real-time PCR is an advanced
SYBR Green Realtime PCR Master Mix -Plus-
 Instruction manual SYBR Green Realtime PCR Master Mix -Plus- 0810 F0925K SYBR Green Realtime PCR Master Mix -Plus- Contents QPK-212T 1mLx1 QPK-212 1mLx5 Store at -20 C, protected from light [1] Introduction
Instruction manual SYBR Green Realtime PCR Master Mix -Plus- 0810 F0925K SYBR Green Realtime PCR Master Mix -Plus- Contents QPK-212T 1mLx1 QPK-212 1mLx5 Store at -20 C, protected from light [1] Introduction
VLLM0421c Medical Microbiology I, practical sessions. Protocol to topic J10
 Topic J10+11: Molecular-biological methods + Clinical virology I (hepatitis A, B & C, HIV) To study: PCR, ELISA, your own notes from serology reactions Task J10/1: DNA isolation of the etiological agent
Topic J10+11: Molecular-biological methods + Clinical virology I (hepatitis A, B & C, HIV) To study: PCR, ELISA, your own notes from serology reactions Task J10/1: DNA isolation of the etiological agent
ChIP TROUBLESHOOTING TIPS
 ChIP TROUBLESHOOTING TIPS Creative Diagnostics Abstract ChIP dissects the spatial and temporal dynamics of the interactions between chromatin and its associated factors CD Creative Diagnostics info@creative-
ChIP TROUBLESHOOTING TIPS Creative Diagnostics Abstract ChIP dissects the spatial and temporal dynamics of the interactions between chromatin and its associated factors CD Creative Diagnostics info@creative-
Your partner in immunology
 Your partner in immunology Expertise Expertise Reactivity Reactivity Quality Quality Advice Advice Who are we? Specialist of antibody engineering Covalab is a French biotechnology company, specialised
Your partner in immunology Expertise Expertise Reactivity Reactivity Quality Quality Advice Advice Who are we? Specialist of antibody engineering Covalab is a French biotechnology company, specialised
BacReady TM Multiplex PCR System
 BacReady TM Multiplex PCR System Technical Manual No. 0191 Version 10112010 I Description.. 1 II Applications 2 III Key Features.. 2 IV Shipping and Storage. 2 V Simplified Procedures. 2 VI Detailed Experimental
BacReady TM Multiplex PCR System Technical Manual No. 0191 Version 10112010 I Description.. 1 II Applications 2 III Key Features.. 2 IV Shipping and Storage. 2 V Simplified Procedures. 2 VI Detailed Experimental
PyroPhage 3173 DNA Polymerase, Exonuclease Minus (Exo-)
 PyroPhage 3173 DNA Polymerase, Exonuclease Minus (Exo-) FOR RESEARCH USE ONLY. NOT FOR HUMAN OR DIAGNOSTIC USE Lucigen Corporation 2905 Parmenter St, Middleton, WI 53562 USA Toll Free: (888) 575-9695 (608)
PyroPhage 3173 DNA Polymerase, Exonuclease Minus (Exo-) FOR RESEARCH USE ONLY. NOT FOR HUMAN OR DIAGNOSTIC USE Lucigen Corporation 2905 Parmenter St, Middleton, WI 53562 USA Toll Free: (888) 575-9695 (608)
PROTOCOL. Immunocytochemistry (ICC) MATERIALS AND EQUIPMENT REQUIRED
 PROTOCOL Immunocytochemistry (ICC) 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 11-07 MATERIALS AND EQUIPMENT REQUIRED Materials: MitoSciences primary monoclonal antibody/antibodies Fluorophore-conjugated
PROTOCOL Immunocytochemistry (ICC) 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 11-07 MATERIALS AND EQUIPMENT REQUIRED Materials: MitoSciences primary monoclonal antibody/antibodies Fluorophore-conjugated
How To Get A Cell Print
 QUICK CELL CAPTURE AND CHARACTERIZATION GUIDE FOR CELLSEARCH CUSTOMERS CellSave EDTA Blood sample Rare cell capture Enumeration Single protein marker Cell capture for molecular characterization CELLSEARCH
QUICK CELL CAPTURE AND CHARACTERIZATION GUIDE FOR CELLSEARCH CUSTOMERS CellSave EDTA Blood sample Rare cell capture Enumeration Single protein marker Cell capture for molecular characterization CELLSEARCH
SUPPLEMENTARY DATA 1
 SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
Reverse Transcription System
 TECHNICAL BULLETIN Reverse Transcription System Instruc ons for use of Product A3500 Revised 1/14 TB099 Reverse Transcription System All technical literature is available on the Internet at: www.promega.com/protocols/
TECHNICAL BULLETIN Reverse Transcription System Instruc ons for use of Product A3500 Revised 1/14 TB099 Reverse Transcription System All technical literature is available on the Internet at: www.promega.com/protocols/
HuCAL Custom Monoclonal Antibodies
 HuCAL Custom Monoclonal HuCAL Custom Monoclonal Antibodies Highly Specific, Recombinant Antibodies in 8 Weeks Highly Specific Monoclonal Antibodies in Just 8 Weeks HuCAL PLATINUM (Human Combinatorial Antibody
HuCAL Custom Monoclonal HuCAL Custom Monoclonal Antibodies Highly Specific, Recombinant Antibodies in 8 Weeks Highly Specific Monoclonal Antibodies in Just 8 Weeks HuCAL PLATINUM (Human Combinatorial Antibody
Pharmaceutical Biotechnology. Recombinant DNA technology Western blotting and SDS-PAGE
 Pharmaceutical Biotechnology Recombinant DNA technology Western blotting and SDS-PAGE Recombinant DNA Technology Protein Synthesis Western Blot Western blots allow investigators to determine the molecular
Pharmaceutical Biotechnology Recombinant DNA technology Western blotting and SDS-PAGE Recombinant DNA Technology Protein Synthesis Western Blot Western blots allow investigators to determine the molecular
All-in-One mirna qrt-pcr Reagent Kits For quantitative detection of mature mirna
 All-in-One mirna qrt-pcr Reagent Kits For quantitative detection of mature mirna All-in-One TM mirna First-Strand cdna Synthesis Kit AMRT-0020 (20 RT reactions), AMRT-0060 (60 RT reactions) Used in combination
All-in-One mirna qrt-pcr Reagent Kits For quantitative detection of mature mirna All-in-One TM mirna First-Strand cdna Synthesis Kit AMRT-0020 (20 RT reactions), AMRT-0060 (60 RT reactions) Used in combination
Anti-ATF6 α antibody, mouse monoclonal (1-7)
 Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
Immunophenotyping peripheral blood cells
 IMMUNOPHENOTYPING Attune Accoustic Focusing Cytometer Immunophenotyping peripheral blood cells A no-lyse, no-wash, no cell loss method for immunophenotyping nucleated peripheral blood cells using the Attune
IMMUNOPHENOTYPING Attune Accoustic Focusing Cytometer Immunophenotyping peripheral blood cells A no-lyse, no-wash, no cell loss method for immunophenotyping nucleated peripheral blood cells using the Attune
Global MicroRNA Amplification Kit
 Global MicroRNA Amplification Kit Store kit at -20 C on receipt (ver. 3-060901) A limited-use label license covers this product. By use of this product, you accept the terms and conditions outlined in
Global MicroRNA Amplification Kit Store kit at -20 C on receipt (ver. 3-060901) A limited-use label license covers this product. By use of this product, you accept the terms and conditions outlined in
Introduction to flow cytometry
 Introduction to flow cytometry Flow cytometry is a popular laser-based technology. Discover more with our introduction to flow cytometry. Flow cytometry is now a widely used method for analyzing the expression
Introduction to flow cytometry Flow cytometry is a popular laser-based technology. Discover more with our introduction to flow cytometry. Flow cytometry is now a widely used method for analyzing the expression
Title: Mapping T cell epitopes in PCV2 capsid protein - NPB #08-159. Date Submitted: 12-11-09
 Title: Mapping T cell epitopes in PCV2 capsid protein - NPB #08-159 Investigator: Institution: Carol Wyatt Kansas State University Date Submitted: 12-11-09 Industry summary: Effective circovirus vaccines
Title: Mapping T cell epitopes in PCV2 capsid protein - NPB #08-159 Investigator: Institution: Carol Wyatt Kansas State University Date Submitted: 12-11-09 Industry summary: Effective circovirus vaccines
Superior TrueMAB TM monoclonal antibodies for the recognition of proteins native epitopes
 Superior TrueMAB TM monoclonal antibodies for the recognition of proteins native epitopes Outlines Brief introduction of OriGene s mission on gene-centric product solution. TrueMAB monoclonal antibody
Superior TrueMAB TM monoclonal antibodies for the recognition of proteins native epitopes Outlines Brief introduction of OriGene s mission on gene-centric product solution. TrueMAB monoclonal antibody
ab185916 Hi-Fi cdna Synthesis Kit
 ab185916 Hi-Fi cdna Synthesis Kit Instructions for Use For cdna synthesis from various RNA samples This product is for research use only and is not intended for diagnostic use. Version 1 Last Updated 1
ab185916 Hi-Fi cdna Synthesis Kit Instructions for Use For cdna synthesis from various RNA samples This product is for research use only and is not intended for diagnostic use. Version 1 Last Updated 1
RPCI 004 v.002 Staining Procedure For all Directly Conjugated Reagents (Whole Blood Method)
 Immune Tolerance Network RPCI 004 v.002 Staining Procedure For all Directly Conjugated Reagents (Whole Blood Method) Author: Paul Wallace, Director, RPCI Laboratory of Flow Cytometry Approved by: Paul
Immune Tolerance Network RPCI 004 v.002 Staining Procedure For all Directly Conjugated Reagents (Whole Blood Method) Author: Paul Wallace, Director, RPCI Laboratory of Flow Cytometry Approved by: Paul
How To Make A Tri Reagent
 TRI Reagent For processing tissues, cells cultured in monolayer or cell pellets Catalog Number T9424 Store at room temperature. TECHNICAL BULLETIN Product Description TRI Reagent is a quick and convenient
TRI Reagent For processing tissues, cells cultured in monolayer or cell pellets Catalog Number T9424 Store at room temperature. TECHNICAL BULLETIN Product Description TRI Reagent is a quick and convenient
MAB Solut. MABSolys Génopole Campus 1 5 rue Henri Desbruères 91030 Evry Cedex. www.mabsolut.com. is involved at each stage of your project
 Mabsolus-2015-UK:Mise en page 1 03/07/15 14:13 Page1 Services provider Department of MABSolys from conception to validation MAB Solut is involved at each stage of your project Creation of antibodies Production
Mabsolus-2015-UK:Mise en page 1 03/07/15 14:13 Page1 Services provider Department of MABSolys from conception to validation MAB Solut is involved at each stage of your project Creation of antibodies Production
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
Design of conditional gene targeting vectors - a recombineering approach
 Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
RNA Viruses. A Practical Approac h. Alan J. Cann
 RNA Viruses A Practical Approac h Alan J. Cann List of protocols page xiii Abbreviations xvii Investigation of RNA virus genome structure 1 A j. Easton, A.C. Marriott and C.R. Pringl e 1 Introduction-the
RNA Viruses A Practical Approac h Alan J. Cann List of protocols page xiii Abbreviations xvii Investigation of RNA virus genome structure 1 A j. Easton, A.C. Marriott and C.R. Pringl e 1 Introduction-the
Western BLoT Immuno Booster
 Cat. # T7111A For Research Use Western BLoT Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 3 IV. Materials Required but Not Provided... 3 V. Precautions... 3 VI.
Cat. # T7111A For Research Use Western BLoT Product Manual Table of Contents I. Description... 3 II. Components... 3 III. Storage... 3 IV. Materials Required but Not Provided... 3 V. Precautions... 3 VI.
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College
 Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
1.Gene Synthesis. 2.Peptide & Phospho-P. Assembly PCR. Design & Synthesis. Advantages. Specifications. Advantages
 1.Gene Synthesis Assembly PCR Looking for a cdna for your research but could not fish out the gene through traditional cloning methods or a supplier? Abnova provides a gene synthesis service via assembly
1.Gene Synthesis Assembly PCR Looking for a cdna for your research but could not fish out the gene through traditional cloning methods or a supplier? Abnova provides a gene synthesis service via assembly
PicoMaxx High Fidelity PCR System
 PicoMaxx High Fidelity PCR System Instruction Manual Catalog #600420 (100 U), #600422 (500 U), and #600424 (1000 U) Revision C Research Use Only. Not for Use in Diagnostic Procedures. 600420-12 LIMITED
PicoMaxx High Fidelity PCR System Instruction Manual Catalog #600420 (100 U), #600422 (500 U), and #600424 (1000 U) Revision C Research Use Only. Not for Use in Diagnostic Procedures. 600420-12 LIMITED
Microarray Technology
 Microarrays And Functional Genomics CPSC265 Matt Hudson Microarray Technology Relatively young technology Usually used like a Northern blot can determine the amount of mrna for a particular gene Except
Microarrays And Functional Genomics CPSC265 Matt Hudson Microarray Technology Relatively young technology Usually used like a Northern blot can determine the amount of mrna for a particular gene Except
Table of Contents. I. Description... 2. II. Kit Components... 2. III. Storage... 2. IV. 1st Strand cdna Synthesis Reaction... 3
 Table of Contents I. Description... 2 II. Kit Components... 2 III. Storage... 2 IV. 1st Strand cdna Synthesis Reaction... 3 V. RT-PCR, Real-time RT-PCR... 4 VI. Application... 5 VII. Preparation of RNA
Table of Contents I. Description... 2 II. Kit Components... 2 III. Storage... 2 IV. 1st Strand cdna Synthesis Reaction... 3 V. RT-PCR, Real-time RT-PCR... 4 VI. Application... 5 VII. Preparation of RNA
PrimeSTAR HS DNA Polymerase
 Cat. # R010A For Research Use PrimeSTAR HS DNA Polymerase Product Manual Table of Contents I. Description...3 II. III. IV. Components...3 Storage...3 Features...3 V. General Composition of PCR Reaction
Cat. # R010A For Research Use PrimeSTAR HS DNA Polymerase Product Manual Table of Contents I. Description...3 II. III. IV. Components...3 Storage...3 Features...3 V. General Composition of PCR Reaction
Cloning Blunt-End Pfu DNA Polymerase- Generated PCR Fragments into pgem -T Vector Systems
 Promega Notes Number 71, 1999, p. 10 Blunt-End Pfu DNA Polymerase- Generated PCR Fragments into pgem -T Vector Systems By Kimberly Knoche, Ph.D., and Dan Kephart, Ph.D. Promega Corporation Corresponding
Promega Notes Number 71, 1999, p. 10 Blunt-End Pfu DNA Polymerase- Generated PCR Fragments into pgem -T Vector Systems By Kimberly Knoche, Ph.D., and Dan Kephart, Ph.D. Promega Corporation Corresponding
Chapter 3 Contd. Western blotting & SDS PAGE
 Chapter 3 Contd. Western blotting & SDS PAGE Western Blot Western blots allow investigators to determine the molecular weight of a protein and to measure relative amounts of the protein present in different
Chapter 3 Contd. Western blotting & SDS PAGE Western Blot Western blots allow investigators to determine the molecular weight of a protein and to measure relative amounts of the protein present in different
GenScript BloodReady TM Multiplex PCR System
 GenScript BloodReady TM Multiplex PCR System Technical Manual No. 0174 Version 20040915 I Description.. 1 II Applications 2 III Key Features.. 2 IV Shipping and Storage. 2 V Simplified Procedures. 2 VI
GenScript BloodReady TM Multiplex PCR System Technical Manual No. 0174 Version 20040915 I Description.. 1 II Applications 2 III Key Features.. 2 IV Shipping and Storage. 2 V Simplified Procedures. 2 VI
Using Digital Photography to Supplement Learning of Biotechnology. Methods
 RESEARCH ON LEARNING Using Digital Photography to Supplement Learning of Biotechnology Fran n orf l u s AbstrAct The author used digital photography to supplement learning of biotechnology by students
RESEARCH ON LEARNING Using Digital Photography to Supplement Learning of Biotechnology Fran n orf l u s AbstrAct The author used digital photography to supplement learning of biotechnology by students
TaqMan Fast Advanced Master Mix. Protocol
 TaqMan Fast Advanced Master Mix Protocol For Research Use Only. Not intended for any animal or human therapeutic or diagnostic use. Information in this document is subject to change without notice. APPLIED
TaqMan Fast Advanced Master Mix Protocol For Research Use Only. Not intended for any animal or human therapeutic or diagnostic use. Information in this document is subject to change without notice. APPLIED
Supplementary Figure 1.
 Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Identification of the VTEC serogroups mainly associated with human infections by conventional PCR amplification of O-associated genes
 Identification of the VTEC serogroups mainly associated with human infections by conventional PCR amplification of O-associated genes 1. Aim and field of application The present method concerns the identification
Identification of the VTEC serogroups mainly associated with human infections by conventional PCR amplification of O-associated genes 1. Aim and field of application The present method concerns the identification
User Manual/Hand book. qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse)
 User Manual/Hand book qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse) Kit Components Cat. No. MA003...Human Whole Genome mirna qpcr Profiling Kit (-20 C) The following components are sufficient
User Manual/Hand book qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse) Kit Components Cat. No. MA003...Human Whole Genome mirna qpcr Profiling Kit (-20 C) The following components are sufficient
CompleteⅡ 1st strand cdna Synthesis Kit
 Instruction Manual CompleteⅡ 1st strand cdna Synthesis Kit Catalog # GM30401, GM30402 Green Mountain Biosystems. LLC Web: www.greenmountainbio.com Tel: 800-942-1160 Sales: Sales@ greenmountainbio.com Support:
Instruction Manual CompleteⅡ 1st strand cdna Synthesis Kit Catalog # GM30401, GM30402 Green Mountain Biosystems. LLC Web: www.greenmountainbio.com Tel: 800-942-1160 Sales: Sales@ greenmountainbio.com Support:
Cloning GFP into Mammalian cells
 Protocol for Cloning GFP into Mammalian cells Studiepraktik 2013 Molecular Biology and Molecular Medicine Aarhus University Produced by the instructors: Tobias Holm Bønnelykke, Rikke Mouridsen, Steffan
Protocol for Cloning GFP into Mammalian cells Studiepraktik 2013 Molecular Biology and Molecular Medicine Aarhus University Produced by the instructors: Tobias Holm Bønnelykke, Rikke Mouridsen, Steffan
Real-time quantitative RT -PCR (Taqman)
 Real-time quantitative RT -PCR (Taqman) Author: SC, Patti Lab, 3/03 This is performed as a 2-step reaction: 1. cdna synthesis from DNase 1-treated total RNA 2. PCR 1. cdna synthesis (Advantage RT-for-PCR
Real-time quantitative RT -PCR (Taqman) Author: SC, Patti Lab, 3/03 This is performed as a 2-step reaction: 1. cdna synthesis from DNase 1-treated total RNA 2. PCR 1. cdna synthesis (Advantage RT-for-PCR
Analysis of gene expression data. Ulf Leser and Philippe Thomas
 Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
Hypoxyprobe -1 Plus Kit Kit contents:
 Updated 2015 1 PRODUCT INSERT Hypoxyprobe, Inc 121 Middlesex Turnpike Burlington, MA 01803 USA www.hypoxyprobe.com Hypoxyprobe -1 Plus Kit Kit contents: Solid pimonidazole HCl (Hypoxyprobe -1) FITC conjugated
Updated 2015 1 PRODUCT INSERT Hypoxyprobe, Inc 121 Middlesex Turnpike Burlington, MA 01803 USA www.hypoxyprobe.com Hypoxyprobe -1 Plus Kit Kit contents: Solid pimonidazole HCl (Hypoxyprobe -1) FITC conjugated
EdU Flow Cytometry Kit. User Manual
 User Manual Ordering information: (for detailed kit content see Table 2) EdU Flow Cytometry Kits for 50 assays: Product number EdU Used fluorescent dye BCK-FC488-50 10 mg 6-FAM Azide BCK-FC555-50 10 mg
User Manual Ordering information: (for detailed kit content see Table 2) EdU Flow Cytometry Kits for 50 assays: Product number EdU Used fluorescent dye BCK-FC488-50 10 mg 6-FAM Azide BCK-FC555-50 10 mg
