Ribosomal protein 6S mrna is a biomarker upregulated in multiple sclerosis, downregulated by interferon treatment, and affected by season
|
|
|
- Adam Garrett
- 5 years ago
- Views:
Transcription
1 Ribosomal protein 6S mrna is a biomarker upregulated in multiple sclerosis, downregulated by interferon treatment, and affected by season Grant P. Parnell 1, Prudence N. Gatt 1, Fiona C. McKay 1, Stephen Schibeci 1, Malgorzata Krupa 2, Joseph E. Powell 3,4, Peter M. Visscher 3,4, Grant W. Montgomery 4, Jeannette Lechner-Scott 5, Simon Broadley 6, Christopher Liddle 1, Mark Slee 2, Steve Vucic 7, Graeme J. Stewart 1, David R. Booth 1 1 Westmead Millennium Institute, University of Sydney, Sydney, New South Wales, 2145, Australia 2 School of Medicine, Flinders University of South Australia, South Australia, 5042, Australia 3 University of Queensland Diamantina Institute, Translational Research Institute, Brisbane, Queensland, 4102, Australia 4 The Queensland Brain Institute, The University of Queensland, Brisbane, Queensland, 4072, Australia 5 University of Newcastle, New South Wales, 2300, Australia 6 Griffith University, Gold Coast, Queensland, 4222, Australia 7 Westmead Clinical School, University of Sydney, Westmead Hospital, Sydney, New South Wales, 2145, Australia Running Title: RPS6 is dysregulated in MS Keywords: Multiple sclerosis, biomarkers, RPS6, season, interferon beta Corresponding Author: A/Prof David Booth, Darcy Rd., Westmead Millennium Institute, University of Sydney, Sydney, Australia, david.booth@sydney.edu.au, ph fax Abstract Background: Multiple Sclerosis (MS) is an immune-mediated disease of the central nervous system which responds to therapies targeting circulating immune cells. 1
2 Objective: To test if the T cell activation gene expression pattern (TCAGE) we had previously described from whole blood was replicated in an independent cohort. Methods: We used RNA-seq to interrogate the whole blood transcriptomes of 72 individuals (40 healthy control, 32 untreated MS). A cohort of 862 control individuals from the Brisbane Systems Genetics Study (BSGS) was used to assess heritability and seasonal expression. The effect of interferon beta (IFNB) therapy on expression was evaluated. Results: The MS/TCAGE association was replicated and rationalised to a single marker, ribosomal protein S6 (RPS6). Expression of RPS6 was higher in MS than controls (p<0.0004), and lower in winter than summer (p<4.6e-06). The seasonal pattern correlated with monthly UV light index (R=0.82, p<0.002), and was also identified in the BSGS cohort (p<0.0016). Variation in expression of RPS6 was not strongly heritable. RPS6 expression was reduced by IFNB therapy. Conclusions: These data support investigation of RPS6 as a potential therapeutic target and candidate biomarker for measuring clinical response to IFNB and other MS therapies, and of MS disease heterogeneity. 2
3 Introduction MS is an autoimmune disease of the central nervous system caused, at least in part, by pathogenic leukocytes which transit from lymphoid tissue to the brain via the peripheral circulation. 1 This paradigm is supported by the success of immunomodulatory therapies such as Natalizumab, which prevents activated T cell migration across the blood-brain barrier from the peripheral circulation 2 ; and of Fingolimod, which blocks egress of activated T cells from the lymph nodes. 3 It is also supported by studies on genetic susceptibility 4, since the majority of identified genes associated with MS are predominantly expressed in immune cells, especially the dendritic cell/ T cell axis. 4 Monitoring changes in the numbers and states of these cells may identify pathogenic processes, useful biomarkers of susceptibility, molecular subtypes of diseases, response to therapy, and new therapeutic targets. Earlier, we had reported mrna expression for all known genes in whole blood from 144 individuals, 99 of whom had untreated MS. 5 We had used PAXgene blood collections, which capture transcriptomes immediately on blood draw, rather than peripheral blood mononuclear cell (PBMCs) methods, which are less reproducible. 6 Analysis of gene expression and genotype indicated the dominant pattern of dysregulation of T cell gene expression in MS in these individuals is matched with the genetic associations from the GWAS 4 pointing to T cell dysregulation in MS. We identified a T cell activation gene expression (TCAGE) signature in whole blood that predicted the likelihood of MS. This signature reflects excess expression of genes upregulated on T cell activation and proliferation, mainly from translation and oxidative phosphorylation metabolic 3
4 pathways. 5 This TCAGE signature needed to be confirmed in an independent cohort, and potentially simplified. The effect of therapy on the TCAGE is currently unknown, and the heritability of this signature is also not established. In this study, we have first tested if next generation sequencing of messenger RNA (RNA-seq) could be used to interrogate gene expression in whole blood samples by comparing expression measured by both techniques (RNA-seq and microarrays) for the same samples. We then used RNA-seq to assay whole blood transcriptomes for 40 healthy controls and 32 people with untreated MS. The TCAGE signature could be rationalized to a single gene, ribosomal protein S6 (RPS6). We have tested if RPS6 expression is affected by genotype. Finally, interferon beta (IFNB thought to function by immunomodulation. 7 We have tested if RPS6 expression is altered by therapy. 4
5 Materials and Methods Sample Collection RNA-seq Cohort: For the RNA-seq experiment, a single PAXgene blood RNA tube (PreAnalytiX, Switzerland) was collected from people with MS who were not receiving any treatment, and had not received any immunomodulatory therapy in the previous three months, from clinics in Sydney and Adelaide. Blood was also collected from healthy controls of matching ages and not receiving immunomodulatory therapy. Demographics for patients and controls are described in Table 1. Brisbane Systems Genetics Study (BSGS) cohort: This cohort comprises 862 individuals of European descent from 312 independent families 8 located within the greater Brisbane region. For each cohort informed written consent was obtained from each donor, and the study was approved by the Human Research Ethics Committee (HREC) from Sydney, Adelaide and Brisbane. IFNB therapy cohort: of anonymised donors who were identified as MxA bioactivity positive or negative by bioactivity testing. 9 Approval to use these samples was also granted by SWAHS HRECS. RNA-seq Whole blood: Total RNA was isolated from PAXgene RNA blood tubes using the PAXgene Blood RNA Kit (Qiagen, Germany), and subsequently treated with Globinclear (Life Technologies, NY, USA), to deplete the whole blood sample of alpha and beta globin mrna. RNA was then prepared for sequencing on Illumina 5
6 HiSeq 2000 using the Illumina TruSeq RNA sample preparation kit V1 (Illumina, CA, USA). Samples were indexed and 6 run per lane, generating at least 10 million 50bp reads per sample. Raw sequence data was aligned to the UCSC human reference genome (hg19) using the Tophat software package. 10 Aligned sequencing reads were summarised to counts per gene using the Read Assignment via Expectation Maximiation (RAEM) procedure 11 and reads per kilobase per million mapped reads (RPKM) values calculated in SAMMate (v 2.6.1). RPKM values were transformed by quantile normalisation prior to visualization using BRB-Array Tools. 12 Visualisation was performed by performing hierarchical clustering using 1 minus correlation and average linkage metrics. The microarray gene expression analysis has been previously described in 5 for the ANZgene cohort, and in 13 for the Brisbane Systems Genetic Study. Quantitative RTPCR Primers were designed to amplify across the last two exons of RPS6 - RPS6F:GCGGCGTATTGCTCTGAAGA, RPS6R: GCAGAGAGGAAAGTCTGCGT. cdna was made using SuperScript III Reverse Transcriptase (Invitrogen, San Diego, CAL), and RTPCR was performed using Power SYBER Green PCR Master Mix (Applied Biosystems, Carlsbad, CAL) at an initial 30sec 95 C denaturation, 30sec 64 C annealing and 30sec 72 C extension for 5 cycles, then 30sec 95 C denaturation, 30sec 60 C annealing and 30sec 72 C extension for 40 cycles, followed by a melt curve of 15sec 95 C, 15sec 55 C and 15sec 95 C. Quantification was by comparison to GAPDHF:ACGCATTTGGTCGTATTGGG, GAPDHR: TGATTTTGGAGGGATCTCGC, as previously described. 14 6
7 Statistical Analysis Firstly, both the pooled ANZgene cohort and individual RNA-seq data were analysed using the statistical package, EdgeR. 15 EdgeR was used to assess differences in gene expression between MS and controls. Genes expressed at >25 reads per million total reads and calculated to be differentially expressed with FDR<10% were identified (Supplementary Table 1). Secondly, using the data obtained by RNA-seq, all genes with a minimum RPKM value greater than 100 were selected for further analysis (344 genes). To account for differences in expression levels, for each gene the expression level was ranked across individuals, and ranked data used to test correlations between genes. Further details are provided in figure legends and the supplementary information section. 7
8 Results Replication of the TCAGE in an independent cohort using RNA-seq We interrogated whole blood gene expression using mrna from blood collected into PAXgene tubes, and RNA-seq to assess gene expression. To test the RNA-seq protocol, we first prepared two cdna libraries from pooled mrna from 20 MS patients and 20 healthy controls (HC) used in the ANZgene microarray experiment reported earlier, 5 and measured expression of the transcriptome using Hiseq2000 to 30 million reads per pool. All 38 TCAGE genes expressed at >25 reads per million reads were more highly expressed in MS, 16 of these at p<0.001 (Fig. 1, Supplementary Table 1A, Fig. S1). Half of these 16 were genes encoding ribosomal proteins. We then prepared RNA-seq libraries from an independent collection of whole blood from 40 HC and 32 untreated MS patients (cohort traits detailed in Table 1), and measured transcriptomes to a depth of >10 million reads per sample. We used EdgeR to assess the relative expression of all genes expressed with at least 25 reads per million reads in all samples. Of the 26 TCAGE genes in this list 12 were ribosomal protein genes, all of which were more highly expressed in MS (12/12, p<0.0005). This low p value is indicative only, since these genes are likely to be coordinately regulated. Since our second study was smaller than the first, we used a permissive FDR of 10% to define a dysregulated gene set (genes listed in Supplementary Table 1B). Ribosomal protein genes are again over-represented. We then tested the cellular pathways that were over-represented, and the immune cell subsets tagged, by these genes; and compared these with those 8
9 identified in our previously published microarray study. Again, over-expressed genes were predominantly expressed in T cells (Fig. 2a), underexpressed genes predominantly expressed in other immune cell subsets (Fig. 2b), and the most dysregulated pathways were those involving the ribosomal protein genes (Fig. 2c). RPS6 is a surrogate marker for ribosomal protein and other TCAGE genes If the ribosomal protein (RP) genes are coordinately expressed, measurement of one should provide a proxy for regulation of all. From these data, we chose RPS6 as the most highly associated with MS (p< , t test on RPKM ranks), and compared the rank of its expression with the sum of the ranks of all TCAGE genes in each individual. Expression of RPS6 was tightly correlated with the expression of all 12 RP genes (R= 0.93, p< 1.96E-25, Fig. 3a), with all 26 TCAGE genes (R=0.85, p<1.65e-18), and with the 14 nonrp genes in the TCAGE list (R=0.70, p<1.87e-11). By comparing the aligned reads from the RNA-seq analysis of pooled MS and HC samples, it can be seen that exon usage for RPS6 was the same in MS and HC (Fig. 3b inset). We designed primers from across the exon 4-5 boundary and compared RPS6 expression by RTPCR with the RPKM measurement from RNA-seq. Measurements from both techniques were highly correlated (R=0.93, p=2.73e-24, Fig. 3b). RPS6 expression is seasonal 9
10 In the RNA-seq cohort age and gender were not associated with RPS6 expression (Fig S6), but season was (Fig. 4b). We analysed the data using month of sample collection, with months classified as Summer or Winter on the basis of RPS6 expression: In Summer - October-March RPS6 expression was highest ; Winter - April-September, lowest expression. Overall, in the RNA-seq cohort RPS6 expression was higher in MS than HC (Fig. 4a, p<0.003). However, this difference was only evident in winter (p< ) with a trend in summer (Fig. 4b, p=0.19). The seasonal difference for HC (Fig. 4, p<7.2e-8) and MS patients (p<0.004) was also highly significant. Similar differences were seen in the ANZgene microarray cohort (Fig. 4a), with a significant difference between MS and controls in summer (p<0.0002). The RPS6 seasonal pattern was confirmed in a third large cohort of HC from the subtropical city of Brisbane (p<0.0016, Fig. S2). In both the RNA-seq cohort (R=0.82, p< 0.002) Fig. 5b) and the Brisbane cohort (p<0.01, Fig. S2) the RPS6 expression correlated with UV radiation. Although RPS6 expression in the RNA-seq cohort did not correlate with Vitamin D levels (R=0.27, ns, Fig5), correlation increases if two months out of phase, with RPS6 increasing two months earlier than Vitamin D (R=0.71, p<0.011). No diurnal effect on RPS6 expression was detected in 5 individuals assayed at 4 time points (Supplementary Methods, Figure S3). Using Receiver Operator Curves, where the expression of RPS6 was 95% sensitive to detect MS, the specificity was 43%, with an area under the curve of 0.72, p= (Fig. 6). In winter, this specificity improves to 67%, and the area under the curve to 0.92, p< In summer RPS6 expression was not predictive of MS. 10
11 Because some studies but not others 16 have indicated month of birth affects on MS risk, and RPS6 regulation might be set by imprinting, we tested if expression was associated with month of birth. There was no association. Relative expression of ribosomal protein genes in immune cell subsets To identify the cell subsets contributing to the difference in the RP signature between MS and HC, we interrogated transcriptomes from 23 immune cell subsets in whole blood using RNA-seq (Fig. 7, Supplementary Methods), and compared this to in silico data for the subsets. 17, 18 The expression level between subsets was consistent (Figure S4). RPS6 and other ribosomal protein genes were most highly expressed in CD4 and CD8 T cells. RPS6 expression is altered by interferon beta We measured RPS6 expression 9-15 hours after IFNB injection in 32 MS patients, 16 of which were classified as responders to IFNB, as indicated by MxA upregulation, whilst 16 were classified as non-responders. The difference between the two is considered to be due to the development of neutralizing antibodies in the latter. 9 RPS6 expression was lower in the IFNB responsive group when compared to the non-responsive MS patients in both summer (p=0.018) and winter (p=0.006). In addition, the RPS6 expression was lower in MS patients, classified as IFNB responders when compared to controls in summer (p=0.046) and winter (p=0.079). The non-responders exhibited higher 11
12 RPS6 expression than controls in summer (p=0.003) and winter (p=0.006), and they had similar expression of RPS6 to untreated MS (Fig. 8). RPS6 expression is not strongly heritable Expression profiles were collected for the 862 individuals from extended MZ and DZ twin families in Brisbane System Genetics Cohort (BSGS) using Illumina HT12 v.40 arrays. 13 Genotypes for each of these individuals were also determined (Illumina 610-Quad Beadchip). Family relationships among individuals within BSGS were used to estimate the additive genetic component of expression variation for the two probes in RPS6 using ASREML. 19 Heritabilities (h 2 ) for the two probes were estimated by the ratio of additive genetic variation to total phenotypic variation. Estimates of h 2 for the two RPS6 probes on this chip were zero, even after correcting for month of sample collection, suggesting that any variation in the expression levels of the two probes is due to environmental factors, at least for whole blood. None of the 528,509 SNPs on this chip were significantly correlated with RPS6 expression. 12
13 Discussion Using an independent cohort and alternative method of transcriptome interrogation, we have replicated our original finding that a set genes predominantly expressed in T cells, notably the ribosomal protein (RP) genes, are overexpressed in MS. Measurement of expression of one gene, RPS6, could be used as a surrogate for the expression of the other RP and TCAGE genes. As well as disease state, RP gene expression was highly affected by season. Higher expression was seen in the late spring, summer months than winter and early spring, mirroring the UV light seasonal pattern. We found no evidence that the expression pattern was heritable, suggesting that the difference in expression between MS and HC was caused by disease and/or environmental factors. RPS6 whole blood expression was also reduced on IFNB therapy. Ribosomes are made up of equimolar amounts of each of the 80 ribosomal proteins, so that co-ordinate regulation of RPs is to be expected, 20 and is observed in this study. However, there are extra-ribosomal functions, and variable phosphorylation of the ribosome constituent proteins, so that cell subsets have subtle but important differences in their RP transcriptomes. 21 RPS6 is essential for ribosome biogenesis, is phosphorylated at evolutionarily conserved residues, and regulates protein synthesis, cell growth and proliferation, but also has specific features which have led to its intensive investigation as a biomarker. 22 It is most highly expressed in lymphocytes, especially naïve T cells (Fig 8 17, 18 ). It controls metabolic signaling in CD8 T cells; 23 has been reported to be a predictor of resistance to therapeutic inhibition of FLT3 targeting leukemic blasts; 24 and predicts sensitivity to the inhibitor of 13
14 immune cell proliferation, mtor. 25 Higher expression of RPS6 in PBMCs was observed in people with MS more likely to relapse. 26 A gene regulating RPS6 phosphorylation, RPS6KB1, is associated with MS susceptibility. Phosphorylation of RPS6 is downstream of mtor activation, and therapeutic inhibition of mtor by drugs such as rapamycin increase the proportion of Tregs to Th1 and Th17 cells. 27 These data support a role for RPS6 in MS susceptibility, pathogenesis, therapeutic response, and as a target for novel therapies. Immune cell expression of RPS6 is higher throughout the year in untreated MS patients, suggesting a greater T cell proliferation/activation, which may be enough to increase risk of development of MS and relapses. MS onset and relapses are most common in spring/summer in both the northern and southern hemispheres. 28, 29 This is when UV light and RPS6 expression start to increase and before serum vitamin D increases. Seasonally corrected serum vitamin D levels correlate with reduced risk of onset and relapses in MS, in both hemispheres 29. Demands on the immune system are clearly seasonal, and immune cell subsets numbers and proportions in whole blood fluctuate. 30, 31 Although the regulators of the seasonal pattern are not known, they could include melatonin, light sensitive hormones such as vitamin D, and UV light factors independent of vitamin D. 32 The seasonal fluctuation in serum 25D levels is some two months out of phase with the RPS6 and UV light fluctuation. This could indicate RPS6 expression is immediately responsive to UV light, or that immediate vitamin D 14
15 fluctuation due to UV light affects immune cells more than serum vitamin D levels. IFNB has been the most commonly prescribed drug for MS, with many possible mechanisms of action proposed. 33 These include regulation of the balance of immune cell subsets. We have shown that the RPS6 signal is greatly reduced on IFNB therapy. This may be affected by a number of factors: reduced T cell activation and proliferation as a therapeutic response, reduced RPS6 transcription on therapy, or to altered T cell representation in whole blood after injection due to trafficking of immune cell subsets. Reduced lymphocyte and neutrophil numbers, but greatly increased monocyte numbers were seen in whole blood after injection, 34 and altered immune cell state might also be expected. RPS6 whole blood levels would be expected to drop, as observed in the current cohort. Whether this reduction is correlated, or even underpins clinical response needs to be determined. A major limitation of using whole blood is that particular immune cell subset differences are difficult to identify. In this case, although RPS6 expression is predominantly from T cells (Fig. 6), the cell subset source of the RPS6 difference between MS and controls will now need to be established using flow cytometry. Cell subsets which vary by season, and in interferon response, have previously been described, 31,32,34 but which of these, or of other subsets, contribute to the RPS6 seasonal fluctuation and IFNB response will also need to be established through flow cytometric methods. Another limitation of this study is that the RNAseq cohort controls and some MS patients were from Sydney, and many MS 15
16 patients but no controls were from Adelaide. No confounding factors between the two regions are known eg they have similar UV light levels (Fig 4), but its possible. High RPS6 expression, fold change between seasons, or discrepancy between RPS6 and other physiological parameters such as serum vitamin D, may indicate MS susceptibility and state, but also a molecular subtype of MS particularly likely to respond to certain therapies. Testing such possibilities is vital to developing cost-effective therapies in autoimmune diseases. 35 Funding This work was supported by project grants from Multiple Sclerosis Research Australia (MSRA), the National Health and Medical Research Council Australia, and Merck Serono. DB is funded by the Hunt Family MSRA Senior Research Fellowship. Author Contribution Design: DB, GP Laboratory Work: GP,PG,SS Sample Collection: MS: SV,PG,FM,MK,MS,JL, SB; HC:PG, FM,PV, GM,JP Funding: DB, GS Draft: DB, GP, GS 16
17 Final Version: all authors Conflict of Interests None Acknowledgements We would like to thank the people with MS and controls for donating blood to support this research. This work was funded by grants from Multiple Sclerosis Research Australia and Merck Serono. David Booth is the Hunt Family Senior MS Research Fellow. 17
18 References 1. Compston A and Coles A. Multiple sclerosis. Lancet. 2008; 372: Bielekova B and Becker BL. Monoclonal antibodies in MS: mechanisms of action. Neurology. 2010; 74 Suppl 1: S Mehling M, Kappos L and Derfuss T. Fingolimod for multiple sclerosis: mechanism of action, clinical outcomes, and future directions. Current neurology and neuroscience reports. 2011; 11: Sawcer S, Hellenthal G, Pirinen M, et al. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature. 2011; 476: Gandhi KS, McKay FC, Cox M, et al. The multiple sclerosis whole blood mrna transcriptome and genetic associations indicate dysregulation of specific T cell pathways in pathogenesis. Human Molecular Genetics Joehanes R, Johnson AD, Barb JJ, et al. Gene expression analysis of whole blood, peripheral blood mononuclear cells, and lymphoblastoid cell lines from the Framingham Heart Study. Physiological genomics. 2012; 44: Stuve O and Ransohoff RM. Immunotherapy for multiple sclerosis: the curious case of interferon beta. Archives of neurology. 2009; 66: Powell JE, Henders AK, McRae AF, et al. Genetic control of gene expression in whole blood and lymphoblastoid cell lines is largely independent. Genome research. 2012; 22: McKay F, Schibeci S, Heard R, Stewart G and Booth D. Analysis of neutralizing antibodies to therapeutic interferon-beta in multiple sclerosis patients: a comparison of three methods in a large Australasian cohort. Journal of immunological methods. 2006; 310: Trapnell C, Pachter L and Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 2009; 25: Xu G, Deng N, Zhao Z, Judeh T, Flemington E and Zhu D. SAMMate: a GUI tool for processing short read alignments in SAM/BAM format. Source code for biology and medicine. 2011; 6: Simon R, Lam A, Li M-C, Ngan M, Menenzes S and Zhao Y. Analysis of Gene Expression Data Using BRB-Array Tools. Cancer Informatics. 2007; 3: Powell JE, Henders AK, McRae AF, et al. The Brisbane Systems Genetics Study: genetical genomics meets complex trait genetics. PLoS ONE. 2012; 7: e Livak KJ and Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001; 25: Robinson MD, McCarthy DJ and Smyth GK. edger: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010; 26: Hintzen R. Multiple sclerosis: Month of birth effect in MS-fact or artefact? Nature reviews Neurology Liu SM, Xavier R, Good KL, et al. Immune cell transcriptome datasets reveal novel leukocyte subset-specific genes and genes associated with allergic processes. The Journal of allergy and clinical immunology. 2006; 118:
19 18. Su AI, Wiltshire T, Batalov S, et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc Natl Acad Sci U S A. 2004; 101: Gilmour AR, Thompson R and Cullis BR. Average information REML: An efficient algorithm for variance parameter estimation in linear mixed models. Biometrics. 1995; 51: Perry RP. Balanced production of ribosomal proteins. Gene. 2007; 401: Zimmermann RA. The double life of ribosomal proteins. Cell. 2003; 115: Rosner M, Schipany K and Hengstschlager M. Spatial consequences of blocking mtor/s6k: relevance for therapy. Cell Cycle. 2012; 11: Salmond RJ, Emery J, Okkenhaug K and Zamoyska R. MAPK, phosphatidylinositol 3-kinase, and mammalian target of rapamycin pathways converge at the level of ribosomal protein S6 phosphorylation to control metabolic signaling in CD8 T cells. J Immunol. 2009; 183: Perl AE, Kasner MT, Shank D, Luger SM and Carroll M. Single-cell pharmacodynamic monitoring of S6 ribosomal protein phosphorylation in AML blasts during a clinical trial combining the mtor inhibitor sirolimus and intensive chemotherapy. Clinical cancer research : an official journal of the American Association for Cancer Research. 2012; 18: Boulay A, Zumstein-Mecker S, Stephan C, et al. Antitumor efficacy of intermittent treatment schedules with the rapamycin derivative RAD001 correlates with prolonged inactivation of ribosomal protein S6 kinase 1 in peripheral blood mononuclear cells. Cancer research. 2004; 64: Achiron A, Feldman A, Magalashvili D, Dolev M and Gurevich M. Suppressed RNA-polymerase 1 pathway is associated with benign multiple sclerosis. PLoS ONE. 2012; 7: e Cobbold SP, Adams E, Nolan KF, Regateiro FS and Waldmann H. Connecting the mechanisms of T-cell regulation: dendritic cells as the missing link. Immunological reviews. 2010; 236: Jin Y, de Pedro-Cuesta J, Soderstrom M, Stawiarz L and Link H. Seasonal patterns in optic neuritis and multiple sclerosis: a meta-analysis. Journal of the neurological sciences. 2000; 181: Simpson S, Jr., Taylor B, Blizzard L, et al. Higher 25-hydroxyvitamin D is associated with lower relapse risk in multiple sclerosis. Annals of neurology. 2010; 68: Maes M, Stevens W, Scharpe S, et al. Seasonal variation in peripheral blood leukocyte subsets and in serum interleukin-6, and soluble interleukin-2 and -6 receptor concentrations in normal volunteers. Experientia. 1994; 50: Khoo AL, Koenen HJ, Chai LY, et al. Seasonal variation in vitamin D(3) levels is paralleled by changes in the peripheral blood human T cell compartment. PLoS ONE. 2012; 7: e Hart PH. Vitamin D supplementation, moderate sun exposure, and control of immune diseases. Discovery medicine. 2012; 13: Dhib-Jalbut S and Marks S. Interferon-beta mechanisms of action in multiple sclerosis. Neurology. 2010; 74 Suppl 1: S Kantor AB, Deng J, Waubant E, et al. Identification of short-term pharmacodynamic effects of interferon-beta-1a in multiple sclerosis subjects 19
20 with broad- based phenotypic profiling. Journal of neuroimmunology. 2007; 188: Blumberg RS, Dittel B, Hafler D, von Herrath M and Nestle FO. Unraveling the autoimmune translational research process layer by layer. Nature medicine. 2012; 18: Boyages S and Bilinski K. Seasonal reduction in vitamin D level persists into spring in NSW Australia: implications for monitoring and replacement therapy. Clinical endocrinology. 2012; 77:
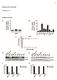
21 Figure Legends Figure 1. Heat map of TCAGE gene expression in RNA-seq and ANZgene cohorts. Rows are expression of the most highly expressed T cell activation genes (TCAGE) identified in Gandhi et al (2010) 6 plotted against the healthy controls and MS patients, sorted by season, for the ANZgene cohort (from pooled RNA-seq) and the RNA-seq cohort. For the latter, each individual is a column. Columns sorted by month of collection. Orange is high expression, blue is low. Figure 2. Heat map of Immune Cell Subset Expression of Genes Dyregulated in MS in the RNA-seq Cohort. Rows are the genes identified as dysregulated in the RNA-seq cohort (expression > 25 reads per million read, FDR 10%), columns the immune cell subsets (detailed in the Methods section). A. Genes overexpressed in MS, B. Genes underexpressed in MS, C. Pathways overrepresented in the overexpressed genes. This is determined for the 552 genes with the lowest p values (corresponding to the number identified as differentially expressed at 10% FDR in the microarray analysis by 5 ) in both cohorts, and the 10% FDR list from the RNA-seq cohort. These lists were uploaded into Metacore (GeneGo, USA) and tested for overrepresentation in curated biological pathways (Figure 2C). Figure 3. RPS6 expression is correlated with expression of other ribosomal genes. A. Rank of RPS6 expression in 72 individuals, measured by RNA-seq, and Sum of Ranks of expression of all other Ribosomal Genes for those individuals. Correlation=0.93 B. Primers designed from exon usage in MS and Controls (inset) and the linear correlation of RTPCR and RNA-seq measurement of RPS6 is Figure 4. RPS6 expression in whole blood is dependent on season and disease state. A. Comparison of RPS6 expression in the RNA-seq cohort (this study) and the ANZgene cohort, published in Gandhi et al, RPS6 is measured by reads per kilobase per million mapped reads (RPKM) for RNA-seq, and by fluorescence on microarrays (ANZgene). Data are from normalised expression, compared using twotailed T tests. B. Effect of season on RPS6 expression in the RNA-seq cohort. Figure 5. RPS6 expression is affected by season and is correlated with UV radiation and Vitamin D levels. A. Top graph is RPS6 expression in each month, middle graph is UV index for different localities used as the source of samples (Australian Bureau of Meteorology website: lowest graph is Vitamin D levels in NSW. 48 B. Correlation of RPS6 expression with UV radiation (R=0.83), Vitamin D (R=0.25), and Vitamin D, two months out of phase (R=0.70). R is the correlation co-efficient, determined from a linear correlation. 21
22 Figure 6. Receiver operator curves to predict MS from RPS6 expression Figure 7. RPS6 is predominantly expressed in T cells. Heat map shown for immune cell subsets (columns) and most expressed ribosomal protein genes, including RPS6 (rows). Colour is proportional to expression, with orange high expression and blue low expression. Immune cell subsets are defined in Supplementary Material and Methods. Figure 8. Effect of Interferon Treatment on RPS6 Expression. People with MS on interferon beta without neutralizing antibodies (IFN (R)), had lower RPS6 expression than healthy controls (HC), those treated with interferon but with neutralizing antibodies (IFN (NR)), and people with MS (MS). The latter did not differ from IFN (NR). Each comparison for the IFNB treatment was assessed using the two tailed T test, with no correction for multiple testing. 22
Autoimmunity and immunemediated. FOCiS. Lecture outline
 1 Autoimmunity and immunemediated inflammatory diseases Abul K. Abbas, MD UCSF FOCiS 2 Lecture outline Pathogenesis of autoimmunity: why selftolerance fails Genetics of autoimmune diseases Therapeutic
1 Autoimmunity and immunemediated inflammatory diseases Abul K. Abbas, MD UCSF FOCiS 2 Lecture outline Pathogenesis of autoimmunity: why selftolerance fails Genetics of autoimmune diseases Therapeutic
Gene Expression Assays
 APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
Understanding West Nile Virus Infection
 Understanding West Nile Virus Infection The QIAGEN Bioinformatics Solution: Biomedical Genomics Workbench (BXWB) + Ingenuity Pathway Analysis (IPA) Functional Genomics & Predictive Medicine, May 21-22,
Understanding West Nile Virus Infection The QIAGEN Bioinformatics Solution: Biomedical Genomics Workbench (BXWB) + Ingenuity Pathway Analysis (IPA) Functional Genomics & Predictive Medicine, May 21-22,
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS)
 Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
Introduction to transcriptome analysis using High Throughput Sequencing technologies (HTS) A typical RNA Seq experiment Library construction Protocol variations Fragmentation methods RNA: nebulization,
New Technologies for Sensitive, Low-Input RNA-Seq. Clontech Laboratories, Inc.
 New Technologies for Sensitive, Low-Input RNA-Seq Clontech Laboratories, Inc. Outline Introduction Single-Cell-Capable mrna-seq Using SMART Technology SMARTer Ultra Low RNA Kit for the Fluidigm C 1 System
New Technologies for Sensitive, Low-Input RNA-Seq Clontech Laboratories, Inc. Outline Introduction Single-Cell-Capable mrna-seq Using SMART Technology SMARTer Ultra Low RNA Kit for the Fluidigm C 1 System
Introduction To Real Time Quantitative PCR (qpcr)
 Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Supplemental Material. Methods
 Supplemental Material Methods Measurement of lncrnas expression Total RNA was extracted from PAXgene TM tubes using the PAXgene blood RNA kit (Qiagen, Venlo, Netherlands) as described by the manufacturer.
Supplemental Material Methods Measurement of lncrnas expression Total RNA was extracted from PAXgene TM tubes using the PAXgene blood RNA kit (Qiagen, Venlo, Netherlands) as described by the manufacturer.
ONLINE SUPPLEMENTAL MATERIAL. Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue
 ONLINE SUPPLEMENTAL MATERIAL Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue Sungmi Park 1, Ko-Ting Lu 1, Xuebo Liu 1, Tapan K. Chatterjee 2, Steven M. Rudich 3, Neal
ONLINE SUPPLEMENTAL MATERIAL Allele-Specific Expression of Angiotensinogen in Human Subcutaneous Adipose Tissue Sungmi Park 1, Ko-Ting Lu 1, Xuebo Liu 1, Tapan K. Chatterjee 2, Steven M. Rudich 3, Neal
Compositional Changes of B and T Cell Subtypes during Fingolimod Treatment in Multiple Sclerosis Patients: A 12-Month Follow-Up Study
 Compositional Changes of B and T Cell Subtypes during Fingolimod Treatment in Multiple Sclerosis Patients: A 12-Month Follow-Up Study Nele Claes 1 *; Tessa Dhaeze 1 *; Judith Fraussen PhD 1 ; Bieke Broux
Compositional Changes of B and T Cell Subtypes during Fingolimod Treatment in Multiple Sclerosis Patients: A 12-Month Follow-Up Study Nele Claes 1 *; Tessa Dhaeze 1 *; Judith Fraussen PhD 1 ; Bieke Broux
G E N OM I C S S E RV I C ES
 GENOMICS SERVICES THE NEW YORK GENOME CENTER NYGC is an independent non-profit implementing advanced genomic research to improve diagnosis and treatment of serious diseases. capabilities. N E X T- G E
GENOMICS SERVICES THE NEW YORK GENOME CENTER NYGC is an independent non-profit implementing advanced genomic research to improve diagnosis and treatment of serious diseases. capabilities. N E X T- G E
Materials and Methods. Blocking of Globin Reverse Transcription to Enhance Human Whole Blood Gene Expression Profiling
 Application Note Blocking of Globin Reverse Transcription to Enhance Human Whole Blood Gene Expression Profi ling Yasmin Beazer-Barclay, Doug Sinon, Christopher Morehouse, Mark Porter, and Mike Kuziora
Application Note Blocking of Globin Reverse Transcription to Enhance Human Whole Blood Gene Expression Profi ling Yasmin Beazer-Barclay, Doug Sinon, Christopher Morehouse, Mark Porter, and Mike Kuziora
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
Gene Expression Analysis
 Gene Expression Analysis Jie Peng Department of Statistics University of California, Davis May 2012 RNA expression technologies High-throughput technologies to measure the expression levels of thousands
Gene Expression Analysis Jie Peng Department of Statistics University of California, Davis May 2012 RNA expression technologies High-throughput technologies to measure the expression levels of thousands
treatments) worked by killing cancerous cells using chemo or radiotherapy. While these techniques can
 Shristi Pandey Genomics and Medicine Winter 2011 Prof. Doug Brutlag Chronic Myeloid Leukemia: A look into how genomics is changing the way we treat Cancer. Until the late 1990s, nearly all treatment methods
Shristi Pandey Genomics and Medicine Winter 2011 Prof. Doug Brutlag Chronic Myeloid Leukemia: A look into how genomics is changing the way we treat Cancer. Until the late 1990s, nearly all treatment methods
Analysis of gene expression data. Ulf Leser and Philippe Thomas
 Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
Analysis of Illumina Gene Expression Microarray Data
 Analysis of Illumina Gene Expression Microarray Data Asta Laiho, Msc. Tech. Bioinformatics research engineer The Finnish DNA Microarray Centre Turku Centre for Biotechnology, Finland The Finnish DNA Microarray
Analysis of Illumina Gene Expression Microarray Data Asta Laiho, Msc. Tech. Bioinformatics research engineer The Finnish DNA Microarray Centre Turku Centre for Biotechnology, Finland The Finnish DNA Microarray
Identification of rheumatoid arthritis and osteoarthritis patients by transcriptome-based rule set generation
 Identification of rheumatoid arthritis and osterthritis patients by transcriptome-based rule set generation Bering Limited Report generated on September 19, 2014 Contents 1 Dataset summary 2 1.1 Project
Identification of rheumatoid arthritis and osterthritis patients by transcriptome-based rule set generation Bering Limited Report generated on September 19, 2014 Contents 1 Dataset summary 2 1.1 Project
Head of College Scholars List Scheme. Summer Studentship. Report Form
 Head of College Scholars List Scheme Summer Studentship Report Form This report should be completed by the student with his/her project supervisor. It should summarise the work undertaken during the project
Head of College Scholars List Scheme Summer Studentship Report Form This report should be completed by the student with his/her project supervisor. It should summarise the work undertaken during the project
ncounter Leukemia Fusion Gene Expression Assay Molecules That Count Product Highlights ncounter Leukemia Fusion Gene Expression Assay Overview
 ncounter Leukemia Fusion Gene Expression Assay Product Highlights Simultaneous detection and quantification of 25 fusion gene isoforms and 23 additional mrnas related to leukemia Compatible with a variety
ncounter Leukemia Fusion Gene Expression Assay Product Highlights Simultaneous detection and quantification of 25 fusion gene isoforms and 23 additional mrnas related to leukemia Compatible with a variety
Uses of Flow Cytometry
 Uses of Flow Cytometry 1. Multicolour analysis... 2 2. Cell Cycle and Proliferation... 3 a. Analysis of Cellular DNA Content... 4 b. Cell Proliferation Assays... 5 3. Immunology... 6 4. Apoptosis... 7
Uses of Flow Cytometry 1. Multicolour analysis... 2 2. Cell Cycle and Proliferation... 3 a. Analysis of Cellular DNA Content... 4 b. Cell Proliferation Assays... 5 3. Immunology... 6 4. Apoptosis... 7
From Reads to Differentially Expressed Genes. The statistics of differential gene expression analysis using RNA-seq data
 From Reads to Differentially Expressed Genes The statistics of differential gene expression analysis using RNA-seq data experimental design data collection modeling statistical testing biological heterogeneity
From Reads to Differentially Expressed Genes The statistics of differential gene expression analysis using RNA-seq data experimental design data collection modeling statistical testing biological heterogeneity
micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved
 microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
RT-PCR: Two-Step Protocol
 RT-PCR: Two-Step Protocol We will provide both one-step and two-step protocols for RT-PCR. We recommend the twostep protocol for this class. In the one-step protocol, the components of RT and PCR are mixed
RT-PCR: Two-Step Protocol We will provide both one-step and two-step protocols for RT-PCR. We recommend the twostep protocol for this class. In the one-step protocol, the components of RT and PCR are mixed
Clinical Trials of Disease Modifying Treatments
 MS CENTER CLINICAL RESEARCH The UCSF MS Center is an internationally recognized leader in multiple sclerosis clinical research. We conduct clinical trials involving the use of experimental treatments,
MS CENTER CLINICAL RESEARCH The UCSF MS Center is an internationally recognized leader in multiple sclerosis clinical research. We conduct clinical trials involving the use of experimental treatments,
FlipFlop: Fast Lasso-based Isoform Prediction as a Flow Problem
 FlipFlop: Fast Lasso-based Isoform Prediction as a Flow Problem Elsa Bernard Laurent Jacob Julien Mairal Jean-Philippe Vert September 24, 2013 Abstract FlipFlop implements a fast method for de novo transcript
FlipFlop: Fast Lasso-based Isoform Prediction as a Flow Problem Elsa Bernard Laurent Jacob Julien Mairal Jean-Philippe Vert September 24, 2013 Abstract FlipFlop implements a fast method for de novo transcript
Frequently Asked Questions Next Generation Sequencing
 Frequently Asked Questions Next Generation Sequencing Import These Frequently Asked Questions for Next Generation Sequencing are some of the more common questions our customers ask. Questions are divided
Frequently Asked Questions Next Generation Sequencing Import These Frequently Asked Questions for Next Generation Sequencing are some of the more common questions our customers ask. Questions are divided
The Immunopathogenesis of Relapsing MS
 The Immunopathogenesis of Relapsing MS Olaf Stüve, M.D., Ph.D. Neurology Section VA North Texas Health Care System Dallas VA Medical Center Departments of Neurology and Neurotherapeutics University of
The Immunopathogenesis of Relapsing MS Olaf Stüve, M.D., Ph.D. Neurology Section VA North Texas Health Care System Dallas VA Medical Center Departments of Neurology and Neurotherapeutics University of
Case Report Early Bilateral Cystoid Macular Oedema Secondary to Fingolimod in Multiple Sclerosis
 Case Reports in Medicine Volume 2012, Article ID 134636, 4 pages doi:10.1155/2012/134636 Case Report Early Bilateral Cystoid Macular Oedema Secondary to Fingolimod in Multiple Sclerosis Lei Liu and Fiona
Case Reports in Medicine Volume 2012, Article ID 134636, 4 pages doi:10.1155/2012/134636 Case Report Early Bilateral Cystoid Macular Oedema Secondary to Fingolimod in Multiple Sclerosis Lei Liu and Fiona
Relative Quantification of mirna Target mrnas by Real-Time qpcr. 1 Introduction. Gene Expression Application Note No. 4
 Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
PreciseTM Whitepaper
 Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
Targeting Specific Cell Signaling Pathways for the Treatment of Malignant Peritoneal Mesothelioma
 The Use of Kinase Inhibitors: Translational Lab Results Targeting Specific Cell Signaling Pathways for the Treatment of Malignant Peritoneal Mesothelioma Sheelu Varghese, Ph.D. H. Richard Alexander, M.D.
The Use of Kinase Inhibitors: Translational Lab Results Targeting Specific Cell Signaling Pathways for the Treatment of Malignant Peritoneal Mesothelioma Sheelu Varghese, Ph.D. H. Richard Alexander, M.D.
Supplemental Information. McBrayer et al. Supplemental Data
 1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
Standards, Guidelines and Best Practices for RNA-Seq V1.0 (June 2011) The ENCODE Consortium
 Standards, Guidelines and Best Practices for RNA-Seq V1.0 (June 2011) The ENCODE Consortium I. Introduction: Sequence based assays of transcriptomes (RNA-seq) are in wide use because of their favorable
Standards, Guidelines and Best Practices for RNA-Seq V1.0 (June 2011) The ENCODE Consortium I. Introduction: Sequence based assays of transcriptomes (RNA-seq) are in wide use because of their favorable
BIOINF 525 Winter 2016 Foundations of Bioinformatics and Systems Biology http://tinyurl.com/bioinf525-w16
 Course Director: Dr. Barry Grant (DCM&B, bjgrant@med.umich.edu) Description: This is a three module course covering (1) Foundations of Bioinformatics, (2) Statistics in Bioinformatics, and (3) Systems
Course Director: Dr. Barry Grant (DCM&B, bjgrant@med.umich.edu) Description: This is a three module course covering (1) Foundations of Bioinformatics, (2) Statistics in Bioinformatics, and (3) Systems
RNAseq / ChipSeq / Methylseq and personalized genomics
 RNAseq / ChipSeq / Methylseq and personalized genomics 7711 Lecture Subhajyo) De, PhD Division of Biomedical Informa)cs and Personalized Biomedicine, Department of Medicine University of Colorado School
RNAseq / ChipSeq / Methylseq and personalized genomics 7711 Lecture Subhajyo) De, PhD Division of Biomedical Informa)cs and Personalized Biomedicine, Department of Medicine University of Colorado School
Expression Quantification (I)
 Expression Quantification (I) Mario Fasold, LIFE, IZBI Sequencing Technology One Illumina HiSeq 2000 run produces 2 times (paired-end) ca. 1,2 Billion reads ca. 120 GB FASTQ file RNA-seq protocol Task
Expression Quantification (I) Mario Fasold, LIFE, IZBI Sequencing Technology One Illumina HiSeq 2000 run produces 2 times (paired-end) ca. 1,2 Billion reads ca. 120 GB FASTQ file RNA-seq protocol Task
A Primer of Genome Science THIRD
 A Primer of Genome Science THIRD EDITION GREG GIBSON-SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc. Publishers Sunderland, Massachusetts USA Contents Preface xi 1 Genome Projects:
A Primer of Genome Science THIRD EDITION GREG GIBSON-SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc. Publishers Sunderland, Massachusetts USA Contents Preface xi 1 Genome Projects:
Essentials of Real Time PCR. About Sequence Detection Chemistries
 Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Disclosures. Consultant and Speaker for Biogen Idec, TEVA Neuroscience, EMD Serrono, Mallinckrodt, Novartis, Genzyme, Accorda Therapeutics
 Mitzi Joi Williams, MD Neurologist MS Center of Atlanta, Atlanta, GA Disclosures Consultant and Speaker for Biogen Idec, TEVA Neuroscience, EMD Serrono, Mallinckrodt, Novartis, Genzyme, Accorda Therapeutics
Mitzi Joi Williams, MD Neurologist MS Center of Atlanta, Atlanta, GA Disclosures Consultant and Speaker for Biogen Idec, TEVA Neuroscience, EMD Serrono, Mallinckrodt, Novartis, Genzyme, Accorda Therapeutics
How To Get A Cell Print
 QUICK CELL CAPTURE AND CHARACTERIZATION GUIDE FOR CELLSEARCH CUSTOMERS CellSave EDTA Blood sample Rare cell capture Enumeration Single protein marker Cell capture for molecular characterization CELLSEARCH
QUICK CELL CAPTURE AND CHARACTERIZATION GUIDE FOR CELLSEARCH CUSTOMERS CellSave EDTA Blood sample Rare cell capture Enumeration Single protein marker Cell capture for molecular characterization CELLSEARCH
Validating Microarray Data Using RT 2 Real-Time PCR Products
 Validating Microarray Data Using RT 2 Real-Time PCR Products Introduction: Real-time PCR monitors the amount of amplicon as the reaction occurs. Usually, the amount of product is directly related to the
Validating Microarray Data Using RT 2 Real-Time PCR Products Introduction: Real-time PCR monitors the amount of amplicon as the reaction occurs. Usually, the amount of product is directly related to the
Effects of Herceptin on circulating tumor cells in HER2 positive early breast cancer
 Effects of Herceptin on circulating tumor cells in HER2 positive early breast cancer J.-L. Zhang, Q. Yao, J.-H. Chen,Y. Wang, H. Wang, Q. Fan, R. Ling, J. Yi and L. Wang Xijing Hospital Vascular Endocrine
Effects of Herceptin on circulating tumor cells in HER2 positive early breast cancer J.-L. Zhang, Q. Yao, J.-H. Chen,Y. Wang, H. Wang, Q. Fan, R. Ling, J. Yi and L. Wang Xijing Hospital Vascular Endocrine
ALLIANCE FOR LUPUS RESEARCH AND PFIZER S CENTERS FOR THERAPEUTIC INNOVATION CHALLENGE GRANT PROGRAM PROGRAM GUIDELINES
 ALLIANCE FOR LUPUS RESEARCH AND PFIZER S CENTERS FOR THERAPEUTIC INNOVATION CHALLENGE GRANT PROGRAM PROGRAM GUIDELINES DESCRIPTION OF GRANT MECHANISM The Alliance for Lupus Research (ALR) is an independent,
ALLIANCE FOR LUPUS RESEARCH AND PFIZER S CENTERS FOR THERAPEUTIC INNOVATION CHALLENGE GRANT PROGRAM PROGRAM GUIDELINES DESCRIPTION OF GRANT MECHANISM The Alliance for Lupus Research (ALR) is an independent,
Real-time PCR: Understanding C t
 APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
LEUKEMIA LYMPHOMA MYELOMA Advances in Clinical Trials
 LEUKEMIA LYMPHOMA MYELOMA Advances in Clinical Trials OUR FOCUS ABOUT emerging treatments Presentation for: Judith E. Karp, MD Advancements for Acute Myelogenous Leukemia Supported by an unrestricted educational
LEUKEMIA LYMPHOMA MYELOMA Advances in Clinical Trials OUR FOCUS ABOUT emerging treatments Presentation for: Judith E. Karp, MD Advancements for Acute Myelogenous Leukemia Supported by an unrestricted educational
Investigating the genetic basis for intelligence
 Investigating the genetic basis for intelligence Steve Hsu University of Oregon and BGI www.cog-genomics.org Outline: a multidisciplinary subject 1. What is intelligence? Psychometrics 2. g and GWAS: a
Investigating the genetic basis for intelligence Steve Hsu University of Oregon and BGI www.cog-genomics.org Outline: a multidisciplinary subject 1. What is intelligence? Psychometrics 2. g and GWAS: a
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual
 Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Single-Cell DNA Sequencing with the C 1. Single-Cell Auto Prep System. Reveal hidden populations and genetic diversity within complex samples
 DATA Sheet Single-Cell DNA Sequencing with the C 1 Single-Cell Auto Prep System Reveal hidden populations and genetic diversity within complex samples Single-cell sensitivity Discover and detect SNPs,
DATA Sheet Single-Cell DNA Sequencing with the C 1 Single-Cell Auto Prep System Reveal hidden populations and genetic diversity within complex samples Single-cell sensitivity Discover and detect SNPs,
Targeted Therapy What the Surgeon Needs to Know
 Targeted Therapy What the Surgeon Needs to Know AATS Focus in Thoracic Surgery 2014 David R. Jones, M.D. Professor & Chief, Thoracic Surgery Memorial Sloan Kettering Cancer Center I have no disclosures
Targeted Therapy What the Surgeon Needs to Know AATS Focus in Thoracic Surgery 2014 David R. Jones, M.D. Professor & Chief, Thoracic Surgery Memorial Sloan Kettering Cancer Center I have no disclosures
Real-Time PCR Vs. Traditional PCR
 Real-Time PCR Vs. Traditional PCR Description This tutorial will discuss the evolution of traditional PCR methods towards the use of Real-Time chemistry and instrumentation for accurate quantitation. Objectives
Real-Time PCR Vs. Traditional PCR Description This tutorial will discuss the evolution of traditional PCR methods towards the use of Real-Time chemistry and instrumentation for accurate quantitation. Objectives
REAL TIME PCR USING SYBR GREEN
 REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
ALPHA (TNFa) IN OBESITY
 THE ROLE OF TUMOUR NECROSIS FACTOR ALPHA (TNFa) IN OBESITY Alison Mary Morris, B.Sc (Hons) A thesis submitted to Adelaide University for the degree of Doctor of Philosophy Department of Physiology Adelaide
THE ROLE OF TUMOUR NECROSIS FACTOR ALPHA (TNFa) IN OBESITY Alison Mary Morris, B.Sc (Hons) A thesis submitted to Adelaide University for the degree of Doctor of Philosophy Department of Physiology Adelaide
Cancer Immunotherapy: Can Your Immune System Cure Cancer? Steve Emerson, MD, PhD Herbert Irving Comprehensive Cancer Center
 Cancer Immunotherapy: Can Your Immune System Cure Cancer? Steve Emerson, MD, PhD Herbert Irving Comprehensive Cancer Center Bodnar s Law Simple Things are Important Very Simple Things are Very Important
Cancer Immunotherapy: Can Your Immune System Cure Cancer? Steve Emerson, MD, PhD Herbert Irving Comprehensive Cancer Center Bodnar s Law Simple Things are Important Very Simple Things are Very Important
Using Illumina BaseSpace Apps to Analyze RNA Sequencing Data
 Using Illumina BaseSpace Apps to Analyze RNA Sequencing Data The Illumina TopHat Alignment and Cufflinks Assembly and Differential Expression apps make RNA data analysis accessible to any user, regardless
Using Illumina BaseSpace Apps to Analyze RNA Sequencing Data The Illumina TopHat Alignment and Cufflinks Assembly and Differential Expression apps make RNA data analysis accessible to any user, regardless
Microarray Data Analysis. A step by step analysis using BRB-Array Tools
 Microarray Data Analysis A step by step analysis using BRB-Array Tools 1 EXAMINATION OF DIFFERENTIAL GENE EXPRESSION (1) Objective: to find genes whose expression is changed before and after chemotherapy.
Microarray Data Analysis A step by step analysis using BRB-Array Tools 1 EXAMINATION OF DIFFERENTIAL GENE EXPRESSION (1) Objective: to find genes whose expression is changed before and after chemotherapy.
MOLECULAR PHARMACOLOGY AND EXPERIMENTAL THERAPEUTICS
 MOLECULAR PHARMACOLOGY AND EXPERIMENTAL THERAPEUTICS R. M. Weinshilboum, M.D., Program Director L. Wang, M.D., Ph.D., Program Co-Director D. C. Mays, Ph.D., Associate Program Director Ph.D. Degree Course
MOLECULAR PHARMACOLOGY AND EXPERIMENTAL THERAPEUTICS R. M. Weinshilboum, M.D., Program Director L. Wang, M.D., Ph.D., Program Co-Director D. C. Mays, Ph.D., Associate Program Director Ph.D. Degree Course
Human Genome Organization: An Update. Genome Organization: An Update
 Human Genome Organization: An Update Genome Organization: An Update Highlights of Human Genome Project Timetable Proposed in 1990 as 3 billion dollar joint venture between DOE and NIH with 15 year completion
Human Genome Organization: An Update Genome Organization: An Update Highlights of Human Genome Project Timetable Proposed in 1990 as 3 billion dollar joint venture between DOE and NIH with 15 year completion
News on modifying diseases therapies. Michel CLANET CHU Toulouse France ECTRIMS
 News on modifying diseases therapies Michel CLANET CHU Toulouse France ECTRIMS Current treatment strategies Future oral treatments Future non oral treatments Drug safety and risks CIS at risk of MS Active
News on modifying diseases therapies Michel CLANET CHU Toulouse France ECTRIMS Current treatment strategies Future oral treatments Future non oral treatments Drug safety and risks CIS at risk of MS Active
Appendix 2 Molecular Biology Core Curriculum. Websites and Other Resources
 Appendix 2 Molecular Biology Core Curriculum Websites and Other Resources Chapter 1 - The Molecular Basis of Cancer 1. Inside Cancer http://www.insidecancer.org/ From the Dolan DNA Learning Center Cold
Appendix 2 Molecular Biology Core Curriculum Websites and Other Resources Chapter 1 - The Molecular Basis of Cancer 1. Inside Cancer http://www.insidecancer.org/ From the Dolan DNA Learning Center Cold
Modelling and analysis of T-cell epitope screening data.
 Modelling and analysis of T-cell epitope screening data. Tim Beißbarth 2, Jason A. Tye-Din 1, Gordon K. Smyth 1, Robert P. Anderson 1 and Terence P. Speed 1 1 WEHI, 1G Royal Parade, Parkville, VIC 3050,
Modelling and analysis of T-cell epitope screening data. Tim Beißbarth 2, Jason A. Tye-Din 1, Gordon K. Smyth 1, Robert P. Anderson 1 and Terence P. Speed 1 1 WEHI, 1G Royal Parade, Parkville, VIC 3050,
SYBR Green Realtime PCR Master Mix -Plus-
 Instruction manual SYBR Green Realtime PCR Master Mix -Plus- 0810 F0925K SYBR Green Realtime PCR Master Mix -Plus- Contents QPK-212T 1mLx1 QPK-212 1mLx5 Store at -20 C, protected from light [1] Introduction
Instruction manual SYBR Green Realtime PCR Master Mix -Plus- 0810 F0925K SYBR Green Realtime PCR Master Mix -Plus- Contents QPK-212T 1mLx1 QPK-212 1mLx5 Store at -20 C, protected from light [1] Introduction
Clinical Research Infrastructure
 Clinical Research Infrastructure Enhancing UK s Clinical Research Capabilities & Technologies At least 150m to establish /develop cutting-edge technological infrastructure, UK wide. to bring into practice
Clinical Research Infrastructure Enhancing UK s Clinical Research Capabilities & Technologies At least 150m to establish /develop cutting-edge technological infrastructure, UK wide. to bring into practice
Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Biochemistry
 Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Biochemistry The Master Degree in Medical Laboratory Sciences /Clinical Biochemistry, is awarded by the Faculty of Graduate Studies
Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Biochemistry The Master Degree in Medical Laboratory Sciences /Clinical Biochemistry, is awarded by the Faculty of Graduate Studies
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
Validation and Replication
 Validation and Replication Overview Definitions of validation and replication Difficulties and limitations Working examples from our group and others Why? False positive results still occur. even after
Validation and Replication Overview Definitions of validation and replication Difficulties and limitations Working examples from our group and others Why? False positive results still occur. even after
RT 2 Profiler PCR Array: Web-Based Data Analysis Tutorial
 RT 2 Profiler PCR Array: Web-Based Data Analysis Tutorial Samuel J. Rulli, Jr., Ph.D. qpcr-applications Scientist Samuel.Rulli@QIAGEN.com Pathway Focused Research from Sample Prep to Data Analysis! -2-
RT 2 Profiler PCR Array: Web-Based Data Analysis Tutorial Samuel J. Rulli, Jr., Ph.D. qpcr-applications Scientist Samuel.Rulli@QIAGEN.com Pathway Focused Research from Sample Prep to Data Analysis! -2-
Services. Updated 05/31/2016
 Updated 05/31/2016 Services 1. Whole exome sequencing... 2 2. Whole Genome Sequencing (WGS)... 3 3. 16S rrna sequencing... 4 4. Customized gene panels... 5 5. RNA-Seq... 6 6. qpcr... 7 7. HLA typing...
Updated 05/31/2016 Services 1. Whole exome sequencing... 2 2. Whole Genome Sequencing (WGS)... 3 3. 16S rrna sequencing... 4 4. Customized gene panels... 5 5. RNA-Seq... 6 6. qpcr... 7 7. HLA typing...
Lecture 6: Single nucleotide polymorphisms (SNPs) and Restriction Fragment Length Polymorphisms (RFLPs)
 Lecture 6: Single nucleotide polymorphisms (SNPs) and Restriction Fragment Length Polymorphisms (RFLPs) Single nucleotide polymorphisms or SNPs (pronounced "snips") are DNA sequence variations that occur
Lecture 6: Single nucleotide polymorphisms (SNPs) and Restriction Fragment Length Polymorphisms (RFLPs) Single nucleotide polymorphisms or SNPs (pronounced "snips") are DNA sequence variations that occur
Micro RNAs: potentielle Biomarker für das. Blutspenderscreening
 Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Tutorial for proteome data analysis using the Perseus software platform
 Tutorial for proteome data analysis using the Perseus software platform Laboratory of Mass Spectrometry, LNBio, CNPEM Tutorial version 1.0, January 2014. Note: This tutorial was written based on the information
Tutorial for proteome data analysis using the Perseus software platform Laboratory of Mass Spectrometry, LNBio, CNPEM Tutorial version 1.0, January 2014. Note: This tutorial was written based on the information
Personalised Medicine in MS
 Personalised Medicine in MS Supportive Evidence from Therapeutic Trials Ludwig Kappos Neurology and Department of Biomedicine University Hospital CH-4031 Basel LKappos@uhbs.ch Established partially effective
Personalised Medicine in MS Supportive Evidence from Therapeutic Trials Ludwig Kappos Neurology and Department of Biomedicine University Hospital CH-4031 Basel LKappos@uhbs.ch Established partially effective
Chapter 43: The Immune System
 Name Period Our students consider this chapter to be a particularly challenging and important one. Expect to work your way slowly through the first three concepts. Take particular care with Concepts 43.2
Name Period Our students consider this chapter to be a particularly challenging and important one. Expect to work your way slowly through the first three concepts. Take particular care with Concepts 43.2
Gene Therapy. The use of DNA as a drug. Edited by Gavin Brooks. BPharm, PhD, MRPharmS (PP) Pharmaceutical Press
 Gene Therapy The use of DNA as a drug Edited by Gavin Brooks BPharm, PhD, MRPharmS (PP) Pharmaceutical Press Contents Preface xiii Acknowledgements xv About the editor xvi Contributors xvii An introduction
Gene Therapy The use of DNA as a drug Edited by Gavin Brooks BPharm, PhD, MRPharmS (PP) Pharmaceutical Press Contents Preface xiii Acknowledgements xv About the editor xvi Contributors xvii An introduction
MystiCq microrna cdna Synthesis Mix Catalog Number MIRRT Storage Temperature 20 C
 microrna cdna Synthesis Mix Catalog Number MIRRT Storage Temperature 20 C Product Description The microrna cdna Synthesis Mix has been designed to easily convert micrornas into cdna templates for qpcr
microrna cdna Synthesis Mix Catalog Number MIRRT Storage Temperature 20 C Product Description The microrna cdna Synthesis Mix has been designed to easily convert micrornas into cdna templates for qpcr
Diabetes and Drug Development
 Diabetes and Drug Development Metabolic Disfunction Leads to Multiple Diseases Hypertension ( blood pressure) Metabolic Syndrome (Syndrome X) LDL HDL Lipoproteins Triglycerides FFA Hyperinsulinemia Insulin
Diabetes and Drug Development Metabolic Disfunction Leads to Multiple Diseases Hypertension ( blood pressure) Metabolic Syndrome (Syndrome X) LDL HDL Lipoproteins Triglycerides FFA Hyperinsulinemia Insulin
Using TaqMan Endogenous Control Assays to select an endogenous control for experimental studies
 APPLICATION NOTE TaqMan Gene Expression Assays Using TaqMan Endogenous Control Assays to select an endogenous control for experimental studies Introduction Quantitative real-time PCR (qpcr) allows for
APPLICATION NOTE TaqMan Gene Expression Assays Using TaqMan Endogenous Control Assays to select an endogenous control for experimental studies Introduction Quantitative real-time PCR (qpcr) allows for
Vad är bioinformatik och varför behöver vi det i vården? a bioinformatician's perspectives
 Vad är bioinformatik och varför behöver vi det i vården? a bioinformatician's perspectives Dirk.Repsilber@oru.se 2015-05-21 Functional Bioinformatics, Örebro University Vad är bioinformatik och varför
Vad är bioinformatik och varför behöver vi det i vården? a bioinformatician's perspectives Dirk.Repsilber@oru.se 2015-05-21 Functional Bioinformatics, Örebro University Vad är bioinformatik och varför
Gene Mapping Techniques
 Gene Mapping Techniques OBJECTIVES By the end of this session the student should be able to: Define genetic linkage and recombinant frequency State how genetic distance may be estimated State how restriction
Gene Mapping Techniques OBJECTIVES By the end of this session the student should be able to: Define genetic linkage and recombinant frequency State how genetic distance may be estimated State how restriction
Haematopoietic Chimerism Analysis after Allogeneic Stem Cell Transplantation
 Haematopoietic Chimerism Analysis after Allogeneic Stem Cell Transplantation Dr Ros Ganderton, Ms Kate Parratt, Dr Debbie Richardson, Dr Kim Orchard and Dr Liz Hodges Departments of Molecular Pathology
Haematopoietic Chimerism Analysis after Allogeneic Stem Cell Transplantation Dr Ros Ganderton, Ms Kate Parratt, Dr Debbie Richardson, Dr Kim Orchard and Dr Liz Hodges Departments of Molecular Pathology
The Prospect of Stem Cell Therapy in Multiple Sclerosis. Multiple sclerosis is a multifocal inflammatory disease of the central
 The Prospect of Stem Cell Therapy in Multiple Sclerosis Multiple sclerosis is a multifocal inflammatory disease of the central nervous system that generally affects young individuals, causing paralysis
The Prospect of Stem Cell Therapy in Multiple Sclerosis Multiple sclerosis is a multifocal inflammatory disease of the central nervous system that generally affects young individuals, causing paralysis
CyTOF2. Mass cytometry system. Unveil new cell types and function with high-parameter protein detection
 CyTOF2 Mass cytometry system Unveil new cell types and function with high-parameter protein detection DISCOVER MORE. IMAGINE MORE. MASS CYTOMETRY. THE FUTURE OF CYTOMETRY TODAY. Mass cytometry resolves
CyTOF2 Mass cytometry system Unveil new cell types and function with high-parameter protein detection DISCOVER MORE. IMAGINE MORE. MASS CYTOMETRY. THE FUTURE OF CYTOMETRY TODAY. Mass cytometry resolves
Quantitative proteomics background
 Proteomics data analysis seminar Quantitative proteomics and transcriptomics of anaerobic and aerobic yeast cultures reveals post transcriptional regulation of key cellular processes de Groot, M., Daran
Proteomics data analysis seminar Quantitative proteomics and transcriptomics of anaerobic and aerobic yeast cultures reveals post transcriptional regulation of key cellular processes de Groot, M., Daran
Microarray Technology
 Microarrays And Functional Genomics CPSC265 Matt Hudson Microarray Technology Relatively young technology Usually used like a Northern blot can determine the amount of mrna for a particular gene Except
Microarrays And Functional Genomics CPSC265 Matt Hudson Microarray Technology Relatively young technology Usually used like a Northern blot can determine the amount of mrna for a particular gene Except
Introduction To Epigenetic Regulation: How Can The Epigenomics Core Services Help Your Research? Maria (Ken) Figueroa, M.D. Core Scientific Director
 Introduction To Epigenetic Regulation: How Can The Epigenomics Core Services Help Your Research? Maria (Ken) Figueroa, M.D. Core Scientific Director Gene expression depends upon multiple factors Gene Transcription
Introduction To Epigenetic Regulation: How Can The Epigenomics Core Services Help Your Research? Maria (Ken) Figueroa, M.D. Core Scientific Director Gene expression depends upon multiple factors Gene Transcription
Overview of Phase 1 Oncology Trials of Biologic Therapeutics
 Overview of Phase 1 Oncology Trials of Biologic Therapeutics Susan Jerian, MD ONCORD, Inc. February 28, 2008 February 28, 2008 Phase 1 1 Assumptions and Ground Rules The goal is regulatory approval of
Overview of Phase 1 Oncology Trials of Biologic Therapeutics Susan Jerian, MD ONCORD, Inc. February 28, 2008 February 28, 2008 Phase 1 1 Assumptions and Ground Rules The goal is regulatory approval of
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE E15
 INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE DEFINITIONS FOR GENOMIC BIOMARKERS, PHARMACOGENOMICS,
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE DEFINITIONS FOR GENOMIC BIOMARKERS, PHARMACOGENOMICS,
Immune dysregulation in MS
 Immune dysregulation in MS N. Muls, A. Dang, V. Van Pesch, C. Sindic Multiple sclerosis (MS) is considered to be a chronic autoimmune disease resulting in inflammation and demyelination in the central
Immune dysregulation in MS N. Muls, A. Dang, V. Van Pesch, C. Sindic Multiple sclerosis (MS) is considered to be a chronic autoimmune disease resulting in inflammation and demyelination in the central
How To Change Medicine
 P4 Medicine: Personalized, Predictive, Preventive, Participatory A Change of View that Changes Everything Leroy E. Hood Institute for Systems Biology David J. Galas Battelle Memorial Institute Version
P4 Medicine: Personalized, Predictive, Preventive, Participatory A Change of View that Changes Everything Leroy E. Hood Institute for Systems Biology David J. Galas Battelle Memorial Institute Version
Measuring gene expression (Microarrays) Ulf Leser
 Measuring gene expression (Microarrays) Ulf Leser This Lecture Gene expression Microarrays Idea Technologies Problems Quality control Normalization Analysis next week! 2 http://learn.genetics.utah.edu/content/molecules/transcribe/
Measuring gene expression (Microarrays) Ulf Leser This Lecture Gene expression Microarrays Idea Technologies Problems Quality control Normalization Analysis next week! 2 http://learn.genetics.utah.edu/content/molecules/transcribe/
Deep profiling of multitube flow cytometry data Supplemental information
 Deep profiling of multitube flow cytometry data Supplemental information Kieran O Neill et al December 19, 2014 1 Table S1: Markers in simulated multitube data. The data was split into three tubes, each
Deep profiling of multitube flow cytometry data Supplemental information Kieran O Neill et al December 19, 2014 1 Table S1: Markers in simulated multitube data. The data was split into three tubes, each
specific B cells Humoral immunity lymphocytes antibodies B cells bone marrow Cell-mediated immunity: T cells antibodies proteins
 Adaptive Immunity Chapter 17: Adaptive (specific) Immunity Bio 139 Dr. Amy Rogers Host defenses that are specific to a particular infectious agent Can be innate or genetic for humans as a group: most microbes
Adaptive Immunity Chapter 17: Adaptive (specific) Immunity Bio 139 Dr. Amy Rogers Host defenses that are specific to a particular infectious agent Can be innate or genetic for humans as a group: most microbes
The Functional but not Nonfunctional LILRA3 Contributes to Sex Bias in Susceptibility and Severity of ACPA-Positive Rheumatoid Arthritis
 The Functional but not Nonfunctional LILRA3 Contributes to Sex Bias in Susceptibility and Severity of ACPA-Positive Rheumatoid Arthritis Yan Du Peking University People s Hospital 100044 Beijing CHINA
The Functional but not Nonfunctional LILRA3 Contributes to Sex Bias in Susceptibility and Severity of ACPA-Positive Rheumatoid Arthritis Yan Du Peking University People s Hospital 100044 Beijing CHINA
J D R F R E Q U E S T S L E T T E R S O F I N T E N T F O R : B I O M AR K E R S O F P AN C R E A T I C B E T A C E L L S T R E S S AN D H E AL T H
 J D R F R E Q U E S T S L E T T E R S O F I N T E N T F O R : B I O M AR K E R S O F P AN C R E A T I C B E T A C E L L S T R E S S AN D H E AL T H PURPOSE JDRF, the world s leading non-profit organization
J D R F R E Q U E S T S L E T T E R S O F I N T E N T F O R : B I O M AR K E R S O F P AN C R E A T I C B E T A C E L L S T R E S S AN D H E AL T H PURPOSE JDRF, the world s leading non-profit organization
AP Biology Essential Knowledge Student Diagnostic
 AP Biology Essential Knowledge Student Diagnostic Background The Essential Knowledge statements provided in the AP Biology Curriculum Framework are scientific claims describing phenomenon occurring in
AP Biology Essential Knowledge Student Diagnostic Background The Essential Knowledge statements provided in the AP Biology Curriculum Framework are scientific claims describing phenomenon occurring in
Basic processing of next-generation sequencing (NGS) data
 Basic processing of next-generation sequencing (NGS) data Getting from raw sequence data to expression analysis! 1 Reminder: we are measuring expression of protein coding genes by transcript abundance
Basic processing of next-generation sequencing (NGS) data Getting from raw sequence data to expression analysis! 1 Reminder: we are measuring expression of protein coding genes by transcript abundance
Application Guide... 2
 Protocol for GenomePlex Whole Genome Amplification from Formalin-Fixed Parrafin-Embedded (FFPE) tissue Application Guide... 2 I. Description... 2 II. Product Components... 2 III. Materials to be Supplied
Protocol for GenomePlex Whole Genome Amplification from Formalin-Fixed Parrafin-Embedded (FFPE) tissue Application Guide... 2 I. Description... 2 II. Product Components... 2 III. Materials to be Supplied
Sequencing Library qpcr Quantification Guide
 Sequencing Library qpcr Quantification Guide FOR RESEARCH USE ONLY Introduction 3 Quantification Workflow 4 Best Practices 5 Consumables and Equipment 6 Select Control Template 8 Dilute qpcr Control Template
Sequencing Library qpcr Quantification Guide FOR RESEARCH USE ONLY Introduction 3 Quantification Workflow 4 Best Practices 5 Consumables and Equipment 6 Select Control Template 8 Dilute qpcr Control Template
The immune system. Bone marrow. Thymus. Spleen. Bone marrow. NK cell. B-cell. T-cell. Basophil Neutrophil. Eosinophil. Myeloid progenitor
 The immune system Basophil Neutrophil Bone marrow Eosinophil Myeloid progenitor Dendritic cell Pluripotent Stem cell Lymphoid progenitor Platelets Bone marrow Thymus NK cell T-cell B-cell Spleen Cancer
The immune system Basophil Neutrophil Bone marrow Eosinophil Myeloid progenitor Dendritic cell Pluripotent Stem cell Lymphoid progenitor Platelets Bone marrow Thymus NK cell T-cell B-cell Spleen Cancer
Mir-X mirna First-Strand Synthesis Kit User Manual
 User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
