1 Protein Structure and Its Physical Basis Physical Forces and Principles Underlying Protein Folding and Structure... 3
|
|
|
- Audra Hoover
- 10 years ago
- Views:
Transcription
1 Protein Folding A. Szilágyi. J. Kardos. S. Osváth. L. Barna. P. Závodszky 1 Protein Structure and Its Physical Basis Physical Forces and Principles Underlying Protein Folding and Structure The Protein Folding Problem Folding Mechanisms and Kinetics Molten Globules and Other Compact Denatured States The Structure of the Molten Globule The Role of the Molten Globule in Folding Simulations of Protein Folding and Unfolding All Atom Models Simplified Models Multiscale Modeling Free Energy Landscapes of Proteins Traps on the Folding Pathway Backbone Isomerization Formation of Disulfide Bridges Transition States on the Folding Pathway Folding of Multidomain and Multi Subunit Proteins Multidomain Proteins Multi Subunit Proteins Protein Folding in the Cell The Hsp70 Family The Folding Cage of Chaperonins (Hsp60 Family) The Hsp90 Chaperone System Chaperone Assisted Assembly of Cellular Complexes Cooperation Between the Different Chaperone Systems Chaperones and Neurodegenerative Diseases Protein Misfolding and Aggregation Degenerative Diseases Associated with Amyloid Deposition The Structure and Morphology of Amyloid Fibrils Mechanism of Amyloid Fibril Formation Amyloid Formation of Globular Proteins Physicochemical and Sequence Determinants of Amyloid Formation Factors Inducing or Inhibiting Amyloid Formation Under Physiological Conditions # Springer-Verlag Berlin Heidelberg 2007
2 2 Protein folding 12 New Biophysical Techniques for the Study of Protein Folding Rapid Mixing Methods Real Time NMR Spectroscopy Chemically Induced Nuclear Polarization High Pressure NMR Spectroscopy Protein Folding and Dynamics Studied by Mass Spectrometry Mechanical Unfolding of Proteins Conclusions... 30
3 Protein folding 3 Abstract: Since Anfinsen s famous experiments in the 1960s, it has been known that the complex threedimensional structure of protein molecules is encoded in their amino acid sequences, and the chains autonomously fold under proper conditions. Cracking this code, which is sometimes called the second part of the genetic code, has been one of the greatest challenges of molecular biology. Although a full understanding of how proteins fold remains elusive, theoretical and experimental studies of protein folding have come a long way since Anfinsen s findings. In the living cell, folding occurs in a complex and crowded environment, often involving helper proteins, and in some cases it can go awry: the protein can misfold, aggregate, or form amyloid fibers. It is increasingly recognized that misfolded proteins and amyloid formation are the root cause of a number of serious illnesses including several neurodegenerative diseases. Therefore, the study of protein folding remains a key area of biomedical research. List of Abbreviations: AFM, atomic force microscopy; CCP, chaperonin containing TCP 1; CIDNP, chemically induced nuclear polarization; DMD, discrete molecular dynamics; EM, electron microscopy; ESIMS, electrospray ionization mass spectrometry; FIS, factor for inversion stimulation; FMN, flavin mononucleotide; NOE, nuclear Overhauser effect; TF, trigger factor; TCP 1, tailless complex polypeptide 1 1 Protein Structure and Its Physical Basis The function of a protein can only be interpreted from its structure. The nervous system is a network of cells, and the peculiar functional properties of these cells can be derived from the properties and interactions of their proteins. Proteins are involved in all stages of neural activity. Those embedded wholly or partly in membranes regulate the transport of ions and molecules as a means of signal exchange with other cells and the external medium. Some of them have enzymatic functions to catalyze the chemical processes essential for function. The diverse and highly specific function of proteins is a consequence of their sophisticated, individual surface pattern regarding shape, charge, and hydrophobicity. The surface pattern is a consequence of the unique three dimensional structure of the polypeptide chain. Proteins are linear polymers with nonrepetitive, specific covalent structure. The covalent structure is determined by the order of amino acids in which they are linked together. Since Anfinsen s famous experiments (1973) in the 1960s, it has been believed and today generally accepted that folding and the resulting native structure of proteins are autonomously governed and determined by the amino acid sequence of a particular protein and its natural solvent environment. 1.1 Physical Forces and Principles Underlying Protein Folding and Structure A linear polypeptide chain is autonomously organized into a space filling, compact, and well defined threedimensional structure. In a globular protein, the internal core is mostly formed by hydrophobic amino acid residues, held together by van der Waals forces, and the surface of the globule is formed by mostly charged and polar side chains. Proteins exist in this state of condensed matter while the specific conformation is largely determined by the flexibility of the polypeptide backbone and by the specific, consistent intermolecular interactions of the side chains. The monomeric unit in a polypeptide chain is the peptide group. The sequence of amino acids is the primary structure of the protein. The C, O, N, and H atoms lie in the same plane; successive planes define angles f and c. The conformation of a chain of n amino acids can be defined by 2n parameters. The restricted flexibility of the polypeptide chain is a major factor among those determining protein structure and folding. The native conformation must be energetically stable. From a thermodynamic point of view, the free energy of a protein molecule is influenced by the following major contributions: (1) the hydrophobic effect, (2) the energy of hydrogen bonds, (3) the energy of electrostatic interactions, and (4) the conformational entropy due to the restricted motion of the main chain and the side chains.
4 4 Protein folding The hydrophobic effect used to be explained as a primarily entropic effect arising from the rearrangement of hydrogen bonds between solvent molecules around an apolar solute. This hydration process is energetically unfavorable, and therefore drives apolar solutes together, thereby decreasing their solventexposed surface area. Today, the hydrophobic effect is usually viewed as a combined effect of hydration (an entropic effect) and van der Waals interactions between solute molecules (an enthalpic effect) (Makhatadze and Privalov, 1995). It is therefore entropic at low temperatures and enthalpic at high temperatures, which results in a complex temperature dependence of its strength (Schellman, 1997). Nevertheless, the hydrophobic force has long been considered as the major driving force of protein folding (Dill, 1990) as it leads to a rapid collapse of the polypeptide chain, thereby largely reducing the configurational space to explore. Without doubt, the hydrophobic interaction is also a major stabilizing force contributing to the thermodynamic stability of the folded state. The role of hydrogen bonds in folding and stability used to be underestimated based on the argument that intramolecular hydrogen bonds can be replaced by hydrogen bonds between the protein and the solvent. After a number of mutational studies, however, hydrogen bonds have now been recognized as having a contribution to protein stability as important as the hydrophobic effect (Pace et al., 1996). This contribution was estimated to be kcal/mol per buried intramolecular hydrogen bond (Pace et al., 1996). Electrostatic interactions such as ion pairs and salt bridges in proteins have been an area of active research (Kumar and Nussinov, 2002). While hydrogen bonds and hydrophobic forces are essentially nonspecific, electrostatic interactions are largely specific, and therefore play an important role in specifying the fold of a protein as well as in protein flexibility and function. Computational and experimental evidence shows that salt bridges can be stabilizing or destabilizing. On the other hand, genome wide and structural comparisons of thermophilic and mesophilic proteins indicate that salt bridges may significantly contribute to the enhanced thermal stability of proteins from thermophilic organisms (Szilagyi and Zavodszky, 2000; Li et al., 2005; Razvi and Scholtz, 2006). The major destabilizing contribution to the stability of the folded state is the conformational entropy of the polypeptide chain. Folding a long chain into a specific, compact structure obviously results in a significant entropy decrease. This is counterbalanced by the various intrachain interactions described above. The resulting overall stability of the protein (the free energy difference between the folded and the unfolded state) is marginal, being on the order of 5 kcal/mol. This number is a small difference between huge stabilizing and destabilizing contributions. We qualitatively know that the hydrophobic effect and hydrogen bonds are the major stabilizing contributions and the conformational entropy is the major destabilizing one. However, due to the compensatory effects in the total energy balance, a quantitative prediction with respect to the significance of any specific type of interaction cannot be made with confidence (Jaenicke, 2000). 2 The Protein Folding Problem The central dogma of molecular biology states that the flow of sequential information from nucleic acid to protein is unidirectional: nucleic acid sequences encode the sequence of proteins but once translation occurs, information cannot flow back from protein to nucleic acid. A possible extension of the central dogma would be to add that the sequence of the protein encodes its three dimensional structure. Indeed, this coding is sometimes called the second half of the genetic code. Cracking this code would be equivalent to solving the protein folding problem. To see what this problem is all about, let us look at Anfinsen s classic experiment (1973) (> Figure -1). Ribonuclease, an enzyme with 124 amino acid residues, contains four disulfide bridges. Treatment of ribonuclease with 8 M urea in the presence of the reducing agent b mercaptoethanol causes a complete unfolding of the ribonuclease molecule, yielding an essentially random form. In this process, the four disulfide bridges get cleaved, resulting in eight free SH groups. The enzymatic activity of the molecule is completely lost. Allowing the cysteines to reoxidize under denaturing conditions results in a mixture of scrambled species where the eight SH groups randomly pair to form four disulfide bridges (it is easy to
5 Protein folding 5. Figure -1 Scheme of Anfinsen s experiment. See text for details calculate that there are 5 possible pairings). However, when urea is slowly removed by dialysis and a small amount of b mercaptoethanol is added, disulfide interchange takes place and the mixture of scrambled ribonuclease molecules is eventually converted to a homogeneous product, which is fully enzymatically active and indistinguishable from native ribonuclease. This crucial experiment supports the idea known as the thermodynamic hypothesis, which states that the three dimensional structure of a native protein in its normal physiological state is the one in which the Gibbs free energy of the whole system is lowest. Therefore, in a given environment, the native conformation of the protein is fully determined by its amino acid sequence. How the information encoded in the sequence gets translated into the three dimensional structure? This is the protein folding problem. Assuming that the polypeptide chain randomly samples all possible configurations, we can estimate the time required for a protein to fold. If each bond connecting two neighboring amino acids can have, say, three possible states then a protein of 1 amino acids could exist in 3 0 ¼ 5 47 different configurations. Only one of these configurations corresponds to the native state. Even if the protein is able to sample new configurations at a rate of 13 per second, it will take 27 years to try them all. Nevertheless, proteins do fold, and in a timescale of seconds or less. This contradiction was first pointed out by Cyrus Levinthal in 1969 (Levinthal, 1969) and has become known as Levinthal s paradox. To resolve the paradox, Levinthal argued that the protein cannot fold by random search and there must be specific folding pathways. The concept of folding pathways motivated a large number of experimental studies aimed at finding the specific folding intermediates and also gave rise to a number of models describing the folding process. For example, the nucleation/growth model (Wetlaufer, 1973) tried to resolve Levinthal s paradox by assuming that the rate limiting step of the folding process is a nucleation event, presumably the formation of smaller structural units, and once nucleation occurs the nuclei grow fast and the folding process rapidly completes. This model is not consistent with the large number of observations where folding intermediates were observed. According to the diffusion collision adhesion model (Karplus and Weaver, 1976), fluctuating microdomains (portions of secondary structure or hydrophobic clusters) move diffusively and repeatedly collide with each other. Collisions can lead to a coalescence of the microdomains into larger units (adhesion). The rate limiting stage is assumed to be the diffusion process. This model is well supported by many experiments (Karplus and Weaver, 1994). The framework model (Baldwin, 1989) states that the
6 6 Protein folding folding process is hierarchical, starting with the formation of the secondary structure elements, and the docking of the preformed substructures is the rate limiting step. The hydrophobic collapse model (Dill, 1985) is based on the view that the hydrophobic effect is the main driving force of folding, and the process starts with a rapid collapse of the chain, followed by the formation of the secondary structure. In fact, whether hydrophobic collapse or secondary structure formation occurs first has remained a largely undecided issue even to this day. Finally, the jigsaw puzzle model (Harrison and Durbin, 1985) denied the necessity of a unique, directed folding pathway and stated that each protein molecule can follow a different route to the native structure, just like there are multiple ways to solve a jigsaw puzzle. This idea is actually consistent with a new view of protein folding, which gained popularity in the 1990s: the energy landscape view. The energy landscape view likens the energy landscape of a protein to a funnel, with the native structure at its global minimum, and each molecule may follow a different microscopic route from the top to the bottom (energy landscapes are discussed in more detail later). The many models of protein folding are not mutually exclusive; they try to grasp different aspects of folding, and experimental results give some support to each model we mentioned. A newer model, named nucleation condensation model, is an attempt to unite the features of both the framework and the hydrophobic collapse mechanisms (Fersht, 1995; Fersht, 1997). In this model, long range and other native hydrophobic interactions form in the transition state to stabilize the otherwise weak secondary structure. The framework and the hydrophobic collapse models are viewed as two extremes of the nucleation condensation mechanism; most proteins fold by a mechanism that is somewhere between the two extremes, i.e., secondary structure and hydrophobic interactions form nearly simultaneously and synergistically (Daggett and Fersht, 2003). 3 Folding Mechanisms and Kinetics A unified view of protein folding should be general enough to interpret the diverse experimental findings of the field. Thermodynamics offers such a universal approach. Thermodynamic systems in equilibrium occupy the states with lowest Gibbs free energy at constant pressure and temperature. The Gibbs free energy (G) consists of an enthalpy and an entropic term Gq ðþ¼hq ðþ Tq ðþ; where H is the enthalpy, T the absolute temperature, and S the entropy of the protein, and q represents the reaction coordinate used to describe the progress of the protein advancing from the unfolded toward the native state. Under physiological conditions, proteins maintain their native structure because the favorable enthalpic term arising from the solvent and protein interactions exceeds in magnitude the unfavorable entropic term, and therefore the native state has a smaller Gibbs free energy than the denatured state. The stability of the protein depends on the solvent solvent, protein solvent, and protein protein interactions. These interactions depend on the intensive parameters that describe the thermodynamic state of the system. The enthalpic and the entropic terms are large, but of opposite sign, and almost cancel each other. The Gibbs free energy difference between the biologically active and denatured states of the proteins is rather small (Scharnagl et al., 2005). Proteins are stable only within a narrow range of conditions and can be denatured by changing virtually any of the intensive parameters (Shortle, 1996). Experiments prove that proteins can be unfolded by heat (Tsai et al., 2002; Prabhu and Sharp, 2005), cold (Franks, 1995; Kunugi and Tanaka, 2002), high pressure (Smeller, 2002; Meersman et al., 2006), extreme ph (Puett, 1973; Fitch et al., 2006), and addition of salts (Pfeil, 1981). Studies of protein stability and folding systematically change one or more of the intensive parameters and follow the kinetics of the change and/or the shift of equilibrium. There is a broad selection of methods that can be used to follow the structural changes of the proteins, including fluorescence (Isenman et al., 1979; Vanhove et al., 1998), phosphorescence (Mersol et al., 1993; Mazhul et al., 2003), circular dichroism (Kelly and Price, 2000), infrared spectroscopy (Fabian and Naumann, 2004; Ma and Gruebele, 2005),
7 AU1 Protein folding 7 nuclear magnetic resonance (Englander and Mayne, 1992; Kamatari et al., 2004), and mass spectroscopy (Miranker et al., 1996; Konermann and Simmons, 2003). Both theoretical and experimental results indicate that a single reaction coordinate in general is not enough to describe protein folding, and multiple reaction coordinates must be used. (Becker and Karplus, 1997; Ma and Gruebele, 2005). Finding the adequate reaction coordinates for protein folding is not straightforward. Several kinetic and thermodynamic coordinates have been used to describe the nativeness of a given protein state. Thermodynamic reaction coordinates use a thermodynamic parameter, e.g., Gibbs free energy and/or entropy, to define the distance between the native state and the actual state of a protein. The kinetic reaction coordinate measures the time needed for the protein to reach the native state from a given starting state. An important thermodynamic reaction coordinate often used to describe the folding process is the number of native contacts present in the conformation, which proved useful in interpreting simple folding processes. Thermodynamic reaction coordinates, however, are inadequate to describe folding dominated by kinetic traps because they completely ignore the Gibbs free energy barriers separating the different states (Sali et al., 1994; Wolynes et al., 1995; Chan and Dill, 1998). The Gibbs free energy barrier to folding is determined by the unfavorable loss in configurational entropy upon folding and the gain in stabilizing native interactions. Starting from the unfolded protein, the polypeptide chain has to fold partially in order to bring together the residues that need to form the contacts stabilizing the native structure. The constrained polypeptide chain has smaller entropy, which means higher Gibbs free energy. As native contacts form, the enthalpy term decreases, the protein is stabilized. The rate limiting step in the folding process is the formation of the transition state, i.e., the conformation that has the highest Gibbs free energy on the folding pathway (Chan and Dill, 1998; Lindorff Larsen et al., 2005). The simplest model for unfolding and refolding involves a single cooperative folding step, in which the unfolded (U) and folded (F) states of the protein interconvert: U F. This simple mechanism well describes the folding of several small proteins (Gillespie and Plaxco, 2004). The formation of a contact between two residues in the transition state involves an entropic cost which depends on the sequence separation of the two residues: the longer the chain between them the greater the entropic cost, and this entropic cost contributes to the height of the Gibbs free energy barrier between the unfolded and the folded state. If nonnative interactions play a marginal role in the transition state, it is possible to estimate the folding kinetics from the average sequence separation of the contacts in the native structure (Plaxco et al., 1998; Grantcharova et al., 2001; Zarrine Afsar et al., 2005). Intermediate structures were observed to accumulate during the folding of many proteins (Englander, 2000). Such intermediate states are trapped structures that have low Gibbs free energy. Mass action models that involve one or more intermediate states were constructed to explain more complex folding kinetics. Mass action models distinguish between on pathway and off pathway intermediates depending on whether the intermediate is on the folding pathway between the unfolded and native states (Baldwin, 1996). Off pathway intermediates often correspond to misfolded structures that must completely or partially unfold to allow formation of the native fold (Evans et al., 2005). The most general theory of protein folding is a statistical mechanical model that uses the concept of the energy landscape, which is discussed in more detail later. Here we only want to clarify that in the energy landscape view, there is no clear distinction between on and offpathway intermediates. Folding mechanisms involving these two types of intermediates only differ in the distribution of traps on a Gibbs free energy landscape (Onuchic and Wolynes, 2004; Jahn and Radford, 2005). Energy landscape theory of protein folding predicts that the enthalpic and the entropic term of the transition state Gibbs free energy can cancel each other, leading to folding that lacks an activation barrier (Bryngelson et al., 1995). Although such downhill folding has indeed been found experimentally, it seems to be atypical, probably because such proteins are evolutionary unfavorable (Yang and Gruebele, 2004a). The probable reason for this is that proteins that fold downhill lack the barrier that prevents partial unfolding, and thus become more prone to aggregation (Yang and Gruebele, 2003; Gruebele, 2005). The folding process is usually not restricted to a narrow path in conformational space. Each molecule may follow a different path, and the molecule population can be very heterogeneous. Mass action models are inadequate to describe these folding processes, and the more complex energy landscape view must be used.
8 8 Protein folding Heterogeneous folding ensembles give rise to stretched folding kinetics and/or probe dependent observation of the kinetics (Sabelko et al., 1999; Osvath et al., 2003; Ma and Gruebele, 2005; Osvath et al., 2006). 4 Molten Globules and Other Compact Denatured States As mentioned previously, Levinthal (1969) postulated the existence of folding pathways in In an effort to find the specific intermediates along the folding pathway, folding studies were performed on several globular proteins (Wong and Tanford, 1973; Kuwajima et al., 1976; Robson and Pain, 1976; Nozaka et al., 1978). Equilibrium intermediates were reported but it was not clear whether these are related to the specific folding intermediates. However, the equilibrium intermediates characterized in different proteins were found to be remarkably similar to each other: all of them had native like secondary structure and were compact, but lacked a specific tertiary structure. In 1983, Ohgushi and Wada (Ohgushi and Wada, 1983) proposed that the equilibrium intermediates belong to a common physical state of globular proteins, and they termed this the molten globule state. After Kuwajima and coworkers (1985) and Ikeguchi and coworkers (1986) had shown that the molten globule state of a lactalbumin is identical to its transient folding intermediate, researchers started to study molten globules with renewed interest. Molten globule states could be generated using mild denaturing conditions (low or high ph, moderate concentrations of denaturants, high temperature, and various salts) in about different proteins such as a lactalbumin, carbonic anhydrase B, b lactamase, ribonuclease A, T4 lysozyme, cytochrome c, apomyoglobin, and staphylococcal nuclease (Ptitsyn, 1995). 4.1 The Structure of the Molten Globule The common characteristics of the molten globule state as described by Kuwajima (Arai and Kuwajima, 2000) are (1) the presence of a significant amount of secondary structure, (2) the absence of most of the specific tertiary structure produced by the tight packing of side chains, (3) compactness of the protein molecule with a radius of gyration % 30% larger than that of the native state, and (4) the presence of a loosely packed hydrophobic core that increases the hydrophobic surface accessible to solvent. The experimental techniques typically used to detect molten globules are (1) far and near UV CD spectra that detect the secondary and tertiary structures of a protein, (2) hydrodynamic methods such as viscosity measurement and molecular sieve chromatography that determine the molecular size of the protein, and (3) hydrophobic dye (typically ANS, 8 anilino naphtalene 1 sulfonate) binding experiments that detect the formation of a loose hydrophobic core and estimate the extent of hydrophobic area exposed to the solvent. These techniques, however, only provide information about the average structural properties of the protein molecule. More advanced techniques that provide a more detailed picture about the molten globule were not available until the 1990s. Therefore, the exact nature of molten globule structure had been a matter of some debate. > Figure -2 shows an illustration of two different models for the molten globule. The traditional view of the molten globule (Shakhnovich and Finkelstein, 1989; Ptitsyn, 1992) assumes that the backbone of the polypeptide chain essentially has a (fluctuating) native like fold and the disorder is in the side chains. However, this view was criticized on the basis of thermodynamic arguments and energy landscape theory (Dill et al., 1995; Privalov, 1996). Privalov (1996) argued that molten globule states are either misfolded structures or states where one portion of the structure is already folded and another one is still unfolded. Dill and coworkers (1995) stated on the basis of simulations of simplified models that backbone and side chain degrees of freedom are strongly coupled, therefore a state where the backbone is ordered and the side chains are disordered is unlikely. The new experimental techniques developed in the 1990s, such as stopped flow circular dichroism, pulsed hydrogen exchange, X ray scattering, and mutational approaches, have allowed characterization of the molten globule states in great detail (Hughson et al., 1990; Dobson, 1994; Carlsson and Jonsson, 1995; Carra and Privalov, 1996; Dabora et al., 1996; Dyson and Wright, 1996; Kataoka and Goto, 1996; Kuwajima, 1996; Song et al., 1998; Chakraborty et al., 2001; Demarest et al., 2001; Ramboarina and
9 Protein folding 9. Figure -2 Illustration of different models for the structure of the molten globule. The backbone of the protein is represented by a thick line and side chains are shown as pieces of various shapes hanging from the backbone. (a) The native state of the protein, with all the buried side chains fitting closely together like the pieces of a jigsaw puzzle. (b) The side chain molten globule model: the fold of the backbone is native like but the side chains are only loosely packed. (c) One half of the molecule is fully folded, with the specific side chain interactions present, while the other half is completely unfolded Redfield, 2003; Redfield, 2004). These studies confirmed the view that the molten globule state is indeed more heterogeneous than previously thought: one part of the structure is more organized and native like while other portions are less organized. However, there is a great deal of variety among different proteins in the exact structure of the molten globule state. 4.2 The Role of the Molten Globule in Folding The study of molten globules was strongly motivated by the idea that the molten globule state could be identical to the transient intermediate during folding. This notion was supported by kinetic circular dichroism measurements on a lactalbumin (Ikeguchi et al., 1986). However, later theoretical studies
10 Protein folding suggested that the experimentally observed folding intermediates may not be productive but kinetically trapped, misfolded species (Dill and Chan, 1997). Careful kinetic measurements during the refolding of several proteins including interleukin 1b, staphylococcal nuclease, and apomyoglobin (Heidary et al., 1997; Walkenhorst et al., 1997; Jamin and Baldwin, 1998; Maki et al., 1999) have provided firm evidence that, contrary to the theoretical predictions, the molten globule state is the productive on pathway folding intermediate in most cases. An attempt to resolve the apparent contradiction between theory and experiment introduced the hierarchical folding model (Arai and Kuwajima, 2000) where the folding of a protein occurs in two stages: (1) formation of the molten globule state from the fully folded state and (2) formation of the native state from the molten globule state. 5 Simulations of Protein Folding and Unfolding Computational models and simulations have greatly advanced our understanding of protein folding. The information from such studies is complementary to experiments. In fact, there is a synergy between theory and experiment: theory provides testable models and experiments provide the means to test and validate the models. The outcome from this combination is a much richer view of the system in question than what either approach could provide alone. In particular, simulations can help identify or predict transition and intermediate states along the folding pathway, provide predictions of the rate of folding and in some cases, predict the final, folded structure. Simulating protein folding presents a significant challenge. Small proteins typically fold in the several microseconds to seconds timescale; detailed atomistic simulations, however, are currently limited to the nanosecond to microsecond regime. Therefore, simulation of folding requires either simplified models or special sampling methods, both of which introduce new approximations. 5.1 All Atom Models AU2 The most straightforward approach to simulating protein folding and unfolding is to use an all atom model with a force field like AMBER or CHARMM and apply traditional molecular dynamics simulation. These force fields describe the energies of the deformations of covalent bonds as well as van der Waals interactions, charge charge interactions, hydrogen bonds, and so on. Traditional molecular dynamics numerically solves Newton s equations of motion by calculating the forces acting on atoms and computing accelerations, velocities, and atomic displacements. Temperature is assigned to the system by assigning appropriate velocities to the atoms. To simulate unfolding, the simplest method of increasing sampling is to increase the temperature of the simulation to 498 K or more. At these temperatures, the native structure of the protein is usually lost within a few nanoseconds. This technique has been applied to numerous examples, including bovine pancreatic trypsin inhibitor (Kazmirski and Daggett, 1998a), lysozyme (Kazmirski and Daggett, 1998b), myoglobin (Tirado Rives and Jorgensen, 1993), barnase (Wong et al., 2000), ubiquitin (Alonso and Daggett, 1998), the SH3 domain (Tsai et al., 1999), etc. Features of the unfolding process such as the transition state ensemble or the unfolded ensemble have shown remarkable agreement with experimental results (Day and Daggett, 2003). A direct simulation of folding is a much harder problem. The longest continuous all atom molecular dynamics simulation so far is still the 1 ms simulation of villin headpiece subdomain performed by Duan and Kollman in 1998 (Duan and Kollman, 1998). In this simulation, the chain sampled a large number of conformations after initial collapse, and near native structures appeared but the true native conformation was not reached. Although processor speeds have dramatically increased since 1998, computational power is still insufficient to allow a meaningful all atom simulation of the entire folding process. Even if the native state could be reached in a single trajectory, multiple simulations would have to be performed to construct a believable folding pathway. An interesting alternative is the massively distributed method employed in the Folding at Home project (Pande et al., 2003). A large number of computers run the simulation in parallel. As soon as a
11 Protein folding 11 transition is detected (as a momentary surge in the heat capacity) in one of the simulations, all computers receive a copy of the posttransition conformation and the simulation is continued until the native conformation is reached. Although this approach has been criticized as being flawed (Fersht and Daggett, 2002), it has been successfully applied to fold several small, fast folding proteins (Snow et al., 2004; Sorin and Pande, 2005). 5.2 Simplified Models In simplified (coarse grained) models (Dokholyan, 2006), effective particles (beads) represent amino acids or groups of atoms. An empirical potential function, usually derived from protein structures, is used to describe the interaction between these beads. The shape of this potential is often very simple, such as a square well function. In many simplified models, the positions of the beads are restricted to points on a lattice. Perhaps the most minimal model is the one where there are only two types of amino acids: hydrophobic and polar, and the chain is restricted to a two dimensional lattice (Dill et al., 1995). Smaller on lattice model proteins allow an exhaustive enumeration of all possible states of the given system. This approach allows a complete thermodynamic description of the phase space and has greatly enhanced our understanding of protein folding. The funnel view and the concept of energy landscapes (see > Sect. -6) arose directly from the exhaustive sampling allowed by these minimal models (Bryngelson et al., 1995). In the case of larger, more complex simplified models, exhaustive enumeration of states is not possible. Monte Carlo is a common choice for simulating simplified models. In Monte Carlo simulation, small moves are generated randomly and accepted or rejected based on the energy of the new conformation. This is often performed in the framework of advanced sampling schemes such as Replica Exchange Monte Carlo, where several replicas of the system are simulated at various temperatures (Kihara et al., 2001; Pokarowski et al., 2003). A more recent simulation approach, termed discrete (or discontinuous) molecular dynamics (DMD) (Smith and Hall, 2001; Ding and Dokholyan, 2005), extends the accessible simulation time by using long integration time steps with approximated energy functions. Simple models like this start showing remarkable success. Recently, Trp cage, a 20 residue miniprotein has been folded to a conformation very close to the experimental structure (Ding et al., 2005a). It is believed that this technique will be applicable to larger proteins. 5.3 Multiscale Modeling Approaches using simplified, coarse grained models can be combined with fine grained, all atom simulations in what is called multiscale molecular modeling. Bradley and coworkers (2005) reported highresolution structure prediction for proteins up to 85 residues using a multiscale approach that sampled low and high resolution conformations. DMD simulations combined with all atom, traditional molecular dynamics have been used to simulate the formation of a b helix (Khare et al., 2005) and to identify the transition state of the SH3 domain (Ding et al., 2005b). Although it remains to be seen whether the force fields used are transferable to larger proteins, multiscale modeling has the potential to break the 1 ms barrier of direct folding simulations. 6 Free Energy Landscapes of Proteins Energy landscapes are mathematical devices that help us understand the microscopic behavior of a molecular system (Bryngelson et al., 1995). An energy landscape of a system with n degrees of freedom is an energy function F(x) ¼ F(x 1, x 2,..., x n ) where x 1,..., x n are variables specifying the microscopic state of the system (Dill, 1999). In the case of a protein, x 1, x 2,..., x n can be, for instance, all the dihedral angles of the chain, thus specifying a single conformation of the protein. F(x) then is usually defined as the free
12 12 Protein folding energy of the protein in the given conformation, where the entropic part of the free energy comes from all possible solvent configurations. Thus, F(x) is the free energy of a microstate, not a macrostate, because it does not include the chain conformational entropy. The stable conformation of the protein can be found by determining the set of values x 1, x 2,..., x n (i.e., the conformation) that gives the minimum value of the freeenergy function. Although energy landscapes are, by definition, high dimensional surfaces, they are often pictured as a surface in three dimensions. In these pictures, the vertical axis represents the free energy and the horizontal axes represent the conformational degrees of freedom of the polypeptide chain. Random heteropolymers, such as a random sequence of amino acids, have a very rugged energy landscape with many local minima (Plotkin et al., 1996). Systems like this easily get trapped in one of the local minima and usually do not have a well defined, single, stable conformation. Real proteins, however, are not random sequences; evolution has optimized their sequences so that they quickly and efficiently fold into a well defined three dimensional conformation (Onuchic and Wolynes, 2004). In a real protein, most of the interactions that can form between parts of the chain are mutually supportive and cooperatively lead to a low energy structure which is therefore minimally frustrated. This principle of minimal frustration (Bryngelson and Wolynes, 1987), gleaned from simplified models of proteins and the theory of spin glasses, led to the realization that the energy landscape of a real protein should be shaped like a funnel (> Figure -3).. Figure -3 Schematic representation of a funnel shaped energy landscape. The width of the funnel represents the conformational freedom of the chain. The vertical axis represents the free energy; as free energy decreases, the nativeness of the chain increases. Denatured (unfolded) states are at the top of the funnel while the native state is the global minimum. There is some ruggedness in the energy landscape near the native state A funnel shaped energy landscape means that the free energy of a structure depends on how close it is to the native state: the closer it is to the native structure the lower its free energy. Also, the top of the funnel, representing the nonnative states, is wide (the conformational entropy is high), and it narrows as one gets closer to the bottom: near native states represent more compact conformations, and therefore have low conformational entropy. The fact that the energy landscape of a real protein is essentially funnel shaped has several important consequences (Wolynes, 2005). First, the native structure should be robust against mutations (Nelson and Onuchic, 1998). A point mutation represents a small perturbation to the energy landscape; therefore, the basic shape of the funnel and the location of the global minimum cannot change much: the mutant protein will fold into essentially the same structure as the wild type protein. This structural robustness underlies the most commonly used method of predicting structures: homology modeling (Zhang and Skolnick, 2004).
13 Protein folding 13 If sequence analysis shows two sequences to be evolutionarily related, one can be almost certain that the two structures are essentially the same. Another important consequence of the funnel shape is that the folding rate and folding mechanism of a protein is largely determined by its native state topology (Baker, 2000). Assuming that the energy landscape is a perfect funnel, its shape is completely determined by the topology of the native state, and this shape completely determines the folding mechanism. Of course, the energy landscape of a real protein is never a perfect funnel: it is always somewhat rugged, and the exact layout of the small heaps and valleys on the landscape will influence the details of the folding mechanism. Still, for many proteins, the landscape is close enough to a perfect funnel to allow the development of methods that can successfully predict various features of the folding mechanism from just the native structure as input (Alm and Baker, 1999; Galzitskaya and Finkelstein, 1999; Munoz and Eaton, 1999). The folding rate can be predicted from just the contact order (the average sequence separation between residues that make contacts in the native structure). Using a perfectly smooth, funnel shaped energy function based on the Go model (Go, 1983) (where only native contacts contribute to the free energy), transition state structures, folding nuclei, and F values have been predicted for several small proteins such as CheY, CI 2, barnase (Galzitskaya and Finkelstein, 1999), l repressor, and SH3 domain (Alm and Baker, 1999), with surprisingly good agreement with experiments. The concept of energy landscapes also helps us understand some of the more complex protein folding scenarios. The folding of many proteins involves slow steps, bottlenecks, and multiple, kinetically distinguishable stages (Wolynes et al., 1995). These phenomena can be rationalized by noticing that energy landscapes are often not perfect, smooth funnels but are rugged and bumpy (Dill, 1999). Slow steps in the folding process can arise from climbing an uphill slope after being trapped in local minima. This scenario may appropriately describe the folding of b lactoglobulin, a predominantly b sheet protein, which passes through a helical phase as it folds (Hamada et al., 1996). Because the landscape view is a microscopic view of the folding process, each molecule may follow a different route on the energy landscape and may encounter different obstacles in its way. For example, in the folding of hen egg white lysozyme, a subpopulation of the molecules folds fast while another subpopulation folds quite slowly (Radford et al., 1992). In the landscape view, it is easy to see how molecules starting from one region of the top of the funnel may ski down unhindered while molecules starting from another region may get trapped behind a mountain range. Bottlenecks do not always involve uphill climbing though, they can be entropic barriers too; in this case, the aimless search for a downhill route on a large, level field will limit the rate of folding (Dill and Chan, 1997). This example shows that bottlenecks are typically ensembles of widely different conformations rather than well defined, single conformations (Chan and Dill, 1998). The landscape view of protein folding also gives us a clue to chaperone function: to get a misfolded protein to fold correctly, no specific recognition is needed; it is sufficient just to move it to the top of the funnel where it can restart the downhill search for the folded conformation (Chan and Dill, 1996; Todd et al., 1996). 7 Traps on the Folding Pathway The folding protein faces several obstacles in translating the information encoded in the amino acid sequence into the three dimensional construct of the biologically active structure. The cis trans isomerization of peptide bonds and the formation of disulfide bridges are slow steps that can form bottlenecks in the protein folding reaction (Balbach and Schmid, 2000; Creighton, 2000). Under certain circumstances, the two processes can be linked and the formation of the correct disulfide bonds is facilitated in the presence of peptidyl prolyl cis trans isomerase (Schonbrunner and Schmid, 1992). 7.1 Backbone Isomerization The peptide bond between nonproline amino acids is much more stable in the trans than in the cis conformation (> Figure -4a). The difference in stability arises from interactions between the C a i and H a i atoms with the C a i þ 1 and H a i þ 1 atoms, from electrostatic interactions between the O i and C i þ 1 atoms
14 14 Protein folding. Figure -4 (a) The cis and trans states of nonprolyl peptide bonds. (b) The cis and trans states of prolyl peptide bonds and from conformational entropy (Zimmerman and Scheraga, 1976; Wedemeyer et al., 2002). The large difference in the stability of the two isomers keeps the trans form 0 to 00 times more populated than the cis isomer (Ramachandran and Mitra, 1976; Jorgensen and Gao, 1988; Scherer et al., 1998). Because of the high number of peptide bonds in the protein, the fraction of protein molecules that have cis peptide bonds in the denatured ensemble can be significant. Since isomerization of peptide bonds is a slow process, nonnative isomers of peptide bonds can cause slow phases in protein folding. It has been shown that cis trans isomerization of nonprolyl peptide bonds can give rise to significant slow folding phases (Eyles, 2001). Nonprolyl cis peptide bonds are energetically unfavorable and are very rare in the native structures of proteins (Stewart et al., 1990). Isomerization of the trans peptide bond accumulated in the denatured state into the native cis conformation of these proteins can become the rate limiting step of the folding reaction (Odefey et al., 1995). Proline residues are a much more common source of kinetic complications during folding. The X Pro peptide bond (where X can be any amino acid) is the only peptide bond for which the stability of the cis and trans conformations is comparable. The cis trans isomerization of X Pro peptide bonds is a widely recognized rate limiting factor, which can induce additional slow phases in protein folding (Brandts et al., 1975; Wedemeyer et al., 2002). The otherwise strong preference of the peptide bond to be in trans state does not apply to the X Pro bond, mostly because of the steric symmetry between the C a and the C g atoms of proline (> Figure -4b). The entropy change accompanying the cis trans transition and the electrostatic interactions between the O i and C i þ 1 atoms are also different in the X Pro bonds than in other peptide bonds (Zimmerman and Scheraga, 1976; Wedemeyer et al., 2002). As a result, the trans conformation of X Pro bonds is only marginally more stable than the cis conformation in unfolded polypeptides. In the unfolded protein, a mixture containing both conformations is present, with % 30% of the X Pro bonds in the cis state (Cheng and Bovey, 1977; Grathwohl and Wuthrich, 1981). In the native conformation of a protein, intraprotein interactions will stabilize either the cis or the trans isomer for most of the X Pro bonds. The molecules that contain incorrect isomers in the unfolded ensemble must undergo isomerization during the folding reaction (Kiefhaber et al., 1990a, b; Texter et al., 1992). Consequently, the presence of proline residues leads to a number of proline isomerization reactions that must occur before folding can complete. Both experimental and theoretical findings show that there is a high energy barrier for isomerization (16 20 kcal/mol in model compounds) of X Pro bonds (Balbach and Schmid, 2000; Kang and Choi, 2004). This results in characteristic times of 00 s for the
15 Protein folding 15 conformational flipping at room temperature (Brandts et al., 1975). Mutagenesis studies have indeed shown that specific proline residues can often be assigned to slow recovery phases from misfolded states (Evans et al., 1987; Kelley and Richards, 1987; Wood et al., 1988; Herning et al., 1991; Kiefhaber et al., 1992; Wu and Matthews, 2002; Street et al., 2005). The high energy barrier for isomerization reflects a partial double bond character of the X Pro bond (Balbach and Schmid, 2000). In vivo, enzymes (e.g., peptidyl prolyl cis trans isomerases) facilitate folding to the native structure by lowering the energy barrier for isomerization (Schmid et al., 1993; Stein, 1993; Gothel and Marahiel, 1999; Shaw, 2002). Laser induced temperature jump measurements indicate that the role of proline residues in protein folding is more complex. Prolines can have opposite effects on the slow and fast steps of the folding kinetics. The presence of proline residues leads not only to additional slow phases, but also modifies the millisecond and sub millisecond dynamics of the protein. The X Pro bonds do not isomerize on the millisecond timescale. This increased backbone rigidity can speed up the fast folding steps of the ensembles that contain the proline in the native like isomerization state (Osvath and Gruebele, 2003). 7.2 Formation of Disulfide Bridges Formation of the correct native like disulfide bridges can also hinder protein folding. Disulfide bridges between pairs of cysteines are part of the native structure for many proteins. The formation of disulfide bridges is a prerequisite of the proper folding and biological function in these proteins. Disulfide bonds increase the thermodynamic stability of the native structure by establishing conformational constraints within the protein (Creighton, 2000). It has been shown that several proteins start to fold during synthesis and disulfide bond formation begins in the emerging chain (Bergman and Kuehl, 1979; Peters and Davidson, 1982; Braakman et al., 1991; Braakman et al., 1992). Folding and disulfide bridge formation is completed after the end of the translation (Wedemeyer et al., 2002). Depending on the number of the cysteine residues in a polypeptide sequence and on the native structure, misfolding can occur owing to errors in disulfide pairing. Disulfides formed randomly in early folding intermediates may cause kinetic complications during the later folding steps (Creighton, 1979; Konishi et al., 1982). Repairing of the errors is part of the folding process and occurs in a trial and error manner (Chatrenet and Chang, 1992; Schwaller et al., 2003). Since disulfide bonds constrain the free movement of the polypeptide chain, the formation of native disulfide pairs can also prevent correct folding. It has been shown that native like disulfide bridges formed too early during folding must be broken up to allow the folding process to continue, and they can reform at a later stage (Creighton and Goldenberg, 1984). Formation of disulfide bonds requires the presence of an oxidizing agent or a disulfide reagent such as glutathione or dithiothreitol. When a disulfide reagent is present, disulfide bond formation occurs by a thiol disulfide exchange reaction. This is a two electron redox reaction in which a disulfide reagent takes one electron from each cysteine thiolate, and a disulfide bond is formed (> Figure -5) (Wedemeyer et al., 2000). The first step is the result of a nucleophilic attack by a cysteine thiolate on the disulfide reagent. The rate of this reaction is determined by the reactivity of the cysteine thiolate, the presence and nature of the disulfide reagent, ph, temperature, ionic strength, and cosolvents. In the second step, the mixed disulfide is broken up by another nucleophilic attack by the second cysteine thiolate. As a result, the external thiol is replaced by a protein thiolate and an intraprotein disulfide bond is formed. The prerequisite of the second step is a conformational change in the protein that brings the two cysteine residues together, thus the redox reaction is coupled to the folding of the protein (Creighton, 1997; Bulaj, 2005). The disulfide pairing during in vivo folding is helped by protein disulfide isomerases (Bulleid and Freedman, 1988; Freedman, 1989; Ellgaard and Ruddock, 2005). These enzymes correct the disulfide pairing mistakes by catalyzing the reshuffling of disulfide bridges so that the native pairing can emerge (Gilbert, 1997). The process leads to a biologically active structure, which is resistant to further rearrangement (Walker and Gilbert, 1997).
16 16 Protein folding. Figure -5 Scheme of the redox reaction that leads to the formation of disulfide bonds in proteins 8 Transition States on the Folding Pathway To gain insight into the folding mechanism, it is essential to characterize the energy landscape of the protein. This includes the determination of the structures and energies of the protein in the traps and the transition states of the landscape (Gruebele, 2002). The local minima of the energy landscape act as traps that are responsible for the buildup of kinetic intermediate states. Since the protein accumulates in measurable quantities in the intermediate states, the intermediate structures can be experimentally studied (Roder et al., 2000). Transition states are usually not populated in detectable quantities. The only way to gain information about the structural and energetic properties of the transition states is through kinetic studies of the folding reaction (Daggett and Fersht, 2000). The kinetics of the transition between different states is determined by the escape rates from local minima of the landscape. In simple cases, protein folding can be described as a diffusive process over a barrier determined by the energy landscape. For barriers that are larger than the thermal energy k B T, the folding rate predicted by transition state theory can be calculated by a Kramer like equation (Onuchic et al., 1996) k ¼ n exp DG {. k B T : Here k denotes the rate of the conformational transition, n is a coefficient that depends on a number of things including the shape of the barrier and solvent viscosity, DG { is the Gibbs free energy height of the transition state, k B is Boltzmann s constant, and T is the absolute temperature. Transition state theory thus allows us to determine the Gibbs free energy of the transition state using simple kinetic measurements. Gaining information about the structure of the transition state is a more intricate problem. To date, the only way to learn about the structure of the transition state is F value analysis, a method that uses site directed mutagenesis to map out the residue residue contacts present in the transition state (Matthews and Hurle, 1987; Fersht et al., 1992). The calculated F value compares the change in the folding rate and the change in the stability of the protein caused by a specific mutation (Clementi et al., 2000) F ¼ ð RT lnðk mut =k wt ÞÞ DGmut DG wt : Here R is the gas constant, T is the absolute temperature, and k mut and k wt are the folding rates of the mutant and wild type protein, respectively. DGmut and DGwt represent the stabilities of the mutant and of the wild type protein, i.e., the Gibbs free energy difference between the folded and the unfolded state. If the unfolded state is assumed to be a randomly fluctuating chain, its free energy can be taken as the solvation free energy. However, it has been shown that denatured proteins can have some residual structure (Smith et al., 1996; McCarney et al., 2005), and a determination of their free energies becomes more complicated. For the purposes of F value analysis, however, we can just use the Gibbs free energy of the unfolded state as a reference, i.e., define it as zero. The expression for F can be simplified if the following two conditions are met: (1) The folding rate can be calculated from the activation energy using a Kramer like exponential dependence, thus folding can be treated as a simple cross barrier diffusive reaction and (2) The folding
17 Protein folding 17 mechanism is not altered significantly by the mutation, thus the parameter n reflecting the barrier shape and the configurational diffusion of the protein is insensitive to the mutation. If the above assumptions hold, the new expression for F is:. F ¼ DG mut { DG wt { DGmut DG wt ; where DG { mut and DG { wt denote the activation Gibbs free energies for the mutant and the wild type protein, respectively. This way, the F value compares the Gibbs free energy change introduced by the mutation in the transition state and in the native state (> Figure -6).. Figure -6 Schematic representation of the Gibbs free energies important in the F value analysis In order to be able to extract structural information from F value analysis, two more conditions have to be met (Fersht et al., 1992; Clementi et al., 2000): 1. The folding pathway is not altered significantly by the mutation, thus the intermediate and transition states are the same for the mutant and the wild type protein. 2. The folded region of the transition state has a native like structure. The above conditions were found to be true for several proteins, and F value analysis was used to characterize transition states and short lived intermediates of several folding pathways (Matouschek et al., 1990; Serrano et al., 1992; Grantcharova et al., 1998; Villegas et al., 1998; Fulton et al., 1999; Goldenberg, 1999; Raschke et al., 1999; Crane et al., 2000; Bulaj and Goldenberg, 2001; Capaldi et al., 2002; Paci et al., 2003; Hubner et al., 2004; Lindorff Larsen et al., 2004; Hubner et al., 2005). Usually, a detailed analysis of the mutation is necessary to identify the contacts disrupted in the mutant structure. The use of multiple mutations of the same residue can increase the accuracy of the prediction of the transition state structure (Matouschek et al., 1995). F values are normally between 0 and 1, but negative F values corresponding to mutations that speed up the folding kinetics have also been observed (Yang and Gruebele, 2003; Yang and Gruebele, 2004b). The F value is a valuable tool in detecting native contacts in transition state structures, but in general, it is not proportional to the extent or strength of the native contact appearing in the transition state. This means that structural information can unambiguously be assigned only to F ¼ 0 and F ¼ 1 (Fersht and Sato, 2004). A F value close to 1 indicates that the free energy change introduced by the mutation is almost identical for the transition state and the native state. This implies that the mutated residue already forms its native
18 18 Protein folding contacts in the transition state. A F value close to 0 indicates that the height of the Gibbs free energy barrier of the transition state was not altered by the mutation. In this case, the mutated residue does not form native contacts in the transition state, and the environment of the mutated residue is probably denaturedlike (Fersht and Sato, 2004). It has been shown that for deletions of small hydrophobic residues, the F value is roughly proportional to the extent of the native contact formation. Therefore, these residues are preferred as targets for mutations in F value analysis (Fersht and Sato, 2004). 9 Folding of Multidomain and Multi Subunit Proteins 9.1 Multidomain Proteins A domain is a part of the polypeptide chain that forms a compact globular substructure with more interactions within itself than with other parts of the chain (Janin and Wodak, 1983). Most proteins longer than about 200 to 250 residues consist of several domains. When investigating the role of domains in folding, the first question to answer is whether a domain can fold by itself. Isolated domains can be produced by limited proteolysis or genetic engineering, and their folding can be studied by the same experimental methods as used for the whole proteins. Autonomous folding was demonstrated for the domains of several multidomain proteins including tryptophan synthase, b lactamase, aspartokinase homoserine dehydrogenase, plasminogen, phosphoglycerate kinase (Jaenicke, 1987), and several other domains (Sharma et al., 1990; Herold et al., 1991; Shoelson et al., 1993; Williams and Shoelson, 1993; Jecht et al., 1994). In many cases, it was shown that the stability of the domains in isolation is close to the stability when the other domains are also present (Garel and Dautry Varsat, 1980; Muller and Garel, 1984; Novokhatny et al., 1984; Jaenicke, 1987; Tsunenaga et al., 1987; Rudolph et al., 1990; Missiakas et al., 1992). Interestingly, the rate of folding of isolated domains was found to be greater or the same as that of the same domains integrated within the intact protein (Teale and Benjamin, 1977; Dautry Varsat and Garel, 1981; Blond and Goldberg, 1986; Tsunenaga et al., 1987; Missiakas et al., 1992). This suggests that the presence of the rest of the chain slows down the folding of a domain, e.g., by forming unfavorable interactions with it. The folding kinetics of multidomain proteins is usually complex, showing an initial rapid phase characterized by large changes in several physical parameters (fluorescence, UV absorption, circular dichroism, etc.), followed by a second, slower phase by much smaller changes in the physical parameters (Jaenicke, 1987; Jaenicke, 1999). The intermediate that accumulates after the initial rapid phase appears largely folded but lacks some properties of the native structure: it is more labile to proteolysis, is not recognized by some antinative antibodies, and lacks catalytic activity. These properties only appear after the second, slower stage of folding. The findings suggest that the individual domains fold in the rapid phase of the folding process, and the second, slow step is the pairing or association of the already folded domains. This is also supported by the fact that in several cases the rate of folding was found to be inversely proportional to solvent viscosity, implying that movement of large, globular species is involved in the rate limiting step of the folding process (Vaucheret et al., 1987; Chrunyk and Matthews, 1990). Also, in tryptophan synthase, single mutations in each domain were found to decrease the folding rate while the double mutant was able to fold at the same rate as the wild type protein, suggesting that the rate limiting step of folding involves the formation of the interface between the two domains (Tsuji et al., 1993). Thus, the folding process can be described by the following general scheme Fast domain folding Unfolded! Slow domain pairing Intermediate with folded but unpaired domains! Native The existence of an intermediate with folded but unpaired domains has important consequences. If the protein concentration is sufficiently high, the domain pairing step may occur between domains from two identical molecules, leading to the formation of a dimer instead of a monomer. The domains make the same interactions in the dimer as in the monomer, but the interactions are formed intermolecularly rather than intramolecularly. Higher order oligomers may also form and the process may lead to fibril formation or
19 Protein folding 19. Figure -7 The folding of a hypothetical two domain protein. The unfolded chain folds into two domains that are not yet paired. The fate of this intermediate depends on the protein concentration and the solvent conditions: it may either form the native state; it may associate with another chain to form a domain swapped dimer; or several chains may aggregate or form a fibril aggregation (see > Figure -7). The exchange of domains between identical molecules is actually a special case of a range of phenomena termed domain swapping (Bennett et al., 1994). The term describes situations where two or more proteins exchange part of their structure (not necessarily whole domains) to form intertwined oligomers (Rousseau et al., 2003). Domain swapping has been implicated in amyloid fibril formation, although not all models of amyloid formation involve domain swapping (Nelson and Eisenberg, 2006). 9.2 Multi Subunit Proteins Almost all large proteins are formed by the association of subunits. There are several evolutionary advantages to forming multimers: regulation of catalytic activity through cooperativity between the subunits, generation of new functions and enzymatic activities, formation of large structures, increasing
20 20 Protein folding protein stability, and facilitating the folding process. Subunit subunit interfaces are very diverse regarding the nature and distribution of inter subunit interactions. About one third of the interfaces have a large and contiguous hydrophobic patch surrounded by a ring of inter subunit polar interactions; the remaining two thirds show a mixture of small hydrophobic patches, polar interactions, and water molecules scattered over the interface area. There are several experimental techniques to study the folding and association of multi subunit proteins. Depending on the strength of the binding between the subunits, the experimentalist can attempt to produce folded but dissociated subunits under equilibrium conditions, using techniques such as dilution, cold dissociation, chemical modification, ligand induced dissociation, mildly denaturing conditions, or elevated pressure (Jaenicke and Lilie, 2000). If unfolded monomers cannot be obtained, folding and association should be studied as coupled processes. Several methods can be used to monitor the association state of the protein after starting reassociation of the (folded or unfolded) subunits. By rapid chemical cross linking, snapshots can be taken during the reassociation process and investigated by gel electrophoresis. Other methods include hybridization with isoenzymes or modified subunits and measurement of relative reactivation. There is also a wide range of biophysical methods to monitor and/ or measure the thermodynamic parameters of protein protein interactions (Lakey and Raggett, 1998), including surface plasmon resonance, isothermal titration calorimetry, fluorescence energy transfer, mass spectrometry, light scattering, and high pressure liquid chromatography. After collecting the timedependent data, a reaction equation can be fit and a reaction scheme established. The assembly of multi subunit proteins can usually be described as a series of unimolecular (isomerization) and bimolecular (association) steps 4M u! 4M! 2D 0! 2D! T 0! T; where M represents a monomeric, D a dimeric, and Ta tetrameric state, and the subscript u represents the unfolded state. Are folding and association separate events, or are they more or less coupled? The traditional view is that monomers assume a near native conformation before binding to their partners. In recent years, however, several proteins have been found that are intrinsically unstructured as monomers and only fold upon binding to DNA, RNA, a membrane, or another protein (Dyson and Wright, 2002). Homodimers whose monomers only fully fold upon dimerization include troponin C site III (Monera et al., 1992), Arc repressor (Robinson and Sauer, 1996), FIS (factor for inversion stimulation) (Hobart et al., 2002), Trp repressor (Gloss et al., 2001), and the dimeric form of p53 (Mateu et al., 1999). The HIV gp41 protein is a homotrimer with intrinsically unfolded monomers (Marti et al., 2004). It has been shown that the binding mechanism (whether monomer folding is coupled to binding or there are folded monomeric intermediates) is determined by the native topology, especially the number of inter and intramolecular contacts and the hydrophobicity of the interface (Levy et al., 2004). Flexibility of the chain seems to play a major role in the binding mechanism (Levy et al., 2005); one important manifestation of this is the fly casting effect (Shoemaker et al., 2000): a relatively unstructured chain can have a greater capture radius to catch its binding partner before fully folding. Protein Folding in the Cell Under carefully chosen in vitro conditions, small single domain proteins fold in a cooperative and reversible manner. Usually, the experiment is carried out in dilute solutions at low temperatures where the folding reaction is not complicated by off pathway reactions such as aggregation. Contrarily, the interior of a cell is a highly crowded environment with an estimated protein concentration of 300 mg/ml (Zimmerman and Trach, 1991), and in extreme cases, such as in a thermophilic archaeon, the temperature can exceed 80 C. Another difference is that in vivo the folding process is not separated from the relatively slow synthesis of the polypeptide chain (Creighton, 1990; Jaenicke, 1991). Folding cannot complete before a folding unit is completely synthesized. Nascent chains emerging from the ribosome should avoid formation of misfolded intermediates and aggregation. To ensure efficient folding, cells have evolved a large and
21 Protein folding 21 diverse group of proteins that assist the formation of the native structure. These proteins form the complex machinery of molecular chaperones, whose function is to guide other proteins to their proper folding and unfolding routes and help the assembly or disassembly of macromolecular structures, without becoming permanent components of these structures. Some chaperones are also stress or heat shock proteins because the need for chaperone function increases under conditions of stress that cause proteins to unfold or misassemble (Ellis, 2005). Although chaperone systems in eukaryotic cells are more complex than in prokaryotes, several homologous classes of chaperones have been identified in these different cell types. Two major classes of chaperones, the Hsp70s and the cylindrical chaperonin complexes, protect nonnative polypeptide chains from misfolding and aggregation in the cytosol of prokaryotes and eukaryotes, and inside the chloroplasts and mitochondria (Ostermann et al., 1989; Hartl, 1996), in an ATP dependent manner. The two classes exhibit distinct structural and functional properties and show entirely different mechanisms of action. Other protein classes, such as the Hsp40s, nucleotide exchange factors, or ADP destabilizing factors, act as cofactors to the chaperones. Here, we briefly summarize what is known about the major chaperone classes..1 The Hsp70 Family Hsp70s are distributed in all types of cells and in all cellular compartments. The Hsp70 family is a highly conserved protein family consisting of numerous homologs with distinct cellular functions (Frydman, 2001). These chaperones have a molecular mass of approximately 70 kda. The molecules consist of two domains, the 44 kda N terminal domain, which mediates ATP binding (Flaherty et al., 1990) and the small C terminal domain, which binds the protein substrate (Zhu et al., 1996). Substrate binding and release is modulated by ATP binding and hydrolysis. When ATP is bound, substrate binding and release occur rapidly, while with ADP bound, both substrate binding and release are slow. The function of Hsp70s is tuned by cofactors modulating substrate and nucleotide binding. Members of the Hsp70 family are DnaK in bacteria, Ssa and Ssb in yeast, and Hsc70 and Hsp70 in mammals. Hsp70s have a substrate binding cleft, which recognizes extended stretches of polypeptide chains rich in hydrophobic residues (Flynn et al., 1991; Rudiger et al., 1997), such as segments of partially unfolded proteins in nonnative conformations. Bound substrates are protected from aggregation; however, folding is obviously not possible in the bound state. Repeated binding and release might keep substrate proteins in extended, monomeric conformation giving them the chance to assume their native structure. However, many of the Hsp70 bound substrates are transferred to the real folding machine, the chaperonin system. Hsp40s are cofactors that stimulate ATP hydrolysis by Hsp70s. Hsp40s are also capable of binding substrates and can pass the bound substrates on to a Hsp70 molecule. The molecule usually consists of an N terminal J domain, which is responsible for Hsp70 binding and a C terminal chaperone domain containing hydrophobic patches for substrate binding (Sha et al., 2000). Representatives of the family are DnaJ in E. coli, Ydj1 and Sis1 (ribosome associated) in yeast, and Hdj1 2 and Hsp40 in mammals. Several eukaryotic homologs contain only a J domain, which may have a role in the localization of Hsp70s. Nucleotide exchange factors promote the release of ADP from members of the Hsp70 family. This group includes GrpE, a 23 kda protein in E.coli, and its homologs in mitochondria and chloroplasts, as well as several 60 kda proteins including Sti1 in yeast and Hop in mammals. In mammals, the Bag1 protein (Hohfeld and Jentsch, 1997; Takayama et al., 1997) also promotes the release of ADP from Hsp70 and the subsequent substrate release. These are modular proteins with domains that might have a role in connecting different chaperone systems. Hop may link Hsp70 to the Hsp90 system (Gross and Hessefort, 1996), and Bag1 contains an ubiquitin homology domain, suggesting a possible direction of Hsp70 bound substrates to the S26 proteasome system (Luders et al., 2000; Terada and Mori, 2000). ADP stabilizing factors, such as the 48 kda Hip protein in mammals, bind Hsp70 and stabilize the ADP Hsp70 complex.
22 22 Protein folding Small chaperones exhibiting Hsp70 like activity have a function similar to that of Hsp70, but unlike Hsp70, their function is independent from nucleotide binding. The trigger factor (TF) in E. coli has an overlapping function with DnaK. Additionally, it binds to the ribosome and displays prolyl isomerase activity (Hesterkamp et al., 1996). Hsp70 can be functionally substituted by the ubiquitous prefoldin (GimC) chaperone. Prefoldin is a heterohexamer that can interact with nascent polypeptides in vitro. The crystal structure of prefoldin reveals a unique quaternary structure forming a novel class of chaperones (Siegert et al., 2000)..2 The Folding Cage of Chaperonins (Hsp60 Family) AU3 Chaperonins are barrel like multi subunit complexes that primarily promote ATP dependent protein folding (Farr et al., 2000; Brinker et al., 2001). The unique mechanism of chaperonins involves the capture and isolation of substrate polypeptide chains inside the chamber of the complex. In the case of group I chaperonins such as GroEL in E. coli, the mitochondrial Hsp60, and the Rubiscobinding subunit (RBP) in plants, the system requires a Hsp type co chaperonin that acts as a cap or lid. The GroEL system with GroES as co chaperonin is capable of correctly folding proteins of sizes up to kda and with multiple domains (Houry et al., 1999). GroEL is a homooligomer of 14 subunits (see > Figure -8). These subunits consist of an apical, an intermediate, and an equatorial domain and are arranged in two stacked rings forming two chambers. The apical domains in the open conformation. Figure -8 (a) The structure of the GroEL complex with the GroES lid on (closed state). (b) The structure of the GroEL complex in the open state (no GroES lid present). One subunit of both GroEL and GroES is shown in a darker color provide hydrophobic side chains that can interact nonspecifically with the exposed hydrophobic surface of the unfolded substrate chain. Subsequent binding of the GroES cap and seven ATP molecules to GroEL triggers a conformational change resulting in an increased volume of the central cavity, a separation of the hydrophobic residues, and an exposure of hydrophilic residues in the apical region (Shtilerman et al., 1999). This may induce partial unfolding of the substrate molecule and its release into the inside of the central cavity of the chaperonin. The isolated environment of the central cavity is ideal for the substrate molecule for folding up into its native structure. Upon completion of ATP hydrolysis, which may take approximately s, binding of seven new ATP molecules to the trans ring triggers the dissociation of the GroES cap and the substrate molecule is released. In the case of incomplete folding, the substrate molecule can be recaptured and the annealing and relaxation cycle can be repeated until the molecule correctly folds.
23 Protein folding 23 Group II chaperonins are found in eukaryotes and in archaea and are homologous to group I chaperonins with a sequence identity up to 40%, showing similar double ring architecture. The eukaryotic group II chaperonin named TCP 1 (for tailless complex polypeptide 1) or CCP (for chaperonin containing TCP 1) is a heterooctamer consisting different subunits of kda (Kubota et al., 1995). Group II chaperonins have a helical protrusion on the apical domain that takes the place of the co chaperonin GroES. The crystal structure of the thermosome from Thermoplasma acidophilum reveals that this protrusion can assume different conformations in the open state including a helix and b sheet, which can increase the plasticity for binding different types of substrates..3 The Hsp90 Chaperone System Hsp90 plays a central role in eukaryotes in the regulation of the components of signal transduction systems such as tyrosine kinases and steroid hormone receptors. Hsp90 is ATP dependent and requires interaction with several cofactors, some of them having chaperone activity. An example is the substrate transfer from Hsp70 to Hsp90, which is facilitated by Hop (Hsp70 Hsp90 organizing protein). Hop contains binding sites for both chaperones and contributes to the rapid transfer of receptor molecules from Hsp70 to Hsp90. A detailed review on Hsp90 function has been published recently (Pearl and Prodromou, 2006)..4 Chaperone Assisted Assembly of Cellular Complexes Recent work has shown the role of nuclear chaperones in the assembly of nucleosomes and has led to the discovery of a cytosolic chaperone required for mammalian proteasome assembly, suggesting that besides the folding of individual proteins, the formation of oligomeric complexes may also be assisted by chaperones (Ellis, 2006)..5 Cooperation Between the Different Chaperone Systems Newly synthesized polypeptide chains, proteins losing their native state upon stress conditions, and other proteins requiring translocation in the cell may interact with several different chaperone systems. Chaperones having overlapping functions may compete for their substrate molecules as well. Hsp70 binds polypeptides and prevents aggregation. This may be sufficient for the correct folding of some proteins. Others may require further transfer from Hsp70 to the more efficient folding machine: the Hsp60 system. In E. coli, the cooperation between DnaK and GroEL has been proven in vivo (Teter et al., 1999). TF, which may substitute for the function of DnaK, also appears to cooperate with GroEL in substrate binding (Kandror et al., 1997). In eukaryotic cells, Hsp70 and TCP 1 associate, indicating a close functional relationship (Lewis et al., 1992). As mentioned above, Hsp70 and Hsp90 may also interact in receptor transfer. > Figure -9 shows a schematic representation of the general view of de novo protein folding in the cytosol..6 Chaperones and Neurodegenerative Diseases The pathogenesis of several neurodegenerative diseases is associated with protein misfolding, aggregation, and deposition of the protein, which may be manifested in cell degeneration and loss of function of the affected cells or organs. These degenerative disorders include polyglutamine (polyq) tract diseases, Alzheimer s and Parkinson s disease, amyotrophic lateral sclerosis, and Creutzfeldt Jakob syndrome (Chiti and Dobson, 2006). Chaperones whose function is to prevent protein aggregation in the cell may play a crucial role in the onset or progress of neurodegenerative diseases.
24 24 Protein folding. Figure -9 A schematic representation of the general view of de novo protein folding in the cytosol
25 11 Protein Misfolding and Aggregation Protein folding 25 Native states of proteins almost always represent the thermodynamically most stable conformation under physiological conditions (Vendruscolo et al., 2003). All the information regarding the native structure is hidden in the amino acid sequence. However, as we have seen, correct folding is a challenge for proteins in a living cell and only a part of the proteins can assume their native structure spontaneously. In the crowded milieu of the cell, efficient protein folding and transport depend on the presence of a complex machinery of chaperones, chaperonins, and cofactors. The primary mission of this machinery is to prevent the aggregation of nascent polypeptide chains and proteins that unfold upon environmental stress (Frydman, 2001). The failure of a specific protein to adopt or maintain its native functional conformation may result in pathological conditions referred to as protein misfolding diseases. Misfolding diseases include a wide range of diseases with different pathological mechanisms. The loss of the normal cellular function because of a reduction in the number of functional protein molecules is responsible for diseases such as cystic fibrosis (Amaral, 2004) and early onset emphysema (Lomas and Carrell, 2002). The major group of misfolding diseases, however, is associated with aggregation and deposition of proteins in the human body in the form of organized, fibrillar aggregates, generally termed amyloid fibrils. On one hand, unwanted protein aggregation may be caused by malfunctioning of the cellular protein quality control machinery. It may occur when the ubiquitin proteasome protein degradation system cannot eliminate misfolded, aggregation prone molecules (Ross and Pickart, 2004; Mandel et al., 2005); when the chaperone machinery performs insufficiently (Lee and Tsai, 2005); if the normal cellular transport route of a protein is damaged; or when inappropriate protease activity produces amyloidogenic protein fragments. On the other hand, aggregation and amyloid formation of a protein may be promoted by an increased expression level or by pathological mutations destabilizing the structure and inducing intermediate amyloidogenic conformations (Chiti and Dobson, 2006) Degenerative Diseases Associated with Amyloid Deposition Amyloid deposition is associated with more than 20 human degenerative diseases. We distinguish different groups of diseases by the location of amyloid deposits in the body. Neurodegenerative conditions such as Alzheimer s, Parkinson s, and Huntington s disease as well as spongiform encephalopathies affect the central nervous system. In nonneuropathic localized amyloidoses, protein deposition occurs in a certain type of tissue such as Langerhans islands in type II diabetes. In systemic amyloidoses such as AL amyloidosis, which involves the deposition of immunoglobulin light chain fragments, protein deposition is not limited to a single tissue. Alzheimer s and Parkinson s disease are sporadic and usually develop with aging, which suggests the role of the protein quality control machinery in the disease, although hereditary forms are also documented. Other conditions are hereditary, arising from specific mutations, such as lysozyme and fibrinogen amyloidosis. The special property of spongiform encephalopathy is that it can be transmissible in humans and mammals (Chiti and Dobson, 2006). Hemodialysis related amyloidosis represents the first amyloid disease that is a complication of a medical therapy (Gejyo et al., 1985). Protein aggregates can accumulate both extracellularly and intracellularly. The term intracellular inclusion has been suggested as more appropriate for amyloid like aggregates depositing inside the cell (Westermark et al., 2005). However, herein we use the term amyloid fibril The Structure and Morphology of Amyloid Fibrils A common characteristic feature of amyloid fibrils formed from different proteins is the well ordered structure with high b structure content. X ray fiber diffraction studies have shown that the b strands are oriented perpendicularly to the fibril axis (Sunde and Blake, 1997). Electron microscopy (EM) and atomic
26 26 Protein folding force microscopy (AFM) revealed that amyloid fibrils possess diverse morphologies at the fibrillar level. Single protofilaments can be straight or curved, with a diameter of 2 5 nm, showing no helical twist. Fibrils usually consist of 2 6 protofilaments, twisting together in a rope like or ribbon form with a diameter of 7 15 nm (Serpell et al., 2000). High resolution structure determination of amyloid fibrils is a grand challenge. Traditional spectroscopic methods fail because of the insoluble nature or the large size of the fibrils. Crystallization for X ray is complicated because fibrils favor growth in one direction. The crystal structure of the amyloid form of the GNNQQNY peptide has recently been solved (Nelson and Eisenberg, 2006). An extended b sheet is formed in the crystal, with each of the b strands consisting of a single peptide. The structure revealed a tight packing of the side chains between two b sheets, excluding water molecules. Solid state NMR spectroscopy may provide high resolution information on amyloid structure. Tycko (2006) and coworkers built a model structure of the Ab(1 40) amyloid b peptide, associated with Alzheimer s disease, based on solid state NMR constraints. In the model, the different Ab molecules are stacked on each other in a parallel arrangement and in register, forming two b sheets. Every single Ab molecule contributes to two b strands, one in each b sheet. Without providing detailed structural information, hydrogen deuterium exchange methods combined with NMR spectroscopy, as well as limited proteolysis with mass spectrometry, are capable of collecting site specific information on the extent of the rigid amyloid core (Hoshino et al., 2002; Myers et al., 2006a). The morphology of amyloid fibrils grown in vitro highly depends on solution conditions such as buffer composition, ph, temperature, and protein concentration. Even under the same conditions, fibrils with different morphologies can be formed from the same polypeptide. Structural studies revealed that this polymorphism of amyloid fibrils is a reflection of different underlying structures at the molecular level (Petkova et al., 2005). AFM and EM have shown a wide variety of protein aggregates depending on the conditions. Proteins may form disordered aggregates, oligomers, spherical aggregates, prefibrillar aggregates, and fibrils with various morphologies (Kad et al., 2003; Stine et al., 2003). > Figure - shows a unified view of the various types of structures that can be formed by polypeptide chains in vivo or in vitro Mechanism of Amyloid Fibril Formation The kinetics of amyloid formation and the amyloid content of a protein solution can be studied by light scattering, thioflavin T fluorescence, or other spectroscopic methods. Amyloid formation is usually a nucleation dependent reaction showing an initial lag phase, followed by rapid growth. Clearing the solution of any aggregated material by ultracentrifugation prior to the reaction may significantly increase the lag phase. Oppositely, addition of a small amount of preformed aggregated material can reduce or eliminate the lag phase. These observations suggest that the lag phase is the time required for proper nucleation. Under some conditions, no lag time is observable, suggesting that nucleation is not always the rate limiting step (Uversky et al., 2002; Pedersen et al., 2004). To understand the in vivo mechanism of amyloidoses, it is important to understand the nucleation process and carefully examine the reaction during the lag phase. Oligomers, spherical or chain like aggregates, sometimes termed protofibrils, forming prior to fibril formation have been observed in many systems (Chiti and Dobson, 2006). It is extremely important to understand the mechanism of the formation of these prefibrillar species. Cell culture and in vivo studies have revealed their toxicity for living cells, and they may be involved in the pathogenesis of diseases such as Alzheimer s (Lue et al., 1999; McLean et al., 1999) Amyloid Formation of Globular Proteins One group of amyloid forming systems consists of peptides or protein fragments that are natively unfolded in the monomeric form (Alzheimer s Ab, Parkinson s a synuclein). It is generally accepted that stable globular proteins need to partially unfold to an amyloidogenic intermediate for fibril formation. This is
27 Protein folding 27. Figure - A unified view of the major types of structure that can be formed by polypeptide chains supported by experimental data showing increased amyloidogenicity under conditions that destabilize the native state. Destabilizing mutations may also promote amyloid formation, which explains the mechanism of some hereditary diseases (Raffen et al., 1999; Canet et al., 2002). Moreover, under carefully chosen denaturing conditions, most proteins are capable of aggregation and amyloid formation in vitro, suggesting that the amyloid state is a general property of polypeptide chains (Stefani and Dobson, 2003). However, out of thousands of different proteins in the human body, only some two dozen are responsible for diseases, indicating that protein sequences and the cellular machinery have evolved to avoid unwanted protein aggregation Physicochemical and Sequence Determinants of Amyloid Formation Hydrophobicity of the peptide chain has been shown to influence its aggregation propensity (Otzen et al., 2000). Clusters of consecutive hydrophobic residues are avoided by evolution (Schwartz et al., 2001). Another crucial factor is the charge of the polypeptide chain. A high net charge may prevent aggregation of the polypeptide (Chiti et al., 2002). Secondary structure propensity may also affect amyloid formation: a high propensity to form b sheet structure and low a helix propensity are likely to increase the probability of aggregation (Chiti and Dobson, 2006). Alternating patterns of polar and nonpolar residues, which promote b sheet formation, are less frequent in natural protein sequences than expected on a random basis (Broome and Hecht, 2000). Proline residues break b sheet structures, and hence inhibit aggregation (Steward et al., 2002; Parrini et al., 2005). The presence of b bulges on b strands exposed on the protein surface has been found to protect against the formation of intermolecular b sheets (Richardson and Richardson, 2002).
28 28 Protein folding 11.6 Factors Inducing or Inhibiting Amyloid Formation Under Physiological Conditions Amyloid formation may be facilitated under proper in vitro conditions, but in vivo, it always occurs under physiological conditions. Some disease related proteins such as b 2 microglobulin cannot form amyloid fibrils at physiological ph in vitro, suggesting the presence of unknown factors contributing to the aggregation process in vivo. Collagen, apolipoprotein E, heparin, serum amyloid P component, and low concentrations of sodium dodecyl sulfate have been reported to promote the aggregation of b 2 microglobulin under physiological conditions (Yamamoto et al., 2004; Myers et al., 2006b). Other molecules binding to and stabilizing the native state may prevent oligomerization or amyloid formation of diseaserelated proteins in the human body, and might serve as effective therapeutic agents in the future. 12 New Biophysical Techniques for the Study of Protein Folding In the last decade a wide range of physical and chemical methods complemented the fundamental techniques to study protein folding. We give a brief summary of the advances of these methods Rapid Mixing Methods The mechanism of protein folding can be studied by two different groups of approaches. Equilibrium methods provide information about possible folding intermediate states or deduce rate constants from the molecular fluctuations or dynamic properties of the system. Relaxation methods follow the change of the system evolving toward a new equilibrium after a rapid perturbation of its extrinsic variables, such as temperature, ph, pressure, or solvent composition (Roder et al., 2004). The time required for folding varies greatly among proteins, ranging from microseconds to minutes (Roder and Shastry, 1999; Roder et al., 2006). The smallest protein molecules with no folding intermediates fold on the microsecond timescale, which, on one hand, might make them suitable for in silico folding simulation studies. On the other hand, such fast reactions make the experimental detection of events during the folding process difficult. Basic techniques in the study of folding kinetics are stopped flow fluorescence spectroscopy and stopped flow circular dichroism. These techniques are capable of monitoring the formation of secondary and tertiary structures during the folding reaction with millisecond time resolution. In such experiments, rapid processes occurring within the dead time of the measurement were observed for many proteins. The challenge to resolve this initial burst phase and to reveal the structural changes taking place during the first millisecond stimulated the development of new, rapid kinetic techniques capable of triggering and monitoring the folding process on the submillisecond timescale. While the conventional stopped flow apparatus, in which a small volume of a freshly made mixture containing the reacting components is injected into the measurement cell, is quite economical and offers a wide range of applications (Gibson and Milnes, 1964), its dead time is usually about 1 millisecond or longer. Continuous flow methods extend the time resolution to the microsecond time range (Chan et al., 1997; Takahashi et al., 1997; Shastry et al., 1998; Akiyama et al., 2002). In the continuous flow cell, solutions are mixed under highly turbulent conditions to achieve complete mixing. The kinetics of the reaction is monitored under steady state flow conditions as a function of the distance downstream from the mixer by using relatively simple and inexpensive detection methods. Using this technique, it has become possible to study the initial collapse and formation of intermediates in the early stage of the folding reaction, during the burst phase (Uzawa et al., 2004; Welker et al., 2004; Kimura et al., 2005) Real Time NMR Spectroscopy Nuclear magnetic resonance (NMR) spectroscopy has greatly contributed to our understanding of the protein folding problem. NMR provides high spatial resolution and a broad timescale ranging from
29 Protein folding 29 picoseconds to days. Hydrogen deuterium exchange experiments, revealing dynamical events at an atomic level, have illuminated the process of unfolding from the native state and the structure of folding intermediates (Wagner and Wuthrich, 1982; Wand et al., 1986; Bai et al., 1995). The quenched flow pulselabeling technique has enabled researchers to study the early stages of protein folding using a conventional NMR instrument (Roder et al., 1988; Udgaonkar and Baldwin, 1988; Radford et al., 1992). NMR studies have characterized the properties of denaturant induced equilibrium folding intermediate states such as the molten globule (Arai and Kuwajima, 2000). In equilibrium systems, the rates of conversion between distinct conformational states can be calculated from a line shape analysis of the NMR resonances (Huang and Oas, 1995), and therefore can provide kinetic data on folding. Slow folding reactions such as cis trans prolyl isomerization can be directly followed by sequential recording of one dimensional (1D) NMR spectra (Balbach et al., 1999). This method is particularly useful for discovering intermediates formed at the late stages of the folding process. Using a stopped flow device for injection of the protein solution into the NMR tube that already contains the denaturant or the refolding buffer pushes the dead time of mixing below 1 s (Zeeb and Balbach, 2004). One of the first proteins studied by real time NMR was a lactalbumin (Balbach et al., 1995). 1D NOE (nuclear Overhauser effect) experiments revealed the native like compactness of the transient molten globule state of a lactalbumin (Forge et al., 1999). These experiments also demonstrated that the transient intermediate closely resembles the well characterized stable molten globule state formed at low ph. While 1D NMR spectra have limited resolution, multidimensional NMR can provide high spatial resolution information on the folding process. Because recording multidimensional spectra is time consuming, only slow processes could be followed directly by sequential recording (Liu et al., 1996). Balbach and coworkers (1996, 1999) developed new methods to reconstruct the kinetic history of folding reactions from a single two dimensional NMR spectrum recorded during the entire time course of the reaction. The basis of these methods is that the line widths and intensities reflect the history of the folding events occurring during spectral accumulation. When applied to a lactalbumin, the technique demonstrated the cooperative nature of the folding of the main chain Chemically Induced Nuclear Polarization Chemically induced nuclear polarization (CIDNP) can be used to probe the solvent accessibility of certain aromatic residues in proteins (Mok et al., 2003; Mok and Hore, 2004). The reactive collision of polarizable amino acids such as tryptophan, tyrosine, and histidine with a photoexcited dye such as flavin mononucleotide (FMN) results in an electron transfer (in the case of Trp and Tyr) or proton transfer (His) reaction forming a pair of radicals. Electron nuclear hyperfine interactions between the two radicals result in a significant enhancement of NMR signals. The photosensitizer flavin molecule can be excited by laser as light source. For the photoreaction to take place, the aromatic side chains must be accessible to the photosensitizer, e.g., located on the surface of the protein molecule. The CIDNP spectrum is recorded immediately after the laser flash and corrected by a dark spectrum recorded without irradiation. Besides the equilibrium studies of protein surfaces, the technique can be combined with a stopped flow apparatus and in this way it can be used to study folding intermediates. Using CIDNP pulse labeling technique, the exposed tryptophan and tyrosine residues in a molten globule state can be identified (Lyon et al., 2002; Mok et al., 2003) High Pressure NMR Spectroscopy When high pressure is applied to a protein solution, it shifts the conformational equilibrium of the protein molecules toward lower volume conformers, thereby decreasing the partial molar volume of the protein. The combination of high pressure with heteronuclear two dimensional NMR spectroscopy provides atomic resolution information on the structure of the protein molecule at different stages of the folding process (Kamatari et al., 2004). By varying the pressure, one can explore the conformational space from the folded to the unfolded conformer. In recent years, numerous studies using high pressure NMR spectroscopy have
30 30 Protein folding been carried out (Akasaka and Yamada, 2001) on locally disordered (Kuwata et al., 2001; Kuwata et al., 2002; Kitahara et al., 2005), molten globule (Kitahara et al., 2002; Lassalle et al., 2003), unfolded (Kamatari et al., 2001; Arnold et al., 2002; Refaee et al., 2003) as well as oligomeric or aggregated states of proteins (Niraula et al., 2004; Silva et al., 2006) Protein Folding and Dynamics Studied by Mass Spectrometry Mass spectrometry of protein molecules has become a rapidly developing field in the last decade (Konermann and Simmons, 2003; Eyles and Kaltashov, 2004). In comparison with NMR spectroscopy, which provides site specific information averaged in time, mass spectrometry is capable of detecting different conformers coexisting in the protein solution. This method is especially useful for the study of lowpopulated intermediate states and is free of the molecular size limitation of NMR spectroscopy. Because of its high sensitivity, a protein concentration in the femtomolar range is sufficient for analysis. Structural and dynamic properties of various conformational states can be studied by hydrogen/deuterium exchange (HDX) combined with mass spectrometry. Recently, Kaltashov and coworkers investigated the conformational ensemble of the molten globule state of ubiquitin (Hoerner et al., 2005). Using protein ion fragmentation in the gas phase, they evaluated the stability of various segments of the protein in the molten globular state. By the method of pulse labeling HDX MS, it is possible to study the kinetics of folding and to explore complex folding scenarios with parallel pathways (Konermann and Simmons, 2003). Co populated protein conformers can be detected and characterized directly by electrospray ionization mass spectrometry (ESIMS) (Mohimen et al., 2003; Borysik et al., 2004). Protein surface areas in solution may be determined by ESIMS (Kaltashov and Mohimen, 2005). Limited proteolysis with ESIMS provides site specific structural information on different conformational states of the protein molecules including protein aggregates and the amyloid state (Myers et al., 2006a) Mechanical Unfolding of Proteins In the first studies of the mechanical unfolding of single protein molecules using AFM, the giant sarcomeric protein titin, consisting of a large number of immunoglobulin segments, was used (Erickson, 1997; Rief et al., 1997; Rounsevell et al., 2004). Because of the heterogeneity of titin domains, it was not possible to assign the individual force peaks to specific domains. Using tandem repeats of a single domain, constructed by protein engineering techniques, it was possible to explain the mechanical characteristics of single domains in terms of their specific structures (Carrion Vazquez et al., 1999; Carrion Vazquez et al., 2000). Using force measuring optical tweezers, it is possible to induce mechanical unfolding and refolding of individual molecules (Kellermayer et al., 1997). In a recent work, Cecconi and coworkers (2005) showed that E. coli ribonuclease H molecule unfolds in a two state manner and refolds through a transient molten globule like intermediate. We may expect significant progress in the application of other new techniques such as the study of single molecule folding kinetics by optical techniques in the near future (Lipman et al., 2003). 13 Conclusions In spite of the great advances in experimental technique and the tremendous boost in computational power we have witnessed in the past decades, the protein folding problem is still far from solved. We already have a good understanding of some of the simpler protein folding mechanisms, and we can simulate the folding of a few, very small proteins. However, we are still at a loss when the folding behavior of more complex, multidomain proteins is to be explained, especially when protein protein interactions, misfolding, aggregation, and amyloid formation complicate the situation such as in a living cell. Although we have known
31 Protein folding 31 since Anfinsen that sequence determines structure (in a given environment), and the field of protein structure prediction has made great progress, we are still not, in general, capable of predicting the structure from a sequence. The research of protein folding, however, is no longer an exotic field with only theoretical significance. Diseases such as Alzheimer s or Parkinson s have taught us that protein folding can go awry, and misfolded or aggregated proteins can cause a lot of trouble. In order to successfully defend us against these and other folding diseases, we should reach a level of understanding of folding phenomena that is not just descriptive but allows us to influence the folding behavior of proteins in the way we want. Therefore, the research of folding continues, and in all likelihood will bring about great advances in the development of new experimental techniques and new theoretical approaches. Acknowledgments A.S. was supported by a Bolyai János fellowship. A.S. and P.Z. were supported by grants from the Hungarian Scientific Research Fund (OTKA T and NI 61915). S.O. was supported by a grant from the Hungarian Scientific Research Fund (OTKA D 38480). References Akasaka K, Yamada H On line cell high pressure nuclear magnetic resonance technique: Application to protein studies. Methods Enzymol 338: Akiyama S, Takahashi S, Kimura T, Ishimori K, Morishima I, et al Conformational landscape of cytochrome c folding studied by microsecond resolved small angle x ray scattering. Proc Natl Acad Sci USA 99: Alm E, Baker D Prediction of protein folding mechanisms from free energy landscapes derived from native structures. Proc Natl Acad Sci USA 96: Alonso DO, Daggett V Molecular dynamics simulations of hydrophobic collapse of ubiquitin. Protein Sci 7: Amaral MD Cftr and chaperones: Processing and degradation. J Mol Neurosci 23: Anfinsen CB Principles that govern the folding of protein chains. Science 181: Arai M, Kuwajima K Role of the molten globule state in protein folding. Adv Protein Chem 53: Arnold MR, Kremer W, Ludemann HD, Kalbitzer HR H NMR parameters of common amino acid residues measured in aqueous solutions of the linear tetrapeptides gly gly x ala at pressures between 0.1 and 200 MPa. Biophys Chem 96: Bai Y, Sosnick TR, Mayne L, Englander SW Protein folding intermediates: Native state hydrogen exchange. Science 269: Baker D A surprising simplicity to protein folding. Nature 405: Balbach J, Schmid FX Proline isomerization and its catalysis in protein folding. Mechanisms of Protein Folding. Pain RH, editor. Oxford: Oxford University Press; pp Balbach J, Forge V, Lau WS, van Nuland NA, Brew K, et al Protein folding monitored at individual residues during a two dimensional NMR experiment. Science 274: Balbach J, Forge V, van Nuland NA, Winder SL, Hore PJ, et al Following protein folding in real time using NMR spectroscopy. Nat Struct Biol 2: Balbach J, Steegborn C, Schindler T, Schmid FX A protein folding intermediate of ribonuclease T1 characterized at high resolution by 1D and 2D real time NMR spectroscopy. J Mol Biol 285: Baldwin RL How does protein folding get started? Trends Biochem Sci 14: Baldwin RL On pathway versus off pathway folding intermediates. Fold Des 1: R1-R8. Becker OM, Karplus M The topology of multidimensional potential energy surfaces: Theory and application to peptide structure and kinetics. J Chem Phys 6: Bennett MJ, Choe S, Eisenberg D Domain swapping: Entangling alliances between proteins. Proc Natl Acad Sci USA 91: Bergman LW, Kuehl WM Formation of an intrachain disulfide bond on nascent immunoglobulin light chains. J Biol Chem 254: Blond S, Goldberg ME Kinetic characterization of early intermediates in the folding of E. coli tryptophan synthase b2 subunit. Proteins 1: Borysik AJH, Radford SE, Ashcroft AE Co populated conformational ensembles of b 2 microglobulin uncovered
32 32 Protein folding quantitatively by electrospray ionization mass spectrometry. J Biol Chem 279: Braakman I, Helenius J, Helenius A Manipulating disulfide bond formation and protein folding in the endoplasmic reticulum. EMBO J 11: Braakman I, Hoover Litty H, Wagner KR, Helenius A Folding of influenza hemagglutinin in the endoplasmic reticulum. J Cell Biol 114: Bradley P, Misura KMS, Baker D Toward high resolution de novo structure prediction for small proteins. Science 309: Brandts JF, Halvorson HR, Brennan M Consideration of the possibility that the slow step in protein denaturation reactions is due to cis trans isomerism of proline residues. Biochemistry 14: Brinker A, Pfeifer G, Kerner MJ, Naylor DJ, Hartl FU, et al Dual function of protein confinement in chaperoninassisted protein folding. Cell 7: Broome BM, Hecht MH Nature disfavors sequences of alternating polar and non polar amino acids: Implications for amyloidogenesis. J Mol Biol 296: Bryngelson JD, Onuchic JN, Socci ND, Wolynes PG Funnels, pathways, and the energy landscape of protein folding: A synthesis. Proteins 21: Bryngelson JD, Wolynes PG Spin glasses and the statistical mechanics of protein folding. Proc Natl Acad Sci USA 84: Bulaj G Formation of disulfide bonds in proteins and peptides. Biotechnol Adv 23: Bulaj G, Goldenberg DP F values for bpti folding intermediates and implications for transition state analysis. Nat Struct Biol 8: Bulleid NJ, Freedman RB Defective co translational formation of disulphide bonds in protein disulphide isomerase deficient microsomes. Nature 335: Canet D, Last AM, Tito P, Sunde M, Spencer A, et al Local cooperativity in the unfolding of an amyloidogenic variant of human lysozyme. Nat Struct Biol 9: Capaldi AP, Kleanthous C, Radford SE Im7 folding mechanism: Misfolding on a path to the native state. Nat Struct Biol 9: Carlsson U, Jonsson BH Folding of b sheet proteins. Curr Opin Struct Biol 5: Carra JH, Privalov PL Thermodynamics of denaturation of staphylococcal nuclease mutants: An intermediate state in protein folding. FASEB J : Carrion Vazquez M, Oberhauser AF, Fisher TE, Marszalek PE, Li H, et al Mechanical design of proteins studied by single molecule force spectroscopy and protein engineering. Prog Biophys Mol Biol 74: Carrion Vazquez M, Oberhauser AF, Fowler SB, Marszalek PE, Broedel SE, et al Mechanical and chemical unfolding of a single protein: A comparison. Proc Natl Acad Sci USA 96: Cecconi C, Shank EA, Bustamante C, Marqusee S Direct observation of the three state folding of a single protein molecule. Science 309: Chakraborty S, Ittah V, Bai P, Luo L, Haas E, et al Structure and dynamics of the a lactalbumin molten globule: Fluorescence studies using proteins containing a single tryptophan residue. Biochemistry 40: Chan CK, Hu Y, Takahashi S, Rousseau DL, Eaton WA, et al Submillisecond protein folding kinetics studied by ultrarapid mixing. Proc Natl Acad Sci USA 94: Chan HS, Dill KA A simple model of chaperoninmediated protein folding. Proteins 24: Chan HS, Dill KA Protein folding in the landscape perspective: Chevron plots and non Arrhenius kinetics. Proteins 30: Chatrenet B, Chang JY The folding of hirudin adopts a mechanism of trial and error. J Biol Chem 267: Cheng HN, Bovey FA Cis trans equilibrium and kinetic studies of acetyl L proline and glycyl L proline. Biopolymers 16: Chiti F, Dobson CM Protein misfolding, functional amyloid, and human disease. Annu Rev Biochem 75: Chiti F, Calamai M, Taddei N, Stefani M, Ramponi G, et al Studies of the aggregation of mutant proteins in vitro provide insights into the genetics of amyloid diseases. Proc Natl Acad Sci USA 99 (Suppl. 4): Chrunyk BA, Matthews CR Role of diffusion in the folding of the a subunit of tryptophan synthase from Escherichia coli. Biochemistry 29: Clementi C, Nymeyer H, Onuchic JN Topological and energetic factors: What determines the structural details of the transition state ensemble and en route intermediates for protein folding? An investigation for small globular proteins. J Mol Biol 298: Crane JC, Koepf EK, Kelly JW, Gruebele M Mapping the transition state of the ww domain b sheet. J Mol Biol 298: Creighton TE Intermediates in the refolding of reduced ribonuclease A. J Mol Biol 129: Creighton TE Protein folding. Biochem J 270: Creighton TE Protein folding coupled to disulphide bond formation. Biol Chem 378: Creighton TE Protein folding coupled to disulphidebond formation. Mechanisms of protein folding. Pain RH, editor. Oxford: Oxford University Press; pp Creighton TE, Goldenberg DP Kinetic role of a metastable native like two disulphide species in the folding transition of bovine pancreatic trypsin inhibitor. J Mol Biol 179: AU5
33 Protein folding 33 Dabora JM, Pelton JG, Marqusee S Structure of the acid state of Escherichia coli ribonuclease HI. Biochemistry 35: Daggett V, Fersht AR Transition states in protein folding. Mechanisms of protein folding. Pain RH, editor. Oxford: Oxford University Press; pp Daggett V, Fersht AR Is there a unifying mechanism for protein folding? Trends Biochem Sci 28: Dautry Varsat A, Garel JR Independent folding regions in aspartokinase homoserine dehydrogenase. Biochemistry 20: Day R, Daggett V All atom simulations of protein folding and unfolding. Adv Protein Chem 66: Demarest SJ, Horng JC, Raleigh DP A protein dissection study demonstrates that two specific hydrophobic clusters play a key role in stabilizing the core structure of the molten globule state of human a lactalbumin. Proteins 42: Dill KA Theory for the folding and stability of globular proteins. Biochemistry 24: Dill KA Dominant forces in protein folding. Biochemistry 29: Dill KA Polymer principles and protein folding. Protein Sci 8: Dill KA, Chan HS From levinthal to pathways to funnels. Nat Struct Biol 4: -19. Dill KA, Bromberg S, Yue K, Fiebig KM, Yee DP, et al Principles of protein folding a perspective from simple exact models. Protein Sci 4: Ding F, Dokholyan NV Simple but predictive protein models. Trends Biotechnol 23: Ding F, Buldyrev SV, Dokholyan NV. 2005a. Folding trp cage to NMR resolution native structure using a coarse grained protein model. Biophys J 88: Ding F, Guo W, Dokholyan NV, Shakhnovich EI, Shea J. 2005b. Reconstruction of the src sh3 protein domain transition state ensemble using multiscale molecular dynamics simulations. J Mol Biol 350: Dobson CM Protein folding. Solid evidence for molten globules. Curr Biol 4: Dokholyan NV Studies of folding and misfolding using simplified models. Curr Opin Struct Biol 16: Duan Y, Kollman PA Pathways to a protein folding intermediate observed in a 1 ms simulation in aqueous solution. Science 282: Dyson HJ, Wright PE Insights into protein folding from NMR. Annu Rev Phys Chem 47: Dyson HJ, Wright PE Coupling of folding and binding for unstructured proteins. Curr Opin Struct Biol 12: Ellgaard L, Ruddock LW The human protein disulphide isomerase family: Substrate interactions and functional properties. EMBO Rep 6: Ellis RJ Chaperone function: The orthodox view. Molecular Chaperones and Cell Signalling. Henderson B, Pockley AG, editors. Cambridge University Press; pp Ellis RJ Molecular chaperones: Assisting assembly in addition to folding. Trends Biochem Sci 31: Englander SW Protein folding intermediates and pathways studied by hydrogen exchange. Annu Rev Biophys Biomol Struct 29: Englander SW, Mayne L Protein folding studied using hydrogen exchange labeling and two dimensional NMR. Annu Rev Biophys Biomol Struct 21: Erickson HP Stretching single protein molecules: Titin is a weird spring. Science 276: Evans MS, Clarke TF, Clark PL Conformations of cotranslational folding intermediates. Protein Pept Lett 12: Evans PA, Dobson CM, Kautz RA, Hatfull G, Fox RO Proline isomerism in staphylococcal nuclease characterized by NMR and site directed mutagenesis. Nature 329: Eyles SJ Proline not the only culprit? Nat Struct Biol 8: Eyles SJ, Kaltashov IA Methods to study protein dynamics and folding by mass spectrometry. Methods 34: Fabian H, Naumann D Methods to study protein folding by stopped flow FT IR. Methods 34: Farr GW, Furtak K, Rowland MB, Ranson NA, Saibil HR, et al Multivalent binding of nonnative substrate proteins by the chaperonin GroEL. Cell 0: Fersht AR Optimization of rates of protein folding: The nucleation condensation mechanism and its implications. Proc Natl Acad Sci USA 92: Fersht AR Nucleation mechanisms in protein folding. Curr Opin Struct Biol 7: 3-9. Fersht AR, Daggett V Protein folding and unfolding at atomic resolution. Cell 8: Fersht AR, Sato S F value analysis and the nature of protein folding transition states. Proc Natl Acad Sci USA 1: Fersht AR, Matouschek A, Serrano L The folding of an enzyme. I. Theory of protein engineering analysis of stability and pathway of protein folding. J Mol Biol 224: Fitch CA, Whitten ST, Hilser VJ, Garcia Moreno EB Molecular mechanisms of ph driven conformational transitions of proteins: Insights from continuum electrostatics calculations of acid unfolding. Proteins 63: Flaherty KM, DeLuca Flaherty C, McKay DB Threedimensional structure of the ATPase fragment of a 70k heat shock cognate protein. Nature 346: Flynn GC, Pohl J, Flocco MT, Rothman JE Peptidebinding specificity of the molecular chaperone bip. Nature 353:
34 34 Protein folding Forge V, Wijesinha RT, Balbach J, Brew K, Robinson CV, et al Rapid collapse and slow structural reorganisation during the refolding of bovine a lactalbumin. J Mol Biol 288: Franks F Protein destabilization at low temperatures. Adv Protein Chem 46: Freedman RB Protein disulfide isomerase: Multiple roles in the modification of nascent secretory proteins. Cell 57: Frydman J Folding of newly translated proteins in vivo: The role of molecular chaperones. Annu Rev Biochem 70: Fulton KF, Main ER, Daggett V, Jackson SE Mapping the interactions present in the transition state for unfolding/folding of fkbp12. J Mol Biol 291: Galzitskaya OV, Finkelstein AV A theoretical search for folding/unfolding nuclei in three dimensional protein structures. Proc Natl Acad Sci USA 96: Garel JR, Dautry Varsat A Sequential folding of a bifunctional allosteric protein. Proc Natl Acad Sci USA 77: Gejyo F, Yamada T, Odani S, Nakagawa Y, Arakawa M, et al A new form of amyloid protein associated with chronic hemodialysis was identified as b 2 microglobulin. Biochem Biophys Res Commun 129: Gibson QH, Milnes L Apparatus for rapid and sensitive spectrophotometry. Biochem J 91: Gilbert HF Protein disulfide isomerase and assisted protein folding. J Biol Chem 272: Gillespie B, Plaxco KW Using protein folding rates to test protein folding theories. Annu Rev Biochem 73: Gloss LM, Simler BR, Matthews CR Rough energy landscapes in protein folding: Dimeric E. coli trp repressor folds through three parallel channels. J Mol Biol 312: Go N Theoretical studies of protein folding. Annu Rev Biophys Bioeng 12: Goldenberg DP Finding the right fold. Nat Struct Biol 6: Gothel SF, Marahiel MA Peptidyl prolyl cis trans isomerases, a superfamily of ubiquitous folding catalysts. Cell Mol Life Sci 55: Grantcharova V, Alm EJ, Baker D, Horwich AL Mechanisms of protein folding. Curr Opin Struct Biol 11: Grantcharova VP, Riddle DS, Santiago JV, Baker D Important role of hydrogen bonds in the structurally polarized transition state for folding of the src SH3 domain. Nat Struct Biol 5: Grathwohl C, Wuthrich K NMR studies of the rates of proline cis trans isomerization in oligopeptides. Biopolymers 20: Gross M, Hessefort S Purification and characterization of a 66 kda protein from rabbit reticulocyte lysate which promotes the recycling of hsp 70. J Biol Chem 271: Gruebele M Protein folding: The free energy surface. Curr Opin Struct Biol 12: Gruebele M Downhill protein folding: Evolution meets physics. C R Biol 328: Hamada D, Segawa S, Goto Y Non native a helical intermediate in the refolding of b lactoglobulin, a predominantly b sheet protein. Nat Struct Biol 3: Harrison SC, Durbin R Is there a single pathway for the folding of a polypeptide chain?. Proc Natl Acad Sci USA 82: Hartl FU Molecular chaperones in cellular protein folding. Nature 381: Heidary DK, Gross LA, Roy M, Jennings PA Evidence for an obligatory intermediate in the folding of interleukin 1b. Nat Struct Biol 4: Herning T, Yutani K, Taniyama Y, Kikuchi M Effects of proline mutations on the unfolding and refolding of human lysozyme: The slow refolding kinetic phase does not result from proline cis trans isomerization. Biochemistry 30: Herold M, Leistler B, Hage A, Luger K, Kirschner K Autonomous folding and coenzyme binding of the excised pyridoxal 5 0 phosphate binding domain of aspartate aminotransferase from Escherichia coli. Biochemistry 30: Hesterkamp T, Hauser S, Lutcke H, Bukau B Escherichia coli trigger factor is a prolyl isomerase that associates with nascent polypeptide chains. Proc Natl Acad Sci USA 93: Hobart SA, Ilin S, Moriarty DF, Osuna R, Colon W Equilibrium denaturation studies of the Escherichia coli factor for inversion stimulation: Implications for in vivo function. Protein Sci 11: Hoerner JK, Xiao H, Kaltashov IA Structural and dynamic characteristics of a partially folded state of ubiquitin revealed by hydrogen exchange mass spectrometry. Biochemistry 44: Hohfeld J, Jentsch S GrpE like regulation of the hsc70 chaperone by the anti apoptotic protein bag 1. EMBO J 16: Hoshino M, Katou H, Hagihara Y, Hasegawa K, Naiki H, et al Mapping the core of the b 2 microglobulin amyloid fibril by h/d exchange. Nat Struct Biol 9: Houry WA, Frishman D, Eckerskorn C, Lottspeich F, Hartl FU Identification of in vivo substrates of the chaperonin GroEL. Nature 402: Huang GS, Oas TG Submillisecond folding of monomeric l repressor. Proc Natl Acad Sci USA 92:
35 Protein folding 35 Hubner IA, Edmonds KA, Shakhnovich EI Nucleation and the transition state of the SH3 domain. J Mol Biol 349: Hubner IA, Shimada J, Shakhnovich EI Commitment and nucleation in the protein G transition state. J Mol Biol 336: Hughson FM, Wright PE, Baldwin RL Structural characterization of a partly folded apomyoglobin intermediate. Science 249: Ikeguchi M, Kuwajima K, Mitani M, Sugai S Evidence for identity between the equilibrium unfolding intermediate and a transient folding intermediate: A comparative study of the folding reactions of a lactalbumin and lysozyme. Biochemistry 25: Isenman DE, Lancet D, Pecht I Folding pathways of immunoglobulin domains. The folding kinetics of the cg3 domain of human IgG1. Biochemistry 18: Jaenicke R Folding and association of proteins. Prog Biophys Mol Biol 49: Jaenicke R Protein folding: Local structures, domains, subunits, and assemblies. Biochemistry 30: Jaenicke R Stability and folding of domain proteins. Prog Biophys Mol Biol 71: Jaenicke R Stability and stabilization of globular proteins in solution. J Biotechnol 79: Jaenicke R, Lilie H Folding and association of oligomeric and multimeric proteins. Adv Protein Chem 53: Jahn TR, Radford SE The yin and yang of protein folding. FEBS J 272: Jamin M, Baldwin RL Two forms of the ph 4 folding intermediate of apomyoglobin. J Mol Biol 276: Janin J, Wodak SJ Structural domains in proteins and their role in the dynamics of protein function. Prog Biophys Mol Biol 42: Jecht M, Tomschy A, Kirschner K, Jaenicke R Autonomous folding of the excised coenzyme binding domain of D glyceraldehyde 3 phosphate dehydrogenase from Thermotoga maritima. Protein Sci 3: Jorgensen WL, Gao J Cis trans energy difference for the peptide bond in the gas phase and in aqueous solution. J Am Chem Soc 1: Kad NM, Myers SL, Smith DP, Smith DA, Radford SE, et al Hierarchical assembly of b 2 microglobulin amyloid in vitro revealed by atomic force microscopy. J Mol Biol 330: Kaltashov IA, Mohimen A Estimates of protein surface areas in solution by electrospray ionization mass spectrometry. Anal Chem 77: Kamatari YO, Kitahara R, Yamada H, Yokoyama S, Akasaka K High pressure NMR spectroscopy for characterizing folding intermediates and denatured states of proteins. Methods 34: Kamatari YO, Yamada H, Akasaka K, Jones JA, Dobson CM, et al Response of native and denatured hen lysozyme to high pressure studied by 15 N/ 1 H NMR spectroscopy. Eur J Biochem 268: Kandror O, Sherman M, Moerschell R, Goldberg AL Trigger factor associates with GroEL in vivo and promotes its binding to certain polypeptides. J Biol Chem 272: Kang YK, Choi HY Cis trans isomerization and puckering of proline residue. Biophys Chem 111: Karplus M, Weaver DL Protein folding dynamics. Nature 260: Karplus M, Weaver DL Protein folding dynamics: The diffusion collision model and experimental data. Protein Sci 3: Kataoka M, Goto Y X ray solution scattering studies of protein folding. Fold Des 1: R7-R114. Kazmirski SL, Daggett V. 1998a. Simulations of the structural and dynamical properties of denatured proteins: The molten coil state of bovine pancreatic trypsin inhibitor. J Mol Biol 277: Kazmirski SL, Daggett V. 1998b. Non native interactions in protein folding intermediates: Molecular dynamics simulations of hen lysozyme. J Mol Biol 284: Kellermayer MS, Smith SB, Granzier HL, Bustamante C Folding unfolding transitions in single titin molecules characterized with laser tweezers. Science 276: Kelley RF, Richards FM Replacement of proline 76 with alanine eliminates the slowest kinetic phase in thioredoxin folding. Biochemistry 26: Kelly SM, Price NC The use of circular dichroism in the investigation of protein structure and function. Curr Protein Pept Sci 1: Khare SD, Ding F, Gwanmesia KN, Dokholyan NV Molecular origin of polyglutamine aggregation in neurodegenerative diseases. PLoS Comput Biol 1: Kiefhaber T, Grunert HP, Hahn U, Schmid FX Folding of RNase t1 is decelerated by a specific tertiary contact in a folding intermediate. Proteins 12: Kiefhaber T, Quaas R, Hahn U, Schmid FX. 1990a. Folding of ribonuclease T1. 1. Existence of multiple unfolded states created by proline isomerization. Biochemistry 29: Kiefhaber T, Quaas R, Hahn U, Schmid FX. 1990b. Folding of ribonuclease T1. 2. Kinetic models for the folding and unfolding reactions. Biochemistry 29: Kihara D, Lu H, Kolinski A, Skolnick J Touchstone: An ab initio protein structure prediction method that uses threading based tertiary restraints. Proc Natl Acad Sci USA 98: Kimura T, Akiyama S, Uzawa T, Ishimori K, Morishima I, et al Specifically collapsed intermediate in the early stageofthefoldingofribonucleasea.j Mol Biol 350:
36 36 Protein folding Kitahara R, Yamada H, Akasaka K, Wright PE High pressure NMR reveals that apomyoglobin is an equilibrium mixture from the native to the unfolded. J Mol Biol 320: Kitahara R, Yokoyama S, Akasaka K NMR snapshots of a fluctuating protein structure: Ubiquitin at 30 bar 3 kbar. J Mol Biol 347: Konermann L, Simmons DA Protein folding kinetics and mechanisms studied by pulse labeling and mass spectrometry. Mass Spectrom Rev 22: Konishi Y, Ooi T, Scheraga HA Regeneration of ribonuclease A from the reduced protein. Rate limiting steps. Biochemistry 21: Kubota H, Hynes G, Willison K The chaperonin containing t complex polypeptide 1 (tcp 1). Multi subunit machinery assisting in protein folding and assembly in the eukaryotic cytosol. Eur J Biochem 230: Kumar S, Nussinov R Close range electrostatic interactions in proteins. Chembiochem 3: Kunugi S, Tanaka N Cold denaturation of proteins under high pressure. Biochim Biophys Acta 1595: Kuwajima K The molten globule state of a lactalbumin. FASEB J : 2-9. Kuwajima K, Hiraoka Y, Ikeguchi M, Sugai S Comparison of the transient folding intermediates in lysozyme and a lactalbumin. Biochemistry 24: Kuwata K, Li H, Yamada H, Batt CA, Goto Y, et al High pressure NMR reveals a variety of fluctuating conformers in b lactoglobulin. J Mol Biol 305: Kuwata K, Li H, Yamada H, Legname G, Prusiner SB, et al Locally disordered conformer of the hamster prion protein: A crucial intermediate to Prpsc? Biochemistry 41: Kuwajima K, Nitta K, Yoneyama M, Sugai S Three state denaturation of a lactalbumin by guanidine hydrochloride. J Mol Biol 6: Lakey JH, Raggett EM Measuring protein protein interactions. Curr Opin Struct Biol 8: Lassalle MW, Li H, Yamada H, Akasaka K, Redfield C Pressure induced unfolding of the molten globule of all Ala a lactalbumin. Protein Sci 12: Lee S, Tsai FTF Molecular chaperones in protein quality control. J Biochem Mol Biol 38: Levinthal C How to fold graciously. DeBrunner P, Tsibris JCM, Munck E, editors. Mössbauer spectroscopy in biological systems: Proceedings of a meeting held at Allerton house, Monticello, Illinois. University of Illinois Press; pp Levy Y, Cho SS, Onuchic JN, Wolynes PG A survey of flexible protein binding mechanisms and their transition states using native topology based energy landscapes. J Mol Biol 346: Levy Y, Wolynes PG, Onuchic JN Protein topology determines binding mechanism. Proc Natl Acad Sci USA 1: Lewis VA, Hynes GM, Zheng D, Saibil H, Willison K T complex polypeptide 1 is a subunit of a heteromeric particle in the eukaryotic cytosol. Nature 358: Li WF, Zhou XX, Lu P Structural features of thermozymes. Biotechnol Adv 23: Lindorff Larsen K, Rogen P, Paci E, Vendruscolo M, Dobson CM Protein folding and the organization of the protein topology universe. Trends Biochem Sci 30: Lindorff Larsen K, Vendruscolo M, Paci E, Dobson CM Transition states for protein folding have native topologies despite high structural variability. Nat Struct Mol Biol 11: Lipman EA, Schuler B, Bakajin O, Eaton WA Singlemolecule measurement of protein folding kinetics. Science 301: Liu X, Siegel DL, Fan P, Brodsky B, Baum J Direct NMR measurement of folding kinetics of a trimeric peptide. Biochemistry 35: Lomas DA, Carrell RW Serpinopathies and the conformational dementias. Nat Rev Genet 3: Luders J, Demand J, Hohfeld J The ubiquitin related bag 1 provides a link between the molecular chaperones hsc70/hsp70 and the proteasome. J Biol Chem 275: Lue LF, Kuo YM, Roher AE, Brachova L, Shen Y, et al Soluble amyloid b peptide concentration as a predictor of synaptic change in Alzheimer s disease. Am J Pathol 155: Lyon CE, Suh E, Dobson CM, Hore PJ Probing the exposure of tyrosine and tryptophan residues in partially folded proteins and folding intermediates by CIDNP pulselabeling. J Am Chem Soc 124: Ma H, Gruebele M Kinetics are probe dependent during downhill folding of an engineered l6 85 protein. Proc Natl Acad Sci USA 2: Makhatadze GI, Privalov PL Energetics of protein structure. Adv Protein Chem 47: Maki K, Ikura T, Hayano T, Takahashi N, Kuwajima K Effects of proline mutations on the folding of staphylococcal nuclease. Biochemistry 38: Mandel S, Grunblatt E, RiedererP, Amariglio N, Jacob Hirsch J, et al Gene expression profiling of sporadic Parkinson s disease substantia nigra pars compacta reveals impairment of ubiquitin proteasome subunits, skp1a, aldehyde dehydrogenase, and chaperone hsc 70. Ann N Y Acad Sci 53: Marti DN, Bjelic S, Lu M, Bosshard HR, Jelesarov I Fast folding of the HIV 1 and SIV gp41 six helix bundles. J Mol Biol 336: 1-8.
37 Protein folding 37 Mateu MG, Sanchez Del Pino MM, Fersht AR Mechanism of folding and assembly of a small tetrameric protein domain from tumor suppressor p53. Nat Struct Biol 6: Matouschek A, Kellis JTJ, Serrano L, Bycroft M, Fersht AR Transient folding intermediates characterized by protein engineering. Nature 346: Matouschek A, Otzen DE, Itzhaki LS, Jackson SE, Fersht AR Movement of the position of the transition state in protein folding. Biochemistry 34: Matthews CR, Hurle MR Mutant sequences as probes of protein folding mechanisms. Bioessays 6: Mazhul VM, Zaitseva EM, Shavlovsky MM, Stepanenko OV, Kuznetsova IM, et al Monitoring of actin unfolding by room temperature tryptophan phosphorescence. Biochemistry 42: McCarney ER, Kohn JE, Plaxco KW Is there or isn t there? The case for (and against) residual structure in chemically denatured proteins. Crit Rev Biochem Mol Biol 40: McLean CA, Cherny RA, Fraser FW, Fuller SJ, Smith MJ, et al Soluble pool of Ab amyloid as a determinant of severity of neurodegeneration in Alzheimer s disease. Ann Neurol 46: Meersman F, Smeller L, Heremans K Protein stability and dynamics in the pressure temperature plane. Biochim Biophys Acta 1764: Mersol JV, Steel DG, Gafni A Detection of intermediate protein conformations by room temperature tryptophan phosphorescence spectroscopy during denaturation of Escherichia coli alkaline phosphatase. Biophys Chem 48: Miranker A, Robinson CV, Radford SE, Dobson CM Investigation of protein folding by mass spectrometry. FASEB J : Missiakas D, Betton JM, Chaffotte A, Minard P, Yon JM Kinetic studies of the refolding of yeast phosphoglycerate kinase: Comparison with the isolated engineered domains. Protein Sci 1: Mohimen A, Dobo A, Hoerner JK, Kaltashov IA A chemometric approach to detection and characterization of multiple protein conformers in solution using electrospray ionization mass spectrometry. Anal Chem 75: Mok KH, Hore PJ Photo CIDNP NMR methods for studying protein folding. Methods 34: Mok KH, Nagashima T, Day IJ, Jones JA, Jones CJV, et al Rapid sample mixing technique for transient NMR and photo CIDNP spectroscopy: Applications to real time protein folding. J Am Chem Soc 125: Monera OD, Shaw GS, Zhu BY, Sykes BD, Kay CM, et al Role of interchain a helical hydrophobic interactions in Ca 2þ affinity, formation, and stability of a two site domain in troponin c. Protein Sci 1: Muller K, Garel JR Folding of aspartokinase homoserine dehydrogenase I is dominated by tertiary interactions. Biochemistry 23: Munoz V, Eaton WA A simple model for calculating the kinetics of protein folding from three dimensional structures. Proc Natl Acad Sci USA 96: Myers SL, Thomson NH, Radford SE, Ashcroft AE. 2006a. Investigating the structural properties of amyloid like fibrils formed in vitro from b 2 microglobulin using limited proteolysis and electrospray ionisation mass spectrometry. Rapid Commun Mass Spectrom 20: Myers SL, Jones S, Jahn TR, Morten IJ, Tennent GA, et al. 2006b. A systematic study of the effect of physiological factors on b 2 microglobulin amyloid formation at neutral ph. Biochemistry 45: Nelson ED, Onuchic JN Proposed mechanism for stability of proteins to evolutionary mutations. Proc Natl Acad Sci USA 95: Nelson R, Eisenberg D Recent atomic models of amyloid fibril structure. Curr Opin Struct Biol 16: Niraula TN, Konno T, Li H, Yamada H, Akasaka K, et al Pressure dissociable reversible assembly of intrinsically denatured lysozyme is a precursor for amyloid fibrils. Proc Natl Acad Sci USA 1: Novokhatny VV, Kudinov SA, Privalov PL Domains in human plasminogen. J Mol Biol 179: Nozaka M, Kuwajima K, Nitta K, Sugai S Detection and characterization of the intermediate on the folding pathway of human a lactalbumin. Biochemistry 17: Odefey C, Mayr LM, Schmid FX Non prolyl cis trans peptide bond isomerization as a rate determining step in protein unfolding and refolding. J Mol Biol 245: Ohgushi M, Wada A Molten globule state : A compact form of globular proteins with mobile side chains. FEBS Lett 164: Onuchic JN, Wolynes PG Theory of protein folding. Curr Opin Struct Biol 14: Onuchic JN, Socci ND, Luthey Schulten Z, Wolynes PG Protein folding funnels: The nature of the transition state ensemble. Fold Des 1: Ostermann J, Horwich AL, Neupert W, Hartl FU Protein folding in mitochondria requires complex formation with hsp60 and ATP hydrolysis. Nature 341: Osvath S, Gruebele M Proline can have opposite effects on fast and slow protein folding phases. Biophys J 85: Osvath S, Herenyi L, Zavodszky P, Fidy J, Kohler G Hierarchic finite level energy landscape model to describe AU4
38 38 Protein folding the refolding kinetics of phosphoglycerate kinase. J Biol Chem. Osvath S, Sabelko JJ, Gruebele M Tuning the heterogeneous early folding dynamics of phosphoglycerate kinase. J Mol Biol 333: Otzen DE, Kristensen O, Oliveberg M Designed protein tetramer zipped together with a hydrophobic Alzheimer homology: A structural clue to amyloid assembly. Proc Natl Acad Sci USA 97: Pace CN, Shirley BA, McNutt M, Gajiwala K Forces contributing to the conformational stability of proteins. FASEB J : Paci E, Clarke J, Steward A, Vendruscolo M, Karplus M Self consistent determination of the transition state for protein folding: Application to a fibronectin type III domain. Proc Natl Acad Sci USA 0: Pande VS, Baker I, Chapman J, Elmer SP, Khaliq S, et al Atomistic protein folding simulations on the submillisecond time scale using worldwide distributed computing. Biopolymers 68: Parrini C, Taddei N, Ramazzotti M, Degl Innocenti D, Ramponi G, et al Glycine residues appear to be evolutionarily conserved for their ability to inhibit aggregation. Structure (Camb) 13: Pearl LH, Prodromou C Structure and mechanism of the hsp90 molecular chaperone machinery. Annu Rev Biochem 75: Pedersen JS, Christensen G, Otzen DE Modulation of s6 fibrillation by unfolding rates and gatekeeper residues. J Mol Biol 341: Peters TJ, Davidson LK The biosynthesis of rat serum albumin. In vivo studies on the formation of the disulfide bonds. J Biol Chem 257: Petkova AT, Leapman RD, Guo Z, Yau W, Mattson MP, et al Self propagating, molecular level polymorphism in Alzheimer s b amyloid fibrils. Science 307: Pfeil W The problem of the stability globular proteins. Mol Cell Biochem 40: Plaxco KW, Simons KT, Baker D Contact order, transition state placement and the refolding rates of single domain proteins. J Mol Biol 277: Plotkin S, Wang J, Wolynes P Correlated energy landscape model for finite, random heteropolymers. Phys Rev E Stat Nonlin Soft Matter Phys 53: Pokarowski P, Kolinski A, Skolnick J A minimal physically realistic protein like lattice model: Designing an energy landscape that ensures all or none folding to a unique native state. Biophys J 84: Prabhu NV, Sharp KA Heat capacity in proteins. Annu Rev Phys Chem 56: Privalov PL Intermediate states in protein folding. J Mol Biol 258: Ptitsyn OB The molten globule state. Protein folding. Creighton TE, editor. New York: W.H. Freeman & Co.; pp Ptitsyn OB Molten globule and protein folding. Adv Protein Chem 47: Puett D The equilibrium unfolding parameters of horse and sperm whale myoglobin. Effects of guanidine hydrochloride, urea, and acid. J Biol Chem 248: Radford SE, Dobson CM, Evans PA The folding of hen lysozyme involves partially structured intermediates and multiple pathways. Nature 358: Raffen R, Dieckman LJ, Szpunar M, Wunschl C, Pokkuluri PR, et al Physicochemical consequences of amino acid variations that contribute to fibril formation by immunoglobulin light chains. Protein Sci 8: Ramachandran GN, Mitra AK An explanation for the rare occurrence of cis peptide units in proteins and polypeptides. J Mol Biol 7: Ramboarina S, Redfield C Structural characterisation of the human a lactalbumin molten globule at high temperature. J Mol Biol 330: Raschke TM, Kho J, Marqusee S Confirmation of the hierarchical folding of RNase H: A protein engineering study. Nat Struct Biol 6: Razvi A, Scholtz JM Lessons in stability from thermophilic proteins. Protein Sci 15: Redfield C Using nuclear magnetic resonance spectroscopy to study molten globule states of proteins. Methods 34: Refaee M, Tezuka T, Akasaka K, Williamson MP Pressure dependent changes in the solution structure of hen egg white lysozyme. J Mol Biol 327: Richardson JS, Richardson DC Natural b sheet proteins use negative design to avoid edge to edge aggregation. Proc Natl Acad Sci USA 99: Rief M, Gautel M, Oesterhelt F, Fernandez JM, Gaub HE Reversible unfolding of individual titin immunoglobulin domains by AFM. Science 276: Robinson CR, Sauer RT Equilibrium stability and submillisecond refolding of a designed single chain arc repressor. Biochemistry 35: Robson B, Pain RH The mechanism of folding of globular proteins. Equilibria and kinetics of conformational transitions of penicillinase from Staphylococcus aureus involving a state of intermediate conformation. Biochem J 155: Roder H, Elove GA, Englander SW Structural characterization of folding intermediates in cytochrome c by H exchange labelling and proton NMR. Nature 335: Roder H, Elöve GA, Shastry MC. R Early stages of protein folding. Mechanisms of Protein Folding. Pain RH, editor. Oxford: Oxford University Press; pp
39 Protein folding 39 Roder H, Shastry MR Methods for exploring early events in protein folding. Curr Opin Struct Biol 9: Roder H, Maki K, Cheng H Early events in protein folding explored by rapid mixing methods. Chem Rev 6: Roder H, Maki K, Cheng H, Shastry MCR Rapid mixing methods for exploring the kinetics of protein folding. Methods 34: Ross CA, Pickart CM The ubiquitin proteasome pathway in Parkinson s disease and other neurodegenerative diseases. Trends Cell Biol 14: Rounsevell R, Forman JR, Clarke J Atomic force microscopy: Mechanical unfolding of proteins. Methods 34: Rousseau F, Schymkowitz JWH, Itzhaki LS The unfolding story of three dimensional domain swapping. Structure 11: Rudiger S, Germeroth L, Schneider Mergener J, Bukau B Substrate specificity of the DNAk chaperone determined by screening cellulose bound peptide libraries. EMBO J 16: Rudolph R, Siebendritt R, NesslauerG, Sharma AK, Jaenicke R Folding of an all b protein: Independent domain folding in g II crystallin from calf eye lens. Proc Natl Acad Sci USA 87: Sabelko J, Ervin J, Gruebele M Observation of strange kinetics in protein folding. Proc Natl Acad Sci USA 96: Sali A, Shakhnovich E, Karplus M How does a protein fold? Nature 369: Scharnagl C, Reif M, Friedrich J Stability of proteins: Temperature, pressure and the role of the solvent. Biochim Biophys Acta 1749: Schellman JA Temperature, stability, and the hydrophobic interaction. Biophys J 73: Scherer G, Kramer ML, Schutkowski M, Reimer U, Fischer G Barriers to rotation of secondary amide peptide bonds. J Am Chem Soc 120: Schmid FX, Mayr LM, Mucke M, Schonbrunner ER Prolyl isomerases: Role in protein folding. Adv Protein Chem 44: Schonbrunner ER, Schmid FX Peptidyl prolyl cis trans isomerase improves the efficiency of protein disulfide isomerase as a catalyst of protein folding. Proc Natl Acad Sci USA 89: Schwaller M, Wilkinson B, Gilbert HF Reductionreoxidation cycles contribute to catalysis of disulfide isomerization by protein disulfide isomerase. J Biol Chem 278: Schwartz R, Istrail S, King J Frequencies of amino acid strings in globular protein sequences indicate suppression of blocks of consecutive hydrophobic residues. Protein Sci : Serpell LC, Sunde M, Benson MD, Tennent GA, Pepys MB, et al The protofilament substructure of amyloid fibrils. J Mol Biol 300: Serrano L, Matouschek A, Fersht AR The folding of an enzyme. III. structure of the transition state for unfolding of barnase analysed by a protein engineering procedure. J Mol Biol 224: Sha B, Lee S, Cyr DM The crystal structure of the peptide binding fragment from the yeast Hsp40 protein sis1. Structure 8: Shakhnovich EI, Finkelstein AV Theory of cooperative transitions in protein molecules. I. Why denaturation of globular protein is a first order phase transition. Biopolymers 28: Sharma AK, Minke Gogl V, Gohl P, Siebendritt R, Jaenicke R, et al Limited proteolysis of g II crystallin from calf eye lens. Physicochemical studies on the N terminal domain and the intact two domain protein. Eur J Biochem 194: Shastry MC, Luck SD, Roder H A continuous flow capillary mixing method to monitor reactions on the microsecond time scale. Biophys J 74: Shaw PE Peptidyl prolyl isomerases: A new twist to transcription. EMBO Rep 3: Shoelson SE, Sivaraja M, Williams KP, Hu P, Schlessinger J, et al Specific phosphopeptide binding regulates a conformational change in the PI 3 kinase SH2 domain associated with enzyme activation. EMBO J 12: Shoemaker BA, Portman JJ, Wolynes PG Speeding molecular recognition by using the folding funnel: The fly casting mechanism. Proc Natl Acad Sci USA 97: Shortle D The denatured state (the other half of the folding equation) and its role in protein stability. FASEB J : Shtilerman M, Lorimer GH, Englander SW Chaperonin function: Folding by forced unfolding. Science 284: Siegert R, Leroux MR, Scheufler C, Hartl FU, Moarefi I Structure of the molecular chaperone prefoldin: Unique interaction of multiple coiled coil tentacles with unfolded proteins. Cell 3: Silva JL, Cordeiro Y, Foguel D Protein folding and aggregation: Two sides of the same coin in the condensation of proteins revealed by pressure studies. Biochim Biophys Acta 1764: Smeller L Pressure temperature phase diagrams of biomolecules. Biochim Biophys Acta 1595: Smith AV, Hall CK Assembly of a tetrameric a helical bundle: Computer simulations on an intermediateresolution protein model. Proteins 44:
40 40 Protein folding Smith LJ, Fiebig KM, Schwalbe H, Dobson CM The concept of a random coil. Residual structure in peptides and denatured proteins. Fold Des 1: R95-6. Snow CD, Qiu L, Du D, Gai F, Hagen SJ, et al Trp zipper folding kinetics by molecular dynamics and temperaturejump spectroscopy. Proc Natl Acad Sci USA 1: Song J, Bai P, Luo L, Peng ZY Contribution of individual residues to formation of the native like tertiary topology in the a lactalbumin molten globule. J Mol Biol 280: Sorin EJ, Pande VS Exploring the helix coil transition via all atom equilibrium ensemble simulations. Biophys J 88: Stefani M, Dobson CM Protein aggregation and aggregate toxicity: New insights into protein folding, misfolding diseases and biological evolution. J Mol Med 81: Stein RL Mechanism of enzymatic and nonenzymatic prolyl cis trans isomerization. Adv Protein Chem 44: Steward A, Adhya S, Clarke J Sequence conservation in Ig like domains: The role of highly conserved proline residues in the fibronectin type III superfamily. J Mol Biol 318: Stewart DE, Sarkar A, Wampler JE Occurrence and role of cis peptide bonds in protein structures. J Mol Biol 214: Stine WBJ, Dahlgren KN, Krafft GA, LaDu MJ In vitro characterization of conditions for amyloid b peptide oligomerization and fibrillogenesis. J Biol Chem 278: Street TO, Bradley CM, Barrick D An improved experimental system for determining small folding entropy changes resulting from proline to alanine substitutions. Protein Sci 14: Sunde M, Blake C The structure of amyloid fibrils by electron microscopy and X ray diffraction. Adv Protein Chem 50: Szilagyi A, Zavodszky P Structural differences between mesophilic, moderately thermophilic and extremely thermophilic protein subunits: Results of a comprehensive survey. Structure 8: Takahashi S, Yeh SR, Das TK, Chan CK, Gottfried DS, et al Folding of cytochrome c initiated by submillisecond mixing. Nat Struct Biol 4: Takayama S, Bimston DN, Matsuzawa S, Freeman BC, Aime Sempe C, et al Bag 1 modulates the chaperone activity of hsp70/hsc70. EMBO J 16: Teale JM, Benjamin DC Antibody as immunological probe for studying refolding of bovine serum albumin. Refolding within each domain. J Biol Chem 252: Terada K, Mori M Human dnaj homologs dj2 and dj3, and bag 1 are positive cochaperones of hsc70. J Biol Chem 275: Teter SA, Houry WA, Ang D, Tradler T, Rockabrand D, et al Polypeptide flux through bacterial hsp70: Dnak cooperates with trigger factor in chaperoning nascent chains. Cell 97: Texter FL, Spencer DB, Rosenstein R, Matthews CR Intramolecular catalysis of a proline isomerization reaction in the folding of dihydrofolate reductase. Biochemistry 31: Tirado Rives J, Jorgensen WL Molecular dynamics simulations of the unfolding of apomyoglobin in water. Biochemistry 32: Todd MJ, Lorimer GH, Thirumalai D Chaperoninfacilitated protein folding: Optimization of rate and yield by an iterative annealing mechanism. Proc Natl Acad Sci USA 93: Tsai C, Maizel JVJ, Nussinov R The hydrophobic effect: A new insight from cold denaturation and a two state water structure. Crit Rev Biochem Mol Biol 37: Tsai J, Levitt M, Baker D Hierarchy of structure loss in MD simulations of src SH3 domain unfolding. J Mol Biol 291: Tsuji T, Chrunyk BA, Chen X, Matthews CR Mutagenic analysis of the interior packing of an a/b barrel protein. Effects on the stabilities and rates of interconversion of the native and partially folded forms of the a subunit of tryptophan synthase. Biochemistry 32: Tsunenaga M, Goto Y, Kawata Y, Hamaguchi K Unfolding and refolding of a type k immunoglobulin light chain and its variable and constant fragments. Biochemistry 26: Tycko R Molecular structure of amyloid fibrils: Insights from solid state NMR. Q Rev Biophys: Udgaonkar JB, Baldwin RL NMR evidence for an early framework intermediate on the folding pathway of ribonuclease A. Nature 335: Uversky VN, Li J, Souillac P, Millett IS, Doniach S, et al Biophysical properties of the synucleins and their propensities to fibrillate: Inhibition of a synuclein assembly by b and g synucleins. J Biol Chem 277: Uzawa T, Akiyama S, Kimura T, Takahashi S, Ishimori K, et al Collapse and search dynamics of apomyoglobin folding revealed by submillisecond observations of a helical content and compactness. Proc Natl Acad Sci USA 1: Vanhove M, Lejeune A, Pain RH b lactamases as models for protein folding studies. Cell Mol Life Sci 54: Vaucheret H, Signon L, Le Bras G, Garel JR Mechanism of renaturation of a large protein, aspartokinase homoserine dehydrogenase. Biochemistry 26:
41 Protein folding 41 Vendruscolo M, Zurdo J, MacPhee CE, Dobson CM Protein folding and misfolding: A paradigm of self assembly and regulation in complex biological systems. Philos Transact A Math Phys Eng Sci 361: Villegas V, Martinez JC, Aviles FX, Serrano L Structure of the transition state in the folding process of human procarboxypeptidase a2 activation domain. J Mol Biol 283: Wagner G, Wuthrich K Amide protein exchange and surface conformation of the basic pancreatic trypsin inhibitor in solution. Studies with two dimensional nuclear magnetic resonance. J Mol Biol 160: Walkenhorst WF, Green SM, Roder H Kinetic evidence for folding and unfolding intermediates in staphylococcal nuclease. Biochemistry 36: Walker KW, Gilbert HF Scanning and escape during protein disulfide isomerase assisted protein folding. J Biol Chem 272: Wand AJ, Roder H, Englander SW Two dimensional 1 H NMR studies of cytochrome c: Hydrogen exchange in the N terminal helix. Biochemistry 25: Wedemeyer WJ, Welker E, Narayan M, Scheraga HA Disulfide bonds and protein folding. Biochemistry 39: Wedemeyer WJ, Welker E, Scheraga HA Proline cistrans isomerization and protein folding. Biochemistry 41: Welker E, Maki K, Shastry MCR, Juminaga D, Bhat R, et al Ultrarapid mixing experiments shed new light on the characteristics of the initial conformational ensemble during the folding of ribonuclease A. Proc Natl Acad Sci USA 1: Westermark P, Benson MD, Buxbaum JN, Cohen AS, Frangione B, et al Amyloid: Toward terminology clarification. Report from the nomenclature committee of the international society of amyloidosis. Amyloid 12: 1-4. Wetlaufer DB Nucleation, rapid folding, and globular intrachain regions in proteins. Proc Natl Acad Sci USA 70: Williams KP, Shoelson SE Cooperative self assembly of SH2 domain fragments restores phosphopeptide binding. Biochemistry 32: Wolynes PG Energy landscapes and solved proteinfolding problems. Philos Transact A Math Phys Eng Sci 363: ; discussion Wolynes PG, Onuchic JN, Thirumalai D Navigating the folding routes. Science 267: Wong KB, Clarke J, Bond CJ, Neira JL, Freund SM, et al Towards a complete description of the structural and dynamic properties of the denatured state of barnase and the role of residual structure in folding. J Mol Biol 296: Wong KP, Tanford C Denaturation of bovine carbonic anhydrase B by guanidine hydrochloride. A process involving separable sequential conformational transitions. J Biol Chem 248: Wood LC, White TB, Ramdas L, Nall BT Replacement of a conserved proline eliminates the absorbance detected slow folding phase of iso 2 cytochrome c. Biochemistry 27: Wu Y, Matthews CR Parallel channels and rate limiting steps in complex protein folding reactions: Prolyl isomerization and the a subunit of trp synthase, a TIM barrel protein. J Mol Biol 323: Yamamoto S, Hasegawa K, Yamaguchi I, Tsutsumi S, Kardos J, et al Low concentrations of sodium dodecyl sulfate induce the extension of b 2 microglobulin related amyloid fibrils at a neutral ph. Biochemistry 43: Yang WY, Gruebele M Folding at the speed limit. Nature 423: Yang WY, Gruebele M. 2004a. Detection dependent kinetics as a probe of folding landscape microstructure. J Am Chem Soc 126: Yang WY, Gruebele M. 2004b. Folding l repressor at its speed limit. Biophys J 87: Zarrine Afsar A, Larson SM, Davidson AR The family feud: Do proteins with similar structures fold via the same pathway? Curr Opin Struct Biol 15: Zeeb M, Balbach J Protein folding studied by real time NMR spectroscopy. Methods 34: Zhang Y, Skolnick J Automated structure prediction of weakly homologous proteins on a genomic scale. Proc Natl Acad Sci USA 1: Zhu X, Zhao X, Burkholder WF, Gragerov A, Ogata CM, et al Structural analysis of substrate binding by the molecular chaperone Dnak. Science 272: Zimmerman SB, Trach SO Estimation of macromolecule concentrations and excluded volume effects for the cytoplasm of Escherichia coli. J Mol Biol 222: Zimmerman SS, Scheraga HA Stability of cis, trans, and nonplanar peptide groups. Macromolecules 9:
42 Neural Protein Metabolism and Function Chapter No.: Author Query Form Query Refs. Details Required Author s response AU1 AU2 AU3 AU4 AU5 Should native fold be changed to native state? Can...numerous examples, including... be changed to...numerous proteins, examples of which include...? Should Rubisco be changed to Ru- BisCO? Please provide page range and vol. no. for Osvath et al. (2006). Please provide publisher location.
Protein Folding. The resulting three-dimensional structure is determined by the amino acid sequence (Anfinsen's dogma).
 Protein Folding Protein folding is the physical process by which a polypeptide folds into its characteristic and functional three-dimensional structure from random coil. Each protein exists as an unfolded
Protein Folding Protein folding is the physical process by which a polypeptide folds into its characteristic and functional three-dimensional structure from random coil. Each protein exists as an unfolded
Hydrogen Bonds The electrostatic nature of hydrogen bonds
 Hydrogen Bonds Hydrogen bonds have played an incredibly important role in the history of structural biology. Both the structure of DNA and of protein a-helices and b-sheets were predicted based largely
Hydrogen Bonds Hydrogen bonds have played an incredibly important role in the history of structural biology. Both the structure of DNA and of protein a-helices and b-sheets were predicted based largely
Peptide bonds: resonance structure. Properties of proteins: Peptide bonds and side chains. Dihedral angles. Peptide bond. Protein physics, Lecture 5
 Protein physics, Lecture 5 Peptide bonds: resonance structure Properties of proteins: Peptide bonds and side chains Proteins are linear polymers However, the peptide binds and side chains restrict conformational
Protein physics, Lecture 5 Peptide bonds: resonance structure Properties of proteins: Peptide bonds and side chains Proteins are linear polymers However, the peptide binds and side chains restrict conformational
(c) How would your answers to problem (a) change if the molecular weight of the protein was 100,000 Dalton?
 Problem 1. (12 points total, 4 points each) The molecular weight of an unspecified protein, at physiological conditions, is 70,000 Dalton, as determined by sedimentation equilibrium measurements and by
Problem 1. (12 points total, 4 points each) The molecular weight of an unspecified protein, at physiological conditions, is 70,000 Dalton, as determined by sedimentation equilibrium measurements and by
Disulfide Bonds at the Hair Salon
 Disulfide Bonds at the Hair Salon Three Alpha Helices Stabilized By Disulfide Bonds! In order for hair to grow 6 inches in one year, 9 1/2 turns of α helix must be produced every second!!! In some proteins,
Disulfide Bonds at the Hair Salon Three Alpha Helices Stabilized By Disulfide Bonds! In order for hair to grow 6 inches in one year, 9 1/2 turns of α helix must be produced every second!!! In some proteins,
Built from 20 kinds of amino acids
 Built from 20 kinds of amino acids Each Protein has a three dimensional structure. Majority of proteins are compact. Highly convoluted molecules. Proteins are folded polypeptides. There are four levels
Built from 20 kinds of amino acids Each Protein has a three dimensional structure. Majority of proteins are compact. Highly convoluted molecules. Proteins are folded polypeptides. There are four levels
18.2 Protein Structure and Function: An Overview
 18.2 Protein Structure and Function: An Overview Protein: A large biological molecule made of many amino acids linked together through peptide bonds. Alpha-amino acid: Compound with an amino group bonded
18.2 Protein Structure and Function: An Overview Protein: A large biological molecule made of many amino acids linked together through peptide bonds. Alpha-amino acid: Compound with an amino group bonded
Protein Physics. A. V. Finkelstein & O. B. Ptitsyn LECTURE 1
 Protein Physics A. V. Finkelstein & O. B. Ptitsyn LECTURE 1 PROTEINS Functions in a Cell MOLECULAR MACHINES BUILDING BLOCKS of a CELL ARMS of a CELL ENZYMES - enzymatic catalysis of biochemical reactions
Protein Physics A. V. Finkelstein & O. B. Ptitsyn LECTURE 1 PROTEINS Functions in a Cell MOLECULAR MACHINES BUILDING BLOCKS of a CELL ARMS of a CELL ENZYMES - enzymatic catalysis of biochemical reactions
Advanced Medicinal & Pharmaceutical Chemistry CHEM 5412 Dept. of Chemistry, TAMUK
 Advanced Medicinal & Pharmaceutical Chemistry CHEM 5412 Dept. of Chemistry, TAMUK Dai Lu, Ph.D. [email protected] Tel: 361-221-0745 Office: RCOP, Room 307 Drug Discovery and Development Drug Molecules Medicinal
Advanced Medicinal & Pharmaceutical Chemistry CHEM 5412 Dept. of Chemistry, TAMUK Dai Lu, Ph.D. [email protected] Tel: 361-221-0745 Office: RCOP, Room 307 Drug Discovery and Development Drug Molecules Medicinal
Nafith Abu Tarboush DDS, MSc, PhD [email protected] www.facebook.com/natarboush
 Nafith Abu Tarboush DDS, MSc, PhD [email protected] www.facebook.com/natarboush α-keratins, bundles of α- helices Contain polypeptide chains organized approximately parallel along a single axis: Consist
Nafith Abu Tarboush DDS, MSc, PhD [email protected] www.facebook.com/natarboush α-keratins, bundles of α- helices Contain polypeptide chains organized approximately parallel along a single axis: Consist
Combinatorial Biochemistry and Phage Display
 Combinatorial Biochemistry and Phage Display Prof. Valery A. Petrenko Director - Valery Petrenko Instructors Galina Kouzmitcheva and I-Hsuan Chen Auburn 2006, Spring semester COMBINATORIAL BIOCHEMISTRY
Combinatorial Biochemistry and Phage Display Prof. Valery A. Petrenko Director - Valery Petrenko Instructors Galina Kouzmitcheva and I-Hsuan Chen Auburn 2006, Spring semester COMBINATORIAL BIOCHEMISTRY
Peptide Bonds: Structure
 Peptide Bonds: Structure Peptide primary structure The amino acid sequence, from - to C-terminus, determines the primary structure of a peptide or protein. The amino acids are linked through amide or peptide
Peptide Bonds: Structure Peptide primary structure The amino acid sequence, from - to C-terminus, determines the primary structure of a peptide or protein. The amino acids are linked through amide or peptide
Carbohydrates, proteins and lipids
 Carbohydrates, proteins and lipids Chapter 3 MACROMOLECULES Macromolecules: polymers with molecular weights >1,000 Functional groups THE FOUR MACROMOLECULES IN LIFE Molecules in living organisms: proteins,
Carbohydrates, proteins and lipids Chapter 3 MACROMOLECULES Macromolecules: polymers with molecular weights >1,000 Functional groups THE FOUR MACROMOLECULES IN LIFE Molecules in living organisms: proteins,
IV. -Amino Acids: carboxyl and amino groups bonded to -Carbon. V. Polypeptides and Proteins
 IV. -Amino Acids: carboxyl and amino groups bonded to -Carbon A. Acid/Base properties 1. carboxyl group is proton donor! weak acid 2. amino group is proton acceptor! weak base 3. At physiological ph: H
IV. -Amino Acids: carboxyl and amino groups bonded to -Carbon A. Acid/Base properties 1. carboxyl group is proton donor! weak acid 2. amino group is proton acceptor! weak base 3. At physiological ph: H
Amino Acids. Amino acids are the building blocks of proteins. All AA s have the same basic structure: Side Chain. Alpha Carbon. Carboxyl. Group.
 Protein Structure Amino Acids Amino acids are the building blocks of proteins. All AA s have the same basic structure: Side Chain Alpha Carbon Amino Group Carboxyl Group Amino Acid Properties There are
Protein Structure Amino Acids Amino acids are the building blocks of proteins. All AA s have the same basic structure: Side Chain Alpha Carbon Amino Group Carboxyl Group Amino Acid Properties There are
Proteins the primary biological macromolecules of living organisms
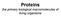 Proteins the primary biological macromolecules of living organisms Protein structure and folding Primary Secondary Tertiary Quaternary structure of proteins Structure of Proteins Protein molecules adopt
Proteins the primary biological macromolecules of living organisms Protein structure and folding Primary Secondary Tertiary Quaternary structure of proteins Structure of Proteins Protein molecules adopt
The peptide bond is rigid and planar
 Level Description Bonds Primary Sequence of amino acids in proteins Covalent (peptide bonds) Secondary Structural motifs in proteins: α- helix and β-sheet Hydrogen bonds (between NH and CO groups in backbone)
Level Description Bonds Primary Sequence of amino acids in proteins Covalent (peptide bonds) Secondary Structural motifs in proteins: α- helix and β-sheet Hydrogen bonds (between NH and CO groups in backbone)
Papers listed: Cell2. This weeks papers. Chapt 4. Protein structure and function
 Papers listed: Cell2 During the semester I will speak of information from several papers. For many of them you will not be required to read these papers, however, you can do so for the fun of it (and it
Papers listed: Cell2 During the semester I will speak of information from several papers. For many of them you will not be required to read these papers, however, you can do so for the fun of it (and it
A. A peptide with 12 amino acids has the following amino acid composition: 2 Met, 1 Tyr, 1 Trp, 2 Glu, 1 Lys, 1 Arg, 1 Thr, 1 Asn, 1 Ile, 1 Cys
 Questions- Proteins & Enzymes A. A peptide with 12 amino acids has the following amino acid composition: 2 Met, 1 Tyr, 1 Trp, 2 Glu, 1 Lys, 1 Arg, 1 Thr, 1 Asn, 1 Ile, 1 Cys Reaction of the intact peptide
Questions- Proteins & Enzymes A. A peptide with 12 amino acids has the following amino acid composition: 2 Met, 1 Tyr, 1 Trp, 2 Glu, 1 Lys, 1 Arg, 1 Thr, 1 Asn, 1 Ile, 1 Cys Reaction of the intact peptide
Topic 2: Energy in Biological Systems
 Topic 2: Energy in Biological Systems Outline: Types of energy inside cells Heat & Free Energy Energy and Equilibrium An Introduction to Entropy Types of energy in cells and the cost to build the parts
Topic 2: Energy in Biological Systems Outline: Types of energy inside cells Heat & Free Energy Energy and Equilibrium An Introduction to Entropy Types of energy in cells and the cost to build the parts
Ionization of amino acids
 Amino Acids 20 common amino acids there are others found naturally but much less frequently Common structure for amino acid COOH, -NH 2, H and R functional groups all attached to the a carbon Ionization
Amino Acids 20 common amino acids there are others found naturally but much less frequently Common structure for amino acid COOH, -NH 2, H and R functional groups all attached to the a carbon Ionization
Recap. Lecture 2. Protein conformation. Proteins. 8 types of protein function 10/21/10. Proteins.. > 50% dry weight of a cell
 Lecture 2 Protein conformation ecap Proteins.. > 50% dry weight of a cell ell s building blocks and molecular tools. More important than genes A large variety of functions http://www.tcd.ie/biochemistry/courses/jf_lectures.php
Lecture 2 Protein conformation ecap Proteins.. > 50% dry weight of a cell ell s building blocks and molecular tools. More important than genes A large variety of functions http://www.tcd.ie/biochemistry/courses/jf_lectures.php
Lecture 19: Proteins, Primary Struture
 CPS260/BGT204.1 Algorithms in Computational Biology November 04, 2003 Lecture 19: Proteins, Primary Struture Lecturer: Pankaj K. Agarwal Scribe: Qiuhua Liu 19.1 The Building Blocks of Protein [1] Proteins
CPS260/BGT204.1 Algorithms in Computational Biology November 04, 2003 Lecture 19: Proteins, Primary Struture Lecturer: Pankaj K. Agarwal Scribe: Qiuhua Liu 19.1 The Building Blocks of Protein [1] Proteins
CSC 2427: Algorithms for Molecular Biology Spring 2006. Lecture 16 March 10
 CSC 2427: Algorithms for Molecular Biology Spring 2006 Lecture 16 March 10 Lecturer: Michael Brudno Scribe: Jim Huang 16.1 Overview of proteins Proteins are long chains of amino acids (AA) which are produced
CSC 2427: Algorithms for Molecular Biology Spring 2006 Lecture 16 March 10 Lecturer: Michael Brudno Scribe: Jim Huang 16.1 Overview of proteins Proteins are long chains of amino acids (AA) which are produced
Chapter 12 - Proteins
 Roles of Biomolecules Carbohydrates Lipids Proteins 1) Catalytic 2) Transport 3) Regulatory 4) Structural 5) Contractile 6) Protective 7) Storage Nucleic Acids 12.1 -Amino Acids Chapter 12 - Proteins Amino
Roles of Biomolecules Carbohydrates Lipids Proteins 1) Catalytic 2) Transport 3) Regulatory 4) Structural 5) Contractile 6) Protective 7) Storage Nucleic Acids 12.1 -Amino Acids Chapter 12 - Proteins Amino
Chemical Bonds. Chemical Bonds. The Nature of Molecules. Energy and Metabolism < < Covalent bonds form when atoms share 2 or more valence electrons.
 The Nature of Molecules Chapter 2 Energy and Metabolism Chapter 6 Chemical Bonds Molecules are groups of atoms held together in a stable association. Compounds are molecules containing more than one type
The Nature of Molecules Chapter 2 Energy and Metabolism Chapter 6 Chemical Bonds Molecules are groups of atoms held together in a stable association. Compounds are molecules containing more than one type
The peptide bond Peptides and proteins are linear polymers of amino acids. The amino acids are
 Introduction to Protein Structure Proteins are large heteropolymers usually comprised of 50 2500 monomer units, although larger proteins are observed 7. The monomer units of proteins are amino acids. The
Introduction to Protein Structure Proteins are large heteropolymers usually comprised of 50 2500 monomer units, although larger proteins are observed 7. The monomer units of proteins are amino acids. The
http://faculty.sau.edu.sa/h.alshehri
 http://faculty.sau.edu.sa/h.alshehri Definition: Proteins are macromolecules with a backbone formed by polymerization of amino acids. Proteins carry out a number of functions in living organisms: - They
http://faculty.sau.edu.sa/h.alshehri Definition: Proteins are macromolecules with a backbone formed by polymerization of amino acids. Proteins carry out a number of functions in living organisms: - They
Chapter 3. Protein Structure and Function
 Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
CHEMISTRY STANDARDS BASED RUBRIC ATOMIC STRUCTURE AND BONDING
 CHEMISTRY STANDARDS BASED RUBRIC ATOMIC STRUCTURE AND BONDING Essential Standard: STUDENTS WILL UNDERSTAND THAT THE PROPERTIES OF MATTER AND THEIR INTERACTIONS ARE A CONSEQUENCE OF THE STRUCTURE OF MATTER,
CHEMISTRY STANDARDS BASED RUBRIC ATOMIC STRUCTURE AND BONDING Essential Standard: STUDENTS WILL UNDERSTAND THAT THE PROPERTIES OF MATTER AND THEIR INTERACTIONS ARE A CONSEQUENCE OF THE STRUCTURE OF MATTER,
Determination of Equilibrium Constants using NMR Spectrscopy
 CHEM 331L Physical Chemistry Laboratory Revision 1.0 Determination of Equilibrium Constants using NMR Spectrscopy In this laboratory exercise we will measure a chemical equilibrium constant using key proton
CHEM 331L Physical Chemistry Laboratory Revision 1.0 Determination of Equilibrium Constants using NMR Spectrscopy In this laboratory exercise we will measure a chemical equilibrium constant using key proton
Quaternary structure
 Quaternary structure Assembly of multiple polypeptide chains in one integral structure The arrangement of the subunits gives rise to a stable structure Subunits may be identical or different A common shorthand
Quaternary structure Assembly of multiple polypeptide chains in one integral structure The arrangement of the subunits gives rise to a stable structure Subunits may be identical or different A common shorthand
8/20/2012 H C OH H R. Proteins
 Proteins Rubisco monomer = amino acids 20 different amino acids polymer = polypeptide protein can be one or more polypeptide chains folded & bonded together large & complex 3-D shape hemoglobin Amino acids
Proteins Rubisco monomer = amino acids 20 different amino acids polymer = polypeptide protein can be one or more polypeptide chains folded & bonded together large & complex 3-D shape hemoglobin Amino acids
Lecture 13-14 Conformation of proteins Conformation of a protein three-dimensional structure native state. native condition
 Lecture 13-14 Conformation of proteins Conformation of a protein refers to the three-dimensional structure in its native state. There are many different possible conformations for a molecule as large as
Lecture 13-14 Conformation of proteins Conformation of a protein refers to the three-dimensional structure in its native state. There are many different possible conformations for a molecule as large as
This class deals with the fundamental structural features of proteins, which one can understand from the structure of amino acids, and how they are
 This class deals with the fundamental structural features of proteins, which one can understand from the structure of amino acids, and how they are put together. 1 A more detailed view of a single protein
This class deals with the fundamental structural features of proteins, which one can understand from the structure of amino acids, and how they are put together. 1 A more detailed view of a single protein
Invariant residue-a residue that is always conserved. It is assumed that these residues are essential to the structure or function of the protein.
 Chapter 6 The amino acid side chains have polar and nonpolar properties, and the relative hydrophobicity of the amino acid side chains is critical for the folding and stability of a protein. The more hydrophobic
Chapter 6 The amino acid side chains have polar and nonpolar properties, and the relative hydrophobicity of the amino acid side chains is critical for the folding and stability of a protein. The more hydrophobic
1 The water molecule and hydrogen bonds in water
 The Physics and Chemistry of Water 1 The water molecule and hydrogen bonds in water Stoichiometric composition H 2 O the average lifetime of a molecule is 1 ms due to proton exchange (catalysed by acids
The Physics and Chemistry of Water 1 The water molecule and hydrogen bonds in water Stoichiometric composition H 2 O the average lifetime of a molecule is 1 ms due to proton exchange (catalysed by acids
Molecular Dynamics Simulations
 Molecular Dynamics Simulations Yaoquan Tu Division of Theoretical Chemistry and Biology, Royal Institute of Technology (KTH) 2011-06 1 Outline I. Introduction II. Molecular Mechanics Force Field III. Molecular
Molecular Dynamics Simulations Yaoquan Tu Division of Theoretical Chemistry and Biology, Royal Institute of Technology (KTH) 2011-06 1 Outline I. Introduction II. Molecular Mechanics Force Field III. Molecular
Paper: 6 Chemistry 2.130 University I Chemistry: Models Page: 2 of 7. 4. Which of the following weak acids would make the best buffer at ph = 5.0?
 Paper: 6 Chemistry 2.130 University I Chemistry: Models Page: 2 of 7 4. Which of the following weak acids would make the best buffer at ph = 5.0? A) Acetic acid (Ka = 1.74 x 10-5 ) B) H 2 PO - 4 (Ka =
Paper: 6 Chemistry 2.130 University I Chemistry: Models Page: 2 of 7 4. Which of the following weak acids would make the best buffer at ph = 5.0? A) Acetic acid (Ka = 1.74 x 10-5 ) B) H 2 PO - 4 (Ka =
Structure of proteins
 Structure of proteins Primary structure: is amino acids sequence or the covalent structure (50-2500) amino acids M.Wt. of amino acid=110 Dalton (56 110=5610 Dalton). Single chain or more than one polypeptide
Structure of proteins Primary structure: is amino acids sequence or the covalent structure (50-2500) amino acids M.Wt. of amino acid=110 Dalton (56 110=5610 Dalton). Single chain or more than one polypeptide
1. A covalent bond between two atoms represents what kind of energy? a. Kinetic energy b. Potential energy c. Mechanical energy d.
 1. A covalent bond between two atoms represents what kind of energy? a. Kinetic energy b. Potential energy c. Mechanical energy d. Solar energy A. Answer a is incorrect. Kinetic energy is the energy of
1. A covalent bond between two atoms represents what kind of energy? a. Kinetic energy b. Potential energy c. Mechanical energy d. Solar energy A. Answer a is incorrect. Kinetic energy is the energy of
ENZYME SCIENCE AND ENGINEERING PROF. SUBHASH CHAND DEPARTMENT OF BIOCHEMICAL ENGINEERING AND BIOTECHNOLOGY IIT DELHI LECTURE 4 ENZYMATIC CATALYSIS
 ENZYME SCIENCE AND ENGINEERING PROF. SUBHASH CHAND DEPARTMENT OF BIOCHEMICAL ENGINEERING AND BIOTECHNOLOGY IIT DELHI LECTURE 4 ENZYMATIC CATALYSIS We will continue today our discussion on enzymatic catalysis
ENZYME SCIENCE AND ENGINEERING PROF. SUBHASH CHAND DEPARTMENT OF BIOCHEMICAL ENGINEERING AND BIOTECHNOLOGY IIT DELHI LECTURE 4 ENZYMATIC CATALYSIS We will continue today our discussion on enzymatic catalysis
Disaccharides consist of two monosaccharide monomers covalently linked by a glycosidic bond. They function in sugar transport.
 1. The fundamental life processes of plants and animals depend on a variety of chemical reactions that occur in specialized areas of the organism s cells. As a basis for understanding this concept: 1.
1. The fundamental life processes of plants and animals depend on a variety of chemical reactions that occur in specialized areas of the organism s cells. As a basis for understanding this concept: 1.
Science Standard Articulated by Grade Level Strand 5: Physical Science
 Concept 1: Properties of Objects and Materials Classify objects and materials by their observable properties. Kindergarten Grade 1 Grade 2 Grade 3 Grade 4 PO 1. Identify the following observable properties
Concept 1: Properties of Objects and Materials Classify objects and materials by their observable properties. Kindergarten Grade 1 Grade 2 Grade 3 Grade 4 PO 1. Identify the following observable properties
A disaccharide is formed when a dehydration reaction joins two monosaccharides. This covalent bond is called a glycosidic linkage.
 CH 5 Structure & Function of Large Molecules: Macromolecules Molecules of Life All living things are made up of four classes of large biological molecules: carbohydrates, lipids, proteins, and nucleic
CH 5 Structure & Function of Large Molecules: Macromolecules Molecules of Life All living things are made up of four classes of large biological molecules: carbohydrates, lipids, proteins, and nucleic
AP BIOLOGY 2008 SCORING GUIDELINES
 AP BIOLOGY 2008 SCORING GUIDELINES Question 1 1. The physical structure of a protein often reflects and affects its function. (a) Describe THREE types of chemical bonds/interactions found in proteins.
AP BIOLOGY 2008 SCORING GUIDELINES Question 1 1. The physical structure of a protein often reflects and affects its function. (a) Describe THREE types of chemical bonds/interactions found in proteins.
CHAPTER 6 AN INTRODUCTION TO METABOLISM. Section B: Enzymes
 CHAPTER 6 AN INTRODUCTION TO METABOLISM Section B: Enzymes 1. Enzymes speed up metabolic reactions by lowering energy barriers 2. Enzymes are substrate specific 3. The active site in an enzyme s catalytic
CHAPTER 6 AN INTRODUCTION TO METABOLISM Section B: Enzymes 1. Enzymes speed up metabolic reactions by lowering energy barriers 2. Enzymes are substrate specific 3. The active site in an enzyme s catalytic
The Organic Chemistry of Amino Acids, Peptides, and Proteins
 Essential rganic Chemistry Chapter 16 The rganic Chemistry of Amino Acids, Peptides, and Proteins Amino Acids a-amino carboxylic acids. The building blocks from which proteins are made. H 2 N C 2 H Note:
Essential rganic Chemistry Chapter 16 The rganic Chemistry of Amino Acids, Peptides, and Proteins Amino Acids a-amino carboxylic acids. The building blocks from which proteins are made. H 2 N C 2 H Note:
FTIR Analysis of Protein Structure
 FTIR Analysis of Protein Structure Warren Gallagher A. Introduction to protein structure The first structures of proteins at an atomic resolution were determined in the late 1950 s. 1 From that time to
FTIR Analysis of Protein Structure Warren Gallagher A. Introduction to protein structure The first structures of proteins at an atomic resolution were determined in the late 1950 s. 1 From that time to
Myoglobin and Hemoglobin
 Myoglobin and Hemoglobin Myoglobin and hemoglobin are hemeproteins whose physiological importance is principally related to their ability to bind molecular oxygen. Myoglobin (Mb) The oxygen storage protein
Myoglobin and Hemoglobin Myoglobin and hemoglobin are hemeproteins whose physiological importance is principally related to their ability to bind molecular oxygen. Myoglobin (Mb) The oxygen storage protein
Indiana's Academic Standards 2010 ICP Indiana's Academic Standards 2016 ICP. map) that describe the relationship acceleration, velocity and distance.
 .1.1 Measure the motion of objects to understand.1.1 Develop graphical, the relationships among distance, velocity and mathematical, and pictorial acceleration. Develop deeper understanding through representations
.1.1 Measure the motion of objects to understand.1.1 Develop graphical, the relationships among distance, velocity and mathematical, and pictorial acceleration. Develop deeper understanding through representations
Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Two Forms of Energy
 Module 2D - Energy and Metabolism Objective # 19 All living organisms require energy for survival. In this module we will examine some general principles about chemical reactions and energy usage within
Module 2D - Energy and Metabolism Objective # 19 All living organisms require energy for survival. In this module we will examine some general principles about chemical reactions and energy usage within
Chemical Basis of Life Module A Anchor 2
 Chemical Basis of Life Module A Anchor 2 Key Concepts: - Water is a polar molecule. Therefore, it is able to form multiple hydrogen bonds, which account for many of its special properties. - Water s polarity
Chemical Basis of Life Module A Anchor 2 Key Concepts: - Water is a polar molecule. Therefore, it is able to form multiple hydrogen bonds, which account for many of its special properties. - Water s polarity
Chapter 8: An Introduction to Metabolism
 Chapter 8: An Introduction to Metabolism Name Period Concept 8.1 An organism s metabolism transforms matter and energy, subject to the laws of thermodynamics 1. Define metabolism. The totality of an organism
Chapter 8: An Introduction to Metabolism Name Period Concept 8.1 An organism s metabolism transforms matter and energy, subject to the laws of thermodynamics 1. Define metabolism. The totality of an organism
Lecture 11 Enzymes: Kinetics
 Lecture 11 Enzymes: Kinetics Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 8, pp. 216-225 Key Concepts Kinetics is the study of reaction rates (velocities). Study of enzyme kinetics is useful for
Lecture 11 Enzymes: Kinetics Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 8, pp. 216-225 Key Concepts Kinetics is the study of reaction rates (velocities). Study of enzyme kinetics is useful for
--not necessarily a protein! (all proteins are polypeptides, but the converse is not true)
 00Note Set 5b 1 PEPTIDE BONDS AND POLYPEPTIDES OLIGOPEPTIDE: --chain containing only a few amino acids (see tetrapaptide, Fig 5.9) POLYPEPTIDE CHAINS: --many amino acids joined together --not necessarily
00Note Set 5b 1 PEPTIDE BONDS AND POLYPEPTIDES OLIGOPEPTIDE: --chain containing only a few amino acids (see tetrapaptide, Fig 5.9) POLYPEPTIDE CHAINS: --many amino acids joined together --not necessarily
Introduction, Noncovalent Bonds, and Properties of Water
 Lecture 1 Introduction, Noncovalent Bonds, and Properties of Water Reading: Berg, Tymoczko & Stryer: Chapter 1 problems in textbook: chapter 1, pp. 23-24, #1,2,3,6,7,8,9, 10,11; practice problems at end
Lecture 1 Introduction, Noncovalent Bonds, and Properties of Water Reading: Berg, Tymoczko & Stryer: Chapter 1 problems in textbook: chapter 1, pp. 23-24, #1,2,3,6,7,8,9, 10,11; practice problems at end
Lecture 15: Enzymes & Kinetics Mechanisms
 ROLE OF THE TRANSITION STATE Lecture 15: Enzymes & Kinetics Mechanisms Consider the reaction: H-O-H + Cl - H-O δ- H Cl δ- HO - + H-Cl Reactants Transition state Products Margaret A. Daugherty Fall 2004
ROLE OF THE TRANSITION STATE Lecture 15: Enzymes & Kinetics Mechanisms Consider the reaction: H-O-H + Cl - H-O δ- H Cl δ- HO - + H-Cl Reactants Transition state Products Margaret A. Daugherty Fall 2004
Helices From Readily in Biological Structures
 The α Helix and the β Sheet Are Common Folding Patterns Although the overall conformation each protein is unique, there are only two different folding patterns are present in all proteins, which are α
The α Helix and the β Sheet Are Common Folding Patterns Although the overall conformation each protein is unique, there are only two different folding patterns are present in all proteins, which are α
Previous lecture: Today:
 Previous lecture: The energy requiring step from substrate to transition state is an energy barrier called the free energy of activation G Transition state is the unstable (10-13 seconds) highest energy
Previous lecture: The energy requiring step from substrate to transition state is an energy barrier called the free energy of activation G Transition state is the unstable (10-13 seconds) highest energy
Modern Construction Materials Prof. Ravindra Gettu Department of Civil Engineering Indian Institute of Technology, Madras
 Modern Construction Materials Prof. Ravindra Gettu Department of Civil Engineering Indian Institute of Technology, Madras Module - 2 Lecture - 2 Part 2 of 2 Review of Atomic Bonding II We will continue
Modern Construction Materials Prof. Ravindra Gettu Department of Civil Engineering Indian Institute of Technology, Madras Module - 2 Lecture - 2 Part 2 of 2 Review of Atomic Bonding II We will continue
Proteins. Proteins. Amino Acids. Most diverse and most important molecule in. Functions: Functions (cont d)
 Proteins Proteins Most diverse and most important molecule in living i organisms Functions: 1. Structural (keratin in hair, collagen in ligaments) 2. Storage (casein in mother s milk) 3. Transport (HAEMOGLOBIN!)
Proteins Proteins Most diverse and most important molecule in living i organisms Functions: 1. Structural (keratin in hair, collagen in ligaments) 2. Storage (casein in mother s milk) 3. Transport (HAEMOGLOBIN!)
Role of Hydrogen Bonding on Protein Secondary Structure Introduction
 Role of Hydrogen Bonding on Protein Secondary Structure Introduction The function and chemical properties of proteins are determined by its three-dimensional structure. The final architecture of the protein
Role of Hydrogen Bonding on Protein Secondary Structure Introduction The function and chemical properties of proteins are determined by its three-dimensional structure. The final architecture of the protein
Molecular Cell Biology
 Harvey Lodish Arnold Berk Paul Matsudaira Chris A. Kaiser Monty Krieger Matthew P. Scott Lawrence Zipursky James Darnell Molecular Cell Biology Fifth Edition Chapter 2: Chemical Foundations Copyright 2004
Harvey Lodish Arnold Berk Paul Matsudaira Chris A. Kaiser Monty Krieger Matthew P. Scott Lawrence Zipursky James Darnell Molecular Cell Biology Fifth Edition Chapter 2: Chemical Foundations Copyright 2004
Amino Acids and Proteins
 Amino Acids and Proteins Proteins are composed of amino acids. There are 20 amino acids commonly found in proteins. All have: N2 C α R COO Amino acids at neutral p are dipolar ions (zwitterions) because
Amino Acids and Proteins Proteins are composed of amino acids. There are 20 amino acids commonly found in proteins. All have: N2 C α R COO Amino acids at neutral p are dipolar ions (zwitterions) because
MULTIPLE CHOICE QUESTIONS
 MULTIPLE CHOICE QUESTIONS 1. Most components of energy conversion systems evolved very early; thus, the most fundamental aspects of energy metabolism tend to be: A. quite different among a diverse group
MULTIPLE CHOICE QUESTIONS 1. Most components of energy conversion systems evolved very early; thus, the most fundamental aspects of energy metabolism tend to be: A. quite different among a diverse group
Biological Molecules
 Biological Molecules I won t lie. This is probably the most boring topic you have ever done in any science. It s pretty much as simple as this: learn the material deal with it. Enjoy don t say I didn t
Biological Molecules I won t lie. This is probably the most boring topic you have ever done in any science. It s pretty much as simple as this: learn the material deal with it. Enjoy don t say I didn t
Pipe Cleaner Proteins. Essential question: How does the structure of proteins relate to their function in the cell?
 Pipe Cleaner Proteins GPS: SB1 Students will analyze the nature of the relationships between structures and functions in living cells. Essential question: How does the structure of proteins relate to their
Pipe Cleaner Proteins GPS: SB1 Students will analyze the nature of the relationships between structures and functions in living cells. Essential question: How does the structure of proteins relate to their
Non-Covalent Bonds (Weak Bond)
 Non-Covalent Bonds (Weak Bond) Weak bonds are those forces of attraction that, in biological situations, do not take a large amount of energy to break. For example, hydrogen bonds are broken by energies
Non-Covalent Bonds (Weak Bond) Weak bonds are those forces of attraction that, in biological situations, do not take a large amount of energy to break. For example, hydrogen bonds are broken by energies
CHM333 LECTURE 13 14: 2/13 15/13 SPRING 2013 Professor Christine Hrycyna
 INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
Enzymes and Metabolic Pathways
 Enzymes and Metabolic Pathways Enzyme characteristics Made of protein Catalysts: reactions occur 1,000,000 times faster with enzymes Not part of reaction Not changed or affected by reaction Used over and
Enzymes and Metabolic Pathways Enzyme characteristics Made of protein Catalysts: reactions occur 1,000,000 times faster with enzymes Not part of reaction Not changed or affected by reaction Used over and
PROTEINS THE PEPTIDE BOND. The peptide bond, shown above enclosed in the blue curves, generates the basic structural unit for proteins.
 Ca 2+ The contents of this module were developed under grant award # P116B-001338 from the Fund for the Improvement of Postsecondary Education (FIPSE), United States Department of Education. However, those
Ca 2+ The contents of this module were developed under grant award # P116B-001338 from the Fund for the Improvement of Postsecondary Education (FIPSE), United States Department of Education. However, those
Figure 5. Energy of activation with and without an enzyme.
 Biology 20 Laboratory ENZYMES & CELLULAR RESPIRATION OBJECTIVE To be able to list the general characteristics of enzymes. To study the effects of enzymes on the rate of chemical reactions. To demonstrate
Biology 20 Laboratory ENZYMES & CELLULAR RESPIRATION OBJECTIVE To be able to list the general characteristics of enzymes. To study the effects of enzymes on the rate of chemical reactions. To demonstrate
Review of Chemical Equilibrium 7.51 September 1999. free [A] (µm)
![Review of Chemical Equilibrium 7.51 September 1999. free [A] (µm) Review of Chemical Equilibrium 7.51 September 1999. free [A] (µm)](/thumbs/40/21435474.jpg) Review of Chemical Equilibrium 7.51 September 1999 Equilibrium experiments study how the concentration of reaction products change as a function of reactant concentrations and/or reaction conditions. For
Review of Chemical Equilibrium 7.51 September 1999 Equilibrium experiments study how the concentration of reaction products change as a function of reactant concentrations and/or reaction conditions. For
Peptide Bond Amino acids are linked together by peptide bonds to form polypepetide chain.
 Peptide Bond Peptide Bond Amino acids are linked together by peptide bonds to form polypepetide chain. + H 2 O 2 Peptide bonds are strong and not broken by conditions that denature proteins, such as heating.
Peptide Bond Peptide Bond Amino acids are linked together by peptide bonds to form polypepetide chain. + H 2 O 2 Peptide bonds are strong and not broken by conditions that denature proteins, such as heating.
Lecture Overview. Hydrogen Bonds. Special Properties of Water Molecules. Universal Solvent. ph Scale Illustrated. special properties of water
 Lecture Overview special properties of water > water as a solvent > ph molecules of the cell > properties of carbon > carbohydrates > lipids > proteins > nucleic acids Hydrogen Bonds polarity of water
Lecture Overview special properties of water > water as a solvent > ph molecules of the cell > properties of carbon > carbohydrates > lipids > proteins > nucleic acids Hydrogen Bonds polarity of water
BIO 361 Biochemistry. Oficina: CABD Building 20 Room 133 First Floor Fall 2015 Email: [email protected] Thursday 16.00-17-00 Office Hours:
 Centro Universitario Internacional BIO 361 Biochemistry Carlos Santos Ocaña Course Information: Oficina: CABD Building 20 Room 133 First Floor Fall 2015 Email: [email protected] Thursday 16.00-17-00 Office
Centro Universitario Internacional BIO 361 Biochemistry Carlos Santos Ocaña Course Information: Oficina: CABD Building 20 Room 133 First Floor Fall 2015 Email: [email protected] Thursday 16.00-17-00 Office
4. Which carbohydrate would you find as part of a molecule of RNA? a. Galactose b. Deoxyribose c. Ribose d. Glucose
 1. How is a polymer formed from multiple monomers? a. From the growth of the chain of carbon atoms b. By the removal of an OH group and a hydrogen atom c. By the addition of an OH group and a hydrogen
1. How is a polymer formed from multiple monomers? a. From the growth of the chain of carbon atoms b. By the removal of an OH group and a hydrogen atom c. By the addition of an OH group and a hydrogen
NMR SPECTROSCOPY. Basic Principles, Concepts, and Applications in Chemistry. Harald Günther University of Siegen, Siegen, Germany.
 NMR SPECTROSCOPY Basic Principles, Concepts, and Applications in Chemistry Harald Günther University of Siegen, Siegen, Germany Second Edition Translated by Harald Günther JOHN WILEY & SONS Chichester
NMR SPECTROSCOPY Basic Principles, Concepts, and Applications in Chemistry Harald Günther University of Siegen, Siegen, Germany Second Edition Translated by Harald Günther JOHN WILEY & SONS Chichester
Part A: Amino Acids and Peptides (Is the peptide IAG the same as the peptide GAI?)
 ChemActivity 46 Amino Acids, Polypeptides and Proteins 1 ChemActivity 46 Part A: Amino Acids and Peptides (Is the peptide IAG the same as the peptide GAI?) Model 1: The 20 Amino Acids at Biological p See
ChemActivity 46 Amino Acids, Polypeptides and Proteins 1 ChemActivity 46 Part A: Amino Acids and Peptides (Is the peptide IAG the same as the peptide GAI?) Model 1: The 20 Amino Acids at Biological p See
Protein Melting Curves
 Protein Melting Curves In previous classes we talked a lot about what aspects of a protein structure stabilize a protein and what aspects destabilize it. But how would one actually test such predictions?
Protein Melting Curves In previous classes we talked a lot about what aspects of a protein structure stabilize a protein and what aspects destabilize it. But how would one actually test such predictions?
Introduction to Protein Folding
 Introduction to Protein Folding Chapter 4 Proteins: Three Dimensional Structure and Function Conformation - three dimensional shape Native conformation - each protein folds into a single stable shape (physiological
Introduction to Protein Folding Chapter 4 Proteins: Three Dimensional Structure and Function Conformation - three dimensional shape Native conformation - each protein folds into a single stable shape (physiological
CNAS ASSESSMENT COMMITTEE CHEMISTRY (CH) DEGREE PROGRAM CURRICULAR MAPPINGS AND COURSE EXPECTED STUDENT LEARNING OUTCOMES (SLOs)
 CNAS ASSESSMENT COMMITTEE CHEMISTRY (CH) DEGREE PROGRAM CURRICULAR MAPPINGS AND COURSE EXPECTED STUDENT LEARNING OUTCOMES (SLOs) DEGREE PROGRAM CURRICULAR MAPPING DEFINED PROGRAM SLOs Course No. 11 12
CNAS ASSESSMENT COMMITTEE CHEMISTRY (CH) DEGREE PROGRAM CURRICULAR MAPPINGS AND COURSE EXPECTED STUDENT LEARNING OUTCOMES (SLOs) DEGREE PROGRAM CURRICULAR MAPPING DEFINED PROGRAM SLOs Course No. 11 12
H H N - C - C 2 R. Three possible forms (not counting R group) depending on ph
 Amino acids - 0 common amino acids there are others found naturally but much less frequently - Common structure for amino acid - C, -N, and functional groups all attached to the alpha carbon N - C - C
Amino acids - 0 common amino acids there are others found naturally but much less frequently - Common structure for amino acid - C, -N, and functional groups all attached to the alpha carbon N - C - C
Methods for Protein Analysis
 Methods for Protein Analysis 1. Protein Separation Methods The following is a quick review of some common methods used for protein separation: SDS-PAGE (SDS-polyacrylamide gel electrophoresis) separates
Methods for Protein Analysis 1. Protein Separation Methods The following is a quick review of some common methods used for protein separation: SDS-PAGE (SDS-polyacrylamide gel electrophoresis) separates
Steffen Lindert, René Staritzbichler, Nils Wötzel, Mert Karakaş, Phoebe L. Stewart, and Jens Meiler
 Structure 17 Supplemental Data EM-Fold: De Novo Folding of α-helical Proteins Guided by Intermediate-Resolution Electron Microscopy Density Maps Steffen Lindert, René Staritzbichler, Nils Wötzel, Mert
Structure 17 Supplemental Data EM-Fold: De Novo Folding of α-helical Proteins Guided by Intermediate-Resolution Electron Microscopy Density Maps Steffen Lindert, René Staritzbichler, Nils Wötzel, Mert
2012 HORIBA Scientific. All rights reserved. 2012 HORIBA Scientific. All rights reserved.
 Raman Spectroscopy for proteins Catalina DAVID Ph.D. application scientist Outline Raman spectroscopy in few words What is Raman spectroscopy? What is the information we can get? Basics of Raman analysis
Raman Spectroscopy for proteins Catalina DAVID Ph.D. application scientist Outline Raman spectroscopy in few words What is Raman spectroscopy? What is the information we can get? Basics of Raman analysis
Translation Study Guide
 Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
BIOLOGICAL MEMBRANES: FUNCTIONS, STRUCTURES & TRANSPORT
 BIOLOGICAL MEMBRANES: FUNCTIONS, STRUCTURES & TRANSPORT UNIVERSITY OF PNG SCHOOL OF MEDICINE AND HEALTH SCIENCES DISCIPLINE OF BIOCHEMISTRY AND MOLECULAR BIOLOGY BMLS II / B Pharm II / BDS II VJ Temple
BIOLOGICAL MEMBRANES: FUNCTIONS, STRUCTURES & TRANSPORT UNIVERSITY OF PNG SCHOOL OF MEDICINE AND HEALTH SCIENCES DISCIPLINE OF BIOCHEMISTRY AND MOLECULAR BIOLOGY BMLS II / B Pharm II / BDS II VJ Temple
Study the following diagrams of the States of Matter. Label the names of the Changes of State between the different states.
 Describe the strength of attractive forces between particles. Describe the amount of space between particles. Can the particles in this state be compressed? Do the particles in this state have a definite
Describe the strength of attractive forces between particles. Describe the amount of space between particles. Can the particles in this state be compressed? Do the particles in this state have a definite
Structure Determination
 5 Structure Determination Most of the protein structures described and discussed in this book have been determined either by X-ray crystallography or by nuclear magnetic resonance (NMR) spectroscopy. Although
5 Structure Determination Most of the protein structures described and discussed in this book have been determined either by X-ray crystallography or by nuclear magnetic resonance (NMR) spectroscopy. Although
Introduction to Proteins and Enzymes
 Introduction to Proteins and Enzymes Basics of protein structure and composition The life of a protein Enzymes Theory of enzyme function Not all enzymes are proteins / not all proteins are enzymes Enzyme
Introduction to Proteins and Enzymes Basics of protein structure and composition The life of a protein Enzymes Theory of enzyme function Not all enzymes are proteins / not all proteins are enzymes Enzyme
Polymers: Introduction
 Chapter Outline: Polymer Structures Hydrocarbon and Polymer Molecules Chemistry of Polymer Molecules Molecular Weight and Shape Molecular Structure and Configurations Copolymers Polymer Crystals Optional
Chapter Outline: Polymer Structures Hydrocarbon and Polymer Molecules Chemistry of Polymer Molecules Molecular Weight and Shape Molecular Structure and Configurations Copolymers Polymer Crystals Optional
Chemistry. CHEMISTRY SYLLABUS, ASSESSMENT and UNIT PLANNERS GENERAL AIMS. Students should be able to
 i CHEMISTRY SYLLABUS, ASSESSMENT and UNIT PLANNERS GENERAL AIMS Students should be able to - apply and use knowledge and methods that are typical to chemistry - develop experimental and investigative skills,
i CHEMISTRY SYLLABUS, ASSESSMENT and UNIT PLANNERS GENERAL AIMS Students should be able to - apply and use knowledge and methods that are typical to chemistry - develop experimental and investigative skills,
博 士 論 文 ( 要 約 ) A study on enzymatic synthesis of. stable cyclized peptides which. inhibit protein-protein interactions
 博 士 論 文 ( 要 約 ) 論 文 題 目 A study on enzymatic synthesis of stable cyclized peptides which inhibit protein-protein interactions ( 蛋 白 質 間 相 互 作 用 を 阻 害 する 安 定 な 環 状 化 ペプチドの 酵 素 合 成 に 関 する 研 究 ) 氏 名 張 静 1
博 士 論 文 ( 要 約 ) 論 文 題 目 A study on enzymatic synthesis of stable cyclized peptides which inhibit protein-protein interactions ( 蛋 白 質 間 相 互 作 用 を 阻 害 する 安 定 な 環 状 化 ペプチドの 酵 素 合 成 に 関 する 研 究 ) 氏 名 張 静 1
CHM333 LECTURE 13 14: 2/13 15/12 SPRING 2012 Professor Christine Hrycyna
 INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
DO PHYSICS ONLINE FROM QUANTA TO QUARKS QUANTUM (WAVE) MECHANICS
 DO PHYSICS ONLINE FROM QUANTA TO QUARKS QUANTUM (WAVE) MECHANICS Quantum Mechanics or wave mechanics is the best mathematical theory used today to describe and predict the behaviour of particles and waves.
DO PHYSICS ONLINE FROM QUANTA TO QUARKS QUANTUM (WAVE) MECHANICS Quantum Mechanics or wave mechanics is the best mathematical theory used today to describe and predict the behaviour of particles and waves.
How To Understand Enzyme Kinetics
 Chapter 12 - Reaction Kinetics In the last chapter we looked at enzyme mechanisms. In this chapter we ll see how enzyme kinetics, i.e., the study of enzyme reaction rates, can be useful in learning more
Chapter 12 - Reaction Kinetics In the last chapter we looked at enzyme mechanisms. In this chapter we ll see how enzyme kinetics, i.e., the study of enzyme reaction rates, can be useful in learning more
