TOWARDS UNDERSTANDING THE EPIGENETIC CONTRIBUTION TO NEURAL DEVELOPMENT AND NEURAL TUBE DEFECTS IN THE MOUSE MODEL SYSTEM LAURA HARMACEK
|
|
|
- Elijah Dean
- 9 years ago
- Views:
Transcription
1 TOWARDS UNDERSTANDING THE EPIGENETIC CONTRIBUTION TO NEURAL DEVELOPMENT AND NEURAL TUBE DEFECTS IN THE MOUSE MODEL SYSTEM by LAURA HARMACEK B.A, Smith College, 2006 A thesis submitted to the Faculty of the Graduate School of the University of Colorado in partial fulfillment of the requirements for the degree of Doctor of Philosophy Molecular Biology Program 2014
2 This thesis for the Doctor of Philosophy degree by Laura Diane Harmacek has been approved for the Molecular Biology Program by Trevor Williams, Chair Lee Niswander, Advisor Heide Ford Jay Hesselberth Pepper Schedin Date 01/03/2014 ii
3 Harmacek, Laura, Diane (Ph.D., Molecular Biology) Towards Understanding the Epigenetic Contribution to Neural Development and Neural Tube Defects in the Mouse Model System Thesis directed by Professor Lee Niswander ABSTRACT Failure of embryonic neural tube closure results in the second most common class of birth defects known as neural tube defects (NTDs). While NTDs are likely the result of complex multigenic dysfunction, it is not known whether polymorphisms in epigenetic regulators may be risk factors for NTDs in humans. Here we used a mouse model of NTDs caused by mutation of an epigenetic regulator to determine the mechanistic contribution of this key factor to neural tube closure. We characterized Baf155 msp3, a unique ENU-induced allele in mice. Homozygous Baf155 msp3 embryos exhibit highly penetrant cranial neural tube defects called exencephaly, allowing us to investigate the roles of an assembled, but malfunctional BAF chromatin remodeling complex in vivo at the time of neural tube closure. Evidence of defects in proliferation and apoptosis were found within the neural tube. RNA-Seq analysis revealed relatively few genes that showed altered expression in Baf155 msp3 mutant neural tissue, which was surprising given the broad epigenetic role of the BAF complex. Dysregulated genes included those involved in neural development and cell survival. Another unexpected finding was that, even though the NTD was consistently observed, the gene expression changes between individual mutants were variable. This suggests that inconsistent gene regulation contributes to failed neural tube closure. These results shed light on the role of the BAF complex in the process of neural tube closure and iii
4 highlight the importance of studying missense alleles to understand epigenetic regulation during critical phases of development. The form and content of this abstract are approved. I recommend its publication. Approved: Lee A. Niswander iv
5 I dedicate this work to my husband, Benjamin Warren, my family (Marilyn, Joe and Grant) and my friends who helped me throughout my time in graduate school.. v
6 ACKNOWLEDGMENTS I would like to thank all the members of the Niswander Lab. I thank Lee Niswander for being eternally patient, flexible and a truly fantastic mentor. I thank Jianfu (Jeff) Chen for training me and helping me get started in the lab. I thank Jonathan Wilde for insightful discussions and help with techniques in the lab. I thank Claire Tsai for helping me throughout school and being a true friend. I thank Martin Gartz Hanson for bringing the broader neuroscience perspective to my research. Thanks to Heather Ray for mentoring everyone in the lab, including me. Also, Heather always made the lab an enjoyable place to be. A special thanks to Juliette Petersen for always being there for me, helping me with protocols, and keeping the lab running. Last, I thank Lori Bulwith without whose assistance in the moues room and friendship I could not have been as successful in graduate school. vi
7 TABLE OF CONTENTS CHAPTER I. INTRODUCTION... 1 FORMATION OF THE NEURAL TUBE AND DEFECTS THAT OCCUR DUE TO FAILED NEURAL TUBE CLOSURE... 1 Neural Tube Defects in Humans... 1 The Mouse as a Model System... 3 Overview of Neurulation... 4 Cellular and Molecular Mechanisms of Neurulation... 5 Anterior/Posterior Patterning Dorsal/Ventral Patterning Apoptosis Proliferation and Differentiation NEURAL TUBE DEFECTS IN CHROMATIN MAINTENANCE MUTANTS Chromatin Structure Epigenetics DNA Methylation Histone Modification Chromatin Remodeling II. THE MSP3 MUTANT MOUSE EXHIBITS EXENCEPHALY AND OTHER PHENOTYPIC DEFECTS INTRODUCTION RESULTS Baf155msp3 MUTANTS EXHIBIT CRANIAL NEURAL TUBE DEFECTS CONCLUSION MATERIALS AND METHODS vii
8 Mouse Strains Genotyping III. MOLECUALR AND CELLULAR CHARACTERIZATION OF THE Baf155 msp3 MUTANT SHOWS A DECREASE IN CELL PROLIFERATION AND CELL SURVIVAL INTRODUCTION RESULTS Mapping the ENU-Induced Mutation to Baf BAF155 msp3 Mutant Protein Can Still Associate with the Core Remodeling Complex Cell Proliferation Is Decreased and Cell Death Is Increased in Baf155 msp3/msp3 Neural Progenitor Cells CONCLUSION MATERIALS AND METHODS Analysis of Mutant Phenotype Antibodies Used Yeast Two-Hybrid Assay Protocol Immunoprecipitation Assay and Western Blots IV. GENE EXPRESSION ANALYSIS OF Baf155 msp3/ msp3 EMBRYOS SHOWS GENE EXPRESSION VARIABILITY AND OVERALL UPREGULATION OF GENES INTRODUCTION RESULTS Gene Expression Levels Are More Variable in Baf155 msp3 Mutant Cranial Tissue: Averaged Sample Analysis Gene Expression Levels Are More Variable in Baf155 msp3 Mutant Cranial Tissue: Individual Sample Analysis CONCLUSION MATERIALS AND METHODS viii
9 RNA-Seq Library Preparation and Analysis Data Analysis and Interpretation V. DISCUSSION REFERENCES APPENDIX A. FOLATE SUPPLEMENTATION, TESTS FOR POSSIBLE GENETIC INTERACTIONS, AND ADDITIONAL PHENOTYPIC ANALYSES SHOW MINIMAL EFFECTS IN Baf155 msp3 MICE INTRODUCTION RESULTS Baf155 msp3 Mice Do Not Respond to Folic Acid Supplementation Test for Genetic Interaction between Baf155 msp3 and Other NTD Mutant Lines Additional Phenotype Characterization of the Baf155 msp3/msp3 embryos CONCLUSION MATERIALS AND METHODS High Folic Acid Diet Protocol Genetic Interactions Protocols Neurofilament Staining of Whole Mount Embryos Bone and Cartilage Staining (Alcian Blue/Alizarin Red Staining) B. GCN5 ACETYLTRANSFERASE ACTIVITY FUNCTIONS IN NEURAL CREST CELL BIOLOGY INTRODUCTION RESULTS Gcn5 hat/hat Mutant NCCs Migrate Farther than WT ix
10 Sox10 Expression Is Disrupted in the Gcn5 hat/hat Mutant Craniofacial Structures Are Mislocalized and Malformed in the Gcn5 hat/hat Mutant CONCLUSION MATERIALS AND METHODS Neural Tube Explants Sox10 Staining Bone and Cartilage Staining (Alcian Blue/Alizarin Red Staining) C. TABLE OF MISREGULATED GENES FROM RNA ISOLATED FROM E9.5 CRANIAL NEURAL TISSUE OF BAF155 MSP3/MSP3 EMBRYOS D. TABLE OF THE OVERLAP OF MISREGULATED GENES IN CRANIAL NEURAL TISSUE E. LAURA HARMACEK S JOURNAL ARTICLE FROM JOURNAL OF DEVELOPMENTAL NEUROBIOLOGY, WILEY ONLINE LIBRARY, 2013: x
11 LIST OF TABLES Table II.1 Phenotype and genotype of embryos resulting from cross between heterozygous Baf155 +/msp3 mice III.1 Functional list of potential variants of the Msp3 phenotype A.1 Phenotype and genotype of embryos resulting from cross between heterozygous Baf155 +/msp3 mice after 9.5 days on a high FA diet A.2 Phenotype and genotype of embryos resulting from crosses between heterozygous Baf155 +/msp3 female mice and other heterozygous male mice C.1 List of misregulated genes in cranial tissue from averaged WT and averaged Mutant E9.5 embryos based on EdgeR analysis D.1 Misexpressed genes in individual Baf155 msp3/msp3 mutants xi
12 LIST OF FIGURES Figure I.1 Schematic for mouse neural tube closure sites and where defects can arise I.2 Overview of neurulation I.3 Anterior aspect of the neural plate I.4 Patterning in the neural tube is set up by Sonic Hedgehog (Shh) and Bone Morphogenetic Protein (BMP) I.5 Molecular mechanisms of nucleosome remodeling in vitro I.6 Schematic of the BAF chromatin remodeling complex II.1 Baf155 msp3/msp3 embryos show neural tube defects III.1 BAF155 msp3 associates with other core BAF complex proteins III.2 BAF155 is disordered in the region containing the Msp3 mutation III.3 BAF155 msp3 interacts with BAF60a, BAF60b and BAF III.4 BAF155 function is not necessary for early neural patterning III.5 BAF155 function is necessary to maintain proliferation and cell survival IV.1 Variable gene expression in Baf155 msp3/msp3 mutant cranial tissue A.1 Mice show a range of phenotypes on both regular and high folic acid diet A.2 Baf155 function is not necessary for skeletal patterning A.3 Baf155 function is not necessary for skeletal patterning or neuron outgrowth B.1 Gcn5 hat/hat mutants show NCC migration and specification defects B.2 Skeletal stainings of E16.5 Gcn5 hat/hat mutants xii
13 CHAPTER I INTRODUCTION FORMATION OF THE NEURAL TUBE AND DEFECTS THAT OCCUR DUE TO FAILED NEURAL TUBE CLOSURE Neural Tube Defects in Humans Neural Tube Defects (NTDs) are the second most common birth defect in humans, ranging in incidence from 0.5 to 2 in 1000 live births worldwide, with some variation in the incidence depending on the country and region studied [1]. NTDs result when the precursor of the brain and spinal cord, the neural tube, fails to form properly during development. NTDs include caudal neural tube closure defects such as spina bifida, and cranial neural tube defects such as exencephaly. The most severe form of NTD is craniorachischisis, failure of the neural tube to close along the cranial to caudal regions (Figure I.1). Figure I.1 Schematic for mouse neural tube closure sites and where defects can arise. Neural tube defects can arise when the neural tube does not close properly. These include exencephaly, craniorachischisis and spina bifida. There are three closure points in mice; closure point 3 (CP3) is most anterior, CP2 can vary depending on the strain, and CP1 occurs at the hindbrain/cervical spine border. Image reproduced from [2]. 1
14 Fetuses that develop an open cranial neural tube are stillborn or die shortly after birth, while those with spina bifida have a wide range of health outcomes. For example, 78% of infants with spina bifida survive into adulthood [3] but they require on average five surgeries in the first year of life and experience diminished quality of life. Secondary consequences of spina bifida include reduced bladder control, lower-body malfunction, paraplegia, cognitive disabilities, and hydrocephaly [4]. A patient with spina bifida is estimated to incur lifetime health care costs totaling $1.4 million (adjusted to 2001 dollars) while only reaching an average of 20 years of age [5]. Living with spina bifida is an emotional burden to the individuals affected as well as their families, and this strain cannot be measured monetarily. Hence, NTDs have a long-term impact on quality of life for the patients and their families. Human population studies indicate there is a genetic component to NTD incidence. NTDs are associated with several genetic syndromes including Meckel Syndrome and chromosomal rearrangements such as trisomy 13 and trisomy 18. The recurrence risk of NTDs for siblings of affected individuals is approximately 50-times greater than that of the general population, with a further increase in risk after two or more affected pregnancies [6]. Also, several twin studies have shown that both identical twins are % more likely to have spina bifida than both fraternal twins [7]. Furthermore, epidemiological studies estimate that genetic factors may account for up to 70% of NTD prevalence [8]. Counter to these data, relatively few studies report multi-generational occurrences, indicating the genetic contribution does not follow straightforward Mendelian inheritance. Thus, NTDs have been described as being multi-factorial or oligogenic rather than a model of dominant or recessive inheritance with partial penetrance [5, 9]. 2
15 Targeted genetic screens of individuals with spina bifida and their families provide further evidence for a genetic component to NTDs. Genome-wide studies of patients with spina bifida and their extended families indicate chromosomes 2, 7, and 10 might contain risk loci contributing to the deformities [10, 11]. Despite the ample evidence for genetic factors, little is known about specific genes that play a role in human neural tube closure defects. There is evidence for involvement of a few genes, including Pax1, Vangl1, Disheveled 2 and Disheveled 3 in human NTDs [12-15]. In one compelling study, a Vangl1 (VNGL1) genomic variant was identified in a patient population with spina bifida. In vitro studies of the same variant showed evidence that the mutation impairs protein binding functions of VANGL1, suggesting that the variant might be a risk allele for NTDs [14]. Despite that fact that over 130 epidemiological studies have been conducted to identify risk factors for NTDs (including whole-genome analysis, single gene analyses in large NTD populations, and family-based association studies), the VNGL1 example is one of only a few epidemiological studies that has pinpointed a single gene variant as a potential risk factor for NTDs [16]. The single gene approach to identify genetic risk factors has thus far been only modestly successful in the case of human NTDs. This may be due to the fact that human NTDs are suggested to be multigenic disorders with a complex genetic etiology. A better understanding of the genetic causes of NTDs is crucial for designing preventive strategies and better counseling for couples at risk. The Mouse as a Model System Due to ethical concerns, many of the molecular and developmental studies that are necessary to uncover the causes of NTDs cannot be conducted in humans. The mouse is a useful model system to understand the embryological mechanisms of neural tube closure 3
16 and, when it fails, NTDs. First, the mouse and human undergo a similar process of neurulation, which includes closure of the neural tube. Second, NTDs in mice are morphologically similar to NTDs in humans. Third, over 200 mutations in the mouse have been identified that result in NTDs [17-19], indicating the multigenic nature of NTDs, and these will be discussed below. Fourth, similar to humans, the mouse develops in utero. Last, gene conservation is extremely high between humans and mice [20]. The genomes are each estimated to contain approximately 30,000 protein-coding genes. Moreover, 99% of genes in the mouse genome have a human homologue, and vice versa [20]. Therefore, the mouse is an effective model for studying NTDs. Overview of Neurulation Neurulation is the embryonic process that gives rise to the central nervous system in the adult. The process begins early in development during the gastrulation phase, at E6.0 in the mouse embryo, and is completed by E10.5. Neurulation occurs in several overlapping steps. There are two types of neurulation: Primary neurulation and secondary neurulation [21]. Secondary neurulation occurs in embryos just below the tail bud, and proliferation of stem cells is the mechanism primarily responsible for creating this tissue. After proliferation, a cluster of mesenchymal cells just under the ectoderm form a lumen in an initially solid core in a process known as cavitation. Secondary neurulation occurs at the posterior neural pore and does not appear to contribute to open NTDs. This thesis will focus on primary neurulation because it occurs in the cranial region where exencephaly results, so secondary neurulation will not be discussed here; however both primary and secondary neurulation occur in all amniotes, including humans [22]. 4
17 Initially during primary neurulation, cells in the dorsal ectoderm thicken and are specified to become the neural plate by signals from the mesoderm below. The flat neural plate is found on the dorsal aspect of the embryo, along the anterior/posterior (A/P) axis. Cells along the midline of the neural plate just above the notochord, an important structure that releases signaling factors essential during neurulation, begin to constrict along their apical aspects, causing the neural plate to bend, forming the neural folds. The constriction of the cells forms hinge points and the neural folds begin to rise. These neural folds continue to lift and bend closer together. Finally, the neural folds appose each other to form an elongated rounded structure on the dorsal side of the embryo. Neurulation is complete when the neural folds fuse to form the neural tube (Figure I.2). Coordination of physical, cellular, and molecular events is required for the neural tube to correctly form. Cellular and Molecular Mechanisms of Neurulation The first stage of neurulation is neural induction, or specification of the ectodermal cells that will give rise to the neural ectoderm. Initially, early ectodermal cells have two potential fates, epidermis (skin) or neural ectoderm (nervous system). In classic studies of mouse neural induction, the molecular basis of neurulation was shown to be signaling from the node, an important organizer for neural cell fate. For example, explant studies of stage matched embryos showed the node, which gives rise to the notochord, is sufficient to induce a secondary site of neural induction in non-neural cells [23]. This suggests that inductive signals are secreted by the node to determine neural cell fate. Classic studies were also conducted in Xenopus to elucidate the mechanisms underlying neural induction. 5
18 Figure I.2 Overview of neurulation. The neural plate begins as a flat sheet of cells. A) The cells in the neural palate thicken. B) These cells receive signals from the notochord to begin bending at the medial hinge point just above the notochord. Cells also bend at the dorsolateral hinge points. C) The neural folds appose each other after lifting and bending. D) The folds fuse into two monolayers of cells; the non-neural ectoderm and the neural ectoderm. Image adapted from [24]. The dorsal cap, also known as the animal cap, is a region on the developing Xenopus embryo at the blastula stage that gives rise to the dorsal anterior structures, or ectoderm and neural tissues. These tissues have an intrinsic neural cell fate. Bone morphogenetic proteins (BMPs) were shown to block this intrinsic neural cell fate and instead generate epidermal fate in the dorsal cap of Xenopus, indicating BMP antagonizes neural character and promotes the formation of epidermis [25]. In Xenopus, the inductive signals for epidermis were shown to be BMP-4 [26] and inductive signals of neural fate were shown to be antagonists of BMP-4, which are secreted by the node and the paraxial mesoderm [27]. In summary, the node organizer and paraxial mesoderm secretes antagonists of BMP signaling and thereby serves to induce neural character of the cells of the neural ectoderm, which form neural plate cells. 6
19 After neural induction, the cells of the neural plate begin to change their shape, and rearrange their positions, in order to change shape from a flat sheet of cells to a 3- dimensional tube of cells. Cells in the neural plate extend along their medial-lateral axes, making them wider than they are long. They simultaneously converge towards the midline of the embryo/neural plate. These movements cause the embryo to elongate along the rostral-caudal axis and extend the neural plate. This process is called convergent extension. An important genetic and molecular pathway that controls convergent extension movements is the Planar Cell Polarity (PCP) pathway. PCP is a signal transduction pathway involving the extracellular ligands Wnts, the cell-surface receptors Frizzleds, and the cytoplasmic factors Disheveleds (Dvl). In Xenopus, Disheveled-dependent convergent extension is required for neural tube formation [28]. In mice, disruption of several genes in the PCP pathway, including Disheveled 1, Disheveled 2, and Vangl2, results in NTDs, suggesting that the PCP pathway is essential for NT formation. Importantly, live imaging studies of Looptail (Lp; Vangl2 mutation) mutant mouse embryos show a reduction in convergent extension of the A/P axis [29]. This study directly connects convergent extension defects with PCP mouse mutants, and therefore closes a longtime gap in knowledge within the field. While the cellular process of convergent extension and the PCP signaling pathway are important for extending the embryo along the rostral-caudal axis, different molecular and cellular processes control neurulation along the dorsal-ventral axis. While the neural plate is elongating through convergent extension movements, the notochord (Figure I.2) secretes a still unidentified signal which acts on the neural ectoderm and induces formation of the medial hinge point (also called the floor plate) just 7
20 above the notochord. Following this, the dorsolateral hinge points are formed approximately midway along dorsal neural folds. Cellular reorganization of the cytoskeleton called apical constriction occurs and, as a consequence, the hinge points form and the neural folds elevate. Briefly, the actomyosin filaments on the apical side of the cells contract so the cells become wedge-shaped instead of columnar. The constriction of the cytoskeleton along the apical aspect of cells in the neural folds increases the apical curvature and brings the neural folds together. The molecular basis for this constriction is not well understood, however SHROOM3, which localizes to the cytoskeleton and binds F-actin, is required for apical constriction in the neural tube in both Xenopus [30] and mouse [31]. Live imaging studies during neural tube closure of the Shroom3 m1nisw/m1nisw mutant mouse found that the rate and dynamics of neural fold bending (or inflection) was slower in mutant compared to Wild Type (WT) [32], supporting previous findings that Shroom3 plays a role in neural tube formation and apical constriction. Bending and constriction of the neural folds brings them close to one another along the dorsal midline, positioning the neural tissue for the final step of fusion. The final step of neural tube formation occurs as the apposing neural folds meet and undergo a cellular fusion event to form two complete tissue layers. The overlying nonneural ectoderm separates from the neural ectoderm and fuses together to form a monolayer of non-neural ectoderm, which overlies the closed neural ectoderm tube (Figure I.2). The molecular basis of fusion is not well established; however, a class of receptor tyrosine kinase (RTK) signaling molecules known as Ephrins may play a role. Mice with mutations in Efna5, encoding the Ephrin A5 ligand, and EphA7, encoding the Ephrin Receptor A7, experience an open cranial neural tube [33]. Usually, the Ephrin 8
21 receptor and Ephrin ligand, both of which are attached to the cell surface, causes cells to be repulsed away from each other through inducing bidirectional signaling cascades. This study highlights that Ephrin signaling is required for neural tube fusion via a mechanism where they can carry out an adhesive rather than a repressive role through a splice variant of EphA7 which suppresses tyrosine phosphorylation of the full-length EphA7 in vitro. Other genes required for neural fold fusion are the Grainy-head like (Grhl) family of transcription factors. Grhl2 and Grhl3 are localized to the neural folds and control the transcription of many genes known to encode cell adhesion proteins [34]. Deletion or mutations in the Grhl genes in mice result in neural tube defects [34-36]. There still remains a great deal of work to be done before we fully understand the genetic pathways and cellular mechanisms that are required for neural tube fusion. The fusion of the neural tube occurs at distinct times along the rostral-caudal axis. In the mouse there are three initial fusion points, known as closure points. Closure point 1 (Figure I.1) initiates early at E8.5 when an embryo has 6-7 pairs of somites. Fusion begins at the hindbrain-cervical boundary and continues both rostrally towards the head and caudally towards the tail. Closure point 2 occurs in the midbrain-forebrain region and precedes both rostrally and caudally. Closure point 3 occurs near the anterior neural boundary located at the most rostral extremity of the embryo and proceeds caudally. Closure points 2 and 3 occur simultaneously as the process of neurulation continues. Through the bidirectional fusion of ectoderm tissue between the closure points, the entire neural tube fuses along the rostral caudal axis to complete neural tube closure [37]. The exact location and occurrence of closure point 2 is variable in the mouse and has been observed at different points in different genetic backgrounds [38]. It is most 9
22 commonly found between the midbrain-forebrain boundary, however in some backgrounds it has been observed initiating from rostrally within the midbrain, caudally within the forebrain, or in some cases, there are only 2 fusion points, and closure point 2 is not observed. Based on live imaging studies, the initiation of closure point 2 appears to be variable on a mixed genetic background of C57BL/6J and 129/SvImJ studied [32]. This variance in presence and location of closure point 2 both between mouse strains and within a mixed background mouse strain might explain variation in exencephaly penetrance observed in different mutant mouse strains. Anterior/Posterior Patterning At E8.0, the elongated embryo has a clear morphology and the embryonic structure of the neural tissue is easily observed (Figure I.3). From anterior to posterior, the embryonic neural tissue is divided into the following regions: the prosencephalon (Forebrain), mesencephalon (Midbrain) and rhombencephalon (Hindbrain), and the spinal cord. Anterior/Posterior (A/P) patterning specifies each of these structural regions in the developing brain with a combination of transcription factors (OTX, OTX2 and GBX2) and secreted ligands (FGF8 and WNT). Each structural region is induced to express a unique set of genes which specifies the fate of that functional region. Even as early as E8.5 in the developing mouse embryo, A/P patterning genes are expressed in distinct regions, indicating patterning information is already being imparted on the neural tissues before the neural tube has completely closed. At this stage, Otx is expressed in the forebrain, Wnt/Otx2 is expressed in the midbrain and Ffg8/Gbx2 is expressed in the anterior hindbrain [39]. Developmental organizers induce A/P patterning in the forebrain and the mid/hind brain by releasing diffusible signaling factors. There are different developmental 10
23 organizers along the A/P axis of the developing embryo that contribute to overall A/P patterning. Figure I.3 Anterior aspect of the neural plate. Early in development at E8.0, the developing mouse embryonic neural tissue can be divided into 4 distinct regions- the prosencephalon, the mesencephalon, the rhombencephalon and the spinal cord. Image adapted from [40]. Forebrain specification is established by a combination of transcription factors and secreted molecules, some from the Anterior Neural Boundary (ANB). The ANB is the organizer in the most anterior pole of the neuroaxis, and establishes forebrain patterning by repression of WNT signaling. There are several lines of evidence that suggest repression of WNT signaling is essential for forebrain patterning. First, the transcription factor SIX3, which has been implicated in repression of WNT signaling, is required for forebrain development. For example, Six3 -/- mice have a severely truncated prosencephalon [41], indicating repression of WNT signaling by SIX3 is necessary for forebrain development. Next, other WNT antagonists secreted from the ANB can locally induce Fgf8 expression, which is also important for forebrain specification [42]. The organizer is not the only 11
24 origin of signals that are important for forebrain patterning. SHH is not secreted from the ANB organizer, but is an important molecular signal in rostral development. Shh -/- embryos have multiple defects, including abnormal forebrain morphology [43]. The Shh -/- embryos also are unable to establish and maintain structures important in neurulation including the notochord and floor plate. In summary, forebrain specification results from modulation of SHH signaling, as well as WNT signaling cascades controlled by the ANB organizer. Midbrain and hindbrain (MHB) patterning also depend on an important organizing center, the isthmus organizer, located between the mid and hindbrain. Cells anterior to the isthmus are specified as midbrain while cells posterior to the isthmus are specified as hindbrain. FGF8 is expressed in the isthmus, as well as the ANB, and FGF8 is generally considered to be an important molecule in isthmus organizing activity [44]. Murine and avian studies have indicated that FGF8 can largely provide the functional equivalent of the isthmus tissue. For example, mice that are trans-heterozygous for the null and hypomorphic alleles of FGF8 lack most of the midbrain and cerebellum, indicating FGF8 is required for midbrain and hindbrain development [45]. FGF8 soaked beads can induce the expression of important midbrain/hindbrain patterning genes generally expressed by the isthmus and neighboring neural cells including En1, En2, Pax5, and Gbx2, but not Fgf8, in chick embryos and mouse brain explants [39], suggesting FGF8 is a strong inductive signal for MHB patterning. In addition to signals from the isthmus, transcription factors can interact to contribute to MHB patterning. For example, OTX and GBX2 mutually repress one another, creating a binary cell fate in the midbrain/hindbrain region [39]. Overall, the anterior-posterior fate of the midbrain and hindbrain regions are 12
25 dependent on signaling from the isthmus organizer, particularly FGF8, expression of WNTs, as well as mutual repression between other transcription factors. Specification of the eight hindbrain rhombomeres and the more caudal spinal cord is promoted by varying concentrations of the signaling molecule Retinoic Acid (RA) [46]. Low concentrations of RA are required for successively anterior rhombomere fates. One mechanism by which the rhombomere cell fate is determined is by Hox gene expression and RA signaling. RA controls the expression of Hox genes, which correlates with specific rhombomere fate, via retinoic acid response elements (RAREs) in the Hox gene cluster. RA binds to re0..0tinoic acid receptors (RARs) in the nucleus, which transduce the RA signal [47]. RARs form a heterodimer with retinoid receptors (RXRs) and bind to RAREs in the gene promoter to control the transcription of target genes, including Hox genes. Hox genes control the A/P body plan in almost all metazoans. Hox genes are generally arranged on the chromosome in the same order to which they are expressed along the body, referred to as collinear expression [48]. This is controlled by extensive feedback inhibition and regulation by the HOX proteins themselves, micrornas, and other cofactors. Hox gene expression along the A/P axis is dependent on binding of the HOX proteins to their gene promoters, which can control either activating and repressive functions, along with other DNA binding proteins that act as HOX cofactors, including RA. There is extensive negative feedback that allows specific HOX genes to bind to specific regions on the promoter. This control is evidenced by genetic studies in the mouse. Mutations in RAREs in the promoters of Hox1a and Hox1b result in reduced expression of these Hox genes and hindbrain defects [49, 50], suggesting that lack of RA binding at the RAREs in Hox promoters can change the expression of Hox genes. The complex control of A/P patterning 13
26 comprising rhombomere fate is better understood compared with A/P patterning in the spinal cord. While relatively little is known about A/P patterning in the spinal cord, it is known that the paraxial (adjacent) mesoderm plays a role in secreting RA and other signals that act on the neural ectoderm. High concentrations of RA secreted by the paraxial mesoderm are required for the differentiation of cells at the posterior end of the spinal cord [51]. This is similar to the specification of the hindbrain, in that higher concentrations of RA are required for specification of more posterior structures. Overall, the cell fate in the hindbrain and spinal cord are tightly controlled by secreted RA and subsequent RA signaling has been shown to promote HOX gene expression in the rhombomeres of the hindbrain. Dorsal/Ventral Patterning Establishing reliable mechanisms of neural progenitor specification is important in proper neural tube closure. Dorsal/ventral (D/V) patterning in the spinal cord organizes cell fates to result in the formation of 12 distinct neural progenitor domains. From ventral to dorsal, these domains are pv3, pmn, pv2, pv1, pv0, pd6, pd5, pd4, pd3, pd2 and pd1 (Figure I.4). These progenitor domains give rise to the ventral interneuron progenitor domains and motor neuron domains in the ventral aspect of the spinal cord, and dorsal interneuron domains in the dorsal aspect of the spinal cord. Dorsalization of the spinal cord utilizes a cascade of inductive signaling molecules derived from the cells of the non-neural ectoderm and roof plate (RP) in the neural tube (Figure I.4). Transforming Growth Factorβ (TGF-β) family members are the predominant signaling factors that mediate dorsal cell fates. Ventralization signals of the vertebrate spinal cord system are induced and mediated by SHH secreted from the notochord and floor plate (FP) in the developing embryo. These 14
27 factors and signaling cascades regulate expression of a specific combination of transcription factors in each population of neural progenitor cells, which then help to specify the fate of the distinct subsets of post mitotic neurons. Figure I.4 Patterning in the neural tube is set up by Sonic Hedgehog (Shh) and Bone Morphogenetic Protein (BMP). Expression of SHH, induced by the notochord and secreted from the floor plate (FP) of the neural tube, creates a soluble gradient which induces formation of various proneural domains in the ventral neural tube. Secretion of BMP from the roof plate (RP) forms a signaling gradient that induces pro-neural domains in the dorsal neural tube. These progenitor domains give rise to the dorsal and ventral interneurons (denoted in the colored circles). Image adapted from [52]. Ventral Patterning of the Spinal Cord Molecular signals for ventral patterning begin early in development. The notochord, the rod-like structure derived from the mesoderm, secretes Sonic Hedgehog (SHH), a strong morphogen. SHH signals through the Patched Receptor and Gli activators, and has the potential to influence the transcription of itself and of other transcription factors. SHH released from the notochord induces SHH secretion from the floor plate, which diffuses dorsally in a gradient that extends along the ventral to dorsal axis. [53]. The SHH gradient induces cells of the spinal cord to express different transcription factors based on the concentration, time and duration of exposure to SHH [54]. Some transcription 15
28 factors are induced and some are repressed by SHH signaling. For example, Class 1 genes, defined as being repressed at certain concentrations of SHH, include the homeodomain genes Pax6, Pax7, and Irx3. Class 2 genes are induced by SHH- Gli signaling and include Foxa2, Nkx2.2, Olig2, Nkx6.1, and Dbx2 [55]. However, the SHH morphogen gradient alone cannot account for the distinct pattern and refinement of the proneural transcription factor expression domains in the ventral neural tube. This highly ordered patterning system requires feedback between the transcription factors themselves. For example, Pax6 and Nxk2.2 can mutually repress each other in the ventral neural tube and this allows the refinement of an initially indistinct border between two cells that initially express both genes, into two differently patterned cells within neighboring domains [55]. Mechanisms such as these fine-tune the expression pattern of transcription factors in the neural tube to define specific neural progenitor domains. Dorsal Patterning of the Spinal Cord The dorsal half of the neural tube is also patterned by a secreted morphogen gradient, and subsequent activation and repression of transcription factors, in this case, members of the basic-helix-loop-helix (bhlh) family. The signaling molecules are members of the TGF- superfamily, which are secreted by the overlying non-neural ectoderm and the roof plate to induce six dorsal neural progenitor populations (pd1-6), and in turn give rise to dorsal interneurons di1-6. Dorsal patterning is less well understood than ventral spinal cord patterning, due in part to the fact that multiple TGF- family members are expressed in the dorsal spinal cord and they have redundant functions in neural progenitor induction. The TGF- family members BMP2, BMP4, and BMP7, are secreted from the non-neural ectoderm and can mediate dorsalization signals of the spinal 16
29 cord in Xenopus [56, 57] and are likely mediators in mice. The roof plate is also important for secreting signals to mediate dorsal spinal cord patterning. Roof plate ablation studies show the roof plate is necessary for the expression of the bhlh transcription factors Math1, Neruog1 and Neurog2, which are necessary for formation of d1-d3 interneurons [58]. Formation of the pd3-pd6 progenitor domains is independent of roof plate signaling [59]. While progenitor domains have varying responses to non-neural ectoderm and roof plate signals, complete spinal NT patterning relies on both dorsalization signals and ventralization signals. Together BMPs secreted from the RP and SHH secreted from the FP refine the expression of various transcription factors in the neural ectoderm. For example, Pax3, Pax7, Msx1 and Msx2 are initially expressed along the entire D/V axis of the neural ectoderm, but these genes are then repressed ventrally by SHH signaling. The expression of these transcription factors is also elevated dorsally by exposure to BMPs [57]. Overall, DV patterning in the neural tube is established via secreted morphogen gradients which can either induce or suppress transcription factors which can themselves induce or suppress one another, in order to specify distinct neural progenitor domains in the spinal cord. Apoptosis Neural tube closure (NTC) is regulated by other types of signal cascades, including the cascade that ultimately results in programmed cell death or apoptosis. Over 50 years ago, ultrastructural studies showed the existence of dying cells in the neuroepithelium, but it was unclear how these cells were dying [60]. Relatively recently, it was discovered that those cells were undergoing apoptosis [61]. Apoptosis occurs most frequently at the tip of the neural folds just before fusion, but can occur elsewhere in the neural epithelium. For 17
30 example, mouse embryos null for Mdm4 [62], an important gene that functions to inhibit the apoptosis cascade, exhibit an open cranial neural tube and have increased apoptosis in the neural tube compared to the WT littermates. These phenotypes are attributed to a reduced number of cells that can participate in the closure of the neural tube, suggesting proper cell number is important in neural tube closure. Mouse strains that are deficient for pro-apoptotic genes like p53 [63] and Caspase 9 [64] exhibit reduced apoptotic death and exencephaly. The excess number of cells, which would otherwise be reduced by apoptosis, appears to interrupt normal shaping, bending or fusion of the neural plate. Taken together, these studies suggest that changes in the extent of apoptosis can detrimentally affect neural tube closure. Despite the dramatic phenotypes caused by either decreasing or increasing apoptosis, blocking apoptosis with pharmacological agents during neurulation in the mouse does not appear to affect NT closure [65], suggesting apoptosis may not be necessary for neural tube closure. A live imaging study showed that inhibiting caspase activation delayed the kinetics of neural tube closure but did not result in NTD [66]. These authors observed cell fragmentation typical of apoptosis but they also discovered atypical caspase-positive but non-apoptotic cells that actively move, or dance, in the neural and non-neural ectoderm during neural tube closure. This study suggests there are more dynamic mechanisms of apoptosis that may contribute to the closing neural tube [66]. The studies outlined here use several different methods to analyze apoptosis during NTC and in NTDs, but the role of apoptosis during this critical phase is still controversial. Proliferation and Differentiation During cranial neural tube closure, all cells in the neural plate are proliferative; with differentiation occurring only after neurulation is complete. Cell proliferation is 18
31 linked with differentiation because as cells become post-mitotic they begin to differentiate into neuronal cells. The cell cycle is a short 4-6 hours and proliferation is tightly regulated along the dorsal and ventral axes of the neural tube, with the dorsal half proliferating more than the ventral half [67]. If cells differentiate early, they exit the cell cycle and no longer proliferate, resulting in reduced numbers of neural progenitors and neurons. For example, inactivation of Hes1 leads to early appearance of neuronal markers during neurulation, preventing cranial neural tube closure [68]. The tight control of cell proliferation is necessary for the neural folds to meet properly at the dorsal midline. Too much or too little cell proliferation can abrogate neural tube closure. Excess proliferation has been implicated as increasing the risk for NTD in some mutants. For example, overexpression of Notch3 in mice results in increased numbers of neuronal progenitor cells and exencephaly [69], suggesting an increase in cell proliferation within the neural tissue may result in too many cells to allow proper apposition and fusion of the neural folds, resulting in an NTD. Hypoproliferation, on the other hand, is thought to generate insufficient tissue growth to allow for appropriate bending of the neural plate and subsequent fusion. The tight control of cell proliferation in tissues outside the neuroepithelium is also critical for NT closure. For example, the Twist knockout mice show an NTD associated with a cellular deficiency in the cranial mesenchyme due to reduced proliferation [70]. These studies indicate that loss of proliferative control in several different locations can lead to NTDs. 19
32 NEURAL TUBE DEFECTS IN CHROMATIN MAINTENANCE MUTANTS Chromatin Structure DNA exists in the nucleus as a highly ordered structure called chromatin. In this state, DNA is largely inaccessible to transcription machinery. The basic organizational unit of chromatin is the nucleosome: 147 base pairs of DNA wrapped around a histone protein octomer [71]. Histone proteins are important in maintaining the structure of chromatin. The nucleosome stability on the DNA is maintained by the interaction of the positively charged histones cores interacting with the negatively charged DNA phosphate backbone. In vivo studies indicate nucleosome positioning may be encoded in the genome, suggesting some sequence specificity for nucleosome binding [72]. Each histone protein has a flexible amino acid region at the N- terminus, termed a histone tail, which is free from the wrapped DNA. While the core structure of a histone is highly conserved, there are many histone variants that differ greatly in primary sequence [73]. A nucleosome can be composed of different histone variants. Specific compositions of histone variants are found in different contexts and are thought to confer different functions. For example, histone variant H3.3 is found in active chromatin sites and may promote transcription [74]. Additionally, higher eukaryotes possess a unique histone known as a linker histone, H1. This histone can connect two nucleosomes and can increase compaction of chromosomes [75]. Chromatin structure can be modified in several ways discussed below to allow for flexible organization of DNA in the nucleus, recruitment of specific transcription factors, and changes in gene expression. 20
33 Epigenetics Epigenetics is the study of heritable changes in gene transcription that are independent of underlying DNA sequence [76]. For example, in a developing organism, gene transcription must be significantly altered for cells to become restricted to specific linages. This is important because, as an organism develops, different cell types must express lineage specific genes, even though each cell contains the entire genome and is therefore genetically identical. These changes in expression between different cells in a given organism or given tissue are carried out by epigenetic mechanisms. Molecularly, there are three main types of epigenetic processes: 1. DNA methylation, which is a covalent modification to the DNA. 2. Histone modifications, which include methyl or acetyl groups added to histone tails. 3. Chromatin Remodeling or the movement of nucleosomes along the DNA. NTDs are thought to be multigenic because approaches to correlate single gene mutations have thus far revealed only a few gene variants that may contribute to human NTDs. Thus, studying epigenetic processes in the context of neural tube defects may uncover causes for these congenital anomalies. NTDs in mice can arise as a consequence of mutations in genes that mediate epigenetic regulation [19, 77]. Here I will review the mechanisms of epigenetic regulation and how they relate to neural tube defects, with particular emphasis on chromatin remodeling. DNA Methylation DNA methylation can restrict access of transcription factors to regions of the genome. Covalent attachment of a methyl group to DNA occurs on the Guanine of a Cytosine and Guanine residing together, known as the CpG dinucleotide. These 21
34 modifications inhibit the binding of transcription factors and are generally associated with gene silencing. There are two classes of methyltransferases that catalyze this modification. DNMT1 is a maintenance enzyme that adds methylation to hemi-methylated regions of DNA after replication [78]. This function is essential for maintaining methylation patterns in proliferating cells. DNMT3a and DNMT3b methylate CpG islands de novo to establish DNA-methylation patterns throughout development and thus allow for cells to change transcriptional programs [79]. Evidence indicates that DNA methylation is important in proper closure of the neural tube. For example, Dnmt3b -/- embryos exhibit exencephaly at variable penetrance [79]. Additionally, embryos cultured in the methylation inhibitor 5- azacytodine develop NTDs [80]. The mechanisms underlying these NTDs are poorly understood, however in theory; inhibiting methylation would prevent silencing of critical genes during development, thus disrupting the normal developmental pattern of gene expression which may contribute to failure of neural tube closure. Histone Modification Addition of covalent modifications to histone tails is an epigenetic mechanism to indicate and modulate active or repressive transcriptional signals. Positively charged lysine, arginine and polar neutral serine residues on histone tails can be covalently modified by addition of acetyl groups, methyl groups, and others [81]. Unlike DNA methylation, which usually results in gene silencing, these modifications can confer activation or repression of a given locus. A single modification does not determine the transcriptional outcome; rather it is the combination of all the acetyl and methyl residues on all histone tails in a region that refine gene expression levels. A recent report suggests there may be 51 different chromatin states based on enrichment of histone modifications 22
35 that lead to different transcription levels [82]. Acetyl groups are added to lysine residues by histone acetyl acetyltransferases (HAT) enzymes and removed by histone deacetylases (HDACs) [81]. Genome wide association studies have found that enzymes with HAT activity are generally associated with transcriptional activation and enzymes with HDAC activity are located in both active as well as inactive genes [83]. HATs are found within protein complexes called co-activators, while HDACs are found within complexes called co-repressors. Mice with mutations in several genes encoding histone acetyltransferases and histone deacetylases exhibit neural tube defects. These include the HATS Kat2a (also known as Gcn5) [84, 85], Cbp [86] and p300 [87], as well as the HDACS Hdac4 [88] and Sirt1 [89]. Embryos null for CBP or P300 show a severe NTD from the midbrain to the hindbrain. CBP and P300 are transcriptional co-activators that each contains a HAT domain. They can form a complex with the activator TFIIb or with RNA polymerse II to promote transcription [90], suggesting that disruption of transcriptional activation may be responsible for these neural tube defects. These two genes are part of the same Kat3 histone acetylation family and do not share significant homology to any other histone acetyltransferase domains [91], which might explain why these mouse mutants have similar defects at this critical time of neural tube closure. Both addition and removal of histone modifications by their respective enzymes, and the complexes they form appear to be an epigenetic mechanism that is important in neural tube closure and overall transcriptional control. Chromatin remodeling is the epigenetic mechanism that changes the chromatin overall structure to affect the dynamics of transcription [92]. This remodeling is carried out by disrupting the contacts between the DNA and the histone proteins. Early in vitro studies 23
36 using histones assembled on a DNA template to form chromatin have indicated nucleosomes are predominantly remodeled by 4 different mechanisms: 1. nucleosome sliding along the DNA to an alternate position, 2. DNA-histone contacts breaking and reforming, but the nucleosome remains on the DNA [93] 3. nucleosome removed from the DNA [94], and 4. nucleosome replacement for other histone variants [95] (Figure I.5). It should be noted that these processes are not mutually exclusive and can occur simultaneously. Chromatin Remodeling Chromatin remodeling requires energy in the form of ATP to break the histone- DNA contacts. All chromatin remolding ATPases contain both the Dead/H ATP binding domain and the HELICc domain. There are 4 families of ATPases classified by the other functional domains that they contain. These four families are ISWI, CHD, INO80 and SWI/SNF. Each family is conserved in yeast through humans; however each species can contain different orthologs of these families. Although the four families are distinguished by different structural domains, they are all able to perform all four versions of nucleosome remodeling in vitro [96]. In vivo, the chromatin remodeling complexes form large multisubunit machines to carry out their functions. The chromatin remodeling complexes are characterized by which ATPase they contain [92]. ISWI Family The ISWI family of ATPases has unique functions when associating with distinct complexes that affect both developmental and transcriptional pathways. ISWI contains the Dead/H-ATPase cassette as well as the SANT and SLIDE domains which contribute to functional differences between the other families of ATPases. The ISWI ATPase was 24
37 originally discovered in an in vitro assay for nucleosome remodeling activities using Drosophila embryo extracts [98]. Subsequent in vitro assays indicated the single ATPase was necessary for transcriptional activation [98]. Several ISWI complexes are essential for organismal viability. D. melanogaster has only a single ISWI ATPase and it is required for embryogenesis, demonstrating its functional importance. Figure I.5 Molecular mechanisms of nucleosome remodeling in vitro. Nucleosome remodeling in vitro can occur in 4 different ways as a consequence of ATP-dependent remodeling activity. A. The nucleosome can slide along the DNA to another position. B. DNA-histone contacts can be broken to increase accessibility to DNA but the histone remains in the same position. C. Nucleosomes can be displaced from the DNA to open chromatin structure. D. The core nucleosome can be replaced by a nucleosome containing histone variants. Image reproduced from [97]. In mammals, there are two ISWI ATPases- SNF2H and SNF2L. Deletion of the SNF2H ATPase in mice is embryonic lethal, after embryonic implantation, possibly due to impaired proliferation [99]. Mice deficient for another ISWI complex containing the SNF2L ATPase die between E7.5 - E8.5. At this time in development, the A/P axis and primitive streak are developing. The mutant mice lack mesoderm and endoderm 25
38 differentiation [100]. Therefore, the ISWI ATPases and the complexes they form are essential in embryonic development; however, the differing mutant phenotypes show they have non-redundant roles. ISWI and its chromatin remodeling subunits are also essential in neurulation and neurogenesis. SNF2L forms a chromatin remodeling complex with the transcription factor CECR2. During development this transcription factor is expressed in the primitive central nervous system. Cecr2 homozygous null animals show exencephaly with variable strainspecific penetrance [101]. This suggests CECR2 and by extension the chromatin remodeling complex it forms with SNF2L are involved in neural tube closure. SNF2L is also involved in other aspects of neurulation. SNF2L knockdown in human cells reduces expression of Engrailed, which is essential in mid-hindbrain formation [102]. In addition to roles in neurulation and patterning, ISWI complexes function later in neurogenesis. Consistent with roles in neurogenesis, ATPase SNF2L expression in a neuroblastoma cell line can initiate neurite outgrowth. These studies suggest ISWI complexes and ATPases may initiate expression of developmental genes and gene networks during neurogenesis. Therefore, ISWI ATPases and the complexes they form in vivo function in the morphogenetic process of neural tube closure and transcriptional activation during neurogenesis. INO80 Family INO80 family of ATPases has distinct functions in cellular maintenance and development. It was originally purified in yeast as a large, 15-subunit complex. The INO80 family of ATPases is characterized by a split ATPase domain, about 200 amino acids between the DEXD/H and HELICc domains. This spacing allows incorporation of 26
39 additional subunits into the complex that bind to sites of DNA damage, known as Holliday Junctions [103]. Functionally, the INO80 complex is implicated in the cell cycle checkpoints, DNA damage response, and H2A.Z deposition [104], roles that may be mediated by the split ATPase domain and the binding of specific complex proteins. In mammals, the INO80 complex TIP60-p400 includes p400, which is an INO80 related ATPase, and TIP60, an enzyme with HAT activity. The combination of these two enzymes as well as the other 14 subunits in the complex have functions in Embryonic Stem (ES) cell maintenance [105]. For example, the Tip60-p400 complex suppresses differentiation in ES cells through controlling transcriptional programs. Depletion of TIP60 or p400 results in an altered ES cell morphology, premature differentiation and cell cycle arrest. Microarray analysis of the Tip60 and p400 depleted ES cells showed differentiation genes were activated upon depletion of the two complex proteins [106]. Neither TIP60 nor p400 has been deleted in mice, so the function in mammalian development is yet unknown, but the studies in ES cells suggest these proteins may be essential to repress differentiation of ES cells. Overall, INO80 ATPase family is necessary for many aspects of cell survival and for ES cell self-renewal and suppression of differentiation. CHD Family The CHD family is the least well-characterized in mammals of the four ATPase families. It contains 9 family members defined by their tandem chromodomain repeats. Chromodomains are able to interact with methylated histones [107]. The CHD-related ATPases are classified into 3 broad subfamilies. Subfamily I (with ATPases CHD1 and CHD2); Subfamily II (CHD3 and CHD4); and subfamily III (CHD5, CHD6, and CHD7)[108]. Subfamily II consists of CHD3 and CHD4 subunits which are members of 27
40 the NuRD (Nucleosomal remodeling and histone deacetylase) repressive complex. NuRD is thought to be recruited to transcriptionally active DNA by binding of its chromodomains to chromatin. The ATPase activity increases the accessibility of the complex to the acetylated histones by remodeling nucleosomes, and the associated HDAC deacetylates the histones tails to repress the previously activated genes [109]. This complex contains 6-7 interchangeable subunits, including an HDAC subunit, which is responsible for the repressive functions in vivo [110]. The diverse subunit combinations can result in opposing functions of the NuRD complex. For example, the NuRD complex with an MTA3 subunit, not MTA1, and a BCL6 subunit, maintains normal differentiated, noninvasive phenotype of breast epithelial cells, repressing the expression of metastasis-inducing gene Snail. Expression of MTA1 in breast epithelial cells represses MTA3, resulting in expression of Snail, and a cancer-like mesenchymal phenotype [111]. These studies highlight the fact that subunits of the NuRD complex may be exchanged for context specific partners which confer various functions. Another CHD family member, CHD7, has developmental roles in humans as well as mice. Mutations in CHD7 in humans results in CHARGE syndrome, characterized by eye defects, heart defects, growth retardation, and ear abnormalities [112]. Chd7 +/- mice recapitulate some of these phenotypes, including inner ear dysfunction [113]. This suggests that CHD7 is functionally important in inner ear development in mice and humans, and that CHD7 function may be conserved in the mouse and human. Overall, the CHD family of ATPases can form complexes that associate in a combinatorial manner with cell-type-specific functions, and have general, non-redundant functions in organismal development. 28
41 SWI/SNF Family The Swi/Snf family of chromatin remodelers was originally described in yeast [114]. It is characterized by the presence of a bromodomain which can bind acetylated histones and bring epigenetic interacting proteins closer to chromatin [115, 116]. The bromo and ATPase domains provide a means of interaction between the Swi/Snf complex and the chromatin. The Swi/Snf ATPase is not essential for yeast survival and is estimated to only activate transcription of approximately 6% of the genes in the yeast genome [117]. Within multicellular organisms, compared to unicellular organisms, the function of the Swi/Snf complex has become more diverse and also necessary for organismal survival. In Drosophila, there is one Swi/Snf related ATPase called Brahma (Brm). Proteins within the Swi/Snf related complex in flies, known as the Brahma Associated Proteins (BAP) complex are essential for survival. The Brm ATPase, as well as other members of the BAP complex regulate gene expression in the developmental processes of wing venation and in the larval-pupal transition [118]. In humans and mice, there are two SWI/SNF ATPases: BRG1 and BRM. These two ATPases are interchangeable subunits in the mammalian BRG1/BRM Associated Factor ATP-dependent chromatin remodeling complex known as the BAF complex. An active mammalian BAF complex consists of at least 15 subunits, 3 of which have been recently characterized [119]. Several of the 15 subunits have interchangeable isoforms, and a total 26 genes that encode subunits have been identified [120] (Figure I.6). Most of these subunits can be replaced by one or more isoforms. In different cell types, the BAF complex is composed of different subunit isoforms. These cell-type specific BAF complexes are crucial for maintaining the appropriate developmental state of the cell. 29
42 Figure I.6 Schematic of the BAF chromatin remodeling complex. The ATP-dependent BAF Chromatin Remodeling Complex is a large 2MB molecular machine composed of at least 15 subunits. Three subunits are not depicted here: BCL7, BCL11 and SS18. The subunits with more than one isoform contribute only one protein per isoform to each individual complex. For example, a BAF complex contains one protein of the BAF60 isoforms: BAF60A, BAF60B or BAF60C. The BAF complex with different protein isoform compositions has been shown to have cell-type specific roles. Image adapted from [121] The BAF chromatin remodeling complex is essential in many aspects of mammalian development, and some subunits are necessary for viability. Mice lacking one of the two ATPases of the BAF complex, BRG1 and BRM, have distinct phenotypes. Brg1 -/- mice are not viable, whereas mice deficient for BRM develop normally but exhibit a small body mass [122], demonstrating BRG1 is essential for the developing embryo but BRM is dispensable. Mice deficient in BAF155 [123] or BAF47 [124] exhibit periimplantation embryonic death, due to the lack of the placental trophoblast layer. This and the data below provide further evidence that BAF complex proteins, and by extension the BAF chromatin remodeling complex, functions are essential for viability. Embryonic stem (ES) cells are pluripotent and have the capacity for limitless selfrenewal. If these characteristics are not maintained, ES cells differentiate. The specific assembly of the BAF complex in ES cells has been called esbaf because this set of proteins are not found in other cell types and the esbaf complex has specific roles in ES cell function [125]. The esbaf complex is composed of distinct subunits which include a 30
43 homodimer of BAF155 and the exclusion of BAF170. Specifically, BAF155 is functionally necessary for ES cell proliferation and self-renewal capacity, and cannot be replaced by BAF170 [125]. Molecularly, esbaf has been described as a transcriptional refiner because it reduces the expression of pluripotency genes like Oct4 and Sox2, but does not silence them, thus regulating the expression levels of these critical genes [126]. The esbaf complex also has a repressive role in ES cells. BRG1 and BAF155 suppress the expression of differentiation genes including Pax6 and Fgf5. BRG1 synergizes with PRC2 to maintain the H3K27me3 repressive mark on several pro-differentiation genes in the genome. Conversely, BRG1 antagonizes PRC2 repression at specific target genes induced by the cytokine leukocyte inhibitory factor (LIF), a strong pluripotency factor that represses differentiation [127]. This indicates that BRG1 can prime the ES cell genome for differentiation by localizing to repressed differentiation genes in the LIF/Stat3 pathway. Overall, esbaf helps maintain ES cells through its roles in both refining the expression of important pluripotency genes and repressing subsets of differentiation genes, while promoting others for differentiation. This suggests BAF s function is extended from remodeling nucleosomes to acting as an overall gene regulator by interacting at specific loci to refine the pluripotency transcriptional network. EsBAF is an example of the complexity of functions that can be specified by the combinatorial assembly of the BAF complex. For an ES cell to differentiate, it must exit the pluripotency state and silence selfrenewal genes. RNAi knockdown of the esbaf subunits BAF155 and BAF57 prevents silencing of the critical self-renewal gene Nanog upon addition of the differentiation factor Retinoic Acid (RA) in ES cells. This knockdown prevents Nanog silencing even in the 31
44 presence of self-renewal factors like Oct4 [128], indicating these esbaf subunits are strong regulators of the exit from the self-renewal of ES cells. Additionally, BAF155 is necessary for heterochromatin formation globally and locally at pluripotency genes during differentiation induced by RA [128]. In the context of induced neuronal differentiation, BAF155 and BAF57 act to promote differentiation by both silencing self-renewal factors and promoting chromatin compaction, two functions that may be related. This is an example of the diverse and cell-type specific functions that the mammalian BAF complex has at both the cellular and molecular level. Heart development is another embryonic process that requires specific signals for specification of cell fate and to promote correct morphology. The tissue-specific roles for subunits of the BAF complex have been defined in cardiac development. Baf60c is expressed specifically in the heart and somites of the developing mouse embryo. sirna knock-down studies suggest Baf60c is required for heart morphogenesis, cardiac and skeletal muscle differentiation [129], and for establishing left-right asymmetry [130]. Cardiac development requires precise expression and recruitment of transcription factors. Baf60c mediates BAF complex interactions with cardiac transcription factors including GATA4 and TBX5 and can ectopically specify cardiomyocytes outside of the developing heart regions in mouse embryos [131]. These studies highlight the importance of BAF60c in heart morphology and cardiac specification. Overall, the BAF complex subunits show cardiac-specific functions during an essential period in development. The BAF complex has a critical role in neural development as well. The BAF complex subunits are different in neuronal progenitor cells compared to ES cells. Moreover, the BAF complex subunits change as the neural progenitor cells undergo 32
45 differentiation. Elegant studies in the mouse suggest a switch of specific subunits of BAF45 and BAF53 in the BAF complex are sufficient for cellular specification and transition from neural progenitors to post-mitotic neurons [132]. The composition of these subunits are termed the neural progenitor BAF (npbaf) and the neuronal BAF (nbaf). Molecularly, the switch between the neural progenitor subunit BAF53A and the neuronal subunit BAF53B is mediated by mirnas and the repression transcription factor known as REST [133]. Repression of mir-9* and mir-124 by REST results in expression of the BAF53A subunit and proliferation characteristic of neural progenitor cells, suggesting BAF53A is essential for self-renewal in neural progenitors. Neuronal cells that have exited the cell cycle also rely on a switch in the BAF53 subunit. Expression of the same two mirnas (mir-9* and mir0124) results in repression of BAF53A, and then the expression of the BAF53B subunit. BAF53B increases dendritic growth in post-mitotic neurons in the E12.5 neural tube. These studies may suggest how other subunits are exchanged within the complex. Overall, specific neural signaling or neuronal functions are mediated by different subunits of the BAF complex through interaction with transcription factors or mirnas. The BAF complex also has general neural developmental roles mediated by BRG1. BRG1 is required for the self-renewal of neural progenitors. Conditional inactivation of Brg1 by Cre recombinase specifically expressed in neural progenitors results in thinning of the neocortex and absence of the cerebellum [132]. This suggests that BRG1 maintains the neural progenitor population in order to generate sufficient numbers of progenitors to ultimately give rise to the correct number of neurons to form essential brain structures. The BAF complex is also required for regulating signaling pathways in the developing brain. BRG1 controls expression of several genes in the SHH and Notch signaling pathway at 33
46 E12.5, suggesting neurogenesis might be directed transcriptionally by BRG1 [132]. This data suggests BRG1 refines expression of genes in two important signaling pathways during neurogenesis. BRG1 also functions in neurulation at E9.5 as Brg1 +/- mice show neural tube defects at a 15-30% penetrance [134]. These studies indicate the importance of the BAF chromatin remodeling complex in the development of the early CNS. As highlighted above, BAF155 is important during early development [123, 135] and ES cell differentiation [125, 128]. In addition to a developmental role, BAF155 is also implicated in early B-cell [136] development, thymocyte survival [137], as a tumor suppressor in mouse models [138], and as a modulator of Androgen transactivation in mouse prostate cells [139]. However, it has been difficult to study the function of the protein specifically during later stages of development and in particular tissues and organs due to the early lethality of Baf155 null/null mouse embryos. It is also established that Baf155 has a role in neural tube closure; as 20% of Baf155 +/null heterozygous embryos exhibit exencephaly with increased cranial proliferation observed 4 days after neural tube closure [135]. However, the role of BAF155 has not been determined during the time of neural tube closure. The focus of this thesis is on the role of the BAF chromatin remodeling complex during neural tube closure, and why neural tube defects arise upon loss of BAF function. Specifically, I have studied a unique Baf155 mouse mutant exhibiting an NTD and characterized its affect molecularly and cellularly and its role in transcriptional control. 34
47 CHAPTER II THE MSP3 MUTANT MOUSE EXHIBITS EXENCEPHALY AND OTHER PHENOTYPIC DEFECTS INTRODUCTION Neural tube defects (NTDs) result when the embryonic neural tube, which gives rise to the adult brain and spinal cord, fails to close completely. NTDs are the second most common birth defect in humans, occurring in ~1 in 1000 live births worldwide [5, 37]. NTDs include caudal neural tube closure defects such as spina bifida, and cranial neural tube defects such as exencephaly. These devastating birth defects occur due to disruption in the intricate process of neural tube closure, which requires the coordination of very precise cellular and molecular mechanisms. The complexity of this process is highlighted by the fact that over 200 genetic mutations in the mouse have been identified that result in NTDs [17-19]. These affected genes play roles in several different processes during neural tube closure, including cell cycle regulation, neurogenesis, cell viability, developmental signaling, and epigenetic regulation of transcription [9]. NTD mutants also often show a wide range of other phenotypes, implicating the mutated gene in the regulation of other developmental processes. Ethylnitrosourea (ENU) mutagenesis and forward genetic screen is an unbiased method that can be used to identify genes and molecular pathways involved in a given developmental process [140]. ENU mutagenesis followed by a forward genetic screen was conducted in the lab of Dr. William Pavan in an effort to identify phenotypic abnormalities and misexpression of the neural crest marker Sox10 in the mouse, referred to as modifiers of Sox10 expression pattern (msp). [141]. One abnormality identified in the screen was an 35
48 open cranial neural tube at embryonic day 11.5 (E11.5), and this was called the Msp3 phenotype. Dr. Pavan generously gifted the mouse line containing the Msp3 phenotype to Dr. Niswander s lab, as her lab specializes in characterizing the mechanisms of murine neural tube defects. In the original publication [141], the region carrying the mutation that causes the Msp3 phenotype was narrowed to an 8 Mb region of chromosome 9; however, the specific gene mutation was not identified. Later, the gene which harbored the ENUinduced mutation was identified as Smarcc1 encoding the BAF155 protein. For the remainder of the thesis, I will refer to the mutant gene as Baf155 msp3 and the protein as BAF155 msp3. The discovery of the causative mutation in Smarcc1 will be discussed in Chapter III. The majority of this thesis focuses on characterizing this mouse line and the molecular consequences of the mutation. The first task of this thesis is to characterize the phenotypic abnormalities which manifest in the Baf155 msp3 mouse line. RESULTS 1 Baf155 msp3 MUTANTS EXHIBIT CRANIAL NEURAL TUBE DEFECTS The Msp3 phenotype was originally found in an ENU mutagenesis screen in mice [141], and the allele was subsequently identified as the Baf155 msp3 mutation, discussed below. Mice heterozygous for the Baf155 +/msp3 mutation are indistinguishable from wildtype littermates in terms of size, cranial structure, and behavior based on observation. They are viable and fertile, producing an average of 7-8 pups per litter. Homozygous Baf155 msp3/msp3 mutant embryos display cranial neural tube defects and developmental delay (combined phenotypes at 92% penetrance, exencephaly at 81% frequency, and developmental delay with and without exencephaly at 30% frequency) 1 Parts of chapters 2-5 and all of Appendix E are taken with permission from Harmacek, L., D. E. Watkins-Chow, J. Chen, K. L. Jones, W. J. Pavan, J. M. Salbaum and L. Niswander (2013). "A unique missense allele of BAF155, a core BAF chromatin remodeling complex protein, causes neural tube closure defects in mice." Dev Neurobiol. 36
49 (Figure II.1A-C and Table II.1). The embryonic phenotypes range in severity and include exencephaly, coloboma, vascular defects, and developmental delay, defined as having at least 2 somites less than the number of somites in the non-mutant littermate with the fewest somites. For example, Figure II.1A shows embryos from a single litter at E9.5, just after the time of neural tube closure. Mutant embryo 1 (MUT 1) is exencephalic from the forebrain through the hindbrain but not delayed (25 somites), MUT 2 shows midbrain and hindbrain exencephaly and developmental delay (21 somites), MUT 3 is severely delayed (14-15 somites) and hence cranial neural closure cannot be scored. Additionally, embryos from a single litter at E15.5 exhibit different phenotypes (Figure II.1C). Mutant embryo 1 (MUT 1) shows midbrain and hindbrain exencephaly with the eye defect coloboma, or a hole in the eye structure, whereas mutant embryo 2 (MUT2) exhibits midbrain exencephaly, vascular defects and epidermal edema. Coloboma presented only in MUT 1 and variable vascular defect phenotypes were observed in 2 of 5 mutant embryos at E14.5 and E15.5. Another observed defect was an enlarged pericardium at E9.5 (~12% of mutants) (data not shown). Most homozygous mutants can survive until E15.5 but no homozygotes are found at the time of weaning (Table II.1), suggesting the mutation is embryonic lethal at E16.5 or later. Preliminary data on additional observed defects and phenotypes are described in appendix A. No other morphological defects beyond the ones outlined in this thesis were observed. CONCLUSION This chapter characterizes the Msp3 phenotype in the Baf155 +/msp3 heterozygous and homozygous mutants. The most apparent defect in the Baf155 msp3/msp3 homozygous mice is a cranial NTD, which is 80% penetrant. This defect occurs in a spectrum of 37
50 Figure II.1 Baf155 msp3/msp3 embryos show neural tube defects. A-C: Lateral views of embryos at the indicated stages. A) Within a single litter, three individual homozygous Baf155 msp3 mutant (MUT1-3) embryos show a range of phenotypes including failure of neural tube closure and developmental delay compared to wildtype (WT). B) E12.5 Baf155 msp3/msp3 mutant shows exencephaly without other apparent defects. C) E15.5 mutants show a range of phenotypes including exencephaly and coloboma, and vascular defects. Red scale bar=1 mm, blue scale bars=2 cm Table II.1 Phenotype and genotype of embryos resulting from cross between heterozygous Baf155 +/msp3 mice. Phenotype of Baf155msp3/msp3 embryos dev delay exencephaly (unable to [with dev score normal delay] exencephaly) morphology +/+ +/- -/- E [9] 5 3 E [8] 3 4 E [4] - 1 E [1] 2 - E [0] 2 - E [0] 1 - E [0] - - E Adult
51 severity, and therefore is variable, the meaning of which will be discussed in chapter IV. Additionally there are potential defects in the pericardial sac and vasculature. No Baf msp3/msp3 homozygous embryos are found after E15.5, this suggests that the Baf155 msp3 allele may be homozygous recessive embryonic lethal. However, only one litter was examined at E16.5 and later embryonic stages were not examined. While exencephaly is the most prominent phenotype in the homozygous mutant embryos, NTD itself does not cause embryonic lethality in mice or humans. Nonetheless, exencephalic neonates would die soon after birth and would not be observed at weaning. Thus, lethality may be due to a different cause, and this is frequently due to cardiovascular or placental defects [19]. Vascular leakage, variable edema and hemorrhage were observed in a subset of the Baf155 msp3/msp3 embryos at E15.5. Whether these phenotypes are a direct consequence of the BAF155 msp3 mutation or an indirect effect remains to be determined. The phenotypes observed in the Baf155 msp3/msp3 mutants are consistent with organismal death but the cause of death is currently unclear and not studied here. The vascular defects observed in the Baf155 msp3/msp3 homozygous mutants may result from a malfunction of mutant BAF155 msp3. BAF155 has been implicated in yolk sac development, and embryos carrying a rescue mutant allele of Baf155 show edema and vascular defects [142]. In rescue experiments in which a Baf155 transgene was inserted into the Baf155 null/null background, there was a lack of pro-angiogenesis factors such as Angiopoetin1, Tie2, and EphrinB2 in the yolk sac, suggesting BAF155 controls the expression of these factors during yolk sac development. Embryos in these studies died at E10.5, so later phenotypes could not be scored in this transgenic model [142]. Additional evidence supports the possibility that BAF155 may function in vascular development. 39
52 Embryos lacking the microrna mir-126, which was suggested to control endothelial integrity, show similar phenotypes to the E15.5 Baf155 msp3/msp3 mutant embryos of edema and vascular leakage [143], suggesting that BAF155 may control endothelial integrity. The vascular defects in Baf155 msp3/msp3 mutant described here were only observed in a few individual embryos and not studied in detail; therefore BAF functions with respect to vascular development cannot be definitively ascertained. MATERIALS AND METHODS Mouse Strains Baf155 msp3 (Smarcc1 msp3 ) was originally identified on a mixed genetic background (BALB/cJ; C57BL/6J) in an ENU screen [141]. The mutation in Smarcc1 was identified as described in the results in Chapter III. Mice have been maintained on a mixed C57BL/6J:129S1/SvlmJ background as a 3 rd generation cross and heterozygous carriers were mated to produce the embryos and results presented here. Further crossing into C57BL/6J results in more penetrant developmental delay. For timed pregnancies, noon of the day of an observed vaginal plug was designated E0.5. At dissection, the embryonic phenotype was recorded and a portion of the yolk sac used for genotyping. DNA was isolated from the yolk sac by lysing in PCR Lysis Buffer (50 mm Tris (ph 8.8, 1mM EDTA, 0.5% Tween-20 and 0.2 mg/ml proteinase K). Samples were digested overnight at 56 C and heat inactivated at 95 C for 10 minutes. Genotyping DNA samples were genotyped using a custom TaqMan assay (Applied Biosystems) with Taqman probes designed across the site of the msp3 mutation specific for both the wildtype allele (Vic-CTC-CTG-TTG-TAA-CTG-C) and the ENU induced mutant allele 40
53 (Fam-CTC-CTG-TTT-TAA-CTG-C). The following primers were used, forward primer: TTT-GCA-GAT-GAG-CAG-GAT-GAA-GAA and reverse primer: TCT-CAT-TTC-AGG- CCT-AAA-TAA-ACT-TTT-ACC-T. PCR reactions were carried out in 2x Taqman Universal Fast PCR Master Mix (Applied Biosystems, ), 10 M each genic primer, and 100 nm of each allele-specific probe. Cycling conditions were 95 C for 10 minutes, 40 cycles of 95 C for 3 seconds, and 60 C for 30 minutes. Reactions were analyzed on a Roche Light Cycler 480 in the Niswander Lab. Relative quantitation of the two alleles was determined in an endpoint assay for genotyping on the Light Cycler machine. 41
54 CHAPTER III MOLECUALR AND CELLULAR CHARACTERIZATION OF THE Baf155 msp3 MUTANT SHOWS A DECREASE IN CELL PROLIFERATION AND CELL SURVIVAL INTRODUCTION The causative mutation giving rise to the Msp3 phenotype characterized in Chapter II is localized to the Smarcc1 gene encoding the BAF155 protein. This protein is a member of mammalian BRG1/BRM Associated Factor ATP-dependent chromatin remodeling complex (BAF complex), which contains an estimated 15 protein subunits encoded by 26 genes [120] and is part of the Swi/Snf family of chromatin remodelers originally described in yeast [114]. In many organisms, including mice and humans, investigation of the BAF chromatin remodeling complex in different cell types indicates significant heterogeneity in subunit association. For example, the BAF complex is composed of different protein isoforms in embryonic stem (ES) cells, developing cardiomyocytes, and neural progenitor cells, suggesting there are tissue and cell-type specific roles for the complex during development [121]. The core components of the complex, ATPase BRG1 or BRM, along with BAF155, BAF170, and BAF47/INI5, have been isolated from all cell types studied to date and can remodel nucleosomes in vitro at the same efficiency as the fully intact BAF chromatin remodeling complex [144]. This core set of BAF proteins is particularly important in vivo, as complete loss of BRG1, BAF155 or BAF47 all result in periimplantation embryonic death and heterozygous loss of expression can lead to variable penetrance cranial NTDs [124, 134, 135]. 42
55 BAF155 assembles with the mammalian BAF chromatin remodeling complex to energetically disrupt histone-dna interactions to ultimately translocate histones on the DNA template, or remodel chromatin. In addition to being a core component of the BAF complex, BAF155 also plays an important role in maintaining the BAF complex. For example, BAF155 protects BAF complex proteins from degradation and maintains the nuclear localization of individual BAF proteins [145, 146]. Moreover, BAF155 shows a near-perfect overlap of association with BRG1 on the ES cell genome [147], indicating BAF155 represents assembled BAF complexes on chromatin in vivo. Overall, BAF155 is a critical protein for the molecular integrity and function of the BAF complex. Despite this critical role, the functions of BAF155 and the BAF complex have not been studied during neural tube closure. The purpose of this chapter is to verify the causative mutation that results in the Msp3 phenotype and characterize the molecular and cellular consequences of the Baf155 msp3 allele in vivo and in vitro. RESULTS Mapping the ENU-Induced Mutation to Baf155 The discovery of the ENU-induced allele and the initial characterization of the Msp3 phenotype, which consists of an open neural tube, were completed in the lab of Dr. William Pavan by Dawn Watkins-Chow at NIH. The Msp3 phenotype was identified in an ENU mutagenesis screen on a mixed genetic background (BALB/cJ; C57BL/6J) and localized to mouse chromosome 9 using traditional linkage mapping [141]. During subsequent outcrossing to C57BL/6J, the critical interval was narrowed to a 2Mb region (rs D9Mit37; NCBI Build36 chr9: 108,655, ,607,406). Resequencing of 43
56 exons in the interval was completed using an exon-based hybrid selection strategy (Illumina Genome Analyzer, Broad Institute, Mouse Mutant Resequencing Initiative). Filtering of coding or splicing homozygous variants not present in dbsnp (database of SNP variants, found at revealed a single variant in Smarcc1 that was not present in the parental BALB/cJ strain. This C to A variant was confirmed with Sanger sequencing and a TaqMan SNP genotyping assay. After this identification, the Msp3 mouse strain was kindly gifted to Dr. Niswander for further characterization. At this point, I took over the project and conducted the remaining experiments in this section. Due to the random nature of ENU mutagenesis, there is a possibility that more than one mutagenized locus exists in a narrowed critical interval and may be the causative mutation. A second unbiased sequencing and bioinformatics analysis was used to identify the location of the ENU induced mutation. At the time the reanalysis, mice were backcrossed for at least 10 generations to generate a largely congenic strain in which the genome is a mixed C57BL/6J;129SvlmJ background, with each generation selected for the NTD phenotype and the C to A transversion in the Smarcc1 gene in heterozygotes. Cranial tissue from wild type (WT) and mutant embryos was collected for RNA-Seq to analyze differential gene expression (discussed in Chapter IV) and to analyze the resulting data for a re-assessment of the ENU-induced mutation. RNA-Seq (Illumina Hi Seq 2000) was performed and the resulting gene expression data was analyzed for variants. Dr. Kathryn Gowan assisted with the variant analysis. Variants were filtered similar to the first analysis overlapping the linkage disequilibrium (LD) region between Mb on chromosome 9 (defined by the variant calling program used). Nineteen nonsynonymous SNPs were 44
57 identified in the LD region (Table III.1), but only two variants were novel and not present in the dbsnp as indicated by a lack of a reference SNP ID (rsid), Dusp7 nonframeshift deletion and the Smarcc1 C to A variant. Because the nonframeshift deletion in Dusp7 was not identified in the first set of variants, and was only present in the second set of variants, it was not considered as the causative mutation. These two independent genomic and exon sequencing analyses identify the causative ENU-induced mutation in Smarcc1, encoding BAF155, which we called Baf155 msp3. For the remainder of the thesis we refer to the mutant gene as Baf155 msp3 and the mutant protein as BAF155 msp3. This C to A substitution identified as the mutation in the Msp3 mouse genome is predicted to cause a threonine to lysine substitution at amino acid 416 of BAF155 (NP_033237; Figure III.1A) The missense mutation site and surrounding sequences are conserved between H. sapien, M. musculus, R. norvegicus and D. rerio, but not in S. cerevisiae (Figure III.1A) and this, coupled with the embryonic phenotypes in Baf155 msp/3msp3 mutants, indicates this residue is important for BAF155 protein function. Mutant Baf155 RNA and Protein Are Expressed, and the Protein Is Localized to the Nucleus Missense mutations within protein-coding genes may affect mrna stability, as well as protein stability or function. To understand the molecular consequences of the C to A missense mutation in the Baf155 gene, Dr. Jianfu Chen in the Niswander lab analyzed mrna and protein expression from E11.5 mutant and WT embryos. RT-PCR analysis showed the presence of Baf155 mrna in Baf155 msp/3msp3 embryos indicating that the transcript is not subject to nonsense-mediated decay (Figure III.1B). 45
58 46 Table III.1 Functional list of potential variants of the Msp3 phenotype. The causative Baf155 SNP is highlighted in yellow and does not have an rsid, indicating the variant has not been submitted to the database and therefore is a novel mutation. The nonframeshift deletion in Dusp7 does not have an rsid, but was not found in the first set of variants and therefore was not considered as the causative mutation. Function Gene chr start stop ref var rsid nonframeshift deletion Dusp7:NM_153459:exon chr GGC -. splicing Rpl29 chr A G rs splicing Abhd14a chr T C rs nonsynonymous SNV Parp3:NM_145619:exon5 chr A C rs nonsynonymous SNV Grm2:NM_ :exon4 chr T G rs nonsynonymous SNV Dock3:NM_153413:exon32 chr A T rs nonsynonymous SNV Celsr3:NM_080437:exon1 chr C G rs nonsynonymous SNV Celsr3:NM_080437:exon1 chr A G rs unknown UNKNOWN chr G A rs nonsynonymous SNV Col7a1:NM_007738:exon15 chr A G rs splicing Trex1 chr A G rs nonsynonymous SNV Atrip:NM_172774:exon12 chr C T rs nonsynonymous SNV Atrip:NM_172774:exon9 chr A G rs nonsynonymous SNV Atrip:NM_172774:exon8 chr G A rs nonsynonymous SNV Plxnb1:NM_172775:exon3 chr G A rs nonsynonymous SNV Cdc25a:NM_007658:exon5 chr C A rs nonsynonymous SNV Cdc25a:NM_007658:exon12 chr G A rs nonsynonymous SNV Dhx30:NM_ :exon8 chr G A rs nonsynonymous SNV Baf155/Smarcc1:NM_009211:exon13 chr C A. nonsynonymous SNV Cspg5:NM_013884:exon2:c.C950A chr C A rs nonsynonymous SNV Als2cl:NM_146228:exon27 chr T C rs nonsynonymous SNV Als2cl:NM_146228:exon27 chr A T rs
59 Figure III.1 BAF155 msp3 associates with other core BAF complex proteins. A) Schematic of the protein domain structure of BAF155. The msp3 ENU-induced mutation causes a C to A change at Chr9 bp (Build 36) in the Smarcc1 and leads to a threonine to lysine conversion of amino acid 416 in the BAF155 protein. This missense mutation alters an evolutionarily conserved amino acid in a conserved region, but not in a known structural domain. B) RT-PCR (top) and Western blot (bottom) analysis of Baf155 expression from E11.5 cranial tissue lysate. Gapdh and -Actin serve as internal standards. C) Confocal microscope images of neuroepithelial sections of E9.5 WT and mutant cranial neural tubes. Panels represent wildtype and mutant nuclei stained with antibodies against BAF155 (red) and BRG1 (green). Hoechst stains nuclei (blue). White scale bars=2 um. D) Immunoprecipitation of protein extracts from E11.5 mutant and wildtype embryos using anti-baf155 antibody followed by western blotting with anti- BRG1, anti-baf155, anti-baf170 and anti-baf60a. E) Schematic of the BAF complex proteins associated with the mutant BAF155 protein. BAF155 T416K associates with BRG1, BAF170 and BAF60a (IP), and BAF47 and BAF60a/b (yeast two-hybrid). BRG1, BAF170, BAF47 and BAF155 are considered core complex proteins. 47
60 Furthermore, Western blot analysis confirmed the presence of full length BAF155 in homozygous mutant embryos at similar levels to wild-type (Figure III.1B). Together, these data suggest that Baf155 mrna and protein stability are not compromised in the Baf155 mutant. The BAF155 protein contains four conserved domains- the Chromo, SWIRM, and SANT domains and a leucine zipper motif. These domains are predicted to allow BAF155 to bind to histone tail modifications, DNA and other proteins. Point mutations occurring in a conserved domain could disrupt the specific function of that domain by abolishing or changing intermolecular interactions, and the overall secondary structure of the protein. To understand the molecular consequences of this point mutation on the secondary structure of the mutant Baf155 msp3 protein, I used the protein structure analysis program Phyre (Protein Homology/analogY Recognition Engine) [148]. Phyre predicts secondary structure within a specific protein sequence based on the Structural Classification of Proteins (SCOP) database and the Protein Data Bank (PDB) of known protein structures. Using this data, the program can identify regions of order, indicating conserved secondary structure that may be a defined biological domain. The corresponding blocks of amino acids in input sequences that match predicted structures are tagged as intrinsically ordered, and have a low disordered number on a scale of 1-9 of disorder probability. Blocks of amino acids that do not have significant homology with other proteins in the databases containing known secondary structure would have a high disordered number. Using Phyre, BAF155 is not predicted to have a known conserved domain or region of homology in the 30 amino acid sequence surrounding the T416K substitution (Figure III.2). Further, this sequence is predicted by Phyre to have the highest probability of disorder, a disorder 48
61 probability of 9. These data suggest the affected amino acid is not within a conserved domain. Disordered regions within proteins are known to occur in regions of Protein- Protein or Protein-Nucleic Acid interactions as well as sites of posttranslational modification [149, 150]. This then suggests that the disordered region containing the affected amino acid may be involved in other protein-protein interactions, including transient associations or strong binding. However, the mutation is located N-terminal to the conserved SWIRM domain in the BAF155 protein, which could affect protein-protein interactions. Figure III.2 BAF155 is disordered in the region containing the Msp3 mutation Using the online secondary structure prediction tool Phyre, secondary structure of the BAF155 protein was analyzed. The mutated amino acid is T416, denoted by the purple arrow and box. The region containing the Mps3 mutation is in an intrinsically disordered region of the protein with a disorder probability score of 9 out of 9. Immediately to the right of the disordered region is the intrinsically ordered SWIRM domain, denoted by a disordered probability between 0-3. The Msp3 phenotypes in the Baf155 msp3/msp3 embryos might be explained by a misfunctional protein or complex. The BAF complex is normally localized to the nucleus [134]. Mutations in the BAF155 nuclear localization signal (NLS) result in incorrect cytoplasmic localization of BAF155, BAF47 and the ATPase BRG1, indicating that BAF155 maintains the subcellular localization of the BAF complex in the nucleus [146]. To test the possibility that subcellular localization of BAF155 and BRG1 may be compromised in the Baf155 msp3/msp3 mutants, we examined their localization in cryosections 49
62 from wildtype and mutant embryos. In Baf155 msp3/msp3 mutants, BRG1 and BAF155 are detected in the nucleus at high levels and are not detected in the cytoplasm, comparable to wildtype (Figure III.1C). This suggests that subcellular mislocalization of BAF155 or other BAF proteins does not account for the phenotype seen in Baf155 msp3/msp3 mutants. BAF155 msp3 Mutant Protein Can Still Associate with the Core Remodeling Complex BAF155, along with ATPase BRG1, BAF170, and BAF47, are considered core components in the BAF remodeling complex because they are sufficient to remodel nucleosomes in vitro at the same efficiency as the fully intact BAF remodeling complex. BAF155 interacts directly with two core complex proteins (BRG1 and BAF47) in the SANT and SWIRM domain [146]. It also interacts with BAF60a in the region that encompasses the BAF155 msp3 missense mutation, T416K [146]. To test the possibility that the interaction between BAF155 and the BAF core complex is disrupted by the T416K mutation, Dr. Jianfu Chen in the lab performed co-immunoprecipitation assays using E11.5 mutant and WT embryos. These studies revealed that BRG1, BAF170 and BAF60a still associate with BAF155 msp3 in vivo and in a similar ratio as compared to the WT, despite the amino acid change (Figure III.1D). Additionally, yeast two-hybrid analyses suggest that BAF155 msp3 can still interact directly with BAF47, BAF60a and BAF60b in vitro (Figure III.3). These experiments indicate that the mutant BAF155 msp3 protein still interacts with the core components of the BAF remodeling complex and suggest the mutant protein could interact with other BAF complex proteins in vivo as well (summarized in Figure III.1E). These data suggest that the BAF155 msp3 mutant protein is present and can interact molecularly in the remodeling complex. 50
63 Figure III.3 BAF155 msp3 interacts with BAF60a, BAF60b and BAF47. Yeast two-hybrid assay. Full length cdna of Baf155 and Baf155 msp3 were cloned into pdest22 (Binding domain, BD), and Baf60a, Baf60b, and Baf47 were cloned into pdest32 (Activation domain, AD) vectors for the yeast two-hybrid assay. Reciprocal experiments were done in which the Binding and Activation domains were switched. The transformed cultures from the indicated strains were streaked onto synthetic complete medium plates lacking leucine, tryptophan, and histidine and containing 0.5 mm 3-amino- 1,2,4-triazole (-His +0.5mM AT), or the cultures were streaked onto SC-Trp-Leu (+His) plates. The bottom panels represent negative controls using some of the single plasmid strains. The plates were incubated at 32 C for 2-4 days for interaction analysis. These protein interaction data, as well as the longer survival of Baf155 msp3/msp3 embryos relative to Baf155 null/null embryos, suggest that the BAF155 msp3 mutant protein is present and can interact molecularly in the remodeling complex, but its function may be partially compromised. Thus, it appears that Baf155 msp3 is a missense allele and the mouse may exhibit a hypomorphic phenotype. 51
64 Cell Proliferation Is Decreased and Cell Death Is Increased in Baf155 msp3/msp3 Neural Progenitor Cells NTDs can result when the spatiotemporal control of neural tube closure is disrupted [9] due to any of several different mechanisms including: reduced neural progenitor cell proliferation, changes in cell fate, decreased cell survival, or defects in patterning. To investigate if the NTD might arise because of improper patterning, we examined a battery of molecular markers that reflect the dorsal-ventral or anterior-posterior axis patterning of E9.5 embryos, just after closure of the neural tube. Antibody staining and RNA in situ hybridization showed no change in the expression pattern of Shh, PAX6, PAX3, Fgf8 and FoxG1 in mutant neural tubes when compared to wildtype somite matched embryos (Figure III.4A-F). Neural crest cells are specified during neural tube closure and abnormal expression of neural crest genes could also reflect altered patterning [151]. Therefore, we examined markers characteristic of early neural crest specification including p75ntr, Tfap2a, and Sox10 (Figure III.4G-I) and again found no change in the expression pattern compared to WT. These results suggest that neural patterning is not disrupted in the Baf155 msp3/msp3 mutant embryos. The neural tissue grows significantly during neural tube closure by proliferation of neural progenitor cells. The tight control of cell proliferation is necessary for the neural folds to meet properly at the dorsal midline. At E9.5, the vast majority of the neural ectoderm is comprised of proliferating neural progenitor cells. Too much or too little cell proliferation can abrogate neural tube closure. Previous studies noted an increase in cell proliferation in the telencephalic striatum region of exencephalic hemizygous Baf155 +/null 52
65 embryos at E13.5 [135]. Thus, we asked if the proliferation rate might be altered in Baf155 msp3/msp3 mutant embryos at the time of neural tube closure. Figure III.4 BAF155 function is not necessary for early neural patterning. A-C) Transverse sections of E9.5 embryos followed by immunofluorescence for dorsal/ventral markers Sonic Hedgehog (Shh) (A), PAX6 (B) and PAX3 (C). D and E) Lateral and F) frontal views of whole mount RNA in situ hybridization on E10.0 WT and Baf155 msp3/msp3 mutant embryos with probes to anterior/posterior markers Fgf8 (D) and FoxG1 (E and F). G) Confocal images of transverse sections of E10.5 WT and mutant embryos incubated with anti-p75 ntr antibody to visualize the enteric neural crest cells. Hoechst stains nuclei. H) Transverse sections of E10.5 rostral spinal cords hybridized with TFAP2α probe marking early migrating neural crest cells. I) Lateral view of E10.5 WT and mutant embryos hybridized with Sox10 probe to mark early neural crest cells. White scale bars=100um, Red scale bars= 1mm The mitotic rate of neuroepithelial cells in somite-matched E9.5 Baf155 +/+ and mutant Baf155 msp3/msp3 embryos was compared using phospho-histone 3 (ph3). In the hindbrain, there was a statistically significant decrease in the mitotic index (ph3+ cells/1000 neuroepithelial cells) of Baf155 msp3/msp3 mutants compared to WT (20±9 vs 71±10) (Figure III.5A,D). We also quantified the number of cells in S phase by performing bromodeoxyuridine (BrdU) incorporation experiments (Figure III.5A,E). 53
66 There was no apparent difference in number of cells in S phase in mutant compared to WT neural tubes at E9.5. These studies suggest the cells undergo DNA replication, but there may be a mitosis defect in the neuroepithelium. These findings do not fit with traditional ideas about cell cycle and will be discussed in the conclusions of this chapter. A decrease in cell survival within the neural tissue may result in too few cells to allow proper apposition and fusion of the neural folds, resulting in a NTD. To determine if neural progenitor cell survival was affected in mutant embryos we used a TUNEL assay to visualize apoptotic cells in somite-matched E9.5 WT and mutant neural epithelium. This showed a statistically significant increase in TUNEL positive cells per 1000 nuclei in the hindbrain at E9.5 in the Baf155 msp3/msp3 mutant compared to WT (7.2±1.0 vs 3.7±0.5) (Figure III.5B,F). Together, these studies suggest that BAF155 function is necessary to maintain the normal rates of proliferation and cell survival in the neuroepithelium. As neural precursors begin to differentiate starting around E10.5, they exit the cell cycle and leave the ventricular zone to reside at the periphery of the neural tube. If differentiation does not occur at the correct time, this can contribute to an excess or a deficiency in cell number for effective neural fold juxtaposition and ultimate neural tube closure. To determine whether or not neural differentiation was compromised in the homozygous mutants we evaluated Tuj1 expression, a known neural differentiation marker, in the spinal cord and hindbrain of E10.5 somite matched Baf155 +/+ and Baf155 msp3/msp3 embryos. Mutant and WT embryos exhibit the same characteristic expression pattern of Tuj1 (Figure III.5C) and therefore neural differentiation does not appear to be affected in the homozygous mutant. 54
67 Figure III.5 BAF155 function is necessary to maintain proliferation and cell survival. A) Transverse sections of E9.5 hindbrains of WT and Baf155 msp3 mutant embryos followed by immunofluorescence for proliferation marker p-h3 (green), S phase marker BrdU (½ hr BrdU incorporation) (red) and merged image with Hoechst stained nuclei. B) Transverse sections of E9.5 hindbrains in WT and Mutant embryos to visualize apoptotic cells by TUNEL staining. C) Confocal images of transverse sections through rostral spinal cords. Anti-Tuj1 antibody was used to visualize neural differentiation in neuroepithelial cells. D) Quantification of anti-phospho-h3 positive cells in WT and Mutant hindbrain neural tube. E) Quantification of anti-brdu positive cells in WT and Mutant hindbrain neural tube. F) Quantification of apoptosis by TUNEL staining in WT and Mutant hindbrain neural tube. Ratio of p-h3, BrdU or TUNEL positive cells were calculated by dividing the number of positive stained cells by the total number of neuroepithelial cells, indicated by Hoechst staining, then multiplying by Error bars indicate SEM of at least 3 sections from 3 biological replicates. (*) P<0.01, Student s t-test. The neural tube is outlined by yellow or white dots. 55
68 CONCLUSION While it is known that BAF155 is important during early development, it has been difficult to study the function of the protein due to the early lethality of Baf155 null/null mouse embryos. Twenty percent of BAF155 heterozygous embryos exhibit exencephaly with increased cranial proliferation observed 4 days after the time of neural tube closure [135]; however, the role of BAF155 has not been determined during the time of neural tube closure. Two separate analyses of the mouse line exhibiting the Msp3 phenotype have identified the causative mutation of the Msp3 phenotype as a missense mutation in the 13 th exon of the Baf155 gene. The gene and protein are expressed, and the protein is properly localized to the nucleus in the mutant embryos. Molecular integrity of protein complexes is essential for their function. Overall, these data show the BAF complex isolated from the Baf155 msp3/msp3 embryos is stable and able to associate with the core complex members. This allele represents a unique tool to investigate the function of BAF155 as well as the BAF complex in a biological context. This chapter provides a cellular and molecular characterization of the novel SNP T416K in the Baf155 msp3/msp3 mouse. These data indicate that while there is no change in patterning, there is an increase in apoptosis and a decrease in proliferation in mutant neuroepithelial cells at the time of neural tube closure. The BAF complex clearly has a role in remodeling nucleosomes in genomic chromatin but it also can interact with many partners, so the BAF complex likely functions in a broader context as well. One possibility is that the BAF155 msp3 mutation disrupts the interaction of BAF155 with transcription factors or chromatin maintenance proteins. For example, it has been suggested that intrinsic disorder within the BAF protein structures may explain diverse roles of chromatin remodeling proteins [152]. The point mutation 56
69 within our allele occurs in an intrinsically disordered region based on Phyre analysis (Figure III.2), suggesting this region may be involved in other protein-protein interactions, including transient associations or strong binding. However, I observed no difference between the mutant and WT protein for association to BAF60a and BAF60b, which directly interacts within the region of the BAF155 T416K mutation. It remains possible that another protein may interact within this disordered region and disruption of this interaction may lead to NTDs. An experiment to test which proteins are not able to associate with BAF155 would be to perform IP-Mass Spec in mutant and WT samples. I would expect fewer proteins to associate with the BAF155 msp3 protein than with the WT BAF155 protein, suggesting a loss of function in the mutant protein. The currently defined BAF proteins associated with the BAF complex do not account for the large, 2 Mb size of the complex when purified. This indicates that peripherally associated complex members or transiently associated transcription factors must also associate with the complex. Because I have shown that the core BAF proteins associate in the mutant complex, I would expect there is a change in association with some non-baf proteins in the mutant BAF complex. However, it remains possible that other identified BAF complex members may not associate with the mutant BAF complex. I tried to examine this possibility by testing BAF45C, BAF57, BAF180, BAF250A (ARID1A), and BAF250B (ARID1B) in immunoprecipitation experiments but antibodies against these proteins did not detect the mouse proteins. It is also possible that the BAF155 msp3 mutation does not disrupt the protein-protein associations, but may instead create a gain or loss of function in the mutant BAF complex. This function could be tested with various biochemical assays. Because the BAF complex utilizes ATP for its functions, an ATP 57
70 Assay could be conducted using a colorimetric ATP Assay Kit (Abcam, cat #ab83355). The WT and mutant complex could be isolated from MEFs using a tagged BAF155 protein. The rate of ATP usage could be measured colorimetrically. The WT and mutant ATP activity can then be compared. As another possibility, the point mutation in the complex could affect the dissociation rate of the complex. Crabtree and colleagues have noted the BAF complex can interchange its subunits as cells undergo differentiation, particularly in the switch from neural progenitors to neurons [133]. It is possible that the association rate and/or dissociation rate of the complex may be altered, resulting in a change of function. This rate is difficult to test, but may explain a change in function between the mutant BAF complex and the WT complex. Additional potential assays to test function will be addressed in Chapter V. Cell cycle control is critical for proper neural tube closure. In the BAF155 msp3/msp3 mouse mutants, we found that the number of M-phase cells was decreased, but the number of S-phase cells measured by BrdU incorporation was not statistically different in the neuroepithelium compared to WT. These data are not consistent with the traditional model of cell cycle progression. In the traditional model, it would be expected that if the number of cells in M-phase were decreased, then the number of cells in S-phase would also be decreased because the total number of cells progressing through the cell cycle would be diminished. To explain this discordance between what was observed and what was expected, it is possible that the cells are arresting in G2/M, and thus not progressing through the cell cycle to reach M-phase again. An ideal experiment to test where in the cell cycle the neuroepithelial cells are slowing would be a propidium iodide staining of DNA to test DNA content within the cells by flow cytometry. An arrest in G2/M in the mutant cells 58
71 compared with a normal progression in the WT cells might explain how similar number cells can progress through S phase in the mutant vs. WT, but that fewer cells might progress through mitosis. Because of the small number of cells in the neuroepithelium, flow cytometry analysis might not be possible. Another experiment to test the cell cycle progression and ultimately cell cycle exit in the neuroepithelium would be to stain sections with other cell cycle progression markers including Ki-67. Ki-67 marks cells in the proliferation phase, as opposed to cells that have exited the cell cycle, although its exact mechanism is not well understood [153]. Comparing the number of cells in the proliferation phase may suggest a defect cell cycle progression. It is also possible that some cells are not exiting the cell cycle properly. To test this, animals are pulsed with BrdU for 24 hours, embryos are fixed and sectioned, and resulting sections are stained with anti-brdu and anti-ki67 antibodies. Cells that are not co-labeled with Ki67 and BrdU suggest those cells have exited the cell cycle. These studies would provide evidence that some neuroepithelial progenitor cells may be not exiting the cell cycle properly, or may be arresting in the cell cycle. Proper control of proliferation and cell survival are critical for neural tube closure. In the Baf155 msp3/msp3 mouse mutants we identified both a decrease in cell proliferation and an increase in apoptosis. This would significantly reduce neural ectoderm number and this may explain the underlying defect in neural tube closure. It is possible that this regulation of cell number by proliferation and apoptosis may occur by a single common mechanism. Two possible scenarios are outlined below. There is no evidence in the literature that BAF155 may regulate both cell death and proliferation, but recently the BAF complex has been suggested to regulate cell cycle progression in cancer cells [154]. This study suggests 59
72 that the BAF complex can affect both cell cycle progression and cell death via the association with Topoisomerase II and regulation of the progression through G2/M checkpoint [154]. Another possible link between apoptosis and proliferation could be due to BAF-mediated transcription of centrosomal genes, whose inhibition can lead to cell cycle defects and cell death [155]. For example, inhibition of the centrosome protein Nde1 was shown to delay cell cycle reentry in cell lines, and to reduce mitosis in zebrafish [156]. Specifically in the Baf155 msp3/msp3 mutant, there is decreased expression of the centrosomal gene Poc1a, which may lead to the observed decreased cell proliferation and survival phenotype. Here are just two possible mechanisms that may control cell survival in the Baf155 msp3/msp3 homozygous mutant. It is also possible that the observed defects in cell death and cell proliferation may occur independently of one another. BAF155 can be phosphorylated by and interact with Cyclin E [157] in the Rb pathway as well as AKT [158] in the PI3K/AKT pathway. However, we did not detect misregulation of Rb or PI3K pathway genes in the mutant by RNA-Seq analysis (discussed in Chapter IV), suggesting these signaling pathways are not disrupted and BAF155 may play other transcriptional roles in controlling proliferation. While many genes known to control proliferation were misregulated in the mutant cranial tissue, no specific cyclins or cell cycle genes were differentially expressed. The RNA-Seq studies in chapter IV also revealed several misregulated genes that contribute to cell survival, including the downregulated genes Abl1and ApoB, and the upregulated genes Adora3, Fam72a, SST, and Map3k12. Mouse embryos mutant for Apolipoprotein B (ApoB) exhibit exencephaly (30% incidence) and excessive cell death in the hindbrain [159]. Knock-out of Abl1 kinase in combination with Abl related gene (ARG) results in 60
73 NTDs and disruption of the actin cytoskeleton of the neuroepithelium [160]. Although the mechanism underlying the increased apoptosis in Baf155 msp3/msp3 mutants is unclear, altered expression of these genes may contribute to the increase of cell death in the neural epithelium of the Baf155 msp3/msp3 mutant. Together these data suggest that BAF155 plays a role cell proliferation and cell death via transcriptional control, and these may be common or separate processes controlled by BAF155. Limitations to these experiments include the difficulty of obtaining adequate quantities of neuroepithelial cells to conduct flow cell cycle analysis experiments, addressed above. Additionally, it would be interesting to determine whether cells that do not progress through the cell cycle properly would later undergo apoptosis. To test this, it would be ideal to conduct live imaging experiments of cells in the ventricular zone and within the neural tube. However a significant limitation of the mouse model system is that these imaging experiments cannot be conducted over the period of time required to capture the cell cycle defect through the point where cells may undergo apoptosis. MATERIALS AND METHODS Analysis of Mutant Phenotype Embryos were fixed in 4% paraformaldehyde (PFA) in PBS and processed for whole mount or cryosection (8 m) in situ hybridization. Whole-mount and section RNA in situ hybridizations were performed as described (Holmes & Niswander 2001) with digoxygenin -labeled antisense riboprobes. In conducting the whole mount in situ protocol, E9.5 embryos were digested with PK for 5 minutes, E10.5 embryos were digested for 7 minutes. For both sections and whole mount, the probes were hybridized with the tissue at 65 C overnight. Sections were imaged on a Nikon Eclipse 80i microscope. For 61
74 immunohistochemistry, embryos were dissected and fixed in 4% PFA in PBS, washed 3 times with PBS, cryopreserved in 30% sucrose, embedded in OCT and sectioned (8 m). For cell cycle analysis, BrdU (50 mg/kg) in PBS was injected intraperitoneally into pregnant dams 30 minutes prior to euthanasia and embryo processing. For apoptosis, the Apop-tag In Situ Apoptosis Detection Kit (Millipore, S7160) was used following the protocol to indirectly detect apoptotic cells by the TUNEL method. For immunostaining, tissue sections were treated with PBS/3% goat serum for 1 hour at room temperature. Primary antibody was added and incubated at 4 C overnight. For detection, Alexaconjugated secondary antibodies (Molecular Probes) were incubated for 1 hour at room temperature. Hoechst was added along with the secondary antibodies as a nuclear stain. Stained sections were imaged with Zeiss LSM510 META confocal microscope. Cells were counted using the image J cell counter application. The total Hoechst stained nuclei were divided by the positive nuclei stained with Ph3 or TUNEL staining. CONCLUSION While NTDs in mice have been studied using primarily a single gene approach, epigenetic proteins can alter the expression of many genes in a given biological context, and may inform our understanding of NTDs in both mice and humans. RNA-Seq is a robust method to measure global gene expression in biological samples. The RNA-Seq studies conducted here on E9.5 mutant and WT cranial tissue show that gene expression in the Baf155 msp3/ msp3 embryos is mostly upregulated, and that mutant tissue exhibits extensive variability in gene expression compared to the WT. The individual genes that are misregulated cluster into the categories of neural differentiation, cell death and apoptosis. 62
75 These studies suggest that the defects seen in the mutant Baf155 msp3/msp3 embryos can be partially accounted for by changes in transcriptional control. The BAF complex can act as both a repressor and activator of gene expression, and has been found to refine the gene transcriptional network in ES cells to maintain pluripotency [128, 147]. Using RNA-Seq, I analyzed transcriptional control in the cranial tissue of the mutant embryo compared to WT. Most of the differentially expressed genes in the Baf155 msp3 mutant embryo are upregulated, including neural development genes discussed below, suggesting that BAF155 and the BAF complex largely play a repressive role during neurulation. Consistent with the expectation that the BAF complex provides more global, rather than gene-specific, transcriptional regulation, we did not find strong single gene candidates to explain the underlying mechanism of the NTD. Instead, we observed a broader dysregulation in expression of genes and gene families that are associated with NTDs when disrupted. While the fold change in gene expression was not particularly striking across all the mutants, it is not unreasonable to suggest that small gene expression changes in combination may result in an NTD. As previously noted, neural tube closure is a highly dynamic process that requires precise coordination of complex cellular functions to proceed to completion. Moreover, many of the misregulated genes in Table C.1 are associated with NTDs: mutations in Abl [160] and Frem2 [162, 163], as well as other genes discussed below result in NTDs. Interaction with specific genomic targets that regulate the developmental program of neural tube closure may be disrupted in the BAF155 msp3/ msp3 mutant embryos. In ES cells, BAF155 is bound to promoter or genic regions of ~5,630 distinct genes [147]. Moreover, esbaf facilitates Stat3 genomic binding, and can act either synergistically or 63
76 antagonistically with the Polycomb Repressive Complex Group (PcG) in ES cells [127]. In HeLa cells, BAF proteins associate with up to 77% of genic DNA [164]. Given the potentially large set of genomic targets, there are surprisingly few genes misregulated overall in Baf155 mps3/ msp3 mutant cranial tissue at E9.5, especially considering that at least some of these misexpressed genes are likely indirect targets. There is also very little overlap in the datasets of differentially expressed genes between our point mutant and knock-out/knock-down of Brg1 or other BAF members in ES cells [147], neural progenitors or neurons [132]. This lack of overlap likely reflects the differences between mutant alleles: knock-down of one component presumably results in incorrect ratios of complex proteins, whereas our point mutant does not apparently change the levels of complex proteins and the complex is largely intact. Thus, the point mutation may be expected to reveal more specific targets relative to neural tube closure. BAF subunits are heterogeneously expressed and assembled in various cell types and these different complexes can have distinct functions. BAF subunit mutations have been reported in many human neurologic diseases including Autism, Schizophrenia, Coffin-Siris syndrome, sporadic mental retardation [120], however these neurological diseases are not accompanied by NTDs according to the Online Mendelian inheritance of Man database (OMIM). Different protein assemblies of the BAF complex exist in the developing central nervous system, including the neural progenitor BAF complex (npbaf) and neuronal BAF complex (nbaf) [132]. These have specific functions in the regulation of proliferation and differentiation of mammalian neural stem cells. Later loss of BRG1 function in neurons (Brg1fl/fl;Nestin-Cre) and transcriptional analysis of E12.5 mutant brains revealed an upregulation of genes within the Notch and Shh pathways. While it is 64
77 believed that BAF155 is present in BAF complexes in all cell types, we did not see gene expression changes in the Baf155 msp3/msp3 mutant at E9.5 that might reflect alterations in Notch or Shh signaling. This difference in gene control between the Brg1fl/fl;Nestin-Cre and the Baf155 msp3/ msp3 embryos could reflect temporal differences in BAF complex function during brain development. Many of the upregulated genes in Baf155 msp3/msp3 mutants at the time of neural tube closure contribute to neural development, including Spon1, Doc3, Topb2, Map3k12, Cdk18, Sema3c, Sst, Igta4, Epha3, Fzd2, Fzd3, Ablim3, and Plcg2. While no differentiation defect was observed in the Baf155 msp3/ msp3 mutants at and shortly after the time of neural tube closure, the modest misregulation of many neural development genes may contribute to the NTD phenotype. Particularly interesting are the genes associated with NTDs, namely the Wnt receptors Fzd2 and Fzd3. The Wnt/Fzd (PCP) pathway controls aspects of neural development, including neural tube closure, in mice and humans [165]. Although the Fzd2 -/- mouse does not show NTD, in a double heterozygous cross with another PCP component, Fzd +/- :Vangl Lp/+, fully penetrant cranial NTDs are observed [166]. Also, Fzd3 -/- ; Fzd6 -/- mutants present with the most severe NTD, craniorachischisis [167]. This supports the idea that NTDs can arise due to aberrant expression of multiple risk alleles. As more data become available from global sequencing studies at the time of neural tube closure, ultimately a gene interaction network can be developed. Because statistically significant genes found in my studies showed little overlap with genes found in other transcriptional analysis studies of the BAF complex at different times and different tissues, my data suggest it is necessary to carefully select the appropriate time point and tissue type to identify genes that are temporally disrupted. Here we have selected cranial tissue at the 65
78 time of neural tube closure. Mutations in over 200 individual genes have been discovered to show NTDs, and the function of those individual genes identified; however the genetic networks remain poorly understood. So it will be informative to identify and then compare the gene expression profiles in mouse models that carry mutations in global regulators as well as specific transcription factors, cilia related genes, and signaling pathways implicated in NTDs such as PCP and Shh. From these data, the NTD community can identify whether there are specific genes that are commonly misregulated in mice exhibiting NTD. This information should help to inform studies of human NTD cases. My work also suggests that greater variability in overall gene expression may itself contribute to NTD incidence, as has been suggested by Kappen and Salbaum (submitted, 2013). Gene variability will be examined in greater depth in Chapter V of this thesis. Limitations to the RNA-Seq data collected include both technical and experimental. Inconsistencies between the samples could have arisen during tissue processing, library preparation, and mechanical differences during collection of the RNA-Seq data. The current RNA-Seq technology relies on synthesizing cdna from the transcripts, and an inherent efficiency bias can be introduced by the RNA structure [168]. Another source of bias is that selection of RNA transcripts was completed using an Oligo dt column, which only selects for transcripts with a poly A tail. This eliminates all transcripts without a polya tail, including long non-coding RNAs. Only 3 WT and 3 mutant BAF155 msp3/mpp3 embryos were collected, which is a low N value. If more embryos were collected, the gene expression data could reveal a decrease in variability, or show that the variability seen was well representative of the entire population. The embryos were matched based on somite 66
79 number and morphology in order to try to minimize variability in gene expression due to differences in the age and phenotype of the embryos. Antibodies Used Antibodies against the following proteins were used: TUJ1 (Covance, MMS425P, 1:500), phospho-histone H3 (Cell Signaling, 9701, 1:200), BrdU (Novus Biologicals, NB , 1:200), p75 NTR (Promega, G3231, 1:100), BRG1 (Santa Cruz, SC :200), BAF155 (Santa Cruz, SC-10756, 1:200), BAF170, (Santa Cruz, SC17838, 1:200) and BAF60a (BD Transduction Laboratories, , 1:500). Antibodies from the Developmental Studies Hybridoma Bank were used at a 1:10 dilution including PAX3 and PAX6. Yeast Two-Hybrid Assay Protocol Full length cdna of Baf155 and Baf155 msp3 were cloned into pdest22 (Binding domain, BD), and Baf60a, Baf60b, and Baf47 were cloned into pdest32 (Activation domain, AD) vectors for the yeast two-hybrid assay. Reciprocal experiments were done in which the Binding and Activation domains were switched. The transformed cultures from the indicated strains were streaked onto synthetic complete medium plates lacking leucine, tryptophan, and histidine and containing 0.5 mm 3-amino-1,2,4-triazole (-His +0.5mM AT), or the cultures were streaked onto SC-Trp-Leu (+His) plates. Immunoprecipitation Assay and Western Blots Embryonic day E11.5 embryos were placed in lysis buffer (50 mm Tris-HCl at ph7.4, 150 mm NaCl, 1 mm EDTA, 1% Triton X-100, including 1 tablet protease inhibitor (Roche Complete) per 10 mls lysate) and incubated on ice for 10 min. Tissue was sonicated and tissue debris was pelleted at 12,500 rpm for 10 min at 4 C, and the 67
80 supernatant was incubated with primary antibodies overnight at 4 C. The lysates were incubated with Protein A or G Sepharose beads for 2 h, followed by washing of the immunoprecipitates three times with lysis buffer and elution of bound proteins in SDS- PAGE sampling buffer for 10 min at 100 C. Western blots were performed as described previously (Chen et al. 2005) using primary antibodies as listed. Secondary antibodies used were goat anti-rabbit ( , Bio-Rad), and goat anti-mouse ( , Bio-Rad). 68
81 CHAPTER IV GENE EXPRESSION ANALYSIS OF Baf155 msp3/ msp3 EMBRYOS SHOWS GENE EXPRESSION VARIABILITY AND OVERALL UPREGULATION OF GENES INTRODUCTION Neural tube defects are likely the result of complex multigenic dysfunction; however, most of the NTD studies in both mouse and human focus on analysis of a mutation in a single gene or a few genes. Epigenetic regulators alter the expression of hundreds of genes in a given biological context. Mutations in some of these regulators have been linked to neural tube defects in mice. Of the >200 mouse NTD mutants, ~5% contain disruptions in genes encoding epigenetic regulators. Mutations in two of the BAF chromatin remodeling complex subunits show neural tube defects, but these NTDs have not been characterized in detail at the time of neural tube closure. Global gene expression analysis, especially of an epigenetic regulator that could have wide-spread effects, would greatly enhance an understanding of the multigenic nature of NTDs [35]. RNA-Seq analysis is an unbiased method to discover global transcriptional differences between biological samples [161]. This technology allows a comparison of transcripts in any number of experimental conditions including WT versus mutant, or treated versus untreated biological samples. Several million short sequencing reads are generated per sample. The reads are mapped back to the genome bioinformatically to identify quantitative levels of all genes expressed under the experimental and control conditions. These results are then analyzed to discover genes that are differentially expressed and may be involved in a given biological process. 69
82 The BAF155 msp3 mutant protein contributes to the formation of an intact BAF complex and the mutant embryos show proliferation and apoptosis defects in the developing neural tube. We predict that, even though the complex is intact, the complex does not function normally and global gene expression will be different in mutant embryos compared to wild type. Therefore, we conducted RNA-Seq experiments to analyze the differential gene regulation in cranial tissue from mutant and WT to help elucidate the differences in transcriptional control that may help to explain the neural tube defect. RESULTS Gene Expression Levels Are More Variable in Baf155 msp3 Mutant Cranial Tissue: Averaged Sample Analysis BAF155, as a component of a chromatin remodeling complex, is thought to be involved in the regulation of gene expression. BAF155 knockdown in ES cells results in misregulation of hundreds of genes [147]. The Baf155 msp3 allele presents a unique case for gene expression analysis wherein the BAF complex appears to be intact, in contrast to disruption of the BAF complex when BAF155 function is lost [145, 146]. To examine gene expression in the Baf155 msp3/ msp3 mutant at the time of neural tube closure, I performed RNA-Seq and compared gene expression levels between homozygous WT and homozygous mutant embryos. RNA was isolated from somite matched (21-23 somites) and phenotype matched cranial tissue of homozygous wildtype and homozygous mutant embryos (3 of each genotype) at E9.5 just after the time of neural tube closure. At E9.5, the vast majority of the embryonic neural tube is comprised of proliferating neural progenitor cells, constituting a relatively homogeneous cell population. Our collaborator, Dr. Michael Salbaum from the Pennington Biomedical Research Center in Baton Rouge, Louisiana, 70
83 conducted extensive analysis of the RNA-Seq data comparing all 3 mutants to all 3 WT replicates, and identified 78 genes as being upregulated on average in the mutant samples while 22 genes were downregulated. The full list of differentially expressed genes in all 3 mutant replicates collectively, as well as p-values, FDR and fold change, is shown in Table C.1. To facilitate an understanding of global gene expression differences, I used Ingenuity Pathway Analysis (IPA) to assign functional categories to the up- and downregulated genes in the mutants. The most significant molecular and cellular functions of these genes were: 1. cellular movement (1.46E-04>p>4.12E-02), 2. Cell death and survival (2.69E-04>p>4.12E-02), and 3. Cell growth and proliferation (2.69E-04>p>3.41E-02). Many neuronal genes were significantly misregulated as well. Gene Expression Levels Are More Variable in Baf155 msp3 Mutant Cranial Tissue: Individual Sample Analysis This analysis was done in collaboration with Dr. Michael Salbaum. We used multidimensional scaling (MDS) analysis to conduct an unsupervised examination of the relationship between the global gene expression in each individual mutant and the wildtype cranial tissue samples. As seen in Figure IV.1A, a relationship between the genotypic subtype and gene expression was clearly observed. The wildtype replicates were tightly grouped in the MDS plot and showed little dimensional variance. In contrast, the mutant replicates did not group together and showed increased variance in both dimensions 1 and 2. Overall, there was a trend towards higher variance of gene expression in the mutant samples compared to the wildtype samples, leading to a more dispersed MDS plot where the mutants do not cluster with the wildtype controls and each mutant does not cluster with the other mutants. 71
84 Figure IV.1 Variable gene expression in Baf155 msp3/msp3 mutant cranial tissue. A) A multidimensional scaling (MDS) plot of differentially expressed genes from cranial tissue of three somite matched WT and three somite matched Baf155 msp3/msp3 mutants. B) Genes were ranked in ascending fashion according to their coefficient of variation observed in wild type samples, and for each gene, the distance from mean was plotted as the log2 of the ratio of the expression level of a gene in an individual sample normalized to the mean of the expression level of the respective gene in the wildtype samples. Top: All three wild type samples; bottom: all three mutant samples; right: individual samples, from top: wild type 1 (blue), wild type 2 (red), wild type 3(green), mutant 1 (purple), mutant 2 (turquoise), mutant 3 (orange). Based on the lack of clustering between the individual mutants when global gene expression was compared, the averaged gene expression analysis did not illustrate the nuances of gene expression in the Baf155 msp3/ msp3 mutant samples. Therefore, in a more exploratory analysis, we asked if the observed transcriptional changes were uniformly 72
85 distributed in the 3 biological replicates between the homozygous mutant group and compared to the homozygous wildtype group. For each sample, we established for each gene a distance from mean parameter by normalizing the expression level of any given gene to the arithmetic mean of the expression of the respective gene in the wild type samples; this ratio underwent logarithmic transformation to obtain the distance value. All genes were ranked by their coefficient of variation in the wildtype samples, and the distance value was plotted as shown in Figure IV.1B (wildtype in top panel, mutants in bottom panel). The average wildtype plot showed a relatively tight spacing of all data points in a 'trumpet' shape. This coincides well with the low degree of variation we identified in the differential expression test. The same plot for the mutant samples showed a larger spread of gene expression ratios, and mutant replicate 2 (Mut2 in turquoise) seems to have the greatest difference in the ratio of gene expression compared to both remaining mutants and wild type controls. Again, this correlates well with the differential expression tests. To identify whether there are groups of genes that may contribute to this gene expression variability more than others, we generated three separate mutant differential gene expression lists by directly comparing each mutant gene count to the average of the wildtype gene counts. This analysis shows collections of genes from each individual mutant as opposed to the genes misregulated in all of the mutants. Over 2149 genes were differentially expressed in Mutant replicate 2 (Mut2) as opposed to 188 genes in Mutant replicate 1 (Mut1) or 215 genes in Mutant replicate 3 (Mut3) (based on DESeq testing, and counting genes with padj<0.05). When we directly compared these three differentially expressed gene lists, we identified a subset of 27 genes that was significantly misregulated 73
86 in all of the mutants (Table D.1). Interestingly, many of the genes on this list do not overlap with Table C.1, comparing all mutant samples to all WT samples, because there is often one gene outlier that is not significantly misregulated in each individual. Pathway analysis using IPA suggests that no canonical pathways are shared between these 27 genes, but 6 of these genes are predicted to be within a nucleic acid metabolism network. Overall, these results suggest that gene expression in the mutants is more varied than in the wildtype samples, and there is little overlap in the functionality of the most variable genes between individuals in the Baf155 msp3/ msp3 mutant group. MATERIALS AND METHODS RNA-Seq Library Preparation and Analysis Three somite matched E9.5 embryos were collected of both Baf155 +/+ and Baf155 Msp3/Msp3 alleles. Cranial tissue above the pharyngeal arches and otic vesicles was harvested and stored at -80 C until all samples were collected. RNA was isolated using the RNeasy mini kit from Qiagen (74106). On average, 1400 ng of total RNA was obtained per sample. RNA concentration and A260/A280 ratio was analyzed using the Nanodrop ND-1000 spectrophotometer (Thermo Fisher Scientific). Libraries were made using the Illumina TruSeq RNA Sample Preparation Kit V2 (RS ). Libraries were analyzed on the 2100 Bioanalyzer (Agilent Technologies, Inc) for proper size and integrity. For high-throughput RNA-Sequencing analysis, the UC Denver Genomics Core used an Illumina High Seq Mapping and bioinformatics analysis was done in collaboration with the Bioinformatics Shared Resources. The sequenced libraries generated 804,339,023 unfiltered reads of ~100 bps in length. Each replicate represents at least 96.9 million reads, a density sufficient for qualitative analysis of gene expression. The ~100 bp reads were 74
87 aligned to the MM9 mouse genome using Gsnap [169], allowing a mismatch of 4% and an indel cost of 2.0. The mapped transcripts were visualized in the UCSC genome browser to bioinformatically verify differential expression in transcript counts and/or transcript length, as well as splicing of introns. Data Analysis and Interpretation For measurements of differential gene expression edger software [170] was used. Raw read counts were established for each RefSeq gene. In cases where more than one transcript was annotated, the transcript with the highest overall counts was used as proxy for the gene in question, yielding a data table with a single count entry for each gene per sample. The data table was trimmed to remove all genes where zero counts were reported in one or more of the 6 samples, with the exception of genes where either all the mutant samples, or all the wildtype samples showed zero counts, and the opposing side of the paradigm showed counts in each sample. This resulted in a gene data table with entries. The data set was used for differential expression analyses using edger. Comparisons of all three mutant samples versus all three wildtype samples were performed, as well as separate comparisons of each individual mutant sample against all three wildtype samples; resulting gene lists were used for Ingenuity Pathway Analysis to reveal biological features. Overall sample relationship was visualized using the multidimensional scaling feature of edger. Raw gene count numbers were normalized using DESeq [171], and normalized counts were used to calculate the coefficient of variation for each gene in the normal samples, as well as determine for each gene the relationship of the counts in each sample relative to the mean of the normalized counts derived from the three wildtype samples for the pertinent gene. To obtain distance from 75
88 mean, the ratio of normalized counts divided by mean of the normal underwent logarithmic transformation. All genes were ranked according to their coefficient of variation, and distance from mean was plotted for each gene for normal as well as for mutant samples. 76
89 CHAPTER V DISCUSSION Neural Tube Defects in humans have a complex etiology; it is likely that multiple genes contribute to the incidence of this common birth defect, but the specific genetic causes of NTDs are still largely unknown. Moreover, it is unknown whether polymorphisms in epigenetic regulators may be risk factors for NTDs in humans. Studies in model organisms like the mouse can provide insight into the genetics underlying NTDs. Indeed, mutations and deletions in a number of epigenetic regulators can cause NTDs in mice. In this study we focused on the BAF chromatin remodeling complex, as mutations in two members of the BAF complex cause NTDs in mice, yet the functions of the BAF complex at the time of neural tube closure have not been studied. Here we characterized the first missense allele of the Baf155 gene, Baf155 msp3/ msp3, which exhibits a cranial NTD. We find that the mutant protein can still interact with the core BAF complex proteins BRG1, BAF170, and BAF47 and properly localize to the nucleus. This unique allele allows an investigation of the cause of the NTD in the presence of an assembled core BAF complex. In Baf155 msp3 mutant embryos, we discovered a reduction in neural progenitor cell proliferation and an increase in cell death, revealing a possible mechanism underlying the NTD. We analyzed transcriptional control in the Baf155 msp3 mutants using RNA-Seq and discovered misregulation of genes involved in proliferation and apoptosis. We also found increased variability in gene expression in the mutant compared to WT, which will be discussed below as a possible cause for NTDs. 77
90 The Baf155 msp3 mutant exhibited increased variability of gene expression compared to the WT. This was shown by a multidimensional scaling plot where the mutants did not cluster with the WT. Additionally, the mutants did not cluster with each other. This suggests that there is a trend towards higher variance of global gene expression in the Baf msp3 mutants. Additionally the gene expression in individual mutant samples exhibits increased variability compared to the individual WT samples. This was analyzed by calculating the coefficient of variance in each individual sample compared to normalized gene expression derived from the mean of WT samples. Overall gene expression in the WTs showed tightly clustered coefficients of variance, suggesting low variability of gene expression in each of the three WT samples. In contrast, the mutants exhibited increased variability as the coefficients of variance were more dispersed in the individual Baf155 msp3/ msp3 samples compared to the individual WT samples. These data suggest that gene expression is more variable in each individual Baf155 msp3 mutant sample compared to each individual WT sample. Another way of looking at this is that the BAF155 msp3 mutant protein may cause misregulation of genes in an inconsistent way. Most current methods of transcriptional analysis, microarrays and RNA-Seq, are aimed at reducing variability by pooling and averaging samples. This approach masks individual variation in transcriptional control between samples. Conceptually, my studies required analyzing each mutant individually, and nonstandard statistical analyses to allow for discovery of variably transcribed genes. With that said, we were also able to find some connections with various potential mechanistic pathways that show misregulation in the mutant. However, disruptions of single genes and pathways are likely not representative of most birth defects, and the added complexity of transcriptional variability has recently been implicated in 78
91 diabetic model of NTDs in mice [172]. There are many mechanisms that may contribute to transcriptional variability, and several will be outlined here. First, it is necessary to outline the current evidence implicating variability of expression in birth defects and cancers. Recent studies have indicated the molecular basis of developmental defects and cancers may, at least in part, include an overall increase in noise of gene expression due to changes in epigenetic regulation [173]. Some scholars define noise as increased transcription of biologically irrelevant DNA [174]. However, as the field of biology progresses, it is difficult to definitively ascertain if a transcript lacks biological significance. It has become clear that many non-mrna transcripts, once originally thought of as junk, are actually important for other functions such as gene regulation [175]. It is tempting to suggest that this variability is derived from stochasticity in expression, or variation that cannot be explained by any known or measurable factor [176]. But knowing how much of the observed variation is in fact stochastic has been long debated. Early studies from Gartner on the quantitative traits in inbred rats and mice indicated that 20-30% of variation he found could be explained by genetic or environmental conditions, and the other 70%-80% may be explained by a third component [177]. This third component is likely a combination of many separate components, one of which is epigenetic control. Another estimate from early studies of morphological variation of the mouse skeleton is that only 10% of the observed variation cannot be explained [178]. This estimate seems more realistic considering stochasticity excludes all known and measurable variables. These two studies evaluated animals after the final phenotype has been reached; however, transcriptional control may be expected to 79
92 contain a similar amount of unexplained variation. With current advances in genetic technologies, several factors contributing to genetic regulation that were once undiscovered are now easily measurable. For example, single cell studies in E. coli suggest the precision of gene regulation can be limited by both intrinsic noise of low intracellular transcript levels and extrinsic noise of varying levels of inducible transcription. [179]. Remarkably, we can now measure the intracellular copy number of gene transcripts, and this will allow an analysis of gene expression variability at a single cell level. As this field progresses, it should be possible to account for and measure the effects of more of the unknowns that were once thought to contribute to genetic variability. Gene expression variability may not only give rise to single cell phenotypes, but also to organismal phenotypes. Given that our RNA-Seq results show that most differentially expressed genes are upregulated in the Baf155 msp3/msp3 homozygous mutant, it is probable that BAF155 msp3 contributed to the increased gene expression, whether directly or indirectly. Therefore, it is possible that the BAF complex normally suppresses transcriptional variability in the developing embryo. To extrapolate further, the mutant phenotypes may, in fact, be a result of variable gene expression. Attributing a mutant phenotype to variable gene expression is a relatively new concept in the field, however several studies suggest this may be the case. Intestinal cell specification of C. elegans was found to be highly dependent on the number of gene transcripts in the intestinal cell specification gene network expressed in each cell. Moreover, perturbations in the expression of specific genes were suggested to underlie the phenotypic variation [180]. This study found a threshold of gene expression could specifically give rise to some of the 20 alternative cell fates in the intestinal cells. If this finding is applicable to other model 80
93 systems, these data suggest that very small gene expression variability may contribute to gross phenotypic variation. Considering the BAF155 mutant, this may suggest that the NTD phenotype seen in 80% of the mutant embryos may be a consequence of, or at least impacted by, the gene expression variation discovered in the cranial tissue at E9.5. The phenotypic variation may be modulated by the gene expression variation, which in turn can be controlled by global gene regulators. There are increasingly more examples that disruption of global regulators contributes to increased variability within the cell, suggesting these proteins normally function to suppress transcriptional variability. For example, global regulators like HSP90 can play a role in limiting phenotypic variation by buffering transcriptional variation in Drosophila and Arabidopsis [181]. BAF155 can be considered an epigenetic regulatory protein because of its role within the chromatin remodeling complex. As an epigenetic regulatory protein, it has the potential to regulate genes throughout the entire genome and hence act as a global regulator. The BAF complex associates with the promoter or genic regions of approximately 3,380 distinct genes in ES cells [147], suggesting it has the potential to regulate thousands of genes. However, in our mutant, we found that only about 150 genes were misregulated using a stringent analysis, suggesting that the intact but disabled complex results in the misregulation of relatively few genes compared to WT. As suggested above and in other analyses of the BAF complex [121], BAF155 may be a suppressor of gene transcription, and perhaps many of its target genes remained repressed due to other retained functions of the BAF155 msp3 mutant protein. Conditional loss of BAF155 function in the embryonic brain tissue would be necessary to determine the overall number of genes that are regulated by BAF155 during this developmental period. 81
94 While mutations in the BAF complex itself may globally misregulate genes, there are other globally acting proteins whose altered expression in the Baf155 msp3 mutant may contribute to the observed variable gene expression. For example, expression of the HDAC SIRT7 gene was found to be misregulated in the Baf155 msp3 mutants. This could indirectly contribute to the gene expression variability seen in the mutant embryos. SIRT7 localizes to the nucleolus and has been implicated as an activator of RNA Pol I [182]. These studies suggest SIRT7 can increase transcription via increased number of ribosomes. Moreover, SIRT7 associates with protein subunits in the BAF complex [183]. Therefore, it is possible that misregulation of the mutant BAF complex containing the BAF155 msp3 protein, affects the activity and association of SIRT7, leading to increased variability in overall gene expression in the mutant. The Baf155 msp3 mutant embryos show variability in gene expression based on the RNA-Seq data. While these studies lend insight into the transcriptional regulation functions of BAF155, there are caveats to understanding the specific role of BAF155 in transcriptional control. Specifically, it is unclear which gene targets are directly controlled by the BAF chromatin remodeling complex and which are not. The best method to test if transcription may be directly controlled by the complex is to perform ChIP-Seq experiments. From studies such as these, it can be determined if BAF155 msp3 directly associates with genes that are found to be misregulated in the mutant. Additionally, these studies would suggest which genes are transcriptionally controlled by secondary or indirect mechanisms and not by direct association with the BAF complex. Because previous characterization of the BAF complex targets were based on depletion of BAF155 or BRG1 82
95 [125, 135, 147], the data presented in chapters IV may be more reliable in indicating targets that are specific for the intact complex. The proposed ChIP-Seq experiments in the context of the Baf155 msp3 mutant may reveal several outcomes that each can assist in revealing the functions of BAF155 in the context of development, and of the BAF155 msp3 protein in the context of NTDs. One possible outcome of a ChIP- Seq assay on the Baf155 msp3 mutant and WT embryos, may be variable association with gene promoters and gene bodies within the individual mutants, similar to the variable gene expression seen in individual mutants. This might manifest as differences in genes where BAF155 is bound, as well as the peak height corresponding to BAF155 enrichment at each gene, in individual Baf155 msp3 mutant embryos compared to WT embryos. This would suggest that the mutant BAF complex is inconsistent in its associations with DNA, which could contribute to the variable gene expression observed in the Baf155 msp3 mutant embryos. This outcome would support a hypothesis that the BAF chromatin remodeling complex associates at different gene promoters in different mutants compared to the WT, resulting in variable transcription of genes. One possible mechanism for altered association is a change in nuclear architecture. Evidence for this is that BRG1 is required for DNA looping at some gene promoter control regions to control transcription in vivo [184]. It is possible that the chromatin structure and nuclear architecture is altered in the Baf155 msp3 mutants, and this might be revealed by BAF155 ChIP-Seq experiments and could be correlated with variable gene association. Another possible outcome of Chip-Seq experiments on Baf155 msp3 mutant and WT E9.5 embryos is that the mutant BAF155 msp3 protein may associate with the same gene promoters and at a similar enrichment as compared to the WT. This would suggest the 83
96 genomic association of the mutant protein is not altered, nor is it variable between the individual mutants compared to the WT, despite the fact that the overall gene expression in the mutant embryos is variable. This would suggest that the differences in gene expression in individual mutants are not due to the association of mutant BAF155 msp3 on the gene promoters and must be due to some other mechanism. One mechanism may be that the nucleosome translocation function of the BAF complex is altered in the mutant compared to the WT. The WT BAF complex in ES cells is poised at differentiation genes [147]. In the E9.5 neural tissue, it is possible that the BAF complex in the WT embryos associates with the genome at genes that have been repressed. If the nucleosome translocation function is altered, the genes that are repressed in the WT may not be turned off in the mutant. These experiments would provide more insight into the origin of gene expression variability in the Baf155 msp3/msp3 embryo and whether it is a consequence of genomic associations or some other function of BAF155. The variability in gene expression in the Baf155 msp3/msp3 embryos might be a consequence of a gain or loss of function of the BAF complex. It is possible that BAF155 msp3 associates with more genes than the WT proteins. If so, this could suggest that the BAF155 msp3 may have an increased affinity for genes not normally bound by the complex, which would be consistent with a gain of function mutation. Because the domains within the BAF155 protein are suggested to interact with other proteins, and BAF155 has been shown to directly interact with proteins in the BAF complex [146], the mutation may disrupt a functional protein-protein interaction. This could change the affinity for other complex proteins or other transcription factors, and result in increased and/or prolonged genomic association of the BAF155 msp3 protein or BAF complex. While 84
97 no secondary structure is predicted to be disrupted by Phyre, the T to K mutation does change the threonine, which is a neutral polar amino acid, to lysine, which is a basic polar amino acid. This slightly alters the molecular environment at that location of the mutation and therefore could alter protein-protein interactions. BAF155 maintains tight associations within the complex; the mutation could change the overall structure of the complex, and thereby impede the manner in which the complex as a whole associates with the genome or remodels chromatin. Alternatively, BAF155 msp3 may associate with fewer genes compared to the WT BAF155. If so, then the upregulation of genes that we observed in the RNA-Seq experiment in the Baf155 msp3 mutant is consistent with a repressive function of BAF155. Additionally, this would be consistent with a loss of function mutation in BAF155 msp3. This loss of function may be attributed to altered chromatin remodeling functions in the msp3/ msp3 mutant BAF complex. Because it is unknown if the variability in the Baf155 mutant is due to gain or loss of direct interaction of the mutant BAF complex with gene promoters, it is yet unclear if the variability in the Baf15 msp3/msp3 mutant is by caused by malfunction of the mutant BAF complex or the functional consequence of genes that have become misregulated in the Baf155 msp3 mutant. It is possible the malfunction of the BAF complex in the Baf155 msp3 mutant embryos is due to chromatin remodeling dysfunction. As a chromatin remodeling machine, the BAF complex functions to remodel nucleosomes, thus changing the chromatin structure to increase accessibility of DNA to transcriptional machinery. This function was not tested in the studies outlined in this work and thus it is possible there is a defect in the chromatin remodeling function of BAF155 msp3. To test for this defect, an in vitro nucleosome remodeling assay can be performed. Briefly, the complex can be isolated from 85
98 mutant and WT cranial tissue by immunoprecipitation. Nucleosome movement in this assay can be visualized on an agarose gel and would appear as a shift in a small sized band, indicating nucleosomes have become more compact in the chromatin. Possible outcomes for the in vitro nucleosome remodeling assay are as follows: 1. The BAF complex containing the mutant BAF155 msp3 protein shows no movement of the nucleosomes in the array compared to the WT; 2. A reduced amount of movement on the nucleosomes in the array compared to the WT; 3. The same movement of nucleosomes with the mutant isolated complex compared to the WT; or 4. Increased movement compared to the WT. No nucleosome movement would imply that the mutant BAF complex is defective for in vitro chromatin remodeling function and likely could not remodel nucleosomes in vivo. While complete lack of chromatin remodeling activity cannot be ruled out, it is unlikely given that our Baf155 msp3/msp3 mutant embryos live longer than the Baf155 null/null embryos, suggesting BAF155 msp3, and possibly the BAF complex in our mutant, is partially functional. Therefore, reduced chromatin remodeling function, or no change in chromatin remodeling function, are more likely outcomes. The in vitro nucleosome remodeling assay will clarify whether the mutant complex has the ability to remodel nucleosomes in an artificial environment. In vivo, chromatin remodeling is more complex and requires more factors, and hence the in vitro assay cannot replicate this complexity but may be informative in understanding how BAF function may be compromised. Another effective test to determine the remodeling activity of the BAF complex is an in vivo DNaseI sensitivity assay. This assay tests whether the nucleosome pattern in the promoter region of the genes is altered, by assaying DNA accessibility. It would be ideal to assay the same tissue in which the RNA-Seq and follow up ChIP-Seq were conducted, 86
99 namely cranial tissue from E9.5 heads, but this requires significant quantities of tissue samples, or approximately 2 mg of extracted protein, from at least 8 pooled E9.5 heads. It is important to use the correct cell type to ascertain if the genes identified by ChIP Seq and RNA-Seq experiments are also associated with changes in nucleosome remodeling in the mutant relative to WT. Alternatively, the question of nucleosome remodeling in vivo can also be addressed in ES cells derived from Baf155 msp3/msp3 mutants. A DNaseI sensitivity assay indicates if the DNA is prone to digestion, presumably due to movement of nucleosomes from the promoter regions. Upon the addition of the differentiation factor Retinoic Acid (RA), the chromatin becomes more compact at Nanog and Oct4 loci and this is thought to reduce the accessibility of transcription factors to the DNA in the promoter region which then ultimately turns off the expression of these two pluripotency genes [128]. A DNaseI sensitivity assay will test if there is a change in accessibility of the chromatin to the enzyme DNaseI in gene promoters in WT and Baf155 msp3/msp3 ES cells. After isolating chromatin from the WT and mutant ES cell lines, the DNA is digested with the DNaseI enzyme. Next, associated chromatin proteins are digested away from the DNA template and then the DNA is separated on a gel. In this scenario, a Southern blot assay is used to probe for genes whose expression is decreased after differentiation such as Nanog or whose expression does not change after differentiation, such as Gapdh. In the absence of RA addition to WT and Baf155 msp3/msp3 ES cells, I would expect to see lower molecular weight bands that resemble a ladder on a Southern blot using a probe to the Nanog promoter. The low molecular weight bands correspond to DNA that is accessible to DNAaseI in the promoter region of Nanog and thus the chromatin has not been condensed via chromatin remodeling because Nanog is 87
100 expressed in WT ES cells prior to differentiation. The smallest molecular weight band corresponds to the DNA wrapped around one nucleosome. Upon addition of RA to WT ES cells, I would expect the Southern blot for the Nanog promoter to have fewer lower molecular weight bands, corresponding to reduced DNaseI accessibility to the DNA presumably because of tightly compacted chromatin around the Nanog promoter which serves to turn off its expression. Upon addition of RA to Baf155 msp3/msp3 mutant ES cells, if the mutant BAF chromatin remodeling function is impaired, then I would expect to see no change in the band pattern on the Southern blot compared to WT or to the Baf155 msp3 mutant before RA addition. This would suggest that the accessibility of the Nanog promoter cannot be changed upon differentiation due to dysfunction of the mutant BAF chromatin remodeling complex. Moreover, this could suggest that the mutant BAF complex cannot function to remodel nucleosomes with the same efficiency as the WT remodeling complex, and thus would be visualized on the Southern Blot as more intense lower molecular weight bands compared to WT after RA addition. This assay only tests DNA compaction in ES cells at a critical transition in development, but could be a proxy for understanding mutant BAF function during neural tube closure. Several lines of evidence indicate that the BAF chromatin remodeling complex can remodel nucleosomes in the Baf155 msp3 mutants. The core complex, defined as the minimum composition of proteins that remodels nucleosomes as efficiently as the whole complex, consists of 4 BAF proteins, BAF155, BAF170, BRG1 and BAF47 [144]. All of these proteins associate with mutant BAF155 msp3, suggesting the complex is intact and can maintain the chromatin remodeling function in vivo. Additionally, several studies that have depleted BRG1 from mouse do not show significant changes in nucleosome position in ES 88
101 cells and neural progenitor cells [96], suggesting BRG1 has other function in vivo and may not exclusively remodel nucleosomes. Therefore, the BAF chromatin remodeling complex may have other functions in addition to chromatin remodeling. Despite the varying degree of gene expression changes between the Baf155 msp3 mutants and WT, our mouse model shows a consistent NTD phenotype. Neural tube closure is a complex, multigenic process and our data indicate that inconsistent regulation of genes by a chromatin remodeling complex with impaired function can lead to developmental defects during this highly dynamic phase of embryonic development. 89
102 REFERENCES 1. Mitchell LE: Epidemiology of neural tube defects. In: American Journal of Medical Genetics Part C: Seminars in Medical Genetics: Wiley Online Library: Kibar Z, Capra V, Gros P: Toward understanding the genetic basis of neural tube defects. Clin Genet 2007, 71(4): Mitchell LE, Adzick NS, Melchionne J, Pasquariello PS, Sutton LN, Whitehead AS: Spina bifida. The Lancet 2004, 364(9448): Simeonsson RJ, McMillen JS, Huntington GS: Secondary conditions in children with disabilities: spina bifida as a case example. Mental retardation and developmental disabilities research reviews 2002, 8(3): Detrait ER, George TM, Etchevers HC, Gilbert JR, Vekemans M, Speer MC: Human neural tube defects: Developmental biology, epidemiology, and genetics. Neurotoxicol Teratol 2005, 27(3): Rampersaud E, Melvin E, Speer M: Nonsyndromic neural tube defects: genetic basis and genetic investigations. Neural tube defects: from origin to treatment 2005, 42(12): Elwood JM, Little J, Elwood JH: Epidemiology and control of neural tube defects, vol. 20: Oxford University Press New York; Leck I: Causation of neural tube defects: clues from epidemiology. Br Med Bull 1974, 30(2): Copp AJ, Greene ND: Genetics and development of neural tube defects. The Journal of pathology 2010, 220(2): Rampersaud E, Bassuk AG, Enterline DS, George T, Siegel DG, Melvin EC, Aben J, Allen J, Aylsworth A, Brei T: Whole genomewide linkage screen for neural tube defects reveals regions of interest on chromosomes 7 and 10. J Med Genet 2005, 42(12): Stamm DS, Rampersaud E, Slifer SH, Mehltretter L, Siegel DG, Xie J, Hu Lince D, Craig DW, Stephan DA, George TM: High density single nucleotide polymorphism screen in a large multiplex neural tube defect family refines linkage to loci at 7p21. 1 pter and 2q33. 1 q35. Birth Defects Research Part A: Clinical and Molecular Teratology 2006, 76(6):
103 12. Hol FA, Geurds MP, Chatkupt S, Shugart YY, Balling R, Schrander-Stumpel CT, Johnson WG, Hamel BC, Mariman EC: PAX genes and human neural tube defects: an amino acid substitution in PAX1 in a patient with spina bifida. J Med Genet 1996, 33(8): Volcik K, Blanton S, Kruzel M, Townsend I, Tyerman G, Mier R, Northrup H: Testing for genetic associations with the PAX gene family in a spina bifida population. Am J Med Genet 2002, 110(3): Kibar, Zoha, Torban PD, Elena, McDearmid PD, Jonathan, Reynolds PD, Annie, Berghout MS, Joanne et al: Mutations in VANGL1 Associated with Neural- Tube Defects. N Engl J Med 2007: De Marco P, Merello E, Consales A, Piatelli G, Cama A, Kibar Z, Capra V: Genetic analysis of disheveled 2 and disheveled 3 in human neural tube defects. J Mol Neurosci 2013, 49(3): Greene ND, Stanier P, Copp AJ: Genetics of human neural tube defects. Hum Mol Genet 2009, 18(R2):R113-R Harris MJ, Juriloff DM: Mini-review: toward understanding mechanisms of genetic neural tube defects in mice. Teratology 1999, 60(5): Harris M, Juriloff D: Mouse mutants with neural tube closure defects and their role in understanding human neural tube defects. Birth Defects Research Part A: Clinical and Molecular Teratology 2007, 79(3): Harris MJ, Juriloff DM: An update to the list of mouse mutants with neural tube closure defects and advances toward a complete genetic perspective of neural tube closure. Birth Defects Research Part A: Clinical and Molecular Teratology 2010, 88(8): Waterston RH, Lindblad-Toh K, Birney E, Rogers J, Abril JF, Agarwal P, Agarwala R, Ainscough R, Alexandersson M, An P et al: Initial sequencing and comparative analysis of the mouse genome. Nature 2002, 420(6915): Harrington MJ, Hong E, Brewster R: Comparative analysis of neurulation: First impressions do not count. Mol Reprod Dev 2009, 76(10): Saitsu H, Yamada S, Uwabe C, Ishibashi M, Shiota K: Development of the posterior neural tube in human embryos. Anat Embryol (Berl) 2004, 209(2): Beddington R: Induction of a second neural axis by the mouse node. Development 1994, 120(3):
104 24. Ray HJ, Niswander L: Mechanisms of tissue fusion during development. Development 2012, 139(10): De Robertis EM, Kuroda H: Dorsal-ventral patterning and neural induction in Xenopus embryos. Annu Rev Cell Dev Biol 2004, 20: Wilson PA, Hemmati-Brivanlou A: Induction of epidermis and inhibition of neural fate by Bmp-4. Nature 1995, 376(6538): Hemmati-Brivanlou A, Stewart RM, Harland RM: Region-specific neural induction of an engrailed protein by anterior notochord in Xenopus. Science 1990, 250(4982): Wallingford JB, Harland RM: Neural tube closure requires Dishevelleddependent convergent extension of the midline. Development 2002, 129(24): Ybot-Gonzalez P, Savery D, Gerrelli D, Signore M, Mitchell CE, Faux CH, Greene ND, Copp AJ: Convergent extension, planar-cell-polarity signalling and initiation of mouse neural tube closure. Development 2007, 134(4): Haigo SL, Hildebrand JD, Harland RM, Wallingford JB: Shroom induces apical constriction and is required for hingepoint formation during neural tube closure. Curr Biol 2003, 13(24): Hildebrand JD, Soriano P: Shroom, a PDZ domain containing actin-binding protein, is required for neural tube morphogenesis in mice. Cell 1999, 99(5): Massarwa Ra, Niswander L: In toto live imaging of mouse morphogenesis and new insights into neural tube closure. Development 2013, 140(1): Holmberg J, Clarke DL, Frisén J: Regulation of repulsion versus adhesion by different splice forms of an Eph receptor. Nature 2000, 408(6809): Rifat Y, Parekh V, Wilanowski T, Hislop NR, Auden A, Ting SB, Cunningham JM, Jane SM: Regional neural tube closure defined by the< i> Grainy head</i>-like transcription factors. Dev Biol 2010, 345(2): Pyrgaki C, Liu A, Niswander L: < i> Grainyhead-like 2</i> regulates neural tube closure and adhesion molecule expression during neural fold fusion. Dev Biol 2011, 353(1):
105 36. Werth M, Walentin K, Aue A, Schönheit J, Wuebken A, Pode-Shakked N, Vilianovitch L, Erdmann B, Dekel B, Bader M: The transcription factor grainyhead-like 2 regulates the molecular composition of the epithelial apical junctional complex. Development 2010, 137(22): Copp A: Neurulation in the cranial region normal and abnormal. J Anat 2005, 207: Juriloff D, Harris M, Tom C, MacDonald K: Normal mouse strains differ in the site of initiation of closure of the cranial neural tube. Teratology 1991, 44(2): Liu A, Joyner AL: Early anterior/posterior patterning of the midbrain and cerebellum. Annu Rev Neurosci 2001, 24(1): Accessed June [ Lagutin OV, Zhu CC, Kobayashi D, Topczewski J, Shimamura K, Puelles L, Russell HR, McKinnon PJ, Solnica-Krezel L, Oliver G: Six3 repression of Wnt signaling in the anterior neuroectoderm is essential for vertebrate forebrain development. Genes Dev 2003, 17(3): Houart C, Caneparo L, Heisenberg C-P, Barth KA, Take-Uchi M, Wilson SW: Establishment of the telencephalon during gastrulation by local antagonism of Wnt signaling. Neuron 2002, 35(2): Chiang C, Ying LTT, Lee E, Young KE, Corden JL, Westphal H, Beachy PA: Cyclopia and defective axial patterning in mice lacking Sonic hedgehog gene function. Nature 1996, 383(6599): Nakamura H, Katahira T, Matsunaga E, Sato T: Isthmus organizer for midbrain and hindbrain development. Brain research reviews 2005, 49(2): Meyers EN, Lewandoski M, Martin GR: An Fgf8 mutant allelic series generated by Cre-and Flp-mediated recombination. Nat Genet 1998, 18(2): Ross SA, McCaffery PJ, Drager UC, De Luca LM: Retinoids in embryonal development. Physiol Rev 2000, 80(3): Mark M, Ghyselinck NB, Chambon P: Function of retinoid nuclear receptors: lessons from genetic and pharmacological dissections of the retinoic acid signaling pathway during mouse embryogenesis. Annu Rev Pharmacol Toxicol 2006, 46:
106 48. Lemons D, McGinnis W: Genomic evolution of Hox gene clusters. Science 2006, 313(5795): Studer M, Gavalas A, Marshall H, Ariza-McNaughton L, Rijli FM, Chambon P, Krumlauf R: Genetic interactions between Hoxa1 and Hoxb1 reveal new roles in regulation of early hindbrain patterning. Development 1998, 125(6): Dupé V, Davenne M, Brocard J, Dollé P, Mark M, Dierich A, Chambon P, Rijli FM: In vivo functional analysis of the Hoxa-1 3 retinoic acid response element (3 RARE). Development 1997, 124(2): Muhr J, Graziano E, Wilson S, Jessell TM, Edlund T: Convergent inductive signals specify midbrain, hindbrain, and spinal cord identity in gastrula stage chick embryos. Neuron 1999, 23(4): Avilés EC, Wilson NH, Stoeckli ET: Sonic hedgehog and Wnt: antagonists in morphogenesis but collaborators in axon guidance. Front Cell Neurosci 2013, 7: Chamberlain CE, Jeong J, Guo C, Allen BL, McMahon AP: Notochord-derived Shh concentrates in close association with the apically positioned basal body in neural target cells and forms a dynamic gradient during neural patterning. Development 2008, 135(6): Briscoe J, Pierani A, Jessell T, Ericson J: A homeodomain protein code specifies progenitor cell identity and neuronal fate in the ventral neural tube. Cell 2000, 101: Briscoe J, Ericson J: Specification of neuronal fates in the ventral neural tube. Curr Opin Neurobiol 2001, (11): Liem Jr KF, Tremml G, Jessell TM: A role for the roof plate and its resident TGFβ-related proteins in neuronal patterning in the dorsal spinal cord. Cell 1997, 91(1): Liem Jr KF, Tremml G, Roelink H, Jessell TM: Dorsal differentiation of neural plate cells induced by BMP-mediated signals from epidermal ectoderm. Cell 1995, 82(6): Lee KJ, Dietrich P, Jessell TM: Genetic ablation reveals that the roof plate is essential for dorsal interneuron specification. Nature 2000, 403(6771):
107 59. Müller T, Brohmann H, Pierani A, Heppenstall PA, Lewin GR, Jessell TM, Birchmeier C: The homeodomain factor lbx1 distinguishes two major programs of neuronal differentiation in the dorsal spinal cord. Neuron 2002, 34(4): Schlüter G: Ultrastructural observations on cell necrosis during formation of the neural tube in mouse embryos. Z Anat Entwicklungsgesch 1973, 141(3): Lawson A, Schoenwolf GC, England MA, Addai FK, Ahima RS: Programmed cell death and the morphogenesis of the hindbrain roof plate in the chick embryo. Anat Embryol (Berl) 1999, 200(5): Migliorini D, Denchi E, Danovi D, Jochemsen A, Capillo M, Gobbi A, Helin K, Pelicci P, Marine J: Mdm4 (Mdmx) regulates p53-induced growth arrest and neuronal cell death during early embryonic mouse development. Molecular and cellular biology 2002, 22(15): Sah V, Attardi L, Mulligan G, Williams B: A subset of p53-deficient embryos exhibit exencephaly. Nature 1995, 10: Kuida K, Haydar TF, Kuan C-Y, Gu Y, Taya C, Karasuyama H, Su MS-S, Rakic P, Flavell RA: Reduced apoptosis and cytochrome c mediated caspase activation in mice lacking caspase 9. Cell 1998, 94(3): Massa V, Savery D, Ybot-Gonzalez P, Ferraro E, Rongvaux A, Cecconi F, Flavell R, Greene ND, Copp AJ: Apoptosis is not required for mammalian neural tube closure. Proc Natl Acad Sci U S A 2009, 106(20): Yamaguchi Y, Shinotsuka N, Nonomura K, Takemoto K, Kuida K, Yosida H, Miura M: Live imaging of apoptosis in a novel transgenic mouse highlights its role in neural tube closure. The Journal of Cell Biology 2011, 195(6): Copp AJ, Greene ND, Murdoch JN: The genetic basis of mammalian neurulation. Nat Rev Genet 2003, 4(10): Ishibashi M, Ang S-L, Shiota K, Nakanishi S, Kageyama R, Guillemot F: Targeted disruption of mammalian hairy and Enhancer of split homolog-1 (HES-1) leads to up-regulation of neural helix-loop-helix factors, premature neurogenesis, and severe neural tube defects. Genes Dev 1995, 9(24): Lardelli M, Williams R, Mitsiadis T, Lendahl U: Expression of the Notch 3 intracellular domain in mouse central nervous system progenitor cells is lethal and leads to disturbed neural tube development. Mech Dev 1996, 59(2):
108 70. Chen Z, Behringer R: twist is required in head mesenchyme for cranial neural tube morphogenesis. Genes & Development 1995, 9(6): Luger K, Mäder A, Richmond R, Sargent D: Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 1997, 386: Segal E, Fondufe-Mittendorf Y, Chen L, Thåström A, Field Y, Moore IK, Wang J- PZ, Widom J: A genomic code for nucleosome positioning. Nature 2006, 442(7104): Kamakaka RT, Biggins S: Histone variants: deviants? Genes Dev 2005, 19(3): Ahmad K, Henikoff S: The histone variant H3. 3 marks active chromatin by replication-independent nucleosome assembly. Mol Cell 2002, 9(6): Kornberg RD: Chromatin structure: a repeating unit of histones and DNA. Science 1974, 184(4139): Gosden, Roger, Ph.D., Feinberg DS, Andrew: Genetics and Epigenetics Nature's Pen-and-Pencil Set. N Engl J Med 2007: Juriloff DM, Harris MJ: Mouse models for neural tube closure defects. Hum Mol Genet 2000, 9(6): Li E, Bestor TH, Jaenisch R: Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell 1992, 69(6): Okano M, Bell DW, Haber DA, Li E: DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 1999, 99(3): Dunlevy LP, Burren KA, Chitty LS, Copp AJ, Greene ND: Excess methionine suppresses the methylation cycle and inhibits neural tube closure in mouse embryos. FEBS Lett 2006, 580(11): Kouzarides T: Chromatin modifications and their function. Cell 2007, 128(4): Ernst J, Kellis M: Discovery and characterization of chromatin states for systematic annotation of the human genome. Nat Biotechnol 2010, 28(8): Wang Z, Zang C, Cui K, Schones DE, Barski A, Peng W, Zhao K: Genome-wide mapping of HATs and HDACs reveals distinct functions in active and inactive genes. Cell 2009, 138(5):
109 84. Bu P, Evrard YA, Lozano G, Dent SY: Loss of Gcn5 Acetyltransferase Activity Leads to Neural Tube Closure Defects and Exencephaly in Mouse Embryos. Mol Cell Biol 2007, 27(9): Xu W, Edmondson D, Evrard Y, Wakamiya M: Loss of Gcn5l2 leads to increased apoptosis and mesodermal defects during mouse development. Nature 2000, 26: Tanaka Y, Naruse I, Hongo T, Xu M: Extensive brain hemorrhage and embryonic lethality in a mouse null mutant of CREB-binding protein. Mech Dev 2000, 95(1-2): Yao T, Oh S, Fuchs M, Zhou N, Ch'ng: Gene dosage dependent embryonic development and proliferation defects in mice lacking the transcriptional integrator p300. Cell 1998, 93: Vega RB, Matsuda K, Oh J, Barbosa AC, Yang X, Meadows E, McAnally J, Pomajzl C, Shelton JM, Richardson JA et al: Histone Deacetylase 4 Controls Chondrocyte Hypertrophy during Skeletogenesis. Cell 2004, 119(4): Cheng H-: Developmental defects and p53 hyperacetylation in Sir2 homolog (SIRT1)-deficient mice. Proceedings of the National Academy of Sciences 2003, 100(19): Kalkhoven E: CBP and p300: HATs for different occasions. Biochem Pharmacol 2004, 68(6): Marmorstein R: Structure of histone acetyltransferases. J Mol Biol 2001, 311(3): Clapier CR, Cairns BR: The Biology of Chromatin Remodeling Complexes. Annu Rev Biochem 2009, 78(1): Narlikar G, Fan H, Kingston R: Cooperation between complexes that regulate chromatin structure and transcription. Cell 2002, 108(4): Lorch Y, Zhang M, Kornberg R: Histone octamer transfer by a chromatinremodeling complex. Cell 1999, 96(3): Mizuguchi G, Shen X, Landry J, Wu W, Sen S, Wu C: ATP-driven exchange of histone H2AZ variant catalyzed by SWR1 chromatin remodeling complex. Science 2004, 303(5656): Hargreaves DC, Crabtree GR: ATP-dependent chromatin remodeling: genetics, genomics and mechanisms. Cell Res 2011, 21(3):
110 97. Mohrmann L, Verrijzer CP: Composition and functional specificity of SWI2/SNF2 class chromatin remodeling complexes. Biochimica et Biophysica Acta (BBA) - Gene Structure and Expression 2005, 1681(2-3): Tsukiyama T, Daniel C, Tamkun J, Wu C: ISWI, a member of the SWl2/SNF2 ATPase family, encodes the 140 kda subunit of the nucleosome remodeling factor. Cell 1995, 83(6): Stopka T, Skoultchi AI: The ISWI ATPase Snf2h is required for early mouse development. Proceedings of the National Academy of Sciences 2003, 100(24): Landry J, Sharov AA, Piao Y, Sharova LV, Xiao H, Southon E, Matta J, Tessarollo L, Zhang YE, Ko MS: Essential role of chromatin remodeling protein Bptf in early mouse embryos and embryonic stem cells. PLoS genetics 2008, 4(10):e Banting GS: CECR2, a protein involved in neurulation, forms a novel chromatin remodeling complex with SNF2L. Hum Mol Genet 2004, 14(4): Barak O, Lazzaro MA, Lane WS, Speicher DW, Picketts DJ, Shiekhattar R: Isolation of human NURF: a regulator of Engrailed gene expression. The EMBO journal 2003, 22(22): Shen X, Mizuguchi G, Hamiche A, Wu C: A chromatin remodelling complex involved in transcription and DNA processing. Nature 2000, 406(6795): Morrison AJ, Shen X: Chromatin remodelling beyond transcription: the INO80 and SWR1 complexes. Nature Reviews Molecular Cell Biology 2009, 10(6): Sapountzi V, Logan IR, Robson CN: Cellular functions of TIP60. The international journal of biochemistry & cell biology 2006, 38(9): Fazzio TG, Huff JT, Panning B: An RNAi screen of chromatin proteins identifies Tip60-p400 as a regulator of embryonic stem cell identity. Cell 2008, 134(1): Jacobs SA, Khorasanizadeh S: Structure of HP1 chromodomain bound to a lysine 9-methylated histone H3 tail. Science 2002, 295(5562): Marfella CG, Imbalzano AN: The Chd family of chromatin remodelers. Mutation Research/Fundamental and Molecular Mechanisms of Mutagenesis 2007, 618(1):
111 109. Tong JK, Hassig CA, Schnitzler GR, Kingston RE, Schreiber SL: Chromatin deacetylation by an ATP-dependent nucleosome remodelling complex. Nature 1998, 395(6705): Xue Y, Wong J, Moreno GT, Young MK, Côté J, Wang W: NURD, a novel complex with both ATP-dependent chromatin-remodeling and histone deacetylase activities. Mol Cell 1998, 2(6): Toh Y, Nicolson GL: The role of the MTA family and their encoded proteins in human cancers: molecular functions and clinical implications. Clin Exp Metastasis 2009, 26(3): Vissers LE, van Ravenswaaij CM, Admiraal R, Hurst JA, de Vries BB, Janssen IM, van der Vliet WA, Huys EH, de Jong PJ, Hamel BC: Mutations in a new member of the chromodomain gene family cause CHARGE syndrome. Nat Genet 2004, 36(9): Hurd EA, Capers PL, Blauwkamp MN, Adams ME, Raphael Y, Poucher HK, Martin DM: Loss of Chd7 function in gene-trapped reporter mice is embryonic lethal and associated with severe defects in multiple developing tissues. Mamm Genome 2007, 18(2): Nasmyth K: The determination of mother cell-specific mating type switching in yeast by a specific regulator of HO transcription. The EMBO journal 1986, 6(1): Zeng L, Zhou M-M: Bromodomain: an acetyl-lysine binding domain. FEBS Lett 2002, 513(1): Hassan AH, Prochasson P, Neely KE, Galasinski SC, Chandy M, Carrozza MJ, Workman JL: Function and selectivity of bromodomains in anchoring chromatin-modifying complexes to promoter nucleosomes. Cell 2002, 111(3): Sudarsanam P, Iyer V, Brown P, Winston F: Whole-genome expression analysis of snf/swi mutants of Saccharomycescerevisiae. Proceedings of the National Academy of Sciences of the United States of America 2000, 97(7): Brown E, Malakar S, Krebs JE: How many remodelers does it take to make a brain? Diverse and cooperative roles of ATP-dependent chromatinremodeling complexes in development This paper is one of a selection of papers published in this Special Issue, entitled 28th International West Coast Chromatin and Chromosome Conference, and has undergone the Journal's usual peer review process. Biochem Cell Biol 2007, 85(4):
112 119. Kadoch C, Hargreaves DC, Hodges C, Elias L, Ho L, Ranish J, Crabtree GR: Proteomic and bioinformatic analysis of mammalian SWI/SNF complexes identifies extensive roles in human malignancy. Nat Genet 2013, 45(6): Ronan JL, Wu W, Crabtree GR: From neural development to cognition: unexpected roles for chromatin. Nature reviews Genetics 2013, 14(5): Ho L, Crabtree GR: Chromatin remodelling during development. Nature 2010, 463(7280): Reyes J, Barra J, Muchardt C, Camus A, Babinet C, Yaniv M: Altered control of cellular proliferation in the absence of mammalian brahma (SNF2α). The EMBO journal 1998, 17(23): Sun F, Tang F, Yan AY, Fang HY, Sheng HZ: Expression of SRG3, a chromatinremodelling factor, in the mouse oocyte and early preimplantation embryos. Zygote (Cambridge, England) 2007, 15(2): Mandel S, Gozes I: Activity-dependent Neuroprotective Protein Constitutes a Novel Element in the SWI/SNF Chromatin Remodeling Complex. J Biol Chem 2007, 282(47): Ho L, Ronan JL, Wu J, Staahl BT, Chen L, Kuo A, Lessard J, Nesvizhskii AI, Ranish J, Crabtree GR: An embryonic stem cell chromatin remodeling complex, esbaf, is essential for embryonic stem cell self-renewal and pluripotency. Proceedings of the National Academy of Sciences 2009, 106(13): Schwartz YB, Pirrotta V: Polycomb silencing mechanisms and the management of genomic programmes. Nature Reviews Genetics 2007, 8(1): Ho L, Miller EL, Ronan JL, Ho WQ, Jothi R, Crabtree GR: esbaf facilitates pluripotency by conditioning the genome for LIF/STAT3 signalling and by regulating polycomb function. Nat Cell Biol 2011, 13(8): Schaniel C, Ang Y, Ratnakumar K, Cormier C, James T, Bernstein E, Lemischka IR, Paddison PJ: Smarcc1/Baf155 Couples Self-Renewal Gene Repression with Changes in Chromatin Structure in Mouse Embryonic Stem Cells. Stem Cells 2009, 27: Takeuchi JK, Lickert H, Bisgrove BW, Sun X, Yamamoto M, Chawengsaksophak K, Hamada H, Yost HJ, Rossant J, Bruneau BG: Baf60c is a nuclear Notch signaling component required for the establishment of left-right asymmetry. Proc Natl Acad Sci U S A 2007, 104(3): Lickert H, Takeuchi J, Both I, Walls J: Baf60c is essential for function of BAF chromatin remodelling complexes in heart development. Nature 2004, 432:
113 131. Takeuchi JK, Bruneau BG: Directed transdifferentiation of mouse mesoderm to heart tissue by defined factors. Nature 2009, 459(7247): Lessard J, Wu JI, Ranish JA, Wan M, Winslow MM, Staahl BT, Wu H, Aebersold R, Graef IA, Crabtree GR: An Essential Switch in Subunit Composition of a Chromatin Remodeling Complex during Neural Development. Neuron 2007, 55(2): Yoo AS, Staahl BT, Chen L, Crabtree GR: MicroRNA-mediated switching of chromatin-remodelling complexes in neural development. Nature 2009, 460(7255): Bultman S, Gebuhr T, Yee D, La Mantia C, Nicholson J, Gilliam A, Randazzo F, Metzger D, Chambon P, Crabtree G: A Brg1 Null Mutation in the Mouse Reveals Functional Differences among Mammalian SWI/SNF Complexes. Mol Cell 2000, 6(6): Kim JK, Huh S-, Choi H, Lee K-, Shin D, Lee C, Nam J-, Kim H, Chung H, Lee HW et al: Srg3, a Mouse Homolog of Yeast SWI3, Is Essential for Early Embryogenesis and Involved in Brain Development. Mol Cell Biol 2001, 21(22): Choi J, Ko M, Jeon S, Jeon Y, Park K, Lee C, Lee H, Seong RH: The SWI/SNFlike BAF complex is essential for early B cell development. The Journal of Immunology 2012, 188(8): Jang J, Choi YI, Choi J, Lee KY, Chung H, Jeon SH, Seong RH: Notch1 confers thymocytes a resistance to GC-induced apoptosis through Deltex1 by blocking the recruitment of p300 to the SRG3 promoter. Cell Death Differ 2005, 13(9): Ahn J, Ko M, Lee C, Kim J, Yoon H, Seong RH: Srg3, a mouse homolog of BAF155, is a novel p53 target and acts as a tumor suppressor by modulating p21waf1/cip1 expression. Oncogene 2010, 30(4): Hong CY, Suh JH, Kim K, Gong E-, Jeon SH, Ko M, Seong RH, Kwon HB, Lee K: Modulation of Androgen Receptor Transactivation by the SWI3-Related Gene Product (SRG3) in Multiple Ways. Mol Cell Biol 2005, 25(12): Kasarskis A, Manova K, Anderson KV: A phenotype-based screen for embryonic lethal mutations in the mouse. Proceedings of the National Academy of Sciences 1998, 95(13):
114 141. Buac K, Watkins-Chow DE, Loftus SK, Larson DM, Incao A, Gibney G, Pavan WJ: A Sox10 Expression Screen Identifies an Amino Acid Essential for Erbb3 Function. PLoS Genetics 2008, 4(9):e Han D, Jeon S, Sohn DH, Lee C, Ahn S, Kim WK, Chung H, Seong RH: SRG3, a core component of mouse SWI/SNF complex, is essential for extra-embryonic vascular development. Dev Biol 2008, 315(1): Wang S, Aurora AB, Johnson BA, Qi X, McAnally J, Hill JA, Richardson JA, Bassel-Duby R, Olson EN: The endothelial-specific microrna mir-126 governs vascular integrity and angiogenesis. Dev Cell 2008, 15(2): Phelan ML, Sif S, Narlikar GJ, Kingston RE: Reconstitution of a core chromatin remodeling complex from SWI/SNF subunits. Mol Cell 1999, 3(2): Chen J, Archer TK: Regulating SWI/SNF Subunit Levels via Protein-Protein Interactions and Proteasomal Degradation: BAF155 and BAF170 Limit Expression of BAF57. Mol Cell Biol 2005, 25(20): Sohn DH, Lee KY, Lee C, Oh J, Chung H, Jeon SH, Seong RH: SRG3 Interacts Directly with the Major Components of the SWI/SNF Chromatin Remodeling Complex and Protects Them from Proteasomal Degradation. J Biol Chem 2007, 282(14): Ho L, Jothi R, Ronan JL, Cui K, Zhao K, Crabtree GR: An embryonic stem cell chromatin remodeling complex, esbaf, is an essential component of the core pluripotency transcriptional network. Proceedings of the National Academy of Sciences 2009, 106(13): Kelley LA, Sternberg MJ: Protein structure prediction on the Web: a case study using the Phyre server. Nat Protoc 2009, 4(3): Zhang Y, Stec B, Godzik A: Between order and disorder in protein structures: analysis of dual personality fragments in proteins. Structure 2007, 15(9): Fuxreiter M, Tompa P, Simon I: Local structural disorder imparts plasticity on linear motifs. Bioinformatics 2007, 23(8): Trainor PA: Specification of neural crest cell formation and migration in mouse embryos. Semin Cell Dev Biol 2005, 16(6): Sandhu KS: Intrinsic disorder explains diverse nuclear roles of chromatin remodeling proteins. J Mol Recognit 2009, 22(1):
115 153. Scholzen T, Gerdes J: The Ki 67 protein: from the known and the unknown. J Cell Physiol 2000, 182(3): Dykhuizen EC, Hargreaves DC, Miller EL, Cui K, Korshunov A, Kool M, Pfister S, Cho Y, Zhao K, Crabtree GR: BAF complexes facilitate decatenation of DNA by topoisomerase IIα. Nature 2013, 497(7451): Doxsey S, Zimmerman W, Mikule K: Centrosome control of the cell cycle. Trends Cell Biol 2005, 15(6): Kim S, Zaghloul NA, Bubenshchikova E, Oh EC, Rankin S, Katsanis N, Obara T, Tsiokas L: Nde1-mediated inhibition of ciliogenesis affects cell cycle re-entry. Nat Cell Biol 2011, 13(4): Shanahan F, Seghezzi W, Parry D, Mahony D, Lees E: Cyclin E associates with BAF155 and BRG1, components of the mammalian SWI-SNF complex, and alters the ability of BRG1 to induce growth arrest. Mol Cell Biol 1999, 19(2): Foster KS, McCrary WJ, Ross JS, Wright CF: Members of the hswi/snf chromatin remodeling complex associate with and are phosphorylated by protein kinase B/Akt. Oncogene 2006, 25(33): Huang L, Voyiaziakis E, Markenson D: apo B gene knockout in mice results in embryonic lethality in homozygotes and neural tube defects, male infertility, and reduced HDL cholesterol ester and apo AI transport. Journal of Clinical 1995, 96: Koleske A, Gifford A, Scott M, Nee M, Bronson R: Essential roles for the Abl and Arg tyrosine kinases in neurulation. Neuron 1998, 21: Wang Z, Gerstein M, Snyder M: RNA-Seq: a revolutionary tool for transcriptomics. Nature Reviews Genetics 2009, 10(1): Jadeja S, Smyth I, Pitera JE, Taylor MS, Haelst M, Bentley E, McGregor L, Hopkins J, Chalepakis G, Philip N et al: Identification of a new gene mutated in Fraser syndrome and mouse myelencephalic blebs. Nat Genet 2005, 37(5): Timmer JR: Tissue morphogenesis and vascular stability require the Frem2 protein, product of the mouse myelencephalic blebs gene. Proceedings of the National Academy of Sciences 2005, 102(33):
116 164. Euskirchen GM, Auerbach RK, Davidov E, Gianoulis TA, Zhong G, Rozowsky J, Bhardwaj N, Gerstein MB, Snyder M: Diverse Roles and Interactions of the SWI/SNF Chromatin Remodeling Complex Revealed Using Global Approaches. PLoS Genetics 2011, 7(3):e Juriloff DM, Harris MJ: A consideration of the evidence that genetic defects in planar cell polarity contribute to the etiology of human neural tube defects. Birth Defects Res A 2012, 94(10): Yu H, Smallwood PM, Wang Y, Vidaltamayo R, Reed R, Nathans J: Frizzled 1 and frizzled 2 genes function in palate, ventricular septum and neural tube closure: general implications for tissue fusion processes. Development 2010, 137(21): Wang Y: The Role of Frizzled3 and Frizzled6 in Neural Tube Closure and in the Planar Polarity of Inner-Ear Sensory Hair Cells. J Neurosci 2006, 26(8): Ozsolak F, Milos PM: RNA sequencing: advances, challenges and opportunities. Nature Reviews Genetics 2010, 12(2): Wu TD, Nacu S: Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 2010, 26(7): Robinson MD, McCarthy DJ, Smyth GK: edger: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26(1): Anders S, Huber W: Differential expression analysis for sequence count data. Genome Biol 2010, 11(10):R Salbaum JM, Kappen C: Diabetic embryopathy: a role for the epigenome? Birth Defects Res A Clin Mol Teratol 2011, 91(8): Pujadas E, Feinberg AP: Regulated noise in the epigenetic landscape of development and disease. Cell 2012, 148(6): Struhl K: Transcriptional noise and the fidelity of initiation by RNA polymerase II. Nat Struct Mol Biol 2007, 14(2): Mercer TR, Dinger ME, Mattick JS: Long non-coding RNAs: insights into functions. Nature Reviews Genetics 2009, 10(3): Zernicka-Goetz M, Huang S: Stochasticity versus determinism in development: a false dichotomy? Nature Reviews Genetics 2010, 11(11):
117 177. Gärtner K: A third component causing random variability beside environment and genotype. A reason for the limited success of a 30 year long effort to standardize laboratory animals? Lab Anim 1990, 24(1): Grüneberg H: Genetical studies on the skeleton of the mouse. Journal of Genetics 1952, 51(1): Elowitz MB, Levine AJ, Siggia ED, Swain PS: Stochastic gene expression in a single cell. Science 2002, 297(5584): Raj A, Rifkin SA, Andersen E, van Oudenaarden A: Variability in gene expression underlies incomplete penetrance. Nature 2010, 463(7283): Queitsch C, Sangster TA, Lindquist S: Hsp90 as a capacitor of phenotypic variation. Nature 2002, 417(6889): Ford E, Voit R, Liszt G, Magin C, Grummt I, Guarente L: Mammalian Sir2 homolog SIRT7 is an activator of RNA polymerase I transcription. Genes Dev 2006, 20(9): Tsai Y-C, Greco TM, Boonmee A, Miteva Y, Cristea IM: Functional proteomics establishes the interaction of SIRT7 with chromatin remodeling complexes and expands its role in regulation of RNA polymerase I transcription. Mol Cell Proteomics 2012, 11(5): Kim S-I, Bultman SJ, Kiefer CM, Dean A, Bresnick EH: BRG1 requirement for long-range interaction of a locus control region with a downstream promoter. Proceedings of the National Academy of Sciences 2009, 106(7): Group VSR: Prevention of neural tube defects: results of the medical research council vitamin study. MRC vitamin study research group. Lancet 1991, 338: Honein M, Paulozzi L, Mathews T, Erickson J, Wong L: Impact of folic acid fortification of the US food supply on the occurrence of neural tube defects. Jama 2001, 285(23): Harris MJ: Insights into prevention of human neural tube defects by folic acid arising from consideration of mouse mutants. Birth Defects Research Part A: Clinical and Molecular Teratology 2009, 85(4): Marean A, Graf A, Zhang Y, Niswander L: Folic acid supplementation can adversely affect murine neural tube closure and embryonic survival. Hum Mol Genet 2011, 20(18):
118 189. Kelley R, Ideker T: Systematic interpretation of genetic interactions using protein networks. Nat Biotechnol 2005, 23(5): Etheridge SL, Ray S, Li S, Hamblet NS, Lijam N, Tsang M, Greer J, Kardos N, Wang J, Sussman DJ: Murine dishevelled 3 functions in redundant pathways with dishevelled 1 and 2 in normal cardiac outflow tract, cochlea, and neural tube development. PLoS genetics 2008, 4(11):e Kim T, Goodman J, Anderson KV, Niswander L: Phactr4 Regulates Neural Tube and Optic Fissure Closure by Controlling PP1-, Rb-, and E2F1-Regulated Cell-Cycle Progression. Dev Cell 2007, 13(1): Li B, Carey M, Workman JL: The Role of Chromatin during Transcription. Cell 2007, 128(4): Dunaief J, Strober B, Guha S, Khavari P, Ålin K: The retinoblastoma protein and BRG1 form a complex and cooperate to induce cell cycle arrest. Cell 1994, 79: Régnier CH, Masson R, Kedinger V, Textoris J, Stoll I, Chenard M-P, Dierich A, Tomasetto C, Rio M-C: Impaired neural tube closure, axial skeleton malformations, and tracheal ring disruption in TRAF4-deficient mice. Proceedings of the National Academy of Sciences 2002, 99(8): Stottmann RW, Berrong M, Matta K, Choi M, Klingensmith J: The BMP antagonist Noggin promotes cranial and spinal neurulation by distinct mechanisms. Dev Biol 2006, 295(2): Kramer ER, Knott L, Su F, Dessaud E, Krull CE, Helmbacher F, Klein R: Cooperation between GDNF/Ret and ephrina/epha4 signals for motor-axon pathway selection in the limb. Neuron 2006, 50(1): Betancur P, Bronner-Fraser M, Sauka-Spengler T: Assembling neural crest regulatory circuits into a gene regulatory network. Annu Rev Cell Dev Biol 2010, 26: Jin Q, Yu LR, Wang L, Zhang Z, Kasper LH, Lee JE, Wang C, Brindle PK, Dent SY, Ge K: Distinct roles of GCN5/PCAF-mediated H3K9ac and CBP/p300- mediated H3K18/27ac in nuclear receptor transactivation. EMBO J, 30(2): Lin W, Zhang Z, Chen CH, Behringer RR, Dent SY: Proper Gcn5 histone acetyltransferase expression is required for normal anteroposterior patterning of the mouse skeleton. Dev Growth Differ 2008, 50(5):
119 200. Xu W, Edmondson DG, Evrard YA, Wakamiya M, Behringer RR, Roth SY: Loss of Gcn5l2 leads to increased apoptosis and mesodermal defects during mouse development. Nat Genet 2000, 26(2): Lin W, Srajer G, Evrard YA, Phan HM, Furuta Y, Dent SY: Developmental potential of Gcn5(-/-) embryonic stem cells in vivo and in vitro. Dev Dyn 2007, 236(6): Lin W, Zhang Z, Srajer G, Chen YC, Huang M, Phan HM, Dent SY: Proper expression of the Gcn5 histone acetyltransferase is required for neural tube closure in mouse embryos. Dev Dyn 2008, 237(4): Martínez-Cerdeño V, Lemen JM, Chan V, Wey A, Lin W, Dent SR, Knoepfler PS: N-Myc and GCN5 Regulate Significantly Overlapping Transcriptional Programs in Neural Stem Cells. PLoS One 2012, 7(6):e Coles EG, Gammill LS, Miner JH, Bronner-Fraser M: Abnormalities in neural crest cell migration in laminin α5 mutant mice. Dev Biol 2006, 289(1): Fogerty F, Akiyama S, Yamada K: Inhibition of binding of fibronectin to matrix assembly sites by anti-integrin (alpha 5 beta 1) antibodies. The Journal of cell 1990, 111: Patel JH, Du Y, Ard PG, Phillips C, Carella B, Chen C-J, Rakowski C, Chatterjee C, Lieberman PM, Lane WS: The c-myc oncoprotein is a substrate of the acetyltransferases hgcn5/pcaf and TIP60. Mol Cell Biol 2004, 24(24): Mundell NA, Labosky PA: Neural crest stem cell multipotency requires Foxd3 to maintain neural potential and repress mesenchymal fates. Development 2011, 138(4):
120 APPENDIX A FOLATE SUPPLEMENTATION, TESTS FOR POSSIBLE GENETIC INTERACTIONS, AND ADDITIONAL PHENOTYPIC ANALYSES SHOW MINIMAL EFFECTS IN Baf155 msp3 MICE INTRODUCTION Neural tube defects are thought to be multigenic and multifactorial traits suggesting there may be genetic interactions or genetic-environmental interactions that contribute to the incidence of NTD. In this light, it has been found that environmental factors such as diet play a prominent role in NTD prevention. Primary prevention studies of human NTDs in the 1990s found that folic acid consumed periconceptually significantly reduced the incidence of spina bifida in humans [185]. This led to fortification of grains with folic acid (FA) which began in 1998 in the USA, reducing the burden of NTDs by 26% or more [186]; however, still an estimated 2,500 infants annually in the US suffer from NTDs. Mice have been intensely studied as a model system to understand the genetic and environmental interactions and how these factors influence the risk for neural tube defects and etiology [19]. At least 19 mouse lines that exhibit NTD have been tested for FA responsiveness, and these studies have shown varying levels of responsiveness [187, 188]. However, the mechanisms underlying these gene-environment interactions are poorly understood. Genetic interaction can be defined as two mutations that have a combined phenotypic effect not seen by either mutation alone. This powerful genetic experiment can establish functional linkages between genes [189]. Neural tube defects are thought to arise due to disruptions in multiple genetic events and not necessarily a single affected allele. 108
121 Genetic interactions may explain the oligogenic etiology of NTDs as well as functional linkages between the more than 200 genes associated with NTDs in mice. Genetic interactions have been best demonstrated for members of the PCP pathway. Some genetic crosses between heterozygous animals carrying different PCP mutations to generate different allelic combinations, including double heterozygous animals, showed NTDs, whereas single heterozygous mutants may not. This includes Dvl3 +/- ; Ltap Lp/+ at 32% NTD penetrance [190], Fzd2 +/- ;/Vngl2 Lp/+ [166], and, Fzd3 -/- ;Fzd6 -/- [167]. Despite the information gained from these PCP mutants and their role in NTDs, there is still a gap in understanding the genetic combinations that may contribute to NTD. Epigenetic factors may contribute to gene-gene and gene-environment interactions to give rise to NTD etiology by altering the expression of several genes through their roles in regulating genomic structure. The purpose of these studies was to ascertain if the Baf155 msp3 mouse line is responsive to FA, if the Baf155 msp3 allele interacts genetically with other NTD alleles in mice, and to present preliminary analysis on other possible phenotypes of the Baf155 msp3/msp3 mutant. These answers would determine whether future follow-up studies in the Niswander Lab would be warranted. RESULTS Baf155 msp3 Mice Do Not Respond to Folic Acid Supplementation Folic acid (FA) has been shown to reduce NTD prevalence in both mice and humans [187]. To establish whether Baf155 msp3 mice are responsive to FA supplementation and may show prevention of NTDs, I examined the effect of FA supplementation on 109
122 homozygous recessive mice. On regular mouse chow, the Baf155 msp3/msp3 mice display a 81% penetrance of NTDs (92/114) (Table II.1). Table A.1 Phenotype and genotype of embryos resulting from cross between heterozygous Baf155 +/msp3 mice after 9.5 days on a high FA diet Phenotype of Baf155 msp3/msp3 embryos (high FA diet) +/+ +/- -/- exencephaly dev delay normal morphology E E Total On the morning following successful mating between Baf155 +/msp3 heterozygous male and female, the female was placed on a high folic acid diet. After 9 days of gestation, pregnant dams were dissected and embryos were scored for phenotype and genotype. In all cases, the genotype of the embryos correlated with the phenotype, indicating the Baf155 msp3/msp3 homozygous mutant embryos showed 100% penetrant (8/10, with two embryos that were unscoreable) NTDs (Table A.1). From this small sample size, there was no apparent rescue or detrimental effect observed in the Baf155 msp3/msp3 homozygous mutants on the high folic acid diet compared to normal diet. There might be a trend towards a slight detrimental response (100% versus 81% on normal chow), however, this is based on very low sample numbers and only 5 litters examined. Using the Fisher s exact test, the difference in the penetrance of the NTDs on regular and high folic acid diet are not significantly different, with a p-value of 1.0. This suggests that Baf155 msp3 mice are not responsive to dietary FA supplementation after short term exposure. Additionally, a range of phenotypes was observed in mutant mice on both the regular diet and the high folic acid 110
123 diet (Figure A.1 A versus B), again suggesting the mice are not responsive to FA supplementation. Figure A.1 Mice show a range of phenotypes on both regular and high folic acid diet. Lateral views of embryos from pregnant dams fed regular mouse chow (A) or high FA chow (B, 10ppm) at the time of plugging. A) Within a single litter dissected at E9.5, two individual homozygous Baf155 msp3 mutants (MUT1 and MUT2) embryos show a range of phenotypes including failure of neural tube closure and developmental delay compared to wildtype (WT). B) The E9.5 Baf155 msp3/msp3 mutants from mothers fed high FA show exencephaly and developmental delay (MUT 1-MUT3) compared to WT. The range of phenotypic variability is similar in mutant embryos from mothers on either diet. Test for Genetic Interaction between Baf155 msp3 and Other NTD Mutant Lines Crossing a mouse that is heterozygous for recessive mutation in one gene with a mouse heterozygous for a recessive mutation in a different gene and analyzing the phenotype of progeny that carry both mutant alleles provides a test for possible genetic interactions. This can contribute to the understanding of functional linkages between genes. To gain possible new insight into BAF155 function, I tested whether the Baf155 msp3 allele genetically interacts with mutations in three other genes previously associated with NTDs. In all of these cases, single heterozygotes do not exhibit NTDs, so this assay would determine whether double or compound heterozygous embryos display abnormalities. The gene functions tested are as follows: Gcn5/Kat2a is an epigenetic regulator via histone acetyltransferase activity [85], Phactr4 is involved in cell cycle progression [191], and L3P 111
124 is involved in neural patterning and ciliogenesis (although the affected gene has not been identified; Niswander, unpublished). While BAF155 has not been specifically implicated in ciliogenesis or neural patterning, I tested for a possible genetic interaction between L3P and Baf155 msp3. A male carrying the L3P allele and a female carrying the Baf155 msp3 allele were timed-mated to test if there is a genetic interaction between these two alleles. Embryos from two litters were analyzed here, and one double heterozygous mutant embryo recovered which was phenotypically normal. Additionally, three resorbed decidua were recovered, two of which contained tissue from double heterozygous embryos and one of which contained tissue from a Baf155 +/msp3 heterozygote. Although the sample size is too small to draw definitive conclusions, these data might mean that L3P +/- ;Baf155 +/msp3 embryos implant into the uterus but then embryonic development fails. If so, the embryonic lethal phenotype would be partially penetrant. If both embryos and resorptions are considered, there is no significant difference in expected Mendelian ratios using the Chi-squared test. A targeted allele of Gcn5, Gcn5 hat/hat, disrupts histone acetyltransferase activity [84]. It is known that HATs and chromatin remodelers form large complexes that can each associate with the same gene locus to control transcription via their respective epigenetic mechanisms [192]. Thus, it is possible that BAF155 and GCN5 proteins may normally function on the same gene or a similar subset of genes, and if so, Baf155 +/msp3 ; Gcn5 +/hat double heterozygous embryos may show exencephaly or other abnormalities. Three litters were dissected from this genetic cross, and five Gcn5 +/hat ;Baf155 +/msp3 double heterozygous embryos were recovered (Table A.2). Of the five mutant embryos, none exhibited altered phenotypes compared to single heterozygous embryos, which have been 112
125 documented to be viable and fertile [84, 141]. Moreover, no resorptions were identified at E9.5 and the number of single heterozygous and double heterozygous embryos compared to the WT is not statistically different based on the Chi-squared test, suggesting there was no loss of embryos prior to dissection. These data suggest that Baf155 msp3 and Gcn5 hat do not genetically interact prior to or at E9.5. Table A.2 Phenotype and genotype of embryos resulting from crosses between heterozygous Baf155 +/msp3 female mice and other heterozygous male mice. Lines crossed (Heterozygotes) Stage recovered Observed embryo ratios Resorptions Observed het/het phenotype L3p/Baf155 E9.5 +/+ het/+ +/het het/het non-phenotypic, 2 resorbed Gcn5/Baf155 E9.5 +/+ het/+ +/het het/het non-phenotypic Phactr4/Baf155 adult +/+ het/+ +/het het/het n/a non-phenotypic Phactr4 encodes a phosphatase that interacts with retinoblastoma (RB) in promoting cell cycle progression during neural tube closure [191]. BRG1, the ATPase in the BAF chromatin remodeling complex, has also been implicated in RB interactions and inducing cell cycle arrest [193]. Therefore, the Phactr4 phosphatase and the Baf155 chromatin remodeling protein may genetically interact in mice. A Phactr4 humdy male (ENU-induced allele[191]) was mated to two Baf155 mps3 females. This yielded 8 pups that survived until weaning. Out of the 8 pups, 4 were genotyped as double heterozygous at the Phactr4 and Baf155 loci. Three mutants out of 8 total pups is within the expected 113
126 Mendelian ratio based on the chi-squared test, suggesting there was not a loss of double heterozygous embryos during pregnancy or prior to weaning. Despite the small sample size, the fact that Baf155 and Phactr4 double heterozygotes are viable suggests these genes do not genetically interact to affect embryonic development. Additional Phenotype Characterization of the Baf155 msp3/msp3 embryos Skeletal malformations are commonly associated with NTDs [77, 194, 195], therefore we examined skeletal formation with Alcian Blue/Alizarin Red (AB/AR) staining for cartilage and bone in WT and Baf155 msp3/msp3 embryos at E15.5. There are no fully formed frontal and parietal bones in the cranium of the mutant embryo compared to the WT, presumably a consequence of the exencephaly phenotype (Figure A.2). Other than this obvious defect, there are no other apparent skeletal formation defects in the mutant. This suggests that the Msp3 mutation does not affect skeletal patterning. This staining was performed on one WT and two phenotypic mutants, and therefore should be completed in at least one more stage-matched biological replicate to validate these findings. In chapter 4, RNA-Seq revealed that several genes known to regulate neurogenesis at stages later than E9.5 are misregulated in the Baf155 msp3 mutant embryos. To investigate if neuron growth is altered in the mutant embryos at later stages, we examined axon projections using neurofilament staining in mutant and WT embryos at E11.5. The WT embryo show organized dorsal root ganglia (DRG) and enteric plexus (EP). (Figure A.3B). The Baf155 msp3/msp3 homozygous mutants show no apparent difference in the DRG, enteric plexus or axon projections in general compared to the WT. This study at E11.5 suggests that there may not be later defects in neuronal outgrowth in the Baf155 msp3 114
127 mutant. This study was performed on one biological replicate, and therefore needs to be repeated. Figure A.2 Baf155 function is not necessary for skeletal patterning Lateral views of WT (left) and Baf155 msp3/msp3 mutant (right) E15.5 embryos stained with Alcian Blue for cartilage and Alizarin Red for bone. Insets: WT (left) and mutant (right) embryos prior to staining. FB= Frontal Bone; MX= Maxilla; PB= Parietal Bone; OB= Occipital Bone CONCLUSION NTDs are the second most common birth defect in humans. FA supplementation has been shown to reduce the incidence of NTDs in humans and mice; however, there is a lack of understanding of how epigenetic mechanisms inform the gene-environment and gene-gene interactions. A hypomorphic mutation in the BAF155 chromatin remodeling protein, which in mice results in 81% penetrant NTD, can contribute to the understanding of gene-environment and gene-gene interactions. The data presented here suggests that the Baf155 msp3 is not responsive to dietary folic acid supplementation. Additionally, I conducted genetic experiments to determine whether Baf155 msp3 interacts with three other NTD mouse lines in the Niswander Lab: Gcn5 hat, L3P and, Phactr4 humdy. 115
128 Figure A.3 Baf155 function is not necessary for neuron outgrowth. Lateral views of whole mount neurofilament staining in the trunk region of E11.5 WT (left) and Mutant (right) FL= Forelimb; HL= Hindlimb; EP= Enteric Plexus; DRG= Dorsal Root Ganglia Of these mouse lines tested for genetic interactions with Baf155 msp3, only L3P +/- ;Baf155 +/msp3 double heterozygotes showed a potential interaction in the form of early embryonic lethality. My studies were small in scale to provide preliminary evidence to determine if more extensive experiments were warranted. However, the small sample size causes difficulty in interpreting some of the experiments, the implications of which will be discussed below. In the studies conducted between Baf155 msp3 and L3P, there might be a trend towards a slight detrimental response. Using the powers sample size calculator, verification of a 19% increase in penetrance from 81% to 100% difference, or a slight detrimental response, requires recovering at least 49 embryos. It was determined that using this number 116
129 of mice to identify at best a minor detrimental effect would not be a productive use of resources. There have been at least 19 mouse lines tested for FA supplementation responsiveness. The responses range from a beneficial response, no response or a detrimental response. NTD prevention (or beneficial response) ranged from 12% to 85% decrease in NTD incidence in the tested mouse lines [187, 188]. Given the small numbers obtained in this thesis, a beneficial effect of FA supplementation in Baf155 msp3 mice cannot be ruled out. However, in order to ascertain if there is in fact a beneficial effect, or a decrease in NTD penetrance between mice on regular diet and those on high FA diet, a 15% difference between the two sample populations would be needed to indicate statistical significance. Using a power sample size calculator, verification of a 15% decrease in penetrance, or a 15% rescue, requires recovering at least 46 embryos. At this time, the actual penetrance cannot be determined but if there was a strong beneficial response (85%), we may have observed Baf155 msp3 homozygous mutants without NTD, which we did not in our small sample size. While a beneficial response cannot be ruled out, my preliminary studies do not suggest this avenue would be productive. Therefore, these preliminary studies suggest Baf155 msp3 mice show no difference in NTD rates after a short term of exposure to high FA diet compared to regular diet. Genetic interaction experiments suggest the genetic component of L3P and BAF155 may share common pathways or potential genetic targets that are not immediately apparent, but resorptions seen at E9.5 suggest possible defects in gastrulation. L3P mutants are defective in Shh signaling, but microarray experiments do not reveal other pathways as significantly misregulated. RNA-Seq of BAF155 cranial tissue does not show 117
130 misregulation of genes in the Shh pathway. It is possible that the subsets of genes that may contribute to the early embryonic death phenotype are not misregulated in the single homozygous mutants, but rather misregulation is compounded in the double heterozygous embryos. Given that L3P shows defects in ciliogenesis, an informative study would be to cross Baf155 msp3 with other cilia mutants to test if BAF155 genetically interacts with ciliogenesis pathways. Surprisingly, there was no genetic interaction seen in the Baf155 +/msp3 ;Gcn5 +/hat double heterozygotes given they are both epigenetic regulators or in the Baf155 +/msp3 ;Phactr4 +/humdy double heterozygotes given a common RB pathway interaction. Despite the lack of interaction with GCN5, this does not indicate that the BAF complex and GCN5 do not interact at the same gene targets during neural tube closure. However, it does suggest that the subset of genes affected by the BAF155 msp3 mutation and the GCN5 hat mutation may not overlap. Phactr4 +/humdy ;Baf155 +/msp3 double heterozygotes are viable, but fertility was not tested in these mice. Exencephaly is usually embryonic lethal due to secondary affects, or pups are cannibalized upon birth by the mother, and therefore not viable. Therefore, the Phactr4 +/humdy ;Baf155 +/msp3 mice likely do not exhibit exencephaly or other double heterozygous embryonic lethal phenotypes, but this scenario cannot be ruled out considering the small sample size tested. It is unlikely in light of the normal Mendelian ratios and high incidence of viable double heterozygotes. The low number of embryos tested makes it difficult to conclude whether BAF155 interacts with other genes during the process of neural tube closure. Moreover, our studies do not test whether the genes may genetically interact to control later embryonic or adult processes. This would require in-depth analysis of later embryonic and adult stages in the 118
131 double heterozygous animals. Technical limitations of the bone and cartilage staining in the WT and BAF155 msp3/msp3 embryos are that the tissue was not cleared sufficiently so a detailed analysis of the skeletal structure in the mutant compared to the WT could not be conducted. MATERIALS AND METHODS High Folic Acid Diet Protocol For high folic acid diet exposure, heterozygous Baf155 +/msp3 males and females were mated. After a vaginal plug was observed, the female was placed on a high folic acid diet (10ppm; Research Diets, Inc. D ; ppm is equivalent to mg/kg of chow and is not based on body weight). The normal diet contains 4 ppm (Teklad Global Soy Protein- Free Extruded Rodent Diet, 2020X). After 9 days of gestation, pregnant dams were dissected and embryos scored for phenotype and genotype. NTD penetrance in embryos on high FA diet was compared to embryos on Teckland Rodent chow. Fishers exact test was used to evaluate differences in the rate of NTDs observed in mutant embryos. Chi squared goodness-of-fit-test was used to assess significance between the expected and observed Mendelian ratios in embryo loss. Genetic Interactions Protocols Females from one line and males from a different line were mated. Briefly, heterozygous males from the L3P, Gcn5 hat and Phactr4 humdy lines were time-mated with Baf155 +/msp3 heterozygous females. Embryos were recovered after 9.5 days of gestation, scored for phenotype and genotype. To isolate DNA, a portion of the yolk sac was harvested from embryos or tail tip was harvested from adult mice and lysed in PCR Lysis 119
132 Buffer (50 mm Tris (ph 8.8, 1mM EDTA, 0.5% Tween-20 and 0.2 mg/ml proteinase K). Samples were digested overnight at 56 C and heat inactivated at 95 C for 10 minutes. Phactr4 mice genotyping: Genomic DNA from mouse tails was sequenced by the University of Colorado DNA Sequencing and Analysis Core ( using traditional Sanger sequencing with the following primer: 5 TCTTGCAGCTCAGTC 3. Briefly, after the primer annealed to the DNA, 3 primer extension was used to elongate the DNA with fluorescently labeled bases terminally labeling each strand of newly synthesized DNA. The newly synthesized DNA was separated using capillary electrophoresis and the DNA sequence was then called based on the fluorescently labeled terminal DNA bases. The wild type Phactr4 gene contained a G at locus 1949, and could be identified. The G to C transversion was identified in heterozygous embryos at base 1949 of the Phactr4 gene [191] shown by a peak that was half as tall as the WT peak, and yielded both G and C in the trace. Gcn5 hat/hat embryo genotyping: Gcn5 hat/hat primers and PCR program to identify the D609A mutation in exon 13 are listed below. DNA was isolated as above, however, the genomic DNA was phenol-chloroform extracted and diluted in 500ul of pure molecular grade water. Standard Niswander genotyping for the mix was then used- 5 ul diluted DNA, 10mM primers, 6 mm MgCl, 1x PCR buffer. 5 ACTCAAGTGCTGAGTCTGGC 3 and 5 CCCCCAAAGGACCTTCCAAT 3. Gcn5 hat/hat PCR reaction for genotyping: C for 5 minutes C for 1 minute C for 1 minute C for 1 minute 5. Repeat steps 2 through 4 for 35 cycles total C for 5 minutes 7. 4 C hold 120
133 PCR products are digested for 1 hour with the restriction enzyme Pst1 and visualized on a 1% agarose gel. Pst1 cuts at CTGCAG, which creates a smaller product for mutant alleles as it only cuts mutant genomic DNA. WT yields a single band at 424 base pairs, while mutant alleles yielding bands at 223bp and 202bp [84]. L3P primers and PCR program: D4dmm35 Forward: D4dmm 35 Reverse: D4dmm139.5a Forward: D4dmm139.5a Reverse: CGACTTCAGGGAGTGTGACC CGGTCACATGGTATCTGCTG CGTGTGTATGAGCCACACATT GTCCAGGTCCTTCTCCCTTC L3P PCR reaction for genotyping: C for 12 minutes C for 20 seconds C for 30 seconds C for 45 seconds 5. Repeat steps 2 through 4 for 56 cycles total C for 7 minutes 7. 4 C hold PCR products are visualized on a 4% agarose gel. D4dmm35 primers are specific for the WT allele and yield a PCR product with a size of 273bp, while D4dmm139.5a primers are specific for the mutant allele and yield a size of 146bp. Neurofilament Staining of Whole Mount Embryos This protocol was revised from Kramer et al. [196]. Briefly, embryos were fixed in 4% Paraformaldehyde (PFA) for 20 minutes and postfixed in methanol in a stepwise manner to dehydrate the embryos. Then, embryos were fixed in Dent s fixative [4 parts 121
134 100% methanol (MeOH), 1 part DMSO] for 16 hours at 4ºC. Embryos were dehydrated twice in 100% MeOH for 30 minutes and subsequently subjected to five 30 minute cycles of freezing at -80ºC and thawing to room temperature (RT) in 100% MeOH. Embryos were then incubated at -80ºC for 3 days in fresh 100% MeOH and rehydrated for 90 minutes each in 50% MeOH, 15% MeOH, and PBS at RT. Three wash steps for one hour each at room temperature were then followed by incubation with the primary neurofilament antibody (Abcam, ab65845)(1:500 in blocking serum) for 16 hours at RT. After washing 5 times in TBS/1% Triton X-100/0.2% gelatin for one hour each, the embryos were incubated with a Bio-Rad goat anti-rabbit HRP secondary antibody for 16 hours at RT. The secondary antibody was diluted 1:500 in blocking serum (100 mm Tris- HCl/150 mm NaCl/5% Blocking Reagent (Perkin Elmer)). Finally, embryos were washed as above and developed in diaminobenzidine working solution (DAB, Sigma) followed by sequential dehydration in MeOH and clearing in 1:2 benzyl alcohol to benzyl benzoate (BABB). Bone and Cartilage Staining (Alcian Blue/Alizarin Red Staining) Briefly, embryos were desensitized in ice-cold 95% ethanol for 1 hour and dehydrated in 95% ethanol over night at room temperature. Embryos were then stained overnight rocking at room temperature in freshly prepared stain solution of.05% alcian Blue, 10% glacial acetic acid, and 65% ethanol and 0.005% alizarin red. Next, embryos were rinsed in water and tissue was dissolved in 2% KOH overnight. Embryos were cleared in 20% glycerol/1 % KOH for 1 hour, 33% glycerol/1% KOH for 1 hour, and finally 50% glycerol/1% KOH for 3 days. 122
135 APPENDIX B GCN5 ACETYLTRANSFERASE ACTIVITY FUNCTIONS IN NEURAL CREST CELL BIOLOGY INTRODUCTION During embryonic neural tube formation, many neural cell types become specified and begin to differentiate. One of these cell types, the neural crest cells (NCCs), are specified within the early neuroectoderm by expression of SOX10, BMP, and WNT signaling molecules in the mouse [197]. The NCCs undergo an epithelial-to-mesenchymal transition (EMT), delaminate, and migrate out from the dorsal neural tube to specific sites in the embryo [151]. NCCs in the rostral trunk migrate out of the dorsal neural tube at various times and in different waves, giving rise to structures that include the dorsal root ganglia, sympathetic ganglia, and craniofacial structures. While the transcriptional regulation of NCC specification and EMT is relatively well studied, there is little known as to the epigenetic control of NCC specification and behavior. GCN5 (KAT2a) is a lysine acetyltransferase known to acetylate histone H3 lysine 9 (H3K9) and lysine 18 (H3K18) [198, 199]. Depletion of GCN5 causes embryonic lethality at E7.5 due to severe defects during gastrulation [200]. Mice that harbor hypomorphic alleles for Gcn5 have defects in neural tube closure and patterning of the trunk skeleton [201, 202]. Catalytically inactive (Gnc5 hat/hat ) mutant embryos exhibit neural tube defects at a high penetrance. It is also known that GCN5 controls neural stemness [203] and exhibits p53-dependent activities [84]. This chapter focuses on a mouse model with a catalytically inactive histone acetyltransferase domain. Data presented here indicates there are potential NCC defects relating to specification and migration. 123
136 These studies are an extension of the RNA microarray analysis of the cranial tissue of E9.5 Gcn5 hat/hat embryos that Jonathan Wilde performed in the Niswander lab. He discovered dysregulation of several genes involved in neural crest cell biology. Many of the genes were validated by q-pcr including Lama5, Cxcl12 (Sdf1), Cxcl13, EphA3, Evi1 (PRDM3), Prdm16, and Prmt8. Additionally, he found that expression of the neural differentiation marker Tuj1 is decreased in the dorsal root ganglia. Moreover, dorsal root ganglia in the rostral trunk are approximately 47% smaller than WT after normalization to correct for the growth restriction seen in the mutants. Using the Gcn5 hat/hat mouse line, I tested the hypothesis that GCN5 acetylation activity regulates NCC specification, migration and craniofacial development. RESULTS Gcn5 hat/hat Mutant NCCs Migrate Farther than WT Neural crest cells are specified and migrate out of the neural tube as it is forming between E8.5-E10.5 [134]. To understand the functional consequences of ablating HAT activity on NCC migration, I used neural tube explants from E10.5 mutant and WT embryos. I analyzed the number of cells and distance the cells migrated in Gcn5 hat/hat neural tube explants compared to WT explants. I found that mutant NCCs showed a greater number of cells that migrated out of the neural tube, and the distance migrated was farther compared to WT neural tube explants. This indicates that the migratory capacity of Gcn5 mutant NCCs is greater ex vivo than WT (Figure B.1A, B). This experiment was only completed on two neighboring regions of the neural tube from one mutant and one WT, so more biological replicates are needed to confirm these results. 124
137 Figure B.1 Gcn5 hat/hat mutants show NCC migration and specification defects. A) WT neural tube explant (darker dense tissue) in which the extent of NCC migration is indicated by the red dotted line. B) Mutant neural tube explant shows a significant increase in both NCC number and the extent of migration. C) Quantification of the number of cells leaving the NT, n=2. D) Quantification of the linear distance traveled from the NT measured at 4 perpendicular locations in each of two replicates. E and F) Lateral views of whole mount RNA in situ hybridization on E10.5 WT (E) and Gcn5 mutant (F) embryos with probe to NC marker Sox10. Red arrows show broadening domains in jugal ganglia (left) and trigeminal ganglia (right). (*) P<0.01, (**) P<0.05 Student s t-test. Sox10 Expression Is Disrupted in the Gcn5 hat/hat Mutant In mouse, NCCs must be specified prior to migrating from the neural tube. One molecule that is known to be required for specification, and to persist during migration and differentiation of NCCs is Sox10 [197]. To understand if the specification, migration or differentiation of neural crest cells was altered, we examined expression of Sox10 in mutant and WT in whole mount E10.5 embryos. Expression in the trigeminal region is expanded posteriorly in the mutant compared to the WT embryo (Figure B.1 E, F). Additionally, expression in the developing brachial plexus nerve bundle was expanded in the Gcn5 hat mutant compared to the WT. These domains that are expanded should 125
138 represent the mesencephalic migration stream and the 4 th rhombomere migration stream, or the cranial neural crest population and the vagal neural crest population, respectively. The Sox10 experiments were repeated in two total biological replicates. These data combined with the increased migration seen in neural tube explant experiments suggest that NCC migration and/or differentiation may be disrupted in the Gcn5 hat/hat embryos. Figure B.2 Skeletal stainings of E16.5 Gcn5 hat/hat mutants A) Lateral view of WT skeleton stained for bone (red) and cartilage (blue). B) Gcn5 hat/hat mutants exhibit a lack of the jugal bone and mislocalization of the squamosal (*). C) Ventral view of the WT mandible. D) Ventral view of the mutant mandible shows a medial-lateral thickening and lateral processes that are not present in the WT. j=jugal; md=mandible; mc= Meckel s cartilage; mx=maxilla; s=squamosal Craniofacial Structures Are Mislocalized and Malformed in the Gcn5 hat/hat Mutant Neural crest cells give rise to facial structures including the jugal and the squamosal bones. Thus, I asked if the craniofacial structures might be altered in Gcn5 hat/hat mutant embryos at the latest stage that they survive, E16.5. Skeletal structures were stained 126
139 for bone using Alizarin Red (AR) and stained for cartilage using Alcian Blue (AB). They were then compared in stage-matched E16.5 Gcn5 +/+ and mutant Gcn5 hat/hat embryos. Mutants show loss of the jugal bone and mislocalization of the squamosal bone compared to the WT. Gcn5 hat/hat mutant mandibles at E16.5 exhibit overall thickening with an ectopic process on the lateral edge (Figure B.2). Taken together, these data suggest GCN5 acetylation activity may function in migration or differentiation of NCCs that give rise to craniofacial structures. CONCLUSION The epigenetic roles in NCC specification, migration and differentiation are not well understood. An epigenetic regulator, GCN5, is a lysine acetyltransferase that interacts in several large complexes in the nucleus, and preliminary data suggests it may play a role in aspects of NCC function. Microarray data generated in the Niswander lab by Jonathan Wilde, which was derived from E9.5 WT and Gcn5 hat/hat cranial tissue, indicated that abolished GCN5 acetylation activity leads to misregulation of several genes involved in the NCC gene regulatory network [197]. The data presented here indicates that NCCs from Gcn5 hat/hat mutants have altered migration capacity compared to WT, altered Sox10 domain in the cranial NCC and vagal NCC populations, and malformed craniofacial structures later in development. While these defects strongly suggest that migration and differentiation of NCCs are functionally altered, the molecular mechanisms are not clear. The Sox10 stained whole mount embryos show a broadened expression domain in the 4 th rhombomere migration stream and mesencephalic migration stream. The localization and extent of Sox10 staining can vary significantly even in WT embryos (T. Williams, personal communication), and therefore two biological replicates may not be 127
140 sufficient to conclude that there is a defect in Sox10 localization in the Gcn5 hat/hat mutant. The studies should be repeated to verify the results. In addition to repeating the Sox10 staining, it is necessary to understand which molecular cues are aberrant and may be contributing to the NCC defect. The craniofacial abnormalities suggest that differentiation may also be disrupted in GCN5 mutant. Thus, staining with more molecular markers of neural crest cells will lend insight into what stage of NCC development might be disrupted in the GCN5 mutant. To this end, one could characterize markers of neural crest cells both temporally and spatially using Tfap2, Bmp2 and Wnt1for specification; N-Cadherin and Cadherin 6b for epithelial to mesenchymal transition (EMT); Sema3a for repulsive NCC signaling; and Snail for motility genes in E9.5 and E10.5 tissue sections of Gcn5 hat/hat mutants compared to WT. For differentiation, one can characterize Dlx 2, 3 5 and Hand2 for genes that contribute to differentiation into craniofacial structures. These studies will aid in understanding the gene regulatory network in the NCCs in the GCN5 mutant and contribute to understanding the molecular basis for how GCN5 HAT activity controls NCC biology. The increase in cell migration from the neural tube in explant cultures may be due to misregulation of genes discovered in the microarray study, specifically the increase in Lama5, encoding the LAMININ 5 receptor, which is a cell adhesion molecule expressed on neural crest cells [204]. To test if the increased migration distance is due to an increase in LAMININ 5, one could block LAMININ 5 receptor function using an antibody [205] in the media of the mutant NT explants. If this were the case, I would expect that inhibiting the function of LAMININ would decrease the migration of mutant NCC. If LAMININ function does not play a role in migration, I would expect that inhibition of LAMININ 5 128
141 would have no effect on the number of cells migrating from WT or mutant neural tube explants, nor on the distance migrated. Future directions also include identifying the molecular targets of GCN5 in the specification and migration of the NCCs. Some of these direct or indirect targets have been suggested in the microarray analysis completed in E9.5 mutant and WT heads, which included a small population of NCCs. ChIP would be an informative experiment to indicate if the molecular change in expression is due to a direct interaction between GCN5 and the gene of interest or an indirect interaction. This would indicate the genomic interactions of GCN5. However, it is known that GCN5 not only acetylates histones, but can also acetylate many proteins including c-myc [206] and p53 (S. Dent, unpublished studies, [84]). Therefore, GCN5 may function posttranslationally to acetylate a transcription factor or signaling molecule, and loss of GCN5 HAT activity could disrupt the function of the protein and lead to NCC dysfunction that is unrelated to chromatin epigenetic function of GCN5. In summary, GCN5 HAT activity has functional roles in NCC biology; however, the cellular and molecular basis for those functions requires more characterization. MATERIALS AND METHODS Neural Tube Explants NT explant protocol was based on Mundell, 2011 [207]. Briefly, the vagal and trunk regions of the neural tube (otic placode to somite 4, and somite 5- somite 12) was micro-dissected from mouse embryos at day E10.5 and isolated from surrounding tissue using Dispase at 1mg/ml (Roche). Isolated neural tube tissue was plated on a dish coated with 30 ug/ml fibronectin and cultured in self-renewal media at 3-4% oxygen and 5% CO 2 129
142 for 2 days. Self-renewal media: Dulbecco s modified Eagles Media low glucose (Invitrogen) 30% neurobasal media (Invitrogen), 15% chick embryo extract (CEE, Gemini Bio Products), 2% B27 (Invitrogen), 1% N2 (Invitrogen) 117 nm retinoic acid (Sigma), 50uM 2-mercaptoethanol (Sigma), 20ng/ul Insulin-like Growth Factor (IGF), and 20 ng/ml basic Fibroblast Growth Factor (bfgf). Sox10 Staining See Chapter II, Materials and Methods- Analysis of Mutant Phenotypes Bone and Cartilage Staining (Alcian Blue/Alizarin Red Staining) See Appendix A, Materials and Methods- Bone and Cartilage Staining (Alcian Blue/ Alizarin Red Staining) 130
143 131 APPENDIX C TABLE OF MISREGULATED GENES FROM RNA ISOLATED FROM E9.5 CRANIAL NEURAL TISSUE OF BAF155 MSP3/MSP3 EMBRYOS Table C.1 List of misregulated genes in cranial tissue from averaged WT and averaged Mutant E9.5 embryos based on EdgeR analysis. Top list comprises upregulated genes, bottom list comprises downregulated genes. FC=Fold change >1.5; FDR=False Discovery Rate >0.05. Nr=Normalized Category: Neural development= ND, Apoptosis= AP Cell survival/cell Death=CS Centrosome/cilia=CN Global regulators= GR Adhesion/Cell polarity=ad Growth and proliferation, cellular development, and cell morphology= GP Upregulated Genes: Gene FC PValue FDR Category WT1_Nr WT2_Nr WT3_Nr Mut1_Nr Mut2_Nr Mut3_Nr Top2b E ND, GR, GP Kif E E-11 ND, CN Metrnl E E Spon E E-13 ND
144 132 Abcg E E-08 GP Cxcl E Fzd E E-05 ND Fam72a AP, CS Sirt E E-14 AP, GR Sema3c E ND, CS Coil E Nfkb E E-07 ND, CS Fam71d E E Sst E E-14 ND, AP, CS Pcbp E E-20 CS Stxbp ND Cand GP Dgkd E E Plcg E ND, CS Map3k E E-16 ND, GP, CS Tspan E E Lmo E CS Zfp Sez6l Ctse E AP, CS Ppat E E Ctnna E E-05 ND, CS Adora E AP, CS Ndfip ND, AD Bcl2a1d CS Srgap E ND Samd
145 133 Tsta ND Slc22a Glyctk E Adipor E AD Kcnmb E Cyb5r E E Epha E E-06 AD, GP Oca E CS Ccdc E Rassf E E Ryk ND Nat E CS Dnajc E E-05 GP Maf Ccdc E Cnga E Sult6b E Cdk ND Nipal E Fzd E ND Frem ND, AD Ablim E ND H2-DMa E E-05 CS, AD St E GP Hsf E E-09 ND, GP Thtpa E GP Enthd E Tpm E CS Wdfy E Pla2g12b E E-07 AP, GP
146 134 Nts E CS Itga E E-05 ND, CS AD Nars Creb Card E GP Secisbp E GP Ube2e GP Erbb E AD, GP Hfm E Utp11l E Faf E AP, GP Gjc E AD Dock E ND, CS Pisd-ps E Iqcf
147 135 Downregulated Genes: GeneID FC PValue FDR Category WT1_Nr WT2_Nr WT3_Nr Mut1_Nr Mut2_Nr Mut3_Nr Chd E ND Cldn E AD, GP Abca8b E Vsig E Hbp GP Svep E Cyp4f E E Ints Abl E AD, GP Elac ND Pla2g E E Rnf E CS Dhx E E-14 ND Rassf E E Ms4a Apob E CS Gatad2b E E-06 ND Col15a E E Mep1a E CS Tusc E AP Aqp E E Pcp4l E Poc1a E E-53 CN
148 APPENDIX D TABLE OF THE OVERLAP OF MISREGULATED GENES IN CRANIAL NEURAL TISSUE Table D.1 Misexpressed genes in individual Baf155 msp3/msp3 mutants Twenty-seven overlapping genes misexpressed in cranial tissue of each individual mutant sample compared to the average of all 3 WT samples expressed as a log2 fold change. 136
149 APPENDIX E LAURA HARMACEK S JOURNAL ARTICLE FROM JOURNAL OF DEVELOPMENTAL NEUROBIOLOGY, WILEY ONLINE LIBRARY, 2013: A unique missense allele of baf155, a core BAF chromatin remodeling complex protein, causes neural tube closure defects in mice Laura Harmacek 1, Dawn E. Watkins-Chow 2, Jianfu Chen 1, Kenneth L. Jones 3, William J. Pavan 2, J. Michael Salbaum 4, and Lee Niswander 1,* 1. Howard Hughes Medical Institute, Department of Pediatrics and Graduate Program in Molecular Biology, University of Colorado Anschutz Medical Campus, Children s Hospital Colorado, Aurora, CO 80045, USA 2. Genetic Disease Research Branch, National Human Genome Research Institute, National Institutes of Health, Bethesda, MD, USA 3. Department of Biochemistry and Molecular Genetics, University of Denver Anschutz Medical Campus, Aurora CO, USA 4. Department of Regulation of Gene Expression, Pennington Biomedical Research Center, 6400 Perkins Road, Baton Rouge, LA 70808, USA *Correspondence: [email protected] Mailstop 8133, Building RC1 South, Room L , East 17th Avenue, Aurora, CO (phone); (FAX) KEY WORDS Neural tube defect; epigenetic regulation; Baf155 msp3 ; apoptosis; proliferation. 137
150 RUNNING TITLE Baf155 regulation of neural tube closure ACKNOWLDEGMENTS We thank members of the Niswander lab for helpful discussions throughout this project. We thank Dr. Jim Huntley (UC Boulder) for assistance in RNA-Seq library preparation. Antibodies obtained from the Developmental Studies Hybridoma Bank were developed under the auspices of the NICHD and maintained by The University of Iowa, Department of Biology, Iowa City, IA Thank you to Jennifer Moran and the Mouse Mutant Resequencing Initiative for help with exon-based hybrid selection. Grants: The research was supported by the Department of Pediatrics, a Cancer Center Support Grant (P30CA046934), the Bioinformatics Shared Resource, and the CCTSI. L.N. is an Investigator of the Howard Hughes Medical Institute. ABSTRACT Failure of embryonic neural tube closure results in the second most common class of birth defects known as neural tube defects (NTDs). While NTDs are likely the result of complex multigenic dysfunction, it is not known whether polymorphisms in epigenetic regulators may be risk factors for NTDs. Here we characterized Baf155 msp3, a unique ENU-induced allele in mice. Homozygous Baf155 mps3 embryos exhibit highly penetrant exencephaly, allowing us to investigate the roles of an assembled, but malfunctional BAF chromatin remodeling complex in vivo at the time of neural tube closure. Evidence of defects in proliferation and apoptosis were found within the neural tube. RNA-Seq analysis revealed that surprisingly few genes showed altered expression in Baf155 mutant neural tissue, given the broad epigenetic role of the BAF complex, but included genes involved in neural development and cell survival. Moreover, gene expression changes between individual mutants were variable even though the NTD was consistently observed. This suggests 138
151 that inconsistent gene regulation contributes to failed neural tube closure. These results shed light on the role of the BAF complex in the process of neural tube closure and highlight the importance of studying missense alleles to understand epigenetic regulation during critical phases of development. INTRODUCTION Neural tube defects (NTDs) result when the embryonic neural tube, which gives rise to the adult brain and spinal cord, fails to close completely. NTDs are the second most common birth defect in humans, occurring in 1 in 1000 live births worldwide (A. Copp, 2005; Detrait et al., 2005). NTDs include caudal neural tube closure defects such as spina bifida, and cranial neural tube defects such as exencephaly. These devastating birth defects occur due to disruption in the intricate process of neural tube closure, which requires the coordination of many molecular and cellular functions. The complexity of this process is illustrated by the fact that over 200 mutations in the mouse have been identified that result in NTDs (M. Harris and D. Juriloff, 1999; M. J. Harris and D. M. Juriloff, 2010). These affected genes play roles in multiple processes, including cell cycle regulation, neurogenesis, cell viability, developmental signaling, and epigenetic regulation of transcription (A. J. Copp and Greene, 2010). Despite the body of knowledge from studies in animal models, very few gene mutations have been identified in human NTDs using a single gene approach. Moreover, based on twin studies and epidemiological studies, NTDs in humans are likely the result of multiple genetic alterations and therefore many genes would be expected to contribute to this complex disease etiology (Elwood et al., 1992; Leck, 1974). It is possible that studying global gene modifiers might contribute to our understanding of the causation of NTDs. Epigenetic mediators represent a good candidate for global gene expression modifiers, and their role in the NTD disease etiology has been poorly characterized. 139
152 DNA exists in the nucleus as chromatin, a highly ordered structure which consists of organized units called nucleosomes - DNA wrapped around a histone protein octomer (Luger et al., 1997). Epigenetics refers to heritable changes in gene transcription that are independent of DNA sequence, most commonly thought of at the level of chromatin. There are two main types of epigenetic processes covalent modifications that are directly added to the chromatin (DNA methylation and histone modifications) and movement of nucleosomes along the DNA (chromatin remodeling). Chromatin remodeling requires an ATPase to hydrolyze ATP for energy to disrupt histone-dna contacts and move chromatin proteins along the DNA. Overall, this process alters chromatin structure and changes DNA accessibility to transcriptional regulatory factors (Clapier and Cairns, 2009). The dynamic process of neural tube closure requires rapid changes in gene expression which might be achieved through chromatin remodeling. NTDs can arise as a consequence of mutations in genes that mediate epigenetic regulation (M. J. Harris and D. M. Juriloff, 2010). Of the >200 mouse NTD mutants, ~5% involve disruptions in genes encoding epigenetic regulators, which are involved in all facets of epigenetic regulation. This includes DNA methylation (Dnmt3l (Hata et al., 2002), Dnmt3b (Okano et al., 1999)), histone methylation or acetylation (Kat2a [Gcn5] (Bu et al., 2007), Cbp (Tanaka et al., 2000), p300 (Yao et al., 1998), Hdac4 (Vega et al., 2004), Sirt1 (Cheng et al., 2003)), and chromatin remodeling (Smarcc1 (J. K. Kim et al., 2001), Smarca4 (Bultman et al., 2000), Cited2 (Bamforth et al., 2001; Dunwoodie et al., 1998), Cerc2 (Banting, 2004)). Smarcc1 and Smarca4 encode BAF155 and BRG1, respectively, core components of an ATPdependent chromatin remodeling complex. The mammalian BRG1/BRM Associated Factor ATP-dependent chromatin remodeling complex (BAF complex) contains 15 protein subunits encoded by 26 genes (Ronan et al., 2013)and is part of the Swi/Snf family of chromatin remodelers originally described in 140
153 yeast (Nasmyth, 1987). In many organisms, including mice and humans, investigation of the BAF chromatin remodeling complex in different cell types indicates significant heterogeneity in subunit association. For example, the BAF complex is composed of different protein isoforms in embryonic stem (ES) cells, developing cardiomyocytes, and neural progenitor cells, suggesting there are tissue and cell-type specific roles for the complex during development (Ho and Crabtree, 2010). However, the core components of the complex, ATPase BRG1 or BRM, along with BAF155, BAF170, and BAF47/INI5, have been isolated from all cell types studied to date and can remodel nucleosomes in vitro at the same efficiency as the fully intact BAF chromatin remodeling complex (Phelan et al., 1999). This core set of BAF proteins is particularly important in vivo, as complete loss of BRG1, BAF155 or BAF47 all result in peri-implantation embryonic death and heterozygous loss of expression can lead to variable penetrance cranial NTDs (Bultman et al., 2000; Ho et al., 2009; J. K. Kim et al., 2001; Mandel and Gozes, 2007). The core component BAF155 (encoded by the Smarcc1 gene) also plays an important role in maintenance of the BAF complex. BAF155 protects BAF complex proteins from degradation and maintains their nuclear localization (Chen and Archer, 2005; Sohn et al., 2007). BAF155 and BRG1 show a near-perfect overlap of association on the ES cell genome (Ho et al PNAS 2009), suggesting BAF155 association indicates an intact complex comprising the ATPase. While it is known that BAF155 is important during early development (J. K. Kim et al., 2001; Sun et al., 2007) it has been difficult to study the function of the protein during this time due to the early lethality of Baf155 null mouse embryos. Twenty percent of BAF155 heterozygous embryos exhibit exencephaly with increased cranial proliferation observed 4 days after the time of neural tube closure (J. K. Kim et al., 2001). However, the role of BAF155 has not been determined during the time of neural tube closure. 141
154 Here, we characterize a missense mutation allele of the Smarcc1 gene (called Smarcc1 msp3 or Baf155 msp3 ). Mice homozygous for the Baf155 msp3 allele show 81% incidence of exencephaly. The BAF155 msp3 protein is expressed and the BAF core complex can assemble in vivo, providing a unique means to study the function of the BAF chromatin remodeling complex during embryogenesis. The molecular basis of the NTD is not due to aberrant BAF complex localization within the cell, disruption of neural tube patterning, or premature differentiation. However, proliferation and apoptosis are misregulated in the mutants, suggesting a potential mechanism underlying the NTD. Gene expression analysis of cranial tissue at the time of neural tube closure showed that genes involved in cell survival and neuronal development were misregulated and that mutant tissue showed significantly more variability in gene expression than wildtype. Overall, these studies lend new insight into the functions of the BAF chromatin remodeling complex during the critical stage of neural tube closure. MATERIALS AND METHODS Mouse Strains Baf155 msp3 (Smarcc1 msp3 ) was originally identified on a mixed genetic background (BALB/cJ; C57BL/6J) in an ENU screen (Buac et al., 2008).The mutation in Smarcc1 was identified as described in the results. Mice have been maintained on a mixed C57BL/6J:129S1/SvlmJ background as a 3 rd generation cross and heterozygous carriers were mated to produce the embryos and results presented here. Further crossing into C57BL/6J results in more penetrant developmental delay. For timed pregnancies, noon of the day of an observed vaginal plug was designated E0.5. At dissection, the embryonic phenotype was recorded and a portion of the yolk sac used for genotyping. Genotyping 142
155 DNA samples were genotyped using a custom TaqMan assay (Applied Biosystems) with Taqman probes designed across the site of the msp3 mutation specific for both the wildtype allele (Vic-CTC-CTG-TTG-TAA-CTG-C) and the ENU induced mutant allele (Fam-CTC-CTG-TTT-TAA-CTG-C). The following primers were used, forward primer: TTT-GCA-GAT-GAG-CAG-GAT-GAA-GAA and reverse primer: TCT-CAT-TTC-AGG- CCT-AAA-TAA-ACT-TTT-ACC-T. PCR reactions were carried out in 2x Taqman Universal Fast PCR Master Mix (Applied Biosystems, ), 10uM each genic primer, and 100 nm of each allele-specific probe. Cycling conditions were 95 C for 10 minutes, 40 cycles of 95 C for 3 seconds, and 60 C for 30 minutes. Relative quantitation of the two alleles was determined in an endpoint assay for genotyping. Analysis of mutant phenotype Embryos were fixed in 4% paraformaldehyde (PFA) in PBS and processed for whole mount or cryosection (8 um) in situ hybridization. Whole-mount and section RNA in situ hybridizations were performed as described(holmes and Niswander, 2001) with digoxygenin-labeled antisense riboprobes. Sections were imaged on a Nikon Eclipse 80i microscope. For immunohistochemistry, embryos were dissected and fixed in 4% PFA in PBS, washed 3 times with PBS, cryopreserved in 30% sucrose, embedded in OCT and sectioned (8uM). For cell cycle analysis, BrdU (50 mg/kg) in PBS was injected intraperitoneally into pregnant dams 30 minutes prior to euthanasia and embryo processing. For apoptosis, the Apop-tag In Situ Apoptosis Detection Kit (Milipore, S7160) was used following the protocol to indirectly detect apoptotic cells by the TUNEL method. For immunostaining, tissue sections were treated with PBS/3% goat serum for 1 hour at room temperature. Primary antibody was added and incubated at 4 C overnight. For detection, Alexa-conjugated secondary antibodies (Molecular Probes) were incubated for 1 hour at room temperature. Hoechst was added along with the secondary antibodies as a 143
156 nuclear stain. Stained sections were imaged with Zeiss LSM510 META confocal microscope. Antibodies used Antibodies against the following proteins were used: TUJ1 (Covance, MMS425P, 1:500), phospho-histone H3 (Cell Signaling, 9701, 1:200), BrdU (Novus Biologicals, NB , 1:200), p75 NTR (Promega, G3231, 1:100), BRG1 (Santa Cruz, SC :200), BAF155 (Santa Cruz, SC-10756, 1:200) and BAF170, (Santa Cruz, SC17838, 1:200). Antibodies from the Developmental Studies Hybridoma Bank were used at a 1:10 dilution including PAX3 and PAX6. Immunoprecipitation assay and Western Blots Embryonic day E11.5 embryos were placed in lysis buffer (50 mm Tris-HCl at ph7.4, 150 mm NaCl, 1 mm EDTA, 1% Triton X-100, including 1 tablet protease inhibitor (Roche Complete) per 10 mls lysate) and incubated on ice for 10 min. Tissue debris was pelleted at 12,500 rpm for 10 min at 4 C, and the supernatant was incubated with primary antibodies overnight at 4 C. The lysates were incubated with Protein A or G Sepharose beads for 2 h, followed by washing of the immunoprecipitates three times with lysis buffer and elution of bound proteins in SDS-PAGE sampling buffer for 10 min at 100 C. Western blots were performed as described previously (Chen et al., 2005) using primary antibodies as listed. Secondary antibodies used were goat anti-rabbit ( , Bio-Rad), and goat anti-mouse ( , Bio-Rad). RNA-Seq library preparation and analysis Three somite matched E9.5 embryos were collected of both Baf155 +/+ and Baf155 Msp3/Msp3 alleles. Cranial tissue above the pharyngeal arches and otic vesicles was harvested and 144
157 stored at -80 C until all samples were collected. RNA was isolated using the RNEasy mini kit from Qiagen (74106). On average, 1400 ng of total RNA was obtained per sample. RNA concentration and A260/A280 ratio was analyzed using the Nanodrop ND-1000 spectrophotometer (Thermo Fisher Scientific). Libraries were made using the Illumina TruSeq RNA Sample Preparation Kit V2 (RS ). Libraries were analyzed on the 2100 Bioanalyzer (Agilent Technologies, Inc) for proper size and integrity. For highthroughput RNA-Sequencing analysis, the UC Denver Genomics Core used an Illumina High Seq Mapping and bioinformatics analysis was done in collaboration with the Bioinformatics Shared Resources. The sequenced libraries generated 804,339,023 unfiltered reads of ~100 bps in length. Each replicate represents at least 96.9 million reads, a density sufficient for qualitative analysis of gene expression (Phelan et al., 1999). The ~100 bp reads were aligned to the MM9 mouse genome using Gsnap,(Wu and Nacu, 2010) allowing a mismatch of 4% and an indel cost of 2.0. The mapped transcripts were visualized in the UCSC genome browser to bioinformatically verify differential expression in transcript counts and/or transcript length, as well as splicing of introns. Data Analysis and interpretation For measurements of differential gene expression edger software (Robinson et al., 2010) was used. Raw read counts were established for each RefSeq gene. In cases where more than one transcript was annotated, the transcript with the highest overall counts was used as proxy for the gene in question, yielding a data table with a single count entry for each gene per sample. The data table was trimmed to remove all genes where zero counts were reported in one or more of the 6 samples, with the exception of genes where either all the mutant samples, or all the wildtype samples showed zero counts, and the opposing side of the paradigm showed counts in each sample. This resulted in a gene data table 145
158 with entries. The data set was used for differential expression analyses using edger. Comparisons of all three mutant samples versus all three wildtype samples were performed, as well as separate comparisons of each individual mutant sample against all three wildtype samples; resulting gene lists were used for Ingenuity Pathway Analysis to reveal biological features. Overall sample relationship was visualized using the multidimensional scaling feature of edger. Raw gene count numbers were normalized using DESeq (Anders and Huber, 2010), and normalized counts were used to calculate the coefficient of variation for each gene in the normal samples, as well as determine for each gene the relationship of the counts in each sample relative to the mean of the normalized counts derived from the three wildtype samples for the pertinent gene. To obtain distance from mean, the ratio of normalized counts divided by mean of the normal underwent logarithmic transformation. All genes were ranked according to their coefficient of variation, and distance from mean was plotted for each gene for normal as well as for mutant samples. RESULTS Baf155 msp3 mutants exhibit cranial neural tube defects The Msp3 phenotype was originally discovered in a mouse ENU mutagenesis screen (Buac et al., 2008), however the specific gene mutation was not identified. As outlined below, we identified the ENU-induced mutation in the Smarcc1 gene, encoding BAF155, a component of the Brg1/Brama-associated factor (BAF) ATP-dependent chromatin remodeling complex (BAF complex). For the remainder of the paper we refer to the mutant gene as Baf155 msp3 and the protein as BAF155 msp3. Baf155 msp3/+ heterozygotes are indistinguishable from wild-type littermates and they are viable and fertile. Homozygous Baf155 msp3/msp3 mutant embryos display cranial neural tube defects and developmental 146
159 delay (combined phenotypes at 92% penetrance, exencephaly at 81% frequency, and developmental delay with and without exencephaly at 30% frequency) on a mixed C57BL/6J; 129SvlmJ background (Figure 1A-C). The embryonic phenotypes range in severity and include exencephaly and developmental delay, defined as being at least 2 somites less than the least somite number of the non-mutant littermates (Figure 1 and Table 1). For example, Figure 1A shows embryos from a single litter at embryonic day 9.5 (E9.5), just after the time of neural tube closure. Mutant embryo 1 (MUT 1) is exencephalic from the forebrain through the hindbrain but not delayed (25 somites), MUT 2 shows midbrain and hindbrain exencephaly and developmental delay (21 somites), MUT 3 is severely delayed (14-15 somites) and hence cranial neural closure cannot be scored. Other observed defects are an enlarged pericardium (~12% of mutants) and, at later stages, variable epidermal edema. Most homozygous mutants can survive until E15.5 but no homozygotes are found at the time of weaning (Table 1). The cause of death in the Baf155 msp3/msp3 mutants is currently unclear and was not explored here. Mapping the ENU-induced mutation to Baf155 Baf155 msp3 was originally identified on a mixed genetic background (BALB/cJ; C57BL/6J) in an ENU mutagenesis screen and localized to mouse chromosome 9 using traditional linkage mapping (Buac et al., 2008). During subsequent outcrossing to C57BL/6J, the critical interval was narrowed to a 2Mb region (rs D9Mit37; NCBI Build36 chr9: 108,655, ,607,406). Resequencing of exons in the interval was completed using an exon-based hybrid selection strategy (Illumina Genome Analyzer, Broad Institute, Mouse Mutant Resequencing Initiative). Filtering of coding or splicing homozygous variants not present in dbsnp revealed a single variant in Smarcc1 that was not present in the parental BALB/cJ strain. This C to A variant was confirmed with Sanger sequencing and a TaqMan SNP genotyping assay. Subsequently, mice were crossed with 129SvlmJ 147
160 to generate a mixed C57BL/6J;129SvlmJ background and RNA-Seq (Illumina Hi Seq 2000) was performed on wildtype (WT) and mutant cranial tissue (results discussed below), and the resulting data was analyzed for variants. Variants were filtered similar to the first analysis overlapping the linkage disequilibrium region between Mb on chromosome 9 (defined by the variant calling program). Nineteen nonsynonymous SNPs were identified, but only the Smarcc1 C to A variant was novel and not present in dbsnp. These two independent genomic analyses identify the causative ENU-induced mutation in Smarcc1, encoding BAF155. This C to A substitution is predicted to cause a threonine to lysine substitution at amino acid 416 of BAF155 (NP_033237; Figure 2A). The BAF155 protein is comprised of one Chromo domain, one SWIRM domain, and one SANT domain. The affected amino acid is not within a conserved domain but is located N-terminal to the SWIRM domain in the BAF155 protein. The missense mutation site and surrounding sequences are conserved between H. sapien, M. musculus, R. norvegicus and D. rerio, but not in S. cerevisiae (Figure 2A), suggesting this residue may be functionally important. Mutant Baf155 RNA and protein are present and the protein is localized to the nucleus Missense mutations within protein-coding genes may affect mrna stability, as well as protein stability or function. To understand the molecular consequences of the C to A missense mutation in the Baf155 gene, we analyzed mrna and protein expression from E11.5 mutant and WT embryos. RT-PCR analysis showed the presence of Baf155 mrna in Baf155 msp/3msp3 embryos indicating that the transcript is not subject to nonsensemediated decay (Figure 2B). Furthermore, Western blot analysis confirmed the presence of full length BAF155 in homozygous mutant embryos at similar levels to wild-type (Figure 2B). Together, these data suggest that BAF155 mrna and protein stability are not compromised in the Baf155 mutant. 148
161 The BAF complex is normally localized to the nucleus. Mutations in the BAF155 nuclear localization signal (NLS) result in incorrect cytoplasmic localization of BAF155, BAF47 and the ATPase BRG1, indicating that BAF155 maintains the subcellular localization of the BAF complex in the nucleus (Sohn et al., 2007). To test the possibility that subcellular localization of BAF155 and BRG1 may be compromised in the Baf155 msp3 mutants, we examined their localization in cryosections from wildtype and mutant embryos. In mutants, BRG1 and BAF155 are present in the nucleus at high levels while they are not detected in the cytoplasm, comparable to wildtype (Figure 2C). This suggests that subcellular mislocalization of BAF155 or other BAF proteins does not account for the functional deficit seen in Baf155 msp3/msp3 mutants. BAF155 msp3 mutant protein can still associate with the core remodeling complex BAF155, along with ATPase BRG1, BAF170, and BAF47, are considered core components in the BAF remodeling complex because they are sufficient to remodel nucleosomes in vitro at the same efficiency as the fully intact BAF remodeling complex (Phelan et al., 1999). BAF155 interacts directly with two core complex proteins (BRG1 and BAF47) as well as with BAF60a (Sohn et al., 2007) in the region that encompasses the BAF155 msp3 missense mutation, T416K. To test the possibility that the interaction between BAF155 and the BAF core complex is disrupted by the T416K mutation we performed coimmunoprecipitation assays using E11.5 mutant and WT embryos. These studies revealed that BRG1 and BAF170 still associate with BAF155 msp3 in vivo and in a similar ratio as compared to the WT, despite the amino acid change (Figure 2D). Additionally, yeast two-hybrid analysis indicated that BAF155 msp3 can still interact directly with BAF47 and BAF60a in vitro (Supplemental Figure 1). These experiments indicate that the mutant BAF155 msp3 protein still interacts with the core components of the BAF remodeling complex and suggest the mutant protein could interact with other BAF complex proteins in 149
162 vivo as well (summarized in Figure 2E). These protein interaction data, as well as the longer survival of Baf155 msp3/msp3 embryos relative to Baf155 null embryos (J. K. Kim et al., 2001; Sun et al., 2007) suggest that the BAF155 msp3 mutant protein is present and can interact molecularly in the remodeling complex, but its function may be partially compromised. Thus, it appears that Baf155 msp3 is a missense allele and the mouse may exhibit a hypomorphic phenotype. Cell proliferation is decreased and cell death is increased in Baf155 msp3/msp3 neural progenitor cells NTDs can result when the spatiotemporal control of neural tube closure is disrupted (A. J. Copp and Greene, 2010) due to any of several different mechanisms including: reduced neural progenitor cell proliferation, changes in cell fate, decreased cell survival, or defects in patterning. To investigate if the NTD might arise because of improper patterning, we examined a battery of molecular markers that reflect the dorsal-ventral or anteriorposterior axis patterning of E9.5 embryos, just after closure of the neural tube. Antibody staining and RNA in situ hybridization showed no change in the expression pattern of Shh, PAX6, PAX3, Fgf8 and FoxG1 in mutant neural tubes when compared to wildtype somite matched embryos (Figure 3A-F). Neural crest cells are specified during neural tube closure and abnormal expression of neural crest genes could also reflect altered patterning (Trainor, 2005). Therefore, we examined markers characteristic of early neural crest specification including p75 NTR, Tfap2a, and Sox10 (Figure 3G-I) and again found no change in the expression pattern compared to WT. These results suggest that neural patterning is not disrupted in the Baf155 msp3 mutant. The final step in neural tube closure is the apposition of the two neural folds, which grow and bend closer together partially by increasing the number of neural progenitor cells, and finally, when the two folds meet, tissue fusion. The tight control of cell proliferation is 150
163 necessary for the neural folds to meet properly at the fusion point. At E9.5, the vast majority of the neural ectoderm is comprised of proliferating neural progenitor cells. Too much or too little cell proliferation can abrogate neural tube closure. Previous studies noted an increase in cell proliferation in the telencephalic striatum region of exencephalic heterozygous Baf155 null embryos at E13.5 (J. K. Kim et al., 2001). Thus, we asked if the proliferation rate might be altered in Baf155 msp3/msp3 mutant embryos at the time of neural tube closure. The mitotic rate of neuroepithelial cells in somite-matched E9.5 Baf155 +/+ and mutant Baf155 msp3/msp3 embryos was compared using phospho-histone 3 (ph3). In the hindbrain, there was a statistically significant decrease in the mitotic index (ph3+ cells/1000 neuroepithelial cells) of Baf155 msp3/msp3 mutants compared to WT (20±9 vs 71±10) (Figure 4A,D). We also quantified the number of cells in S phase by performing bromodeoxyuridine (BrdU) incorporation experiments (Figure 4A,E). There was no apparent difference in number of cells in S phase in mutant compared to WT neural tubes at E9.5. These studies suggest that cells are cycling in the Baf155 msp3 mutant, but there may be a mitosis defect in the neuroepithelium. A decrease in cell survival within the neural tissue may result in too few cells to allow proper apposition and fusion of the neural folds, resulting in a NTD. To determine if neural progenitor cell survival was affected in mutant embryos we used a TUNEL assay to visualize apoptotic cells in somite-matched E9.5 WT and mutant neural epithelium. This showed a statistically significant increase in TUNEL positive cells per 1000 nuclei in the hindbrain at E9.5 in the Baf155 msp3/msp3 mutant compared to WT (7.2±1.0 vs 3.7±0.5) (Figure 4B,F). Together, these studies suggest that BAF155 function is necessary to maintain the normal rates of proliferation and cell survival in the neuroepithelium. 151
164 As neural precursors begin to differentiate starting around E10.5, they exit the cell cycle and leave the ventricular zone to reside at the periphery of the neural tube. If differentiation does not occur at the correct time, this can contribute to an excess or a deficiency in cell number for effective neural fold juxtaposition and ultimate fusion to form the neural tube. To determine whether or not neural differentiation was compromised in the homozygous mutants we evaluated Tuj1 expression, a known neural differentiation marker, in the spinal cord and hindbrain of E10.5 somite matched Baf155 +/+ and Baf155 msp3/msp3 embryos. Mutant and WT embryos exhibit the same characteristic expression pattern of Tuj1 (Figure 4C) and therefore neural differentiation does not appear to be affected in the homozygous mutant. Gene expression levels are more variable in Baf155 msp3 mutant cranial tissue Averaged sample analysis BAF155, as a component of a chromatin remodeling complex, is thought to be involved in the regulation of gene expression. Baf155 knockdown in ES cells results in misregulation of hundreds of genes (Ho et al., 2009). The Baf155 msp3 allele presents a unique case for gene expression analysis wherein the BAF complex appears to be intact, in contrast to disruption of the BAF complex when BAF155 function is lost (Chen and Archer, 2005; Sohn et al., 2007). To examine gene expression in the Baf155 msp3 mutant at the time of neural tube closure, we performed RNA-Seq and compared gene expression levels between homozygous wildtype and homozygous mutant embryos. RNA was isolated from somite matched (21-23 somites) and phenotype matched cranial tissue of homozygous wildtype and homozygous mutant embryos (3 of each genotype) at E9.5 just after the time of neural tube closure. At E9.5, the vast majority of the embryonic neural tube is comprised of proliferating neural progenitor cells, constituting a relatively homogeneous cell population. After extensive analysis of the RNA-Seq data comparing all 3 mutants to 152
165 all 3 WT replicates, 78 genes were identified as being upregulated on average in the mutant samples while 22 genes were downregulated. The full list of differentially expressed genes in all 3 mutant replicates collectively, as well as p-values, FDR and fold change, is shown in Supplemental Table 1. To facilitate an understanding of global gene expression differences, Ingenuity Pathway Analysis (IPA) was used to assign functional categories to the up- and down-regulated genes in the mutants. The most significant molecular and cellular functions of these genes were: 1. cellular movement (1.46E- 04>p>4.12E-02), 2. Cell death and survival (2.69E-04>p>4.12E-02), and 3. Cell growth and proliferation (2.69E-04>p>3.41E-02). Many neuronal genes were significantly misregulated as well. Individual sample analysis We used multidimensional scaling (MDS) analysis to conduct an unsupervised examination of the relationship between the global gene expression in each individual mutant and the wildtype cranial tissue samples. As seen in Figure 5A, a relationship between the genotypic subtype and gene expression was clearly observed. The wildtype replicates were tightly grouped in the MDS plot and showed little dimensional variance. In contrast, the mutant replicates did not group together and showed increased variance in both dimensions 1 and 2. Overall, there was a trend towards higher variance of gene expression in the mutant samples compared to the wildtype samples, leading to a more dispersed MDS plot where the mutants do not cluster with the wildtype controls and each mutant does not cluster with the other mutants. Based on the lack of clustering between the individual mutants when global gene expression was compared, the averaged gene expression analysis did not illustrate the nuances of gene expression in the Baf155 msp3 mutant samples. Therefore, in a more exploratory analysis, we asked if the observed transcriptional changes were uniformly 153
166 distributed in the 3 biological replicates between the homozygous mutant group and compared to the homozygous wildtype groups. For each sample, we established for each gene a distance from mean parameter by normalizing the expression level of any given gene to the arithmetic mean of the expression of the respective gene in the wild type samples; this ratio underwent logarithmic transformation to obtain the distance value. All genes were ranked by their coefficient of variation in the wildtype samples, and the distance value was plotted as shown in Figure 5B (wildtype in top panel, mutants in bottom panel). The average wildtype plot showed a relatively tight spacing of all data points in a 'trumpet' shape. This coincides well with the low degree of variation we identified in the differential expression test. The same plot for the mutant samples showed a larger spread of gene expression ratios, and mutant replicate 2 (Mut2 in turquoise) seems to have the greatest difference in the ratio of gene expression compared to both remaining mutants and wild type controls. Again, this correlates well with the differential expression tests. To identify whether there are groups of genes that may contribute to this gene expression variability more than others, we generated three separate mutant differential gene expression lists by directly comparing each mutant gene count to the average of the wildtype gene counts. This analysis shows collections of genes from each individual mutant as opposed to the genes misregulated in all of the mutants. Over 2149 genes were differentially expressed in Mutant replicate 2 (Mut2) as opposed to 188 genes in Mutant replicate 1 (Mut1) or 215 genes in Mutant replicate 3 (Mut3) (based on DESeq testing, and counting genes with padj<0.05). When we directly compared these three differentially expressed gene lists, we identified a subset of 27 genes that was significantly misregulated in all of the mutants (Supplemental Table 2). Interestingly, many of the genes on this list do not overlap with Supplemental Table 1, comparing all mutant 154
167 samples to all WT samples, because there is often one gene outlier that is not significantly misregulated in each individual. Pathway analysis using IPA suggests that no canonical pathways are shared between these 27 genes, but 6 of these genes are predicted to be within a nucleic acid metabolism network. Overall, these results suggest that gene expression in the mutants is more varied than in the wildtype samples, and there is little overlap in the functionality of the most variable genes between individuals in the Baf155 msp3 mutant group. DISCUSSION Neural Tube Defects in humans have a complex etiology and it is likely that multiple genes contribute to the incidence of this common birth defect, but the specific genetic causes of NTDs are still largely unknown. Moreover, it is unknown whether polymorphisms in epigenetic regulators may be risk factors for NTDs in humans. Studies in model organisms like the mouse can provide insight into the genetics underlying NTDs. Indeed, mutations and deletions in a number of epigenetic regulators can cause NTDs in mice. In this study we focused on the BAF chromatin remodeling complex, as mutations in two members of the BAF complex cause NTDs in mice, yet the functions of the BAF complex at the time of neural tube closure have not been studied. Here we characterized the first missense allele of the Baf155 gene, Baf155 msp3, which exhibits a cranial NTD. We found that the mutant protein can still interact with the core BAF complex proteins BRG1, BAF170 and BAF47 and properly localize to the nucleus. This unique allele allowed an investigation of the cause of the NTD in the presence of an assembled core BAF complex. In Baf155 msp3 mutant embryos, we discovered a reduction in neural progenitor cell proliferation and an increase in cell death, revealing a possible mechanism underlying the NTD. We analyzed transcriptional control in the Baf155 mutant using RNA-Seq and discovered misregulation of genes involved in apoptosis and proliferation. We also found 155
168 increased variability in gene expression in the mutant compared to wild type, the meaning of which will be speculated on below. The BAF complex can act as both a repressor and activator of genes, and has been found to refine the gene transcriptional network in ES cells to maintain pluripotency (Ho et al., 2009; Schaniel et al., 2009; Zhang et al., 2007). Using RNA-Seq, we analyzed transcriptional control in the cranial tissue of the mutant embryo compared to wild type. Most of the differentially expressed genes in the Baf155 msp3 mutant embryo are upregulated, including neural development genes discussed below, suggesting that BAF155 and the BAF complex largely play a repressive role during neurulation. Consistent with the expectation that the BAF complex provides more global, rather than gene-specific, transcriptional regulation, we did not find strong single gene candidates to explain the underlying mechanism of the NTD. Instead, we observed a broader dysregulation in expression of genes and gene families that are associated with NTDs when disrupted. While the fold change in gene expression was not particularly striking across all the mutants, it is not unreasonable to suggest that small gene expression changes in combination may result in an NTD. As previously noted, neural tube closure is a highly dynamic process that requires precise coordination of complex cellular functions to proceed to completion. Moreover, many of the misregulated genes in Supplemental Table 1 are associated with NTDs: mutations in Abl ( Koleske et al., 1998), Frem2 (Jadeja et al., 2005; Timmer, 2005), as well as other genes discussed below result in NTDs. Baf155 regulation of cell survival during neural development Proper control of proliferation and cell survival are critical for neural tube closure. In the Baf155 msp3 mouse mutants we identified both a decrease in cell proliferation and an increase in apoptosis. RNA-Seq revealed several misregulated genes that contribute to cell survival, including the downregulated genes Abl1and ApoB, and the upregulated 156
169 genes Adora3, Fam72a, SST, and Map3k12. Mouse embryos mutant for Apolipoprotein B (ApoB) exhibit exencephaly (30% incidence) and excessive cell death in the hindbrain (Huang et al., 1995). Knock-out of Abl1 kinase in combination with Abl related gene (ARG) results in NTDs and disruption of the actin cytoskeleton of the neuroepithelium (Koleske et al., 1998). Although the mechanism underlying the increased apoptosis in Baf155 msp3 mutants is unclear, altered expression of these genes may contribute to the increase of cell death in the neural epithelium of the Baf155 mutant. BAF155 can be phosphorylated by and interact with CyclinE (Shanahan et al., 1999) in the Rb pathway as well as Akt (Foster et al., 2006) in the PI3K/AKT pathway. However, we did not detect misregulation of Rb or PI3K pathway genes in the mutant, suggesting these signaling pathways are not disrupted and BAF155 may play other transcriptional roles in controlling proliferation. While many genes known to control proliferation were misregulated in the mutant cranial tissue, no specific cyclins or cell cycle genes were differentially expressed that could account for the observed reduction in proliferation. However, there is decreased expression of the centrosomal gene Poc1a. Inhibition of centrosomal genes can lead to cell cycle defects and cell death (Doxsey et al., 2005). For example, inhibition of the centrosome protein Nde1 was shown to delay cell cycle reentry in cell lines, and to reduce mitosis in zebrafish (S. Kim et al., 2011). Together these data suggest that Baf155 may directly or indirectly regulate cell proliferation and cell death via transcriptional control. Regulation of neural development genes by Baf155 BAF subunits are heterogeneously expressed and assembled in various cell types and these different complexes can have distinct functions. Different protein assemblies of the BAF complex exist in the developing central nervous system, including the neural progenitor BAF complex (npbaf) and neuronal BAF complex (nbaf) (Lessard et al., 157
170 2007). These have specific functions in the regulation of proliferation and differentiation of mammalian neural stem cells. Later loss of BRG1 function in neurons (Brg1 fl/fl ;Nestin-Cre) and transcriptional analysis of E12.5 mutant brains revealed an upregulation of genes within the Notch and Shh pathways. While it is believed that BAF155 is present in BAF complexes in all cell types, we did not see gene expression changes in the Baf155 msp3 mutant at E9.5 that might reflect alterations in Notch or Shh signaling. This difference in gene control between the Brg1 fl/fl ;Nestin-Cre and the Baf155 msp3 mice could reflect temporal differences in BAF complex function during brain development. Many of the upregulated genes in Baf155 msp3 mutants at the time of neural tube closure contribute to neural development, including Spon1, Doc3, Topb2, Map3k12, Cdk18, Sema3c, Sst, Igta4, Epha3, Fzd2, Fzd3, Ablim3, and Plcg2. BAF subunit mutations have been reported in many human neurologic diseases including Autism, Schizophrenia, Coffin-Siris syndrome, sporadic mental retardation (Ronan et al., 2013), however these neurological diseases are not accompanied by NTDs according to the Online Mendelian inheritance of Man database (OMIM). While no differentiation defect was observed in the Baf155 msp3 mutants at and shortly after the time of neural tube closure, the modest misregulation of many neural development genes may contribute to the NTD phenotype. Particularly interesting are the genes associated with NTDs, namely the Wnt receptors Fzd2 and Fzd3. The Wnt/Fzd (PCP) pathway controls aspects of neural development, including neural tube closure, in mice and humans (D. M. Juriloff and M. J. Harris, 2012). Although the Fzd2 -/- mouse does not show NTD, in a double heterozygous cross with another PCP component, Fzd +/- :Vangl Lp/+, fully penetrant cranial NTDs are observed (Yu et al., 2010). Also, Fzd3 -/- ; Fzd6 -/- mutants present with the most severe NTD, craniorachischisis(y. Wang, 2006). This supports the idea that NTDs can arise due to aberrant expression of multiple risk alleles. 158
171 Potential roles of Baf155 in the Baf155 msp3 mutant Hemizygous Brg1 or Baf155 mice show defects in neural tube closure, suggesting a dosage-sensitive role for BAF complexes in neural development (Bultman et al., 2000; J. K. Kim et al., 2001). In the Baf155 msp3/msp3 mutant, the core components of the complex are intact and the RNA and protein levels are not significantly changed, yet the neural tube defect is highly penetrant. Our studies suggest an alternative mechanism for epigenetic control of neural tube closure, in addition to dosage of the BAF proteins. One possibility is that the BAF155 msp3 mutation disrupts the interaction of BAF155 with transcription factors or chromatin maintenance proteins. For example, it has been suggested that intrinsic disorder within the BAF protein structures may explain diverse roles of chromatin remodeling proteins (Sandhu, 2009), and the point mutation within our allele occurs in an intrinsically disordered region based on Phyre analysis (data not shown). However, we observed no difference between the mutant and WT protein for association to BAF60, which directly interacts within the region of the Baf155 T416K mutation. It remains possible that another protein may interact within this disordered region and disruption of this interaction may lead to NTDs. Another possibility is that interaction with specific genomic targets that regulate the developmental program of neural tube closure may be disrupted. In ES cells, BAF155 is bound to promoter or genic regions of ~5,630 distinct genes (Ho et al., 2009). Moreover, esbaf facilitates Stat3 genomic binding, and can act either synergistically or antagonistically with the Polycomb Repressive Complex Group (PcG) in ES cells (Ho et al., 2011). In HeLa cells, BAF proteins associate with up to 77% of genic DNA (Euskirchen et al., 2011). Given the potentially large set of genomic targets, there are surprisingly few genes misregulated overall in Baf155 msp3 mutant cranial tissue at E9.5, especially considering that at least some of these misexpressed genes are likely indirect targets. 159
172 There is also very little overlap in the datasets of differentially expressed genes between our point mutant and knock-out/knock-down of Brg1 or other BAF members in ES cells (Ho et al., 2009), neural progenitors or neurons (Lessard et al., 2007). This lack of overlap likely reflects the differences between mutant alleles: knock-down of one component presumably results in incorrect ratios of complex proteins, whereas our point mutant does not apparently change the levels of complex proteins and the complex is largely intact. Thus, the point mutation may be expected to reveal more specific targets relative to neural tube closure. Gene expression is more variable in Baf155 msp3 cranial tissue The genes highlighted above show strong statistical significance in terms of changes in gene expression in the averaged mutant samples versus averaged wildtype samples. However, the RNA-Seq analysis also raised another interesting finding in that there is an increase in the degree of variability of gene expression between individual Baf155 msp3/msp3 embryos. This appears to be unique to the Baf155 msp3 embryos, and perhaps other epigenetic regulators, because in gene expression studies of other NTD mutants, high gene expression variability was not observed (Pyrgaki et al., 2011). Based on EdgeR and IPA analysis, there is little overlap in the most variable genes between individual mutants, and there does not appear to be a group of genes that mediates this variability. However, Topoisomerase 2β(Top2b) and Sirtuin7(Sirt7) (a histone deacetylase) are differently expressed in all the mutant embryos and stand out as globally acting proteins that may contribute to further differences in gene expression between the individual mutants. Top2b plays a critical role in axonal development in mice (Yang, 2000) and it has been suggested to be involved in DNA repair (Yamane et al., 1997). Sirt7 can preferentially deacetylate H3K18, interact with BRG1/BRM, and is suggested to regulate RNA Polymerase I transcription in many cell types (Barber et al., 2012; Tsai et al., 2012). It is 160
173 possible that global alterations in histone acetylation could contribute to a greater variability in gene expression. Variable gene expression might also be due to differential occupancy of the BAF complex on the genome, or disruption of other proteins that interact with the BAF complex to modulate its function. Despite the lack of overlap of gene expression changes between Baf155 msp3 and other BAF complex gene knockouts and the varying degree of gene expression changes between Baf155 msp3 mutants, our mouse model shows a consistent NTD phenotype. Neural tube closure is a complex, multigenic process and our data indicate that inconsistent regulation of genes by a chromatin remodeling complex with reduced function can lead to developmental defects during this highly dynamic phase of embryonic development. REFERENCES Anders, S., Huber, W., Differential expression analysis for sequence count data. Genome Biol. 11:R106. Bamforth, S.D., Bragança, J., Eloranta, J.J., Murdoch, J.N., Marques, F.I., Kranc, K.R., Farza, H., Henderson, D.J., Hurst, H.C., Bhattacharya, S., Cardiac malformations, adrenal agenesis, neural crest defects and exencephaly in mice lacking Cited2, a new Tfap2 co-activator. Nat. Genet. 29: Banting, G.S., CECR2, a protein involved in neurulation, forms a novel chromatin remodeling complex with SNF2L. Hum Mol Genet 14: Barber, M.F., Michishita-Kioi, E., Xi, Y., Tasselli, L., Kioi, M., Moqtaderi, Z., Tennen, R.I., Paredes, S., Young, N.L., Chen, K., Struhl, K., Garcia, B.A., Gozani, O., Li, W., Chua, K.F., SIRT7 links H3K18 deacetylation to maintenance of oncogenic transformation. Nature 487: Bu, P., Evrard, Y.A., Lozano, G., Dent, S.Y.R., Loss of Gcn5 acetyltransferase 161
174 activity leads to neural tube closure defects and exencephaly in mouse embryos. Mol Cell Biol 27: Buac, K., Watkins-Chow, D.E., Loftus, S.K., Larson, D.M., Incao, A., Gibney, G., Pavan, W.J., A Sox10 Expression Screen Identifies an Amino Acid Essential for Erbb3 Function. PLoS Genet. 4:e Bultman, S., Gebuhr, T., Yee, D., La Mantia, C., Nicholson, J., Gilliam, A., Randazzo, F., Metzger, D., Chambon, P., Crabtree, G., A Brg1 Null Mutation in the Mouse Reveals Functional Differences among Mammalian SWI/SNF Complexes. Molecular Cell 6: Chen, J., Archer, T.K., Regulating SWI/SNF Subunit Levels via Protein-Protein Interactions and Proteasomal Degradation: BAF155 and BAF170 Limit Expression of BAF57. Mol Cell Biol 25: Chen, J.-F., Mandel, E.M., Thomson, J.M., Wu, Q., Callis, T.E., Hammond, S.M., Conlon, F.L., Wang, D.-Z., The role of microrna-1 and microrna-133 in skeletal muscle proliferation and differentiation. Nat. Genet. 38: Cheng, H.-L., Mostoslavsky, R., Saito, S., Manis, J.P., Gu, Y., Patel, P., Bronson, R., Appella, E., Alt, F.W., Chua, K.F., Developmental defects and p53 hyperacetylation in Sir2 homolog (SIRT1)-deficient mice. 100: Clapier, C.R., Cairns, B.R., The Biology of Chromatin Remodeling Complexes. Annu. Rev. Biochem. 78: Copp, A., Neurulation in the cranial region normal and abnormal 207, Copp, A.J., Greene, N.D.E., Genetics and development of neural tube defects. J. Pathol. 220: Detrait, E.R., George, T.M., Etchevers, H.C., Gilbert, J.R., Vekemans, M., Speer, M.C., Human neural tube defects: developmental biology, epidemiology, and 162
175 genetics. Neurotoxicol Teratol 27: Doxsey, S., Zimmerman, W., Mikule, K., Centrosome control of the cell cycle. Trends in Cell Biology 15: Dunwoodie, S.L., Rodriguez, T.A., Beddington, R.S., Msg1 and Mrg1, founding members of a gene family, show distinct patterns of gene expression during mouse embryogenesis. 72: Elwood, J., Little, J., Elwood, J., Epidemiology and control of neural tube defects. Euskirchen, G.M., Auerbach, R.K., Davidov, E., Gianoulis, T.A., Zhong, G., Rozowsky, J., Bhardwaj, N., Gerstein, M.B., Snyder, M., Diverse Roles and Interactions of the SWI/SNF Chromatin Remodeling Complex Revealed Using Global Approaches. PLoS Genet. 7:e Foster, K.S.J., McCrary, W.J., Ross, J.S., Wright, C.F., Members of the hswi/snf chromatin remodeling complex associate with and are phosphorylated by protein kinase B/Akt. Oncogene 25: Harris, M., Juriloff, D., Mini-review: toward understanding mechanisms of genetic neural tube defects in mice 60: Harris, M.J., Juriloff, D.M., An update to the list of mouse mutants with neural tube closure defects and advances toward a complete genetic perspective of neural tube closure. Birth Defects Res. Part A Clin. Mol. Teratol. 88: Hata, K., Okano, M., Lei, H., Li, E., Dnmt3L cooperates with the Dnmt3 family of de novo DNA methyltransferases to establish maternal imprints in mice. Development 129: Ho, L., Crabtree, G., Chromatin remodelling during development 463: Ho, L., Jothi, R., Ronan, J., Cui, K., Zhao, K., Crabtree, G., An embryonic stem cell chromatin remodeling complex, esbaf, is an essential component of the core 163
176 pluripotency transcriptional network 106:5187. Ho, L., Miller, E.L., Ronan, J.L., Ho, W.Q., Jothi, R., Crabtree, G.R., esbaf facilitates pluripotency by conditioning the genome for LIF/STAT3 signalling and by regulating polycomb function. Nat. Cell Biol. 13: Holmes, G., Niswander, L., Expression ofslit-2 andslit-3 during chick development. Dev. Dyn. 222: Huang, L.S., Voyiaziakis, E., Markenson, D.F., Sokol, K.A., Hayek, T., Breslow, J.L., apo B gene knockout in mice results in embryonic lethality in homozygotes and neural tube defects, male infertility, and reduced HDL cholesterol ester and apo A-I transport rates in heterozygotes. J. Clin. Invest. 96: Ito, T., Bulger, M., Pazin, M.J., Kobayashi, R., Kadonaga, J.T., ACF, an ISWIcontaining and ATP-utilizing chromatin assembly and remodeling factor 90: Jadeja, S., Smyth, I., Pitera, J.E., Taylor, M.S., van Haelst, M., Bentley, E., McGregor, L., Hopkins, J., Chalepakis, G., Philip, N., Perez Aytes, A., Watt, F.M., Darling, S.M., Jackson, I., Woolf, A.S., Scambler, P.J., Identification of a new gene mutated in Fraser syndrome and mouse myelencephalic blebs. Nat. Genet. 37: Juriloff, D.M., Harris, M.J., A consideration of the evidence that genetic defects in planar cell polarity contribute to the etiology of human neural tube defects. Birth Defects Res. Part A Clin. Mol. Teratol. 94: Kim, J.K., Huh, S.O., Choi, H., Lee, K.S., Shin, D., Lee, C., Nam, J.S., Kim, H., Chung, H., Lee, H.W., Park, S.D., Seong, R.H., Srg3, a Mouse Homolog of Yeast SWI3, Is Essential for Early Embryogenesis and Involved in Brain Development. Mol Cell Biol 21: Kim, S., Zaghloul, N.A., Bubenshchikova, E., Oh, E.C., Rankin, S., Katsanis, N., Obara, T., Tsiokas, L., Nature cell biology 2011 KimNde1-mediated inhi-2. Nat. Cell 164
177 Biol. 13: Koleske, A.J., Gifford, A.M., Scott, M.L., Nee, M., Bronson, R.T., Miczek, K.A., Baltimore, D., Essential roles for the Abl and Arg tyrosine kinases in neurulation. 21: Leck, I., Causation of neural tube defects: clues from epidemiology. Br. Med. Bull. 30, Lessard, J., Wu, J.I., Ranish, J.A., Wan, M., Winslow, M.M., Staahl, B.T., Wu, H., Aebersold, R., Graef, I.A., Crabtree, G.R., An Essential Switch in Subunit Composition of a Chromatin Remodeling Complex during Neural Development. Neuron 55: Luger, K., Mäder, A., Richmond, R., Sargent, D., Richmond, T., Crystal structure of the nucleosome core particle at 2.8 A resolution 389:251. Mandel, S., Gozes, I., Activity-dependent neuroprotective protein constitutes a novel element in the SWI/SNF chromatin remodeling complex 282: Nasmyth, K., The determination of mother cell-specific mating type switching in yeast by a specific regulator of HO transcription. 6: Okano, M., Bell, D.W., Haber, D.A., Li, E., DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99: Phelan, M.L., Sif, S., Narlikar, G.J., Kingston, R.E., Reconstitution of a core chromatin remodeling complex from SWI/SNF subunits. 3: Pyrgaki, C., Liu, A., Niswander, L., Grainyhead-like 2 regulates neural tube closure and adhesion molecule expression during neural fold fusion. Dev. Biol. 353: Robinson, M.D., McCarthy, D.J., Smyth, G.K., edger: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 165
178 26: Ronan, J.L., Wu, W., Crabtree, G.R., From neural development to cognition: unexpected roles for chromatin. Nat Rev Genet. 14: Sandhu, K.S., Intrinsic disorder explains diverse nuclear roles of chromatin remodeling proteins. J. Mol. Recognit. 22:1 8. Schaniel, C., Ang, Y.-S., Ratnakumar, K., Cormier, C., James, T., Bernstein, E., Lemischka, I.R., Paddison, P.J., Smarcc1/Baf155 Couples Self-Renewal Gene Repression with Changes in Chromatin Structure in Mouse Embryonic Stem Cells. Stem Cells 27: Shanahan, F., Seghezzi, W., Parry, D., Mahony, D., Lees, E., Cyclin E associates with BAF155 and BRG1, components of the mammalian SWI-SNF complex, and alters the ability of BRG1 to induce growth arrest. 19: Sohn, D., Lee, K., Lee, C., Oh, J., Chung, H., Jeon, S., Seong, R., SRG3 interacts directly with the major components of the SWI/SNF chromatin remodeling complex and protects them from proteasomal degradation. 282: Sun, F., Tang, F., Yan, A.Y., Fang, H.Y., Sheng, H.Z., Expression of SRG3, a chromatin-remodelling factor, in the mouse oocyte and early preimplantation embryos. 15: Tanaka, Y., Naruse, I., Hongo, T., Xu, M., Nakahata, T., Maekawa, T., Ishii, S., Extensive brain hemorrhage and embryonic lethality in a mouse null mutant of CREBbinding protein. 95: Timmer, J.R., Tissue morphogenesis and vascular stability require the Frem2 protein, product of the mouse myelencephalic blebs gene 102: Trainor, P.A., Specification of neural crest cell formation and migration in mouse embryos. Seminars in Cell & Developmental Biology 16:
179 Tsai, Y.C., Greco, T.M., Boonmee, A., Miteva, Y., Cristea, I.M., Functional Proteomics Establishes the Interaction of SIRT7 with Chromatin Remodeling Complexes and Expands Its Role in Regulation of RNA Polymerase I Transcription. Molecular & Cellular Proteomics 11: Vega, R.B., Matsuda, K., Oh, J., Barbosa, A.C., Yang, X., Meadows, E., McAnally, J., Pomajzl, C., Shelton, J.M., Richardson, J.A., Karsenty, G., Olson, E.N., Histone Deacetylase 4 Controls Chondrocyte Hypertrophy during Skeletogenesis 119: Wang, W., Xue, Y., Zhou, S., Kuo, A., Cairns, B.R., Crabtree, G.R., Diversity and specialization of mammalian SWI/SNF complexes. Genes Dev. 10: Wang, Y., The Role of Frizzled3 and Frizzled6 in Neural Tube Closure and in the Planar Polarity of Inner-Ear Sensory Hair Cells. J. Neurosci. 26: Wu, T.D., Nacu, S., Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26: Yamane, K., Kawabata, M., Tsuruo, T., A DNA-topoisomerase-II-binding protein with eight repeating regions similar to DNA-repair enzymes and to a cell-cycle regulator. 25: Yang, X., DNA Topoisomerase II and Neural Development. Science 287, Yao, T.P., Oh, S.P., Fuchs, M., Zhou, N.D., Ch'ng, L.E., Newsome, D., Bronson, R.T., Li, E., Livingston, D.M., Eckner, R., Gene dosage-dependent embryonic development and proliferation defects in mice lacking the transcriptional integrator p : Yu, H., Smallwood, P.M., Wang, Y., Vidaltamayo, R., Reed, R., Nathans, J., Frizzled 1 and frizzled 2 genes function in palate, ventricular septum and neural tube closure: general implications for tissue fusion processes. Development 137:
180 3717. Zhang, B., Chambers, K.J., Faller, D.V., Wang, S., Reprogramming of the SWI/SNF complex for co-activation or co-repression in prohibitin-mediated estrogen receptor regulation. Oncogene 26: FIGURE LEGENDS Figure 1. Baf155 msp3/msp3 embryos show neural tube defects. A-C: Lateral views of embryos at the indicated stages. A) Within a single litter, three individual homozygous Baf155 msp3 mutant (MUT1-3) embryos show a range of phenotypes including failure of neural tube closure and developmental delay compared to wildtype (WT). B) E12.5 Baf155 msp3/msp3 mutant shows exencephaly without other apparent defects. C) E15.5 mutant with exencephaly and coloboma. Red scale bar=1mm, blue scale bars=2cm Figure 2. BAF155 msp3 associates with other core BAF complex proteins. A) Schematic of the protein domain structure of BAF155. The msp3 ENU-induced mutation causes a C to A change at Chr9 bp (Build 36) in the Smarcc1 gene and leads to a threonine to lysine conversion of amino acid 416 in the BAF155 protein. This missense mutation alters an evolutionarily conserved amino acid in a conserved region, but not in a known structural domain. B) RT-PCR (top) and Western blot (bottom) analysis of Baf155 expression from E11.5 cranial tissue lysate. Gapdh -Actin serve as internal standards. C) Confocal microscope images of neuroepithelial sections of E9.5 WT and mutant cranial neural tubes. Panels represent wildtype and mutant nuclei stained with antibodies against BAF155 (red) and BRG1 (green). Hoechst stains nuclei (blue). White scale bars=2 um D) Immunoprecipitation of protein extracts from E11.5 mutant and wildtype embryos using anti-baf155 antibody followed by western blotting with anti- 168
181 BRG1, anti-baf155 and anti-baf170. E) Schematic of the BAF complex proteins associated with the mutant BAF155 protein. BAF155 T416K associates with BRG1 and BAF170 (IP), and BAF47 and BAF60a/b (yeast two-hybrid, supplemental data). BRG1, BAF170, BAF47 and BAF155 are considered core complex proteins. Figure 3. BAF155 function is not necessary for early neural patterning. A-C) Transverse sections of E9.5 embryos followed by immunofluorescence for dorsal/ventral markers Sonic Hedgehog (Shh) (A), NKX2.2 (B) and PAX3 (C). D and E) Lateral and F) frontal views of whole mount RNA in situ hybridization on E10.0 WT and Baf155 msp3/msp3 mutant embryos with probes to anterior/posterior markers Fgf8 (D) and FoxG1 (E and F). G) Confocal images of transverse sections of E10.5 WT and mutant embryos incubated with anti-p75 ntr antibody to visualize the enteric neural crest cells. Hoechst stains nuclei. H) Transverse sections of E10.5 rostral spinal cords hybridized with TFAP2α probe marking early migrating neural crest cells. I) Lateral view of E10.5 WT and mutant embryos hybridized with Sox10 probe to mark early neural crest cells. White scale bars=100 um, Red scale bars= 1mm. Figure 4. BAF155 function is necessary to maintain proliferation and cell survival. A) Transverse sections of E9.5 hindbrains of WT and Baf155 msp3 mutant embryos followed by immunofluorescence for proliferation marker p-h3 (green), S phase marker BrdU (½ hr BrdU incorporation) (red) and merged image with Hoechst stained nuclei. B) Transverse sections of E9.5 hindbrains in WT and Mutant embryos to visualize apoptotic cells by TUNEL staining. C) Confocal images of transverse sections through rostral spinal cords. Anti-Tuj1 antibody was used to visualize neural differentiation in neuroepithelial cells. D) Quantification of anti-phospho-h3 positive cells in WT and Mutant hindbrain neural tube. 169
182 E) Quantification of anti-brdu positive cells in WT and Mutant hindbrain neural tube. F) Quantification of apoptosis by TUNEL staining in WT and Mutant hindbrain neural tube. Ratio of p-h3, BrdU or TUNEL positive cells were calculated by dividing the number of positive stained cells by the total number of neuroepithelial cells, indicated by Hoechst staining, then multiplying by Error bars indicate SEM of at least 3 sections from 3 biological replicates. (*) P<0.01, Student s t-test. The neural tube is outlined by yellow or white dots. Figure 5. Variable gene expression in Baf155 msp3/msp3 mutant cranial tissue. A) A multidimensional scaling (MDS) plot of differentially expressed genes from cranial tissue of three somite matched WT and three somite matched Baf155 msp3/msp3 mutants. B) B) Genes were ranked in ascending fashion according to their coefficient of variation observed in wild type samples, and for each gene, the distance from mean was plotted as the log2 of the ratio of the expression level of a gene in an individual sample normalized to the mean of the expression level of the respective gene in the wildtype samples. Top: All three wild type samples; bottom: all three mutant samples; right: individual samples, from top: wild type 1 (blue), wild type 2 (red), wild type 3 (green), mutant 1 (purple), mutant 2 (turquoise), mutant 3 (orange). Supplementary Figure 1. BAF155 msp3 interacts with BAF60a, BAF60b and BAF47. Yeast two-hybrid assay. Full length cdna of Baf155 and Baf155 msp3 were cloned into pdest22 (Binding domain, BD), and Baf60a, Baf60b, and Baf47 were cloned into pdest32 (Activation domain, AD) vectors for the yeast two-hybrid assay. Reciprocal experiments were done in which the Binding and Activation domains were switched. The transformed cultures from the indicated strains were streaked onto synthetic complete medium plates 170
183 lacking leucine, tryptophan, and histidine and containing 0.5 mm 3-amino-1,2,4-triazole (- His +0.5mM AT), or the cultures were streaked onto SC-Trp-Leu (+His) plates. The bottom panels represent negative controls using some of the single plasmid strains. The plates were incubated at 32 C for 2-4 days for interaction analysis. TABLES Table 1. Phenotype and genotype of embryos resulting from cross between heterozygous Baf155 msp3/+ mice. Supplemental Table 1. List of misregulated genes in cranial tissue from averaged WT and averaged Mutant E9.5 embryos based on EdgeR analysis. Top list comprises upregulated genes, bottom list comprises downregulated genes. FC=Fold change >1.5; FDR=False Discovery Rate>0.05. Supplemental Table 2. Twenty-seven overlapping genes misexpressed in cranial tissue of each individual mutant sample compared to the average of all 3 WT samples. 171
184 172
185 173
186 174
187 175
188 Table 1. Phenotype of Baf155msp3/msp3 embryos exencephaly [with dev delay] dev delay (unable to score exencephaly) normal morphology +/+ +/- -/- E [9] 5 3 E [8] 3 4 E [4] - 1 E [1] 2 - E [0] 2 - E [0] 1 - E [0] - - E Adult
189 Supplemental Table 1. Neural development= ND, Apoptosis= AP Cell survival/cell Death=CS Centrosome/cilia=CN Global regulators= GR Adhesion/Cell polarity=ad Cellular development, growth and proliferation, cell morphology= GP Upregulated GeneID FC P Value FDR Category Top2b E ND, GR, GP Kif E E-11 ND, CN Metrnl E E-13 Spon E E-13 ND Abcg E E-08 GP Cxcl E Fzd E E-05 ND Fam72a AP, CS Sirt E E-14 AP, GR Sema3c E ND, CS Coil E Nfkb E E-07 ND, CS Fam71d E E-07 Sst E E-14 ND, AP, CS Pcbp E E-20 CS Stxbp ND Cand GP Dgkd E E-05 Plcg E ND, CS Map3k E E-16 ND, GP, CS Tspan E E-06 Lmo E CS Zfp Sez6l Ctse E AP, CS Ppat E E-09 Ctnna E E-05 ND, CS Adora E AP, CS Ndfip ND, AD Bcl2a1d CS Srgap E ND Samd
190 Tsta ND Slc22a Glyctk E Adipor E AD Kcnmb E Cyb5r E E-06 Epha E E-06 AD, GP Oca E CS Ccdc E Rassf E E-06 Ryk ND Nat E CS Dnajc E E-05 GP Maf Ccdc E Cnga E Sult6b E Cdk ND Nipal E Fzd E ND Frem ND, AD Ablim E ND H2-DMa E E-05 CS, AD St E GP Hsf E E-09 ND, GP Thtpa E GP Enthd E Tpm E CS Wdfy E Pla2g12b E E-07 AP, GP Nts E CS Itga E E-05 ND, CS AD Nars Creb Card E GP Secisbp E GP Ube2e GP Erbb E AD, GP Hfm E Utp11l E Faf E AP, GP Gjc E AD 178
191 Dock E ND, CS Pisd-ps E Iqcf Chd E ND Downregulated: Gene ID FC P Value FDR Category Cldn E AD, GP Abca8b E Vsig E Hbp GP Svep E Cyp4f E E-05 Ints Abl E AD, GP Elac ND Pla2g E E-05 Rnf E CS Dhx E E-14 ND Rassf E E-05 Ms4a Apob E CS Gatad2b E E-06 ND Col15a E E-12 Mep1a E CS Tusc E AP Aqp E E-05 Pcp4l E Poc1a E E-53 CN 179
192 Supplemental Table 2. Log2 Fold Change Mut 2 vs 3 Mut 1 vs 3 Mut 1 vs 2 Col15a Col25a Creb Cyb5r Dab Dhx Dock Gatad2b H2-DMa Hfm Ift Loh12cr Map3k Mreg Nmu Pcbp Pdcl Pla2g12b Pla2g Poc1a Ppat Rasgrf Secisbp Sst Svep Ten Vsig
Limb Development. Limb Development. Limb Axes. Limb Development. Clinical Terms
 Limb Development Overview of Limb Formation Initiation of Limb Development Limb Field Outgrowth of the Mesoderm Morphogenetic Signaling Development of Limb Tissues Skeleton Musculature Innervation Vasculature
Limb Development Overview of Limb Formation Initiation of Limb Development Limb Field Outgrowth of the Mesoderm Morphogenetic Signaling Development of Limb Tissues Skeleton Musculature Innervation Vasculature
Module 3 Questions. 7. Chemotaxis is an example of signal transduction. Explain, with the use of diagrams.
 Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Lecture 3: Mutations
 Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
Histone modifications. and ChIP. G. Valle - Università di Padova
 Histone modifications and ChIP Histone acetylation It has been shown that the acetylation of lysines is highly dynamic and regulated by the opposing action of two families of enzymes, histone acetyltransferases
Histone modifications and ChIP Histone acetylation It has been shown that the acetylation of lysines is highly dynamic and regulated by the opposing action of two families of enzymes, histone acetyltransferases
Chapter 47: Animal Development
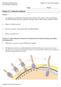 Name Period Overview 1. An organism s development is controlled by the genome of the zygote as well as by molecules from the mother that are in the cytoplasm of the egg. What are these proteins and RNAs
Name Period Overview 1. An organism s development is controlled by the genome of the zygote as well as by molecules from the mother that are in the cytoplasm of the egg. What are these proteins and RNAs
http://www.springer.com/3-540-23372-5
 http://www.springer.com/3-540-23372-5 Chromatin Remodeling Factors and DNA Replication Patrick Varga-Weisz Abstract Chromatin structures have to be precisely duplicated during DNA replication to maintain
http://www.springer.com/3-540-23372-5 Chromatin Remodeling Factors and DNA Replication Patrick Varga-Weisz Abstract Chromatin structures have to be precisely duplicated during DNA replication to maintain
Trasposable elements: P elements
 Trasposable elements: P elements In 1938 Marcus Rhodes provided the first genetic description of an unstable mutation, an allele of a gene required for the production of pigment in maize. This instability
Trasposable elements: P elements In 1938 Marcus Rhodes provided the first genetic description of an unstable mutation, an allele of a gene required for the production of pigment in maize. This instability
GENE REGULATION. Teacher Packet
 AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
1 Mutation and Genetic Change
 CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
Genetics Lecture Notes 7.03 2005. Lectures 1 2
 Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Psychoonkology, Sept. 2010 lifestyle factors and epigenetics
 Psychoonkology, Sept. 2010 lifestyle factors and epigenetics Alexander G. Haslberger Dep. für Ernährungswissenschaften Univ. of Vienna Working group: Food, GI-Microbiology, Epigenetics Content Health:
Psychoonkology, Sept. 2010 lifestyle factors and epigenetics Alexander G. Haslberger Dep. für Ernährungswissenschaften Univ. of Vienna Working group: Food, GI-Microbiology, Epigenetics Content Health:
EPIGENETICS DNA and Histone Model
 EPIGENETICS ABSTRACT A 3-D cut-and-paste model depicting how histone, acetyl and methyl molecules control access to DNA and affect gene expression. LOGISTICS TIME REQUIRED LEARNING OBJECTIVES DNA is coiled
EPIGENETICS ABSTRACT A 3-D cut-and-paste model depicting how histone, acetyl and methyl molecules control access to DNA and affect gene expression. LOGISTICS TIME REQUIRED LEARNING OBJECTIVES DNA is coiled
Cancer SBL101. James Gomes School of Biological Sciences Indian Institute of Technology Delhi
 Cancer SBL101 James Gomes School of Biological Sciences Indian Institute of Technology Delhi All Figures in this Lecture are taken from 1. Molecular biology of the cell / Bruce Alberts et al., 5th ed.
Cancer SBL101 James Gomes School of Biological Sciences Indian Institute of Technology Delhi All Figures in this Lecture are taken from 1. Molecular biology of the cell / Bruce Alberts et al., 5th ed.
AP Biology Essential Knowledge Student Diagnostic
 AP Biology Essential Knowledge Student Diagnostic Background The Essential Knowledge statements provided in the AP Biology Curriculum Framework are scientific claims describing phenomenon occurring in
AP Biology Essential Knowledge Student Diagnostic Background The Essential Knowledge statements provided in the AP Biology Curriculum Framework are scientific claims describing phenomenon occurring in
Control of Gene Expression
 Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
Control of Gene Expression (Learning Objectives) Explain the role of gene expression is differentiation of function of cells which leads to the emergence of different tissues, organs, and organ systems
The first steps to forming a new organism Descriptive embryology 2. Cleavage, Gastrulation, Neurulation and Organogenesis
 Developmental Biology BY1101 P. Murphy Lectures 4 and 5 The first steps to forming a new organism Descriptive embryology 2 Cleavage, Gastrulation, Neurulation and Organogenesis Early animal development
Developmental Biology BY1101 P. Murphy Lectures 4 and 5 The first steps to forming a new organism Descriptive embryology 2 Cleavage, Gastrulation, Neurulation and Organogenesis Early animal development
The Developing Person Through the Life Span 8e by Kathleen Stassen Berger
 The Developing Person Through the Life Span 8e by Kathleen Stassen Berger Chapter 3 Heredity and Environment PowerPoint Slides developed by Martin Wolfger and Michael James Ivy Tech Community College-Bloomington
The Developing Person Through the Life Span 8e by Kathleen Stassen Berger Chapter 3 Heredity and Environment PowerPoint Slides developed by Martin Wolfger and Michael James Ivy Tech Community College-Bloomington
Summary of Ph.D. thesis. Tamás Matusek. Supervisor: József Mihály Ph.D.
 Summary of Ph.D. thesis GENETIC, CELL BIOLOGICAL AND BIOCHEMICAL ANALYSIS OF THE TISSUE SPECIFIC FUNCTIONS OF A NEW DROSOPHILA FORMIN Tamás Matusek Supervisor: József Mihály Ph.D. Biological Research Center
Summary of Ph.D. thesis GENETIC, CELL BIOLOGICAL AND BIOCHEMICAL ANALYSIS OF THE TISSUE SPECIFIC FUNCTIONS OF A NEW DROSOPHILA FORMIN Tamás Matusek Supervisor: József Mihály Ph.D. Biological Research Center
CONTRACTING ORGANIZATION: University of Alabama at Birmingham Birmingham, AL 35294
 AD Award Number: W81XWH-08-1-0030 TITLE: Regulation of Prostate Cancer Bone Metastasis by DKK1 PRINCIPAL INVESTIGATOR: Gregory A. Clines, M.D., Ph.D. CONTRACTING ORGANIZATION: University of Alabama at
AD Award Number: W81XWH-08-1-0030 TITLE: Regulation of Prostate Cancer Bone Metastasis by DKK1 PRINCIPAL INVESTIGATOR: Gregory A. Clines, M.D., Ph.D. CONTRACTING ORGANIZATION: University of Alabama at
Control of Gene Expression
 Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Neural Tube Defects - NTDs
 Neural Tube Defects - NTDs Introduction Neural tube defects are also known as NTDs. They happen when the spine and brain do not fully develop while the fetus is forming in the uterus. Worldwide, there
Neural Tube Defects - NTDs Introduction Neural tube defects are also known as NTDs. They happen when the spine and brain do not fully develop while the fetus is forming in the uterus. Worldwide, there
Chapter 7. Physical interaction between Jamb and Jamc is necessary for myoblast fusion. Summary
 Chapter 7 Physical interaction between Jamb and Jamc is necessary for myoblast fusion Summary In this chapter I describe a series of transplant experiments designed to further elucidate the function and
Chapter 7 Physical interaction between Jamb and Jamc is necessary for myoblast fusion Summary In this chapter I describe a series of transplant experiments designed to further elucidate the function and
Reproductive System & Development: Practice Questions #1
 Reproductive System & Development: Practice Questions #1 1. Which two glands in the diagram produce gametes? A. glands A and B B. glands B and E C. glands C and F D. glands E and F 2. Base your answer
Reproductive System & Development: Practice Questions #1 1. Which two glands in the diagram produce gametes? A. glands A and B B. glands B and E C. glands C and F D. glands E and F 2. Base your answer
Sickle cell anemia: Altered beta chain Single AA change (#6 Glu to Val) Consequence: Protein polymerizes Change in RBC shape ---> phenotypes
 Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
CELL MEMBRANES, TRANSPORT, and COMMUNICATION. Teacher Packet
 AP * BIOLOGY CELL MEMBRANES, TRANSPORT, and COMMUNICATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production
AP * BIOLOGY CELL MEMBRANES, TRANSPORT, and COMMUNICATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production
Cell Division Mitosis and the Cell Cycle
 Cell Division Mitosis and the Cell Cycle A Chromosome and Sister Chromatids Key Points About Chromosome Structure A chromosome consists of DNA that is wrapped around proteins (histones) and condensed Each
Cell Division Mitosis and the Cell Cycle A Chromosome and Sister Chromatids Key Points About Chromosome Structure A chromosome consists of DNA that is wrapped around proteins (histones) and condensed Each
Genetics Module B, Anchor 3
 Genetics Module B, Anchor 3 Key Concepts: - An individual s characteristics are determines by factors that are passed from one parental generation to the next. - During gamete formation, the alleles for
Genetics Module B, Anchor 3 Key Concepts: - An individual s characteristics are determines by factors that are passed from one parental generation to the next. - During gamete formation, the alleles for
Appendix C DNA Replication & Mitosis
 K.Muma Bio 6 Appendix C DNA Replication & Mitosis Study Objectives: Appendix C: DNA replication and Mitosis 1. Describe the structure of DNA and where it is found. 2. Explain complimentary base pairing:
K.Muma Bio 6 Appendix C DNA Replication & Mitosis Study Objectives: Appendix C: DNA replication and Mitosis 1. Describe the structure of DNA and where it is found. 2. Explain complimentary base pairing:
Developmental Disorders
 Developmental Disorders General Principles Causes: Genetic Environmental (Maternal, Physical, Chemical) Mechanisms (Retinoic Acid) General Principles Teratology Study of Monsters Teratogen agent that produces
Developmental Disorders General Principles Causes: Genetic Environmental (Maternal, Physical, Chemical) Mechanisms (Retinoic Acid) General Principles Teratology Study of Monsters Teratogen agent that produces
MUTATION, DNA REPAIR AND CANCER
 MUTATION, DNA REPAIR AND CANCER 1 Mutation A heritable change in the genetic material Essential to the continuity of life Source of variation for natural selection New mutations are more likely to be harmful
MUTATION, DNA REPAIR AND CANCER 1 Mutation A heritable change in the genetic material Essential to the continuity of life Source of variation for natural selection New mutations are more likely to be harmful
Control of Gene Expression
 Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Cell Biology Questions and Learning Objectives
 Cell Biology Questions and Learning Objectives (with hypothetical learning materials that might populate the objective) The topics and central questions listed here are typical for an introductory undergraduate
Cell Biology Questions and Learning Objectives (with hypothetical learning materials that might populate the objective) The topics and central questions listed here are typical for an introductory undergraduate
STEM CELL FELLOWSHIP
 Module I: The Basic Principles of Stem Cells 1. Basics of Stem Cells a. Understanding the development of embryonic stem cells i. Embryonic stem cells ii. Embryonic germ cells iii. Differentiated stem cell
Module I: The Basic Principles of Stem Cells 1. Basics of Stem Cells a. Understanding the development of embryonic stem cells i. Embryonic stem cells ii. Embryonic germ cells iii. Differentiated stem cell
The Making of the Fittest: Evolving Switches, Evolving Bodies
 OVERVIEW MODELING THE REGULATORY SWITCHES OF THE PITX1 GENE IN STICKLEBACK FISH This hands-on activity supports the short film, The Making of the Fittest:, and aims to help students understand eukaryotic
OVERVIEW MODELING THE REGULATORY SWITCHES OF THE PITX1 GENE IN STICKLEBACK FISH This hands-on activity supports the short film, The Making of the Fittest:, and aims to help students understand eukaryotic
Chapter 5: Organization and Expression of Immunoglobulin Genes
 Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
How To Understand How Gene Expression Is Regulated
 What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
Influence of Sex on Genetics. Chapter Six
 Influence of Sex on Genetics Chapter Six Humans 23 Autosomes Chromosomal abnormalities very severe Often fatal All have at least one X Deletion of X chromosome is fatal Males = heterogametic sex XY Females
Influence of Sex on Genetics Chapter Six Humans 23 Autosomes Chromosomal abnormalities very severe Often fatal All have at least one X Deletion of X chromosome is fatal Males = heterogametic sex XY Females
2006 7.012 Problem Set 6 KEY
 2006 7.012 Problem Set 6 KEY ** Due before 5 PM on WEDNESDAY, November 22, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You create an artificial
2006 7.012 Problem Set 6 KEY ** Due before 5 PM on WEDNESDAY, November 22, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You create an artificial
B Cell Generation, Activation & Differentiation. B cell maturation
 B Cell Generation, Activation & Differentiation Naïve B cells- have not encountered Ag. Have IgM and IgD on cell surface : have same binding VDJ regions but different constant region leaves bone marrow
B Cell Generation, Activation & Differentiation Naïve B cells- have not encountered Ag. Have IgM and IgD on cell surface : have same binding VDJ regions but different constant region leaves bone marrow
Sample Questions for Exam 3
 Sample Questions for Exam 3 1. All of the following occur during prometaphase of mitosis in animal cells except a. the centrioles move toward opposite poles. b. the nucleolus can no longer be seen. c.
Sample Questions for Exam 3 1. All of the following occur during prometaphase of mitosis in animal cells except a. the centrioles move toward opposite poles. b. the nucleolus can no longer be seen. c.
An Overview of Cells and Cell Research
 An Overview of Cells and Cell Research 1 An Overview of Cells and Cell Research Chapter Outline Model Species and Cell types Cell components Tools of Cell Biology Model Species E. Coli: simplest organism
An Overview of Cells and Cell Research 1 An Overview of Cells and Cell Research Chapter Outline Model Species and Cell types Cell components Tools of Cell Biology Model Species E. Coli: simplest organism
12.1 The Role of DNA in Heredity
 12.1 The Role of DNA in Heredity Only in the last 50 years have scientists understood the role of DNA in heredity. That understanding began with the discovery of DNA s structure. In 1952, Rosalind Franklin
12.1 The Role of DNA in Heredity Only in the last 50 years have scientists understood the role of DNA in heredity. That understanding began with the discovery of DNA s structure. In 1952, Rosalind Franklin
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Name Class Date. Figure 13 1. 2. Which nucleotide in Figure 13 1 indicates the nucleic acid above is RNA? a. uracil c. cytosine b. guanine d.
 13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
Gene mutation and molecular medicine Chapter 15
 Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
The correct answer is c A. Answer a is incorrect. The white-eye gene must be recessive since heterozygous females have red eyes.
 1. Why is the white-eye phenotype always observed in males carrying the white-eye allele? a. Because the trait is dominant b. Because the trait is recessive c. Because the allele is located on the X chromosome
1. Why is the white-eye phenotype always observed in males carrying the white-eye allele? a. Because the trait is dominant b. Because the trait is recessive c. Because the allele is located on the X chromosome
Heredity - Patterns of Inheritance
 Heredity - Patterns of Inheritance Genes and Alleles A. Genes 1. A sequence of nucleotides that codes for a special functional product a. Transfer RNA b. Enzyme c. Structural protein d. Pigments 2. Genes
Heredity - Patterns of Inheritance Genes and Alleles A. Genes 1. A sequence of nucleotides that codes for a special functional product a. Transfer RNA b. Enzyme c. Structural protein d. Pigments 2. Genes
Antigenic variation in Plasmodium falciparum : Erythrocyte invasion and immune escape mechanisms
 Antigenic variation in Plasmodium falciparum : Erythrocyte invasion and immune escape mechanisms Introduction Why does immunity to malaria take so long to develop? The parasite s survival depends on its
Antigenic variation in Plasmodium falciparum : Erythrocyte invasion and immune escape mechanisms Introduction Why does immunity to malaria take so long to develop? The parasite s survival depends on its
AP Biology 2015 Free-Response Questions
 AP Biology 2015 Free-Response Questions College Board, Advanced Placement Program, AP, AP Central, and the acorn logo are registered trademarks of the College Board. AP Central is the official online home
AP Biology 2015 Free-Response Questions College Board, Advanced Placement Program, AP, AP Central, and the acorn logo are registered trademarks of the College Board. AP Central is the official online home
5. MESODERM FORMATION / SEGMENTATION
 5. MESODERM FORMATION / SEGMENTATION Dr. Ann -Judith Silverman Department of Anatomy & Cell Biology Telephone: 305-3540 E-mail: [email protected] READING ASSIGNMENT: Larsen Human Embryology, 3rd Edition,
5. MESODERM FORMATION / SEGMENTATION Dr. Ann -Judith Silverman Department of Anatomy & Cell Biology Telephone: 305-3540 E-mail: [email protected] READING ASSIGNMENT: Larsen Human Embryology, 3rd Edition,
Complex multicellular organisms are produced by cells that switch genes on and off during development.
 Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Neurotrophic factors and Their receptors
 Neurotrophic factors and Their receptors Huang Shu-Hong Institute of neurobiology 1 For decades, scientists believed that brain cells of the central nervous system could not regrow following damage due
Neurotrophic factors and Their receptors Huang Shu-Hong Institute of neurobiology 1 For decades, scientists believed that brain cells of the central nervous system could not regrow following damage due
What is Cancer? Cancer is a genetic disease: Cancer typically involves a change in gene expression/function:
 Cancer is a genetic disease: Inherited cancer Sporadic cancer What is Cancer? Cancer typically involves a change in gene expression/function: Qualitative change Quantitative change Any cancer causing genetic
Cancer is a genetic disease: Inherited cancer Sporadic cancer What is Cancer? Cancer typically involves a change in gene expression/function: Qualitative change Quantitative change Any cancer causing genetic
www.njctl.org PSI Biology Mitosis & Meiosis
 Mitosis and Meiosis Mitosis Classwork 1. Identify two differences between meiosis and mitosis. 2. Provide an example of a type of cell in the human body that would undergo mitosis. 3. Does cell division
Mitosis and Meiosis Mitosis Classwork 1. Identify two differences between meiosis and mitosis. 2. Provide an example of a type of cell in the human body that would undergo mitosis. 3. Does cell division
CHAPTER 15 THE CHROMOSOMAL BASIS OF INHERITANCE. Section B: Sex Chromosomes
 CHAPTER 15 THE CHROMOSOMAL BASIS OF INHERITANCE Section B: Sex Chromosomes 1. The chromosomal basis of sex varies with the organism 2. Sex-linked genes have unique patterns of inheritance 1. The chromosomal
CHAPTER 15 THE CHROMOSOMAL BASIS OF INHERITANCE Section B: Sex Chromosomes 1. The chromosomal basis of sex varies with the organism 2. Sex-linked genes have unique patterns of inheritance 1. The chromosomal
Autoimmunity and immunemediated. FOCiS. Lecture outline
 1 Autoimmunity and immunemediated inflammatory diseases Abul K. Abbas, MD UCSF FOCiS 2 Lecture outline Pathogenesis of autoimmunity: why selftolerance fails Genetics of autoimmune diseases Therapeutic
1 Autoimmunity and immunemediated inflammatory diseases Abul K. Abbas, MD UCSF FOCiS 2 Lecture outline Pathogenesis of autoimmunity: why selftolerance fails Genetics of autoimmune diseases Therapeutic
Replication Study Guide
 Replication Study Guide This study guide is a written version of the material you have seen presented in the replication unit. Self-reproduction is a function of life that human-engineered systems have
Replication Study Guide This study guide is a written version of the material you have seen presented in the replication unit. Self-reproduction is a function of life that human-engineered systems have
Question Points Score. Total for Final: 200 1 16 2 17 3 18 4 15 5 10 6 10 7 8 8 12 9 28 10 10 11 12 12 12 13 10 14 16 15 6.
 Question Points Score Total for Final: 200 1 16 2 17 3 18 4 15 5 10 6 10 7 8 8 12 9 28 10 10 11 12 12 12 13 10 14 16 15 6 200 Page 1 of 19 Question 1. You are given four slides that contain cuticle preparations
Question Points Score Total for Final: 200 1 16 2 17 3 18 4 15 5 10 6 10 7 8 8 12 9 28 10 10 11 12 12 12 13 10 14 16 15 6 200 Page 1 of 19 Question 1. You are given four slides that contain cuticle preparations
Plant Growth & Development. Growth Stages. Differences in the Developmental Mechanisms of Plants and Animals. Development
 Plant Growth & Development Plant body is unable to move. To survive and grow, plants must be able to alter its growth, development and physiology. Plants are able to produce complex, yet variable forms
Plant Growth & Development Plant body is unable to move. To survive and grow, plants must be able to alter its growth, development and physiology. Plants are able to produce complex, yet variable forms
Algorithms in Computational Biology (236522) spring 2007 Lecture #1
 Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
UNIT 13 (OPTION) Genetic Abnormalities
 Unit 13 Genetic Abnormailities 1 UNIT 13 (OPTION) Genetic Abnormalities Originally developed by: Hildur Helgedottir RN, MN Revised (2000) by: Marlene Reimer RN, PhD, CCN (C) Associate Professor Faculty
Unit 13 Genetic Abnormailities 1 UNIT 13 (OPTION) Genetic Abnormalities Originally developed by: Hildur Helgedottir RN, MN Revised (2000) by: Marlene Reimer RN, PhD, CCN (C) Associate Professor Faculty
Chromosomes, Mapping, and the Meiosis Inheritance Connection
 Chromosomes, Mapping, and the Meiosis Inheritance Connection Carl Correns 1900 Chapter 13 First suggests central role for chromosomes Rediscovery of Mendel s work Walter Sutton 1902 Chromosomal theory
Chromosomes, Mapping, and the Meiosis Inheritance Connection Carl Correns 1900 Chapter 13 First suggests central role for chromosomes Rediscovery of Mendel s work Walter Sutton 1902 Chromosomal theory
CHROMOSOMES Dr. Fern Tsien, Dept. of Genetics, LSUHSC, NO, LA
 CHROMOSOMES Dr. Fern Tsien, Dept. of Genetics, LSUHSC, NO, LA Cytogenetics is the study of chromosomes and their structure, inheritance, and abnormalities. Chromosome abnormalities occur in approximately:
CHROMOSOMES Dr. Fern Tsien, Dept. of Genetics, LSUHSC, NO, LA Cytogenetics is the study of chromosomes and their structure, inheritance, and abnormalities. Chromosome abnormalities occur in approximately:
Cystic Fibrosis Webquest Sarah Follenweider, The English High School 2009 Summer Research Internship Program
 Cystic Fibrosis Webquest Sarah Follenweider, The English High School 2009 Summer Research Internship Program Introduction: Cystic fibrosis (CF) is an inherited chronic disease that affects the lungs and
Cystic Fibrosis Webquest Sarah Follenweider, The English High School 2009 Summer Research Internship Program Introduction: Cystic fibrosis (CF) is an inherited chronic disease that affects the lungs and
Becker Muscular Dystrophy
 Muscular Dystrophy A Case Study of Positional Cloning Described by Benjamin Duchenne (1868) X-linked recessive disease causing severe muscular degeneration. 100 % penetrance X d Y affected male Frequency
Muscular Dystrophy A Case Study of Positional Cloning Described by Benjamin Duchenne (1868) X-linked recessive disease causing severe muscular degeneration. 100 % penetrance X d Y affected male Frequency
FGF-1 as Cosmetic Supplement
 FGF-1 as Cosmetic Supplement Ing-Ming Chiu, Ph.D. Professor, Internal Medicine and Molecular and Cellular Biochemistry The Ohio State University Columbus, Ohio, U.S.A. GENTEON USA Fibroblast Growth Factor
FGF-1 as Cosmetic Supplement Ing-Ming Chiu, Ph.D. Professor, Internal Medicine and Molecular and Cellular Biochemistry The Ohio State University Columbus, Ohio, U.S.A. GENTEON USA Fibroblast Growth Factor
Role Of Mitochondria In Mesenchymal Stem Cells
 Role Of Mitochondria In Mesenchymal Stem Cells Roman Eliseev, MD, PhD, Tamara Raymond, Regis O'Keefe, MD, PhD. University of Rochester, Rochester, NY, USA. Disclosures: R. Eliseev: None. T. Raymond: None.
Role Of Mitochondria In Mesenchymal Stem Cells Roman Eliseev, MD, PhD, Tamara Raymond, Regis O'Keefe, MD, PhD. University of Rochester, Rochester, NY, USA. Disclosures: R. Eliseev: None. T. Raymond: None.
Lecture 6: Single nucleotide polymorphisms (SNPs) and Restriction Fragment Length Polymorphisms (RFLPs)
 Lecture 6: Single nucleotide polymorphisms (SNPs) and Restriction Fragment Length Polymorphisms (RFLPs) Single nucleotide polymorphisms or SNPs (pronounced "snips") are DNA sequence variations that occur
Lecture 6: Single nucleotide polymorphisms (SNPs) and Restriction Fragment Length Polymorphisms (RFLPs) Single nucleotide polymorphisms or SNPs (pronounced "snips") are DNA sequence variations that occur
T Cell Maturation,Activation and Differentiation
 T Cell Maturation,Activation and Differentiation Positive Selection- In thymus, permits survival of only those T cells whose TCRs recognize self- MHC molecules (self-mhc restriction) Negative Selection-
T Cell Maturation,Activation and Differentiation Positive Selection- In thymus, permits survival of only those T cells whose TCRs recognize self- MHC molecules (self-mhc restriction) Negative Selection-
Cellular Reproduction
 9 Cellular Reproduction section 1 Cellular Growth Before You Read Think about the life cycle of a human. On the lines below, write some of the stages that occur in the life cycle of a human. In this section,
9 Cellular Reproduction section 1 Cellular Growth Before You Read Think about the life cycle of a human. On the lines below, write some of the stages that occur in the life cycle of a human. In this section,
Hormones & Chemical Signaling
 Hormones & Chemical Signaling Part 2 modulation of signal pathways and hormone classification & function How are these pathways controlled? Receptors are proteins! Subject to Specificity of binding Competition
Hormones & Chemical Signaling Part 2 modulation of signal pathways and hormone classification & function How are these pathways controlled? Receptors are proteins! Subject to Specificity of binding Competition
LECTURE 6 Gene Mutation (Chapter 16.1-16.2)
 LECTURE 6 Gene Mutation (Chapter 16.1-16.2) 1 Mutation: A permanent change in the genetic material that can be passed from parent to offspring. Mutant (genotype): An organism whose DNA differs from the
LECTURE 6 Gene Mutation (Chapter 16.1-16.2) 1 Mutation: A permanent change in the genetic material that can be passed from parent to offspring. Mutant (genotype): An organism whose DNA differs from the
Neural tube defects: open spina bifida (also called spina bifida cystica)
 Screening Programmes Fetal Anomaly Neural tube defects: open spina bifida (also called spina bifida cystica) Information for health professionals Publication date: April 2012 Review date: April 2013 Version
Screening Programmes Fetal Anomaly Neural tube defects: open spina bifida (also called spina bifida cystica) Information for health professionals Publication date: April 2012 Review date: April 2013 Version
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE Q5B
 INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
Biology 1406 - Notes for exam 5 - Population genetics Ch 13, 14, 15
 Biology 1406 - Notes for exam 5 - Population genetics Ch 13, 14, 15 Species - group of individuals that are capable of interbreeding and producing fertile offspring; genetically similar 13.7, 14.2 Population
Biology 1406 - Notes for exam 5 - Population genetics Ch 13, 14, 15 Species - group of individuals that are capable of interbreeding and producing fertile offspring; genetically similar 13.7, 14.2 Population
The Nucleus: DNA, Chromatin And Chromosomes
 The Nucleus: DNA, Chromatin And Chromosomes Professor Alfred Cuschieri Department of Anatomy, University of Malta. Objectives By the end of this unit the student should be able to: 1. List the major structural
The Nucleus: DNA, Chromatin And Chromosomes Professor Alfred Cuschieri Department of Anatomy, University of Malta. Objectives By the end of this unit the student should be able to: 1. List the major structural
Optional Tests Offered Before and During Pregnancy
 Plano Women s Healthcare Optional Tests Offered Before and During Pregnancy Alpha-Fetoprotein Test (AFP) and Quad Screen These are screening tests that can assess your baby s risk of having such birth
Plano Women s Healthcare Optional Tests Offered Before and During Pregnancy Alpha-Fetoprotein Test (AFP) and Quad Screen These are screening tests that can assess your baby s risk of having such birth
Structure and Function of DNA
 Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Skeletal Development Multiple Cellular Origins
 Skeletal Development Multiple Cellular Origins 1 - Paraxial Mesoderm Somite, Sclerotome Axial Skeleton (e.g. vertebra) 2 - Lateral Plate Mesoderm Appendicular Skeleton (e.g. limb) 3 - Neural Crest Head
Skeletal Development Multiple Cellular Origins 1 - Paraxial Mesoderm Somite, Sclerotome Axial Skeleton (e.g. vertebra) 2 - Lateral Plate Mesoderm Appendicular Skeleton (e.g. limb) 3 - Neural Crest Head
Lab # 12: DNA and RNA
 115 116 Concepts to be explored: Structure of DNA Nucleotides Amino Acids Proteins Genetic Code Mutation RNA Transcription to RNA Translation to a Protein Figure 12. 1: DNA double helix Introduction Long
115 116 Concepts to be explored: Structure of DNA Nucleotides Amino Acids Proteins Genetic Code Mutation RNA Transcription to RNA Translation to a Protein Figure 12. 1: DNA double helix Introduction Long
Interaktionen von Nukleinsäuren und Proteinen
 Sonja Prohaska Computational EvoDevo Universitaet Leipzig June 9, 2015 DNA is never naked in a cell DNA is usually in association with proteins. In all domains of life there are small, basic chromosomal
Sonja Prohaska Computational EvoDevo Universitaet Leipzig June 9, 2015 DNA is never naked in a cell DNA is usually in association with proteins. In all domains of life there are small, basic chromosomal
Vocabulary & General Concepts of Brain Organization
 Vocabulary & General Concepts of Brain Organization Jeanette J. Norden, Ph.D. Professor Emerita Vanderbilt University School of Medicine Course Outline Lecture 1: Vocabulary & General Concepts of Brain
Vocabulary & General Concepts of Brain Organization Jeanette J. Norden, Ph.D. Professor Emerita Vanderbilt University School of Medicine Course Outline Lecture 1: Vocabulary & General Concepts of Brain
Genetic Mutations. Indicator 4.8: Compare the consequences of mutations in body cells with those in gametes.
 Genetic Mutations Indicator 4.8: Compare the consequences of mutations in body cells with those in gametes. Agenda Warm UP: What is a mutation? Body cell? Gamete? Notes on Mutations Karyotype Web Activity
Genetic Mutations Indicator 4.8: Compare the consequences of mutations in body cells with those in gametes. Agenda Warm UP: What is a mutation? Body cell? Gamete? Notes on Mutations Karyotype Web Activity
Biological Sciences Initiative. Human Genome
 Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Uses of Flow Cytometry
 Uses of Flow Cytometry 1. Multicolour analysis... 2 2. Cell Cycle and Proliferation... 3 a. Analysis of Cellular DNA Content... 4 b. Cell Proliferation Assays... 5 3. Immunology... 6 4. Apoptosis... 7
Uses of Flow Cytometry 1. Multicolour analysis... 2 2. Cell Cycle and Proliferation... 3 a. Analysis of Cellular DNA Content... 4 b. Cell Proliferation Assays... 5 3. Immunology... 6 4. Apoptosis... 7
Cell Division CELL DIVISION. Mitosis. Designation of Number of Chromosomes. Homologous Chromosomes. Meiosis
 Cell Division CELL DIVISION Anatomy and Physiology Text and Laboratory Workbook, Stephen G. Davenport, Copyright 2006, All Rights Reserved, no part of this publication can be used for any commercial purpose.
Cell Division CELL DIVISION Anatomy and Physiology Text and Laboratory Workbook, Stephen G. Davenport, Copyright 2006, All Rights Reserved, no part of this publication can be used for any commercial purpose.
School of Nursing. Presented by Yvette Conley, PhD
 Presented by Yvette Conley, PhD What we will cover during this webcast: Briefly discuss the approaches introduced in the paper: Genome Sequencing Genome Wide Association Studies Epigenomics Gene Expression
Presented by Yvette Conley, PhD What we will cover during this webcast: Briefly discuss the approaches introduced in the paper: Genome Sequencing Genome Wide Association Studies Epigenomics Gene Expression
Genetics 1. Defective enzyme that does not make melanin. Very pale skin and hair color (albino)
 Genetics 1 We all know that children tend to resemble their parents. Parents and their children tend to have similar appearance because children inherit genes from their parents and these genes influence
Genetics 1 We all know that children tend to resemble their parents. Parents and their children tend to have similar appearance because children inherit genes from their parents and these genes influence
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
Chapter 13: Meiosis and Sexual Life Cycles
 Name Period Chapter 13: Meiosis and Sexual Life Cycles Concept 13.1 Offspring acquire genes from parents by inheriting chromosomes 1. Let s begin with a review of several terms that you may already know.
Name Period Chapter 13: Meiosis and Sexual Life Cycles Concept 13.1 Offspring acquire genes from parents by inheriting chromosomes 1. Let s begin with a review of several terms that you may already know.
BCOR101 Midterm II Wednesday, October 26, 2005
 BCOR101 Midterm II Wednesday, October 26, 2005 Name Key Please show all of your work. 1. A donor strain is trp+, pro+, met+ and a recipient strain is trp-, pro-, met-. The donor strain is infected with
BCOR101 Midterm II Wednesday, October 26, 2005 Name Key Please show all of your work. 1. A donor strain is trp+, pro+, met+ and a recipient strain is trp-, pro-, met-. The donor strain is infected with
BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN. Third Examination April 2, 2009
 BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Third Examination April 2, 2009 Please put your name at the top of each page. Read each question carefully before answering- i.e. read the instructions. Be
BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Third Examination April 2, 2009 Please put your name at the top of each page. Read each question carefully before answering- i.e. read the instructions. Be
1.1.2. thebiotutor. AS Biology OCR. Unit F211: Cells, Exchange & Transport. Module 1.2 Cell Membranes. Notes & Questions.
 thebiotutor AS Biology OCR Unit F211: Cells, Exchange & Transport Module 1.2 Cell Membranes Notes & Questions Andy Todd 1 Outline the roles of membranes within cells and at the surface of cells. The main
thebiotutor AS Biology OCR Unit F211: Cells, Exchange & Transport Module 1.2 Cell Membranes Notes & Questions Andy Todd 1 Outline the roles of membranes within cells and at the surface of cells. The main
AP BIOLOGY 2012 SCORING GUIDELINES
 AP BIOLOGY 2012 SCORING GUIDELINES Question 1 Note: At least 1 point must be earned from each of parts (a), (b), (c), and (d) in order to earn a maximum score of 10. The ability to reproduce is a characteristic
AP BIOLOGY 2012 SCORING GUIDELINES Question 1 Note: At least 1 point must be earned from each of parts (a), (b), (c), and (d) in order to earn a maximum score of 10. The ability to reproduce is a characteristic
Title: Author(s): Source: Document Type: Copyright : http://naturalhistorymag.com/ Full Text: [ILLUSTRATION OMITTED]
![Title: Author(s): Source: Document Type: Copyright : http://naturalhistorymag.com/ Full Text: [ILLUSTRATION OMITTED] Title: Author(s): Source: Document Type: Copyright : http://naturalhistorymag.com/ Full Text: [ILLUSTRATION OMITTED]](/thumbs/33/16056031.jpg) Title: Author(s): Source: Document Type: Copyright : http://naturalhistorymag.com/ Full Text: Epigenetics in action: Part 2: turn-ons and turn-offs Nessa Carey Natural History. 120.5 (May 2012): p30. Article
Title: Author(s): Source: Document Type: Copyright : http://naturalhistorymag.com/ Full Text: Epigenetics in action: Part 2: turn-ons and turn-offs Nessa Carey Natural History. 120.5 (May 2012): p30. Article
Given these characteristics of life, which of the following objects is considered a living organism? W. X. Y. Z.
 Cell Structure and Organization 1. All living things must possess certain characteristics. They are all composed of one or more cells. They can grow, reproduce, and pass their genes on to their offspring.
Cell Structure and Organization 1. All living things must possess certain characteristics. They are all composed of one or more cells. They can grow, reproduce, and pass their genes on to their offspring.
