TuBe or not TuBe? dimanche 6 novembre 2011
|
|
|
- Dorcas Shaw
- 8 years ago
- Views:
Transcription
1 TuBe or not TuBe? 1
2 2
3 Early Exciting news came out: Discovery of a new communication channel between cells BACTERIAL NANOTUBES 2
4 Early Exciting news came out: Discovery of a new communication channel between cells BACTERIAL NANOTUBES Can we characterize them? What could a synthetic biology approach bring to this problem? 2
5 3
6 TuBe or not TuBe? harnessing bacterial nanotubes by and for synthetic biology 3
7 Harnessing the possibilities of the nanotube network Factory Pattern formation Amorphous computing 4
8 Back to reality Dubey, G.P. & Ben-Yehuda, S. Intercellular Nanotubes Mediate Bacterial Communication. Cell (2011). 5
9 Back to reality Dubey, G.P. & Ben-Yehuda, S. Intercellular Nanotubes Mediate Bacterial Communication. Cell (2011). 5
10 rescence microscopy 15 hr after plating, when small colonies were visible. The dashed line indicates the border between the two populations. (a) Phase contrast image (blue). (b) GFP fluorescence image (green). (c) Overlay of phase and GFP fluorescence images. The scale bar represents 10 mm. (B) Average fluorescence intensity of the gfp! population (as indicated in Aa) as a function of the distance from the gfp+ population. The gfp! region was divided into identical sub-regions and the average fluorescence signal was defined in arbitrary units (AU). (C) Exponentially growing PY79 (gfp!) and SB444 (gfp+) cells were mixed, plated on an LB agarose pad, and incubated in a temperature controlled chamber at 37" C. Cells were visualized by time-lapse fluorescence microscopy and phase contrast (blue) and fluorescence (green) images collected at 10 min intervals. Select overlay images are shown from the following time points: (a) t0 min, (b) t30 min, and (c) t60 min. Each pair of colored arrows (red and yellow) indicates different locations where transfer of fluorescent molecules between neighboring cells is increasing over time. Larger fields of the same Redregion arrows: are shown in Figure S1. The scale bar represents gfp-1gaining mm. fluorescence (D) Average fluorescence intensity of the gfp! cells as a function of their distance from the gfp+ cells at t0 min (light blue bars) and at t60 min (dark blue bars) of the coincubation experiment as described in (C) (see Extended R was Experimental Procedures). NoR detectible signal Dubey, G.P. & Ben-Yehuda, S. Intercellular Nanotubes Mediate Bacterial Communication. Cell (2011). measured when cells were located beyond 1mm at t0 min. Back to reality Cm + Kan 5
11 Appearance of GFP in originally gfp cells t = 0 min t = 45 min Confirmation of GFP transfer (from B. subtilis 3610 gfp + to 3610 gfp strains) 6
12 Alternative explanation for antibiotic resistance transfer 1 Verification of published results CmR KanR CmR +KanR B. subtilis strains on LBA + Cm & Kan plate 2 Additionnal control experiment: Separating cells by filter Filter KanR expressing GFP CmR Fluorescence image 7
13 BACTERIAL NANOTUBES Episode 2 8
14 Links between teams: collaboration map AMERICA 2011 JAPAN JAPAN USA ASIA Paris EUROPE 9
15 Links between teams: collaboration map AMERICA What do igemers 2011 share? JAPAN JAPAN USA ASIA Paris EUROPE 9
16 Assisted diffusion model through nanotubes Cell 1 Cell 2 Modeling results: Time < 1µs Volume transfer 0.1% 10
17 Assisted diffusion model through nanotubes Cell 1 Cell 2 Modeling results: Time < 1µs Volume transfer 0.1% 10
18 Passive diffusion model through nanotubes Molecule name 1st molecule transfer (s) T7 trna insulin GFP glucose 4.46E E E E E-4 11
19 Amplified & robust detection of nanotube Amplified & robust detection of nanotube communication Emitter 12
20 Amplified & robust detection of nanotube Amplified & robust detection of nanotube communication Receptor Amplifier Emitter 12
21 Amplified & robust detection of nanotube Amplified & robust detection of nanotube communication Receptor Amplifier Emitter Reporter 12
22 Amplified & robust detection of nanotube Amplified & robust detection of nanotube communication Receptor Amplifier Emitter Reporter 12
23 Amplified & robust detection of nanotube Amplified & robust detection of nanotube communication Receptor Amplifier Emitter Reporter 12
24 Amplified & robust detection of nanotube Amplified & robust detection of nanotube communication Receptor Amplifier Emitter Reporter Emitter transient signal 12
25 Amplified & robust detection of nanotube Amplified & robust detection of nanotube communication Receptor Amplifier Emitter Reporter Receiver response Emitter transient signal 12
26 Our designs Concentrator Positive feedback loop YFP concentrator T7 RNA polymerase diffusion trna Amber ComS diffusion system Bistable switches λ switch Sporulation induction by KinA 13
27 trna amber diffusion T7 pol. amber trna RFP T7 GFP Emitter 14
28 trna amber diffusion mrna T7 pol. amber trna RFP T7 GFP Emitter 14
29 trna amber diffusion mrna T7 pol. amber trna RFP T7 GFP Emitter Receptor 14
30 trna amber diffusion mrna T7 pol. amber trna RFP T7 GFP Emitter Receptor Amplifier 14
31 trna amber characterization in E.coli IPTG trna Amber GFP Amber GFP amber GFP amber + trna amber 15
32 T7 polymerase amber characterization in E.coli pt7 11 GFP Fluorescence/OD 8,25 5,5 2,75 0 pt7 pveg GFP T7 amber pt7 pveg phs GFP T7 amber trna Amber 16
33 YFP concentrator TetR-YFP TetO array Emitter Receptor + Concentrator 17
34 YFP concentrator TetR-YFP TetO array Emitter Receptor + Concentrator 17
35 YFP concentrator: characterization in E.coli TetR-YFP TetR-YFP/TetO Arrows:Bright YFP foci TetR-YFP/TetO Red:ibpA-mCherry Green: tetr-yfp spot 18
36 Testing nanotubes formation: YFP concentrator Emitter + Receiver Mix E.coli TetR-YFP/B.Subtilis TetO Mix B.Subtilis TetR-YFP/B.Subtilis TetO No TetO foci observed after dozens of repeats 19
37 T7 autoloop design T7 pol pt7 T7 pol GFP Expression of GFP as monitor Autoloop 20
38 T7 autoloop design T7 pol pt7 T7 pol GFP Expression of GFP as monitor Autoloop External input 20
39 T7 autoloop design T7 pol pt7 T7 pol GFP Expression of GFP as monitor Autoloop Autoloop response External input 20
40 T7 autoloop characterization in E.coli T7 autoloop in T7 + E.coli cells T7 autoloop in T7 B.subtilis Fluorescence/OD Memory of autoloop-amplified signal over time in E.coli Time(min) 21
41 T7 RNA polymerase diffusion design B. subtilis IPTG Induction T7 pol pt7 T7 pol GFP Expression of GFP as monitor phs T7 pol RFP Autoloop B. subtilis 22
42 T7 diffusion design modeling Delayed differential equations Genetic network model Number of molecules GFP T7 autoloop mrna T7 pol from emitter 1h Time vfvr 2h 23
43 T7 RNA polymerase diffusion experiments Emitter/Receiver mix Plasmidic TRANS RFP GFP 24
44 T7 RNA polymerase diffusion experiments Emitter/Receiver mix Chromosomal Plasmidic TRANS RFP GFP 24
45 Microfluidic chambers for nanotube formation Why use microfluidic device? Monolayer Exponential phase Long-time observation Mix of B.subtilis Mondragón-Palomino, O., Danino, T., Selimkhanov, J., Tsimring, L. & Hasty, J. Entrainment of a population of synthetic genetic oscillators. Science (2011). 25
46 Achievements Successfully reproduced and improved the original experiments, proposed an alternative hypothesis. Designed, modeled and characterized 6 emitter/receivers in E.coli and B.subtilis. Developed 2 original computational diffusion models accounting for transport through nanotubes. Provided proofs of principle of 5 working emitter/receiver devices. Created 49 new BioBricks and characterized 25 BioBricks for B. subtilis. Collaboration: -Grenoble igem team for the human practice -Fatih Turkey team for the rewriting of the B.Subtilis page of the Parts Registry -Dundee, Edinbourgh, Freibourg, Pekin 26
47 27
48 Conclusion 27
49 Conclusion Smaller molecules Microfluidic conditions Statistical methods EM microscopy 27
50 The question remains! TuBe or not TuBe? 27
51 The Team Hovannes Agopyan Adrien Basso-Blandin Ouriel Caën Baptiste Couly Laura Da Silva Mathias Toulouze Kévin Yauy Camille Huet de Froberville Edward Kwarteng Danyel Lee Adrien Lhomme-Duchadeuil Oleg Mikhajlov Babak Nichabouri Cyrille Pauthenier Axel Séguret Hosting laboratory The mentors Ariel Lindner, Yifan Yang, Aleksandra Nivina, Antoine Decrulle,Raphaël Pantier, Thomas Lombès 28
52 Acknowledgments A special thanks to the Grenoble s igem team for our great collaboration! Help from labs all around the world M. Elowitz, Caltech S. Serror, Orsay University L. A. Sonenshein, Tufts University D. Lane, Toulouse II University J. V. Veening, Gröningen Usiversity P. Bassereau, Institut Curie H. Putzer and C. Condon, from IBPC Y. Chai, Harvard University P.Dubey and S.Ben-Yehuda, Hebrew University of Jerusalem 29
53 Penetrance of human practice questioning within igem % of teams with HP projects Total number of igem teams Number of igem teams having a human practice project 30
54 31
55 FACS experiments 32
56 T7 RNA polymerase diffusion design B. Subtilis IPTG Induction T7 pol pt7 T7 pol GFP Expression of GFP as monitor phs T7 pol RFP Autoloop B. Subtilis 33
57 trna amber design modeling Random walker model Functional polymerases produced Time (s) Genetic network model Number of molecules T7 RNAp translated amber mrna trna amber Time (hours) 19
58 Check up: where are we now? Working & Available biobricks for B. subtilis Our project: 15 working and characterized parts submitted Before 2011 With our contribution 35
59 Sporulation induction by KinA Phosphorelay phs KinA KinA peps SpoA GFP 36
60 Hypothesis for nanotube formation Local instability Bulge formation Nanotube formation 37
61 Testing nanotubes formation: YFP concentrator Control TetR-YFP E.coli Mix E.coli TetR-YFP/B.Subtilis TetO E.coli/B.subtilis Emitter only Control TetR-YFP B.Subtilis Emitter + Receiver Mix B.Subtilis TetR-YFP/B.Subtilis TetO B.subtilis/B.subtilis 13
Modeling and Simulation of Gene Regulatory Networks
 Modeling and Simulation of Gene Regulatory Networks Hidde de Jong INRIA Grenoble - Rhône-Alpes Hidde.de-Jong@inria.fr http://ibis.inrialpes.fr INRIA Grenoble - Rhône-Alpes and IBIS IBIS: systems biology
Modeling and Simulation of Gene Regulatory Networks Hidde de Jong INRIA Grenoble - Rhône-Alpes Hidde.de-Jong@inria.fr http://ibis.inrialpes.fr INRIA Grenoble - Rhône-Alpes and IBIS IBIS: systems biology
Bacterial Transformation with Green Fluorescent Protein. Table of Contents Fall 2012
 Bacterial Transformation with Green Fluorescent Protein pglo Version Table of Contents Bacterial Transformation Introduction..1 Laboratory Exercise...3 Important Laboratory Practices 3 Protocol...... 4
Bacterial Transformation with Green Fluorescent Protein pglo Version Table of Contents Bacterial Transformation Introduction..1 Laboratory Exercise...3 Important Laboratory Practices 3 Protocol...... 4
Chapter 6: Biological Networks
 Chapter 6: Biological Networks 6.4 Engineering Synthetic Networks Prof. Yechiam Yemini (YY) Computer Science Department Columbia University Overview Constructing regulatory gates A genetic toggle switch;
Chapter 6: Biological Networks 6.4 Engineering Synthetic Networks Prof. Yechiam Yemini (YY) Computer Science Department Columbia University Overview Constructing regulatory gates A genetic toggle switch;
GENETIC TRANSFORMATION OF BACTERIA WITH THE GENE FOR GREEN FLUORESCENT PROTEIN (GFP)
 GENETIC TRANSFORMATION OF BACTERIA WITH THE GENE FOR GREEN FLUORESCENT PROTEIN (GFP) LAB BAC3 Adapted from "Biotechnology Explorer pglo Bacterial Transformation Kit Instruction Manual". (Catalog No. 166-0003-EDU)
GENETIC TRANSFORMATION OF BACTERIA WITH THE GENE FOR GREEN FLUORESCENT PROTEIN (GFP) LAB BAC3 Adapted from "Biotechnology Explorer pglo Bacterial Transformation Kit Instruction Manual". (Catalog No. 166-0003-EDU)
Performance Metrics for a Biomolecular Step Response
 Submitted, 212 Conference on Decision and Control (CDC) http://www.cds.caltech.edu/~murray/papers/sm12-cdc.html Performance Metrics for a Biomolecular Step Response Shaunak Sen and Richard M. Murray Abstract
Submitted, 212 Conference on Decision and Control (CDC) http://www.cds.caltech.edu/~murray/papers/sm12-cdc.html Performance Metrics for a Biomolecular Step Response Shaunak Sen and Richard M. Murray Abstract
Determine Osmolarity Activity Using RFP under PompC
 Determine Osmolarity Activity Using RFP under PompC Materials and Equipment: E. coli : o containing RFP under Plac o (blank) Isogenic strains containing mrfp under PompC: o 21153 o 3367-3 ΔenvZ Isogenic
Determine Osmolarity Activity Using RFP under PompC Materials and Equipment: E. coli : o containing RFP under Plac o (blank) Isogenic strains containing mrfp under PompC: o 21153 o 3367-3 ΔenvZ Isogenic
Green Fluorescent Protein (GFP): Genetic Transformation, Synthesis and Purification of the Recombinant Protein
 Green Fluorescent Protein (GFP): Genetic Transformation, Synthesis and Purification of the Recombinant Protein INTRODUCTION Green Fluorescent Protein (GFP) is a novel protein produced by the bioluminescent
Green Fluorescent Protein (GFP): Genetic Transformation, Synthesis and Purification of the Recombinant Protein INTRODUCTION Green Fluorescent Protein (GFP) is a novel protein produced by the bioluminescent
A and B are not absolutely linked. They could be far enough apart on the chromosome that they assort independently.
 Name Section 7.014 Problem Set 5 Please print out this problem set and record your answers on the printed copy. Answers to this problem set are to be turned in to the box outside 68-120 by 5:00pm on Friday
Name Section 7.014 Problem Set 5 Please print out this problem set and record your answers on the printed copy. Answers to this problem set are to be turned in to the box outside 68-120 by 5:00pm on Friday
Transformation of the bacterium E. coli. using a gene for Green Fluorescent Protein
 Transformation of the bacterium E. coli using a gene for Green Fluorescent Protein Background In molecular biology, transformation refers to a form of genetic exchange in which the genetic material carried
Transformation of the bacterium E. coli using a gene for Green Fluorescent Protein Background In molecular biology, transformation refers to a form of genetic exchange in which the genetic material carried
Molecular Biology Techniques: A Classroom Laboratory Manual THIRD EDITION
 Molecular Biology Techniques: A Classroom Laboratory Manual THIRD EDITION Susan Carson Heather B. Miller D.Scott Witherow ELSEVIER AMSTERDAM BOSTON HEIDELBERG LONDON NEW YORK OXFORD PARIS SAN DIEGO SAN
Molecular Biology Techniques: A Classroom Laboratory Manual THIRD EDITION Susan Carson Heather B. Miller D.Scott Witherow ELSEVIER AMSTERDAM BOSTON HEIDELBERG LONDON NEW YORK OXFORD PARIS SAN DIEGO SAN
Induction of Enzyme Activity in Bacteria:The Lac Operon. Preparation for Laboratory: Web Tutorial - Lac Operon - submit questions
 Induction of Enzyme Activity in Bacteria:The Lac Operon Preparation for Laboratory: Web Tutorial - Lac Operon - submit questions I. Background: For the last week you explored the functioning of the enzyme
Induction of Enzyme Activity in Bacteria:The Lac Operon Preparation for Laboratory: Web Tutorial - Lac Operon - submit questions I. Background: For the last week you explored the functioning of the enzyme
Appendix 2 Molecular Biology Core Curriculum. Websites and Other Resources
 Appendix 2 Molecular Biology Core Curriculum Websites and Other Resources Chapter 1 - The Molecular Basis of Cancer 1. Inside Cancer http://www.insidecancer.org/ From the Dolan DNA Learning Center Cold
Appendix 2 Molecular Biology Core Curriculum Websites and Other Resources Chapter 1 - The Molecular Basis of Cancer 1. Inside Cancer http://www.insidecancer.org/ From the Dolan DNA Learning Center Cold
Lab 10: Bacterial Transformation, part 2, DNA plasmid preps, Determining DNA Concentration and Purity
 Lab 10: Bacterial Transformation, part 2, DNA plasmid preps, Determining DNA Concentration and Purity Today you analyze the results of your bacterial transformation from last week and determine the efficiency
Lab 10: Bacterial Transformation, part 2, DNA plasmid preps, Determining DNA Concentration and Purity Today you analyze the results of your bacterial transformation from last week and determine the efficiency
DNA CAN BE TRANSFERRED BETWEEN BACTERIA GENETIC ENGINEERING USING RECOMBINANT DNA TECHNOLOGY
 Bacterial Transformation DNA CAN BE TRANSFERRED BETWEEN BACTERIA Background Information Plasmid Transformed Cell Figure 1: Bacterial Transformation Quick Reference Abbreviations GFP pgfp gfp Green fl uorescent
Bacterial Transformation DNA CAN BE TRANSFERRED BETWEEN BACTERIA Background Information Plasmid Transformed Cell Figure 1: Bacterial Transformation Quick Reference Abbreviations GFP pgfp gfp Green fl uorescent
HCS604.03 Exercise 1 Dr. Jones Spring 2005. Recombinant DNA (Molecular Cloning) exercise:
 HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
A Mathematical Model of a Synthetically Constructed Genetic Toggle Switch
 BENG 221 Mathematical Methods in Bioengineering Project Report A Mathematical Model of a Synthetically Constructed Genetic Toggle Switch Nick Csicsery & Ricky O Laughlin October 15, 2013 1 TABLE OF CONTENTS
BENG 221 Mathematical Methods in Bioengineering Project Report A Mathematical Model of a Synthetically Constructed Genetic Toggle Switch Nick Csicsery & Ricky O Laughlin October 15, 2013 1 TABLE OF CONTENTS
Recombinant DNA & Genetic Engineering. Tools for Genetic Manipulation
 Recombinant DNA & Genetic Engineering g Genetic Manipulation: Tools Kathleen Hill Associate Professor Department of Biology The University of Western Ontario Tools for Genetic Manipulation DNA, RNA, cdna
Recombinant DNA & Genetic Engineering g Genetic Manipulation: Tools Kathleen Hill Associate Professor Department of Biology The University of Western Ontario Tools for Genetic Manipulation DNA, RNA, cdna
Tools and reference standards supporting the engineering and evolution of synthetic biological systems
 Tools and reference standards supporting the engineering and evolution of synthetic biological systems By Jason R. Kelly Bachelor of Science, Biology and Chemical Engineering Massachusetts Institute of
Tools and reference standards supporting the engineering and evolution of synthetic biological systems By Jason R. Kelly Bachelor of Science, Biology and Chemical Engineering Massachusetts Institute of
DNA Fingerprinting. Unless they are identical twins, individuals have unique DNA
 DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
Bacterial Transformation and Plasmid Purification. Chapter 5: Background
 Bacterial Transformation and Plasmid Purification Chapter 5: Background History of Transformation and Plasmids Bacterial methods of DNA transfer Transformation: when bacteria take up DNA from their environment
Bacterial Transformation and Plasmid Purification Chapter 5: Background History of Transformation and Plasmids Bacterial methods of DNA transfer Transformation: when bacteria take up DNA from their environment
Transfection-Transfer of non-viral genetic material into eukaryotic cells. Infection/ Transduction- Transfer of viral genetic material into cells.
 Transfection Key words: Transient transfection, Stable transfection, transfection methods, vector, plasmid, origin of replication, reporter gene/ protein, cloning site, promoter and enhancer, signal peptide,
Transfection Key words: Transient transfection, Stable transfection, transfection methods, vector, plasmid, origin of replication, reporter gene/ protein, cloning site, promoter and enhancer, signal peptide,
Agrobacterium tumefaciens-mediated transformation of Colletotrichum graminicola and Colletotrichum sublineolum
 Agrobacterium tumefaciens-mediated transformation of Colletotrichum graminicola and Colletotrichum sublineolum Flowers and Vaillancourt, 2005. Current Genetics 48: 380-388 NOTE added by L. Vaillancourt:
Agrobacterium tumefaciens-mediated transformation of Colletotrichum graminicola and Colletotrichum sublineolum Flowers and Vaillancourt, 2005. Current Genetics 48: 380-388 NOTE added by L. Vaillancourt:
Molecular Genetics: Challenges for Statistical Practice. J.K. Lindsey
 Molecular Genetics: Challenges for Statistical Practice J.K. Lindsey 1. What is a Microarray? 2. Design Questions 3. Modelling Questions 4. Longitudinal Data 5. Conclusions 1. What is a microarray? A microarray
Molecular Genetics: Challenges for Statistical Practice J.K. Lindsey 1. What is a Microarray? 2. Design Questions 3. Modelling Questions 4. Longitudinal Data 5. Conclusions 1. What is a microarray? A microarray
Process of Science: Using Diffusion and Osmosis
 Process of Science: Using Diffusion and Osmosis OBJECTIVES: 1. To understand one way to approach the process of science through an investigation of diffusion and osmosis. 2. To explore how different molecules
Process of Science: Using Diffusion and Osmosis OBJECTIVES: 1. To understand one way to approach the process of science through an investigation of diffusion and osmosis. 2. To explore how different molecules
Recombinant DNA and Biotechnology
 Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Analysis of gene expression data. Ulf Leser and Philippe Thomas
 Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
PRESTWICK ACADEMY NATIONAL 5 BIOLOGY CELL BIOLOGY SUMMARY
 Name PRESTWICK ACADEMY NATIONAL 5 BIOLOGY CELL BIOLOGY SUMMARY Cell Structure Identify animal, plant, fungal and bacterial cell ultrastructure and know the structures functions. Plant cell Animal cell
Name PRESTWICK ACADEMY NATIONAL 5 BIOLOGY CELL BIOLOGY SUMMARY Cell Structure Identify animal, plant, fungal and bacterial cell ultrastructure and know the structures functions. Plant cell Animal cell
SUPPLEMENTARY DATA 1
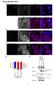 SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
Stochastic Gene Expression in Prokaryotes: A Point Process Approach
 Stochastic Gene Expression in Prokaryotes: A Point Process Approach Emanuele LEONCINI INRIA Rocquencourt - INRA Jouy-en-Josas ASMDA Mataró June 28 th 2013 Emanuele LEONCINI (INRIA) Stochastic Gene Expression
Stochastic Gene Expression in Prokaryotes: A Point Process Approach Emanuele LEONCINI INRIA Rocquencourt - INRA Jouy-en-Josas ASMDA Mataró June 28 th 2013 Emanuele LEONCINI (INRIA) Stochastic Gene Expression
How To Understand How Gene Expression Is Regulated
 What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
Thymine = orange Adenine = dark green Guanine = purple Cytosine = yellow Uracil = brown
 1 DNA Coloring - Transcription & Translation Transcription RNA, Ribonucleic Acid is very similar to DNA. RNA normally exists as a single strand (and not the double stranded double helix of DNA). It contains
1 DNA Coloring - Transcription & Translation Transcription RNA, Ribonucleic Acid is very similar to DNA. RNA normally exists as a single strand (and not the double stranded double helix of DNA). It contains
Gene Switches Teacher Information
 STO-143 Gene Switches Teacher Information Summary Kit contains How do bacteria turn on and turn off genes? Students model the action of the lac operon that regulates the expression of genes essential for
STO-143 Gene Switches Teacher Information Summary Kit contains How do bacteria turn on and turn off genes? Students model the action of the lac operon that regulates the expression of genes essential for
Physics 42 Lab 4 Fall 2012 Cathode Ray Tube (CRT)
 Physics 42 Lab 4 Fall 202 Cathode Ray Tube (CRT) PRE-LAB Read the background information in the lab below and then derive this formula for the deflection. D = LPV defl 2 SV accel () Redraw the diagram
Physics 42 Lab 4 Fall 202 Cathode Ray Tube (CRT) PRE-LAB Read the background information in the lab below and then derive this formula for the deflection. D = LPV defl 2 SV accel () Redraw the diagram
LAB 16 Rapid Colony Transformation of E. coli with Plasmid DNA
 LAB 16 Rapid Colony Transformation of E. coli with Plasmid DNA Objective: In this laboratory investigation, plasmids containing fragments of foreign DNA will be used to transform Escherichia coli cells,
LAB 16 Rapid Colony Transformation of E. coli with Plasmid DNA Objective: In this laboratory investigation, plasmids containing fragments of foreign DNA will be used to transform Escherichia coli cells,
AP Biology Essential Knowledge Student Diagnostic
 AP Biology Essential Knowledge Student Diagnostic Background The Essential Knowledge statements provided in the AP Biology Curriculum Framework are scientific claims describing phenomenon occurring in
AP Biology Essential Knowledge Student Diagnostic Background The Essential Knowledge statements provided in the AP Biology Curriculum Framework are scientific claims describing phenomenon occurring in
Cloning GFP into Mammalian cells
 Protocol for Cloning GFP into Mammalian cells Studiepraktik 2013 Molecular Biology and Molecular Medicine Aarhus University Produced by the instructors: Tobias Holm Bønnelykke, Rikke Mouridsen, Steffan
Protocol for Cloning GFP into Mammalian cells Studiepraktik 2013 Molecular Biology and Molecular Medicine Aarhus University Produced by the instructors: Tobias Holm Bønnelykke, Rikke Mouridsen, Steffan
FRET Basics and Applications an EAMNET teaching module
 FRET Basics and Applications an EAMNET teaching module Timo Zimmermann + Stefan Terjung Advanced Light Microscopy Facility European Molecular Biology Laboratory, Heidelberg http://www.embl.de/almf/ http://www.embl.de/eamnet/
FRET Basics and Applications an EAMNET teaching module Timo Zimmermann + Stefan Terjung Advanced Light Microscopy Facility European Molecular Biology Laboratory, Heidelberg http://www.embl.de/almf/ http://www.embl.de/eamnet/
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College
 Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
Name: Date: Period: DNA Unit: DNA Webquest
 Name: Date: Period: DNA Unit: DNA Webquest Part 1 History, DNA Structure, DNA Replication DNA History http://www.dnaftb.org/dnaftb/1/concept/index.html Read the text and answer the following questions.
Name: Date: Period: DNA Unit: DNA Webquest Part 1 History, DNA Structure, DNA Replication DNA History http://www.dnaftb.org/dnaftb/1/concept/index.html Read the text and answer the following questions.
20.309: Biological Instrumentation and Measurement. Heejin Choi Rumi Chunara Yuri Matsumoto
 20.309: Biological Instrumentation and Measurement Instructors: Laboratory Instructor: Teaching Assistants: Scott Manalis and Peter So Steve Wasserman Jaewon Cha Heejin Choi Rumi Chunara Yuri Matsumoto
20.309: Biological Instrumentation and Measurement Instructors: Laboratory Instructor: Teaching Assistants: Scott Manalis and Peter So Steve Wasserman Jaewon Cha Heejin Choi Rumi Chunara Yuri Matsumoto
Transformation Kit BACTERIAL TRANSFORMATION: GREEN FLUORESCENT PROTEIN. Partnership for Biotechnology and Genomics Education
 Transformation Kit BACTERIAL TRANSFORMATION: GREEN FLUORESCENT PROTEIN Partnership for Biotechnology and Genomics Education Barbara Soots Linda Curro Education Coordinator University of California Davis
Transformation Kit BACTERIAL TRANSFORMATION: GREEN FLUORESCENT PROTEIN Partnership for Biotechnology and Genomics Education Barbara Soots Linda Curro Education Coordinator University of California Davis
Algorithms in Computational Biology (236522) spring 2007 Lecture #1
 Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
Scottish Qualifications Authority
 National Unit specification: general information Unit code: FH2G 12 Superclass: RH Publication date: March 2011 Source: Scottish Qualifications Authority Version: 01 Summary This Unit is a mandatory Unit
National Unit specification: general information Unit code: FH2G 12 Superclass: RH Publication date: March 2011 Source: Scottish Qualifications Authority Version: 01 Summary This Unit is a mandatory Unit
Radius 24-Well Cell Migration Assay (Laminin Coated)
 Product Manual Radius 24-Well Cell Migration Assay (Laminin Coated) Catalog Number CBA-125-LN 24 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Cell migration is a highly
Product Manual Radius 24-Well Cell Migration Assay (Laminin Coated) Catalog Number CBA-125-LN 24 assays FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Cell migration is a highly
The E. coli Insulin Factory
 The E. coli Insulin Factory BACKGROUND Bacteria have not only their normal DNA, they also have pieces of circular DNA called plasmids. Plasmids are a wonderfully ally for biologists who desire to get bacteria
The E. coli Insulin Factory BACKGROUND Bacteria have not only their normal DNA, they also have pieces of circular DNA called plasmids. Plasmids are a wonderfully ally for biologists who desire to get bacteria
EPIGENETICS DNA and Histone Model
 EPIGENETICS ABSTRACT A 3-D cut-and-paste model depicting how histone, acetyl and methyl molecules control access to DNA and affect gene expression. LOGISTICS TIME REQUIRED LEARNING OBJECTIVES DNA is coiled
EPIGENETICS ABSTRACT A 3-D cut-and-paste model depicting how histone, acetyl and methyl molecules control access to DNA and affect gene expression. LOGISTICS TIME REQUIRED LEARNING OBJECTIVES DNA is coiled
Stochastic Gene Expression in Prokaryotes: A Point Process Approach
 Stochastic Gene Expression in Prokaryotes: A Point Process Approach Emanuele LEONCINI INRIA Rocquencourt - INRA Jouy-en-Josas Mathematical Modeling in Cell Biology March 27 th 2013 Emanuele LEONCINI (INRIA)
Stochastic Gene Expression in Prokaryotes: A Point Process Approach Emanuele LEONCINI INRIA Rocquencourt - INRA Jouy-en-Josas Mathematical Modeling in Cell Biology March 27 th 2013 Emanuele LEONCINI (INRIA)
Mureva Assembly surface mounted wiring devices
 1.0 Waterproof Assembly surface Switches P124543 P124547 External dimensions : 72 x 72 x 47 mm.. Assembly surface rockers become illuminated using bulbs and indicators in option. They can carry a label
1.0 Waterproof Assembly surface Switches P124543 P124547 External dimensions : 72 x 72 x 47 mm.. Assembly surface rockers become illuminated using bulbs and indicators in option. They can carry a label
Mir-X mirna First-Strand Synthesis Kit User Manual
 User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
RAINBOW ELECTROPHORESIS 1 An Introduction to Gel Electrophoresis
 RAINBOW ELECTROPHORESIS 1 An Introduction to Gel Electrophoresis INTRODUCTION This laboratory will demonstrate the basics of electrophoresis and the theory behind the separation of molecules on an agarose
RAINBOW ELECTROPHORESIS 1 An Introduction to Gel Electrophoresis INTRODUCTION This laboratory will demonstrate the basics of electrophoresis and the theory behind the separation of molecules on an agarose
Introduction To Real Time Quantitative PCR (qpcr)
 Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Synthetic Biology: DNA Digital Storage, Computation and the Organic Computer
 Synthetic Biology: DNA Digital Storage, Computation and the Organic Computer Alex Widdel University of Minnesota, Morris 1 / 27 Outline Overview of Synthetic Biology 1 Overview of Synthetic Biology 2 3
Synthetic Biology: DNA Digital Storage, Computation and the Organic Computer Alex Widdel University of Minnesota, Morris 1 / 27 Outline Overview of Synthetic Biology 1 Overview of Synthetic Biology 2 3
Objectives: Vocabulary:
 Introduction to Agarose Gel Electrophoresis: A Precursor to Cornell Institute for Biology Teacher s lab Author: Jennifer Weiser and Laura Austen Date Created: 2010 Subject: Molecular Biology and Genetics
Introduction to Agarose Gel Electrophoresis: A Precursor to Cornell Institute for Biology Teacher s lab Author: Jennifer Weiser and Laura Austen Date Created: 2010 Subject: Molecular Biology and Genetics
sensors Miniature Sensors - S100 The universal miniature photoelectric sensor
 Miniature Sensors - S1 The universal miniature photoelectric sensor Two threaded front mounting holes Two slotted rear mounting holes Anti-tampering sensor (no adjustment) Standard optic functions and
Miniature Sensors - S1 The universal miniature photoelectric sensor Two threaded front mounting holes Two slotted rear mounting holes Anti-tampering sensor (no adjustment) Standard optic functions and
Supplementary Figure 1.
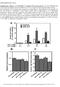 Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Biology Behind the Crime Scene Week 4: Lab #4 Genetics Exercise (Meiosis) and RFLP Analysis of DNA
 Page 1 of 5 Biology Behind the Crime Scene Week 4: Lab #4 Genetics Exercise (Meiosis) and RFLP Analysis of DNA Genetics Exercise: Understanding how meiosis affects genetic inheritance and DNA patterns
Page 1 of 5 Biology Behind the Crime Scene Week 4: Lab #4 Genetics Exercise (Meiosis) and RFLP Analysis of DNA Genetics Exercise: Understanding how meiosis affects genetic inheritance and DNA patterns
Lab Exercise 3: Media, incubation, and aseptic technique
 Lab Exercise 3: Media, incubation, and aseptic technique Objectives 1. Compare the different types of media. 2. Describe the different formats of media, plate, tube etc. 3. Explain how to sterilize it,
Lab Exercise 3: Media, incubation, and aseptic technique Objectives 1. Compare the different types of media. 2. Describe the different formats of media, plate, tube etc. 3. Explain how to sterilize it,
The Making of the Fittest: Evolving Switches, Evolving Bodies
 OVERVIEW MODELING THE REGULATORY SWITCHES OF THE PITX1 GENE IN STICKLEBACK FISH This hands-on activity supports the short film, The Making of the Fittest:, and aims to help students understand eukaryotic
OVERVIEW MODELING THE REGULATORY SWITCHES OF THE PITX1 GENE IN STICKLEBACK FISH This hands-on activity supports the short film, The Making of the Fittest:, and aims to help students understand eukaryotic
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation
 OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
Student Manual. pglo Transformation
 Student Manual pglo Transformation Lesson 1 Introduction to Transformation In this lab you will perform a procedure known as genetic transformation. Remember that a gene is a piece of DNA which provides
Student Manual pglo Transformation Lesson 1 Introduction to Transformation In this lab you will perform a procedure known as genetic transformation. Remember that a gene is a piece of DNA which provides
United States Department of Agriculture Agricultural Marketing Service, Science & Technology Microbiological Data Program
 SOP No.: MDP-QA-03 Page 1 of 6 1. Purpose To provide minimum requirements and standard procedures for quality assurance (QA) controls used in the USDA/AMS (MDP). 2. Scope This standard operating procedure
SOP No.: MDP-QA-03 Page 1 of 6 1. Purpose To provide minimum requirements and standard procedures for quality assurance (QA) controls used in the USDA/AMS (MDP). 2. Scope This standard operating procedure
WAMLocal. Wireless Asset Monitoring - Local Food Safety Software. Software Installation and User Guide BA/WAM-L-F
 Wireless Asset Monitoring - Local Food Safety Software BA/WAM-L-F Software Installation and User Guide System Overview The BAPI Wireless Asset Monitoring Local (WAM Local) Software receives temperature
Wireless Asset Monitoring - Local Food Safety Software BA/WAM-L-F Software Installation and User Guide System Overview The BAPI Wireless Asset Monitoring Local (WAM Local) Software receives temperature
Application Note. Single Cell PCR Preparation
 Application Note Single Cell PCR Preparation From Automated Screening to the Molecular Analysis of Single Cells The AmpliGrid system is a highly sensitive tool for the analysis of single cells. In combination
Application Note Single Cell PCR Preparation From Automated Screening to the Molecular Analysis of Single Cells The AmpliGrid system is a highly sensitive tool for the analysis of single cells. In combination
SmartFlare RNA Detection Probes: Principles, protocols and troubleshooting
 Technical Guide SmartFlare RNA Detection Probes: Principles, protocols and troubleshooting Principles of SmartFlare technology RNA detection traditionally requires transfection, laborious sample prep,
Technical Guide SmartFlare RNA Detection Probes: Principles, protocols and troubleshooting Principles of SmartFlare technology RNA detection traditionally requires transfection, laborious sample prep,
fxmiracle.com WINNING PIPS SYSTEM
 WINNING PIPS SYSTEM Thank you for taking the time to download this free guide. In your hands now is one of the best forex trading systems you might have ever come across. The key to winning with this profitable
WINNING PIPS SYSTEM Thank you for taking the time to download this free guide. In your hands now is one of the best forex trading systems you might have ever come across. The key to winning with this profitable
Outline. 1. Experiment. 2. Sample analysis and storage. 3. Image analysis and presenting data. 4. Probemaker
 Tips and tricks Note: this is just an informative document with general recommendations. Please contact support@olink.com should you have any queries. Document last reviewed 2011-11-17 Outline 1. Experiment
Tips and tricks Note: this is just an informative document with general recommendations. Please contact support@olink.com should you have any queries. Document last reviewed 2011-11-17 Outline 1. Experiment
Disc Diffusion Susceptibility Methods
 Disc Diffusion Susceptibility Methods Introduction When a filter paper disc impregnated with a chemical is placed on agar the chemical will diffuse from the disc into the agar. This diffusion will place
Disc Diffusion Susceptibility Methods Introduction When a filter paper disc impregnated with a chemical is placed on agar the chemical will diffuse from the disc into the agar. This diffusion will place
restriction enzymes 350 Home R. Ward: Spring 2001
 restriction enzymes 350 Home Restriction Enzymes (endonucleases): molecular scissors that cut DNA Properties of widely used Type II restriction enzymes: recognize a single sequence of bases in dsdna, usually
restriction enzymes 350 Home Restriction Enzymes (endonucleases): molecular scissors that cut DNA Properties of widely used Type II restriction enzymes: recognize a single sequence of bases in dsdna, usually
APPLICATION INFORMATION
 DRAFT: Rev. D A-2045A APPLICATION INFORMATION Flow Cytometry 3-COLOR COMPENSATION Raquel Cabana,* Mark Cheetham, Jay Enten, Yong Song, Michael Thomas,* and Brendan S. Yee Beckman Coulter, Inc., Miami FL
DRAFT: Rev. D A-2045A APPLICATION INFORMATION Flow Cytometry 3-COLOR COMPENSATION Raquel Cabana,* Mark Cheetham, Jay Enten, Yong Song, Michael Thomas,* and Brendan S. Yee Beckman Coulter, Inc., Miami FL
PALLETS ROLLER CONVEYOR LOADING CONVEYOR CHAIN TRANSFER TURNTABLE ROLLER STOP
 AUGUST 12, 2014 INDEX 04 07 04 06 EMITTER REMOVER 07 08 10 12 14 BOXES BELT CONVEYOR BELT CONVEYOR GATE STRAIGHT SPUR CONVEYOR CONVEYOR SCALE 16 17 18 19 20 22 24 26 ALIGNERS WHEEL ALIGNER BRACKET CHUTE
AUGUST 12, 2014 INDEX 04 07 04 06 EMITTER REMOVER 07 08 10 12 14 BOXES BELT CONVEYOR BELT CONVEYOR GATE STRAIGHT SPUR CONVEYOR CONVEYOR SCALE 16 17 18 19 20 22 24 26 ALIGNERS WHEEL ALIGNER BRACKET CHUTE
Transfection reagent for visualizing lipofection with DNA. For ordering information, MSDS, publications and application notes see www.biontex.
 METAFECTENE FluoR Transfection reagent for visualizing lipofection with DNA For ordering information, MSDS, publications and application notes see www.biontex.com Description Cat. No. Size METAFECTENE
METAFECTENE FluoR Transfection reagent for visualizing lipofection with DNA For ordering information, MSDS, publications and application notes see www.biontex.com Description Cat. No. Size METAFECTENE
CCR Biology - Chapter 9 Practice Test - Summer 2012
 Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Effects of Antibiotics on Bacterial Growth and Protein Synthesis: Student Laboratory Manual
 Effects of Antibiotics on Bacterial Growth and Protein Synthesis: Student Laboratory Manual I. Purpose...1 II. Introduction...1 III. Inhibition of Bacterial Growth Protocol...2 IV. Inhibition of in vitro
Effects of Antibiotics on Bacterial Growth and Protein Synthesis: Student Laboratory Manual I. Purpose...1 II. Introduction...1 III. Inhibition of Bacterial Growth Protocol...2 IV. Inhibition of in vitro
BINARY OPTIONS MENTOR USER MANUAL
 BINARY OPTIONS MENTOR USER MANUAL Version 1.4 Introduction : Welcome to Binary Options Mentor!! We provide you herewith a system of alerts with arrows for binary options. This system works for all timeframes
BINARY OPTIONS MENTOR USER MANUAL Version 1.4 Introduction : Welcome to Binary Options Mentor!! We provide you herewith a system of alerts with arrows for binary options. This system works for all timeframes
Wiring diagrams for luminous 1P 1-way switches with 110 V~ or 250 V~ signalling units
 Devices Switches - TECHICA CHARACTERISTICS Scope Switching on and off of ohm-inductive loads: lighting circuits: - lighting fittings (luminaires) for use incandescent lamps - lighting fittings (luminaires)
Devices Switches - TECHICA CHARACTERISTICS Scope Switching on and off of ohm-inductive loads: lighting circuits: - lighting fittings (luminaires) for use incandescent lamps - lighting fittings (luminaires)
DNA-functionalized hydrogels for confined membrane-free in vitro transcription/translation
 Electronic Supplementary Material (ESI) for Lab on a Chip. This journal is The Royal Society of Chemistry 2014 DNA-functionalized hydrogels for confined membrane-free in vitro transcription/translation
Electronic Supplementary Material (ESI) for Lab on a Chip. This journal is The Royal Society of Chemistry 2014 DNA-functionalized hydrogels for confined membrane-free in vitro transcription/translation
Procedures For DNA Sequencing
 Procedures For DNA Sequencing Plant-Microbe Genomics Facility (PMGF) Ohio State University 420 Biological Sciences Building 484 W. 12th Ave., Columbus OH 43210 Telephone: 614/247-6204 FAX: 614/292-6337
Procedures For DNA Sequencing Plant-Microbe Genomics Facility (PMGF) Ohio State University 420 Biological Sciences Building 484 W. 12th Ave., Columbus OH 43210 Telephone: 614/247-6204 FAX: 614/292-6337
Crime Scenes and Genes
 Glossary Agarose Biotechnology Cell Chromosome DNA (deoxyribonucleic acid) Electrophoresis Gene Micro-pipette Mutation Nucleotide Nucleus PCR (Polymerase chain reaction) Primer STR (short tandem repeats)
Glossary Agarose Biotechnology Cell Chromosome DNA (deoxyribonucleic acid) Electrophoresis Gene Micro-pipette Mutation Nucleotide Nucleus PCR (Polymerase chain reaction) Primer STR (short tandem repeats)
Enzymes: Amylase Activity in Starch-degrading Soil Isolates
 Enzymes: Amylase Activity in Starch-degrading Soil Isolates Introduction This week you will continue our theme of industrial microbiologist by characterizing the enzyme activity we selected for (starch
Enzymes: Amylase Activity in Starch-degrading Soil Isolates Introduction This week you will continue our theme of industrial microbiologist by characterizing the enzyme activity we selected for (starch
Related topics: Application Note 27 Data Analysis of Tube Formation Assays.
 Tube Formation Assays in µ-slide Angiogenesis Related topics: Application Note 27 Data Analysis of Tube Formation Assays. Contents 1. General Information... 1 2. Material... 2 3. Work Flow Overview...
Tube Formation Assays in µ-slide Angiogenesis Related topics: Application Note 27 Data Analysis of Tube Formation Assays. Contents 1. General Information... 1 2. Material... 2 3. Work Flow Overview...
Supplementary Information: Chemical communication between bacteria and cell- free gene expression systems within linear chains of emulsion droplets
 Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2016 Supplementary Information: Chemical communication between bacteria and cell- free gene
Electronic Supplementary Material (ESI) for Integrative Biology. This journal is The Royal Society of Chemistry 2016 Supplementary Information: Chemical communication between bacteria and cell- free gene
Product: Expression Arrest TM egfp control shrna vector
 Product: Expression Arrest TM egfp control vector Catalog #: RHS1702 Product Description The laboratory of Dr. Greg Hannon at Cold Spring Harbor Laboratory (CSHL) has created an RNAi Clone Library comprised
Product: Expression Arrest TM egfp control vector Catalog #: RHS1702 Product Description The laboratory of Dr. Greg Hannon at Cold Spring Harbor Laboratory (CSHL) has created an RNAi Clone Library comprised
Practice Questions 1: Scientific Method
 Practice Questions 1: Scientific Method 1. A student divided some insect larvae into four equal groups, each having the same amount of food. Each group was kept at a different temperature, and the average
Practice Questions 1: Scientific Method 1. A student divided some insect larvae into four equal groups, each having the same amount of food. Each group was kept at a different temperature, and the average
How To Run A Factory I/O On A Microsoft Gpu 2.5 (Sdk) On A Computer Or Microsoft Powerbook 2.3 (Powerpoint) On An Android Computer Or Macbook 2 (Powerstation) On
 User Guide November 19, 2014 Contents 3 Welcome 3 What Is FACTORY I/O 3 How Does It Work 4 I/O Drivers: Connecting To External Technologies 5 System Requirements 6 Run Mode And Edit Mode 7 Controls 8 Cameras
User Guide November 19, 2014 Contents 3 Welcome 3 What Is FACTORY I/O 3 How Does It Work 4 I/O Drivers: Connecting To External Technologies 5 System Requirements 6 Run Mode And Edit Mode 7 Controls 8 Cameras
Arndt & Voß GmbH Elektronik - Meßtechnik
 8 channel measuring unit Contents: Page 1. Displays and control elements 2-4 2. Technical Data 4 3. Power Supply 4 4. Profibus Interface X312 4 5. Connection schematics 4-6 Security comments according
8 channel measuring unit Contents: Page 1. Displays and control elements 2-4 2. Technical Data 4 3. Power Supply 4 4. Profibus Interface X312 4 5. Connection schematics 4-6 Security comments according
Modeling DNA Replication and Protein Synthesis
 Skills Practice Lab Modeling DNA Replication and Protein Synthesis OBJECTIVES Construct and analyze a model of DNA. Use a model to simulate the process of replication. Use a model to simulate the process
Skills Practice Lab Modeling DNA Replication and Protein Synthesis OBJECTIVES Construct and analyze a model of DNA. Use a model to simulate the process of replication. Use a model to simulate the process
Name Date Period. 2. When a molecule of double-stranded DNA undergoes replication, it results in
 DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
Cable Discharge Event
 Cable Discharge Event 1.0 Introduction The widespread use of electronic equipment in various environments exposes semiconductor devices to potentially destructive Electro Static Discharge (ESD). Semiconductor
Cable Discharge Event 1.0 Introduction The widespread use of electronic equipment in various environments exposes semiconductor devices to potentially destructive Electro Static Discharge (ESD). Semiconductor
Biotechnology: DNA Technology & Genomics
 Chapter 20. Biotechnology: DNA Technology & Genomics 2003-2004 The BIG Questions How can we use our knowledge of DNA to: diagnose disease or defect? cure disease or defect? change/improve organisms? What
Chapter 20. Biotechnology: DNA Technology & Genomics 2003-2004 The BIG Questions How can we use our knowledge of DNA to: diagnose disease or defect? cure disease or defect? change/improve organisms? What
The Steps. 1. Transcription. 2. Transferal. 3. Translation
 Protein Synthesis Protein synthesis is simply the "making of proteins." Although the term itself is easy to understand, the multiple steps that a cell in a plant or animal must go through are not. In order
Protein Synthesis Protein synthesis is simply the "making of proteins." Although the term itself is easy to understand, the multiple steps that a cell in a plant or animal must go through are not. In order
Long-Term Effects of Drug Addiction
 Long-Term Effects of Drug Addiction Part 1: Addiction is a chronic disease Drug addiction is considered a chronic brain disease because drugs cause long-lasting changes in brain structure and function.
Long-Term Effects of Drug Addiction Part 1: Addiction is a chronic disease Drug addiction is considered a chronic brain disease because drugs cause long-lasting changes in brain structure and function.
Automating Cell Biology Annual general meeting, September 7, 2015. Phase Holographic Imaging www.phiab.se
 Automating Cell Biology Annual general meeting, September 7, 2015 www.phiab.se 1 PHASE HOLOGRAPHIC IMAGING (PHI) Began as a research project at Lund University, Sweden, in 2000 Founded in 2004 Sales in
Automating Cell Biology Annual general meeting, September 7, 2015 www.phiab.se 1 PHASE HOLOGRAPHIC IMAGING (PHI) Began as a research project at Lund University, Sweden, in 2000 Founded in 2004 Sales in
PREPARED FOR: U.S. Army Medical Research and Materiel CommandFort Detrick, Maryland 21702 5012
 AD Award Number: W81XWH 07 1 0542 TITLE: Is Nuclear Structure Altered in Breast Cancer Cells? PRINCIPAL INVESTIGATOR: Han Htun, Ph.D. CONTRACTING ORGANIZATION: University of California Los Angeles, CA
AD Award Number: W81XWH 07 1 0542 TITLE: Is Nuclear Structure Altered in Breast Cancer Cells? PRINCIPAL INVESTIGATOR: Han Htun, Ph.D. CONTRACTING ORGANIZATION: University of California Los Angeles, CA
Application of Automated Data Collection to Surface-Enhanced Raman Scattering (SERS)
 Application Note: 52020 Application of Automated Data Collection to Surface-Enhanced Raman Scattering (SERS) Timothy O. Deschaines, Ph.D., Thermo Fisher Scientific, Madison, WI, USA Key Words Array Automation
Application Note: 52020 Application of Automated Data Collection to Surface-Enhanced Raman Scattering (SERS) Timothy O. Deschaines, Ph.D., Thermo Fisher Scientific, Madison, WI, USA Key Words Array Automation
THE ENVIRONMENTAL TECHNOLOGY VERIFICATION PROGRAM ANALYSIS OF TOTAL COLIFORMS AND E. COLI IN DRINKING WATER
 THE ENVIRONMENTAL TECHNOLOGY VERIFICATION PROGRAM ETV Verification Statement TECHNOLOGY TYPE: APPLICATION: COLIFORM DETECTION ANALYSIS OF TOTAL COLIFORMS AND E. COLI IN DRINKING WATER TECHNOLOGY NAME:
THE ENVIRONMENTAL TECHNOLOGY VERIFICATION PROGRAM ETV Verification Statement TECHNOLOGY TYPE: APPLICATION: COLIFORM DETECTION ANALYSIS OF TOTAL COLIFORMS AND E. COLI IN DRINKING WATER TECHNOLOGY NAME:
Activity 4 Long-Term Effects of Drug Addiction
 Activity 4 Long-Term Effects of Drug Addiction Core Concept: Addictive drugs may lead to long-term changes in brain function. Class time required: Approximately 60-80 minutes Teacher Provides: Copy of
Activity 4 Long-Term Effects of Drug Addiction Core Concept: Addictive drugs may lead to long-term changes in brain function. Class time required: Approximately 60-80 minutes Teacher Provides: Copy of
Quick Start Contents
 Quick Start Guide 1 Quick Start Contents Contact Information.. 3 Create an Account 3 Order Saliva Collection Supplies...3 Collect a Specimen..4 Order a Test..5 Package and Ship Specimens...8 Schedule FedEx
Quick Start Guide 1 Quick Start Contents Contact Information.. 3 Create an Account 3 Order Saliva Collection Supplies...3 Collect a Specimen..4 Order a Test..5 Package and Ship Specimens...8 Schedule FedEx
Simultaneously Monitoring Gene Expression Kinetics and Genetic Noise in Single Cells by Optical Well Arrays
 Anal. Chem. 2004, 76, 6282-6286 Simultaneously Monitoring Gene Expression Kinetics and Genetic Noise in Single Cells by Optical Well Arrays Yina Kuang, Israel Biran, and David R. Walt* Department of Chemistry,
Anal. Chem. 2004, 76, 6282-6286 Simultaneously Monitoring Gene Expression Kinetics and Genetic Noise in Single Cells by Optical Well Arrays Yina Kuang, Israel Biran, and David R. Walt* Department of Chemistry,
Improved methods for site-directed mutagenesis using Gibson Assembly TM Master Mix
 CLONING & MAPPING DNA CLONING DNA AMPLIFICATION & PCR EPIGENETICS RNA ANALYSIS Improved methods for site-directed mutagenesis using Gibson Assembly TM Master Mix LIBRARY PREP FOR NET GEN SEQUENCING PROTEIN
CLONING & MAPPING DNA CLONING DNA AMPLIFICATION & PCR EPIGENETICS RNA ANALYSIS Improved methods for site-directed mutagenesis using Gibson Assembly TM Master Mix LIBRARY PREP FOR NET GEN SEQUENCING PROTEIN
MEF Starter Nucleofector Kit
 page 1 of 7 MEF Starter Nucleofector Kit for Mouse Embryonic Fibroblasts (MEF) MEF display significant phenotypic variations which depend on the strain, the genetic background of the mice they are isolated
page 1 of 7 MEF Starter Nucleofector Kit for Mouse Embryonic Fibroblasts (MEF) MEF display significant phenotypic variations which depend on the strain, the genetic background of the mice they are isolated
