Wild-type and mutant p53 differentially regulate transcription of
|
|
|
- Marilynn Nichols
- 8 years ago
- Views:
Transcription
1 Proc. Natl. Acad. Sci. USA Vol. 93, pp , August 1996 Biochemistry Wild-type and mutant p53 differentially regulate transcription of the insulin-like growth factor I receptor gene HAIM WERNER*t, EDDY KARNIELI*t, FRANK J. RAUSCHER III, AND DEREK LERoITH* *Section on Molecular and Cellular Physiology, Diabetes Branch, National Institute of Diabetes, Digestive and Kidney Diseases, National Institutes of Health, Bethesda, MD 20892; tinstitute of Endocrinology and Metabolism, Rambam Medical Center and B. Rappaport Faculty of Medicine, Technion-Israel Institute of Technology, Haifa 31096, Israel; and Wistar Institute of Anatomy and Biology, Philadelphia, PA Communicated by J. Edward Rall, National Institute of Diabetes, Digestive and Kidney Diseases, Bethesda, MD, May 13, 1996 (received for review March 26, 1996) ABSTRACT The insulin-like growth factor I receptor (IGF- I-R) plays a critical role in transformation events. It is highly overexpressed in most malignant tissues where it functions as an anti-apoptotic agent by enhancing cell survival. Tumor suppressor p53 is a nuclear transcription factor that blocks cell cycle progression and induces apoptosis. p53 is the most frequently mutated gene in human cancer. Cotransfection of Saos-2 (osteosarcoma-derived cells) and RD (rhabdomyosarcoma-derived cells) cells with IGF-I-R promoter constructs driving luciferase reporter genes and with wild-type p53 expression vectors suppressed promoter activity in a dose-dependent manner. This effect of p53 is mediated at the level of transcription and it involves interaction with TBP, the TATA box-binding component of TFIID. On the other hand, three tumor-derived mutant forms of p53 (mut 143, mut 248, and mut 273) stimulated the activity of the IGF-I-R promoter and increased the levels of IGF-I-R/ luciferase fusion mrna. These results suggest that wild-type p53 has the potential to suppress the IGF-I-R promoter in the postmitotic, fully differentiated cell, thus resulting in low levels of receptor gene expression in adult tissues. Mutant versions of p53 protein, usually associated with malignant states, can derepress the IGF-I-R promoter, with ensuing mitogenic activation by locally produced or circulating IGFs. The insulin-like growth factor I receptor (IGF-I-R) is a membrane-bound heterotetramer with ligand-induced tyrosine kinase activity (1, 2). IGF-I-R is constitutively expressed by most tissues, where it mediates the trophic and differentiative actions of the IGFs, IGF-I and IGF-II (3-5). The central role of the IGF-I-R during the cell cycle is demonstrated by the fact that overexpression of this receptor in BALB/c3T3 fibroblasts abrogates all requirements for exogenous growth factors (6). Furthermore, deletion of the IGF-I-R in mice by homologous recombination results in nonviable offspring (7, 8). In addition, there is compelling evidence that the IGF-I-R has a pivotal role in malignancy (9, 10). Thus, it is highly overexpressed in most tumors and cancer cell lines and, furthermore, fibroblast cell lines established from mouse embryos lacking the IGF-I-R cannot be transformed by any of a number of oncogenes, including the simian virus 40 large T antigen, activated ras, and others (11, 12). The regulatory region of the IGF-I-R gene comprises a unique "initiator" motif, from which transcription is initiated in vivo, and that acts in concert with upstream Spl sites (13, 14). The region flanking the transcription start site is extremely G+C rich, with no obvious TATA or CAAT elements (14-17). When measured in transient transfection assays, the IGF-I-R promoter displays very high basal activity (14). Paradoxically, the expression of the receptor gene in normal adult tissues is extremely low (18), suggesting that in the postmitotic, fully differentiated cell the IGF-I-R promoter is under constitutive inhibitory control. The publication costs of this article were defrayed in part by page charge payment. This article must therefore be hereby marked "advertisement" in accordance with 18 U.S.C solely to indicate this fact. Wild-type (wt) p53 is a tumor suppressor gene product that, in its hyperphosphorylated state, blocks progression of cells through the cell cycle (19). p53 is localized to the nucleus, where it functions as a DNA sequence-specific transcription factor (20, 21). It has been shown that p53 can either activate or suppress the activity of a number of target genes. Gene activation usually involves interaction of p53 with a specific consensus sequence, whereas it is thought that gene suppression involves interaction of p53 with the basal transcription machinery (22-24). Mutations in the p53 gene are the most frequent mutations in human cancer (25, 26). The vast majority of these mutations occur in the central domain of the p53 molecule, which is the region involved in DNA binding. Because the IGF-I-R gene is highly overexpressed in most malignancies in which p53 is mutated, we investigated the potential regulation of the IGF- I-R gene by wt and mutant p53. The results obtained indicate that mutant p53 proteins have a stimulatory effect on promoter activity, whereas wt p53 suppresses the activity of the IGF-I-R promoter. These effects of p53 seem to involve its interaction with components of the basal transcription machinery. Due to the central role of the IGF-I-R in cell cycle progression and transformation, de-repression of IGF-I-R promoter by mutant p53 may constitute an important paradigm in tumorigenesis. MATERIALS AND METHODS Cell Cultures, Plasmids, and DNA Transfections. Saos-2 cells were obtained from the American Type Culture Collection. Saos-2 is a human osteogenic sarcoma-derived cell line in which both p53 alleles are deleted (27). RD is a human rhabdomyosarcoma cell line that exhibits a mutant p53 gene (Arg -* Trp transition at codon 248) (28). RD cells were kindly provided by Lee Helman (National Cancer Institute, Bethesda, MD). Saos-2 cells were grown in Dulbecco's modified Eagle's medium (DMEM) supplemented with 10% fetal bovine serum (FBS) and 5% calf serum. RD cells were grown in DMEM plus 10% FBS. For transient cotransfection experiments, the following fragments of the IGF-I-R gene were subcloned upstream of a promoterless firefly luciferase reporter gene (poluc): -2350/+640, -476/+640, -455/+30, and -40/+640 (nucleotide 1 corresponds to the transcription initiation site). The basal promoter activity of these plasmids has been previously reported (14, 29). A wt p53 expression vector (pcb6.p53) was constructed by inserting an kb XbaI fragment of the human p53 into the cytomegalovirus (CMV)-containing plasmid pcb6+ (30). An additional wt p53 (pc53-sn3) and three mutant expression vectors were kindly provided by Edward Mercer (Thomas Abbreviations: IGF, insulin-like growth factor; IGF-I-R, insulin-like growth factor I receptor; wt, wild type; CMV, cytomegalovirus; X3-gal,,3-galactosidase; GST, glutathione S-transferase; TBP, TATA boxbinding protein; f.p.u., footprint unit. tpresent address: Department of Clinical Biochemistry, Sackler Faculty of Medicine, Tel Aviv University, Ramat Aviv 69978, Israel. To whom reprint requests should be addressed. 8318
2 Biochemistry: Werner et al. Jefferson University, Philadelphia). pc53-sn3 encodes wt p53 in the pcmv-neo-bam vector (31). pc53-scx3 encodes a mutant p53 harboring a Val Ala mutation at position 143. pc53-248w contains an Arg Trp mutation at position 248 and pc53-273h is a mutant p53 in which an Arg residue at position 273 was mutated to His. All three mutant p53 are in the pcmv-neo-bam expression vector. Both cell lines were transiently transfected by the calcium phosphate method. For Saos-2 cells, we used a kit from 5 Prime-3 Prime Inc.; each 100-mm dish received 10,tg of reporter plasmid and 2.5,ug of expression vector. RD cells were transfected following the protocol described by Chen and Okayama (32), using 5,ug each of reporter and expression plasmids and 15,ug of salmon sperm DNA. Cells were harvested 48 h (Saos-2) or 72 h (RD) after transfection, and luciferase activities were measured as described (14). In preliminary experiments, cells were cotransfected with a CMV-,-gal vector (13-gal, f3-galactosidase), but since expression from the CMV promoter was found to be affected by p53, subsequent experiments were normalized to total protein, which was measured using a Bio-Rad kit. In a number of pilot experiments, normalization for transfection efficiency was done using a RAS-,B-gal plasmid (33), kindly provided by Ronald Evans (The Salk Institute, San Diego). The levels of (3-gal generated by this plasmid were not affected by p53 and the results obtained were essentially the same as those obtained using protein normalization. Luciferase mrna Measurements. After (48 h) transient transfection, Saos-2 cells were lysed in 4 M guanidinium isothiocyanate containing 0.01% 2-mercaptoethanol, and total RNA was prepared according to Chirgwin et al. (34). The integrity of the RNA was assessed by ethidium bromide staining of the 28S and 18S ribosomal RNA bands after gel electrophoresis. The levels of luciferase mrna were determined by solution hybridization/ RNase protection assay using 25,ug of total RNA as previously described (18). An antisense luciferase RNA probe was generated by linearization of the pgem-luc DNA vector (Promega) with EcoRV and transcription with T7 RNA polymerase in the presence of [a-32p]utp. Hybridization of this 413-b probe with luciferase RNA results in a protected band of -365 bp. As an intemal control, an 18S ribosomal antisense RNA probe that was labeled using the MEGAshortscript T7 kit (Ambion) was included in the hybridization reaction. GST.p53 Preparation. Purified p53 protein was prepared as a glutathione S-transferase (GST) fusion protein. GST.p53 and GST plasmids (in pgex-2 vector) were transformed into Escherichia coli XA-90 strain, and recombinant proteins were induced by addition of 1 mm isopropyl f3-d-thiogalactoside (IPTG). Cells were harvested after 3 h, washed with phosphate-buffered saline, and resuspended in 10 ml of extraction buffer containing 50 mm Hepes (ph 7.4), 250 mm NaCl, 1 mm EDTA, 5 mm NaF, 1 mm dithiothreitol (DTT), 5 p,g/ml each of aprotinin, leupeptin, and pepstatin A, 100,tg/ml phenylmethylsulfonyl fluoride (PMSF), and 1% Nonidet P-40. After sonication and centrifugation, the supernatant was mixed with GST-agarose beads for 2 h at 4 C. Beads were spun down, washed with extraction buffer, and proteins eluted with 25 mm reduced glutathione in extraction buffer. Peak fractions were dialyzed against 20 mm Hepes (ph 7.4), 150 mm NaCl, 1 mm EDTA, 10% glycerol, and 0.1% Nonidet P-40. Gel Retardation Assays. A fragment of the IGF-I-R promoter (-40 to + 115) encompassing the in vivo transcription initiation site was isolated by digestion of a genomic DNA clone with PmlI andxhol. After purification from agarose gels, the fragment was dephosphorylated using calf intestinal phosphatase, and end-labeled with [y-32p]atp using T4 polynucleotide kinase. The labeled probe was separated from unincorporated nucleotides using Elu-Tip columns (Schleicher and Schuell). Recombinant TATA box-binding protein (TBP) was Proc. Natl. Acad. Sci. USA 93 (1996) 8319 purchased from Promega and used at a concentration of 1 footprint unit (f.p.u.) per reaction. Gel retardation assays were performed by preincubating TBP protein, GST.p53 protein (100 ng), or a combination of both proteins, in 9,ul of 20 mm Hepes (ph 7.5), 70 mm KCl, 12% glycerol, 0.05% Nonidet P-40, 100,uM ZnSO4, 0.05 M DTT, 1 mg/ml bovine serum albumin, and 0.1 mg/ml poly(di-dc), in the presence or absence of the indicated unlabeled DNA competitor, on ice. After 15 min, 75,000 dpm ( pg) of the labeled fragment was added, and the reaction was incubated for an additional 10 min. Changes in mobility were assessed by electrophoresis through a 5% polyacrylamide gel that was run at 250 V for 2 h at 4 C. After fixation in 10% acetic acid, the gels were autoradiographed at -70 C. In Vitro Transcription Assays. The DNA template used in in vitro transcription reactions includes 476 bp of 5'-flanking region and 640 bp of 5'-untranslated region. The fragment was isolated from vector DNA by HindlIl digestion and purified by agarose gel electrophoresis. In vitro transcriptions were performed as described (35). Briefly, 500 ng of the DNA template were incubated with HeLa whole-cell extract (12.5 to 44,tg protein) (36) in the presence of 420,tm each of ATP, GTP, and CTP, and 15,uCi of [a-32p]utp (400 Ci/mmol; 1 Ci = 37 GBq), in a final volume of 18,lI. The composition of the reaction buffer was as follows: 8.33 mm Hepes (ph 7.9), 42 mm KCl, 5.2 mm MgCl2, 42,tM EDTA, 7% glycerol, and 1 mm DTT. Where indicated, a-amanitin was added to a final concentration of 10,tg/ml. Purified GST.p53 fusion protein (or GST control) was added at a concentration of 25 to 150 ng per reaction. Transcription reactions were incubated at 30 C for 1 h, and were terminated by the addition of 20 mm Tris HCl (ph 7.8), 150 mm NaCl, and 0.2% SDS. Following extraction with phenol-chloroform and precipitation with ethanol, the transcription products were resolved on a 4% polyacrylamide gel containing 7 M urea, which was run at 350 V for 2 h. Wet gels were autoradiographed at -70 C. RESULTS Suppression of IGF-I-R Promoter by wt p53. Activation of the IGF-I-R constitutes a basic requirement for progression through the cell cycle. Because wt p53 specifically blocks this process, whereas mutant p53 proteins are unable to halt cell proliferation, we examined whether wt p53 can suppress the activity of the IGF-I-R promoter. For this purpose, coexpression studies were performed by using an IGF-I-R promoter fused to a luciferase reporter gene [p(-2350/+640)luc], together with wt p53 expression vectors. Transfections were performed in the human osteosarcoma cell line Saos-2 that, due to the lack of endogenous p53, provides a "clean" background for this type of study. As shown in Fig. L4, increasing amounts of pcb6.p53 suppressed promoter activity in a dosedependent manner. Maximal suppression was seen with 2.5,ug of expression vector, at which concentration promoter activity was 11.7 ± 1.6% (mean ± SEM of six experiments, each in duplicate) of control levels. Likewise, expression vector pc53- SN3 suppressed promoter activity, although to a lesser extent. Thus, at 2.5 jig of input DNA, the activity of the IGF-I-R promoter was 46.9 ± 5.7% (mean + SEM; n = 3 experiments, each in duplicate) of control values (Fig. 1A). Experiments were also performed in the rhabdomyosarcoma cell line, RD (Fig. IB). In this cell line, pcb6.p53 reduced promoter activity to 25.4 ± 3.1% of control levels (mean ± SEM, n = 6), whereas pc53-sn3 reduced it to 58.8 ± 5.2% (n = 2). Transcriptional Repression of IGF-I-R/Luciferase Fusion mrna by wt p53 in Vivo. To establish whether the specific repressive effect of wt p53 in vivo was indeed mediated at the level of transcription, RNA was prepared from Saos-2 cells that were transiently transfected with pcb6.p53, and the levels of luciferase mrna were measured by means of a sensitive
3 8320 Biochemistry: Werner et al. Proc. Natl. Acad. Sci. USA 93 (1996) A Saos-2 B RD C Control wt p53 ; Luciferase mrna --- a 4-4) wt p53 (ug) U 4 4) O- E L 0 Ezl_ Control pcb6.p53 pc53-sn3 18S rrna -_- E 25- ML-.1w *,#,..,. FIG. 1. Regulation of IGF-I-R promoter activity bywt p53. (A) The reporter plasmid p(-2350/+640)luc (10,ug) was cotransfected into Saos-2 cells with increasing amounts of the wt p53 expression vector pcb6.p53 (m) or with 2.5 jig of pc53-sn3 (0) using the calcium phosphate method. The values of luciferase activity shown are expressed as a percentage of the levels seen in the absence of p53. Experiments were performed between three and six times, each time in duplicate. Where not shown, the SEM bars are smaller than the symbol size. (B) p(-2350/+640)luc (5 jig) and pcb6.p53 or pc53-sn3 (5 pg) were cotransfected into RD cells, and luciferase activity was measured after 72 h. The results are expressed as percentage of the luciferase levels generated by cotransfecting the empty expression vectors (pcb6+ or pcmv-neo-bam, respectively). Experiments were done between two and six times, each in duplicate. (C) Saos-2 cells were transiently cotransfected with p(-2350/+640)luc and pcb6.p53 as indicated above, lysed in 4 M guanidinium thiocyanate after 48 h, and the levels of the IGF-I-R/luciferase fusion mrna were measured by solution hybridization/rnase protection assay. The autoradiogram was exposed for 17 days. solution hybridization/rnase protection assay. As shown in Fig. 1C, wt p53 reduced the levels of luciferase mrna by -70%. Because wt p53 did not affect the levels of the constitutively expressed 18S ribosomal RNA (Fig. 1C), we can infer that the effect of p53 on luciferase mrna is not the result of a generalized transcriptional "switch-off." Localization of the Promoter Region Responsible for Transcriptional Repression. A consensus DNA binding half site for wt p53 has been identified (Pu-Pu-Pu-C-A/T-T/A-G-Py-Py- Py) and shown to mediate most of the gene stimulatory effects of wt p53 (37). Sequencing analysis of the IGF-I-R promoter region (including 2.3 kb of 5'-flanking region and the complete 943-bp 5'-untranslated region) revealed the presence of multiple sites highly related to the consensus sequence. Fig. 2 Upper shows the location of sites that conform to the consensus sequence at least at 8 of 10 nucleotides in each half site. To examine whether the presence of those potential p53 binding sites correlated with the effect of p53, coexpression studies were performed using IGF-I-R promoter/reporter plasmids containing different portions of 5'-flanking and 5'- untranslated sequences, together with the pcb6.p53 expression vector (Fig. 2). Wt p53 suppressed promoter activity of all four constructs assayed (constructs 1-4), regardless of the various combinations of 5'-flanking and 5'-untranslated regions involved. The basal promoter activity of p(-40/+640)luc, which lacks most of the 5'-flanking region, was extremely low, though p53 was still able to reduce those levels. Wt p53 did not affect the luciferase levels generated by poluc (construct 5). The results obtained indicate that the suppressive effect of p53 on IGF-I-R promoter activity was independent of the potential binding sites present in this region. Because the only region in common to all of the constructs is the fragment surrounding the "initiator" element, we sought to characterize the interactions of p53 with this region. Interaction of p53 with Components of the Basal Transcription Machinery. TBP, in addition to being essential for transcription from TATA-containing promoters, is required for transcription initiation of promoters that, like the IGF-I-R gene, contain Spl binding sites and an "initiator" element but lack a TATA box (38). To study the interaction between TBP and p53 in regulation of IGF-I-R promoter, gel retardation assays were performed using a labeled DNA fragment (-40/+115) comprising the initiator region, together with purified TBP (1 f.p.u.) or GST.p53 (100 ng), or a combination of both proteins. Incubation of TBP with the labeled probe generated two retarded bands (Fig. 3A) whose appearance was prevented by addition of an -600-fold molar excess of the unlabeled probe (Fig. 3C). Furthermore, the formation of the DNA-TBP complexes was abolished by addition of GST.p53 to the binding reaction. In addition, incubation of this fragment with GST.p53 generated one retarded band, both in the absence and presence of TBP (Fig. 3A). To more precisely map the sites of interaction of TBP with the IGF-I-R promoter fragment, competition was performed using six 24-mer oligonucleotides covering the region between ONSTRUCTS C:) EL oj 0 0 or a: r1 rrna A AlUG EI---- it../t,...,.:. 3 F. 7, LU -640 r7 777r LU CONSTRUCTS -40,640 LUG FIG. 2. Suppression of IGF-I-R promoter activity by wt p53. (Upper) The 5'-flanking (open bar) and 5'-untranslated (hatched bar) regions of the IGF-I-R gene contain multiple sites that are highly related to the putative p53 consensus site. Each bar denotes a potential site composed of two half motifs (Pu-Pu-Pu-C-A/T-T/A-G-Py-Py-Py), each containing at least eight of ten nucleotides, separated by 0-13 bp. The arrow indicates the transcription start site. Saos-2 cells were cotransfected with 10,g of IGF-I-R reporter plasmids [or poluc (construct 5)] and 2.5 Ag of pcb6.p53 (or empty pcb6+, results designated as basal). LUC, luciferase cdna (not shown to scale). (Lower) Values are luciferase units normalized per total protein (x103).
4 Biochemistry: Wemer et al. A. LO C m 6) ( b0 co o I LO 0. B. -40 I Proc. Natl. Acad. Sci. USA 93 (1996) 8321 _ l U 5 -~l 6 DNA-TBP Complex L Qliaomorx LoQ:i2aon Free Probe - 06 #go --- DNA-p53 Complex 1-37 / / / / /+ 113 cotcaoccacc;armmzwc-wa GGOCOOCQCrGOCTGAO0GTTC TOrrTACCAOCATTAACTCOCTGA GAAAAAAOAGAOGAOGoOAACCG AOGAOGAOCGA&OCOCACCGOOCGA C. Protein Competitor I -B ~~ Ollgom.r 2 Free probe FIG. 3. Gel retardation analysis of the IGF-I-R transcription start site with TBP and GST.p53. (A) A DNA fragment extending from -40 to +115 was end-labeled with [y-32p]atp and used in binding reactions with TBP (1 fp.u.), GST.p53 (100 ng), or both proteins. (B) Location of oligomers used in competition assays. The underlined bases in oligomer 2 correspond to the "initiator" motif, and the asterisk denotes the first transcribed nucleotide, as determined by primer extension assay (15). (C) Competition experiments were performed by incubating the labeled -40/+115 fragment with TBP (1 f.p.u.) in the absence (lane 2) or presence of excess unlabeled probe (60 ng, lane 3) or oligomer 2 ( ng, lanes 4 and 5, respectively). nucleotides -40 and +115 (Fig. 3B). The only oligomer that was able to prevent the-formation of DNA-TBP complexes was oligomer 2, which encompasses the "initiator" element (Fig. 3C). None of the other five oligomers had a significant effect on TBP binding (data not shown), suggesting that TBP binds specifically to the transcription start site. In addition, none of the six oligomers was able to compete out the retarded band generated by p53 (data not shown), indicating that p53 binds nonspecifically to this DNA fragment. In Vtro Transcription Assays. To demonstrate that the action of wt p53 on IGF-I-R promoter was a direct effect at the transcriptional level, in vitro transcription reactions were performed using a DNA template (-476 to +640) comprising the proximal promoter region. This fragment has been previously shown to exhibit high levels of promoter activity in functional assays (29). Furthermore, this fragment contains a number of Spl sites that are required for transcription initiation (14). Incubation of HeLa cell extracts, which contain low endogenous levels of p53 (39), with the IGF-I-R promoter template, resulted in transcription initiation that was inhibited by a-amanitin (10 i,g/ml), a specific RNA polymerase II inhibitor. The levels of transcription observed decreased with increasing amounts of HeLa extract, consistent with transcriptional suppression of the IGF-I-R gene by endogenous p53 and/or other tumor suppressors (Fig. 4A). Addition of exogenous purified GST.p53 (25 to 150 ng) abolished transcription in a dose-dependent fashion (Fig. 4B). Intriguingly, under the in vitro conditions assayed, transcription started from a site located -100 bp downstream from the initiator element. The location of this alternative site was corroborated using a number of overlapping DNA templates in in vitro transcription assays (data not shown). Stimulation of IGF-I-R Gene Transcription by Mutant p53. To address the question whether overexpression of the IGF-I-R gene in human malignancies can result from lack of inhibition by mutant p53, Saos-2 cells were cotransfected with an IGF-I-R promoter/luciferase reporter plasmid together with expression vectors encoding mutant versions of p53. Whereas wt p53 in the same vector (pc53-sn3) suppressed promoter activity to 47% of control levels (Fig. 1A), pc53-scx3, pc53-248w, and pc53-273h mutants stimulated its activity to 227%, 319%, and 406% of control values, respectively (Fig. SA). Increased levels of luciferase activity were associated with increased concentrations of IGF-I-R/luciferase fusion mrna, (200% to 570% of control), as detected by solution hybridization/rnase protection assays (Fig. S B and C). These results thus indicate that mutant p53 proteins can induce transcription from the IGF-I-R gene in vivo.
5 8322 Biochemistry: Werner et al Proc. Natl. Acad. Sci. USA 93 (1996) A. B. Template DNA HeLa extract (ug) -amanitin IGF-I-R mrnal Protein (ng) IGF-I-RmRNA M t * * t GST GST.p A 4 4.*. -~~~~~~~404 M _ FIG. 4. In vitro transcription assays. (A) Dose-dependent suppression of IGF-I-R gene transcription by HeLa cell extracts. Increasing amounts of whole-cell HeLa extracts were incubated with a purified DNA template extending from -476 to +640, and with ATP, GTP, CTP, and [a-32p]utp for 1 h at 30 C, in the absence or presence of a-amanitin (10 jig/ml). The labeled transcription products were resolved on a 4% denaturing gel, which was autoradiographed for 8 days. (B) Suppression of IGF-I-R gene transcription by exogenous GST.p53. In vitro transcription assays were performed using 12.5,ug of HeLa extract and increasing amounts of purified GST or GST.p53 protein. M, pbr322/mspi molecular mass marker. DISCUSSION We have identified the IGF-I-R promoter as a molecular target for tumor suppressor p53. The IGF-I-R promoter is a highly regulated, TATA-less, "initiator"-containing promoter that directs transcription of very high levels of receptor mrna at embryonic and early postnatal stages (18). The abundance of this transcript significantly decreases at adult stages, whereas malignant states associated with augmented cellular proliferation are characterized by a rebound in gene expression (9, 10). The results of this study demonstrate that, in spite of the presence of potential p53 binding sites both upstream and downstream of the IGF-I-R gene transcription start site (a finding which is usually associated with genes stimulated by p53), wt p53 suppresses transcription from the IGF-I-R promoter both in vivo and in vitro. The difference in the extent of suppression between Saos-2 and RD cells may be due to the different p53 backgrounds in both cell lines. Thus, overexpression of wt p53 in Saos-2, a cell A.- U 041 E T I B Control m143 m248 m273 Luciterase mrna.. qinmu _M 40_A* 11P_ C. 1BS rrna _ line that lacks any endogenous p53, resulted in inhibition of promoter activity by 88%, whereas the effect of p53 transfection in RD cells, which express an endogenous p53 mutated at codon 248, was comparatively lower (75% inhibition). The mechanism of action of p53 on the IGF-I-R promoter seems to involve its interaction with TBP, the TATA-box binding subunit of the general initiation factor, TFIID. Results of gel retardation assays indicate that TBP binds specifically to the "initiator" element of the IGF-I-R promoter, whereas p53 displays a nonspecific interaction with this DNA fragment. However, p53 precludes binding of TBP to the promoter region, most probably through protein-protein interaction. As a result, TBP is no longer able to assemble a functional transcription initiation complex. Direct contact between wt p53 and TBP was previously demonstrated using affinity chromatography (40). The region of p53 involved in this interaction is the highly acidic N-terminal 73 amino acid fragment that functions as a transcriptional activation domain. Oncogenic versions of p53 mutated in their DNAbinding core domain are impaired in their ability to bind TBP, possibly due to overall conformational changes. Specific repression of transcription by wt p53 has been previously postulated to be limited to TATA-containing promoters (41). This proposal was based on the observation that a number of "initiator"-containing promoters, including the proliferating cell nuclear antigen (PCNA), c-ha-ras, and the epidermal growth factor receptor, are immune to the effects of p53 (42). In addition, a synthetic promoter containing an "initiator" element downstream of a simian virus bp activator element was unaffected by coexpression with wt p53, whereas an homologous promoter in which the "initiator" was replaced by the adenovirus major late promoter TATA box was significantly repressed (41). The results of the present study provide evidence for a novel class of "initiator" element-containing promoters that are susceptible to inhibitory regulation by p53, though it is still unknown what elements in the IGF-I-R promoter confer upon it sensitivity to p53. Interestingly, results of in vitro transcription assays point to an alternative initiation site, which differs by -100 bp from the site previously shown to function in vivo (14, 15). The reason for this discrepancy is presently unknown, though it may reflect the preference of members of the basal transcription complex to contact specific DNA sequences in the 5'-untranslated region under in vitro conditions. Thus, it is conceivable that the mechanism for p53 regulation of IGF-I-R gene transcription in vitro may differ from the in vivo mechanism, though with identical end results, i.e., suppression of IGF-I-R promoter. Alternatively, the C z Zr 600- I.,t En co 40C - < 300- z E 200- U - L-100- w cj Control m143 m248 m273 Control m143 m248 m273 FIG. 5. Stimulation of IGF-I-R gene transcription by mutant p53 proteins in vivo. (A) Activation of IGF-I-R promoter activity. Saos-2 cells were cotransfected with 10,tg of the reporter plasmid p(-2350/+640)luc and 2.5,ug of the expression vectors pc53-scx3, pc53-248w, and pc53-273h (or empty pcmv-neo-bam plasmid, as a control). The levels of luciferase activity are expressed as percentage of control values. Experiments were performed between three and four times, each in duplicate. (B) Transcriptional stimulation by mutant p53. RNA was prepared from Saos-2 cells that were transiently cotransfected with reporter and expression vectors, as indicated above, and the levels of IGF-I-R/luciferase fusion mrna were determined by solution hybridization/rnase protection assay. The autoradiogram was exposed for 28 days. (C) Quantitation of IGF-I-R/luciferase fusion mrna. The autoradiogram shown in B was scanned using National Institutes of Health IMAGE software (version 1.55). The levels of luciferase mrna, normalized to those of 18S rrna, are expressed as percentage of controls.
6 Biochemistry: Werner et al. discrepancy between in vivo and in vitro results may be due to technical reasons in in vitro transcription assays. The IGF-I-R has an important role as an inhibitor of apoptosis, both in vivo and in vitro (43). Activation of this receptor by IGFs protects cells from apoptotic death in a number of models. For example, IGF-I was shown to inhibit the etoposide-induced apoptosis in BALB/c3T3 fibroblasts overexpressing the IGF-I-R. This protective effect of IGF-I was less marked in parental BALB/c3T3 cells and it was totally absent in fibroblasts lacking the IGF-I-R (44). Similar results were seen in vivo using a biodiffusion chamber that allows passage of nutrients and proteins but excludes the inward or outward flow of intact cells. In this model, a decrease in the number of IGF-I-Rs was associated with massive cell death. Furthermore, overexpression of the receptor protects cells from apoptosis (45). Because wt p53 is a potent inducer of apoptosis, we may speculate that the effect of p53 on apoptosis is mediated, at least partially, through suppression of the IGF-I-R promoter. Lack of inhibition of the IGF-I-R gene by mutant p53 in malignant states may help expand a cell population that is otherwise destined to die. Furthermore, additional components of the IGF-signaling system have been shown to be modulated by p53. Thus, the expression of IGF-II P3 transcripts is reduced by wt p53 (46). On the other hand, the IGF-binding protein 3 (IGF-BP3) has been shown to be induced by wt, but not mutant, p53 (47). Because IGF-BP3 is an inhibitor of mitogenic signaling by IGFs, it follows that p53 can regulate the IGF system both at the level of availability of IGF ligands and at the level of activity of the IGF receptor. However, suppression of the IGF-I-R promoter is not limited to p53. We have recently demonstrated that WT1, a tumor suppressor involved in the etiology of Wilms tumor, binds both upstream and downstream of the IGF-I-R gene transcription start site by means of its zinc finger domain and suppresses promoter activity in functional assays (29, 48). In addition, overexpression of WT1 in G401 cells was associated with a decrease in the endogenous levels of IGF-I-R and with a reduction in IGF-I-mediated cellular proliferation and anchorage-independent growth (49). In conclusion, we have presented evidence for the suppression of IGF-I-R gene transcription by wt p53, and for its stimulation by mutant p53. When combined with our previous results on tumor suppressor WT1, a novel paradigm for tumorigenesis emerges. According to this model, the expression of the IGF-I-R gene is constitutively inhibited in the terminally differentiated cell by local (i.e., WT1) as well as by more ubiquitous (i.e., p53) tumor suppressors. As a result of this negative control, cells remain at a postmitotic state and out of the cell cycle. Mutation of tumor suppressors or activation of oncogenes, two events usually associated with malignancy, can de-repress the IGF-I-R gene promoter with increased production of cell-surface receptors and augmented mitogenic activation by locally produced or circulating IGFs. We thank Drs. Vicky Blakesley and Lee Helman for critical reading of the manuscript, Dr. Fatah Kashanchi for help with in vitro transcription assays, Drs. Edward Mercer and Ronald Evans for the p53 expression vectors and the Ras-,B-gal plasmid, respectively, and Dr. Lee Helman for RD cells. We are grateful to Violet Katz for expert secretarial assistance. 1. LeRoith, D., Werner, H., Beitner-Johnson, D. & Roberts, C. T., Jr. (1995) Endocr. Rev. 16, Werner, H., Woloschak, M., Stannard, B., Shen-Orr, Z., Roberts, C. T., Jr., & LeRoith, D. (1991) Insulin-Like Growth Factors: Molecular and Cellular Aspects (CRC, Boca Raton, FL), pp Jones, J. I. & Clemmons, D. R. (1995) Endocr. Rev. 16, Daughaday, W. H. & Rotwein, P. (1989) Endocr. Rev. 10, Werner, H., Adamo, M., Roberts, C. T., Jr., & LeRoith, D. (1994) Vitamins Hormones 48, Proc. Natl. Acad. Sci. USA 93 (1996) Pietrzkowski, Z. R., Lammers, R., Carpenter, G., Soderquist, A. M., Limardo, M., Phillips, P. D., Ullrich, A. & Baserga, R. (1992) Cell Growth Differ. 3, Baker, J., Liu, J.-P., Robertson, E. J. & Efstratiadis, A. (1993) Cell 75, Liu, J.-P., Baker, J., Perkins, A. S., Robertson, E. J. & Efstratiadis, A. (1993) Cell 75, Baserga, R. (1995) Cancer Res. 55, Werner, H. & LeRoith, D. (1996) Adv. Cancer Res. 68, Sell, C., Rubini, M., Rubin, R., Lin, J.-P., Efstratiadis, A. & Baserga, R. (1993) Proc. Natl. Acad. Sci. USA 90, Sell, C., Dumenil, G., Deveaud, C., Miura, M., Coppola, D., DeAngelis, T., Rubin, R., Efstratiadis, A. & Baserga, R. (1994) Mol. CeU Biol. 14, Smale, S. T. & Baltimore, D. (1989) Cell 57, Werner, H., Bach, M. A., Stannard, B., Roberts, C. T., Jr., & LeRoith, D. (1992) Mot. Endocrinol. 6, Werner, H., Stannard, B., Bach, M. A., LeRoith, D. & Roberts, C. T., Jr. (1990) Biochem. Biophys. Res. Commun. 169, Cooke, D. W., Bankert, L. A., Roberts, C. T., Jr., LeRoith, D. & Casella, S. J. (1991) Biochem. Biophys. Res. Commun. 177, Mamula, P. W. & Goldfine, I. D. (1992) DNA Cell Biol. 11, Werner, H., Woloschak, M., Adamo, M. L., Shen-Orr, Z., Roberts, C. T., Jr., & LeRoith, D. (1989) Proc. Natl. Acad. Sci. USA 86, Oren, M. (1992) FASEB J. 6, Kern, S. E., Kinzler, K. W., Bruskin, A., Jarosz, D., Friedman, P., Prives, C. & Vogelstein, B. (1991) Science 252, Pietenpol, J. A., Tokino, T., Thiagalingam, S., El-Deiry, W. S., Kinzler, K. W. & Vogelstein, B. (1994) Proc. Natl. Acad. Sci. USA 91, El-Deiry, W. S., Tokino, T., Velculescu, V. E., Levy, D. B., Parsons, R., Trent, J. M., Lin, D., Mercer, W. E., Kinzler, K. W. & Vogelstein, B. (1993) Cell 75, Farmer, G., Bargonetti, J., Zhu, H., Friedman, P., Prywes, R. & Prives, C. (1992) Nature (London) 358, Seto, E., Usheva, A., Zambetti, G. P., Momand, J., Horikoshi, N., Weinmann, R., Levine, A. J. & Shenk, T. (1992) Proc. Natl. Acad. Sci. USA 89, Hollstein, M., Sidransky, D., Vogelstein, B. & Harris, C. C. (1991) Science 253, Harris, C. C. & Hollstein, M. (1993) N. Engl. J. Med. 329, Fogh, J., Wright, W. C. & Loveless, J. D. (1977) J. Natl. Cancer Inst. 58, Felix, C. A., Kappel, C. C., Mitsudomi, T., Nau, M. M., Tsokos, M., Crouch, G. D., Nisen, P. D., Winick, N. J. & Helman, L. J. (1992) Cancer Res. 52, Werner, H., Rauscher, F. J., III, Sukhatme, V. P., Drummond, I. A., Roberts, C. T., Jr., & LeRoith, D. (1994) J. Biol. Chem. 269, Patwardhan, S., Gashler, A., Siegel, M. G., Chang, L C., Joseph, L. J., Shows, T. B., LeBeau, M. M. & Sukhatme, V. P. (1991) Oncogene 6, Baker, S. J., Markowitz, S., Fearon, E. R., Willson, J. K. V. & Vogelstein, B. (1990) Science 249, Chen, C. & Okayama, H. (1987) Mol. Cell Biol. 7, Ueba, T., Nosaka, T., Takahashi, J. A., Shibata, F., Florkiewicz, R. Z., Vogelstein, B., Oda, Y., Kikuchi, H. & Hatanaka, M. (1994) Proc. Natl. Acad. Sci. USA 91, Chirgwin, J. M., Przbyla, A. E., MacDonald, R. J. & Rutter, W. J. (1979) Biochemistry 24, Bohan, C. A., Kashanchi, F., Ensoli, B., Buonaguro, L., Boris-Lawrie, K. A. & Brady, J. N. (1992) Gene Exp. 2, Manley, J. L., Fire, A., Cano, A., Sharp, P. A. & Gefter, M. L. (1980) Proc. Natl. Acad. Sci. USA 77, El-Deiry, W. S., Kern, S. E., Pietenpol, J. A., Kinzler, K W. & Vogelstein, B. (1992) Nat. Genet. 1, Zenzie-Gregory, B., Khachi, A., Garraway, I. P. & Smale, S. T. (1993) Mol. Cell BioL 13, Matlashewski, G., Banks, L., Pim, D. & Crawford, L. (1986) Eur. J. Biochem. 154, Truant, R., Xiao, H., Ingles, C. J. & Greenblatt, J. (1993)J. Biol. Chem. 268, Mack, D. H., Vartikar, J., Pipas, J. M. & Laimins, L. A. (1993) Nature (London) 363, Chin, K.-V., Ueda, K., Pastan, I. & Gottesman, M. M. (1992) Science 255, Baserga, R. (1994) Cell 79, Sell, C., Baserga, R. & Rubin, R. (1995) Cancer Res. 55, Resnicoff, M., Abraham, D., Yutanawiboonchai, W., Rotman, H. L., Kajstura, J., Rubin, R., Zoltick, P. & Baserga, R. (1995) Cancer Res. 55, Zhang, L., Kashanchi, F., Zhan, Q., Zhan, S., Brady, J. N., Fornace, A. J., Seth, P. & Helman, L. J. (1996) Cancer Res. 56, Buckbinder, L., Talbott, R., Velasco-Miguel, S., Takenaka, I., Faha, B., Seizinger, B. R. & Kley, N. (1995) Nature (London) 377, Werner, H., Re, G. G., Drummond, I. A., Sukhatme, V. P., Rauscher, F. J., III, Sens, D. A., Garvin, A. J., LeRoith, D. & Roberts, C. T., Jr. (1993) Proc. Natl. Acad. Sci. USA 90, Werner, H., Shen-Orr, Z., Rauscher, F. J., III, Morris, J. F., Roberts, C. T., Jr., & LeRoith, D. (1993) Mol. Cell. Biol. 15,
2.1.2 Characterization of antiviral effect of cytokine expression on HBV replication in transduced mouse hepatocytes line
 i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
In vitro analysis of pri-mirna processing. by Drosha-DGCR8 complex. (Narry Kim s lab)
 In vitro analysis of pri-mirna processing by Drosha-DGCR8 complex (Narry Kim s lab) 1-1. Preparation of radiolabeled pri-mirna transcript The RNA substrate for a cropping reaction can be prepared by in
In vitro analysis of pri-mirna processing by Drosha-DGCR8 complex (Narry Kim s lab) 1-1. Preparation of radiolabeled pri-mirna transcript The RNA substrate for a cropping reaction can be prepared by in
QUANTITATIVE RT-PCR. A = B (1+e) n. A=amplified products, B=input templates, n=cycle number, and e=amplification efficiency.
 QUANTITATIVE RT-PCR Application: Quantitative RT-PCR is used to quantify mrna in both relative and absolute terms. It can be applied for the quantification of mrna expressed from endogenous genes, and
QUANTITATIVE RT-PCR Application: Quantitative RT-PCR is used to quantify mrna in both relative and absolute terms. It can be applied for the quantification of mrna expressed from endogenous genes, and
RevertAid Premium First Strand cdna Synthesis Kit
 RevertAid Premium First Strand cdna Synthesis Kit #K1651, #K1652 CERTIFICATE OF ANALYSIS #K1651 Lot QUALITY CONTROL RT-PCR using 100 fg of control GAPDH RNA and GAPDH control primers generated a prominent
RevertAid Premium First Strand cdna Synthesis Kit #K1651, #K1652 CERTIFICATE OF ANALYSIS #K1651 Lot QUALITY CONTROL RT-PCR using 100 fg of control GAPDH RNA and GAPDH control primers generated a prominent
Understanding the immune response to bacterial infections
 Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
4.1 3T12 and 312 are immortalized cell lines with transforming potential:
 DISCUSSION CHAPTER 4 4.1 3T12 and 312 are immortalized cell lines with transforming potential: Transformation of a normal cell with finite number of divisions into a tumorigenic cell of potentially infinite
DISCUSSION CHAPTER 4 4.1 3T12 and 312 are immortalized cell lines with transforming potential: Transformation of a normal cell with finite number of divisions into a tumorigenic cell of potentially infinite
Rapid GST Inclusion Body Solubilization and Renaturation Kit
 Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
Molecular Biology Techniques: A Classroom Laboratory Manual THIRD EDITION
 Molecular Biology Techniques: A Classroom Laboratory Manual THIRD EDITION Susan Carson Heather B. Miller D.Scott Witherow ELSEVIER AMSTERDAM BOSTON HEIDELBERG LONDON NEW YORK OXFORD PARIS SAN DIEGO SAN
Molecular Biology Techniques: A Classroom Laboratory Manual THIRD EDITION Susan Carson Heather B. Miller D.Scott Witherow ELSEVIER AMSTERDAM BOSTON HEIDELBERG LONDON NEW YORK OXFORD PARIS SAN DIEGO SAN
The F Box Protein Fbx6 Regulates Chk1 Stability and Cellular Sensitivity to Replication Stress
 Molecular Cell, Volume 35 Supplemental Data The F Box Protein Fbx6 Regulates Chk1 Stability and Cellular Sensitivity to Replication Stress You-Wei Zhang, John Brognard, Chris Coughlin, Zhongsheng You,
Molecular Cell, Volume 35 Supplemental Data The F Box Protein Fbx6 Regulates Chk1 Stability and Cellular Sensitivity to Replication Stress You-Wei Zhang, John Brognard, Chris Coughlin, Zhongsheng You,
RNA Viruses. A Practical Approac h. Alan J. Cann
 RNA Viruses A Practical Approac h Alan J. Cann List of protocols page xiii Abbreviations xvii Investigation of RNA virus genome structure 1 A j. Easton, A.C. Marriott and C.R. Pringl e 1 Introduction-the
RNA Viruses A Practical Approac h Alan J. Cann List of protocols page xiii Abbreviations xvii Investigation of RNA virus genome structure 1 A j. Easton, A.C. Marriott and C.R. Pringl e 1 Introduction-the
The p53-mdm-2 autoregulatory
 The p53-mdm-2 autoregulatory feedl ack loop Xiangwei Wu, J. Henri Bayle, David Olson, and Arnold J. Levine 1 Department of Molecular Biology, Princeton University, Princeton, New Jersey 08544-1014 USA
The p53-mdm-2 autoregulatory feedl ack loop Xiangwei Wu, J. Henri Bayle, David Olson, and Arnold J. Levine 1 Department of Molecular Biology, Princeton University, Princeton, New Jersey 08544-1014 USA
Notch 1 -dependent regulation of cell fate in colorectal cancer
 Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
Application Guide... 2
 Protocol for GenomePlex Whole Genome Amplification from Formalin-Fixed Parrafin-Embedded (FFPE) tissue Application Guide... 2 I. Description... 2 II. Product Components... 2 III. Materials to be Supplied
Protocol for GenomePlex Whole Genome Amplification from Formalin-Fixed Parrafin-Embedded (FFPE) tissue Application Guide... 2 I. Description... 2 II. Product Components... 2 III. Materials to be Supplied
Angelo Peschiaroli, N. Valerio Dorrello, Daniele Guardavaccaro, Monica Venere, Thanos Halazonetis, Nicholas E. Sherman, and Michele Pagano
 Molecular Cell, Volume 23 Supplemental Data SCF βtrcp -Mediated Degradation of Claspin Regulates Recovery from the DNA Replication Checkpoint Response Angelo Peschiaroli, N. Valerio Dorrello, Daniele Guardavaccaro,
Molecular Cell, Volume 23 Supplemental Data SCF βtrcp -Mediated Degradation of Claspin Regulates Recovery from the DNA Replication Checkpoint Response Angelo Peschiaroli, N. Valerio Dorrello, Daniele Guardavaccaro,
The Need for a PARP in vivo Pharmacodynamic Assay
 The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D., Chief Scientific Officer, Trevigen, Inc., Gaithersburg, MD For further infomation, please contact: William Booth, Ph.D. Tel: +44 (0)1235
The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D., Chief Scientific Officer, Trevigen, Inc., Gaithersburg, MD For further infomation, please contact: William Booth, Ph.D. Tel: +44 (0)1235
Anti-ATF6 α antibody, mouse monoclonal (1-7)
 Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
DP419 RNAsimple Total RNA Kit. RNAprep pure Series. DP501 mircute mirna Isolation Kit. DP438 MagGene Viral DNA / RNA Kit. DP405 TRNzol Reagent
 Overview of TIANGEN Products DP419 RNAsimple Total RNA Kit DP430 RNAprep pure Kit(For Cell/Bacteria) DP315/DP315-R TIANamp Virus DNA/RNA Kit DP431 RNAprep pure Kit (For Tissue) Silica-membrane Technology
Overview of TIANGEN Products DP419 RNAsimple Total RNA Kit DP430 RNAprep pure Kit(For Cell/Bacteria) DP315/DP315-R TIANamp Virus DNA/RNA Kit DP431 RNAprep pure Kit (For Tissue) Silica-membrane Technology
First Strand cdna Synthesis
 380PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name First Strand cdna Synthesis (Cat. # 786 812) think proteins! think G-Biosciences
380PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name First Strand cdna Synthesis (Cat. # 786 812) think proteins! think G-Biosciences
STOP. Before using this product, please read the Limited Use License statement below:
 STOP Before using this product, please read the Limited Use License statement below: Important Limited Use License information for pdrive5lucia-rgfap The purchase of the pdrive5lucia-rgfap vector conveys
STOP Before using this product, please read the Limited Use License statement below: Important Limited Use License information for pdrive5lucia-rgfap The purchase of the pdrive5lucia-rgfap vector conveys
Gene Expression Assays
 APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
Chromatin Immunoprecipitation
 Chromatin Immunoprecipitation A) Prepare a yeast culture (see the Galactose Induction Protocol for details). 1) Start a small culture (e.g. 2 ml) in YEPD or selective media from a single colony. 2) Spin
Chromatin Immunoprecipitation A) Prepare a yeast culture (see the Galactose Induction Protocol for details). 1) Start a small culture (e.g. 2 ml) in YEPD or selective media from a single colony. 2) Spin
Integrated Protein Services
 Integrated Protein Services Custom protein expression & purification Version DC04-0012 Expression strategy The first step in the recombinant protein generation process is to design an appropriate expression
Integrated Protein Services Custom protein expression & purification Version DC04-0012 Expression strategy The first step in the recombinant protein generation process is to design an appropriate expression
MEF Starter Nucleofector Kit
 page 1 of 7 MEF Starter Nucleofector Kit for Mouse Embryonic Fibroblasts (MEF) MEF display significant phenotypic variations which depend on the strain, the genetic background of the mice they are isolated
page 1 of 7 MEF Starter Nucleofector Kit for Mouse Embryonic Fibroblasts (MEF) MEF display significant phenotypic variations which depend on the strain, the genetic background of the mice they are isolated
Molecular Cloning, Product Brochure
 , Product Brochure Interest in any of the products, request or order them at Bio-Connect. Bio-Connect B.V. T NL +31 (0)26 326 44 50 T BE +32 (0)2 503 03 48 Begonialaan 3a F NL +31 (0)26 326 44 51 F BE
, Product Brochure Interest in any of the products, request or order them at Bio-Connect. Bio-Connect B.V. T NL +31 (0)26 326 44 50 T BE +32 (0)2 503 03 48 Begonialaan 3a F NL +31 (0)26 326 44 51 F BE
DNA Fingerprinting. Unless they are identical twins, individuals have unique DNA
 DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
DNA Fingerprinting Unless they are identical twins, individuals have unique DNA DNA fingerprinting The name used for the unambiguous identifying technique that takes advantage of differences in DNA sequence
Supplementary Figure 1.
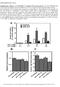 Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Essentials of Real Time PCR. About Sequence Detection Chemistries
 Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
PCR was carried out in a reaction volume of 20 µl using the ABI AmpliTaq GOLD kit (ABI,
 Supplemental Text/Tables PCR Amplification and Sequencing PCR was carried out in a reaction volume of 20 µl using the ABI AmpliTaq GOLD kit (ABI, Foster City, CA). Each PCR reaction contained 20 ng genomic
Supplemental Text/Tables PCR Amplification and Sequencing PCR was carried out in a reaction volume of 20 µl using the ABI AmpliTaq GOLD kit (ABI, Foster City, CA). Each PCR reaction contained 20 ng genomic
Genomic DNA Extraction Kit INSTRUCTION MANUAL
 Genomic DNA Extraction Kit INSTRUCTION MANUAL Table of Contents Introduction 3 Kit Components 3 Storage Conditions 4 Recommended Equipment and Reagents 4 Introduction to the Protocol 4 General Overview
Genomic DNA Extraction Kit INSTRUCTION MANUAL Table of Contents Introduction 3 Kit Components 3 Storage Conditions 4 Recommended Equipment and Reagents 4 Introduction to the Protocol 4 General Overview
The cell lines used in this study were obtained from the American Type Culture
 Supplementary materials and methods Cell culture and drug treatments The cell lines used in this study were obtained from the American Type Culture Collection (ATCC) and grown as previously described.
Supplementary materials and methods Cell culture and drug treatments The cell lines used in this study were obtained from the American Type Culture Collection (ATCC) and grown as previously described.
How To Make A Tri Reagent
 TRI Reagent For processing tissues, cells cultured in monolayer or cell pellets Catalog Number T9424 Store at room temperature. TECHNICAL BULLETIN Product Description TRI Reagent is a quick and convenient
TRI Reagent For processing tissues, cells cultured in monolayer or cell pellets Catalog Number T9424 Store at room temperature. TECHNICAL BULLETIN Product Description TRI Reagent is a quick and convenient
HCS604.03 Exercise 1 Dr. Jones Spring 2005. Recombinant DNA (Molecular Cloning) exercise:
 HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
Genomic DNA Clean & Concentrator Catalog Nos. D4010 & D4011
 Page 0 INSTRUCTION MANUAL Catalog Nos. D4010 & D4011 Highlights Quick (5 minute) spin column recovery of large-sized DNA (e.g., genomic, mitochondrial, plasmid (BAC/PAC), viral, phage, (wga)dna, etc.)
Page 0 INSTRUCTION MANUAL Catalog Nos. D4010 & D4011 Highlights Quick (5 minute) spin column recovery of large-sized DNA (e.g., genomic, mitochondrial, plasmid (BAC/PAC), viral, phage, (wga)dna, etc.)
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation
 OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
Southern Blot Analysis (from Baker lab, university of Florida)
 Southern Blot Analysis (from Baker lab, university of Florida) DNA Prep Prepare DNA via your favorite method. You may find a protocol under Mini Yeast Genomic Prep. Restriction Digest 1.Digest DNA with
Southern Blot Analysis (from Baker lab, university of Florida) DNA Prep Prepare DNA via your favorite method. You may find a protocol under Mini Yeast Genomic Prep. Restriction Digest 1.Digest DNA with
PyroPhage 3173 DNA Polymerase, Exonuclease Minus (Exo-)
 PyroPhage 3173 DNA Polymerase, Exonuclease Minus (Exo-) FOR RESEARCH USE ONLY. NOT FOR HUMAN OR DIAGNOSTIC USE Lucigen Corporation 2905 Parmenter St, Middleton, WI 53562 USA Toll Free: (888) 575-9695 (608)
PyroPhage 3173 DNA Polymerase, Exonuclease Minus (Exo-) FOR RESEARCH USE ONLY. NOT FOR HUMAN OR DIAGNOSTIC USE Lucigen Corporation 2905 Parmenter St, Middleton, WI 53562 USA Toll Free: (888) 575-9695 (608)
Introduction. Preparation of Template DNA
 Procedures and Recommendations for DNA Sequencing at the Plant-Microbe Genomics Facility Ohio State University Biological Sciences Building Room 420, 484 W. 12th Ave., Columbus OH 43210 Telephone: 614/247-6204;
Procedures and Recommendations for DNA Sequencing at the Plant-Microbe Genomics Facility Ohio State University Biological Sciences Building Room 420, 484 W. 12th Ave., Columbus OH 43210 Telephone: 614/247-6204;
Pure-IP Western Blot Detection Kit
 Product Manual Pure-IP Western Blot Detection Kit Catalog Number PRB-5002 20 blots FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction The technique of immunoprecipitation (IP) is used
Product Manual Pure-IP Western Blot Detection Kit Catalog Number PRB-5002 20 blots FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction The technique of immunoprecipitation (IP) is used
TABLE OF CONTENT. Page ACKNOWLEDGEMENTS. iii ENGLISH ABSTRACT THAI ABSTRACT. vii LIST OF TABLES LIST OF FIGURES. xvi ABBREVIATIONS.
 x TABLE OF CONTENT ACKNOWLEDGEMENTS ENGLISH ABSTRACT THAI ABSTRACT LIST OF TABLES LIST OF FIGURES ABBREVIATIONS iii iv vii xv xvi xviii CHAPTER I: INTRODUCTION 1.1 Statement of problems 1 1.2 Literature
x TABLE OF CONTENT ACKNOWLEDGEMENTS ENGLISH ABSTRACT THAI ABSTRACT LIST OF TABLES LIST OF FIGURES ABBREVIATIONS iii iv vii xv xvi xviii CHAPTER I: INTRODUCTION 1.1 Statement of problems 1 1.2 Literature
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual
 Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
KMS-Specialist & Customized Biosimilar Service
 KMS-Specialist & Customized Biosimilar Service 1. Polyclonal Antibody Development Service KMS offering a variety of Polyclonal Antibody Services to fit your research and production needs. we develop polyclonal
KMS-Specialist & Customized Biosimilar Service 1. Polyclonal Antibody Development Service KMS offering a variety of Polyclonal Antibody Services to fit your research and production needs. we develop polyclonal
CompleteⅡ 1st strand cdna Synthesis Kit
 Instruction Manual CompleteⅡ 1st strand cdna Synthesis Kit Catalog # GM30401, GM30402 Green Mountain Biosystems. LLC Web: www.greenmountainbio.com Tel: 800-942-1160 Sales: Sales@ greenmountainbio.com Support:
Instruction Manual CompleteⅡ 1st strand cdna Synthesis Kit Catalog # GM30401, GM30402 Green Mountain Biosystems. LLC Web: www.greenmountainbio.com Tel: 800-942-1160 Sales: Sales@ greenmountainbio.com Support:
Chromatin Immunoprecipitation (ChIP)
 Chromatin Immunoprecipitation (ChIP) Day 1 A) DNA shearing 1. Samples Dissect tissue (One Mouse OBs) of interest and transfer to an eppendorf containing 0.5 ml of dissecting media (on ice) or PBS but without
Chromatin Immunoprecipitation (ChIP) Day 1 A) DNA shearing 1. Samples Dissect tissue (One Mouse OBs) of interest and transfer to an eppendorf containing 0.5 ml of dissecting media (on ice) or PBS but without
Module 3 Questions. 7. Chemotaxis is an example of signal transduction. Explain, with the use of diagrams.
 Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
pcas-guide System Validation in Genome Editing
 pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
Protein Expression and Analysis. Vijay Yajnik, MD, PhD GI Unit MGH
 Protein Expression and Analysis Vijay Yajnik, MD, PhD GI Unit MGH Identify your needs Antigen production Biochemical studies Cell Biology Protein interaction studies including proteomics Structural studies
Protein Expression and Analysis Vijay Yajnik, MD, PhD GI Unit MGH Identify your needs Antigen production Biochemical studies Cell Biology Protein interaction studies including proteomics Structural studies
Chapter 18: Applications of Immunology
 Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
ptune Inducible Vector
 ptune Inducible Vector Application Guide Table of Contents Package contents and Storage Conditions:...2 Related products:...2 Introduction...2 Figure 1. Schematic Diagrams of ptune Inducible vector...3
ptune Inducible Vector Application Guide Table of Contents Package contents and Storage Conditions:...2 Related products:...2 Introduction...2 Figure 1. Schematic Diagrams of ptune Inducible vector...3
Transcription in prokaryotes. Elongation and termination
 Transcription in prokaryotes Elongation and termination After initiation the σ factor leaves the scene. Core polymerase is conducting the elongation of the chain. The core polymerase contains main nucleotide
Transcription in prokaryotes Elongation and termination After initiation the σ factor leaves the scene. Core polymerase is conducting the elongation of the chain. The core polymerase contains main nucleotide
Analysis of the DNA Methylation Patterns at the BRCA1 CpG Island
 Analysis of the DNA Methylation Patterns at the BRCA1 CpG Island Frédérique Magdinier 1 and Robert Dante 2 1 Laboratory of Molecular Biology of the Cell, Ecole Normale Superieure, Lyon, France 2 Laboratory
Analysis of the DNA Methylation Patterns at the BRCA1 CpG Island Frédérique Magdinier 1 and Robert Dante 2 1 Laboratory of Molecular Biology of the Cell, Ecole Normale Superieure, Lyon, France 2 Laboratory
Classic Immunoprecipitation
 292PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Classic Immunoprecipitation Utilizes Protein A/G Agarose for Antibody Binding (Cat.
292PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Classic Immunoprecipitation Utilizes Protein A/G Agarose for Antibody Binding (Cat.
Cloning Blunt-End Pfu DNA Polymerase- Generated PCR Fragments into pgem -T Vector Systems
 Promega Notes Number 71, 1999, p. 10 Blunt-End Pfu DNA Polymerase- Generated PCR Fragments into pgem -T Vector Systems By Kimberly Knoche, Ph.D., and Dan Kephart, Ph.D. Promega Corporation Corresponding
Promega Notes Number 71, 1999, p. 10 Blunt-End Pfu DNA Polymerase- Generated PCR Fragments into pgem -T Vector Systems By Kimberly Knoche, Ph.D., and Dan Kephart, Ph.D. Promega Corporation Corresponding
Reverse Transcription System
 TECHNICAL BULLETIN Reverse Transcription System Instruc ons for use of Product A3500 Revised 1/14 TB099 Reverse Transcription System All technical literature is available on the Internet at: www.promega.com/protocols/
TECHNICAL BULLETIN Reverse Transcription System Instruc ons for use of Product A3500 Revised 1/14 TB099 Reverse Transcription System All technical literature is available on the Internet at: www.promega.com/protocols/
BUFFERS and MEDIAS Coomassie Blue Staining Solution Coomassie blue Destaining Solution DMEM Normal Cell Culture Media
 BUFFERS and MEDIAS Coomassie Blue Staining Solution 2 g Coomassie Blue 2 L Methanol or Ethanol * 1.6 L 400 ml Glacial acetic acid *If you will be microwaving the gel in staining solution for rapid staining
BUFFERS and MEDIAS Coomassie Blue Staining Solution 2 g Coomassie Blue 2 L Methanol or Ethanol * 1.6 L 400 ml Glacial acetic acid *If you will be microwaving the gel in staining solution for rapid staining
ab185916 Hi-Fi cdna Synthesis Kit
 ab185916 Hi-Fi cdna Synthesis Kit Instructions for Use For cdna synthesis from various RNA samples This product is for research use only and is not intended for diagnostic use. Version 1 Last Updated 1
ab185916 Hi-Fi cdna Synthesis Kit Instructions for Use For cdna synthesis from various RNA samples This product is for research use only and is not intended for diagnostic use. Version 1 Last Updated 1
Martin D. Smith, Sally J. Dawson, Linda M. Boxer 1 and David S. Latchman* 4100 4107 Nucleic Acids Research, 1998, Vol. 26, No. 18
 4100 4107 Nucleic Acids Research, 1998, Vol. 26, No. 18 1998 Oxford University Press The N-terminal domain unique to the long form of the Brn-3a transcription factor is essential to protect neuronal cells
4100 4107 Nucleic Acids Research, 1998, Vol. 26, No. 18 1998 Oxford University Press The N-terminal domain unique to the long form of the Brn-3a transcription factor is essential to protect neuronal cells
Updated: July 2005. 5' End label RNA markers 18.113 (18mer) and 44.12 (24mer) with Kinase and 32P-gamma-ATP. Gel purify labeled markers.
 RNA Cloning Method Flowchart Extract total RNA containing small RNAs. Check quality on denaturing gel with EtBr staining 5' End label RNA markers 18.113 (18mer) and 44.12 (24mer) with Kinase and 32P-gamma-ATP.
RNA Cloning Method Flowchart Extract total RNA containing small RNAs. Check quality on denaturing gel with EtBr staining 5' End label RNA markers 18.113 (18mer) and 44.12 (24mer) with Kinase and 32P-gamma-ATP.
2006 7.012 Problem Set 6 KEY
 2006 7.012 Problem Set 6 KEY ** Due before 5 PM on WEDNESDAY, November 22, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You create an artificial
2006 7.012 Problem Set 6 KEY ** Due before 5 PM on WEDNESDAY, November 22, 2006. ** Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. You create an artificial
7.012 Quiz 3 practice
 MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel 7.012 Quiz 3 practice Quiz 3 on Friday, November 12th
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel 7.012 Quiz 3 practice Quiz 3 on Friday, November 12th
DNA Core Facility: DNA Sequencing Guide
 DNA Core Facility: DNA Sequencing Guide University of Missouri-Columbia 216 Life Sciences Center Columbia, MO 65211 http://biotech.missouri.edu/dnacore/ Table of Contents 1. Evaluating Sequencing Data..
DNA Core Facility: DNA Sequencing Guide University of Missouri-Columbia 216 Life Sciences Center Columbia, MO 65211 http://biotech.missouri.edu/dnacore/ Table of Contents 1. Evaluating Sequencing Data..
Procedures For DNA Sequencing
 Procedures For DNA Sequencing Plant-Microbe Genomics Facility (PMGF) Ohio State University 420 Biological Sciences Building 484 W. 12th Ave., Columbus OH 43210 Telephone: 614/247-6204 FAX: 614/292-6337
Procedures For DNA Sequencing Plant-Microbe Genomics Facility (PMGF) Ohio State University 420 Biological Sciences Building 484 W. 12th Ave., Columbus OH 43210 Telephone: 614/247-6204 FAX: 614/292-6337
Supplemental Fig. S1. The schematic diagrams of the expression constructs used in this study.
 1 Supplemental data Supplemental Fig. S1. The schematic diagrams of the expression constructs used in this study. Supplemental Fig. S2. Ingenuity Pathway Analysis (IPA) of the 56 putative caspase substrates
1 Supplemental data Supplemental Fig. S1. The schematic diagrams of the expression constructs used in this study. Supplemental Fig. S2. Ingenuity Pathway Analysis (IPA) of the 56 putative caspase substrates
Hepatitis B Virus Genemer Mix
 Product Manual Hepatitis B Virus Genemer Mix Primer Pair for amplification of HBV Specific DNA Fragment Includes Internal Negative Control Primers and Template Catalog No.: 60-2007-12 Store at 20 o C For
Product Manual Hepatitis B Virus Genemer Mix Primer Pair for amplification of HBV Specific DNA Fragment Includes Internal Negative Control Primers and Template Catalog No.: 60-2007-12 Store at 20 o C For
Mouse ES Cell Nucleofector Kit
 page 1 of 7 Mouse ES Cell Nucleofector Kit for Mouse Embryonic Stem Cells Cell type Origin Cells derived from mouse blastocysts. Morphology Round cells growing in clumps. Important remarks! 1. This protocol
page 1 of 7 Mouse ES Cell Nucleofector Kit for Mouse Embryonic Stem Cells Cell type Origin Cells derived from mouse blastocysts. Morphology Round cells growing in clumps. Important remarks! 1. This protocol
Protein Expression. A Practical Approach J. HIGGIN S
 Protein Expression A Practical Approach S. J. HIGGIN S B. D. HAMES List of contributors Abbreviations xv Xvi i 1. Protein expression in mammalian cell s Marlies Otter-Nilsson and Tommy Nilsso n 1. Introduction
Protein Expression A Practical Approach S. J. HIGGIN S B. D. HAMES List of contributors Abbreviations xv Xvi i 1. Protein expression in mammalian cell s Marlies Otter-Nilsson and Tommy Nilsso n 1. Introduction
PNA BRAF Mutation Detection Kit
 - PNA BRAF Mutation Detection Kit Catalog Number KA2102 50 tests/kit Version: 01 Intended for research use only www.abnova.com Introduction and Background Intended use The PNA BRAF Mutation Detection Kit
- PNA BRAF Mutation Detection Kit Catalog Number KA2102 50 tests/kit Version: 01 Intended for research use only www.abnova.com Introduction and Background Intended use The PNA BRAF Mutation Detection Kit
CUSTOM ANTIBODIES. Fully customised services: rat and murine monoclonals, rat and rabbit polyclonals, antibody characterisation, antigen preparation
 CUSTOM ANTIBODIES Highly competitive pricing without compromising quality. Rat monoclonal antibodies for the study of gene expression and proteomics in mice and in mouse models of human diseases available.
CUSTOM ANTIBODIES Highly competitive pricing without compromising quality. Rat monoclonal antibodies for the study of gene expression and proteomics in mice and in mouse models of human diseases available.
PrimeSTAR HS DNA Polymerase
 Cat. # R010A For Research Use PrimeSTAR HS DNA Polymerase Product Manual Table of Contents I. Description...3 II. III. IV. Components...3 Storage...3 Features...3 V. General Composition of PCR Reaction
Cat. # R010A For Research Use PrimeSTAR HS DNA Polymerase Product Manual Table of Contents I. Description...3 II. III. IV. Components...3 Storage...3 Features...3 V. General Composition of PCR Reaction
RT31-020 20 rxns. RT31-100 100 rxns TRANSCRIPTME Enzyme Mix (1) 40 µl 2 x 50 µl 5 x 40 µl
 Components RT31-020 20 rxns RT31-050 50 rxns RT31-100 100 rxns TRANSCRIPTME Enzyme Mix (1) 40 µl 2 x 50 µl 5 x 40 µl 2x RT Master Mix (2) 200 µl 2 x 250 µl 5 x 200 µl RNase H (E. coli) 20 µl 2 x 25 µl
Components RT31-020 20 rxns RT31-050 50 rxns RT31-100 100 rxns TRANSCRIPTME Enzyme Mix (1) 40 µl 2 x 50 µl 5 x 40 µl 2x RT Master Mix (2) 200 µl 2 x 250 µl 5 x 200 µl RNase H (E. coli) 20 µl 2 x 25 µl
Chapter 5: Organization and Expression of Immunoglobulin Genes
 Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE Q5B
 INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
Troubleshooting Guide for DNA Electrophoresis
 Troubleshooting Guide for Electrophoresis. ELECTROPHORESIS Protocols and Recommendations for Electrophoresis electrophoresis problem 1 Low intensity of all or some bands 2 Smeared bands 3 Atypical banding
Troubleshooting Guide for Electrophoresis. ELECTROPHORESIS Protocols and Recommendations for Electrophoresis electrophoresis problem 1 Low intensity of all or some bands 2 Smeared bands 3 Atypical banding
Integrated Protein Services
 Integrated Protein Services Custom protein expression & purification Last date of revision June 2015 Version DC04-0013 www.iba-lifesciences.com Expression strategy The first step in the recombinant protein
Integrated Protein Services Custom protein expression & purification Last date of revision June 2015 Version DC04-0013 www.iba-lifesciences.com Expression strategy The first step in the recombinant protein
Exploiting science for engineering: BRCA2 targeted therapies
 20.109 MOD1 DNA ENGINEERING Fall 2010 Exploiting science for engineering: BRCA2 targeted therapies Orsi Kiraly Engelward lab Homologous recombination is important No HR chromosomal aberrations cell death
20.109 MOD1 DNA ENGINEERING Fall 2010 Exploiting science for engineering: BRCA2 targeted therapies Orsi Kiraly Engelward lab Homologous recombination is important No HR chromosomal aberrations cell death
Publikationsliste Claudia Götz
 Publikationsliste Claudia Götz 1. Reinhard,B., Götz, C., and Faillard, H.: Synthesis of N-Acetyl-9-Oacetylneuraminic acid α-p-aminophenylthioketoside and its application as ligand in the affinity chromatography
Publikationsliste Claudia Götz 1. Reinhard,B., Götz, C., and Faillard, H.: Synthesis of N-Acetyl-9-Oacetylneuraminic acid α-p-aminophenylthioketoside and its application as ligand in the affinity chromatography
Rapid Acquisition of Unknown DNA Sequence Adjacent to a Known Segment by Multiplex Restriction Site PCR
 Rapid Acquisition of Unknown DNA Sequence Adjacent to a Known Segment by Multiplex Restriction Site PCR BioTechniques 25:415-419 (September 1998) ABSTRACT The determination of unknown DNA sequences around
Rapid Acquisition of Unknown DNA Sequence Adjacent to a Known Segment by Multiplex Restriction Site PCR BioTechniques 25:415-419 (September 1998) ABSTRACT The determination of unknown DNA sequences around
2. Are there any chaperones such as GroES, GroEL, and DnaK..etc added into the solution A or solution B in the kit?
 GENERAL 1. What are the MW limits of proteins that can be produced by PURExpress? 2. Are there any chaperones such as GroES, GroEL, and DnaK..etc added into the solution A or solution B in the kit? 3.
GENERAL 1. What are the MW limits of proteins that can be produced by PURExpress? 2. Are there any chaperones such as GroES, GroEL, and DnaK..etc added into the solution A or solution B in the kit? 3.
Rubisco; easy Purification and Immunochemical Determination
 Rubisco; easy Purification and Immunochemical Determination Ulrich Groß Justus-Liebig-Universität Gießen, Institute of Plant Nutrition, Department of Tissue Culture, Südanlage 6, D-35390 Giessen e-mail:
Rubisco; easy Purification and Immunochemical Determination Ulrich Groß Justus-Liebig-Universität Gießen, Institute of Plant Nutrition, Department of Tissue Culture, Südanlage 6, D-35390 Giessen e-mail:
Chapter 11. Rapid Screening of Gli2/3 Mutants Using the Flp-In System. Pawel Niewiadomski and Rajat Rohatgi. Abstract.
 Chapter 11 Rapid Screening of Gli2/3 Mutants Using the Flp-In System Abstract Gli2 and Gli3 respond to the Hedgehog (Hh) signal in mammals by undergoing posttranslational modifications and moving to the
Chapter 11 Rapid Screening of Gli2/3 Mutants Using the Flp-In System Abstract Gli2 and Gli3 respond to the Hedgehog (Hh) signal in mammals by undergoing posttranslational modifications and moving to the
TECHNICAL BULLETIN. HIS-Select Nickel Affinity Gel. Catalog Number P6611 Storage Temperature 2 8 C
 HIS-Select Nickel Affinity Gel Catalog Number P6611 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description HIS-Select Nickel Affinity Gel is an immobilized metalion affinity chromatography (IMAC)
HIS-Select Nickel Affinity Gel Catalog Number P6611 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description HIS-Select Nickel Affinity Gel is an immobilized metalion affinity chromatography (IMAC)
MICB ABI PRISM 310 SEQUENCING GUIDE SEQUENCING OF PLASMID DNA
 Plasmid DNA Preparation MICB ABI PRISM 310 SEQUENCING GUIDE SEQUENCING OF PLASMID DNA Introduction: I have always used the classic Alkaline Lysis miniprep method to isolate plasmid DNA. (See below) If
Plasmid DNA Preparation MICB ABI PRISM 310 SEQUENCING GUIDE SEQUENCING OF PLASMID DNA Introduction: I have always used the classic Alkaline Lysis miniprep method to isolate plasmid DNA. (See below) If
HighPure Maxi Plasmid Kit
 HighPure Maxi Plasmid Kit For purification of high pure plasmid DNA with high yields www.tiangen.com PP120109 HighPure Maxi Plasmid Kit Kit Contents Storage Cat.no. DP116 Contents RNaseA (100 mg/ml) Buffer
HighPure Maxi Plasmid Kit For purification of high pure plasmid DNA with high yields www.tiangen.com PP120109 HighPure Maxi Plasmid Kit Kit Contents Storage Cat.no. DP116 Contents RNaseA (100 mg/ml) Buffer
Troubleshooting Sequencing Data
 Troubleshooting Sequencing Data Troubleshooting Sequencing Data No recognizable sequence (see page 7-10) Insufficient Quantitate the DNA. Increase the amount of DNA in the sequencing reactions. See page
Troubleshooting Sequencing Data Troubleshooting Sequencing Data No recognizable sequence (see page 7-10) Insufficient Quantitate the DNA. Increase the amount of DNA in the sequencing reactions. See page
GENOTYPING ASSAYS AT ZIRC
 GENOTYPING ASSAYS AT ZIRC A. READ THIS FIRST - DISCLAIMER Dear ZIRC user, We now provide detailed genotyping protocols for a number of zebrafish lines distributed by ZIRC. These protocols were developed
GENOTYPING ASSAYS AT ZIRC A. READ THIS FIRST - DISCLAIMER Dear ZIRC user, We now provide detailed genotyping protocols for a number of zebrafish lines distributed by ZIRC. These protocols were developed
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College
 Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
STUDIES ON SEED STORAGE PROTEINS OF SOME ECONOMICALLY MINOR PLANTS
 STUDIES ON SEED STORAGE PROTEINS OF SOME ECONOMICALLY MINOR PLANTS THESIS SUBMITTED FOR THE DEGREB OF DOCTOR OF PHILOSOPHY (SCIENCE) OF THE UNIVERSITY OF CALCUTTA 1996 NRISINHA DE, M.Sc DEPARTMENT OF BIOCHEMISTRY
STUDIES ON SEED STORAGE PROTEINS OF SOME ECONOMICALLY MINOR PLANTS THESIS SUBMITTED FOR THE DEGREB OF DOCTOR OF PHILOSOPHY (SCIENCE) OF THE UNIVERSITY OF CALCUTTA 1996 NRISINHA DE, M.Sc DEPARTMENT OF BIOCHEMISTRY
'LVFXVVLRQ $UUD\HGF'1$H[SUHVVLRQOLEUDULHV 5RERWWHFKQRORJ\DQGDUUD\HGOLEUDULHV
 Discussion 74 'LVFXVVLRQ This study describes arrayed cdna libraries as a source of clonally expressed recombinant proteins which can be directly linked to clones characterised and identified by DNA hybridisation
Discussion 74 'LVFXVVLRQ This study describes arrayed cdna libraries as a source of clonally expressed recombinant proteins which can be directly linked to clones characterised and identified by DNA hybridisation
Expresión de la información genética en eucariotas. similitudes y diferencias con procariotas. metodologías de estudio 2
 Expresión de la información genética en eucariotas similitudes y diferencias con procariotas metodologías de estudio 2 Víctor Romanowski, 2013 Polymerase chain reaction (PCR) The polymerase chain reaction
Expresión de la información genética en eucariotas similitudes y diferencias con procariotas metodologías de estudio 2 Víctor Romanowski, 2013 Polymerase chain reaction (PCR) The polymerase chain reaction
Troubleshooting the Single-step PCR Site-directed Mutagenesis Procedure Intended to Create a Non-functional rop Gene in the pbr322 Plasmid
 Troubleshooting the Single-step PCR Site-directed Mutagenesis Procedure Intended to Create a Non-functional rop Gene in the pbr322 Plasmid Lina Jew Department of Microbiology & Immunology, University of
Troubleshooting the Single-step PCR Site-directed Mutagenesis Procedure Intended to Create a Non-functional rop Gene in the pbr322 Plasmid Lina Jew Department of Microbiology & Immunology, University of
pcmv6-neo Vector Application Guide Contents
 pcmv6-neo Vector Application Guide Contents Package Contents and Storage Conditions... 2 Product Description... 2 Introduction... 2 Production and Quality Assurance... 2 Methods... 3 Other required reagents...
pcmv6-neo Vector Application Guide Contents Package Contents and Storage Conditions... 2 Product Description... 2 Introduction... 2 Production and Quality Assurance... 2 Methods... 3 Other required reagents...
Genetics Test Biology I
 Genetics Test Biology I Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Avery s experiments showed that bacteria are transformed by a. RNA. c. proteins.
Genetics Test Biology I Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Avery s experiments showed that bacteria are transformed by a. RNA. c. proteins.
Contents. molecular biology techniques. - Mutations in Factor II. - Mutations in MTHFR gene. - Breast cencer genes. - p53 and breast cancer
 Contents Introduction: biology and medicine, two separated compartments What we need to know: - boring basics in DNA/RNA structure and overview of particular aspects of molecular biology techniques - How
Contents Introduction: biology and medicine, two separated compartments What we need to know: - boring basics in DNA/RNA structure and overview of particular aspects of molecular biology techniques - How
MagExtractor -Genome-
 Instruction manual MagExtractor-Genome-0810 F0981K MagExtractor -Genome- NPK-101 100 preparations Store at 4 C Contents [1] Introduction [2] Components [3] Materials required [4] Protocol 1. Purification
Instruction manual MagExtractor-Genome-0810 F0981K MagExtractor -Genome- NPK-101 100 preparations Store at 4 C Contents [1] Introduction [2] Components [3] Materials required [4] Protocol 1. Purification
Aurora Forensic Sample Clean-up Protocol
 Aurora Forensic Sample Clean-up Protocol 106-0008-BA-D 2015 Boreal Genomics, Inc. All rights reserved. All trademarks are property of their owners. http://www.borealgenomics.com support@borealgenomics.com
Aurora Forensic Sample Clean-up Protocol 106-0008-BA-D 2015 Boreal Genomics, Inc. All rights reserved. All trademarks are property of their owners. http://www.borealgenomics.com support@borealgenomics.com
Nucleic Acid Techniques in Bacterial Systematics
 Nucleic Acid Techniques in Bacterial Systematics Edited by Erko Stackebrandt Department of Microbiology University of Queensland St Lucia, Australia and Michael Goodfellow Department of Microbiology University
Nucleic Acid Techniques in Bacterial Systematics Edited by Erko Stackebrandt Department of Microbiology University of Queensland St Lucia, Australia and Michael Goodfellow Department of Microbiology University
Aviva Systems Biology
 Aviva Custom Antibody Service and Price Mouse Monoclonal Antibody Service Package Number Description Package Contents Time Price Customer provides antigen protein $6,174 Monoclonal package1 (From protein
Aviva Custom Antibody Service and Price Mouse Monoclonal Antibody Service Package Number Description Package Contents Time Price Customer provides antigen protein $6,174 Monoclonal package1 (From protein
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
A role for TFIIH in controlling the activity of early RNA polymerase II elongation complexes
 Proc. Natl. Acad. Sci. USA Vol. 94, pp. 9006 9010, August 1997 Biochemistry A role for TFIIH in controlling the activity of early RNA polymerase II elongation complexes ARIK DVIR*, RONALD C. CONAWAY*,
Proc. Natl. Acad. Sci. USA Vol. 94, pp. 9006 9010, August 1997 Biochemistry A role for TFIIH in controlling the activity of early RNA polymerase II elongation complexes ARIK DVIR*, RONALD C. CONAWAY*,
Design of conditional gene targeting vectors - a recombineering approach
 Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
The fastest spin-column based procedure for purifying up to 10 mg of ultra-pure endotoxin-free transfection-grade plasmid DNA.
 INSTRUCTION MANUAL ZymoPURE Plasmid Gigaprep Kit Catalog Nos. D4204 (Patent Pending) Highlights The fastest spin-column based procedure for purifying up to 10 mg of ultra-pure endotoxin-free transfection-grade
INSTRUCTION MANUAL ZymoPURE Plasmid Gigaprep Kit Catalog Nos. D4204 (Patent Pending) Highlights The fastest spin-column based procedure for purifying up to 10 mg of ultra-pure endotoxin-free transfection-grade
