The Medical Exploration Toolkit - An efficient support for visual computing in surgical planning and training
|
|
|
- Evan Turner
- 8 years ago
- Views:
Transcription
1 Nr.: FIN The Medical Exploration Toolkit - An efficient support for visual computing in surgical planning and training Konrad Mühler, Christian Tietjen, Felix Ritter, Bernhard Preim Arbeitsgruppe Visualisierung (ISG) Fakultät für Informatik Otto-von-Guericke-Universität Magdeburg
2 Impressum ( 10 MDStV): Herausgeber: Otto-von-Guericke-Universität Magdeburg Fakultät für Informatik Der Dekan Verantwortlich für diese Ausgabe: Otto-von-Guericke-Universität Magdeburg Fakultät für Informatik Konrad Mühler Postfach Magdeburg muehler@isg.cs.uni-magdeburg.de Auflage: 50 Redaktionsschluss: Juni 2008 Herstellung: Dezernat Allgemeine Angelegenheiten, Sachgebiet Reproduktion Bezug: Universitätsbibliothek/Hochschulschriften- und Tauschstelle
3 The Medical Exploration Toolkit An efficient support for visual computing in surgical planning and training Konrad Mühler, Christian Tietjen, Felix Ritter, and Bernhard Preim Abstract Application development is often guided by the usage of software libraries and toolkits. For medical applications, the currently available toolkits focus on image analysis and volume rendering. Advanced interactive visualizations and user interface issues are not adequately supported. Hence, we present a toolkit for application development in the field of medical intervention planning, training and presentation the MEDICALEXPLORATIONTOOLKIT (METK). The METK is based on rapid prototyping platform MeVisLab and offers a large variety of facilities for an easy and efficient application development process. We present dedicated techniques for advanced medical visualizations, exploration, standardized documentation and interface widgets for common tasks. These include, e.g., advanced animation facilities, viewpoint selection, several illustrative rendering techniques, and new techniques for object selection in 3d surface models. No extended programming skills are needed for application building, since a graphical programming approach can be used. The toolkit is freely available. Well documented interfaces facilitate the extension of the toolkit. 1 INTRODUCTION Software assistants for intervention planning, e.g., for surgery, interventional radiology or radiation treatment planning are a relatively recent development. Surgical applications have special demands on visualization and interaction. It is not sufficient to display and analyze slice data and to create volume-rendered images. Instead, an indepth analysis of the image data needs to be supported with appropriate 3D interaction techniques and advanced visualization techniques. With the MEDICALEXPLORATIONTOOLKIT (METK), we present a widely applicable library for application development that closes the gap between image analysis, processing, and basic visualizations on the one hand, and the surgical needs concerning visualization and interaction on the other hand. The METK is based on the image processing and development environment MeVisLab [MeVisLab, 2008] [Rexilius et al., 2006]. Visual computing in surgical applications requires to provide comprehensive patient related information, including visualizations of the relevant anatomic and pathologic structures, and enabling a faithful representation of the area around the pathologies. Moreover, measurements, annotations, resection lines and other information may be important to directly support pre-interventional decisions. On the one hand, flexible visualization and interaction is needed to cope with the peculiarities of individual cases, but on the other hand, strong guidance is desirable to avoid that surgeons are overwhelmed by the facilities. Most of these information, e.g., measurements of the structure s extent can only be derived after the segmentation of relevant structures. Visualizations based on pre-segmented structures are mandatory for operation planning in many fields, due to a high density of soft tissue structures with overlapping image intensity values. Thus, operation planning in the abdominal region (e.g., liver, pancreas or kidney), the neck region, and orthopedic interventions is preferably performed using isosurfaces in combination with the original 2d slices, while in neuro surgery or in emergency cases still volume rendering of the original image data is preferred. Therefore, the METK development focuses on surface-based visualizations of segmented structures. The METK does not support the segmentation process, since many established segmentation techniques are included in the underlying MeVisLab. Furthermore, several applications (e.g., HepaVision for liver surgery [Bourquain et al., 2002] or Neck- Vision for neck surgery [Cordes et al., 2006]) and service providers [MeVis Medical Solutions - Distant Services, 2008] are available to perform this task. The METK can import segmentation masks as well Konrad Mühler, Christian Tietjen, and Bernhard Preim, University of Magdeburg, muehler@isg.cs.uni-magdeburg.de Felix Ritter, MeVis Research, felix.ritter@mevis.de as surface nets of structures (e.g., in Open Inventor or STL format). Open source and freely available toolkits are widespread in the research application development. Using toolkits, application prototypes can be build up quickly, reverting on ready-to-use basic functions. In the medical domain, toolkits and libraries for image analysis and volume rendering are widely available [Caban et al., 2007], e.g., the MITK [Wolf et al., 2005], 3dSlicer [Pieper et al., 2004] or VolumeShop [Bruckner and Groeller, 2005]. However, they are difficult to program, since substantial C++ knowledge is required. The METK supports an easier application building process for surgical applications, where no extended programming skills are needed, since graphical programming in combination with script-based interface design and many visualization and interaction techniques (like automatic viewpoint selection and animations) are employed. The METK is a turn-key environment because all described functions are fully implemented and basic applications in form of example networks and datasets are provided with the METK. The METK provides capabilities for application developers as well as techniques that directly support the end user of such applications the surgeon. The METK differs from other toolkits in the following ways: 1. Providing a large variety of relatively recent visualization and interaction techniques as well as interface widgets and templates for surgical operation planning. 2. Faster building of ready-to-use applications with more user friendly interfaces than with most other available platforms, while no extended programming skills are needed. 3. The open source and well documented data interface enables developers to easily extend the METK by new modules. Outline In Section 2 we present different surgical application scenarios and derive requirements for surgical applications concerning visualization, interaction, case management, and interface design. In Section 3, we review related toolkits and work. In Section 4, we give an overview of the key concepts of the METK and present the techniques that we integrated in the METK. Furthermore, details with respect to new techniques for 2d overlay visualizations, of advanced facilities for selecting objects in a scene with multiple semi-transparent objects, and of facilities to reuse once defined visualization parameters are presented. In Section 5 we describe how application building is achieved using the METK and present some applications where the METK has been successfully applied. In Section 6 we close with a summary and a view on future developments.
4 2 REQUIREMENT ANALYSIS Our development is guided by experiences in several fields of surgical application development. Thus, we will present selected application scenarios and derive requirements for surgical applications and their development in this section. Scenario 1: Neck dissections must be planned carefully. They are carried out for patients with malignant tumors in the neck or head region to remove lymph node metastases. Depending on the broadening of enlarged lymph nodes, only a few of them or a large part of the neck including muscles must be resected. A lot of structures has to be taken into account (e.g., vessels, muscles and up to 60 lymph nodes). The surgeons must explore the distances of all larger lymph nodes to vital structures of risk in order to judge if they are infiltrating vitally important structures or if there is enough space to safely resect them individually [Krueger et al., 2005]. Scenario 2: One application scenario in abdominal surgery is the resection of tumors in the liver, kidney or pancreas which are rather similar with respect to the demands on software support. Here, a tumor or several metastases need to be resected out of the liver volume with a specific safety margin. In difficult cases, this intervention requires an intensive and detailed computer-based operation planning process. The tumor and especially the surrounding vessels must be inspected carefully in 2d as well as in 3d. The remaining liver volume must be calculated with respect to vessel supply and drainage and under supplied tissue needs to be adequately visualized in 3d as well as in 2d [Preim et al., 2000]. Scenario 3: In spine surgery, small changes of the spine s anatomy can evoke symptomatic disorders for the patient. Hence, the spine surgeon must inspect the spatial relations between nerval and spinal structures as well as the relation of the spine to surrounding muscles, vessels and glands. The surgeons need to place virtual needles and implants in the spine region to plan different strategies for the access route later in the intervention. Dedicated 3d visualizations can help the planning surgeons to find such access routes without injuring important structures. For all scenarios, the exploration of the dataset must be as fast as possible in the clinical routine. Applications for presentations are of utmost importance for collaborative intervention planning like tumor board discussions, where a complex case is presented by one medical doctor to initiate an interdisciplinary discussion to finally come to treatment decisions. Due to the importance of such interventions for patients, their consultation is essential and special applications for demonstration purposes are crucial. In general, surgeons are medical experts, usually with only modest computer experience. They benefit from faithful spatial renditions of the patient s individual anatomy, but they usually have no special abilities to explore and handle 2d and 3d data. From our experience with surgical applications, we derive basic requirements for such applications: 1. Surgical applications must primarily support the surgeon s decision-making process. 2. Measurement capabilities must be provided to support, e.g., distance, volume, and angle measurements, since these measurements are often closely related to surgical decisions. 3. Due to the importance of 2d slice data, 2d and 3d views of data should be coherent and synchronized, while the exploration of 3d data must be supported in particular. 4. Important structures need to be emphasized, preserving the context. 5. Dedicated techniques for special surgical fields should be provided (e.g., resection techniques for abdominal surgery, DTI visualization for neuro surgery, and multi-modal data visualization for cardio surgery). 6. Since surgeons use computers only occasionally, they dislike complex interfaces [Cordes et al., 2007]. Therefore, Commercial companies provide applications with carefully designed, highly visual user interfaces. Thus, pleasant user interfaces should be provided. Besides those requirements that an end user application must fulfill, special requirements are necessary to support an efficient development process: Ready-to-use applications should be created quickly. Thereby, essential feedback of end users can be obtained in an early development stage. The application building process should not require extended programming, like implementing a lot of classes for application behavior and interfaces in C++. Existing applications as well as new applications should be able to be extended by new techniques, e.g., for special surgical requirements, obtained from feedback or techniques from newer visualization developments. Even though there are many similarities of surgical applications, every field of use has its special needs. Nearly all surgical applications need an efficient case management and inter-application communication, advanced 3d visualizations of segmented structures and guidance for their exploration. Measurement facilities are less important for patient s consultation, while resection techniques are primarily necessary for abdominal surgery planning. Thus, a modular approach is desirable where only required features are integrated in an application. 3 RELATED WORK Since medical visualization and medical image analysis is a very active field of research, several toolkits are available. We will discuss them in this section with respect to the presented requirements. For the fast generation of visualizations, [Bavoil et al., 2005] presented VisTrails. VisTrails is a publicly available pipeline-based environment, where visualizations for many fields of use, like time-varying data or diffusion tensor, can be created. The system also provides the ability to reuse several parts of previous visualization pipelines using visualization by analogy [Scheidegger et al., 2007]. VisTrails focuses on the generation and exploration of singular datasets and does not support an application building process quite well. VolumeShop [Bruckner and Groeller, 2005] is a standalone prototype for the interactive direct volume illustration of single datasets. Providing impressive facilities to create illustrations, it is very useful for presentation purposes, where an artist creates singular aesthetic visualizations. VolumeShop also enables the testing and development of new visualization techniques, because it is an open source system. But extensive programming skills are needed. The main drawback is the lack of support to build applications based on the presented visualization and interaction techniques. The 3dSlicer [Pieper et al., 2004] was used in a variety of research applications, mainly in the field of neuro imaging. As the name implies, the focus of the 3dSlicer lies on slicing 2d volume data. It only provides some basic 3d visualization techniques. The Medical Imaging Interaction Toolkit (MITK) [Wolf et al., 2005] is a C++ framework that is build up on ITK and VTK. It focuses on image analysis algorithms and interaction support for the segmentation and registration of medical images. The developers themselves state the MITK not intended to be an application framework. JULIUS[Jansen et al., 2001] is a software framework that consists of a core application which can be extended by plug-ins. It focuses on the medical image analysis and provides algorithms for segmentation, registration and intra-operative navigation. For visualizations VTK is used, while no special visualizations for surgical applications were mentioned.
5 A C++ toolkit specializing in intra-operative support is the Image- Guided Surgery Toolkit (IGSTK) [Enquobahrie et al., 2007]. It supports the development of applications for interventional radiology procedures and image-guided surgery, where external tracking devices are applied. A closed framework for the visualization of multi-modal medical images is presented by Manssour et al. [Manssour et al., 2001]. They focus on segmentation and registration, but do not demonstrate how individual applications can be created with their framework. Another framework for medical image analysis is CAVASS [Grevera et al., 2007]. It provides simple volume rendering and isosurface visualizations, while there is no dedicated support for application development, since CANVASS is a fixed system that is only extendable by its open source interface. A recent development environment for fast visualization prototyping is the DEVIDE system [Botha and Post, 2008], which provides substantial capabilities in accessing and changing code and underlying dataflow networks during runtime. DEVIDE supports the development of new visualization and segmentation techniques well. SciRun [Weinstein et al., 2005] is a problem solving environment that is based on dataflow networks consisting of single modules, e.g., for scalar and vector visualization, simulation, and image processing. Also end user applications can be created, called PowerApps. Amira [Stalling et al., 2005] is a commercially available objectoriented extensible toolkit for scientific visualization. It provides a wide range of analysis, simulation and visualization techniques. Amira offers a visual programming approach where the dataflow metaphor is used to connect several modules to a network. Even though there are scripting and rudimental GUI facilities available in Amira, one main drawback is the lack of support for application development. Using Amira, no end user applications can be created. MeVisLab [Rexilius et al., 2006] is a freely available rapid prototyping platform with many similarities to Amira and a broad overlapping of functionality (see Bitter et al. [Bitter et al., 2007] for a comparison of MeVisLab, Amira, SciRun and MITK), since all offer visual programming and Amira and MeVisLab use Open Inventor. However, MeVisLab focuses on the medical domain and provides facilities for application development, like higher definition language to design application interfaces. ITK and VTK are also integrated in MeVis- Lab and a memory efficient way of handling large image data volumes is provided. Using a demand-driven scheduling, all calculations and time and memory expensive operations are only performed for the part of the data that is currently needed or visible. MeVisLab focuses on medical image analysis and provides an extensive library for those purposes. However, the provided features aim at handling single images and objects, but have no special capabilities to handle whole cases. The METK reuses virtually everything from MeVisLab, including the MeVis library, the graphical network development paradigm and the efficient memory management for handling large volume datasets. In addition to modular toolkits for individual setups, there are dedicated commercial volume rendering systems, such as Barco s VOXAR3D [Barco, 2008], the VolumeGraphicsLibrary [Volume Graphics, 2008] or Kitware s VolView [Kitware, 2008]. In essence, there are many supportive toolkits and frameworks for medical image analysis and visualization. But some of them are focused on the creation of singular impressive visualizations (e.g., VisTrails [Bavoil et al., 2005], VolumeShop [Bruckner and Groeller, 2005]), some of them focusing on medical image analysis (e.g., MITK [Wolf et al., 2005], 3dSlicer [Pieper et al., 2004]), and only some of them allowing to build up own standalone applications (e.g., SciRun [Weinstein et al., 2005], MeVis- Lab [Rexilius et al., 2006]). To the best of our knowledge there is no toolkit or framework to create efficient medical applications with highend visualizations, adequate interaction techniques and user interfaces guidance. A very short and preliminary report on the METK presented here will be presented as a software demo on a national workshop [Tietjen et al., 2008]. Techniques which have only mentioned in the short workshop paper are described here in detail for the first time. 4 THE ARCHITECTURE OF THE METK In the MEDICALEXPLORATIONTOOLKIT each function is encapsulated in a module. Using MeVisLab s visual programming environment, modules can be freely combined in a dataflow network to build up individual applications with an individual feature profile. This allows the developer to design applications that support the individual workflow of different surgical intervention planning processes in an efficient and fast manner. All functions are organized in three layers: the data management and communication layer, the visualization layer, and the exploration layer (see Figure 1). The lowest layer imports the case data and provides data management functions. The visualization layer comprises viewer classes and special rendering modules, whereas basic viewer classes and the volume rendering are reused from MeVisLab. All highend interaction and exploration techniques that are necessary to create powerful surgical applications are available from the exploration layer. The layers and their provided functions will be described in the next sections. Key states and Undo Facilities Widgets and Layout Templates Viewer Volume Rendering Data Management and Communication Layer Smooth Isosurfaces and Vessel Visualizations MRI and CT Images Segmentation Masks of Structrues Exploration Layer Object Selection and Fast Object Manipulation Animations Visualization Layer Colored 2d Overlays using MCSM Illustrative Visualizations Measurement Tools Viewpoint Selection Lift Chart Colored Isosurfaces Multi Coded Segmentation Masks Basic MeVisLab Functions Fig. 1. Layer structure of the METK. A large variety of different visualization and interaction techniques as well as case management facilities and user interface widgets can be freely combined to build up individual surgical applications.
6 4.1 Data Management and Communication Layer The database layer contains functions for case data management, interapplication communication and 3d isosurfaces generation. The METK can load and handle a wide range of medical CT and MRI image formats, including RAW and DICOM. Additionally, segmentation masks can be loaded to automatically generate 3d isosurfaces. This operation must be performed only once, since the isosurfaces are stored for further loadings of a case. Depending on the type of structures, different algorithms for generating the isosurfaces are used. In most cases, marching cube in combination with a constrained elastic surface smoothing, which was determined to be the best algorithm for most medical structures [Bade et al., 2007], is applied. For vessels we use a model-based surface reconstruction that respects the thin and branching structure of vessel trees [Hahn et al., 2001]. Besides segmentation masks, also surface nets of structures and secondary objects (like medical probes) can be imported. The METK provides facilities to convert cases segmented with other tools like ITK. Case and cache management. To reduce the memory requirements and to speed up the whole process of loading and exploring a case, we integrated an efficient case management in the METK. Each structure and each image stack is only loaded once and distributed virtually in the application network. Even if the structures are visualized with different techniques in different viewers at the same time or if an image stack is browsed in a 2d viewer and a 3d viewer simultaneously (see Figure 8), it will only be maintained once in the application cache. Communication and synchronization. Besides the data management, the METK provides a communication structure between all modules. Events can be sent between specific modules for a direct inter-module communication and be broadcasted to reach all modules. Therefore, all changes of underlying data and parameters are communicated to all modules listening to those parameters, so they can adjust their own parameters, data, and visualizations. This leads to identical visual properties of all structures in all viewers and widgets and thus to a consistent view of all data. Another information, communicated automatically in the METK, is the currently selected structure. Hence, a synchronized view in different viewers can be provided. If the user selects a structure in a 3d viewer, all 2d viewers can display the suitable slice for this structure and vice versa. If the user picks a structure in a 2d slice, it can be emphasized in all 3d viewers, moving the camera automatically to a good viewpoint on this structure (see Section 4.3). The synchronization of parameters also enables the usage of synchronized 3d viewers. Different 3d viewers can be automatically synchronized in the METK by synchronizing its camera parameters (position, orientation etc.). This can be used to explore different datasets from the same viewpoints, to compare different intervention strategies for one patient or to compare pre- and post-operative data. Multi-coded segmentations. Usually, each segmentation mask, representing one structure, is stored in a single file. This is inefficient in case of memory and performance. Storing all segmentation masks in one image stack can overcome this problem. However, one voxel of an image may belong to more than one structure, when structures overlap each other. For example a voxel in the liver tissue may belong to a tumor and the liver tissue itself. Thus, we cannot assign one label to each structure for the resulting segmentation mask of all structures. A straightforward approach is to assign each structure to one bit of a 8 byte voxel value. But this approach is limited by the number of bits: in our example, only 64 labels could be stored in a 8 byte value. Since in real data, only a small subset of all possible combinations of overlapping structures occur, we developed a more efficient solution: the multi-coded segmentation masks (MCSM). An MCSM contains all segmentation masks of all structures of a case. Each combination of labels of a voxel that appears in the data is encoded with a distinct voxel value (see Figure 2). For example all voxels simultaneously belonging to the liver tissue and the hepatic vein (and to no other structure) are assigned one unique voxel value. The mapping of voxel values to structure lists is stored separately in the case data. An MCSM is created by sequentially adding one segmentation mask after another. If a new combination of voxel labels occurs Liver Liver, Tumor Liver, Hepatic Artery Hepatic Artery Liver, Hepatic Vein Liver, Tumor, Hepatic Vein Fig. 2. Multi-Coded segmentation masks. Segmentation masks of single structures are sequently added to the MCSM. For each structure all values of voxels that belongs to the structure are stored separately. Thus, many segmentation masks of overlapping structures can be saved in one MCSM. at a voxel position, a new number is assigned to this combination. After all single segmentation masks were added to the MCSM, it can be used e.g., as an efficient base for colored overlays. The upper bound of 2 64 labels will never be reached with medical datasets, since even the theoretical case that in a dataset of voxels each voxel represents another combination of structures is covered by the MCSM. 4.2 Visualization Layer In the visualization layer, all actions are performed that are necessary to provide basic and advanced 3d visualization techniques to the user. Based on the isosurfaces stored in the cache, a material is assigned to each structure to achieve appealing isosurface visualizations. Important structures are visualized with a high opacity or with a silhouette. Thus, they are still visible as context structures, but do not hide the view onto important structures. For volume rendering the METK employs the GigaVoxelRenderer [Link et al., 2006], which is integrated in MeVisLab. It enables the tagged volume rendering of segmented structures. Thus, different structures can be visualized with different transfer functions. By reason of consistency, the colors of structures are the same as in its isosurface visualization. To visualize unsegmented tissue, a global transfer function can be applied and the volume rendering can be combined with the isosurface visualizations. Advanced 2d visualizations. The basic problem of the slice-based visualization, namely the lack of an overview in cross-sectional images, has been tackled with a 2.5D approach to provide the essential information, the LIFTCHART [Tietjen et al., 2006a]. A narrow frame attached next to the cross sectional image represents the overall extent of slices in the volume dataset. The top and bottom boundary of the frame correspond to the top and bottom slice of the dataset (see Figure 3). Each segmented structure is displayed as a bar at the equivalent vertical position inside this frame. Upper bars correspond to higher structures in the body. Different arrangements of the bars are possible, e.g., condensing all structures of the same type into one column. The currently displayed slice of the volume dataset is depicted by a horizontal line in the LIFTCHART widget (see Figure 3). To support the correlation between structures in 3d scenes and 2d slices, structures can be visualized in 2d slice data as colored and semitransparent overlays, so the underlying gray values are still visible. If more than one structure should be displayed at the same voxel position, the combined color can be calculated in different ways: 1. Only the color of the most important structure is chosen. 2. A weighted mixture of all colors of the overlapping structures is calculated.
7 (a) Fig. 3. LIFTCHART. 3(a) LIFTCHART in a 2d viewer and the corresponding dataset in 3d 3(b). The location of different structures in the slice stack can be identified by their color. Selecting a structure in the LIFTCHART selects the corresponding slice in the viewer. Furthermore, safety margins are depicted with red and yellow. 3. Application-dependent overlapping regions can be emphasized separately in dependency of involved structures, e.g., the infiltration of lymph nodes in a muscle can be colored red with a silhouette, even if this visualization style does not appear in one of the two structures. The calculation of the overlays is performed based on our multicoded segmentation masks. Safety margins are essential for the intervention planning and intraoperative navigation. Therefore for all structures at risk, a 3d Euclidean distance transform is performed. Depicting important distance thresholds (e.g., yellow and red) turned out to be appropriate. The distances may be displayed in 2d as well as in 3d. In the 2d view, silhouette lines as halos are drawn at the important distances. In 3d, unicolored surfaces are drawn on structures visualizing the range of distance to critical structures, e.g., lymph nodes to vessels and muscles (see Figure 3(b)). Illustrative visualizations. As known from many anatomic books, context structures are mostly drawn stippled or with a silhouette. Those techniques were adapted for medical structures [Baer et al., 2007] and we provide it in the METK. The visualization layer also provides several viewers. 2d viewers can display slices in many ways: singular or in a multi-slice view, where axial, sagittal, and coronal as well as free multi planar reformations can be shown. 3d and 2d viewers can be freely combined in an arbitrary number and arrangement. 4.3 Exploration Layer To support the exploration process, we provide several techniques, interaction facilities and interface widgets. Animation. To guide the user as well as to provide smooth transitions between different viewpoints and visualization styles, we integrated the animation framework developed by [Muehler et al., 2006]. Using an adaptive script language, one script with an animation description can be reused for many similar cases in an application, providing that segmentations results are stored and named in a standardized way. The script-based animations can also be used in an interactive application to guide the user s exploration process. Selected structures can be approached due to automatic camera flights, and appearance changes can be smoothly translated to preserve the user s orientation. The flexibility of the animation facilities enables the usage of advanced animation techniques like story telling principles [Wohlfart and Hauser, 2007]. Thus, applications based on the METK can provide both interactive animations in real time for exploration support, and rendered videos for presentation and interdisciplinary discussions like in a tumor board. Viewpoint selection. Animations are enhanced by a dedicated viewpoint selection technique for multi-object 3d scenes [Muehler et al., 2007] that has been applied in several intervention planning tasks. Finding good views on single structures or groups (b) of structures is essential for an automatic guided exploration. After selecting a structure, the camera position is transformed automatically to a good view on this structure. The quality of a viewpoint is affected by many parameters. The structure should be visible to a maximum extend. A good viewpoint should be stable, i.e., minor rotations must not completely hide the selected structure. Medical doctors of different medical professions have different preferred regions to look at a 3d scene of segmented structures. Thus, the preferred region is also an important parameter for viewpoint estimation. These and other parameters are considered by our viewpoint selection technique. As discussed in [Muehler et al., 2007], relatively good presets for certain application scenarios may be defined. Thus, no further justification is necessary during the usage. Good viewpoints are employed under several circumstances in the METK. They are used to generate standardized views for documentation (in combination with other standardized visualization parameters). If the user picks a structure from a list or from the viewer, the camera can be automatically moved to the best viewpoint of the structure. Camera paths. To produce appealing and non-destructive camera paths by moving the camera from one viewpoint to another, we developed and integrated a set of different path algorithms in the METK. To preserve the orientation on long distance movements of the camera, we zoom out to a global view on the scene in the first part of the panning process and zoom into the target structure at the end of the flight. We also make camera movements more appealing by slow acceleration at the beginning and the end instead of abrupt speed changes. Measurement tools [Preim et al., 2002] are also integrated, e.g., for distances measurements and its appropriate visualization by means of arrows. The proposed measurement tools are extended by automatic measurement facilities for computing minimal distances between two structures and by calculation of a structure s volume. Key states. For presentation purposes, interdisciplinary discussions, or patient consultation several views and visualizations of the explored data needs to be saved. Instead of only saving screenshots, we introduce the concept of key states. In a key state, all information about a scene and its visualization is stored. This includes camera parameters as well as visualization properties. Thus, a complete state of a visualization can be restored later for further explorations or demonstrations. Since key states are stored in the case data, they can be transfered from one application (e.g., a surgical planning software) to another (e.g., an application for patient consultation). Naturally, key states can also be exported as screenshots for usage in documents or presentations. Usually, a surgeon creates a couple of key states during a planning process (see Figure 4). In combination with our animation facilities, videos can be created automatically from a set of key states, where smooth transitions between the key states are computed automatically. Those videos are used to teach other surgeons for example. Key states can also be used to define presets. Applying a once defined key state to a new case with similar structures, those structures are visualized with the same properties as they where stored in the key state. Undo. One important feature, especially for surgeons who are inexperienced in 3d exploration and mouse-based navigation, is an undo function for 3d scene manipulations. In the METK, after every performed action (e.g., a camera movement or a visualization change) the state of the whole scene is stored in a key state. Changes performed in a very narrow time range (e.g., automatic changes of the visualization) are combined in one key state. The user can return to arbitrary steps. In combination with the animation facilities of the METK, from a set of (undo) key states videos can be generated automatically. Those videos can be used for demonstration purposes or for the illustration of the exploration process for educational purposes. Object selection. We provide several new techniques to select objects in 3d scenes with many objects of different transparencies. In such scenes the selection is ambiguous, if there is more than one object in the picking ray. The simplest approach to disambiguate the selection is always to select the first object in the pick ray. However, if this object has a high transparency, the user probably wants to select
8 Title: Overview Comment: Liver tissue with portal veins and suspected metastases Title: Liver territories Comment: Affected territories for two central metastases Title: Resection volume (a) Fig. 5. Object selection. In 5(a) the selection of inner structures like vessels is made possible in the first place, while in 5(b) the transparent oesophagus in front of the opaque spine cannot be selected automatically. (b) Comment: Suggested resection volume for one metastasis Fig. 4. Key states. Several key states that were created for planning a liver operation. The surgeon stored the key states together with a title and a short comment. another object behind. In complex medical scenes, some objects are completely enclosed by others, so they are never the first object in the pick ray. For example in liver surgery, the liver tissue always encloses nearly all other structures like vessels and tumors (see Figure 5(a)). We developed a procedure to automatically select an object, after the user clicked on the scene. It is assumed that the user points the mouse consciously. That means, when the mouse cursor is placed over a very large and a very small object, he or she placed it deliberately over the small object. Furthermore, it is assumed that the perception of very transparent objects is less than the perception of more opaque objects. To identify the desired object, the algorithm proceeds as follows: All objects hit by the pick ray are determined and sorted by the depth-distance of their intersection point. Only objects, which are locally visible by at least 10% are taken into account. As a next step, the size of the projected bounding box on the viewport of all objects and the transparency degree of the single objects is determined. The impact of transparency and the projected bounding box size are adjustable, to consider different types of scenes (e.g., scenes with objects of rather equal size). The object with the highest rating is selected at the end. Thus, for example opaque structures behind structures with a high transparency are selected. The algorithm reveals its limitations when a transparent structure is totally underlaid by an opaque structure with a similar size (see Figure 5(b)). In such cases, at all points the user picks the transparent sturcture, the opaque one in the background is selected. Therefore, we provide two interaction techniques in the METK. The first allows the user to scroll between all structures in the pick ray, using the mouse wheel or the cursor keys. Starting with the most presumably structure the user can scroll stepwise back and forth. The currently selected structure will be clearly emphasized, using a large silhouette and an opaque color. The second selection technique offers a list of all structures in the pick ray directly at the cursor position, like a popup menu, so the user does not need to defocus from the scene. We extend the list of textual structure names by pictorial representations of the structures. However, this technique aims at experienced users who know all structures by name (see Figure 6(a)). Object manipulation. Even if users adjust the appearance of a visualization globally by selecting a preset, they might want to adapt the appearance of single structures individually. We provide some GUI widgets that can be integrated in a panel or window to adjust all visualization parameters like color, transparency or silhouette width. But we also provide an exploration technique, where the user can easily (a) Fig. 6. Fast exploration popup menus. 6(a) All structures in the pick ray are presented as a fast accessible list to the user. 6(b) For quick parameter manipulation, a context popup is presented right to the cursor position. adjust the most important parameters directly in the viewer. By right clicking on a structure, a popup menu appears at the cursor s position in the focus of the user and provides a slider for transparency, a color selector as well as the possibility to hide the picked structure. The provided list of parameters can be adapted and extended for individual application requirements. In some cases it might be useful to provide a button to request further information of the structure or a button to add this structure to a list (see Figure 6(b)). Graphical application interface. For a fast and efficient application development, predefined widgets for common and recurrent tasks are provided, e.g., lists to select structures or to change their visibility. We provide panels to change the visualization parameters of structures like color, transparency or silhouette width, efficiently. Feedback from surgeons using our applications clearly revealed that a rather low level of flexibility is needed and guidance is considered essential. Surgeons want clear and easy to understand interfaces [Cordes et al., 2007] instead of interfaces, overloaded with parameter sliders and value inputs. They want to get the best visualization for the current task or medical question automatically or at the utmost selecting a well defined preset from a small list of choices. We support this, e.g., with our key states and the animation facilities. Furthermore, we also provide ready designed sample interfaces as templates for surgical applications that were approved by evaluation [Cordes et al., 2007] and by many interviews with our medical partners. 5 APPLICATION DEVELOPMENT WITH THE METK Since the METK is an extension of MeVisLab, all METK applications can be build up by creating networks of modules in a visual programming environment. The interfaces are defined script-based. Thus, no extended programming skills are needed to create applications using (b)
9 Fig. 8. NeckSurgeryPlanner. The NECKSURGERYPLANNER is an application to support the operation planning for neck dissections. To provide deep insight in the original 2d data as well as in the segmented 3d structures, 3d and 2d views are used synchronized. On the left, a browser for enabling and disabling structures and key state previews are provided. Fig. 9. LiverSurgeryTrainer. The LIVERSURGERYTRAINER is an application to teach abdominal surgeons the planning workflow for liver resection and living liver donor transplantations. The application layout contains only a few widgets. the METK features. This quickly yields to applications that a surgeon can use (see Figure 7). Our experiences show that important feedback is given by surgeons only after they tried first functional prototypes. Thus, this stage should be achieved fast. To extend the functionalities of the METK or to supplement existing functions, there are basically two options for developers: They can implement simple functions, like a patient data management or widget panels in modules, written in the script language Python. Advanced and especially performance critical issues can be implemented in C++ libraries. Since this is, for example, only necessary for new visualization techniques like DTI visualization, this does not contradict the supposed low programming skills that are needed to build up readyto-use applications with the METK. Several full-fledged applications were designed with our toolkit. Using the METK, a training system for liver surgeons was developed, the LIVERSURGERYTRAINER [Bade et al., 2006] (see Figure 9). The LIVERSURGERYTRAINER was evaluated in a comprehensive evaluation [Cordes et al., 2007]. The feedback from the surgeons inspired several refinements of the METK. For neck surgery planning the NECKSURGERYPLANNER was developed [Tietjen et al., 2006b] (see Figure 8). It supports the decision-making process for neck dissections. 6 CONCLUSION AND FUTURE WORK We presented an extensive toolkit for surgical application development the MEDICALEXPLORATIONTOOLKIT. It was shown that, using the METK, applications that fulfill surgical requirements of exploration support and visualization techniques, can be build up quickly. Introducing the multi-coded segmentation masks, we provide an efficient way to store multiple overlapping segmentation masks in one mask, supporting colored overlays in 2d. With our animation facilities, the viewpoint selection as well as our new support for object selection, we provide a substantial guidance for the exploration of 3d scenes. Currently, we are working on bridging the gap between preoperative planning software and intra-operative usage of the planning results. We plan to adapt selected techniques like the automatic viewpoint selection for an intra-operative use, since there more guidance in 3d exploration is needed, due to the special surrounding. Our experiences with automatic techniques like the object and viewpoint selection showed that more semantic information about the importance of structures and their relations into the visualizations needs to be integrated. Therewith, the presented context to structures of interest or the intended user interactions can be adjusted in a more appropriate manner. Due to the open structure and the free availability of the METK, many existing visualization and interaction techniques can be integrated, e.g., more sophisticated vessel visualizations [Schumann et al., 2007]. The METK does currently not provide special techniques for neuro surgery like DTI or for cardio vascular diagnostics like multi-modal visualizations. One aim for the future is to detach the functionalities for animation, viewpoint selection and illustrative visualization to offer them to a wider community and other fields beyond surgical applications. REFERENCES [Bade et al., 2007] Bade, R., Konrad, O., and Preim, B. (2007). Reducing Artifacts in Surface Meshes Extracted from Binary Volumes. Journal of WSCG, 15: [Bade et al., 2006] Bade, R., Riedel, I., Schmidt, L., Oldhafer, K. J., and Preim, B. (2006). Combining Training and Computer-assisted Planning of Oncologic Liver Surgery. In Bildverarbeitung fuer die Medizin 2006 (BVM 2006), pages [Baer et al., 2007] Baer, A., Tietjen, C., Bade, R., and Preim, B. (2007). Hardware-Accelerated Stippling of Surfaces Derived from Medical Volume Data. In Museth, K., Moeller, T., and Ynnerman, A., editors, IEEE/Eurographics Symposium on Visualization (EuroVis), pages [Barco, 2008] Barco (2008). [Bavoil et al., 2005] Bavoil, L., Callahan, S. P., Crossno, P. J., Freire, J., Scheidegger, C. E., Silva, C. T., and Vo, H. T. (2005). Vistrails: enabling interactive multiple-view visualizations. In Proceedings of IEEE Visualization, pages [Bitter et al., 2007] Bitter, I., Uitert, R. V., Wolf, I., Ibanez, L., and Kuhnigk, J.-M. (2007). Comparison of Four Freely Available Frameworks for Image Processing and Visualization That Use ITK. Transactions on Visualization and Computer Graphics, 13(3): [Botha and Post, 2008] Botha, C. P. and Post, F. H. (2008). Hybrid Scheduling in the DeVIDE Dataflow Visualisation Environment. In Hauser, H., Strassburger, S., and Theisel, H., editors, Proceedings of Simulation and Visualization, pages SCS Publishing House Erlangen. [Bourquain et al., 2002] Bourquain, H., Schenk, A., Link, F., Preim, B., Prause, G., and Peitgen, H.-O. (2002). HepaVision2: A software assistant for preoperative planning in living-related liver transplantation and oncologic liver surgery. In Lemke, H. U., editor, Computer Assisted Radiology and Surgery (CARS 2002), pages [Bruckner and Groeller, 2005] Bruckner, S. and Groeller, M. E. (2005). Volumeshop: An interactive system for direct volume illustration. In C. T. Silva, E. Groeller, H. R., editor, Proceedings of IEEE Visualization 2005, pages [Caban et al., 2007] Caban, J. J., Joshi, A., and Nagy, P. (2007). Rapid Development of Medical Imaging Tools with Open-Source Libraries. Journal of Digital Imaging, 20: [Cordes et al., 2006] Cordes, J., Dornheim, J., Preim, B., Hertel, I., and Strauß, G. (2006). Preoperative Segmentation of Neck CT Datasets for the Planning of Neck Dissections. In SPIE Medical Imaging. Spie Press.
10 (a) (b) Fig. 7. Sample METK networks applications.7(a) A network to present segmented datasets in a 3d viewer with a GUI widget to change visibility of structures. A spine surgery dataset is displayed.7(b) A network of an application to synchronally explore 3d and 2d data, extended with silhouettes and a colored distance transformation in 2d and 3d. A neck surgery dataset is displayed. [Cordes et al., 2007] Cordes, J., Muehler, K., Oldhafer, K., Stavrou, G., Hillert, C., and Preim, B. (2007). Evaluation of a training system of the computer-based planning of liver surgery. In Curac, pages , Karlsruhe. [Enquobahrie et al., 2007] Enquobahrie, A., Cheng, P., Gary, K., Ibanez, L., Gobbi, D., Lindseth, F., Yaniv, Z., Aylward, S., Jomier, J., and Cleary, K. (2007). The Image-Guided Surgery Toolkit IGSTK: An Open Source C++ Software Toolkit. Journal of Digital Imaging, 20: [Grevera et al., 2007] Grevera, G., Udupa, J., Odhner, D., Zhuge, Y., Souza, A., Iwanaga, T., and Mishra, S. (2007). CAVASS: A Computer-Assisted Visualization and Analysis Software System. Journal of Digital Imaging, 20: [Hahn et al., 2001] Hahn, H., Preim, B., Selle, D., and Peitgen, H.-O. (2001). Visualization and Interaction Techniques for the Exploration of Vascular Structures. [Jansen et al., 2001] Jansen, T., von Rymon-Lipinski, B., Krol, Z., Ritter, L., and Keeve, E. (2001). An Extendable Application Framework for Medical Visualization and Surgical Planning. In SPIE Medical Imaging, pages , San Diego, CA. [Kitware, 2008] Kitware (2008). VolView. products/volview.html. [Krueger et al., 2005] Krueger, A., Tietjen, C., Hintze, J., Preim, B., Hertel, I., and Strauss, G. (2005). Interactive Visualization for Neck Dissection Planning. In IEEE/Eurographics Symposium on Visualization (EuroVis), pages [Link et al., 2006] Link, F., Koenig, M., and Peitgen, H.-O. (2006). Multiresolution volume rendering with per object shading. In Vision Modelling and Visualization, pages Aka GmbH Berlin. [Manssour et al., 2001] Manssour, I. H., Furuie, S. S., Nedel, L. P., and Freitas, C. M. D. S. (2001). A framework to visualize and interact with multimodal medical images. In VG01 - Joint IEEE TCVG and Eurographics Workshop on Volume Graphics, pages [MeVis Medical Solutions - Distant Services, 2008] MeVis Medical Solutions - Distant Services (2008). Distant_Services.html. [MeVisLab, 2008] MeVisLab (2008). [Muehler et al., 2006] Muehler, K., Bade, R., and Preim, B. (2006). Adaptive script based animations for intervention planning. In Proc. of Medical Image Computing and Computer-Assisted Intervention (MICCAI), pages Springer. [Muehler et al., 2007] Muehler, K., Neugebauer, M., Tietjen, C., and Preim, B. (2007). Viewpoint Selection for Intervention Planning. In Museth, K., Moeller, T., and Ynnerman, A., editors, IEEE/Eurographics Symposium on Visualization (EuroVis), pages [Pieper et al., 2004] Pieper, S., Halle, M., and Kikinis, R. (2004). 3D Slicer. In IEEE International Symposium on Biomedical Imaging: Nano to Macro. [Preim et al., 2000] Preim, B., Selle, D., Spindler, W., Oldhafer, K. J., and Peitgen, H.-O. (2000). Interaction Techniques and Vessel Analysis for Preoperative Planning in Liver Surgery. In Medical Imaging and Computer- Assisted Intervention, MICCAI [Preim et al., 2002] Preim, B., Tietjen, C., Spindler, W., and Peitgen, H.-O. (2002). Integration of Measurement Tools in Medical Visualizations. In IEEE Visualization 2002, pages 21 28, Boston. [Rexilius et al., 2006] Rexilius, J., Kuhnigk, J.-M., Hahn, H. K., and Peitgen, H.-O. (2006). An Application Framework for Rapid Prototyping of Clinically Applicable Software Assistants. In Hochberger, C. and Liskowsky, R., editors, Informatik fuer Menschen - Band 1, pages GI-Edition - Lecture Notes in Informatics (LNI), P-93. [Scheidegger et al., 2007] Scheidegger, C., Vo, H., Koop, D., Freire, J., and Silva, C. (2007). Querying and creating visualizations by analogy. In IEEE Transactions on Visualization and Computer Graphics, volume 13, pages IEEE Educational Activities Department. [Schumann et al., 2007] Schumann, C., Oeltze, S., Bade, R., and Preim, B. (2007). Model-free Surface Visualization of Vascular Trees. In Museth, K., Moeller, T., and Ynnerman, A., editors, IEEE/Eurographics Symposium on Visualization (EuroVis), pages [Stalling et al., 2005] Stalling, D., Westerhoff, M., and Hege, H.-C. (2005). Amira: A Highly Interactive System for Visual Data Analysis. In Hansen, C. D. and Johnson, C. R., editors, The Visualization Handbook, chapter 38, pages Elsevier. [Tietjen et al., 2006a] Tietjen, C., Meyer, B., Schlechtweg, S., Preim, B., Hertel, I., and Strauß, G. (2006a). Enhancing Slice-based Visualizations of Medical Volume Data. In Santos, B. S., Ertl, T., and Joy, K., editors, EUROVIS - Eurographics /IEEE VGTC Symposium on Visualization, pages , Lisbon, Portugal. Eurographics Association. [Tietjen et al., 2008] Tietjen, C., Mühler, K., Ritter, F., Konrad, O., Hindennach, M., and Preim, B. (2008). METK - The Medical Exploration Toolkit. In Bildverarbeitung für die Medizin (BVM), pages [Tietjen et al., 2006b] Tietjen, C., Preim, B., Hertel, I., and Strauß, G. (2006b). A Software-Assistant for Pre-operative Planning and Visualization of Neck Dissections. In CURAC, pages [Volume Graphics, 2008] Volume Graphics (2008). volumegraphics.com. [Weinstein et al., 2005] Weinstein, D., Parker, S., Simpson, J., Zimmerman, K., and Jones, G. (2005). Visualization in the scirun problem-solving environment. In Hansen, C. and Johnson, C., editors, The Visualization Handbook, pages Elsevier. [Wohlfart and Hauser, 2007] Wohlfart, M. and Hauser, H. (2007). Story Telling for Presentation in Volume Visualization. In Museth, K., Moeller, T., and Ynnerman, A., editors, IEEE/Eurographics Symposium on Visualization (EuroVis), pages [Wolf et al., 2005] Wolf, I., Vetter, M., Wegner, I., Boettger, T., Nolden, M., Schoebinger, M., Hastenteufel, M., Kunert, T., and Meinzer, H.-P. (2005). The medical imaging interaction toolkit. Medical Image Analysis, 9:
Adaptive script based animations for intervention planning
 Adaptive script based animations for intervention planning Konrad Muehler, Ragnar Bade, and Bernhard Preim Department of Simulation and Graphics, Department of Computer Science, Otto-von-Guericke University
Adaptive script based animations for intervention planning Konrad Muehler, Ragnar Bade, and Bernhard Preim Department of Simulation and Graphics, Department of Computer Science, Otto-von-Guericke University
Impact of Model-based Risk Analysis for Liver Surgery Planning
 Impact of Model-based Risk Analysis for Liver Surgery Planning C. Hansen¹, S.Zidowitz 1, B. Preim 2, G. Stavrou 3, K. J. Oldhafer 3, H. K. Hahn 1 1 Fraunhofer MEVIS, Institute for Medical Image Computing,
Impact of Model-based Risk Analysis for Liver Surgery Planning C. Hansen¹, S.Zidowitz 1, B. Preim 2, G. Stavrou 3, K. J. Oldhafer 3, H. K. Hahn 1 1 Fraunhofer MEVIS, Institute for Medical Image Computing,
Computer Aided Liver Surgery Planning Based on Augmented Reality Techniques
 Computer Aided Liver Surgery Planning Based on Augmented Reality Techniques Alexander Bornik 1, Reinhard Beichel 1, Bernhard Reitinger 1, Georg Gotschuli 2, Erich Sorantin 2, Franz Leberl 1 and Milan Sonka
Computer Aided Liver Surgery Planning Based on Augmented Reality Techniques Alexander Bornik 1, Reinhard Beichel 1, Bernhard Reitinger 1, Georg Gotschuli 2, Erich Sorantin 2, Franz Leberl 1 and Milan Sonka
How To Develop An Image Guided Software Toolkit
 IGSTK: a software toolkit for image-guided surgery applications Kevin Cleary a,*, Luis Ibanez b, Sohan Ranjan a, Brian Blake c a Imaging Science and Information Systems (ISIS) Center, Department of Radiology,
IGSTK: a software toolkit for image-guided surgery applications Kevin Cleary a,*, Luis Ibanez b, Sohan Ranjan a, Brian Blake c a Imaging Science and Information Systems (ISIS) Center, Department of Radiology,
AnatomyBrowser: A Framework for Integration of Medical Information
 In Proc. First International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI 98), Cambridge, MA, 1998, pp. 720-731. AnatomyBrowser: A Framework for Integration of Medical
In Proc. First International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI 98), Cambridge, MA, 1998, pp. 720-731. AnatomyBrowser: A Framework for Integration of Medical
Twelve. Figure 12.1: 3D Curved MPR Viewer Window
 Twelve The 3D Curved MPR Viewer This Chapter describes how to visualize and reformat a 3D dataset in a Curved MPR plane: Curved Planar Reformation (CPR). The 3D Curved MPR Viewer is a window opened from
Twelve The 3D Curved MPR Viewer This Chapter describes how to visualize and reformat a 3D dataset in a Curved MPR plane: Curved Planar Reformation (CPR). The 3D Curved MPR Viewer is a window opened from
QAV-PET: A Free Software for Quantitative Analysis and Visualization of PET Images
 QAV-PET: A Free Software for Quantitative Analysis and Visualization of PET Images Brent Foster, Ulas Bagci, and Daniel J. Mollura 1 Getting Started 1.1 What is QAV-PET used for? Quantitative Analysis
QAV-PET: A Free Software for Quantitative Analysis and Visualization of PET Images Brent Foster, Ulas Bagci, and Daniel J. Mollura 1 Getting Started 1.1 What is QAV-PET used for? Quantitative Analysis
A novel Real-Time Web3D Surgical Teaching Tool based on WebGL
 A novel Real-Time Web3D Surgical Teaching Tool based on WebGL Steven Birr 1, Jeanette Mönch 1, Dirk Sommerfeld 1, Bernhard Preim 1 1 Institut für Simulation und Graphik, Otto-von-Guericke-Universität Magdeburg
A novel Real-Time Web3D Surgical Teaching Tool based on WebGL Steven Birr 1, Jeanette Mönch 1, Dirk Sommerfeld 1, Bernhard Preim 1 1 Institut für Simulation und Graphik, Otto-von-Guericke-Universität Magdeburg
3D Viewer. user's manual 10017352_2
 EN 3D Viewer user's manual 10017352_2 TABLE OF CONTENTS 1 SYSTEM REQUIREMENTS...1 2 STARTING PLANMECA 3D VIEWER...2 3 PLANMECA 3D VIEWER INTRODUCTION...3 3.1 Menu Toolbar... 4 4 EXPLORER...6 4.1 3D Volume
EN 3D Viewer user's manual 10017352_2 TABLE OF CONTENTS 1 SYSTEM REQUIREMENTS...1 2 STARTING PLANMECA 3D VIEWER...2 3 PLANMECA 3D VIEWER INTRODUCTION...3 3.1 Menu Toolbar... 4 4 EXPLORER...6 4.1 3D Volume
Virtual Resection with a Deformable Cutting Plane
 Virtual Resection with a Deformable Cutting Plane Olaf Konrad-Verse 1, Bernhard Preim 2, Arne Littmann 1 1 Center for Medical Diagnostic Systems and Visualization, Universitaetsallee 29, 28359 Bremen,
Virtual Resection with a Deformable Cutting Plane Olaf Konrad-Verse 1, Bernhard Preim 2, Arne Littmann 1 1 Center for Medical Diagnostic Systems and Visualization, Universitaetsallee 29, 28359 Bremen,
Clinical Training for Visage 7 Cardiac. Visage 7
 Clinical Training for Visage 7 Cardiac Visage 7 Overview Example Usage 3 Cardiac Workflow Examples 4 Remove Chest Wall 5 Edit Chest Wall Removal 6 Object Display Popup 7 Selecting Optimal Phase 8 Thick
Clinical Training for Visage 7 Cardiac Visage 7 Overview Example Usage 3 Cardiac Workflow Examples 4 Remove Chest Wall 5 Edit Chest Wall Removal 6 Object Display Popup 7 Selecting Optimal Phase 8 Thick
VisIt Visualization Tool
 The Center for Astrophysical Thermonuclear Flashes VisIt Visualization Tool Randy Hudson hudson@mcs.anl.gov Argonne National Laboratory Flash Center, University of Chicago An Advanced Simulation and Computing
The Center for Astrophysical Thermonuclear Flashes VisIt Visualization Tool Randy Hudson hudson@mcs.anl.gov Argonne National Laboratory Flash Center, University of Chicago An Advanced Simulation and Computing
Intro to 3D Animation Using Blender
 Intro to 3D Animation Using Blender Class Instructor: Anthony Weathersby Class Objectives A primer in the areas of 3D modeling and materials An introduction to Blender and Blender s toolset Course Introduction
Intro to 3D Animation Using Blender Class Instructor: Anthony Weathersby Class Objectives A primer in the areas of 3D modeling and materials An introduction to Blender and Blender s toolset Course Introduction
OPENNESS IN (TELE-) RADIOLOGY WORKSTATIONS: THE CHILI PLUGIN CONCEPT
 Reprint of :.Lemke HU, Vannier MW, Inamura K, Farman A (Eds). CAR 98 - Computer Assisted Radiology and Surgery. Amsterdam: Elsevier (1998) 437-442. OPENNESS IN (TELE-) RADIOLOGY WORKSTATIONS: THE CHILI
Reprint of :.Lemke HU, Vannier MW, Inamura K, Farman A (Eds). CAR 98 - Computer Assisted Radiology and Surgery. Amsterdam: Elsevier (1998) 437-442. OPENNESS IN (TELE-) RADIOLOGY WORKSTATIONS: THE CHILI
Project Setup and Data Management Tutorial
 Project Setup and Heavy Construction Edition Version 1.20 Corporate Office Trimble Navigation Limited Engineering and Construction Division 5475 Kellenburger Road Dayton, Ohio 45424-1099 U.S.A. Phone:
Project Setup and Heavy Construction Edition Version 1.20 Corporate Office Trimble Navigation Limited Engineering and Construction Division 5475 Kellenburger Road Dayton, Ohio 45424-1099 U.S.A. Phone:
Design document Goal Technology Description
 Design document Goal OpenOrienteering Mapper is a program to draw orienteering maps. It helps both in the surveying and the following final drawing task. Support for course setting is not a priority because
Design document Goal OpenOrienteering Mapper is a program to draw orienteering maps. It helps both in the surveying and the following final drawing task. Support for course setting is not a priority because
Introduction to MeVisLab Visual Programming Image Processing / VIsualization Examples VTK / ITK Integration MeVisLab SDK Features GUI Scripting
 Felix Ritter, MeVis Research Bremen, Germany Introduction to MeVisLab Visual Programming Image Processing / VIsualization Examples VTK / ITK Integration MeVisLab SDK Features GUI Scripting 2 Innovation
Felix Ritter, MeVis Research Bremen, Germany Introduction to MeVisLab Visual Programming Image Processing / VIsualization Examples VTK / ITK Integration MeVisLab SDK Features GUI Scripting 2 Innovation
A Tool for creating online Anatomical Atlases
 A Tool for creating online Anatomical Atlases Summer Scholarship Report Faculty of Science University of Auckland Summer 2003/2004 Daniel Rolf Wichmann UPI: dwic008 UID: 9790045 Supervisor: Burkhard Wuensche
A Tool for creating online Anatomical Atlases Summer Scholarship Report Faculty of Science University of Auckland Summer 2003/2004 Daniel Rolf Wichmann UPI: dwic008 UID: 9790045 Supervisor: Burkhard Wuensche
Building 3D anatomical scenes on the Web
 Building 3D anatomical scenes on the Web F. Evesque, S. Gerlach, R. D. Hersch Ecole Polytechnique Fédérale de Lausanne (EPFL) http://visiblehuman.epfl.ch Abstract We propose a new service for building
Building 3D anatomical scenes on the Web F. Evesque, S. Gerlach, R. D. Hersch Ecole Polytechnique Fédérale de Lausanne (EPFL) http://visiblehuman.epfl.ch Abstract We propose a new service for building
Introduction to Autodesk Inventor for F1 in Schools
 Introduction to Autodesk Inventor for F1 in Schools F1 in Schools Race Car In this course you will be introduced to Autodesk Inventor, which is the centerpiece of Autodesk s digital prototyping strategy
Introduction to Autodesk Inventor for F1 in Schools F1 in Schools Race Car In this course you will be introduced to Autodesk Inventor, which is the centerpiece of Autodesk s digital prototyping strategy
Visualization of 2D Domains
 Visualization of 2D Domains This part of the visualization package is intended to supply a simple graphical interface for 2- dimensional finite element data structures. Furthermore, it is used as the low
Visualization of 2D Domains This part of the visualization package is intended to supply a simple graphical interface for 2- dimensional finite element data structures. Furthermore, it is used as the low
3D-GIS in the Cloud USER MANUAL. August, 2014
 3D-GIS in the Cloud USER MANUAL August, 2014 3D GIS in the Cloud User Manual August, 2014 Table of Contents 1. Quick Reference: Navigating and Exploring in the 3D GIS in the Cloud... 2 1.1 Using the Mouse...
3D-GIS in the Cloud USER MANUAL August, 2014 3D GIS in the Cloud User Manual August, 2014 Table of Contents 1. Quick Reference: Navigating and Exploring in the 3D GIS in the Cloud... 2 1.1 Using the Mouse...
Cassandra 2.0: Tutorial
 Cassandra 2.0 Tutorial V1.0 Sébastien Jourdain, Fatiha Zeghir 2005/06/01 1 / 16 Abstract Cassandra is a generic VTK data viewer written in Java which provides native multiplatform support. Cassandra is
Cassandra 2.0 Tutorial V1.0 Sébastien Jourdain, Fatiha Zeghir 2005/06/01 1 / 16 Abstract Cassandra is a generic VTK data viewer written in Java which provides native multiplatform support. Cassandra is
Tutorial: Biped Character in 3D Studio Max 7, Easy Animation
 Tutorial: Biped Character in 3D Studio Max 7, Easy Animation Written by: Ricardo Tangali 1. Introduction:... 3 2. Basic control in 3D Studio Max... 3 2.1. Navigating a scene:... 3 2.2. Hide and Unhide
Tutorial: Biped Character in 3D Studio Max 7, Easy Animation Written by: Ricardo Tangali 1. Introduction:... 3 2. Basic control in 3D Studio Max... 3 2.1. Navigating a scene:... 3 2.2. Hide and Unhide
The File-Card-Browser View for Breast DCE-MRI Data
 The File-Card-Browser View for Breast DCE-MRI Data Sylvia Glaßer 1, Kathrin Scheil 1, Uta Preim 2, Bernhard Preim 1 1 Department of Simulation and Graphics, University of Magdeburg 2 Department of Radiology,
The File-Card-Browser View for Breast DCE-MRI Data Sylvia Glaßer 1, Kathrin Scheil 1, Uta Preim 2, Bernhard Preim 1 1 Department of Simulation and Graphics, University of Magdeburg 2 Department of Radiology,
The main imovie window is divided into six major parts.
 The main imovie window is divided into six major parts. 1. Project Drag clips to the project area to create a timeline 2. Preview Window Displays a preview of your video 3. Toolbar Contains a variety of
The main imovie window is divided into six major parts. 1. Project Drag clips to the project area to create a timeline 2. Preview Window Displays a preview of your video 3. Toolbar Contains a variety of
3D Interactive Information Visualization: Guidelines from experience and analysis of applications
 3D Interactive Information Visualization: Guidelines from experience and analysis of applications Richard Brath Visible Decisions Inc., 200 Front St. W. #2203, Toronto, Canada, rbrath@vdi.com 1. EXPERT
3D Interactive Information Visualization: Guidelines from experience and analysis of applications Richard Brath Visible Decisions Inc., 200 Front St. W. #2203, Toronto, Canada, rbrath@vdi.com 1. EXPERT
Data Visualization. Brief Overview of ArcMap
 Data Visualization Prepared by Francisco Olivera, Ph.D., P.E., Srikanth Koka and Lauren Walker Department of Civil Engineering September 13, 2006 Contents: Brief Overview of ArcMap Goals of the Exercise
Data Visualization Prepared by Francisco Olivera, Ph.D., P.E., Srikanth Koka and Lauren Walker Department of Civil Engineering September 13, 2006 Contents: Brief Overview of ArcMap Goals of the Exercise
multimodality image processing workstation Visualizing your SPECT-CT-PET-MRI images
 multimodality image processing workstation Visualizing your SPECT-CT-PET-MRI images InterView FUSION InterView FUSION is a new visualization and evaluation software developed by Mediso built on state of
multimodality image processing workstation Visualizing your SPECT-CT-PET-MRI images InterView FUSION InterView FUSION is a new visualization and evaluation software developed by Mediso built on state of
Maya 2014 Basic Animation & The Graph Editor
 Maya 2014 Basic Animation & The Graph Editor When you set a Keyframe (or Key), you assign a value to an object s attribute (for example, translate, rotate, scale, color) at a specific time. Most animation
Maya 2014 Basic Animation & The Graph Editor When you set a Keyframe (or Key), you assign a value to an object s attribute (for example, translate, rotate, scale, color) at a specific time. Most animation
Interactive Voting System. www.ivsystem.nl. IVS-Basic IVS-Professional 4.4
 Interactive Voting System www.ivsystem.nl IVS-Basic IVS-Professional 4.4 Manual IVS-Basic 4.4 IVS-Professional 4.4 1213 Interactive Voting System The Interactive Voting System (IVS ) is an interactive
Interactive Voting System www.ivsystem.nl IVS-Basic IVS-Professional 4.4 Manual IVS-Basic 4.4 IVS-Professional 4.4 1213 Interactive Voting System The Interactive Voting System (IVS ) is an interactive
CREATING A 3D VISUALISATION OF YOUR PLANS IN PLANSXPRESS AND CORTONA VRML CLIENT
 CREATING A 3D VISUALISATION OF YOUR PLANS IN PLANSXPRESS AND CORTONA VRML CLIENT 20-25 Minutes This topic is for users of PlansXpress Total Toolkit Edition. To upgrade to PlansXpress Total Toolkit, call
CREATING A 3D VISUALISATION OF YOUR PLANS IN PLANSXPRESS AND CORTONA VRML CLIENT 20-25 Minutes This topic is for users of PlansXpress Total Toolkit Edition. To upgrade to PlansXpress Total Toolkit, call
Visualization of Phylogenetic Trees and Metadata
 Visualization of Phylogenetic Trees and Metadata November 27, 2015 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com
Visualization of Phylogenetic Trees and Metadata November 27, 2015 Sample to Insight CLC bio, a QIAGEN Company Silkeborgvej 2 Prismet 8000 Aarhus C Denmark Telephone: +45 70 22 32 44 www.clcbio.com support-clcbio@qiagen.com
ArcGIS. Tips and Shortcuts. for Desktop
 ArcGIS Tips and Shortcuts for Desktop Map Navigation Refresh and redraw the display. F5 Suspend the map s drawing. F9 Zoom in and out. Center map. Roll the mouse wheel backward and forward. Hold down Ctrl
ArcGIS Tips and Shortcuts for Desktop Map Navigation Refresh and redraw the display. F5 Suspend the map s drawing. F9 Zoom in and out. Center map. Roll the mouse wheel backward and forward. Hold down Ctrl
The Design and Implementation of a C++ Toolkit for Integrated Medical Image Processing and Analyzing
 The Design and Implementation of a C++ Toolkit for Integrated Medical Image Processing and Analyzing Mingchang Zhao, Jie Tian 1, Xun Zhu, Jian Xue, Zhanglin Cheng, Hua Zhao Medical Image Processing Group,
The Design and Implementation of a C++ Toolkit for Integrated Medical Image Processing and Analyzing Mingchang Zhao, Jie Tian 1, Xun Zhu, Jian Xue, Zhanglin Cheng, Hua Zhao Medical Image Processing Group,
SOFTWARE FOR 3D IMAGE VISUALISATION, ANALYSIS AND MODEL GENERATION
 SOFTWARE FOR 3D IMAGE VISUALISATION, ANALYSIS AND MODEL GENERATION Solutions for Life Sciences SIMPLEWARE SOFTWARE Simpleware provides easy-to-use software solutions for the processing of 3D image data
SOFTWARE FOR 3D IMAGE VISUALISATION, ANALYSIS AND MODEL GENERATION Solutions for Life Sciences SIMPLEWARE SOFTWARE Simpleware provides easy-to-use software solutions for the processing of 3D image data
Technical What s New. Autodesk Alias Product Line
 Autodesk Alias Product Line Purpose-built for industrial designers and creative professionals, digital modelers/sculptors, and automotive/transportation designers, the Autodesk Alias 2010 product line
Autodesk Alias Product Line Purpose-built for industrial designers and creative professionals, digital modelers/sculptors, and automotive/transportation designers, the Autodesk Alias 2010 product line
Data Visualization. Prepared by Francisco Olivera, Ph.D., Srikanth Koka Department of Civil Engineering Texas A&M University February 2004
 Data Visualization Prepared by Francisco Olivera, Ph.D., Srikanth Koka Department of Civil Engineering Texas A&M University February 2004 Contents Brief Overview of ArcMap Goals of the Exercise Computer
Data Visualization Prepared by Francisco Olivera, Ph.D., Srikanth Koka Department of Civil Engineering Texas A&M University February 2004 Contents Brief Overview of ArcMap Goals of the Exercise Computer
Welcome to Corel DESIGNER, a comprehensive vector-based drawing application for creating technical graphics.
 Importing 3D models Welcome to Corel DESIGNER, a comprehensive vector-based drawing application for creating technical graphics. In this tutorial, you will modify a three-dimensional model of a transmission
Importing 3D models Welcome to Corel DESIGNER, a comprehensive vector-based drawing application for creating technical graphics. In this tutorial, you will modify a three-dimensional model of a transmission
anatomage table Interactive anatomy study table
 anatomage table Interactive anatomy study table Anatomage offers a unique, life-size interactive anatomy visualization table for the medical community. Anatomage Table offers an unprecedented realistic
anatomage table Interactive anatomy study table Anatomage offers a unique, life-size interactive anatomy visualization table for the medical community. Anatomage Table offers an unprecedented realistic
Outline. Fundamentals. Rendering (of 3D data) Data mappings. Evaluation Interaction
 Outline Fundamentals What is vis? Some history Design principles The visualization process Data sources and data structures Basic visual mapping approaches Rendering (of 3D data) Scalar fields (isosurfaces
Outline Fundamentals What is vis? Some history Design principles The visualization process Data sources and data structures Basic visual mapping approaches Rendering (of 3D data) Scalar fields (isosurfaces
Visualization with ParaView
 Visualization with ParaView Before we begin Make sure you have ParaView 4.1.0 installed so you can follow along in the lab section http://paraview.org/paraview/resources/software.php Background http://www.paraview.org/
Visualization with ParaView Before we begin Make sure you have ParaView 4.1.0 installed so you can follow along in the lab section http://paraview.org/paraview/resources/software.php Background http://www.paraview.org/
Introduction to Autodesk Inventor for F1 in Schools
 F1 in Schools race car Introduction to Autodesk Inventor for F1 in Schools In this course you will be introduced to Autodesk Inventor, which is the centerpiece of Autodesk s Digital Prototyping strategy
F1 in Schools race car Introduction to Autodesk Inventor for F1 in Schools In this course you will be introduced to Autodesk Inventor, which is the centerpiece of Autodesk s Digital Prototyping strategy
Sweet Home 3D user's guide
 1 de 14 08/01/2013 13:08 Features Download Online Gallery Blog Documentation FAQ User's guide Video tutorial Developer's guides History Reviews Support 3D models Textures Translations Forum Report a bug
1 de 14 08/01/2013 13:08 Features Download Online Gallery Blog Documentation FAQ User's guide Video tutorial Developer's guides History Reviews Support 3D models Textures Translations Forum Report a bug
CTvox for Android. Version 1.5
 CTvox for Android Version 1.5 Contents Introduction... 1 Background... 1 Volume rendering... 1 Multi-volume rendering... 2 Transfer function... 2 Installing CTvox... 3 Creating and transferring volume
CTvox for Android Version 1.5 Contents Introduction... 1 Background... 1 Volume rendering... 1 Multi-volume rendering... 2 Transfer function... 2 Installing CTvox... 3 Creating and transferring volume
Graphic Design. Background: The part of an artwork that appears to be farthest from the viewer, or in the distance of the scene.
 Graphic Design Active Layer- When you create multi layers for your images the active layer, or the only one that will be affected by your actions, is the one with a blue background in your layers palette.
Graphic Design Active Layer- When you create multi layers for your images the active layer, or the only one that will be affected by your actions, is the one with a blue background in your layers palette.
WebSphere Business Monitor
 WebSphere Business Monitor Dashboards 2010 IBM Corporation This presentation should provide an overview of the dashboard widgets for use with WebSphere Business Monitor. WBPM_Monitor_Dashboards.ppt Page
WebSphere Business Monitor Dashboards 2010 IBM Corporation This presentation should provide an overview of the dashboard widgets for use with WebSphere Business Monitor. WBPM_Monitor_Dashboards.ppt Page
The LiverSurgeryTrainer - Training of Computer-Based Planning in Liver Resection Surgery
 The LiverSurgeryTrainer - Training of Computer-Based Planning in Liver Resection Surgery Jeanette Mönch 1, Konrad Mühler 2, Christian Hansen 3, Karl-Jürgen Oldhafer 4, Gregor Stavrou 4, Christian Hillert
The LiverSurgeryTrainer - Training of Computer-Based Planning in Liver Resection Surgery Jeanette Mönch 1, Konrad Mühler 2, Christian Hansen 3, Karl-Jürgen Oldhafer 4, Gregor Stavrou 4, Christian Hillert
DataPA OpenAnalytics End User Training
 DataPA OpenAnalytics End User Training DataPA End User Training Lesson 1 Course Overview DataPA Chapter 1 Course Overview Introduction This course covers the skills required to use DataPA OpenAnalytics
DataPA OpenAnalytics End User Training DataPA End User Training Lesson 1 Course Overview DataPA Chapter 1 Course Overview Introduction This course covers the skills required to use DataPA OpenAnalytics
Autodesk Fusion 360: Assemblies. Overview
 Overview In this module you will learn how different components can be put together to create an assembly. We will use several tools in Fusion 360 to make sure that these assemblies are constrained appropriately
Overview In this module you will learn how different components can be put together to create an assembly. We will use several tools in Fusion 360 to make sure that these assemblies are constrained appropriately
MicroStrategy Analytics Express User Guide
 MicroStrategy Analytics Express User Guide Analyzing Data with MicroStrategy Analytics Express Version: 4.0 Document Number: 09770040 CONTENTS 1. Getting Started with MicroStrategy Analytics Express Introduction...
MicroStrategy Analytics Express User Guide Analyzing Data with MicroStrategy Analytics Express Version: 4.0 Document Number: 09770040 CONTENTS 1. Getting Started with MicroStrategy Analytics Express Introduction...
Software Packages The following data analysis software packages will be showcased:
 Analyze This! Practicalities of fmri and Diffusion Data Analysis Data Download Instructions Weekday Educational Course, ISMRM 23 rd Annual Meeting and Exhibition Tuesday 2 nd June 2015, 10:00-12:00, Room
Analyze This! Practicalities of fmri and Diffusion Data Analysis Data Download Instructions Weekday Educational Course, ISMRM 23 rd Annual Meeting and Exhibition Tuesday 2 nd June 2015, 10:00-12:00, Room
Tizian TM Creativ RT. Abutment Designer. for the CAD-software version 3.932. Instruction Manual
 Tizian TM Creativ RT Abutment Designer for the CAD-software version 3.932 Instruction Manual Schütz Dental GmbH 61191 Rosbach, Germany Tel.: +49 6003 / 814-0 Fax: +49 6003 / 814-906 Version: 12/2010 Contents
Tizian TM Creativ RT Abutment Designer for the CAD-software version 3.932 Instruction Manual Schütz Dental GmbH 61191 Rosbach, Germany Tel.: +49 6003 / 814-0 Fax: +49 6003 / 814-906 Version: 12/2010 Contents
imc FAMOS 6.3 visualization signal analysis data processing test reporting Comprehensive data analysis and documentation imc productive testing
 imc FAMOS 6.3 visualization signal analysis data processing test reporting Comprehensive data analysis and documentation imc productive testing imc FAMOS ensures fast results Comprehensive data processing
imc FAMOS 6.3 visualization signal analysis data processing test reporting Comprehensive data analysis and documentation imc productive testing imc FAMOS ensures fast results Comprehensive data processing
M-Files Gantt View. User Guide. App Version: 1.1.0 Author: Joel Heinrich
 M-Files Gantt View User Guide App Version: 1.1.0 Author: Joel Heinrich Date: 02-Jan-2013 Contents 1 Introduction... 1 1.1 Requirements... 1 2 Basic Use... 1 2.1 Activation... 1 2.2 Layout... 1 2.3 Navigation...
M-Files Gantt View User Guide App Version: 1.1.0 Author: Joel Heinrich Date: 02-Jan-2013 Contents 1 Introduction... 1 1.1 Requirements... 1 2 Basic Use... 1 2.1 Activation... 1 2.2 Layout... 1 2.3 Navigation...
Visualization methods for patent data
 Visualization methods for patent data Treparel 2013 Dr. Anton Heijs (CTO & Founder) Delft, The Netherlands Introduction Treparel can provide advanced visualizations for patent data. This document describes
Visualization methods for patent data Treparel 2013 Dr. Anton Heijs (CTO & Founder) Delft, The Netherlands Introduction Treparel can provide advanced visualizations for patent data. This document describes
ACE: After Effects CS6
 Adobe Training Services Exam Guide ACE: After Effects CS6 Adobe Training Services provides this exam guide to help prepare partners, customers, and consultants who are actively seeking accreditation as
Adobe Training Services Exam Guide ACE: After Effects CS6 Adobe Training Services provides this exam guide to help prepare partners, customers, and consultants who are actively seeking accreditation as
How to use PGS: Basic Services Provision Map App
 How to use PGS: Basic Services Provision Map App The PGS: Basic Services Provision Map App The main features of the PGP Basic Services web application includes: Navigation Tools Map Tools Main Map Links
How to use PGS: Basic Services Provision Map App The PGS: Basic Services Provision Map App The main features of the PGP Basic Services web application includes: Navigation Tools Map Tools Main Map Links
TABLE OF CONTENTS. INTRODUCTION... 5 Advance Concrete... 5 Where to find information?... 6 INSTALLATION... 7 STARTING ADVANCE CONCRETE...
 Starting Guide TABLE OF CONTENTS INTRODUCTION... 5 Advance Concrete... 5 Where to find information?... 6 INSTALLATION... 7 STARTING ADVANCE CONCRETE... 7 ADVANCE CONCRETE USER INTERFACE... 7 Other important
Starting Guide TABLE OF CONTENTS INTRODUCTION... 5 Advance Concrete... 5 Where to find information?... 6 INSTALLATION... 7 STARTING ADVANCE CONCRETE... 7 ADVANCE CONCRETE USER INTERFACE... 7 Other important
GCE APPLIED ICT A2 COURSEWORK TIPS
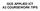 GCE APPLIED ICT A2 COURSEWORK TIPS COURSEWORK TIPS A2 GCE APPLIED ICT If you are studying for the six-unit GCE Single Award or the twelve-unit Double Award, then you may study some of the following coursework
GCE APPLIED ICT A2 COURSEWORK TIPS COURSEWORK TIPS A2 GCE APPLIED ICT If you are studying for the six-unit GCE Single Award or the twelve-unit Double Award, then you may study some of the following coursework
Turtle Beach Grip 500 Laser Gaming Mouse. User Guide
 Turtle Beach Grip 500 Laser Gaming Mouse User Guide Table of Contents Table of Contents... 4 Introduction... 5 Installation... 5 Opening and Closing Grip 500 Configuration Software... 6 Configuring Your
Turtle Beach Grip 500 Laser Gaming Mouse User Guide Table of Contents Table of Contents... 4 Introduction... 5 Installation... 5 Opening and Closing Grip 500 Configuration Software... 6 Configuring Your
ACE: After Effects CC
 Adobe Training Services Exam Guide ACE: After Effects CC Adobe Training Services provides this exam guide to help prepare partners, customers, and consultants who are actively seeking accreditation as
Adobe Training Services Exam Guide ACE: After Effects CC Adobe Training Services provides this exam guide to help prepare partners, customers, and consultants who are actively seeking accreditation as
Volume visualization I Elvins
 Volume visualization I Elvins 1 surface fitting algorithms marching cubes dividing cubes direct volume rendering algorithms ray casting, integration methods voxel projection, projected tetrahedra, splatting
Volume visualization I Elvins 1 surface fitting algorithms marching cubes dividing cubes direct volume rendering algorithms ray casting, integration methods voxel projection, projected tetrahedra, splatting
Help. Contents Back >>
 Contents Back >> Customizing Opening the Control Panel Control Panel Features Tabs Control Panel Lists Control Panel Buttons Customizing Your Tools Pen and Airbrush Tabs 2D Mouse and 4D Mouse Tabs Customizing
Contents Back >> Customizing Opening the Control Panel Control Panel Features Tabs Control Panel Lists Control Panel Buttons Customizing Your Tools Pen and Airbrush Tabs 2D Mouse and 4D Mouse Tabs Customizing
Visualization with ParaView. Greg Johnson
 Visualization with Greg Johnson Before we begin Make sure you have 3.8.0 installed so you can follow along in the lab section http://paraview.org/paraview/resources/software.html http://www.paraview.org/
Visualization with Greg Johnson Before we begin Make sure you have 3.8.0 installed so you can follow along in the lab section http://paraview.org/paraview/resources/software.html http://www.paraview.org/
Content Author's Reference and Cookbook
 Sitecore CMS 6.2 Content Author's Reference and Cookbook Rev. 091019 Sitecore CMS 6.2 Content Author's Reference and Cookbook A Conceptual Overview and Practical Guide to Using Sitecore Table of Contents
Sitecore CMS 6.2 Content Author's Reference and Cookbook Rev. 091019 Sitecore CMS 6.2 Content Author's Reference and Cookbook A Conceptual Overview and Practical Guide to Using Sitecore Table of Contents
Appointment Scheduler
 EZClaim Appointment Scheduler User Guide Last Update: 11/19/2008 Copyright 2008 EZClaim This page intentionally left blank Contents Contents... iii Getting Started... 5 System Requirements... 5 Installing
EZClaim Appointment Scheduler User Guide Last Update: 11/19/2008 Copyright 2008 EZClaim This page intentionally left blank Contents Contents... iii Getting Started... 5 System Requirements... 5 Installing
Provenance for Visualizations
 V i s u a l i z a t i o n C o r n e r Editors: Claudio Silva, csilva@cs.utah.edu Joel E. Tohline, tohline@rouge.phys.lsu.edu Provenance for Visualizations Reproducibility and Beyond By Claudio T. Silva,
V i s u a l i z a t i o n C o r n e r Editors: Claudio Silva, csilva@cs.utah.edu Joel E. Tohline, tohline@rouge.phys.lsu.edu Provenance for Visualizations Reproducibility and Beyond By Claudio T. Silva,
How To Compress Video For Real Time Transmission
 University of Edinburgh College of Science and Engineering School of Informatics Informatics Research Proposal supervised by Dr. Sethu Vijayakumar Optimized bandwidth usage for real-time remote surveillance
University of Edinburgh College of Science and Engineering School of Informatics Informatics Research Proposal supervised by Dr. Sethu Vijayakumar Optimized bandwidth usage for real-time remote surveillance
IntelliSpace PACS 4.4. Image Enabled EMR Workflow and Enterprise Overview. Learning Objectives
 IntelliSpace PACS 4.4 Image Enabled EMR Workflow and Enterprise Overview Learning Objectives Use the Cerner EMR to view images and reports via IntelliSpace PACS: Image Launch Workflow (SUBI Image Viewer)
IntelliSpace PACS 4.4 Image Enabled EMR Workflow and Enterprise Overview Learning Objectives Use the Cerner EMR to view images and reports via IntelliSpace PACS: Image Launch Workflow (SUBI Image Viewer)
IV3Dm provides global settings which can be set prior to launching the application and are available through the device settings menu.
 ImageVis3D Mobile This software can be used to display and interact with different kinds of datasets - such as volumes or meshes - on mobile devices, which currently includes iphone and ipad. A selection
ImageVis3D Mobile This software can be used to display and interact with different kinds of datasets - such as volumes or meshes - on mobile devices, which currently includes iphone and ipad. A selection
FACILITIES INVENTORY MANAGEMENT SYSTEM:
 FACILITIES INVENTORY MANAGEMENT SYSTEM: FM:INTERACT USER S GUIDE University of North Texas Office of Facilities Management and Construction P.O. Box 311040 Denton, TX 76203-1040 TEL: (940)565.4974 FAX:
FACILITIES INVENTORY MANAGEMENT SYSTEM: FM:INTERACT USER S GUIDE University of North Texas Office of Facilities Management and Construction P.O. Box 311040 Denton, TX 76203-1040 TEL: (940)565.4974 FAX:
HOW TO LINK AND PRESENT A 4D MODEL USING NAVISWORKS. Timo Hartmann t.hartmann@ctw.utwente.nl
 Technical Paper #1 HOW TO LINK AND PRESENT A 4D MODEL USING NAVISWORKS Timo Hartmann t.hartmann@ctw.utwente.nl COPYRIGHT 2009 VISICO Center, University of Twente visico@utwente.nl How to link and present
Technical Paper #1 HOW TO LINK AND PRESENT A 4D MODEL USING NAVISWORKS Timo Hartmann t.hartmann@ctw.utwente.nl COPYRIGHT 2009 VISICO Center, University of Twente visico@utwente.nl How to link and present
Working With Animation: Introduction to Flash
 Working With Animation: Introduction to Flash With Adobe Flash, you can create artwork and animations that add motion and visual interest to your Web pages. Flash movies can be interactive users can click
Working With Animation: Introduction to Flash With Adobe Flash, you can create artwork and animations that add motion and visual interest to your Web pages. Flash movies can be interactive users can click
Switching from PC SAS to SAS Enterprise Guide Zhengxin (Cindy) Yang, inventiv Health Clinical, Princeton, NJ
 PharmaSUG 2014 PO10 Switching from PC SAS to SAS Enterprise Guide Zhengxin (Cindy) Yang, inventiv Health Clinical, Princeton, NJ ABSTRACT As more and more organizations adapt to the SAS Enterprise Guide,
PharmaSUG 2014 PO10 Switching from PC SAS to SAS Enterprise Guide Zhengxin (Cindy) Yang, inventiv Health Clinical, Princeton, NJ ABSTRACT As more and more organizations adapt to the SAS Enterprise Guide,
Intraoperative Adaptation and Visualization of Preoperative Risk Analyses for Oncologic Liver Surgery
 Intraoperative Adaptation and Visualization of Preoperative Risk Analyses for Oncologic Liver Surgery Christian Hansen *a, Stefan Schlichting b, Stephan Zidowitz a, Alexander Köhn a, Milo Hindennach a,
Intraoperative Adaptation and Visualization of Preoperative Risk Analyses for Oncologic Liver Surgery Christian Hansen *a, Stefan Schlichting b, Stephan Zidowitz a, Alexander Köhn a, Milo Hindennach a,
Introduction to Visualization with VTK and ParaView
 Introduction to Visualization with VTK and ParaView R. Sungkorn and J. Derksen Department of Chemical and Materials Engineering University of Alberta Canada August 24, 2011 / LBM Workshop 1 Introduction
Introduction to Visualization with VTK and ParaView R. Sungkorn and J. Derksen Department of Chemical and Materials Engineering University of Alberta Canada August 24, 2011 / LBM Workshop 1 Introduction
Presented by Peiqun (Anthony) Yu
 Presented by Peiqun (Anthony) Yu A Multi-Scale, Multi-Layer, Translucent Virtual Space Henry Lieberman, IEEE International Conference on Information Visualization, London, September 1997. Constant Information
Presented by Peiqun (Anthony) Yu A Multi-Scale, Multi-Layer, Translucent Virtual Space Henry Lieberman, IEEE International Conference on Information Visualization, London, September 1997. Constant Information
ANIMATION a system for animation scene and contents creation, retrieval and display
 ANIMATION a system for animation scene and contents creation, retrieval and display Peter L. Stanchev Kettering University ABSTRACT There is an increasing interest in the computer animation. The most of
ANIMATION a system for animation scene and contents creation, retrieval and display Peter L. Stanchev Kettering University ABSTRACT There is an increasing interest in the computer animation. The most of
S. Hartmann, C. Seiler, R. Dörner and P. Grimm
 &DVH6WXG\9LVXDOL]DWLRQRI0HFKDQLFDO3URSHUWLHVDQG 'HIRUPDWLRQVRI/LYLQJ&HOOV S. Hartmann, C. Seiler, R. Dörner and P. Grimm Fraunhofer Anwendungszentrum für Computergraphik in Chemie und Pharmazie Varrentrappstraße
&DVH6WXG\9LVXDOL]DWLRQRI0HFKDQLFDO3URSHUWLHVDQG 'HIRUPDWLRQVRI/LYLQJ&HOOV S. Hartmann, C. Seiler, R. Dörner and P. Grimm Fraunhofer Anwendungszentrum für Computergraphik in Chemie und Pharmazie Varrentrappstraße
imc FAMOS 6.3 visualization signal analysis data processing test reporting Comprehensive data analysis and documentation imc productive testing
 imc FAMOS 6.3 visualization signal analysis data processing test reporting Comprehensive data analysis and documentation imc productive testing www.imcfamos.com imc FAMOS at a glance Four editions to Optimize
imc FAMOS 6.3 visualization signal analysis data processing test reporting Comprehensive data analysis and documentation imc productive testing www.imcfamos.com imc FAMOS at a glance Four editions to Optimize
Intro to Excel spreadsheets
 Intro to Excel spreadsheets What are the objectives of this document? The objectives of document are: 1. Familiarize you with what a spreadsheet is, how it works, and what its capabilities are; 2. Using
Intro to Excel spreadsheets What are the objectives of this document? The objectives of document are: 1. Familiarize you with what a spreadsheet is, how it works, and what its capabilities are; 2. Using
Intellect Platform - Tables and Templates Basic Document Management System - A101
 Intellect Platform - Tables and Templates Basic Document Management System - A101 Interneer, Inc. 4/12/2010 Created by Erika Keresztyen 2 Tables and Templates - A101 - Basic Document Management System
Intellect Platform - Tables and Templates Basic Document Management System - A101 Interneer, Inc. 4/12/2010 Created by Erika Keresztyen 2 Tables and Templates - A101 - Basic Document Management System
Big Data: Rethinking Text Visualization
 Big Data: Rethinking Text Visualization Dr. Anton Heijs anton.heijs@treparel.com Treparel April 8, 2013 Abstract In this white paper we discuss text visualization approaches and how these are important
Big Data: Rethinking Text Visualization Dr. Anton Heijs anton.heijs@treparel.com Treparel April 8, 2013 Abstract In this white paper we discuss text visualization approaches and how these are important
Imaris Quick Start Tutorials
 Imaris 1 Introduction Why should you read and practice the Imaris? They provide you with the basic information how-to-use Imaris but may also show yet unrecognized new features of the software to the advanced
Imaris 1 Introduction Why should you read and practice the Imaris? They provide you with the basic information how-to-use Imaris but may also show yet unrecognized new features of the software to the advanced
WebFOCUS BI Portal: S.I.M.P.L.E. as can be
 WebFOCUS BI Portal: S.I.M.P.L.E. as can be Author: Matthew Lerner Company: Information Builders Presentation Abstract: This hands-on session will introduce attendees to the new WebFOCUS BI Portal. We will
WebFOCUS BI Portal: S.I.M.P.L.E. as can be Author: Matthew Lerner Company: Information Builders Presentation Abstract: This hands-on session will introduce attendees to the new WebFOCUS BI Portal. We will
Ansur Test Executive. Users Manual
 Ansur Test Executive Users Manual April 2008 2008 Fluke Corporation, All rights reserved. All product names are trademarks of their respective companies Table of Contents 1 Introducing Ansur... 4 1.1 About
Ansur Test Executive Users Manual April 2008 2008 Fluke Corporation, All rights reserved. All product names are trademarks of their respective companies Table of Contents 1 Introducing Ansur... 4 1.1 About
Vodafone Business Product Management Group. Hosted Services EasySiteWizard Pro 8 User Guide
 Vodafone Business Product Management Group Hosted Services EasySiteWizard Pro 8 User Guide Vodafone Group 2010 Other than as permitted by law, no part of this document may be reproduced, adapted, or distributed,
Vodafone Business Product Management Group Hosted Services EasySiteWizard Pro 8 User Guide Vodafone Group 2010 Other than as permitted by law, no part of this document may be reproduced, adapted, or distributed,
VISUALIZATION STRATEGIES AND TECHNIQUES FOR HIGH-DIMENSIONAL SPATIO- TEMPORAL DATA
 VISUALIZATION STRATEGIES AND TECHNIQUES FOR HIGH-DIMENSIONAL SPATIO- TEMPORAL DATA Summary B. Schmidt, U. Streit and Chr. Uhlenküken University of Münster Institute of Geoinformatics Robert-Koch-Str. 28
VISUALIZATION STRATEGIES AND TECHNIQUES FOR HIGH-DIMENSIONAL SPATIO- TEMPORAL DATA Summary B. Schmidt, U. Streit and Chr. Uhlenküken University of Münster Institute of Geoinformatics Robert-Koch-Str. 28
Files Used in this Tutorial
 Generate Point Clouds Tutorial This tutorial shows how to generate point clouds from IKONOS satellite stereo imagery. You will view the point clouds in the ENVI LiDAR Viewer. The estimated time to complete
Generate Point Clouds Tutorial This tutorial shows how to generate point clouds from IKONOS satellite stereo imagery. You will view the point clouds in the ENVI LiDAR Viewer. The estimated time to complete
m ac romed ia Fl a s h Curriculum Guide
 m ac romed ia Fl a s h Curriculum Guide 1997 1998 Macromedia, Inc. All rights reserved. Macromedia, the Macromedia logo, Dreamweaver, Director, Fireworks, Flash, Fontographer, FreeHand, and Xtra are trademarks
m ac romed ia Fl a s h Curriculum Guide 1997 1998 Macromedia, Inc. All rights reserved. Macromedia, the Macromedia logo, Dreamweaver, Director, Fireworks, Flash, Fontographer, FreeHand, and Xtra are trademarks
Processing Data with rsmap3d Software Services Group Advanced Photon Source Argonne National Laboratory
 Processing Data with rsmap3d Software Services Group Advanced Photon Source Argonne National Laboratory Introduction rsmap3d is an application for producing 3D reciprocal space maps from x-ray diffraction
Processing Data with rsmap3d Software Services Group Advanced Photon Source Argonne National Laboratory Introduction rsmap3d is an application for producing 3D reciprocal space maps from x-ray diffraction
A Novel Multitouch Interface for 3D Object Manipulation
 A Novel Multitouch Interface for 3D Object Manipulation Oscar Kin-Chung Au School of Creative Media City University of Hong Kong kincau@cityu.edu.hk Chiew-Lan Tai Department of Computer Science & Engineering
A Novel Multitouch Interface for 3D Object Manipulation Oscar Kin-Chung Au School of Creative Media City University of Hong Kong kincau@cityu.edu.hk Chiew-Lan Tai Department of Computer Science & Engineering
Structure Tools and Visualization
 Structure Tools and Visualization Gary Van Domselaar University of Alberta gary.vandomselaar@ualberta.ca Slides Adapted from Michel Dumontier, Blueprint Initiative 1 Visualization & Communication Visualization
Structure Tools and Visualization Gary Van Domselaar University of Alberta gary.vandomselaar@ualberta.ca Slides Adapted from Michel Dumontier, Blueprint Initiative 1 Visualization & Communication Visualization
Advanced Volume Rendering Techniques for Medical Applications
 Advanced Volume Rendering Techniques for Medical Applications Verbesserte Darstellungsmethoden für Volumendaten in medizinischen Anwendungen J. Georgii 1, J. Schneider 1, J. Krüger 1, R. Westermann 1,
Advanced Volume Rendering Techniques for Medical Applications Verbesserte Darstellungsmethoden für Volumendaten in medizinischen Anwendungen J. Georgii 1, J. Schneider 1, J. Krüger 1, R. Westermann 1,
4D Interactive Model Animations
 Animation Using 4D Interactive Models MVSand EVS-PRO have two distinctly different animation concepts. Our traditional animations consist of a sequence of bitmap images that have been encoded into an animation
Animation Using 4D Interactive Models MVSand EVS-PRO have two distinctly different animation concepts. Our traditional animations consist of a sequence of bitmap images that have been encoded into an animation
Using MindManager 14
 Using MindManager 14 Susi Peacock, Graeme Ferris, Susie Beasley, Matt Sanders and Lindesay Irvine Version 4 September 2014 2011 Queen Margaret University 1. Navigating MindManager 14... 3 Tool Bars and
Using MindManager 14 Susi Peacock, Graeme Ferris, Susie Beasley, Matt Sanders and Lindesay Irvine Version 4 September 2014 2011 Queen Margaret University 1. Navigating MindManager 14... 3 Tool Bars and
Video, film, and animation are all moving images that are recorded onto videotape,
 See also Data Display (Part 3) Document Design (Part 3) Instructions (Part 2) Specifications (Part 2) Visual Communication (Part 3) Video and Animation Video, film, and animation are all moving images
See also Data Display (Part 3) Document Design (Part 3) Instructions (Part 2) Specifications (Part 2) Visual Communication (Part 3) Video and Animation Video, film, and animation are all moving images
ParaVision 6. Innovation with Integrity. The Next Generation of MR Acquisition and Processing for Preclinical and Material Research.
 ParaVision 6 The Next Generation of MR Acquisition and Processing for Preclinical and Material Research Innovation with Integrity Preclinical MRI A new standard in Preclinical Imaging ParaVision sets a
ParaVision 6 The Next Generation of MR Acquisition and Processing for Preclinical and Material Research Innovation with Integrity Preclinical MRI A new standard in Preclinical Imaging ParaVision sets a
CENTRICITY WEB VERSION 3.0. Updated 11/24/2009 Medical Imaging IT Technical Support please call the Help Desk 617-732-5927
 TABLE OF CONTENTS PAGE 1 CENTRICITY QUICK REFERENCE GUIDE PAGE 2-3 ACCESSING CENTRICITY FROM LMR PAGES 4-5 ACCESSING CENTRICITY FROM CAS PAGES 6-7 3 1 2 9 1. Worklist Views Allows the user to view studies
TABLE OF CONTENTS PAGE 1 CENTRICITY QUICK REFERENCE GUIDE PAGE 2-3 ACCESSING CENTRICITY FROM LMR PAGES 4-5 ACCESSING CENTRICITY FROM CAS PAGES 6-7 3 1 2 9 1. Worklist Views Allows the user to view studies
