A Supervised Named-Entity Extraction System for Medical Text
|
|
|
- Brittney Barton
- 10 years ago
- Views:
Transcription
1 A Supervised Named-Entity Extraction System for Medical Text Andreea Bodnari 1,2, Louise Deléger 2, Thomas Lavergne 2, Aurélie Névéol 2, and Pierre Zweigenbaum 2 1 MIT, CSAIL, Cambridge, Massachusetts, USA 2 LIMSI-CNRS, rue John von Neumann, F Orsay, France Abstract. We present our participation in Task 1a of the 2013 CLEFeHEALTH Challenge, whose goal was the identification of disorder named entities from electronic medical records. We developed a supervised CRF model that based on a rich set of features learns to predict disorder named entities. The CRF system uses external knowledge from specialized biomedical terminologies and Wikipedia. Our system performance was evaluated at F-measure in the context of strict evaluation and F-measure in the context of relaxed evaluation. Keywords: Named-entity recognition, Natural Language Processing, Medical records, Machine learning 1 Introduction Electronic medical records (EMRs) represent rich data repositories loaded with valuable patient information. Automated tools are required to process this patient information and make it available to medical professionals and specialized medical systems. These automated tools take as input the plain text of EMRs and output data of interest. For example, named-entity extraction tools process the plain text of EMRs and extract instances of named entities (i.e., noun phrases) that can be classified into a certain semantic category. The 2013 CLEF-eHEALTH challenge aims to develop methods and resources that make EMRs more understandable by both patients and health professionals. The challenge spans over three tasks. We participate in the first task, which focuses on the identification of disorder named entities in electronic medical records, and develop a system that can perform NER on medical text. We propose a solution that combines a rich feature set with external knowledge gathered from both specialized and general domain knowledge repositories. We present a named-entity recognition system specialized in the medical domain. Our system learns a CRF model from the training data, based on a rich feature set that combines external knowledge sources with information gathered from the EMR text. We discuss the system design, present the system results on the 2013 CLEF-eHEALTH [10] training and test data, and discuss the specific features that helped most with the system performance.
2 2 Andreea Bodnari et al. 2 Related work The natural language processing (NLP) community has organized specialized competitions to evaluate and help improve the state of the art in various NLP domains. Competitions in the general domain include the Text Retrieval Evaluation Conferences [2], the Semantic Evaluation (SemEval) [1], and the Conference on Natural Language Learning (CoNLL) shared-tasks [5]. In the medical domain, the Informatics for Integrating Biology and the Bedside (i2b2) center organized a series of NLP competitions focused on information extraction from unstructured clinical documents. The NLP competitions tried to support advancement in a series of NLP tasks like information extraction [13], information retrieval [11], semantic textual similarity, co-reference resolution [12]. Tasks like information retrieval, dependency parsing, and named entity recognition exhibit relatively well performing solutions that can be applied in real life settings. Yet, results of NLP competitions showed that the research community is still struggling in the specialized domain (i.e., medical, biomedical) compared to the general domain. 3 Materials and methods 3.1 Data The corpus used for the 2013 CLEF-eHEALTH challenge consists of de-identified plain text EMRs from the MIMIC II database, version 2.5 [8]. The EMR documents were extracted from the intensive-care unit setting and included discharge summaries, electrocardiography reports, echo reports, and radiology reports. The training set contained 200 documents and a total of 94,243 words, while the test set contained 100 documents and a total of 87,799 words (see Table 2). Annotation of disorder noun phrases (NPs) was carried out as part of the Shared Annotated Resources project [7]. The text of each EMR document was annotated by two professional coders trained for this task, followed by an open adjudication step. A disorder noun phrase is defined as any span of text which can be mapped to a concept in the SNOMED-CT terminology and which belongs to the Disorder semantic group. A concept is in the Disorder semantic group if it belongs to one of the following Unified Medical Language System (UMLS) [4] semantic types: congenital abnormality, acquired abnormality, injury or poisoning, pathological function, disease or syndrome, mental or behavioral dysfunction, experimental model of disease, anatomical abnormality, neoplastic process, signs and symptoms. The training set contained 5, 874 annotations, while the test set contained 5, 351 annotations (see Table 2). The non-contiguous entities accounted for approx. 10% of the training and test data. 3.2 System design We developed a supervised linear-chain Conditional Random Fields (CRF) model using 10-fold cross validation on the training set data. We first present the preprocessing steps we performed on the datasets. We then describe the model
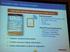
3 Supervised Named-Entity Extraction 3 Table 1. Description of training and test data sets Training Test Word count 94,243 87,799 Annotation count 5,874 5,351 Non-Contiguous annotation count Documents feature set together with the CRF feature patterns. The feature production architecture is schematized in Figure 1. Data pre-processing Before using the challenge corpora for training and testing, we performed several pre-processing steps: the training and test corpora provided by the challenge organizers were deidentified and thus contained special de-identification marks; to turn deidentification code into more normal phrases, we performed re-identification with pseudonyms on the input text. EMR documents present in general a well-structured form, with a header, document body, and a footer. The header and footer contain information relevant to clinical administration, but the disorder NPs are only encountered inside the document body. We thus removed the header and footer from the EMR documents and performed analysis on the document body only. Sections Section labels Full text Tokenize, Re-identify Re-identified, tokenized text Detect document type and split Header Body Tokenized sentences ctakes MetaMap Exact match POS-tagged, parsed, UMLS information (1) UMLS information (2) Footer Wikipedia UMLS information (3) Wikipedia semantic classes Fig. 1. Diagram of feature production.
4 4 Andreea Bodnari et al. System features Given a sentence s =... w 2 w 1 w 0 w 1 w 2... and a token of interest w k, we define features over w k and n-grams centered at w k. 1. Lexical and morphological features: we include information on the token and on the token lemma in the form of unigrams over w k 1, w k, w k+1 and bigrams over w k 1 w k. We also include as unigram features the following token characteristics: token contains only upper case characters, token is a digit, is capitalized, or is a punctuation. Additionally we include as unigram features over w k token suffixes ranging from 1 to 4 characters. Finally we add a 5-gram feature which detects patterns containing two slashes, such as m/r/g, which may reveal up to three disorders (such constructs are split into 5 tokens by our tokenizer), and apply it over w k 4, w k 2, w k (the non-slash positions of the pattern). 2. Syntactic features: we tokenize and parse the EMR plain text using the ctakes [9] system. We include as features the part of speech information in the form of unigrams over w k 3, w k 2,..., w k+3 and bigrams over w k 3,..., w k+2. We further include the parse-tree dependency information for the current token w k, the bigram w k 1 w k, and trigram w k 1 w k w k Document structure features: the feature set contains as features the document type (e.g., radiology report, discharge summary) and the section type (e.g., Findings, Laboratory data, Social History). We extract the section type using a rule-based section extraction tool that identifies the occurrence of section names within the EMR. The section extraction tool uses a list of 58 manually defined section names. Both document type and section type are unigram features over w k. 4. UMLS features: we include UMLS information from three sources. We first use the UMLS information provided by ctakes: the semantic type unique identifier (defined over unigrams w k 1, w k, w k+1 and bigram w k 1 w k ), and semantic group information (defined over unigrams w k 1, w k, w k+1 and bigrams w k 1 w k and w k w k+1 ). Secondly, we process the EMR plain text using MetaMap [3] and include the semantic group it identifies. We use an additional UMLS mapping where we directly search for UMLS noun phrases within the EMR text through exact match and include the semantic group and concept unique identifier (CUI) of the identified phrase. The MetaMap and the direct UMLS mapping features are unigram features over w k 1, w k, w k+1. We also define a unigram binary feature for being a member of the Disorder semantic group and one for being a member of the Anatomy semantic group. 5. Wikipedia features: we make use of the Wikipedia Category information in order to classify the noun-phrases contained in EMRs. We group the Wikipedia categories into nine semantic groups: disorder, body part, living being, chemicals, phenomenon, object, geographical location, devices, and other. The other category contains the Wikipedia categories not included in any of the defined categories. We use the article titles from the English Wikipedia and search for their occurrence within the EMR plain text. Once an article title is found we map its Wikipedia category to one of the categories
5 Supervised Named-Entity Extraction 5 previously defined. We define the Wikipedia system feature as unigram over w k 1, w k, w k+1. All features pertaining to multi-token expressions instead of only single tokens (for instance, being a UMLS term with a given semantic group, or being an abbreviation) are encoded with the begin inside outside (B-I-O) scheme: given a label L, the first token is labeled B-L, the next tokens are labeled I-L, and tokens not having this feature are labeled O. All features can use both unigrams and bigrams of classes: this leverages the specific capabilities of linear-chain CRFs to label sequences instead of isolated tokens. Problem formulation We model the problem as a supervised classification task with three labels: B-Disorder, I-Disorder, and O (outside). We include in the gold standard of the training set the contiguous entities and only partially took into account the non-contiguous entities. Observing that non-contiguous entities generally follow the general SNOMED v3.5 model, with a morphology / dysfunction part, which is akin to a disorder, and another (generally, anatomy) part, we only include the morphology / dysfunction part of non-contiguous entities, based on their UMLS semantic types, labelling them with B-Disorder and I-Disorder classes. This allows us to handle them as though they were contiguous entities without polluting the gold standard labels with disorder-labeled anatomy segments that would perturb training and classification. We used the Wapiti [6] 1 implementation of CRFs because it is fast and offers convenient patterns (e.g., patterns using regular expressions on field values). 4 Results and discussion 4.1 Evaluation metrics We evaluate the system s ability to correctly identify the spans of disorder NPs. The evaluation measures are precision, recall, and F-measure, defined as Precision = Recall = T P T P + F P T P T P + F N (1) (2) F-measure = 2 P recision Recall P recision + Recall where T P = count of system NPs presenting same span as gold standard NPs; F P = count of system NPs presenting divergent span from gold standard NPs; F N = count of gold standard NPs not present in the system NPs. 1 (3)
6 6 Andreea Bodnari et al. We compute the system NP span overlap to the gold standard NP under two settings: the strict evaluation setting, where the system NP span is identical to the gold standard NP span, and relaxed evaluation setting, where the system NP span overlaps the gold standard NP span. 4.2 Results Our best run evaluated at F-measure on the training data and F- measure on the test data under the strict evaluation setting; under the relaxed evaluation setting, it obtained an F-measure on the training data and F-measure on the test data. In general, the system precision was higher than the system recall (0.814 precision on the test data vs recall under strict evaluation, and precision vs recall on the test data under relaxed evaluation). The system recall is lower as we did not handle the non-contiguous NPs that accounted for approx. 10% of the training and test data. Table 2. System results on training and test data sets under strict and relaxed evaluation settings. Training Test Precision Recall F measure Precision Recall F measure Strict Relaxed Discussion The 2013 CLEF-eHEALTH Task 1a is the second challenge after the 2010 i2b2/va Shared-Task aiming to identify disorder named entities in clinical text. Even though the final goals of the two challenges were similar, they differed in several structural points. First, the 2010 i2b2/va corpus had token-based annotations and was already segmented into tokens, thus requiring no further pre-processing. In contrast, the 2013 CLEF-eHEALTH corpus marked annotations at the character level and pre-processing was desirable, which motivated the first steps of our pipeline. Secondly, entity boundaries were defined differently in the two challenges. A first example is by the determiners which were included as part of the annotation in the 2010 i2b2/va Shared-Task but were excluded in the 2013 CLEF-eHEALTH challenge (e.g., a brain tumor vs brain tumor). A second example is the non-contiguous entities included only in the 2013 CLEF-eHEALTH challenge (e.g., given the EMR sentence The pain reported by the patient is occurring in lower back, the annotated entity is pain... in lower back ). A final difference between the two challenges is the corpus size: approx. 18,550 problem entities in the i2b2 training corpus, compared to 5,874 disorder entities in CLEF-eHEALTH training corpus.
7 Supervised Named-Entity Extraction 7 Error analysis on the training data revealed some systematic mistakes performed by our NER model. The majority of the incorrectly predicted entities were NPs with morphological structure resembling the one of the disorder NPs (e.g., anastomosis contains the Greek suffix -osis meaning abnormal condition and resembles the name of several disorders like necrosis, osteoporosis, but can be both a disorder and a procedure), disorder names used as findings (e.g., the noun phrase esophageal varices is a finding based on the context which showed non-bleeding grade III esophageal varices), and disorder entities used in a negated context (e.g., NP fasciculations inside the context tongue midline without fasciculations). The system also predicted parts of NPs it was trained on as being standalone entities; for example, the phrase left ventricular was predicted as a disorder entity after the system encountered the phrase left ventricular aneurysm during training. In general, our NER system failed to identify the non-contiguous named entities (it could at best identify some of their parts), abbreviations or NPs rarely encountered in the training set (e.g., aaa, 3vd, inability to walk), misspelled disorder entities (e.g., hematochezeia) and the full span of several NPs longer than 2 tokens (e.g., chronic subdural hematoma). Out-of-vocabulary tokens, i.e., terms not found in the UMLS because of lack of coverage, variants, or misspellings, were an important source of lack of recall. 5 Conclusion and perspectives We present a clinical NER system designed for participation in Task 1a of the 2013 CLEF-eHEALTH challenge. We design our system as a CRF with a rich feature set and external knowledge gathered from specialized terminologies and general domain knowledge repositories. Our system evaluates at F-measure in the strict evaluation context and F-measure in the relaxed evaluation context, obtaining a mid-range position. Our entity detection system presents good precision but performs worse in terms of recall. In order to improve system recall, additional textual data can be integrated into the CRF model. We expect that including the Brown word clustering information as part of the feature vector would provide a fallback for some out-of-vocabulary tokens and help increase accuracy based on its unsupervised word classes. The non-contiguous entities are only partially handled by our system, thus better handling of all parts of non-contiguous entities, for instance through syntactic dependencies, should result in improved recall. 6 Acknowledgments This work was partly funded through project Accordys 2 funded by ANR under grant number ANR-12-CORD The first author was funded by the 2 Accordys: Agrégation de Contenus et de COnnaissances pour Raisonner à partir de cas de DYSmorphologie fœtale, Content and Knowledge Aggregation for Case-based Reasoning in the field of Fetal Dysmorphology (ANR ).
8 8 Andreea Bodnari et al. Châteaubriand Science Fellowship We acknowledge the Shared Annotated Resources (ShARe) project funded by the United States National Institutes of Health with grant number R01GM References 1. ACM. SemEval Portal. Portal. Accessed July 19, Timothy G. Armstrong, Alistair Moffat, William Webber, and Justin Zobel. Improvements that don t add up: ad-hoc retrieval results since In Proceedings of the 18th ACM conference on Information and knowledge management, CIKM 09, pages , New York, NY, USA, ACM. 3. Alan A. Aronson. Effective mapping of biomedical text to the UMLS Metathesaurus: the Metamap program. In Proceedings of the AMIA Annual Symposium, pages American Medical Informatics Association, Olivier Bodenreider. The Unified Medical Language System (UMLS): integrating biomedical terminology. Nucleic Acid Res, 32:D267D270, CoNLL. CoNLL: the conference of SIGNLL. Accessed July 19, Thomas Lavergne, Olivier Cappé, and François Yvon. Practical very large scale CRFs. In ACL Proc, pages , Shared Annotated Resources. Shared Annotated Resources. clinicalnlpannotation.org/index.php/main_page. Accessed May 24, M. Saeed, M. Villarroel, A.T. Reisner, G. Clifford, L. Lehman, G.B. Moody, T. Heldt, T.H. Kyaw, B.E. Moody, and R.G. Mark. Multiparameter intelligent monitoring in intensive care II (MIMIC-II): A public-access ICU database. Clinical Care Medicine, 39: , Guergana K Savova, James J Masanz, Philip V Ogren, Jiaping Zheng, Sunghwan Sohn, Karin C Kipper-Schuler, and Christopher G Chute. Mayo clinical Text Analysis and Knowledge Extraction System (ctakes): architecture, component evaluation and applications. Journal of American Medical Informatics Association, 17: , Hanna Suominen, Sanna Salanterä, Wendy W. Sumitra Velupillai Chapman, Guergana Savova, Noemie Elhadad, Sameer Pradhan, Brett R. South, Danielle L. Mowery, Gareth J.F. Jones, Johannes Leveling, Liadh Kelly, Lorraine Goeuriot, David Martinez, and Guido Zuccon. Overview of the ShARe/CLEF ehealth evaluation lab In Proceedings of CLEF 2013, Lecture Notes in Computer Science, Berlin Heidelberg, Springer. To appear. 11. Özlem Uzuner. Recognizing obesity and comorbidities in sparse data. Journal of American Medical Informatics Association, 16(4): , Aug Özlem Uzuner, Andreea Bodnari, Shuying Shen, Tyler Forbush, John Pestian, and Brett South. Evaluating the state of the art in coreference resolution for electronic medical records. Journal of American Medical Informatics Association, 17: , February Özlem Uzuner, Brett R. South, Shuying Shen, and Scott L. DuVall i2b2/va challenge on concepts, assertions, and relations in clinical text. Journal of the American Medical Informatics Association, 18(5): , 2011.
A Supervised Abbreviation Resolution System for Medical Text
 A Supervised Abbreviation Resolution System for Medical Text Pierre Zweigenbaum 1, Louise Deléger 1, Thomas Lavergne 1, Aurélie Névéol 1, and Andreea Bodnari 1,2 1 LIMSI-CNRS, rue John von Neumann, F-91400
A Supervised Abbreviation Resolution System for Medical Text Pierre Zweigenbaum 1, Louise Deléger 1, Thomas Lavergne 1, Aurélie Névéol 1, and Andreea Bodnari 1,2 1 LIMSI-CNRS, rue John von Neumann, F-91400
TMUNSW: Identification of disorders and normalization to SNOMED-CT terminology in unstructured clinical notes
 TMUNSW: Identification of disorders and normalization to SNOMED-CT terminology in unstructured clinical notes Jitendra Jonnagaddala a,b,c Siaw-Teng Liaw *,a Pradeep Ray b Manish Kumar c School of Public
TMUNSW: Identification of disorders and normalization to SNOMED-CT terminology in unstructured clinical notes Jitendra Jonnagaddala a,b,c Siaw-Teng Liaw *,a Pradeep Ray b Manish Kumar c School of Public
Identify Disorders in Health Records using Conditional Random Fields and Metamap
 Identify Disorders in Health Records using Conditional Random Fields and Metamap AEHRC at ShARe/CLEF 2013 ehealth Evaluation Lab Task 1 G. Zuccon 1, A. Holloway 1,2, B. Koopman 1,2, A. Nguyen 1 1 The Australian
Identify Disorders in Health Records using Conditional Random Fields and Metamap AEHRC at ShARe/CLEF 2013 ehealth Evaluation Lab Task 1 G. Zuccon 1, A. Holloway 1,2, B. Koopman 1,2, A. Nguyen 1 1 The Australian
Recognizing and Encoding Disorder Concepts in Clinical Text using Machine Learning and Vector Space Model *
 Recognizing and Encoding Disorder Concepts in Clinical Text using Machine Learning and Vector Space Model * Buzhou Tang 1,2, Yonghui Wu 1, Min Jiang 1, Joshua C. Denny 3, and Hua Xu 1,* 1 School of Biomedical
Recognizing and Encoding Disorder Concepts in Clinical Text using Machine Learning and Vector Space Model * Buzhou Tang 1,2, Yonghui Wu 1, Min Jiang 1, Joshua C. Denny 3, and Hua Xu 1,* 1 School of Biomedical
Generating Patient Problem Lists from the ShARe Corpus using SNOMED CT/SNOMED CT CORE Problem List
 Generating Patient Problem Lists from the ShARe Corpus using SNOMED CT/SNOMED CT CORE Problem List Danielle Mowery Janyce Wiebe University of Pittsburgh Pittsburgh, PA [email protected] [email protected]
Generating Patient Problem Lists from the ShARe Corpus using SNOMED CT/SNOMED CT CORE Problem List Danielle Mowery Janyce Wiebe University of Pittsburgh Pittsburgh, PA [email protected] [email protected]
Travis Goodwin & Sanda Harabagiu
 Automatic Generation of a Qualified Medical Knowledge Graph and its Usage for Retrieving Patient Cohorts from Electronic Medical Records Travis Goodwin & Sanda Harabagiu Human Language Technology Research
Automatic Generation of a Qualified Medical Knowledge Graph and its Usage for Retrieving Patient Cohorts from Electronic Medical Records Travis Goodwin & Sanda Harabagiu Human Language Technology Research
Integrating Public and Private Medical Texts for Patient De-Identification with Apache ctakes
 Integrating Public and Private Medical Texts for Patient De-Identification with Apache ctakes Presented By: Andrew McMurry & Britt Fitch (Apache ctakes committers) Co-authors: Guergana Savova, Ben Reis,
Integrating Public and Private Medical Texts for Patient De-Identification with Apache ctakes Presented By: Andrew McMurry & Britt Fitch (Apache ctakes committers) Co-authors: Guergana Savova, Ben Reis,
A Method for Automatic De-identification of Medical Records
 A Method for Automatic De-identification of Medical Records Arya Tafvizi MIT CSAIL Cambridge, MA 0239, USA [email protected] Maciej Pacula MIT CSAIL Cambridge, MA 0239, USA [email protected] Abstract
A Method for Automatic De-identification of Medical Records Arya Tafvizi MIT CSAIL Cambridge, MA 0239, USA [email protected] Maciej Pacula MIT CSAIL Cambridge, MA 0239, USA [email protected] Abstract
Word Completion and Prediction in Hebrew
 Experiments with Language Models for בס"ד Word Completion and Prediction in Hebrew 1 Yaakov HaCohen-Kerner, Asaf Applebaum, Jacob Bitterman Department of Computer Science Jerusalem College of Technology
Experiments with Language Models for בס"ד Word Completion and Prediction in Hebrew 1 Yaakov HaCohen-Kerner, Asaf Applebaum, Jacob Bitterman Department of Computer Science Jerusalem College of Technology
Automated Problem List Generation from Electronic Medical Records in IBM Watson
 Proceedings of the Twenty-Seventh Conference on Innovative Applications of Artificial Intelligence Automated Problem List Generation from Electronic Medical Records in IBM Watson Murthy Devarakonda, Ching-Huei
Proceedings of the Twenty-Seventh Conference on Innovative Applications of Artificial Intelligence Automated Problem List Generation from Electronic Medical Records in IBM Watson Murthy Devarakonda, Ching-Huei
Annotating Medical Forms using UMLS
 Annotating Medical Forms using UMLS Victor Christen 1, Anika Groß 1, Julian Varghese 2, Martin Dugas 2, Erhard Rahm 1 1 Department of Computer Science, Universität Leipzig, Germany 2 Institute of Medical
Annotating Medical Forms using UMLS Victor Christen 1, Anika Groß 1, Julian Varghese 2, Martin Dugas 2, Erhard Rahm 1 1 Department of Computer Science, Universität Leipzig, Germany 2 Institute of Medical
Micro blogs Oriented Word Segmentation System
 Micro blogs Oriented Word Segmentation System Yijia Liu, Meishan Zhang, Wanxiang Che, Ting Liu, Yihe Deng Research Center for Social Computing and Information Retrieval Harbin Institute of Technology,
Micro blogs Oriented Word Segmentation System Yijia Liu, Meishan Zhang, Wanxiang Che, Ting Liu, Yihe Deng Research Center for Social Computing and Information Retrieval Harbin Institute of Technology,
Open Domain Information Extraction. Günter Neumann, DFKI, 2012
 Open Domain Information Extraction Günter Neumann, DFKI, 2012 Improving TextRunner Wu and Weld (2010) Open Information Extraction using Wikipedia, ACL 2010 Fader et al. (2011) Identifying Relations for
Open Domain Information Extraction Günter Neumann, DFKI, 2012 Improving TextRunner Wu and Weld (2010) Open Information Extraction using Wikipedia, ACL 2010 Fader et al. (2011) Identifying Relations for
Free Text Phrase Encoding and Information Extraction from Medical Notes. Jennifer Shu
 Free Text Phrase Encoding and Information Extraction from Medical Notes by Jennifer Shu Submitted to the Department of Electrical Engineering and Computer Science in partial fulfillment of the requirements
Free Text Phrase Encoding and Information Extraction from Medical Notes by Jennifer Shu Submitted to the Department of Electrical Engineering and Computer Science in partial fulfillment of the requirements
POSBIOTM-NER: A Machine Learning Approach for. Bio-Named Entity Recognition
 POSBIOTM-NER: A Machine Learning Approach for Bio-Named Entity Recognition Yu Song, Eunji Yi, Eunju Kim, Gary Geunbae Lee, Department of CSE, POSTECH, Pohang, Korea 790-784 Soo-Jun Park Bioinformatics
POSBIOTM-NER: A Machine Learning Approach for Bio-Named Entity Recognition Yu Song, Eunji Yi, Eunju Kim, Gary Geunbae Lee, Department of CSE, POSTECH, Pohang, Korea 790-784 Soo-Jun Park Bioinformatics
A De-identifier For Electronic Medical Records Based On A Heterogeneous Feature Set. Arya Tafvizi
 A De-identifier For Electronic Medical Records Based On A Heterogeneous Feature Set by Arya Tafvizi S.B., Physics, MIT, 2010 S.B., Computer Science and Engineering, MIT, 2011 Submitted to the Department
A De-identifier For Electronic Medical Records Based On A Heterogeneous Feature Set by Arya Tafvizi S.B., Physics, MIT, 2010 S.B., Computer Science and Engineering, MIT, 2011 Submitted to the Department
Natural Language Processing for Clinical Informatics and Translational Research Informatics
 Natural Language Processing for Clinical Informatics and Translational Research Informatics Imre Solti, M. D., Ph. D. [email protected] K99 Fellow in Biomedical Informatics University of Washington Background
Natural Language Processing for Clinical Informatics and Translational Research Informatics Imre Solti, M. D., Ph. D. [email protected] K99 Fellow in Biomedical Informatics University of Washington Background
A Knowledge-Poor Approach to BioCreative V DNER and CID Tasks
 A Knowledge-Poor Approach to BioCreative V DNER and CID Tasks Firoj Alam 1, Anna Corazza 2, Alberto Lavelli 3, and Roberto Zanoli 3 1 Dept. of Information Eng. and Computer Science, University of Trento,
A Knowledge-Poor Approach to BioCreative V DNER and CID Tasks Firoj Alam 1, Anna Corazza 2, Alberto Lavelli 3, and Roberto Zanoli 3 1 Dept. of Information Eng. and Computer Science, University of Trento,
ANNLOR: A Naïve Notation-system for Lexical Outputs Ranking
 ANNLOR: A Naïve Notation-system for Lexical Outputs Ranking Anne-Laure Ligozat LIMSI-CNRS/ENSIIE rue John von Neumann 91400 Orsay, France [email protected] Cyril Grouin LIMSI-CNRS rue John von Neumann 91400
ANNLOR: A Naïve Notation-system for Lexical Outputs Ranking Anne-Laure Ligozat LIMSI-CNRS/ENSIIE rue John von Neumann 91400 Orsay, France [email protected] Cyril Grouin LIMSI-CNRS rue John von Neumann 91400
Ngram Search Engine with Patterns Combining Token, POS, Chunk and NE Information
 Ngram Search Engine with Patterns Combining Token, POS, Chunk and NE Information Satoshi Sekine Computer Science Department New York University [email protected] Kapil Dalwani Computer Science Department
Ngram Search Engine with Patterns Combining Token, POS, Chunk and NE Information Satoshi Sekine Computer Science Department New York University [email protected] Kapil Dalwani Computer Science Department
Annotation and Extraction of Relations from Italian Medical Records
 Annotation and Extraction of Relations from Italian Medical Records Giuseppe Attardi, Vittoria Cozza, Daniele Sartiano Dipartimento di Informatica Università di Pisa Largo B. Pontecorvo, 3 I-56127 Pisa,
Annotation and Extraction of Relations from Italian Medical Records Giuseppe Attardi, Vittoria Cozza, Daniele Sartiano Dipartimento di Informatica Università di Pisa Largo B. Pontecorvo, 3 I-56127 Pisa,
Extracting Clinical entities and their assertions from Chinese Electronic Medical Records Based on Machine Learning
 3rd International Conference on Materials Engineering, Manufacturing Technology and Control (ICMEMTC 2016) Extracting Clinical entities and their assertions from Chinese Electronic Medical Records Based
3rd International Conference on Materials Engineering, Manufacturing Technology and Control (ICMEMTC 2016) Extracting Clinical entities and their assertions from Chinese Electronic Medical Records Based
Automated Content Analysis of Discussion Transcripts
 Automated Content Analysis of Discussion Transcripts Vitomir Kovanović [email protected] Dragan Gašević [email protected] School of Informatics, University of Edinburgh Edinburgh, United Kingdom [email protected]
Automated Content Analysis of Discussion Transcripts Vitomir Kovanović [email protected] Dragan Gašević [email protected] School of Informatics, University of Edinburgh Edinburgh, United Kingdom [email protected]
Structuring Unstructured Clinical Narratives in OpenMRS with Medical Concept Extraction
 Structuring Unstructured Clinical Narratives in OpenMRS with Medical Concept Extraction Ryan M Eshleman, Hui Yang, and Barry Levine [email protected], [email protected], [email protected] Department
Structuring Unstructured Clinical Narratives in OpenMRS with Medical Concept Extraction Ryan M Eshleman, Hui Yang, and Barry Levine [email protected], [email protected], [email protected] Department
Terminology Extraction from Log Files
 Terminology Extraction from Log Files Hassan Saneifar 1,2, Stéphane Bonniol 2, Anne Laurent 1, Pascal Poncelet 1, and Mathieu Roche 1 1 LIRMM - Université Montpellier 2 - CNRS 161 rue Ada, 34392 Montpellier
Terminology Extraction from Log Files Hassan Saneifar 1,2, Stéphane Bonniol 2, Anne Laurent 1, Pascal Poncelet 1, and Mathieu Roche 1 1 LIRMM - Université Montpellier 2 - CNRS 161 rue Ada, 34392 Montpellier
11-792 Software Engineering EMR Project Report
 11-792 Software Engineering EMR Project Report Team Members Phani Gadde Anika Gupta Ting-Hao (Kenneth) Huang Chetan Thayur Suyoun Kim Vision Our aim is to build an intelligent system which is capable of
11-792 Software Engineering EMR Project Report Team Members Phani Gadde Anika Gupta Ting-Hao (Kenneth) Huang Chetan Thayur Suyoun Kim Vision Our aim is to build an intelligent system which is capable of
Architecture of an Ontology-Based Domain- Specific Natural Language Question Answering System
 Architecture of an Ontology-Based Domain- Specific Natural Language Question Answering System Athira P. M., Sreeja M. and P. C. Reghuraj Department of Computer Science and Engineering, Government Engineering
Architecture of an Ontology-Based Domain- Specific Natural Language Question Answering System Athira P. M., Sreeja M. and P. C. Reghuraj Department of Computer Science and Engineering, Government Engineering
Predicting Chief Complaints at Triage Time in the Emergency Department
 Predicting Chief Complaints at Triage Time in the Emergency Department Yacine Jernite, Yoni Halpern New York University New York, NY {jernite,halpern}@cs.nyu.edu Steven Horng Beth Israel Deaconess Medical
Predicting Chief Complaints at Triage Time in the Emergency Department Yacine Jernite, Yoni Halpern New York University New York, NY {jernite,halpern}@cs.nyu.edu Steven Horng Beth Israel Deaconess Medical
Combining structured data with machine learning to improve clinical text de-identification
 Combining structured data with machine learning to improve clinical text de-identification DT Tran Scott Halgrim David Carrell Group Health Research Institute Clinical text contains Personally identifiable
Combining structured data with machine learning to improve clinical text de-identification DT Tran Scott Halgrim David Carrell Group Health Research Institute Clinical text contains Personally identifiable
Interactive Dynamic Information Extraction
 Interactive Dynamic Information Extraction Kathrin Eichler, Holmer Hemsen, Markus Löckelt, Günter Neumann, and Norbert Reithinger Deutsches Forschungszentrum für Künstliche Intelligenz - DFKI, 66123 Saarbrücken
Interactive Dynamic Information Extraction Kathrin Eichler, Holmer Hemsen, Markus Löckelt, Günter Neumann, and Norbert Reithinger Deutsches Forschungszentrum für Künstliche Intelligenz - DFKI, 66123 Saarbrücken
Special Topics in Computer Science
 Special Topics in Computer Science NLP in a Nutshell CS492B Spring Semester 2009 Jong C. Park Computer Science Department Korea Advanced Institute of Science and Technology INTRODUCTION Jong C. Park, CS
Special Topics in Computer Science NLP in a Nutshell CS492B Spring Semester 2009 Jong C. Park Computer Science Department Korea Advanced Institute of Science and Technology INTRODUCTION Jong C. Park, CS
Workshop. Neil Barrett PhD, Jens Weber PhD, Vincent Thai MD. Engineering & Health Informa2on Science
 Engineering & Health Informa2on Science Engineering NLP Solu/ons for Structured Informa/on from Clinical Text: Extrac'ng Sen'nel Events from Pallia've Care Consult Le8ers Canada-China Clean Energy Initiative
Engineering & Health Informa2on Science Engineering NLP Solu/ons for Structured Informa/on from Clinical Text: Extrac'ng Sen'nel Events from Pallia've Care Consult Le8ers Canada-China Clean Energy Initiative
LABERINTO at ImageCLEF 2011 Medical Image Retrieval Task
 LABERINTO at ImageCLEF 2011 Medical Image Retrieval Task Jacinto Mata, Mariano Crespo, Manuel J. Maña Dpto. de Tecnologías de la Información. Universidad de Huelva Ctra. Huelva - Palos de la Frontera s/n.
LABERINTO at ImageCLEF 2011 Medical Image Retrieval Task Jacinto Mata, Mariano Crespo, Manuel J. Maña Dpto. de Tecnologías de la Información. Universidad de Huelva Ctra. Huelva - Palos de la Frontera s/n.
Terminology Extraction from Log Files
 Terminology Extraction from Log Files Hassan Saneifar, Stéphane Bonniol, Anne Laurent, Pascal Poncelet, Mathieu Roche To cite this version: Hassan Saneifar, Stéphane Bonniol, Anne Laurent, Pascal Poncelet,
Terminology Extraction from Log Files Hassan Saneifar, Stéphane Bonniol, Anne Laurent, Pascal Poncelet, Mathieu Roche To cite this version: Hassan Saneifar, Stéphane Bonniol, Anne Laurent, Pascal Poncelet,
Document Image Retrieval using Signatures as Queries
 Document Image Retrieval using Signatures as Queries Sargur N. Srihari, Shravya Shetty, Siyuan Chen, Harish Srinivasan, Chen Huang CEDAR, University at Buffalo(SUNY) Amherst, New York 14228 Gady Agam and
Document Image Retrieval using Signatures as Queries Sargur N. Srihari, Shravya Shetty, Siyuan Chen, Harish Srinivasan, Chen Huang CEDAR, University at Buffalo(SUNY) Amherst, New York 14228 Gady Agam and
Automated assignment of ICD-9-CM codes to radiology reports
 Automated assignment of ICD-9-CM codes to radiology reports Richárd Farkas University of Szeged Filip Ginter University of Turku Overview Why clinical coding? Importance, use of automated coding Challenge
Automated assignment of ICD-9-CM codes to radiology reports Richárd Farkas University of Szeged Filip Ginter University of Turku Overview Why clinical coding? Importance, use of automated coding Challenge
Clinic + - A Clinical Decision Support System Using Association Rule Mining
 Clinic + - A Clinical Decision Support System Using Association Rule Mining Sangeetha Santhosh, Mercelin Francis M.Tech Student, Dept. of CSE., Marian Engineering College, Kerala University, Trivandrum,
Clinic + - A Clinical Decision Support System Using Association Rule Mining Sangeetha Santhosh, Mercelin Francis M.Tech Student, Dept. of CSE., Marian Engineering College, Kerala University, Trivandrum,
Semantic annotation of requirements for automatic UML class diagram generation
 www.ijcsi.org 259 Semantic annotation of requirements for automatic UML class diagram generation Soumaya Amdouni 1, Wahiba Ben Abdessalem Karaa 2 and Sondes Bouabid 3 1 University of tunis High Institute
www.ijcsi.org 259 Semantic annotation of requirements for automatic UML class diagram generation Soumaya Amdouni 1, Wahiba Ben Abdessalem Karaa 2 and Sondes Bouabid 3 1 University of tunis High Institute
An Information Extraction Framework for Cohort Identification Using Electronic Health Records
 An Information Extraction Framework for Cohort Identification Using Electronic Health Records Hongfang Liu PhD 1, Suzette J. Bielinski PhD 1, Sunghwan Sohn PhD 1, Sean Murphy 1, Kavishwar B. Wagholikar
An Information Extraction Framework for Cohort Identification Using Electronic Health Records Hongfang Liu PhD 1, Suzette J. Bielinski PhD 1, Sunghwan Sohn PhD 1, Sean Murphy 1, Kavishwar B. Wagholikar
Medical-Miner at TREC 2011 Medical Records Track
 Medical-Miner at TREC 2011 Medical Records Track 1 J.M. Córdoba, 1 M.J. Maña, 1 N.P. Cruz, 1 J. Mata, 2 F. Aparicio, 2 M. Buenaga, 3 D. Glez-Peña, 3 F. Fdez-Riverola 1 Universidad de Huelva 2 Universidad
Medical-Miner at TREC 2011 Medical Records Track 1 J.M. Córdoba, 1 M.J. Maña, 1 N.P. Cruz, 1 J. Mata, 2 F. Aparicio, 2 M. Buenaga, 3 D. Glez-Peña, 3 F. Fdez-Riverola 1 Universidad de Huelva 2 Universidad
BANNER: AN EXECUTABLE SURVEY OF ADVANCES IN BIOMEDICAL NAMED ENTITY RECOGNITION
 BANNER: AN EXECUTABLE SURVEY OF ADVANCES IN BIOMEDICAL NAMED ENTITY RECOGNITION ROBERT LEAMAN Department of Computer Science and Engineering, Arizona State University GRACIELA GONZALEZ * Department of
BANNER: AN EXECUTABLE SURVEY OF ADVANCES IN BIOMEDICAL NAMED ENTITY RECOGNITION ROBERT LEAMAN Department of Computer Science and Engineering, Arizona State University GRACIELA GONZALEZ * Department of
Find the signal in the noise
 Find the signal in the noise Electronic Health Records: The challenge The adoption of Electronic Health Records (EHRs) in the USA is rapidly increasing, due to the Health Information Technology and Clinical
Find the signal in the noise Electronic Health Records: The challenge The adoption of Electronic Health Records (EHRs) in the USA is rapidly increasing, due to the Health Information Technology and Clinical
SVM Based Learning System For Information Extraction
 SVM Based Learning System For Information Extraction Yaoyong Li, Kalina Bontcheva, and Hamish Cunningham Department of Computer Science, The University of Sheffield, Sheffield, S1 4DP, UK {yaoyong,kalina,hamish}@dcs.shef.ac.uk
SVM Based Learning System For Information Extraction Yaoyong Li, Kalina Bontcheva, and Hamish Cunningham Department of Computer Science, The University of Sheffield, Sheffield, S1 4DP, UK {yaoyong,kalina,hamish}@dcs.shef.ac.uk
Protein-protein Interaction Passage Extraction Using the Interaction Pattern Kernel Approach for the BioCreative 2015 BioC Track
 Protein-protein Interaction Passage Extraction Using the Interaction Pattern Kernel Approach for the BioCreative 2015 BioC Track Yung-Chun Chang 1,2, Yu-Chen Su 3, Chun-Han Chu 1, Chien Chin Chen 2 and
Protein-protein Interaction Passage Extraction Using the Interaction Pattern Kernel Approach for the BioCreative 2015 BioC Track Yung-Chun Chang 1,2, Yu-Chen Su 3, Chun-Han Chu 1, Chien Chin Chen 2 and
Unsupervised Extraction of Diagnosis Codes from EMRs Using Knowledge-Based and Extractive Text Summarization Techniques
 Unsupervised Extraction of Diagnosis Codes from EMRs Using Knowledge-Based and Extractive Text Summarization Techniques Ramakanth Kavuluru 1,2, Sifei Han 2, and Daniel Harris 2 1 Division of Biomedical
Unsupervised Extraction of Diagnosis Codes from EMRs Using Knowledge-Based and Extractive Text Summarization Techniques Ramakanth Kavuluru 1,2, Sifei Han 2, and Daniel Harris 2 1 Division of Biomedical
Identifying Focus, Techniques and Domain of Scientific Papers
 Identifying Focus, Techniques and Domain of Scientific Papers Sonal Gupta Department of Computer Science Stanford University Stanford, CA 94305 [email protected] Christopher D. Manning Department of
Identifying Focus, Techniques and Domain of Scientific Papers Sonal Gupta Department of Computer Science Stanford University Stanford, CA 94305 [email protected] Christopher D. Manning Department of
A Mixed Trigrams Approach for Context Sensitive Spell Checking
 A Mixed Trigrams Approach for Context Sensitive Spell Checking Davide Fossati and Barbara Di Eugenio Department of Computer Science University of Illinois at Chicago Chicago, IL, USA [email protected], [email protected]
A Mixed Trigrams Approach for Context Sensitive Spell Checking Davide Fossati and Barbara Di Eugenio Department of Computer Science University of Illinois at Chicago Chicago, IL, USA [email protected], [email protected]
Software Architecture Document
 Software Architecture Document Natural Language Processing Cell Version 1.0 Natural Language Processing Cell Software Architecture Document Version 1.0 1 1. Table of Contents 1. Table of Contents... 2
Software Architecture Document Natural Language Processing Cell Version 1.0 Natural Language Processing Cell Software Architecture Document Version 1.0 1 1. Table of Contents 1. Table of Contents... 2
Efficient Techniques for Improved Data Classification and POS Tagging by Monitoring Extraction, Pruning and Updating of Unknown Foreign Words
 , pp.290-295 http://dx.doi.org/10.14257/astl.2015.111.55 Efficient Techniques for Improved Data Classification and POS Tagging by Monitoring Extraction, Pruning and Updating of Unknown Foreign Words Irfan
, pp.290-295 http://dx.doi.org/10.14257/astl.2015.111.55 Efficient Techniques for Improved Data Classification and POS Tagging by Monitoring Extraction, Pruning and Updating of Unknown Foreign Words Irfan
PPInterFinder A Web Server for Mining Human Protein Protein Interaction
 PPInterFinder A Web Server for Mining Human Protein Protein Interaction Kalpana Raja, Suresh Subramani, Jeyakumar Natarajan Data Mining and Text Mining Laboratory, Department of Bioinformatics, Bharathiar
PPInterFinder A Web Server for Mining Human Protein Protein Interaction Kalpana Raja, Suresh Subramani, Jeyakumar Natarajan Data Mining and Text Mining Laboratory, Department of Bioinformatics, Bharathiar
Search and Data Mining: Techniques. Text Mining Anya Yarygina Boris Novikov
 Search and Data Mining: Techniques Text Mining Anya Yarygina Boris Novikov Introduction Generally used to denote any system that analyzes large quantities of natural language text and detects lexical or
Search and Data Mining: Techniques Text Mining Anya Yarygina Boris Novikov Introduction Generally used to denote any system that analyzes large quantities of natural language text and detects lexical or
Introduction to Data Mining
 Introduction to Data Mining Jay Urbain Credits: Nazli Goharian & David Grossman @ IIT Outline Introduction Data Pre-processing Data Mining Algorithms Naïve Bayes Decision Tree Neural Network Association
Introduction to Data Mining Jay Urbain Credits: Nazli Goharian & David Grossman @ IIT Outline Introduction Data Pre-processing Data Mining Algorithms Naïve Bayes Decision Tree Neural Network Association
Ask your Database: Natural Language Processing using In-Memory Technology
 Enterprise Platform and Integration Concepts Master Project Summer Term 2015 Ask your Database: Natural Language Processing using In-Memory Technology Dr. Mariana Neves April 10th, 2015 Question Answering
Enterprise Platform and Integration Concepts Master Project Summer Term 2015 Ask your Database: Natural Language Processing using In-Memory Technology Dr. Mariana Neves April 10th, 2015 Question Answering
Stanford s Distantly-Supervised Slot-Filling System
 Stanford s Distantly-Supervised Slot-Filling System Mihai Surdeanu, Sonal Gupta, John Bauer, David McClosky, Angel X. Chang, Valentin I. Spitkovsky, Christopher D. Manning Computer Science Department,
Stanford s Distantly-Supervised Slot-Filling System Mihai Surdeanu, Sonal Gupta, John Bauer, David McClosky, Angel X. Chang, Valentin I. Spitkovsky, Christopher D. Manning Computer Science Department,
MIRACLE at VideoCLEF 2008: Classification of Multilingual Speech Transcripts
 MIRACLE at VideoCLEF 2008: Classification of Multilingual Speech Transcripts Julio Villena-Román 1,3, Sara Lana-Serrano 2,3 1 Universidad Carlos III de Madrid 2 Universidad Politécnica de Madrid 3 DAEDALUS
MIRACLE at VideoCLEF 2008: Classification of Multilingual Speech Transcripts Julio Villena-Román 1,3, Sara Lana-Serrano 2,3 1 Universidad Carlos III de Madrid 2 Universidad Politécnica de Madrid 3 DAEDALUS
Transformation of Free-text Electronic Health Records for Efficient Information Retrieval and Support of Knowledge Discovery
 Transformation of Free-text Electronic Health Records for Efficient Information Retrieval and Support of Knowledge Discovery Jan Paralic, Peter Smatana Technical University of Kosice, Slovakia Center for
Transformation of Free-text Electronic Health Records for Efficient Information Retrieval and Support of Knowledge Discovery Jan Paralic, Peter Smatana Technical University of Kosice, Slovakia Center for
Clustering Connectionist and Statistical Language Processing
 Clustering Connectionist and Statistical Language Processing Frank Keller [email protected] Computerlinguistik Universität des Saarlandes Clustering p.1/21 Overview clustering vs. classification supervised
Clustering Connectionist and Statistical Language Processing Frank Keller [email protected] Computerlinguistik Universität des Saarlandes Clustering p.1/21 Overview clustering vs. classification supervised
Processing Electronic Medical Records. Master thesis. Lars-Erik Bruce. Ontology-Driven Information Extraction and Structuring in the Clinical Domain
 UNIVERSITY OF OSLO Department of Informatics Processing Electronic Medical Records Ontology-Driven Information Extraction and Structuring in the Clinical Domain Master thesis Lars-Erik Bruce August 1,
UNIVERSITY OF OSLO Department of Informatics Processing Electronic Medical Records Ontology-Driven Information Extraction and Structuring in the Clinical Domain Master thesis Lars-Erik Bruce August 1,
Wikipedia and Web document based Query Translation and Expansion for Cross-language IR
 Wikipedia and Web document based Query Translation and Expansion for Cross-language IR Ling-Xiang Tang 1, Andrew Trotman 2, Shlomo Geva 1, Yue Xu 1 1Faculty of Science and Technology, Queensland University
Wikipedia and Web document based Query Translation and Expansion for Cross-language IR Ling-Xiang Tang 1, Andrew Trotman 2, Shlomo Geva 1, Yue Xu 1 1Faculty of Science and Technology, Queensland University
Parsing Software Requirements with an Ontology-based Semantic Role Labeler
 Parsing Software Requirements with an Ontology-based Semantic Role Labeler Michael Roth University of Edinburgh [email protected] Ewan Klein University of Edinburgh [email protected] Abstract Software
Parsing Software Requirements with an Ontology-based Semantic Role Labeler Michael Roth University of Edinburgh [email protected] Ewan Klein University of Edinburgh [email protected] Abstract Software
Sentiment analysis on tweets in a financial domain
 Sentiment analysis on tweets in a financial domain Jasmina Smailović 1,2, Miha Grčar 1, Martin Žnidaršič 1 1 Dept of Knowledge Technologies, Jožef Stefan Institute, Ljubljana, Slovenia 2 Jožef Stefan International
Sentiment analysis on tweets in a financial domain Jasmina Smailović 1,2, Miha Grčar 1, Martin Žnidaršič 1 1 Dept of Knowledge Technologies, Jožef Stefan Institute, Ljubljana, Slovenia 2 Jožef Stefan International
Term extraction for user profiling: evaluation by the user
 Term extraction for user profiling: evaluation by the user Suzan Verberne 1, Maya Sappelli 1,2, Wessel Kraaij 1,2 1 Institute for Computing and Information Sciences, Radboud University Nijmegen 2 TNO,
Term extraction for user profiling: evaluation by the user Suzan Verberne 1, Maya Sappelli 1,2, Wessel Kraaij 1,2 1 Institute for Computing and Information Sciences, Radboud University Nijmegen 2 TNO,
PoS-tagging Italian texts with CORISTagger
 PoS-tagging Italian texts with CORISTagger Fabio Tamburini DSLO, University of Bologna, Italy [email protected] Abstract. This paper presents an evolution of CORISTagger [1], an high-performance
PoS-tagging Italian texts with CORISTagger Fabio Tamburini DSLO, University of Bologna, Italy [email protected] Abstract. This paper presents an evolution of CORISTagger [1], an high-performance
Developing VA GDx: An Informatics Platform to Capture and Integrate Genetic Diagnostic Testing Data into the VA Electronic Medical Record
 Developing VA GDx: An Informatics Platform to Capture and Integrate Genetic Diagnostic Testing Data into the VA Electronic Medical Record Scott L. DuVall Jun 27, 2014 1 Julie Lynch Vickie Venne Dawn Provenzale
Developing VA GDx: An Informatics Platform to Capture and Integrate Genetic Diagnostic Testing Data into the VA Electronic Medical Record Scott L. DuVall Jun 27, 2014 1 Julie Lynch Vickie Venne Dawn Provenzale
Probabilistic topic models for sentiment analysis on the Web
 University of Exeter Department of Computer Science Probabilistic topic models for sentiment analysis on the Web Chenghua Lin September 2011 Submitted by Chenghua Lin, to the the University of Exeter as
University of Exeter Department of Computer Science Probabilistic topic models for sentiment analysis on the Web Chenghua Lin September 2011 Submitted by Chenghua Lin, to the the University of Exeter as
VCU-TSA at Semeval-2016 Task 4: Sentiment Analysis in Twitter
 VCU-TSA at Semeval-2016 Task 4: Sentiment Analysis in Twitter Gerard Briones and Kasun Amarasinghe and Bridget T. McInnes, PhD. Department of Computer Science Virginia Commonwealth University Richmond,
VCU-TSA at Semeval-2016 Task 4: Sentiment Analysis in Twitter Gerard Briones and Kasun Amarasinghe and Bridget T. McInnes, PhD. Department of Computer Science Virginia Commonwealth University Richmond,
The American Academy of Ophthalmology Adopts SNOMED CT as its Official Clinical Terminology
 The American Academy of Ophthalmology Adopts SNOMED CT as its Official Clinical Terminology H. Dunbar Hoskins, Jr., M.D., P. Lloyd Hildebrand, M.D., Flora Lum, M.D. The road towards broad adoption of electronic
The American Academy of Ophthalmology Adopts SNOMED CT as its Official Clinical Terminology H. Dunbar Hoskins, Jr., M.D., P. Lloyd Hildebrand, M.D., Flora Lum, M.D. The road towards broad adoption of electronic
Tibetan-Chinese Bilingual Sentences Alignment Method based on Multiple Features
 , pp.273-280 http://dx.doi.org/10.14257/ijdta.2015.8.4.27 Tibetan-Chinese Bilingual Sentences Alignment Method based on Multiple Features Lirong Qiu School of Information Engineering, MinzuUniversity of
, pp.273-280 http://dx.doi.org/10.14257/ijdta.2015.8.4.27 Tibetan-Chinese Bilingual Sentences Alignment Method based on Multiple Features Lirong Qiu School of Information Engineering, MinzuUniversity of
Building a Question Classifier for a TREC-Style Question Answering System
 Building a Question Classifier for a TREC-Style Question Answering System Richard May & Ari Steinberg Topic: Question Classification We define Question Classification (QC) here to be the task that, given
Building a Question Classifier for a TREC-Style Question Answering System Richard May & Ari Steinberg Topic: Question Classification We define Question Classification (QC) here to be the task that, given
Data Deduplication in Slovak Corpora
 Ľ. Štúr Institute of Linguistics, Slovak Academy of Sciences, Bratislava, Slovakia Abstract. Our paper describes our experience in deduplication of a Slovak corpus. Two methods of deduplication a plain
Ľ. Štúr Institute of Linguistics, Slovak Academy of Sciences, Bratislava, Slovakia Abstract. Our paper describes our experience in deduplication of a Slovak corpus. Two methods of deduplication a plain
The Prolog Interface to the Unstructured Information Management Architecture
 The Prolog Interface to the Unstructured Information Management Architecture Paul Fodor 1, Adam Lally 2, David Ferrucci 2 1 Stony Brook University, Stony Brook, NY 11794, USA, [email protected] 2 IBM
The Prolog Interface to the Unstructured Information Management Architecture Paul Fodor 1, Adam Lally 2, David Ferrucci 2 1 Stony Brook University, Stony Brook, NY 11794, USA, [email protected] 2 IBM
Computer-assisted coding and natural language processing
 Computer-assisted coding and natural language processing Without changes to current coding technology and processes, ICD-10 adoption will be very difficult for providers to absorb, due to the added complexity
Computer-assisted coding and natural language processing Without changes to current coding technology and processes, ICD-10 adoption will be very difficult for providers to absorb, due to the added complexity
