topics about enzyme function:
|
|
|
- Jody Flowers
- 7 years ago
- Views:
Transcription
1 6 Enzymes
2 CHAPTER 6 Enzymes topics about enzyme function: Physiological significance of enzymes Origin of catalytic power of enzymes Chemical mechanisms of catalysis Mechanisms of chymotrypsin and lysozyme Description of enzyme kinetics and inhibition
3 What are enzymes? Enzymes are catalysts Increase reaction rates without being used up Most enzymes are globular proteins However, some RNA (ribozymes and ribosomal RNA) also catalyze reactions We will celebrate my inspiration, the Biochemist Louis Pasteur.
4 Why do cells evolve biocatalysis over inorganic catalysts? Greater reaction specificity: avoids side products Milder reaction conditions: conducive to conditions in cells Higher reaction rates: in a biologically useful timeframe Capacity for regulation: control of biological pathways COO - - COO OH - COO NH 2 O - COO Metabolites have many potential pathways of decomposition - COO NH 2 OH O - COO Chorismate mutase - OOC OH - COO O Enzymes make the desired one most favorable
5 Enzyme-Substrate Complex Enzymes act by binding substrates The noncovalent enzyme substrate complex is known as the Michaelis complex Description of chemical interactions Development of kinetic equations v k cat K [ E][ S] m [ S]
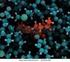
6 Enzyme-Substrate Complex Binding of a substrate to an enzyme at the active site. The enzyme chymotrypsin with bound substrate (PDB ID 7GCH). Some key active-site amino acid residues appear as a red splotch on the enzyme surface.
7 Enzymatic Catalysis Enzymes do not affect equilibrium (ΔG) Slow reactions face significant activation barriers (ΔG ) that must be surmounted during the reaction Enzymes increase reaction rates (k) by decreasing ΔG k k BT h G exp RT
8 Reaction Coordinate Diagram Reaction coordinate diagram. The free energy of the system is plotted against the progress of the reaction S P. A diagram of this kind is a description of the energy changes during the reaction, and the horizontal axis (reaction coordinate) reflects the progressive chemical changes (e.g., bond breakage or formation) as S is converted to P. The activation energies, G, for the S P and P S reactions are indicated. G is the overall standard freeenergy change in the direction S P.
9 Enzymes Decrease ΔG Reaction coordinate diagram comparing enzyme-catalyzed and uncatalyzed reactions. In the reaction S P, the ES and EP intermediates occupy minima in the energy progress curve of the enzymecatalyzed reaction. The terms G uncat and G cat correspond to the activation energy for the uncatalyzed reaction and the overall activation energy for the catalyzed reaction, respectively. The activation energy is lower when the enzyme catalyzes the reaction.
10 Rate Enhancement by Enzymes
11 How to Lower G Enzymes organize reactive groups into close proximity and proper orientation Uncatalyzed bimolecular reactions two free reactants single restricted transition state conversion is entropically unfavorable Uncatalyzed unimolecular reactions flexible reactant rigid transition state conversion is entropically unfavorable for flexible reactants Catalyzed reactions Enzyme uses the binding energy of substrates to organize the reactants to a fairly rigid ES complex Entropy cost is paid during binding Rigid reactant complex transition state conversion is entropically OK
12 Support for the Proximity Model The rate of anhydride formation from esters and carboxylates shows a strong dependence on proximity of two reactive groups.
13 From previous slide; Shown here are reactions of an ester with a carboxylate group to form an anhydride. The R group is the same in each case. (a) For this bimolecular reaction, the rate constant k is second-order, with units of M -1 s -1. (b) When the two reacting groups are in a single molecule, and thus have less freedom of motion, the reaction is much faster. For this unimolecular reaction, k has units of s -1. Dividing the rate constant for (b) by the rate constant for (a) gives a rate enhancement of about 10 5 M. (The enhancement has units of molarity because we are comparing a unimolecular and a bimolecular reaction.) Put another way, if the reactant in (b) were present at a concentration of 1 M, the reacting groups would behave as though they were present at a concentration of 10 5 M. Note that the reactant in (b) has freedom of rotation about three bonds (shown with curved arrows), but this still represents a substantial reduction of entropy over (a). If the bonds that rotate in (b) are constrained as in (c), the entropy is reduced further and the reaction exhibits a rate enhancement of 10 8 M relative to (a).
14 How to Lower G Enzymes bind transition states best The idea was proposed by Linus Pauling in 1946 Enzyme active sites are complimentary to the transition state of the reaction Enzymes bind transition states better than substrates Stronger/additional interactions with the transition state as compared to the ground state lower the activation barrier
15 Illustration of TS Stabilization Idea: Imaginary Stickase
16 An imaginary enzyme (stickase) designed to catalyze breakage of a metal stick. (a) Before the stick is broken, it must first be bent (the transition state). In both stickase examples, magnetic interactions take the place of weak bonding interactions between enzyme and substrate. (b) A stickase with a magnet-lined pocket complementary in structure to the stick (the substrate) stabilizes the substrate. Bending is impeded by the magnetic attraction between stick and stickase. (c) An enzyme with a pocket complementary to the reaction transition state helps to destabilize the stick, contributing to catalysis of the reaction. The binding energy of the magnetic interactions compensates for the increase in free energy required to bend the stick. Reaction coordinate diagrams (right) show the energy consequences of complementarity to substrate versus complementarity to transition state (EP complexes are omitted). G M, the difference between the transition-state energies of the uncatalyzed and catalyzed reactions, is contributed by the magnetic interactions between the stick and stickase. When the enzyme is complementary to the substrate (b), the ES complex is more stable and has less free energy in the ground state than substrate alone. The result is an increase in the activation energy.
17 Catalytic Mechanisms acid-base catalysis: give and take protons covalent catalysis: change reaction paths metal ion catalysis: use redox cofactors, pk a shifters electrostatic catalysis: preferential interactions with TS
18 General Acid-Base Catalysis How a catalyst circumvents unfavorable charge development during cleavage of an amide. The hydrolysis of an amide bond, shown here, is the same reaction as that catalyzed by chymotrypsin and other proteases. Charge development is unfavorable and can be circumvented by donation of a proton by H 3 O + (specific acid catalysis) or HA (general acid catalysis), where HA represents any acid. Similarly, charge can be neutralized by proton abstraction by OH (specific base catalysis) or B: (general base catalysis), where B: represents any base.
19
20
21
22 Amino Acids in General Acid-Base Catalysis Amino acids in general acidbase catalysis. Many organic reactions that are used to model biochemical processes are promoted by proton donors (general acids) or proton acceptors (general bases). The active sites of some enzymes contain amino acid functional groups, such as those shown here, that can participate in the catalytic process as proton donors or proton acceptors.
23 Covalent Catalysis A transient covalent bond between the enzyme and the substrate Changes the reaction Pathway Uncatalyzed: A B H2O A B Catalyzed: A B X : A X H2O B A X : B Requires a nucleophile on the enzyme Can be a reactive serine, thiolate, amine, or carboxylate
24 Metal Ion Catalysis Involves a metal ion bound to the enzyme Interacts with substrate to facilitate binding Stabilizes negative charges Participates in oxidation reactions
25 Chymotrypsin uses most of the enzymatic mechanisms Structure of chymotrypsin. (PDB ID 7GCH) (c) The polypeptide backbone as a ribbon structure. Disulfide bonds are yellow; the three chains are colored as in part (a).
26 Active Site of Chymotrypsin with Substrate Structure of chymotrypsin. (PDB ID 7GCH) (d) A close-up of the active site with a substrate (white and yellow) bound. The hydroxyl of Ser 195 attacks the carbonyl group of the substrate (the oxygens are red); the developing negative charge on the oxygen is stabilized by the oxyanion hole (amide nitrogens from Ser 195 and Gly 193, in blue), as explained in Figure The aromatic amino acid side chain of the substrate (yellow) sits in the hydrophobic pocket. The amide nitrogen of the peptide bond to be cleaved (protruding toward the viewer and projecting the path of the rest of the substrate polypeptide chain) is shown in white.
27 Chymotrypsin Mechanism Step 1: Substrate Binding (step 1) Hydrolytic cleavage of a peptide bond by chymotrypsin. The reaction has two phases. In the acylation phase (steps 1 to 4), formation of a covalent acyl-enzyme intermediate is coupled to cleavage of the peptide bond. In the deacylation phase (steps 5 to 7), deacylation regenerates the free enzyme; this is essentially the reverse of the acylation phase, with water mirroring, in reverse, the role of the amine component of the substrate.
28 Chymotrypsin Mechanism Step 2: Nucleophilic Attack
29 Chymotrypsin Mechanism Step 3: Substrate Cleavage
30 Chymotrypsin Mechanism Step 4: Water Comes In
31 Chymotrypsin Mechanism Step 5: Water Attacks
32 Chymotrypsin Mechanism Step 6: Break-off from the Enzyme
33 Chymotrypsin Mechanism Step 7: Product Dissociates
34 Peptidoglycan and Lysozyme Peptidoglycan is a polysaccharide found in many bacterial cell walls Cleavage of the cell wall leads to the lysis of bacteria Lysozyme is an antibacterial enzyme
35 Peptidoglycan and Lysozyme Hen egg white lysozyme and the reaction it catalyzes. (b) Reaction catalyzed by hen egg white lysozyme. A segment of a peptidoglycan polymer is shown, with the lysozyme binding sites A through F shaded. The glycosidic C O bond between sugar residues bound to sites D and E is cleaved, as indicated by the red arrow. The hydrolytic reaction is shown in the inset, with the fate of the oxygen in the H 2 O traced in red. Mur2Ac is N-acetylmuramic acid; GlcNAc, N-acetylglucosamine. RO represents a lactyl (lactic acid) group; NAc and AcN, an N-acetyl group (see key).
36 General Acid-Base + Covalent Catalysis: Cleavage of Peptidoglycan by Lysozyme X-ray structures of lysozyme with bound substrate analogs show that the C-1 carbon is located between Glu 35 and Asp 52 residues.
37 From previous slide; Lysozyme reaction. In this reaction, the water introduced into the product at C-1 of Mur2Ac is in the same configuration as the original glycosidic bond. The reaction is thus a molecular substitution with retention of configuration. (b) A surface rendering of the lysozyme active site with the covalent enzyme-substrate intermediate shown as a ball-and-stick structure. Side-chains of active-site residues are shown as ball-and-stick structures protruding from ribbons (PDB ID 1H6M).
38 Cleavage of Peptidoglycan by Lysozyme: Two Successive S N 2 Steps Model Asp 52 acts as a nucleophile to attack the anomeric carbon in the first S N 2 step Glu 35 acts as a general acid and protonates the leaving group in the transition state Water hydrolyzes the covalent glycosyl-enzyme intermediate Glu 35 acts as a general base to deprotonate water in the second S N 2 step
39
40
41
42 Experimental work I am no longer allowed to enjoy!
43 What are enzymes? Enzymes are catalysts Increase reaction rates without being used up Most enzymes are globular proteins However, some RNA (ribozymes and ribosomal RNA) also catalyze reactions We will celebrate my inspiration, the Biochemist Louis Pasteur.
44 Why biocatalysis over inorganic catalysts? Greater reaction specificity: avoids side products Milder reaction conditions: conducive to conditions in cells Higher reaction rates: in a biologically useful timeframe Capacity for regulation: control of biological pathways COO - - COO OH - COO NH 2 O - COO Metabolites have many potential pathways of decomposition - COO NH 2 OH O - COO Chorismate mutase - OOC OH - COO O Enzymes make the desired one most favorable
45 What is enzyme kinetics? Kinetics is the study of the rate at which compounds react Rate of enzymatic reaction is affected by: enzyme substrate effectors temperature
46 Why study enzyme kinetics? Quantitative description of biocatalysis Determine the order of binding of substrates Elucidate acid-base catalysis Understand catalytic mechanism Find effective inhibitors Understand regulation of activity
47 How to Do Kinetic Measurements Experiment: 1)Mix enzyme + substrate 2)Record rate of substrate disappearance/product formation as a function of time (the velocity of reaction) 3)Plot initial velocity versus substrate concentration. 4)Change substrate concentration and repeat
48 From previous slide; Initial velocities of enzymecatalyzed reactions A theoretical enzyme catalyzes the reaction S P, and is present at a concentration sufficient to catalyze the reaction at a maximum velocity, V max, of 1 μm/min. The Michaelis constant, K m (explained in the text), is 0.5 μm. Progress curves are shown for substrate concentrations below, at, and above the K m. The rate of an enzymecatalyzed reaction declines as substrate is converted to product. A tangent to each curve taken at time = 0 defines the initial velocity, V 0, of each reaction.
49 Effect of Substrate Concentration Ideal rate: v V K max [ m S] S Deviations due to: limitation of measurements substrate inhibition substrate prep contains inhibitors enzyme prep contains inhibitors
50 Effect of Substrate Concentration Effect of substrate concentration on the initial velocity of an enzyme-catalyzed reaction. The maximum velocity, V max, is extrapolated from the plot because V 0 approaches but never quite reaches V max. The substrate concentration at which V 0 is half maximal is K m, the Michaelis constant. The concentration of enzyme in an experiment such as this is generally so low that [S] >> [E] even when [S] is described as low or relatively low. The units shown are typical for enzyme-catalyzed reactions and are given only to help illustrate the meaning of V 0 and [S]. (Note that the curve describes part of a rectangular hyperbola, with one asymptote at V max. If the curve were continued below [S] = 0, it would approach a vertical asymptote at [S] = K m.)
51 Saturation Kinetics: At high [S] velocity does not depend on [S] Dependence of initial velocity on substrate concentration. This graph shows the kinetic parameters that define the limits of the curve at high and low [S]. At low [S], K m >> [S] and the [S] term in the denominator of the Michaelis-Menten equation (Eqn 6-9) becomes insignificant. The equation simplifies to V 0 = V max [S]/K m and V 0 exhibits a linear dependence on [S], as observed here. At high [S], where [S] >> K m, the K m term in the denominator of the Michaelis-Menten equation becomes insignificant and the equation simplifies to V 0 = V max ; this is consistent with the plateau observed at high [S]. The Michaelis-Menten equation is therefore consistent with the observed dependence of V 0 on [S], and the shape of the curve is defined by the terms V max /K m at low [S] and V max at high [S].
52 Determination of Kinetic Parameters Nonlinear Michaelis-Menten plot should be used to calculate parameters K m and V max. Linearized double-reciprocal plot is good for analysis of two-substrate data or inhibition.
53 Lineweaver-Burk Plot: Linearized, Double-Reciprocal
54 Derivation of Enzyme Kinetics Equations Start with a model mechanism Identify constraints and assumptions Carry out algebra or graph theory for complex reactions Simplest Model Mechanism: E + S ES E + P One reactant, one product, no inhibitors
55 Identify Constraints and Assumptions Total enzyme concentration is constant Mass balance equation for enzyme: E Tot = [E] + [ES] It is also implicitly assumed that: S Tot = [S] + [ES] [S] Steady state assumption d[ ES] dt rateof formationof ES rateof breakdown of ES 0 What is the observed rate? Rate of product formation v net dp dt k[es]
56 Carry out the algebra The final form in case of a single substrate is v k cat K [ E m tot ][ S] [ S] k cat (turnover number): how many substrate molecules can one enzyme molecule convert per second K m (Michaelis constant): an approximate measure of substrate s affinity for enzyme Microscopic meaning of K m and k cat depends on the details of the mechanism
57 Enzyme efficiency is limited by diffusion: k cat /K M Can gain efficiency by having high velocity or affinity for substrate Catalase vs. acetylcholinesterase
58 Two-Substrate Reactions Kinetic mechanism: the order of binding of substrates and release of products When two or more reactants are involved, enzyme kinetics allows to distinguish between different kinetic mechanisms Sequential mechanism Ping-Pong mechanism
59 Common mechanisms for enzymecatalyzed bisubstrate reactions. (a) The enzyme and both substrates come together to form a ternary complex. In ordered binding, substrate 1 must bind before substrate 2 can bind productively. In random binding, the substrates can bind in either order. (b) An enzymesubstrate complex forms, a product leaves the complex, the altered enzyme forms a second complex with another substrate molecule, and the second product leaves, regenerating the enzyme. Substrate 1 may transfer a functional group to the enzyme (to form the covalently modified E ), which is subsequently transferred to substrate 2. This is called a Ping-Pong or double-displacement mechanism.
60 Sequential Kinetic Mechanism We cannot easily distinguish random from ordered Random mechanisms in equilibrium will give intersection point at y-axis Lineweaver-Burk: lines intersect Steady-state kinetic analysis of bisubstrate reactions. In these double-reciprocal plots (see Box 6-1), the concentration of substrate 1 is varied while the concentration of substrate 2 is held constant. This is repeated for several values of [S 2 ], generating several separate lines. (a) Intersecting lines indicate that a ternary complex is formed in the reaction.
61 Ping-Pong Kinetic Mechanism Steady-state kinetic analysis of bisubstrate reactions. In these doublereciprocal plots (see Box 6-1), the concentration of substrate 1 is varied while the concentration of substrate 2 is held constant. This is repeated for several values of [S 2 ], generating several separate lines. (b) parallel lines indicate a Ping-Pong (doubledisplacement) pathway. Lineweaver-Burk: lines are parallel
62 Enzyme Inhibition Inhibitors are compounds that decrease enzyme s activity Irreversible inhibitors (inactivators) react with the enzyme One inhibitor molecule can permanently shut off one enzyme molecule They are often powerful toxins but also may be used as drugs Reversible inhibitors bind to and can dissociate from the enzyme They are often structural analogs of substrates or products They are often used as drugs to slow down a specific enzyme Reversible inhibitor can bind: to the free enzyme and prevent the binding of the substrate to the enzyme-substrate complex and prevent the reaction
63 Competitive Inhibition Competes with substrate for binding Binds active site Does not affect catalysis No change in V max ; apparent increase in K M Lineweaver-Burk: lines intersect at the y-axis
64 Competitive Inhibition
65 Competitive Inhibition
66 Uncompetitive Inhibition Only binds to ES complex Does not affect substrate binding Inhibits catalytic function Decrease in V max ; apparent decrease in K M No change in K M /V max Lineweaver-Burk: lines are parallel
67 Uncompetitive Inhibition
68 Uncompetitive Inhibition
69 Mixed Inhibition Binds enzyme with or without substrate Binds to regulatory site Inhibits both substrate binding and catalysis Decrease in V max ; apparent change in K M Lineweaver-Burk: lines intersect left from the y-axis Noncompetitive inhibitors are mixed inhibitors such that there is no change in K M
70 Mixed Inhibition
71 Mixed Inhibition
72 Enzyme activity can be regulated Regulation can be: noncovalent modification covalent modification irreversible reversible
73 Noncovalent Modification: Allosteric Regulators The kinetics of allosteric regulators differ from Michaelis-Menten kinetics.
74 Aspartate transcarbamylase;
75 From previous slide Substrate-activity curves for representative allosteric enzymes. Three examples of complex responses of allosteric enzymes to their modulators. (a) The sigmoid curve of a homotropic enzyme, in which the substrate also serves as a positive (stimulatory) modulator, or activator. Note the resemblance to the oxygen-saturation curve of hemoglobin. The sigmoidal curve is a hybrid curve in which the enzyme is present primarily in the relatively inactive T state at low substrate concentration, and primarily in the more active R state at high substrate concentration. The curves for the pure T and R states are plotted separately in color. ATCase exhibits a kinetic pattern similar to this. (b) The effects of several different concentrations of a positive modulator (+) or a negative modulator (-) on an allosteric enzyme in which K 0.5 is altered without a change in V max. The central curve shows the substrate-activity relationship without a modulator. For ATCase, CTP is a negative modulator and ATP is a positive modulator.
76 Some Reversible Covalent Modifications
77 Zymogens are activated by irreversible covalent modification
78 From previous slide; Activation of zymogens by proteolytic cleavage. Shown here is the formation of chymotrypsin and trypsin from their zymogens, chymotrypsinogen and trypsinogen. The bars represent the amino acid sequences of the polypeptide chains, with numbers indicating the positions of the residues (the aminoterminal residue is number 1). Residues at the termini of the polypeptide fragments generated by cleavage are indicated below the bars. Note that in the final active forms, some numbered residues are missing. Recall that the three polypeptide chains (A, B, and C) of chymotrypsin are linked by disulfide bonds (see Fig. 6 19).
79 The blood coagulation cascade uses irreversible covalent modification The coagulation cascades. The interlinked intrinsic and extrinsic pathways leading to the cleavage of fibrinogen to form active fibrin are shown. Active serine proteases in the pathways are shown in blue. Green arrows denote activating steps, and red arrows indicate inhibitory processes.
80 Some enzymes use multiple types of regulation
81 Regulation of muscle glycogen phosphorylase activity by phosphorylation. The activity of glycogen phosphorylase in muscle is subjected to a multilevel system of regulation involving much more than the covalent modification (phosphorylation) shown in Figure Allosteric regulation, and a regulatory cascade sensitive to hormonal status that acts on the enzymes involved in phosphorylation and dephosphorylation, also play important roles. The activity of both forms of the enzyme is allosterically regulated by an activator (AMP) and by inhibitors (glucose 6-phosphate and ATP) that bind to separate sites on the enzyme. The activities of phosphorylase kinase and phosphorylase phosphatase 1 (PP1) are also regulated by covalent modification, via a short pathway that responds to the hormones glucagon and epinephrine. One path leads to the phosphorylation of phosphorylase kinase and phosphoprotein phosphatase inhibitor 1 (PPI-1). The phosphorylated phosphorylase kinase is activated and in turn phosphorylates and activates glycogen phosphorylase. At the same time, the phosphorylated PPI-1 interacts with and inhibits PP1. PPI-1 also keeps itself active (phosphorylated) by inhibiting phosphoprotein phosphatase 2B (PP2B), the enzyme that dephosphorylates (inactivates) it. In this way, the equilibrium between the a and b forms of glycogen phosphorylase is shifted decisively toward the more active glycogen phosphorylase a. Note that the two forms of phosphorylase kinase are both activated to a degree by Ca 2+ ion (not shown). This pathway is discussed in more detail in Chapters 14, 15, and 23.
Lecture 15: Enzymes & Kinetics Mechanisms
 ROLE OF THE TRANSITION STATE Lecture 15: Enzymes & Kinetics Mechanisms Consider the reaction: H-O-H + Cl - H-O δ- H Cl δ- HO - + H-Cl Reactants Transition state Products Margaret A. Daugherty Fall 2004
ROLE OF THE TRANSITION STATE Lecture 15: Enzymes & Kinetics Mechanisms Consider the reaction: H-O-H + Cl - H-O δ- H Cl δ- HO - + H-Cl Reactants Transition state Products Margaret A. Daugherty Fall 2004
Enzymes reduce the activation energy
 Enzymes reduce the activation energy Transition state is an unstable transitory combination of reactant molecules which occurs at the potential energy maximum (free energy maximum). Note - the ΔG of the
Enzymes reduce the activation energy Transition state is an unstable transitory combination of reactant molecules which occurs at the potential energy maximum (free energy maximum). Note - the ΔG of the
Lecture 11 Enzymes: Kinetics
 Lecture 11 Enzymes: Kinetics Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 8, pp. 216-225 Key Concepts Kinetics is the study of reaction rates (velocities). Study of enzyme kinetics is useful for
Lecture 11 Enzymes: Kinetics Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 8, pp. 216-225 Key Concepts Kinetics is the study of reaction rates (velocities). Study of enzyme kinetics is useful for
Enzymes. Enzyme Structure. Enzyme Classification. CHEM464/Medh, J.D. Reaction Rate and Enzyme Activity
 Enzymes Enzymes are biological catalysts They are not consumed or altered during the reaction They do not change the equilibrium, just reduce the time required to reach equilibrium. They increase the rate
Enzymes Enzymes are biological catalysts They are not consumed or altered during the reaction They do not change the equilibrium, just reduce the time required to reach equilibrium. They increase the rate
How To Understand Enzyme Kinetics
 Chapter 12 - Reaction Kinetics In the last chapter we looked at enzyme mechanisms. In this chapter we ll see how enzyme kinetics, i.e., the study of enzyme reaction rates, can be useful in learning more
Chapter 12 - Reaction Kinetics In the last chapter we looked at enzyme mechanisms. In this chapter we ll see how enzyme kinetics, i.e., the study of enzyme reaction rates, can be useful in learning more
Chemistry 20 Chapters 15 Enzymes
 Chemistry 20 Chapters 15 Enzymes Enzymes: as a catalyst, an enzyme increases the rate of a reaction by changing the way a reaction takes place, but is itself not changed at the end of the reaction. An
Chemistry 20 Chapters 15 Enzymes Enzymes: as a catalyst, an enzyme increases the rate of a reaction by changing the way a reaction takes place, but is itself not changed at the end of the reaction. An
Enzymes. Enzymes are characterized by: Specificity - highly specific for substrates
 Enzymes Enzymes are characterized by: Catalytic Power - rates are 10 6-10 12 greater than corresponding uncatalyzed reactions Specificity - highly specific for substrates Regulation - acheived in many
Enzymes Enzymes are characterized by: Catalytic Power - rates are 10 6-10 12 greater than corresponding uncatalyzed reactions Specificity - highly specific for substrates Regulation - acheived in many
Chapter 8: An Introduction to Metabolism
 Chapter 8: An Introduction to Metabolism Name Period Concept 8.1 An organism s metabolism transforms matter and energy, subject to the laws of thermodynamics 1. Define metabolism. The totality of an organism
Chapter 8: An Introduction to Metabolism Name Period Concept 8.1 An organism s metabolism transforms matter and energy, subject to the laws of thermodynamics 1. Define metabolism. The totality of an organism
Lecture 3: Enzyme kinetics
 Computational Systems Biology Lecture 3: Enzyme kinetics Fri 19 Jan 2009 1 Images from: D. L. Nelson, Lehninger Principles of Biochemistry, IV Edition, W. H. Freeman ed. A. Cornish-Bowden Fundamentals
Computational Systems Biology Lecture 3: Enzyme kinetics Fri 19 Jan 2009 1 Images from: D. L. Nelson, Lehninger Principles of Biochemistry, IV Edition, W. H. Freeman ed. A. Cornish-Bowden Fundamentals
ENZYMES. Serine Proteases Chymotrypsin, Trypsin, Elastase, Subtisisin. Principle of Enzyme Catalysis
 ENZYMES Serine Proteases Chymotrypsin, Trypsin, Elastase, Subtisisin Principle of Enzyme Catalysis Linus Pauling (1946) formulated the first basic principle of enzyme catalysis Enzyme increase the rate
ENZYMES Serine Proteases Chymotrypsin, Trypsin, Elastase, Subtisisin Principle of Enzyme Catalysis Linus Pauling (1946) formulated the first basic principle of enzyme catalysis Enzyme increase the rate
Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Two Forms of Energy
 Module 2D - Energy and Metabolism Objective # 19 All living organisms require energy for survival. In this module we will examine some general principles about chemical reactions and energy usage within
Module 2D - Energy and Metabolism Objective # 19 All living organisms require energy for survival. In this module we will examine some general principles about chemical reactions and energy usage within
Previous lecture: Today:
 Previous lecture: The energy requiring step from substrate to transition state is an energy barrier called the free energy of activation G Transition state is the unstable (10-13 seconds) highest energy
Previous lecture: The energy requiring step from substrate to transition state is an energy barrier called the free energy of activation G Transition state is the unstable (10-13 seconds) highest energy
Regulation of enzyme activity
 1 Regulation of enzyme activity Regulation of enzyme activity is important to coordinate the different metabolic processes. It is also important for homeostasis i.e. to maintain the internal environment
1 Regulation of enzyme activity Regulation of enzyme activity is important to coordinate the different metabolic processes. It is also important for homeostasis i.e. to maintain the internal environment
1. A covalent bond between two atoms represents what kind of energy? a. Kinetic energy b. Potential energy c. Mechanical energy d.
 1. A covalent bond between two atoms represents what kind of energy? a. Kinetic energy b. Potential energy c. Mechanical energy d. Solar energy A. Answer a is incorrect. Kinetic energy is the energy of
1. A covalent bond between two atoms represents what kind of energy? a. Kinetic energy b. Potential energy c. Mechanical energy d. Solar energy A. Answer a is incorrect. Kinetic energy is the energy of
CHM333 LECTURE 13 14: 2/13 15/13 SPRING 2013 Professor Christine Hrycyna
 INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
Enzymes Enzyme Mechanism
 Mechanisms of Enzymes BCMB 3100 Chapters 6, 7, 8 Enzymes Enzyme Mechanism 1 Energy diagrams Binding modes of enzyme catalysis Chemical modes of enzyme catalysis Acid-Base catalysis Covalent catalysis Binding
Mechanisms of Enzymes BCMB 3100 Chapters 6, 7, 8 Enzymes Enzyme Mechanism 1 Energy diagrams Binding modes of enzyme catalysis Chemical modes of enzyme catalysis Acid-Base catalysis Covalent catalysis Binding
Biochemistry - I. Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture-11 Enzyme Mechanisms II
 Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture-11 Enzyme Mechanisms II In the last class we studied the enzyme mechanisms of ribonuclease A
Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture-11 Enzyme Mechanisms II In the last class we studied the enzyme mechanisms of ribonuclease A
Energy & Enzymes. Life requires energy for maintenance of order, growth, and reproduction. The energy living things use is chemical energy.
 Energy & Enzymes Life requires energy for maintenance of order, growth, and reproduction. The energy living things use is chemical energy. 1 Energy exists in two forms - potential and kinetic. Potential
Energy & Enzymes Life requires energy for maintenance of order, growth, and reproduction. The energy living things use is chemical energy. 1 Energy exists in two forms - potential and kinetic. Potential
MULTIPLE CHOICE QUESTIONS
 MULTIPLE CHOICE QUESTIONS 1. Most components of energy conversion systems evolved very early; thus, the most fundamental aspects of energy metabolism tend to be: A. quite different among a diverse group
MULTIPLE CHOICE QUESTIONS 1. Most components of energy conversion systems evolved very early; thus, the most fundamental aspects of energy metabolism tend to be: A. quite different among a diverse group
ENZYME SCIENCE AND ENGINEERING PROF. SUBHASH CHAND DEPARTMENT OF BIOCHEMICAL ENGINEERING AND BIOTECHNOLOGY IIT DELHI LECTURE 4 ENZYMATIC CATALYSIS
 ENZYME SCIENCE AND ENGINEERING PROF. SUBHASH CHAND DEPARTMENT OF BIOCHEMICAL ENGINEERING AND BIOTECHNOLOGY IIT DELHI LECTURE 4 ENZYMATIC CATALYSIS We will continue today our discussion on enzymatic catalysis
ENZYME SCIENCE AND ENGINEERING PROF. SUBHASH CHAND DEPARTMENT OF BIOCHEMICAL ENGINEERING AND BIOTECHNOLOGY IIT DELHI LECTURE 4 ENZYMATIC CATALYSIS We will continue today our discussion on enzymatic catalysis
CHAPTER 4: Enzyme Structure ENZYMES
 CHAPTER 4: ENZYMES Enzymes are biological catalysts. There are about 40,000 different enzymes in human cells, each controlling a different chemical reaction. They increase the rate of reactions by a factor
CHAPTER 4: ENZYMES Enzymes are biological catalysts. There are about 40,000 different enzymes in human cells, each controlling a different chemical reaction. They increase the rate of reactions by a factor
CHAPTER 6 AN INTRODUCTION TO METABOLISM. Section B: Enzymes
 CHAPTER 6 AN INTRODUCTION TO METABOLISM Section B: Enzymes 1. Enzymes speed up metabolic reactions by lowering energy barriers 2. Enzymes are substrate specific 3. The active site in an enzyme s catalytic
CHAPTER 6 AN INTRODUCTION TO METABOLISM Section B: Enzymes 1. Enzymes speed up metabolic reactions by lowering energy barriers 2. Enzymes are substrate specific 3. The active site in an enzyme s catalytic
What affects an enzyme s activity? General environmental factors, such as temperature and ph. Chemicals that specifically influence the enzyme.
 CH s 8-9 Respiration & Metabolism Metabolism A catalyst is a chemical agent that speeds up a reaction without being consumed by the reaction. An enzyme is a catalytic protein. Hydrolysis of sucrose by
CH s 8-9 Respiration & Metabolism Metabolism A catalyst is a chemical agent that speeds up a reaction without being consumed by the reaction. An enzyme is a catalytic protein. Hydrolysis of sucrose by
Chapter 8: Energy and Metabolism
 Chapter 8: Energy and Metabolism 1. Discuss energy conversions and the 1 st and 2 nd law of thermodynamics. Be sure to use the terms work, potential energy, kinetic energy, and entropy. 2. What are Joules
Chapter 8: Energy and Metabolism 1. Discuss energy conversions and the 1 st and 2 nd law of thermodynamics. Be sure to use the terms work, potential energy, kinetic energy, and entropy. 2. What are Joules
Enzymes: Introduction
 Enzymes: Introduction Firefly bioluminescence is produced by an oxidation reaction catalyzed by the enzyme firefly luciferase. The oxidized substrate (product of the reaction) is in an electronically excited
Enzymes: Introduction Firefly bioluminescence is produced by an oxidation reaction catalyzed by the enzyme firefly luciferase. The oxidized substrate (product of the reaction) is in an electronically excited
Lecture 10 Enzymes: Introduction
 Lecture 10 Enzymes: Introduction Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 8, pp. 205-217 (These pages in textbook are very important -- concepts of thermodynamics are fundamental to all of biochemistry.)
Lecture 10 Enzymes: Introduction Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 8, pp. 205-217 (These pages in textbook are very important -- concepts of thermodynamics are fundamental to all of biochemistry.)
How To Understand The Chemistry Of An Enzyme
 Chapt. 8 Enzymes as catalysts Ch. 8 Enzymes as catalysts Student Learning Outcomes: Explain general features of enzymes as catalysts: Substrate -> Product Describe nature of catalytic sites general mechanisms
Chapt. 8 Enzymes as catalysts Ch. 8 Enzymes as catalysts Student Learning Outcomes: Explain general features of enzymes as catalysts: Substrate -> Product Describe nature of catalytic sites general mechanisms
CHM333 LECTURE 13 14: 2/13 15/12 SPRING 2012 Professor Christine Hrycyna
 INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
Enzymes. Enzyme Classification
 Enzyme Classification Simple Enzymes: composed of whole proteins Complex Enzymes: composed of protein plus a relatively small organic molecule holoenzyme = apoenzyme + prosthetic group / coenzyme A prosthetic
Enzyme Classification Simple Enzymes: composed of whole proteins Complex Enzymes: composed of protein plus a relatively small organic molecule holoenzyme = apoenzyme + prosthetic group / coenzyme A prosthetic
Enzymes and Metabolic Pathways
 Enzymes and Metabolic Pathways Enzyme characteristics Made of protein Catalysts: reactions occur 1,000,000 times faster with enzymes Not part of reaction Not changed or affected by reaction Used over and
Enzymes and Metabolic Pathways Enzyme characteristics Made of protein Catalysts: reactions occur 1,000,000 times faster with enzymes Not part of reaction Not changed or affected by reaction Used over and
A. A peptide with 12 amino acids has the following amino acid composition: 2 Met, 1 Tyr, 1 Trp, 2 Glu, 1 Lys, 1 Arg, 1 Thr, 1 Asn, 1 Ile, 1 Cys
 Questions- Proteins & Enzymes A. A peptide with 12 amino acids has the following amino acid composition: 2 Met, 1 Tyr, 1 Trp, 2 Glu, 1 Lys, 1 Arg, 1 Thr, 1 Asn, 1 Ile, 1 Cys Reaction of the intact peptide
Questions- Proteins & Enzymes A. A peptide with 12 amino acids has the following amino acid composition: 2 Met, 1 Tyr, 1 Trp, 2 Glu, 1 Lys, 1 Arg, 1 Thr, 1 Asn, 1 Ile, 1 Cys Reaction of the intact peptide
green B 1 ) into a single unit to model the substrate in this reaction. enzyme
 Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
8/20/2012 H C OH H R. Proteins
 Proteins Rubisco monomer = amino acids 20 different amino acids polymer = polypeptide protein can be one or more polypeptide chains folded & bonded together large & complex 3-D shape hemoglobin Amino acids
Proteins Rubisco monomer = amino acids 20 different amino acids polymer = polypeptide protein can be one or more polypeptide chains folded & bonded together large & complex 3-D shape hemoglobin Amino acids
ENZYMES (DR. NUGENT)
 ENZYMES (DR. NUGENT) I. INTRODUCTION TO ENZYMES/CATALYSTS a. Definition: enzyme = biological catalyst (catalyst = speeds rate of reaction) b. Features of enzymes: i. Enzymes speeds up the rate of a reaction
ENZYMES (DR. NUGENT) I. INTRODUCTION TO ENZYMES/CATALYSTS a. Definition: enzyme = biological catalyst (catalyst = speeds rate of reaction) b. Features of enzymes: i. Enzymes speeds up the rate of a reaction
Lecture 4 Enzymes Catalytic proteins. Enzymes. Enzymes 10/21/10. What enzymes do therefore is:
 Lecture 4 Catalytic proteins Are a type of protein that acts as a catalyst-speeding up chemical reactions A catalyst is defined as a chemical agent that changes the rate of a reaction without being consumed
Lecture 4 Catalytic proteins Are a type of protein that acts as a catalyst-speeding up chemical reactions A catalyst is defined as a chemical agent that changes the rate of a reaction without being consumed
18.2 Protein Structure and Function: An Overview
 18.2 Protein Structure and Function: An Overview Protein: A large biological molecule made of many amino acids linked together through peptide bonds. Alpha-amino acid: Compound with an amino group bonded
18.2 Protein Structure and Function: An Overview Protein: A large biological molecule made of many amino acids linked together through peptide bonds. Alpha-amino acid: Compound with an amino group bonded
Catalysis by Enzymes. Enzyme A protein that acts as a catalyst for a biochemical reaction.
 Catalysis by Enzymes Enzyme A protein that acts as a catalyst for a biochemical reaction. Enzymatic Reaction Specificity Enzyme Cofactors Many enzymes are conjugated proteins that require nonprotein portions
Catalysis by Enzymes Enzyme A protein that acts as a catalyst for a biochemical reaction. Enzymatic Reaction Specificity Enzyme Cofactors Many enzymes are conjugated proteins that require nonprotein portions
ENZYMES - EXTRA QUESTIONS
 ENZYMES - EXTRA QUESTIONS 1. A chemical reaction has a G o = -60 kj/mol. If this were an enzyme-catalyzed reaction what can you predict about the kinetics? A. It will exhibit very rapid kinetics. B. It
ENZYMES - EXTRA QUESTIONS 1. A chemical reaction has a G o = -60 kj/mol. If this were an enzyme-catalyzed reaction what can you predict about the kinetics? A. It will exhibit very rapid kinetics. B. It
PHYSIOLOGY AND MAINTENANCE Vol. II Enzymes: The Biological Catalysts of Life - Pekka Mäntsälä and Jarmo Niemi
 ENZYMES: THE BIOLOGICAL CATALYSTS OF LIFE Pekka Mäntsälä and Jarmo Niemi University of Turku, Department of Biochemistry, Finland Keywords: enzymes, specificity, catalysis, cofactors, enzyme turnover,
ENZYMES: THE BIOLOGICAL CATALYSTS OF LIFE Pekka Mäntsälä and Jarmo Niemi University of Turku, Department of Biochemistry, Finland Keywords: enzymes, specificity, catalysis, cofactors, enzyme turnover,
Enzymes and Metabolism
 Enzymes and Metabolism Enzymes and Metabolism Metabolism: Exergonic and Endergonic Reactions Chemical Reactions: Activation Every chemical reaction involves bond breaking and bond forming A chemical reaction
Enzymes and Metabolism Enzymes and Metabolism Metabolism: Exergonic and Endergonic Reactions Chemical Reactions: Activation Every chemical reaction involves bond breaking and bond forming A chemical reaction
Keystone Review Practice Test Module A Cells and Cell Processes. 1. Which characteristic is shared by all prokaryotes and eukaryotes?
 Keystone Review Practice Test Module A Cells and Cell Processes 1. Which characteristic is shared by all prokaryotes and eukaryotes? a. Ability to store hereditary information b. Use of organelles to control
Keystone Review Practice Test Module A Cells and Cell Processes 1. Which characteristic is shared by all prokaryotes and eukaryotes? a. Ability to store hereditary information b. Use of organelles to control
AP BIOLOGY 2008 SCORING GUIDELINES
 AP BIOLOGY 2008 SCORING GUIDELINES Question 1 1. The physical structure of a protein often reflects and affects its function. (a) Describe THREE types of chemical bonds/interactions found in proteins.
AP BIOLOGY 2008 SCORING GUIDELINES Question 1 1. The physical structure of a protein often reflects and affects its function. (a) Describe THREE types of chemical bonds/interactions found in proteins.
IV. -Amino Acids: carboxyl and amino groups bonded to -Carbon. V. Polypeptides and Proteins
 IV. -Amino Acids: carboxyl and amino groups bonded to -Carbon A. Acid/Base properties 1. carboxyl group is proton donor! weak acid 2. amino group is proton acceptor! weak base 3. At physiological ph: H
IV. -Amino Acids: carboxyl and amino groups bonded to -Carbon A. Acid/Base properties 1. carboxyl group is proton donor! weak acid 2. amino group is proton acceptor! weak base 3. At physiological ph: H
1. The diagram below represents a biological process
 1. The diagram below represents a biological process 5. The chart below indicates the elements contained in four different molecules and the number of atoms of each element in those molecules. Which set
1. The diagram below represents a biological process 5. The chart below indicates the elements contained in four different molecules and the number of atoms of each element in those molecules. Which set
Figure 5. Energy of activation with and without an enzyme.
 Biology 20 Laboratory ENZYMES & CELLULAR RESPIRATION OBJECTIVE To be able to list the general characteristics of enzymes. To study the effects of enzymes on the rate of chemical reactions. To demonstrate
Biology 20 Laboratory ENZYMES & CELLULAR RESPIRATION OBJECTIVE To be able to list the general characteristics of enzymes. To study the effects of enzymes on the rate of chemical reactions. To demonstrate
Copyright 2000-2003 Mark Brandt, Ph.D. 54
 Pyruvate Oxidation Overview of pyruvate metabolism Pyruvate can be produced in a variety of ways. It is an end product of glycolysis, and can be derived from lactate taken up from the environment (or,
Pyruvate Oxidation Overview of pyruvate metabolism Pyruvate can be produced in a variety of ways. It is an end product of glycolysis, and can be derived from lactate taken up from the environment (or,
Carbohydrates, proteins and lipids
 Carbohydrates, proteins and lipids Chapter 3 MACROMOLECULES Macromolecules: polymers with molecular weights >1,000 Functional groups THE FOUR MACROMOLECULES IN LIFE Molecules in living organisms: proteins,
Carbohydrates, proteins and lipids Chapter 3 MACROMOLECULES Macromolecules: polymers with molecular weights >1,000 Functional groups THE FOUR MACROMOLECULES IN LIFE Molecules in living organisms: proteins,
Disaccharides consist of two monosaccharide monomers covalently linked by a glycosidic bond. They function in sugar transport.
 1. The fundamental life processes of plants and animals depend on a variety of chemical reactions that occur in specialized areas of the organism s cells. As a basis for understanding this concept: 1.
1. The fundamental life processes of plants and animals depend on a variety of chemical reactions that occur in specialized areas of the organism s cells. As a basis for understanding this concept: 1.
NO CALCULATORS OR CELL PHONES ALLOWED
 Biol 205 Exam 1 TEST FORM A Spring 2008 NAME Fill out both sides of the Scantron Sheet. On Side 2 be sure to indicate that you have TEST FORM A The answers to Part I should be placed on the SCANTRON SHEET.
Biol 205 Exam 1 TEST FORM A Spring 2008 NAME Fill out both sides of the Scantron Sheet. On Side 2 be sure to indicate that you have TEST FORM A The answers to Part I should be placed on the SCANTRON SHEET.
Cellular Energy: ATP & Enzymes. What is it? Where do organism s get it? How do they use it?
 Cellular Energy: ATP & Enzymes What is it? Where do organism s get it? How do they use it? Where does Energy come from? Ultimately, from the sun. It is transferred between organisms in the earth s lithosphere,
Cellular Energy: ATP & Enzymes What is it? Where do organism s get it? How do they use it? Where does Energy come from? Ultimately, from the sun. It is transferred between organisms in the earth s lithosphere,
Name: Hour: Elements & Macromolecules in Organisms
 Name: Hour: Elements & Macromolecules in Organisms Most common elements in living things are carbon, hydrogen, nitrogen, and oxygen. These four elements constitute about 95% of your body weight. All compounds
Name: Hour: Elements & Macromolecules in Organisms Most common elements in living things are carbon, hydrogen, nitrogen, and oxygen. These four elements constitute about 95% of your body weight. All compounds
Experiment 10 Enzymes
 Experiment 10 Enzymes Enzymes are proteins that act as catalysts for biological reactions. Enzymes, like all catalysts, speed up reactions without being used up themselves. They do this by lowering the
Experiment 10 Enzymes Enzymes are proteins that act as catalysts for biological reactions. Enzymes, like all catalysts, speed up reactions without being used up themselves. They do this by lowering the
Oxygen-Binding Proteins
 Oxygen-Binding Proteins Myoglobin, Hemoglobin, Cytochromes bind O 2. Oxygen is transported from lungs to various tissues via blood in association with hemoglobin In muscle, hemoglobin gives up O 2 to myoglobin
Oxygen-Binding Proteins Myoglobin, Hemoglobin, Cytochromes bind O 2. Oxygen is transported from lungs to various tissues via blood in association with hemoglobin In muscle, hemoglobin gives up O 2 to myoglobin
Name Date Period. Keystone Review Enzymes
 Name Date Period Keystone Review Enzymes 1. In order for cells to function properly, the enzymes that they contain must also function properly. What can be inferred using the above information? A. Cells
Name Date Period Keystone Review Enzymes 1. In order for cells to function properly, the enzymes that they contain must also function properly. What can be inferred using the above information? A. Cells
Enzymes. A. a lipid B. a protein C. a carbohydrate D. a mineral
 Enzymes 1. All cells in multicellular organisms contain thousands of different kinds of enzymes that are specialized to catalyze different chemical reactions. Given this information, which of the following
Enzymes 1. All cells in multicellular organisms contain thousands of different kinds of enzymes that are specialized to catalyze different chemical reactions. Given this information, which of the following
Helices From Readily in Biological Structures
 The α Helix and the β Sheet Are Common Folding Patterns Although the overall conformation each protein is unique, there are only two different folding patterns are present in all proteins, which are α
The α Helix and the β Sheet Are Common Folding Patterns Although the overall conformation each protein is unique, there are only two different folding patterns are present in all proteins, which are α
Anabolic and Catabolic Reactions are Linked by ATP in Living Organisms
 Chapter 5: Microbial Metabolism Microbial Metabolism Metabolism refers to all chemical reactions that occur within a living a living organism. These chemical reactions are generally of two types: Catabolic:
Chapter 5: Microbial Metabolism Microbial Metabolism Metabolism refers to all chemical reactions that occur within a living a living organism. These chemical reactions are generally of two types: Catabolic:
The Kinetics of Enzyme Reactions
 The Kinetics of Enzyme Reactions This activity will introduce you to the chemical kinetics of enzyme-mediated biochemical reactions using an interactive Excel spreadsheet or Excelet. A summarized chemical
The Kinetics of Enzyme Reactions This activity will introduce you to the chemical kinetics of enzyme-mediated biochemical reactions using an interactive Excel spreadsheet or Excelet. A summarized chemical
Enzymes Enzymes Enzymes: proteins ( metabolic pathways & biological Lysozyme Purification of Enzymes:
 Enzymes: Enzymes More than 2000 different enzymes are currently known. The function of enzymes and other catalysts is to lower the activation energy, ΔG, for a reaction and thereby enhance the reaction
Enzymes: Enzymes More than 2000 different enzymes are currently known. The function of enzymes and other catalysts is to lower the activation energy, ΔG, for a reaction and thereby enhance the reaction
Elements & Macromolecules in Organisms
 Name: Date: Per: Table # Elements & Macromolecules in rganisms Most common elements in living things are carbon, hydrogen, nitrogen, and oxygen. These four elements constitute about 95% of your body weight.
Name: Date: Per: Table # Elements & Macromolecules in rganisms Most common elements in living things are carbon, hydrogen, nitrogen, and oxygen. These four elements constitute about 95% of your body weight.
Chapter 3 Molecules of Cells
 Bio 100 Molecules of cells 1 Chapter 3 Molecules of Cells Compounds containing carbon are called organic compounds Molecules such as methane that are only composed of carbon and hydrogen are called hydrocarbons
Bio 100 Molecules of cells 1 Chapter 3 Molecules of Cells Compounds containing carbon are called organic compounds Molecules such as methane that are only composed of carbon and hydrogen are called hydrocarbons
1. Enzymes. Biochemical Reactions. Chapter 5: Microbial Metabolism. 1. Enzymes. 2. ATP Production. 3. Autotrophic Processes
 Chapter 5: Microbial Metabolism 1. Enzymes 2. ATP Production 3. Autotrophic Processes 1. Enzymes Biochemical Reactions All living cells depend on biochemical reactions to maintain homeostasis. All of the
Chapter 5: Microbial Metabolism 1. Enzymes 2. ATP Production 3. Autotrophic Processes 1. Enzymes Biochemical Reactions All living cells depend on biochemical reactions to maintain homeostasis. All of the
A disaccharide is formed when a dehydration reaction joins two monosaccharides. This covalent bond is called a glycosidic linkage.
 CH 5 Structure & Function of Large Molecules: Macromolecules Molecules of Life All living things are made up of four classes of large biological molecules: carbohydrates, lipids, proteins, and nucleic
CH 5 Structure & Function of Large Molecules: Macromolecules Molecules of Life All living things are made up of four classes of large biological molecules: carbohydrates, lipids, proteins, and nucleic
Chapter 3: Biological Molecules. 1. Carbohydrates 2. Lipids 3. Proteins 4. Nucleic Acids
 Chapter 3: Biological Molecules 1. Carbohydrates 2. Lipids 3. Proteins 4. Nucleic Acids Elements in Biological Molecules Biological macromolecules are made almost entirely of just 6 elements: Carbon (C)
Chapter 3: Biological Molecules 1. Carbohydrates 2. Lipids 3. Proteins 4. Nucleic Acids Elements in Biological Molecules Biological macromolecules are made almost entirely of just 6 elements: Carbon (C)
Computational Systems Biology. Lecture 2: Enzymes
 Computational Systems Biology Lecture 2: Enzymes 1 Images from: David L. Nelson, Lehninger Principles of Biochemistry, IV Edition, Freeman ed. or under creative commons license (search for images at http://search.creativecommons.org/)
Computational Systems Biology Lecture 2: Enzymes 1 Images from: David L. Nelson, Lehninger Principles of Biochemistry, IV Edition, Freeman ed. or under creative commons license (search for images at http://search.creativecommons.org/)
Chapter 2 Polar Covalent Bonds; Acids and Bases
 John E. McMurry http://www.cengage.com/chemistry/mcmurry Chapter 2 Polar Covalent Bonds; Acids and Bases Javier E. Horta, M.D., Ph.D. University of Massachusetts Lowell Polar Covalent Bonds: Electronegativity
John E. McMurry http://www.cengage.com/chemistry/mcmurry Chapter 2 Polar Covalent Bonds; Acids and Bases Javier E. Horta, M.D., Ph.D. University of Massachusetts Lowell Polar Covalent Bonds: Electronegativity
Paper: 6 Chemistry 2.130 University I Chemistry: Models Page: 2 of 7. 4. Which of the following weak acids would make the best buffer at ph = 5.0?
 Paper: 6 Chemistry 2.130 University I Chemistry: Models Page: 2 of 7 4. Which of the following weak acids would make the best buffer at ph = 5.0? A) Acetic acid (Ka = 1.74 x 10-5 ) B) H 2 PO - 4 (Ka =
Paper: 6 Chemistry 2.130 University I Chemistry: Models Page: 2 of 7 4. Which of the following weak acids would make the best buffer at ph = 5.0? A) Acetic acid (Ka = 1.74 x 10-5 ) B) H 2 PO - 4 (Ka =
Bioenergetics. Free Energy Change
 Bioenergetics Energy is the capacity or ability to do work All organisms need a constant supply of energy for functions such as motion, transport across membrane barriers, synthesis of biomolecules, information
Bioenergetics Energy is the capacity or ability to do work All organisms need a constant supply of energy for functions such as motion, transport across membrane barriers, synthesis of biomolecules, information
Preliminary MFM Quiz
 Preliminary MFM Quiz 1. The major carrier of chemical energy in all cells is: A) adenosine monophosphate B) adenosine diphosphate C) adenosine trisphosphate D) guanosine trisphosphate E) carbamoyl phosphate
Preliminary MFM Quiz 1. The major carrier of chemical energy in all cells is: A) adenosine monophosphate B) adenosine diphosphate C) adenosine trisphosphate D) guanosine trisphosphate E) carbamoyl phosphate
Enzymes: Practice Questions #1
 Enzymes: Practice Questions #1 1. Compound X increases the rate of the reaction below. Compound X is most likely A. an enzyme B. a lipid molecule C. an indicator D. an ADP molecule 2. The equation below
Enzymes: Practice Questions #1 1. Compound X increases the rate of the reaction below. Compound X is most likely A. an enzyme B. a lipid molecule C. an indicator D. an ADP molecule 2. The equation below
The purpose of this lab is to investigate the impact of temperature, substrate concentration,
 Lee 1 Jessica Lee AP Biology Mrs. Kingston 23 October 2013 Abstract: The purpose of this lab is to investigate the impact of temperature, substrate concentration, enzyme concentration, and the presence
Lee 1 Jessica Lee AP Biology Mrs. Kingston 23 October 2013 Abstract: The purpose of this lab is to investigate the impact of temperature, substrate concentration, enzyme concentration, and the presence
The Citric Acid Cycle
 The itric Acid ycle February 14, 2003 Bryant Miles I. itrate Synthase + 3 SoA The first reaction of the citric acid cycle is the condensation of acetyloa and oxaloacetate to form citrate and oas. The enzyme
The itric Acid ycle February 14, 2003 Bryant Miles I. itrate Synthase + 3 SoA The first reaction of the citric acid cycle is the condensation of acetyloa and oxaloacetate to form citrate and oas. The enzyme
Introduction, Noncovalent Bonds, and Properties of Water
 Lecture 1 Introduction, Noncovalent Bonds, and Properties of Water Reading: Berg, Tymoczko & Stryer: Chapter 1 problems in textbook: chapter 1, pp. 23-24, #1,2,3,6,7,8,9, 10,11; practice problems at end
Lecture 1 Introduction, Noncovalent Bonds, and Properties of Water Reading: Berg, Tymoczko & Stryer: Chapter 1 problems in textbook: chapter 1, pp. 23-24, #1,2,3,6,7,8,9, 10,11; practice problems at end
(c) How would your answers to problem (a) change if the molecular weight of the protein was 100,000 Dalton?
 Problem 1. (12 points total, 4 points each) The molecular weight of an unspecified protein, at physiological conditions, is 70,000 Dalton, as determined by sedimentation equilibrium measurements and by
Problem 1. (12 points total, 4 points each) The molecular weight of an unspecified protein, at physiological conditions, is 70,000 Dalton, as determined by sedimentation equilibrium measurements and by
Amino Acids, Peptides, Proteins
 Amino Acids, Peptides, Proteins Functions of proteins: Enzymes Transport and Storage Motion, muscle contraction Hormones Mechanical support Immune protection (Antibodies) Generate and transmit nerve impulses
Amino Acids, Peptides, Proteins Functions of proteins: Enzymes Transport and Storage Motion, muscle contraction Hormones Mechanical support Immune protection (Antibodies) Generate and transmit nerve impulses
Biological molecules:
 Biological molecules: All are organic (based on carbon). Monomers vs. polymers: Monomers refer to the subunits that, when polymerized, make up a larger polymer. Monomers may function on their own in some
Biological molecules: All are organic (based on carbon). Monomers vs. polymers: Monomers refer to the subunits that, when polymerized, make up a larger polymer. Monomers may function on their own in some
Recognizing Organic Molecules: Carbohydrates, Lipids and Proteins
 Recognizing Organic Molecules: Carbohydrates, Lipids and Proteins Oct 15 8:05 PM What is an Organic Molecule? An Organic Molecule is a molecule that contains carbon and hydrogen and oxygen Carbon is found
Recognizing Organic Molecules: Carbohydrates, Lipids and Proteins Oct 15 8:05 PM What is an Organic Molecule? An Organic Molecule is a molecule that contains carbon and hydrogen and oxygen Carbon is found
Chapter 3. Protein Structure and Function
 Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
CHAPTER 11 Mechanism of Enzyme Action
 CHAPTER 11 Mechanism of Enzyme Action 1. General properties of enzymes 2. Activation energy and the reaction coordinate 3. Catalytic mechanism 4. Lysozyme 5. Serine proteases Enzyme act with great speed
CHAPTER 11 Mechanism of Enzyme Action 1. General properties of enzymes 2. Activation energy and the reaction coordinate 3. Catalytic mechanism 4. Lysozyme 5. Serine proteases Enzyme act with great speed
Myoglobin and Hemoglobin
 Myoglobin and Hemoglobin Myoglobin and hemoglobin are hemeproteins whose physiological importance is principally related to their ability to bind molecular oxygen. Myoglobin (Mb) The oxygen storage protein
Myoglobin and Hemoglobin Myoglobin and hemoglobin are hemeproteins whose physiological importance is principally related to their ability to bind molecular oxygen. Myoglobin (Mb) The oxygen storage protein
Chapter 5: The Structure and Function of Large Biological Molecules
 Name Period Concept 5.1 Macromolecules are polymers, built from monomers 1. The large molecules of all living things fall into just four main classes. Name them. 2. Circle the three classes that are called
Name Period Concept 5.1 Macromolecules are polymers, built from monomers 1. The large molecules of all living things fall into just four main classes. Name them. 2. Circle the three classes that are called
Overview of Glycolysis Under anaerobic conditions, the glycolytic pathway present in most species results in a balanced reaction:
 Glycolysis Glucose is a valuable molecule. It can be used to generate energy (in red blood cells and in brain under normal conditions, glucose is the sole energy source), and it can be used to generate
Glycolysis Glucose is a valuable molecule. It can be used to generate energy (in red blood cells and in brain under normal conditions, glucose is the sole energy source), and it can be used to generate
Amino Acids. Amino acids are the building blocks of proteins. All AA s have the same basic structure: Side Chain. Alpha Carbon. Carboxyl. Group.
 Protein Structure Amino Acids Amino acids are the building blocks of proteins. All AA s have the same basic structure: Side Chain Alpha Carbon Amino Group Carboxyl Group Amino Acid Properties There are
Protein Structure Amino Acids Amino acids are the building blocks of proteins. All AA s have the same basic structure: Side Chain Alpha Carbon Amino Group Carboxyl Group Amino Acid Properties There are
Chemical Basis of Life Module A Anchor 2
 Chemical Basis of Life Module A Anchor 2 Key Concepts: - Water is a polar molecule. Therefore, it is able to form multiple hydrogen bonds, which account for many of its special properties. - Water s polarity
Chemical Basis of Life Module A Anchor 2 Key Concepts: - Water is a polar molecule. Therefore, it is able to form multiple hydrogen bonds, which account for many of its special properties. - Water s polarity
Chapter 16 Amino Acids, Proteins, and Enzymes
 Chapter 16 Amino Acids, Proteins, and Enzymes 1 Functions of Proteins Proteins in the body are polymers made from 20 different amino acids differ in characteristics and functions that depend on the order
Chapter 16 Amino Acids, Proteins, and Enzymes 1 Functions of Proteins Proteins in the body are polymers made from 20 different amino acids differ in characteristics and functions that depend on the order
ph: Measurement and Uses
 ph: Measurement and Uses One of the most important properties of aqueous solutions is the concentration of hydrogen ion. The concentration of H + (or H 3 O + ) affects the solubility of inorganic and organic
ph: Measurement and Uses One of the most important properties of aqueous solutions is the concentration of hydrogen ion. The concentration of H + (or H 3 O + ) affects the solubility of inorganic and organic
Introduction to Proteins and Enzymes
 Introduction to Proteins and Enzymes Basics of protein structure and composition The life of a protein Enzymes Theory of enzyme function Not all enzymes are proteins / not all proteins are enzymes Enzyme
Introduction to Proteins and Enzymes Basics of protein structure and composition The life of a protein Enzymes Theory of enzyme function Not all enzymes are proteins / not all proteins are enzymes Enzyme
Combinatorial Biochemistry and Phage Display
 Combinatorial Biochemistry and Phage Display Prof. Valery A. Petrenko Director - Valery Petrenko Instructors Galina Kouzmitcheva and I-Hsuan Chen Auburn 2006, Spring semester COMBINATORIAL BIOCHEMISTRY
Combinatorial Biochemistry and Phage Display Prof. Valery A. Petrenko Director - Valery Petrenko Instructors Galina Kouzmitcheva and I-Hsuan Chen Auburn 2006, Spring semester COMBINATORIAL BIOCHEMISTRY
Summary of Metabolism. Mechanism of Enzyme Action
 Summary of Metabolism Mechanism of Enzyme Action 1. The substrate contacts the active site 2. The enzyme-substrate complex is formed. 3. The substrate molecule is altered (atoms are rearranged, or the
Summary of Metabolism Mechanism of Enzyme Action 1. The substrate contacts the active site 2. The enzyme-substrate complex is formed. 3. The substrate molecule is altered (atoms are rearranged, or the
Molecular Models in Biology
 Molecular Models in Biology Objectives: After this lab a student will be able to: 1) Understand the properties of atoms that give rise to bonds. 2) Understand how and why atoms form ions. 3) Model covalent,
Molecular Models in Biology Objectives: After this lab a student will be able to: 1) Understand the properties of atoms that give rise to bonds. 2) Understand how and why atoms form ions. 3) Model covalent,
H H N - C - C 2 R. Three possible forms (not counting R group) depending on ph
 Amino acids - 0 common amino acids there are others found naturally but much less frequently - Common structure for amino acid - C, -N, and functional groups all attached to the alpha carbon N - C - C
Amino acids - 0 common amino acids there are others found naturally but much less frequently - Common structure for amino acid - C, -N, and functional groups all attached to the alpha carbon N - C - C
Regulation of the Citric Acid Cycle
 Regulation of the itric Acid ycle I. hanges in Free Energy February 17, 2003 Bryant Miles kj/mol 40 20 0 20 40 60 80 Reaction DGo' DG TA Free Energy hanges 1 2 3 4 5 6 7 8 9 1.) itrate Synthase 2.) Aconitase
Regulation of the itric Acid ycle I. hanges in Free Energy February 17, 2003 Bryant Miles kj/mol 40 20 0 20 40 60 80 Reaction DGo' DG TA Free Energy hanges 1 2 3 4 5 6 7 8 9 1.) itrate Synthase 2.) Aconitase
CHEMISTRY STANDARDS BASED RUBRIC ATOMIC STRUCTURE AND BONDING
 CHEMISTRY STANDARDS BASED RUBRIC ATOMIC STRUCTURE AND BONDING Essential Standard: STUDENTS WILL UNDERSTAND THAT THE PROPERTIES OF MATTER AND THEIR INTERACTIONS ARE A CONSEQUENCE OF THE STRUCTURE OF MATTER,
CHEMISTRY STANDARDS BASED RUBRIC ATOMIC STRUCTURE AND BONDING Essential Standard: STUDENTS WILL UNDERSTAND THAT THE PROPERTIES OF MATTER AND THEIR INTERACTIONS ARE A CONSEQUENCE OF THE STRUCTURE OF MATTER,
Ionization of amino acids
 Amino Acids 20 common amino acids there are others found naturally but much less frequently Common structure for amino acid COOH, -NH 2, H and R functional groups all attached to the a carbon Ionization
Amino Acids 20 common amino acids there are others found naturally but much less frequently Common structure for amino acid COOH, -NH 2, H and R functional groups all attached to the a carbon Ionization
PROTEINS THE PEPTIDE BOND. The peptide bond, shown above enclosed in the blue curves, generates the basic structural unit for proteins.
 Ca 2+ The contents of this module were developed under grant award # P116B-001338 from the Fund for the Improvement of Postsecondary Education (FIPSE), United States Department of Education. However, those
Ca 2+ The contents of this module were developed under grant award # P116B-001338 from the Fund for the Improvement of Postsecondary Education (FIPSE), United States Department of Education. However, those
Review of Chemical Equilibrium 7.51 September 1999. free [A] (µm)
![Review of Chemical Equilibrium 7.51 September 1999. free [A] (µm) Review of Chemical Equilibrium 7.51 September 1999. free [A] (µm)](/thumbs/40/21435474.jpg) Review of Chemical Equilibrium 7.51 September 1999 Equilibrium experiments study how the concentration of reaction products change as a function of reactant concentrations and/or reaction conditions. For
Review of Chemical Equilibrium 7.51 September 1999 Equilibrium experiments study how the concentration of reaction products change as a function of reactant concentrations and/or reaction conditions. For
Lecture 6. Regulation of Protein Synthesis at the Translational Level
 Regulation of Protein Synthesis (6.1) Lecture 6 Regulation of Protein Synthesis at the Translational Level Comparison of EF-Tu-GDP and EF-Tu-GTP conformations EF-Tu-GDP EF-Tu-GTP Next: Comparison of GDP
Regulation of Protein Synthesis (6.1) Lecture 6 Regulation of Protein Synthesis at the Translational Level Comparison of EF-Tu-GDP and EF-Tu-GTP conformations EF-Tu-GDP EF-Tu-GTP Next: Comparison of GDP
Chapter 6 An Overview of Organic Reactions
 John E. McMurry www.cengage.com/chemistry/mcmurry Chapter 6 An Overview of Organic Reactions Why this chapter? To understand organic and/or biochemistry, it is necessary to know: -What occurs -Why and
John E. McMurry www.cengage.com/chemistry/mcmurry Chapter 6 An Overview of Organic Reactions Why this chapter? To understand organic and/or biochemistry, it is necessary to know: -What occurs -Why and
Biochemistry 462a Hemoglobin Structure and Function Reading - Chapter 7 Practice problems - Chapter 7: 1-6; Proteins extra problems
 Biochemistry 462a Hemoglobin Structure and Function Reading - Chapter 7 Practice problems - Chapter 7: 1-6; Proteins extra problems Myoglobin and Hemoglobin Oxygen is required for oxidative metabolism
Biochemistry 462a Hemoglobin Structure and Function Reading - Chapter 7 Practice problems - Chapter 7: 1-6; Proteins extra problems Myoglobin and Hemoglobin Oxygen is required for oxidative metabolism
Exam 4 Outline CH 105 Spring 2012
 Exam 4 Outline CH 105 Spring 2012 You need to bring a pencil and your ACT card. Chapter 24: Lipids 1. Describe the properties and types of lipids a. All are hydrophobic b. Fatty acid-based typically contain
Exam 4 Outline CH 105 Spring 2012 You need to bring a pencil and your ACT card. Chapter 24: Lipids 1. Describe the properties and types of lipids a. All are hydrophobic b. Fatty acid-based typically contain
