Why do we have linear chromosomes? A matter of Adam and Eve
|
|
|
- Baldric Poole
- 9 years ago
- Views:
Transcription
1 Mutation Research 434 Ž Community address: Minireview Why do we have linear chromosomes? A matter of Adam and Eve Fuyuki Ishikawa ), Taku Naito Laboratory of Molecular and Cellular Assembly, Graduate School of Biological Information, Tokyo Institute of Technology, 4259 Nagatsuta, Midori-ku, Yokohama , Japan Accepted 30 March 1999 Keywords: Linear chromosome; Circular chromosome; Telomere; Meiosis 1. Introduction It is usually assumed that prokaryotic cells have circular chromosomes, whereas eukaryotic cells have linear chromosomes. One of the consequences of linear chromosomes is the presence of chromosomal ends called telomeres. Simple physical ends of DNA, such as those produced by DNA double-strand breaks Ž DSB. by ionizing radiation, are genetically unstable, mutagenic, and sometimes oncogenic Žreviewed in Ref. wx. 1. Telomeres are a complex composed of telomeric DNA and a number of telomere-specific and non-specific proteins. This large molecular assembly that forms the telomeres protects the genomic ends from end-to-end fusion or exonucleolytic erosion Žreviewed in Ref. wx. 2. Due to the end-replication problem, telomeric DNA is shortened as the cell divides wx 3. In most eukaryotes, this shortening of telomeric DNA is compensated by the activity of an enzyme called telomerase that synthesizes telomeric DNA de novo wx 4. However, telomerase is strictly regulated to be inactive in most human somatic cells, and telomere lengths decline as an individual ages wx 5. This results ) Corresponding author. fishikaw@bio.titech.ac.jp in cellular senescence and cancer development due to telomere insufficiencies Že.g., Ref. wx 6, and rewx. 7. Therefore, telomeres are some- viewed in Ref. times referred to as the the Achilles heel of the chromosome wx 8. Why do we have linear chromosomes that lead to senescence and cancers, instead of circular chromosomes? In this article, we review as to what extent different chromosome configurations are conserved among different kingdoms, and propose a hypothesis to explain why this remarkable conservation has evolved. 2. Chromosome configurations of prokaryotes and eukaryotes In this article, we operatively define chromosomes as genetic materials containing house-keeping genes essential for the cell s survival that replicate synchronously with cell division to distinguish them from extra-chromosomal genetic elements, such as plasmids, bacteriophage and transposons wx 9. The recent invention of molecular biological tools to analyze large DNA structures, especially pulse field gel electrophoresis Ž PFGE., in addition to classical r99r$ - see front matter q 1999 Elsevier Science B.V. All rights reserved. Ž. PII: S
2 100 F. Ishikawa, T. NaitorMutation Research tools, such as genetic linkage studies, have greatly enriched our knowledge about the chromosome configuration in different species, phyla and kingdoms. Given the very large number of species on the earth, both known and unknown, it is far from possible to make a complete catalogue of the chromosome structures existing among the diverse range of species. However, recent studies have identified the presence of several exceptional cases to the general rule that one circular chromosome is present in prokaryotic cells and more than one linear chromosome are present in eukaryotic cells. Nevertheless, these exceptional cases indeed further strengthen the notion that prokaryotes and eukaryotes have maintained circular and linear chromosomes, respectively, throughout their evolution Prokaryotes Generally, the small sizes of prokaryotic genomes Ž typically 1 10 Mb. make it feasible to carry out physical analyses, such as PFGE and total nucleotide sequencing Žreviewed in Ref. w10 x.. Genome analyses of Archea have lagged behind that of eubacteria. For example, the genome sequencing of only six archebacteria had been completed, whereas 17 eubacteria had been sequenced at the time of this review Ž All archebacteria examined so far contain one circular chromosome. Therefore, we will focus on several exceptional cases in eubacteria, where this general rule is not applied. Borrelia, Treponema and Leptospira are members of the spirochete group Ž Phylum Spirochaetae.. They are pathogenic bacteria, and Borrelia causes Lyme disease. Borrelia burgdorferi has been shown to have one 910-kb linear chromosome w11 13 x. This unique feature is conserved in the other members of the Borrelia genus w14 x. In contrast, other two closely related genera, Treponema and Leptospira, which also belong to the Spirochaetae, contain circular chromosomes w15 17 x. This specific distribution of linear chromosomes in the Borrelia genus suggests that Borrelia s linear chromosomes were evolutionarily derived from the ancestral circular chromosomes after the three genera had diverged. Streptomyces is another genus that has been shown to possess linear chromosomes w18 x. It should be noted that Streptomyces, a member of Phylum Actinomycete, is phylogenically distant from Borrelia. The closely related genus, Mycobacterium, which is within the same phylum, contains a conventional single circular chromosome w19 x, again suggesting the relatively recent origin of the linear chromosome in Streptomyces. Finally, Agrobacterium tumefaciens, a member of Proteobacteria has one 2.1-Mb linear and three 3- Mb, 450-kb and 200-kb circular replicons. It is known that at least the two 1-Mb and 3-Mb replicons contain metabolically essential genes, which classifies them as chromosomes w20 x. The conventional DNA replication mechanism does not replicate the very ends of linear DNA, because all DNA polymerases need primers for initiating synthesis. The diverse range of linear genomes solves this end-replication problem by different strategies. Adenovirus initiates the replication of its linear genome using a protein primer Žterminal protein, TP. w21 x. TP forms a covalent bond with the 5 X -OH of dnmp, and the DNA polymerase starts synthesis using this base as the first nucleotide to be incorporated. As a result, adenovirus DNA has a covalently associated TP at its 5 X -ends. On the other hand, vaccinia virus has a hairpin structure at both ends of its linear genome w22 x. One strand is continuous to the other strand, and DNA synthesis continues onto the next strand after completing one strand. A palindromic sequence is left after nicking the hairpin DNA to resolve the two daughter duplex DNAs. The terminal structures of the linear genomic DNAs of Borrelia and Streptomyces have also been reported. Streptomyces has 5 X -end associated proteins, suggesting that the telomeres are replicated by TP primers in this bacterium w23,24 x. In contrast, Borrelia has hairpin structures with 26-bp inverted repeats at both telomeres, suggesting that its telomeres are replicated in a way similar to vaccinia virus w25 x. These studies indicate that at least three prokaryotic genera possess linear chromosomes, instead of circular ones. However, the appearance of linear chromosomes in prokaryotes seems rather sporadic. First, these three genera are distantly related to each other. Second, closely related genera belonging to the same phylum contain conventional circular chromosomes. Finally, the solutions for the end-replication problems differ between Borrelia and Strep-
3 F. Ishikawa, T. NaitorMutation Research tomyces. Accordingly, it is suggested that prokaryote linear chromosomes have not been inherited directly from one ancient prokaryote that had linear chromosomes. Instead, they most likely have developed recently from the circular chromosomes of an ancestor species. Therefore, it may be concluded that the prokaryote genomes have been maintained phylogenically in circular forms Eukaryotes Eukaryotes contain larger genomes than prokaryotes Ž typically larger than 10 Mb.. Accordingly, in many cases, the chromosomes can be visualized by microscopy to analyze the gross structures. However, large chromosome sizes are a disadvantage in another respect, since it is usually difficult to construct a physical map of the genome. As will be reviewed here, many reports have been published showing the presence of circular chromosomes. However, the evidence for covalently linked circular chromosomes is not available in most cases. To avoid possible confusion, circular chromosomes judged solely on morphological criteria will be called ring chromosomes in this review. Circular chromosomes have been reported both in budding yeast w26,27x and fission yeast w28,29 x. They were isolated spontaneously or artificially, and have been shown to be circular by either a genetic or a physical approach. In each case, only one circular chromosome was identified, and it was unstable mitotically and meiotically. As there are many opportunities to examine karyotypes in a variety of medical settings, many cases of ring chromosomes have been reported to be associated with a variety of clinical manifestations. In most cases, the ring chromosomes have been found in somatic cells, either normal or cancerous, and either constitutively or in mosaicism. However, few notable cases in which one ring chromosome had apparently been inherited from one of the parents who also had the same ring chromosomes have been reported Že.g., Ref. w30 x.. In these cases, the parents generally showed mosaic ring chromosomes, suggesting that an individual who has a ring chromosome constitutively is infertile. There has been no report describing the inheritance of more than one ring chromosomes. These results have suggested that in some rare cases, a single ring chromosome may be normally segregated in meiosis, and fertilized. In summary, circular or ring chromosomes have been found sporadically in eukaryotes. With some rare exceptions, they are not usually inherited. However, there have been no reports describing eukaryotic cells having more than one circular or ring chromosome that have been meiotically transmitted. These results suggest that ring chromosomes face some difficulty in sexual reproduction. Since mitochondrial DNAs are circular in most eukaryotes, chromosome circularity itself is obviously not incompatible with inheritance. Circular chromosomes may be incompatible with a process specific to sexual reproduction, such as meiosis. This section has shown that two chromosome configurations, circular and linear, are remarkably conserved in prokaryotes and eukaryotes, respectively. In an evolutionary sense, linear chromosomes require extra energy to maintain intact telomeres, which is not a requirement for circular chromosomes. This reasoning suggests that there must be some advantages to eukaryotes that have been acquired in a trade-off for this extra burden. One of the most direct experiments to test this hypothesis would be to construct a eukaryotic cell that maintains its genome in a circular form, and to see what biological functions this cell has lost. However, until recently, there has been no report that describes the existence of eukaryotic cells that maintain completely circular genomes. 3. ATM family genes and telomeres Telomeres are comprised of many components to accomplish its functions. Proteins involved in telomere maintenance are now being studied in some detail, especially in simple eukaryotic cells such as yeast Žreviewed in Ref. w31 x.. One group of interesting proteins thus identified is the ATM family. The Saccharomyces cereõisiae TEL1 gene was originally identified by screening for mutants with short telomw32 x. When this gene was cloned, it ere phenotypes turned out to have a significant level of homology with the human ATM gene, whose mutations cause the hereditary disease, ataxia telangiectasia Ž A-T.
4 102 F. Ishikawa, T. NaitorMutation Research w33,34 x. Interestingly, telomeres in A-T cells also showed excessive shortening w35,36 x, suggesting that the ATM family genes are involved in telomere maintenance in all cells from yeast to human. There is at least more than one member of the ATM family genes present in one species w37 x. The budding yeast Saccharomyces has TEL1 and MEC1 genes, and human has ATM, ATR and the relatively distant member DNA-PK. All these proteins have a PI3 Ž phosphatidylinositol 3. -kinase-like domain at the C-termini, yet they have protein kinase activity. Fission yeast, Schizosaccharomyces pombe, also has two ATM family genes, named rad3 q and tel1 q w x q q 38,39. The mutant defective for rad3 or tel1 shows moderate to minimal telomere shortenings w39,40 x. However, when both ATM family genes were mutated, there was an additive effect, and the fission yeast chromosomes essentially lost all telomw39 x. Thus, the ATM family genes are ere sequences redundant but essential for stable telomere maintenance. The rad3 tel1 double mutant cells grow very slowly with a low viability, and showed aberrantly irregular colony shapes, as expected for cells suffering from extensive telomere shortening. However, derivative cells that showed apparently normal colony shapes appeared spontaneously among these double mutants at a relatively high frequency. Surprisingly, these derivative cells contain three self-circularized chromosomes Žfission yeast contains three linear chromosomes. w39 x. This case is the first report describing the existence of eukaryotic cells that maintain the genome exclusively in circular forms. Previously, it had been postulated that eukaryotic cells do not have circular chromosomes because if an odd number of crossing-over events occurs between the two sister chromatids by SCE, this would result in the formation of dicentric circular chromosomes. Thus, formed circular dimer chromosomes would be eventually broken during mitosis by the two spindles pulling them apart Že.g., see discussions in Refs. w41,42 x.. Since yeasts undergo efficient homologous recombination and SCEs, we would expect that complete circular genomes in yeast should be highly unstable and lead to cell death. Indeed, the fission yeast rad3 tel1 mutant with the three circular chromosomes showed anaphase bridges and some degree of aneuploidy Ž Naito and Ishikawa, unpublished.. However, the fact that this mutant grew well mitotically as a mass suggests that the SCE of dicentric circular chromosomes may have a relatively small effect. Indeed, SCE happens in prokaryotic cells to produce circular dimers. In Escherichia coli, these circular dimers are known to be resolved by both the reca-independent resolvases, XerC and XerD, that act on a specific locus called dif located at the replication terminus, and the reca-dependent recombination pathway Žreviewed in Refs. w43,44 x.. In higher eukaryotes, genomic DNA is organized into multiple loops by tight association of matrix-associated region Ž MAR. on DNA with nuclear scaffolds. In a topological sense, each loop can be assumed microscopically to be a closed circle. Therefore, closed circular oligomers may also be formed by SCE in linear eukaryotic genomes, making the hypothesis that the circular dimer formation prohibits circular chromosomes in eukaryotes unlikely. Eukaryotes may have mechanisms similar to XerCD and reca-dependent recombination in E. coli to resolve these microscopic oligomers. Indeed, RAD51-deficient chicken cells, the eukaryote recahomologue, are shown to be arrested in G2rM phase and to accumulate chromosome breaks, suggesting the possible involvement of the Rad51 protein in resolving SCE-intermediates w45 x. 4. Sister chromatid exchanges SCEs and circular dimer formation 5. Meiosis and telomeres Telomeres perform a number of important functions in different biological situations. Recently, the role of the telomeres in meiosis has come into light Žreviewed in Ref. w46 x.. The first hint came from the cytological observation that telomeres are closely clustered with each other at a specific stage of meiosis. There have been ample observations that telomeres and centromeres are positioned asymmetrically in nuclei Žreviewed in Ref. w47 x.. In mitotic interphase, the centromeres have a tendency to cluster around the centriole, presumably reflecting the
5 F. Ishikawa, T. NaitorMutation Research association between these two structures during the last anaphase. This centromere clustering is named the Rabl w48x orientation after the German scientist who first described it. However, during meiotic prophase, this relative distribution of centromeres and telomeres is reversed: In the leptotenerzygotene stage, telomeres, instead of centromeres, are clustered at the inner surface of the nuclear envelope. The centromeres are distributed randomly in the nucleus at this stage. This polarized chromosomal distribution is called the bouquet arrangement w49 x, and is found during meiosis in many species Žre- w50 x.. The functional significance of viewed in Ref. this peculiar conformation has been recently revealed. In fission yeast, two haploid cells of opposite mating types conjugate to produce diploid cells Ž karyogamy., and enter meiosis Ž zygotic meiosis.. Immediately after karyogamy, the fused nucleus forms an elongated shape, called a horse-tail w51 x. Using time-elapsed image recording, the fused horse-tail nuclei were found to undergo a dynamic to-and-fro oscillating movement w52 x. Moreover, by a combination of telomere-specific fluorescence in situ hybridization Ž FISH. and immunostaining of the spindle pole body ŽSPB, a centriole-counterpart in yeast., it was shown that six telomeres of the three fission yeast chromosomes are closely associated with SPB, and they lead the front edge of this horse-tail movement. In fission yeast, the bouquet arrangement is established by an association between the telomeres and SPB. In this way, the microtubule enucleated from the SPB promotes the dynamic nuclear movement by pulling the telomeres and dragging the chromosomes behind as a mass Žreviewed in Ref. w53 x.. Recently, three additional genetic studies have further indicated the importance of telomeres in the meiotic process. Telomere DNA consists of wellconserved G-rich simple tandem repeats. Telomere DNA-specific binding proteins are known to exist in several species. These include TRF1 and TRF2 in human w54 x, Rap1 in budding yeast w55 x, and taz1p in fission yeast w56 x. Fission yeast mutants defective for taz1 q failed to form the telomere clustering at the horse-tail stage of pre-meiosis, and showed reduced spore viability w57,58 x. Taz1p is presumably involved in the SPB-telomere association w53 x. The rad3 tel1 fission mutant with the three circular chromosomes was examined for spore viability after azygotic meiosis w39 x. In this case, diploid cells derived from two haploid cells harboring circular chromosomes produced no viable spores, a phenotype more profound than that of the taz1 mutant. These studies clearly indicated that telomeres are essential for a productive meiotic process. 6. How telomeres are essential for meiosis Several scenarios can be proposed to explain why functional telomeres are essential for meiosis. Meiosis consists of two successive cell divisions, called meiosis I and meiosis II. Meiosis II is similar to mitotic cell division, but meiosis I is unique. Meiosis is a process that produces four haploid cells from one diploid cell. Meiosis I is responsible for this reduction of ploidy by segregating the two homologous chromosomes to the two daughter cells. To accomplish this reductional segregation, each pair of homologous chromosomes needs to be paired before the onset of meiosis I Žreviewed in Ref. w59 x.. Homologous chromosome pairing has another important role in inducing homologous recombination between the two homologues. This homologous recombination shuffles the two alleles originally derived from different individuals Ž father and mother., and ensures that the haploid cells contain chimeric genetic information. Moreover, the recombination and segregation are interdependent, since the covalent associations formed by the recombinational Holiday junction between the two homologues Ž chiasmata. are thought to be essential for stable homologue pairing, and ensuring proper segregation. Therefore, homologue-pairing is at the heart of the mechanism of meiosis. Circular chromosomes potentially undergo more than one pathway during meiosis, and in all cases, they have very small probabilities of proper segregation Ž Fig. 1.. In normal meiotic prophase, linear chromosomes gather together by telomere-clustering Ž Fig. 1A.. Telomere associations of homologous chromosomes may help the homologue pairing by aligning the two chromosomes that are now tethered at both ends. After successful pairing, homologous recombination occurs between the two homologues,
6 104 F. Ishikawa, T. NaitorMutation Research Ž. Ž. Fig. 1. Meiosis I of linear chromosomes A and possible pathways in meiosis I of circular chromosomes B. For details, see the text.
7 F. Ishikawa, T. NaitorMutation Research and this covalent association further contributes to the stable chromosome pairing. In anaphase I, the Holiday junctions are resolved and the two homologues are segregated to different daughter cells. In contrast, circular chromosomes may undergo several different pathways Ž Fig. 1B.. As circular chromosomes lack functional telomeres, two homologues cannot be positioned in proximity. In this case, no homologue pairing and recombination occur, and the homologues are randomly segregated to daughter cells Ž pathway 1.. Two homologues may be positioned closely by chance, and somehow may pair and undergo recombination Ž pathway 2.. However, if an odd number of crossing-over events occurs between two homologues, this results in the formation of dicentric circular chromosomes Ž pathway 3.. If resolvase fails to resolve this form into monomers, the dicentric circle enters anaphase. When the spindles of different origins attach to each of the two kinetochores, the chromosome will be pulled apart and tear Ž pathway 4.. When a common spindle attaches to both of the two kinetochores, the chromosome is segregated to only one cell, with the other cell receiving no homologue. In either case, daughter cells will lose a significant amount of genetic information. When an even number of crossing-over events occurs Ž pathway 7., or the dicentric circles are resolved into monomeric circles Ž pathway 6., the two homologues may be segregated properly to the two daughter cells. However, even in this case, it is not known if spindles correctly attach to the kinetochores of chromosomes that have not been associated with telomeres during the meiotic prophase. Overall, the chance that one particular circular chromosome is segregated properly in meiosis I is very small. All eukaryotes contain more than one and usually many chromosomes. The chance, that one daughter cell will have all chromosomes properly segregated in circular forms, is the multiple of these small probabilities for each circular chromosome, and should be negligible. In conclusion, there is essentially no chance that all circular chromosomes are properly segregated during meiosis. This essential role of telomeres in accomplishing reductional chromosome segregation in meiosis must be the major reason that linear chromosomes are strikingly conserved in eukaryotes, which are characterized by the presence of sexual reproduction in most cases. 7. Conclusion We have stated that linear chromosomes are essential for productive meiosis. Meiosis Ža mechanism to generate haploid cells. is a prerequisite for shuffling the genetic information between individuals. It has been proposed that the production of genetically diverse offspring is advantageous in an ever-changw60 x. Indeed, the num- ing or saturated environment ber of absolutely asexual eukaryotic organisms is very small Žreviewed in Refs. w61,62 x., and the conservation of the potential of sexual reproduction seems to be as strong as the conservation of linear chromosomes in eukaryotes. Recent studies have indicated that chromosome linearity is important for meiosis, and we would like to propose that the correlation between sex and linear chromosomes is based on a mechanistic reason, and not on a superficial parallelism. Once upon a time, two groups of living creatures emerged from a common ancestor. One group decided to maintain genomes in circular forms, because this form is more economical without the need to maintain telomeres. However, the progeny of this group Ž Bacteria. is not able to exchange genomic information by meiosis and fertilization, and thus, needs to grow faster and keep the genome size as small as possible. The other group decided to maintain the genomes in linear forms. Although this strategy requires extra energy to maintain telomeres, these organisms have enjoyed the dynamic flow of genomic information by sexual reproduction. This process has allowed this group Ž Eukaryota. the chance to produce a variety of offspring. Accordingly, eukaryotes have complicated systems, and grow less rapidly than prokaryotes. Acknowledgements We thank E.A. Kamei Ž Gunma University. and H. Niki Ž Kumamoto University. for critical reading of and comments on the manuscript. The excellent secretarial works of F. Nishizaki, K. Saito and K. Yokoyama are acknowledged. This work was supported by a grant-in-aid from the Organization for Pharmaceutical Safety and Research, Japan.
8 106 F. Ishikawa, T. NaitorMutation Research References wx 1 T. de Lange, Telomere dynamics and genome instability in human cancer, in: E.H. Blackburn, C.W. Greider Ž Eds.., Telomeres, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 1995, pp wx 2 E.H. Blackburn, Structure and function of telomeres, Nature 350 Ž wx 3 J.D. Watson, Origin of concatemeric T7 DNA, Nature of New Biology 239 Ž wx 4 E.H. Blackburn, Telomerase, Annu. Rev. Biochem. 61 Ž wx 5 C.B. Harley, B. Villeponteau, Telomeres and telomerase in aging and cancer, Curr. Opin. Genet. Dev. 5 Ž wx 6 S. Hatakeyama, K. Fujita, M. Omine, F. Ishikawa, The jumping translocation at 1q21 involves shortened telomeres, Blood 91 Ž wx 7 F. Ishikawa, Telomere crisis, the driving force in cancer cell evolution, Biochem. Biophys. Res. Commun. 230 Ž wx 8 C.B. Harley, A.B. Futcher, C.W. Greider, Telomeres shorten during ageing of human fibroblasts, Nature 345 Ž wx 9 S. Krawiec, M. Riley, Organization of the bacterial chromosome, Microbiol. Rev. 54 Ž w10x S. Casjens, The diverse and dynamic structure of bacterial genomes, Annu. Rev. Genet. 32 Ž w11x M.S. Ferdows, A.G. Barbour, Megabase-sized linear DNA in the bacterium, Borrelia burgdorferi, the Lyme disease agent, Proc. Natl. Acad. Sci. USA 86 Ž w12x B.E. Davidson, J. MacDougall, I. Saint Girons, Physical map of the linear chromosome of the bacterium, Borrelia burgdorferi 212, a causative agent of Lyme disease, and localization of rrna genes, J. Bacteriol. 174 Ž w13x C.M. Fraser, S. Casjens, W.M. Huang, G.G. Sutton, R. Clayton, R. Lathigra, O. White, K.A. Ketchum, R. Dodson, E.K. Hickey, M. Gwinn, B. Dougherty, J.F. Tomb, R.D. Fleischmann, D. Richardson, J. Peterson, A.R. Kerlavage, J. Quackenbush, S. Salzberg, M. Hanson, R. van Vugt, N. Palmer, M.D. Adams, J. Gocayne, J.C. Venter, Genomic sequence of a Lyme disease spirochaete, Borrelia burgdorferi, Nature 390 Ž w14x S. Casjens, M. Delange, H.L.r. Ley, P. Rosa, W.M. Huang, Linear chromosomes of Lyme disease agent spirochetes: genetic diversity and conservation of gene order, J. Bacteriol. 177 Ž w15x R.L. Zuerner, Physical map of chromosomal and plasmid DNA comprising the genome of Leptospira interrogans, Nucleic Acids Res. 19 Ž w16x E.M. Walker, J.K. Howell, Y. You, A.R. Hoffmaster, H. JD, G.M. Weinstock, S.J. Norris, Physical map of the genome of Treponema pallidum subsp. pallidum Ž Nichols., J. Bacteriol. 177 Ž w17x J. MacDougall, I. Saint Girons, Physical map of the Treponema denticola circular chromosome, J. Bacteriol. 177 Ž w18x Y.S. Lin, H.M. Kieser, D.A. Hopwood, C.W. Chen, The chromosomal DNA of Streptomyces liõidans 66 is linear, Mol. Microbiol. 10 Ž w19x K. Eiglmeier, N. Honore, S.A. Woods, B. Caudron, S.T. Cole, Use of an ordered cosmid library to deduce the genomic organization of Mycobacterium leprae, Mol. Microbiol. 7 Ž w20x A. Allardet-Servent, S. Michaux-Charachon, E. Jumas-Bilak, L. Karayan, M. Ramuz, Presence of one linear and one circular chromosome in the Agrobacterium tumefaciens C58 genome, J. Bacteriol. 175 Ž w21x M.D. Challberg, S.V. Desiderio, T.J.J. Kelly, Adenovirus DNA replication in vitro: characterization of a protein covalently linked to nascent DNA strands, Proc. Natl. Acad. Sci. USA 77 Ž w22x B.M. Baroudy, S. Venkatesan, B. Moss, Incompletely basepaired flip-flop terminal loops link the two DNA strands of the vaccinia virus genome into one uninterrupted polynucleotide chain, Cell 28 Ž w23x A. Lezhava, T. Mizukami, T. Kajitani, D. Kameoka, M. Redenbach, H. Shinkawa, O. Nimi, H. Kinashi, Physical map of the linear chromosome of Streptomyces griseus, J. Bacteriol. 177 Ž w24x C.W. Chen, Complications and implications of linear bacterial chromosomes, Trends Genet. 12 Ž w25x S. Casjens, M. Murphy, M. DeLange, L. Sampson, R. van Vugt, W.M. Huang, Telomeres of the linear chromosomes of Lyme disease spirochetes: nucleotide sequence and possible exchange with linear plasmid telomeres, Mol. Microbiol. 26 Ž w26x J.E. Haber, P.C. Thorburn, D. Rogers, Meiotic and mitotic behavior of dicentric chromosomes in Saccharomyces cere- Õisiae, Genetics 106 Ž w27x S.A. Greenfeder, C.S. Newlon, A replication map of a 61-kb circular derivative of Saccharomyces cereõisiae chromosome III, Mol. Biol. Cell 3 Ž w28x O. Niwa, T. Toda, M. Yanagida, Triploid meiosis and aneuploidy in Schizosaccharomyces pombe: an unstable disomic for chromosome III, Curr. Genet. 9 Ž w29x J.B. Fan, M. Rochet, C. Gaillardin, C.L. Smith, Detection and characterization of a ring chromosome in the fission yeast Schizosaccharomyces pombe, Nucleic Acids Res. 20 Ž w30x J. Jenderny, A. Caliebe, C. Beyer, W. Grote, Transmission of a ring chromosome 18 from a mother with 46,XXr47,XX,q rž 18. mosaicism to her daughter, resulting in a 46,XX,rŽ 18. karyotype, J. Med. Genet. 30 Ž w31x D. Shore, Telomeres unsticky ends, Science 281 Ž w32x A.J. Lustig, T.D. Petes, Identification of yeast mutants with altered telomere structure, Proc. Natl. Acad. Sci. USA 83 Ž w33x D.M. Morrow, D.A. Tagle, Y. Shiloh, F.S. Collins, P. Hieter, TEL1, an S. cereõisiae homolog of the human gene mutated in ataxia telangiectasia, is functionally related to the yeast checkpoint gene MEC1, Cell 82 Ž w34x K. Savitsky, A. Bar-Shira, S. Gilad, G. Rotman, Y. Ziv, L.
9 F. Ishikawa, T. NaitorMutation Research Vanagaite, D.A. Tagle, S. Smith, T. Uziel, S. Sfez, M. Ashkenazi, I. Pecker, M. Frydman, R. Harnik, S.R. Patanjali, A. Simmons, G.A. Clines, A. Sartiel, R.A. Gatti, L. Chessa, O. Sanal, M.F. Lavin, A single ataxia telangiectasia gene with a product similar to PI-3 kinase, Science 268 Ž w35x J.A. Metcalfe, J. Parkhill, L. Campbell, M. Stacey, P. Biggs, P.J. Byrd, A.M. Taylor, Accelerated telomere shortening in ataxia telangiectasia, Nat. Genet. 13 Ž w36x S.J. Xia, M.A. Shammas, R.J. Shmookler Reis, Reduced telomere length in ataxia telangiectasia fibroblasts, Mutat. Res. 364 Ž w37x V.A. Zakian, ATM-related genes: what do they tell us about functions of the human gene?, Cell 82 Ž w38x N.J. Bentley, D.A. Holtzman, G. Flaggs, K.S. Keegan, A. DeMaggio, J.C. Ford, M. Hoekstra, A.M. Carr, The Schizosaccharomyces pombe rad3 checkpoint gene, EMBO J. 15 Ž w39x T. Naito, A. Matsuura, F. Ishikawa, Circular chromosome formation in a fission yeast mutant defective in two ATM homologues, Nat. Genet. 20 Ž w40x M. Dahlen, T. Olsson, G. Kanter-Smoler, A. Ramne, P. Sunnerhagen, Regulation of telomere length by checkpoint genes in Schizosaccharomyces pombe, Mol. Biol. Cell 9 Ž w41x S.A. Endow, D.J. Komma, K.C. Atwood, Ring chromosomes and rdna magnification in Drosophila, Genetics 108 Ž w42x S. Sutou, Reversal of DNA polarity as revealed by sister chromatid exchanges in ring chromosomes, Mutat. Res. 394 Ž w43x A. Lobner-Olesen, P.L. Kuempel, Chromosome partitioning in Escherichia coli, J. Bacteriol. 174 Ž w44x L.I. Rothfield, Bacterial chromosome segregation, Cell 77 Ž w45x E. Sonoda, M.S. Sasaki, J.M. Buerstedde, O. Bezzubova, A. Shinohara, H. Ogawa, M. Takata, Y. Yamaguchi-Iwai, S. Takeda, Rad51-deficient vertebrate cells accumulate chromosomal breaks prior to cell death, EMBO J. 17 Ž w46x T. de Lange, Ending up with the right partner, Nature 392 Ž w47x A.F. Dernburg, J.W. Sedat, W.Z. Cande, H.W. Bass, Cytology of telomeres, in: E.H. Blackburn, C.W. Greider Ž Eds.., Telomeres, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 1995, pp w48x C. Rabl, Uber zelltheilung, Morphol. Jahrb. 10 Ž w49x G. Eisen, The spermatogenesis of Batrachoseps, J. Morphol. 17 Ž w50x D. Zickler, N. Kleckner, The leptotene zygotene transition of meiosis, Annu. Rev. Genet. 32 Ž w51x C. Robinow, The number of chromosomes in Schizosaccharomyces pombe: light microscopy of stained preparations, Genetics 87 Ž w52x Y. Chikashige, D.Q. Ding, H. Funabiki, T. Haraguchi, S. Mashiko, M. Yanagida, Y. Hiraoka, Telomere-led premeiotic chromosome movement in fission yeast, Science 264 Ž w53x Y. Hiraoka, Meiotic telomeres: a matchmaker for homologous chromosomes, Genes Cells 3 Ž w54x D. Broccoli, A. Smogorzewska, L. Chong, T. de Lange, Human telomeres contain two distinct Myb-related proteins, TRF1 and TRF2, Nat. Genet. 17 Ž w55x A.J. Lustig, S. Kurtz, D. Shore, Involvement of the silencer and UAS binding protein RAP1 in regulation of telomere length, Science 250 Ž w56x J.P. Cooper, E.R. Nimmo, R.C. Allshire, T.R. Cech, Regulation of telomere length and function by a Myb-domain protein in fission yeast, Nature 385 Ž w57x E.R. Nimmo, A.L. Pidoux, P.E. Perry, R.C. Allshire, Defective meiosis in telomere-silencing mutants of Schizosaccharomyces pombe, Nature 392 Ž w58x J.P. Cooper, Y. Watanabe, P. Nurse, Fission yeast Taz1 protein is required for meiotic telomere clustering and recombination, Nature 392 Ž w59x G.S. Roeder, Meiotic chromosomes: it takes two to tango, Genes Dev. 11 Ž w60x G.C. Williams, Sex and Evolution, Princeton University Press, Princeton, w61x G. Bell, The Masterpiece of Nature, University of California Press, Berkeley, w62x O.P. Judson, B.B. Normark, Ancient asexual scandals, TREE 11 Ž
Chapter 13: Meiosis and Sexual Life Cycles
 Name Period Chapter 13: Meiosis and Sexual Life Cycles Concept 13.1 Offspring acquire genes from parents by inheriting chromosomes 1. Let s begin with a review of several terms that you may already know.
Name Period Chapter 13: Meiosis and Sexual Life Cycles Concept 13.1 Offspring acquire genes from parents by inheriting chromosomes 1. Let s begin with a review of several terms that you may already know.
Lecture 7 Mitosis & Meiosis
 Lecture 7 Mitosis & Meiosis Cell Division Essential for body growth and tissue repair Interphase G 1 phase Primary cell growth phase S phase DNA replication G 2 phase Microtubule synthesis Mitosis Nuclear
Lecture 7 Mitosis & Meiosis Cell Division Essential for body growth and tissue repair Interphase G 1 phase Primary cell growth phase S phase DNA replication G 2 phase Microtubule synthesis Mitosis Nuclear
1. Why is mitosis alone insufficient for the life cycle of sexually reproducing eukaryotes?
 Chapter 13: Meiosis and Sexual Life Cycles 1. Why is mitosis alone insufficient for the life cycle of sexually reproducing eukaryotes? 2. Define: gamete zygote meiosis homologous chromosomes diploid haploid
Chapter 13: Meiosis and Sexual Life Cycles 1. Why is mitosis alone insufficient for the life cycle of sexually reproducing eukaryotes? 2. Define: gamete zygote meiosis homologous chromosomes diploid haploid
List, describe, diagram, and identify the stages of meiosis.
 Meiosis and Sexual Life Cycles In this topic we will examine a second type of cell division used by eukaryotic cells: meiosis. In addition, we will see how the 2 types of eukaryotic cell division, mitosis
Meiosis and Sexual Life Cycles In this topic we will examine a second type of cell division used by eukaryotic cells: meiosis. In addition, we will see how the 2 types of eukaryotic cell division, mitosis
Cell Growth and Reproduction Module B, Anchor 1
 Cell Growth and Reproduction Module B, Anchor 1 Key Concepts: - The larger a cell becomes, the more demands the cell places on its DNA. In addition, a larger cell is less efficient in moving nutrients
Cell Growth and Reproduction Module B, Anchor 1 Key Concepts: - The larger a cell becomes, the more demands the cell places on its DNA. In addition, a larger cell is less efficient in moving nutrients
Sample Questions for Exam 3
 Sample Questions for Exam 3 1. All of the following occur during prometaphase of mitosis in animal cells except a. the centrioles move toward opposite poles. b. the nucleolus can no longer be seen. c.
Sample Questions for Exam 3 1. All of the following occur during prometaphase of mitosis in animal cells except a. the centrioles move toward opposite poles. b. the nucleolus can no longer be seen. c.
Chapter 13: Meiosis and Sexual Life Cycles
 Name Period Concept 13.1 Offspring acquire genes from parents by inheriting chromosomes 1. Let s begin with a review of several terms that you may already know. Define: gene locus gamete male gamete female
Name Period Concept 13.1 Offspring acquire genes from parents by inheriting chromosomes 1. Let s begin with a review of several terms that you may already know. Define: gene locus gamete male gamete female
CHROMOSOME STRUCTURE CHROMOSOME NUMBERS
 CHROMOSOME STRUCTURE 1. During nuclear division, the DNA (as chromatin) in a Eukaryotic cell's nucleus is coiled into very tight compact structures called chromosomes. These are rod-shaped structures made
CHROMOSOME STRUCTURE 1. During nuclear division, the DNA (as chromatin) in a Eukaryotic cell's nucleus is coiled into very tight compact structures called chromosomes. These are rod-shaped structures made
The cell cycle, mitosis and meiosis
 The cell cycle, mitosis and meiosis Learning objective This learning material is about the life cycle of a cell and the series of stages by which genetic materials are duplicated and partitioned to produce
The cell cycle, mitosis and meiosis Learning objective This learning material is about the life cycle of a cell and the series of stages by which genetic materials are duplicated and partitioned to produce
LAB 8 EUKARYOTIC CELL DIVISION: MITOSIS AND MEIOSIS
 LAB 8 EUKARYOTIC CELL DIVISION: MITOSIS AND MEIOSIS Los Angeles Mission College Biology 3 Name: Date: INTRODUCTION BINARY FISSION: Prokaryotic cells (bacteria) reproduce asexually by binary fission. Bacterial
LAB 8 EUKARYOTIC CELL DIVISION: MITOSIS AND MEIOSIS Los Angeles Mission College Biology 3 Name: Date: INTRODUCTION BINARY FISSION: Prokaryotic cells (bacteria) reproduce asexually by binary fission. Bacterial
1. When new cells are formed through the process of mitosis, the number of chromosomes in the new cells
 Cell Growth and Reproduction 1. When new cells are formed through the process of mitosis, the number of chromosomes in the new cells A. is half of that of the parent cell. B. remains the same as in the
Cell Growth and Reproduction 1. When new cells are formed through the process of mitosis, the number of chromosomes in the new cells A. is half of that of the parent cell. B. remains the same as in the
Workshop: Cellular Reproduction via Mitosis & Meiosis
 Workshop: Cellular Reproduction via Mitosis & Meiosis Introduction In this workshop you will examine how cells divide, including how they partition their genetic material (DNA) between the two resulting
Workshop: Cellular Reproduction via Mitosis & Meiosis Introduction In this workshop you will examine how cells divide, including how they partition their genetic material (DNA) between the two resulting
Bio EOC Topics for Cell Reproduction: Bio EOC Questions for Cell Reproduction:
 Bio EOC Topics for Cell Reproduction: Asexual vs. sexual reproduction Mitosis steps, diagrams, purpose o Interphase, Prophase, Metaphase, Anaphase, Telophase, Cytokinesis Meiosis steps, diagrams, purpose
Bio EOC Topics for Cell Reproduction: Asexual vs. sexual reproduction Mitosis steps, diagrams, purpose o Interphase, Prophase, Metaphase, Anaphase, Telophase, Cytokinesis Meiosis steps, diagrams, purpose
Milestones of bacterial genetic research:
 Milestones of bacterial genetic research: 1944 Avery's pneumococcal transformation experiment shows that DNA is the hereditary material 1946 Lederberg & Tatum describes bacterial conjugation using biochemical
Milestones of bacterial genetic research: 1944 Avery's pneumococcal transformation experiment shows that DNA is the hereditary material 1946 Lederberg & Tatum describes bacterial conjugation using biochemical
Chapter 12: The Cell Cycle
 Name Period Chapter 12: The Cell Cycle Overview: 1. What are the three key roles of cell division? State each role, and give an example. Key Role Example 2. What is meant by the cell cycle? Concept 12.1
Name Period Chapter 12: The Cell Cycle Overview: 1. What are the three key roles of cell division? State each role, and give an example. Key Role Example 2. What is meant by the cell cycle? Concept 12.1
Lecture 2: Mitosis and meiosis
 Lecture 2: Mitosis and meiosis 1. Chromosomes 2. Diploid life cycle 3. Cell cycle 4. Mitosis 5. Meiosis 6. Parallel behavior of genes and chromosomes Basic morphology of chromosomes telomere short arm
Lecture 2: Mitosis and meiosis 1. Chromosomes 2. Diploid life cycle 3. Cell cycle 4. Mitosis 5. Meiosis 6. Parallel behavior of genes and chromosomes Basic morphology of chromosomes telomere short arm
1 Mutation and Genetic Change
 CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
Sexual Reproduction and Meiosis
 12 Sexual Reproduction and Meiosis Concept Outline 12.1 Meiosis produces haploid cells from diploid cells. Discovery of Reduction Division. Sexual reproduction does not increase chromosome number because
12 Sexual Reproduction and Meiosis Concept Outline 12.1 Meiosis produces haploid cells from diploid cells. Discovery of Reduction Division. Sexual reproduction does not increase chromosome number because
Chapter 6 DNA Replication
 Chapter 6 DNA Replication Each strand of the DNA double helix contains a sequence of nucleotides that is exactly complementary to the nucleotide sequence of its partner strand. Each strand can therefore
Chapter 6 DNA Replication Each strand of the DNA double helix contains a sequence of nucleotides that is exactly complementary to the nucleotide sequence of its partner strand. Each strand can therefore
www.njctl.org PSI Biology Mitosis & Meiosis
 Mitosis and Meiosis Mitosis Classwork 1. Identify two differences between meiosis and mitosis. 2. Provide an example of a type of cell in the human body that would undergo mitosis. 3. Does cell division
Mitosis and Meiosis Mitosis Classwork 1. Identify two differences between meiosis and mitosis. 2. Provide an example of a type of cell in the human body that would undergo mitosis. 3. Does cell division
Chapter 12: The Cell Cycle
 Name Period Chapter 12: The Cell Cycle Overview: 1. What are the three key roles of cell division? State each role, and give an example. Key Role Reproduction Growth and development Tissue removal Example
Name Period Chapter 12: The Cell Cycle Overview: 1. What are the three key roles of cell division? State each role, and give an example. Key Role Reproduction Growth and development Tissue removal Example
Chapter 3. Cell Division. Laboratory Activities Activity 3.1: Mock Mitosis Activity 3.2: Mitosis in Onion Cells Activity 3.
 Chapter 3 Cell Division Laboratory Activities Activity 3.1: Mock Mitosis Activity 3.2: Mitosis in Onion Cells Activity 3.3: Mock Meiosis Goals Following this exercise students should be able to Recognize
Chapter 3 Cell Division Laboratory Activities Activity 3.1: Mock Mitosis Activity 3.2: Mitosis in Onion Cells Activity 3.3: Mock Meiosis Goals Following this exercise students should be able to Recognize
4.2 Meiosis. Meiosis is a reduction division. Assessment statements. The process of meiosis
 4.2 Meiosis Assessment statements State that meiosis is a reduction division of a diploid nucleus to form haploid nuclei. Define homologous chromosomes. Outline the process of meiosis, including pairing
4.2 Meiosis Assessment statements State that meiosis is a reduction division of a diploid nucleus to form haploid nuclei. Define homologous chromosomes. Outline the process of meiosis, including pairing
Cell Division CELL DIVISION. Mitosis. Designation of Number of Chromosomes. Homologous Chromosomes. Meiosis
 Cell Division CELL DIVISION Anatomy and Physiology Text and Laboratory Workbook, Stephen G. Davenport, Copyright 2006, All Rights Reserved, no part of this publication can be used for any commercial purpose.
Cell Division CELL DIVISION Anatomy and Physiology Text and Laboratory Workbook, Stephen G. Davenport, Copyright 2006, All Rights Reserved, no part of this publication can be used for any commercial purpose.
5. The cells of a multicellular organism, other than gametes and the germ cells from which it develops, are known as
 1. True or false? The chi square statistical test is used to determine how well the observed genetic data agree with the expectations derived from a hypothesis. True 2. True or false? Chromosomes in prokaryotic
1. True or false? The chi square statistical test is used to determine how well the observed genetic data agree with the expectations derived from a hypothesis. True 2. True or false? Chromosomes in prokaryotic
Practice Problems 4. (a) 19. (b) 36. (c) 17
 Chapter 10 Practice Problems Practice Problems 4 1. The diploid chromosome number in a variety of chrysanthemum is 18. What would you call varieties with the following chromosome numbers? (a) 19 (b) 36
Chapter 10 Practice Problems Practice Problems 4 1. The diploid chromosome number in a variety of chrysanthemum is 18. What would you call varieties with the following chromosome numbers? (a) 19 (b) 36
Appendix C DNA Replication & Mitosis
 K.Muma Bio 6 Appendix C DNA Replication & Mitosis Study Objectives: Appendix C: DNA replication and Mitosis 1. Describe the structure of DNA and where it is found. 2. Explain complimentary base pairing:
K.Muma Bio 6 Appendix C DNA Replication & Mitosis Study Objectives: Appendix C: DNA replication and Mitosis 1. Describe the structure of DNA and where it is found. 2. Explain complimentary base pairing:
Forensic DNA Testing Terminology
 Forensic DNA Testing Terminology ABI 310 Genetic Analyzer a capillary electrophoresis instrument used by forensic DNA laboratories to separate short tandem repeat (STR) loci on the basis of their size.
Forensic DNA Testing Terminology ABI 310 Genetic Analyzer a capillary electrophoresis instrument used by forensic DNA laboratories to separate short tandem repeat (STR) loci on the basis of their size.
Biology Behind the Crime Scene Week 4: Lab #4 Genetics Exercise (Meiosis) and RFLP Analysis of DNA
 Page 1 of 5 Biology Behind the Crime Scene Week 4: Lab #4 Genetics Exercise (Meiosis) and RFLP Analysis of DNA Genetics Exercise: Understanding how meiosis affects genetic inheritance and DNA patterns
Page 1 of 5 Biology Behind the Crime Scene Week 4: Lab #4 Genetics Exercise (Meiosis) and RFLP Analysis of DNA Genetics Exercise: Understanding how meiosis affects genetic inheritance and DNA patterns
Cellular Reproduction
 9 Cellular Reproduction section 1 Cellular Growth Before You Read Think about the life cycle of a human. On the lines below, write some of the stages that occur in the life cycle of a human. In this section,
9 Cellular Reproduction section 1 Cellular Growth Before You Read Think about the life cycle of a human. On the lines below, write some of the stages that occur in the life cycle of a human. In this section,
The Somatic Cell Cycle
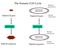 The Somatic Cell Cycle Maternal chromosome Diploid Zygote Diploid Zygote Paternal chromosome MITOSIS MITOSIS Maternal chromosome Diploid organism Diploid organism Paternal chromosome Int terpha ase The
The Somatic Cell Cycle Maternal chromosome Diploid Zygote Diploid Zygote Paternal chromosome MITOSIS MITOSIS Maternal chromosome Diploid organism Diploid organism Paternal chromosome Int terpha ase The
If and when cancer cells stop dividing, they do so at random points, not at the normal checkpoints in the cell cycle.
 Cancer cells have escaped from cell cycle controls Cancer cells divide excessively and invade other tissues because they are free of the body s control mechanisms. Cancer cells do not stop dividing when
Cancer cells have escaped from cell cycle controls Cancer cells divide excessively and invade other tissues because they are free of the body s control mechanisms. Cancer cells do not stop dividing when
Genetics Test Biology I
 Genetics Test Biology I Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Avery s experiments showed that bacteria are transformed by a. RNA. c. proteins.
Genetics Test Biology I Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Avery s experiments showed that bacteria are transformed by a. RNA. c. proteins.
Chromosomes, Karyotyping, and Abnormalities (Learning Objectives) Learn the components and parts of a metaphase chromosome.
 Chromosomes, Karyotyping, and Abnormalities (Learning Objectives) Learn the components and parts of a metaphase chromosome. Define the terms karyotype, autosomal and sex chromosomes. Explain how many of
Chromosomes, Karyotyping, and Abnormalities (Learning Objectives) Learn the components and parts of a metaphase chromosome. Define the terms karyotype, autosomal and sex chromosomes. Explain how many of
CHAPTER 10 CELL CYCLE AND CELL DIVISION
 CHAPTER 10 CELL CYCLE AND CELL DIVISION Cell division is an inherent property of living organisms. It is a process in which cells reproduce their own kind. The growth, differentiation, reproduction and
CHAPTER 10 CELL CYCLE AND CELL DIVISION Cell division is an inherent property of living organisms. It is a process in which cells reproduce their own kind. The growth, differentiation, reproduction and
AS Biology Unit 2 Key Terms and Definitions. Make sure you use these terms when answering exam questions!
 AS Biology Unit 2 Key Terms and Definitions Make sure you use these terms when answering exam questions! Chapter 7 Variation 7.1 Random Sampling Sampling a population to eliminate bias e.g. grid square
AS Biology Unit 2 Key Terms and Definitions Make sure you use these terms when answering exam questions! Chapter 7 Variation 7.1 Random Sampling Sampling a population to eliminate bias e.g. grid square
Sexual Reproduction. The specialized cells that are required for sexual reproduction are known as. And come from the process of: GAMETES
 Sexual Reproduction Sexual Reproduction We know all about asexual reproduction 1. Only one parent required. 2. Offspring are identical to parents. 3. The cells that produce the offspring are not usually
Sexual Reproduction Sexual Reproduction We know all about asexual reproduction 1. Only one parent required. 2. Offspring are identical to parents. 3. The cells that produce the offspring are not usually
Gene Mapping Techniques
 Gene Mapping Techniques OBJECTIVES By the end of this session the student should be able to: Define genetic linkage and recombinant frequency State how genetic distance may be estimated State how restriction
Gene Mapping Techniques OBJECTIVES By the end of this session the student should be able to: Define genetic linkage and recombinant frequency State how genetic distance may be estimated State how restriction
BioSci 2200 General Genetics Problem Set 1 Answer Key Introduction and Mitosis/ Meiosis
 BioSci 2200 General Genetics Problem Set 1 Answer Key Introduction and Mitosis/ Meiosis Introduction - Fields of Genetics To answer the following question, review the three traditional subdivisions of
BioSci 2200 General Genetics Problem Set 1 Answer Key Introduction and Mitosis/ Meiosis Introduction - Fields of Genetics To answer the following question, review the three traditional subdivisions of
4. DNA replication Pages: 979-984 Difficulty: 2 Ans: C Which one of the following statements about enzymes that interact with DNA is true?
 Chapter 25 DNA Metabolism Multiple Choice Questions 1. DNA replication Page: 977 Difficulty: 2 Ans: C The Meselson-Stahl experiment established that: A) DNA polymerase has a crucial role in DNA synthesis.
Chapter 25 DNA Metabolism Multiple Choice Questions 1. DNA replication Page: 977 Difficulty: 2 Ans: C The Meselson-Stahl experiment established that: A) DNA polymerase has a crucial role in DNA synthesis.
Name Date Period. 2. When a molecule of double-stranded DNA undergoes replication, it results in
 DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
Cell Division Mitosis and the Cell Cycle
 Cell Division Mitosis and the Cell Cycle A Chromosome and Sister Chromatids Key Points About Chromosome Structure A chromosome consists of DNA that is wrapped around proteins (histones) and condensed Each
Cell Division Mitosis and the Cell Cycle A Chromosome and Sister Chromatids Key Points About Chromosome Structure A chromosome consists of DNA that is wrapped around proteins (histones) and condensed Each
Genetics Module B, Anchor 3
 Genetics Module B, Anchor 3 Key Concepts: - An individual s characteristics are determines by factors that are passed from one parental generation to the next. - During gamete formation, the alleles for
Genetics Module B, Anchor 3 Key Concepts: - An individual s characteristics are determines by factors that are passed from one parental generation to the next. - During gamete formation, the alleles for
Lecture 11 The Cell Cycle and Mitosis
 Lecture 11 The Cell Cycle and Mitosis In this lecture Cell division Chromosomes The cell cycle Mitosis PPMAT Apoptosis What is cell division? Cells divide in order to reproduce themselves The cell cycle
Lecture 11 The Cell Cycle and Mitosis In this lecture Cell division Chromosomes The cell cycle Mitosis PPMAT Apoptosis What is cell division? Cells divide in order to reproduce themselves The cell cycle
Biology 1406 Exam 4 Notes Cell Division and Genetics Ch. 8, 9
 Biology 1406 Exam 4 Notes Cell Division and Genetics Ch. 8, 9 Ch. 8 Cell Division Cells divide to produce new cells must pass genetic information to new cells - What process of DNA allows this? Two types
Biology 1406 Exam 4 Notes Cell Division and Genetics Ch. 8, 9 Ch. 8 Cell Division Cells divide to produce new cells must pass genetic information to new cells - What process of DNA allows this? Two types
CCR Biology - Chapter 5 Practice Test - Summer 2012
 Name: Class: Date: CCR Biology - Chapter 5 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. If a cell cannot move enough material
Name: Class: Date: CCR Biology - Chapter 5 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. If a cell cannot move enough material
FINDING RELATION BETWEEN AGING AND
 FINDING RELATION BETWEEN AGING AND TELOMERE BY APRIORI AND DECISION TREE Jieun Sung 1, Youngshin Joo, and Taeseon Yoon 1 Department of National Science, Hankuk Academy of Foreign Studies, Yong-In, Republic
FINDING RELATION BETWEEN AGING AND TELOMERE BY APRIORI AND DECISION TREE Jieun Sung 1, Youngshin Joo, and Taeseon Yoon 1 Department of National Science, Hankuk Academy of Foreign Studies, Yong-In, Republic
CCR Biology - Chapter 9 Practice Test - Summer 2012
 Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
CELL DIVISION. STAGES OF MITOTIC DIVISION (Diag. C1)
 1 CELL DIVISION Cell division is the process by which cells replicate in order to replace cell loss, repair tissue damage and reproduce the organism. Two types of cell division are encountered in the Eukaryotic
1 CELL DIVISION Cell division is the process by which cells replicate in order to replace cell loss, repair tissue damage and reproduce the organism. Two types of cell division are encountered in the Eukaryotic
CHAPTER 9 CELLULAR REPRODUCTION P. 243-257
 CHAPTER 9 CELLULAR REPRODUCTION P. 243-257 SECTION 9-1 CELLULAR GROWTH Page 244 ESSENTIAL QUESTION Why is it beneficial for cells to remain small? MAIN IDEA Cells grow until they reach their size limit,
CHAPTER 9 CELLULAR REPRODUCTION P. 243-257 SECTION 9-1 CELLULAR GROWTH Page 244 ESSENTIAL QUESTION Why is it beneficial for cells to remain small? MAIN IDEA Cells grow until they reach their size limit,
by the PCR-mediated method (Krawchuk and Wahls, 1999). The construction of Ams2-null and conditional ams2-shut-off strains was previously described
 Table S1. Fission yeast strains used in this study. Gene disruption was performed by the PCR-mediated method (Krawchuk and Wahls, 1999). The construction of Ams2-null and conditional ams2-shut-off strains
Table S1. Fission yeast strains used in this study. Gene disruption was performed by the PCR-mediated method (Krawchuk and Wahls, 1999). The construction of Ams2-null and conditional ams2-shut-off strains
From DNA to Protein
 Nucleus Control center of the cell contains the genetic library encoded in the sequences of nucleotides in molecules of DNA code for the amino acid sequences of all proteins determines which specific proteins
Nucleus Control center of the cell contains the genetic library encoded in the sequences of nucleotides in molecules of DNA code for the amino acid sequences of all proteins determines which specific proteins
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA
 CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
Guided Notes: Chapter 9 Cellular Reproduction
 Guided Notes: Cellular Reproduction When do cells divide? Cells grow and function normally until they become too. Cell size is because increases faster than This means that there is not enough area on
Guided Notes: Cellular Reproduction When do cells divide? Cells grow and function normally until they become too. Cell size is because increases faster than This means that there is not enough area on
The Huntington Library, Art Collections, and Botanical Gardens
 The Huntington Library, Art Collections, and Botanical Gardens Rooting for Mitosis Overview Students will fix, stain, and make slides of onion root tips. These slides will be examined for the presence
The Huntington Library, Art Collections, and Botanical Gardens Rooting for Mitosis Overview Students will fix, stain, and make slides of onion root tips. These slides will be examined for the presence
Gene mutation and molecular medicine Chapter 15
 Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
Structure and Function of DNA
 Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Genetics 301 Sample Final Examination Spring 2003
 Genetics 301 Sample Final Examination Spring 2003 50 Multiple Choice Questions-(Choose the best answer) 1. A cross between two true breeding lines one with dark blue flowers and one with bright white flowers
Genetics 301 Sample Final Examination Spring 2003 50 Multiple Choice Questions-(Choose the best answer) 1. A cross between two true breeding lines one with dark blue flowers and one with bright white flowers
MCAS Biology. Review Packet
 MCAS Biology Review Packet 1 Name Class Date 1. Define organic. THE CHEMISTRY OF LIFE 2. All living things are made up of 6 essential elements: SPONCH. Name the six elements of life. S N P C O H 3. Elements
MCAS Biology Review Packet 1 Name Class Date 1. Define organic. THE CHEMISTRY OF LIFE 2. All living things are made up of 6 essential elements: SPONCH. Name the six elements of life. S N P C O H 3. Elements
MITOSIS IN ONION ROOT TIP CELLS: AN INTRODUCTION TO LIGHT MICROSCOPY
 MITOSIS IN ONION ROOT TIP CELLS: AN INTRODUCTION TO LIGHT MICROSCOPY Adapted from Foundations of Biology I; Lab 6 Introduction to Microscopy Dr. John Robertson, Westminster College Biology Department,
MITOSIS IN ONION ROOT TIP CELLS: AN INTRODUCTION TO LIGHT MICROSCOPY Adapted from Foundations of Biology I; Lab 6 Introduction to Microscopy Dr. John Robertson, Westminster College Biology Department,
Biology Final Exam Study Guide: Semester 2
 Biology Final Exam Study Guide: Semester 2 Questions 1. Scientific method: What does each of these entail? Investigation and Experimentation Problem Hypothesis Methods Results/Data Discussion/Conclusion
Biology Final Exam Study Guide: Semester 2 Questions 1. Scientific method: What does each of these entail? Investigation and Experimentation Problem Hypothesis Methods Results/Data Discussion/Conclusion
Genetics Lecture Notes 7.03 2005. Lectures 1 2
 Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Classify chromosomes in a karyotype according to size and centromere position. Identify metacentric, submetacentric and acrocentric chromosomes
 Mitosis, Meiosis and the Cell Cycle Prof. Alfred Cuschieri University of Malta Department of Anatomy Objectives By the end of the session the student shoud be able to: Define the meaning of chromosomes
Mitosis, Meiosis and the Cell Cycle Prof. Alfred Cuschieri University of Malta Department of Anatomy Objectives By the end of the session the student shoud be able to: Define the meaning of chromosomes
Given these characteristics of life, which of the following objects is considered a living organism? W. X. Y. Z.
 Cell Structure and Organization 1. All living things must possess certain characteristics. They are all composed of one or more cells. They can grow, reproduce, and pass their genes on to their offspring.
Cell Structure and Organization 1. All living things must possess certain characteristics. They are all composed of one or more cells. They can grow, reproduce, and pass their genes on to their offspring.
I. Genes found on the same chromosome = linked genes
 Genetic recombination in Eukaryotes: crossing over, part 1 I. Genes found on the same chromosome = linked genes II. III. Linkage and crossing over Crossing over & chromosome mapping I. Genes found on the
Genetic recombination in Eukaryotes: crossing over, part 1 I. Genes found on the same chromosome = linked genes II. III. Linkage and crossing over Crossing over & chromosome mapping I. Genes found on the
The Human Genome Project
 The Human Genome Project Brief History of the Human Genome Project Physical Chromosome Maps Genetic (or Linkage) Maps DNA Markers Sequencing and Annotating Genomic DNA What Have We learned from the HGP?
The Human Genome Project Brief History of the Human Genome Project Physical Chromosome Maps Genetic (or Linkage) Maps DNA Markers Sequencing and Annotating Genomic DNA What Have We learned from the HGP?
Germ cell formation / gametogenesis And Fertilisation
 Developmental Biology BY1101 P. Murphy Lecture 3 The first steps to forming a new organism Descriptive embryology I Germ cell formation / gametogenesis And Fertilisation Why bother with sex? In terms of
Developmental Biology BY1101 P. Murphy Lecture 3 The first steps to forming a new organism Descriptive embryology I Germ cell formation / gametogenesis And Fertilisation Why bother with sex? In terms of
Mitosis, Meiosis and Fertilization 1
 Mitosis, Meiosis and Fertilization 1 I. Introduction When you fall and scrape the skin off your hands or knees, how does your body make new skin cells to replace the skin cells that were scraped off? How
Mitosis, Meiosis and Fertilization 1 I. Introduction When you fall and scrape the skin off your hands or knees, how does your body make new skin cells to replace the skin cells that were scraped off? How
The correct answer is c A. Answer a is incorrect. The white-eye gene must be recessive since heterozygous females have red eyes.
 1. Why is the white-eye phenotype always observed in males carrying the white-eye allele? a. Because the trait is dominant b. Because the trait is recessive c. Because the allele is located on the X chromosome
1. Why is the white-eye phenotype always observed in males carrying the white-eye allele? a. Because the trait is dominant b. Because the trait is recessive c. Because the allele is located on the X chromosome
Biology 3A Laboratory MITOSIS Asexual Reproduction
 Biology 3A Laboratory MITOSIS Asexual Reproduction OBJECTIVE To study the cell cycle and understand how, when and why cells divide. To study and identify the major stages of cell division. To relate the
Biology 3A Laboratory MITOSIS Asexual Reproduction OBJECTIVE To study the cell cycle and understand how, when and why cells divide. To study and identify the major stages of cell division. To relate the
1.1 Introduction. 1.2 Cells CHAPTER. 1.2.1 Prokaryotic Cells. 1.2.2 Eukaryotic Cells
 C HAPTER 1CELLS AND CELL DIVISION CHAPTER 1.1 Introduction In genetics, we view cells as vessels for the genetic material. Our main interest is in the chromosomes and their environment. This being said,
C HAPTER 1CELLS AND CELL DIVISION CHAPTER 1.1 Introduction In genetics, we view cells as vessels for the genetic material. Our main interest is in the chromosomes and their environment. This being said,
Meiosis is a special form of cell division.
 Page 1 of 6 KEY CONCEPT Meiosis is a special form of cell division. BEFORE, you learned Mitosis produces two genetically identical cells In sexual reproduction, offspring inherit traits from both parents
Page 1 of 6 KEY CONCEPT Meiosis is a special form of cell division. BEFORE, you learned Mitosis produces two genetically identical cells In sexual reproduction, offspring inherit traits from both parents
Mitochondrial DNA Analysis
 Mitochondrial DNA Analysis Lineage Markers Lineage markers are passed down from generation to generation without changing Except for rare mutation events They can help determine the lineage (family tree)
Mitochondrial DNA Analysis Lineage Markers Lineage markers are passed down from generation to generation without changing Except for rare mutation events They can help determine the lineage (family tree)
Test Two Study Guide
 Test Two Study Guide 1. Describe what is happening inside a cell during the following phases (pictures may help but try to use words): Interphase: : Consists of G1 / S / G2. Growing stage, cell doubles
Test Two Study Guide 1. Describe what is happening inside a cell during the following phases (pictures may help but try to use words): Interphase: : Consists of G1 / S / G2. Growing stage, cell doubles
Basic Concepts Recombinant DNA Use with Chapter 13, Section 13.2
 Name Date lass Master 19 Basic oncepts Recombinant DN Use with hapter, Section.2 Formation of Recombinant DN ut leavage Splicing opyright lencoe/mcraw-hill, a division of he Mcraw-Hill ompanies, Inc. Bacterial
Name Date lass Master 19 Basic oncepts Recombinant DN Use with hapter, Section.2 Formation of Recombinant DN ut leavage Splicing opyright lencoe/mcraw-hill, a division of he Mcraw-Hill ompanies, Inc. Bacterial
GENE REGULATION. Teacher Packet
 AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
Genetic Mutations. Indicator 4.8: Compare the consequences of mutations in body cells with those in gametes.
 Genetic Mutations Indicator 4.8: Compare the consequences of mutations in body cells with those in gametes. Agenda Warm UP: What is a mutation? Body cell? Gamete? Notes on Mutations Karyotype Web Activity
Genetic Mutations Indicator 4.8: Compare the consequences of mutations in body cells with those in gametes. Agenda Warm UP: What is a mutation? Body cell? Gamete? Notes on Mutations Karyotype Web Activity
somatic cell egg genotype gamete polar body phenotype homologous chromosome trait dominant autosome genetics recessive
 CHAPTER 6 MEIOSIS AND MENDEL Vocabulary Practice somatic cell egg genotype gamete polar body phenotype homologous chromosome trait dominant autosome genetics recessive CHAPTER 6 Meiosis and Mendel sex
CHAPTER 6 MEIOSIS AND MENDEL Vocabulary Practice somatic cell egg genotype gamete polar body phenotype homologous chromosome trait dominant autosome genetics recessive CHAPTER 6 Meiosis and Mendel sex
Chapter 8: Variation in Chromosome Structure and Number
 Chapter 8: Variation in Chromosome Structure and Number Student Learning Objectives Upon completion of this chapter you should be able to: 1. Know the principles and terminology associated with variations
Chapter 8: Variation in Chromosome Structure and Number Student Learning Objectives Upon completion of this chapter you should be able to: 1. Know the principles and terminology associated with variations
BioBoot Camp Genetics
 BioBoot Camp Genetics BIO.B.1.2.1 Describe how the process of DNA replication results in the transmission and/or conservation of genetic information DNA Replication is the process of DNA being copied before
BioBoot Camp Genetics BIO.B.1.2.1 Describe how the process of DNA replication results in the transmission and/or conservation of genetic information DNA Replication is the process of DNA being copied before
Cell Division and Mitosis DNA. Sexual Reproduction and Meiosis. 2. Meiosis occurs in the reproductive organs, producing four haploid sex cells.
 ell Division and Mitosis 1. he life cycle of a cell has two parts growth and development, and cell division. 2. In mitosis, the nucleus divides to form two identical nuclei. Mitosis occurs in four continuous
ell Division and Mitosis 1. he life cycle of a cell has two parts growth and development, and cell division. 2. In mitosis, the nucleus divides to form two identical nuclei. Mitosis occurs in four continuous
Answer: 2. Uracil. Answer: 2. hydrogen bonds. Adenine, Cytosine and Guanine are found in both RNA and DNA.
 Answer: 2. Uracil Adenine, Cytosine and Guanine are found in both RNA and DNA. Thymine is found only in DNA; Uracil takes its (Thymine) place in RNA molecules. Answer: 2. hydrogen bonds The complementary
Answer: 2. Uracil Adenine, Cytosine and Guanine are found in both RNA and DNA. Thymine is found only in DNA; Uracil takes its (Thymine) place in RNA molecules. Answer: 2. hydrogen bonds The complementary
How Cancer Begins???????? Chithra Manikandan Nov 2009
 Cancer Cancer is one of the most common diseases in the developed world: 1 in 4 deaths are due to cancer 1 in 17 deaths are due to lung cancer Lung cancer is the most common cancer in men Breast cancer
Cancer Cancer is one of the most common diseases in the developed world: 1 in 4 deaths are due to cancer 1 in 17 deaths are due to lung cancer Lung cancer is the most common cancer in men Breast cancer
12.1 The Role of DNA in Heredity
 12.1 The Role of DNA in Heredity Only in the last 50 years have scientists understood the role of DNA in heredity. That understanding began with the discovery of DNA s structure. In 1952, Rosalind Franklin
12.1 The Role of DNA in Heredity Only in the last 50 years have scientists understood the role of DNA in heredity. That understanding began with the discovery of DNA s structure. In 1952, Rosalind Franklin
Microscopes and the Metric System
 Microscopes and the Metric System BIO162 Fall 2007 Sizes of Microorganisms: -Viruses: 0.01 0.3 um -Bacteria: 1 3 um -Fungi: 3 30 um -Protozoa: 5 1000 um 1 Measuring Microorganisms Ocular Micrometer The
Microscopes and the Metric System BIO162 Fall 2007 Sizes of Microorganisms: -Viruses: 0.01 0.3 um -Bacteria: 1 3 um -Fungi: 3 30 um -Protozoa: 5 1000 um 1 Measuring Microorganisms Ocular Micrometer The
Trasposable elements: P elements
 Trasposable elements: P elements In 1938 Marcus Rhodes provided the first genetic description of an unstable mutation, an allele of a gene required for the production of pigment in maize. This instability
Trasposable elements: P elements In 1938 Marcus Rhodes provided the first genetic description of an unstable mutation, an allele of a gene required for the production of pigment in maize. This instability
Name: Class: Date: ID: A
 Name: Class: _ Date: _ Meiosis Quiz 1. (1 point) A kidney cell is an example of which type of cell? a. sex cell b. germ cell c. somatic cell d. haploid cell 2. (1 point) How many chromosomes are in a human
Name: Class: _ Date: _ Meiosis Quiz 1. (1 point) A kidney cell is an example of which type of cell? a. sex cell b. germ cell c. somatic cell d. haploid cell 2. (1 point) How many chromosomes are in a human
1.5 page 3 DNA Replication S. Preston 1
 AS Unit 1: Basic Biochemistry and Cell Organisation Name: Date: Topic 1.5 Nucleic Acids and their functions Page 3 l. DNA Replication 1. Go through PowerPoint 2. Read notes p2 and then watch the animation
AS Unit 1: Basic Biochemistry and Cell Organisation Name: Date: Topic 1.5 Nucleic Acids and their functions Page 3 l. DNA Replication 1. Go through PowerPoint 2. Read notes p2 and then watch the animation
Compiled and/or written by Amy B. Vento and David R. Gillum
 Fact Sheet Describing Recombinant DNA and Elements Utilizing Recombinant DNA Such as Plasmids and Viral Vectors, and the Application of Recombinant DNA Techniques in Molecular Biology Compiled and/or written
Fact Sheet Describing Recombinant DNA and Elements Utilizing Recombinant DNA Such as Plasmids and Viral Vectors, and the Application of Recombinant DNA Techniques in Molecular Biology Compiled and/or written
A and B are not absolutely linked. They could be far enough apart on the chromosome that they assort independently.
 Name Section 7.014 Problem Set 5 Please print out this problem set and record your answers on the printed copy. Answers to this problem set are to be turned in to the box outside 68-120 by 5:00pm on Friday
Name Section 7.014 Problem Set 5 Please print out this problem set and record your answers on the printed copy. Answers to this problem set are to be turned in to the box outside 68-120 by 5:00pm on Friday
Biological Sciences Initiative. Human Genome
 Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
DNA Replication & Protein Synthesis. This isn t a baaaaaaaddd chapter!!!
 DNA Replication & Protein Synthesis This isn t a baaaaaaaddd chapter!!! The Discovery of DNA s Structure Watson and Crick s discovery of DNA s structure was based on almost fifty years of research by other
DNA Replication & Protein Synthesis This isn t a baaaaaaaddd chapter!!! The Discovery of DNA s Structure Watson and Crick s discovery of DNA s structure was based on almost fifty years of research by other
CHAPTER 6 GRIFFITH/HERSHEY/CHASE: DNA IS THE GENETIC MATERIAL IDENTIFICATION OF DNA DNA AND HEREDITY DNA CAN GENETICALLY TRANSFORM CELLS
 CHAPTER 6 GRIFFITH/HERSHEY/CHASE: DNA IS THE GENETIC MATERIAL In 1928, Frederick Griffith was able to transform harmless bacteria into virulent pathogens with an extract that Oswald Avery proved, in 1944,
CHAPTER 6 GRIFFITH/HERSHEY/CHASE: DNA IS THE GENETIC MATERIAL In 1928, Frederick Griffith was able to transform harmless bacteria into virulent pathogens with an extract that Oswald Avery proved, in 1944,
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
The illustrations below reflect other scientists results in identifying and counting the stages of the onion root tip and the whitefish blastula.
 Abstract: The purpose of this laboratory experiment was to identify in what stage of mitosis viewed cells were in. The stages of mitosis include prophase, metaphase, anaphase and telophase. Although the
Abstract: The purpose of this laboratory experiment was to identify in what stage of mitosis viewed cells were in. The stages of mitosis include prophase, metaphase, anaphase and telophase. Although the
The following chapter is called "Preimplantation Genetic Diagnosis (PGD)".
 Slide 1 Welcome to chapter 9. The following chapter is called "Preimplantation Genetic Diagnosis (PGD)". The author is Dr. Maria Lalioti. Slide 2 The learning objectives of this chapter are: To learn the
Slide 1 Welcome to chapter 9. The following chapter is called "Preimplantation Genetic Diagnosis (PGD)". The author is Dr. Maria Lalioti. Slide 2 The learning objectives of this chapter are: To learn the
AP Biology Essential Knowledge Student Diagnostic
 AP Biology Essential Knowledge Student Diagnostic Background The Essential Knowledge statements provided in the AP Biology Curriculum Framework are scientific claims describing phenomenon occurring in
AP Biology Essential Knowledge Student Diagnostic Background The Essential Knowledge statements provided in the AP Biology Curriculum Framework are scientific claims describing phenomenon occurring in
DNA: Structure and Replication
 7 DNA: Structure and Replication WORKING WITH THE FIGURES 1. In Table 7-1, why are there no entries for the first four tissue sources? For the last three entries, what is the most likely explanation for
7 DNA: Structure and Replication WORKING WITH THE FIGURES 1. In Table 7-1, why are there no entries for the first four tissue sources? For the last three entries, what is the most likely explanation for
What is Cancer? Cancer is a genetic disease: Cancer typically involves a change in gene expression/function:
 Cancer is a genetic disease: Inherited cancer Sporadic cancer What is Cancer? Cancer typically involves a change in gene expression/function: Qualitative change Quantitative change Any cancer causing genetic
Cancer is a genetic disease: Inherited cancer Sporadic cancer What is Cancer? Cancer typically involves a change in gene expression/function: Qualitative change Quantitative change Any cancer causing genetic
