THE Tor kinases are key components of an evolutionarily
|
|
|
- Jerome Pitts
- 8 years ago
- Views:
Transcription
1 Copyright Ó 2007 by the Genetics Society of America DOI: /genetics Efficient Tor Signaling Requires a Functional Class C Vps Protein Complex in Saccharomyces cerevisiae Sara A. Zurita-Martinez,*,1 Rekha Puria,*,1 Xuewen Pan, Jef D. Boeke and Maria E. Cardenas*,2 *Department of Molecular Genetics and Microbiology, Duke University Medical Center, Durham, North Carolina 27710, Department of Biochemistry and Molecular Biology, Baylor College of Medicine, Houston, Texas and Department of Molecular Biology and Genetics, The Johns Hopkins University School of Medicine, Baltimore, Maryland Manuscript received March 2, 2007 Accepted for publication May 25, 2007 ABSTRACT The Tor kinases regulate responses to nutrients and control cell growth. Unlike most organisms that only contain one Tor protein, Saccharomyces cerevisiae expresses two, Tor1 and Tor2, which are thought to share all of the rapamycin-sensitive functions attributable to Tor signaling. Here we conducted a genetic screen that defined the global TOR1 synthetic fitness or lethal interaction gene network. This screen identified mutations in distinctive functional categories that impaired vacuolar function, including components of the EGO/Gse and PAS complexes that reduce fitness. In addition, tor1 is lethal in combination with mutations in class C Vps complex components. We find that Tor1 does not regulate the known function of the class C Vps complex in protein sorting. Instead class C vps mutants fail to recover from rapamycin-induced growth arrest or to survive nitrogen starvation and have low levels of amino acids. Remarkably, addition of glutamate or glutamine restores viability to a tor1 pep3 mutant strain. We conclude that Tor1 is more effective than Tor2 at providing rapamycin-sensitive Tor signaling under conditions of amino acid limitation, and that an intact class C Vps complex is required to mediate intracellular amino acid homeostasis for efficient Tor signaling. THE Tor kinases are key components of an evolutionarily conserved nutrient-responsive pathway that regulates cell growth and proliferation in eukaryotic organisms. The Tor kinases were first identified in yeast cells as the targets of the antiproliferative drug rapamycin (Heitman et al. 1991). Thereafter, rapamycin has been instrumental in elucidating biological events governed by Tor signaling, including complex transcriptional and translational programs (reviewed in Rohde et al. 2001; Crespo and Hall 2002). When yeast cells are grown in ample nutrient conditions, Tor activity promotes the expression of genes encoding trnas, ribosomal proteins, and rrna, while inactivating genes required for utilization of poor nitrogen and carbon sources, and stress responses (Zaragoza et al. 1998; Beck and Hall 1999; Cardenas et al. 1999; Hardwick et al. 1999; Powers and Walter 1999; Komeili et al. 2000). Tor activity also supports translation, in large part by suppressing the general amino acid control response regulated by the Gcn2 kinase, and possibly by also affecting the stability of eif4g (Berset et al. 1998; Valenzuela et al. 2001; Cherkasova and Hinnebusch 2003; Kubota et al. 2003; Rohde et al. 2004). Inhibition of Tor by rapamycin elicits many of the 1 These authors contributed equally to this work. 2 Corresponding author: Department of Molecular Genetics and Microbiology, Duke University Medical Center, 322 CARL Bldg., Box 3546, Research Dr., Durham, NC carde004@mc.duke.edu cellular responses that are triggered by starvation for nutrients, such as inhibition of protein synthesis, downregulation of amino acid permeases, protein degradation, autophagy, and cell cycle arrest (reviewed by Rohde et al. 2001; De Virgilio and Loewith 2006). Unlike most organisms that express only one Tor protein, Saccharomyces cerevisiae has two highly homologous Tor proteins, Tor1 and Tor2, which are thought to share all of the rapamycin-sensitive functions attributable to Tor signaling, while only Tor2 serves a unique and essential rapamycin-insensitive role (reviewed in Crespo and Hall 2002). A recent study has suggested that Tor1 and Tor2 differ in providing a rapamycinsensitive function in certain class C vps mutants (Xie et al. 2005); however, the exact nature of this function remains to be determined. The Tor proteins form two distinct multiprotein complexes: TORC1 and TORC2. Tor1 (and to a lesser extent Tor2) is a component of the TORC1 complex, which includes Lst8, Kog1, Tco89, and Bit61. TORC2 consists of Tor2, Lst8, and the Avo1, Avo2, and Avo3 proteins (Loewith et al. 2002; Wedaman et al. 2003; Reinke et al. 2004). FKBP12-rapamycin physically associates with TORC1 but not with TORC2, suggesting that this second complex mediates the rapamycin-insensitive Tor2 role in controlling polarization of the actin cytoskeleton (Loewith et al. 2002). To understand why yeast cells express two functional Tor proteins, we sought to define novel Tor1- or Tor2- specific functions. Here we performed a genomewide Genetics 176: (August 2007)
2 2140 Zurita-Martinez et al. screen searching for genes that when mutated in combination with tor1 reduce fitness or render cells inviable. These genes identified distinct functional networks including those involved in protein sorting, vacuolar inheritance, and microautophagy. In particular, we find that tor1 shows synthetic lethality or synthetic reduced fitness with mutations in different components of the class C Vps complex, which includes the Pep3, Pep5, Vps16, Vps33, Vps39, and Vps41 proteins (Banta et al. 1988; Raymond et al. 1992; Rieder and Emr 1997; Wurmser et al. 2000). The class C Vps complex plays a central role in protein sorting by regulating vesicle docking and fusion at the endosome and between the endosome and the vacuole (Srivastava et al. 2000; Peterson and Emr 2001). Viability of the tor1 pep3 double mutant was restored by expression of Tor1 but not Tor2, indicating that the function linking Tor1 and the class C Vps complex is unique to the rapamycin-sensitive TORC1 complex. Mutants lacking components of the class C Vps complex fail to recover from rapamycin-induced growth arrest and to survive nitrogen starvation, have low levels of amino acids, in particular glutamate, and show growth defects at 37 (this study; Kitamoto et al. 1988; Robinson et al. 1991). Remarkably, addition of glutamate or glutamine rescues the growth defect of class C single vps mutants and of a tor1 pep3 ts conditional mutant at 37. Our studies suggest that, in contrast to Tor2, Tor1 is specialized to support growth under conditions where intracellular amino acid concentrations are drastically reduced. We also conclude that an intact class C Vps complex is required to provide intracellular amino acid homeostasis for proper Tor1 signaling. These findings provide a physiological foundation to understand the duplication and divergence of Tor1 and Tor2 functions in S. cerevisiae. MATERIALS AND METHODS Yeast strains and media: Strains used in this study are listed in Table 1. All the strains are isogenic derivatives of BY4741 or BY4742 and unless otherwise indicated were constructed by the Saccharomyces Genome Deletion Project (distributed by Invitrogen, Carlsbad, CA). Strains SZY25 and SZY21 were obtained by deletion of TOR1 with URA3 in strains BY4741 and 13652, respectively. Strains SZY26, 27, 28, 29, 30, 31, 32, and 33 as well as RPY50 and RPY51 were created by crossing strain SZY25 to the pep3, pep5, vps15, vps16, vps34, vac7, vac8, vac17, vps39, and vps41 MATa haploid strains, respectively. Strain SZY36 was obtained from strain BY4742 by deletion of the VPS33 gene with KanMX4. Strain SZY37 was constructed by replacing the URA3 gene in strain SZY25 with LEU2. Strain SZY40 was obtained by crossing strain SZY37 to the pep3 haploid mutant strain #14105 and the resulting diploid transformed with plasmid pbj9113 expressing the pep3 ts -108 (Srivastava et al. 2000) was sporulated and dissected to obtain strain SZY40-4. SZY43 is a meiotic segregant of strain SZY32. Strain SZY49 was constructed by crossing strains SZY25 and SZY36. Yeast synthetic medium (YNB) with ammonium was always supplemented with 2% glucose (SG). SG medium was supplemented with amino acids to satisfy any auxotrophic requirements (SC) and for the experiment presented in Figure 6C with 0.2% of the indicated amino acid. Sporulation medium was 1.5% potassium acetate (KAc) (ph 7.5), supplemented with any required amino acids. SLAD, YPD, and all other media were prepared as described previously (Gimeno et al. 1992; Sherman 2002). Rapamycin was added to the media from concentrated stock solutions in 90% ethanol, 10% Tween-20. Yeast transformations were performed by the lithium acetate method (Schiestl and Gietz 1989). Unless noted otherwise, mutant yeast strains were constructed by PCR-mediated gene disruption, replacing the entire open reading frame of the targeted gene with the indicated genes (Longtine et al. 1998; Goldstein and McCusker 1999). All gene deletions were confirmed by PCR. Plasmids: Low-copy centromeric plasmids, prs315-tor1 expressing TOR1, and pml40-3 expressing TOR2, as well as plasmids expressing Tor2-Tor1 hybrid proteins were described previously (Lorenz and Heitman 1995; Alarcon et al. 1996). Plasmid psz12 (Hybrid 4) was created by gap repair; briefly, a 173-bp (from nucleotide 5316 to 5488 of TOR2) PCR product was generated with primers SZ CATAATTGG GCCT TAGCTAATTTTGAAGTAATATCCATGCTAACATCTGTCTC TAAAAAGAAACAGGAAG 39 and SZ GAAAAAAGC CCTTGATCGCTGGAACAACATGTCTTTGAATAAGATTAG AAGAGTAATGAACTTC 39 and plasmid pml40-3 as template. The PCR product was cotransformed with NcoI-digested prs315-tor1 into strain All hybrids were confirmed by sequencing. Plasmid pmep2-gfp was kindly provided by J. Rutherford and will be published elsewhere. Plasmids pbj9113 bearing the pep3ts-108 (Srivastava et al. 2000) and p188 containing the GCN4-lacZ reporter gene were provided by E. Jones and A. Hinnebusch, respectively. Western blotting and a-factor processing: Cell extracts from exponentially growing cultures in YEPD were prepared as previously described except that the breakage buffer consisted of 100 mm Tris-HCl, 50 mm KCl, 1 mm EDTA, 5% glycerol, and the protease inhibitors leupeptin, aprotinin, and pepstatin added at 1 mg/ml and 0.5 mm phenylmethyl sulfonyl fluoride (Rohde et al. 2004). Western blot analysis with 40 mg of protein for Ape1 and Alp1 and 10 mg for CpY was performed by standard techniques with antibodies specific for Ape1 and Alp1 (kindly provided by Y. Ohsumi) and CpY (Molecular Probes, Eugene, OR). The antisera recognizing amino acids and of Tor1 and Tor2, respectively, were previously described (Cardenas and Heitman 1995; Alarcon et al. 1996). For metabolic labeling cultures were grown to early exponential phase in SC medium. Cells were washed and resuspended in SC without methionine medium and treated with drug vehicle alone or 100 nm rapamycin and incubated for 20 min. Metabolic labeling of yeast cells with Trans 35 S-LABEL (ICN), pulse chase, cell extract preparation, immunoprecipitation with specific antibodies for a-factor (a generous gift of T. Graham), and CpY were performed at 30 according to published protocols (Graham 1998). Amino acid determination, Northern blot analysis, and b-galactosidase assays: Amino acid extraction from exponentially growing cells in YPD medium was performed as described (Chen and Kaiser 2002). Amino acid analyses were carried out in duplicate by anion exchange chromatography employing a Beckman 6300 Li citrate-based analyzer followed by post-column ninhydrin reaction detection system at the Molecular Structure Facility, University of California at Davis. Northern blot analysis as well as b-galactosidase assays to determine GCN4-lacZ reporter gene activity were previously described (Cardenas et al. 1999; Rohde et al. 2004). Fluorescent microscopy: Cells were collected, spotted onto microscope slides, and imaged in a Nikon Eclipse E400
3 Tor and Class C Vps Protein Complex 2141 TABLE 1 Yeast strains used in this study Strain Genotype Reference/source BY4741 MATa his3d1 leu2d0 met15d0 ura3d0 Research Genetics BY4742 MATahis3D1 leu2d0 lys2d0 ura3d0 Research Genetics #13652 BY4742 ssd1tkanmx4 Research Genetics #14105 BY4742 pep3tkanmx4 Research Genetics #10817 BY4742 pep5tkanmx4 Research Genetics #13236 BY4742 vps15tkanmx4 Research Genetics #12783 BY4742 vps16tkanmx4 Research Genetics #15149 BY4742 vps34tkanmx4 Research Genetics #13774 BY4742 vps39tkanmx4 Research Genetics #14015 BY4742 vps41tkanmx4 Research Genetics #13021 BY4742 vac7tkanmx4 Research Genetics #10253 BY4742 vac8tkanmx4 Research Genetics #13470 BY4742 vac17tkanmx4 Research Genetics #16864 BY4742 tor1tkanmx4 Research Genetics #16522 BY4742 gtr1tkanmx4 Research Genetics #13214 BY4742 ego3tkanmx4 Research Genetics SZY21 MATahis3D1 leu2d0 lys2d0 ura3d0 ssd1tkanmx4 tor1tura3 SZY25 MATa his3d1 leu2d0 met15d0 ura3d0 tor1tura3 SZY26 pep3tkanmx4 tor1tura3 SZY27 PEP5/pep5TKanMX4 tor1tura3/tor1 SZY28 VPS15/vps15TKanMX4 tor1tura3/tor1 SZY29 MATa/ahis3D1/his3D1 leu2d/leu2d0 LYS2/lys2D0 met15d0/met15 VPS16/vps16TKanMX4 tor1tura3/tor1 SZY30 VPS34/vps34TKanMX4 tor1tura3/tor1 SZY31 VAC7/vac7TKanMX4 tor1tura3/tor1 SZY32 VAC8/vac8TKanMX4 tor1tura3/tor1 SZY33 VAC17/vac17TKanMX4 tor1tura3/tor1 SZY36 BY4742 vps33tkanmx4 SZY37 MATa his3d1 leu2d0 met15d0 ura3d0 tor1tleu2 SZY40 ura3d0/ura3d0 PEP3/pep3TKanMX4 tor1tleu2/tor1 SZY40-4 MATa his3d1 leu2d0 ura3d0 met15d0 tor1tleu2 pep3tkanmx4 [pep3 ts -108] SZY43 MATahis3D1 leu2d0 met15d0 ura3d0 tor1tura3 vac8tkanmx4 SZY49 VPS33/vps33TKanMX4 tor1tura3/tor1 RPY48 VPS39/vps39TKanMX4 tor1tura3/tor1 RPY49 VPS41/vps41TKanMX4 tor1tura3/tor1 RPY50 MATahis3D1 leu2d0 met15d0 ura3d0 tor1tura3 gtr1tkanmx4 RPY51 MATahis3D1 leu2d0 met15d0 ura3d0 tor1tura3 ego3tkanmx4 microscope equipped for epifluorescence and with a Nikon DXM1200F digital camera. RESULTS Mutation of TOR1 is synthetically lethal in combination with mutations in the class C VPS genes: To understand the functional divergence between Tor1 and Tor2, we performed a genomewide scale screen to identify genes that when mutated exhibit a reduced fitness or synthetic lethal interaction with a tor1 mutation. This screen, performed by diploid-based synthetic lethality analysis on microarrays (dslam) (Pan et al. 2004), yielded 261 interactions that met the cut-off control/experimental hybridization ratio (C/E ratio) of $2 (Table 2). Remarkably, this set of genes comprises distinct clusters that share common functions, including transcriptional
4 2142 Zurita-Martinez et al. TABLE 2 Synthetic lethal and reduced fitness interaction gene network of TOR1 ORF Gene Phenotype Vacuolar function/protein sorting YLR240W VPS34 SF YLR396C VPS33 SL YPL045W VPS16 SL YCL063W VAC17 SF YNL045W VAC7 SF YBR097W VPS15 SF YLR148W PEP3 SL YEL013W VAC8 SF YMR231W PEP5 SL YDL077C VPS39 SF YDR080W VPS41 SF YBR077C EGO3 SF YGR163W GTR2 SF YML121W GTR1 SF YKR007W EGO1 NC Ribosomal function YBR191W RPL21A NC YNL302C RPS19B NC YBL027W RPL19B NC YMR143W RPS16A NC YDR025W RPS11A NC YGL135W RPl1B NC YJR145C RPS4A NC YGL076C RPL7A NC YBL087C RPL23A NC YNL248C RPA49 NC Uncharacterized ORFs YDR417C YDR417C NC YNL086W YNL086W NC YAL011W SWC1 NC YDR161W YDR161W NC YJL022W YJL022W NC Mitochondrial function YFL016C MDJ1 NC YPL118W MRP51 NC YMR193W MRPL24 NC YPL013C MRPS16 NC YJL063C MRPL8 NC YCR046C IMG1 NC YLR426W YLR426W NC YGR171C MSM1 NC YPR100W MRPL51 NC YNL284C MRPL10 NC YNL073W MSK1 NC YMR158W MRPS8 NC YOR150W MRPl23 NC YHR091C MSR1 NC YKL170W MRPL38 NC YBR146W MRPS9 NC YNR037C RSM19 NC YIL093C RSM25 NC YBR268W MRPL37 NC YGR220C MRPL9 NC YKL138C MRPL31 NC YNL284C MRPL10 NC (continued ) TABLE 2 (Continued) ORF Gene Phenotype Other functions YIL086W CST6 NC YGL107C RMD9 NC YHR067W RMD12 NC YLR226W BUR2 NC YPR101W SNT309 NC YDL090C RAM1 NC YMR125W CBC1 NC YLR369W SSQ1 NC YPL059W GRX5 NC YPL178W CBC2 NC Genes are organized in functional categories according to the published literature. Note that only the tor1 interactions with genes involved in vacuolar function and protein sorting (except for ego1) were confirmed by tetrad dissection and the observed phenotypes, synthetic lethal (SL) or synthetic fitness defect (SF), are indicated. The rest of the interactions have not been confirmed (NC) and should be considered as candidate interactions. regulation, mrna processing, ribosomal and mitochondrial functions, vesicle docking and fusion, protein transport, microautophagy, and vacuolar inheritance (Table 2, Figure 1A, and data not shown). In this study only the genetic interactions involved in vacuolar functions and protein trafficking were validated by classic tetrad analysis and the rest should be considered as potential synthetic interactions until subject to further analysis. Tetrad analysis was conducted in mating crosses between the pep3, pep5, vps16, vps33, vps15, vps34, vac7, vac8, vac17, gtr1, gtr2, and ego3 deletion mutants and strain SZY25, in which the entire TOR1 open reading frame was replaced with the URA3 selectable marker. As shown in Figure 1B the haploid meiotic progeny of the tor1 and pep3, pep5, vps16, and vps33 crosses were wild type (WT), ura 1, or G418 resistant but no meiotic segregants with the ability to grow on both SD-ura and G418 selective media were recovered, confirming that these double mutants are synthetically lethal. In this analysis the rest of the genes examined exhibited a synthetic reduced fitness interaction when mutated in combination with tor1, as defined by the smaller size of the double mutant colony and a slow growth phenotype (illustrated in Figure 1C for the tor1 vps15, tor1 vps34, tor1 vac7, tor1 vac8, and tor1 vac17 and data not shown for tor1 gtr1, tor1 gtr2, and tor1 ego3 double mutants). The class C VPS genes also include VPS39 and VPS41; importantly, these genes were identified by the tor1 dslam screen but scored just below the C/E ratio of $2. On the basis of tetrad analysis, mutation of these vps genes in combination with tor1 also resulted in a synthetic reduced fitness phenotype (Figure 1C). These results validate the dslam screen and indicate that mutations in the class C VPS genes
5 Tor and Class C Vps Protein Complex 2143 Figure 1. TOR1 exhibits synthetically lethal and reduced fitness interactions with genes involved in protein sorting and vacuolar functions. (A) Schematic of the distinctive functional categories, according to the published literature, identified by the tor1 dslam screen. (B) Heterozygous diploid strains tor1 pep3 (SZY26), tor1 pep5 (SZY27), tor1 vps16 (SZY29), and tor1 vps33 (SZY49) (see Table 1 for complete genotypes) were sporulated and dissected on YPD solid medium. After 3 days of incubation at 30, colonies were replica plated onto YPD containing G418 (200 mg/ml) and SC-ura media. Plates were incubated for 2 days and photographed. (C) Heterozygous diploid strains tor1 vps15 (SZY28), tor1 vps34 (SZY30), tor1 vac7 (SZY31), tor1 vac17 (SZY33), tor1 vac8 (SZY32), tor1 vps39 (RPY50), and tor1 vps41 (RPY51) were sporulated and dissected as indicated above. Photographs show the colony size on the YPD plate of representative segregants from individual tetrads. show a synthetic lethal interaction with tor1. For the remainder of this study, the focus is on understanding the basis for the synthetic lethality and reduced fitness defect exhibited by the tor1 class C vps double mutants. Class C vps mutants fail to recover from rapamycininduced growth arrest: One hallmark of the genes linked to Tor signaling is that their mutation alters sensitivity to rapamycin. Mutation of 18 genes (including 14 involved in vacuolar functions and protein trafficking) out of the 62 shown in Table 2 resulted in rapamycin hypersensitivity as compared with the WT strain (Figure 2A) (Chan et al. 2000; Xie et al. 2005). In particular, mutants lacking components of the proteinsorting apparatus, including the class C Vps complex, the pre-autophagosomal (PAS) complex, as well as the vac8 mutant, are all extremely sensitive to rapamycin as compared to either the WT or the tor1 strains (Figure 2A). In addition, the class C vps mutants were tested for their ability to resume growth after rapamycin exposure. Actively growing cells were treated with rapamycin for 6 hr, washed, and spotted on YPD medium without drug. In contrast to the WT or tor1 strains, which readily recovered from rapamycin-induced growth arrest, mutations in class C VPS genes abolished recovery (Figure 2B). Interestingly, a similar phenotype was observed with strains harboring mutations in the EGO/Gse protein complex (Figure 1A), which was shown recently to play a role in combination with TORC1 in exit from rapamycininduced quiescence (Dubouloz et al. 2005). Expression of TOR1 but not TOR2 restores growth of a tor1 pep5 double mutant: The synthetic lethality of tor1 class C vps double mutants is surprising as it has been proposed that either Tor1 or Tor2 can provide all of the rapamycin-sensitive essential functions attributable to the TORC1 signaling complex. The Tor proteins share a high degree of amino acid sequence identity with a few discrete stretches of dissimilarity, particularly at the extreme N-terminus and, to a lesser extent, toward the C-terminal region. To understand the structural basis for the differential functions of the two TOR genes, we created TOR2-TOR1 chimeric hybrid alleles cloned into low-copy, centromeric plasmids and examined their ability to provide TOR1 function and restore viability in tor1 pep5 meiotic segregants. First the TOR2-TOR1 hybrids were verified to be efficiently expressed, based on Western blot analysis with antibodies specific for Tor1 or Tor2 (Figure 3A). Second, we made use of the fact that a tor1 ssd1 strain is unable to grow at 39 (Alarcon et al.
6 2144 Zurita-Martinez et al. Figure 2. Class C vps mutants are hypersensitive to rapamycin and fail to recover from rapamycin-induced growth arrest. (A) The WT (BY4742), tor1 (#16864), and isogenic strains bearing mutations in different components of the class C Vps and PAS protein complexes as well as in genes involved in vacuolar inheritance, were grown overnight in liquid YPD medium. Equivalent numbers of cells were serially diluted and aliquots were spotted onto YPD and YPD containing 20 nm rapamycin. Plates were photographed following 3 days incubation at 30. (B) Cultures of isogenic WT (BY4742), tor1 (#16864), pep5 (#10817), vps16 (#12783), and vps33 (SZY36) were grown to exponential phase. Cultures were divided in half and treated with drug vehicle alone or with 100 nm rapamycin for 6 hr. Cells were pelleted by centrifugation, washed twice, and equivalent numbers of cells were spotted on YPD medium. After incubation photographs were taken at 48 hr for both the untreated (YPD) and rapamycin-treated cells and 72 hr for the rapamycin-treated cells. 1999), apparently due to inability of TOR2 to support cell integrity in the absence of the TOR1 and SSD1 genes (Reinke et al. 2004). Modest overexpression of TOR1 or TOR2 as well as of the TOR2-TOR1 hybrids efficiently rescued growth of the tor1 ssd1 mutant strain at 39, indicating that the TOR2-TOR1 hybrids function to restore plasma membrane integrity (Figure 3B). While ectopic expression of plasmid-borne copies of TOR1 effectively rescued the growth of tor1 pep5 segregants, overexpression of TOR2 (also from a low-copy centromeric plasmid) failed to do so (Figure 3C). This result indicates that TOR1 has evolved to provide a function linked to the class C Vps complex which is not shared by TOR2. TOR2-TOR1 hybrids 1, 2, and 3 were Figure 3. Expression of TOR1 but not TOR2 rescues viability of tor1 pep5 segregants. (A) The tor1 ssd1 mutant strain SZY21 was transformed with vector prs315 and its derivatives encoding TOR1, TOR2, and the following TOR2 TOR1 hybrids (depicted in C): hybrid 1, Tor2 amino acids fused to Tor1 amino acids ; hybrid 2, Tor2 amino acids fused to Tor1 amino acids ; hybrid 3, Tor2 amino acids fused to Tor1 amino acids ; hybrid 4, the Tor1 amino acid sequence from was replaced by the homologous amino acid sequence of Tor2 from Protein extracts of the different transformants were analyzed by Western blot with antibodies specific to an N-terminal region of Tor1 or Tor2. In these extracts, Cpr1 was also detected and serve as a loading control. (B) Equivalent cell numbers of the SZY21 transformants as indicated above were serially diluted and spotted in SD-leu medium. Plates were photographed after incubation at 30 or 39 and Tor1 function was scored by the ability to complement the conditional synthetic lethal phenotype of the tor1 ssd1 strain at 39. (C) The tor1 pep5 diploid strain SZY27 transformed with the plasmids depicted at the left was sporulated and dissected on YPD medium. Following incubation at 30 for 3 days the plates were photographed and replica plated to YPD containing 200 mg/ml G418, SC-ura, and SC-leu media to score genotypes. The numbers on the right indicate the percent of tetrads (from a total of 10 scored) with a 4:0, 3:1, and 2:2 viable:inviable ratio.
7 Tor and Class C Vps Protein Complex 2145 able to suppress the synthetic lethality of tor1 pep5 segregants, albeit with an apparent differential efficiency. However, it should be noted that this assay examines both the increase in 4:0 segregation events and the relative growth of the rescued segregants. Based on an increase in 4:0 segregation events, hybrids 1, 2, and 3 all provide TOR1 function but it would be wrong to infer that hybrid 2 affords the best rescue because more 4:0 events were observed. Given the known segregation pattern of CEN-based plasmids (usually 2:2), the proportion of 4:0 segregation events seen with hybrids 1, 2, and 3 need not reflect a difference in rescue efficiency. In fact, hybrids 1, 3, and 4 rescue better based on colony growth. These results largely map this function to the C-terminal 788 amino acids of Tor1 (Figure 3C). Closer examination of this region in the Tor proteins revealed a discrete domain, from amino acid 1772 to 1815, which is highly dissimilar. However, replacement of this Tor1 region with the corresponding Tor2 sequence (hybrid 4) did not affect the ability to restore growth of tor1 pep5 segregants (Figure 3C). Thus, we conclude that the functional-structural differences between Tor1 and Tor2 map elsewhere in the C-terminal domain. Tor1 does not regulate the functions of the class C Vps complex: To gain insight into the molecular mechanisms that underlie the synthetic lethal interaction between TOR1 and class C VPS genes, we considered the hypothesis that Tor1 regulates the functions of this complex. This model is particularly attractive since Tor1 has been localized to internal membranes that resemble those associated with the endocytic pathway (Wedaman et al. 2003). To test this hypothesis, we examined if Tor1 mutation confers defects in class C Vps complex functions. The class C Vps protein complex is required for non-endosomal Golgi-to-vacuole transport, cytoplasmto-vacuole targeting (Cvt), recycling from endosomes back to the late Golgi, endocytosis, and autophagy. The tor1 mutant did not show defects in the maturation of the vacuolar hydrolases CpY, Ape1, or Alp1 as compared to the WTstrain or the pep3 mutant, which is defective in the proteolytic processing of these proteins (Figure 4A). Furthermore, CpY, Ape1, and Alp1 maturation was not altered by rapamycin treatment (Figure 4A). Moreover, mutation of TOR1 did not have any effect on endocytosis of the ammonium permease Mep2 elicited by ammonium addition to cells growing in ammonium-limiting medium (Figure 4B). These results indicate that the endosomal-to-vacuole, the Cvt, and the non-endosomalto-vacuole protein-sorting routes are not regulated by Tor signaling and that Tor1 does not regulate endocytosis. This result is in accord with a previous study that identified a rapamycin-insensitive role for Tor2, but not for Tor1, in regulating endocytosis (dehart et al. 2003). Earlier studies have shown that treatment of yeast cells with rapamycin results in autophagy, indicating a role for Tor signaling in regulating this process (Noda and Ohsumi 1998). To test if mutation of TOR1 had any Figure 4. Tor1 signaling does not control class C Vps complex functions. (A) Tor1 does not control the maturation of vacuolar hydrolases. Exponentially growing cultures of the WT (BY4742), tor1 (#16864), and pep3 (#14105) strains in YPD medium were treated with either drug vehicle ( ) or 100 nm rapamycin (1) for 1 hr. Protein extracts were prepared and analyzed by Western blot with antibodies specific for CpY, Ape1, and Alp1. (B) Tor 1 does not regulate endocytosis of the high-affinity ammonium permease Mep2. The WT, tor1, and pep3 strains indicated in A were transformed with plasmid pmep2-gfp. Transformants were grown to early exponential phase in SC-ura, washed twice, and resuspended in SLAD medium supplemented with the required amino acids to satisfy auxotrophic requirements. After 4 hr incubation in this medium, 50 mm ammonium sulfate was added to the cultures and incubation continued for 30 min. Cell samples were collected prior to and after addition of ammonium sulfate (NH 1 4 ) and imaged for direct epifluorescence as indicated under experimental procedures. (C) Effects of TOR1 mutation in autophagy-mediated maturation of Ape1. The WT (BY4742), and the tor1 (#16864), pep3 (#14105), vac8 (#10253), and the tor1 vac8 (SZY43) mutant strains were grown and treated with rapamycin as indicated in A. Protein extracts were prepared and analyzed by Western blot with specific Ape1 antibodies. effect on autophagy, we made use of the fact that a vac8 mutant is defective in the Cvt pathway. Therefore, in a vac8 mutant, maturation of proape1 in the vacuole is defective; however, this defect can be bypassed by induction of autophagy (Abeliovich et al. 2000). Maturation of Ape1 is enhanced and induced by rapamycin
8 2146 Zurita-Martinez et al. Figure 5. Tor1 does not regulate protein sorting from the ER to Golgi complex or protein cycling between the Golgi complex and endosomes. (A) Schematic of a fraction of the genes (grouped in functional categories according to the published literature) that when mutated alter rapamycin sensitivity and showed reduced fitness and synthetically lethal interactions in combination with mutation of PEP3. Interactions of the genes indicated in bold were confirmed by tetrad analysis. Genes enclosed by a box were synthetically lethal in combination with the pep3 mutation. (B) The WT (BY4742), tor1 (#16864), and pep3 (#14105) mutants were grown to early exponential phase and treated with drug vehicle ( )or100nm rapamycin for 20 min (1). Cell were pulse labeled with Trans 35 S-LABEL for 8 min (min) and chased with unlabeled amino acids for 0, 5, 6, 14, 16, and 20 min as indicated in the figure (for further detail, see materials and methods). The mature (m) and processed (p) forms of a-factor (top) and CpY (bottom) were immunoprecipitated with a-factor and CpY-specific antibodies, respectively. Immunoprecipitated proteins were separated by SDS-PAGE and visualized by autoradiography. The migration of precursor forms characteristic for the Golgi complex and the ER compartment are indicated. in the WT and vac8 strains, respectively (Figure 4C). Interestingly, mutation of TOR1 restores Ape1 processing in the vac8 cells, and rapamycin treatment of the tor1 vac8 double mutant dramatically enhances Ape1 processing (Figure 4C). As expected, mutation of PEP3 efficiently blocked the rapamycin-induced maturation of Ape1. These results show that mutation of TOR1 alone results in a low level of autophagy, whereas mutation of PEP3 blocks this process. We reasoned that if the basis for the synthetic lethality of the tor1 class C vps double mutants arose from a defect in effective execution of autophagy, tor1 mutation should also exhibit synthetic lethality in combination with mutations in other genes required for autophagy. Although our screen detected a synthetic reduced fitness interaction between tor1 and vps15 or vps34, two genes required for autophagy (Figure 1A) (Kihara et al. 2001), we found that a tor1 apg13 double-mutant strain that should be defective in autophagy was fully viable (data not shown). Synthetic lethal interaction can arise between two genes whose products act in parallel or compensating pathways. Accordingly, we considered the hypothesis that TOR1 regulates a pathway that functions in parallel with the class C complex for protein sorting. A prediction of this model is that key genes working within this pathway should be synthetically lethal in combination with a pep3 mutation. A dslam screen with pep3 revealed 38 genes that are known when mutated to alter the sensitivity of cells to rapamycin (Figure 5A, data not shown) (Chan et al. 2000; Xie et al. 2005). Within this set, 13 genes share distinctive roles in protein trafficking between the ER to Golgi complex and protein cycling between the Golgi complex and endosomes (Figure 5A). To test if Tor1 has a role in protein trafficking between these cell compartments, we tested the effects of TOR1 mutation and rapamycin treatment on a-factor maturation. The mating pheromone a factor is synthesized as a precursor and subjected to extensive glycosylation as it transverses the ER and Golgi complex, and then is cleaved at the late Golgi complex by the Kex2 protease to produce mature pheromone. Importantly, Kex2 normally cycles between the late Golgi complex and endosomal compartments. Thus, a-factor maturation serves as a reporter of protein trafficking between the ER to Golgi complex and protein cycling between the Golgi complex and endosomes. In this experiment we also monitored the processing of CpY in more detail. While mutation of PEP3 resulted in accumulation of glycosylated forms of a factor in the Golgi complex, and also blocked CpY processing, neither mutation of TOR1 nor rapamycin treatment of the WT strain perturbed a-factor or CpY processing to any significant extent (Figure 5B). Furthermore, rapamycin treatment in the pep3 strain did not alter the ratio of mature to immature a-factor forms observed in the pep3 untreated cells (Figure 5B). In addition, mutation of TOR1 or rapamycin treatment had no effect on the normal cycling of the SNARE protein Snc1 from early endosomes to the Golgi compartment and back to the plasma membrane (data not shown). Collectively, these results exclude models
9 Tor and Class C Vps Protein Complex 2147 Figure 6. Class C vps mutants show severe defects in nitrogen metabolism. (A) The class C Vps complex is required for adaptation to nitrogen limitation. Isogenic WT (BY4742), and pep5 (#10817), vps16 (#12783), vps33 (SZY36), and tor1 (#16864) mutant strains were grown and spotted on YPD medium as indicated in the legend to Figure 2A. Following overnight incubation, the plate was photographed (see overnight image) and then starved for nitrogen by replica-plating to YNB without nitrogen medium. After incubating for 10 days, cells were replica plated back to YPD, incubated for 24 hr, and photographed. (B) Class C Vps complex mutations induce the general amino acid control response. Exponentially growing cultures of the WT (BY4742), and the pep5 (#10817), pep3 (#14105) mutant strains harboring the GCN4-lacZ reporter plasmid p180 were grown to exponential phase in SC-ura medium. Cultures were treated with 100 nm rapamycin for 0, 1, and 2 hr and analyzed for b-galactosidase activity. (C) Growth of the class C vps mutants at 37 is rescued by addition of glutamine, glutamate, and to a lesser extent by arginine. Isogenic WT (BY4742), and tor1 (#16864), pep3 (#14105), tor1 pep3 (strain SZY40-4 transformed with plasmid pbj9113 expressing the pep3-108ts allele), pep5 (#10817), vps16 (#12783), vps33 (SZY36), vps39 (#13774), and vps41 (#14015) mutant strains were grown and spotted on SC medium ( ) or SC medium supplemented with the indicated amino acid. Plates were incubated at 37 for 2 days and photographed. (D) Supplementation with glutamine or glutamate does not rescue the growth defect of the tor1 vac8, tor1 gtr1, and tor1 ego3 double mutants. Isogenic WT (BY4742), tor1 vac8 (SZY43), tor1 gtr1(rpy50), and tor1 ego3 (RPY51) double mutant strains were assayed as indicated in C except that plates were incubated at 30. that evoke a role for Tor1 in regulating the class C Vps complex, or acting in a parallel compensating pathway to provide a known class C Vps complex function. The synthetic lethality between TOR1 and PEP3 mutations is suppressed by amino acids: Next, we entertained a modelin which the class C Vps complexpromotes Tor activity. Because Tor signaling is activated by nutrients, we tested the ability of the class C vps mutants to respond to nitrogen starvation. In contrast to the WT and tor1 strains, which effectively survived a 10-day period of incubation on ammonium-starvation medium, the class C vps mutants failed to resume growth (Figure 6A). Previous reports have shown that class C vps mutants contain severely fragmented vacuoles and are unable to store normal levels of basic amino acids in the vacuole (Kitamoto et al. 1988; Raymond et al. 1992). Low levels of intracellular amino acids, or rapamycin exposure, are known to trigger the general amino acid control response by inactivation of eif2a, which results in a block to general protein synthesis and preferential translation of Gcn4. We examined if the low amino acid levels in class C vps mutants would suffice to activate Gcn4 translation. In accord with the above observation, pep3 or pep5 mutations resulted in robust Gcn4 translation, comparable to that observed in the WT strain treated with rapamycin (Figure 6B). It has been proposed that glutamine and possibly glutamate, which are amino acids central to nitrogen metabolism, regulate nutrient signaling pathways, including the Tor pathway (Crespo et al. 2002; reviewed by Liu and Butow 2006). These observations prompted us to examine in more detail the intracellular amino acid content in the class C vps mutants. Under the growth conditions analyzed (active growth in YPD medium) the pep3, pep5, and vps33 mutants showed a lower level of the basic amino acids lysine, histidine, and arginine than that detected in the WT strain, in accord with an earlier report (Table 3) (Kitamoto et al. 1988). Remarkably, in contrast to the WTand tor1 strains, glutamate levels were significantly reduced (1.5-fold) in the class C vps mutants. The class C Vps complex is also required to support growth at 37 (Figure 6C) (Robinson et al.
10 2148 Zurita-Martinez et al. TABLE 3 The class C vps mutants show dramatically reduced levels of amino acids at 37 Amino acid content (nmol/od 600 ) Glutamate Glutamine Lysine Histidine Arginine Strain WT tor pep pep ND vps ND ND, not detected. 1991, Koning et al. 2002). We sought to test if this growth defect is linked to amino acid levels. Exponentially growing cultures of the different strains were shifted from 30 to 37 and the amino acid levels were determined. Interestingly, shift to 37 for 3 hr caused a modest decline in glutamate and a near twofold decrease in glutamine in both the WT and the tor1 strains. In the pep3, pep5, and vps33 mutants the concentration of these amino acids declined more markedly, two- to threefold depending on the strain, and glutamate reached a threefold lower level than the one observed in the WT strain under similar conditions (Table 3). Moreover, supplementation of the growth media with glutamate, or with glutamine and to lesser extent with arginine, restored growth of the pep3, pep5, vps33, vps39, and vps41 mutant strains at 37. This effect was specific and was not observed when the individual basic amino acids lysine or histidine were added as supplements (Figure 6C). The vps16 strain has a more severe growth defect at 37 than the rest of the class C vps mutants and this is only marginally alleviated by glutamine supplementation (see below for discussion). Strikingly, glutamate or glutamine and less efficiently arginine also supported the growth of a tor1 pep3 double-mutant strain carrying a partial loss of function pep3 ts allele on a plasmid (Figure 6C). In contrast, supplementation with glutamine or glutamate did not rescue the growth defect of the tor1 vac8, tor1 gtr1,or tor1 ego3 double mutants (Figure 6D). Finally, supplementation with glutamine, glutamate, arginine, lysine, and histidine partially rescued the viability of tor1 pep5 segregants; however, these segregants grew poorly and rapidly accumulated suppressor mutations (data not shown). These results argue that the synthetic lethality of the tor1 pep3 double mutant derives from the metabolic derangement that underlies the reduced amino acid concentrations observed in the class C vps mutants. DISCUSSION Our genomewide search to define synthetic fitness or lethal interaction partners of TOR1 identified sets of genes that provide distinct functions. Among the synthetically lethal genes identified, four encode components of the class C Vps complex that functions in the recognition and fusion of vesicles with vacuolar and secretory membranes. This screen also identified a synthetic reduced fitness interaction between tor1 and mutations in genes encoding the components of the EGO/ Gse complex, confirming and extending a previous study (Dubouloz et al. 2005). The EGO/Gse complex localizes to pre-vacuolar and vacuolar membranes and is required for sorting of the Gap1 amino acid permease to the plasma membrane in response to intracellular amino acids (Gao and Kaiser 2006). Rapamycin exposure results in autophagy and thereby in a massive influx of membranes into the vacuolar membrane (Noda and Ohsumi 1998). In combination with TORC1, the EGO/Gse complex has been proposed to play a role in recovery from rapamycin-induced cell cycle arrest by enabling recycling of vacuolar membranes via microautophagy (Dubouloz et al. 2005). We have shown here that class C vps mutants are unable to recover from prolonged exposure to rapamycin. Although it is possible that the class C Vps complex is required for microautophagy, we were unable to address this question since class C vps mutants fail to complete autophagy and contain severely fragmented vacuoles. We find that the structural difference between Tor1 and Tor2 to support viability of tor1 pep5 segregants maps to the C-terminal 788 amino acid region of Tor1. This result is in contrast with a previous study which concluded that the ability of a Tor1SR (rapamycin-resistant mutant) to confer rapamycin resistance in a vps16 mutant lies in the N-terminal 120 amino acid region of Tor1 (Xie et al. 2005). At present we do not have a good explanation for this discrepancy other than the biological assays to test for Tor1 function in the two studies were different. Our screen also identified the Vps34 phosphatidylinositol 3-kinase and the Vps15 protein kinase, members of the PAS protein complex which functions in autophagy and protein sorting (Stack et al. 1995; Kihara et al. 2001), and genes involved in vacuolar inheritance (Figure 1A). Taken together these results reveal a prominent link between Tor signaling and vacuolar function. This
11 Tor and Class C Vps Protein Complex 2149 view is further underscored by studies that have localized different components of the Tor pathway to the vacuolar membrane, including Tor2, Kog1, and Tco89 (Cardenas and Heitman 1995; Reinke et al. 2004; Araki et al. 2005). An important function of the vacuole is to preserve amino acid homeostasis by sequestering basic amino acids, which are toxic when present at high concentrations in the cytosol (reviewed by Klionsky et al. 1990). Our results and those of others have shown that mutants lacking components of the class C Vps complex are unable to recover from ammonium starvation, they have low concentrations of intracellular amino acids, particularly glutamate and glutamine, and this concentration is further reduced upon shift of these mutants from 30 to 37. Moreover, class C vps mutants have growth defects at 37 (Figure 6C) (Robinson et al. 1991, Koning et al. 2002). Remarkably, addition of glutamate or glutamine and to lesser extent arginine restored the growth at 37 of individual class C vps mutants and a tor1 pep3 mutant carrying a pep3 ts allele, and partially rescued the viability of tor1 pep5 segregants. The effect of arginine could be explained by the ability of this amino acid to be converted into glutamate by the concerted actions of arginase and ornithine transaminase (reviewed by Davis 1986). We found that the vps16 strain was only marginally rescued by glutamine supplementation, suggesting that Vps16 has an additional function(s) different from that shared with the other members of the class C Vps complex. In this regard a role for Vps16 in regulating mrna decapping and stability has been demonstrated (Zhang et al. 1999). Interestingly, genes encoding two subunits of the cap-binding complex, CBC1 and CBC2, were also identified by the tor1 synthetic screen and a rapamycin-sensitive role for Tor in controlling mrna stability has been described (Albig and Decker 2001). Thus, it is possible that Tor1 function becomes compromised in the vps16 mutant via these activities independent of the class C Vps complex function. In summary, our results indicate that the class C Vps complex is required to maintain amino acid homeostasis for effective Tor activity and lend further support to the proposal that glutamate/glutamine activates Tor signaling (Crespo et al. 2002). These studies also demonstrate that Tor1 is more efficient than Tor2 at providing rapamycin-sensitive Tor signaling under conditions where intracellular amino acids are drastically reduced. In support of this model, whereas Tor1 is a bona fide member of the rapamycin-sensitive TORC1 complex, Tor2 weakly interacts with this complex only in cells lacking Tor1 and no stable association of Tor2 with TORC1 has been detected in wild-type cells (Loewith et al. 2002; Reinke et al. 2004). Strikingly, although both humans and many fungi express a single Tor kinase, the Tor genes have been independently duplicated in both the budding yeast S. cerevisiae and the fission yeast Schizosaccharomyces pombe (reviewed by Weisman 2004). Interestingly, in both cases one of the two paralogs is essential (TOR2) and the other is not under standard conditions (TOR1). Early studies in S. pombe revealed a role for Tor1 in amino acid uptake (Weisman et al. 2005). A more recent study has proposed a shared function for Tor1 and Tor2 in enabling cell proliferation and survival under both normal and adverse conditions and a positive and negative role for Tor1 and Tor2, respectively, in regulating G1 arrest and sexual differentiation (Uritani et al. 2006). Taken together, our findings provide a physiological basis for understanding the functional differences that distinguish Tor1 and Tor2 in S. cerevisiae and yield insight as to why the two genes are among the few (2 8%) retained following the genome duplication and massive gene loss event that marked the evolution of hemiascomycetous yeasts. We thank Todd Graham for advice and suggestions with the a factor processing experiments. We thank Todd Graham, Elizabeth Jones, Yoshinori Ohsumi, Alan Hinnebusch, and Julian Rutherford for generous gifts of plasmids and antisera and Joseph Heitman and John McCusker for critical reading of the manuscript. This work was supported by R01CA from the National Cancer Institute (to M.E.C); Xuewen Pan was supported by the Leukemia and Lymphoma Society and by National Institute of Health grant HG (to J.D.B). LITERATURE CITED Abeliovich, H., W. A. Dunn, Jr., J. Kim and D. J. Klionsky, 2000 Dissection of autophagosome biogenesis into distinct nucleation and expansion steps. J. Cell Biol. 151: Alarcon, C. M., M. E. Cardenas and J. Heitman, 1996 Mammalian RAFT1 kinase domain provides rapamycin-sensitive TOR function in yeast. Genes Dev. 10: Alarcon, C. M., J. Heitman and M. E. Cardenas, 1999 Protein kinase activity and identification of a toxic effector domain of the target of rapamycin TOR proteins in yeast. Mol. Biol. Cell 10: Albig, A. R., and C. J. Decker, 2001 The target of rapamycin signaling pathway regulates mrna turnover in the yeast Saccharomyces cerevisiae. Mol. Biol. Cell 12: Araki, T., Y. Uesono, T. Oguchi and E. A. Toh, 2005 LAS24/KOG1, a component of the TOR complex 1 (TORC1), is needed for resistance to local anesthetic tetracaine and normal distribution of actin cytoskeleton in yeast. Genes Genet. Syst. 80: Banta, L. M., J. S. Robinson, D. J. Klionsky and S. D. Emr, 1988 Organelle assembly in yeast: characterization of yeast mutants defective in vacuolar biogenesis and protein sorting. J. Cell Biol. 107: Beck, T., and M. N. Hall, 1999 The TOR signalling pathway controls nuclear localization of nutrient-regulated transcription factors. Nature 402: Berset, C., H. Trachsel and M. Altmann, 1998 The TOR (target of rapamycin) signal transduction pathway regulates the stability of translation initiation factor eif4g in the yeast Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 95: Cardenas, M. E., and J. Heitman, 1995 FKBP12-rapamycin target TOR2 is a vacuolar protein with an associated phosphatidylinositol-4 kinase activity. EMBO J. 14: Cardenas, M. E., N. S. Cutler, M. C. Lorenz, C. J. Di Como and J. Heitman, 1999 The TOR signaling cascade regulates gene expression in response to nutrients. Genes Dev. 13: Chan, T. F., J. Carvalho, L.Riles and X. F. Zheng, 2000 A chemical genomics approach toward understanding the global functions of the target of rapamycin protein (TOR). Proc. Natl. Acad. Sci. USA 97: Chen, E. J., and C. A. Kaiser, 2002 Amino acids regulate the intracellular trafficking of the general amino acid permease of Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 99:
12 2150 Zurita-Martinez et al. Cherkasova, V. A., and A. G. Hinnebusch, 2003 Translational control by TOR and TAP42 through dephosphorylation of eif2alpha kinase GCN2. Genes Dev. 17: Crespo, J. L., and M. N. Hall, 2002 Elucidating TOR signaling and rapamycin action: lessons from Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 66: Crespo, J. L., T. Powers, B.Fowler and M. N. Hall, 2002 The TOR-controlled transcription activators GLN3, RTG1, and RTG3 are regulated in response to intracellular levels of glutamine. Proc. Natl. Acad. Sci. USA 99: Davis, R. H., 1986 Compartmental and regulatory mechanisms in the arginine pathways of Neurospora crassa and Saccharomyces cerevisiae. Microbiol. Rev. 50: dehart, A. K., J. D. Schnell, D. A. Allen, J. Y. Tsai and L. Hicke, 2003 Receptor internalization in yeast requires the Tor2-Rho1 signaling pathway. Mol. Biol. Cell 14: De Virgilio, C., and R. Loewith, 2006 Cell growth control: little eukaryotes make big contributions. Oncogene 25: Dubouloz, F., O. Deloche, V. Wanke, E. Cameroni and C. De Virgilio, 2005 The TOR and EGO protein complexes orchestrate microautophagy in yeast. Mol. Cell 19: Gao, M., and C. A. Kaiser, 2006 A conserved GTPase-containing complex is required for intracellular sorting of the general amino-acid permease in yeast. Nat. Cell Biol. 8: Gimeno, C. J., P. O. Ljungdahl, C. A. Styles and G. R. Fink, 1992 Unipolar cell divisions in the yeast S. cerevisiae lead to filamentous growth: regulation by starvation and RAS. Cell 68: Goldstein, A. L., and J. H. McCusker, 1999 Three new dominant drug resistance cassettes for gene disruption in Saccharomyces cerevisiae. Yeast 15: Graham, T. R., 1998 Metabolic labeling and immunoprecipitation of yeast proteins. John Wiley & Sons, New York. Hardwick, J. S., F. G. Kuruvilla, J. K. Tong, A. F. Shamji and S. L. Schreiber, 1999 Rapamycin-modulated transcription defines the subset of nutrient-sensitive signaling pathways directly controlled by the Tor proteins. Proc. Natl. Acad. Sci. USA 96: Heitman, J., N. R. Movva and M. N. Hall, 1991 Targets for cell cycle arrest by the immunosuppressant rapamycin in yeast. Science 253: Kihara, A., T. Noda, N. Ishihara and Y. Ohsumi, 2001 Two distinct Vps34 phosphatidylinositol 3-kinase complexes function in autophagy and carboxypeptidase Y sorting in Saccharomyces cerevisiae. J. Cell Biol. 152: Kitamoto,K.,K.Yoshizawa,Y.Ohsumi and Y. Anraku, 1988 Mutants of Saccharomyces cerevisiae with defective vacuolar function. J. Bacteriol. 170: Klionsky, D. J., P. K. Herman and S. D. Emr, 1990 The fungal vacuole: composition, function, and biogenesis. Microbiol. Rev. 54: Komeili, A., K. P. Wedaman, E. K. O Shea and T. Powers, 2000 Mechanism of metabolic control. Target of rapamycin signaling links nitrogen quality to the activity of the Rtg1 and Rtg3 transcription factors. J. Cell Biol. 151: Koning, A. J., L. L. Larson, E.J.Cadera, M.L.Parrish and R. L. Wright, 2002 Mutations that affect vacuole biogenesis inhibit proliferation of the endoplasmic reticulum in Saccharomyces cerevisiae. Genetics 160: Kubota, H., T. Obata, K. Ota, T. Sasaki and T. Ito, 2003 Rapamycin-induced translational derepression of GCN4 mrna involves a novel mechanism for activation of the eif2[a] Kinase GCN2. J. Biol. Chem. 278: Liu, Z., and R. A. Butow, 2006 Mitochondrial retrograde signaling. Annu. Rev. Genet. 40: Loewith, R., E. Jacinto, S. Wullschleger, A. Lorberg, J. L. Crespo et al., 2002 Two TOR complexes, only one of which is rapamycin sensitive, have distinct roles in cell growth control. Mol. Cell 10: Longtine, M. S., A. McKenzie, III, D. J. Demarini, N. G. Shah, A. Wach et al., 1998 Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14: Lorenz,M.C.,andJ.Heitman, 1995 TOR mutations confer rapamycin resistance by preventing interaction with FKBP12-rapamycin. J. Biol. Chem. 270: Noda, T., andy. Ohsumi, 1998 Tor, a phosphatidylinositol kinase homologue, controls autophagy in yeast. J. Biol. Chem. 273: Pan, X., D. S. Yuan, D. Xiang, X. Wang, S. Sookhai-Mahadeo et al., 2004 A robust toolkit for functional profiling of the yeast genome. Mol. Cell 16: Peterson, M. R., and S. D. Emr, 2001 The class C Vps complex functions at multiple stages of the vacuolar transport pathway. Traffic 2: Powers, T., and P. Walter, 1999 Regulation of ribosome biogenesis by the rapamycin-sensitive TOR-signaling pathway in Saccharomyces cerevisiae. Mol. Biol. Cell 10: Raymond, C. K., I. Howald-Stevenson, C. A. Vater and T. H. Stevens, 1992 Morphological classification of the yeast vacuolar protein sorting mutants: evidence for a prevacuolar compartment in class E vps mutants. Mol. Biol. Cell 3: Reinke, A., S. Anderson, J. M. McCaffery, J. Yates, III, S. Aronova et al., 2004 TOR complex 1 includes a novel component, Tco89p (YPL180w), and cooperates with Ssd1p to maintain cellular integrity in Saccharomyces cerevisiae. J.Biol.Chem.279: Rieder, S. E., and S. D. Emr, 1997 A novel RING finger protein complex essential for a late step in protein transport to the yeast vacuole. Mol. Biol. Cell 8: Robinson, J. S., T. R. Graham ands. D. Emr, 1991 A putative zinc finger protein, Saccharomyces cerevisiae Vps18p, affects late Golgi functions required for vacuolar protein sorting and efficient alpha-factor prohormone maturation. Mol. Cell. Biol. 11: Rohde, J., J. Heitman and M. E. Cardenas, 2001 The TOR kinases link nutrient sensing to cell growth. J. Biol. Chem. 276: Rohde, J. R., S. Campbell, S. A. Zurita-Martinez, N. S. Cutler, M. Ashe et al., 2004 TOR controls transcriptional and translational programs via Sap-Sit4 protein phosphatase signaling effectors. Mol. Cell. Biol. 24: Schiestl, R. H., and R. D. Gietz, 1989 High efficiency transformation of intact yeast cells using single stranded nucleic acids as a carrier. Curr. Genet. 16: Sherman, F., 2002 Getting started with yeast. Methods Enzymol. 350: Srivastava, A., C. A. Woolford and E. W. Jones, 2000 Pep3p/ Pep5p complex: a putative docking factor at multiple steps of vesicular transport to the vacuole of Saccharomyces cerevisiae. Genetics 156: Stack, J. H., B. Horazdovsky and S. D. Emr, 1995 Receptor-mediated protein sorting to the vacuole in yeast: roles for a protein kinase, a lipid kinase and GTP-binding proteins. Annu. Rev. Cell Dev. Biol 11: Uritani, M., H. Hidaka, Y. Hotta, M. Ueno, T. Ushimaru et al., 2006 Fission yeast Tor2 links nitrogen signals to cell proliferation and acts downstream of the Rheb GTPase. Genes Cells 11: Valenzuela, L., C. Aranda and A. Gonzalez, 2001 TOR modulates GCN4-dependent expression of genes turned on by nitrogen limitation. J. Bacteriol. 183: Wedaman,K.P.,A.Reinke,S.Anderson,J.Yates, III, J. M. McCaffery et al., 2003 Tor kinases are in distinct membrane-associated protein complexes in Saccharomyces cerevisiae. Mol. Biol. Cell 14: Weisman, R., 2004 The fission yeast TOR proteins and the rapamycin response: an unexpected tale. Curr. Top. Microbiol. Immunol. 279: Weisman, R., I. Roitburg, T. Nahari and M. Kupiec, 2005 Regulation of leucine uptake by tor11 in Schizosaccharomyces pombe is sensitive to rapamycin. Genetics 169: Wurmser, A. E., T. K. Sato and S. D. Emr, 2000 New component of the vacuolar class C Vps complex couples nucleotide exchange on the Ypt7 GTPase to SNARE-dependent docking and fusion. J. Cell Biol. 151: Xie, M. W., F. Jin, H. Hwang, S. Hwang, V. Anand et al., 2005 Insights into TOR function and rapamycin response: chemical genomic profiling by using a high-density cell array method. Proc. Natl. Acad.Sci.USA102: Zaragoza, D., A. Ghavidel, J. Heitman and M. C. Schultz, 1998 Rapamycin induces the G0 program of transcriptional repression in yeast by interfering with the TOR signaling pathway. Mol. Cell. Biol. 18: Zhang, S., C. J. Williams, K. Hagan ands. W. Peltz, 1999 Mutations in VPS16 and MRT1 stabilize mrnas by activating an inhibitor of the decapping enzyme. Mol. Cell. Biol. 19: Communicating editor: A. P. Mitchell
Genetics Lecture Notes 7.03 2005. Lectures 1 2
 Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Lecture 8. Protein Trafficking/Targeting. Protein targeting is necessary for proteins that are destined to work outside the cytoplasm.
 Protein Trafficking/Targeting (8.1) Lecture 8 Protein Trafficking/Targeting Protein targeting is necessary for proteins that are destined to work outside the cytoplasm. Protein targeting is more complex
Protein Trafficking/Targeting (8.1) Lecture 8 Protein Trafficking/Targeting Protein targeting is necessary for proteins that are destined to work outside the cytoplasm. Protein targeting is more complex
The Awesome Power of Yeast Genetics: Spontaneous and Induced Mutagenesis and Complementation Analysis using Saccharomyces cerevisiae.
 The Awesome Power of Yeast Genetics: Spontaneous and Induced Mutagenesis and Complementation Analysis using Saccharomyces cerevisiae. Mutations occur as a consequence of normal cellular physiology and
The Awesome Power of Yeast Genetics: Spontaneous and Induced Mutagenesis and Complementation Analysis using Saccharomyces cerevisiae. Mutations occur as a consequence of normal cellular physiology and
Molecular Biology. Yeast Transformation. Yeast Plasmids. Gene Disruption, tagging. Cloning by Complementation. Epistasis
 Molecular Biology Yeast Transformation Yeast Plasmids Gene Disruption, tagging Cloning by Complementation Epistasis Transformation Transformation: introduction of DNA 1978, ca 1000x less efficient than
Molecular Biology Yeast Transformation Yeast Plasmids Gene Disruption, tagging Cloning by Complementation Epistasis Transformation Transformation: introduction of DNA 1978, ca 1000x less efficient than
Sickle cell anemia: Altered beta chain Single AA change (#6 Glu to Val) Consequence: Protein polymerizes Change in RBC shape ---> phenotypes
 Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
Chromatin Immunoprecipitation
 Chromatin Immunoprecipitation A) Prepare a yeast culture (see the Galactose Induction Protocol for details). 1) Start a small culture (e.g. 2 ml) in YEPD or selective media from a single colony. 2) Spin
Chromatin Immunoprecipitation A) Prepare a yeast culture (see the Galactose Induction Protocol for details). 1) Start a small culture (e.g. 2 ml) in YEPD or selective media from a single colony. 2) Spin
Anti-ATF6 α antibody, mouse monoclonal (1-7)
 Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
Molecular and Cell Biology Laboratory (BIOL-UA 223) Instructor: Ignatius Tan Phone: 212-998-8295 Office: 764 Brown Email: ignatius.tan@nyu.
 Molecular and Cell Biology Laboratory (BIOL-UA 223) Instructor: Ignatius Tan Phone: 212-998-8295 Office: 764 Brown Email: ignatius.tan@nyu.edu Course Hours: Section 1: Mon: 12:30-3:15 Section 2: Wed: 12:30-3:15
Molecular and Cell Biology Laboratory (BIOL-UA 223) Instructor: Ignatius Tan Phone: 212-998-8295 Office: 764 Brown Email: ignatius.tan@nyu.edu Course Hours: Section 1: Mon: 12:30-3:15 Section 2: Wed: 12:30-3:15
Chapter 8. Summary and Perspectives
 Chapter 8 Summary and Perspectives 131 Chapter 8 Summary Overexpression of the multidrug resistance protein MRP1 confer multidrug resistance (MDR) to cancer cells. The contents of this thesis describe
Chapter 8 Summary and Perspectives 131 Chapter 8 Summary Overexpression of the multidrug resistance protein MRP1 confer multidrug resistance (MDR) to cancer cells. The contents of this thesis describe
Recombinant DNA Unit Exam
 Recombinant DNA Unit Exam Question 1 Restriction enzymes are extensively used in molecular biology. Below are the recognition sites of two of these enzymes, BamHI and BclI. a) BamHI, cleaves after the
Recombinant DNA Unit Exam Question 1 Restriction enzymes are extensively used in molecular biology. Below are the recognition sites of two of these enzymes, BamHI and BclI. a) BamHI, cleaves after the
Cell Biology Questions and Learning Objectives
 Cell Biology Questions and Learning Objectives (with hypothetical learning materials that might populate the objective) The topics and central questions listed here are typical for an introductory undergraduate
Cell Biology Questions and Learning Objectives (with hypothetical learning materials that might populate the objective) The topics and central questions listed here are typical for an introductory undergraduate
BME 42-620 Engineering Molecular Cell Biology. Lecture 02: Structural and Functional Organization of
 BME 42-620 Engineering Molecular Cell Biology Lecture 02: Structural and Functional Organization of Eukaryotic Cells BME42-620 Lecture 02, September 01, 2011 1 Outline A brief review of the previous lecture
BME 42-620 Engineering Molecular Cell Biology Lecture 02: Structural and Functional Organization of Eukaryotic Cells BME42-620 Lecture 02, September 01, 2011 1 Outline A brief review of the previous lecture
Lecture 6: Single nucleotide polymorphisms (SNPs) and Restriction Fragment Length Polymorphisms (RFLPs)
 Lecture 6: Single nucleotide polymorphisms (SNPs) and Restriction Fragment Length Polymorphisms (RFLPs) Single nucleotide polymorphisms or SNPs (pronounced "snips") are DNA sequence variations that occur
Lecture 6: Single nucleotide polymorphisms (SNPs) and Restriction Fragment Length Polymorphisms (RFLPs) Single nucleotide polymorphisms or SNPs (pronounced "snips") are DNA sequence variations that occur
(1-p) 2. p(1-p) From the table, frequency of DpyUnc = ¼ (p^2) = #DpyUnc = p^2 = 0.0004 ¼(1-p)^2 + ½(1-p)p + ¼(p^2) #Dpy + #DpyUnc
 Advanced genetics Kornfeld problem set_key 1A (5 points) Brenner employed 2-factor and 3-factor crosses with the mutants isolated from his screen, and visually assayed for recombination events between
Advanced genetics Kornfeld problem set_key 1A (5 points) Brenner employed 2-factor and 3-factor crosses with the mutants isolated from his screen, and visually assayed for recombination events between
to 370C in a matter of seconds. Materials and Methods.-Yeast spheroplasts were prepared, resuspended to a concentration
 A MUTANT OF YEAST APPARENTLY DEFECTIVE IN THE INITIATION OF PROTEIN SYNTHESIS* BT LELAND H. HARTWELLt AND CALVIN S. MCLAUGHLIN DEPARTMENT OF MOLECULAR AND CELL BIOLOGY, UNIVERSITY OF CALIFORNIA (IRVINE)
A MUTANT OF YEAST APPARENTLY DEFECTIVE IN THE INITIATION OF PROTEIN SYNTHESIS* BT LELAND H. HARTWELLt AND CALVIN S. MCLAUGHLIN DEPARTMENT OF MOLECULAR AND CELL BIOLOGY, UNIVERSITY OF CALIFORNIA (IRVINE)
ISTEP+: Biology I End-of-Course Assessment Released Items and Scoring Notes
 ISTEP+: Biology I End-of-Course Assessment Released Items and Scoring Notes Page 1 of 22 Introduction Indiana students enrolled in Biology I participated in the ISTEP+: Biology I Graduation Examination
ISTEP+: Biology I End-of-Course Assessment Released Items and Scoring Notes Page 1 of 22 Introduction Indiana students enrolled in Biology I participated in the ISTEP+: Biology I Graduation Examination
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE Q5B
 INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
by the PCR-mediated method (Krawchuk and Wahls, 1999). The construction of Ams2-null and conditional ams2-shut-off strains was previously described
 Table S1. Fission yeast strains used in this study. Gene disruption was performed by the PCR-mediated method (Krawchuk and Wahls, 1999). The construction of Ams2-null and conditional ams2-shut-off strains
Table S1. Fission yeast strains used in this study. Gene disruption was performed by the PCR-mediated method (Krawchuk and Wahls, 1999). The construction of Ams2-null and conditional ams2-shut-off strains
MOL.911 HNL Expression
 1 W I S S E N T E C H N I K L E I D E N S C H A F T MOL.911 HNL Expression www.tugraz.at 2 Hydroxynitrile lyase (Hnl) R 1 HCN R 1 OH R 2 C O R 2 C * CN S selective: Hevea brasiliensis R selective: Prunus
1 W I S S E N T E C H N I K L E I D E N S C H A F T MOL.911 HNL Expression www.tugraz.at 2 Hydroxynitrile lyase (Hnl) R 1 HCN R 1 OH R 2 C O R 2 C * CN S selective: Hevea brasiliensis R selective: Prunus
Expression and Purification of Recombinant Protein in bacteria and Yeast. Presented By: Puspa pandey, Mohit sachdeva & Ming yu
 Expression and Purification of Recombinant Protein in bacteria and Yeast Presented By: Puspa pandey, Mohit sachdeva & Ming yu DNA Vectors Molecular carriers which carry fragments of DNA into host cell.
Expression and Purification of Recombinant Protein in bacteria and Yeast Presented By: Puspa pandey, Mohit sachdeva & Ming yu DNA Vectors Molecular carriers which carry fragments of DNA into host cell.
Methods for Protein Analysis
 Methods for Protein Analysis 1. Protein Separation Methods The following is a quick review of some common methods used for protein separation: SDS-PAGE (SDS-polyacrylamide gel electrophoresis) separates
Methods for Protein Analysis 1. Protein Separation Methods The following is a quick review of some common methods used for protein separation: SDS-PAGE (SDS-polyacrylamide gel electrophoresis) separates
Name Date Period. 2. When a molecule of double-stranded DNA undergoes replication, it results in
 DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
MCAS Biology. Review Packet
 MCAS Biology Review Packet 1 Name Class Date 1. Define organic. THE CHEMISTRY OF LIFE 2. All living things are made up of 6 essential elements: SPONCH. Name the six elements of life. S N P C O H 3. Elements
MCAS Biology Review Packet 1 Name Class Date 1. Define organic. THE CHEMISTRY OF LIFE 2. All living things are made up of 6 essential elements: SPONCH. Name the six elements of life. S N P C O H 3. Elements
Induction of Enzyme Activity in Bacteria:The Lac Operon. Preparation for Laboratory: Web Tutorial - Lac Operon - submit questions
 Induction of Enzyme Activity in Bacteria:The Lac Operon Preparation for Laboratory: Web Tutorial - Lac Operon - submit questions I. Background: For the last week you explored the functioning of the enzyme
Induction of Enzyme Activity in Bacteria:The Lac Operon Preparation for Laboratory: Web Tutorial - Lac Operon - submit questions I. Background: For the last week you explored the functioning of the enzyme
hydrocortisone (5 mg/ml), EGF (10 µg/ml) and Heparin (5000 U/ml). Antibodies against the N-terminal peptide of MEK1 (MEK1-N) and against Flotillin 1
 SUPPLEMENTARY METHODS Cells HUVEC (Human Umbilical Vein Endothelial Cells) were grown in complete Medium 199 (Gibco) supplemented with glutamax, 10% foetal calf serum, BBE (9 mg/ml), hydrocortisone (5
SUPPLEMENTARY METHODS Cells HUVEC (Human Umbilical Vein Endothelial Cells) were grown in complete Medium 199 (Gibco) supplemented with glutamax, 10% foetal calf serum, BBE (9 mg/ml), hydrocortisone (5
European Medicines Agency
 European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
BCOR101 Midterm II Wednesday, October 26, 2005
 BCOR101 Midterm II Wednesday, October 26, 2005 Name Key Please show all of your work. 1. A donor strain is trp+, pro+, met+ and a recipient strain is trp-, pro-, met-. The donor strain is infected with
BCOR101 Midterm II Wednesday, October 26, 2005 Name Key Please show all of your work. 1. A donor strain is trp+, pro+, met+ and a recipient strain is trp-, pro-, met-. The donor strain is infected with
How To Understand How Gene Expression Is Regulated
 What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
GeneCopoeia Genome Editing Tools for Safe Harbor Integration in. Mice and Humans. Ed Davis, Liuqing Qian, Ruiqing li, Junsheng Zhou, and Jinkuo Zhang
 G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
Lecture Series 7. From DNA to Protein. Genotype to Phenotype. Reading Assignments. A. Genes and the Synthesis of Polypeptides
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Chromatin Immunoprecipitation (ChIP)
 Chromatin Immunoprecipitation (ChIP) Day 1 A) DNA shearing 1. Samples Dissect tissue (One Mouse OBs) of interest and transfer to an eppendorf containing 0.5 ml of dissecting media (on ice) or PBS but without
Chromatin Immunoprecipitation (ChIP) Day 1 A) DNA shearing 1. Samples Dissect tissue (One Mouse OBs) of interest and transfer to an eppendorf containing 0.5 ml of dissecting media (on ice) or PBS but without
Hydroxynitrile lyase (Hnl)
 Hydroxynitrile lyase (Hnl) R 1 HCN R 1 OH R 2 C O R 2 C * CN S-selective: Hevea brasiliensis R-selective: Prunus spp. (S)-Hnl of Hevea brasiliensis and (R)-Hnl of Prunus amygdalus Hb_Hnl Type II Hnl intracellular
Hydroxynitrile lyase (Hnl) R 1 HCN R 1 OH R 2 C O R 2 C * CN S-selective: Hevea brasiliensis R-selective: Prunus spp. (S)-Hnl of Hevea brasiliensis and (R)-Hnl of Prunus amygdalus Hb_Hnl Type II Hnl intracellular
Synthetic Genetic Array (SGA) Analysis in Saccharomyces cerevisiae and Schizosaccharomyces pombe
 CHAPTER SEVEN Synthetic Genetic Array (SGA) Analysis in Saccharomyces cerevisiae and Schizosaccharomyces pombe Anastasia Baryshnikova,* Michael Costanzo,* Scott Dixon, Franco J. Vizeacoumar,* Chad L. Myers,
CHAPTER SEVEN Synthetic Genetic Array (SGA) Analysis in Saccharomyces cerevisiae and Schizosaccharomyces pombe Anastasia Baryshnikova,* Michael Costanzo,* Scott Dixon, Franco J. Vizeacoumar,* Chad L. Myers,
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation
 OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
GENOTYPING ASSAYS AT ZIRC
 GENOTYPING ASSAYS AT ZIRC A. READ THIS FIRST - DISCLAIMER Dear ZIRC user, We now provide detailed genotyping protocols for a number of zebrafish lines distributed by ZIRC. These protocols were developed
GENOTYPING ASSAYS AT ZIRC A. READ THIS FIRST - DISCLAIMER Dear ZIRC user, We now provide detailed genotyping protocols for a number of zebrafish lines distributed by ZIRC. These protocols were developed
BCH401G Lecture 39 Andres
 BCH401G Lecture 39 Andres Lecture Summary: Ribosome: Understand its role in translation and differences between translation in prokaryotes and eukaryotes. Translation: Understand the chemistry of this
BCH401G Lecture 39 Andres Lecture Summary: Ribosome: Understand its role in translation and differences between translation in prokaryotes and eukaryotes. Translation: Understand the chemistry of this
Protein Expression. A Practical Approach J. HIGGIN S
 Protein Expression A Practical Approach S. J. HIGGIN S B. D. HAMES List of contributors Abbreviations xv Xvi i 1. Protein expression in mammalian cell s Marlies Otter-Nilsson and Tommy Nilsso n 1. Introduction
Protein Expression A Practical Approach S. J. HIGGIN S B. D. HAMES List of contributors Abbreviations xv Xvi i 1. Protein expression in mammalian cell s Marlies Otter-Nilsson and Tommy Nilsso n 1. Introduction
Essentials of Real Time PCR. About Sequence Detection Chemistries
 Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Lecture 3: Mutations
 Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
Lecture 3: Mutations Recall that the flow of information within a cell involves the transcription of DNA to mrna and the translation of mrna to protein. Recall also, that the flow of information between
Design of conditional gene targeting vectors - a recombineering approach
 Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
Gene Regulation -- The Lac Operon
 Gene Regulation -- The Lac Operon Specific proteins are present in different tissues and some appear only at certain times during development. All cells of a higher organism have the full set of genes:
Gene Regulation -- The Lac Operon Specific proteins are present in different tissues and some appear only at certain times during development. All cells of a higher organism have the full set of genes:
Chapter 5: Organization and Expression of Immunoglobulin Genes
 Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
A. 'Hypersensitive' peptide bonds and autodegradation of proteins
 ABSTRACT A. 'Hypersensitive' peptide bonds and autodegradation of proteins Several pure proteins, which gave a single band on electrophoretic analysis, when stored for a long time, were found to be partially
ABSTRACT A. 'Hypersensitive' peptide bonds and autodegradation of proteins Several pure proteins, which gave a single band on electrophoretic analysis, when stored for a long time, were found to be partially
LECTURE 6 Gene Mutation (Chapter 16.1-16.2)
 LECTURE 6 Gene Mutation (Chapter 16.1-16.2) 1 Mutation: A permanent change in the genetic material that can be passed from parent to offspring. Mutant (genotype): An organism whose DNA differs from the
LECTURE 6 Gene Mutation (Chapter 16.1-16.2) 1 Mutation: A permanent change in the genetic material that can be passed from parent to offspring. Mutant (genotype): An organism whose DNA differs from the
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
Classic Immunoprecipitation
 292PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Classic Immunoprecipitation Utilizes Protein A/G Agarose for Antibody Binding (Cat.
292PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Classic Immunoprecipitation Utilizes Protein A/G Agarose for Antibody Binding (Cat.
CHAPTER 40 The Mechanism of Protein Synthesis
 CHAPTER 40 The Mechanism of Protein Synthesis Problems: 2,3,6,7,9,13,14,15,18,19,20 Initiation: Locating the start codon. Elongation: Reading the codons (5 3 ) and synthesizing protein amino carboxyl.
CHAPTER 40 The Mechanism of Protein Synthesis Problems: 2,3,6,7,9,13,14,15,18,19,20 Initiation: Locating the start codon. Elongation: Reading the codons (5 3 ) and synthesizing protein amino carboxyl.
Bottlenecks in Clinical Source Material Acquisition. Aby J. Mathew, PhD May 5, 2009 ISCT Annual Meeting San Diego, CA amathew@biolifesolutions.
 Bottlenecks in Clinical Source Material Acquisition Aby J. Mathew, PhD May 5, 2009 ISCT Annual Meeting San Diego, CA amathew@biolifesolutions.com Biopreservation What s the issue? Biopreservation considerations
Bottlenecks in Clinical Source Material Acquisition Aby J. Mathew, PhD May 5, 2009 ISCT Annual Meeting San Diego, CA amathew@biolifesolutions.com Biopreservation What s the issue? Biopreservation considerations
Western Blot Analysis with Cell Samples Grown in Channel-µ-Slides
 Western Blot Analysis with Cell Samples Grown in Channel-µ-Slides Polyacrylamide gel electrophoresis (PAGE) and subsequent analyses are common tools in biochemistry and molecular biology. This Application
Western Blot Analysis with Cell Samples Grown in Channel-µ-Slides Polyacrylamide gel electrophoresis (PAGE) and subsequent analyses are common tools in biochemistry and molecular biology. This Application
Chapter 13: Meiosis and Sexual Life Cycles
 Name Period Chapter 13: Meiosis and Sexual Life Cycles Concept 13.1 Offspring acquire genes from parents by inheriting chromosomes 1. Let s begin with a review of several terms that you may already know.
Name Period Chapter 13: Meiosis and Sexual Life Cycles Concept 13.1 Offspring acquire genes from parents by inheriting chromosomes 1. Let s begin with a review of several terms that you may already know.
CD3/TCR stimulation and surface detection Determination of specificity of intracellular detection of IL-7Rα by flow cytometry
 CD3/TCR stimulation and surface detection Stimulation of HPB-ALL cells with the anti-cd3 monoclonal antibody OKT3 was performed as described 3. In brief, antibody-coated plates were prepared by incubating
CD3/TCR stimulation and surface detection Stimulation of HPB-ALL cells with the anti-cd3 monoclonal antibody OKT3 was performed as described 3. In brief, antibody-coated plates were prepared by incubating
Review of the Cell and Its Organelles
 Biology Learning Centre Review of the Cell and Its Organelles Tips for most effective learning of this material: Memorize the names and structures over several days. This will help you retain what you
Biology Learning Centre Review of the Cell and Its Organelles Tips for most effective learning of this material: Memorize the names and structures over several days. This will help you retain what you
RNA & Protein Synthesis
 RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
Modeling and Simulation of Gene Regulatory Networks
 Modeling and Simulation of Gene Regulatory Networks Hidde de Jong INRIA Grenoble - Rhône-Alpes Hidde.de-Jong@inria.fr http://ibis.inrialpes.fr INRIA Grenoble - Rhône-Alpes and IBIS IBIS: systems biology
Modeling and Simulation of Gene Regulatory Networks Hidde de Jong INRIA Grenoble - Rhône-Alpes Hidde.de-Jong@inria.fr http://ibis.inrialpes.fr INRIA Grenoble - Rhône-Alpes and IBIS IBIS: systems biology
Practice Problems 4. (a) 19. (b) 36. (c) 17
 Chapter 10 Practice Problems Practice Problems 4 1. The diploid chromosome number in a variety of chrysanthemum is 18. What would you call varieties with the following chromosome numbers? (a) 19 (b) 36
Chapter 10 Practice Problems Practice Problems 4 1. The diploid chromosome number in a variety of chrysanthemum is 18. What would you call varieties with the following chromosome numbers? (a) 19 (b) 36
Recombinant DNA Technology
 Recombinant DNA Technology Dates in the Development of Gene Cloning: 1965 - plasmids 1967 - ligase 1970 - restriction endonucleases 1972 - first experiments in gene splicing 1974 - worldwide moratorium
Recombinant DNA Technology Dates in the Development of Gene Cloning: 1965 - plasmids 1967 - ligase 1970 - restriction endonucleases 1972 - first experiments in gene splicing 1974 - worldwide moratorium
Lecture 4. Polypeptide Synthesis Overview
 Initiation of Protein Synthesis (4.1) Lecture 4 Polypeptide Synthesis Overview Polypeptide synthesis proceeds sequentially from N Terminus to C terminus. Amino acids are not pre-positioned on a template.
Initiation of Protein Synthesis (4.1) Lecture 4 Polypeptide Synthesis Overview Polypeptide synthesis proceeds sequentially from N Terminus to C terminus. Amino acids are not pre-positioned on a template.
Translation Study Guide
 Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Supplemental Fig. S1. The schematic diagrams of the expression constructs used in this study.
 1 Supplemental data Supplemental Fig. S1. The schematic diagrams of the expression constructs used in this study. Supplemental Fig. S2. Ingenuity Pathway Analysis (IPA) of the 56 putative caspase substrates
1 Supplemental data Supplemental Fig. S1. The schematic diagrams of the expression constructs used in this study. Supplemental Fig. S2. Ingenuity Pathway Analysis (IPA) of the 56 putative caspase substrates
CHAPTER 9 IMMUNOGLOBULIN BIOSYNTHESIS
 CHAPTER 9 IMMUNOGLOBULIN BIOSYNTHESIS Although the process by which a functional gene for immunoglobulin HEAVY and LIGHT CHAINS is formed is highly unusual, the SYNTHESIS, POST- TRANSLATIONAL PROCESSING
CHAPTER 9 IMMUNOGLOBULIN BIOSYNTHESIS Although the process by which a functional gene for immunoglobulin HEAVY and LIGHT CHAINS is formed is highly unusual, the SYNTHESIS, POST- TRANSLATIONAL PROCESSING
1.1.2. thebiotutor. AS Biology OCR. Unit F211: Cells, Exchange & Transport. Module 1.2 Cell Membranes. Notes & Questions.
 thebiotutor AS Biology OCR Unit F211: Cells, Exchange & Transport Module 1.2 Cell Membranes Notes & Questions Andy Todd 1 Outline the roles of membranes within cells and at the surface of cells. The main
thebiotutor AS Biology OCR Unit F211: Cells, Exchange & Transport Module 1.2 Cell Membranes Notes & Questions Andy Todd 1 Outline the roles of membranes within cells and at the surface of cells. The main
mirnaselect pep-mir Cloning and Expression Vector
 Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
Instructions. Torpedo sirna. Material. Important Guidelines. Specifications. Quality Control
 is a is a state of the art transfection reagent, specifically designed for the transfer of sirna and mirna into a variety of eukaryotic cell types. is a state of the art transfection reagent, specifically
is a is a state of the art transfection reagent, specifically designed for the transfer of sirna and mirna into a variety of eukaryotic cell types. is a state of the art transfection reagent, specifically
Given these characteristics of life, which of the following objects is considered a living organism? W. X. Y. Z.
 Cell Structure and Organization 1. All living things must possess certain characteristics. They are all composed of one or more cells. They can grow, reproduce, and pass their genes on to their offspring.
Cell Structure and Organization 1. All living things must possess certain characteristics. They are all composed of one or more cells. They can grow, reproduce, and pass their genes on to their offspring.
Rapid heteromerization and phosphorylation of ligand-activated plant transmembrane receptors and their associated kinase BAK1
 Supporting Online Material for Rapid heteromerization and phosphorylation of ligand-activated plant transmembrane receptors and their associated kinase BAK1 Birgit Schulze, Tobias Mentzel, Anna Jehle,
Supporting Online Material for Rapid heteromerization and phosphorylation of ligand-activated plant transmembrane receptors and their associated kinase BAK1 Birgit Schulze, Tobias Mentzel, Anna Jehle,
ab185915 Protein Sumoylation Assay Ultra Kit
 ab185915 Protein Sumoylation Assay Ultra Kit Instructions for Use For the measuring in vivo protein sumoylation in various samples This product is for research use only and is not intended for diagnostic
ab185915 Protein Sumoylation Assay Ultra Kit Instructions for Use For the measuring in vivo protein sumoylation in various samples This product is for research use only and is not intended for diagnostic
From DNA to Protein. Proteins. Chapter 13. Prokaryotes and Eukaryotes. The Path From Genes to Proteins. All proteins consist of polypeptide chains
 Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
Antibody Function & Structure
 Antibody Function & Structure Specifically bind to antigens in both the recognition phase (cellular receptors) and during the effector phase (synthesis and secretion) of humoral immunity Serology: the
Antibody Function & Structure Specifically bind to antigens in both the recognition phase (cellular receptors) and during the effector phase (synthesis and secretion) of humoral immunity Serology: the
Western Blotting. USA: proteintech@ptglab.com UK & Europe: europe@ptglab.com China: service@ptglab.com. www.ptglab.com
 Western Blotting All steps are carried out at room temperature unless otherwise indicated. Recipes for all solutions highlighted bold are included at the end of the protocol. SDS-PAGE 1. Construct an SDS-PAGE
Western Blotting All steps are carried out at room temperature unless otherwise indicated. Recipes for all solutions highlighted bold are included at the end of the protocol. SDS-PAGE 1. Construct an SDS-PAGE
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
Gene mutation and molecular medicine Chapter 15
 Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
Gene mutation and molecular medicine Chapter 15 Lecture Objectives What Are Mutations? How Are DNA Molecules and Mutations Analyzed? How Do Defective Proteins Lead to Diseases? What DNA Changes Lead to
Structure and Function of DNA
 Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Fighting the Battles: Conducting a Clinical Assay
 Fighting the Battles: Conducting a Clinical Assay 6 Vocabulary: In Vitro: studies in biology that are conducted using components of an organism that have been isolated from their usual biological surroundings
Fighting the Battles: Conducting a Clinical Assay 6 Vocabulary: In Vitro: studies in biology that are conducted using components of an organism that have been isolated from their usual biological surroundings
Concluding lesson. Student manual. What kind of protein are you? (Basic)
 Concluding lesson Student manual What kind of protein are you? (Basic) Part 1 The hereditary material of an organism is stored in a coded way on the DNA. This code consists of four different nucleotides:
Concluding lesson Student manual What kind of protein are you? (Basic) Part 1 The hereditary material of an organism is stored in a coded way on the DNA. This code consists of four different nucleotides:
First Strand cdna Synthesis
 380PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name First Strand cdna Synthesis (Cat. # 786 812) think proteins! think G-Biosciences
380PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name First Strand cdna Synthesis (Cat. # 786 812) think proteins! think G-Biosciences
Mitochondrial DNA Analysis
 Mitochondrial DNA Analysis Lineage Markers Lineage markers are passed down from generation to generation without changing Except for rare mutation events They can help determine the lineage (family tree)
Mitochondrial DNA Analysis Lineage Markers Lineage markers are passed down from generation to generation without changing Except for rare mutation events They can help determine the lineage (family tree)
RNA and Protein Synthesis
 Name lass Date RN and Protein Synthesis Information and Heredity Q: How does information fl ow from DN to RN to direct the synthesis of proteins? 13.1 What is RN? WHT I KNOW SMPLE NSWER: RN is a nucleic
Name lass Date RN and Protein Synthesis Information and Heredity Q: How does information fl ow from DN to RN to direct the synthesis of proteins? 13.1 What is RN? WHT I KNOW SMPLE NSWER: RN is a nucleic
Todd C. Lorenz, Ph.D.
 Todd C. Lorenz, Ph.D. Phone 909.593.3511 1950 3 rd Street, FH 109D tlorenz@laverne.edu La Verne, CA 91750 EDUCATION Ph.D., Biological Chemistry, University of California, Los Angeles (2007) M.S., Chemistry,
Todd C. Lorenz, Ph.D. Phone 909.593.3511 1950 3 rd Street, FH 109D tlorenz@laverne.edu La Verne, CA 91750 EDUCATION Ph.D., Biological Chemistry, University of California, Los Angeles (2007) M.S., Chemistry,
SICKLE CELL ANEMIA & THE HEMOGLOBIN GENE TEACHER S GUIDE
 AP Biology Date SICKLE CELL ANEMIA & THE HEMOGLOBIN GENE TEACHER S GUIDE LEARNING OBJECTIVES Students will gain an appreciation of the physical effects of sickle cell anemia, its prevalence in the population,
AP Biology Date SICKLE CELL ANEMIA & THE HEMOGLOBIN GENE TEACHER S GUIDE LEARNING OBJECTIVES Students will gain an appreciation of the physical effects of sickle cell anemia, its prevalence in the population,
Microarray Technology
 Microarrays And Functional Genomics CPSC265 Matt Hudson Microarray Technology Relatively young technology Usually used like a Northern blot can determine the amount of mrna for a particular gene Except
Microarrays And Functional Genomics CPSC265 Matt Hudson Microarray Technology Relatively young technology Usually used like a Northern blot can determine the amount of mrna for a particular gene Except
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
Bio 102 Practice Problems Genetic Code and Mutation
 Bio 102 Practice Problems Genetic Code and Mutation Multiple choice: Unless otherwise directed, circle the one best answer: 1. Beadle and Tatum mutagenized Neurospora to find strains that required arginine
Bio 102 Practice Problems Genetic Code and Mutation Multiple choice: Unless otherwise directed, circle the one best answer: 1. Beadle and Tatum mutagenized Neurospora to find strains that required arginine
Vpsl0p Cycles between the Late-Golgi and Prevacuolar Compartments in Its Function as the Sorting Receptor for Multiple Yeast Vacuolar Hydrolases
 Vpsl0p Cycles between the Late-Golgi and Prevacuolar Compartments in Its Function as the Sorting Receptor for Multiple Yeast Vacuolar Hydrolases Antony A. Cooper and Tom H. Stevens Institute of Molecular
Vpsl0p Cycles between the Late-Golgi and Prevacuolar Compartments in Its Function as the Sorting Receptor for Multiple Yeast Vacuolar Hydrolases Antony A. Cooper and Tom H. Stevens Institute of Molecular
Cloning GFP into Mammalian cells
 Protocol for Cloning GFP into Mammalian cells Studiepraktik 2013 Molecular Biology and Molecular Medicine Aarhus University Produced by the instructors: Tobias Holm Bønnelykke, Rikke Mouridsen, Steffan
Protocol for Cloning GFP into Mammalian cells Studiepraktik 2013 Molecular Biology and Molecular Medicine Aarhus University Produced by the instructors: Tobias Holm Bønnelykke, Rikke Mouridsen, Steffan
2007 7.013 Problem Set 1 KEY
 2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
Protocol for Western Blotting
 Protocol for Western Blotting Materials Materials used on Day 3 Protease inhibitor stock: 1 μg/μl pepstatin A in DMSO 200 μm leupeptin in OG Buffer 200 mm PMSF: Freshly made. Ex) 34.8 mg PMSF in 1 ml isopropanol
Protocol for Western Blotting Materials Materials used on Day 3 Protease inhibitor stock: 1 μg/μl pepstatin A in DMSO 200 μm leupeptin in OG Buffer 200 mm PMSF: Freshly made. Ex) 34.8 mg PMSF in 1 ml isopropanol
KMS-Specialist & Customized Biosimilar Service
 KMS-Specialist & Customized Biosimilar Service 1. Polyclonal Antibody Development Service KMS offering a variety of Polyclonal Antibody Services to fit your research and production needs. we develop polyclonal
KMS-Specialist & Customized Biosimilar Service 1. Polyclonal Antibody Development Service KMS offering a variety of Polyclonal Antibody Services to fit your research and production needs. we develop polyclonal
Genetic Analysis. Phenotype analysis: biological-biochemical analysis. Genotype analysis: molecular and physical analysis
 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
Rapid GST Inclusion Body Solubilization and Renaturation Kit
 Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
Supplemental Information. McBrayer et al. Supplemental Data
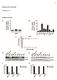 1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
Chapter 18: Applications of Immunology
 Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
they differ from each other when judged in terms of temperature sensitivity,'
 72 GENETICS: LACY AND BONNER.PRoc. N. A. S. 11 Pratt, David, and Gunther S. Stent, these PROCEEDINGS, 45, 1507 (1959). 12 The authors gratefully acknowledge the contributions of Dr. A. W. Kimball of the
72 GENETICS: LACY AND BONNER.PRoc. N. A. S. 11 Pratt, David, and Gunther S. Stent, these PROCEEDINGS, 45, 1507 (1959). 12 The authors gratefully acknowledge the contributions of Dr. A. W. Kimball of the
Chapter 6 DNA Replication
 Chapter 6 DNA Replication Each strand of the DNA double helix contains a sequence of nucleotides that is exactly complementary to the nucleotide sequence of its partner strand. Each strand can therefore
Chapter 6 DNA Replication Each strand of the DNA double helix contains a sequence of nucleotides that is exactly complementary to the nucleotide sequence of its partner strand. Each strand can therefore
Forensic DNA Testing Terminology
 Forensic DNA Testing Terminology ABI 310 Genetic Analyzer a capillary electrophoresis instrument used by forensic DNA laboratories to separate short tandem repeat (STR) loci on the basis of their size.
Forensic DNA Testing Terminology ABI 310 Genetic Analyzer a capillary electrophoresis instrument used by forensic DNA laboratories to separate short tandem repeat (STR) loci on the basis of their size.
Title: Mapping T cell epitopes in PCV2 capsid protein - NPB #08-159. Date Submitted: 12-11-09
 Title: Mapping T cell epitopes in PCV2 capsid protein - NPB #08-159 Investigator: Institution: Carol Wyatt Kansas State University Date Submitted: 12-11-09 Industry summary: Effective circovirus vaccines
Title: Mapping T cell epitopes in PCV2 capsid protein - NPB #08-159 Investigator: Institution: Carol Wyatt Kansas State University Date Submitted: 12-11-09 Industry summary: Effective circovirus vaccines
Notch 1 -dependent regulation of cell fate in colorectal cancer
 Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
Protein Synthesis How Genes Become Constituent Molecules
 Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
The Need for a PARP in vivo Pharmacodynamic Assay
 The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D., Chief Scientific Officer, Trevigen, Inc., Gaithersburg, MD For further infomation, please contact: William Booth, Ph.D. Tel: +44 (0)1235
The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D., Chief Scientific Officer, Trevigen, Inc., Gaithersburg, MD For further infomation, please contact: William Booth, Ph.D. Tel: +44 (0)1235
Benchtop Mitochondria Isolation Protocol
 Benchtop Mitochondria Isolation Protocol Note: Specific protocols are available for the following products: MS850 Mitochondria Isolation Kit for Rodent Tissue MS851 Mitochondria Isolation Kit for Rodent
Benchtop Mitochondria Isolation Protocol Note: Specific protocols are available for the following products: MS850 Mitochondria Isolation Kit for Rodent Tissue MS851 Mitochondria Isolation Kit for Rodent
