Designing an RNAi Experiment using Nucleofection
|
|
|
- Abel Baker
- 7 years ago
- Views:
Transcription
1 Technical Reference Guide Designing an RNAi Experiment using Nucleofection The Nucleofector Technology is well suited for the transfection of sirna duplexes or shrna vectors into both primary cells and difficult-to-transfect cell lines. Choose sirna Select gene target(s) Select control sirna Negative control sirna - e.g. sicontrol Non-Targeting Pool Positive control sirna - e.g. sicontrol GAPDH sirna Choose Cell Type and Transfection Protocol Select cell type(s) to maximize physiological relevance of results Find optimized nucleofection protocols at Select transfection controls Untreated sample (no sirna and transfection) Mock-transfection (no sirna, only transfection) Confirm sirna Delivery Confirm sirna delivery efficiency using: Fluorescently-labeled sirna or Fluorescent expression plasmid (e.g. pmaxgfp TM ) or pmaxgfp TM and maxgfp TM sirna or sirna targeting housekeeping gene Choose Detection Assay Select detection assay(s) mrna - branched-dna, RT-PCR Protein - ELISA, Western, FACS analysis Phenotype - viability, apoptosis Optimize Target Knockdown Determine optimal sirna concentration Nucleofector p pmol (2 nm - 2 µm) 96-well Shuttle p pmol (2 nm - 2 µm) Adapt Assay Conditions Optimize detection assay(s) conditions for specific system Determine optimal cell densities for linear detection range Correlate results from multiple assays Optimize Assay Conditions Perform detection time course or multiple assay mrna p 5-72 hours Protein p hours Confirm specifity of silencing event Confirm gene knockdown results with different sirna reagents If using Standard SMARTpool sirna reagents, follow-up with ON-TARGETplus TM SMARTpool sirna reagents targeting same genes If using individual sigenome sirna, use multiple sirnas targeting same genes If possible, perform rescue experiments amaxa AG, Europe/World amaxa Inc., USA Scientific Support Scientific Support +49 (0) (toll free) scientific-support@amaxa.com scientific-support.us@amaxa.com
2 1. Establish/verify Nucleofection Conditions with pmaxgfp Optimal nucleofection conditions for a particular cell type are identical whether you are transfecting DNA or RNA. We recommend performing a preliminary experiment with pmaxgfp (our positive control plasmid, included in every kit) in order to establish/verify the optimal Nucleofector Solution and program for your cells. Once these conditions have been determined, they remain the same whether you are nucleofecting DNA or RNA (or both together). 2. Identify Appropriate Experimental Controls To ensure that the conclusions drawn from sirna experiments are accurate, it is necessary to include the appropriate experimental controls. We recommend including at least four types of experimental controls in every RNAi experiment. Parallel testing of multiple controls under several conditions can be easily performed using the 96-well Shuttle System. a) Positive sirna control: This should be a validated sirna pool or individual sirna targeting a wellcharacterized house keeping gene, such as cyclophilin B (also known as PPIB), glyceraldehyde- 3-phosphate dehydrogenase (GAPDH), or Lamin. A good positive control reagent targeting a well-expressed but non-essential gene is useful for establishing experimental parameters without affecting cellular viability and can also be used as negative control that is unassociated with any particular pathway under study (i.e., it fails to generate a observable phenotype in the assay being employed). b) Negative sirna control: Negative sirna control reagents are bioinformatically designed and validated to have no known target in the cell line of choice. These reagents are important for distinguishing sequence-specific silencing from sequence-independent effects that are associated with the delivery of sirna into the cell. Such sequence independent effects can include toxicity resulting from the process of transfection in conjunction with nucleic acid delivery or hyper-sensitivity to introduction of double stranded RNA. Investigators are encouraged to test multiple candidates in their own experimental systems to empirically confirm that the negative controls do not result in any observable and unintended off-target effects. For that purpose Dharmacon offers a comprehensive portfolio of multiple negative controls, including the ON-TARGETPLUS Non-Targeting Controls, which have been confirmed by microarray analysis to have little to no off-target signature in HeLa cells. c) Untreated transfection control: The untreated control sample is comprised of cells that have neither been treated with sirna nor subjected to the transfection process. This control serves as an indicator of baseline cellular activity to which all other conditions can be compared. d) Mock-treated control: The mock-treated control sample is one in which the cells are subjected to the transfection procedure in the absence of sirna. In the case of nucleofection, the cells would be exposed to the Nucleofector Solution and subjected to the nucleofection procedure in the absence of sirna. The analysis of page 2 amaxa Leading Transfection Technology
3 mock-treated cells will indicate whether the transfection process results in cytotoxicity or other non-specific effects. 3. Fine Tune Specific sirna Sequence/ concentrations Using the same nucleofection conditions, simply substitute sirna for the pmaxgfp used in the preliminary experiment. We often also include pmaxgfp (either in a separate parallel sample or co-nucleofected with the sirna) as an easy means of comparing relative transfection efficiencies between experiments or selecting transfected cells. The transfection efficiency using DNA is usually substantially lower as compared to sirna. If you are interested in using fluorescently labelled oligos, please first read our suggestions for working with these substrates or contact our Scientific Support Team for specific suggestions. a) Selection of an optimal sirna sequence: If you are using sirna sequences which have not been previously characterized, we recommend investing a considerable amount of time in their selection. The majority of sirna providers offer an oligo optimization service, however, it is still often necessary to test several gene-specific sirna oligos in order to find one which efficiently down-regulates your target gene. b) Determination of the optimal effective sirna concentration: When performing sirna-mediated knockdown experiments it is advisable to conduct a dose-response (concentration) analysis to determine the minimum sirna concentration necessary for sufficient target knockdown on mrna, protein or functional level. For nucleofection, the optimal sirna concentration can range from lower than 2 nm up to 2 µm, depending on multiple factors such as the cell type, and the half-life of the mrna and/or protein of the gene target. To determine the optimal concentration for your cell type and target, we suggest to perform an initial titration of the sirna concentration within the range of 2 nm 2 µm (Nucleofector : pmol in 100 µl; 96-well Shuttle : pmol in 20 µl). Starting concentrations for a minimum titration would be 30 and 300 nm. If looking at concentrations these values may seem higher than with lipid-based methods, but it is important to remember that nucleofection occurs in a 5x to 25x lower volume (20 µl with 96-well Shuttle vs. 100 µl with 96-well lipofection; 100 µl standard Nucleofector vs ml with 6-well lipofection). sirna concentration Cell type Targets/Analysis method Knockdown Reference 2 nm (0.2 pmol*) 7 nm (0.7 pmol*) 1 nm (0.1 pmol*) COS-7 (monkey kidney fibroblast) THP-1 (human monocytic leukaemia) HUVEC (human umbilical vein endothelial cells) Bruton s tyrosine kinase/ FACS Interferon Regulatory Factor (IRF5) / RT-PCR Interferon Integrins 1/ 3 and Akt / Western Blot Migration 96% Lindvall JM et al. (2005) Immunol Rev 203, »strongly reduced«schoenemeyer A et al. (2005) J Biol Chem 280, > 90% Short SM et al. (2005) J Cell Biol 168, * per cuvette (100 ml volume) page 3 amaxa Leading Transfection Technology
4 The optimal effective sirna concentration is dependent on the target and the cell type. Indeed, there are numerous publications in which nucleofection of < 50 nm sirna has been observed to elicit knockdown of the desired genes (see table). Some customers have also reported satisfying results with concentrations higher than 1 µm, but it is important to keep a balance between efficient knockdown and minimizing off-target effects. Although keeping sirnas <30nt avoids activating the protein kinase (PKR) and 2,5 -oligoadenylate synthetase pathways, sirnas have still been demonstrated to elicit non-specific effects, including both stimulation and repression of non-target genes. 4. Determine Optimal Analysis Time Point As the stability and half-life of various mrnas and their protein products varies, it is important to empirically determine the best time points for assessing target knockdown. For example, it has been documented that in mammalian cells, mrna half-life can range from minutes to days while the T1/2 of protein products can range from less than a few minutes to several days. Taking this into consideration, the experimental design should allow sufficient time for the sirna to associate with RISC and deplete mrna/protein concentrations to desired levels. In general, the recommended time course ranges are 5 to 72 hours to deplete target mrna and 24 to 96 hours to adequately knockdown target proteins and assess phenotypic outcomes. 5. Verification of sirna Specificity Keeping sirna concentrations as low as possible helps to minimize non-specific effects, but it is also important to include appropriate controls in all experiments 3. Monitoring gene knockdown at both the mrna and protein levels verifies that the sirna sequence is acting through the»classical«rnai pathway rather than as a microrna (which - at least in part - inhibit translation of target mrna, rather than targeting its destruction). A good way to enhance confidence in RNAi data is to demonstrate a similar effect with two or more sirnas targeted to different sites in the RNA transcript of interest. The rescue experiment as ultimate control: As recently suggested in the Nature Cell Biology editorial, the control of choice for any RNAi experiment is rescue by expression of the target gene in a form refractory to the sirna, usually achieved by utilising one or more silent third-codon point mutations or a deletion in the untranslated region within the targeted sequence. Translational effects can be avoided by using sirnas targeted against 3'-untranslated regions. In practical terms this means either co-transfecting the sirna and a plasmid expressing the sirna-resistant form of the target gene together, or using an shrna-expression vector which co-expresses the sirna-resistant target gene. Fortunately, the ability to transfect DNA and RNA using identical nucleofection conditions means that both of these types of experiments are quite straight-forward and easy to perform using the Nucleofector Technology. 6. Additional Notes: Measuring transfection efficiency: a) Using fluorescently labelled sirna Experiments with fluorescently labeled sirnas have shown transfection efficiencies of up to 99% in some cell types. Unless a confocal microscope or FACS is available, the page 4 amaxa Leading Transfection Technology
5 use of fluorescently labeled sirna for initial set-up experiments is not advisable, as many fluorescent labels fade quickly following nucleofection. Likewise, the amount of fluorescently-labeled sirna needed in order to adequately visualize fluorescent cells is often much higher than would be optimal for functional response, making this both an expensive and not highly informative experiment. Furthermore, microscopic analysis may lead to false positive results as a result of sirna sticking to the membrane and not actually entering the cell. We usually suggest including a sample nucleofecting pmaxgfp in parallel to (or in the same sample as) the sirna in order to provide a general estimate of relative transfection efficiency. Nevertheless it has to be taken into account, the transfection efficiency of sirna molecules is usually much higher. If you wish to use labelled sirna for your experiments, please contact our Scientific Support Team for suggestions to help your experiments run as smoothly as possible. b) sirna Test kit In order to quickly demonstrate general RNAi efficacy for a particular cell type you can use our sirna Test Kit for Cell Lines and Adherent Primary Cells. This provides an sirna against the GFP expressed by our pmaxgfp control and is used in conjunction with the cell-type specific Nucleofector Solution and parameters. For primary human blood cells we recommend examining down-regulation of endogenous genes such as CD2, CD4 and vimentin. An optimized protocol is available which lists the sequences we have used for examining these genes. Enriching for transfected cells One method of enriching for transfected cells is to co-transfect sirna with a plasmid expressing a fluorescent reporter or surface marker, and then sorting for cells expressing the reporter. This approach has been used, for example, in Wu et al. (2005). Using shrna-expressing vectors also allows you to use co-expressed fluorescent or antibiotic resistance markers to select for transfected cells (see below). However, transfection efficiencies for plasmid DNA are generally lower than those for sirna duplexes. Long-term RNAi effects (sirna duplexes vs. shrna-expressing vectors) Chemically synthesized sirna duplexes offer a rapid means for determine the sirna sequences that result in efficient knockdown of your target gene, but this downregulation is transient (generally persisting 2-5 days) and may not be sufficient when silencing targets with low turnover rates or in other applications where a longer duration of effect is required. Consequently a number of different plasmid vectors that express sirna, or shrna (short hairpin RNA), are commercially available. In addition to enabling long-term expression of sirna/shrna, these vectors have the advantages that they can be grown, handled and stored as plasmid DNA, co-express fluorescent markers or antibiotic resistance genes (facilitating identification of transfected cells/selection of stably transfected cells) and can be engineered with inducible promoters to permit switching the knockdown phenotype on and off (such as psuperior). The ability of the Nucleofector Technology to transfect DNA into primary cells (and many cell lines which are difficult or page 5 amaxa Leading Transfection Technology
6 impossible to transfect by other means) makes it possible to now use these vectors in virtually any cell type. Although do keep in mind that transfection efficiencies with sirna oligos are generally higher than with plasmid DNA. Stability of sirna duplexes in the Nucleofector Solutions The Nucleofector Solutions were tested for RNAse activity. Incubation of RNA in the solutions for 2 hours at 37 C did not affect RNA stability. High sirna transfection efficiencies can only be achieved with the amaxa certified cuvettes, included in the cell-type specific kits. For more information (or if you are experiencing any difficulties), please contact our Scientific Support Team. sirna Amount-Concentration Comparison Chart Amount Weight Concentration Molecular weight of a 21 bp Standard Nucleofector (100 µl) sirna ds-oligonucleotide: 21 x 660 g/mol = g/mol = ng/pmol 14 ng/pmol Concentration 96-well Shuttle (20 µl) 1 pmol 14 ng 1 pmol/100 µl = 10 nmol/l = 10 nm 1 pmol/20 µl = 50 nmol/l = 50 nm 5 pmol 69 ng 5 pmol/100 µl = 50 nmol/l = 50 nm 5 pmol/20 µl = 250 nmol/l = 250 nm 10 pmol 140 ng 10 pmol/100 µl = 100 nmol/l = 100 nm 10 pmol/20 µl = 500 nmol/l = 500 nm 20 pmol 277 ng 20 pmol/100 µl = 200 nmol/l = 200 nm 20 pmol/20 µl = 1000 nmol/l = 1 µm 50 pmol 690 ng 50 pmol/100 µl = 500 nmol/l = 500 nm 50 pmol/20 µl = 2500 nmol/l = 2.5 µm 100 pmol 1.4 µg 100 pmol/100 µl = 1000 nmol/l= 1 µm 100 pmol/20 µl = 5000 nmol/l = 5 µm For individual calculation also refer to our website sirna calculator ( References 1 Persengiev SP, et al. RNA Jan;10(1): Ross J, 1995, Microbiol Rev 59: Editorial (2003) Wither RNAi. Nat Cell Biol. 5 (6), Wu et al. (2005) MCB 25(22): WTC-1012_ Further Material Technote»Efficient Delivery of Dharmacon SMARTpool sirna Reagents in Hard-To-Transfect Cell Lines Using amaxa Nucleofector 96-well Shuttle System«(WTA-1003) The Nucleofector Technology, comprising Nucleofection Process, Nucleofector Device, Nucleofector Solutions, Nucleofector 96-well Shuttle System and 96-well Nucleocuvette plates and modules are covered by patent and/or patent pending rights owned by amaxa AG. amaxa, HiFect, Nucleofector, nucleofection, 96-well Shuttle, maxfp and maxgfp are either registered trademarks or trademarks of amaxa AG in the U.S. and/or Germany and/or other countries. Dharmacon, SMARTpool and ON-TARGET PLUS are trademarks of Dharmacon, Inc. All rights reserved, amaxa AG, page 6 amaxa Leading Transfection Technology
TOOLS sirna and mirna. User guide
 TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
TOOLS sirna and mirna User guide Introduction RNA interference (RNAi) is a powerful tool for suppression gene expression by causing the destruction of specific mrna molecules. Small Interfering RNAs (sirnas)
Optimized Protocol sirna Test Kit for Cell Lines and Adherent Primary Cells
 page 1 of 8 sirna Test Kit for Cell Lines and Adherent Primary Cells Kit principle Co-transfection of pmaxgfp, encoding the green fluorescent protein (GFP) from Pontellina p. with an sirna directed against
page 1 of 8 sirna Test Kit for Cell Lines and Adherent Primary Cells Kit principle Co-transfection of pmaxgfp, encoding the green fluorescent protein (GFP) from Pontellina p. with an sirna directed against
Instructions. Torpedo sirna. Material. Important Guidelines. Specifications. Quality Control
 is a is a state of the art transfection reagent, specifically designed for the transfer of sirna and mirna into a variety of eukaryotic cell types. is a state of the art transfection reagent, specifically
is a is a state of the art transfection reagent, specifically designed for the transfer of sirna and mirna into a variety of eukaryotic cell types. is a state of the art transfection reagent, specifically
岑 祥 股 份 有 限 公 司 技 術 專 員 費 軫 尹 20100803
 技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
Thermo Scientific Dharmacon sirna Libraries
 Thermo Scientific Dharmacon sirna Libraries Scientific innovation in sirna potency, specificity and delivery Superior sirna in comprehensive collections for reliable screens Market leading products for
Thermo Scientific Dharmacon sirna Libraries Scientific innovation in sirna potency, specificity and delivery Superior sirna in comprehensive collections for reliable screens Market leading products for
Amaxa Mouse T Cell Nucleofector Kit
 Amaxa Mouse T Cell Nucleofector Kit For T cells isolated from C57BL/6 & BALB/c mice Evaluated for murine T cells isolated from C57BL/6 & BALB/c mice This protocol is designed for murine lymphocytes or
Amaxa Mouse T Cell Nucleofector Kit For T cells isolated from C57BL/6 & BALB/c mice Evaluated for murine T cells isolated from C57BL/6 & BALB/c mice This protocol is designed for murine lymphocytes or
Manual for: sirna-trans Maximo Reagent
 Manual for: sirna-trans Maximo Reagent Contact: info@geneon.net; www.geneon.net GeneON GmbH Germany GeneON GmbH / Protocol sirna-trans reagent vs. 1.3 / www.geneon.net / - 0 - sirna-trans reagent Instruction
Manual for: sirna-trans Maximo Reagent Contact: info@geneon.net; www.geneon.net GeneON GmbH Germany GeneON GmbH / Protocol sirna-trans reagent vs. 1.3 / www.geneon.net / - 0 - sirna-trans reagent Instruction
HiPerFect Transfection Reagent Handbook
 Fifth Edition October 2010 HiPerFect Transfection Reagent Handbook For transfection of eukaryotic cells with sirna and mirna Sample & Assay Technologies QIAGEN Sample and Assay Technologies QIAGEN is the
Fifth Edition October 2010 HiPerFect Transfection Reagent Handbook For transfection of eukaryotic cells with sirna and mirna Sample & Assay Technologies QIAGEN Sample and Assay Technologies QIAGEN is the
Thermo Scientific Dharmacon ON-TARGETplus sirna The Standard for sirna Specificity
 Thermo Scientific Dharmacon sirna The Standard for sirna Specificity Reduce off-targets up to 90% with the market-leading sirna Combine higher target specificity with guaranteed silencing Increase confidence
Thermo Scientific Dharmacon sirna The Standard for sirna Specificity Reduce off-targets up to 90% with the market-leading sirna Combine higher target specificity with guaranteed silencing Increase confidence
MEF Nucleofector Kit 1 and 2
 page 1 of 7 MEF Nucleofector Kit 1 and 2 for Mouse Embryonic Fibroblasts (MEF) MEF display significant phenotypic variations which depend on the strain, the genetic background of the they are isolated
page 1 of 7 MEF Nucleofector Kit 1 and 2 for Mouse Embryonic Fibroblasts (MEF) MEF display significant phenotypic variations which depend on the strain, the genetic background of the they are isolated
Mouse ES Cell Nucleofector Kit
 page 1 of 7 Mouse ES Cell Nucleofector Kit for Mouse Embryonic Stem Cells Cell type Origin Cells derived from mouse blastocysts. Morphology Round cells growing in clumps. Important remarks! 1. This protocol
page 1 of 7 Mouse ES Cell Nucleofector Kit for Mouse Embryonic Stem Cells Cell type Origin Cells derived from mouse blastocysts. Morphology Round cells growing in clumps. Important remarks! 1. This protocol
MEF Starter Nucleofector Kit
 page 1 of 7 MEF Starter Nucleofector Kit for Mouse Embryonic Fibroblasts (MEF) MEF display significant phenotypic variations which depend on the strain, the genetic background of the mice they are isolated
page 1 of 7 MEF Starter Nucleofector Kit for Mouse Embryonic Fibroblasts (MEF) MEF display significant phenotypic variations which depend on the strain, the genetic background of the mice they are isolated
RNA Interference. - Localization - Labeling - Expression Profiling - In Vivo Delivery - Transfection
 RNA Interference - Localization - Labeling - Expression Profiling - In Vivo Delivery - Transfection Rna interference x RNAi Pathways RNA interference x Tools and Multiplexing RNAi Tools pri-mirna Application
RNA Interference - Localization - Labeling - Expression Profiling - In Vivo Delivery - Transfection Rna interference x RNAi Pathways RNA interference x Tools and Multiplexing RNAi Tools pri-mirna Application
INTERFERin in vitro sirna/mirna transfection reagent PROTOCOL. 1 Standard sirna transfection of adherent cells... 2
 INTERFERin in vitro sirna/mirna transfection reagent PROTOCOL DESCRIPTION INTERFERin is a powerful sirna/mirna transfection reagent that ensures efficient gene silencing and reproducible transfection in
INTERFERin in vitro sirna/mirna transfection reagent PROTOCOL DESCRIPTION INTERFERin is a powerful sirna/mirna transfection reagent that ensures efficient gene silencing and reproducible transfection in
TransIT -2020 Transfection Reagent
 Quick Reference Protocol, MSDS and Certificate of Analysis available at mirusbio.com/5400 INTRODUCTION TransIT -2020 Transfection Reagent is a high-performance, animal-origin free, broad spectrum reagent
Quick Reference Protocol, MSDS and Certificate of Analysis available at mirusbio.com/5400 INTRODUCTION TransIT -2020 Transfection Reagent is a high-performance, animal-origin free, broad spectrum reagent
2.1.2 Characterization of antiviral effect of cytokine expression on HBV replication in transduced mouse hepatocytes line
 i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
QIAGEN Transfection Technologies
 QIAGEN Transfection Technologies Efficient and robust transfection for all your applications Sample & Assay Technologies Table of contents QIAGEN solutions for efficient transfection Transfection is a
QIAGEN Transfection Technologies Efficient and robust transfection for all your applications Sample & Assay Technologies Table of contents QIAGEN solutions for efficient transfection Transfection is a
MicroRNA Mike needs help to degrade all the mrna transcripts! Aaron Arvey ISMB 2010
 Target mrna abundance dilutes microrna and sirna activity MicroRNA Mike needs help to degrade all the mrna transcripts! Aaron Arvey ISMB 2010 Target mrna abundance dilutes microrna and sirna activity Erik
Target mrna abundance dilutes microrna and sirna activity MicroRNA Mike needs help to degrade all the mrna transcripts! Aaron Arvey ISMB 2010 Target mrna abundance dilutes microrna and sirna activity Erik
MicroRNA formation. 4th International Symposium on Non-Surgical Contraceptive Methods of Pet Population Control
 MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
MicroRNA formation mirna s are processed from several precursor stages Mammalian genomes seem to have 100 s of mirna s Nucleotides in positions 2-8 of an mirna are considered the mirna seed 5 Methyl-G
sirna Duplexes & RNAi Explorer
 P r o d u c t S p e c i f i c a t i o n s Guaranteed RNAi Explorer kit with Fl/Dabcyl Molecular Beacon Catalog No.:27-6402-01 Guaranteed RNAi Explorer kit with Fluorescein/Tamra TaqMan Catalog No.:27-6402-01
P r o d u c t S p e c i f i c a t i o n s Guaranteed RNAi Explorer kit with Fl/Dabcyl Molecular Beacon Catalog No.:27-6402-01 Guaranteed RNAi Explorer kit with Fluorescein/Tamra TaqMan Catalog No.:27-6402-01
PRODUCT INFORMATION...
 VIROMER RED In vitro plasmid DNA and mrna Standard Transfection PRODUCT INFORMATION... 2 GENERAL... 2 RED OR YELLOW?... 3 PROTOCOL GUIDELINES... 4 GENERAL REMARKS... 4 CELL CULTURE AND PLATING... 4 FORWARD/REVERSE
VIROMER RED In vitro plasmid DNA and mrna Standard Transfection PRODUCT INFORMATION... 2 GENERAL... 2 RED OR YELLOW?... 3 PROTOCOL GUIDELINES... 4 GENERAL REMARKS... 4 CELL CULTURE AND PLATING... 4 FORWARD/REVERSE
Functional Analysis. mircury LNA microrna Mimics
 Functional Analysis mircury LNA microrna Mimics Instruction manual v1.0 April 2014 Table of Contents Product summary......................................................... 3 Content..................................................................
Functional Analysis mircury LNA microrna Mimics Instruction manual v1.0 April 2014 Table of Contents Product summary......................................................... 3 Content..................................................................
Thermo Scientific DharmaFECT Transfection Reagents
 Thermo Scientific Transfection Reagents Maintain cell viability with lipids specially formulated to deliver sirna Achieve robust sirna uptake for dependable gene silencing Expand your RNAi applications
Thermo Scientific Transfection Reagents Maintain cell viability with lipids specially formulated to deliver sirna Achieve robust sirna uptake for dependable gene silencing Expand your RNAi applications
Product: Expression Arrest TM egfp control shrna vector
 Product: Expression Arrest TM egfp control vector Catalog #: RHS1702 Product Description The laboratory of Dr. Greg Hannon at Cold Spring Harbor Laboratory (CSHL) has created an RNAi Clone Library comprised
Product: Expression Arrest TM egfp control vector Catalog #: RHS1702 Product Description The laboratory of Dr. Greg Hannon at Cold Spring Harbor Laboratory (CSHL) has created an RNAi Clone Library comprised
REAL TIME PCR USING SYBR GREEN
 REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
Dicer-Substrate sirna Technology
 As featured in BioRadiations 120 Dicer-Substrate sirna Technology Advances in sirna Designs Improve Gene-Specific Silencing Steve Kulisch, Teresa Esch, Christina Whitman-Guliaev, and Teresa Rubio, Bio-Rad
As featured in BioRadiations 120 Dicer-Substrate sirna Technology Advances in sirna Designs Improve Gene-Specific Silencing Steve Kulisch, Teresa Esch, Christina Whitman-Guliaev, and Teresa Rubio, Bio-Rad
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation
 OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
Supplemental Information. McBrayer et al. Supplemental Data
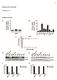 1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
Transfection reagent for visualizing lipofection with DNA. For ordering information, MSDS, publications and application notes see www.biontex.
 METAFECTENE FluoR Transfection reagent for visualizing lipofection with DNA For ordering information, MSDS, publications and application notes see www.biontex.com Description Cat. No. Size METAFECTENE
METAFECTENE FluoR Transfection reagent for visualizing lipofection with DNA For ordering information, MSDS, publications and application notes see www.biontex.com Description Cat. No. Size METAFECTENE
Supplementary Figure 1.
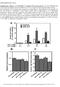 Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Transfection-Transfer of non-viral genetic material into eukaryotic cells. Infection/ Transduction- Transfer of viral genetic material into cells.
 Transfection Key words: Transient transfection, Stable transfection, transfection methods, vector, plasmid, origin of replication, reporter gene/ protein, cloning site, promoter and enhancer, signal peptide,
Transfection Key words: Transient transfection, Stable transfection, transfection methods, vector, plasmid, origin of replication, reporter gene/ protein, cloning site, promoter and enhancer, signal peptide,
Relative Quantification of mirna Target mrnas by Real-Time qpcr. 1 Introduction. Gene Expression Application Note No. 4
 Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
Gene Expression Application Note No. 4 Relative Quantification of mirna Target mrnas by Real-Time qpcr Ute Ernst, Jitao David Zhang, Anja Irsigler, Stefan Wiemann, Ulrich Tschulena Division: Molecular
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE Q5B
 INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
Guideline for Generation of Stable Cell Lines Technical Reference Guide
 BioResearch Guideline for Generation of Stable Cell Lines Technical Reference Guide 1. Background Stable, long-term expression of a gene of interest can be either achieved by eukaryotic vectors that harbor
BioResearch Guideline for Generation of Stable Cell Lines Technical Reference Guide 1. Background Stable, long-term expression of a gene of interest can be either achieved by eukaryotic vectors that harbor
SureSilencing shrna Plasmids
 User Manual BIOMOL GmbH Waidmannstr. 35 22769 Hamburg info@biomol.de www.biomol.de Phone:+49-40-8532600 or 0800-2466651 (D) Fax: +49-40-85326022 or 0800-2466652 (D) SureSilencing shrna Plasmids Genome-Wide
User Manual BIOMOL GmbH Waidmannstr. 35 22769 Hamburg info@biomol.de www.biomol.de Phone:+49-40-8532600 or 0800-2466651 (D) Fax: +49-40-85326022 or 0800-2466652 (D) SureSilencing shrna Plasmids Genome-Wide
Especially developed for the transfection of mammalian cells with sirna or mirna
 METAFECTENE SI Especially developed for the transfection of mammalian cells with sirna or mirna For ordering information, MSDS, publications and application notes see www.biontex.com Description Cat. No.
METAFECTENE SI Especially developed for the transfection of mammalian cells with sirna or mirna For ordering information, MSDS, publications and application notes see www.biontex.com Description Cat. No.
Understanding the immune response to bacterial infections
 Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Essentials of Real Time PCR. About Sequence Detection Chemistries
 Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Recombinant DNA and Biotechnology
 Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
European Medicines Agency
 European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
Notch 1 -dependent regulation of cell fate in colorectal cancer
 Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
RNAi. Martin Latterich
 RNAi Martin Latterich Contributors Abbreviations Preface i x ( xi xiii 1 Methods in RNA interference 1 Martin Latterich and Dalia Halawan i References 2 2 RNAi reagent design 3 Bernd Jagla and Nathalie
RNAi Martin Latterich Contributors Abbreviations Preface i x ( xi xiii 1 Methods in RNA interference 1 Martin Latterich and Dalia Halawan i References 2 2 RNAi reagent design 3 Bernd Jagla and Nathalie
Innate Immunity. Insects rely solely on an innate immune system for defense against infection
 Innate Immunity Insects rely solely on an innate immune system for defense against infection The importance of mammalian innate immunity has only recently become appreciated The innate immune response
Innate Immunity Insects rely solely on an innate immune system for defense against infection The importance of mammalian innate immunity has only recently become appreciated The innate immune response
Lezioni Dipartimento di Oncologia Farmacologia Molecolare. RNA interference. Giovanna Damia 29 maggio 2006
 Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Cell Cycle Phase Determination Kit
 Cell Cycle Phase Determination Kit Item No. 10009349 Customer Service 800.364.9897 * Technical Support 888.526.5351 www.caymanchem.com TABLE OF CONTENTS GENERAL INFORMATION 3 Materials Supplied 3 Safety
Cell Cycle Phase Determination Kit Item No. 10009349 Customer Service 800.364.9897 * Technical Support 888.526.5351 www.caymanchem.com TABLE OF CONTENTS GENERAL INFORMATION 3 Materials Supplied 3 Safety
Amaxa Mouse/Rat Hepatocyte Nucleofector Kit
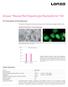 Amaxa Mouse/Rat Hepatocyte Nucleofector Kit For Primary Mouse and Rat Hepatocytes This protocol is designed for freshly isolated mouse and rat hepatocytes; polygonal, adherent cells Example for Nucleofection
Amaxa Mouse/Rat Hepatocyte Nucleofector Kit For Primary Mouse and Rat Hepatocytes This protocol is designed for freshly isolated mouse and rat hepatocytes; polygonal, adherent cells Example for Nucleofection
Name Class Date. Figure 13 1. 2. Which nucleotide in Figure 13 1 indicates the nucleic acid above is RNA? a. uracil c. cytosine b. guanine d.
 13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
Actions of Hormones on Target Cells Page 1. Actions of Hormones on Target Cells Page 2. Goals/ What You Need to Know Goals What You Need to Know
 Actions of Hormones on Target Cells Graphics are used with permission of: Pearson Education Inc., publishing as Benjamin Cummings (http://www.aw-bc.com) Page 1. Actions of Hormones on Target Cells Hormones
Actions of Hormones on Target Cells Graphics are used with permission of: Pearson Education Inc., publishing as Benjamin Cummings (http://www.aw-bc.com) Page 1. Actions of Hormones on Target Cells Hormones
Application Note No. 2 / July 2012. Quantitative Assessment of Cell Quality, Viability and Proliferation. System
 Application Note No. 2 / July 2012 Quantitative Assessment of Cell Quality, Viability and Proliferation System Quantitative Assessment of Cell Quality, Viability and Proliferation Introduction In vitro
Application Note No. 2 / July 2012 Quantitative Assessment of Cell Quality, Viability and Proliferation System Quantitative Assessment of Cell Quality, Viability and Proliferation Introduction In vitro
ncounter Leukemia Fusion Gene Expression Assay Molecules That Count Product Highlights ncounter Leukemia Fusion Gene Expression Assay Overview
 ncounter Leukemia Fusion Gene Expression Assay Product Highlights Simultaneous detection and quantification of 25 fusion gene isoforms and 23 additional mrnas related to leukemia Compatible with a variety
ncounter Leukemia Fusion Gene Expression Assay Product Highlights Simultaneous detection and quantification of 25 fusion gene isoforms and 23 additional mrnas related to leukemia Compatible with a variety
CODICE DESCRIZIONE QUANTITA' PREZZO EURO
 Broad Spectrum Plasmid DNA and sirna/mirna Transfection MIR 6000 TransIT-X2 Dynamic Delivery System 1.5 ml 823,35 MIR 6003 TransIT-X2 Dynamic Delivery System 0.3 ml 196,35 MIR 6004 TransIT-X2 Dynamic Delivery
Broad Spectrum Plasmid DNA and sirna/mirna Transfection MIR 6000 TransIT-X2 Dynamic Delivery System 1.5 ml 823,35 MIR 6003 TransIT-X2 Dynamic Delivery System 0.3 ml 196,35 MIR 6004 TransIT-X2 Dynamic Delivery
GenScript sirna Cassette Protocol
 GenScript sirna Cassette Protocol Technical Manual No. 0168 Version 20040510 I Introduction.... 1 II sirna Cassette.... 1 III Product Description. 2 IV Transfecting Mammalian Cells. 3 V References... 5
GenScript sirna Cassette Protocol Technical Manual No. 0168 Version 20040510 I Introduction.... 1 II sirna Cassette.... 1 III Product Description. 2 IV Transfecting Mammalian Cells. 3 V References... 5
pcas-guide System Validation in Genome Editing
 pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
Dicer Substrate RNAi Design
 INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
Technical Note: Analysis of microrna function using mircury LNA TM microrna Inhibitors
 Technical note: Analysis of microrna function using mircury LNA microrna Inhibitors Introduction The study of micrornas has come a long way since the discovery of these small RNAs some fifteen years ago.
Technical note: Analysis of microrna function using mircury LNA microrna Inhibitors Introduction The study of micrornas has come a long way since the discovery of these small RNAs some fifteen years ago.
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual
 Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Selection of Cell Lines for Manufacturing Therapeutic Antibodies by Flow Cytometric Cell Sorting. Andrew Racher Lonza Biologics
 Selection of Cell Lines for Manufacturing Therapeutic Antibodies by Flow Cytometric Cell Sorting Andrew Racher Lonza Biologics Structure of Talk 1. The Problem 2. Possible Solutions 3. Solution Chosen
Selection of Cell Lines for Manufacturing Therapeutic Antibodies by Flow Cytometric Cell Sorting Andrew Racher Lonza Biologics Structure of Talk 1. The Problem 2. Possible Solutions 3. Solution Chosen
Microarray Technology
 Microarrays And Functional Genomics CPSC265 Matt Hudson Microarray Technology Relatively young technology Usually used like a Northern blot can determine the amount of mrna for a particular gene Except
Microarrays And Functional Genomics CPSC265 Matt Hudson Microarray Technology Relatively young technology Usually used like a Northern blot can determine the amount of mrna for a particular gene Except
1-7 2-8 3-9 4-10 5-11 6-12 7-13 8-14 9-15. Corresponding position of sirna GS
 A C GS Target transcript 5ʼ- - 3ʼ 3ʼ- 21 20 19 18 17 16 15 14 13 12 11 10 9 8 7 6 5 4 3 2 1-5ʼ 10-16 11-17 12-18 13-19 14-20 15-21 sivim-270 3ʼUTR 1-7 2-8 3-9 4-10 5-11 6-12 7-13 8-14 9-15 Corresponding
A C GS Target transcript 5ʼ- - 3ʼ 3ʼ- 21 20 19 18 17 16 15 14 13 12 11 10 9 8 7 6 5 4 3 2 1-5ʼ 10-16 11-17 12-18 13-19 14-20 15-21 sivim-270 3ʼUTR 1-7 2-8 3-9 4-10 5-11 6-12 7-13 8-14 9-15 Corresponding
mirnaselect pep-mir Cloning and Expression Vector
 Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES
 C GUGAAG EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES MicroRNA-adapted shrna (shrna mir ) for increased, specific and consistent knockdown. MicroRNA PROCESSING PATHWAY UTILIZED FOR shrna mir Developed
C GUGAAG EXPRESSION ARREST shrna mir GENOME- WIDE LIBRARIES MicroRNA-adapted shrna (shrna mir ) for increased, specific and consistent knockdown. MicroRNA PROCESSING PATHWAY UTILIZED FOR shrna mir Developed
A role of microrna in the regulation of telomerase? Yuan Ming Yeh, Pei Rong Huang, and Tzu Chien V. Wang
 A role of microrna in the regulation of telomerase? Yuan Ming Yeh, Pei Rong Huang, and Tzu Chien V. Wang Department of Molecular and Cellular Biology, Chang Gung University, Kwei San, Tao Yuan 333, Taiwan
A role of microrna in the regulation of telomerase? Yuan Ming Yeh, Pei Rong Huang, and Tzu Chien V. Wang Department of Molecular and Cellular Biology, Chang Gung University, Kwei San, Tao Yuan 333, Taiwan
Amaxa 4D-Nucleofector Protocol for Mouse Embryonic Stem [ES] Cells For 4D-Nucleofector X Unit Transfection in suspension
![Amaxa 4D-Nucleofector Protocol for Mouse Embryonic Stem [ES] Cells For 4D-Nucleofector X Unit Transfection in suspension Amaxa 4D-Nucleofector Protocol for Mouse Embryonic Stem [ES] Cells For 4D-Nucleofector X Unit Transfection in suspension](/thumbs/30/14702458.jpg) For 4D-Nucleofector X Unit Transfection in suspension Cells derived from mouse blastocysts; round cells, growing in clumps s 1. This protocol is meant to provide an outline for the handling and the Nucleofection
For 4D-Nucleofector X Unit Transfection in suspension Cells derived from mouse blastocysts; round cells, growing in clumps s 1. This protocol is meant to provide an outline for the handling and the Nucleofection
Fast, easy and effective transfection reagent for mammalian cells
 METAFECTENE EASY + Fast, easy and effective transfection reagent for mammalian cells For ordering information, MSDS, publications and application notes see www.biontex.com Description Cat. No. Size METAFECTENE
METAFECTENE EASY + Fast, easy and effective transfection reagent for mammalian cells For ordering information, MSDS, publications and application notes see www.biontex.com Description Cat. No. Size METAFECTENE
SmartFlare RNA Detection Probes: Principles, protocols and troubleshooting
 Technical Guide SmartFlare RNA Detection Probes: Principles, protocols and troubleshooting Principles of SmartFlare technology RNA detection traditionally requires transfection, laborious sample prep,
Technical Guide SmartFlare RNA Detection Probes: Principles, protocols and troubleshooting Principles of SmartFlare technology RNA detection traditionally requires transfection, laborious sample prep,
Functional and Biomedical Aspects of Genome Research
 Functional and Biomedical Aspects of Genome Research 20 11 35 Vorlesung SS 04 Bartsch, Jockusch & Schmitt-John Mi. 9:15-10:00, in W7-135 13 Functional RNAs Thomas Schmitt-John micro RNAs small interfering
Functional and Biomedical Aspects of Genome Research 20 11 35 Vorlesung SS 04 Bartsch, Jockusch & Schmitt-John Mi. 9:15-10:00, in W7-135 13 Functional RNAs Thomas Schmitt-John micro RNAs small interfering
Head of College Scholars List Scheme. Summer Studentship. Report Form
 Head of College Scholars List Scheme Summer Studentship Report Form This report should be completed by the student with his/her project supervisor. It should summarise the work undertaken during the project
Head of College Scholars List Scheme Summer Studentship Report Form This report should be completed by the student with his/her project supervisor. It should summarise the work undertaken during the project
User Manual/Hand book. qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse)
 User Manual/Hand book qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse) Kit Components Cat. No. MA003...Human Whole Genome mirna qpcr Profiling Kit (-20 C) The following components are sufficient
User Manual/Hand book qpcr mirna Arrays ABM catalog # MA003 (human) and MA004 (mouse) Kit Components Cat. No. MA003...Human Whole Genome mirna qpcr Profiling Kit (-20 C) The following components are sufficient
Validating Microarray Data Using RT 2 Real-Time PCR Products
 Validating Microarray Data Using RT 2 Real-Time PCR Products Introduction: Real-time PCR monitors the amount of amplicon as the reaction occurs. Usually, the amount of product is directly related to the
Validating Microarray Data Using RT 2 Real-Time PCR Products Introduction: Real-time PCR monitors the amount of amplicon as the reaction occurs. Usually, the amount of product is directly related to the
How To Get A Cell Print
 QUICK CELL CAPTURE AND CHARACTERIZATION GUIDE FOR CELLSEARCH CUSTOMERS CellSave EDTA Blood sample Rare cell capture Enumeration Single protein marker Cell capture for molecular characterization CELLSEARCH
QUICK CELL CAPTURE AND CHARACTERIZATION GUIDE FOR CELLSEARCH CUSTOMERS CellSave EDTA Blood sample Rare cell capture Enumeration Single protein marker Cell capture for molecular characterization CELLSEARCH
RNA & Protein Synthesis
 RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
MAB Solut. MABSolys Génopole Campus 1 5 rue Henri Desbruères 91030 Evry Cedex. www.mabsolut.com. is involved at each stage of your project
 Mabsolus-2015-UK:Mise en page 1 03/07/15 14:13 Page1 Services provider Department of MABSolys from conception to validation MAB Solut is involved at each stage of your project Creation of antibodies Production
Mabsolus-2015-UK:Mise en page 1 03/07/15 14:13 Page1 Services provider Department of MABSolys from conception to validation MAB Solut is involved at each stage of your project Creation of antibodies Production
Analysis of gene expression data. Ulf Leser and Philippe Thomas
 Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
Analysis of gene expression data Ulf Leser and Philippe Thomas This Lecture Protein synthesis Microarray Idea Technologies Applications Problems Quality control Normalization Analysis next week! Ulf Leser:
Effects of Antibiotics on Bacterial Growth and Protein Synthesis: Student Laboratory Manual
 Effects of Antibiotics on Bacterial Growth and Protein Synthesis: Student Laboratory Manual I. Purpose...1 II. Introduction...1 III. Inhibition of Bacterial Growth Protocol...2 IV. Inhibition of in vitro
Effects of Antibiotics on Bacterial Growth and Protein Synthesis: Student Laboratory Manual I. Purpose...1 II. Introduction...1 III. Inhibition of Bacterial Growth Protocol...2 IV. Inhibition of in vitro
Inhibition of MicroRNA Sensitizes 3D Breast Cancer Microtissues to Radiation Therapy
 CASE STUDY High Content Image Analysis Inhibition of MicroRNA Sensitizes 3D Breast Cancer Microtissues to Radiation Therapy Dr. Nataša Anastasov leads the Personalized Radiation Therapy Group within the
CASE STUDY High Content Image Analysis Inhibition of MicroRNA Sensitizes 3D Breast Cancer Microtissues to Radiation Therapy Dr. Nataša Anastasov leads the Personalized Radiation Therapy Group within the
J. Cell Sci. 128: doi:10.1242/jcs.164566: Supplementary Material
 Figure S1. Microtubule and microfilament drug treatments severely interfere with mitotic spindle morphology, length and positioning. (A) GFP-fused lamin-a/c, LAP2 or BAF1 localizes to the spindle and all
Figure S1. Microtubule and microfilament drug treatments severely interfere with mitotic spindle morphology, length and positioning. (A) GFP-fused lamin-a/c, LAP2 or BAF1 localizes to the spindle and all
Molecular Genetics. RNA, Transcription, & Protein Synthesis
 Molecular Genetics RNA, Transcription, & Protein Synthesis Section 1 RNA AND TRANSCRIPTION Objectives Describe the primary functions of RNA Identify how RNA differs from DNA Describe the structure and
Molecular Genetics RNA, Transcription, & Protein Synthesis Section 1 RNA AND TRANSCRIPTION Objectives Describe the primary functions of RNA Identify how RNA differs from DNA Describe the structure and
Machine Learning Approaches to sirna Efficacy Prediction
 Machine Learning Approaches to sirna Efficacy Prediction by Sahar Abubucker B.E., Madras University, 2000 THESIS Submitted in Partial Fulfillment of the Requirements for the Degree of Master of Science
Machine Learning Approaches to sirna Efficacy Prediction by Sahar Abubucker B.E., Madras University, 2000 THESIS Submitted in Partial Fulfillment of the Requirements for the Degree of Master of Science
Gene Expression Assays
 APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
CD3/TCR stimulation and surface detection Determination of specificity of intracellular detection of IL-7Rα by flow cytometry
 CD3/TCR stimulation and surface detection Stimulation of HPB-ALL cells with the anti-cd3 monoclonal antibody OKT3 was performed as described 3. In brief, antibody-coated plates were prepared by incubating
CD3/TCR stimulation and surface detection Stimulation of HPB-ALL cells with the anti-cd3 monoclonal antibody OKT3 was performed as described 3. In brief, antibody-coated plates were prepared by incubating
Real-time PCR: Understanding C t
 APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
PreciseTM Whitepaper
 Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
RNAi Shooting the Messenger!
 RNAi Shooting the Messenger! Bronya Keats, Ph.D. Department of Genetics Louisiana State University Health Sciences Center New Orleans Email: bkeats@lsuhsc.edu RNA interference (RNAi) A mechanism by which
RNAi Shooting the Messenger! Bronya Keats, Ph.D. Department of Genetics Louisiana State University Health Sciences Center New Orleans Email: bkeats@lsuhsc.edu RNA interference (RNAi) A mechanism by which
HCS604.03 Exercise 1 Dr. Jones Spring 2005. Recombinant DNA (Molecular Cloning) exercise:
 HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
p21 Cell cycle arrest / genetic repair
 Application Note Silencing of expression with Thermo Scientific Dharmacon Accell sirna causes inhibition of the DNA damage response in IMR-32 neuroblastoma cells and protects primary cortical neurons from
Application Note Silencing of expression with Thermo Scientific Dharmacon Accell sirna causes inhibition of the DNA damage response in IMR-32 neuroblastoma cells and protects primary cortical neurons from
Proto col. GoClone Repor ter Construc ts: Sample Protocol for Adherent Cells. Tech support: 877-994-8240. Luciferase Assay System
 Luciferase Assay System Proto col GoClone Repor ter Construc ts: Sample Protocol for Adherent Cells LightSwitch Luciferase Assay System GoClone Reporter Assay Workflow Step 1: Seed cells in plate format.
Luciferase Assay System Proto col GoClone Repor ter Construc ts: Sample Protocol for Adherent Cells LightSwitch Luciferase Assay System GoClone Reporter Assay Workflow Step 1: Seed cells in plate format.
Transcription and Translation of DNA
 Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved
 microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
microrna 2 micrornas Non protein coding, endogenous RNAs of 21-22nt length Evolutionarily conserved Regulate gene expression by binding complementary regions at 3 regions of target mrnas Act as negative
Introduction To Real Time Quantitative PCR (qpcr)
 Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Cloning GFP into Mammalian cells
 Protocol for Cloning GFP into Mammalian cells Studiepraktik 2013 Molecular Biology and Molecular Medicine Aarhus University Produced by the instructors: Tobias Holm Bønnelykke, Rikke Mouridsen, Steffan
Protocol for Cloning GFP into Mammalian cells Studiepraktik 2013 Molecular Biology and Molecular Medicine Aarhus University Produced by the instructors: Tobias Holm Bønnelykke, Rikke Mouridsen, Steffan
SUPPLEMENTARY DATA 1
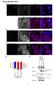 SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
Green Fluorescent Protein (GFP): Genetic Transformation, Synthesis and Purification of the Recombinant Protein
 Green Fluorescent Protein (GFP): Genetic Transformation, Synthesis and Purification of the Recombinant Protein INTRODUCTION Green Fluorescent Protein (GFP) is a novel protein produced by the bioluminescent
Green Fluorescent Protein (GFP): Genetic Transformation, Synthesis and Purification of the Recombinant Protein INTRODUCTION Green Fluorescent Protein (GFP) is a novel protein produced by the bioluminescent
INTERPRETATION INFORMATION SHEET
 Creative Testing Solutions 2424 West Erie Dr. 2205 Highway 121 10100 Martin Luther King Jr. St. No. Tempe, AZ 85282 Bedford, TX 76021 St. Petersburg, FL 33716 INTERPRETATION INFORMATION SHEET Human Immunodeficiency
Creative Testing Solutions 2424 West Erie Dr. 2205 Highway 121 10100 Martin Luther King Jr. St. No. Tempe, AZ 85282 Bedford, TX 76021 St. Petersburg, FL 33716 INTERPRETATION INFORMATION SHEET Human Immunodeficiency
LightSwitch Luciferase Assay System
 Constructs, Assays & Services Assay System Promoter GoClones Synthetic Response Elements Pathway Cell Lines 3 UTR GoClones mirna Mimics & Inhibitors Synthetic mirna Targets Transfection Reagents Assay
Constructs, Assays & Services Assay System Promoter GoClones Synthetic Response Elements Pathway Cell Lines 3 UTR GoClones mirna Mimics & Inhibitors Synthetic mirna Targets Transfection Reagents Assay
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
GeneCopoeia Genome Editing Tools for Safe Harbor Integration in. Mice and Humans. Ed Davis, Liuqing Qian, Ruiqing li, Junsheng Zhou, and Jinkuo Zhang
 G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
Suppl. Figure 1. Cells cultured in oxldl take up Oil Red O in a time-dependent
 Suppl. Figure 1. Cells cultured in oxldl take up Oil Red O in a time-dependent manner. Uptake of Oil Red O was performed as detailed in Materials and Methods and absorbance of samples was expressed as
Suppl. Figure 1. Cells cultured in oxldl take up Oil Red O in a time-dependent manner. Uptake of Oil Red O was performed as detailed in Materials and Methods and absorbance of samples was expressed as
Amaxa Nucleofector 96-well Shuttle System Manual
 Amaxa Nucleofector 96-well Shuttle System Manual For future protocol updates, also check www.lonza.com/cell-database 96-well Shuttle CD-Rom 1 Contents 1.0 The Amaxa Nucleofector Technology 3 2.0 The Nucleofector
Amaxa Nucleofector 96-well Shuttle System Manual For future protocol updates, also check www.lonza.com/cell-database 96-well Shuttle CD-Rom 1 Contents 1.0 The Amaxa Nucleofector Technology 3 2.0 The Nucleofector
G E N OM I C S S E RV I C ES
 GENOMICS SERVICES THE NEW YORK GENOME CENTER NYGC is an independent non-profit implementing advanced genomic research to improve diagnosis and treatment of serious diseases. capabilities. N E X T- G E
GENOMICS SERVICES THE NEW YORK GENOME CENTER NYGC is an independent non-profit implementing advanced genomic research to improve diagnosis and treatment of serious diseases. capabilities. N E X T- G E
Micro RNAs: potentielle Biomarker für das. Blutspenderscreening
 Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
