ARTICLE IN PRESS European Journal of Cell Biology xxx (2010) xxx xxx
|
|
|
- Jocelyn Singleton
- 8 years ago
- Views:
Transcription
1 European Journal of Cell Biology xxx (2010) xxx xxx Contents lists available at ScienceDirect European Journal of Cell Biology journal homepage: Initial receptor ligand interactions modulate gene expression and phagosomal properties during both early and late stages of phagocytosis Eik Hoffmann a,b,,1, Sabrina Marion b,2, Bibhuti Bhusan Mishra c,3, Mathias John d, Ramona Kratzke a, Syed Furquan Ahmad a, Daniela Holzer b, Paras Kumar Anand b, Dieter G. Weiss a, Gareth Griffiths b,4, Sergei A. Kuznetsov a, a Institute of Biological Sciences, Cell Biology and Biosystems Technology Unit, University of Rostock, Germany b European Molecular Biology Laboratory (EMBL), Cell Biology and Biophysics Unit, Heidelberg, Germany c Molecular Pathogenesis Center, Faculty of Pharmacy, University of Lisbon, Portugal d Department of Computer Science, University of Rostock, Germany article info abstract Article history: Received 8 February 2010 Received in revised form 27 April 2010 Accepted 29 April 2010 Keywords: Phagocytosis Receptor ligand interaction Macrophage Phagosome maturation Phagocytic receptor Phagosomal proteome Gene expression Signaling pathways Immune response The receptors engaged during recognition and phagocytic uptake of microorganisms and particles influence signaling events and diverse subcellular responses that occur during phagosome formation and maturation. However, pathogens generally have multiple ligands on their surface, making it difficult to dissect the roles of individual receptors during phagocytosis. Moreover, it remains elusive to which extent receptor ligand interactions and early binding events define the subsequent intracellular fate of phagosomes. Here, we used latex beads coupled to single ligands, focusing on immunoglobulin G, mannan, bacterial lipopolysaccharides and avidin, and monitored: (1) phagocytic uptake rates, (2) fusion of phagosomes with lysosomal compartments, (3) the gene expression profile during phagocytosis, (4) the protein composition of mature phagosomes and (5) time-dependent dynamics of protein association with phagosomes in J774.A1 mouse macrophages. The differently coated latex beads were internalized at different rates and exhibited different kinetics of phagolysosomal fusion events dependent on their specific ligand. Furthermore, less than 60% of identified phagosomal proteins and only 10 15% of changes in gene expression were common to all investigated ligands. These findings demonstrate that each single ligand induced a distinct pattern of genes and a different protein composition of phagosomes. Taken together, our data argue that phagocytic receptor-specific programs of signaling events direct phagosomes to different physiological states and support the existence of a specific receptor ligand signature during the whole process of phagocytosis Elsevier GmbH. All rights reserved. Introduction Corresponding author at: Institute Curie, INSERM U932, Immunity and Cancer, 26 rue d Ulm, Paris Cedex 05, France. Tel.: ; fax: Corresponding author at: University of Rostock, Institute of Biological Sciences, Albert-Einstein-Str. 3, Rostock, Germany. Tel.: ; fax: addresses: eik.hoffmann@curie.fr (E. Hoffmann), sergei.kuznetsov@unirostock.de (S.A. Kuznetsov). 1 Present address: Institute Curie, INSERM U932 - Immunity and Cancer, Paris, France. 2 Present address: Institute Cochin, University Paris Descartes, CNRS UMR 8104, Paris, France. 3 Present address: Department of Molecular Genetics and Microbiology, University of Massachusetts Medical School, Worcester, MA, USA. 4 Present address: Department of Molecular Biosciences, University of Oslo, Norway. Phagocytosis is a process initiated by binding of a particle usually bigger than 0.5 m in diameter to the cell surface of phagocytes followed by its ingestion into a newly formed vesicle, called the phagosome. When extracellular ligands bind to cell surface receptors they induce a receptor-specific program of signaling events, which present a particular time scheme. A well-characterized phagocytic receptor such as the Fc receptor (Fc R), which is engaged during the uptake of particles that have been opsonized with immunoglobulin G (IgG), can be cited as an example. When these particles bind to Fc R, they change the cytoplasmic domains of the receptor, which recruit and activate proteins within seconds initiating immediate responses, such as receptor tyrosine phosphorylation, PI3K/Akt and MAPK activation (Cox and Greenberg, 2001; Swanson and Hoppe, 2004). These signals induce actin polymerization and rapid extension of membrane over the opsonized surface, which engage other IgG molecules that /$ see front matter 2010 Elsevier GmbH. All rights reserved. doi: /j.ejcb
2 2 E. Hoffmann et al. / European Journal of Cell Biology xxx (2010) xxx xxx induce similar responses closing the phagocytic cup and forming the phagosome within a few minutes (Swanson, 2008). At a later stage, a subsequent wave of signaling activates transcription factors that turn on the expression of a receptor-specific set of genes. Therefore, even when internalization is accomplished, a specific signaling program can run for many hours or even days to modulate specific intracellular functions (Ernst, 2000; Aderem, 2003). A major challenge for the innate immune system is to discriminate between a large number of potential pathogens from self antigens, utilizing a finite number of phagocytic receptors. This challenge has been solved by the evolution of a variety of receptors that recognize conserved motifs on pathogens that are not found in higher eukaryotes (Kabelitz and Medzhitov, 2007). These receptors are called pattern-recognition receptors (PRR) and the targets of these receptors pathogen-associated molecular patterns (PAMP; Janeway, 1992). Pathogen-associated motifs include mannans in the yeast cell wall, formylated peptides, lipopolysaccharides and lipoteichoic acids on the surface of Gramnegative and Gram-positive bacteria, respectively. While direct pathogen recognition is a fundamental aspect of innate immunity, opsonization allows additional diversification of the phagocyte recognition repertoire (Stuart and Ezekowitz, 2005). Receptors bearing immunoreceptor tyrosine-based activation motifs (ITAM) or inhibitory motifs (ITIM) are crucial in the immune response (Pasquier et al., 2005). In macrophages, receptors are classified into opsonic phagocytic receptors, like members of the FcR family and complement receptors (CR), and non-opsonic phagocytic receptors, such as lectins (mannose receptor, MR; DC-SIGN, Dectin-1), integrins and scavenger receptors (SR). Many non-opsonic receptors were believed to be unable to induce inflammatory responses on their own, but some C-type lectins, such as Dectin-1 and Mincle, are able to induce gene expression in the absence of other PRR signaling (Brown, 2006; Geijtenbeek and Gringhuis, 2009). A third group contains Toll-like receptors (TLR) which often function as co-receptors in phagocytosis by their discrimination of a broad range of microbial products, including PAMP, lipopolysaccharide, peptidoglycan and bacterial lipopeptides (Underhill et al., 1999; Underhill and Ozinsky, 2002). The role of stimulated TLR in accelerating and modulating phagosome maturation and the hypothesis of TLR-mediated phagosomal autonomy are still matter of debate (Blander and Medzhitov, 2006; Russell and Yates, 2007). Finally, during phagocytosis the activation of macrophages by either pro-inflammatory or anti-inflammatory cytokines contributes to triggering, instructing and terminating the immune response (Martinez et al., 2008). As mentioned before, receptors on the surface of the phagocytic cell orchestrate a set of signaling events that are required for particle internalization. This has also been elegantly demonstrated in non-phagocytic cells, such as CHO cells, which can selectively phagocytose IgG-coated particles after transfection with constructs encoding Fc receptors (Nagarajan et al., 1995). This shows that a single type of ligand on the particle can efficiently induce phagocytic uptake. However, most pathogens possess many different ligands on their surface. Their phagocytic uptake occurs via multi-ligand interactions, which induce the engagement of many receptors at the same time. This makes it very difficult to dissect the contribution of different receptor ligand interactions into the overall process of phagocytosis and in phagosome maturation. Until now, it is unclear whether multiple receptors can signal in parallel in response to phagocytosis or whether a single receptor will dominate the signaling pathways. Here, we introduce a system that can eventually be adapted to answer such questions. We used one of the best-characterized model systems in phagocytosis studies, the latex bead phagosome system (Desjardins and Griffiths, 2003; Griffiths and Mayorga, 2007), and coated these latex beads with single ligands for a global comparison of the characteristics of receptor-mediated uptake pathways involved in phagocytosis. Latex bead phagosomes (LBP), like phagosomes generally, are able to undergo fission and fusion events with endosomes and lysosomes over long time periods (Desjardins et al., 1994; Kjeken et al., 2004), acquire lipids and phosphoinositides (Defacque et al., 2002; Corrotte et al., 2006), bind and move along microtubules (Blocker et al., 1997), assemble actin filaments and bind to them (Jahraus et al., 2001; Al-Haddad et al., 2001) and become acidified (Huynh and Grinstein, 2007). In this study, latex beads were covalently conjugated with the Fc fragment of IgG, which is recognized by Fc receptors (Swanson and Hoppe, 2004) that are expressed in high abundance on the macrophage surface and facilitate the internalization of opsonized targets. For a bead ligand entering macrophages in a non-opsonic pathway, we selected mannan that is recognized predominantly by the mannose receptor (MR; Sheng et al., 2006) and by the mannosebinding lectin (MBL; Weis et al., 1992). Latex beads were also coated with bacterial lipopolysaccharides (LPS), which are recognized by CD14 and TLR4 in macrophages (Wright et al., 1990; Poltorak et al., 1998). The fourth ligand we have selected is avidin, which has been widely used as a non-specfic ligand in phagocytosis studies (Brown et al., 1998; Jahraus et al., 1998, 2001). Avidin is believed to engage multi-ligand receptors, such as members of the scavenger receptor family and integrins (Taylor et al., 2005). Similar to other non-specific bead ligands, such as BSA, the intracellular fate of phagosomes containing this bead type is different compared to Fc R-mediated uptake (Link et al., 2010). Since the applied latex beads are identical for each ligand, we were able to compare cellular functions that are dependent on the particular ligand coupled to the bead. Moreover, LBP can be isolated by sucrose gradient flotation at a very high degree of purity (>95%) allowing one to dissect the molecular composition of these organelles (Desjardins and Griffiths, 2003; Stuart et al., 2007). In this study, our goal was to monitor downstream cellular events of phagocytosis induced by different types of interaction between a single ligand with its receptor over a broad time scale. To achieve this goal, we investigated (1) phagocytic uptake efficiency, (2) phagosomal fusion with lysosomes by confocal microscopy, (3) macrophage gene expression by a microarray approach, (4) the protein composition of phagosomes by mass spectrometry and (5) time-dependent dynamics of association of defined proteins by immunoblotting. All the results obtained indicate that the choice of a particular ligand on the bead has a significant influence on the mentioned processes and strongly supports the notion that each receptor ligand interaction induces its own signaling signature that influences the pattern of macrophage gene expression and the cellular fate of phagosomes. Materials and methods Cells, latex beads and antibodies J774.A1 mouse macrophage-like cells were obtained from the German Resource Center of Biological Material (DSMZ) Braunschweig, Germany, and were cultivated in DMEM (Invitrogen, UK) supplemented with 10% fetal bovine serum (Biochrom, Germany) at 37 C and 5% CO 2. Fluorescent carboxylated latex beads (Invitrogen, USA) and non-fluorescent carboxylated beads (Polysciences, USA), both of 1 m diameter, were conjugated to avidin (Invitrogen, USA), Fc fragment of mouse IgG (Thermo Fisher Scientific, USA), LPS from Klebsiella pneumoniae and mannan from Saccharomyces cerevisae (Sigma Aldrich, USA) as described as supplemental data accompanying this study. Primary antibodies used were anti-actin (clone C4, ICN Biomedicals, USA), anti-lamp-2 (Iowa Hybridoma Bank, USA) and anti-cathepsin-d (clone C-20, Santa Cruz, USA). Secondary anti-
3 E. Hoffmann et al. / European Journal of Cell Biology xxx (2010) xxx xxx 3 bodies were purchased from Dianova, Germany, and Invitrogen, USA. Immunofluorescence microscopy and analysis of phagocytic uptake rates, inhibition of uptake and phagosomal maturation J774.A1 cells were incubated with serum-free DMEM containing 0.01% latex beads (OD 600 = 1.6) for different periods of time (0, 5, 15, 30, 45 and 60 min). Cells were fixed in PBS containing 4% paraformaldehyde and 4% sucrose, ph 7.4 for 20 min followed by quenching in PBS containing 50 mm NH 4 Cl for 10 min, both at room temperature. Cells were permeabilized with 0.2% Triton X- 100 in PBS for 5 min. After blocking of non-specific binding sites with PBS containing 1% gelatin gold (Sigma Aldrich, USA) for 1 h, samples were labeled for F-actin by FITC-conjugated phalloidin (Sigma Aldrich, USA) and for the lysosomal enzyme LAMP-2. In all samples, 50 cells were analyzed for total numbers of internalized beads and the percentage of phagosomes that were positive for LAMP-2 in three independent experiments using a confocal laserscanning microscope (TCS SP2, Leica Microsystems, Germany). Alternatively, uptake of avidin beads and IgG beads was also analyzed by an inside outside staining technique. Macrophages were pre-loaded with rhodamine-conjugated nano gold of 10 nm, a fluid-phase marker that accumulates in lysosomes (Anes et al., 2006). Cells were incubated with beads and fixed as described above. Subsequently, the samples were labeled for 15 min with Alexa 405-conjugated biotin or anti-mouse IgG (Invitrogen, USA) to stain the remaining extracellular beads. After permeabilization, the samples were labeled with Alexa 488-conjugated biotin or antimouse IgG (Invitrogen, USA) to stain all beads. Analysis was carried out with a confocal microscope quantifying beads labeled by Alexa 405 and Alexa 488 (extracellular beads), by Alexa 488 alone (phagosomes before fusion with lysosomal compartments) or by Alexa 488 and rhodamine (phagosomes after fusion with lysosomal compartments). In the histogram shown as supplementary figure S1, internalized beads of 50 analyzed cells for each ligand and time point are shown. For uptake inhibition studies (shown as supplementary figure S2), cells were pre-incubated with the respective ligand in soluble form for 30 min followed by addition of the beads in presence of the soluble ligand. Increasing concentrations of the soluble ligand were tested for inhibition of uptake efficiency compared to control conditions (same bead type in absence of soluble ligand). Average data of three experiments analyzing 50 cells each are shown. Isolation of phagosomes For 2D gel electrophoresis and LC MS/MS, cells were incubated with serum-free DMEM containing 10 mm HEPES, ph 7.4, and 0.02% latex beads coupled to avidin, IgG Fc, mannan and LPS (OD 600 = 3.2) on a shaker for 1 h (pulse) followed by brief washing in PBS and further incubation in complete medium for 1 h (chase). For SDS- PAGE analysis of phagosomes, cells were incubated similarly for 30 min or 1 h pulse followed by a period of 1 or 3 h chase in some experiments. Phagosomes containing latex beads were isolated by sucrose gradient flotation as described elsewhere (Kühnel et al., 2006). Microarray hybridization All gene expression studies with each condition and time point were carried out in triplicates. Cells were seeded into 6-well plates 2 days prior the experiment and were grown as monolayer until they reached 75% confluency. Cells were incubated with serumfree DMEM containing 0.02% latex beads coupled to avidin, IgG Fc, LPS and mannan (OD 600 = 3.2) for 30 and 60 min. Control cells were incubated in serum-free DMEM alone. Subsequently, cells were washed three times in PBS and total RNA was isolated using the RNAeasy Mini kit (Qiagen, Germany) followed by quality control tests using a Bioanalyzer (Agilent Technologies, USA). The preparation of crna and hybridization was carried out using the CodeLink iexpress expression assay reagent kit (Applied Microarrays Inc., USA) following the manufacturers instructions. Subsequently, hybridization to CodeLink mouse whole genome bioarrays (Applied Microarrays Inc., USA) containing 36,000 mouse gene targets was performed. Ten micrograms of crna and 5 l of5 fragmentation buffer were incubated at 94 C for 20 min. Ten micrograms of fragmented crna, 78 l of hybridization buffer component A, 130 l of component B to a final volume of 260 l were incubated at 90 C for 5 min to denature the samples followed by immediate cooling on ice for 30 min to sustain annealing. The hybridization reaction mixture was loaded onto the array input port and sealed with strips to avoid dehydration. The slides were incubated for 20hat37 C on a shaker at 300 rpm. Then, bioarrays were transferred to a bioarray rack containing 0.75 TNT (0.1 M Tris HCl, 0.15 M NaCl and 0.05% Tween 20) followed by an incubation at 46 C for 1 h. Each bioarray was transferred to a chamber containing 3.4 ml of streptavidin-cy5 solution and incubated at room temperature for 30 min. The arrays were washed four times with 1 TNT followed by a brief rinse with 0.1 SSC (0.05% Tween 20). Bioarrays were dried by centrifugation at 9,000 rpm for 10 min and scanned with a GenePix 4000B (Molecular Devices, USA). The complete triplicate data set has been deposited in the GEO database with the accession number GSE17761 using the following URL: Analysis of microarray data The scanned files were analyzed using the CodeLink expression analysis software EXP 4.0. The intensity of each spot was calculated as the difference between mean signal intensity and mean local background intensity. The output CodeLink files were analyzed using GeneSpring GX 7.3 software (Agilent Technologies, USA). Analysis of fold changes in signal intensities of the latex bead samples compared to control conditions were performed using two-way ANOVA with a P-value cut-off <0.01. Genes that exhibited a change in their expression in all three replicates displaying 1.4-fold higher signal intensities were considered significantly upregulated, whereas the ones with 0.6-fold lower signal intensities were considered significantly down-regulated. For analyzing gene regulation data, we used the two-tone table lens (TTTL) visualization tool (John et al., 2008). For gene annotation enrichment analysis we were using the freely accessible database for annotation, visualization and integrated discovery (DAVID; Huang et al., 2009). Functional classification of genes was performed with gene ontology (GO) terms for the biological process using an enrichment P-value cut-off < Two-dimensional SDS-PAGE All chemicals were from Merck, USA, unless otherwise stated. Two-dimensional gel electrophoresis was performed according to the procedures reviewed previously (Görg et al., 2000). The same amount of phagosomes containing latex beads coupled to the Fc fragment of mouse IgG and mannan, respectively, isolated after 2 h, were used. After lysis in 2% Triton X-100, 10 mm DTT, 50 mm Tris, ph 8.0, and protease inhibitors, phagosomal proteins were precipitated in acetone. Precipitates were solubilized in 9 M urea, 16 mm CHAPS, 2 mm TBP, 0.25% Biolyte (BioRad, USA), protease inhibitor cocktail and applied to 11 cm IPG strips (BioRad, USA) at ph 3 10 with rehydration loading overnight as described elsewhere (Bjellqvist et al., 1982). The samples were
4 4 E. Hoffmann et al. / European Journal of Cell Biology xxx (2010) xxx xxx mixed with rehydration solution (8 M urea, 16 mm CHAPS, 2 mm TBP and 0.25% Biolyte) giving a final volume of 200 l. Isoelectric focusing was carried out using a Protean IEF Cell (BioRad, USA) for about 80 kv/h at a maximum voltage of 6,000 V at 20 C. After focusing, each IPG strip was emerged first for 10 min into reduction solution (6 M urea, 30% glycerol, 50 mm Tris, ph 8.8, 173 mm SDS and bromophenolblue) containing 5 mm TBP and then into alkylation solution containing 270 mm 2-iodoacetamide. Subsequently, the IPG strips were loaded on top of 12% SDS polyacrylamide gels in a Protean II xi 2-DE device (BioRad, USA). The gels were run for 45 min at 20 ma per gel and then for 600 V/h at 15 ma/gel in a second step. Visualization was achieved by using the silver staining method according to Shevchenko et al. (1996). Two-dimensional migration of phagosomal proteins was carried out in triplicates and representative silver-stained gels are shown. Liquid chromatography tandem mass spectrometry (LC MS/MS) Purified phagosomes were lysed in 2% Triton X-100, 10 mm DTT, 50 mm Tris, ph 8.0, and protease inhibitor cocktail (Roche, Switzerland) at 4 C for 15 min. The lysate was separated from the beads and then precipitated in acetone for 2 h at 20 C. The pellet was dissolved, reduced with DTT and alkylated with iodoacetamide before being digested with 1 g of Promega-modified trypsin for 24 h at room temperature. A second 1 g of trypsin was added and digestion was allowed to proceed for 24 more hours. The sample was acidified to 5% acetic acid and then desalted on a C18 column. The LC MS/MS procedure was performed by the W.M. Keck Biomedical Mass Spectrometry Laboratory (contact: Dr. Nicholas Sherman, University of Virginia, Charlottesville, USA). Approximately 25% of the digest was introduced into the mass spectrometer for analysis using two run conditions. The LC MS system consisted of a Finnigan LCQ ion trap mass spectrometer system (Thermo Fisher Scientific, USA) with a Protana nanospray ion source interfaced to a self-packed 8 cm 75 m id Phenomenex Jupiter 10 m C18 reversed-phase capillary column. The extracts ( l volumes) were injected and the peptides were eluted from the column by an acetonitrile/0.1 M acetic acid gradient at a flow rate of 0.25 l/min. The nanospray ion source was operated at 2.8 kv. The digests were analyzed using the double play capability of the instrument acquiring full scan mass spectra to determine peptide molecular weights and product ion spectra to determine amino acid sequence in sequential scans. This mode of analysis produces approximately 3000 CAD spectra of ions ranging in abundance over several orders of magnitude. The data were analyzed by database searching using the Sequest search algorithm against mouse IPI accessions and the publicly available UniProtKB/Swiss-Prot database (version 57.13, released January 19, 2010). Identified proteins of one sample per bead ligand were analyzed using Scaffold 2 software (Proteome Software, USA) applying a minimum protein identification probability of 99% with 2 or more unique peptides per identified protein. SDS-PAGE and Western blotting Isolated phagosomes were adjusted to similar concentrations by measuring the optical density at 600 nm (OD 600 ). SDS lysates of phagosome samples were separated by vertical gel electrophoresis using 10% polyacrylamide gels according to the method of Laemmli (1970). After separation, proteins were transferred onto nitrocellulose membranes (Whatman, USA) at constant 30 V overnight using a tank blotting device (BioRad, USA), blocked in 5% non-fat dry milk (Nestlé Co., USA) and labeled according to the method of Towbin et al. (1979). Chemiluminescent signals were developed and visualized on Kodak MR films (Sigma Aldrich, USA). Results Receptor ligand interactions modulate phagocytic particle uptake We used latex beads covalently coated with different ligands to trigger the specific uptake via different receptors. Coating of latex beads with the Fc fragment of mouse IgG (subsequently referred to as IgG Fc beads) induced Fc receptor-mediated uptake, with mannan (mannan beads) induced an internalization predominantly via the mannose receptor, whereas latex beads coated with LPS (LPS beads) entered cells via established CD14- and TLR4-dependent pathways. These specific internalization routes were compared to the non-selective uptake of latex beads coupled to avidin (subsequently referred to as avidin beads). First, we estimated the internalization rates of latex beads coated to different ligands over time. J774.A1 mouse macrophages were incubated with the same numbers of 1 m latex beads coated with IgG Fc, mannan, LPS and avidin and were fixed at different time points. Internalization was studied on cross-sections of images taken by confocal microscopy using FITC-labelled phalloidin to stain the actin cortex in order to confirm full particle enclosure (Fig. 1B). Latex beads only bound to the surface of macrophages or incompletely taken up were not included in this analysis. We observed that all ligands induced an almost linear increase of bead uptake over a time course of 60 min (Fig. 1A), but the rates of uptake were significantly different and dependent on the type of ligand. Mannan beads were internalized at the highest rate and to the highest extent, followed by LPS beads and IgG Fc beads (Fig. 1A). The entry of avidin beads displayed the lowest uptake rate (Fig. 1A). We validated our approach analyzing engulfed beads by labeled actin using an alternative method applying an inside outside staining technique. Macrophages were incubated with IgG Fc beads and avidin beads over a time course of 60 min, respectively. Also by this method, IgG Fc beads were internalized more efficiently compared to avidin beads (Fig. S1, supplementary data) at rates comparable to our previous approach labeling the actin cortex. When avidin beads were applied at 5-times higher concentration, J774.A1 cells internalized these beads much more efficiently (Fig. S1, supplementary data) demonstrating that no defects in phagocytic uptake occurred when lower concentrations were used in previous experiments. In order to confirm that the different ligands are recognized by their expected receptor, competition experiments were performed where cells were first pre-incubated with increasing concentrations of the ligand in soluble form for 30 min followed by a co-incubation of ligand-coated beads together with the soluble ligand (Fig. S2, supplementary data). We found that different concentrations of a given soluble ligand efficiently competed for binding of the same ligand-coated bead at the cell surface. This resulted in a decrease of the corresponding bead uptake to less than 36% of the phagocytosis rate of control conditions for IgG Fc, 31% for mannan, 20% for LPS and 23% for avidin (Fig. S2, supplementary data). In contrast, when cells were co-incubated with beads together with another ligand than the one present on the bead surface, phagocytic uptake rates were not significantly affected (data not shown). This proves that all the studied ligands bound to the latex beads interacted specifically to their expected receptors. Receptor ligand interactions influence phagosome maturation The findings described above showed that phagocytic uptake is regulated efficiently by the specific receptor ligand interaction. This led us to study whether subsequent stages of the phagocytic process are also influenced by the early step of receptor ligand interactions. After engulfment, particles enclosed in phagosomes undergo sequential fusion and fission events with endosomal and
5 E. Hoffmann et al. / European Journal of Cell Biology xxx (2010) xxx xxx 5 amount of phagosomes that acquired this marker protein over a time course of 60 min is shown in Fig. 1C. Strikingly, phagosomes containing mannan beads acquired the marker very rapidly, with 90% already positive for LAMP-2 after 5 min of uptake (Fig. 1C). In contrast, the frequency of fusion events for phagosomes containing IgG Fc beads, LPS beads, and in particular avidin beads, was significantly slower. After 5 min 52% of IgG Fc bead-, 68% of LPS beadand only 9% of avidin bead-containing phagosomes were positive for LAMP-2. After 30 min uptake, phagosomes containing IgG Fc and LPS beads acquired LAMP-2 at rates similar to phagosomes containing mannan beads, whereas 67% of phagosomes containing avidin beads were positive for LAMP-2. These results indicate that the type of ligand on the latex bead surface plays a significant role not only for their uptake efficiency, but also for later stages of phagosome maturation. Gene expression is dependent on the type of receptor ligand interaction during phagocytosis Fig. 1. Latex beads are internalized and acquire lysosomal markers in dependence of their surface ligand. (A) Internalization rates of 1 m latex beads coupled to avidin, to the Fc fragment of IgG (IgG Fc), to lipopolysaccharides (LPS) and to mannan in J774.A1 macrophages over a time course of 60 min. Cells were incubated with the same concentration of beads and 50 cells per time point were analyzed. Shown are averages of three independent experiments. (B) Image of a fixed macrophage incubated with mannan-coated latex beads (blue) for 60 min obtained by confocal microscopy. Maturation of phagosomes was analyzed by co-localization with the late endosomal/lysosomal marker LAMP-2 (red). The actin cytoskeleton was stained with fluorescein-labeled phalloidin (green). Inset shows a detailed view of phagosomes that have fused with late endocytic compartments and lysosomes indicated by their acquisition of LAMP-2. (C) Histogram showing the acquisition kinetics of phagosomes for LAMP-2. Shown are averages of phagosomes that were positive for LAMP-2 analyzed in 50 cells for differently coated beads and different time points in three independent experiments. Bar: 5 m. lysosomal compartments and mature into phagolysosomes. This process is characterized by phagosomal acidification and the acquisition of hydrolases. We analyzed the extent of phagolysosomal fusion by detection on phagosomes of a marker for late endosomes and lysosomes. Macrophages were allowed to internalize latex beads coated with different ligands, then fixed at different time points and labeled using an antibody against the lysosomeassociated membrane protein 2 (LAMP-2; Fig. 1B). The relative We next monitored the pattern of macrophage gene expression in response to the different beads. For this purpose, J774 mouse macrophages were incubated with latex beads conjugated to avidin, IgG Fc, LPS and mannan for 30 and 60 min and total macrophage RNA was isolated and subjected to mouse whole genome bioarrays. This experiment was performed in triplicate for each studied ligand. As a control, macrophages were incubated with the phagocytosis medium in absence of latex beads for 30 and 60 min. Changes in gene expression were considered significant, when they appeared in all three replicates compared to the basal level at control conditions, after performing a two-way ANOVA analysis with a P-value cut-off <0.01. The complete data set of the microarray analysis was deposited at the NCBI GEO database (GSE17761). In Fig. 2A the total number of up-regulated and down-regulated genes for each examined ligand are shown for 30 and 60 min after phagocytic uptake. Strikingly, in all the cases except for IgG Fc beads at 30 min, the number of down-regulated genes was higher than the number of up-regulated genes (Fig. 2A). This result suggests that, independently of the ligand, the process of phagocytosis in macrophages induces a down-regulation of cellular functions normally active in resting cells. This is further supported by the observation that the common and overlapping genes found affected in all ligands or in two or three of the four ligand samples were mostly down-regulated at both time points (64% of all downregulated genes at 30 min and 65% at 60 min; Fig. 2B). In contrast, genes that were uniquely changed for a given ligand were mostly up-regulated (65% of all up-regulated genes at 30 min and 62% at 60 min; Fig. 2B). IgG Fc beads induced the highest number of upregulated genes after 30 min uptake (n = 1009), which decreases to 465 at 60 min. Mannan beads showed an opposite regulation, inducing the up-regulation of a smaller amount of specific genes at 30 min (n = 461), which doubled at 60 min (n = 913). The response induced by LPS beads and avidin beads exhibited less dynamic changes with about 500 unique up-regulated genes at both time points. The finding that the majority of up-regulated genes were specific to a given ligand suggests that binding and internalization of the ligand-coated latex beads induced a unique cellular response rather than stimulating a common pool of genes that would be required for the phagocytic process to proceed irrespective of the ligand. For both the 30 and 60 min time points, the listed genes common to all ligands, or unique to only one of them including their fold change values can be found as supplementary material (Table SI, supplementary data). To gain further insight into the biological function of the genes that were affected during phagocytosis of ligand-coated beads, we mapped genes to their functional annotation according to Gene
6 6 E. Hoffmann et al. / European Journal of Cell Biology xxx (2010) xxx xxx Fig. 2. Gene regulation of J774.A1 macrophages during phagocytosis of coated latex beads analyzed by mouse whole genome bioarrays. Cells were incubated with latex beads coupled to avidin, IgG Fc, LPS and mannan for 30 and 60 min. Gene changes that occurred in all three replicates with a P-value cut-off <0.01 were considered significant. (A) Summary of microarray results showing numbers of up-regulated and down-regulated genes at 30 and 60 min in comparison to untreated control cells at the same time points. (B) Pie charts displaying gene expression changes that were unique to either one ligand (colored), common to all ligands (black) or exhibited overlapping appearance for 2 or 3 ligands (grey). Numbers of up-regulated and down-regulated genes for all categories as well as the overall number of gene changes are shown. Ontology (GO) terms of biological process using the DAVID analysis tool (Huang et al., 2009). In Fig. 3 biological process terms of genes that were changed after 30 min of phagocytic uptake are shown to identify early responses of macrophages during phagocytosis. As expected, genes, which were found affected commonly in all the examined latex bead ligands, mainly belonged to GO terms of metabolic and developmental processes as well as the regulation of cellular and biological processes (Fig. 3A). For example, the inositol polyphosphate-4-phosphatase type I and II were down-regulated 0.04-fold and 0.21-fold, respectively (Table SI, supplementary data). The analysis of functional annotations of genes that were unique to a given ligand revealed several interesting differences (Fig. 3B). First, dominant GO term groups that we found commonly regulated by all ligand types, such as developmental, metabolic and biological processes (Fig. 3A), displayed low percentages of changes (Fig. 3B). Second, we found main differences in the functional groups affected uniquely by a given ligand (Fig. 3B). Avidin beads mainly induced the up-regulation of genes belonging to GO terms of morphogenesis (e.g. laminin: 7.3-fold; Table SI) and signaling pathways (e.g. kinase suppressor of Ras 1: 2.5-fold; Table SI), but also genes involved in cytoskeleton and organelle organization (e.g. scinderin: 4.0-fold; Table SI). In contrast, IgG Fc and LPS beads up-regulated mainly genes important for transcription (e.g. STAT5B by IgG Fc: 1.9-fold; growth factor independent 1 by LPS: 2.5-fold; Table SI) and localization (e.g. syntaxin 18 by IgG Fc: 1.8-fold; epsin 2 by LPS: 3.8-fold; Table SI), the latter including molecules which transport and/or maintain the specific location of organelles. Mannan beads triggered several genes involved in apoptosis (e.g. cell death activator CIDE-A: 2.7-fold; Table SI), intracellular transport (e.g. phosphatidylinositol transfer protein 1: 2.3-fold; Table SI) and localization (e.g. murinoglobulin 1: 2.5-fold; Table SI). These findings indicate once more that specific receptor ligand interactions induce specific changes in gene expression that lead to a unique signature depending on the ligand. Proteomic analysis of mature phagosomes formed after internalization of IgG Fc and mannan beads reveals differences on the organelle level Based on the results obtained so far, we provided evidence that specific interactions between a particular ligand and its receptor at the cell surface are able to modulate phagocytic internalization and maturation kinetics, as well as the gene expression and signaling in macrophages. To couple these findings to functional characteristics at the organelle level, we decided to study the protein composition of mature phagosomes to investigate the impact of receptor ligand interactions at late stages of phagocytosis. Given that each receptor induced to a large extent its own specific set of genes, it was important to test their impact on the protein composition of phagosomes containing different beads. LBP are ideal for these purposes, because they can be purified at very high purities and were applied successfully to proteomic studies in different phagocyte models (Griffiths and Mayorga, 2007). To compare the protein composition of phagosomes during late stages of their maturation and to find out whether initial receptor ligand interactions have an impact on these late characteristics in phagocytosis, we concentrated on phagosomes
7 E. Hoffmann et al. / European Journal of Cell Biology xxx (2010) xxx xxx 7 Fig. 3. Mapping of gene expression changes 30 min after internalization of latex beads coupled to avidin, IgG Fc, LPS and mannan according to their gene ontology (GO) functional annotation. Shown are biological process terms of the most prominent functional groups of up- and down-regulated genes that were common to all ligands (A) as well as the ones, which were unique to only one of the bead ligands (B). containing IgG Fc- and mannan-coated latex beads (subsequently referred to as IgG Fc LBP and mannan LBP). We were interested into these two phagosome types because IgG Fc beads are internalized into macrophages via Fc receptors in opsonic pathways, whereas mannan beads trigger predominantly a non-opsonic uptake via the mannose receptor. Phagosomes made in response to these beads were isolated 2 h after their internalization (1 h pulse followed by 1 h chase). This time point was selected because it has been the most-commonly used time point for many assays investigating mature phagosomes containing latex beads including proteomics (Garin et al., 2001; Jutras et al., 2008). Therefore, we were able to minimize the influence of time-dependent dynamics of proteins involved in phagosome maturation during phagosomal fusion and fission events with endosomal and lysomal compartments and were able to analyze more ligand-dependent differences between the two phagosome types. Equal amounts of phagosomes were lysed and their proteins separated by two-dimensional gel electrophoresis. Silver-stained gels of phagosome extracts were carried out in triplicates and displayed qualitative as well as quantitative differences. Representative examples of IgG Fc and mannan LBP are shown in Fig. 4A. Although many polypeptide spots appeared similar in both samples, a number of other proteins were unique to a given sample (arrows in Fig. 4A), or found in different amounts (arrowheads in Fig. 4A). For a more global analysis of phagosomal proteins we next used LC MS/MS applied to same amounts of IgG Fc LBP and mannan LBP to identify their protein composition. Using a quite stringent cut-off of protein identification probability (99%), we carried out a restrictive analysis of the phagosomal proteomes. Interestingly, only 58% of all 265 identified proteins were common to both phagosome types. Reminiscent of the gene expression results, several proteins in the phagosomal proteomes were unique to either IgG Fc LBP or mannan LBP (Fig. 4B). A selection of identified proteins categorized into functional groups is shown in Table 1, whereas a comprehensive list of all proteins identified by LC MS/MS including numbers of their unique spectra and sequence coverage can be found as supplementary material (Table SII, supplementary data). Many proteins known to be involved in phagosome maturation were found in both LBP types, such as proton ATPase subunits, LAMP-1 and LAMP-2, cathepsin-b and cathepsin-d as well as Rab5 and Rab7 (Table 1). However, many proteins involved in signaling as well as receptor subunits were also present on phagosome membranes and exhibited remarkable differences between IgG Fc LBP and mannan LBP. For example, in IgG Fc LBP we were able to identify the IgG-1 chain as well as the Fc receptor of IgG (Table 1), which provide evidence that IgG Fc beads were internalized by Fc receptor-dependent phagocytosis. In mannan LBP, but not in IgG Fc LBP samples, several signaling molecules were identified, such as beta 2-microglobulin, CD68, CD180, lymphocyte antigen 86, programmed cell death 6-interacting protein, TLR13 and the tyrosine protein kinase Lyn (Table 1). These results suggest that mature
8 8 E. Hoffmann et al. / European Journal of Cell Biology xxx (2010) xxx xxx Fig. 4. Proteomic analysis of phagosomes formed after internalization of latex beads coated with IgG Fc (IgG Fc LBP) and mannan (Mannan LBP) isolated from J774.A1 macrophages 2 h after uptake. (A) Images of phagosomes after two-dimensional gel electrophoresis and silver staining showing proteins, which are unique to IgG Fc and mannan LBP (arrows) as well as proteins that appeared at different quantities (arrowheads). (B) Diagram representing the relative distribution of phagosomal proteins identified by LC MS/MS with a protein identification probability of 99%. 265 proteins were identified in total. phagosomes containing mannan beads, which were formed after non-opsonic internalization, possess the capability to modulate diverse signaling pathways. Since phagosomes rely on intracellular trafficking and interactions with the cytoskeleton to acquire and sustain their degradative properties (Araki, 2006), we also analyzed the presence of small GTPases of the Rab family and cytoskeletal proteins. Interestingly, among the identified Rab proteins found in both phagosome types, such as Rab-1A, -2A, -5B, -5C, -7, -14 and -33B, some were found associated specifically with IgG Fc LBP (Rab-5A, -31) or mannan LBP (Rab-10, -18, -22A, -27A) after 2 h internalization (Table 1). The finding of the association of different isoforms of Rab proteins to different types of phagosomes at a particular time point emphasizes the different composition of these organelles. Additionally, we were also able to detect proteins of all cytoskeleton filament networks on phagosome membranes (cytoplasmic actin, tubulin subunits and vimentin, a member of the intermediate filament system) as well as a high number of identified actin-associated proteins (Table 1), which is further supported by immunoblotting results of phagosomal lysates (see below). The association of cytoskeletal proteins with mature phagosomes did not show many differences between the analyzed LBP types, but points out the importance of intracellular trafficking during phagosome maturation. Receptor ligand interactions also modulate the time-dependent association of proteins with mature phagosomes Since phagosomes acquire proteins during their maturation by fusion with endosomes and lysosomes in a very dynamic way, we finally isolated phagosomes at different time points after uptake to analyze them for their association with different proteins. The same amount of LBP lysates were separated by SDS-PAGE, transferred to nitrocellulose membranes and analyzed by immunoblotting. Our data obtained by confocal microscopy indicated the presence of LAMP-2 at different times depending on the latex bead type (Fig. 1C). We analyzed lysates of phagosomes containing IgG Fc and mannan beads isolated 30, 60, 120 and 240 min after uptake for the presence of LAMP-2, cathepsin-d and actin. As shown in Fig. 5, LAMP-2 was associated with all phagosome types at all analyzed time points with increasing concentration without significant differences between IgG Fc and mannan LBP lysates (Fig. 5). However, when phagosomes were studied for the presence of cathepsin-d and actin, differences were obvious between the two phagosome types. At 60 and 120 min, we detected the immature form of cathepsin-d (52 kd) associated at higher amounts with IgG Fc LBP compared to mannan LBP, whereas it was not present on early phagosomes (30 min). Additionally, we observed a rather striking pattern for actin associated with phagosomes (Fig. 5). Both phagosomes types displayed similar amounts of associated actin early after uptake (30 min), which was absent at 60 min and associates again with mature phagosomes (120 and 240 min). However, actin was much more prominent on mannan LBP at 120 min compared to IgG Fc LBP (Fig. 5). In conclusion, our results show that, in macrophages, receptor ligand interactions during the initial step of phagocytosis dictate gene expression patterns and modulate signaling pathways. Moreover, they also lead to differences on the protein level evident for protein composition, maturation kinetics and dynamics of associated proteins of the analyzed phagosome types. Discussion It is well established that the initial contact of a pathogen with cell surface receptors initiates an immediate signaling response. Since pathogens possess a variety of ligands on their surface, they cause a high complexity of multi-ligand interactions with host receptors inducing extensive overlap and crosstalk between different pathways. It is also clear that a given receptor on phagocytes such as macrophages induce a specific pattern of signaling that show significant differences from other receptors. The question we posed in this study was How do professional phagocytes, such Fig. 5. Receptor ligand interactions modulate the time-dependent association of proteins with mature phagosomes. Phagosomes containing beads coated with IgG Fc (IgG Fc LBP) and mannan (Mannan LBP) were isolated from J774.A1 macrophages after different time points (30, 60, 120 and 240 min). Phagosomal lysates were separated by SDS-PAGE and tested by immunoblotting using specific antibodies against LAMP-2, cathepsin-d and actin.
9 E. Hoffmann et al. / European Journal of Cell Biology xxx (2010) xxx xxx 9 Table 1 Selection of proteins identified by LC MS/MS in phagosome preparations containing IgG Fc-coated or Mannan-coated latex beads isolated from J774.A1 macrophages after 1 h pulse/1 h chase divided into functional categories. Proteins IgG Fc Mannan Phagosomal/lysosomal markers ATPase, H+ transporting, various subunits + + ATP synthase, vacuolar, various subunits + + Calreticulin + + Calnexin + + Cathepsin-B, -D, -L, -Z + + Early endosome antigen Lysosomal acid lipase + + Lysosomal acid phosphatase + + Lysosome-associated membrane protein (LAMP) 1 and Syntaxin Transferrin receptor protein 1 + Vesicle-associated membrane protein (VAMP) 4 + Vesicle-associated membrane protein (VAMP) Signaling molecules Annexin-1 + Annexin-2, -4, -5, Annexin-7 + Beta 2-microglobulin + CD68 (Macrosialin) + CD180 (Toll-like receptor protein RP-105) + + Fc receptor, IgG, IIb isoform + G protein-coupled receptor Heat shock protein 70 (HSP70) + + IgG-1 chain + Lymphocyte antigen 86 + Programmed cell death 6-interacting protein + Toll-like receptor Toll-like receptor 13 + Tyrosine protein kinase Lyn + Rab proteins Rab-1A + + Rab-2A + + Rab-5A + Rab-5B, -5C + + Rab Rab-10 + Rab Rab-18 + Rab-22A + Rab-27A + Rab-31 + Rab-33B + + Cytoskeletal proteins Actin, cytoplasmic + + ADP-ribosylation factor 6 + Arp2/3 complex + + Coronin-1A + Elongation factor 1 alpha + + Elongation factor Moesin + + Profilin Rac Tubulin, alpha and beta chain + + Vimentin + + as macrophages, respond to the binding and phagocytic uptake of latex beads coated with a specific ligand during early and late events of phagocytosis?. The latex bead phagosome offers a system of multivalent interaction of a defined ligand with a defined receptor on the macrophage surface. Here, using different methods we compared the impact of beads coated with the Fc fragment of mouse IgG, mannan, LPS and avidin with their specific receptors on different stages of the phagocytic process and on gene expression. Irrespective of the ligand, J774.A1 cells were able to bind and internalize the beads in a morphologically indistinguishable manner. However, although macrophages were incubated with same numbers of beads, the ones coated with mannan were internalized at faster rates and in higher numbers compared to beads coated with other ligands. In macrophages, mannan and terminal mannose residues are predominantly recognized by the MR, a prototypic pattern-recognition receptor and C-type lectin (Wileman et al., 1986). The MR binds a broad spectrum of pathogens including Candida albicans and Pneumocystis carinii (Ezekowitz et al., 1990, 1991). Although it was shown that serum opsonization enhances the internalization of some pathogens (Balagopal et al., 2006), our observation confirms previous findings in COS cells transfected with different chimeric receptor constructs showing a higher phagocytic index for the MR compared to Fc RI (Kruskal et al., 1992). We also observed that phagosomes containing mannan beads acquired the late endosomal/lysosomal protein LAMP-2 more rapidly than the other phagosome types. In fact, within only 5 min after adding these beads, around 90% of their newly formed phagosomes already exhibited fusion events with late endosomes/lysosomes. Although we did not address this issue, it seems unlikely that receptor-induced newly synthesized proteins are relevant during this short time scale. In murine and rabbit macrophages, several studies support the idea that the MR is coupled to bactericidal functions, such as the production of reactive oxygen intermediates (Klegeris et al., 1996) and cytokines (Shibata et al., 1997). However, in human cells, the data argue that this receptor activates anti-inflammatory signaling (Nigou et al., 2001; Chieppa et al., 2003) and delays the maturation of phagosomes (Astarie-Dequeker et al., 1999). The mechanistic differences between rodents and humans for the same receptor (Paul et al., 1996) remain to be elucidated. Together these data support the idea that interactions of mannan beads with the MR induce a specific signaling, different from the other ligands, which leads to a more efficient bead uptake and more rapid fusion events with lysosomes. The precise nature of these signaling pathways as well as the possible involvement of co-receptors, such as Toll-like receptors, remain to be elucidated. Our study of the gene expression response upon bead binding and uptake showed that the majority of affected genes are down-regulated, independent of the bead ligand, suggesting a general down-regulation of cellular functions during phagocytosis. A similar result was found during phagocytosis of Mycobacterium smegmatis (Gutierrez et al., 2008) and Aspergillus fumigatus (Cortez et al., 2006). We observed that the down-regulated genes common to all ligands encode proteins involved in metabolism, energy homeostasis and development suggesting a decrease of these processes during phagocytosis compared to resting cells. In contrast, around two-third of the up-regulated genes at both investigated time points, 30 and 60 min, were unique to one of the bead ligands. Our analysis mapping altered genes at 30 min to GO functional annotation terms strengthen the observation that specific receptor ligand interactions induce particular cellular responses early after uptake, which differ broadly between the investigated ligands. We found that IgG Fc and LPS beads upregulated many genes involved in transcription and processes of intracellular localization, whereas avidin beads induced predominantly signaling pathways and genes important for cytoskeleton organization. Mannan beads up-regulated genes important for intracellular transport, but also genes involved in apoptotic processes. These also include molecules involved in the recognition of apoptotic cells. Previous findings have demonstrated that particular C-type lectins, such as macrophage galactose-type C-type lectin-1 (MGL-1; Yuita et al., 2005) sense products from dying cells and modulate immunogenic cell death (Cambi and Figdor, 2009). Further analysis of single genes will be necessary to understand the impact of receptor ligand interactions, and coated latex beads provide a useful tool to investigate these processes in more detail.
10 10 E. Hoffmann et al. / European Journal of Cell Biology xxx (2010) xxx xxx The high number of up-regulated genes in response to IgG beads at 30 min and to mannan beads at 60 min also emphasize the differences in time and specificity of the utilized pathways. Signaling mediated through immunoreceptor tyrosine-based activation motifs (ITAM)-bearing Fc receptors during phagocytosis has been investigated in detail (Swanson and Hoppe, 2004) and is known to have an impact on phagosome maturation (Tian et al., 2008), and the immune response (Pasquier et al., 2005). On the other hand, the mannose receptor has a single tyrosine residue in its cytoplasmic tail and different signal transduction pathways are engaged (Kerrigan and Brown, 2009). For some members of the newly emerging family of Syk-coupled C-type lectins, such as Dectin-1, the recognition of PAMP as well as an induction of genes important for an innate immune response have been demonstrated (Rogers et al., 2005; Robinson et al., 2006). We found that 30 min after incubation with mannan beads, macrophages are characterized by a down-regulation of the pro-inflammatory cytokine TNF, whereas the interleukin-13 receptor subunit 1 (IL-13R 1) was up-regulated. IL-13R 1 is known to form a type II receptor after association with IL-4R (Junttila et al., 2008) inducing a cytoplasmic signaling cascade that involves the Janus kinase family and the phosphoinositide 3-kinase (PI3K) pathway (Keegan et al., 1995). Macrophages utilize these pathways to undergo alternative activation pathways that trigger different phenotypes during the immune response to parasites and pathogens (Martinez et al., 2009). Next, our study aimed to analyze the impact of receptor ligand interactions on the organelle level to gain insight into early as well as late events during phagocytosis. Focusing on IgG Fc and mannan LBP to analyze opsonin-dependent and opsonin-independent phagocytic events, we found that around 60% of proteins were common to both LBP types 2 h after internalization. As expected, lysosomal hydrolases important for proteolytic degradation (Brix et al., 2008) as well as cytoskeletal proteins involved in trafficking during phagosome maturation (Araki, 2006) were found at this time point. Previously, a quantitative proteomic analysis of phagosomes containing IgG-coupled latex beads in RAW macrophages has been published (Rogers and Foster, 2007). These data are in good agreement with our proteomic analysis, since 78% of our identified proteins in IgG Fc LBP in J774.A1 macrophages were also identified in that study. Additionally, we also confirmed the presence of many proteins in IgG Fc and mannan LBP in comparison with other proteomic studies of phagosomes containing latex beads (Garin et al., 2001; Jutras et al., 2008; Trost et al., 2009), Francisella tularensis (Kovarova et al., 2002) and Mycobacterium bovis BCG (Lee et al., 2010). Interestingly, some of our identified proteins were specifically found in M. bovis BCG phagosomes but not in phagosomes containing non-coated latex beads in the same study (Lee et al., 2010), such as alpha-mannosidase, Rab-14 and the G protein beta 1 (Gnb1) involved in trans-membrane signaling. Around 40% of the identified proteins from the proteomic analysis were specific for IgG Fc or mannan LBP, which mainly included signaling molecules and Rab proteins, regulators of intracellular trafficking that exhibited an isoform-dependent association with the different phagosome types. Together with our Western blot analysis of phagosomal proteins showing that a lysosomal protein, cathepsin-d, as well as the cytoskeletal protein actin are associated with phagosomes differently during their maturation, it demonstrates that the protein composition of these organelles is also altered by initial receptor ligand interactions occurring upon binding of beads to the macrophage surface. Especially Rab GTPases are reversibly associated with organelle membranes and are responsible for a close interrelationship between receptor signaling and membrane trafficking (Sorkin and von Zastrow, 2009). They function as molecular switches after specific signaling events and effector coupling to determine the type of interactions between phagosomes and other organelles (Kinchen and Ravichandran, 2008). In this respect, Rab-5 and Rab-7 are the key GTPases responsible for the recycling of proteins back to the plasma membrane and as entry points for the acquisition of lysosomal proteases during phagosome maturation. It has been elegantly demonstrated that Rab-5 positive early endocytic organelles switch to Rab-7 positive organelles after a certain threshold of acquisition, a mechanism called Rab conversion (Rink et al., 2005). Many other Rab GTPases are involved during phagosome maturation and organelle trafficking (Stenmark, 2009), but our findings also indicate that their transient appearance on phagosomes is also coupled to the type of phagosome content and signaling during the initial contact between receptor and ligand. Our findings support the idea of the individuality of phagosomes to a certain degree, which allows them to regulate their intracellular fate locally and function as independent entities (Griffiths, 2004). Recent proteomic studies of the response of RAW macrophages upon infection with Candida albicans (Martinez-Solano et al., 2009) and Salmonella enterica (Shi et al., 2009) showed that these cells varied their expression level of proteins involved in cytoskeletal organization and signal transduction resulting in the induction of an anti-inflammatory immune response. In the future, more detailed and connected approaches are necessary to gain insight into the impact of receptor ligand interactions on the inflammatory response of the cell. Our experiments provide proof of the principle that the use of latex beads conjugated to specific ligands can be a powerful tool to follow the fate of different kinds of phagosomes in a phagocytic cell. This system can be adapted to analyze the response of a phagocyte when two, three or more ligands are conjugated to the same bead, or in contrast, when different beads coated with a single ligand are applied simultaneously to these cells. The outcome of such experiments will help to understand, which receptor ligand interactions might dominate signaling events and what their impact on the cellular immune response will be. Acknowledgements This work was supported by grants from the Deutsche Forschungsgemeinschaft (DFG; KU1528/2), the EU commission Marie Curie Research Training Network (MRTN-CT ) and the AXA Research Fund. We are very grateful to the support of Sebastian Amigorena (Institute Curie, Paris, France) and the Genomics core facility and the Proteomics core facility (EMBL Heidelberg, Germany). We acknowledge Nicholas Sherman and the W.M. Keck Biomedical Mass Spectrometry Laboratory, funded by the Pratt Committee of the University of Virginia, USA, for LC MS/MS and help in data analysis. We thank Wolfgang Faigle and the Mass spectrometry platform of Institute Curie, Paris for help with proteomic analysis, Devang Patel and Rhys Allan for experimental help and critical reading of the manuscript, Raphael Zollinger for his comments on the microarray analysis as well as Avin Ramaiya for his help with image processing. Appendix A. Supplementary data Supplementary data associated with this article can be found, in the online version, at doi: /j.ejcb References Aderem, A., Phagocytosis and the inflammatory response. J. Infect. Dis. 187, S340 S345. Al-Haddad, A., Shonn, M.A., Redlich, B., Blocker, A., Burkhardt, J.K., Yu, H., Hammer III, J.A., Weiss, D.G., Steffen, W., Griffiths, G., Kuznetsov, S.A., Myosin Va bound to phagosomes binds to F-actin and delays microtubule-dependent motility. Mol. Biol. Cell 12,
11 E. Hoffmann et al. / European Journal of Cell Biology xxx (2010) xxx xxx 11 Anes, E., Peyron, P., Staali, L., Jordao, L., Gutierrez, M.G., Kress, H., Hagedorn, M., Maridonneau-Parini, I., Skinner, M.A., Wildeman, A.G., Kalamidas, S.A., Kuehnel, M., Griffiths, G., Dynamic life and death interactions between Mycobacterium smegmatis and J774 macrophages. Cell. Microbiol. 8, Araki, N., Role of microtubules and myosins in Fc gamma receptor-mediated phagocytosis. Front. Biosci. 11, Astarie-Dequeker, C., N Diaye, E.N., Le Cabec, V., Rittig, M.G., Prandi, J., Maridonneau- Parini, I., The mannose receptor mediates uptake of pathogenic and nonpathogenic mycobacteria and bypasses bactericidal responses in human macrophages. Infect. Immun. 67, Balagopal, A., MacFarlane, A.S., Mohapatra, N., Soni, S., Gunn, J.S., Schlesinger, L.S., Characterization of the receptor ligand pathways important for entry and survival of Francisella tularensis in human macrophages. Infect. Immun. 74, Bjellqvist, B., Ek, K., Righetti, P.G., Gianazza, E., Görg, A., Westermeier, R., Postel, W., Isoelectric focusing in immobilized ph gradients: principle, methodology and some applications. J. Biochem. Biophys. Methods 6, Blander, J.M., Medzhitov, R., On regulation of phagosome maturation and antigen presentation. Nat. Immunol. 7, Blocker, A., Severin, F.F., Burkhardt, J.K., Bingham, J.B., Yu, H., Olivo, J.C., Schroer, T.A., Hyman, A.A., Griffiths, G., Molecular requirements for bi-directional movement of phagosomes along microtubules. J. Cell Biol. 137, Brix, K., Dunkhorst, A., Mayer, K., Jordans, S., Cysteine cathepsins: cellular roadmap to different functions. Biochimie 90, Brown, G.D., Dectin-1: a signalling non-tlr pattern-recognition receptor. Nat. Rev. Immunol. 6, Brown, M.H., Boles, K., van der Merwe, P.A., Kumar, V., Mathew, P.A., Barclay, A.N., B4, the natural killer and T cell immunoglobulin superfamily surface protein, is a ligand for CD48. J. Exp. Med. 188, Cambi, A., Figdor, C., Necrosis: C-type lectins sense cell death. Curr. Biol. 19, R Chieppa, M., Bianchi, G., Doni, A., Del Prete, A., Sironi, M., Laskarin, G., Monti, P., Piemonti, L., Biondi, A., Mantovani, A., Introna, M., Allavena, P., Cross-linking of the mannose receptor on monocyte-derived dendritic cells activates an anti-inflammatory immunosuppressive program. J. Immunol. 171, Corrotte, M., Chasserot-Golaz, S., Huang, P., Du, G., Ktistakis, N.T., Frohman, M.A., Vitale, N., Bader, M.F., Grant, N.J., Dynamics and function of phospholipase D and phosphatidic acid during phagocytosis. Traffic 7, Cortez, K.J., Lyman, C.A., Kottilil, S., Kim, H.S., Roilides, E., Yang, J., Fullmer, B., Lempicki, R., Walsh, T.J., Functional genomics of innate host defense molecules in normal human monocytes in response to Aspergillus fumigatus. Infect. Immun. 74, Cox, D., Greenberg, S., Phagocytic signaling strategies: Fc(gamma)receptormediated phagocytosis as a model system. Semin. Immunol. 13, Defacque, H., Bos, E., Garvalov, B., Barret, C., Roy, C., Mangeat, P., Shin, H.W., Rybin, V., Griffiths, G., Phosphoinositides regulate membrane-dependent actin assembly by latex bead phagosomes. Mol. Biol. Cell 13, Desjardins, M., Huber, L.A., Parton, R.G., Griffiths, G., Biogenesis of phagolysosomes proceeds through a sequential series of interactions with the endocytic apparatus. J. Cell Biol. 124, Desjardins, M., Griffiths, G., Phagocytosis: latex leads the way. Curr. Opin. Cell Biol. 15, Ernst, J.D., Bacterial inhibition of phagocytosis. Cell. Microbiol. 2, Ezekowitz, R.A., Sastry, K., Bailly, P., Warner, A., Molecular characterization of the human macrophage mannose receptor, demonstration of multiple carbohydrate recognition-like domains and phagocytosis of yeasts in Cos-1 cells. J. Exp. Med. 172, Ezekowitz, R.A., Williams, D.J., Koziel, H., Armstrong, M.Y., Warner, A., Richards, F.F., Rose, R.M., Uptake of Pneumocystis carinii mediated by the macrophage mannose receptor. Nature 351, Garin, J., Diez, R., Kieffer, S., Dermine, J.F., Duclos, S., Gagnon, E., Sadoul, R., Rondeau, C., Desjardins, M., The phagosome proteome: insight into phagosome functions. J. Cell Biol. 152, Geijtenbeek, T.B., Gringhuis, S.I., Signalling through C-type lectin receptors: shaping immune responses. Nat. Rev. Immunol. 9, Görg, A., Obermaier, C., Boguth, G., Harder, A., Scheibe, B., Wildgruber, R., Weiss, W., The current state of two-dimensional electrophoresis with immobilized ph gradients. Electrophoresis 21, Griffiths, G., On phagosome individuality and membrane signalling networks. Trends Cell Biol. 14, Griffiths, G., Mayorga, L., Phagosome proteomes open the way to a better understanding of phagosome function. Genome Biol. 8, 207. Gutierrez, M.G., Mishra, B.B., Jordao, L., Elliott, E., Anes, E., Griffiths, G., NF-kappa B activation controls phagolysosome fusion-mediated killing of mycobacteria by macrophages. J. Immunol. 181, Huang, D.W., Sherman, B.T., Lempicki, R.A., Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, Huynh, K.K., Grinstein, S., Regulation of vacuolar ph and its modulation by some microbial species. Microbiol. Mol. Biol. Rev. 71, Jahraus, A., Tjelle, T.E., Berg, T., Habermann, A., Storrie, B., Ullrich, O., Griffiths, G., In vitro fusion of phagosomes with different endocytic organelles from J774 macrophages. J. Biol. Chem. 273, Jahraus, A., Egeberg, M., Hinner, B., Habermann, A., Sackman, E., Pralle, A., Faulstich, H., Rybin, V., Defacque, H., Griffiths, G., ATP-dependent membrane assembly of F-actin facilitates membrane fusion. Mol. Biol. Cell 12, Janeway Jr., C.A., The immune system evolved to discriminate infectious nonself from noninfectious self. Immunol. Today 13, John, M., Tominski, C., Schumann, H., Visual and analytical extensions for the table lens. In: Börner, K., Gröhn, M.T., Park, J., Roberts, J.C. (Eds.), Visualization and Data Analysis. Proceedings of the SPIE, vol. 6809, p Junttila, I.S., Mizukami, K., Dickensheets, H., Meier-Schellersheim, M., Yamane, H., Donnelly, R.P., Paul, W.E., Tuning sensitivity to IL-4 and IL-13: differential expression of IL-4Ralpha, IL-13Ralpha1, and gammac regulates relative cytokine sensitivity. J. Exp. Med. 205, Jutras, I., Houde, M., Currier, N., Boulais, J., Duclos, S., LaBoissiere, S., Bonneil, E., Kearney, P., Thibault, P., Paramithiotis, E., Hugo, P., Desjardins, M., Modulation of the phagosome proteome by interferon-gamma. Mol. Cell. Proteomics 7, Kabelitz, D., Medzhitov, R., Innate immunity cross-talk with adaptive immunity through pattern recognition receptors and cytokines. Curr. Opin. Immunol. 19, 1 3. Keegan, A.D., Johnston, J.A., Tortolani, P.J., McReynolds, L.J., Kinzer, C., O Shea, J.J., Paul, W.E., Similarities and differences in signal transduction by interleukin 4 and interleukin 13: analysis of Janus kinase activation. Proc. Natl. Acad. Sci. U.S.A. 92, Kerrigan, A.M., Brown, G.D., C-type lectins and phagocytosis. Immunobiology 214, Kinchen, J.M., Ravichandran, K.S., Phagosome maturation: going through the acid test. Nat. Rev. Mol. Cell. Biol. 9, Kjeken, R., Egeberg, M., Habermann, A., Kuehnel, M., Peyron, P., Floetenmeyer, M., Walther, P., Jahraus, A., Defacque, H., Kuznetsov, S.A., Griffiths, G., Fusion between phagosomes, early and late endosomes: a role for actin in fusion between late, but not early endocytic organelles. Mol. Biol. Cell 15, Klegeris, A., Budd, T.C., Greenfield, S.A., Acetylcholinesterase-induced respiratory burst in macrophages: evidence for the involvement of the macrophage mannose fucose receptor. Biochim. Biophys. Acta 1289, Kovarova, H., Halada, P., Man, P., Golovliov, I., Krocova, Z., Spacek, J., Porkertova, S., Necasova, R., Proteome study of Francisella tularensis live vaccine strain-containing phagosome in Bcg/Nramp1 congenic macrophages: resistant allele contributes to permissive environment and susceptibility to infection. Proteomics 2, Kruskal, B.A., Sastry, K., Warner, A.B., Mathieu, C.E., Ezekowitz, R.A., Phagocytic chimeric receptors require both transmembrane and cytoplasmic domains from the mannose receptor. J. Exp. Med. 176, Kühnel, M., Anes, E., Griffiths, G., Isolation of latex bead- and mycobacteriacontaining phagosomes. In: Celis, J. (Ed.), Cell Biology A Laboratory Handbook, 3rd ed. Academic Press, San Diego, pp Laemmli, U.K., Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227, Lee, B.Y., Jethwaney, D., Schilling, B., Clemens, D.L., Gibson, B.W., Horwitz, M.A., The Mycobacterium bovis bacille Calmette-Guerin phagosome proteome. Mol. Cell. Proteomics 9, Link, T.M., Park, U., Vonakis, B.M., Raben, D.M., Soloski, M.J., Caterina, M.J., TRPV2 has a pivotal role in macrophage particle binding and phagocytosis. Nat. Immunol. 11, Martinez, F.O., Sica, A., Mantovani, A., Locati, M., Macrophage activation and polarization. Front. Biosci. 13, Martinez, F.O., Helming, L., Gordon, S., Alternative activation of macrophages: an immunologic functional perspective. Annu. Rev. Immunol. 27, Martinez-Solano, L., Reales-Calderon, J.A., Nombela, C., Molero, G., Gil, C., Proteomics of RAW macrophages upon interaction with heat-inactivated Candida albicans cells unravel an anti-inflammatory response. Proteomics 9, Nagarajan, S., Chesla, S., Cobern, L., Anderson, P., Zhu, C., Selvaraj, P., Ligand binding and phagocytosis by CD16 (Fc gamma receptor III) isoforms. Phagocytic signaling by associated zeta and gamma subunits in Chinese hamster ovary cells. J. Biol. Chem. 270, Nigou, J., Zelle-Rieser, C., Gilleron, M., Thurnher, M., Puzo, G., Mannosylated lipoarabinomannans inhibit IL-12 production by human dendritic cells: evidence for a negative signal delivered through the mannose receptor. J. Immunol. 166, Pasquier, B., Launay, P., Kanamaru, Y., Moura, I.C., Pfirsch, S., Ruffie, C., Henin, D., Benhamou, M., Pretolani, M., Blank, U., Monteiro, R.C., Identification of FcalphaRI as an inhibitory receptor that controls inflammation: dual role of FcRgamma ITAM. Immunity 22, Paul, S., Laochumroonvorapong, P., Kaplan, G., Comparable growth of virulent and avirulent Mycobacterium tuberculosis in human macrophages in vitro. J. Infect. Dis. 174, Poltorak, A., He, X., Smirnova, I., Liu, M.Y., Van Huffel, C., Du, X., Birdwell, D., Alejos, E., Silva, M., Galanos, C., Freudenberg, M., Ricciardi-Castagnoli, P., Layton, B., Beutler, B., Defective LPS signaling in C3H/HeJ and C57BL/10ScCr mice: mutations in Tlr4 gene. Science 282, Rink, J., Ghigo, E., Kalaidzidis, Y., Zerial, M., Rab conversion as a mechanism of progression from early to late endosomes. Cell 122, Robinson, M.J., Sancho, D., Slack, E.C., LeibundGut-Landmann, S., Reis e Sousa, C., Myeloid C-type lectins in innate immunity. Nat. Immunol. 7, Rogers, N.C., Slack, E.C., Edwards, A.D., Nolte, M.A., Schulz, O., Schweighoffer, E., Williams, D.L., Gordon, S., Tybulewicz, V.L., Brown, G.D., Reis e Sousa, C., Syk-dependent cytokine induction by Dectin-1 reveals a novel pattern recognition pathway for C type lectins. Immunity 22,
12 12 E. Hoffmann et al. / European Journal of Cell Biology xxx (2010) xxx xxx Rogers, L.D., Foster, L.J., The dynamic phagosomal proteome and the contribution of the endoplasmic reticulum. Proc. Natl. Acad. Sci. U.S.A. 104, Russell, D.G., Yates, R.M., Toll-like receptors and phagosome maturation. Nat. Immunol. 8, 217. Sheng, K.C., Pouniotis, D.S., Wright, M.D., Tang, C.K., Lazoura, E., Pietersz, G.A., Apostolopoulos, V., Mannan derivatives induce phenotypic and functional maturation of mouse dendritic cells. Immunology 118, Shevchenko, A., Wilm, M., Vorm, O., Mann, M., Mass spectrometric sequencing of proteins silver-stained polyacrylamide gels. Anal. Chem. 68, Shi, L., Chowdhury, S.M., Smallwood, H.S., Yoon, H., Mottaz-Brewer, H.M., Norbeck, A.D., McDermott, J.E., Clauss, T.R., Heffron, F., Smith, R.D., Adkins, J.N., Proteomic investigation of the time course responses of RAW macrophages to infection with Salmonella enterica. Infect. Immun. 77, Shibata, Y., Metzger, W.J., Myrvik, Q.N., Chitin particle-induced cell-mediated immunity is inhibited by soluble mannan: mannose receptor-mediated phagocytosis initiates IL-12 production. J. Immunol. 159, Sorkin, A., von Zastrow, M., Endocytosis and signalling: intertwining molecular networks. Nat. Rev. Mol. Cell. Biol. 10, Stenmark, H., Rab GTPases as coordinators of vesicle traffic. Nat. Rev. Mol. Cell. Biol. 10, Stuart, L.M., Ezekowitz, R.A., Phagocytosis: elegant complexity. Immunity 22, Stuart, L.M., Boulais, J., Charriere, G.M., Hennessy, E.J., Brunet, S., Jutras, I., Goyette, G., Rondeau, C., Letarte, S., Huang, H., Ye, P., Morales, F., Kocks, C., Bader, J.S., Desjardins, M., Ezekowitz, R.A., A systems biology analysis of the Drosophila phagosome. Nature 445, Swanson, J.A., Hoppe, A.D., The coordination of signaling during Fc receptormediated phagocytosis. J. Leukoc. Biol. 76, Swanson, J.A., Shaping cups into phagosomes and macropinosomes. Nat. Rev. Mol. Cell. Biol. 9, Taylor, P.R., Martinez-Pomares, L., Stacey, M., Lin, H.H., Brown, G.D., Gordon, S., Macrophage receptors and immune recognition. Annu. Rev. Immunol. 23, Tian, W., Li, X.J., Stull, N.D., Ming, W., Suh, C.I., Bissonnette, S.A., Yaffe, M.B., Grinstein, S., Atkinson, S.J., Dinauer, M.C., Fc{gamma}R-stimulated activation of the NADPH oxidase: phosphoinositide-binding protein p40phox regulates NADPH oxidase activity after enzyme assembly on the phagosome. Blood 112, Towbin, H., Staehelin, T., Gordon, J., Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc. Natl. Acad. Sci. U.S.A. 76, Trost, M., English, L., Lemieux, S., Courcelles, M., Desjardins, M., Thibault, P., The phagosomal proteome in interferon-gamma-activated macrophages. Immunity 30, Underhill, D.M., Ozinsky, A., Hajjar, A.M., Stevens, A., Wilson, C.B., Bassetti, M., Aderem, A., The Toll-like receptor 2 is recruited to macrophage phagosomes and discriminates between pathogens. Nature 401, Underhill, D.M., Ozinsky, A., Phagocytosis of microbes: complexity in action. Annu. Rev. Immunol. 20, Weis, W.I., Drickamer, K., Hendrickson, W.A., Structure of a C-type mannosebinding protein complexed with an oligosaccharide. Nature 360, Wileman, T.E., Lennartz, M.R., Stahl, P.D., Identification of the macrophage mannose receptor as a 175-kDa membrane protein. Proc. Natl. Acad. Sci. U.S.A. 83, Wright, S.D., Ramos, R.A., Tobias, P.S., Ulevitch, R.J., Mathison, J.C., CD14, a receptor for complexes of lipopolysaccharide (LPS) and LPS binding protein. Science 249, Yuita, H., Tsuiji, M., Tajika, Y., Matsumoto, Y., Hirano, K., Suzuki, N., Irimura, T., Retardation of removal of radiation-induced apoptotic cells in developing neural tubes in macrophage galactose-type C-type lectin-1-deficient mouse embryos. Glycobiology 15,
Hapten - a small molecule that is antigenic but not (by itself) immunogenic.
 Chapter 3. Antigens Terminology: Antigen: Substances that can be recognized by the surface antibody (B cells) or by the TCR (T cells) when associated with MHC molecules Immunogenicity VS Antigenicity:
Chapter 3. Antigens Terminology: Antigen: Substances that can be recognized by the surface antibody (B cells) or by the TCR (T cells) when associated with MHC molecules Immunogenicity VS Antigenicity:
Understanding the immune response to bacterial infections
 Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Methods for Protein Analysis
 Methods for Protein Analysis 1. Protein Separation Methods The following is a quick review of some common methods used for protein separation: SDS-PAGE (SDS-polyacrylamide gel electrophoresis) separates
Methods for Protein Analysis 1. Protein Separation Methods The following is a quick review of some common methods used for protein separation: SDS-PAGE (SDS-polyacrylamide gel electrophoresis) separates
Western Blot Analysis with Cell Samples Grown in Channel-µ-Slides
 Western Blot Analysis with Cell Samples Grown in Channel-µ-Slides Polyacrylamide gel electrophoresis (PAGE) and subsequent analyses are common tools in biochemistry and molecular biology. This Application
Western Blot Analysis with Cell Samples Grown in Channel-µ-Slides Polyacrylamide gel electrophoresis (PAGE) and subsequent analyses are common tools in biochemistry and molecular biology. This Application
The immune response Antibodies Antigens Epitopes (antigenic determinants) the part of a protein antigen recognized by an antibody Haptens small
 The immune response Antibodies Antigens Epitopes (antigenic determinants) the part of a protein antigen recognized by an antibody Haptens small molecules that can elicit an immune response when linked
The immune response Antibodies Antigens Epitopes (antigenic determinants) the part of a protein antigen recognized by an antibody Haptens small molecules that can elicit an immune response when linked
Anti-ATF6 α antibody, mouse monoclonal (1-7)
 Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
Classic Immunoprecipitation
 292PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Classic Immunoprecipitation Utilizes Protein A/G Agarose for Antibody Binding (Cat.
292PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Classic Immunoprecipitation Utilizes Protein A/G Agarose for Antibody Binding (Cat.
CD3/TCR stimulation and surface detection Determination of specificity of intracellular detection of IL-7Rα by flow cytometry
 CD3/TCR stimulation and surface detection Stimulation of HPB-ALL cells with the anti-cd3 monoclonal antibody OKT3 was performed as described 3. In brief, antibody-coated plates were prepared by incubating
CD3/TCR stimulation and surface detection Stimulation of HPB-ALL cells with the anti-cd3 monoclonal antibody OKT3 was performed as described 3. In brief, antibody-coated plates were prepared by incubating
TABLE OF CONTENT. Page ACKNOWLEDGEMENTS. iii ENGLISH ABSTRACT THAI ABSTRACT. vii LIST OF TABLES LIST OF FIGURES. xvi ABBREVIATIONS.
 x TABLE OF CONTENT ACKNOWLEDGEMENTS ENGLISH ABSTRACT THAI ABSTRACT LIST OF TABLES LIST OF FIGURES ABBREVIATIONS iii iv vii xv xvi xviii CHAPTER I: INTRODUCTION 1.1 Statement of problems 1 1.2 Literature
x TABLE OF CONTENT ACKNOWLEDGEMENTS ENGLISH ABSTRACT THAI ABSTRACT LIST OF TABLES LIST OF FIGURES ABBREVIATIONS iii iv vii xv xvi xviii CHAPTER I: INTRODUCTION 1.1 Statement of problems 1 1.2 Literature
THE His Tag Antibody, mab, Mouse
 THE His Tag Antibody, mab, Mouse Cat. No. A00186 Technical Manual No. TM0243 Update date 01052011 I Description.... 1 II Key Features. 2 III Storage 2 IV Applications.... 2 V Examples - ELISA..... 2 VI
THE His Tag Antibody, mab, Mouse Cat. No. A00186 Technical Manual No. TM0243 Update date 01052011 I Description.... 1 II Key Features. 2 III Storage 2 IV Applications.... 2 V Examples - ELISA..... 2 VI
Optimal Conditions for F(ab ) 2 Antibody Fragment Production from Mouse IgG2a
 Optimal Conditions for F(ab ) 2 Antibody Fragment Production from Mouse IgG2a Ryan S. Stowers, 1 Jacqueline A. Callihan, 2 James D. Bryers 2 1 Department of Bioengineering, Clemson University, Clemson,
Optimal Conditions for F(ab ) 2 Antibody Fragment Production from Mouse IgG2a Ryan S. Stowers, 1 Jacqueline A. Callihan, 2 James D. Bryers 2 1 Department of Bioengineering, Clemson University, Clemson,
WESTERN BLOTTING TIPS AND TROUBLESHOOTING GUIDE TROUBLESHOOTING GUIDE
 WESTERN BLOTTING TIPS AND TROUBLESHOOTING GUIDE TIPS FOR SUCCESSFUL WESTERB BLOTS TROUBLESHOOTING GUIDE 1. Suboptimal protein transfer. This is the most common complaint with western blotting and could
WESTERN BLOTTING TIPS AND TROUBLESHOOTING GUIDE TIPS FOR SUCCESSFUL WESTERB BLOTS TROUBLESHOOTING GUIDE 1. Suboptimal protein transfer. This is the most common complaint with western blotting and could
Covalent Conjugation to Cytodiagnostics Carboxylated Gold Nanoparticles Tech Note #105
 Covalent Conjugation to Cytodiagnostics Carboxylated Gold Nanoparticles Tech Note #105 Background Gold nanoparticle conjugates have been widely used in biological research and biosensing applications.
Covalent Conjugation to Cytodiagnostics Carboxylated Gold Nanoparticles Tech Note #105 Background Gold nanoparticle conjugates have been widely used in biological research and biosensing applications.
Running protein gels and detection of proteins
 Running protein gels and detection of proteins 1. Protein concentration determination using the BIO RAD reagent This assay uses a colour change reaction to give a direct measurement of protein concentration.
Running protein gels and detection of proteins 1. Protein concentration determination using the BIO RAD reagent This assay uses a colour change reaction to give a direct measurement of protein concentration.
Chapter 2 Antibodies. Contents. Introduction
 Chapter 2 Antibodies Keywords Immunohistochemistry Antibody labeling Fluorescence microscopy Fluorescent immunocytochemistry Fluorescent immunohistochemistry Indirect immunocytochemistry Immunostaining
Chapter 2 Antibodies Keywords Immunohistochemistry Antibody labeling Fluorescence microscopy Fluorescent immunocytochemistry Fluorescent immunohistochemistry Indirect immunocytochemistry Immunostaining
Investigating the role of a Cryptosporidium parum apyrase in infection
 Investigating the role of a Cryptosporidium parum apyrase in infection David Riccardi and Patricio Manque Abstract This project attempted to characterize the function of a Cryptosporidium parvum apyrase
Investigating the role of a Cryptosporidium parum apyrase in infection David Riccardi and Patricio Manque Abstract This project attempted to characterize the function of a Cryptosporidium parvum apyrase
Peptide synthesis, radiolabelling and radiochemical analysis
 SUPPLEMENTAL DATA MATERIALS AND METHODS Peptide synthesis, radiolabelling and radiochemical analysis Solid phase synthesis of peptides was carried out on using ABI 433A peptide synthesizer, on a preloaded
SUPPLEMENTAL DATA MATERIALS AND METHODS Peptide synthesis, radiolabelling and radiochemical analysis Solid phase synthesis of peptides was carried out on using ABI 433A peptide synthesizer, on a preloaded
Lecture 8. Protein Trafficking/Targeting. Protein targeting is necessary for proteins that are destined to work outside the cytoplasm.
 Protein Trafficking/Targeting (8.1) Lecture 8 Protein Trafficking/Targeting Protein targeting is necessary for proteins that are destined to work outside the cytoplasm. Protein targeting is more complex
Protein Trafficking/Targeting (8.1) Lecture 8 Protein Trafficking/Targeting Protein targeting is necessary for proteins that are destined to work outside the cytoplasm. Protein targeting is more complex
Pure-IP Western Blot Detection Kit
 Product Manual Pure-IP Western Blot Detection Kit Catalog Number PRB-5002 20 blots FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction The technique of immunoprecipitation (IP) is used
Product Manual Pure-IP Western Blot Detection Kit Catalog Number PRB-5002 20 blots FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction The technique of immunoprecipitation (IP) is used
Two-Dimensional Gel Electrophoresis (2-DGE)
 - Introduction - Sample preparation - First dimension: Isoelectric focusing - Second dimension: SDS-PAGE - Detection of protein spots: staining - Imaging analysis & 2D Gel databases - Spot handling: excision,
- Introduction - Sample preparation - First dimension: Isoelectric focusing - Second dimension: SDS-PAGE - Detection of protein spots: staining - Imaging analysis & 2D Gel databases - Spot handling: excision,
KMS-Specialist & Customized Biosimilar Service
 KMS-Specialist & Customized Biosimilar Service 1. Polyclonal Antibody Development Service KMS offering a variety of Polyclonal Antibody Services to fit your research and production needs. we develop polyclonal
KMS-Specialist & Customized Biosimilar Service 1. Polyclonal Antibody Development Service KMS offering a variety of Polyclonal Antibody Services to fit your research and production needs. we develop polyclonal
CHAPTER 2 ANTIGEN/ANTIBODY INTERACTIONS
 CHAPTER 2 ANTIGEN/ANTIBODY INTERACTIONS See APPENDIX (1) THE PRECIPITIN CURVE; (2) LABELING OF ANTIBODIES The defining characteristic of HUMORAL immune responses (which distinguishes them from CELL-MEDIATED
CHAPTER 2 ANTIGEN/ANTIBODY INTERACTIONS See APPENDIX (1) THE PRECIPITIN CURVE; (2) LABELING OF ANTIBODIES The defining characteristic of HUMORAL immune responses (which distinguishes them from CELL-MEDIATED
Protocol for Western Blotting
 Protocol for Western Blotting Materials Materials used on Day 3 Protease inhibitor stock: 1 μg/μl pepstatin A in DMSO 200 μm leupeptin in OG Buffer 200 mm PMSF: Freshly made. Ex) 34.8 mg PMSF in 1 ml isopropanol
Protocol for Western Blotting Materials Materials used on Day 3 Protease inhibitor stock: 1 μg/μl pepstatin A in DMSO 200 μm leupeptin in OG Buffer 200 mm PMSF: Freshly made. Ex) 34.8 mg PMSF in 1 ml isopropanol
Microarray Technology
 Microarrays And Functional Genomics CPSC265 Matt Hudson Microarray Technology Relatively young technology Usually used like a Northern blot can determine the amount of mrna for a particular gene Except
Microarrays And Functional Genomics CPSC265 Matt Hudson Microarray Technology Relatively young technology Usually used like a Northern blot can determine the amount of mrna for a particular gene Except
PROTOCOL 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 www.mitosciences.com
 PROTOCOL Western Blotting Transfer and Detection Procedure 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 02-11 DESCRIPTION Western Blotting Transfer and Detection Procedure ADDITIONAL MATERIALS REQUIRED
PROTOCOL Western Blotting Transfer and Detection Procedure 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 02-11 DESCRIPTION Western Blotting Transfer and Detection Procedure ADDITIONAL MATERIALS REQUIRED
Western Blotting: Mini-gels
 Western Blotting: Mini-gels Materials a Protein Extraction Buffer (for callus or kernel), Solution Stock Final Volume Tris-HCl ph 80 1 M 200 mm 20 ml NaCl 4 M 100 mm 25 ml Sucrose 2 M 400 mm 20 ml EDTA
Western Blotting: Mini-gels Materials a Protein Extraction Buffer (for callus or kernel), Solution Stock Final Volume Tris-HCl ph 80 1 M 200 mm 20 ml NaCl 4 M 100 mm 25 ml Sucrose 2 M 400 mm 20 ml EDTA
Biology 309 Lab Notebook
 Name: Biology 309 Lab Notebook This is a guided lab notebook for you to keep well-organized notes about procedures and record experimental data for experiments as they are performed. It is guided because,
Name: Biology 309 Lab Notebook This is a guided lab notebook for you to keep well-organized notes about procedures and record experimental data for experiments as they are performed. It is guided because,
Protein extraction from Tissues and Cultured Cells using Bioruptor Standard & Plus
 Protein extraction from Tissues and Cultured Cells using Bioruptor Standard & Plus Introduction Protein extraction from tissues and cultured cells is the first step for many biochemical and analytical
Protein extraction from Tissues and Cultured Cells using Bioruptor Standard & Plus Introduction Protein extraction from tissues and cultured cells is the first step for many biochemical and analytical
ELISA BIO 110 Lab 1. Immunity and Disease
 ELISA BIO 110 Lab 1 Immunity and Disease Introduction The principal role of the mammalian immune response is to contain infectious disease agents. This response is mediated by several cellular and molecular
ELISA BIO 110 Lab 1 Immunity and Disease Introduction The principal role of the mammalian immune response is to contain infectious disease agents. This response is mediated by several cellular and molecular
TEMA 10. REACCIONES INMUNITARIAS MEDIADAS POR CÉLULAS.
 TEMA 10. REACCIONES INMUNITARIAS MEDIADAS POR CÉLULAS. The nomenclature of cytokines partly reflects their first-described function and also the order of their discovery. There is no single unified nomenclature,
TEMA 10. REACCIONES INMUNITARIAS MEDIADAS POR CÉLULAS. The nomenclature of cytokines partly reflects their first-described function and also the order of their discovery. There is no single unified nomenclature,
Some terms: An antigen is a molecule or pathogen capable of eliciting an immune response
 Overview of the immune system We continue our discussion of protein structure by considering the structure of antibodies. All organisms are continually subject to attack by microorganisms and viruses.
Overview of the immune system We continue our discussion of protein structure by considering the structure of antibodies. All organisms are continually subject to attack by microorganisms and viruses.
Basics of Immunology
 Basics of Immunology 2 Basics of Immunology What is the immune system? Biological mechanism for identifying and destroying pathogens within a larger organism. Pathogens: agents that cause disease Bacteria,
Basics of Immunology 2 Basics of Immunology What is the immune system? Biological mechanism for identifying and destroying pathogens within a larger organism. Pathogens: agents that cause disease Bacteria,
An In-Gel Digestion Protocol
 An In-Gel Digestion Protocol This protocol describes the digestion of a protein present in an SDS-PAGE gel band with trypsin. The band can be taken from either a 1D or 2D electrophoresis gel. Reagents
An In-Gel Digestion Protocol This protocol describes the digestion of a protein present in an SDS-PAGE gel band with trypsin. The band can be taken from either a 1D or 2D electrophoresis gel. Reagents
TECHNICAL BULLETIN. HIS-Select Nickel Affinity Gel. Catalog Number P6611 Storage Temperature 2 8 C
 HIS-Select Nickel Affinity Gel Catalog Number P6611 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description HIS-Select Nickel Affinity Gel is an immobilized metalion affinity chromatography (IMAC)
HIS-Select Nickel Affinity Gel Catalog Number P6611 Storage Temperature 2 8 C TECHNICAL BULLETIN Product Description HIS-Select Nickel Affinity Gel is an immobilized metalion affinity chromatography (IMAC)
B Cell Generation, Activation & Differentiation. B cell maturation
 B Cell Generation, Activation & Differentiation Naïve B cells- have not encountered Ag. Have IgM and IgD on cell surface : have same binding VDJ regions but different constant region leaves bone marrow
B Cell Generation, Activation & Differentiation Naïve B cells- have not encountered Ag. Have IgM and IgD on cell surface : have same binding VDJ regions but different constant region leaves bone marrow
STANDARD OPERATING PROCEDURE
 STANDARD OPERATING PROCEDURE Title: Evaluation using Western Blot SOP#: M-103 Version #: 1 Author: R. Saul Date Approved: Feb. 5, 2009 Date Modified: 1. PURPOSE The purpose of this document is to describe
STANDARD OPERATING PROCEDURE Title: Evaluation using Western Blot SOP#: M-103 Version #: 1 Author: R. Saul Date Approved: Feb. 5, 2009 Date Modified: 1. PURPOSE The purpose of this document is to describe
Chapter 3. Protein Structure and Function
 Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Student name ID # 2. (4 pts) What is the terminal electron acceptor in respiration? In photosynthesis? O2, NADP+
 1. Membrane transport. A. (4 pts) What ion couples primary and secondary active transport in animal cells? What ion serves the same function in plant cells? Na+, H+ 2. (4 pts) What is the terminal electron
1. Membrane transport. A. (4 pts) What ion couples primary and secondary active transport in animal cells? What ion serves the same function in plant cells? Na+, H+ 2. (4 pts) What is the terminal electron
Dot Blot Analysis. Teacher s Guidebook. (Cat. # BE 502) think proteins! think G-Biosciences www.gbiosciences.com
 PR110 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Dot Blot Analysis Teacher s Guidebook (Cat. # BE 502) think proteins! think G-Biosciences
PR110 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Dot Blot Analysis Teacher s Guidebook (Cat. # BE 502) think proteins! think G-Biosciences
6 Characterization of Casein and Bovine Serum Albumin
 6 Characterization of Casein and Bovine Serum Albumin (BSA) Objectives: A) To separate a mixture of casein and bovine serum albumin B) to characterize these proteins based on their solubilities as a function
6 Characterization of Casein and Bovine Serum Albumin (BSA) Objectives: A) To separate a mixture of casein and bovine serum albumin B) to characterize these proteins based on their solubilities as a function
specific B cells Humoral immunity lymphocytes antibodies B cells bone marrow Cell-mediated immunity: T cells antibodies proteins
 Adaptive Immunity Chapter 17: Adaptive (specific) Immunity Bio 139 Dr. Amy Rogers Host defenses that are specific to a particular infectious agent Can be innate or genetic for humans as a group: most microbes
Adaptive Immunity Chapter 17: Adaptive (specific) Immunity Bio 139 Dr. Amy Rogers Host defenses that are specific to a particular infectious agent Can be innate or genetic for humans as a group: most microbes
2D gel Protocol. 2. Determining Protein Concentration of cell lysates
 2D gel Protocol 1. Lysis and Protein Extraction from cells Prepare cell lysates with Trizol extraction by following Kathleen Lyons s protocol: AfCS Procedure Protocol PP00000155, Version 1, 05/12/03 (Ref.1).
2D gel Protocol 1. Lysis and Protein Extraction from cells Prepare cell lysates with Trizol extraction by following Kathleen Lyons s protocol: AfCS Procedure Protocol PP00000155, Version 1, 05/12/03 (Ref.1).
Product name Company Cat # PowerPac Basic Power supply Bio Rad 165-6019 Mini Protean electrophoresis system Mini trans blot cell Bio Rad 170-3930
 SDS-PAGE and western blot for low molecular weight proteins (2-20kDa) Merav Marom Shamur, Smart Assays Aim: Analysis of low molecular weight proteins by SDS-PAGE and western blot under reducing conditions.
SDS-PAGE and western blot for low molecular weight proteins (2-20kDa) Merav Marom Shamur, Smart Assays Aim: Analysis of low molecular weight proteins by SDS-PAGE and western blot under reducing conditions.
Chapter 16: Innate Immunity
 Chapter 16: Innate Immunity 1. Overview of Innate Immunity 2. Inflammation & Phagocytosis 3. Antimicrobial Substances 1. Overview of Innate Immunity The Body s Defenses The body has 2 types of defense
Chapter 16: Innate Immunity 1. Overview of Innate Immunity 2. Inflammation & Phagocytosis 3. Antimicrobial Substances 1. Overview of Innate Immunity The Body s Defenses The body has 2 types of defense
EZ-Run Protein Gel Solution. EZ-Run Protein Standards. EZ-Run Gel Staining Solution. Traditional SDS-Page Reagents. Protein Electrophoresis
 EZ-Run Protein Gel Solution EZ-Run Protein Standards EZ-Run Gel Staining Solution Traditional SDS-Page Reagents Protein Electrophoresis protein electrophoresis Introduction Sodium dodecyl sulfate polyacrylamide
EZ-Run Protein Gel Solution EZ-Run Protein Standards EZ-Run Gel Staining Solution Traditional SDS-Page Reagents Protein Electrophoresis protein electrophoresis Introduction Sodium dodecyl sulfate polyacrylamide
Molecular Cell Biology. Prof. D. Karunagaran. Department of Biotechnology. Indian Institute of Technology Madras
 Molecular Cell Biology Prof. D. Karunagaran Department of Biotechnology Indian Institute of Technology Madras Module 5 Methods in Cell Biology (Methods to Manipulate Protein, DNA and RNA and Methods to
Molecular Cell Biology Prof. D. Karunagaran Department of Biotechnology Indian Institute of Technology Madras Module 5 Methods in Cell Biology (Methods to Manipulate Protein, DNA and RNA and Methods to
ReadyPrep Protein Extraction Kit (Soluble/Insoluble) Instruction Manual. Catalog #163-2085
 ReadyPrep Protein Extraction Kit (Soluble/Insoluble) Instruction Manual Catalog #163-2085 For technical service, call your local Bio-Rad office, or in the US, call 1-800-4BIORAD (1-800-424-6723) Table
ReadyPrep Protein Extraction Kit (Soluble/Insoluble) Instruction Manual Catalog #163-2085 For technical service, call your local Bio-Rad office, or in the US, call 1-800-4BIORAD (1-800-424-6723) Table
Chapter 18: Applications of Immunology
 Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
In-Depth Qualitative Analysis of Complex Proteomic Samples Using High Quality MS/MS at Fast Acquisition Rates
 In-Depth Qualitative Analysis of Complex Proteomic Samples Using High Quality MS/MS at Fast Acquisition Rates Using the Explore Workflow on the AB SCIEX TripleTOF 5600 System A major challenge in proteomics
In-Depth Qualitative Analysis of Complex Proteomic Samples Using High Quality MS/MS at Fast Acquisition Rates Using the Explore Workflow on the AB SCIEX TripleTOF 5600 System A major challenge in proteomics
Introduction to Proteomics
 Introduction to Proteomics Why Proteomics? Same Genome Different Proteome Black Swallowtail - larvae and butterfly Biological Complexity Yeast - a simple proteome 6,113 proteins = 344,855 tryptic peptides
Introduction to Proteomics Why Proteomics? Same Genome Different Proteome Black Swallowtail - larvae and butterfly Biological Complexity Yeast - a simple proteome 6,113 proteins = 344,855 tryptic peptides
Chapter 3.2» Custom Monoclonal
 198 3 3.2 Custom Monoclonal 199 Mouse monoclonal antibody development Chapter 3.2» Custom Monoclonal 200 In vitro monoclonals expression service 201 Mouse monoclonal antibody additional services 202 Magnetic
198 3 3.2 Custom Monoclonal 199 Mouse monoclonal antibody development Chapter 3.2» Custom Monoclonal 200 In vitro monoclonals expression service 201 Mouse monoclonal antibody additional services 202 Magnetic
RPCI 004 v.002 Staining Procedure For all Directly Conjugated Reagents (Whole Blood Method)
 Immune Tolerance Network RPCI 004 v.002 Staining Procedure For all Directly Conjugated Reagents (Whole Blood Method) Author: Paul Wallace, Director, RPCI Laboratory of Flow Cytometry Approved by: Paul
Immune Tolerance Network RPCI 004 v.002 Staining Procedure For all Directly Conjugated Reagents (Whole Blood Method) Author: Paul Wallace, Director, RPCI Laboratory of Flow Cytometry Approved by: Paul
Proteomics in Practice
 Reiner Westermeier, Torn Naven Hans-Rudolf Höpker Proteomics in Practice A Guide to Successful Experimental Design 2008 Wiley-VCH Verlag- Weinheim 978-3-527-31941-1 Preface Foreword XI XIII Abbreviations,
Reiner Westermeier, Torn Naven Hans-Rudolf Höpker Proteomics in Practice A Guide to Successful Experimental Design 2008 Wiley-VCH Verlag- Weinheim 978-3-527-31941-1 Preface Foreword XI XIII Abbreviations,
Western Blotting. USA: proteintech@ptglab.com UK & Europe: europe@ptglab.com China: service@ptglab.com. www.ptglab.com
 Western Blotting All steps are carried out at room temperature unless otherwise indicated. Recipes for all solutions highlighted bold are included at the end of the protocol. SDS-PAGE 1. Construct an SDS-PAGE
Western Blotting All steps are carried out at room temperature unless otherwise indicated. Recipes for all solutions highlighted bold are included at the end of the protocol. SDS-PAGE 1. Construct an SDS-PAGE
Protein immunoblotting
 Protein immunoblotting (Western blotting) Dr. Serageldeen A. A. Sultan Lecturer of virology Dept. of Microbiology SVU, Qena, Egypt seaas@lycos.com Western blotting -It is an analytical technique used to
Protein immunoblotting (Western blotting) Dr. Serageldeen A. A. Sultan Lecturer of virology Dept. of Microbiology SVU, Qena, Egypt seaas@lycos.com Western blotting -It is an analytical technique used to
Protein transfer from SDS-PAGE to nitrocellulose membrane using the Trans-Blot SD cell (Western).
 Western Blot SOP Protein transfer from SDS-PAGE to nitrocellulose membrane using the Trans-Blot SD cell (Western). Date: 8/16/05, 10/31/05, 2/6/06 Author: N.Oganesyan, R. Kim Edited by: R. Kim Summary:
Western Blot SOP Protein transfer from SDS-PAGE to nitrocellulose membrane using the Trans-Blot SD cell (Western). Date: 8/16/05, 10/31/05, 2/6/06 Author: N.Oganesyan, R. Kim Edited by: R. Kim Summary:
Rubisco; easy Purification and Immunochemical Determination
 Rubisco; easy Purification and Immunochemical Determination Ulrich Groß Justus-Liebig-Universität Gießen, Institute of Plant Nutrition, Department of Tissue Culture, Südanlage 6, D-35390 Giessen e-mail:
Rubisco; easy Purification and Immunochemical Determination Ulrich Groß Justus-Liebig-Universität Gießen, Institute of Plant Nutrition, Department of Tissue Culture, Südanlage 6, D-35390 Giessen e-mail:
Notch 1 -dependent regulation of cell fate in colorectal cancer
 Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
Notch 1 -dependent regulation of cell fate in colorectal cancer Referees: PD Dr. Tobias Dick Prof. Dr. Wilfried Roth http://d-nb.info/1057851272 CONTENTS Summary 1 Zusammenfassung 2 1 INTRODUCTION 3 1.1
CUSTOM ANTIBODIES. Fully customised services: rat and murine monoclonals, rat and rabbit polyclonals, antibody characterisation, antigen preparation
 CUSTOM ANTIBODIES Highly competitive pricing without compromising quality. Rat monoclonal antibodies for the study of gene expression and proteomics in mice and in mouse models of human diseases available.
CUSTOM ANTIBODIES Highly competitive pricing without compromising quality. Rat monoclonal antibodies for the study of gene expression and proteomics in mice and in mouse models of human diseases available.
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
Introduction to Proteomics
 Introduction to Proteomics Åsa Wheelock, Ph.D. Division of Respiratory Medicine & Karolinska Biomics Center asa.wheelock@ki.se In: Systems Biology and the Omics Cascade, Karolinska Institutet, June 9-13,
Introduction to Proteomics Åsa Wheelock, Ph.D. Division of Respiratory Medicine & Karolinska Biomics Center asa.wheelock@ki.se In: Systems Biology and the Omics Cascade, Karolinska Institutet, June 9-13,
Protease Peptide Microarrays Ready-to-use microarrays for protease profiling
 Protocol Protease Peptide Microarrays Ready-to-use microarrays for protease profiling Contact us: InfoLine: +49-30-97893-117 Order per fax: +49-30-97893-299 Or e-mail: peptide@jpt.com www: www.jpt.com
Protocol Protease Peptide Microarrays Ready-to-use microarrays for protease profiling Contact us: InfoLine: +49-30-97893-117 Order per fax: +49-30-97893-299 Or e-mail: peptide@jpt.com www: www.jpt.com
Transfection reagent for visualizing lipofection with DNA. For ordering information, MSDS, publications and application notes see www.biontex.
 METAFECTENE FluoR Transfection reagent for visualizing lipofection with DNA For ordering information, MSDS, publications and application notes see www.biontex.com Description Cat. No. Size METAFECTENE
METAFECTENE FluoR Transfection reagent for visualizing lipofection with DNA For ordering information, MSDS, publications and application notes see www.biontex.com Description Cat. No. Size METAFECTENE
SDS-PAGE Protocol Mutated from the SDS-PAGE protocol written by the Lord of the Flies
 SDS-PAGE Protocol Mutated from the SDS-PAGE protocol written by the Lord of the Flies Pouring the resolving gel 1. Clean glass plates with soap and water, then with ethanol. Assemble the glass plates and
SDS-PAGE Protocol Mutated from the SDS-PAGE protocol written by the Lord of the Flies Pouring the resolving gel 1. Clean glass plates with soap and water, then with ethanol. Assemble the glass plates and
Monoclonal Antibody Fragment Separation and Characterization Using Size Exclusion Chromatography Coupled with Mass Spectrometry
 Monoclonal ntibody Fragment Separation and haracterization Using Size Exclusion hromatography oupled with Mass Spectrometry uthors Haiying hen Katherine McLaughlin Sepax Technologies, Inc. 5 Innovation
Monoclonal ntibody Fragment Separation and haracterization Using Size Exclusion hromatography oupled with Mass Spectrometry uthors Haiying hen Katherine McLaughlin Sepax Technologies, Inc. 5 Innovation
ECL Western Blotting Substrate INSTRUCTIONS FOR USE OF PRODUCTS W1001 AND W1015.
 Technical Manual ECL Western Blotting Substrate INSTRUCTIONS FOR USE OF PRODUCTS W1001 AND W1015. PRINTED IN USA. 6/09 ECL Western Blotting Substrate All technical literature is available on the Internet
Technical Manual ECL Western Blotting Substrate INSTRUCTIONS FOR USE OF PRODUCTS W1001 AND W1015. PRINTED IN USA. 6/09 ECL Western Blotting Substrate All technical literature is available on the Internet
1.1.2. thebiotutor. AS Biology OCR. Unit F211: Cells, Exchange & Transport. Module 1.2 Cell Membranes. Notes & Questions.
 thebiotutor AS Biology OCR Unit F211: Cells, Exchange & Transport Module 1.2 Cell Membranes Notes & Questions Andy Todd 1 Outline the roles of membranes within cells and at the surface of cells. The main
thebiotutor AS Biology OCR Unit F211: Cells, Exchange & Transport Module 1.2 Cell Membranes Notes & Questions Andy Todd 1 Outline the roles of membranes within cells and at the surface of cells. The main
PROTOCOL. Immunocytochemistry (ICC) MATERIALS AND EQUIPMENT REQUIRED
 PROTOCOL Immunocytochemistry (ICC) 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 11-07 MATERIALS AND EQUIPMENT REQUIRED Materials: MitoSciences primary monoclonal antibody/antibodies Fluorophore-conjugated
PROTOCOL Immunocytochemistry (ICC) 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 11-07 MATERIALS AND EQUIPMENT REQUIRED Materials: MitoSciences primary monoclonal antibody/antibodies Fluorophore-conjugated
Western Blot Protocol Protein isolation
 Western Blot Protocol Protein isolation A. Preparation of cell lysates. - Preparation of materials: -Dial the microcentrifuge temperature control setting to 4 C -Prepare a bucket of ice -Prepare lysis
Western Blot Protocol Protein isolation A. Preparation of cell lysates. - Preparation of materials: -Dial the microcentrifuge temperature control setting to 4 C -Prepare a bucket of ice -Prepare lysis
Cell Biology Questions and Learning Objectives
 Cell Biology Questions and Learning Objectives (with hypothetical learning materials that might populate the objective) The topics and central questions listed here are typical for an introductory undergraduate
Cell Biology Questions and Learning Objectives (with hypothetical learning materials that might populate the objective) The topics and central questions listed here are typical for an introductory undergraduate
ab185915 Protein Sumoylation Assay Ultra Kit
 ab185915 Protein Sumoylation Assay Ultra Kit Instructions for Use For the measuring in vivo protein sumoylation in various samples This product is for research use only and is not intended for diagnostic
ab185915 Protein Sumoylation Assay Ultra Kit Instructions for Use For the measuring in vivo protein sumoylation in various samples This product is for research use only and is not intended for diagnostic
Western Blotting. Prepare samples:
 Western Blotting Sive Lab Protocol March 2007 Prepare samples: For zebrafish embryos: Option 1: Take live embryos and put into 1.5 ml tube with E3. Centrifuge gently for 1-2 minutes -yolk lipids will rise
Western Blotting Sive Lab Protocol March 2007 Prepare samples: For zebrafish embryos: Option 1: Take live embryos and put into 1.5 ml tube with E3. Centrifuge gently for 1-2 minutes -yolk lipids will rise
Chromatin Immunoprecipitation (ChIP)
 Chromatin Immunoprecipitation (ChIP) Day 1 A) DNA shearing 1. Samples Dissect tissue (One Mouse OBs) of interest and transfer to an eppendorf containing 0.5 ml of dissecting media (on ice) or PBS but without
Chromatin Immunoprecipitation (ChIP) Day 1 A) DNA shearing 1. Samples Dissect tissue (One Mouse OBs) of interest and transfer to an eppendorf containing 0.5 ml of dissecting media (on ice) or PBS but without
Seahorse XF Cell Mito Stress Test Kit
 Seahorse XF Cell Mito Stress Test Kit Part # 103015-100 User Guide For use with Seahorse XF e and XF Extracellular Flux Analyzers For Research Use Only 1 Table of Contents Product Description... 2 Introduction...
Seahorse XF Cell Mito Stress Test Kit Part # 103015-100 User Guide For use with Seahorse XF e and XF Extracellular Flux Analyzers For Research Use Only 1 Table of Contents Product Description... 2 Introduction...
Name (print) Name (signature) Period. (Total 30 points)
 AP Biology Worksheet Chapter 43 The Immune System Lambdin April 4, 2011 Due Date: Thurs. April 7, 2011 You may use the following: Text Notes Power point Internet One other person in class "On my honor,
AP Biology Worksheet Chapter 43 The Immune System Lambdin April 4, 2011 Due Date: Thurs. April 7, 2011 You may use the following: Text Notes Power point Internet One other person in class "On my honor,
LESSON 3: ANTIBODIES/BCR/B-CELL RESPONSES
 Introduction to immunology. LESSON 3: ANTIBODIES/BCR/B-CELL RESPONSES Today we will get to know: The antibodies How antibodies are produced, their classes and their maturation processes Antigen recognition
Introduction to immunology. LESSON 3: ANTIBODIES/BCR/B-CELL RESPONSES Today we will get to know: The antibodies How antibodies are produced, their classes and their maturation processes Antigen recognition
Genomic DNA Extraction Kit INSTRUCTION MANUAL
 Genomic DNA Extraction Kit INSTRUCTION MANUAL Table of Contents Introduction 3 Kit Components 3 Storage Conditions 4 Recommended Equipment and Reagents 4 Introduction to the Protocol 4 General Overview
Genomic DNA Extraction Kit INSTRUCTION MANUAL Table of Contents Introduction 3 Kit Components 3 Storage Conditions 4 Recommended Equipment and Reagents 4 Introduction to the Protocol 4 General Overview
Western Blot Protocol (updated on 05/20/14)
 Western Blot Protocol (updated on 05/20/14) Required Solutions 10x PBS (1L) 80 g NaCl 2 g KCl 14.4 g Na 2 HPO 4 or 22 g Na 2 HPO 4 7H 2 O 2.4 g KH 2 PO 4 or 2 g KH 2 PO4 Adjust ph to 7.4 Autoclave PBST
Western Blot Protocol (updated on 05/20/14) Required Solutions 10x PBS (1L) 80 g NaCl 2 g KCl 14.4 g Na 2 HPO 4 or 22 g Na 2 HPO 4 7H 2 O 2.4 g KH 2 PO 4 or 2 g KH 2 PO4 Adjust ph to 7.4 Autoclave PBST
Accelerated Electrophoresis with Faster Run Times and Increased Sample Throughput
 ELECTROPHORESIS Criterion TGX Precast Gels Get superior results with convenient, fast Criterion TGX gels: Run in as little as min Blot transfer times as short as min Use standard Laemmli buffers and reagents
ELECTROPHORESIS Criterion TGX Precast Gels Get superior results with convenient, fast Criterion TGX gels: Run in as little as min Blot transfer times as short as min Use standard Laemmli buffers and reagents
Mouse IFN-gamma ELISpot Kit
 Page 1 of 8 Mouse IFN-gamma ELISpot Kit Without Plates With Plates With Sterile Plates Quantity Catalog Nos. 862.031.001 862.031.001P 862.031.001S 1 x 96 tests 862.031.005 862.031.005P 862.031.005S 5 x
Page 1 of 8 Mouse IFN-gamma ELISpot Kit Without Plates With Plates With Sterile Plates Quantity Catalog Nos. 862.031.001 862.031.001P 862.031.001S 1 x 96 tests 862.031.005 862.031.005P 862.031.005S 5 x
First Strand cdna Synthesis
 380PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name First Strand cdna Synthesis (Cat. # 786 812) think proteins! think G-Biosciences
380PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name First Strand cdna Synthesis (Cat. # 786 812) think proteins! think G-Biosciences
Microbiology AN INTRODUCTION EIGHTH EDITION
 TORTORA FUNKE CASE Microbiology AN INTRODUCTION EIGHTH EDITION Differentiate between innate and acquired immunity. Chapter 17 Specific Defenses of the Host: The Immune Response B.E Pruitt & Jane J. Stein
TORTORA FUNKE CASE Microbiology AN INTRODUCTION EIGHTH EDITION Differentiate between innate and acquired immunity. Chapter 17 Specific Defenses of the Host: The Immune Response B.E Pruitt & Jane J. Stein
Approaches that can be used to study expression of specific proteins
 Approaches that can be used to study expression of specific proteins Receptors and transporters Homogenate binding studies Receptor autoradiography Radiochemical Western blotting Immunohistochemistry/cytochemistry
Approaches that can be used to study expression of specific proteins Receptors and transporters Homogenate binding studies Receptor autoradiography Radiochemical Western blotting Immunohistochemistry/cytochemistry
Your partner in immunology
 Your partner in immunology Expertise Expertise Reactivity Reactivity Quality Quality Advice Advice Who are we? Specialist of antibody engineering Covalab is a French biotechnology company, specialised
Your partner in immunology Expertise Expertise Reactivity Reactivity Quality Quality Advice Advice Who are we? Specialist of antibody engineering Covalab is a French biotechnology company, specialised
T Cell Maturation,Activation and Differentiation
 T Cell Maturation,Activation and Differentiation Positive Selection- In thymus, permits survival of only those T cells whose TCRs recognize self- MHC molecules (self-mhc restriction) Negative Selection-
T Cell Maturation,Activation and Differentiation Positive Selection- In thymus, permits survival of only those T cells whose TCRs recognize self- MHC molecules (self-mhc restriction) Negative Selection-
Topic: Serological reactions: the purpose and a principle of reactions. Agglutination test. Precipitation test. CFT, IFT, ELISA, RIA.
 Topic: Serological reactions: the purpose and a principle of reactions. Agglutination test. Precipitation test. CFT, IFT, ELISA, RIA. Serology is the study and use of immunological tests to diagnose and
Topic: Serological reactions: the purpose and a principle of reactions. Agglutination test. Precipitation test. CFT, IFT, ELISA, RIA. Serology is the study and use of immunological tests to diagnose and
APPLICATION FOCUS. Application Solutions for Western Blotting
 APPLICATION FOCUS Application Solutions for Western Blotting WESTERN BLOTTING Companion Products Solutions for consistently better blotting. Companion products from Cell Signaling Technology (CST) are
APPLICATION FOCUS Application Solutions for Western Blotting WESTERN BLOTTING Companion Products Solutions for consistently better blotting. Companion products from Cell Signaling Technology (CST) are
CONTENT. Chapter 1 Review of Literature. List of figures. List of tables
 Abstract Abbreviations List of figures CONTENT I-VI VII-VIII IX-XII List of tables XIII Chapter 1 Review of Literature 1. Vaccination against intracellular pathogens 1-34 1.1 Role of different immune responses
Abstract Abbreviations List of figures CONTENT I-VI VII-VIII IX-XII List of tables XIII Chapter 1 Review of Literature 1. Vaccination against intracellular pathogens 1-34 1.1 Role of different immune responses
Supplemental Information. McBrayer et al. Supplemental Data
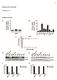 1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
1 Supplemental Information McBrayer et al. Supplemental Data 2 Figure S1. Glucose consumption rates of MM cell lines exceed that of normal PBMC. (A) Normal PBMC isolated from three healthy donors were
STANDARD OPERATING PROCEDURE
 STANDARD OPERATING PROCEDURE Title: Antibody Production at Strategic Diagnostics Inc. SOP#: M-119 Version #: 1 Date Approved: August 6, 2009 Author: Strategic Diagnostic Inc. Date Modified: 1. PURPOSE
STANDARD OPERATING PROCEDURE Title: Antibody Production at Strategic Diagnostics Inc. SOP#: M-119 Version #: 1 Date Approved: August 6, 2009 Author: Strategic Diagnostic Inc. Date Modified: 1. PURPOSE
Protein Analysis. -Detection and quantification. Toby M Holmes
 Protein Analysis -Detection and quantification Toby M Holmes Clinical Research Unit UCD school of Medicine and Medical Sciences Mater Misericordiae University Hospital Dublin Protein structure General
Protein Analysis -Detection and quantification Toby M Holmes Clinical Research Unit UCD school of Medicine and Medical Sciences Mater Misericordiae University Hospital Dublin Protein structure General
Supplementary Materials and Methods (Metabolomics analysis)
 Supplementary Materials and Methods (Metabolomics analysis) Metabolite extraction and mass spectrometry analysis. 12 Vldlr-/- knock out and 12 wild type (WT) retinas were separately frozen at -80 ºC in
Supplementary Materials and Methods (Metabolomics analysis) Metabolite extraction and mass spectrometry analysis. 12 Vldlr-/- knock out and 12 wild type (WT) retinas were separately frozen at -80 ºC in
Introduction to Proteomics 1.0
 Introduction to Proteomics 1.0 CMSP Workshop Tim Griffin Associate Professor, BMBB Faculty Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s
Introduction to Proteomics 1.0 CMSP Workshop Tim Griffin Associate Professor, BMBB Faculty Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s
HUMORAL IMMUNE RE- SPONSES: ACTIVATION OF B CELLS AND ANTIBODIES JASON CYSTER SECTION 13
 SECTION 13 HUMORAL IMMUNE RE- SPONSES: ACTIVATION OF B CELLS AND ANTIBODIES CONTACT INFORMATION Jason Cyster, PhD (Email) READING Basic Immunology: Functions and Disorders of the Immune System. Abbas,
SECTION 13 HUMORAL IMMUNE RE- SPONSES: ACTIVATION OF B CELLS AND ANTIBODIES CONTACT INFORMATION Jason Cyster, PhD (Email) READING Basic Immunology: Functions and Disorders of the Immune System. Abbas,
Workshop 14-16 February 2006
 Theoretical and practical approaches of Hepatocyte primary culture Workshop 14-16 February 2006 Lecture (2) Disaggregation & purification of target cells Coarse organizer Dr. Abo bakr Mohamed Eltayeb General
Theoretical and practical approaches of Hepatocyte primary culture Workshop 14-16 February 2006 Lecture (2) Disaggregation & purification of target cells Coarse organizer Dr. Abo bakr Mohamed Eltayeb General
2007 7.013 Problem Set 1 KEY
 2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
PROTOCOL. Immunostaining for Flow Cytometry. Background. Materials and equipment required.
 PROTOCOL Immunostaining for Flow Cytometry 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 Rev.0 Background The combination of single cell analysis using flow cytometry and the specificity of antibody-based
PROTOCOL Immunostaining for Flow Cytometry 1850 Millrace Drive, Suite 3A Eugene, Oregon 97403 Rev.0 Background The combination of single cell analysis using flow cytometry and the specificity of antibody-based
Biological cell membranes
 Unit 14: Cell biology. 14 2 Biological cell membranes The cell surface membrane surrounds the cell and acts as a barrier between the cell s contents and the environment. The cell membrane has multiple
Unit 14: Cell biology. 14 2 Biological cell membranes The cell surface membrane surrounds the cell and acts as a barrier between the cell s contents and the environment. The cell membrane has multiple
INSTRUCTION Probemaker
 INSTRUCTION Probemaker Instructions for Duolink In Situ Probemaker PLUS (Art. no. 92009-0020) and Duolink In Situ Probemaker MINUS (Art. no. 92010-0020) Table of content 1. Introduction 4 2. Applications
INSTRUCTION Probemaker Instructions for Duolink In Situ Probemaker PLUS (Art. no. 92009-0020) and Duolink In Situ Probemaker MINUS (Art. no. 92010-0020) Table of content 1. Introduction 4 2. Applications
Advanced BioDesign Outlines Solutions. Antibody Overview. by Advanced BioDesign. Project Start. Immunogenicity. Selecting Your Antigen
 Advanced BioDesign Outlines Solutions by Advanced BioDesign Antibody Overview Launching an immunisation programme is an important experimental step that needs care. With Advanced BioDesign, you may develop
Advanced BioDesign Outlines Solutions by Advanced BioDesign Antibody Overview Launching an immunisation programme is an important experimental step that needs care. With Advanced BioDesign, you may develop
