The wheat wcs120 promoter is cold-inducible in both monocotyledonous and dicotyledonous species
|
|
|
- Elmer Boyd
- 8 years ago
- Views:
Transcription
1 FEBS Letters 423 (1998) 324^328 FEBS The wheat wcs120 promoter is cold-inducible in both monocotyledonous and dicotyledonous species Franc ois Ouellet, Alejandro Vazquez-Tello, Fathey Sarhan* Deèpartement des Sciences biologiques, Universiteè du Queèbec aé Montreèal, C.P. 8888, Succ. Centre-ville, Montreal, Que. H3C 3P8, Canada Received 5 December 1997; revised version received 26 January 1998 Abstract The wcs120 gene is specifically induced by low temperature (LT) and encodes a protein that is thought to play an important role in the cold acclimation process in wheat. To identify the regulatory elements involved in its LT responsiveness, the transient expression activity of different promoter regions was determined using the luciferase reporter gene. The data indicate the involvement of putative enhancer elements, negative and positive regulatory regions in the transcriptional regulation of this gene. The promoter was found to be coldinducible in different freezing-tolerant and -sensitive monocot and dicot species, suggesting that universal transcription factors responsive to LT may be present in all plants. This promoter could be used to drive the genes needed for LT tolerance in sensitive species. z 1998 Federation of European Biochemical Societies. Key words: Cold inducibility; Dicotyledon; Freezing tolerance; Monocotyledon; Promoter 1. Introduction *Corresponding author. Fax: (1) (514) sarhan.fathey@uqam.ca Abbreviations: FT, freezing tolerance; FI, fold induction; LT, low temperature; LT 50, temperature required to kill 50% of the plants; WCS, wheat cold-specific The cold acclimation process in plants is genetically programmed and is associated with an increased freezing tolerance (FT) resulting from the expression of many genes [1]. Among these, wcs120 is a speci cally LT-regulated gene encoding a highly abundant protein which appears to play an important role during cold acclimation of wheat [2]. The accumulation of both the mrna and protein was shown to correlate closely with the capacity of wheat cultivars to develop FT. The wcs120 gene copy number and organization are identical in freezing-tolerant and less tolerant wheat cultivars, and the expression is regulated mainly at the transcriptional level. On the other hand, the homologs of wcs120 (and other cold-regulated genes) in chilling-sensitive Gramineae species such as rice and corn are not expressed. This lack of expression may result from ine cient cis-acting elements or from the absence of cold-speci c transcription factors and could explain the inability of the sensitive plants to cold acclimatize. Few cis-acting elements responsive to LT (LTRE) have been identi ed so far, the best known being the A / G CCGAC core motif in the promoters of cor15a/rd29a from Arabidopsis, and bn115 from Brassica napus [3^5]. In our e orts to identify the nuclear events regulating the cold-speci c expression of the wcs120 gene, we have characterized its promoter region. DNA-protein interaction studies revealed the presence of multiple DNA binding proteins in nuclear extracts from non-acclimatized plants which interact with several elements in the promoter [6]. The binding of these factors, which may act as transcriptional repressors, is decreased when they are phosphorylated upon LT exposure. In this report, we present the deletion analysis of the wcs120 promoter region using transient expression in wheat leaves. In addition, the promoter strength was determined in di erent monocot and dicot species. 2. Materials and methods 2.1. Plant material and growth conditions Seeds of winter (Triticum aestivum L. cv Fredrick) and spring wheat (T. aestivum L. cv Glenlea), barley (Hordeum vulgare L. cv Sophie) and winter rye (Secale cereale L. cv Puma) were germinated in moist vermiculite for 7 days. Control plants were maintained under controlled environment at a 24³C/20³C (day/night) regime with a 15 h photoperiod and a 75% relative humidity. For rice (Oryza sativa L. cv Nipponbare), seeds were germinated in a water-saturated mixture of soil, peat and vermiculite at 28³C with a 12 h photoperiod. Rapeseed (Brassica napus L. cv Jet Neuf), alfalfa (Medicago sativa ssp. falcata L. cv Anik), sweet pepper (Capsicum annuum L. cv Supersett), cucumber (Cucumis sativus cv Vertige) and tomato (Lycopersicon esculentum Miller cv Floramerica) were grown at 24³C in the same soil, peat and vermiculite mixture. For the cold treatment, plants were grown for 1 week with a 12 h photoperiod at 4³C for wheat, barley, rye, alfalfa and rapeseed, or at 10³C for rice, cucumber, pepper and tomato. Freezing tolerance of these species, expressed as the LT 50, varies as follows: rye (325³C), Fredrick wheat (316³C), Brassica (316³C), alfalfa (315³C), Glenlea wheat (36³C), barley (34³C), rice (4³C), pepper, tomato and cucumber (5^10³C) Transient expression experiments The 942 bp upstream of the ATG translation initiation codon of wcs120 was cloned and sequenced, and the transcription initiation start site was determined [6]. A deletion series was generated by exonuclease III and exonuclease VII digestion and the subclones were sequenced (T7 sequencing kit, Pharmacia). Constructions bearing transcriptional fusions of the promoter fragments with the luciferase reporter gene and the nos terminator were prepared in pbluescript. For the 3P deletions, the TATA box and transcription start site were provided by the 90 bp proximal fragment of the CaMV 35S promoter. Our results showed that, even though this promoter is much more e cient in dicots, it is active in wheat and its capacity to promote transcription is temperature-independent (Fig. 2B, MIN). All the plasmids used were puri ed from Escherichia coli cultures by alkaline lysis and CsCl centrifugation. For transformation, plasmid mixtures were prepared by mixing equal amounts of each wcs120 promoter-luciferase construct with a ubiquitin promoter-glucuronidase construct (pahc27 [7]). Plasmid DNA was coated on 0.9 Wm tungsten beads (M-10, Bio-Rad) by ethanol precipitation [8] and delivered to the leaf tissues using a microprojectile bombardment apparatus [9]. The tissues were placed on an agar plate and bombarded with a 50 ms 85 PSI helium discharge under a vacuum of 25 inches of Hg. They were then oated on a nutrient solution (0.5 g/l 20:20:20, N:P:K; CIL) in a Petri dish and incubated for 2 days at 24³C or 3 days at 4³ or 10³C, as speci ed for each experiment. Soluble proteins were extracted by grinding each /98/$19.00 ß 1998 Federation of European Biochemical Societies. All rights reserved. PII S (98)
2 F. Ouellet et al./febs Letters 423 (1998) 324^ sample (V50^60 mg) with a mortar and pestle in 400 Wl of ice-cold extraction bu er (50 mm Na-phosphate ph 7.0, 10 mm DTT, 0.1% Triton X-100, 10 mm EDTA) and centrifugation at Ug for 10 min. The supernatant was used directly for enzymatic determination of luciferase (LUC; assay kit from Promega) and L-glucuronidase (GUS [10]) activities. In all cases the results are expressed as LUC to GUS ratios, and are means þ S.D. of at least four independent extracts. Prior to LUC/GUS calculation for the 4³C samples, we found that a correction of the GUS values was absolutely necessary since the GUS activity obtained at 4³C was constantly 2^4-fold lower than the activity at 24³C. A correction factor was thus calculated for each construct by dividing the average GUS activity at 24³C by the average activity at 4³C. The individual GUS values at 4³C were then multiplied by this factor to obtain the corrected values used in the LUC/GUS calculation. If this correction had not been made, the activity of the promoter at LT would have been overestimated. 3. Results and discussion 3.1. Deletion analysis Western analyses were performed to evaluate the capacity of the transformed tissues to express the endogenous wcs120 gene family upon LT exposure. The results show that the sections of untransformed leaves (Fig. 1A) and those of leaves transformed with the beads only (Fig. 1B) accumulate the WCS120 proteins in a similar manner. The conditions tested were those used for the post-bombardment incubation period in the transient expression assays. These results show that the wounding stress caused by the bombardment had no e ect on the level of expression of the endogenous wcs120 genes, and should not a ect the activity of the promoter in the transient assays. Wheat leaf sections were transformed with constructs bearing the di erent deletions of the promoter transcriptionally fused to the luciferase (luc) reporter gene. The chimeric constructs are represented schematically in Fig. 2A. The results show that LT treatment increases LUC activity 8-fold when the full length wcs120 promoter is used (Fig. 2B, FL860). A control experiment was performed using the pahc18 (Ubiluc) vector under the same conditions. The results showed no increase in LUC activity at 4³C when the gene is driven by the Ubi promoter. This con rms that when the wcs120 promoter is used, the increase in activity observed at 4³C is due to the LT inducibility and not to an increased luc transcript stability at this temperature. The 5P deletion up to 3590 leads to an increase of the fold induction (FI) from 8- to 55-fold. This is mainly due to the almost complete loss of basal activity at 24³C. The region between 3860 and 3590 contains a 52 bp direct repeat composed of two elements separated from each other by 4 bp, and four of the ve CGTCGG elements which do not show any homology with known motifs. The loss of activity at both 4³ and 24³C upon deletion of this region suggests that either or both elements could act as transcriptional enhancers. It was reported that repeated sequences of 127^337 bp from Arabidopsis could act as enhancers in tobacco transgenic plants [11]. A further deletion to 3415 increases the activity at both temperatures, suggesting that the region from 3590 to 3415 contains elements that repress transcription. This region contains the two GGGTATA elements of unknown function. The fact that the e ect of the putative negative elements was not apparent in the FL860 construct suggests that the putative enhancers located between 3860 and 3590 may overcome or inactivate the negative factors of the 3590 to 3415 region. If Fig. 1. E ect of bombardment on the expression of the endogenous WCS120 proteins. The proteins were extracted and analyzed by Western blotting using the polyclonal antibody directed against the members of the WCS120 family. Numbers on the right indicate the MW (in kda) of the major members of the family. Three independent samples were analyzed for each condition. A: Protein accumulation in non-transformed wheat leaves sections after 48 h at 24³C and 72 h at 4³C. B: Protein accumulation after bombardment of wheat leaves with tungsten beads without DNA and incubation at 24³C for 48 h or at 4³C for 72 h. the region between 3415 and 3215 is deleted, a decrease in activity is observed at both temperatures, suggesting the presence of enhancer elements. However, the signi cant decrease in activity at 4³C suggests that this region could contain coldinducible positive regulatory elements. In fact, this region contains a CCGAC LTRE previously identi ed in the promoters of cor15a/rd29a from Arabidopsis and bn115 from Brassica napus [3^5]. The presence of a motif identical to this LTRE in the wcs120 promoter may suggest a similar role in wheat. A cdna clone corresponding to a protein from Arabidopsis which can bind the CCGAC motif has been isolated recently [12]. A more detailed characterization of this transcription factor will help to determine the implication of this motif in the LT response. The deletion of the region between 3215 and 372 abolishes almost all promoter activity. The decrease of activity at 4³C suggests that the deleted region contains an LTRE. This region contains the other GCCGAC LTRE and two of the three CACCTGC elements. The latter elements contain the CANNTG motif, which forms the core of several cis-acting elements such as the ABA response elements (ABRE) [13]. This motif was identi ed as the preferred binding site for the bhlh proteins, the common plant regulatory factors (CPRFs) and for the G-box binding factors (GBFs) belonging to the bzip class of proteins [13]. Another element, CACT- CAC, is repeated two times and was identi ed as a binding site for the transcriptional activators GCN4 from yeast and zeste from Drosophila [14,15]. Taken together, these observations suggest that the wcs120 promoter possesses putative cisacting elements that could bind known transcription factors, present in both the plant and animal kingdoms. The promoter region of the wcs120 gene was also analyzed by 3P deletions. The deletion of the proximal 67 bp of the wcs120 promoter leads to a decrease of the FI from 8.0- to 4.0-fold, suggesting the presence of an LTRE in the region between 367 and +1. A further deletion to 3178 decreases
3 326 F. Ouellet et al./febs Letters 423 (1998) 324^328 Fig. 2. Deletion analysis of the wcs120 promoter by transient expression. A: Schematic representation of the promoter fragments and relative positions of the repeated DNA motifs deduced from the sequence analysis (GenBank accession number AF031235). The constructions were named according to the rst (5P deletions) or last (3P deletion) nucleotide of the promoter fragment. B: E ect of cold treatment on the activity of the wcs120 promoter fragments. The leaf sections were transformed with the di erent promoter-luc fragments and Ubi-Gus (pahc27), and incubated at 4³C for 3 days or at 24³C for 2 days. Soluble proteins were extracted and enzymatic activities of LUC and GUS were determined. Numbers at the right of the error bars indicate the induction factors (4³C/24³C relative activity ratio). the FI from 4.0- to 1.8-fold, suggesting the presence of an LTRE. This supports the observation made from the analysis of the 3215 to 372 deletion. Upon removal of the initial 268 bp of the promoter, the activity at 4³C increased whereas that at 24³C decreased compared to 3178 construct. The relative increase of the FI suggests the existence of negative regulatory elements between 3268 and 3178 that would be mainly active at 4³C. Deletion to 3374 completely abolishes the wcs120 promoter activity. Since the most important loss of absolute activity is that at 4³C, an LTRE may be present between 3374 and On the other hand, we cannot rule out the possibility of the existence of an enhancer element in this region. Further deletion to 3501 does not reactivate the promoter, indicating that most of the elements responsible for the promoter activity were removed. Together, the results from the transient expression assays suggest that the transcription of the wcs120 gene is dependent on the presence of multiple regulatory elements. Footprinting experiments are required to identify more precisely the DNA motifs that bind the di erent proteins. It will then be possible to perform the mutation analyses needed to con rm the importance of these motifs in the LT responsiveness Activity of the wcs120 promoter in di erent species The transient activity of the full length promoter (FL860) was determined in di erent freezing-sensitive and -tolerant species. The results show that the promoter activity is similar in the more tolerant Fredrick and less tolerant Glenlea cultivars (Fig. 3). This suggests that even though Glenlea is less tolerant, the plant possesses the trans regulatory elements needed to express this important gene for FT. This result was expected since Northern analysis had shown no signi cant di erences between the levels of expression of the wcs120 gene in Glenlea and Fredrick, during the rst week of acclimation [1]. The kinetic analyses of WCS120 protein accumulation and the close inverse relationship between WCS120 protein levels and LT 50 indicate that the FT of cereals is determined by the degree and duration of LT gene expression. These results and other evidence, such as increased transcript stability and alternative splicing, support a role for post-transcriptional regulation events in the di erential accumulation of proteins at LT [1,16]. The promoter shows a similar LT inducibility in wheat and barley (Fig. 3). Barley, a species closely related to wheat, possesses a gene (dhn5 [17]) that is almost identical to Table 1 Activity of the wcs120 full length promoter in di erent dicotyledonous species after cold exposure Species Family Tolerance to freezing LUC/GUS ratio FI a 24³C cold Alfalfa Leguminosae Tolerant 0 b 1031 þ 109 (ND) b Brassica Cruciferae Tolerant 0 b 39 þ 2 (ND) b Cucumber Cucurbitaceae Sensitive 197 þ þ Tomato Solanaceae Sensitive 73 þ þ 29 1 c Pepper Solanaceae Sensitive þ þ The leaf sections were transformed with the FL860LUC construct and Ubi-Gus (pahc27), and incubated at LT for 3 days or at 24³C for 2 days. The LT treatment was performed at 4³C or at 10³C for the cold-tolerant and -sensitive species, respectively. Soluble proteins were extracted and enzymatic activities of LUC and GUS were determined. a The fold induction (FI) is the cold/24³c relative activity ratio. b The FI was di cult to determine due to the undetectable activity of LUC at 24³C. c The di erence between the two conditions is not signi cant.
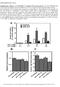
4 F. Ouellet et al./febs Letters 423 (1998) 324^ Fig. 3. Activity of the wcs120 full length promoter in di erent monocotyledonous species after cold exposure. The leaf sections were transformed with the FL860LUC construct and Ubi-Gus (pahc27), and incubated at LT for 3 days or at 24³C for 2 days. The LT treatment was performed at 4³C except for rice, a sensitive species, which was exposed at 10³C. Soluble proteins were extracted and enzymatic activities of LUC and GUS were determined. Numbers above the error bars indicate the fold induction (FI; 4³C/24³C relative activity ratio). In the case of rye and rice, the FI was di cult to estimate due to the undetectable activity of LUC at 24³C. The LUC/GUS ratio was almost zero. wcs120. Winter rye, a highly tolerant Graminea that also possesses homologs of the wcs120 family members, showed a 20- fold increase in activity following exposure to 4³C. Rice, a cold-sensitive monocot species that possesses an inactive gene homologous to wcs120, showed an 11-fold increase at 10³C compared to 24³C. A much higher LT inducibility was observed in the freezing-tolerant dicot species alfalfa and Brassica at 4³C (Table 1). The fold increase was very di cult to estimate due to the undetectable activity of wcs120-luc at 24³C. The wcs120 promoter activity was also evaluated in cold-sensitive dicots (Table 1). In cucumber, a 26-fold induction of the activity at 10³C was observed. On the other hand, in tomato, no di erence in activity was observed between the samples treated at the two temperatures. Pepper is the only species for which a decrease (10-fold) of the promoter activity was observed upon LT exposure. These results indicate that the level of promoter activity is not correlated with the capacity of these species to develop FT. For example, the promoter is less active in rye (the most tolerant species tested) than in wheat, a result that is supported by previous genetic analyses. Amphiploids from rye/wheat crosses show a FT equivalent, but not superior, to that of wheat [1]. Furthermore, the level of expression of the wcs120 family genes in these individuals does not exceed that observed in wheat. This suggests that the elements important for FT in rye are silenced in a predominant wheat background, and would suggest the existence of di erences in the cis- and/or trans-acting elements involved in the transcription of wcs120 and perhaps of other LT-induced genes. Our data indicate that the wcs120 promoter is LT-inducible in the Gramineae (wheat, barley, rice and rye), Cruciferae (Brassica), Leguminosae (alfalfa) and Cucurbitaceae (cucumber), but not in the Solanaceae family (tomato, pepper). The reason for the absence of LT induction in the latter family is not known. The results provide evidence of the existence of common transcription factors in both monocots and dicots, and suggest that these factors have the capacity to recognize the cis-acting elements of a cold-inducible heterologous promoter. The data presented here indicate that the cold sensitivity of some species (such as rice and cucumber) is apparently not due to ine cient or absent transcription factors but may possibly result from the ine ciency of the promoters of the homologous genes. This hypothesis could be veri ed with the characterization of the promoter of the wcs120 homologs in sensitive species. The observations from the transient assays do not, of course, necessarily represent what would happen in transgenic plants. Nevertheless, the identi cation of such a promoter is interesting from an application point of view since it could be a useful tool in the elaboration of strategies aimed at the improvement of FT in sensitive monocot and dicot species. Until now, plants modi ed to overexpress genes potentially implicated in FT have been transformed with constructions bearing the genes under the control of constitutive promoters. The most widely used promoters are those of the maize ubiquitin gene (Ubi) and of the CaMV 35S gene, for monocots and dicots respectively. E orts are now being focused towards the identi cation of promoters that are more e cient in di erent plants. For example, several constructions bearing, in di erent combinations, fragments of the CaMV 35S promoter, an intron of the bean phaseolin gene, the 6 sequence of TMV and terminating sequences from the CaMV 35S or nos genes have been tested in rice (monocot) and tobacco (dicot) [18]. It was shown that the most e cient constructions for rice were not the same as for tobacco, suggesting di erences in the speci city of gene expression between monocot and dicot plants. On the other hand, the overexpression of a gene resulting from the use of a constitutive promoter is not necessarily an objective to achieve in all cases. Indeed, few studies have focused on the physiological consequences related to the constitutive expression of genes that are normally inducible and thus expressed only when needed. The use of a cold-inducible promoter such as that of the wcs120 gene would allow the expression of the genes only when the plant is under cold stress conditions. Acknowledgements: We are grateful to Sylvain Dallaire for the construction of the microprojectile apparatus. This work was supported by a Natural Sciences and Engineering Research Council of Canada research grant (A7199) to F. Sarhan. F. Ouellet was supported by a Ph.D. scholarship from the Fonds pour la Formation de Chercheurs et l'aide aé la recherche, Gouvernement du Queèbec. References [1] Hughes, M.A. and Dunn, M.A. (1996) J. Exp. Bot. 47, 291^305. [2] Sarhan, F., Ouellet, F. and Vazquez-Tello, A. (1997) Physiol. Plant. 101, 439^445. [3] Baker, S.S., Wilhelm, K.S. and Thomashow, M.F. (1994) Plant Mol. Biol. 24, 701^713. [4] Jiang, C., Iu, B. and Singh, J. (1996) Plant Mol. Biol. 30, 679^ 684. [5] Yamaguchi-Shinozaki, K. and Shinozaki, K. (1994) Plant Cell 6, 251^264. [6] Vazquez-Tello, A., Ouellet, F. and Sarhan, F. (1997) Mol. Gen. Genet. (in press). [7] Christensen, A.H. and Quail, P.H. (1996) Transgenic Res. 5, 213^ 218. [8] Godon, C., Caboche, M. and Daniel-Vedele, F. (1993) Biochimie 75, 591^595. [9] Vain, P., Keen, N., Murillo, J., Rathus, C., Nemes, C. and Finer, J.J. (1993) Plant Cell Tiss. Org. 33, 237^246.
5 328 [10] Je erson, R.A., Kavanagh, T.A. and Bevan, M.W. (1987) EMBO J. 6, 3901^3907. [11] Ott, R.W. and Hansen, L.K. (1996) Mol. Gen. Genet. 252, 563^ 571. [12] Stockinger, E.J., Gilmour, S.J. and Thomashow, M.F. (1997) Proc. Natl. Acad. Sci. USA 94, 1035^1040. [13] Meshi, T. and Iwabuchi, M. (1995) Plant Cell Physiol. 36, 1405^ [14] Chen, J.D., Chang, C.S. and Pirrotta, V. (1992) Mol. Cell. Biol. 12, 598^608. [15] Thireos, G., Penn, M.D. and Greer, H. (1984) Proc. Natl. Acad. Sci. USA 81, 5096^5100. F. Ouellet et al./febs Letters 423 (1998) 324^328 [16] Bournay, A.S., Hedley, P.E., Maddison, A., Waugh, R. and Machray, G.C. (1996) Nucleic Acids Res. 24, 2347^2351. [17] Close, T.J., Meyer, N.C. and Radik, J. (1995) Plant Physiol. 107, 289^290. [18] Mitsuhara, I., Ugaki, M., Hirochika, H., Ohshima, M., Murakami, T., Gotoh, Y., Katayose, Y., Nakamura, S., Honkura, R., Nishimiya, S., Ueno, K., Mochizuki, A., Tanimoto, H., Tsugawa, H., Otsuki, Y. and Ohashi, Y. (1996) Plant Cell Physiol. 37, 49^59.
Anti-ATF6 α antibody, mouse monoclonal (1-7)
 Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
How To Understand How Gene Expression Is Regulated
 What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
Integrated Protein Services
 Integrated Protein Services Custom protein expression & purification Version DC04-0012 Expression strategy The first step in the recombinant protein generation process is to design an appropriate expression
Integrated Protein Services Custom protein expression & purification Version DC04-0012 Expression strategy The first step in the recombinant protein generation process is to design an appropriate expression
Rapid GST Inclusion Body Solubilization and Renaturation Kit
 Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
STUDIES ON SEED STORAGE PROTEINS OF SOME ECONOMICALLY MINOR PLANTS
 STUDIES ON SEED STORAGE PROTEINS OF SOME ECONOMICALLY MINOR PLANTS THESIS SUBMITTED FOR THE DEGREB OF DOCTOR OF PHILOSOPHY (SCIENCE) OF THE UNIVERSITY OF CALCUTTA 1996 NRISINHA DE, M.Sc DEPARTMENT OF BIOCHEMISTRY
STUDIES ON SEED STORAGE PROTEINS OF SOME ECONOMICALLY MINOR PLANTS THESIS SUBMITTED FOR THE DEGREB OF DOCTOR OF PHILOSOPHY (SCIENCE) OF THE UNIVERSITY OF CALCUTTA 1996 NRISINHA DE, M.Sc DEPARTMENT OF BIOCHEMISTRY
QUANTITATIVE RT-PCR. A = B (1+e) n. A=amplified products, B=input templates, n=cycle number, and e=amplification efficiency.
 QUANTITATIVE RT-PCR Application: Quantitative RT-PCR is used to quantify mrna in both relative and absolute terms. It can be applied for the quantification of mrna expressed from endogenous genes, and
QUANTITATIVE RT-PCR Application: Quantitative RT-PCR is used to quantify mrna in both relative and absolute terms. It can be applied for the quantification of mrna expressed from endogenous genes, and
Recombinant DNA and Biotechnology
 Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
www.mpiz-koeln.mpg.de Abt. Entwicklungsbiologie de Pflanzen
 WEB ADDRESS: www.mpiz-koeln.mpg.de Forschung Abt. Entwicklungsbiologie de Pflanzen George Coupland How is the transition from vegetative growth to flowering controlled? - How is it regulated by environmental
WEB ADDRESS: www.mpiz-koeln.mpg.de Forschung Abt. Entwicklungsbiologie de Pflanzen George Coupland How is the transition from vegetative growth to flowering controlled? - How is it regulated by environmental
How To Make A Tri Reagent
 TRI Reagent For processing tissues, cells cultured in monolayer or cell pellets Catalog Number T9424 Store at room temperature. TECHNICAL BULLETIN Product Description TRI Reagent is a quick and convenient
TRI Reagent For processing tissues, cells cultured in monolayer or cell pellets Catalog Number T9424 Store at room temperature. TECHNICAL BULLETIN Product Description TRI Reagent is a quick and convenient
SUGAR BEET (Beta vulgaris L.) PROMOTERS FOR DIRECTED TISSUE- SPECIFIC ROOT TRANSCRIPTION
 SUGAR BEET (Beta vulgaris L.) PROMOTERS FOR DIRECTED TISSUE- SPECIFIC ROOT TRANSCRIPTION Senthilkumar Padmanaban, Haiyan Li, David P. Puthoff and Ann C. Smigocki USDA-ARS Molecular Plant Pathology Laboratory,
SUGAR BEET (Beta vulgaris L.) PROMOTERS FOR DIRECTED TISSUE- SPECIFIC ROOT TRANSCRIPTION Senthilkumar Padmanaban, Haiyan Li, David P. Puthoff and Ann C. Smigocki USDA-ARS Molecular Plant Pathology Laboratory,
CUSTOM ANTIBODIES. Fully customised services: rat and murine monoclonals, rat and rabbit polyclonals, antibody characterisation, antigen preparation
 CUSTOM ANTIBODIES Highly competitive pricing without compromising quality. Rat monoclonal antibodies for the study of gene expression and proteomics in mice and in mouse models of human diseases available.
CUSTOM ANTIBODIES Highly competitive pricing without compromising quality. Rat monoclonal antibodies for the study of gene expression and proteomics in mice and in mouse models of human diseases available.
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College
 Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
Biotechnology and Recombinant DNA (Chapter 9) Lecture Materials for Amy Warenda Czura, Ph.D. Suffolk County Community College Primary Source for figures and content: Eastern Campus Tortora, G.J. Microbiology
Hydroxynitrile lyase (Hnl)
 Hydroxynitrile lyase (Hnl) R 1 HCN R 1 OH R 2 C O R 2 C * CN S-selective: Hevea brasiliensis R-selective: Prunus spp. (S)-Hnl of Hevea brasiliensis and (R)-Hnl of Prunus amygdalus Hb_Hnl Type II Hnl intracellular
Hydroxynitrile lyase (Hnl) R 1 HCN R 1 OH R 2 C O R 2 C * CN S-selective: Hevea brasiliensis R-selective: Prunus spp. (S)-Hnl of Hevea brasiliensis and (R)-Hnl of Prunus amygdalus Hb_Hnl Type II Hnl intracellular
GENE CLONING AND RECOMBINANT DNA TECHNOLOGY
 GENE CLONING AND RECOMBINANT DNA TECHNOLOGY What is recombinant DNA? DNA from 2 different sources (often from 2 different species) are combined together in vitro. Recombinant DNA forms the basis of cloning.
GENE CLONING AND RECOMBINANT DNA TECHNOLOGY What is recombinant DNA? DNA from 2 different sources (often from 2 different species) are combined together in vitro. Recombinant DNA forms the basis of cloning.
Module 3 Questions. 7. Chemotaxis is an example of signal transduction. Explain, with the use of diagrams.
 Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Integrated Protein Services
 Integrated Protein Services Custom protein expression & purification Last date of revision June 2015 Version DC04-0013 www.iba-lifesciences.com Expression strategy The first step in the recombinant protein
Integrated Protein Services Custom protein expression & purification Last date of revision June 2015 Version DC04-0013 www.iba-lifesciences.com Expression strategy The first step in the recombinant protein
2.1.2 Characterization of antiviral effect of cytokine expression on HBV replication in transduced mouse hepatocytes line
 i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
i 1 INTRODUCTION 1.1 Human Hepatitis B virus (HBV) 1 1.1.1 Pathogenesis of Hepatitis B 1 1.1.2 Genome organization of HBV 3 1.1.3 Structure of HBV virion 5 1.1.4 HBV life cycle 5 1.1.5 Experimental models
GenScript Antibody Services
 GenScript Antibody Services Scientific experts, innovative technologies, proven performance We specialize in custom antibody production to empower your research Custom Polyclonal Antibody Production Custom
GenScript Antibody Services Scientific experts, innovative technologies, proven performance We specialize in custom antibody production to empower your research Custom Polyclonal Antibody Production Custom
MOL.911 HNL Expression
 1 W I S S E N T E C H N I K L E I D E N S C H A F T MOL.911 HNL Expression www.tugraz.at 2 Hydroxynitrile lyase (Hnl) R 1 HCN R 1 OH R 2 C O R 2 C * CN S selective: Hevea brasiliensis R selective: Prunus
1 W I S S E N T E C H N I K L E I D E N S C H A F T MOL.911 HNL Expression www.tugraz.at 2 Hydroxynitrile lyase (Hnl) R 1 HCN R 1 OH R 2 C O R 2 C * CN S selective: Hevea brasiliensis R selective: Prunus
Classic Immunoprecipitation
 292PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Classic Immunoprecipitation Utilizes Protein A/G Agarose for Antibody Binding (Cat.
292PR 01 G-Biosciences 1-800-628-7730 1-314-991-6034 technical@gbiosciences.com A Geno Technology, Inc. (USA) brand name Classic Immunoprecipitation Utilizes Protein A/G Agarose for Antibody Binding (Cat.
Rubisco; easy Purification and Immunochemical Determination
 Rubisco; easy Purification and Immunochemical Determination Ulrich Groß Justus-Liebig-Universität Gießen, Institute of Plant Nutrition, Department of Tissue Culture, Südanlage 6, D-35390 Giessen e-mail:
Rubisco; easy Purification and Immunochemical Determination Ulrich Groß Justus-Liebig-Universität Gießen, Institute of Plant Nutrition, Department of Tissue Culture, Südanlage 6, D-35390 Giessen e-mail:
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
Engineering of Yellow Mosaic Virus Resistance (YMVR) in Blackgram. Project ID: 1 April 2000 to 31 August 2004. Project Duration:
 Engineering of Yellow Mosaic Virus Resistance (YMVR) in Blackgram ID: Duration: Coordinator in Switzerland: PS1 1 April 2000 to 31 August 2004 Prof. Thomas Hohn University of Basel Botanisches Institut
Engineering of Yellow Mosaic Virus Resistance (YMVR) in Blackgram ID: Duration: Coordinator in Switzerland: PS1 1 April 2000 to 31 August 2004 Prof. Thomas Hohn University of Basel Botanisches Institut
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
Protein Expression. A Practical Approach J. HIGGIN S
 Protein Expression A Practical Approach S. J. HIGGIN S B. D. HAMES List of contributors Abbreviations xv Xvi i 1. Protein expression in mammalian cell s Marlies Otter-Nilsson and Tommy Nilsso n 1. Introduction
Protein Expression A Practical Approach S. J. HIGGIN S B. D. HAMES List of contributors Abbreviations xv Xvi i 1. Protein expression in mammalian cell s Marlies Otter-Nilsson and Tommy Nilsso n 1. Introduction
LightSwitch Dual Assay System DA010 (100 assays)
 LightSwitch Dual Assay System DA010 (100 assays)! Description The LightSwitch Dual Assay System is used to normalize for variation between transfection replicates in cell lines that are particularly difficult
LightSwitch Dual Assay System DA010 (100 assays)! Description The LightSwitch Dual Assay System is used to normalize for variation between transfection replicates in cell lines that are particularly difficult
Plant Growth & Development. Growth Stages. Differences in the Developmental Mechanisms of Plants and Animals. Development
 Plant Growth & Development Plant body is unable to move. To survive and grow, plants must be able to alter its growth, development and physiology. Plants are able to produce complex, yet variable forms
Plant Growth & Development Plant body is unable to move. To survive and grow, plants must be able to alter its growth, development and physiology. Plants are able to produce complex, yet variable forms
DP419 RNAsimple Total RNA Kit. RNAprep pure Series. DP501 mircute mirna Isolation Kit. DP438 MagGene Viral DNA / RNA Kit. DP405 TRNzol Reagent
 Overview of TIANGEN Products DP419 RNAsimple Total RNA Kit DP430 RNAprep pure Kit(For Cell/Bacteria) DP315/DP315-R TIANamp Virus DNA/RNA Kit DP431 RNAprep pure Kit (For Tissue) Silica-membrane Technology
Overview of TIANGEN Products DP419 RNAsimple Total RNA Kit DP430 RNAprep pure Kit(For Cell/Bacteria) DP315/DP315-R TIANamp Virus DNA/RNA Kit DP431 RNAprep pure Kit (For Tissue) Silica-membrane Technology
MagExtractor -Genome-
 Instruction manual MagExtractor-Genome-0810 F0981K MagExtractor -Genome- NPK-101 100 preparations Store at 4 C Contents [1] Introduction [2] Components [3] Materials required [4] Protocol 1. Purification
Instruction manual MagExtractor-Genome-0810 F0981K MagExtractor -Genome- NPK-101 100 preparations Store at 4 C Contents [1] Introduction [2] Components [3] Materials required [4] Protocol 1. Purification
Thesis of Ph.D. Novel cyclin-dependent kinase inhibitor protein from Medicago truncatula. Aladár Pettkó-Szandtner
 Thesis of Ph.D Novel cyclin-dependent kinase inhibitor protein from Medicago truncatula Aladár Pettkó-Szandtner Supervisors: Horváth Gábor and Dudits Dénes Institute of Plant Biology HAS, Biologcal Research
Thesis of Ph.D Novel cyclin-dependent kinase inhibitor protein from Medicago truncatula Aladár Pettkó-Szandtner Supervisors: Horváth Gábor and Dudits Dénes Institute of Plant Biology HAS, Biologcal Research
Integrating DNA Motif Discovery and Genome-Wide Expression Analysis. Erin M. Conlon
 Integrating DNA Motif Discovery and Genome-Wide Expression Analysis Department of Mathematics and Statistics University of Massachusetts Amherst Statistics in Functional Genomics Workshop Ascona, Switzerland
Integrating DNA Motif Discovery and Genome-Wide Expression Analysis Department of Mathematics and Statistics University of Massachusetts Amherst Statistics in Functional Genomics Workshop Ascona, Switzerland
Martin D. Smith, Sally J. Dawson, Linda M. Boxer 1 and David S. Latchman* 4100 4107 Nucleic Acids Research, 1998, Vol. 26, No. 18
 4100 4107 Nucleic Acids Research, 1998, Vol. 26, No. 18 1998 Oxford University Press The N-terminal domain unique to the long form of the Brn-3a transcription factor is essential to protect neuronal cells
4100 4107 Nucleic Acids Research, 1998, Vol. 26, No. 18 1998 Oxford University Press The N-terminal domain unique to the long form of the Brn-3a transcription factor is essential to protect neuronal cells
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE Q5B
 INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
Transcription and Translation of DNA
 Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Induction of Enzyme Activity in Bacteria:The Lac Operon. Preparation for Laboratory: Web Tutorial - Lac Operon - submit questions
 Induction of Enzyme Activity in Bacteria:The Lac Operon Preparation for Laboratory: Web Tutorial - Lac Operon - submit questions I. Background: For the last week you explored the functioning of the enzyme
Induction of Enzyme Activity in Bacteria:The Lac Operon Preparation for Laboratory: Web Tutorial - Lac Operon - submit questions I. Background: For the last week you explored the functioning of the enzyme
Chromatin Immunoprecipitation
 Chromatin Immunoprecipitation A) Prepare a yeast culture (see the Galactose Induction Protocol for details). 1) Start a small culture (e.g. 2 ml) in YEPD or selective media from a single colony. 2) Spin
Chromatin Immunoprecipitation A) Prepare a yeast culture (see the Galactose Induction Protocol for details). 1) Start a small culture (e.g. 2 ml) in YEPD or selective media from a single colony. 2) Spin
Genomic DNA Extraction Kit INSTRUCTION MANUAL
 Genomic DNA Extraction Kit INSTRUCTION MANUAL Table of Contents Introduction 3 Kit Components 3 Storage Conditions 4 Recommended Equipment and Reagents 4 Introduction to the Protocol 4 General Overview
Genomic DNA Extraction Kit INSTRUCTION MANUAL Table of Contents Introduction 3 Kit Components 3 Storage Conditions 4 Recommended Equipment and Reagents 4 Introduction to the Protocol 4 General Overview
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
Luísa Romão. Instituto Nacional de Saúde Dr. Ricardo Jorge Av. Padre Cruz, 1649-016 Lisboa, Portugal. Cooper et al (2009) Cell 136: 777
 Luísa Romão Instituto Nacional de Saúde Dr. Ricardo Jorge Av. Padre Cruz, 1649-016 Lisboa, Portugal Cooper et al (2009) Cell 136: 777 PTC = nonsense or stop codon = UAA, UAG, UGA PTCs can arise in a variety
Luísa Romão Instituto Nacional de Saúde Dr. Ricardo Jorge Av. Padre Cruz, 1649-016 Lisboa, Portugal Cooper et al (2009) Cell 136: 777 PTC = nonsense or stop codon = UAA, UAG, UGA PTCs can arise in a variety
Custom Antibody Services
 prosci-inc.com Custom Antibody Services High Performance Antibodies and More Broad Antibody Catalog Extensive Antibody Services CUSTOM ANTIBODY SERVICES Established in 1998, ProSci Incorporated is a leading
prosci-inc.com Custom Antibody Services High Performance Antibodies and More Broad Antibody Catalog Extensive Antibody Services CUSTOM ANTIBODY SERVICES Established in 1998, ProSci Incorporated is a leading
Name Class Date. Figure 13 1. 2. Which nucleotide in Figure 13 1 indicates the nucleic acid above is RNA? a. uracil c. cytosine b. guanine d.
 13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
PrimeSTAR HS DNA Polymerase
 Cat. # R010A For Research Use PrimeSTAR HS DNA Polymerase Product Manual Table of Contents I. Description...3 II. III. IV. Components...3 Storage...3 Features...3 V. General Composition of PCR Reaction
Cat. # R010A For Research Use PrimeSTAR HS DNA Polymerase Product Manual Table of Contents I. Description...3 II. III. IV. Components...3 Storage...3 Features...3 V. General Composition of PCR Reaction
REAL TIME PCR USING SYBR GREEN
 REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
13.4 Gene Regulation and Expression
 13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
ab185915 Protein Sumoylation Assay Ultra Kit
 ab185915 Protein Sumoylation Assay Ultra Kit Instructions for Use For the measuring in vivo protein sumoylation in various samples This product is for research use only and is not intended for diagnostic
ab185915 Protein Sumoylation Assay Ultra Kit Instructions for Use For the measuring in vivo protein sumoylation in various samples This product is for research use only and is not intended for diagnostic
Twincore - Zentrum für Experimentelle und Klinische Infektionsforschung Institut für Molekulare Bakteriologie
 Twincore - Zentrum für Experimentelle und Klinische Infektionsforschung Institut für Molekulare Bakteriologie 0 HELMHOLTZ I ZENTRUM FÜR INFEKTIONSFORSCHUNG Technische Universität Braunschweig Institut
Twincore - Zentrum für Experimentelle und Klinische Infektionsforschung Institut für Molekulare Bakteriologie 0 HELMHOLTZ I ZENTRUM FÜR INFEKTIONSFORSCHUNG Technische Universität Braunschweig Institut
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation
 OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
OriGene Technologies, Inc. MicroRNA analysis: Detection, Perturbation, and Target Validation -Optimal strategies to a successful mirna research project Optimal strategies to a successful mirna research
Bio-Reagents Gene synthesis Peptide Synthesis Protein Expression Antibody Production. Life Science Products and Services
 Bio-Reagents Gene synthesis Peptide Synthesis Protein Expression Antibody Production Life Science Products and Services Since 2002, Biomatik has provided worldwide researchers in life science discovery
Bio-Reagents Gene synthesis Peptide Synthesis Protein Expression Antibody Production Life Science Products and Services Since 2002, Biomatik has provided worldwide researchers in life science discovery
Molecular Biology Techniques: A Classroom Laboratory Manual THIRD EDITION
 Molecular Biology Techniques: A Classroom Laboratory Manual THIRD EDITION Susan Carson Heather B. Miller D.Scott Witherow ELSEVIER AMSTERDAM BOSTON HEIDELBERG LONDON NEW YORK OXFORD PARIS SAN DIEGO SAN
Molecular Biology Techniques: A Classroom Laboratory Manual THIRD EDITION Susan Carson Heather B. Miller D.Scott Witherow ELSEVIER AMSTERDAM BOSTON HEIDELBERG LONDON NEW YORK OXFORD PARIS SAN DIEGO SAN
Genetomic Promototypes
 Genetomic Promototypes Mirkó Palla and Dana Pe er Department of Mechanical Engineering Clarkson University Potsdam, New York and Department of Genetics Harvard Medical School 77 Avenue Louis Pasteur Boston,
Genetomic Promototypes Mirkó Palla and Dana Pe er Department of Mechanical Engineering Clarkson University Potsdam, New York and Department of Genetics Harvard Medical School 77 Avenue Louis Pasteur Boston,
Production of antigens and antibodies in plants: alternative technology?
 Production of antigens and antibodies in plants: alternative technology? George Lomonossoff John Innes Centre Norwich, UK ECOPA, Alicante 29 th Sept. 2006 Why use Plants as Biofactories? Produce large
Production of antigens and antibodies in plants: alternative technology? George Lomonossoff John Innes Centre Norwich, UK ECOPA, Alicante 29 th Sept. 2006 Why use Plants as Biofactories? Produce large
Automation in Genomics High-throughput purification of nucleic acids from biological samples. Valentina Gualdi Operational Scientist PGP
 Automation in Genomics High-throughput purification of nucleic acids from biological samples Valentina Gualdi Operational Scientist PGP OVERVIEW Nucleic acid purification technologies general aspects Genomic
Automation in Genomics High-throughput purification of nucleic acids from biological samples Valentina Gualdi Operational Scientist PGP OVERVIEW Nucleic acid purification technologies general aspects Genomic
Supplementary Figure 1.
 Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Introduction To Real Time Quantitative PCR (qpcr)
 Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Profiling of non-coding RNA classes Gunter Meister
 Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Recombinant DNA Technology
 Recombinant DNA Technology Dates in the Development of Gene Cloning: 1965 - plasmids 1967 - ligase 1970 - restriction endonucleases 1972 - first experiments in gene splicing 1974 - worldwide moratorium
Recombinant DNA Technology Dates in the Development of Gene Cloning: 1965 - plasmids 1967 - ligase 1970 - restriction endonucleases 1972 - first experiments in gene splicing 1974 - worldwide moratorium
"Fingerprinting" Vegetables DNA-based Marker Assisted Selection
 "Fingerprinting" Vegetables DNA-based Marker Assisted Selection Faster, Cheaper, More Reliable; These are some of the goals that vegetable breeders at seed companies and public institutions desire for
"Fingerprinting" Vegetables DNA-based Marker Assisted Selection Faster, Cheaper, More Reliable; These are some of the goals that vegetable breeders at seed companies and public institutions desire for
Introduction to Genome Annotation
 Introduction to Genome Annotation AGCGTGGTAGCGCGAGTTTGCGAGCTAGCTAGGCTCCGGATGCGA CCAGCTTTGATAGATGAATATAGTGTGCGCGACTAGCTGTGTGTT GAATATATAGTGTGTCTCTCGATATGTAGTCTGGATCTAGTGTTG GTGTAGATGGAGATCGCGTAGCGTGGTAGCGCGAGTTTGCGAGCT
Introduction to Genome Annotation AGCGTGGTAGCGCGAGTTTGCGAGCTAGCTAGGCTCCGGATGCGA CCAGCTTTGATAGATGAATATAGTGTGCGCGACTAGCTGTGTGTT GAATATATAGTGTGTCTCTCGATATGTAGTCTGGATCTAGTGTTG GTGTAGATGGAGATCGCGTAGCGTGGTAGCGCGAGTTTGCGAGCT
Sample Questions for Exam 3
 Sample Questions for Exam 3 1. All of the following occur during prometaphase of mitosis in animal cells except a. the centrioles move toward opposite poles. b. the nucleolus can no longer be seen. c.
Sample Questions for Exam 3 1. All of the following occur during prometaphase of mitosis in animal cells except a. the centrioles move toward opposite poles. b. the nucleolus can no longer be seen. c.
mirnaselect pep-mir Cloning and Expression Vector
 Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
Product Data Sheet mirnaselect pep-mir Cloning and Expression Vector CATALOG NUMBER: MIR-EXP-C STORAGE: -80ºC QUANTITY: 2 vectors; each contains 100 µl of bacterial glycerol stock Components 1. mirnaselect
KMS-Specialist & Customized Biosimilar Service
 KMS-Specialist & Customized Biosimilar Service 1. Polyclonal Antibody Development Service KMS offering a variety of Polyclonal Antibody Services to fit your research and production needs. we develop polyclonal
KMS-Specialist & Customized Biosimilar Service 1. Polyclonal Antibody Development Service KMS offering a variety of Polyclonal Antibody Services to fit your research and production needs. we develop polyclonal
Regulated recruitment and cooperativity in the design of biological regulatory systems
 10.1098/ rsta.2003.1195 Regulated recruitment and cooperativity in the design of biological regulatory systems By M a r k Ptashne Memorial Sloan{Kettering Cancer Center, 1275 York Avenue, Box 595, New
10.1098/ rsta.2003.1195 Regulated recruitment and cooperativity in the design of biological regulatory systems By M a r k Ptashne Memorial Sloan{Kettering Cancer Center, 1275 York Avenue, Box 595, New
Gene Models & Bed format: What they represent.
 GeneModels&Bedformat:Whattheyrepresent. Gene models are hypotheses about the structure of transcripts produced by a gene. Like all models, they may be correct, partly correct, or entirely wrong. Typically,
GeneModels&Bedformat:Whattheyrepresent. Gene models are hypotheses about the structure of transcripts produced by a gene. Like all models, they may be correct, partly correct, or entirely wrong. Typically,
Control of Gene Expression
 Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Effects of Antibiotics on Bacterial Growth and Protein Synthesis: Student Laboratory Manual
 Effects of Antibiotics on Bacterial Growth and Protein Synthesis: Student Laboratory Manual I. Purpose...1 II. Introduction...1 III. Inhibition of Bacterial Growth Protocol...2 IV. Inhibition of in vitro
Effects of Antibiotics on Bacterial Growth and Protein Synthesis: Student Laboratory Manual I. Purpose...1 II. Introduction...1 III. Inhibition of Bacterial Growth Protocol...2 IV. Inhibition of in vitro
In vitro analysis of pri-mirna processing. by Drosha-DGCR8 complex. (Narry Kim s lab)
 In vitro analysis of pri-mirna processing by Drosha-DGCR8 complex (Narry Kim s lab) 1-1. Preparation of radiolabeled pri-mirna transcript The RNA substrate for a cropping reaction can be prepared by in
In vitro analysis of pri-mirna processing by Drosha-DGCR8 complex (Narry Kim s lab) 1-1. Preparation of radiolabeled pri-mirna transcript The RNA substrate for a cropping reaction can be prepared by in
HCS604.03 Exercise 1 Dr. Jones Spring 2005. Recombinant DNA (Molecular Cloning) exercise:
 HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
DNA Isolation Kit for Cells and Tissues
 DNA Isolation Kit for Cells and Tissues for 10 isolations of 00 mg each for tissue or 5 x 10 7 cultured cells Cat. No. 11 81 770 001 Principle Starting material Application Time required Results Benefits
DNA Isolation Kit for Cells and Tissues for 10 isolations of 00 mg each for tissue or 5 x 10 7 cultured cells Cat. No. 11 81 770 001 Principle Starting material Application Time required Results Benefits
Lezioni Dipartimento di Oncologia Farmacologia Molecolare. RNA interference. Giovanna Damia 29 maggio 2006
 Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
Lezioni Dipartimento di Oncologia Farmacologia Molecolare RNA interference Giovanna Damia 29 maggio 2006 RNA INTERFERENCE Sequence-specific gene suppression by dsrnas Gene silencing by dsrna: C. elegans
GenBank, Entrez, & FASTA
 GenBank, Entrez, & FASTA Nucleotide Sequence Databases First generation GenBank is a representative example started as sort of a museum to preserve knowledge of a sequence from first discovery great repositories,
GenBank, Entrez, & FASTA Nucleotide Sequence Databases First generation GenBank is a representative example started as sort of a museum to preserve knowledge of a sequence from first discovery great repositories,
GRS Plasmid Purification Kit Transfection Grade GK73.0002 (2 MaxiPreps)
 1 GRS Plasmid Purification Kit Transfection Grade GK73.0002 (2 MaxiPreps) (FOR RESEARCH ONLY) Sample : Expected Yield : Endotoxin: Format : Operation Time : Elution Volume : 50-400 ml of cultured bacterial
1 GRS Plasmid Purification Kit Transfection Grade GK73.0002 (2 MaxiPreps) (FOR RESEARCH ONLY) Sample : Expected Yield : Endotoxin: Format : Operation Time : Elution Volume : 50-400 ml of cultured bacterial
STOP. Before using this product, please read the Limited Use License statement below:
 STOP Before using this product, please read the Limited Use License statement below: Important Limited Use License information for pdrive5lucia-rgfap The purchase of the pdrive5lucia-rgfap vector conveys
STOP Before using this product, please read the Limited Use License statement below: Important Limited Use License information for pdrive5lucia-rgfap The purchase of the pdrive5lucia-rgfap vector conveys
Using chromosomal laci Q1 to control. high copy number plasmids in Escherichia coli. Weickert; Gene 223; 1998 : 221 231
 Using chromosomal laci Q1 to control expression of genes on high copy number plasmids in Escherichia coli Christopher B Glascock Michael J Christopher B. Glascock, Michael J. Weickert; Gene 223; 1998 :
Using chromosomal laci Q1 to control expression of genes on high copy number plasmids in Escherichia coli Christopher B Glascock Michael J Christopher B. Glascock, Michael J. Weickert; Gene 223; 1998 :
Lecture 13: DNA Technology. DNA Sequencing. DNA Sequencing Genetic Markers - RFLPs polymerase chain reaction (PCR) products of biotechnology
 Lecture 13: DNA Technology DNA Sequencing Genetic Markers - RFLPs polymerase chain reaction (PCR) products of biotechnology DNA Sequencing determine order of nucleotides in a strand of DNA > bases = A,
Lecture 13: DNA Technology DNA Sequencing Genetic Markers - RFLPs polymerase chain reaction (PCR) products of biotechnology DNA Sequencing determine order of nucleotides in a strand of DNA > bases = A,
Instructions. Torpedo sirna. Material. Important Guidelines. Specifications. Quality Control
 is a is a state of the art transfection reagent, specifically designed for the transfer of sirna and mirna into a variety of eukaryotic cell types. is a state of the art transfection reagent, specifically
is a is a state of the art transfection reagent, specifically designed for the transfer of sirna and mirna into a variety of eukaryotic cell types. is a state of the art transfection reagent, specifically
1 Mutation and Genetic Change
 CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
pcas-guide System Validation in Genome Editing
 pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
pcas-guide System Validation in Genome Editing Tagging HSP60 with HA tag genome editing The latest tool in genome editing CRISPR/Cas9 allows for specific genome disruption and replacement in a flexible
About the Kits...2 Description 2 Components 2. Factor Xa Cleavage...3 Small scale optimization 3 Scale-up 3 Monitoring cleavage 4
 Table of Contents About the Kits...2 Description 2 Components 2 Factor Xa Cleavage...3 Small scale optimization 3 Scale-up 3 Monitoring cleavage 4 Factor Xa Capture...5 Capture buffer considerations 5
Table of Contents About the Kits...2 Description 2 Components 2 Factor Xa Cleavage...3 Small scale optimization 3 Scale-up 3 Monitoring cleavage 4 Factor Xa Capture...5 Capture buffer considerations 5
LightSwitch Luciferase Assay System
 Constructs, Assays & Services Assay System Promoter GoClones Synthetic Response Elements Pathway Cell Lines 3 UTR GoClones mirna Mimics & Inhibitors Synthetic mirna Targets Transfection Reagents Assay
Constructs, Assays & Services Assay System Promoter GoClones Synthetic Response Elements Pathway Cell Lines 3 UTR GoClones mirna Mimics & Inhibitors Synthetic mirna Targets Transfection Reagents Assay
Site-Directed Nucleases and Cisgenesis Maria Fedorova, Ph.D.
 Site-Directed Nucleases and Cisgenesis Maria Fedorova, Ph.D. Regulatory Strategy Lead Enabling Technologies DuPont-Pioneer, USA 1 New Plant Breeding Techniques 2007 New Techniques Working Group established
Site-Directed Nucleases and Cisgenesis Maria Fedorova, Ph.D. Regulatory Strategy Lead Enabling Technologies DuPont-Pioneer, USA 1 New Plant Breeding Techniques 2007 New Techniques Working Group established
EU Reference Laboratory for E. coli Department of Veterinary Public Health and Food Safety Unit of Foodborne Zoonoses Istituto Superiore di Sanità
 Identification and characterization of Verocytotoxin-producing Escherichia coli (VTEC) by Real Time PCR amplification of the main virulence genes and the genes associated with the serogroups mainly associated
Identification and characterization of Verocytotoxin-producing Escherichia coli (VTEC) by Real Time PCR amplification of the main virulence genes and the genes associated with the serogroups mainly associated
About the Kits...2 Description 2 Components 2
 Table of Contents About the Kits...2 Description 2 Components 2 Thrombin Cleavage...3 Small scale optimization 3 Scale-up 4 Factors that affect thrombin activity 4 Monitoring cleavage 5 Biotinylated Thrombin
Table of Contents About the Kits...2 Description 2 Components 2 Thrombin Cleavage...3 Small scale optimization 3 Scale-up 4 Factors that affect thrombin activity 4 Monitoring cleavage 5 Biotinylated Thrombin
Genetics 301 Sample Final Examination Spring 2003
 Genetics 301 Sample Final Examination Spring 2003 50 Multiple Choice Questions-(Choose the best answer) 1. A cross between two true breeding lines one with dark blue flowers and one with bright white flowers
Genetics 301 Sample Final Examination Spring 2003 50 Multiple Choice Questions-(Choose the best answer) 1. A cross between two true breeding lines one with dark blue flowers and one with bright white flowers
Outline. interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work?
 Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Outline Outline interfering RNA - What is dat? Brief history of RNA interference. What does it do? How does it work? What is RNA interference? Recently discovered regulatory level. Genome immune system.
Chapter 18: Applications of Immunology
 Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION. Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.
 Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
A quick and simple method for the identi cation of meat species and meat products by PCR assay
 Meat Science 51 (1999) 143±148 A quick and simple method for the identi cation of meat species and meat products by PCR assay T. Matsunaga a, K. Chikuni b *, R. Tanabe b, S. Muroya b, K. Shibata a, J.
Meat Science 51 (1999) 143±148 A quick and simple method for the identi cation of meat species and meat products by PCR assay T. Matsunaga a, K. Chikuni b *, R. Tanabe b, S. Muroya b, K. Shibata a, J.
TIANquick Mini Purification Kit
 TIANquick Mini Purification Kit For purification of PCR products, 100 bp to 20 kb www.tiangen.com TIANquick Mini Purification Kit (Spin column) Cat no. DP203 Kit Contents Contents Buffer BL Buffer PB Buffer
TIANquick Mini Purification Kit For purification of PCR products, 100 bp to 20 kb www.tiangen.com TIANquick Mini Purification Kit (Spin column) Cat no. DP203 Kit Contents Contents Buffer BL Buffer PB Buffer
Chapter 5: Organization and Expression of Immunoglobulin Genes
 Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
Chapter 5: Organization and Expression of Immunoglobulin Genes I. Genetic Model Compatible with Ig Structure A. Two models for Ab structure diversity 1. Germ-line theory: maintained that the genome contributed
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA
 CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
European Medicines Agency
 European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
Towards innovative cropping systems : High Throughput Phenotyping of plant biotic interactions.
 Towards innovative cropping systems : High Throughput Phenotyping of plant biotic interactions. Christophe SALON 1,2 Christian JEUDY 1 Céline BERNARD 2 1 UMR Legumes Genetics and Ecophysiology (LEG), INRA
Towards innovative cropping systems : High Throughput Phenotyping of plant biotic interactions. Christophe SALON 1,2 Christian JEUDY 1 Céline BERNARD 2 1 UMR Legumes Genetics and Ecophysiology (LEG), INRA
Antibody Production Price List
 Antibody Production Price List Presenting Insight Biotechnology s price list for custom polyclonal and monoclonal antibody production services. We are happy to tailor individual packages towards the specific
Antibody Production Price List Presenting Insight Biotechnology s price list for custom polyclonal and monoclonal antibody production services. We are happy to tailor individual packages towards the specific
Nucleic Acid Purity Assessment using A 260 /A 280 Ratios
 Nucleic Acid Purity Assessment using A 260 /A 280 Ratios A common practice in molecular biology is to perform a quick assessment of the purity of nucleic acid samples by determining the ratio of spectrophotometric
Nucleic Acid Purity Assessment using A 260 /A 280 Ratios A common practice in molecular biology is to perform a quick assessment of the purity of nucleic acid samples by determining the ratio of spectrophotometric
MUTATION, DNA REPAIR AND CANCER
 MUTATION, DNA REPAIR AND CANCER 1 Mutation A heritable change in the genetic material Essential to the continuity of life Source of variation for natural selection New mutations are more likely to be harmful
MUTATION, DNA REPAIR AND CANCER 1 Mutation A heritable change in the genetic material Essential to the continuity of life Source of variation for natural selection New mutations are more likely to be harmful
COMMISSION IMPLEMENTING DECISION
 20.3.2014 Official Journal of the European Union L 82/29 COMMISSION IMPLEMENTING DECISION of 18 March 2014 on the organisation of a temporary experiment providing for certain derogations for the marketing
20.3.2014 Official Journal of the European Union L 82/29 COMMISSION IMPLEMENTING DECISION of 18 March 2014 on the organisation of a temporary experiment providing for certain derogations for the marketing
Patent issues in Industrial Biotech:
 Patent issues in Industrial Biotech: Nucleic Acids, Life Forms & Natural Products Konrad Sechley PhD, Vancouver, Canada 18 April, 2016 OVERVIEW Gene patenting Life Forms & Natural Products Conclusions
Patent issues in Industrial Biotech: Nucleic Acids, Life Forms & Natural Products Konrad Sechley PhD, Vancouver, Canada 18 April, 2016 OVERVIEW Gene patenting Life Forms & Natural Products Conclusions
GENE REGULATION. Teacher Packet
 AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
Advanced BioDesign Outlines Solutions. Antibody Overview. by Advanced BioDesign. Project Start. Immunogenicity. Selecting Your Antigen
 Advanced BioDesign Outlines Solutions by Advanced BioDesign Antibody Overview Launching an immunisation programme is an important experimental step that needs care. With Advanced BioDesign, you may develop
Advanced BioDesign Outlines Solutions by Advanced BioDesign Antibody Overview Launching an immunisation programme is an important experimental step that needs care. With Advanced BioDesign, you may develop
