Functional domains and motifs of bacterial type III e ector proteins and their roles in infection
|
|
|
- Kristina Higgins
- 8 years ago
- Views:
Transcription
1 REVIEW ARTICLE Functional domains and motifs of bacterial type III e ector proteins and their roles in infection Paul Dean Institute of Cell and Molecular Bioscience, Medical School, University of Newcastle, Newcastle Upon Tyne, UK Correspondence: Paul Dean, Institute of Cell and Molecular Bioscience, Medical School, University of Newcastle, Newcastle Upon Tyne NE2 4HH, UK. Tel.: ; fax: ; p.dean@ncl.ac.uk Received 13 February 2011; revised 5 April 2011; accepted 7 April DOI: /j x Editor: Miguel Camara Keywords T3SS; TTSS; type three secretion system; pathogens; disease; pathogenesis. Abstract A key feature of the virulence of many bacterial pathogens is the ability to deliver effector proteins into eukaryotic cells via a dedicated type three secretion system (T3SS). Many bacterial pathogens, including species of Chlamydia, Xanthomonas, Pseudomonas, Ralstonia, Shigella, Salmonella, Escherichia and Yersinia, depend on the T3SS to cause disease. T3SS effectors constitute a large and diverse group of virulence proteins that mimic eukaryotic proteins in structure and function. A salient feature of bacterial effectors is their modular architecture, comprising domains or motifs that confer an array of subversive functions within the eukaryotic cell. These domains/motifs therefore represent a fascinating repertoire of molecular determinants with important roles during infection. This review provides a snapshot of our current understanding of bacterial effector domains and motifs where a defined role in infection has been demonstrated. MICROBIOLOGY REVIEWS Introduction Several of the world s most important diseases are caused by bacterial pathogens that deliver effector proteins into eukaryotic host cells using a type three secretion system (T3SS) (Troisfontaines & Cornelis, 2005; Table 1). Bacteria that possess a T3SS cause a wide range of diseases in plants, animals and humans (Troisfontaines & Cornelis, 2005), while others have symbiotic relationships with plant or animal hosts (Preston, 2007; Table 1). The best-studied pathogens to date, which have provided the most knowledge about bacterial effectors, are species of Chlamdyia, Salmonella, Shigella, Yersinia, Pseudomonas, Xanthomonas, Ralstonia and pathogenic Escherichia coli. In all of these organisms, the T3SS is an essential virulence factor, highlighting the central importance of the effector proteins in disease (Coburn et al., 2007). Up to 100 different effector proteins may be delivered into individual host cells by a single bacterium (see Kenny & Valdivia, 2009; Table 1). Effectors are often multifunctional proteins with many overlapping properties and may also cooperate with each other to orchestrate specific responses in the host cell (see Galan, 2009; Fig. 1). In general, although not always, type three effectors display subtle functions inside host cells, often causing more restrained alterations in host cell physiology compared with the more overt effects of bacterial exotoxins. The effector protein family is evolutionarily diverse and exhibits a range of functions within host cells (Fig. 1), targeting most aspects of eukaryotic physiology (Fig. 1). Work over the last decade has been instrumental in defining the commonalities between the functions, molecular mechanisms and structure of bacterial effectors. Indeed, most effectors are viewed as modular proteins, composed of functionally distinct domains or motifs that range from an enzymatic active site to a protein protein interaction or even an organelle-targeting motif. Despite the large size of the bacterial effector family (over 400 predicted proteins), they collectively possess a relatively small subset of domains and motifs that have been shown to play a role in infection (Fig. 2 and Table 2). This review provides a detailed assessment and classification of the effector domains/motifs that have been demonstrated to play a role in the infection process. Where possible, their role in disease has been discussed and is presented in Table 3, although most of the functional studies on effectors to date have been performed on cultured host cells. Throughout the review, motifs are given using the standard amino acid nomenclature, with x referring to any amino acid residue. For brevity, the bacterial origin of the effectors is most often designated according to the genus name of the pathogen.
2 2 P. Dean Table 1. Bacterial pathogens that utilize the T3SS Host Bacterial species Disease caused Repertoires of effectors proven to be secreted or translocated Plant host Animal host Pseudomonas syringae, numerous pathovars Xanthamonas spp. Range of plant diseases, e.g. tomato speck Wide range of plant diseases, e.g. rice bacterial blight and citrus canker HopK1, HopY1, HopAS1, HopU1, HopF2, HopH1, HopC1, HopAT1, HopG1, HopD1, HopQ1, HopR1, HopAM1, HopN1, HopM1, AvrE, AvrB3, HopB1, HopX1, HopZ3, HopAb2, AvrPto, HopE1, HopV1, HopAQ1, HopG1, HopI1, HopA1, HopX1, HopO1, HopT1, AvrRpt2, AvrA, HopW1, HopD1, HopQ1, AvrD1, AvrB2, HopAR1 (see Cunnac et al., 2009 for nomenclature) AvrBs1, AvrBs2, AvrBs3, AvrRxo1, AvrRxv, AvrXccC, AvrXv3, Ecf, HpaA, XopJ, XopX, XopB, XopC, XopD, XopE, XopF, XopN, XopO, XopP, XopQ Ralstonia solancearum Plant wilt on many host species GALA1-7, SKWP1-6, HLK1-3, RipB, PopW, PopP, PopC, RipT, AvrA, PopB, PopA, RipA (many others predicted) Erwinia amylovora Causes fire blight on a range of plant DspE, HrpN, HrpW, HopPtoC, AvrRpt2, EopB species Rhizobium spp. Symbiont; forms nodules on legumes NopL, NopP, NopJ, NopM, NopT, NopB, NopN Pantoea spp. Bacterial wilt on corn and maize WtsE, PthG, HsvG, HsvB Pseudomonas aeruginosa Opportunist pathogenic. Can cause ExoU, ExoY, ExoS, ExoT pneumonia Escherichia coli (EPEC and EHEC), Cirobacter rodentium Diarrhoea (EPEC) or haemorrhagic colitis (EHEC). Cattle commensal (EHEC) Tir, Map, EspF, EspB, EspZ, EspH, EspG, NleA, NleB, NleC, NleD, NleE, NleF, NleG, NleH, NleK, NleL, EspJ, EspK, EspL, EspM, EspY, EspX, EspO, EspW Salmonella enterica serovars Gastroenteritis; typhoid fever AvrA, SipA, SipB, SipC, SipD, SopA, SopB, SopE, SopE2, SptP, SlrP, SopD, SspH1, SteA, SteB, GogB, PipB, SifA, SifB, SopD, SpiC, SseF, SseG, SseI, SseJ, SseK, SspH, SteC, SpvB, SpvC Shigella spp. Bacillary dysentery; shigellosis IpaA, IpaB, IpaC, IpaD, IpaJ, IpgD, IpgB, IcsB, OspC, OspD, OspZ, OspB, OspF, VirA, OspE, OspG, IpaH family Yersinia spp. Bubonic plague (pestis); Gastrointestinal YopE, YopH, YopP/J, YopE, YopM, YopT, YpkA/YopO disease (enterocolitica) Yersinia enterolytica biovar 1B Severe gastrointestinal disease YspA, YspL, YspP, YspF, YspE, YspI, YspK, YspM Photorhabdus spp. Opportunistic pathogen (asymbiotica); insect pathogen (luminescens) LopT Chlamydia spp. Burkholderia spp. Vibrio spp. Obligate intracellular parasites, sexually transmitted disease; can cause blindness Melioidosis (B. pseudomallei); glanders (B. mallei) Gastroenteritis, wound infections (parahaemolyticus); secretory diarrhoea (cholerae) Bordetella spp. Whooping cough BopC/BteA, BopN Aeromonas spp. Opportunistic pathogen; fish/humans AexT, AopB CADD, CT847, tarp, IncA, IncG, CT229, CT813, Cpn0585, Cpn0909, Cpn1020 CHBP, BopE VopL, VopA, VPA450, VopT, VopF, VopS, VopQ Diseases caused by bacteria that are dependent on a T3SS are shown along with a snapshot of the current repertoires of proven effector proteins. Putative effectors based on predictive methods are not given. EPEC, enteropathogenic Escherichia coli; EHEC, enterohaemorrhagic E. coli. Although there are many excellent publications on the functions of effector proteins, this review specifically focuses on examples where effector domains/motifs have a proven role within the host cell during infection, as summarized in Table 2. Evolution of modularity in bacterial effectors Where do bacterial effectors come from? The evolutionary origin of effector genes is open to debate as most bacterial effectors lack significant overall homology to proteins within sequenced genomes. Bacterial effector genes typically exhibit a G/C base composition that is distinctly different from the overall bacterial genome, suggesting that effectors have been acquired by horizontal gene transfer, and much evidence suggests that this is the driving force behind effector evolution (Rohmer et al., 2004). Once acquired by a bacterial strain, effector evolution is then driven by lineage-specific effector functions, dependent on specific protein interactions within the host (Rohmer et al., 2004). A prominent feature shared by bacterial effectors is their modular architecture comprising well-defined regions or
3 Functional domains and motifs of bacterial effector proteins 3 Fig. 1. The cellular functions of bacterial effectors. Activities of bacterial effectors following their translocation into the host cell from a wide range of animal and plant pathogens. Many of the effector functions have been attributed to the domains and motifs described in this review. Major functional classes are subdivided by dotted lines. Effectors are colour-coded depending on species origin (bottom), with host factors given in orange. P, phosphate group; ADPr, ADP-ribose; Ac, acetyl group; Ub., ubiquitin; PI, phosphoinositide. Other common abbreviations are given in the text. domains (Fig. 2) that confer a subversive function. Strikingly, the individual modules within an effector often mediate very different, unrelated functions (see Fig. 2), strongly suggesting that they evolved independent of each other and subsequently combined to form the chimeric effector (Stavrinides et al., 2006). This process of chimerization is a common theme shared by many effectors, as exemplified by the Salmonella effector SptP, which shares a
4 4 P. Dean Fig. 2. The modular architecture of type three effector proteins. Examples of effectors that exhibit high modularity that contain the domains and motifs described in this review. Sec., secretion domain; Chap., chaperonebinding domain; PRR, proline-rich repeats. Abbreviations of common motifs are given in the text. Effectors are not drawn to scale. homologous N-terminal domain with Pseudomonas ExoS and Yersinia YopE, while its C-terminal region is highly homologous to that of another effector, YopH (Kaniga et al., 1996; Fig. 2). This apparent design-by-recombination forms the basis of an attractive hypothesis termed terminal reassortment, proposed by Guttman and colleagues to explain the diversity of bacterial effectors (Stavrinides et al., 2006). In line with this hypothesis, horizontal gene acquisition, followed by genomic shuffling results in the fusion of functional effector modules to the C-termini of different effectors. Providing the newly formed chimeric effector contains an intact N-terminal secretion domain (described below), the new protein is likely to be secreted and translocated into the host cell giving rise to a quick one-step evolutionary mechanism to yield new effectors (Stavrinides et al., 2006). This is evident with the four Salmonella effectors SifA, SspH1, SseJ and SseI, which all share homologous N-termini, but different C-terminal regions (Hansen-Wester et al., 2002). The terminal reassortment hypothesis is strengthened by the finding that 32% of all type three effector families contain chimeric effectors, far greater than that in other analysed protein families, suggesting that this process is important for the evolution of these virulence proteins (Stavrinides et al., 2006).
5 Functional domains and motifs of bacterial effector proteins 5 Table 2. Proven type three effector motifs Subcellular target/activity Effector examples Description of responsible motifs Subcellular targeting Subversive domain/motif Intracellular membrane PipP2, SopD2, YopE, ExoS LFNEF (PipP2), WEK(I/M)xxFF (SopD2); N-terminal Lx 2 Rx 4 L (YopE, ExoS); CLCCFL prenylation motif (SifA) Nucleus (NLS) PopB, PopP2, XopD, AvrBs3 Bipartite NLS: KRKR and KKKKKR (PopB); RRRR and RRQRQ (PopP2). Monopartite NLS: XopD, TAL effectors such as AvrBs3, e.g. KRPR Nucleolus EspF Atypical nuclear/nucleolar targeting motif Plasma membrane ExoU, HopZ, AvrPto, AvrPphB C-terminal MLD containing an essential KAWRN motif (ExoU); PM targeting mediated by Myr and Palm motifs (several plant pathogen effectors) Mitochondrial MTS Map, EspF, SopA N-terminal domain rich in arginine and leucine (Map and EspF); a critical Leu16 (EspF); internal atypical domain (SopA) Golgi STE motif, SseG WEK(I/M)xxFF motif alone; internal golgi targeting domain (SseG) Ser/Thr kinase YpkA Large domain similar to Ser/Thr kinases with critical Asp287 and Lys269 residues RhoGAP AexT, ExoS, ExoT, YopE, SptP Critical arginine finger motif GxLRx 3 T and an essential downstream loop containing the consensus QWGTxGG (AexT, ExoS, ExoT, YopE, SptP); Tir also possesses the arginine finger GxLR RhoGEF SopE, SopE2, BopE Catalytic loop containing the GAGA motif RhoGEF (WxxxE family) Map, SifA, EspM2 Catalytic loop containing the consensus (D/P/E)x 3 AQ; a conserved WxxxE structural motif Lipase SseJ C-terminal domain with a GDSL lipase motif and a SHD catalytic triad Phospholipase ExoU Large cpla 2 domain. SD catalytic dyad of GxSx(G/S) hydrolase motif and a Dx(G/A) active site motif. An essential GGGxxG motif Cysteine protease YopJ, YopT, AvrPphB Catalytic HEC triad: HX 18 EX 44 C (YopJ); catalytic CHD triad (YopT, AvrPphB) and N-terminal Rho-binding domain (YopT), autoproteolytic site GDKG (AvrPphB) E3 Ub ligase SopA, IpaH, SspH1 Active site HECT motif Lx 4 TC (SopA); CxD motif (IpaH and SspH1) ADP-ribosyltransferase HopU1, SpvB, ExoS, ExoT ADPRT conserved region I invariant arginine; region II Gx 9 ST(S/T); region III NAD-binding ExE motif Kinase SteC, YpkA, OspG Kinase subdomain I: nucleotide positioning motif GxGxxG (SteC); subdomain II: invariant lysine residue; subdomain III: invariant glutamic acid Protein phosphatase YopH, SptP PTPase motif HC(x) 5 R(S/T); essential WPD loop PIP phosphatase IpgD, SopB PIPase domain FN(F/V)G(V/I)NE (motif 1) and CKSxKDRTxM (motif 2) py motifs Tir Src kinase phosphorylation motif ExLYxx(I/V) F boxes GALA family The LPx 6 I type F box (all GALA effectors) Lyase OspF, SpvC HopAI1 Lyase motif GDKxH essential for lyase activity WH2 VopL, VopF Homology to WASP WH2; contains the LKKV-like motif D motif SpvC, OspF Docking motif required for specific binding to MAPKs FH1 VopF Highly homologous to mammalian formins (VopF) Actin-related protein binding IpaA, Tir, EspF Phosphotyrosine Nck-binding YDEV motif (Tir), vinculin-binding domain (IpaA), N-WASP-binding motif (EspF) binding site ExoS Mediates binding to the protein FAS to activate the ADPRT domain SH3-binding motif EspF Repeated motif; binds sorting nexin 9 via a conserved RxAPxxP motif J domain HopI1 Hsp70 interaction domain with an invariant HPD motif PDZ1-binding motif Map, NleA, NleH1 Extreme C-terminal motif: -SKI (NleH1), -TRL (Map), -TRV (NleA); mediates effector sorting and protein protein interaction Transcription activation domain AvrBs3 effector family Far C-terminal region of the TAL effectors. Functions in conjunction with the central DNA-binding repeat region Caspase-3 site SipA, SopA Processing site containing the motif DEVD Fic domain VopS Required for AMPylation of Rho GTPases. C-terminal motif: HxFx(D/ E)GNGR The effector activities and subcellular targets are listed together with examples of the effector domains and motifs that have been demonstrated to be responsible. Abbreviations for the domains and motifs are given in the text of the review. Motifs are given in bold. The motif sequence conforms to the annotation used throughout the review.
6 6 P. Dean Table 3. Bacterial effectors and their domains/motifs with proven roles in virulence Bacterial effector Domain or motif with a virulence role References SipA DEVD caspase 3 cleavage site Srikanth et al. (2010) SseI Cys178 unknown function McLaughlin et al. (2009) SseL Cysteine protease site Rytkonen et al. (2007) YopM Leucine-rich repeats (6 15) McPhee et al. (2010) ExoS ADP-ribosyltransferase domain Shaver & Hauser (2004) HopF2Pto Myristoylation motif Wilton et al. (2010) AvrPto Ser149 phosphorylation Anderson et al. (2006) WtsE and AvrE1 WxxxE motif Ham et al. (2009) SifA CAAX box Boucrot et al. (2003) GALA effectors F-box Angot et al. (2006) SseJ Lipase domain SHD triad Ohlson et al. (2005) SpvB ADP-ribosyltransferase domain Lesnick et al. (2001) NleA PDZ domain Lee et al. (2008) YpkA Kinase domain Wiley et al. (2006) YopE GAP domain Black & Bliska (2000) Effector secretion and translocation -- the N-terminus For the majority of effector proteins, the N-terminal region (first residues) encodes the signal sequence required for type three secretion with a more expansive chaperonebinding domain located closely downstream (between residues 50 and 150; for a review, see Ghosh, 2004). The N-terminal secretion signal, while being functionally interchangeable between many effectors (Anderson et al., 1999), does not exhibit a discernable consensus sequence. Some evidence suggests that the effector mrna sequence may be important (Sorg et al., 2005), although this premise is somewhat controversial and appears to depend on the effector in question (Lloyd et al., 2001; Ghosh, 2004). Interestingly, a recently documented prediction method, using a machine-learning approach, analysed the N-termini of 100 effectors and was able to identify putative effectors with over 70% accuracy (Arnold et al., 2009). This was based on the amino acid frequency and the physicochemical properties of the N-terminal residues, suggesting that the amino acid sequence, and not the mrna signal, is most important for secretion (Arnold et al., 2009). Similar approaches have been published recently that collectively show that the amino acid composition of the N-terminal region can be used successfully for effector prediction (reviewed in McDermott et al., 2011). For some effectors, secretion/translocation is also dependent on the C-terminal region. For example, the Salmonella effector SipB requires both the N- and the C-terminal regions for type three secretion as the first 160 residues of SipB are secreted through the flagella secretory system. Only when the C-terminal region of SipB was fused with this N-terminal domain did the protein target the T3SS (Kim et al., 2007). Other examples of effectors that require the C-terminal regions for secretion include SifA (Brown et al., 2006), SipC (Chang et al., 2005) and the E. coli effector Tir (Allen-Vercoe et al., 2005). Subcellular-targeting domains/motifs Mitochondrial-targeting sequences (MTSs) Only a few bacterial effectors are reported to target the mitochondria (Fig. 1). These include Map (Kenny & Jepson, 2000), EspF (Nougayrede & Donnenberg, 2004), SopA (Layton et al., 2005), SipB (Hernandez et al., 2003) and HopG1 (Block et al., 2009). Most mitochondrial proteins in eukaryotes are cytoplasmic and must be imported into the mitochondria via an N-terminal MTS that is cleaved in the mitochondrial matrix (Neupert, 1997). The E. coli effectors EspF and Map bear canonical MTS within their N-termini, with the first 41 and 24 residues of Map and EspF sufficient to target these effectors to host the mitochondria (Fig. 2) (Nagai et al., 2005), where they are subsequently cleaved in the matrix (Kenny & Jepson, 2000; Nougayrede & Donnenberg, 2004; Papatheodorou et al., 2006). Like many eukaryotic mitochondrial proteins, the MTS of EspF contains essential arginine and leucine residues, such as the 16th leucine, which is critical for mitochondrial import (Nagai et al., 2005), and mutation of this residue has been important in identifying the cytoplasmic functions of EspF (Holmes et al., 2010). The two Salmonella effectors SipB and SopA target the mitochondria, but do not have a canonical N-terminal MTS (Hernandez et al., 2003; Layton et al., 2005), although other mechanisms do exist for mitochondrial import (Lutter et al., 2000). Indeed, SopA targets the mitochondria dependent on an internal sequence (residues ) with no apparent role for the N-terminus (residues 1 50) (Layton et al., 2005). Likewise, Pseudomonas
7 Functional domains and motifs of bacterial effector proteins 7 syringae HopG1 does not have a predicted MTS, but green fluorescent protein (GFP) fusions to the C-terminus indicated that the N-terminal residues 1 13 were essential for mitochondrial targeting (Block et al., 2009). However, it is unclear whether native HopG1 targets the mitochondria as ectopic expression of effector fusions can result in anomalous localization patterns. This is exemplified with the E. coli effector Tir, which normally targets the plasma membrane, but when fused to GFP and expressed in host cells, reportedly targets the mitochondria (Malish et al., 2003). Effector- specific membrane-targeting motifs While some effectors encode eukaryotic-like membranetargeting regions, others possess novel sites with no obvious similarity to known eukaryotic proteins. The Salmonella homologues PipB and PipB2 target intracellular membrane compartments and the membranous Salmonella-containing vesicle (SCV) (Knodler et al., 2003). These two effectors share homology mainly due to over 20 pentapeptide repeats (PPR) in their C-terminal domain, bearing the consensus sequence A(N/D)(L/M/F)xx (Knodler et al., 2003). PPR are widely distributed across both pro- and eukaryotes and confer specific structural characteristics such as a superhelical structure (Bateman et al., 1998), but do not confer a common biological function and are likely to be involved in protein protein interactions (Bateman et al., 1998). PipB and PipB2 are considerably less conserved outside of their PPR region, with the C-terminal domain (last 38 residues) of PipB2, but not PipB, targeting this effector to peripheral vesicular compartments (Knodler & Steele-Mortimer, 2005). A C-terminal motif LFNEF of PipB2 is essential for vesicular targeting as alanine substitutions revealed an essential role for each residue in this motif. Subsequent studies have also shown that the LFNEF motif is important for PipB2 function, essential for the recruitment of kinesin-1 to the membrane of the SCV (Henry et al., 2006). SopD2 belongs to a family of Salmonella effectors including SifA, SopD, SseJ and SspH2 termed STE (Salmonella translocated effector) that share a conserved (140 residue) N-terminal region containing the consensus sequence WEK(I/M)xxFF. This motif is required for effector translocation from the bacterium (Miao et al., 1999), but also plays a functional role inside the host cell. Indeed, the STE motif of SopD2 is essential for targeting this effector to late endocytic vesicles as point mutations in this motif (W37P and F44R) abolished endosome targeting of SopD2-GFP (Brown et al., 2006). This indicates that the STE motif is bifunctional, and has been exploited by Salmonella to mediate two distinct functions in different cell types. Interestingly, the WEK(I/M)xxFF motif alone, from a range of STE effectors, is not sufficient to target GFP to endocytic compartments, but instead targets it to the Golgi apparatus (Brown et al., 2006). In addition, the SopD effector, a homologue of SopD2 that also carries the STE motif, was found to reside in the cytoplasm, suggesting that the organelle-targeting function of this motif has been altered in this effector (Brumell et al., 2003). These studies elegantly demonstrate that an effector motif may behave very differently depending on its context and highlight the importance of the entire effector sequence organization in influencing motif function. Pseudomonas aeruginosa ExoS and Yersinia YopE, which share a RhoGAP domain in their C-terminal region (Fig. 2), possess an N-terminally located membrane localization domain (MLD) that targets these effectors to perinuclear vesicles (Pederson et al., 2000; Krall et al., 2004). Although the MLD in YopE shares little similarity to the MLD of ExoS (Krall et al., 2004), both domains are highly hydrophobic and can functionally substitute for each other. Of note, the MLD of YopE and ExoS bear the consensus sequence Lx 2 Rx 4 L (Krall et al., 2004), suggesting possible functional significance, although whether this motif is critical for membrane targeting is unknown. Other effectors that target host cell membranes include SseG, a Salmonella effector that targets the Golgi apparatus dependent on an internal localization domain (residues ) (Salcedo & Holden, 2003). This Golgi-targeting region of SseG has no obvious similarity to eukaryotic proteins, but does share high regional homology to the Edwardsiella effectors EseG and EseF, suggesting functional similarities between them. The E. coli effector NleA also targets the Golgi apparatus, but a targeting motif has not been defined for this protein (Gruenheid et al., 2004). Finally, the P. aeruginosa effector ExoU, a potent phospholipase, targets the plasma membrane via a C-terminal MLD (Fig. 2) containing the essential pentapeptide motif KAWRN (Rabin et al., 2006). This motif was demonstrated to be essential for ubiquitination of this effector on a distal lysine residue, suggesting that membrane targeting of ExoU is a prerequisite for ubiquitination (Stirling et al., 2006). Importantly, the KAWRN motif was shown to be essential for the phospholipase activity of ExoU and its subsequent toxicity in the host cell (Stirling et al., 2006). Membrane-targeting motifs -- prenylation The Salmonella effector SifA is required for the formation of intracellular membranous tubules called Sifs along host cell microtubules (Stein et al., 1996; Beuzon et al., 2000; Brumell et al., 2002). SifA targets intracellular membranes via a C- terminal hexapeptide motif CLCCFL that conforms to the canonical CAAX prenylation box (where AA represents two aliphatic residues) (Reinicke et al., 2005). CAAX boxes target eukaryotic proteins to the plasma membrane by mediating the covalent attachment of a lipid isoprenyl group
8 8 P. Dean to the target CAAX cysteine residue (Gao et al., 2009). Fusion of the last 11 C-terminal residues of SifA to GFP is sufficient to direct GFP to membrane compartments (Boucrot et al., 2003). The removal of the CAAX box from SifA abolished its cellular functions and virulence properties, implying that membrane localization is essential for this effector (Boucrot et al., 2003). Interestingly, the N-terminal region of SifA was also shown to be essential for its function, but only when it was fused to the C-terminal prenylation motif (Boucrot et al., 2003). In addition to prenylation, the cysteine residue adjacent to the SifA CAAX box (CLCCFL) was shown to be acylated following prenylation (Reinicke et al., 2005), although the relevance of this is unclear, but may be linked to protein sorting such as targeting to lipid rafts (Resh, 2006). Membrane-targeting motifs -- myristoylation and palmitoylation Myristoylation (Myr) serves to target proteins to the plasma membrane via the irreversible attachment of a myristoyl group (14-carbon fatty acid) to an extreme N-terminal glycine, which then acts as the membrane anchor. In most cases, the N-terminal methionine must be cleaved off to expose the second residue glycine and therefore almost all Myr motifs begin with a methionine and glycine at the N- terminus and are typically short, approximately six to eight residues in length. In the majority of proteins, Myr sites are also closely associated with a palmitoylation (Palm) site that contains a cysteine at position 3 or 5 to which palmitate or other long-chain fatty acids are attached. Myr and Palm often act in combination to strengthen membrane anchoring (Maurer-Stroh & Eisenhaber, 2004) and both are specific to eukaryotes (Maurer-Stroh & Eisenhaber, 2004), but have also been found in bacterial effectors. All bacterial effectors that have defined Myr and Palm sites are from plant pathogens, particularly P. syringae, and include the HopZ family, HopF2 (Lewis et al., 2008), AvrPhB, AvrRpm1, AvrB, AvrC, AvrPto (Nimchuk et al., 2000; Shan et al., 2000; Maurer-Stroh & Eisenhaber, 2004; Robert-Seilaniantz et al., 2006) and Xanthomonas XopE1/2 (Thieme et al., 2007). Almost all these effectors encode a Myr motif at their extreme N-terminus that targets them to the plant plasma membrane. The consensus motif is not straightforward, but the P. syringae effectors generally conforms to MG(N/C)(I/V)(C/S) noting the palmitoylation cysteines at positions 3 and 5 and the critical Myr glycine residue at position 2. The HopX effector family is slightly different as it displays a highly conserved N-terminal MGLCxSKP Myr motif that differs from the typical pattern (Thieme et al., 2007), but is still crucial in plasma membrane targeting and the subsequent function of the effector (Lewis et al., 2008). Interestingly, the cysteine protease effector AvrPphB from P. syringae undergoes autoproteolysis to expose a new, embedded Myr and Palm site (Nimchuk et al., 2000) that mediates plasma membrane targeting (Dowen et al., 2009). It seems likely that other effectors, with no obvious myristoylation motifs, may utilize similar internal cleavage reactions to promote plasma membrane targeting similar to AvrPphB. Nuclear localization sequences (NLSs) and transcription-related domains A growing number of bacterial effectors (see Fig. 1) target the nucleus including BopN (Nagamatsu et al., 2009), PopP2 (Deslandes et al., 2003), the AvrBs3 family (Szurek et al., 2002), PopB (Gueneron et al., 2000), OspF & OspC (Zurawski et al., 2006), IpaB (Iwai et al., 2007), OspB (Zurawski et al., 2009), YopM (Skrzypek et al., 1998), XopD (Hotson et al., 2003), IpaH9.8 (Toyotome et al., 2001) and SspH1 (Haraga & Miller, 2003). Nuclear import occurs through the nuclear pore complex and may be an active process, dependent on a NLS, or a passive one if the proteins are o 40 kda (Tran & Wente, 2006). Canonical nuclear localization signals are mono- or bipartite, composed of short stretches of basic positively charged amino acids (especially lysine and arginine), and in the case of the bipartite signal, are separated by a stretch of about 9 12 amino acids. The Ralstonia effector PopB contains a bipartite NLS in the centre of the protein, with the sequences KRKR and KKKKKR separated by 13 amino acids (Gueneron et al., 2000). The deletion of either motif prevented PopB from localizing to the nucleus and instead it targeted the cytoplasm (Gueneron et al., 2000). The Ralstonia PopP2 also has a functional bipartite NLS in its N-terminus with the sequence RRRR and RRQRQ separated by 11 amino acids (Deslandes et al., 2003). By contrast, XopD carries a single putative NLS in its C-terminus, although functional studies suggest that this may not be important for its nuclear import, at least when fused to EYFP (Hotson et al., 2003). Interestingly, in addition to being a translocator that inserts into the host plasma membrane, the Shigella effector IpaB has been found to target the nucleus in a cell-cycle-dependent manner, where it interacts with the cell-cycle-regulated protein Mad2L2 (Iwai et al., 2007). IpaB does not possess an NLS, and due to its rather large size (62 kda), has been proposed to enter the nucleus by piggy-back, via its interaction with Mad2L2 (Iwai et al., 2007). This mode of nuclear import may be utilized by other nuclear-targeted effectors that do not possess an NLS. The Xanthomonas AvrBs3 effector families (termed the TAL effectors for transcription activator-like) are highly homologous proteins from plant pathogens that include AvrBs3, PthA, Avrb6, AvrXa7, AvrBs4, PthXo1, AvrHah, Avrxa5 and AvrXa10. These fascinating effectors all mimic
9 Functional domains and motifs of bacterial effector proteins 9 eukaryotic transcription factors and, despite their sequence similarity, induce a diverse array of phenotypes in plants (Kay et al., 2007). The functions of the TAL effectors are dependent on three defined regions (Fig. 2) that are related to their nuclear function: (1) interchangeable NLSs in the C- terminal region (Szurek et al., 2002), (2) highly conserved tandem repeats in the centre of the effector that facilitate effector dimerization essential for nuclear import and DNA binding (Gurlebeck et al., 2005; Kay et al., 2007) and (3) a C-terminal transcription activation domain (TAD) (Zhu et al., 1998). The functional specificity of each TAL effector is determined by the central DNA-binding region, with subtle amino acid changes responsible for binding different host DNA sites (Boch et al., 2009; Moscou & Bogdanove, 2009). By contrast to the central region, the TAD and NLS domains retain functional conservation across the TAL effector, but do not confer lineagespecific functions to the effector as they are functionally interchangeable. Several effectors from different bacterial species display long homologous stretches of leucine-rich repeats (LRR) that bear the consensus sequence Lx 6 Lx 2 I/LPx 3 P. Although LRR are widely distributed in nature, this LPX subtype of LRR is specific to bacterial effectors (Buchanan & Gay, 1996) and is found in YopM, the IpaH family (Fig. 2), SlrP, SspH1 and SspH2 (Miao et al., 1999). Notably, YopM, IpaH9.8 and SspH1 all target the nucleus (Toyotome et al., 2001) (Skrzypek et al., 1998) (Haraga & Miller, 2003), whereas SlrP and SspH2 do not (Haraga & Miller, 2003), suggesting that the LPX domain does not confer this ability alone. The NLS of the LPX effectors has not been studied in depth, with only YopM found to contain an NLS embedded towards the end of its LRR, between the 8th and the 15th LRR, although conflicting reports suggest that a second NLS exists in the C- terminus (Benabdillah et al., 2004; McCoy et al., 2010). Thus, it seems likely that specific residues in the LPX domain of the nuclear-located effectors mediate nuclear targeting. It is therefore interesting to note that the nucleartargeted LPX effectors YopM, SspH1 and IpaH9.8 share the conserved repeating motif NxLTxLPELP, unlike the nonnuclear effectors SlrP and SspH2. Whether this motif is directly related to nuclear targeting or to a common biological function remains to be clarified. Recently, the LRRs of YopM were shown to mediate binding to the host kinase PRK2 and were also shown to be essential for the virulence properties of YopM (McPhee et al., 2010), although the role of nuclear targeting in virulence remains undetermined. Recently, the E. coli effector EspF was found to target the nucleolus, a subnuclear structure involved in ribosome biosynthesis (Dean et al., 2010b). The nuclear/nucleolar localization sequence (NoLS) of EspF resides in the N-terminal region (residues 21 74) between two other cell sorting domains (the MTS and SNX9 motif) and is highly conserved in E. coli strains (Holmes et al., 2010). The NoLS of EspF is unlike other typical nuclear/nucleolar-targeting domains in eukaryotes and viruses, which contain an abundance of arginines and lysines, suggesting that it may represent a novel targeting motif. Alternatively, the NoLS region of EspF may mediate an interaction with a cytoplasmic host protein that then carries EspF into the nucleus/ nucleolus, as suggested above for the nuclear import of IpaB (Iwai et al., 2007). Ubiquitination Ubiquitination is an important eukaryotic process that involves the covalent attachment of ubiquitin to lysine residues of a target protein (Passmore & Barford, 2004). The process occurs in three steps involving (1) activation of ubiquitin by E1-activating enzyme, (2) transferral of ubiquitin to an E2 ubiquitin-conjugating enzyme and (3) E3 ubiquitin ligase catalysing the covalent reaction between ubiquitin and a lysine residue on the substrate (Passmore & Barford, 2004). E3 ligases may bind ubiquitin directly (HECT E3 ligases) or act as a scaffold between E2 and the substrate (RING E3 ligases). As ubiquitination serves to target proteins for degradation by the host proteosome or regulate their function (Passmore & Barford, 2004), it is an obvious target for bacterial pathogens and numerous effectors have evolved mechanisms to manipulate it (Angot et al., 2007; Fig. 1). Effectors that exploit E3 ligases via an F-box F-boxes are well-conserved structural domains (50 residues) in eukaryotic proteins that recruit target proteins to the ubiquitin-conjugating enzyme (Bai et al., 1996). The Agrobacterium effector VirF, although delivered by the type four secretion system, was the first prokaryotic protein found to contain an F-box, which was shown to be essential for virulence (Tzfira et al., 2004). F-boxes have been found in other effectors including Legionella type four effectors (Price et al., 2010) and the Ralstonia type three GALA effector family (Tzfira et al., 2004). GALA effectors, so called because they contain a leucine-rich GAxALA repeat, bear a functional N-terminal F-box that is important for virulence in plant hosts (Angot et al., 2006). Although there is no strict consensus sequence for the F-box, there are several conserved residues along its length as most eukaryotic F-box proteins contain an invariant LPx 10 L motif at the N-terminal region of the F-box that is also conserved in VirF (Tzfira et al., 2004), while the GALA effectors exhibit a less typical LPx 6 I motif (Angot et al., 2006).
10 10 P. Dean Effectors that mimic host E3 ubiquitin ligases The P. syringae effector AvrPtoB suppresses plant programmed cell death (PCD), allowing the pathogen to multiply and cause disease (Abramovitch & Martin, 2005). This inhibitory process is conveyed through a large C-terminal domain (residues ) that has been found to mimic the structure of eukaryotic U-box and RING-finger E3 ubiquitin ligases (Janjusevic et al., 2006), despite no sequence similarity between them. Within the host cell, AvrPtoB interacts with ubiquitin directly (Abramovitch et al., 2006) and causes the ubiquitination of the plant kinase Fen, resulting in its proteosomal degradation (Rosebrock et al., 2007). Remarkably, Fen recognizes the N-terminal region of AvrPtoB leading to PCD in resistant plants, thus revealing that by acquiring the C-terminal E3 ligase domain, AvrPtoB has evolved an effective mechanism to combat its own detection. The Salmonella effector SopA is an E3 ubiquitin ligase (Zhang et al., 2006) with a key role in the induction of inflammation (Wood et al., 2000). Although SopA has weak sequence homology to eukaryotic HECT E3 ligases, it has a highly conserved C-terminal Lx 4 TC motif that is found in the active site of all HECT E3 ligase domains (Huibregtse et al., 1995) (Zhang et al., 2006). The active site cysteine is essential for the E3 ligase activity of SopA (Zhang et al., 2006) and also dictated the ability of SopA to induce inflammation (Zhang et al., 2006). Like AvrPtoB, SopA mimics HECT E3 ligases through convergent evolution, adopting several key structural elements in its C-terminal region in addition to the active site (Diao et al., 2008). The Shigella IpaH effector family defines a new class of E3 ligases (Singer et al., 2008; Zhu et al., 2008), distinct from that mimicked by SopA and AvrPtoB. All IpaH effectors possess a variable N-terminal region containing LRRs and a highly conserved C-terminal region that contains the novel E3 ligase domain (Ashida et al., 2007). An invariant cysteine within this region becomes exposed on an eight-residue loop (Rohde et al., 2007; Singer et al., 2008) that bears the conservedcxdmotifwithboththecysteineandtheaspartic acid residues critical for E3 ligase activity (Singer et al., 2008). IpaH has numerous homologues in other bacterial species, including the Salmonella effector SspH1 that possesses the N- terminal LRR, the C-terminal CxD loop and has been demonstrated to possess E3 ligase activity (Rohde et al., 2007). Effectors that are targets for ubiquitination Ubiquitination increases the molecular mass of target proteins by approximately 8 kda, enabling straightforward identification of many effectors that are ubiquitinated (Marcus et al., 2002; Kubori & Galan, 2003; Ruckdeschel et al., 2006; Stirling et al., 2006). Although single lysines are the target of the ubiquitination reaction, no consensus ubiquitination motifs have been identified to date, and so the preference for the target lysine is difficult to predict (Pickart, 2001). Effectors may be monoubiquitinated such as Salmonella SopB (Marcus et al., 2002) or polyubiquitinated such as Salmonella SopE (Kubori & Galan, 2003), Yersinia YopE (Ruckdeschel et al., 2006) and P. aeruginosa ExoU (Stirling et al., 2006). In most cases, polyubiquitination targets proteins to the proteosome for degradation while monoubiquitination may modulate protein function, as demonstrated for the SopB effector (Knodler et al., 2009). Phosphorylation Protein phosphatases Some of the best-studied bacterial effectors are phosphatases. Yersinia YopH is a protein tyrosine phosphatase (PTPase) that exhibits among the highest-recorded PTPase activity (Zhang et al., 1992). YopH prevents bacteria internalization into macrophages by dephosphorylating a large number of proteins involved in integrin-mediated phagocytosis, such as FAK and paxillin (Persson et al., 1997). The N- terminal region of YopH contains a phosphotyrosine-binding domain that directly targets host tyrosine phosphorylated proteins (Montagna et al., 2001; Ivanov et al., 2005). A proline-rich stretch separates this substrate-binding region from a C-terminal domain that has homology to the catalytic sites of eukaryotic PTPases, with a signature (Ploop) motif HC(x) 5 R(S/T) that is highly conserved in lipid and protein phosphatases (Stuckey et al., 1994). The P- loop is critical for phosphate binding as mutation of the nucleophilic cysteine within this motif abolishes the enzymatic activity of YopH and results in a substrate-trapping protein (Persson et al., 1997). YopH contains a second, more divergent loop (the WPD loop ) found in PTPases that is a 10-residue motif bearing the well-conserved WPD sequence that is also essential for catalysis (Zhang et al., 1994; Hu & Stebbins, 2006). The Salmonella effector SptP, another potent PTPase, shares homology with YopH in its C-terminal region (Kaniga et al., 1996) containing the catalytical HC(x) 5 R(S/T) motif and the WPD loop (Kaniga et al., 1996). Unlike YopH, SptP contains an N-terminal GTPase-activating protein (GAP) domain that, along with the PTPase domain, modulates actin rearrangements in host cells (Fu & Galan, 1999). A substrate of SptP is the ATPase protein VCP (valosincontaining protein), which is dephosphorylated by SptP, promoting bacterial intracellular replication (Humphreys et al., 2009). In addition to YopH and SptP, other effectors that also possess the signature PTPase motifs include the Pseudomonas HopAO1 (Espinosa et al., 2003; Underwood et al., 2007), Chromobacterium CV0974 protein and Aeromonas AopH, although these have not been studied in detail.
11 Functional domains and motifs of bacterial effector proteins 11 Phosphoinositide phosphatases (PiPases) Phosphoinositides are lipid molecules that regulate eukaryotic signalling involved in cytoskeletal reorganization and membrane dynamics (Weber et al., 2009). Because of their pivotal role in these events, phosphoinositide metabolism is a logical target of intracellular bacterial pathogens (Weber et al., 2009). The IpgD effector of the intracellular bacterium Shigella is a PiPase that dephosphorylates phosphatidylinositol-4,5-biphosphate to generate the novel lipid phosphatidylinositol 5-monophosphate (Niebuhr et al., 2002). Salmonella SopB/SigD is another effector PiPase, which, like IpgD, perturbs the actin cytoskeleton and causes the depletion of phosphatidylinositol-4,5-biphosphate (Terebiznik et al., 2002). Both IpgD and SopB possess two domains in their C-terminal region that are conserved in mammalian PiPases (Norris et al., 1998) bearing the consensus sequences FN(F/V)G(V/I)NE (motif 1) and CKSxKDRTxM (motif 2) (see Fig. 2). The conserved cysteine residue in motif 2, which is essential for mammalian inositol 4-phosphatase activity, is also required for the phosphatase activity of SopB and SopB activation of Akt (Steele-Mortimer et al., 2000). A similar finding was obtained when the arginine residue of motif 2 was substituted for an alanine, revealing a critical role for both motifs (Aleman et al., 2005). In addition, SopB also has a C-terminal domain with close homology to the mammalian inositol 5-phosphatase protein synaptojanin and this domain has been shown to play a role in the full phosphatase activity of SopB (Marcus et al., 2001). Effector kinases Effectors that have been reported to possess kinase activity include Salmonella SteC, Yersinia YpkA and Shigella OspG. Like eukaryotic kinases, these effectors contain a series of well-conserved kinase subdomains bearing invariant motifs (see Hanks et al., 1988). SteC for example, which induces F- actin meshwork formation, has the nucleotide positioning motif GxGxxG of kinase subdomain I that has a crucial role in ATP binding (Hanks et al., 1988), an invariant lysine of kinase subdomain II that anchors ATP and an invariant glutamic acid residue in subdomain III that stabilizes the lysine ATP interaction (Poh et al., 2008). Despite SteC containing these subdomains and exhibiting kinase activity in vitro (Poh et al., 2008), it lacks the central catalytic core of many kinases (subdomains VI IX). Other effector kinases, including YpkA (discussed below) and OspG (Kim et al., 2005a), possess the invariant Glu and Lys in subdomains II and III, while the GxGxxG motif of subdomain I is only partially conserved in these effectors. OspG belongs to a larger family of effectors that includes E. coli NleH1/2 (Tobe et al., 2006) and Yersinia YspK (Matsumoto & Young, 2006) that all have the GxGxxG motif and are predicted kinases (Poh et al., 2008). SteC, on the other hand, is not part of the OspG family, but instead shows greater similarity to the eukaryotic kinase Raf-1, which, like SteC, is known to modulate the actin cytoskeleton (Poh et al., 2008). Tyrosine phosphorylated effector proteins A number of effectors, such as Tir and Tarp, are themselves substrates for host kinases (Backert & Selbach, 2005), often evidenced by shift changes in their apparent molecular weight (Kenny et al., 1997; Anderson et al., 2006). The phosphorylation of E. coli Tir on tyrosine 474 is essential for the interaction of Tir with the host protein Nck, leading to actin polymerization (Gruenheid et al., 2001). Interestingly, despite no significant homology between them, the type three effectors Tarp and Tir, and the type four effectors CagA and BepD, share a remarkably similar tyrosine phosphorylation (py) motif with the consensus ExLYxx(I/V) which suggests that they are Src kinases substrates (Backert & Selbach, 2005). The C-terminal residues upstream of the py motif are more divergent between Tarp, Tir, CagA and BepD, and this likely contributes to their different substrate specificities as this region is known to mediate substrate kinase interaction (Backert & Selbach, 2005). It should be noted that bacterial effectors may also be phosphorylated on nontyrosine residues (Anderson et al., 2006) as with the serine phosphorylation of P. syringae AvrPto a step that is required for the virulence function of this effector (Anderson et al., 2006). Effector lipases Lipases are a large group of functionally diverse molecules that include several bacterial effector proteins. The Salmonella effector SseJ, a member of the Salmonella STE effector family that shares similar N-terminal translocation domains (Miao et al., 1999), has a C-terminus that is 29% identical to several members of the GDSL lipase family (Akoh et al., 2004). These lipases are characterized by a GDSL motif (Akoh et al., 2004) and a downstream catalytic triad of SDH residues that are essential for lipolytic activity (Brumlik & Buckley, 1996). SseJ bears both these features, with the catalytic triad spread across the C-terminal region at positions S151, D274 and H384. The SHD triad of SseJ is essential for lipase activity in host cells (Ohlson et al., 2005; Lossi et al., 2008) and contributes to the function of SseJ in Sif formation (Ruiz-Albert et al., 2002). The importance of the lipase activity was revealed in animal studies as the SHD triad played a significant role in SseJ contribution to virulence (Ohlson et al., 2005). The P. aeruginosa effector ExoU is a potent phospholipase with cytotoxic activity and is responsible for epithelial damage in vivo (Phillips et al., 2003; Sato et al., 2003). ExoU shares sequence similarity to a family of widely distributed phospholipases (Schmiel et al., 1998) that include human
12 12 P. Dean cytosolic PLA 2 (cpla 2 ) (Sato et al., 2003), which all have a serine aspartic acid (SD) dyad that is crucial for enzyme activity. Like other bacterial effectors, the overall homology of ExoU to eukaryotic proteins is weak, but there are three well-conserved motifs in ExoU that are important for the activity of many phospholipases (Phillips et al., 2003; Sato et al., 2003). This includes (1) a glycine-rich N-terminal motif that is responsible for stabilizing the tetrahedral intermediate in cpla 2, (2) a serine hydrolase motif GxSx(G/S) and (3) the active site Asp (D) motif Dx(G/A). In addition to the essential requirement of the SD dyad (Sato et al., 2003) that make up the latter two motifs (underlined), the glycines of GxSxG were also essential for ExoU cytotoxicity (Rabin, 2005). Notably, both effector lipases SseJ and ExoU require host factors for their activation (Sato et al., 2003; Lossi et al., 2008), possibly representing an avoidance strategy to restrict the toxicity of these effectors to the host cell. Effector proteases The YopJ effector (also called YopP) is Yersinia s primary antiinflammatory effector (Matsumoto & Young, 2009) that acts by directly binding to mitogen-activated protein kinase (MAPK) kinases and preventing their phosphorylation. It was originally found that the predicted secondary structure of the YopJ family resembled the cysteine protease AVP of adenoviruses (Orth et al., 2000), which also weakly resembled yeast ubiquitin-like protease 1, and it was proposed that YopJ is a ubiquitin-like protein protease. Subsequent experiments revealed that an invariant catalytic triad of histidine, glutamic acid and cysteine residues (H(x)18E(x)44C) was essential for NFkB inhibition by YopJ (Orth et al., 2000). Several other effectors including VopA, SseL, XopJ, AvrA and AvrBst display variations of the catalytic triad (H-(E/N/D)- C), which, at least for AvrA, SseL and VopA, have been shown to be essential for activity (Trosky et al., 2004) (Collier- Hyams et al., 2002; Rytkonen et al., 2007). Indeed, SseL exhibits deubiquitinase activity that is dependent on the catalytic cysteine and was also important for the virulence properties of the effector in vivo (Rytkonen et al., 2007). Recently, YopJ was shown to be an acetyltransferase that acetylates the serine and threonine residues in the activation loop of MAPKK or IKKs preventing their activation by phosphorylation (Mukherjee et al., 2006). Intriguingly, the acetylation activity of YopJ depends on the cysteine protease catalytic triad (Mukherjee et al., 2006). Another family of bacterial effectors, defined by Yersinia YopT and Pseudomonas AvrPphB (Shao et al., 2002), shares an invariant Cys-His-Asp (CHD) triad found in the catalytic core of CA clan cysteine proteases (Zhu et al., 2004). YopT is known to target and cleave prenylated (membrane-bound) Rho GTPases, dependent on its C-terminal cysteine protease domain, preventing them from modulating the cytoskeleton (Aepfelbacher et al., 2003; Shao et al., 2003). In addition to its CHD triad, the proteolytic activity of YopT also depends on its first hundred N-terminal residues and its last eight C- terminal residues. Neither of these regions contains the catalytic triad, but residues of YopT were found to be important in binding to RhoA (Sorg et al., 2003), while the C-terminal domain was proposed to play a direct role in protease activity (Sorg et al., 2003). Although the CHD catalytic residues are invariant between YopT and AvrPphB, the substrate-binding sites are highly divergent and account for the different substrate specificities (Zhu et al., 2004). Other homologues of YopT include the LopT effector from the insect pathogen Photorhabdus, which is known to inhibit phagocytosis (Brugirard-Ricaud et al., 2005). Like YopT, LopT maintains the invariant CHD residues and releases RhoA and Rac from the host plasma membrane (Brugirard- Ricaud et al., 2005). AvrPphB is an interesting effector protease, which, like YopT, contains the catalytic CHD triad, but, when delivered into host cells, is processed to reveal a novel N-terminus (Shao et al., 2002). This exposes embedded Myr and Palm motifs that play an important role in the function of this effector (described above). AvrPphB undergoes autoproteolysis between residues K62 and G63, converting it from a 35- to a 28-kDa protein, dependent on each residue of the CHD triad (Shao et al., 2002; Dowen et al., 2009). A proteaseinactive mutant of the processed form of AvrPphB (28 kda) was unable to induce a plant hypersensitive response, demonstrating the importance of protease activity for this effector (Shao et al., 2002). Interestingly, the autoproteolytic site (GDK-G) of AvrPphB is very similar to the processing site (GDK-S) of its substrate the plant serine threonine kinase PBS1 (Shao et al., 2002). The crystal structure of AvrPphB (Zhu et al., 2004) shows that it resembles the papain-like superfamily of cysteine proteases and as all YopT family members have similar secondary structures, it is likely that they also adopt a similar papain-like tertiary conformation and this is supported by inhibitor studies (Zhu et al., 2004). Finally, the plant pathogen effector, AvrRpt2 from P. syringae, shares structural similarity to staphopain, a CA clan cysteine protease from Staphylococcus (Axtell et al., 2003). AvrRpt2 is known to target the plant host protein RIN4 and cause its elimination (Kim et al., 2005b), dependent on the CA clan catalytic triad of CHD as described above (Axtell et al., 2003). AvrRpt2 requires activation by host cyclophilin (Coaker et al., 2005) and undergoes autocleavage at a VPAFGGW motif that is similar to the two cleavage sites in RIN4 (VPKFGNW and VPKFGDW) (Chisholm et al., 2005; Takemoto & Jones, 2005). Like AvrPphB, mutation of the catalytical triad residues abolished the proteolytic activity of AvrRpt2 (Axtell et al., 2003).
13 Functional domains and motifs of bacterial effector proteins 13 Effector lyases Lyases are a diverse group of enzymes that are able to cleave chemical bonds by mechanisms other than hydrolysis or oxidation and have a wide range of substrates. It was originally reported that the Shigella effector OspF removed a phosphate group from phosphothreonine in MAPK, suggesting that it was acting as a threonine-specific MAPK phosphatase (Li et al., 2007). However, Li et al. (2007) revealed that OspF (and the effectors SpvC and HopAI1) functions as a phosphothreonine lyase by mediating the irreversible removal of phosphate from MAPK (Li et al., 2007), thus representing a novel type of lyase. OspF shares considerable regional homology with SpvC (and HopAI1) including a critical GDKxH motif in the central region of the proteins that is required for phosphothreonine lyase activity (Li et al., 2007). In addition, the N-termini of both SpvC and OspF possess a short canonical D motif found in many MAPK substrates that is required for MAPK docking (Zhu et al., 2007) and is the likely mechanism for MAPK binding by these effectors. Effector adenylate cyclases Cyclic AMP is an important secondary messenger in eukaryotic cells, involved in the regulation of several cellular processes. Numerous pathogens disrupt the levels of camp inside host cells via the deployment of toxins that possess adenylate cyclase activity (Ahuja et al., 2004). ExoY is an effector produced by some strains of P. aeruginosa that shares homology to the calmodulin-activated adenylate cyclase toxins CyaA (Bordetella) and EF (Bacillus). Although ExoY possesses several regions conserved in these toxins, including an ATP/GTP-binding site (Yahr et al., 1998), it does not possess the binding site for calmodulin (Yahr et al., 1998). The ATP-binding site of ExoY is essential for its in vitro adenylate cyclase activity and its ability to cause cell rounding in vivo (Yahr et al., 1998). The absence of the calmodulin-binding site suggests that unlike other bacterial adenylate cyclases, ExoY is not activated by calmodulin and this was indeed the case, although ExoY still requires a host cytosolic factor for activation (Yahr et al., 1998). ADP-ribosyltransferases (ADPRTs) ADP-ribosylation, the transfer of ADP-ribose to a target protein, is used by several bacterial exotoxins and type three effector proteins. HopU1, an effector from P. syringae, is a mono-adprt that suppresses plant immunity dependent on ADP-ribosylation of host RNA-binding proteins (Fu et al., 2007). Like several bacterial ADPRTs (including the toxins VIP2, LT and CT and the effectors ExoS, ExoT and SpvB), HopU1 has three distinct regions with an invariant arginine in region one, a Gx 9 ST(S/T) motif in region two and an ExE biglutamic acid motif in region three (Fu et al., 2007) that is typical of arginine-specific mono-adprts (Domenighini et al., 1994). The SpvB effector ADP-ribosylates actin in infected cells, preventing the conversion of G- into F-actin (Tezcan-Merdol et al., 2001). Importantly, this ADPRT activity of SpvB plays a significant role in disease as mutagenesis of the biglutamic motif attenuated Salmonella virulence in mice (Lesnick et al., 2001). The best-studied ADPRT effectors are the P. aeruginosa homologues ExoT and ExoS. Both proteins are bifunctional due to an N-terminal RhoGAP domain (described below) and a C-terminal ADPRT domain that also bears the critical biglutamic ExE motif described above (Garrity-Ryan et al., 2004). Alanine substitutions revealed that the first glutamic acid in the ExE motif is the active site residue, while the second one mediates the transfer of ADP-ribose to the targeted protein (Garrity-Ryan et al., 2004). ExoT ADPribosylates the SH2 domain of host cytosolic Crk proteins, which are involved in focal adhesion and phagocytosis (Sun & Barbieri, 2003), while ExoS ADP-ribosylates multiple eukaryotic Ras family members such as RhoA, Rac1 and Cdc42 (Henriksson et al., 2002b). ExoS was also shown to depend on the host protein FAS (factor activating ExoS), which bound ExoS through its extreme C-terminal binding site (Fu et al., 1993), and the importance of ExoS activation was demonstrated by removing the last 27 residues of ExoS, which abolished its cytotoxicity (Henriksson et al., 2002a). Interestingly, the arginine finger residue (R146) of the N-terminal GAP domain of ExoS is itself ADP-ribosylated by the C-terminal ADPRT domain, resulting in the inhibition of GAP activity during infection (Riese et al., 2002). This autoregulatory mechanism is an excellent example of how effector domains have coevolved to control effector functions within the host cell. The beauty of selfregulation by ExoS is even more impressive when one considers that the ADPRT activity is itself autoregulated by the domain (Fu et al., 1993) providing a multitiered level of regulation for this effector. Finally, ExoS is an important virulence factor for P. aeruginosa and the virulence properties of ExoS have been assessed by mutational analysis (Shaver & Hauser, 2004), revealing an essential role for the ADPRT domain with a much lesser role in vivo for GAP activity. AMPylation -- a novel effector function One of the benefits of understanding the molecular mechanisms of bacterial effector proteins is that they sometimes reveal themselves as highly novel proteins, mediating previously unseen biochemical properties. VopS of Vibrio parahaemolyticus is one such effector as it was shown to inactivate host Rho, Rac and Cdc42 by the covalent transfer of AMP to a conserved threonine residue on these Rho
14 14 P. Dean GTPases (Yarbrough et al., 2009). The region of VopS that was responsible for AMPylation was a C-terminal region that contained the Fic domain that is widespread in nature. The Fic domain contains an invariant histidine (H) residue within a conserved motif HxFx(D/E)GNGR that was essential for AMPylation by VopS (Yarbrough et al., 2009). Interestingly, a domain with a structure similar to the Fic domain has also been identified in the P. syringae effector AvrB (Kinch et al., 2009); hence, it is likely that AMPylation may be utilized by multiple bacterial effectors. G protein regulatory domains Eukaryotic G proteins bind GTP and cycle between an inactive GDP-bound and an active GTP-bound conformation (Bos et al., 2007). This process is tightly regulated in the host cell by guanine nucleotide exchange factors (GEFs) that activate the G protein, and GAPs that inactivate them (Bos et al., 2007). Several effectors act as GEFs (e.g. SopE, Map and EspM) or GAPs (e.g. AexT, ExoS, ExoT, YopE and SptP) that particularly target G proteins mainly involved in cytoskeletal regulation. Effector GAPs Several well-studied effectors disrupt the actin cytoskeleton via RhoGAP activity (see Fig. 2), including AexT (Fehr et al., 2007), ExoS (Fu & Galan, 1999), ExoT, YopE (Von Pawel- Rammingen et al., 2000) and SptP (Fu & Galan, 1999). Eukaryotic Rho GTPase GAP proteins possess a catalytic arginine carried on an arginine finger that bears the consensus motif GxxRxSG (Juris et al., 2002). The bacterial effector GAPs also possess this critical arginine finger motif with the consensus sequence GxLRx 3 T (Wurtele et al., 2001), for example SptP (GPLRSLMT), ExoS (GALRSLAT), YopE (GPLRGSIT) and AexT (GPLRSLCT). Mutation of the catalytic arginine in this motif abolishes GAP activity for each effector (Fu & Galan, 1999; Black & Bliska, 2000; Von Pawel-Rammingen et al., 2000; Fehr et al., 2007) and was shown to be important in disease as Yersinia carrying a YopE GAP mutant (R144A) was avirulent in mice (Black & Bliska, 2000). Although the arginine finger motif is typical of eukaryotic GAP proteins (Gamblin & Smerdon, 1998), the bacterial GAPs ExoS, ExoT, YopE and SptP all possess a second conserved region comprising a loop 40 residues downstream of the arginine finger. This conserved loop contains the invariant sequence QWGTxGG, which is believed to facilitate hydrogen bond formation between the hydroxyl group of the nucleotide ribose and the effector (Wurtele et al., 2001). Structural elucidation of the GAP domains of SptP, ExoS and YopE reveals that despite diverging at a rapid rate, they share a strikingly high structural similarity (Evdokimov et al., 2002), implying that they have a similar molecular mechanism. This study by Evdokimov et al. (2002) also showed that with the exception of the arginine finger, bacterial GAPs bear little resemblance to their eukaryotic counterparts, suggesting that they arose by convergent evolution. The E. coli Tir effector, while inducing actin polymerization through the Nck/Arp2/3 complex, also possesses GAP activity to downregulate filopodia formation induced by another E. coli effector, Map (Kenny et al., 2002). Like the bacterial GAPs described above, Tir possesses an arginine finger motif, containing the GxLR sequence in its C- terminal region, which was shown to be essential for the downregulation of Map-induced actin polymerization (Kenny et al., 2002). However, unlike the bacterial GAPs, Tir does not possess the second downstream signature motif described above. Effector GEFs The Salmonella effector SopE is a potent GEF protein that directly binds to the Rho GTPases Cdc42 and Rac1 to stimulate GDP/GTP nucleotide exchange (Hardt et al., 1998; Rudolph et al., 1999). SopE is a new type of RhoGEF (Rudolph et al., 1999) as it lacks sequence similarity to known GEF proteins and does not contain the signature DH (Dbl homology) or PH (pleckstrin homology) motifs that are required for catalysis by eukaryotic RhoGEFs (Rossman et al., 2005). Instead, SopE and its effector homologues SopE2 and BopE (Burkholderia) (Upadhyay et al., 2004), which are all confirmed GEFs, share a unique GAGA motif on a catalytic loop that inserts itself between the switch I/II regions of Cdc42 (Schlumberger et al., 2003). Strikingly, the SopE GEF activity is counteracted within the host cell by the GAP activity of the Salmonella effector SptP, providing a fascinating example of the intermolecular crosstalk between these effector domains (Fu & Galan, 1999). G protein mimics or bacterial GEFs? -- the WxxxE family A family of type three effectors that includes EspM, EspT, Map, IpgB1, IpgB2, SifA and SifB were initially believed to be G protein mimics dependent on a WxxxE motif (Alto et al., 2006). However, subsequent work on the crystal structures of Map and SifA revealed a different story implicating these effectors as GEFs that structurally mimic the Salmonella effector SopE, despite sharing no similarity at the sequence level (Ohlson et al., 2008; Huang et al., 2009). In support of this, molecular modelling and nuclear magnetic resonance revealed that EspM2 adopts a SopE-like conformation (Arbeloa et al., 2009). In addition, the ability of Map to induce actin polymerization was dependent on Cdc42 (Kenny et al., 2002) and Map also binds directly to nucleotide-free Cdc42 to promote GTP incorporation
15 Functional domains and motifs of bacterial effector proteins 15 (Huang et al., 2009), strongly suggesting that Map is a RhoGEF protein. The crystal structure of Map in complex with Cdc42 reveals that Map is composed of two bundles of a-helices, forming a V-shaped structure that presents a long catalytic loop (Huang et al., 2009) containing the general consensus sequence (D/P/E)x 3 AQ. Substitution of the conserved alanine or glutamine residues (underlined) abolishes GEF activity (Huang et al., 2009). The essentiality of the WxxxE motif that defines this effector family appears to be a structural one, as this motif was found at the interface of the a-helical bundles, and may thus be essential in maintaining the conformation of the catalytic loop (Huang et al., 2009). Interestingly, despite Map and SopE being unrelated at the sequence levels, these two bacterial GEFs have adopted a conserved functional mechanism for nucleotide exchange and also induce similar conformation changes in Cdc42 (Huang et al., 2009). Several plant pathogen effectors, including most of the AvrE effector family, have recently been reported to contain at least one WxxxE motif (Ham et al., 2009). Indeed, WtsE from Pantoea stewartii and the AvrE1 effector of P. syringae both contain the WxxxE motif, which has been shown to play a central role for the virulence functions of these effectors (Ham et al., 2009). Thus, the WxxxE motif, whether playing a structural role or not, is clearly an important feature of the WxxxE effector family. Microtubule-disrupting effectors Effectors that interfere with microtubules include EseG (Xie et al., 2010), SseG, SseF (Kuhle et al., 2004), EspG (Matsuzawa et al., 2004) and VirA (Yoshida et al., 2002). With the exception of VirA and EspG, little progress has been made regarding the molecular mechanisms of microtubule disruption by these effectors. The Shigella effector VirA binds tubulin subunits directly through a 90 residue region in the centre of the effector (residues ) (Yoshida et al., 2002), resulting in the destabilization of host microtubules (Yoshida et al., 2002). The secondary structure of VirA resembles CA clan cysteine proteases, and Yoshida et al. (2006) showed that degradation of tubulin monomers by VirA is dependent on a critical Cys-34 residue (Yoshida et al., 2006). However, more recent structural data suggest that VirA and its E. coli homologue EspG has no protease active site, but conversely resembles cysteine protease inhibitors, suggesting that VirA may not have direct proteolytic activity (Davis et al., 2008) (Germane et al., 2008). Also, the N-terminal region of VirA that contains the proposed catalytic Cys34 residue (Yoshida et al., 2006) was found to be disordered, like the N-termini of many effectors, making it unlikely to be the active site (Davis et al., 2008). The fact that the VirA structure resembled cysteine protease inhibitors raises the possibility that it may bind to cysteine proteases and, indeed, recent work on the VirA homologue EspG suggests that it can induce the activity of host cysteine proteases (Dean et al., 2010a). These studies on VirA clearly show that although the primary and secondary sequences of an effector may display clues on its identity, structural studies may provide a very different or even a contrary elucidation. Actin nucleation motifs WH2 motifs With the exception of eukaryotic formins, all actin nucleation proteins interact with actin via their WH2 (WASP homology 2) domains (Dominguez, 2009). WH2 domains are short (17 22 residues) regions nearly always found in tandem and forming an N-terminal helix with a conserved downstream LKKV motif (Dominguez, 2009). VopL of Vibrio contains three closely spaced WH2 domains (Liverman et al., 2007) bearing the LKKV-like residues. It induces actin stress fibre formation by directly nucleating actin filaments in vitro, dependent on its three WH2 domains (Liverman et al., 2007). Indeed, mutation of the WH2 domains considerably reduces the actin nucleation activity of VopL, although a single WH2 domain is sufficient to induce this activity (Liverman et al., 2007). The WH2 domains of VopL have high sequence similarity to the WH2 domains of actin-binding proteins WASP, WAVE, Drosophila Spire and the bacterial effectors, Tarp and VopF, which all maintain the LKKV-like motif that is essential for actin binding (Liverman et al., 2007). Of the WH2-containing actin nucleation proteins, the WH2 domain of Chlamydia effector Tarp is the most poorly conserved and it does not encode the LKKV motif, but a more divergent LDDV motif. Tarp binds and nucleates actin directly via a C-terminal actin-binding domain that contains a single WH2 motif that is sufficient for actin nucleation (Jewett et al., 2006). This is unlike all other WH2-containing actin nucleation proteins, which require multiple WH2 motifs, although the finding that Tarp oligomerizes to nucleate actin possibly explains this deviation (Jewett et al., 2006). FH1 domains The Vibrio cholerae effector VopF, like VopL, is able to nucleate actin directly in vitro in the absence of other cellular components (Tam et al., 2007). VopF contains three WH2 domains (discussed above), but also contains a formin homology 1 (FH1) domain, which is typically found in formin family proteins to interact with and nucleate actin (Dominguez, 2009). FH1 domains are highly variable in size, ranging from 15 to 229 residues, and have a high proline content. The VopF FH1 domain is strikingly similar
16 16 P. Dean along its length to the FH1 domain of mammalian formins (Tam et al., 2007) and, while most formins possess both FH1 and FH2 domains, VopF lacks the FH2 domain. It is likely that the FH2 domain of VopF has been substituted by its WH2 domains, and indeed, both types of domains are required for efficient VopF-mediated actin nucleation (Tam et al., 2007). Other actin modulatory domains IpaA is an important Shigella effector that recruits the focal adhesion protein vinculin, an interaction that is believed to promote cell invasion (Tran Van Nhieu et al., 1997). It has been shown that upon binding vinculin, IpaA increases the association of vinculin with F-actin, leading to F-actin depolymerization (Bourdet-Sicard et al., 1999), although this is controversial (Demali et al., 2006). The vinculinbinding site of IpaA was mapped to its last 74 residues (Ramarao et al., 2007), which contains at least two vinculinbinding motifs (Hamiaux et al., 2006). These motifs bind to the N-terminal VD1 domain of vinculin, inducing a conformational change that promotes vinculin binding to F- actin. The IpaA VD1 interaction is very similar to that between VD1 and the host proteins talin and a-actinin, suggesting that IpaA structurally mimics these eukaryotic proteins (Hamiaux et al., 2006). Notably, while the C-terminal region of IpaA shares partial similarity to its Salmonella homologue SipA, SipA does not bind vinculin (Zhou et al., 1999), suggesting no mechanistic relatedness between the two domains (Zhou et al., 1999). Indeed, SipA has a very different function, preventing actin depolymerization by binding actin directly through its C-terminal domain. Other effectors with proven actin modulatory domains include SipC, a Salmonella translocator protein that nucleates and bundles actin through its C- and N-terminal domains, respectively (Hayward & Koronakis, 1999). This dual functionality of SipC is a clear example of functional complementation between effector domains (Hayward & Koronakis, 1999). Interestingly, the SipC homologue IpaC from Shigella possesses the C-terminal actin nucleation domain, but lacks the N-terminal bundling domain, which may explain why Shigella requires the host actin-bundling protein T-fimbrin for efficient uptake (Adam et al., 1995). Finally, the enteropathogenic E. coli effector protein Tir induces actin polymerization following insertion into the host plasma membrane (Kenny et al., 1997). This activity is dependent on Tir binding the eukaryotic adapter protein Nck through a C-terminal Nck-binding site (Gruenheid et al., 2001). The Nck-binding site of Tir bears a conserved YDEV motif, with phosphorylation of the tyrosine essential for Nck binding (Kenny et al., 1997). Nck itself possesses a single SH2 domain and three src homology 3 (SH3) domains, with the former binding to the YDEV Tir motif, while the latter binds to and recruits proteins involved in actin polymerization. Intriguingly, the Nck-binding site of Tir is homologous to that of the vaccinia virus protein A36R that is also tyrosine phosphorylated and is known to recruit Nck (Rottger et al., 1999), implying clear functional similarities between these unrelated proteins. In contrast, the Tir homologue in enterohaemorrhagic E. coli does not possess the Nck-binding site, but instead recruits its own type three effector called Tccp, which substitutes for the Nck protein (Garmendia et al., 2004). The recruitment of Tccp by Tir requires a C-terminal NPY sequence that was found to mediate binding to the host protein IRTKS (insulin receptor tyrosine kinase substrate), which then acts as an adapter between the two effectors (Vingadassalom et al., 2009). Protein--protein interaction elements The J domain J domains are highly conserved modules of about 70 amino acids that are found in Hsp40 and Hsp40-like proteins, including the E. coli chaperone DnaJ (Hennessy et al., 2005). They mediate the interaction of Hsp40-like proteins with their chaperone Hsp70 partner and regulate Hsp70 function (Hennessy et al., 2005). All J domains contain four a-helices with a loop, exposed between helices II and III, encoding a highly conserved HPD motif that is essential for Hsp70 interaction (Kelley, 1998; Hennessy et al., 2005). The P. syringae effector HopI1 contains a C-terminal region with homology to the J domain, containing the invariant HPD motif (Jelenska et al., 2007). HopI1 localizes to the plant chloroplasts, where it suppresses salicylic acid synthesis and disrupts the thylakoid stack structure (Jelenska et al., 2007). These functions are dependent on a functional J-domain as mutation of the HPD motif abolished HopI1 activity (Jelenska et al., 2007). Recently, HopI1 was shown to interact directly with plant Hsp70 proteins via its J domain and cause an increase in Hsp70 abundance and its targeting to the chloroplast (Jelenska et al., 2010). Although Hsp70 was found to be required for HopI1 virulence function, the reason it targets Hsp70 has not yet been determined (Jelenska et al., 2010). SH3-binding motifs The E. coli effector EspF has been shown to recruit and bind the host protein sorting nexin 9 (SNX9), leading to membrane remodelling (Alto et al., 2007). Phage display revealed that EspF binds to the SH3 domain of SNX9 via a repeated motif of RxAPxxP in the C-terminal proline-rich repeat region of EspF (Fig. 2). This repeating motif contains
17 Functional domains and motifs of bacterial effector proteins 17 the canonical PxxP sequence that is known to bind the SH3 domains of eukaryotic proteins (Alto et al., 2007). Interestingly, SH3 motifs are found in numerous effectors, but there are very few documented examples of a functional role for these motifs, the exception being EspF and the E. coli effector Tccp (Aitio et al., 2010). PDZ-binding motifs PDZ (PSD-95/Disk-large/ZO-1) domains are widespread protein protein recognition modules (80 90 residues) found in proteins mainly from multicellular organisms (Hung & Sheng, 2002). They are important in mediating multiprotein complexes at specific subcellular locations within eukaryotic cells. The PDZ domain typically binds to proteins that contain an extreme C-terminal PDZ-binding motif that conforms to the consensus (S/T)x(I/V/L). Several bacterial effectors bear a C-terminal PDZ-binding motif including the E. coli effectors NleH1 (-SKI), Map (-TRL) and NleA (-TRV). All three of these motifs have been shown to be functionally important, acting as cell-sorting signals and mediating the interaction with the host PDZ-containing protein NHERF2 (Martinez et al., 2010). Indeed, the PDZbinding motif of Map allows it to target the host apical plasma membrane while that of NleA mediates trafficking to the Golgi apparatus (Lee et al., 2008; Martinez et al., 2010). This cell sorting function of the PDZ domain was found to be essential for the GEF function of Map (described above) and follows a trend similar to that found in human RhoGEFs, as over 35% possess a C-terminal PDZ-binding motif (Garcia-Mata & Burridge, 2007). Recently, the PDZbinding motif of NleA was shown to be essential for its ability to disrupt intestinal barrier function (Thanabalasuriar et al., 2010) and was also shown to play a role in virulence (Lee et al., 2008). Caspase 3 processing sites Recently, McCormick and colleagues revealed that the Salmonella typhimurium effector SipA possesses a functional caspase-3 cleavage site, containing a DEVD sequence, which lies between the two major functional domains of this effector (Srikanth et al., 2010). Intriguingly, SipA itself induces the activity of caspase-3 within host cells infected with S. typhimurium and mutation of its caspase-3 cleavage site of SipA revealed an important role for this motif in vivo (Srikanth et al., 2010). Remarkably, many other Salmonella effectors possess a centrally located caspase-3 cleavage site, including AvrA, SopB, SifA, SipB and SopA, which, for the latter, has been shown to be functionally important (Srikanth et al., 2010). Thus, caspase-3 cleavage sites may represent a common mechanism to regulate effector function in host cells. YpkA -- a shining example of effector modularity The Yersinia effector YpkA (also called YopO) was purposely excluded from the sections in this review to provide a fitting epilogue of type three effector modularity, although many of the effectors described in this review would have been as suitable. YpkA is an essential Yersinia virulence factor (Galyov et al., 1993) that disrupts the cytoskeleton and inhibits phagocytosis (Wiley et al., 2006). It was originally characterized as a Ser/Thr kinase (Galyov et al., 1993), but subsequent work has shown that it is composed of several important functional domains along its length (Fig. 2), responsible for (1) secretion and chaperone binding, (2) plasma membrane targeting, (3) kinase activity, (4) RhoGDI mimicking and (5) actin binding. YpkA targets the cytoplasmic face of the host plasma membrane (Hakansson et al., 1996) via an N-terminaltargeting domain (residues 20 90) that is essential for the morphological changes induced by this effector (Letzelter et al., 2006). In the same study, the chaperone-binding site (residues 20 77) of YpkA, which directly correlates with the MLD, played no role in bacterial secretion, but was proposed to mask the MLD within the bacteria (Letzelter et al., 2006). Downstream of the MLD is a large kinase domain (residues ; Fig. 2) with homology to eukaryotic Ser/ Thr kinases and in vitro kinase assays have revealed that YpkA autophosphorylates itself (Galyov et al., 1993) with mutation of two critical residues Asp287 and Lys269 abolishing its kinase activity (Juris et al., 2000). The specific kinase activity of YpkA is important for Yersinia disease (Wiley et al., 2006) and plays an intermediate role in cytoskeletal disruption within infected host cells (Juris et al., 2000). YpkA is also known to phosphorylate actin (Juris et al., 2000) and the heterotrimeric G protein Gaq (Navarro et al., 2007), although why YpkA targets Gaq is currently unknown (Navarro et al., 2007). YpkA is synthesized as an inactive kinase, unable to autophosphorylate itself unless specifically activated by monomeric G-, but not F-actin inside the host cell (Trasak et al., 2007), providing an effective mechanism to prevent kinase activity within the bacterium itself. The actin-binding domain of YpkA is found at its extreme C-terminus (Fig. 2) and is critical for the activity of the effector as the removal of the last 20 amino acids abolished YpkA kinase activity (Trasak et al., 2007). The finding that a kinase-deficient YpkA mutant was still able to cause morphological changes in the host cell, similar to the wild-type protein (Dukuzumuremyi et al., 2000), led to the discovery of a second subversive domain of YpkA, in its C-terminal region. This region was found to directly bind the small GTPases RhoA and Rac, but not Cdc42 (Barz et al., 2000; Dukuzumuremyi et al., 2000). The crystal structure of
18 18 P. Dean the C-terminal domain of YpkA in complex with Rac1 finally revealed that this domain of YpkA mimics guanidine nucleotide dissociation inhibitors (GDI) that bind to and lock Rac and RhoA in a GDP-bound off-state (Prehna et al., 2006). Mutations that interfered with GTPase binding completely abolished cytoskeletal disruption by YpkA in infected host cells, demonstrating the importance of this activity during infection (Prehna et al., 2006). Taken together, YpkA is a good example of the multidomain structure of bacterial effector proteins: possessing two prominent, but very different, functional domains that mimic eukaryotic proteins and supported by a smaller cast of essential regulatory or accessory domains. Conclusion The ability of bacterial effectors to take control of their host cell is truly remarkable. These sophisticated molecules are responsible for some of our most devastating diseases, and yet depend on a relatively small set of domains and motifs to do so. As more effectors are uncovered and characterized, this family of molecular determinants will continue to enhance our understanding of host pathogen interactions. I hope this review has provided a glimpse into the ingenuity of bacterial pathogens in evolving such a diverse and fascinating group of virulence proteins. Acknowledgements I am grateful to Profs Brendan Kenny and Martin Embley, Drs David Bulmer, Anjam Khan, Sabrina Muelen and Tom Williams for assistance with this review. I would also like to thank Mrs Philippa Breakspear for collating and organizing all the reading material used in this review. P.D. is supported by a Medical School Faculty Fellowship, University of Newcastle. References Abramovitch RB & Martin GB (2005) AvrPtoB: a bacterial type III effector that both elicits and suppresses programmed cell death associated with plant immunity. FEMS Microbiol Lett 245: 1 8. Abramovitch RB, Janjusevic R, Stebbins CE & Martin GB (2006) Type III effector AvrPtoB requires intrinsic E3 ubiquitin ligase activity to suppress plant cell death and immunity. P Natl Acad Sci USA 103: Adam T, Arpin M, Prevost MC, Gounon P & Sansonetti PJ (1995) Cytoskeletal rearrangements and the functional role of T- plastin during entry of Shigella flexneri into HeLa cells. J Cell Biol 129: Aepfelbacher M, Trasak C, Wilharm G, Wiedemann A, Trulzsch K, Krauss K, Gierschik P & Heesemann J (2003) Characterization of YopT effects on Rho GTPases in Yersinia enterocolitica-infected cells. J Biol Chem 278: Ahuja N, Kumar P & Bhatnagar R (2004) The adenylate cyclase toxins. Crit Rev Microbiol 30: Aitio O, Hellman M, Kazlauskas A, Vingadassalom DF, Leong JM, Saksela K & Permi P (2010) Recognition of tandem PxxP motifs as a unique Src homology 3-binding mode triggers pathogen-driven actin assembly. P Natl Acad Sci USA 107: Akoh CC, Lee GC, Liaw YC, Huang TH & Shaw JF (2004) GDSL family of serine esterases/lipases. Prog Lipid Res 43: Aleman A, Rodriguez-Escudero I, Mallo GV, Cid VJ, Molina M & Rotger R (2005) The amino-terminal non-catalytic region of Salmonella typhimurium SigD affects actin organization in yeast and mammalian cells. Cell Microbiol 7: Allen-Vercoe E, Toh MC, Waddell B, Ho H & DeVinney R (2005) A carboxy-terminal domain of Tir from enterohemorrhagic Escherichia coli O157:H7 (EHEC O157:H7) required for efficient type III secretion. FEMS Microbiol Lett 243: Alto NM, Shao F, Lazar CS et al. (2006) Identification of a bacterial type III effector family with G protein mimicry functions. Cell 124: Alto NM, Weflen AW, Rardin MJ et al. (2007) The type III effector EspF coordinates membrane trafficking by the spatiotemporal activation of two eukaryotic signaling pathways. J Cell Biol 178: Anderson DM, Fouts DE, Collmer A & Schneewind O (1999) Reciprocal secretion of proteins by the bacterial type III machines of plant and animal pathogens suggests universal recognition of mrna targeting signals. P Natl Acad Sci USA 96: Anderson JC, Pascuzzi PE, Xiao F, Sessa G & Martin GB (2006) Host-mediated phosphorylation of type III effector AvrPto promotes Pseudomonas virulence and avirulence in tomato. Plant Cell 18: Angot A, Peeters N, Lechner E, Vailleau F, Baud C, Gentzbittel L, Sartorel E, Genschik P, Boucher C & Genin S (2006) Ralstonia solanacearum requires F-box-like domain-containing type III effectors to promote disease on several host plants. P Natl Acad Sci USA 103: Angot A, Vergunst A, Genin S & Peeters N (2007) Exploitation of eukaryotic ubiquitin signaling pathways by effectors translocated by bacterial type III and type IV secretion systems. PLoS Pathog 3: e3. Arbeloa A, Garnett J, Lillington J, Bulgin RR, Berger CN, Lea SM, Matthews S & Frankel G (2009) EspM2 is a RhoA guanine nucleotide exchange factor. Cell Microbiol 12: Arnold R, Brandmaier S, Kleine F, Tischler P, Heinz E, Behrens S, Niinikoski A, Mewes HW, Horn M & Rattei T (2009) Sequence-based prediction of type III secreted proteins. PLoS Pathog 5: e Ashida H, Toyotome T, Nagai T & Sasakawa C (2007) Shigella chromosomal IpaH proteins are secreted via the type III secretion system and act as effectors. Mol Microbiol 63:
19 Functional domains and motifs of bacterial effector proteins 19 Axtell MJ, Chisholm ST, Dahlbeck D & Staskawicz BJ (2003) Genetic and molecular evidence that the Pseudomonas syringae type III effector protein AvrRpt2 is a cysteine protease. Mol Microbiol 49: Backert S & Selbach M (2005) Tyrosine-phosphorylated bacterial effector proteins: the enemies within. Trends Microbiol 13: Bai C, Sen P, Hofmann K, Ma L, Goebl M, Harper JW & Elledge SJ (1996) SKP1 connects cell cycle regulators to the ubiquitin proteolysis machinery through a novel motif, the F-box. Cell 86: Barz C, Abahji TN, Trulzsch K & Heesemann J (2000) The Yersinia Ser/Thr protein kinase YpkA/YopO directly interacts with the small GTPases RhoA and Rac-1. FEBS Lett 482: Bateman A, Murzin AG & Teichmann SA (1998) Structure and distribution of pentapeptide repeats in bacteria. Protein Sci 7: Benabdillah R, Mota LJ, Lutzelschwab S, Demoinet E & Cornelis GR (2004) Identification of a nuclear targeting signal in YopM from Yersinia spp. Microb Pathogenesis 36: Beuzon CR, Meresse S, Unsworth KE, Ruiz-Albert J, Garvis S, Waterman SR, Ryder TA, Boucrot E & Holden DW (2000) Salmonella maintains the integrity of its intracellular vacuole through the action of SifA. EMBO J 19: Black DS & Bliska JB (2000) The RhoGAP activity of the Yersinia pseudotuberculosis cytotoxin YopE is required for antiphagocytic function and virulence. Mol Microbiol 37: Block A, Guo M, Li G, Elowsky C, Clemente TE & Alfano JR (2009) The Pseudomonas syringae type III effector HopG1 targets mitochondria, alters plant development, and suppresses plant innate immunity. Cell Microbiol 12: Boch J, Scholze H, Schornack S, Landgraf A, Hahn S, Kay S, Lahaye T, Nickstadt A & Bonas U (2009) Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326: Bos JL, Rehmann H & Wittinghofer A (2007) GEFs and GAPs: critical elements in the control of small G proteins. Cell 129: Boucrot E, Beuzon CR, Holden DW, Gorvel JP & Meresse S (2003) Salmonella typhimurium SifA effector protein requires its membrane-anchoring C-terminal hexapeptide for its biological function. J Biol Chem 278: Bourdet-Sicard R, Rudiger M, Jockusch BM, Gounon P, Sansonetti PJ & Nhieu GT (1999) Binding of the Shigella protein IpaA to vinculin induces F-actin depolymerization. EMBO J 18: Brown NF, Szeto J, Jiang X, Coombes BK, Finlay BB & Brumell JH (2006) Mutational analysis of Salmonella translocated effector members SifA and SopD2 reveals domains implicated in translocation, subcellular localization and function. Microbiology 152: Brugirard-Ricaud K, Duchaud E, Givaudan A, Girard PA, Kunst F, Boemare N, Brehelin M & Zumbihl R (2005) Site-specific antiphagocytic function of the Photorhabdus luminescens type III secretion system during insect colonization. Cell Microbiol 7: Brumell JH, Goosney DL & Finlay BB (2002) SifA, a type III secreted effector of Salmonella typhimurium, directs Salmonella-induced filament (Sif) formation along microtubules. Traffic 3: Brumell JH, Kujat-Choy S, Brown NF, Vallance BA, Knodler LA & Finlay BB (2003) SopD2 is a novel type III secreted effector of Salmonella typhimurium that targets late endocytic compartments upon delivery into host cells. Traffic 4: Brumlik MJ & Buckley JT (1996) Identification of the catalytic triad of the lipase/acyltransferase from Aeromonas hydrophila. J Bacteriol 178: Buchanan SG & Gay NJ (1996) Structural and functional diversity in the leucine-rich repeat family of proteins. Prog Biophys Mol Bio 65: Chang J, Chen J & Zhou D (2005) Delineation and characterization of the actin nucleation and effector translocation activities of Salmonella SipC. Mol Microbiol 55: Chisholm ST, Dahlbeck D, Krishnamurthy N, Day B, Sjolander K & Staskawicz BJ (2005) Molecular characterization of proteolytic cleavage sites of the Pseudomonas syringae effector AvrRpt2. P Natl Acad Sci USA 102: Coaker G, Falick A & Staskawicz B (2005) Activation of a phytopathogenic bacterial effector protein by a eukaryotic cyclophilin. Science 308: Coburn B, Sekirov I & Finlay BB (2007) Type III secretion systems and disease. Clin Microbiol Rev 20: Collier-Hyams LS, Zeng H, Sun J, Tomlinson AD, Bao ZQ, Chen H, Madara JL, Orth K & Neish AS (2002) Cutting edge: Salmonella AvrA effector inhibits the key proinflammatory, anti-apoptotic NF-kappa B pathway. J Immunol 169: Cunnac S, Lindeberg M & Collmer A (2009) Pseudomonas syringae type III secretion system effectors: repertoires in search of functions. Curr Opin Microbiol 12: Davis J, Wang J, Tropea JE, Zhang D, Dauter Z, Waugh DS & Wlodawer A (2008) Novel fold of VirA, a type III secretion system effector protein from Shigella flexneri. Protein Sci 17: Dean P, Muhlen S, Quitard S & Kenny B (2010a) The bacterial effectors EspG and EspG2 induce a destructive calpain activity that is kept in check by the co-delivered Tir effector. Cell Microbiol 12: Dean P, Scott JA, Knox AA, Quitard S, Watkins NJ & Kenny B (2010b) The enteropathogenic E. coli effector EspF targets and disrupts the nucleolus by a process regulated by mitochondrial dysfunction. PLoS Pathog 6: e Demali KA, Jue AL & Burridge K (2006) IpaA targets beta1 integrins and rho to promote actin cytoskeleton rearrangements necessary for Shigella entry. J Biol Chem 281:
20 20 P. Dean Deslandes L, Olivier J, Peeters N, Feng DX, Khounlotham M, Boucher C, Somssich I, Genin S & Marco Y (2003) Physical interaction between RRS1-R, a protein conferring resistance to bacterial wilt, and PopP2, a type III effector targeted to the plant nucleus. P Natl Acad Sci USA 100: Diao J, Zhang Y, Huibregtse JM, Zhou D & Chen J (2008) Crystal structure of SopA, a Salmonella effector protein mimicking a eukaryotic ubiquitin ligase. Nat Struct Mol Biol 15: Domenighini M, Magagnoli C, Pizza M & Rappuoli R (1994) Common features of the NAD-binding and catalytic site of ADP-ribosylating toxins. Mol Microbiol 14: Dominguez R (2009) Actin filament nucleation and elongation factors structure function relationships. Crit Rev Biochem Mol 44: Dowen RH, Engel JL, Shao F, Ecker JR & Dixon JE (2009) A family of bacterial cysteine protease type III effectors utilizes acylation-dependent and -independent strategies to localize to plasma membranes. J Biol Chem 284: Dukuzumuremyi JM, Rosqvist R, Hallberg B, Akerstrom B, Wolf- Watz H & Schesser K (2000) The Yersinia protein kinase A is a host factor inducible RhoA/Rac-binding virulence factor. J Biol Chem 275: Espinosa A, Guo M, Tam VC, Fu ZQ & Alfano JR (2003) The Pseudomonas syringae type III-secreted protein HopPtoD2 possesses protein tyrosine phosphatase activity and suppresses programmed cell death in plants. Mol Microbiol 49: Evdokimov AG, Tropea JE, Routzahn KM & Waugh DS (2002) Crystal structure of the Yersinia pestis GTPase activator YopE. Protein Sci 11: Fehr D, Burr SE, Gibert M, d Alayer J, Frey J & Popoff MR (2007) Aeromonas exoenzyme T of Aeromonas salmonicida is a bifunctional protein that targets the host cytoskeleton. J Biol Chem 282: Fu Y & Galan JE (1999) A Salmonella protein antagonizes Rac-1 and Cdc42 to mediate host-cell recovery after bacterial invasion. Nature 401: Fu H, Coburn J & Collier RJ (1993) The eukaryotic host factor that activates exoenzyme S of Pseudomonas aeruginosa is a member of the protein family. P Natl Acad Sci USA 90: Fu ZQ, Guo M, Jeong BR, Tian F, Elthon TE, Cerny RL, Staiger D & Alfano JR (2007) A type III effector ADP-ribosylates RNAbinding proteins and quells plant immunity. Nature 447: Galan JE (2009) Common themes in the design and function of bacterial effectors. Cell Host Microbe 5: Galyov EE, Hakansson S, Forsberg A & Wolf-Watz H (1993) A secreted protein kinase of Yersinia pseudotuberculosis is an indispensable virulence determinant. Nature 361: Gamblin SJ & Smerdon SJ (1998) GTPase-activating proteins and their complexes. Curr Opin Struc Biol 8: Gao J, Liao J & Yang GY (2009) CAAX-box protein, prenylation process and carcinogenesis. Am J Transl Res 1: Garcia-Mata R & Burridge K (2007) Catching a GEF by its tail. Trends Cell Biol 17: Garmendia J, Phillips AD, Carlier MF et al. (2004) TccP is an enterohaemorrhagic Escherichia coli O157:H7 type III effector protein that couples Tir to the actin-cytoskeleton. Cell Microbiol 6: Garrity-Ryan L, Shafikhani S, Balachandran P et al. (2004) The ADP ribosyltransferase domain of Pseudomonas aeruginosa ExoT contributes to its biological activities. Infect Immun 72: Germane KL, Ohi R, Goldberg MB & Spiller BW (2008) Structural and functional studies indicate that Shigella VirA is not a protease and does not directly destabilize microtubules. Biochemistry 47: Ghosh P (2004) Process of protein transport by the type III secretion system. Microbiol Mol Biol R 68: Gruenheid S, DeVinney R, Bladt F, Goosney D, Gelkop S, Gish GD, Pawson T & Finlay BB (2001) Enteropathogenic E. coli Tir binds Nck to initiate actin pedestal formation in host cells. Nat Cell Biol 3: Gruenheid S, Sekirov I, Thomas NA et al. (2004) Identification and characterization of NleA, a non-lee-encoded type III translocated virulence factor of enterohaemorrhagic Escherichia coli O157:H7. Mol Microbiol 51: Gueneron M, Timmers AC, Boucher C & Arlat M (2000) Two novel proteins, PopB, which has functional nuclear localization signals, and PopC, which has a large leucine-rich repeat domain, are secreted through the hrp-secretion apparatus of Ralstonia solanacearum. Mol Microbiol 36: Gurlebeck D, Szurek B & Bonas U (2005) Dimerization of the bacterial effector protein AvrBs3 in the plant cell cytoplasm prior to nuclear import. Plant J 42: Hakansson S, Galyov EE, Rosqvist R & Wolf-Watz H (1996) The Yersinia YpkA Ser/Thr kinase is translocated and subsequently targeted to the inner surface of the HeLa cell plasma membrane. Mol Microbiol 20: Ham JH, Majerczak DR, Nomura K, Mecey C, Uribe F, He SY, Mackey D & Coplin DL (2009) Multiple activities of the plant pathogen type III effector proteins WtsE and AvrE require WxxxE motifs. Mol Plant Microbe In 22: Hamiaux C, van Eerde A, Parsot C, Broos J & Dijkstra BW (2006) Structural mimicry for vinculin activation by IpaA, a virulence factor of Shigella flexneri. EMBO Rep 7: Hanks SK, Quinn AM & Hunter T (1988) The protein kinase family: conserved features and deduced phylogeny of the catalytic domains. Science 241: Hansen-Wester I, Stecher B & Hensel M (2002) Analyses of the evolutionary distribution of Salmonella translocated effectors. Infect Immun 70: Haraga A & Miller SI (2003) A Salmonella enterica serovar typhimurium translocated leucine-rich repeat effector protein inhibits NF-kappa B-dependent gene expression. Infect Immun 71: Hardt WD, Chen LM, Schuebel KE, Bustelo XR & Galan JE (1998) S. typhimurium encodes an activator of Rho GTPases
Lecture 8. Protein Trafficking/Targeting. Protein targeting is necessary for proteins that are destined to work outside the cytoplasm.
 Protein Trafficking/Targeting (8.1) Lecture 8 Protein Trafficking/Targeting Protein targeting is necessary for proteins that are destined to work outside the cytoplasm. Protein targeting is more complex
Protein Trafficking/Targeting (8.1) Lecture 8 Protein Trafficking/Targeting Protein targeting is necessary for proteins that are destined to work outside the cytoplasm. Protein targeting is more complex
Copyright 2000-2003 Mark Brandt, Ph.D. 54
 Pyruvate Oxidation Overview of pyruvate metabolism Pyruvate can be produced in a variety of ways. It is an end product of glycolysis, and can be derived from lactate taken up from the environment (or,
Pyruvate Oxidation Overview of pyruvate metabolism Pyruvate can be produced in a variety of ways. It is an end product of glycolysis, and can be derived from lactate taken up from the environment (or,
Histone modifications. and ChIP. G. Valle - Università di Padova
 Histone modifications and ChIP Histone acetylation It has been shown that the acetylation of lysines is highly dynamic and regulated by the opposing action of two families of enzymes, histone acetyltransferases
Histone modifications and ChIP Histone acetylation It has been shown that the acetylation of lysines is highly dynamic and regulated by the opposing action of two families of enzymes, histone acetyltransferases
Common infection strategies of plant and animal pathogenic bacteria Daniela Büttner and Ulla Bonas
 312 Common infection strategies of plant and animal pathogenic bacteria Daniela Büttner and Ulla Bonas Gram-negative bacterial pathogens use common strategies to invade and colonize plant and animal hosts.
312 Common infection strategies of plant and animal pathogenic bacteria Daniela Büttner and Ulla Bonas Gram-negative bacterial pathogens use common strategies to invade and colonize plant and animal hosts.
2007 7.013 Problem Set 1 KEY
 2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
2007 7.013 Problem Set 1 KEY Due before 5 PM on FRIDAY, February 16, 2007. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. Where in a eukaryotic cell do you
Mechanisms of Hormonal Action Bryant Miles
 Mechanisms of ormonal Action Bryant Miles Multicellular organisms need to coordinate metabolic activities. Complex signaling systems have evolved using chemicals called hormones to regulate cellular activities.
Mechanisms of ormonal Action Bryant Miles Multicellular organisms need to coordinate metabolic activities. Complex signaling systems have evolved using chemicals called hormones to regulate cellular activities.
Regulation of enzyme activity
 1 Regulation of enzyme activity Regulation of enzyme activity is important to coordinate the different metabolic processes. It is also important for homeostasis i.e. to maintain the internal environment
1 Regulation of enzyme activity Regulation of enzyme activity is important to coordinate the different metabolic processes. It is also important for homeostasis i.e. to maintain the internal environment
NO CALCULATORS OR CELL PHONES ALLOWED
 Biol 205 Exam 1 TEST FORM A Spring 2008 NAME Fill out both sides of the Scantron Sheet. On Side 2 be sure to indicate that you have TEST FORM A The answers to Part I should be placed on the SCANTRON SHEET.
Biol 205 Exam 1 TEST FORM A Spring 2008 NAME Fill out both sides of the Scantron Sheet. On Side 2 be sure to indicate that you have TEST FORM A The answers to Part I should be placed on the SCANTRON SHEET.
Biochemistry - I. Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture-11 Enzyme Mechanisms II
 Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture-11 Enzyme Mechanisms II In the last class we studied the enzyme mechanisms of ribonuclease A
Biochemistry - I Prof. S. Dasgupta Department of Chemistry Indian Institute of Technology, Kharagpur Lecture-11 Enzyme Mechanisms II In the last class we studied the enzyme mechanisms of ribonuclease A
Chapter-21b: Hormones and Receptors
 1 hapter-21b: Hormones and Receptors Hormone classes Hormones are classified according to the distance over which they act. 1. Autocrine hormones --- act on the same cell that released them. Interleukin-2
1 hapter-21b: Hormones and Receptors Hormone classes Hormones are classified according to the distance over which they act. 1. Autocrine hormones --- act on the same cell that released them. Interleukin-2
Understanding the immune response to bacterial infections
 Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
IV. -Amino Acids: carboxyl and amino groups bonded to -Carbon. V. Polypeptides and Proteins
 IV. -Amino Acids: carboxyl and amino groups bonded to -Carbon A. Acid/Base properties 1. carboxyl group is proton donor! weak acid 2. amino group is proton acceptor! weak base 3. At physiological ph: H
IV. -Amino Acids: carboxyl and amino groups bonded to -Carbon A. Acid/Base properties 1. carboxyl group is proton donor! weak acid 2. amino group is proton acceptor! weak base 3. At physiological ph: H
BCH401G Lecture 39 Andres
 BCH401G Lecture 39 Andres Lecture Summary: Ribosome: Understand its role in translation and differences between translation in prokaryotes and eukaryotes. Translation: Understand the chemistry of this
BCH401G Lecture 39 Andres Lecture Summary: Ribosome: Understand its role in translation and differences between translation in prokaryotes and eukaryotes. Translation: Understand the chemistry of this
CHAPTER 9 IMMUNOGLOBULIN BIOSYNTHESIS
 CHAPTER 9 IMMUNOGLOBULIN BIOSYNTHESIS Although the process by which a functional gene for immunoglobulin HEAVY and LIGHT CHAINS is formed is highly unusual, the SYNTHESIS, POST- TRANSLATIONAL PROCESSING
CHAPTER 9 IMMUNOGLOBULIN BIOSYNTHESIS Although the process by which a functional gene for immunoglobulin HEAVY and LIGHT CHAINS is formed is highly unusual, the SYNTHESIS, POST- TRANSLATIONAL PROCESSING
Hormones & Chemical Signaling
 Hormones & Chemical Signaling Part 2 modulation of signal pathways and hormone classification & function How are these pathways controlled? Receptors are proteins! Subject to Specificity of binding Competition
Hormones & Chemical Signaling Part 2 modulation of signal pathways and hormone classification & function How are these pathways controlled? Receptors are proteins! Subject to Specificity of binding Competition
RNA and Protein Synthesis
 Name lass Date RN and Protein Synthesis Information and Heredity Q: How does information fl ow from DN to RN to direct the synthesis of proteins? 13.1 What is RN? WHT I KNOW SMPLE NSWER: RN is a nucleic
Name lass Date RN and Protein Synthesis Information and Heredity Q: How does information fl ow from DN to RN to direct the synthesis of proteins? 13.1 What is RN? WHT I KNOW SMPLE NSWER: RN is a nucleic
H H N - C - C 2 R. Three possible forms (not counting R group) depending on ph
 Amino acids - 0 common amino acids there are others found naturally but much less frequently - Common structure for amino acid - C, -N, and functional groups all attached to the alpha carbon N - C - C
Amino acids - 0 common amino acids there are others found naturally but much less frequently - Common structure for amino acid - C, -N, and functional groups all attached to the alpha carbon N - C - C
Diabetes and Insulin Signaling
 Diabetes and Insulin Signaling NATIONAL CENTER FOR CASE STUDY TEACHING IN SCIENCE by Kristy J. Wilson School of Mathematics and Sciences Marian University, Indianapolis, IN Part I Research Orientation
Diabetes and Insulin Signaling NATIONAL CENTER FOR CASE STUDY TEACHING IN SCIENCE by Kristy J. Wilson School of Mathematics and Sciences Marian University, Indianapolis, IN Part I Research Orientation
Copyright 2000-2003 Mark Brandt, Ph.D. 93
 Signal transduction In order to interact properly with their environment, cells need to allow information as well as molecules to cross their cell membranes. Information in many single-celled and all multicellular
Signal transduction In order to interact properly with their environment, cells need to allow information as well as molecules to cross their cell membranes. Information in many single-celled and all multicellular
Name Date Period. 2. When a molecule of double-stranded DNA undergoes replication, it results in
 DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
Biological cell membranes
 Unit 14: Cell biology. 14 2 Biological cell membranes The cell surface membrane surrounds the cell and acts as a barrier between the cell s contents and the environment. The cell membrane has multiple
Unit 14: Cell biology. 14 2 Biological cell membranes The cell surface membrane surrounds the cell and acts as a barrier between the cell s contents and the environment. The cell membrane has multiple
Cell Structure and Function. Eukaryotic Cell: Neuron
 Cell Structure and Function Eukaryotic Cell: Neuron Cell Structure and Function Eukaryotic Cells: Blood Cells Cell Structure and Function Prokaryotic Cells: Bacteria Cell Structure and Function All living
Cell Structure and Function Eukaryotic Cell: Neuron Cell Structure and Function Eukaryotic Cells: Blood Cells Cell Structure and Function Prokaryotic Cells: Bacteria Cell Structure and Function All living
BME 42-620 Engineering Molecular Cell Biology. Lecture 02: Structural and Functional Organization of
 BME 42-620 Engineering Molecular Cell Biology Lecture 02: Structural and Functional Organization of Eukaryotic Cells BME42-620 Lecture 02, September 01, 2011 1 Outline A brief review of the previous lecture
BME 42-620 Engineering Molecular Cell Biology Lecture 02: Structural and Functional Organization of Eukaryotic Cells BME42-620 Lecture 02, September 01, 2011 1 Outline A brief review of the previous lecture
Structure and Function of DNA
 Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Structure and Function of DNA DNA and RNA Structure DNA and RNA are nucleic acids. They consist of chemical units called nucleotides. The nucleotides are joined by a sugar-phosphate backbone. The four
Chem 465 Biochemistry II
 Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Formation of the ribosomal initiation complex for bacterial protein synthesis does not require: A) EF-Tu. B) formylmethionyl
Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Formation of the ribosomal initiation complex for bacterial protein synthesis does not require: A) EF-Tu. B) formylmethionyl
ENZYMES. Serine Proteases Chymotrypsin, Trypsin, Elastase, Subtisisin. Principle of Enzyme Catalysis
 ENZYMES Serine Proteases Chymotrypsin, Trypsin, Elastase, Subtisisin Principle of Enzyme Catalysis Linus Pauling (1946) formulated the first basic principle of enzyme catalysis Enzyme increase the rate
ENZYMES Serine Proteases Chymotrypsin, Trypsin, Elastase, Subtisisin Principle of Enzyme Catalysis Linus Pauling (1946) formulated the first basic principle of enzyme catalysis Enzyme increase the rate
The Steps. 1. Transcription. 2. Transferal. 3. Translation
 Protein Synthesis Protein synthesis is simply the "making of proteins." Although the term itself is easy to understand, the multiple steps that a cell in a plant or animal must go through are not. In order
Protein Synthesis Protein synthesis is simply the "making of proteins." Although the term itself is easy to understand, the multiple steps that a cell in a plant or animal must go through are not. In order
Pipe Cleaner Proteins. Essential question: How does the structure of proteins relate to their function in the cell?
 Pipe Cleaner Proteins GPS: SB1 Students will analyze the nature of the relationships between structures and functions in living cells. Essential question: How does the structure of proteins relate to their
Pipe Cleaner Proteins GPS: SB1 Students will analyze the nature of the relationships between structures and functions in living cells. Essential question: How does the structure of proteins relate to their
Student name ID # 2. (4 pts) What is the terminal electron acceptor in respiration? In photosynthesis? O2, NADP+
 1. Membrane transport. A. (4 pts) What ion couples primary and secondary active transport in animal cells? What ion serves the same function in plant cells? Na+, H+ 2. (4 pts) What is the terminal electron
1. Membrane transport. A. (4 pts) What ion couples primary and secondary active transport in animal cells? What ion serves the same function in plant cells? Na+, H+ 2. (4 pts) What is the terminal electron
ISTEP+: Biology I End-of-Course Assessment Released Items and Scoring Notes
 ISTEP+: Biology I End-of-Course Assessment Released Items and Scoring Notes Page 1 of 22 Introduction Indiana students enrolled in Biology I participated in the ISTEP+: Biology I Graduation Examination
ISTEP+: Biology I End-of-Course Assessment Released Items and Scoring Notes Page 1 of 22 Introduction Indiana students enrolled in Biology I participated in the ISTEP+: Biology I Graduation Examination
Lecture 6. Regulation of Protein Synthesis at the Translational Level
 Regulation of Protein Synthesis (6.1) Lecture 6 Regulation of Protein Synthesis at the Translational Level Comparison of EF-Tu-GDP and EF-Tu-GTP conformations EF-Tu-GDP EF-Tu-GTP Next: Comparison of GDP
Regulation of Protein Synthesis (6.1) Lecture 6 Regulation of Protein Synthesis at the Translational Level Comparison of EF-Tu-GDP and EF-Tu-GTP conformations EF-Tu-GDP EF-Tu-GTP Next: Comparison of GDP
Chapter 8. Summary and Perspectives
 Chapter 8 Summary and Perspectives 131 Chapter 8 Summary Overexpression of the multidrug resistance protein MRP1 confer multidrug resistance (MDR) to cancer cells. The contents of this thesis describe
Chapter 8 Summary and Perspectives 131 Chapter 8 Summary Overexpression of the multidrug resistance protein MRP1 confer multidrug resistance (MDR) to cancer cells. The contents of this thesis describe
Actions of Hormones on Target Cells Page 1. Actions of Hormones on Target Cells Page 2. Goals/ What You Need to Know Goals What You Need to Know
 Actions of Hormones on Target Cells Graphics are used with permission of: Pearson Education Inc., publishing as Benjamin Cummings (http://www.aw-bc.com) Page 1. Actions of Hormones on Target Cells Hormones
Actions of Hormones on Target Cells Graphics are used with permission of: Pearson Education Inc., publishing as Benjamin Cummings (http://www.aw-bc.com) Page 1. Actions of Hormones on Target Cells Hormones
Amino Acids, Peptides, Proteins
 Amino Acids, Peptides, Proteins Functions of proteins: Enzymes Transport and Storage Motion, muscle contraction Hormones Mechanical support Immune protection (Antibodies) Generate and transmit nerve impulses
Amino Acids, Peptides, Proteins Functions of proteins: Enzymes Transport and Storage Motion, muscle contraction Hormones Mechanical support Immune protection (Antibodies) Generate and transmit nerve impulses
Module 3 Questions. 7. Chemotaxis is an example of signal transduction. Explain, with the use of diagrams.
 Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Part A: Amino Acids and Peptides (Is the peptide IAG the same as the peptide GAI?)
 ChemActivity 46 Amino Acids, Polypeptides and Proteins 1 ChemActivity 46 Part A: Amino Acids and Peptides (Is the peptide IAG the same as the peptide GAI?) Model 1: The 20 Amino Acids at Biological p See
ChemActivity 46 Amino Acids, Polypeptides and Proteins 1 ChemActivity 46 Part A: Amino Acids and Peptides (Is the peptide IAG the same as the peptide GAI?) Model 1: The 20 Amino Acids at Biological p See
From DNA to Protein. Proteins. Chapter 13. Prokaryotes and Eukaryotes. The Path From Genes to Proteins. All proteins consist of polypeptide chains
 Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
The Lipid Bilayer Is a Two-Dimensional Fluid
 The Lipid Bilayer Is a Two-Dimensional Fluid The aqueous environment inside and outside a cell prevents membrane lipids from escaping from bilayer, but nothing stops these molecules from moving about and
The Lipid Bilayer Is a Two-Dimensional Fluid The aqueous environment inside and outside a cell prevents membrane lipids from escaping from bilayer, but nothing stops these molecules from moving about and
CSC 2427: Algorithms for Molecular Biology Spring 2006. Lecture 16 March 10
 CSC 2427: Algorithms for Molecular Biology Spring 2006 Lecture 16 March 10 Lecturer: Michael Brudno Scribe: Jim Huang 16.1 Overview of proteins Proteins are long chains of amino acids (AA) which are produced
CSC 2427: Algorithms for Molecular Biology Spring 2006 Lecture 16 March 10 Lecturer: Michael Brudno Scribe: Jim Huang 16.1 Overview of proteins Proteins are long chains of amino acids (AA) which are produced
Cell Biology Questions and Learning Objectives
 Cell Biology Questions and Learning Objectives (with hypothetical learning materials that might populate the objective) The topics and central questions listed here are typical for an introductory undergraduate
Cell Biology Questions and Learning Objectives (with hypothetical learning materials that might populate the objective) The topics and central questions listed here are typical for an introductory undergraduate
Plasma Membrane hydrophilic polar heads
 The Parts of the Cell 3 main parts in ALL cells: plasma membrane, cytoplasm, genetic material this is about the parts of a generic eukaryotic cell Plasma Membrane -is a fluid mosaic model membrane is fluid
The Parts of the Cell 3 main parts in ALL cells: plasma membrane, cytoplasm, genetic material this is about the parts of a generic eukaryotic cell Plasma Membrane -is a fluid mosaic model membrane is fluid
Computational Systems Biology. Lecture 2: Enzymes
 Computational Systems Biology Lecture 2: Enzymes 1 Images from: David L. Nelson, Lehninger Principles of Biochemistry, IV Edition, Freeman ed. or under creative commons license (search for images at http://search.creativecommons.org/)
Computational Systems Biology Lecture 2: Enzymes 1 Images from: David L. Nelson, Lehninger Principles of Biochemistry, IV Edition, Freeman ed. or under creative commons license (search for images at http://search.creativecommons.org/)
Translation. Translation: Assembly of polypeptides on a ribosome
 Translation Translation: Assembly of polypeptides on a ribosome Living cells devote more energy to the synthesis of proteins than to any other aspect of metabolism. About a third of the dry mass of a cell
Translation Translation: Assembly of polypeptides on a ribosome Living cells devote more energy to the synthesis of proteins than to any other aspect of metabolism. About a third of the dry mass of a cell
The Organic Chemistry of Amino Acids, Peptides, and Proteins
 Essential rganic Chemistry Chapter 16 The rganic Chemistry of Amino Acids, Peptides, and Proteins Amino Acids a-amino carboxylic acids. The building blocks from which proteins are made. H 2 N C 2 H Note:
Essential rganic Chemistry Chapter 16 The rganic Chemistry of Amino Acids, Peptides, and Proteins Amino Acids a-amino carboxylic acids. The building blocks from which proteins are made. H 2 N C 2 H Note:
Molecular Cell Biology. Prof. D. Karunagaran. Department of Biotechnology. Indian Institute of Technology Madras. Module 7 Cell Signaling Mechanisms
 Molecular Cell Biology Prof. D. Karunagaran Department of Biotechnology Indian Institute of Technology Madras Module 7 Cell Signaling Mechanisms Lecture 2 GPCR Signaling Receptors - G protein coupled receptors
Molecular Cell Biology Prof. D. Karunagaran Department of Biotechnology Indian Institute of Technology Madras Module 7 Cell Signaling Mechanisms Lecture 2 GPCR Signaling Receptors - G protein coupled receptors
Advanced Medicinal & Pharmaceutical Chemistry CHEM 5412 Dept. of Chemistry, TAMUK
 Advanced Medicinal & Pharmaceutical Chemistry CHEM 5412 Dept. of Chemistry, TAMUK Dai Lu, Ph.D. dlu@tamhsc.edu Tel: 361-221-0745 Office: RCOP, Room 307 Drug Discovery and Development Drug Molecules Medicinal
Advanced Medicinal & Pharmaceutical Chemistry CHEM 5412 Dept. of Chemistry, TAMUK Dai Lu, Ph.D. dlu@tamhsc.edu Tel: 361-221-0745 Office: RCOP, Room 307 Drug Discovery and Development Drug Molecules Medicinal
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Compartmentalization of the Cell. Objectives. Recommended Reading. Professor Alfred Cuschieri. Department of Anatomy University of Malta
 Compartmentalization of the Cell Professor Alfred Cuschieri Department of Anatomy University of Malta Objectives By the end of this session the student should be able to: 1. Identify the different organelles
Compartmentalization of the Cell Professor Alfred Cuschieri Department of Anatomy University of Malta Objectives By the end of this session the student should be able to: 1. Identify the different organelles
How To Understand The Chemistry Of An Enzyme
 Chapt. 8 Enzymes as catalysts Ch. 8 Enzymes as catalysts Student Learning Outcomes: Explain general features of enzymes as catalysts: Substrate -> Product Describe nature of catalytic sites general mechanisms
Chapt. 8 Enzymes as catalysts Ch. 8 Enzymes as catalysts Student Learning Outcomes: Explain general features of enzymes as catalysts: Substrate -> Product Describe nature of catalytic sites general mechanisms
Preliminary MFM Quiz
 Preliminary MFM Quiz 1. The major carrier of chemical energy in all cells is: A) adenosine monophosphate B) adenosine diphosphate C) adenosine trisphosphate D) guanosine trisphosphate E) carbamoyl phosphate
Preliminary MFM Quiz 1. The major carrier of chemical energy in all cells is: A) adenosine monophosphate B) adenosine diphosphate C) adenosine trisphosphate D) guanosine trisphosphate E) carbamoyl phosphate
博 士 論 文 ( 要 約 ) A study on enzymatic synthesis of. stable cyclized peptides which. inhibit protein-protein interactions
 博 士 論 文 ( 要 約 ) 論 文 題 目 A study on enzymatic synthesis of stable cyclized peptides which inhibit protein-protein interactions ( 蛋 白 質 間 相 互 作 用 を 阻 害 する 安 定 な 環 状 化 ペプチドの 酵 素 合 成 に 関 する 研 究 ) 氏 名 張 静 1
博 士 論 文 ( 要 約 ) 論 文 題 目 A study on enzymatic synthesis of stable cyclized peptides which inhibit protein-protein interactions ( 蛋 白 質 間 相 互 作 用 を 阻 害 する 安 定 な 環 状 化 ペプチドの 酵 素 合 成 に 関 する 研 究 ) 氏 名 張 静 1
Gymnázium, Brno, Slovanské nám. 7, WORKBOOK - Biology WORKBOOK. http://agb.gymnaslo.cz
 WORKBOOK http://agb.gymnaslo.cz Biology Subject: Teacher: Iva Kubištová Student:.. School year:../ This material was prepared with using http://biologygmh.com/ Topics: 1. 2. 3. 4. 5. 6. Viruses and Bacteria
WORKBOOK http://agb.gymnaslo.cz Biology Subject: Teacher: Iva Kubištová Student:.. School year:../ This material was prepared with using http://biologygmh.com/ Topics: 1. 2. 3. 4. 5. 6. Viruses and Bacteria
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Cell Structure & Function!
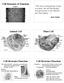 Cell Structure & Function! Chapter 3! The most exciting phrase to hear in science, the one that heralds new discoveries, is not 'Eureka!' but 'That's funny.! -- Isaac Asimov Animal Cell Plant Cell Cell
Cell Structure & Function! Chapter 3! The most exciting phrase to hear in science, the one that heralds new discoveries, is not 'Eureka!' but 'That's funny.! -- Isaac Asimov Animal Cell Plant Cell Cell
MCAS Biology. Review Packet
 MCAS Biology Review Packet 1 Name Class Date 1. Define organic. THE CHEMISTRY OF LIFE 2. All living things are made up of 6 essential elements: SPONCH. Name the six elements of life. S N P C O H 3. Elements
MCAS Biology Review Packet 1 Name Class Date 1. Define organic. THE CHEMISTRY OF LIFE 2. All living things are made up of 6 essential elements: SPONCH. Name the six elements of life. S N P C O H 3. Elements
Transcription in prokaryotes. Elongation and termination
 Transcription in prokaryotes Elongation and termination After initiation the σ factor leaves the scene. Core polymerase is conducting the elongation of the chain. The core polymerase contains main nucleotide
Transcription in prokaryotes Elongation and termination After initiation the σ factor leaves the scene. Core polymerase is conducting the elongation of the chain. The core polymerase contains main nucleotide
Lecture Series 7. From DNA to Protein. Genotype to Phenotype. Reading Assignments. A. Genes and the Synthesis of Polypeptides
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Complex multicellular organisms are produced by cells that switch genes on and off during development.
 Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Lecture 4 Cell Membranes & Organelles
 Lecture 4 Cell Membranes & Organelles Structure of Animal Cells The Phospholipid Structure Phospholipid structure Encases all living cells Its basic structure is represented by the fluidmosaic model Phospholipid
Lecture 4 Cell Membranes & Organelles Structure of Animal Cells The Phospholipid Structure Phospholipid structure Encases all living cells Its basic structure is represented by the fluidmosaic model Phospholipid
Non-standard amino acids. mgr Adrian Jasiński Theoretical Molecular Biophysics/Bioinformatics Group
 Non-standard amino acids mgr Adrian Jasiński Theoretical Molecular Biophysics/Bioinformatics Group Outline Introduction Non-standard amino acids Variations to the standard genetic code Modifications to
Non-standard amino acids mgr Adrian Jasiński Theoretical Molecular Biophysics/Bioinformatics Group Outline Introduction Non-standard amino acids Variations to the standard genetic code Modifications to
Control of Gene Expression
 Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Given these characteristics of life, which of the following objects is considered a living organism? W. X. Y. Z.
 Cell Structure and Organization 1. All living things must possess certain characteristics. They are all composed of one or more cells. They can grow, reproduce, and pass their genes on to their offspring.
Cell Structure and Organization 1. All living things must possess certain characteristics. They are all composed of one or more cells. They can grow, reproduce, and pass their genes on to their offspring.
Chapter 2: Cell Structure and Function pg. 70-107
 UNIT 1: Biochemistry Chapter 2: Cell Structure and Function pg. 70-107 Organelles are internal structures that carry out specialized functions, interacting and complementing each other. Animal and plant
UNIT 1: Biochemistry Chapter 2: Cell Structure and Function pg. 70-107 Organelles are internal structures that carry out specialized functions, interacting and complementing each other. Animal and plant
Sickle cell anemia: Altered beta chain Single AA change (#6 Glu to Val) Consequence: Protein polymerizes Change in RBC shape ---> phenotypes
 Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
Protein Structure Polypeptide: Protein: Therefore: Example: Single chain of amino acids 1 or more polypeptide chains All polypeptides are proteins Some proteins contain >1 polypeptide Hemoglobin (O 2 binding
1 Mutation and Genetic Change
 CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
PRACTICE TEST QUESTIONS
 PART A: MULTIPLE CHOICE QUESTIONS PRACTICE TEST QUESTIONS DNA & PROTEIN SYNTHESIS B 1. One of the functions of DNA is to A. secrete vacuoles. B. make copies of itself. C. join amino acids to each other.
PART A: MULTIPLE CHOICE QUESTIONS PRACTICE TEST QUESTIONS DNA & PROTEIN SYNTHESIS B 1. One of the functions of DNA is to A. secrete vacuoles. B. make copies of itself. C. join amino acids to each other.
Regulation of the Citric Acid Cycle
 Regulation of the itric Acid ycle I. hanges in Free Energy February 17, 2003 Bryant Miles kj/mol 40 20 0 20 40 60 80 Reaction DGo' DG TA Free Energy hanges 1 2 3 4 5 6 7 8 9 1.) itrate Synthase 2.) Aconitase
Regulation of the itric Acid ycle I. hanges in Free Energy February 17, 2003 Bryant Miles kj/mol 40 20 0 20 40 60 80 Reaction DGo' DG TA Free Energy hanges 1 2 3 4 5 6 7 8 9 1.) itrate Synthase 2.) Aconitase
RNA & Protein Synthesis
 RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
RNA & Protein Synthesis Genes send messages to cellular machinery RNA Plays a major role in process Process has three phases (Genetic) Transcription (Genetic) Translation Protein Synthesis RNA Synthesis
Six major functions of membrane proteins: Transport Enzymatic activity
 CH 7 Membranes Cellular Membranes Phospholipids are the most abundant lipid in the plasma membrane. Phospholipids are amphipathic molecules, containing hydrophobic and hydrophilic regions. The fluid mosaic
CH 7 Membranes Cellular Membranes Phospholipids are the most abundant lipid in the plasma membrane. Phospholipids are amphipathic molecules, containing hydrophobic and hydrophilic regions. The fluid mosaic
Specific problems. The genetic code. The genetic code. Adaptor molecules match amino acids to mrna codons
 Tutorial II Gene expression: mrna translation and protein synthesis Piergiorgio Percipalle, PhD Program Control of gene transcription and RNA processing mrna translation and protein synthesis KAROLINSKA
Tutorial II Gene expression: mrna translation and protein synthesis Piergiorgio Percipalle, PhD Program Control of gene transcription and RNA processing mrna translation and protein synthesis KAROLINSKA
Review of the Cell and Its Organelles
 Biology Learning Centre Review of the Cell and Its Organelles Tips for most effective learning of this material: Memorize the names and structures over several days. This will help you retain what you
Biology Learning Centre Review of the Cell and Its Organelles Tips for most effective learning of this material: Memorize the names and structures over several days. This will help you retain what you
Ms. Campbell Protein Synthesis Practice Questions Regents L.E.
 Name Student # Ms. Campbell Protein Synthesis Practice Questions Regents L.E. 1. A sequence of three nitrogenous bases in a messenger-rna molecule is known as a 1) codon 2) gene 3) polypeptide 4) nucleotide
Name Student # Ms. Campbell Protein Synthesis Practice Questions Regents L.E. 1. A sequence of three nitrogenous bases in a messenger-rna molecule is known as a 1) codon 2) gene 3) polypeptide 4) nucleotide
Lecture 15: Enzymes & Kinetics Mechanisms
 ROLE OF THE TRANSITION STATE Lecture 15: Enzymes & Kinetics Mechanisms Consider the reaction: H-O-H + Cl - H-O δ- H Cl δ- HO - + H-Cl Reactants Transition state Products Margaret A. Daugherty Fall 2004
ROLE OF THE TRANSITION STATE Lecture 15: Enzymes & Kinetics Mechanisms Consider the reaction: H-O-H + Cl - H-O δ- H Cl δ- HO - + H-Cl Reactants Transition state Products Margaret A. Daugherty Fall 2004
Quick Hit Activity Using UIL Science Contests For Formative and Summative Assessments of Pre-AP and AP Biology Students
 Quick Hit Activity Using UIL Science Contests For Formative and Summative Assessments of Pre-AP and AP Biology Students Activity Title: Quick Hit Goal of Activity: To perform formative and summative assessments
Quick Hit Activity Using UIL Science Contests For Formative and Summative Assessments of Pre-AP and AP Biology Students Activity Title: Quick Hit Goal of Activity: To perform formative and summative assessments
AP Biology Essential Knowledge Student Diagnostic
 AP Biology Essential Knowledge Student Diagnostic Background The Essential Knowledge statements provided in the AP Biology Curriculum Framework are scientific claims describing phenomenon occurring in
AP Biology Essential Knowledge Student Diagnostic Background The Essential Knowledge statements provided in the AP Biology Curriculum Framework are scientific claims describing phenomenon occurring in
3) There are different types of extracellular signaling molecules. 4) most signaling molecules are secreted by exocytosis
 XIV) Signaling. A) The need for Signaling in multicellular organisms B) yeast need to signal to respond to various factors C) Extracellular signaling molecules bind to receptors 1) most bind to receptors
XIV) Signaling. A) The need for Signaling in multicellular organisms B) yeast need to signal to respond to various factors C) Extracellular signaling molecules bind to receptors 1) most bind to receptors
Chapter 3. Protein Structure and Function
 Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA
 CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
CHAPTER 6: RECOMBINANT DNA TECHNOLOGY YEAR III PHARM.D DR. V. CHITRA INTRODUCTION DNA : DNA is deoxyribose nucleic acid. It is made up of a base consisting of sugar, phosphate and one nitrogen base.the
Kermodul : Angeborene Immunität Innate Immunity. Uwe Sonnewald Email usonne@biologie.uni-erlangen.de
 BC Kermodul : Angeborene Immunität Innate Immunity Pathogens Invasion strategies Pathogen Host Interactions Basal Defence Uwe Sonnewald Email usonne@biologie.uni-erlangen.de 1 Pathogens and diseases Viruses
BC Kermodul : Angeborene Immunität Innate Immunity Pathogens Invasion strategies Pathogen Host Interactions Basal Defence Uwe Sonnewald Email usonne@biologie.uni-erlangen.de 1 Pathogens and diseases Viruses
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE Q5B
 INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
The Aerobic Fate of Pyruvate
 The Aerobic Fate of yruvate February 12, 2003 Bryant Miles I could tell that some of you were not impressed by the mere 2 ATs produced per glucose by glycolysis. The 2 AT s produced are only a small fraction
The Aerobic Fate of yruvate February 12, 2003 Bryant Miles I could tell that some of you were not impressed by the mere 2 ATs produced per glucose by glycolysis. The 2 AT s produced are only a small fraction
TEMA 10. REACCIONES INMUNITARIAS MEDIADAS POR CÉLULAS.
 TEMA 10. REACCIONES INMUNITARIAS MEDIADAS POR CÉLULAS. The nomenclature of cytokines partly reflects their first-described function and also the order of their discovery. There is no single unified nomenclature,
TEMA 10. REACCIONES INMUNITARIAS MEDIADAS POR CÉLULAS. The nomenclature of cytokines partly reflects their first-described function and also the order of their discovery. There is no single unified nomenclature,
Lecture 4: Peptides and Protein Primary Structure [PDF] Key Concepts. Objectives See also posted Peptide/pH/Ionization practice problems.
![Lecture 4: Peptides and Protein Primary Structure [PDF] Key Concepts. Objectives See also posted Peptide/pH/Ionization practice problems. Lecture 4: Peptides and Protein Primary Structure [PDF] Key Concepts. Objectives See also posted Peptide/pH/Ionization practice problems.](/thumbs/25/5655226.jpg) Lecture 4: Peptides and Protein Primary Structure [PDF] Reading: Berg, Tymoczko & Stryer, Chapter 2, pp. 34-37 Practice problems (peptide ionization) [PDF]; problems in textbook: chapter 2, pp. 63-64,
Lecture 4: Peptides and Protein Primary Structure [PDF] Reading: Berg, Tymoczko & Stryer, Chapter 2, pp. 34-37 Practice problems (peptide ionization) [PDF]; problems in textbook: chapter 2, pp. 63-64,
T Cell Maturation,Activation and Differentiation
 T Cell Maturation,Activation and Differentiation Positive Selection- In thymus, permits survival of only those T cells whose TCRs recognize self- MHC molecules (self-mhc restriction) Negative Selection-
T Cell Maturation,Activation and Differentiation Positive Selection- In thymus, permits survival of only those T cells whose TCRs recognize self- MHC molecules (self-mhc restriction) Negative Selection-
http://www.springer.com/3-540-23372-5
 http://www.springer.com/3-540-23372-5 Chromatin Remodeling Factors and DNA Replication Patrick Varga-Weisz Abstract Chromatin structures have to be precisely duplicated during DNA replication to maintain
http://www.springer.com/3-540-23372-5 Chromatin Remodeling Factors and DNA Replication Patrick Varga-Weisz Abstract Chromatin structures have to be precisely duplicated during DNA replication to maintain
Visualizing Cell Processes
 Visualizing Cell Processes A Series of Five Programs produced by BioMEDIA ASSOCIATES Content Guide for Program 3 Photosynthesis and Cellular Respiration Copyright 2001, BioMEDIA ASSOCIATES www.ebiomedia.com
Visualizing Cell Processes A Series of Five Programs produced by BioMEDIA ASSOCIATES Content Guide for Program 3 Photosynthesis and Cellular Respiration Copyright 2001, BioMEDIA ASSOCIATES www.ebiomedia.com
Amino Acids as Acids, Bases and Buffers:
 Amino Acids as Acids, Bases and Buffers: - Amino acids are weak acids - All have at least 2 titratable protons (shown below as fully protonated species) and therefore have 2 pka s o α-carboxyl (-COOH)
Amino Acids as Acids, Bases and Buffers: - Amino acids are weak acids - All have at least 2 titratable protons (shown below as fully protonated species) and therefore have 2 pka s o α-carboxyl (-COOH)
Chapter 20: Antimicrobial Drugs
 Chapter 20: Antimicrobial Drugs 1. Overview of Antimicrobial Drugs 2. Antibacterial Drugs 3. Antiviral Drugs 4. Drugs for Eukaryotic Pathogens 1. Overview of Antimicrobial Drugs Antibiotics An antibiotic
Chapter 20: Antimicrobial Drugs 1. Overview of Antimicrobial Drugs 2. Antibacterial Drugs 3. Antiviral Drugs 4. Drugs for Eukaryotic Pathogens 1. Overview of Antimicrobial Drugs Antibiotics An antibiotic
A. A peptide with 12 amino acids has the following amino acid composition: 2 Met, 1 Tyr, 1 Trp, 2 Glu, 1 Lys, 1 Arg, 1 Thr, 1 Asn, 1 Ile, 1 Cys
 Questions- Proteins & Enzymes A. A peptide with 12 amino acids has the following amino acid composition: 2 Met, 1 Tyr, 1 Trp, 2 Glu, 1 Lys, 1 Arg, 1 Thr, 1 Asn, 1 Ile, 1 Cys Reaction of the intact peptide
Questions- Proteins & Enzymes A. A peptide with 12 amino acids has the following amino acid composition: 2 Met, 1 Tyr, 1 Trp, 2 Glu, 1 Lys, 1 Arg, 1 Thr, 1 Asn, 1 Ile, 1 Cys Reaction of the intact peptide
The Cell Interior and Function
 The Cell Interior and Function 5 5.0 CHAPTER PREVIEW Investigate and understand the organization and function of the cell interior. Define the differences between eukaryotic and prokaryotic cell structure.
The Cell Interior and Function 5 5.0 CHAPTER PREVIEW Investigate and understand the organization and function of the cell interior. Define the differences between eukaryotic and prokaryotic cell structure.
Bacterial (Prokaryotic) Cell. Common features of all cells. Tour of the Cell. Eukaryotic Cell. Plasma Membrane defines inside from outside
 www.denniskunkel.com Tour of the Cell www.denniskunkel.com Today s Topics Properties of all cells Prokaryotes and Eukaryotes Functions of Major Cellular Organelles Information, Synthesis&Transport,, Vesicles
www.denniskunkel.com Tour of the Cell www.denniskunkel.com Today s Topics Properties of all cells Prokaryotes and Eukaryotes Functions of Major Cellular Organelles Information, Synthesis&Transport,, Vesicles
GENE CLONING AND RECOMBINANT DNA TECHNOLOGY
 GENE CLONING AND RECOMBINANT DNA TECHNOLOGY What is recombinant DNA? DNA from 2 different sources (often from 2 different species) are combined together in vitro. Recombinant DNA forms the basis of cloning.
GENE CLONING AND RECOMBINANT DNA TECHNOLOGY What is recombinant DNA? DNA from 2 different sources (often from 2 different species) are combined together in vitro. Recombinant DNA forms the basis of cloning.
Eukaryotes have organelles
 Energy-transducing Eukaryotes have organelles membrane systems An organelle is a discrete membrane bound cellular structure specialized functions. An organelle is to the cell what an organ is to the body
Energy-transducing Eukaryotes have organelles membrane systems An organelle is a discrete membrane bound cellular structure specialized functions. An organelle is to the cell what an organ is to the body
Name Class Date. Figure 13 1. 2. Which nucleotide in Figure 13 1 indicates the nucleic acid above is RNA? a. uracil c. cytosine b. guanine d.
 13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION. Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.
 Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Introduction to Proteins and Enzymes
 Introduction to Proteins and Enzymes Basics of protein structure and composition The life of a protein Enzymes Theory of enzyme function Not all enzymes are proteins / not all proteins are enzymes Enzyme
Introduction to Proteins and Enzymes Basics of protein structure and composition The life of a protein Enzymes Theory of enzyme function Not all enzymes are proteins / not all proteins are enzymes Enzyme
Nucleotides and Nucleic Acids
 Nucleotides and Nucleic Acids Brief History 1 1869 - Miescher Isolated nuclein from soiled bandages 1902 - Garrod Studied rare genetic disorder: Alkaptonuria; concluded that specific gene is associated
Nucleotides and Nucleic Acids Brief History 1 1869 - Miescher Isolated nuclein from soiled bandages 1902 - Garrod Studied rare genetic disorder: Alkaptonuria; concluded that specific gene is associated
An Overview of Cells and Cell Research
 An Overview of Cells and Cell Research 1 An Overview of Cells and Cell Research Chapter Outline Model Species and Cell types Cell components Tools of Cell Biology Model Species E. Coli: simplest organism
An Overview of Cells and Cell Research 1 An Overview of Cells and Cell Research Chapter Outline Model Species and Cell types Cell components Tools of Cell Biology Model Species E. Coli: simplest organism
European Medicines Agency
 European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
Concluding lesson. Student manual. What kind of protein are you? (Basic)
 Concluding lesson Student manual What kind of protein are you? (Basic) Part 1 The hereditary material of an organism is stored in a coded way on the DNA. This code consists of four different nucleotides:
Concluding lesson Student manual What kind of protein are you? (Basic) Part 1 The hereditary material of an organism is stored in a coded way on the DNA. This code consists of four different nucleotides:
