SHORT TECHNICAL REPORTS
|
|
|
- Brianna Gilbert
- 7 years ago
- Views:
Transcription
1 Identification of new fluorescent protein fragments for bimolecular fluorescence complementation analysis under physiological conditions Y. John Shyu 1, Han Liu 1, Xuehong Deng 1, and Chang-Deng Hu 1,2 1 Purdue University, West Lafayette and 2 Walther Cancer Institute, Indianapolis, IN, USA BioTechniques 40:61-66 (January 2006) doi / Protein-protein interactions play a pivotal role in coordinating many cellular processes. Determination of subcellular localization of interacting proteins and visualization of dynamic interactions in living cells are crucial to elucidate cellular functions of proteins. Using fluorescent proteins, we previously developed a bimolecular fluorescence complementation (BiFC) assay and a multicolor BiFC assay to visualize protein-protein interactions in living cells. However, the sensitivity of chromophore maturation of enhanced yellow fluorescent protein (YFP) to higher temperatures requires preincubation at lower temperatures prior to visualizing the BiFC signal. This could potentially limit their applications for the study of many signaling molecules. Here we report the identification of new fluorescent protein fragments derived from Venus and Cerulean for BiFC and multicolor BiFC assays under physiological culture conditions. More importantly, the newly identified combinations exhibit a 13-fold higher BiFC efficiency than originally identified fragments derived from YFP. Furthermore, the use of new combinations reduces the amount of plasmid required for transfection and shortens the incubation time, leading to a 2-fold increase in specific BiFC signals. These newly identified fluorescent protein fragments will facilitate the study of protein-protein interactions in living cells and whole animals under physiological conditions. INTRODUCTION Protein-protein interactions play a pivotal role in mediating signal transduction pathways and executing many cellular functions. Defining how each protein interacts with its partners under physiological conditions provides insight into the cellular functions of the protein. Several methods have been developed to study protein-protein interactions in living cells. These include mammalian two-hybrid assays (1), protein complementation assays (2 10), and fluorescent resonance energy transfer (FRET) (11 14). We previously took the protein complementation approach and used enhanced yellow fluorescent protein (EYFP) and enhanced cyan fluorescent protein (ECFP) to develop a bimolecular fluorescence complementation (BiFC) assay and a multicolor BiFC assay to directly visualize protein-protein interactions and to study the competition of multiple proteins in living cells (15,16). Due to stronger complementation signal and direct readout, the BiFC assay has been widely accepted and used for the study of protein-protein interaction in living cells (17,18, and references therein). However, one limitation of EYFP is the sensitivity of chromophore maturation to higher temperatures (19). This requires preincubation of cells at lower temperatures prior to visualizing the BiFC signal (15,16). Therefore, the availability of new fluorescent protein fragments that can be used under physiological conditions will undoubtedly impact the future applications of the BiFC assay. Here we report the identification of several new combinations for BiFC analysis under physiological conditions. MATERIALS AND METHODS BiFC constructs using fragments derived from three newly engineered fluorescent proteins, Venus (20), Citrine (21), and Cerulean (12) were made. Plasmids pbifc-bjunyn155 and pbifc-bfosyc155, as described previously (15), were used as templates. cdnas encoding the N- or C-terminal fragments of Venus, Citrine, and Cerulean were amplified by PCR and subcloned into pbifc-bjunyn155 and pbifc-bfosyc155 to replace those encoding YN155 and YC155 (Figure 1, A and B). To facilitate the detection of their expression, all N-terminal fragments fused to bjun have a FLAG tag at their N termini, and C-terminal fragments fused to bfos have a hemagglutin (HA) tag at their N termini. COS-1 cells were cultured in Dulbecco s modified Eagle medium (Hyclone, Logan, UT, USA) supplemented with 10% fetal bovine serum with penicillin and streptomycin in 12-well plates at 37 C. The indicated amounts of plasmid encoding bjun or bfos fusion proteins were transfected into COS-1 cells using FuGENE 6 (Roche Applied Science, Indianapolis, IN, USA). For ratio analysis, plasmids encoding ECFP or EYFP were cotransfected as an internal control to measure the BiFC efficiency of fragments derived from YFP or CFP variants, respectively. Images of transfected cells 24 h after transfection were captured using a charge-coupled device (CCD) camera mounted on a TE2000-U inverted fluorescence microscope (Nikon, Melville, NY, USA) with JP4 filters (Chroma, Rockingham, VT, USA). The intensity of more than 100 cells was individually quantified using an automated intensity recognition feature of Metamorph II (Universal Imaging, Downingtown, PA, USA). YFP/CFP ratio analysis was similarly performed as reported previously (15). The median of YFP/CFP ratios was used for comparison of the BiFC efficiency among YFP variants and determination of BiFC specificity, since the distribution of the raw data was highly skewed (15). Since 7.8% of the BiFC signal derived from N173 of Venus paired with C155 of ECFP was Vol. 40, No. 1 (2006) BioTechniques 61
2 read in the CFP channel, we corrected this signal crosstalk by multiplying original CFP images with a correction coefficient. Briefly, cells transfected with plasmids encoding N173 of Venus fused to bjun and C155 of ECFP fused to bfos were used as a control to determine the correction coefficient. Figure 1. Identification of new fluorescent protein fragments for bimolecular fluorescence complementation (BiFC) analysis under physiological conditions. (A) View of enhanced green fluorescent protein (EGFP) and its variants. The indicated residues are substitutions from GFP, and the positions of truncation in fluorescent proteins are indicated with a scissors symbol. (B) View of BiFC principle. N- and C-terminal fragments of fluorescent proteins were fused to bjun and bfos, respectively. The interaction between bjun and bfos brings N- and C-terminal fragments in proximity to reconstitute an intact fluorescent protein (FP). (C) Fluorescent images of COS-1 cells transfected with μg plasmid encoding bjun or bfos fused to the fluorescent protein fragments indicated in each panel. The number indicates the maxima of excitation and emission for each BiFC complex. (D) Quantification of BiFC efficiency among yellow fluorescent protein (YFP) mutants. The BiFC efficiency was determined as described in the main text. The fold increase of median ratios plus standard deviation was determined when compared with N155-C155 of enhanced YFP (EYFP). (E) Immunoblot analysis of fusion proteins. Transfected cells expressing the indicated fusion proteins from Figure 1C were harvested for the detection of fusion proteins using anti-flag and anti-ha antibodies. ECFP, enhanced cyan fluorescent protein. Fluorescent images were captured with both YFP and CFP filters, and the correction coefficient was calculated as described for FRET (22). The original CFP images in the multicolor BiFC experiments were multiplied by the correction coefficient using the FRET function provided in Metamorph II, and a representative of the corrected CFP images was presented here (see Figure 3B). To determine the excitation and emission maxima of each new BiFC complex, COS-1 cells were transfected with 0.25 μg of each plasmid encoding the fusion proteins and incubated at 37 C for 24 h. Harvested cells were resuspended in 1 phosphate-buffered saline (PBS) buffer (Hyclone), and FluoroMax 2 was used in conjunction with the Discovery Scan feature of DataMax for Windows (Horiba Jobin Yvon, Edison, NJ, USA) to determine the excitation and emission maxima of each BiFC complex. To determine BiFC specificity, cells were cotransfected with various amounts of plasmid encoding each fusion protein in combination with 50 ng plasmid encoding ECFP. The median of YFP/CFP ratios was calculated from more than 100 cells. The specificity of BiFC represents the fluorescent signals derived from the reconstituted fluorescent proteins driven by specific bfos-bjun interactions. For the purpose of comparison among different groups transfected with different amounts of plasmid, the BiFC specificity was calculated as fold increase by dividing the median ratio derived from cells coexpressing bfos fusions paired with bjun fusions by the median ratio derived from cells coexpressing bfos(δzip) fusions paired with bjun fusions. Note that bfos(δzip) is a dimerization-deficient mutant and was paired with bjun as a background control (15,16,18). To determine the expression level of various fusion proteins, transfected cells were harvested after images were captured and lysed in 100 μl 1 Laemmli sample loading buffer. The proteins were resolved by 10% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE), followed by the transfer to a nitrocellulose filter. The expression of each fusion protein 62 BioTechniques Vol. 40, No. 1 (2006)
3 was determined using antibodies against the FLAG or HA epitopes (Sigma, St. Louis, MO, USA). RESULTS AND DISCUSSION To identify new fluorescent protein fragments for BiFC analysis under physiological conditions, we examined Figure 2. Comparison of the specificity of bimolecular fluorescence complementation (BiFC) signal using enhanced yellow fluorescent protein (EYFP) and Venus fragments. Cells were transfected with the indicated amount of each plasmid, and the specificity of BiFC was determined as described in the main text. (A) Effect of different amounts of plasmids on the specificity of BiFC using EYFP fragments. (B) Effect of different amounts of plasmids on the specificity of BiFC using Venus fragments. (C) Immunoblot analysis of fusion proteins. Transfected cells expressing the indicated fusion proteins from panels A and B were harvested for the detection of fusion proteins using anti-flag and anti-ha antibodies. BiFC efficiency using Citrine and Venus (20,21), two new YFP mutants engineered to be less sensitive to the environment. Based on our previous findings that fragments derived from EYFP truncated at either position D155 or D173 can support BiFC when fused to the basic region leucine zipper domain of c-jun (bjun) and the basic region leucine zipper domain of c-fos (bfos) (Figure 1, A and B) (15,16), we similarly truncated Citrine and Venus at these two positions and fused the N- and C-terminal fragments to bjun and bfos, respectively (Figure 1B). Coexpression of these fusion proteins in COS-1 cells resulted in fluorescence complementation in all combinations (Figure 1C). Quantification of the BiFC efficiency revealed that all three combinations of Venus showed a 12- to 15-fold increase and Citrine showed a 4- to 5-fold increase when compared with EYFP (Figure 1D). All combinations of fusion proteins were comparably expressed (Figure 1E). These experiments were carried out at 37 C, and images were captured 24 h posttransfection. Preincubation of cells at 30 C for 4 h prior to image capturing significantly increased the BiFC signal for EYFP fragments, whereas the BiFC signals derived from Citrine and Venus fragments were not significantly affected (data not shown). This confirms that the chromophore maturation of Citrine and Venus is indeed less sensitive to higher temperatures. Nevertheless, the average fluorescence intensity derived from EYFP fragments was still only 64% and 40% of that derived from Citrine and Venus, respectively. To determine whether the newly identified fluorescent protein fragments affect the specificity of the BiFC signal, we used bfos(δzip), carrying a small deletion in the Zip region to eliminate specific dimerization with bjun (15), as a background control to compare the BiFC efficiency between wild-type bfos and this mutant bfos when they were paired with bjun fusion proteins (15,16,18). As shown in Figure 2A, when fragments derived from EYFP were used, the BiFC signal derived from wild-type bfos was 3- to 5-fold higher than those derived from bfosδzip within the range of amount of plasmids examined. Lower amount of plasmids for transfection showed relatively higher specificity. Similar tendency was observed when fragments derived from Venus were used (Figure 2B). However, the use of Venus BiFC plasmids exhibited higher specificity when compared with EYFP. In particular, the use of μg plasmids encoding fusion proteins with Venus fragments showed 2-fold higher specificity than the use of EYFP fragments. Comparable expression of the fusion proteins was confirmed by immunoblot analysis (Figure 2C). When the fulllength c-fos and c-jun were examined using the Venus fragments, we observed very bright BiFC signals 12 h after transfection, whereas the use of EYFP fragments required 2 days incubation at 37 C plus at least 2 h preincubation at 30 C (data not shown). These results demonstrate that the use of Venus fragments for BiFC analysis not only eliminates the preincubation at lower temperatures, but also shows higher specificity and requires lower amount of plasmids for transfection, plus a shorter incubation time. Protein-protein interaction is the foundation of signaling networks in cells. Each protein has a few, or even a large number of, alternative interaction partners. These alternative interactions are often selectively induced in response to specific signals. Vol. 40, No. 1 (2006) BioTechniques 63
4 Alternatively, several interaction partners that coexist in the same subcellular compartment may compete for binding to shared interaction partners. Therefore, visualization of multiple interactions in the same cell provides a unique way to study protein interaction networks. Since green fluorescent protein (GFP) variants have virtually identical structure, and the chromophore is formed by three residues at positions 65, 66, and 67 (19), we previously examined all possible combinations among fragments derived from four GFP spectral variants and identified several combinations that showed better spectral resolution and supported the fluorescence complementation (16). The best combination Figure 3. Identification of new combinations for multicolor bimolecular fluorescence complementation (BiFC) analysis. (A) Fluorescent images of COS-1 cells transfected with μg plasmids encoding bjun or bfos fused to the fluorescent protein fragments indicated. The number indicates the maxima of excitation and emission for each BiFC complex. (B) Visualization of differential subnuclear localization of bjun-bfos and Jun-bFos. Plasmids encoding the indicated fusion proteins were cotransfected into COS-1 cells, and images were taken using cyan fluorescent protein (CFP) and yellow fluorescent protein (YFP) filters. Note that the CFP image was corrected as described in the main text. identified previously for multicolor BiFC analysis was the C-terminal fragment of ECFP (ECFP C155) paired with the N-terminal fragments of EYFP (EYFP N173) and ECFP (ECFP N173) (16,23). Recently, it has been reported that Cerulean, an improved ECFP, shows a 2.5-fold increase in fluorescence intensity compared with ECFP (12). This prompted us to explore the possibility that a better multicolor BiFC combination may be identified in conjunction with the newly identified fluorescent protein fragments described above. The C-terminal sequences of EYFP and Citrine are identical, as are those of ECFP and Cerulean (Figure 1A). We compared the complementation efficiency of the C-terminal fragments comprised of residues (C155) derived from EYFP, Venus, and ECFP with the N-terminal fragments comprised of residues (N173) of Cerulean, Citrine, and Venus. C155 of ECFP showed higher complementation efficiency and lower signal bleed-through with N173 derived from Cerulean and Venus than any other combinations (Figure 3A and data not shown). Interestingly, N173 of Citrine complemented with C155 of EYFP and Venus, but did not complement with C155 of ECFP. The only difference among fragments C155 of EYFP, Venus, and ECFP is T203Y (Figure 1A), suggesting the importance of Y203 in fragment complementation of fluorescent proteins. We previously observed that the bzip domain dimers of all bzip proteins were predominantly localized in the nucleoli, whereas the complexes formed by a bzip domain and a fulllength bzip protein were predominantly localized to the nucleoplasm (15). This distinct subcellular localization was visualized simultaneously using our multicolor BiFC assay (16). To validate the usefulness of the newly identified combination for multicolor BiFC assay, we examined the subnuclear localization of bfosbjun heterodimers and bfos-jun heterodimers in the same cell (16). When this combination was used, bright fluorescent signals derived from bfos-bjun heterodimers were predominantly localized in the nucleoli (Figure 3B). In contrast, bfos-jun heterodimers were distinctively localized in the nucleoplasm. Again, transfected cells were only incubated at 37 C for 24 h. In summary, we have identified Venus as the most useful yellow fluorescent protein for BiFC analysis of protein-protein interactions under physiological culture conditions. Furthermore, a new combination between C155 of ECFP and N173 of Cerulean and Venus has proven to be the best combination for multicolor BiFC analysis. The newly identified fragments offer two significant advantages over the previously reported BiFC constructs using EYFP and ECFP. First, the new combinations allow for lower amounts of plasmid for transfection and shorter incubation time, two recommended strategies to reduce nonspecific signals (18). Second, the new combinations eliminate the need for preincubaton of cells at lower temperatures. Because a decrease in temperature itself is a stressful condition, the new combinations should be particularly useful for the study of signaling molecules involved in the stress response. These improvements lead to the increase of both the BiFC signal and the specificity. It is also very likely that the new combinations will be suitable for visualization of protein-protein interactions in whole animals. However, compared with other protein fragment complementation assays using enzymes, such as luciferase (9), β-galactosidase (2), and β-lactamase (4,5), the BiFC assay does not provide signal amplification. In addition, it remains to be determined whether the BiFC assay can be used to visualize transient and inducible interactions (e.g., within minutes) or dynamic interaction in cells, since the chromophore maturation of fluorescent 64 BioTechniques Vol. 40, No. 1 (2006)
5 proteins requires at least 30 min (19), and the BiFC complex is irreversible in vitro (15). One interesting finding in this and our previous BiFC analysis of GFP spectral variants is the unpredictability of fluorescence complementation. We initially thought that the truncation at positions 155 and 173 would be applicable to all GFP variants. However, neither of these positions worked with EGFP (16). Interestingly, N-terminal fragments truncated at either site complemented with the C-terminal fragments derived from EYFP. The only difference between the C-terminal fragments of EYFP and EGFP is the residue at position 203 (Figure 1A). In the case of EYFP, Y203 was used in place of T203. This substitution allows a π π interaction with the chromophore phenol ring (24), suggesting that this interaction not only changes the spectra of the protein but also contributes to the reconstitution of the β-barrel structure. Similar to this Y203, our present analysis suggests that the position 69 also is critical to the fluorescence complementation. Fragments derived from Citrine and Venus all complemented with each other and showed higher BiFC efficiency than fragments derived from EYFP when cells were cultured at 37 C. This is due to the introduction of the substituted residues Q69M for Citrine and F46L for Venus (20,21). These substitutions are thought to facilitate chromophore maturation by increasing protein folding at 37 C. In addition to these two critical substitutions in Citrine and Venus, additional substitutions (F64L, M153T, V163A, and S175G) in Venus are also known to improve protein folding at 37 C (19,20), and these changes might contribute to higher BiFC efficiency for Venus over other YFP variants. Surprisingly, N173 of Citrine did not complement with C155 of ECFP, but N173 of EYFP did (Figure 3A). Since Q69M is the only difference between EYFP and Citrine, this observation suggests that the residue at position 69 not only affects the chromophore maturation but also contributes to the fragment complementation. However, N173 of Citrine complemented well with C155 of EYFP, Venus, and Citrine. Again, this is probably due to the presence of Y203, which might compensate for the presence of M69 in N173 of Citrine. Since the chromophore of fluorescent proteins must be protected from the environment (25), it is important that the reassembly of fluorescent protein fragments is precise. It is therefore likely that complementation of fluorescent protein fragments is determined by a combination of residues from both N- and C-terminal fragments. Consistent with this notion, truncation at neither position 155 nor position 173 of EGFP supports the BiFC in our case (16), whereas truncation at position 157 of EGFP allows the resulting fragments to complement (7). Structural analysis of these reconstituted BiFC complexes will be necessary to provide insight into the mechanisms by which BiFC works. Regardless of the mechanisms, the newly identified fluorescent protein fragments reported here for BiFC analysis will be useful for the study of protein-protein interactions in living cells. ACKNOWLEDGMENTS We would like to thank Dr. Tom Kerppola for his critical reading and comments on the manuscript, as well as Drs. Atsushi Miyawaki and David Piston for kindly providing cdnas of Venus and Cerulean, and the members of the Hu laboratory for helpful discussions. This work was partly supported by the Purdue Cancer Center (NCI-P30CA23168), the Indina Elks Charities, Inc., the Walther Cancer Institute, and a grant from National Science Foundation ( MCB) to C.D.H. COMPETING INTERESTS STATEMENT The authors declare no competing interests. REFERENCES 1. Luo, Y., A. Batalao, H. Zhou, and L. Zhu Mammalian two-hybrid system: a complementary approach to the yeast two-hybrid system. BioTechniques 22: Rossi, F.M., B.T. Blakely, C.A. Charlton, and H.M. Blau Monitoring proteinprotein interactions in live mammalian cells by beta-galactosidase complementation. Methods Enzymol. 328: Johnsson, N. and A. Varshavsky Split ubiquitin as a sensor of protein interactions in vivo. Proc. Natl. Acad. Sci. USA 91: Wehrman, T., B. Kleaveland, J.H. Her, R.F. Balint, and H.M. Blau Protein-protein interactions monitored in mammalian cells via complementation of beta-lactamase enzyme fragments. Proc. Natl. Acad. Sci. USA 99: Galarneau, A., M. Primeau, L.E. Trudeau, and S.W. Michnick Beta-lactamase protein fragment complementation assays as in vivo and in vitro sensors of protein protein interactions. Nat. Biotechnol. 20: Paulmurugan, R. and S.S. Gambhir Firefly luciferase enzyme fragment complementation for imaging in cells and living animals. Anal. Chem. 77: Ghosh, I., A.D. Hamilton, and L. Regan Antiparallel leucine zipper-directed protein reassembly: applications to the green fluorescent protein. J. Am. Chem. Soc. 122: Michnick, S.W., I. Remy, F.X. Campbell- Valois, A. Vallee-Belisle, and J.N. Pelletier Detection of protein-protein interactions by protein fragment complementation strategies. Methods Enzymol. 328: Luker, K.E. and D. Piwnica-Worms Optimizing luciferase protein fragment complementation for bioluminescent imaging of protein-protein interactions in live cells and animals. Methods Enzymol. 385: Rackham, O. and C.M. Brown Visualization of RNA-protein interactions in living cells: FMRP and IMP1 interact on mrnas. EMBO J. 23: Miyawaki, A., J. Llopis, R. Heim, J.M. McCaffery, J.A. Adams, M. Ikura, and R.Y. Tsien Fluorescent indicators for Ca 2+ based on green fluorescent proteins and calmodulin. Nature 388: Rizzo, M.A., G.H. Springer, B. Granada, and D.W. Piston An improved cyan fluorescent protein variant useful for FRET. Nat. Biotechnol. 22: Pollok, B.A. and R. Heim Using GFP in FRET-based applications. Trends Cell Biol. 9: Periasamy, A. and R.N. Day Visualizing protein interactions in living cells using digitized GFP imaging and FRET microscopy. Methods Cell Biol. 58: Hu, C.D., Y. Chinenov, and T.K. Kerppola Visualization of interactions among bzip and Rel family proteins in living cells using bimolecular fluorescence complementation. Mol. Cell 9: Hu, C.D. and T.K. Kerppola Simultaneous visualization of multiple protein interactions in living cells using multicolor fluorescence complementation analysis. Nat. Biotechnol. 21: Wilson, C.G., T.J. Magliery, and L. Regan Detecting protein-protein interactions Vol. 40, No. 1 (2006) BioTechniques 65
6 with GFP-fragment reassembly. Nat. Methods 1: Hu, C.D., A. Grinberg, and T. Kerppola Visualization of protein interaction in living cells using bimolecular fluorescence complementation (BiFC) analysis, p In J.S. Bonifacino, M. Dasso, J.B. Harford, J. Lippincott-Schwartz, and K.M. Yamada (Eds.), Current Protocol in Cell Biology. John Wiley & Sons, New York. 19.Tsien, R.Y The green fluorescent protein. Annu. Rev. Biochem. 67: Nagai, T., K. Ibata, E.S. Park, M. Kubota, K. Mikoshiba, and A. Miyawaki A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat. Biotechnol. 20: Griesbeck, O., G.S. Baird, R.E. Campbell, D.A. Zacharias, and R.Y. Tsien Reducing the environmental sensitivity of yellow fluorescent protein. Mechanism and applications. J. Biol. Chem. 276: Gordon, G.W., G. Berry, X.H. Liang, B. Levine, and B. Herman Quantitative fluorescence resonance energy transfer measurements using fluorescence microscopy. Biophys. J. 74: Grinberg, A.V., C.D. Hu, and T.K. Kerppola Visualization of Myc/Max/ Mad family dimers and the competition for dimerization in living cells. Mol. Cell. Biol. 24: Wachter, R.M., M.A. Elsliger, K. Kallio, G.T. Hanson, and S.J. Remington Structural basis of spectral shifts in the yellow-emission variants of green fluorescent protein. Structure 6: Ormo, M., A.B. Cubitt, K. Kallio, L.A. Gross, R.Y. Tsien, and S.J. Remington Crystal structure of the Aequorea victoria green fluorescent protein. Science 273: Received 11 July 2005; accepted 18 August Address correspondence to Chang-Deng Hu, Department of Medicinal Chemistry & Molecular Pharmacology and the Purdue Cancer Center, Purdue University, West Lafayette, IN 47907, USA. cdhu@pharmacy.purdue.edu To purchase reprints of this article, contact Reprints@BioTechniques.com 66 BioTechniques Vol. 40, No. 1 (2006)
Visualization of Myc/Max/Mad Family Dimers and the Competition for Dimerization in Living Cells
 MOLECULAR AND CELLULAR BIOLOGY, May 2004, p. 4294 4308 Vol. 24, No. 10 0270-7306/04/$08.00 0 DOI: 10.1128/MCB.24.10.4294 4308.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved.
MOLECULAR AND CELLULAR BIOLOGY, May 2004, p. 4294 4308 Vol. 24, No. 10 0270-7306/04/$08.00 0 DOI: 10.1128/MCB.24.10.4294 4308.2004 Copyright 2004, American Society for Microbiology. All Rights Reserved.
FRET Basics and Applications an EAMNET teaching module
 FRET Basics and Applications an EAMNET teaching module Timo Zimmermann + Stefan Terjung Advanced Light Microscopy Facility European Molecular Biology Laboratory, Heidelberg http://www.embl.de/almf/ http://www.embl.de/eamnet/
FRET Basics and Applications an EAMNET teaching module Timo Zimmermann + Stefan Terjung Advanced Light Microscopy Facility European Molecular Biology Laboratory, Heidelberg http://www.embl.de/almf/ http://www.embl.de/eamnet/
Fluorescent dyes for use with the
 Detection of Multiple Reporter Dyes in Real-time, On-line PCR Analysis with the LightCycler System Gregor Sagner, Cornelia Goldstein, and Rob van Miltenburg Roche Molecular Biochemicals, Penzberg, Germany
Detection of Multiple Reporter Dyes in Real-time, On-line PCR Analysis with the LightCycler System Gregor Sagner, Cornelia Goldstein, and Rob van Miltenburg Roche Molecular Biochemicals, Penzberg, Germany
Reversible Dimerization of Aequorea victoria Fluorescent Proteins Increases the Dynamic Range of FRET-Based Indicators
 ARTICLE Reversible Dimerization of Aequorea victoria Fluorescent Proteins Increases the Dynamic Range of FRET-Based Indicators Ippei Kotera,, Takuya Iwasaki,, Hiromi Imamura, Hiroyuki Noji, and Takeharu
ARTICLE Reversible Dimerization of Aequorea victoria Fluorescent Proteins Increases the Dynamic Range of FRET-Based Indicators Ippei Kotera,, Takuya Iwasaki,, Hiromi Imamura, Hiroyuki Noji, and Takeharu
The cell lines used in this study were obtained from the American Type Culture
 Supplementary materials and methods Cell culture and drug treatments The cell lines used in this study were obtained from the American Type Culture Collection (ATCC) and grown as previously described.
Supplementary materials and methods Cell culture and drug treatments The cell lines used in this study were obtained from the American Type Culture Collection (ATCC) and grown as previously described.
Validation & Assay Performance Summary
 Validation & Assay Performance Summary GeneBLAzer RAR alpha DA Assay Kit GeneBLAzer RAR alpha-uas-bla GripTite Cells Cat. no. K1387, K1685 Target Description The retinoic acid receptor alpha (RAR alpha)
Validation & Assay Performance Summary GeneBLAzer RAR alpha DA Assay Kit GeneBLAzer RAR alpha-uas-bla GripTite Cells Cat. no. K1387, K1685 Target Description The retinoic acid receptor alpha (RAR alpha)
BaculoDirect Baculovirus Expression System Free your hands with the BaculoDirect Baculovirus Expression System
 BaculoDirect Baculovirus Expression System Free your hands with the BaculoDirect Baculovirus Expression System The BaculoDirect Baculovirus Expression System gives you: Unique speed and simplicity High-throughput
BaculoDirect Baculovirus Expression System Free your hands with the BaculoDirect Baculovirus Expression System The BaculoDirect Baculovirus Expression System gives you: Unique speed and simplicity High-throughput
Supplemental Fig. S1. The schematic diagrams of the expression constructs used in this study.
 1 Supplemental data Supplemental Fig. S1. The schematic diagrams of the expression constructs used in this study. Supplemental Fig. S2. Ingenuity Pathway Analysis (IPA) of the 56 putative caspase substrates
1 Supplemental data Supplemental Fig. S1. The schematic diagrams of the expression constructs used in this study. Supplemental Fig. S2. Ingenuity Pathway Analysis (IPA) of the 56 putative caspase substrates
Angelo Peschiaroli, N. Valerio Dorrello, Daniele Guardavaccaro, Monica Venere, Thanos Halazonetis, Nicholas E. Sherman, and Michele Pagano
 Molecular Cell, Volume 23 Supplemental Data SCF βtrcp -Mediated Degradation of Claspin Regulates Recovery from the DNA Replication Checkpoint Response Angelo Peschiaroli, N. Valerio Dorrello, Daniele Guardavaccaro,
Molecular Cell, Volume 23 Supplemental Data SCF βtrcp -Mediated Degradation of Claspin Regulates Recovery from the DNA Replication Checkpoint Response Angelo Peschiaroli, N. Valerio Dorrello, Daniele Guardavaccaro,
SUPPLEMENTARY DATA 1
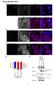 SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
SUPPLEMENTARY DATA 1 Supplementary Figure S1. Overexpression of untagged Ago2 inhibits the nuclear transport of the dotted foci of GFP-signal of myc-gfp-tnrc6a-nes-mut. (A C) HeLa cells expressing myc-gfp-tnrc6a-nes-mut
LightSwitch Dual Assay System DA010 (100 assays)
 LightSwitch Dual Assay System DA010 (100 assays)! Description The LightSwitch Dual Assay System is used to normalize for variation between transfection replicates in cell lines that are particularly difficult
LightSwitch Dual Assay System DA010 (100 assays)! Description The LightSwitch Dual Assay System is used to normalize for variation between transfection replicates in cell lines that are particularly difficult
Chapter 11. Rapid Screening of Gli2/3 Mutants Using the Flp-In System. Pawel Niewiadomski and Rajat Rohatgi. Abstract.
 Chapter 11 Rapid Screening of Gli2/3 Mutants Using the Flp-In System Abstract Gli2 and Gli3 respond to the Hedgehog (Hh) signal in mammals by undergoing posttranslational modifications and moving to the
Chapter 11 Rapid Screening of Gli2/3 Mutants Using the Flp-In System Abstract Gli2 and Gli3 respond to the Hedgehog (Hh) signal in mammals by undergoing posttranslational modifications and moving to the
Product: Expression Arrest TM egfp control shrna vector
 Product: Expression Arrest TM egfp control vector Catalog #: RHS1702 Product Description The laboratory of Dr. Greg Hannon at Cold Spring Harbor Laboratory (CSHL) has created an RNAi Clone Library comprised
Product: Expression Arrest TM egfp control vector Catalog #: RHS1702 Product Description The laboratory of Dr. Greg Hannon at Cold Spring Harbor Laboratory (CSHL) has created an RNAi Clone Library comprised
Fluorescein Isothiocyanate (FITC)- conjugated Antibodies
 USER GUIDE Fluorescein Isothiocyanate (FITC)- conjugated Antibodies Catalog Numbers R933-25, R953-25, R963-25 Document Part Number 25-0376 Publication Number MAN0000194 Revision 2.0 For Research Use Only.
USER GUIDE Fluorescein Isothiocyanate (FITC)- conjugated Antibodies Catalog Numbers R933-25, R953-25, R963-25 Document Part Number 25-0376 Publication Number MAN0000194 Revision 2.0 For Research Use Only.
Sanger Sequencing and Quality Assurance. Zbigniew Rudzki Department of Pathology University of Melbourne
 Sanger Sequencing and Quality Assurance Zbigniew Rudzki Department of Pathology University of Melbourne Sanger DNA sequencing The era of DNA sequencing essentially started with the publication of the enzymatic
Sanger Sequencing and Quality Assurance Zbigniew Rudzki Department of Pathology University of Melbourne Sanger DNA sequencing The era of DNA sequencing essentially started with the publication of the enzymatic
Innate Immunity. Insects rely solely on an innate immune system for defense against infection
 Innate Immunity Insects rely solely on an innate immune system for defense against infection The importance of mammalian innate immunity has only recently become appreciated The innate immune response
Innate Immunity Insects rely solely on an innate immune system for defense against infection The importance of mammalian innate immunity has only recently become appreciated The innate immune response
Annexin V-EGFP Apoptosis Detection Kit
 ab14153 Annexin V-EGFP Apoptosis Detection Kit Instructions for Use For the rapid, sensitive and accurate measurement of apoptosis in various samples This product is for research use only and is not intended
ab14153 Annexin V-EGFP Apoptosis Detection Kit Instructions for Use For the rapid, sensitive and accurate measurement of apoptosis in various samples This product is for research use only and is not intended
MEF Nucleofector Kit 1 and 2
 page 1 of 7 MEF Nucleofector Kit 1 and 2 for Mouse Embryonic Fibroblasts (MEF) MEF display significant phenotypic variations which depend on the strain, the genetic background of the they are isolated
page 1 of 7 MEF Nucleofector Kit 1 and 2 for Mouse Embryonic Fibroblasts (MEF) MEF display significant phenotypic variations which depend on the strain, the genetic background of the they are isolated
ArC Amine Reactive Compensation Bead Kit
 ArC Amine Reactive Compensation Bead Kit Catalog no. A1346 Table 1. Contents and storage information. Material Amount Composition Storage Stability ArC reactive beads (Component A) ArC negative beads (Component
ArC Amine Reactive Compensation Bead Kit Catalog no. A1346 Table 1. Contents and storage information. Material Amount Composition Storage Stability ArC reactive beads (Component A) ArC negative beads (Component
Molecular Spectroscopy
 Molecular Spectroscopy UV-Vis Spectroscopy Absorption Characteristics of Some Common Chromophores UV-Vis Spectroscopy Absorption Characteristics of Aromatic Compounds UV-Vis Spectroscopy Effect of extended
Molecular Spectroscopy UV-Vis Spectroscopy Absorption Characteristics of Some Common Chromophores UV-Vis Spectroscopy Absorption Characteristics of Aromatic Compounds UV-Vis Spectroscopy Effect of extended
Real-Time PCR Vs. Traditional PCR
 Real-Time PCR Vs. Traditional PCR Description This tutorial will discuss the evolution of traditional PCR methods towards the use of Real-Time chemistry and instrumentation for accurate quantitation. Objectives
Real-Time PCR Vs. Traditional PCR Description This tutorial will discuss the evolution of traditional PCR methods towards the use of Real-Time chemistry and instrumentation for accurate quantitation. Objectives
Understanding the immune response to bacterial infections
 Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
Understanding the immune response to bacterial infections A Ph.D. (SCIENCE) DISSERTATION SUBMITTED TO JADAVPUR UNIVERSITY SUSHIL KUMAR PATHAK DEPARTMENT OF CHEMISTRY BOSE INSTITUTE 2008 CONTENTS Page SUMMARY
MEF Starter Nucleofector Kit
 page 1 of 7 MEF Starter Nucleofector Kit for Mouse Embryonic Fibroblasts (MEF) MEF display significant phenotypic variations which depend on the strain, the genetic background of the mice they are isolated
page 1 of 7 MEF Starter Nucleofector Kit for Mouse Embryonic Fibroblasts (MEF) MEF display significant phenotypic variations which depend on the strain, the genetic background of the mice they are isolated
Optimal Conditions for F(ab ) 2 Antibody Fragment Production from Mouse IgG2a
 Optimal Conditions for F(ab ) 2 Antibody Fragment Production from Mouse IgG2a Ryan S. Stowers, 1 Jacqueline A. Callihan, 2 James D. Bryers 2 1 Department of Bioengineering, Clemson University, Clemson,
Optimal Conditions for F(ab ) 2 Antibody Fragment Production from Mouse IgG2a Ryan S. Stowers, 1 Jacqueline A. Callihan, 2 James D. Bryers 2 1 Department of Bioengineering, Clemson University, Clemson,
Genetics Lecture Notes 7.03 2005. Lectures 1 2
 Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Green Fluorescent Protein (GFP): Genetic Transformation, Synthesis and Purification of the Recombinant Protein
 Green Fluorescent Protein (GFP): Genetic Transformation, Synthesis and Purification of the Recombinant Protein INTRODUCTION Green Fluorescent Protein (GFP) is a novel protein produced by the bioluminescent
Green Fluorescent Protein (GFP): Genetic Transformation, Synthesis and Purification of the Recombinant Protein INTRODUCTION Green Fluorescent Protein (GFP) is a novel protein produced by the bioluminescent
Confocal Fluorescence Microscopy
 Chapter 1 Confocal Fluorescence Microscopy 1.1 The principle Confocal fluorescence microscopy is a microscopic technique that provides true three-dimensional (3D) optical resolution. In microscopy, 3D
Chapter 1 Confocal Fluorescence Microscopy 1.1 The principle Confocal fluorescence microscopy is a microscopic technique that provides true three-dimensional (3D) optical resolution. In microscopy, 3D
Cloning GFP into Mammalian cells
 Protocol for Cloning GFP into Mammalian cells Studiepraktik 2013 Molecular Biology and Molecular Medicine Aarhus University Produced by the instructors: Tobias Holm Bønnelykke, Rikke Mouridsen, Steffan
Protocol for Cloning GFP into Mammalian cells Studiepraktik 2013 Molecular Biology and Molecular Medicine Aarhus University Produced by the instructors: Tobias Holm Bønnelykke, Rikke Mouridsen, Steffan
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
www.sciencesignaling.org/cgi/content/full/7/339/ra80/dc1 Supplementary Materials for Manipulation of receptor oligomerization as a strategy to inhibit signaling by TNF superfamily members Julia T. Warren,
Modeling and Simulation of Gene Regulatory Networks
 Modeling and Simulation of Gene Regulatory Networks Hidde de Jong INRIA Grenoble - Rhône-Alpes Hidde.de-Jong@inria.fr http://ibis.inrialpes.fr INRIA Grenoble - Rhône-Alpes and IBIS IBIS: systems biology
Modeling and Simulation of Gene Regulatory Networks Hidde de Jong INRIA Grenoble - Rhône-Alpes Hidde.de-Jong@inria.fr http://ibis.inrialpes.fr INRIA Grenoble - Rhône-Alpes and IBIS IBIS: systems biology
博 士 論 文 ( 要 約 ) A study on enzymatic synthesis of. stable cyclized peptides which. inhibit protein-protein interactions
 博 士 論 文 ( 要 約 ) 論 文 題 目 A study on enzymatic synthesis of stable cyclized peptides which inhibit protein-protein interactions ( 蛋 白 質 間 相 互 作 用 を 阻 害 する 安 定 な 環 状 化 ペプチドの 酵 素 合 成 に 関 する 研 究 ) 氏 名 張 静 1
博 士 論 文 ( 要 約 ) 論 文 題 目 A study on enzymatic synthesis of stable cyclized peptides which inhibit protein-protein interactions ( 蛋 白 質 間 相 互 作 用 を 阻 害 する 安 定 な 環 状 化 ペプチドの 酵 素 合 成 に 関 する 研 究 ) 氏 名 張 静 1
DNA Core Facility: DNA Sequencing Guide
 DNA Core Facility: DNA Sequencing Guide University of Missouri-Columbia 216 Life Sciences Center Columbia, MO 65211 http://biotech.missouri.edu/dnacore/ Table of Contents 1. Evaluating Sequencing Data..
DNA Core Facility: DNA Sequencing Guide University of Missouri-Columbia 216 Life Sciences Center Columbia, MO 65211 http://biotech.missouri.edu/dnacore/ Table of Contents 1. Evaluating Sequencing Data..
Essentials of Real Time PCR. About Sequence Detection Chemistries
 Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
FLUORESCENT PROTEINS - XFPs
 FLUORESCENT PROTEINS - XFPs Marcel Walser; PhD student ETH Zürich Taskforce Kommunikation Presentations Content - Originating organisms - GFP characteristics - Applications - Short protein structure review
FLUORESCENT PROTEINS - XFPs Marcel Walser; PhD student ETH Zürich Taskforce Kommunikation Presentations Content - Originating organisms - GFP characteristics - Applications - Short protein structure review
Quick Guide to AlphaScreen SureFire Assay Optimization
 PPLITION NOT lpha Technology Quick uide to lphascreen Sureire ssay Optimization Introduction Screening kinase activities in cell-based assays offers advantages over the more traditional biochemical approach
PPLITION NOT lpha Technology Quick uide to lphascreen Sureire ssay Optimization Introduction Screening kinase activities in cell-based assays offers advantages over the more traditional biochemical approach
QUANTITATIVE RT-PCR. A = B (1+e) n. A=amplified products, B=input templates, n=cycle number, and e=amplification efficiency.
 QUANTITATIVE RT-PCR Application: Quantitative RT-PCR is used to quantify mrna in both relative and absolute terms. It can be applied for the quantification of mrna expressed from endogenous genes, and
QUANTITATIVE RT-PCR Application: Quantitative RT-PCR is used to quantify mrna in both relative and absolute terms. It can be applied for the quantification of mrna expressed from endogenous genes, and
Technical Note. Roche Applied Science. No. LC 18/2004. Assay Formats for Use in Real-Time PCR
 Roche Applied Science Technical Note No. LC 18/2004 Purpose of this Note Assay Formats for Use in Real-Time PCR The LightCycler Instrument uses several detection channels to monitor the amplification of
Roche Applied Science Technical Note No. LC 18/2004 Purpose of this Note Assay Formats for Use in Real-Time PCR The LightCycler Instrument uses several detection channels to monitor the amplification of
Expresión de la información genética en eucariotas. similitudes y diferencias con procariotas. metodologías de estudio 2
 Expresión de la información genética en eucariotas similitudes y diferencias con procariotas metodologías de estudio 2 Víctor Romanowski, 2013 Polymerase chain reaction (PCR) The polymerase chain reaction
Expresión de la información genética en eucariotas similitudes y diferencias con procariotas metodologías de estudio 2 Víctor Romanowski, 2013 Polymerase chain reaction (PCR) The polymerase chain reaction
Optimized Protocol sirna Test Kit for Cell Lines and Adherent Primary Cells
 page 1 of 8 sirna Test Kit for Cell Lines and Adherent Primary Cells Kit principle Co-transfection of pmaxgfp, encoding the green fluorescent protein (GFP) from Pontellina p. with an sirna directed against
page 1 of 8 sirna Test Kit for Cell Lines and Adherent Primary Cells Kit principle Co-transfection of pmaxgfp, encoding the green fluorescent protein (GFP) from Pontellina p. with an sirna directed against
www.gbo.com/bioscience Tissue Culture 1 Cell/ Microplates 2 HTS- 3 Immunology/ HLA 4 Microbiology/ Bacteriology Purpose Beakers 5 Tubes/Multi-
 11 Cryo 5 Tubes/Multi 2 HTS 3 Immunology / Immunology Technical Information 3 I 2 96 Well ELISA 3 I 4 96 Well ELISA Strip Plates 3 I 6 8 Well Strip Plates 3 I 7 12 Well Strip Plates 3 I 8 16 Well Strip
11 Cryo 5 Tubes/Multi 2 HTS 3 Immunology / Immunology Technical Information 3 I 2 96 Well ELISA 3 I 4 96 Well ELISA Strip Plates 3 I 6 8 Well Strip Plates 3 I 7 12 Well Strip Plates 3 I 8 16 Well Strip
Technical Note. Roche Applied Science. No. LC 19/2004. Color Compensation
 Roche Applied Science Technical Note No. LC 19/2004 Purpose of this Note Color The LightCycler System is able to simultaneously detect and analyze more than one color in each capillary. Due to overlap
Roche Applied Science Technical Note No. LC 19/2004 Purpose of this Note Color The LightCycler System is able to simultaneously detect and analyze more than one color in each capillary. Due to overlap
Chapter 18: Applications of Immunology
 Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
Chapter 18: Applications of Immunology 1. Vaccinations 2. Monoclonal vs Polyclonal Ab 3. Diagnostic Immunology 1. Vaccinations What is Vaccination? A method of inducing artificial immunity by exposing
Colorful choices for your bright ideas
 Living Colors Fluorescent Proteins Reporters and Antibodies for Every Application Colorful choices for your bright ideas Living Colors Fluorescent Proteins Full color spectrum cyan to far red Ideal for
Living Colors Fluorescent Proteins Reporters and Antibodies for Every Application Colorful choices for your bright ideas Living Colors Fluorescent Proteins Full color spectrum cyan to far red Ideal for
BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS
 BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS 1. The Technology Strategy sets out six areas where technological developments are required to push the frontiers of knowledge
BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS 1. The Technology Strategy sets out six areas where technological developments are required to push the frontiers of knowledge
Bacterial Transformation and Plasmid Purification. Chapter 5: Background
 Bacterial Transformation and Plasmid Purification Chapter 5: Background History of Transformation and Plasmids Bacterial methods of DNA transfer Transformation: when bacteria take up DNA from their environment
Bacterial Transformation and Plasmid Purification Chapter 5: Background History of Transformation and Plasmids Bacterial methods of DNA transfer Transformation: when bacteria take up DNA from their environment
Making the switch to a safer CAR-T cell therapy
 Making the switch to a safer CAR-T cell therapy HaemaLogiX 2015 Technical Journal Club May 24 th 2016 Christina Müller - chimeric antigen receptor = CAR - CAR T cells are generated by lentiviral transduction
Making the switch to a safer CAR-T cell therapy HaemaLogiX 2015 Technical Journal Club May 24 th 2016 Christina Müller - chimeric antigen receptor = CAR - CAR T cells are generated by lentiviral transduction
March 19, 2014. Dear Dr. Duvall, Dr. Hambrick, and Ms. Smith,
 Dr. Daniel Duvall, Medical Officer Center for Medicare, Hospital and Ambulatory Policy Group Centers for Medicare and Medicaid Services 7500 Security Boulevard Baltimore, Maryland 21244 Dr. Edith Hambrick,
Dr. Daniel Duvall, Medical Officer Center for Medicare, Hospital and Ambulatory Policy Group Centers for Medicare and Medicaid Services 7500 Security Boulevard Baltimore, Maryland 21244 Dr. Edith Hambrick,
+ - bcma_03. Materiale per uso didattico 1. Kinesin light chain Kinesin heavy chain Kinesin heavy chain Kinesin light chain. tetratricopeptide repeat
 p.959-970 Kinesin light chain Kinesin heavy chain Kinesin heavy chain Kinesin light chain + - tetratricopeptide repeat Zona di possibile interazione con cargo Materiale per uso didattico 1 JIP-1 COOH-term
p.959-970 Kinesin light chain Kinesin heavy chain Kinesin heavy chain Kinesin light chain + - tetratricopeptide repeat Zona di possibile interazione con cargo Materiale per uso didattico 1 JIP-1 COOH-term
HCS604.03 Exercise 1 Dr. Jones Spring 2005. Recombinant DNA (Molecular Cloning) exercise:
 HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
HCS604.03 Exercise 1 Dr. Jones Spring 2005 Recombinant DNA (Molecular Cloning) exercise: The purpose of this exercise is to learn techniques used to create recombinant DNA or clone genes. You will clone
GeneCopoeia Genome Editing Tools for Safe Harbor Integration in. Mice and Humans. Ed Davis, Liuqing Qian, Ruiqing li, Junsheng Zhou, and Jinkuo Zhang
 G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
G e n e C o p o eia TM Expressway to Discovery APPLICATION NOTE Introduction GeneCopoeia Genome Editing Tools for Safe Harbor Integration in Mice and Humans Ed Davis, Liuqing Qian, Ruiqing li, Junsheng
Gene Expression Assays
 APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
MystiCq microrna cdna Synthesis Mix Catalog Number MIRRT Storage Temperature 20 C
 microrna cdna Synthesis Mix Catalog Number MIRRT Storage Temperature 20 C Product Description The microrna cdna Synthesis Mix has been designed to easily convert micrornas into cdna templates for qpcr
microrna cdna Synthesis Mix Catalog Number MIRRT Storage Temperature 20 C Product Description The microrna cdna Synthesis Mix has been designed to easily convert micrornas into cdna templates for qpcr
No Disclosures. Learning Objectives 10/25/13
 No Disclosures Gregory A. Brent, MD Departments of Medicine and Physiology David Geffen School of Medicine at UCLA VA Greater Los Angeles Healthcare System Learning Objectives Describe the pathways that
No Disclosures Gregory A. Brent, MD Departments of Medicine and Physiology David Geffen School of Medicine at UCLA VA Greater Los Angeles Healthcare System Learning Objectives Describe the pathways that
Sonalee Athavankar Daniel D. Clark, Ph.D. Blake R. Peterson, Ph.D. The Pennsylvania State University University Park, PA 16802-7000
 AD Award Number: DAMD17-02-1-0536 TITLE: Discovery of Cyclic Peptide Estrogens and Antiestrogens PRINCIPAL INVESTIGATOR: Sonalee Athavankar Daniel D. Clark, Ph.D. Blake R. Peterson, Ph.D. CONTRACTING ORGANIZATION:
AD Award Number: DAMD17-02-1-0536 TITLE: Discovery of Cyclic Peptide Estrogens and Antiestrogens PRINCIPAL INVESTIGATOR: Sonalee Athavankar Daniel D. Clark, Ph.D. Blake R. Peterson, Ph.D. CONTRACTING ORGANIZATION:
Real-time PCR: Understanding C t
 APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
Introduction APPLICATION NOTE
 2014 Princeton Instruments, Inc. All rights reserved. In the future, as fluorophores with increasingly shorter lifetimes are developed, the
2014 Princeton Instruments, Inc. All rights reserved. In the future, as fluorophores with increasingly shorter lifetimes are developed, the
Molecular Biology. Yeast Transformation. Yeast Plasmids. Gene Disruption, tagging. Cloning by Complementation. Epistasis
 Molecular Biology Yeast Transformation Yeast Plasmids Gene Disruption, tagging Cloning by Complementation Epistasis Transformation Transformation: introduction of DNA 1978, ca 1000x less efficient than
Molecular Biology Yeast Transformation Yeast Plasmids Gene Disruption, tagging Cloning by Complementation Epistasis Transformation Transformation: introduction of DNA 1978, ca 1000x less efficient than
Design of conditional gene targeting vectors - a recombineering approach
 Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
Recombineering protocol #4 Design of conditional gene targeting vectors - a recombineering approach Søren Warming, Ph.D. The purpose of this protocol is to help you in the gene targeting vector design
DNA Integrity Number (DIN) For the Assessment of Genomic DNA Samples in Real-Time Quantitative PCR (qpcr) Experiments
 DNA Integrity Number () For the Assessment of Genomic DNA Samples in Real-Time Quantitative PCR (qpcr) Experiments Application Note Nucleic Acid Analysis Author Arunkumar Padmanaban Agilent Technologies,
DNA Integrity Number () For the Assessment of Genomic DNA Samples in Real-Time Quantitative PCR (qpcr) Experiments Application Note Nucleic Acid Analysis Author Arunkumar Padmanaban Agilent Technologies,
How To Make A Tri Reagent
 TRI Reagent For processing tissues, cells cultured in monolayer or cell pellets Catalog Number T9424 Store at room temperature. TECHNICAL BULLETIN Product Description TRI Reagent is a quick and convenient
TRI Reagent For processing tissues, cells cultured in monolayer or cell pellets Catalog Number T9424 Store at room temperature. TECHNICAL BULLETIN Product Description TRI Reagent is a quick and convenient
Modulating Glucose Uptake in Skeletal Myotubes:
 icell Skeletal Myoblasts Application Protocol Introduction Modulating Glucose Uptake in Skeletal Myotubes: Insulin Induction with Bioluminescent Glucose Uptake Analysis The skeletal muscle is one of the
icell Skeletal Myoblasts Application Protocol Introduction Modulating Glucose Uptake in Skeletal Myotubes: Insulin Induction with Bioluminescent Glucose Uptake Analysis The skeletal muscle is one of the
Design high specificity CRISPR-Cas9 grnas: principles and tools. Heidi Huang, PhD
 Design high specificity CRISPR-Cas9 grnas: principles and tools Heidi Huang, PhD Webinar Agenda 1 2 3 4 Introduction of CRISPR-Cas9 grna Design Resources and Services Q&A 2 What is CRISPR? CRISPR Clustered
Design high specificity CRISPR-Cas9 grnas: principles and tools Heidi Huang, PhD Webinar Agenda 1 2 3 4 Introduction of CRISPR-Cas9 grna Design Resources and Services Q&A 2 What is CRISPR? CRISPR Clustered
Covalent Conjugation to Cytodiagnostics Carboxylated Gold Nanoparticles Tech Note #105
 Covalent Conjugation to Cytodiagnostics Carboxylated Gold Nanoparticles Tech Note #105 Background Gold nanoparticle conjugates have been widely used in biological research and biosensing applications.
Covalent Conjugation to Cytodiagnostics Carboxylated Gold Nanoparticles Tech Note #105 Background Gold nanoparticle conjugates have been widely used in biological research and biosensing applications.
Page finder. 1. Legal 3. 2. Handling 4 2.1. Safety warnings and precautions 4 2.2. Storage 4 2.3. Expiry 4
 GE Healthcare Amersham Low Molecular Weight Calibration Kit for SDS Electrophoresis A lyophilized mixture of six highly purified well-characterized proteins for use in molecular weight determination in
GE Healthcare Amersham Low Molecular Weight Calibration Kit for SDS Electrophoresis A lyophilized mixture of six highly purified well-characterized proteins for use in molecular weight determination in
Cell Biology Questions and Learning Objectives
 Cell Biology Questions and Learning Objectives (with hypothetical learning materials that might populate the objective) The topics and central questions listed here are typical for an introductory undergraduate
Cell Biology Questions and Learning Objectives (with hypothetical learning materials that might populate the objective) The topics and central questions listed here are typical for an introductory undergraduate
PrimePCR Assay Validation Report
 Gene Information Gene Name sorbin and SH3 domain containing 2 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SORBS2 Human Arg and c-abl represent the mammalian
Gene Information Gene Name sorbin and SH3 domain containing 2 Gene Symbol Organism Gene Summary Gene Aliases RefSeq Accession No. UniGene ID Ensembl Gene ID SORBS2 Human Arg and c-abl represent the mammalian
The Need for a PARP in vivo Pharmacodynamic Assay
 The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D., Chief Scientific Officer, Trevigen, Inc., Gaithersburg, MD For further infomation, please contact: William Booth, Ph.D. Tel: +44 (0)1235
The Need for a PARP in vivo Pharmacodynamic Assay Jay George, Ph.D., Chief Scientific Officer, Trevigen, Inc., Gaithersburg, MD For further infomation, please contact: William Booth, Ph.D. Tel: +44 (0)1235
Annexin V-FITC Apoptosis Detection Kit
 ab14085 Annexin V-FITC Apoptosis Detection Kit Instructions for Use For the rapid, sensitive and accurate measurement of Apoptosis in living cells (adherent and suspension). This product is for research
ab14085 Annexin V-FITC Apoptosis Detection Kit Instructions for Use For the rapid, sensitive and accurate measurement of Apoptosis in living cells (adherent and suspension). This product is for research
Milk protein analysis with the Agilent 2100 Bioanalyzer and the Agilent Protein 80 kit
 Milk protein analysis with the Agilent 21 Bioanalyzer and the Agilent Protein 8 kit Application Note Food Analysis Author Rainer Nitsche Agilent Technologies, Inc. Waldbronn, Germany Abstract Protein content
Milk protein analysis with the Agilent 21 Bioanalyzer and the Agilent Protein 8 kit Application Note Food Analysis Author Rainer Nitsche Agilent Technologies, Inc. Waldbronn, Germany Abstract Protein content
Interaktionen von RNAs und Proteinen
 Sonja Prohaska Computational EvoDevo Universitaet Leipzig June 9, 2015 Studying RNA-protein interactions Given: target protein known to bind to RNA problem: find binding partners and binding sites experimental
Sonja Prohaska Computational EvoDevo Universitaet Leipzig June 9, 2015 Studying RNA-protein interactions Given: target protein known to bind to RNA problem: find binding partners and binding sites experimental
MAB Solut. MABSolys Génopole Campus 1 5 rue Henri Desbruères 91030 Evry Cedex. www.mabsolut.com. is involved at each stage of your project
 Mabsolus-2015-UK:Mise en page 1 03/07/15 14:13 Page1 Services provider Department of MABSolys from conception to validation MAB Solut is involved at each stage of your project Creation of antibodies Production
Mabsolus-2015-UK:Mise en page 1 03/07/15 14:13 Page1 Services provider Department of MABSolys from conception to validation MAB Solut is involved at each stage of your project Creation of antibodies Production
REAL TIME PCR USING SYBR GREEN
 REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
Application Note No. 2 / July 2012. Quantitative Assessment of Cell Quality, Viability and Proliferation. System
 Application Note No. 2 / July 2012 Quantitative Assessment of Cell Quality, Viability and Proliferation System Quantitative Assessment of Cell Quality, Viability and Proliferation Introduction In vitro
Application Note No. 2 / July 2012 Quantitative Assessment of Cell Quality, Viability and Proliferation System Quantitative Assessment of Cell Quality, Viability and Proliferation Introduction In vitro
Antibody Function & Structure
 Antibody Function & Structure Specifically bind to antigens in both the recognition phase (cellular receptors) and during the effector phase (synthesis and secretion) of humoral immunity Serology: the
Antibody Function & Structure Specifically bind to antigens in both the recognition phase (cellular receptors) and during the effector phase (synthesis and secretion) of humoral immunity Serology: the
INTERFERin in vitro sirna/mirna transfection reagent PROTOCOL. 1 Standard sirna transfection of adherent cells... 2
 INTERFERin in vitro sirna/mirna transfection reagent PROTOCOL DESCRIPTION INTERFERin is a powerful sirna/mirna transfection reagent that ensures efficient gene silencing and reproducible transfection in
INTERFERin in vitro sirna/mirna transfection reagent PROTOCOL DESCRIPTION INTERFERin is a powerful sirna/mirna transfection reagent that ensures efficient gene silencing and reproducible transfection in
How to Biotinylate with Reproducible Results
 How to Biotinylate with Reproducible Results Introduction The Biotin Streptavidin system continues to be used in many protein based biological research applications including; ELISAs, immunoprecipitation,
How to Biotinylate with Reproducible Results Introduction The Biotin Streptavidin system continues to be used in many protein based biological research applications including; ELISAs, immunoprecipitation,
ab139418 Propidium Iodide Flow Cytometry Kit for Cell Cycle Analysis
 ab139418 Propidium Iodide Flow Cytometry Kit for Cell Cycle Analysis Instructions for Use To determine cell cycle status in tissue culture cell lines by measuring DNA content using a flow cytometer. This
ab139418 Propidium Iodide Flow Cytometry Kit for Cell Cycle Analysis Instructions for Use To determine cell cycle status in tissue culture cell lines by measuring DNA content using a flow cytometer. This
Recombinant DNA Unit Exam
 Recombinant DNA Unit Exam Question 1 Restriction enzymes are extensively used in molecular biology. Below are the recognition sites of two of these enzymes, BamHI and BclI. a) BamHI, cleaves after the
Recombinant DNA Unit Exam Question 1 Restriction enzymes are extensively used in molecular biology. Below are the recognition sites of two of these enzymes, BamHI and BclI. a) BamHI, cleaves after the
Supplementary Figure 1.
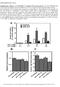 Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
Supplementary Figure 1. (A) MicroRNA 212 enhances IS from pancreatic β-cells. INS-1 832/3 β-cells were transfected with precursors for mirnas 212, 375, or negative control oligonucleotides. 48 hrs after
TransIT -2020 Transfection Reagent
 Quick Reference Protocol, MSDS and Certificate of Analysis available at mirusbio.com/5400 INTRODUCTION TransIT -2020 Transfection Reagent is a high-performance, animal-origin free, broad spectrum reagent
Quick Reference Protocol, MSDS and Certificate of Analysis available at mirusbio.com/5400 INTRODUCTION TransIT -2020 Transfection Reagent is a high-performance, animal-origin free, broad spectrum reagent
Application Guide... 2
 Protocol for GenomePlex Whole Genome Amplification from Formalin-Fixed Parrafin-Embedded (FFPE) tissue Application Guide... 2 I. Description... 2 II. Product Components... 2 III. Materials to be Supplied
Protocol for GenomePlex Whole Genome Amplification from Formalin-Fixed Parrafin-Embedded (FFPE) tissue Application Guide... 2 I. Description... 2 II. Product Components... 2 III. Materials to be Supplied
Fluorescent Proteins: Theory, Applications and Best Practices
 Fluorescent Proteins: Theory, Applications and Best Practices First identified in 1962 in sea creatures, fluorescent proteins have proved versatile and extremely useful, as demonstrated by applications
Fluorescent Proteins: Theory, Applications and Best Practices First identified in 1962 in sea creatures, fluorescent proteins have proved versatile and extremely useful, as demonstrated by applications
Co Extra (GM and non GM supply chains: Their CO EXistence and TRAceability) Outcomes of Co Extra
 GM and non GM supply chains: Their CO EXistence and TRAceability Outcomes of Co Extra Comparison of different real time PCR chemistries and their suitability for detection and quantification of genetically
GM and non GM supply chains: Their CO EXistence and TRAceability Outcomes of Co Extra Comparison of different real time PCR chemistries and their suitability for detection and quantification of genetically
Methionine Sulfoxide Immunoblotting Kit
 Methionine Sulfoxide Immunoblotting Kit Item No. 600160 Customer Service 800.364.9897 * Technical Support 888.526.5351 www.caymanchem.com TABLE OF CONTENTS GENERAL INFORMATION 3 Materials Supplied 4 Precautions
Methionine Sulfoxide Immunoblotting Kit Item No. 600160 Customer Service 800.364.9897 * Technical Support 888.526.5351 www.caymanchem.com TABLE OF CONTENTS GENERAL INFORMATION 3 Materials Supplied 4 Precautions
THE His Tag Antibody, mab, Mouse
 THE His Tag Antibody, mab, Mouse Cat. No. A00186 Technical Manual No. TM0243 Update date 01052011 I Description.... 1 II Key Features. 2 III Storage 2 IV Applications.... 2 V Examples - ELISA..... 2 VI
THE His Tag Antibody, mab, Mouse Cat. No. A00186 Technical Manual No. TM0243 Update date 01052011 I Description.... 1 II Key Features. 2 III Storage 2 IV Applications.... 2 V Examples - ELISA..... 2 VI
Biophysics of Molecules
 Biophysics of Molecules Concepts of cell adhesion and cellular mechanostransduction part II Dr. Carsten Grashoff MPI of Biochemistry E-mail: cgrasho@biochem.mpg.de Measuring mechanotransduction The problem
Biophysics of Molecules Concepts of cell adhesion and cellular mechanostransduction part II Dr. Carsten Grashoff MPI of Biochemistry E-mail: cgrasho@biochem.mpg.de Measuring mechanotransduction The problem
Real-time monitoring of rolling circle amplification using aggregation-induced emission: applications for biological detection
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 215 Supplementary Information Real-time monitoring of rolling circle amplification using aggregation-induced
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 215 Supplementary Information Real-time monitoring of rolling circle amplification using aggregation-induced
Anti-ATF6 α antibody, mouse monoclonal (1-7)
 Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
Anti-ATF6 α antibody, mouse monoclonal (1-7) 73-500 50 ug ATF6 (activating transcription factor 6) is an endoplasmic reticulum (ER) membrane-bound transcription factor activated in response to ER stress.
Nucleic Acid Purity Assessment using A 260 /A 280 Ratios
 Nucleic Acid Purity Assessment using A 260 /A 280 Ratios A common practice in molecular biology is to perform a quick assessment of the purity of nucleic acid samples by determining the ratio of spectrophotometric
Nucleic Acid Purity Assessment using A 260 /A 280 Ratios A common practice in molecular biology is to perform a quick assessment of the purity of nucleic acid samples by determining the ratio of spectrophotometric
Protein immunoblotting
 Protein immunoblotting (Western blotting) Dr. Serageldeen A. A. Sultan Lecturer of virology Dept. of Microbiology SVU, Qena, Egypt seaas@lycos.com Western blotting -It is an analytical technique used to
Protein immunoblotting (Western blotting) Dr. Serageldeen A. A. Sultan Lecturer of virology Dept. of Microbiology SVU, Qena, Egypt seaas@lycos.com Western blotting -It is an analytical technique used to
www.gbo.com/bioscience Tissue Culture 1 Cell/ Microplates 2 HTS- 3 Immunology/ HLA 4 Microbiology/ Bacteriology Purpose Beakers 5 Tubes/Multi-
 11 Cryo 5 Tubes/Multi 2 HTS 3 Immunology / Immunology Technical Information 3 I 2 96 Well ELISA 3 I 4 96 Well ELISA Strip Plates 3 I 6 8 Well Strip Plates 3 I 7 12 Well Strip Plates 3 I 8 16 Well Strip
11 Cryo 5 Tubes/Multi 2 HTS 3 Immunology / Immunology Technical Information 3 I 2 96 Well ELISA 3 I 4 96 Well ELISA Strip Plates 3 I 6 8 Well Strip Plates 3 I 7 12 Well Strip Plates 3 I 8 16 Well Strip
DNA-functionalized hydrogels for confined membrane-free in vitro transcription/translation
 Electronic Supplementary Material (ESI) for Lab on a Chip. This journal is The Royal Society of Chemistry 2014 DNA-functionalized hydrogels for confined membrane-free in vitro transcription/translation
Electronic Supplementary Material (ESI) for Lab on a Chip. This journal is The Royal Society of Chemistry 2014 DNA-functionalized hydrogels for confined membrane-free in vitro transcription/translation
Amersham High Molecular Weight Calibration Kit for native electrophoresis
 Amersham High Molecular Weight Calibration Kit for native electrophoresis A lyophilized mixture of five highly purified well-characterized proteins for use in molecular weight estimation under non-denaturing
Amersham High Molecular Weight Calibration Kit for native electrophoresis A lyophilized mixture of five highly purified well-characterized proteins for use in molecular weight estimation under non-denaturing
LIVE/DEAD Fixable Dead Cell Stain Kits
 USER GUIDE LIVE/DEAD Fixable Dead Cell Stain Kits Pub. No. MAN0002416 (MP34955) Rev. A.0 Table 1. Contents and storage Material Amount Storage Stability Individual Kits: Blue, violet, aqua, yellow-, green,
USER GUIDE LIVE/DEAD Fixable Dead Cell Stain Kits Pub. No. MAN0002416 (MP34955) Rev. A.0 Table 1. Contents and storage Material Amount Storage Stability Individual Kits: Blue, violet, aqua, yellow-, green,
Novel Inhibitors for PRMT1 Discovered by High-Throughput Screening Using Activity-Based Fluorescence Polarization
 Novel Inhibitors for PRMT1 Discovered by High-Throughput Screening Using Activity-Based Fluorescence Polarization Myles B. C. Dillon, Daniel A. Bachovchin, Steven J. Brown, M. G. Finn, Hugh Rosen, Benjamin
Novel Inhibitors for PRMT1 Discovered by High-Throughput Screening Using Activity-Based Fluorescence Polarization Myles B. C. Dillon, Daniel A. Bachovchin, Steven J. Brown, M. G. Finn, Hugh Rosen, Benjamin
Introduction To Real Time Quantitative PCR (qpcr)
 Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
In vitro analysis of pri-mirna processing. by Drosha-DGCR8 complex. (Narry Kim s lab)
 In vitro analysis of pri-mirna processing by Drosha-DGCR8 complex (Narry Kim s lab) 1-1. Preparation of radiolabeled pri-mirna transcript The RNA substrate for a cropping reaction can be prepared by in
In vitro analysis of pri-mirna processing by Drosha-DGCR8 complex (Narry Kim s lab) 1-1. Preparation of radiolabeled pri-mirna transcript The RNA substrate for a cropping reaction can be prepared by in
Rapid GST Inclusion Body Solubilization and Renaturation Kit
 Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
Product Manual Rapid GST Inclusion Body Solubilization and Renaturation Kit Catalog Number AKR-110 FOR RESEARCH USE ONLY Not for use in diagnostic procedures Introduction Bacteria are widely used for His
Module 2 overview SPRING BREAK
 1 Module 2 overview lecture lab 1. Introduction to the module 1. Start-up protein eng. 2. Rational protein design 2. Site-directed mutagenesis 3. Fluorescence and sensors 3. DNA amplification 4. Protein
1 Module 2 overview lecture lab 1. Introduction to the module 1. Start-up protein eng. 2. Rational protein design 2. Site-directed mutagenesis 3. Fluorescence and sensors 3. DNA amplification 4. Protein
Annexin V-FITC Apoptosis Detection Kit
 ab14085 Annexin V-FITC Apoptosis Detection Kit Instructions for Use For the rapid, sensitive and accurate measurement of Apoptosis in living cells (adherent and suspension). This product is for research
ab14085 Annexin V-FITC Apoptosis Detection Kit Instructions for Use For the rapid, sensitive and accurate measurement of Apoptosis in living cells (adherent and suspension). This product is for research
