Mitochondrial DNA and the Peopling of the New World
|
|
|
- Cuthbert Emil Nichols
- 7 years ago
- Views:
Transcription
1 see full issue: May-June 2000 Mitochondrial DNA and the Peopling of the New World Genetic variations among Native Americans provide further clues to who first populated the Americas and when they arrived Theodore G. Schurr On the eve of Christopher Columbus's arrival on San Salvador (now part of the Bahamas) in 1492, there were perhaps several tens of millions of people inhabiting the Americas. Once it became evident that the inhabitants of this New World were not, in fact, East Indians (as Columbus had at first supposed), the existence of the Native American population became a huge puzzle to the Renaissance Europeans. Just who were these people across the ocean, and where did they come from? Various theories were proposed in the centuries that followed, including the notion that the Native Americans (now often called Amerindians) were the descendants of the "lost tribes of Israel." Some scholars even attempted to draw parallels between the Amerindians and the contemporary European Jews of the era. It was not until the 18th century that scholars hit on the notion, now well established, that the Amerindians originated on the Asian continent. (Recent claims including the putative "Caucasian" characteristics of the Kennewick skeleton that European stock may have been present in pre-columbian America do not deny the overwhelming contribution of Asiatic peoples to the ancestry of modern Amerindians.) Anthropologists have been struggling with the question ever since, attempting to explain the diversity of cultures, languages and biological traits among Native Americans. Were there several waves of migrations from Asia, or merely one large movement? Did the ancestral populations reach the Americas only 12,000 years ago (as traditionally believed) or much earlier? Many recent studies have been guided by a tripartite migration model, which envisions three waves of migration corresponding to a three-part division of Native American languages: Amerind, Na-Dene and Eskaleut.
2 However, not all scientists agree that these subgroups represent an accurate partition of the Amerindian languages. Furthermore, the dental and nuclear-dna studies that were used to support a tripartite migration are not unequivocal. The other major question concerning Native American origins asks when people first might have arrived in the Americas. Archeological excavations in North America date the oldest human artifacts to between 11,000 and 14,000 years ago, whereas South American artifacts appear to be at least contemporaneous, if not older. Linguistic analyses also suggest entry dates ranging between 12,000 to 35,000 years ago. Studies of nuclear DNA genes further suggest an arrival time of 30,000 years ago, whereas research on dental variation indicates an emergence of the ancestral Amerindian groups about 18,000 to 20,000 years ago. There is indeed little agreement and much confusion. Over the past 10 years, however, new methodologies have provided a number of important insights into the peopling of the New World. Molecular genetic studies of the variation of mitochondrial DNA (mtdna) in Siberian and Amerindian populations have allowed further inferences to be made about the timing of the colonizations, the number of migrations that reached the New World, and possible regions from which ancestral Native Americans might have originated. Most notably, the new mtdna data suggest not only a very early movement of peoples into the New World but also the genetic contributions of populations originating outside of Siberia, from other parts of Asia. Overall, the mtdna research implies that the colonization of Siberia and the Americas was more complex than previously supposed that there were, in fact, multiple expansions of ancient peoples that contributed to the genetic diversity in aboriginal Siberian and Amerindian populations. The Beauty of mtdna Analysis Mitochondrial DNA is a gift to the molecular anthropologist seeking to sort out the genetic relations between people. Since mtdna is maternally inherited, the analysis of the mitochondrial genome is effectively the study of female genetic history within human populations. Unlike nuclear DNA, mtdna does not appear to undergo a recombination of its nucleotide bases during division and reproduction of the mitochondria. The lack of recombination permits mutations to accumulate in a more or less "linear" or chronological fashion within maternal lineages. As a result, there is minimal ambiguity in reconstructing the branching of female lineages along the accumulated changes in their mtdna. Mitochondrial DNA can also provide a record of human migration. The presence of identical mutations in mtdna from geographically separated human groups is a good indication that there might have been migrations or contacts between the groups. The mtdna also records stochastic processes that can alter the genetic composition of a population. Processes such as genetic drift the natural accumulation of unique genetic changes in a population that is isolated from other populations (and so cannot share its genes) and the founder effect in
3 which a newfound population of migrants contains a purely happenstance sample of genetic variants that are not representative of the parent population were common events in human history that are recorded in the mtdna of living peoples in terms of the frequencies and types of mtdnas present in them. The mtdna has two major regions that are the focus of genetic studies of Native American origins (Figure 2). About 94 percent of the mtdna sequence consists of coding regions, which include genes for ribosomal RNAs (rrnas), transfer RNAs (trnas) and proteins involved in oxidative phosphorylation (the major biochemical process within mitochondria). The remainder of the mtdna genome (about 1,100 nucleotide base pairs) consists of the non-coding control region, which initiates and regulates the replication off mtdna. The control region mutates about two- to ten times faster than the coding region; two hypervariable segments (HVS-I and HVS-II) are the most rapidly evolving portions of it. Two different molecular methods have been employed to take advantage of mtdna's genetic features. Restriction fragment length polymorphism (RFLP) analysis surveys individual mtdnas for sequence variation using a series of DNA-cutting enzymes (restriction endonucleases) that cleave the mtdna molecule at specific nucleotide sequences known as recognition sites. If there are genetic differences between two individuals such that restriction enzymes cut their DNA at different points in the mtdna, the resulting fragments will be of different lengths. In RFLP analysis, the unique combination of fragments detected by a set of restriction enzymes represent genetic polymorphisms within a mtdna haplotype. In turn, each grouping of related haplotypes that is defined by a specific set of shared RFLPs is called a haplogroup or mtdna lineage. Considerable genetic variation within and between human populations has been detected with RFLP analysis. Although a significant portion of this variation is shared among all human populations, a certain amount is only found within geographically circumscribed populations or ethnic groups. The other method of studying mtdna variation involves the direct sequencing of one or both of the hypervariable segments (HVS-I and HVS-II) of the control region. In contrast to RFLP analysis, which can be likened to a general scan of the mtdna genome, direct sequencing provides a nucleotide-by-nucleotide reading of a portion of the mtdna. Because the mutation rate is high in the control region, direct sequencing provides a detailed look at small genetic changes that may have taken place quite recently. Mutations in the control region help to define specific mtdna lineages in human populations (as do certain RFLPs), and they also reveal the genetic differentiation of these lineages in geographically circumscribed areas. By statistically assessing the amount of variation in the control regions within and between mtdna lineages, it is possible to estimate the relative age of genetic variants in
4 a particular geographic region. This process can also be carried out with haplotypes defined by RFLP analysis. Because the mtdna control region mutates rapidly, a single RFLP haplotype may be associated with several control region sequences. Such variations in the control region serve to distinguish subtypes of haplotypes (and may indicate a relatively recent divergence). In addition, the same control region sequence may be observed among different RFLP haplotypes, indicating the evolutionary convergence of control region sequences, the recent accumulation of new RFLPs or the loss of RFLP variation. Consequently, control region sequencing and RFLP haplotyping may not describe the same genotypes. However, in practice this does not necessarily pose a problem because each haplogroup usually has unique RFLPs and control region sequences that distinguish them from other mtdna lineages. The identity of the haplogroups is therefore usually fairly robust, and the relative level of diversity within each mtdna lineage is roughly equivalent. MtDNA in the Founding Americans What does the mtdna genome tell us about the diversity of the original migrations into the Americas? Many studies have now established that the vast majority of modern Native American haplotypes belong to merely four mtdna lineages, the haplogroups designated A, B, C and D. When subjected to the methods of phylogenetic analysis, the RFLP haplotypes usually segregate into their respective haplogroups, revealing their integrity as genealogical units (Figure 3). Moreover, ancient Amerindian samples obtained from different locations in the New World reveal the same general pattern of mtdna diversity. Sequencing studies of the HVS-I region also provide an independent confirmation of four primary mtdna haplogroups in Native Americans. A comparison of Native Americans, Siberians and Asians reveals that the same mtdna lineages in all groups share mutations in the control region that are specific to the haplogroups. The simplest explanation is that the control region mutations arose in Asia in the founding mtdna lineages and were carried to the New World by the ancestral Native Americans. Most of the remaining variants in control region sequences of Native Americans appear to be unique (much like their RFLPs). This suggests that Native American and Asian groups diverged rapidly after the founding New World populations separated from their Asian parental populations. Although most Native American mtdnas fall into four distinct haplogroups, there is less of a consensus about the number of founding haplotypes that accompanied each haplogroup to the New World. To be considered a founding haplotype, candidate mtdnas have to meet several criteria. First, founding haplotypes should be widespread within Amerindians and shared between tribes, since they would have
5 preceded tribal differentiation. Second, founding haplotypes should be central to the branching of their haplogroup in phylogenetic analyses because all new haplotypes would have originated from them. Third, founding haplotypes should be present in East Asian and Siberian populations because they originated in those regions. Conversely, mtdna haplotypes that are derived from the founding haplotypes should have a limited distribution. When these criteria are applied to Native American mtdna, only four to seven RFLP haplotypes clearly meet the standards, and only one (or possibly two) control region sequences are likely to have been in the founding migrants. Thus, both sets of data suggest that a limited number of founding mtdnas were brought to the New World by the founding populations. Another implication is that either some degree of reduction in genetic diversity took place during the colonization of the New World, or a geographically specific subset of Asian mtdnas was brought to the Americas. The geographic and linguistic distribution of haplogroups A-to-D in the Americas also suggests that all four of them were present in the original migration(s). All four haplogroups are observed in populations throughout the Americas and are also found in the three proposed Native American linguistic groups (Amerind, Na-Dene, Eskaleut). However, the original Na-Dene Indians and Eskimo-Aleuts appear to have lacked haplogroup B. Although mtdnas from haplogroups A-to-D are often found together in a single population, many tribes lack haplotypes from at least one of these haplogroups. This trend reflects the fact that genetic drift and founder events have played a significant role in shaping the distribution of mtdna haplotypes in these populations. Such an interpretation is also supported by the high frequency of private mtdna haplotypes in different Amerindian tribes. These results support the idea that early tribal isolation and founder effects led to the divergence of tribal gene pools in different regions (although not all scientists agree on this point). It's also important to mention that the genetic composition of an ancient population may not be the same as the population currently occupying the same geographic region, because of migrations, genetic drift or other stochastic processes. For example, based on haplogroup frequencies, the ancient Stillwater Marsh population in the Great Basin region does not appear to be ancestral to any modern Amerindian population in the same area. On the other hand, ancient Eskimo and Aleut samples have nearly the same haplogroup frequencies as their modern descendents, and the same is true for the ancient Anasazi and Fremont cultures and the modern Pueblo Indian groups. Such results imply that once they became genetically distinct from surrounding groups, many regional Amerindian populations maintained their genetic
6 integrity over a considerable period of time. When Did They Arrive? The antiquity of the founding Native American haplogroups can also provide a temporal yardstick by which to measure human occupancy in the New World. Assuming a particular rate of mutational change for mtdna, it's possible to estimate the number of years that must have passed since two haplogroups or haplotypes diverged. Based on analyses of RFLP haplotypes and control region sequences, haplogroups A, C and D appear to be the oldest mtdna lineages in the New World (averaging about 47,650 to 23,535 years old). The ages of these haplogroups in Siberia are generally comparable to those for the Americas. The considerable age of these haplogroups suggests that the genetic links between Siberian and Native American populations are indeed quite ancient. The RFLP data further suggest that haplogroup B in the Americas is considerably younger (17,700 to 13,500 years old) than American haplogroups A, C and D. However, recent analyses of variation in control region sequences in Native Americans have indicated that haplogroup B may be as diverse (and therefore as old) as haplogroups A, C and D. In addition, previous work suggested that the haplotypes of haplogroup B were present in Central-East Asia at least 24,000 to 30,000 years ago. Overall, it appears that the four primary haplogroups in Native Americans were brought to the New World well before the last glacial maximum (about 18,000 years ago). The age of haplogroup A in Siberia is considerably younger than that in the Americas (averaging only 12,727 to 9,655 years old). This discrepancy arises because the Siberian estimate is based on RFLP data from only two populations, the Chukchi and Siberian Eskimos, whereas the Native American estimate is based on data from 15 to 20 populations. On the other hand, a relatively recent age for haplogroup A has also been estimated for North American Eskimos and Na-Dene Indians based on control region sequences. Thus, it appears that the ancient Beringian populations that gave rise to the Chukchi, the Eskimo-Aleuts and the Na-Dene Indians underwent a recent bottleneck in which variation was reduced (perhaps 13,000 to 7,000 years ago), followed by an expansion of haplogroup A in the Arctic and Subarctic regions of North America. "Other" MtDNA Lineages A number of mtdnas found in Native Americans do not fall into the haplogroups A-to-D. These were originally designated as "other" (OTHER) haplotypes, and the majority were attributed to non-native admixture because of their apparent affinities to European mtdnas. In particular, the OTHER haplotypes detected in the Ojibwa and Navajo resembled the haplogroup X mtdnas seen in French Canadians and other European
7 groups. A single haplotype in the Maya also appeared to belong to the European haplogroup H, the most commonly observed mtdna lineage in Caucasian American and European populations. These findings suggested that most of the OTHER haplotypes seen in Native Americans were likely to be of European origin and, hence, of more recent derivation in New World populations. However, recent work has shown that the OTHER haplotypes that looked similar to the European haplogroup X mtdnas actually belong to a divergent branch of this particular mtdna lineage. All the Amerindian haplogroup X haplotypes share a common core set of RFLP and control region sequence mutations with European haplogroup X mtdnas, but otherwise differ from them by at least several control region mutations. Also, four distinct sublineages of haplogroup X have been identified in Amerindian populations, implying that it has been in the New World long enough to have undergone considerable genetic diversification. In contrast with the distribution of haplogroups A to D, haplogroup X is found nearly exclusively in North American populations. Interestingly, it occurs at its highest frequencies among Algonkian-speaking groups such as the Ojibwa, with all four of the Amerindian sublineages being present with this population. This mtdna lineage has also been detected in two pre-columbian North American populations and may be present in a few ancient Brazilian samples. These data imply that haplogroup X was present in the New World long before Europeans first arrived in the New World. Indeed, in Amerindians haplogroup X ranges between 35,000 to 13,000 years old, depending on the number of founding haplotypes assumed to have been present in the ancestral population and whether RFLP or control region sequence data are used to make the estimates. Haplogroup X is now considered to be a fifth but minor founding mtdna lineage in Native American populations. Non-Native Admixture Aside from haplogroup X, additional OTHER mtdnas have been detected in Amerindian tribes, all of which appear to have different genetic affinities. Because many Native populations exhibit European admixture in nuclear genetic studies, it seems likely that some of their OTHER mtdnas were acquired through gene flow with modern Europeans. Indeed, this does appear to be the case, as European haplogroup H and T mtdnas are found in the Ojibwa, and European haplogroup H and J mtdnas are seen in the Cherokee. It is not yet clear whether the OTHER haplotypes appearing in South American populations such as the Argentinean Mapuche were acquired through non-native admixture (because of limited analyses). There has also been a non-negligible amount of African gene flow into certain Amerindian populations, because African slaves and their descendents intermixed with Native Americans in historical times. Evidence of African-Amerindian admixture has been detected in the
8 Seminoles of Florida and the Narragansett of Massachusetts, in the form of African haplogroup L mtdnas. Similar observations have been made for the Mixatec and Zapotec Indians of Southern Mexico. Judging from these data, African mtdna haplotypes may be present among other Native American groups. The remaining OTHER mtdnas exhibit mutational features similar to those seen in the majority of Asian haplotypes. The great majority of these "Asian" OTHER haplotypes have been observed in South American tribes from the Brazilian Amazon, such as the Makiritare and Yanomami, but a few have also been detected at very low frequencies in a handful of North American groups. These haplotypes possess two mutations that appear in all haplogroup C and D mtdnas, but they lack other characteristic markers of these two mtdna lineages. Because 50 to 75 percent of all Asian mtdnas also have these mutations, these particular OTHER mtdnas were thought to belong to a previously undetected Asian haplogroup that had been brought to the Americas by ancestral populations. However, when subjected to phylogenetic analysis, the control region sequences of these OTHER mtdnas are interspersed with those from Amerindian haplogroups C and D, because they share specific nucleotide substitutions that define these haplogroups. These results suggest that OTHER mtdnas having the two mutations are indigenous haplotypes that arose through reversion mutations after the entry and spread of haplogroups C and D in the Americas. Where Did They Come From? There has been considerable speculation about the areas from which ancestral Paleoindians emerged and expanded into the Americas. Recent studies have suggested that northern China, southeastern Siberia or Mongolia may have been sources because of haplogroup D being present in those regions. The highest frequencies of these four haplogroups occur in the Altai Mountain/Tuva/Lake Baikal region, implying that this general region gave rise to the founders of Native American populations. Otherwise, haplogroup B is absent in the vast majority of native Siberian populations, haplogroup A occurs at very low frequencies outside of Chukotka, and haplogroups C and D are the predominant mtdna lineages in northern Asia. However, the presence of a certain control region mutation in haplogroups C and D may point to alternative source areas for ancestral Native Americans. This mutation appears in the majority of both haplogroup C and D mtdnas in Native American populations, suggesting it is part of the original sequence motifs for both of them. Among all Asian and Siberian mtdnas, however, this mutation only appears in haplogroup C mtdnas from Mongolia and the Amur River region and in haplogroup D mtdnas in the Japanese, Korean and Ainu. This distribution suggests that East Asia as well as southeast Siberia or Mongolia might be source areas (or migration pathways) for these two haplogroups. The origins of haplogroup X mtdnas remain somewhat ambiguous. Judging from its age, haplogroup X could have arrived in the New World
9 either before or after the last glacial maximum (about 18,000 years ago). Regardless of when it was brought to the Americas, the lack of haplogroup X mtdnas in Asian and Siberian groups (and their presence in certain European populations) suggests that these haplotypes originated outside of eastern Siberia, perhaps taken through Beringia by an ancient Eurasian migratory event distinct from th migration(s) that brought the other four mtdna lineages to the Americas. Bibliography Bonatto, S. L., and F. M. Salzano Diversity and age of the four major mtdna haplogroups, and their implications for the peopling of the New World. American Journal of Human Genetics 61: [CrossRef] Brown, M. D., S. H. Hosseini, A. Torroni, J. Allen, T. G. Schurr, R. Scozzari, F. Curciani and D. C. Wallace Haplogroup X: An ancient link between Europe/Western Asia and North America? American Journal of Human Genetics 63: [CrossRef] Kolman, C. J., N. Sambuughin and E. Bermingham Mitochondrial DNA analysis of Mongolian populations and implications for the origins of New World founders. Genetics 142: Lorenz, J. G., and D. G. Smith Distribution of sequence variation in the mtdna control region of Native North Americans. Human Biology 69: Merriwether, D. A., F. Rothhammer and R. E. Ferrell Genetic variation in the New World: Ancient teeth, bone, and tissue as sources of DNA. Experientia 50: Nichols, J The spread of language around the Pacific rim. Evolutionary Anthropology 1: Schurr, T. G., R. I. Sukernik, E. B. Starikovskaya and D. C. Wallace Mitochondrial DNA diversity in Koryaks: Ancient and recent population expansions and dispersals in Okhotsk-Bering Sea region. American Journal of Physical Anthropology 108:1 40. Schurr, T. G., S. W. Ballinger, Y. Y. Gan, J. A. Hodge, D. A. Merriwether, D. N. Lawrence, W. C. Knowler, K. M. Weiss and D. C. Wallace Amerindian mitochondrial DNAs have rare Asian variants at high frequencies, suggesting they derived from four primary maternal lineages. American Journal of Human Genetics 46: Starikovskaya, Y. B., R. I. Sukernik, T. G. Schurr and D. C. Wallace Mitochondrial DNA diversity in Chukchi and Siberian Eskimos: Implications for the genetic prehistory of ancient Beringia. American Journal of Human Genetics 63: [CrossRef] Szathmary, E. J. E Genetics of aboriginal North Americans. Evolutionary Anthropology 1: Torroni, A., J. V. Neel, R. Barrantes, T. G. Schurr and D. C. Wallace A mitochondrial DNA "clock" for the Amerinds and its implications for timing their entry into North America.
10 Proceedings of the National Academy of Sciences, USA. 91: Article Tools printer friendly request classroom permission this article Of Possible Interest book review: An Education in Irrationality feature article: The Molecular Anatomy of an Ancient Adaptive Event book review: Limits of the Genetic Lexicon book review: Tracking Genes Across the Globe book review: Making Hay with Straw Men Sigma Xi, The Scientific Research Society
Rethinking Polynesian Origins: Human Settlement of the Pacific
 LENScience Senior Biology Seminar Series Rethinking Polynesian Origins: Human Settlement of the Pacific Michal Denny, and Lisa Matisoo-Smith Our Polynesian ancestors are renowned as some of the world s
LENScience Senior Biology Seminar Series Rethinking Polynesian Origins: Human Settlement of the Pacific Michal Denny, and Lisa Matisoo-Smith Our Polynesian ancestors are renowned as some of the world s
Lecture 6: Single nucleotide polymorphisms (SNPs) and Restriction Fragment Length Polymorphisms (RFLPs)
 Lecture 6: Single nucleotide polymorphisms (SNPs) and Restriction Fragment Length Polymorphisms (RFLPs) Single nucleotide polymorphisms or SNPs (pronounced "snips") are DNA sequence variations that occur
Lecture 6: Single nucleotide polymorphisms (SNPs) and Restriction Fragment Length Polymorphisms (RFLPs) Single nucleotide polymorphisms or SNPs (pronounced "snips") are DNA sequence variations that occur
From Africa to Aotearoa Part 1: Out of Africa
 From Africa to Aotearoa Part 1: Out of Africa The spread of modern humans out of Africa started around 65,000 years ago, and ended with the settlement of New Zealand 750 years ago. These PowerPoint presentations
From Africa to Aotearoa Part 1: Out of Africa The spread of modern humans out of Africa started around 65,000 years ago, and ended with the settlement of New Zealand 750 years ago. These PowerPoint presentations
Genetic Variation and Human Evolution Lynn B. Jorde, Ph.D. Department of Human Genetics University of Utah School of Medicine.
 Genetic Variation and Human Evolution Lynn B. Jorde, Ph.D. Department of Human Genetics University of Utah School of Medicine. The past two decades have witnessed an explosion of human genetic data. Innumerable
Genetic Variation and Human Evolution Lynn B. Jorde, Ph.D. Department of Human Genetics University of Utah School of Medicine. The past two decades have witnessed an explosion of human genetic data. Innumerable
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
DNA and Forensic Science
 DNA and Forensic Science Micah A. Luftig * Stephen Richey ** I. INTRODUCTION This paper represents a discussion of the fundamental principles of DNA technology as it applies to forensic testing. A brief
DNA and Forensic Science Micah A. Luftig * Stephen Richey ** I. INTRODUCTION This paper represents a discussion of the fundamental principles of DNA technology as it applies to forensic testing. A brief
Principles of Evolution - Origin of Species
 Theories of Organic Evolution X Multiple Centers of Creation (de Buffon) developed the concept of "centers of creation throughout the world organisms had arisen, which other species had evolved from X
Theories of Organic Evolution X Multiple Centers of Creation (de Buffon) developed the concept of "centers of creation throughout the world organisms had arisen, which other species had evolved from X
The Story of Human Evolution Part 1: From ape-like ancestors to modern humans
 The Story of Human Evolution Part 1: From ape-like ancestors to modern humans Slide 1 The Story of Human Evolution This powerpoint presentation tells the story of who we are and where we came from - how
The Story of Human Evolution Part 1: From ape-like ancestors to modern humans Slide 1 The Story of Human Evolution This powerpoint presentation tells the story of who we are and where we came from - how
AP Biology Essential Knowledge Student Diagnostic
 AP Biology Essential Knowledge Student Diagnostic Background The Essential Knowledge statements provided in the AP Biology Curriculum Framework are scientific claims describing phenomenon occurring in
AP Biology Essential Knowledge Student Diagnostic Background The Essential Knowledge statements provided in the AP Biology Curriculum Framework are scientific claims describing phenomenon occurring in
Gene Mapping Techniques
 Gene Mapping Techniques OBJECTIVES By the end of this session the student should be able to: Define genetic linkage and recombinant frequency State how genetic distance may be estimated State how restriction
Gene Mapping Techniques OBJECTIVES By the end of this session the student should be able to: Define genetic linkage and recombinant frequency State how genetic distance may be estimated State how restriction
Algorithms in Computational Biology (236522) spring 2007 Lecture #1
 Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
Practice Questions 1: Evolution
 Practice Questions 1: Evolution 1. Which concept is best illustrated in the flowchart below? A. natural selection B. genetic manipulation C. dynamic equilibrium D. material cycles 2. The diagram below
Practice Questions 1: Evolution 1. Which concept is best illustrated in the flowchart below? A. natural selection B. genetic manipulation C. dynamic equilibrium D. material cycles 2. The diagram below
Forensic DNA Testing Terminology
 Forensic DNA Testing Terminology ABI 310 Genetic Analyzer a capillary electrophoresis instrument used by forensic DNA laboratories to separate short tandem repeat (STR) loci on the basis of their size.
Forensic DNA Testing Terminology ABI 310 Genetic Analyzer a capillary electrophoresis instrument used by forensic DNA laboratories to separate short tandem repeat (STR) loci on the basis of their size.
Biology 1406 - Notes for exam 5 - Population genetics Ch 13, 14, 15
 Biology 1406 - Notes for exam 5 - Population genetics Ch 13, 14, 15 Species - group of individuals that are capable of interbreeding and producing fertile offspring; genetically similar 13.7, 14.2 Population
Biology 1406 - Notes for exam 5 - Population genetics Ch 13, 14, 15 Species - group of individuals that are capable of interbreeding and producing fertile offspring; genetically similar 13.7, 14.2 Population
Mitochondrial DNA Analysis
 Mitochondrial DNA Analysis Lineage Markers Lineage markers are passed down from generation to generation without changing Except for rare mutation events They can help determine the lineage (family tree)
Mitochondrial DNA Analysis Lineage Markers Lineage markers are passed down from generation to generation without changing Except for rare mutation events They can help determine the lineage (family tree)
Globally, about 9.7% of cancers in men are prostate cancers, and the risk of developing the
 Chapter 5 Analysis of Prostate Cancer Association Study Data 5.1 Risk factors for Prostate Cancer Globally, about 9.7% of cancers in men are prostate cancers, and the risk of developing the disease has
Chapter 5 Analysis of Prostate Cancer Association Study Data 5.1 Risk factors for Prostate Cancer Globally, about 9.7% of cancers in men are prostate cancers, and the risk of developing the disease has
Summary. 16 1 Genes and Variation. 16 2 Evolution as Genetic Change. Name Class Date
 Chapter 16 Summary Evolution of Populations 16 1 Genes and Variation Darwin s original ideas can now be understood in genetic terms. Beginning with variation, we now know that traits are controlled by
Chapter 16 Summary Evolution of Populations 16 1 Genes and Variation Darwin s original ideas can now be understood in genetic terms. Beginning with variation, we now know that traits are controlled by
Lab 2/Phylogenetics/September 16, 2002 1 PHYLOGENETICS
 Lab 2/Phylogenetics/September 16, 2002 1 Read: Tudge Chapter 2 PHYLOGENETICS Objective of the Lab: To understand how DNA and protein sequence information can be used to make comparisons and assess evolutionary
Lab 2/Phylogenetics/September 16, 2002 1 Read: Tudge Chapter 2 PHYLOGENETICS Objective of the Lab: To understand how DNA and protein sequence information can be used to make comparisons and assess evolutionary
Name Class Date. binomial nomenclature. MAIN IDEA: Linnaeus developed the scientific naming system still used today.
 Section 1: The Linnaean System of Classification 17.1 Reading Guide KEY CONCEPT Organisms can be classified based on physical similarities. VOCABULARY taxonomy taxon binomial nomenclature genus MAIN IDEA:
Section 1: The Linnaean System of Classification 17.1 Reading Guide KEY CONCEPT Organisms can be classified based on physical similarities. VOCABULARY taxonomy taxon binomial nomenclature genus MAIN IDEA:
The retreat of glaciers and the original people of the Great Lakes
 Subject/target grade: Grade 9-12 Local History, Ecology, or Earth/Environmental Science classes Duration: Four 50-minute class periods; one optional half-day field activity Setting: Classroom Materials
Subject/target grade: Grade 9-12 Local History, Ecology, or Earth/Environmental Science classes Duration: Four 50-minute class periods; one optional half-day field activity Setting: Classroom Materials
14.3 Studying the Human Genome
 14.3 Studying the Human Genome Lesson Objectives Summarize the methods of DNA analysis. State the goals of the Human Genome Project and explain what we have learned so far. Lesson Summary Manipulating
14.3 Studying the Human Genome Lesson Objectives Summarize the methods of DNA analysis. State the goals of the Human Genome Project and explain what we have learned so far. Lesson Summary Manipulating
Evolution (18%) 11 Items Sample Test Prep Questions
 Evolution (18%) 11 Items Sample Test Prep Questions Grade 7 (Evolution) 3.a Students know both genetic variation and environmental factors are causes of evolution and diversity of organisms. (pg. 109 Science
Evolution (18%) 11 Items Sample Test Prep Questions Grade 7 (Evolution) 3.a Students know both genetic variation and environmental factors are causes of evolution and diversity of organisms. (pg. 109 Science
Y Chromosome Markers
 Y Chromosome Markers Lineage Markers Autosomal chromosomes recombine with each meiosis Y and Mitochondrial DNA does not This means that the Y and mtdna remains constant from generation to generation Except
Y Chromosome Markers Lineage Markers Autosomal chromosomes recombine with each meiosis Y and Mitochondrial DNA does not This means that the Y and mtdna remains constant from generation to generation Except
Cultural Anthropology Theories, Perspectives & Methodologies. Different ways of examining and understanding different cultures
 Cultural Anthropology Theories, Perspectives & Methodologies Different ways of examining and understanding different cultures Schools of Thought Cultural anthropologists develop theories to better understand
Cultural Anthropology Theories, Perspectives & Methodologies Different ways of examining and understanding different cultures Schools of Thought Cultural anthropologists develop theories to better understand
Matthew Kaplan and Taylor Edwards. University of Arizona Tucson, Arizona
 Matthew Kaplan and Taylor Edwards University of Arizona Tucson, Arizona Unresolved paternity Consent for testing Ownership of Samples Uncovering genetic disorders SRY Reversal / Klinefelter s Syndrome
Matthew Kaplan and Taylor Edwards University of Arizona Tucson, Arizona Unresolved paternity Consent for testing Ownership of Samples Uncovering genetic disorders SRY Reversal / Klinefelter s Syndrome
Evolution, Natural Selection, and Adaptation
 Evolution, Natural Selection, and Adaptation Nothing in biology makes sense except in the light of evolution. (Theodosius Dobzhansky) Charles Darwin (1809-1882) Voyage of HMS Beagle (1831-1836) Thinking
Evolution, Natural Selection, and Adaptation Nothing in biology makes sense except in the light of evolution. (Theodosius Dobzhansky) Charles Darwin (1809-1882) Voyage of HMS Beagle (1831-1836) Thinking
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE Q5B
 INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
INTERNATIONAL CONFERENCE ON HARMONISATION OF TECHNICAL REQUIREMENTS FOR REGISTRATION OF PHARMACEUTICALS FOR HUMAN USE ICH HARMONISED TRIPARTITE GUIDELINE QUALITY OF BIOTECHNOLOGICAL PRODUCTS: ANALYSIS
Biology Behind the Crime Scene Week 4: Lab #4 Genetics Exercise (Meiosis) and RFLP Analysis of DNA
 Page 1 of 5 Biology Behind the Crime Scene Week 4: Lab #4 Genetics Exercise (Meiosis) and RFLP Analysis of DNA Genetics Exercise: Understanding how meiosis affects genetic inheritance and DNA patterns
Page 1 of 5 Biology Behind the Crime Scene Week 4: Lab #4 Genetics Exercise (Meiosis) and RFLP Analysis of DNA Genetics Exercise: Understanding how meiosis affects genetic inheritance and DNA patterns
European Medicines Agency
 European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
SECOND INTERNATIONAL CONFERENCE ON THE GREAT MIGRATIONS ASIA TO AMERICA
 The Permanent Delegation of Kazakhstan to the UNESCO The Embassy of the Republic of Kazakhstan to the United States of America The Harriman Institute, Columbia University East Central European Center,
The Permanent Delegation of Kazakhstan to the UNESCO The Embassy of the Republic of Kazakhstan to the United States of America The Harriman Institute, Columbia University East Central European Center,
IV. DEMOGRAPHIC PROFILE OF THE OLDER POPULATION
 World Population Ageing 195-25 IV. DEMOGRAPHIC PROFILE OF THE OLDER POPULATION A. AGE COMPOSITION Older populations themselves are ageing A notable aspect of the global ageing process is the progressive
World Population Ageing 195-25 IV. DEMOGRAPHIC PROFILE OF THE OLDER POPULATION A. AGE COMPOSITION Older populations themselves are ageing A notable aspect of the global ageing process is the progressive
Okami Study Guide: Chapter 3 1
 Okami Study Guide: Chapter 3 1 Chapter in Review 1. Heredity is the tendency of offspring to resemble their parents in various ways. Genes are units of heredity. They are functional strands of DNA grouped
Okami Study Guide: Chapter 3 1 Chapter in Review 1. Heredity is the tendency of offspring to resemble their parents in various ways. Genes are units of heredity. They are functional strands of DNA grouped
( 1) Most human populations are a product of mixture of genetically distinct groups that intermixed within the last 4,000 years.
 Frequently asked questions about A Genetic Atlas of Human Admixture History G. Hellenthal, G.B.J. Busby, G. Band, J.F. Wilson, C. Capelli, D. Falush, S. Myers, Science (2014) SUMMARY What is your work
Frequently asked questions about A Genetic Atlas of Human Admixture History G. Hellenthal, G.B.J. Busby, G. Band, J.F. Wilson, C. Capelli, D. Falush, S. Myers, Science (2014) SUMMARY What is your work
A Hands-On Exercise To Demonstrate Evolution
 HOW-TO-DO-IT A Hands-On Exercise To Demonstrate Evolution by Natural Selection & Genetic Drift H ELEN J. YOUNG T RUMAN P. Y OUNG Although students learn (i.e., hear about) the components of evolution by
HOW-TO-DO-IT A Hands-On Exercise To Demonstrate Evolution by Natural Selection & Genetic Drift H ELEN J. YOUNG T RUMAN P. Y OUNG Although students learn (i.e., hear about) the components of evolution by
PRINCIPLES OF POPULATION GENETICS
 PRINCIPLES OF POPULATION GENETICS FOURTH EDITION Daniel L. Hartl Harvard University Andrew G. Clark Cornell University UniversitSts- und Landesbibliothek Darmstadt Bibliothek Biologie Sinauer Associates,
PRINCIPLES OF POPULATION GENETICS FOURTH EDITION Daniel L. Hartl Harvard University Andrew G. Clark Cornell University UniversitSts- und Landesbibliothek Darmstadt Bibliothek Biologie Sinauer Associates,
Worksheet: The theory of natural selection
 Worksheet: The theory of natural selection Senior Phase Grade 7-9 Learning area: Natural Science Strand: Life and living Theme: Biodiversity, change and continuity Specific Aim 1: Acquiring knowledge of
Worksheet: The theory of natural selection Senior Phase Grade 7-9 Learning area: Natural Science Strand: Life and living Theme: Biodiversity, change and continuity Specific Aim 1: Acquiring knowledge of
Biological Sciences Initiative. Human Genome
 Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Biological Sciences Initiative HHMI Human Genome Introduction In 2000, researchers from around the world published a draft sequence of the entire genome. 20 labs from 6 countries worked on the sequence.
Bayesian coalescent inference of population size history
 Bayesian coalescent inference of population size history Alexei Drummond University of Auckland Workshop on Population and Speciation Genomics, 2016 1st February 2016 1 / 39 BEAST tutorials Population
Bayesian coalescent inference of population size history Alexei Drummond University of Auckland Workshop on Population and Speciation Genomics, 2016 1st February 2016 1 / 39 BEAST tutorials Population
Genetics Lecture Notes 7.03 2005. Lectures 1 2
 Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Use of the Agilent 2100 Bioanalyzer and the DNA 500 LabChip in the Analysis of PCR Amplified Mitochondrial DNA Application
 Use of the Agilent 2100 Bioanalyzer and the DNA LabChip in the Analysis of PCR Amplified Mitochondrial DNA Application Homeland Security/Forensics Author Mark Jensen Agilent Technologies, Inc. 2850 Centerville
Use of the Agilent 2100 Bioanalyzer and the DNA LabChip in the Analysis of PCR Amplified Mitochondrial DNA Application Homeland Security/Forensics Author Mark Jensen Agilent Technologies, Inc. 2850 Centerville
Given these characteristics of life, which of the following objects is considered a living organism? W. X. Y. Z.
 Cell Structure and Organization 1. All living things must possess certain characteristics. They are all composed of one or more cells. They can grow, reproduce, and pass their genes on to their offspring.
Cell Structure and Organization 1. All living things must possess certain characteristics. They are all composed of one or more cells. They can grow, reproduce, and pass their genes on to their offspring.
SNP Essentials The same SNP story
 HOW SNPS HELP RESEARCHERS FIND THE GENETIC CAUSES OF DISEASE SNP Essentials One of the findings of the Human Genome Project is that the DNA of any two people, all 3.1 billion molecules of it, is more than
HOW SNPS HELP RESEARCHERS FIND THE GENETIC CAUSES OF DISEASE SNP Essentials One of the findings of the Human Genome Project is that the DNA of any two people, all 3.1 billion molecules of it, is more than
CHAPTER ONE: A CONTINENT OF VILLAGES, TO 1500
 CHAPTER ONE: A CONTINENT OF VILLAGES, TO 1500 SETTLING THE CONTINENT Who Are the Indian People? Migration from Asia Clovis: The First American Technology NEW WAYS OF LIVING ON THE LAND Hunting Traditions
CHAPTER ONE: A CONTINENT OF VILLAGES, TO 1500 SETTLING THE CONTINENT Who Are the Indian People? Migration from Asia Clovis: The First American Technology NEW WAYS OF LIVING ON THE LAND Hunting Traditions
2. True or False? The sequence of nucleotides in the human genome is 90.9% identical from one person to the next. False (it s 99.
 1. True or False? A typical chromosome can contain several hundred to several thousand genes, arranged in linear order along the DNA molecule present in the chromosome. True 2. True or False? The sequence
1. True or False? A typical chromosome can contain several hundred to several thousand genes, arranged in linear order along the DNA molecule present in the chromosome. True 2. True or False? The sequence
Human Genome and Human Genome Project. Louxin Zhang
 Human Genome and Human Genome Project Louxin Zhang A Primer to Genomics Cells are the fundamental working units of every living systems. DNA is made of 4 nucleotide bases. The DNA sequence is the particular
Human Genome and Human Genome Project Louxin Zhang A Primer to Genomics Cells are the fundamental working units of every living systems. DNA is made of 4 nucleotide bases. The DNA sequence is the particular
WORLD. Geographic Trend Report for GMAT Examinees
 2011 WORLD Geographic Trend Report for GMAT Examinees WORLD Geographic Trend Report for GMAT Examinees The World Geographic Trend Report for GMAT Examinees identifies mobility trends among GMAT examinees
2011 WORLD Geographic Trend Report for GMAT Examinees WORLD Geographic Trend Report for GMAT Examinees The World Geographic Trend Report for GMAT Examinees identifies mobility trends among GMAT examinees
America's first people
 America's first people BY PAULINE S. JOHNSON These activities, designed to accompany First Peoples 1 and The Mystery of the First Americans 2, will enable students to explore the origins of human populations
America's first people BY PAULINE S. JOHNSON These activities, designed to accompany First Peoples 1 and The Mystery of the First Americans 2, will enable students to explore the origins of human populations
CURRICULUM GUIDE. When this Forensics course has been completed successfully, students should be able to:
 CURRICULUM GUIDE NAME OF COURSE: FORENSICS COURSE NUMBER: SCI 40 WRITTEN / REVISED: SEPTEMBER, 2011 LEVEL OF COURSE: REPLACMENT NUMBER OF CREDITS: SIX (6) PREREQUISITES: BIOLOGY GRADE LEVELS OFFERED TO:
CURRICULUM GUIDE NAME OF COURSE: FORENSICS COURSE NUMBER: SCI 40 WRITTEN / REVISED: SEPTEMBER, 2011 LEVEL OF COURSE: REPLACMENT NUMBER OF CREDITS: SIX (6) PREREQUISITES: BIOLOGY GRADE LEVELS OFFERED TO:
Chapter 8: Recombinant DNA 2002 by W. H. Freeman and Company Chapter 8: Recombinant DNA 2002 by W. H. Freeman and Company
 Genetic engineering: humans Gene replacement therapy or gene therapy Many technical and ethical issues implications for gene pool for germ-line gene therapy what traits constitute disease rather than just
Genetic engineering: humans Gene replacement therapy or gene therapy Many technical and ethical issues implications for gene pool for germ-line gene therapy what traits constitute disease rather than just
Written by Roberta Estes, www.dnaexplain.com, Copyright 2007-2012
 Proving Your Native American Heritage Written by Roberta Estes, www.dnaexplain.com, Copyright 2007-2012 Many American families carry oral histories of Native American heritage. Most often, we think of
Proving Your Native American Heritage Written by Roberta Estes, www.dnaexplain.com, Copyright 2007-2012 Many American families carry oral histories of Native American heritage. Most often, we think of
The sample is taken with a simple mouth swab no blood is involved. There will be instructions included on how to take the sample.
 DNA testing Thanks for your enquiry about DNA testing. I oversee the Scottish DNA Project on behalf of the University of Strathclyde, Glasgow and act as representative for Family Tree DNA who host our
DNA testing Thanks for your enquiry about DNA testing. I oversee the Scottish DNA Project on behalf of the University of Strathclyde, Glasgow and act as representative for Family Tree DNA who host our
Section 3 Comparative Genomics and Phylogenetics
 Section 3 Section 3 Comparative enomics and Phylogenetics At the end of this section you should be able to: Describe what is meant by DNA sequencing. Explain what is meant by Bioinformatics and Comparative
Section 3 Section 3 Comparative enomics and Phylogenetics At the end of this section you should be able to: Describe what is meant by DNA sequencing. Explain what is meant by Bioinformatics and Comparative
Understanding by Design. Title: BIOLOGY/LAB. Established Goal(s) / Content Standard(s): Essential Question(s) Understanding(s):
 Understanding by Design Title: BIOLOGY/LAB Standard: EVOLUTION and BIODIVERSITY Grade(s):9/10/11/12 Established Goal(s) / Content Standard(s): 5. Evolution and Biodiversity Central Concepts: Evolution
Understanding by Design Title: BIOLOGY/LAB Standard: EVOLUTION and BIODIVERSITY Grade(s):9/10/11/12 Established Goal(s) / Content Standard(s): 5. Evolution and Biodiversity Central Concepts: Evolution
The sequence of bases on the mrna is a code that determines the sequence of amino acids in the polypeptide being synthesized:
 Module 3F Protein Synthesis So far in this unit, we have examined: How genes are transmitted from one generation to the next Where genes are located What genes are made of How genes are replicated How
Module 3F Protein Synthesis So far in this unit, we have examined: How genes are transmitted from one generation to the next Where genes are located What genes are made of How genes are replicated How
Basic Concepts Recombinant DNA Use with Chapter 13, Section 13.2
 Name Date lass Master 19 Basic oncepts Recombinant DN Use with hapter, Section.2 Formation of Recombinant DN ut leavage Splicing opyright lencoe/mcraw-hill, a division of he Mcraw-Hill ompanies, Inc. Bacterial
Name Date lass Master 19 Basic oncepts Recombinant DN Use with hapter, Section.2 Formation of Recombinant DN ut leavage Splicing opyright lencoe/mcraw-hill, a division of he Mcraw-Hill ompanies, Inc. Bacterial
SIXTH GRADE PLATE TECTONICS 1 WEEK LESSON PLANS AND ACTIVITIES
 SIXTH GRADE PLATE TECTONICS 1 WEEK LESSON PLANS AND ACTIVITIES PLATE TECTONIC CYCLE OVERVIEW OF SIXTH GRADE VOLCANOES WEEK 1. PRE: Comparing the structure of different types of volcanoes. LAB: Plotting
SIXTH GRADE PLATE TECTONICS 1 WEEK LESSON PLANS AND ACTIVITIES PLATE TECTONIC CYCLE OVERVIEW OF SIXTH GRADE VOLCANOES WEEK 1. PRE: Comparing the structure of different types of volcanoes. LAB: Plotting
DNA Testing for Genealogy - What Can It Do For You??
 DNA Testing for Genealogy - What Can It Do For You?? Paper courtesy of Roberta Estes, www.dnaexplain.com, e-mail Roberta at Roberta@dnaexplain.com. Graphics courtesy of Family Tree DNA, www.familytreedna.com.
DNA Testing for Genealogy - What Can It Do For You?? Paper courtesy of Roberta Estes, www.dnaexplain.com, e-mail Roberta at Roberta@dnaexplain.com. Graphics courtesy of Family Tree DNA, www.familytreedna.com.
North Carolina Essential Standards Third grade Social Studies
 North Carolina s Third grade Social Studies In third grade, students draw upon knowledge learned in previous grades to develop more sophisticated understandings of how communities may be linked to form
North Carolina s Third grade Social Studies In third grade, students draw upon knowledge learned in previous grades to develop more sophisticated understandings of how communities may be linked to form
Recovering the Romanovs
 Recovering the Romanovs ACTIVITY 1 The Romanov Family: Screen #4 Inheritance of a Sex-linked Trait Key: H=normal allele; h=hemophilia allele; X=X chromosome; Y=Y chromosome 1. Use a Punnett square to show
Recovering the Romanovs ACTIVITY 1 The Romanov Family: Screen #4 Inheritance of a Sex-linked Trait Key: H=normal allele; h=hemophilia allele; X=X chromosome; Y=Y chromosome 1. Use a Punnett square to show
ISTEP+: Biology I End-of-Course Assessment Released Items and Scoring Notes
 ISTEP+: Biology I End-of-Course Assessment Released Items and Scoring Notes Page 1 of 22 Introduction Indiana students enrolled in Biology I participated in the ISTEP+: Biology I Graduation Examination
ISTEP+: Biology I End-of-Course Assessment Released Items and Scoring Notes Page 1 of 22 Introduction Indiana students enrolled in Biology I participated in the ISTEP+: Biology I Graduation Examination
Continuous and discontinuous variation
 Continuous and discontinuous variation Variation, the small differences that exist between individuals, can be described as being either discontinuous or continuous. Discontinuous variation This is where
Continuous and discontinuous variation Variation, the small differences that exist between individuals, can be described as being either discontinuous or continuous. Discontinuous variation This is where
Table of Contents: Introduction. DNA Tribes Digest July 30, 2010
 DNA Tribes Digest July 30, 2010 Copyright 2010 DNA Tribes. All rights reserved. To request an email subscription to DNA Tribes Digest, email digest@dnatribes.com with the subject heading Subscribe. To
DNA Tribes Digest July 30, 2010 Copyright 2010 DNA Tribes. All rights reserved. To request an email subscription to DNA Tribes Digest, email digest@dnatribes.com with the subject heading Subscribe. To
Fifth Grade Native American History. Lesson Plans
 Lesson Plans This unit is an introduction to Native American history in the 19 th and 20 th centuries. The lessons focus on U.S. government policies that have determined the official relationship between
Lesson Plans This unit is an introduction to Native American history in the 19 th and 20 th centuries. The lessons focus on U.S. government policies that have determined the official relationship between
DNA Tribes is on Facebook. Find us at http://facebook.com/dnatribes. DNA Tribes Digest February 1, 2013 Page 1 of 10
 Copyright 2013 DNA Tribes. All rights reserved. DNA Tribes Digest February 1, 2013 To request an email subscription to DNA Tribes Digest, email digest@dnatribes.com with the subject heading Subscribe.
Copyright 2013 DNA Tribes. All rights reserved. DNA Tribes Digest February 1, 2013 To request an email subscription to DNA Tribes Digest, email digest@dnatribes.com with the subject heading Subscribe.
Chapter 3. Chapter Outline. Chapter Outline 9/11/10. Heredity and Evolu4on
 Chapter 3 Heredity and Evolu4on Chapter Outline The Cell DNA Structure and Function Cell Division: Mitosis and Meiosis The Genetic Principles Discovered by Mendel Mendelian Inheritance in Humans Misconceptions
Chapter 3 Heredity and Evolu4on Chapter Outline The Cell DNA Structure and Function Cell Division: Mitosis and Meiosis The Genetic Principles Discovered by Mendel Mendelian Inheritance in Humans Misconceptions
Lecture 10 Friday, March 20, 2009
 Lecture 10 Friday, March 20, 2009 Reproductive isolating mechanisms Prezygotic barriers: Anything that prevents mating and fertilization is a prezygotic mechanism. Habitat isolation, behavioral isolation,
Lecture 10 Friday, March 20, 2009 Reproductive isolating mechanisms Prezygotic barriers: Anything that prevents mating and fertilization is a prezygotic mechanism. Habitat isolation, behavioral isolation,
Innovations in Molecular Epidemiology
 Innovations in Molecular Epidemiology Molecular Epidemiology Measure current rates of active transmission Determine whether recurrent tuberculosis is attributable to exogenous reinfection Determine whether
Innovations in Molecular Epidemiology Molecular Epidemiology Measure current rates of active transmission Determine whether recurrent tuberculosis is attributable to exogenous reinfection Determine whether
Name: Date: Period: DNA Unit: DNA Webquest
 Name: Date: Period: DNA Unit: DNA Webquest Part 1 History, DNA Structure, DNA Replication DNA History http://www.dnaftb.org/dnaftb/1/concept/index.html Read the text and answer the following questions.
Name: Date: Period: DNA Unit: DNA Webquest Part 1 History, DNA Structure, DNA Replication DNA History http://www.dnaftb.org/dnaftb/1/concept/index.html Read the text and answer the following questions.
Basic Principles of Forensic Molecular Biology and Genetics. Population Genetics
 Basic Principles of Forensic Molecular Biology and Genetics Population Genetics Significance of a Match What is the significance of: a fiber match? a hair match? a glass match? a DNA match? Meaning of
Basic Principles of Forensic Molecular Biology and Genetics Population Genetics Significance of a Match What is the significance of: a fiber match? a hair match? a glass match? a DNA match? Meaning of
Culture (from the Encarta Encyclopedia)
 Culture (from the Encarta Encyclopedia) 1. Introduction Culture, in anthropology, is the patterns of behavior and thinking that people living in social groups learn, create, and share. Culture distinguishes
Culture (from the Encarta Encyclopedia) 1. Introduction Culture, in anthropology, is the patterns of behavior and thinking that people living in social groups learn, create, and share. Culture distinguishes
World History: Essential Questions
 World History: Essential Questions Content Standard 1.0: Culture encompasses similarities and differences among people including their beliefs, knowledge, changes, values, and traditions. Students will
World History: Essential Questions Content Standard 1.0: Culture encompasses similarities and differences among people including their beliefs, knowledge, changes, values, and traditions. Students will
Historical Linguistics. Diachronic Analysis. Two Approaches to the Study of Language. Kinds of Language Change. What is Historical Linguistics?
 Historical Linguistics Diachronic Analysis What is Historical Linguistics? Historical linguistics is the study of how languages change over time and of their relationships with other languages. All languages
Historical Linguistics Diachronic Analysis What is Historical Linguistics? Historical linguistics is the study of how languages change over time and of their relationships with other languages. All languages
Reduced-Median-Network Analysis of Complete Mitochondrial DNA Coding-Region Sequences for the Major African, Asian, and European Haplogroups
 Am. J. Hum. Genet. 70:1152 1171, 2002 Reduced-Median-Network Analysis of Complete Mitochondrial DNA Coding-Region Sequences for the Major African, Asian, and European Haplogroups Corinna Herrnstadt, 1
Am. J. Hum. Genet. 70:1152 1171, 2002 Reduced-Median-Network Analysis of Complete Mitochondrial DNA Coding-Region Sequences for the Major African, Asian, and European Haplogroups Corinna Herrnstadt, 1
MCAS Biology. Review Packet
 MCAS Biology Review Packet 1 Name Class Date 1. Define organic. THE CHEMISTRY OF LIFE 2. All living things are made up of 6 essential elements: SPONCH. Name the six elements of life. S N P C O H 3. Elements
MCAS Biology Review Packet 1 Name Class Date 1. Define organic. THE CHEMISTRY OF LIFE 2. All living things are made up of 6 essential elements: SPONCH. Name the six elements of life. S N P C O H 3. Elements
Pacemaker World Geography and Cultures. correlated to. Florida Sunshine State Standards Social Studies Grades 6-8
 Pacemaker World Geography and Cultures correlated to Florida Sunshine State Standards Social Studies Grades 6-8 Pacemaker World Geography and Cultures Pearson Learning Group correlated to Sunshine State
Pacemaker World Geography and Cultures correlated to Florida Sunshine State Standards Social Studies Grades 6-8 Pacemaker World Geography and Cultures Pearson Learning Group correlated to Sunshine State
A Primer of Genome Science THIRD
 A Primer of Genome Science THIRD EDITION GREG GIBSON-SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc. Publishers Sunderland, Massachusetts USA Contents Preface xi 1 Genome Projects:
A Primer of Genome Science THIRD EDITION GREG GIBSON-SPENCER V. MUSE North Carolina State University Sinauer Associates, Inc. Publishers Sunderland, Massachusetts USA Contents Preface xi 1 Genome Projects:
Tracing the evolution of the genus Homo is important for understanding the ancestry of humans; the only living species of Homo.
 Section 3: Tracing the evolution of the genus Homo is important for understanding the ancestry of humans; the only living species of Homo. K What I Know W What I Want to Find Out L What I Learned Essential
Section 3: Tracing the evolution of the genus Homo is important for understanding the ancestry of humans; the only living species of Homo. K What I Know W What I Want to Find Out L What I Learned Essential
Arabidopsis. A Practical Approach. Edited by ZOE A. WILSON Plant Science Division, School of Biological Sciences, University of Nottingham
 Arabidopsis A Practical Approach Edited by ZOE A. WILSON Plant Science Division, School of Biological Sciences, University of Nottingham OXPORD UNIVERSITY PRESS List of Contributors Abbreviations xv xvu
Arabidopsis A Practical Approach Edited by ZOE A. WILSON Plant Science Division, School of Biological Sciences, University of Nottingham OXPORD UNIVERSITY PRESS List of Contributors Abbreviations xv xvu
Some ancestors are difficult to trace. Adoptions illegitimacies, name changes and migrations can present brick walls in our research that seem
 Some ancestors are difficult to trace. Adoptions illegitimacies, name changes and migrations can present brick walls in our research that seem impregnable. In other cases, documentation that would shed
Some ancestors are difficult to trace. Adoptions illegitimacies, name changes and migrations can present brick walls in our research that seem impregnable. In other cases, documentation that would shed
1 Mutation and Genetic Change
 CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Recovering the Romanovs
 Recovering the Romanovs Description of activity Recovering the Romanovs is an excellent opening to any unit on human genetics. To complete the three parts of this module, approximately three 40-minute
Recovering the Romanovs Description of activity Recovering the Romanovs is an excellent opening to any unit on human genetics. To complete the three parts of this module, approximately three 40-minute
Worksheet - COMPARATIVE MAPPING 1
 Worksheet - COMPARATIVE MAPPING 1 The arrangement of genes and other DNA markers is compared between species in Comparative genome mapping. As early as 1915, the geneticist J.B.S Haldane reported that
Worksheet - COMPARATIVE MAPPING 1 The arrangement of genes and other DNA markers is compared between species in Comparative genome mapping. As early as 1915, the geneticist J.B.S Haldane reported that
DAUGHERTY / DOUGHERTY NATIVE AMERICAN DNA
 PiquaShawnee Project DAUGHERTY / DOUGHERTY NATIVE AMERICAN DNA Gayland Eugene Daugherty joined this project to seek additional informaton about his Cherokee Native American ancestry. PiquaShawnee was initiated
PiquaShawnee Project DAUGHERTY / DOUGHERTY NATIVE AMERICAN DNA Gayland Eugene Daugherty joined this project to seek additional informaton about his Cherokee Native American ancestry. PiquaShawnee was initiated
How Did These Ocean Features and Continental Margins Form?
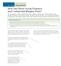 298 10.14 INVESTIGATION How Did These Ocean Features and Continental Margins Form? The terrain below contains various features on the seafloor, as well as parts of three continents. Some general observations
298 10.14 INVESTIGATION How Did These Ocean Features and Continental Margins Form? The terrain below contains various features on the seafloor, as well as parts of three continents. Some general observations
2. Incidence, prevalence and duration of breastfeeding
 2. Incidence, prevalence and duration of breastfeeding Key Findings Mothers in the UK are breastfeeding their babies for longer with one in three mothers still breastfeeding at six months in 2010 compared
2. Incidence, prevalence and duration of breastfeeding Key Findings Mothers in the UK are breastfeeding their babies for longer with one in three mothers still breastfeeding at six months in 2010 compared
CCR Biology - Chapter 9 Practice Test - Summer 2012
 Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Name: Class: Date: CCR Biology - Chapter 9 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Genetic engineering is possible
Elsevier Editorial System(tm) for Forensic Science International: Genetics Manuscript Draft
 Elsevier Editorial System(tm) for Forensic Science International: Genetics Manuscript Draft Manuscript Number: Title: A comment on the Paper: A comparison of Y-chromosomal lineage dating using either resequencing
Elsevier Editorial System(tm) for Forensic Science International: Genetics Manuscript Draft Manuscript Number: Title: A comment on the Paper: A comparison of Y-chromosomal lineage dating using either resequencing
SalarieS of chemists fall
 ACS news SalarieS of chemists fall Unemployment reaches new heights in 2009 as recession hits profession hard The economic recession has taken its toll on chemists. Despite holding up fairly well in previous
ACS news SalarieS of chemists fall Unemployment reaches new heights in 2009 as recession hits profession hard The economic recession has taken its toll on chemists. Despite holding up fairly well in previous
Patient Information. for Childhood
 Patient Information Genetic Testing for Childhood Hearing Loss Introduction This document describes the most common genetic cause of childhood hearing loss and explains the role of genetic testing. Childhood
Patient Information Genetic Testing for Childhood Hearing Loss Introduction This document describes the most common genetic cause of childhood hearing loss and explains the role of genetic testing. Childhood
IIID 14. Biotechnology in Fish Disease Diagnostics: Application of the Polymerase Chain Reaction (PCR)
 IIID 14. Biotechnology in Fish Disease Diagnostics: Application of the Polymerase Chain Reaction (PCR) Background Infectious diseases caused by pathogenic organisms such as bacteria, viruses, protozoa,
IIID 14. Biotechnology in Fish Disease Diagnostics: Application of the Polymerase Chain Reaction (PCR) Background Infectious diseases caused by pathogenic organisms such as bacteria, viruses, protozoa,
School of Nursing. Presented by Yvette Conley, PhD
 Presented by Yvette Conley, PhD What we will cover during this webcast: Briefly discuss the approaches introduced in the paper: Genome Sequencing Genome Wide Association Studies Epigenomics Gene Expression
Presented by Yvette Conley, PhD What we will cover during this webcast: Briefly discuss the approaches introduced in the paper: Genome Sequencing Genome Wide Association Studies Epigenomics Gene Expression
ICFT: An Assault On Biblical Creation (Genesis 1)
 Introduction. ICFT: An Assault On Biblical Creation (Genesis 1) A) In Colleges and Universities around the world, young men and women from Christians homes are challenged with questions by liberal professors
Introduction. ICFT: An Assault On Biblical Creation (Genesis 1) A) In Colleges and Universities around the world, young men and women from Christians homes are challenged with questions by liberal professors
PHYLOGENY AND EVOLUTION OF NEWCASTLE DISEASE VIRUS GENOTYPES
 Eötvös Lóránd University Biology Doctorate School Classical and molecular genetics program Project leader: Dr. László Orosz, corresponding member of HAS PHYLOGENY AND EVOLUTION OF NEWCASTLE DISEASE VIRUS
Eötvös Lóránd University Biology Doctorate School Classical and molecular genetics program Project leader: Dr. László Orosz, corresponding member of HAS PHYLOGENY AND EVOLUTION OF NEWCASTLE DISEASE VIRUS
Evidence for evolution factsheet
 The theory of evolution by natural selection is supported by a great deal of evidence. Fossils Fossils are formed when organisms become buried in sediments, causing little decomposition of the organism.
The theory of evolution by natural selection is supported by a great deal of evidence. Fossils Fossils are formed when organisms become buried in sediments, causing little decomposition of the organism.
AP BIOLOGY 2010 SCORING GUIDELINES (Form B)
 AP BIOLOGY 2010 SCORING GUIDELINES (Form B) Question 2 Certain human genetic conditions, such as sickle cell anemia, result from single base-pair mutations in DNA. (a) Explain how a single base-pair mutation
AP BIOLOGY 2010 SCORING GUIDELINES (Form B) Question 2 Certain human genetic conditions, such as sickle cell anemia, result from single base-pair mutations in DNA. (a) Explain how a single base-pair mutation
POLICY ON HUMAN REMAINS
 Policy document for the strategic development of The Manchester Museum POLICY ON HUMAN REMAINS Endorsed by: The University of Manchester Senior Executive Team Date for review: June 2010 1 Policy Statement
Policy document for the strategic development of The Manchester Museum POLICY ON HUMAN REMAINS Endorsed by: The University of Manchester Senior Executive Team Date for review: June 2010 1 Policy Statement
Mechanisms of Evolution
 page 2 page 3 Teacher's Notes Mechanisms of Evolution Grades: 11-12 Duration: 28 mins Summary of Program Evolution is the gradual change that can be seen in a population s genetic composition, from one
page 2 page 3 Teacher's Notes Mechanisms of Evolution Grades: 11-12 Duration: 28 mins Summary of Program Evolution is the gradual change that can be seen in a population s genetic composition, from one
A Correlation of Miller & Levine Biology 2014
 A Correlation of Miller & Levine Biology To Ohio s New Learning Standards for Science, 2011 Biology, High School Science Inquiry and Application Course Content A Correlation of, to Introduction This document
A Correlation of Miller & Levine Biology To Ohio s New Learning Standards for Science, 2011 Biology, High School Science Inquiry and Application Course Content A Correlation of, to Introduction This document
Can DNA Witness Race? : Forensic Uses of An Imperfect Ancestry Testing Technology
 Can DNA Witness Race? : Forensic Uses of An Imperfect Ancestry Testing Technology Duana Fullwiley Harvard University An intelligent evaluation of facts is often difficult or impossible without the application
Can DNA Witness Race? : Forensic Uses of An Imperfect Ancestry Testing Technology Duana Fullwiley Harvard University An intelligent evaluation of facts is often difficult or impossible without the application
