Conference: From DNA-Inspired Physics to Physics-Inspired Biology. 1-5 June How Proteins Find Their Targets on DNA
|
|
|
- Claribel Gordon
- 8 years ago
- Views:
Transcription
1 Conference: From DNA-Inspired Physics to Physics-Inspired Biology 1-5 June 2009 How Proteins Find Their Targets on DNA Anatoly B. KOLOMEISKY Rice University, Department of Chemistry Houston TX USA
2 Anatoly B. Kolomeisky Department of Chemistry How Proteins Find Their Targets on DNA
3 COLLABORATION: Prof. A.A. Kornyshev, Department of Chemistry, Faculty of Physical Sciences, Imperial College London, UK. Dr. A.G. Cherstvy, Research Center, Juelich, Germany. Rahul K. Das, graduate student in my group
4 Protein-DNA Interactions Play fundamental role in all biological processes Example #1: switching genes on and off in Ecoli bacteria Large numbers of Ecoli bacteria live in human intestines Bacteria needs to produce proteins which are made from aminoacids
5 Gene Switch in Bacteria Production of the amino acid tryptophan DNA low concentration of tryptophan high concentration of tryptophan target
6 Transcription Example #2: transcription proteins must find a special sequence (called TATA box), 25 bp from the binding site of RNA Polymerase before transcription starts Critical step in protein-dna interactions: protein finding and recognizing the target on DNA
7 Protein-DNA Interactions Experimental Observations: Lac repressor protein finds its target with association rate k ass ~10 10 M -1 s -1 (!!!) Riggler et al., J. Mol. Biol (1970) times larger than the maximal rate by threedimensional diffusion-controlled search (from Debye- Smoluchowski theory) The phenomenon is called Facilitated Diffusion Generated huge controversy still unresolved
8 Diffusion-Controlled Rate Constants Compound B reacts with compound A after approaching it within the radius R Debye-Smoluchowski k max 4 ( D D ) R A B For protein-dna interactions it is assumed that DNA does not move, i.e., D A =0, target size is few bases - R~1 nm, for a typical protein D B ~10-11 m 2 s -1 k max ~10 8 M -1 s -1, compare with k exp ~10 10 M -1 s -1
9 Possible Resolution? Biochem Soc. Trans. 2009, 37,
10 Possible Resolution? NO!!! S.E. Halford, Biochem Soc. Trans. 2009, 37, Original experiments have been carried out in low-salt conditions: 10 mm of KCl, Tris/HCl, and magnesium acetate Debye length separates region where electrostatics is important In this system it is ~1-2 nm, comparable with the target size. Electrostatics does not play role in the fast search!
11 Facilitated Diffusion: Current Theoretical Views Protein search for the target is viewed as a sequence of 3D excursions and 1D sliding along the DNA Acceleration due to increase in the effective target length or lowering dimensionality; assumed that D 1 ~D 3 k max 4 ( D D ) R A B S.E. Halford and J.F. Marko, Nucl. Acids Res. 32, 340 (2004) Experiments support facilitated diffusion picture: D.M. Gowers, G.G. Wilson and S.E. Halford, PNAS USA, 102, (2005)
12 Problems: Current theories predict: D 1 ~D 3 ~10-11 m 2 s -1, and (1D)~(3D) Single-molecule experiments: D 1 ~ m 2 s -1, (1D)/(3D)> Phys. Rev. Lett., 97, (2006);Science, 316, 1191 (2007) c 1 c 1 <c 2 c 2 What is wrong with this picture? S.E. Halford and J.F. Marko, Nucl. Acids Res. 32, 340 (2004)
13 Problems: Current theories predict: D 1 ~D 3 ~10-11 m 2 s -1, and t 1 ~t 3 Single-molecule experiments: D 1 ~ m 2 s -1, t 1 /t 3 >10 Phys. Rev. Lett., 97, (2006);Science, 316, 1191 (2007) c 1 c 1 <c 2 c 2 decreasing the concentration of proteins the reaction of association increases! Contradicts Chemistry! Unphysical behavior S.E. Halford and J.F. Marko, Nucl. Acids Res. 32, 340 (2004)
14 Problems: G. Kolesov et al., PNAS USA, 104, (2007) Search time for transcription factors (TF) proteins from theoretical estimates minutes (!!!), while from experiments ~1min Colocalization mechanism: Genes for TF and for their targets are close to enable fast search However, 1) it does not work for eukaryotes; 2) does not work for proteins with many targets; 3) does not explain invitro experiments
15 Our Goal: To develop a simple model of protein search and recognition for targets on DNA consistent with experimental observations and basic laws of Chemistry and Physics Protein-DNA interactions 2 steps: 1) Finding the target sequence 2) Recognizing the target Successful theoretical picture must account for both stages of protein-dna interactions
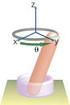
16 Our Theoretical Approach 1) Reaching the target on DNA is viewed as a sequence of searching events (cycles) 2) Each protein on average binds/unbinds several times before finding the target 3) Each cycles consists of 3D and 1D tracks L~1 m length of DNA a~1 nm target size 4) Correlations 5) No assumption of equilibrium
17 Our Theoretical Approach protein One search cycle effective overall 1D motion Mean first-passage time for one searching cycle c x 0 exp[ G( z)] dz D( z) z 0 x 3D segment exp[ G( z' )] dz' D( z) 1D segment position-dependent diffusion constant D3, 0 z x D1, x z z
18 Our Theoretical Approach 0 G(z) search cycle -E eff c x 0 exp[ G( z)] dz D( z) y-equilibrium constant for binding/unbinding z 0 exp[ G( z' )] dz' c p -concentration of proteins in the bulk; c ads adsorbed to DNA x 3D segment y k k 1D segment E ads non-specific binding energy y eff k k on off E exp( k on off c c p ads ads B T ) Eeff exp( k T B )
19 Our Theoretical Approach 0 G(z) search cycle -E eff 3D segment 1D segment Mean first-passage time for one searching cycle time spent by the protein only on 3D segment c 2 x 2 D D 1 time spent by the protein only on 1D segment x x D y 1 eff correlation term reflects forwardbackward trajectories
20 Our Theoretical Approach Consider 1 DNA molecule n ads -number of adsorbed proteins per DNA n ads length of DNA scanned during one cycle Total search time is a sum of one-cycle times c ( L n ads ) L/ n ads average number of cycles before finding the target We assume low concentration of proteins on DNA and no memory in rebinding
21 S j Our Theoretical Approach c ( L n p(1 Average number of search cycles ads p) ) j1 Why average number of search cycles is L/ n ads? p=n ads /L - probability to find a target after binding DNA n ads -average distance scanned by all proteins on DNA in one cycle S j - probability to find the target after j-th cycle j j1 js j 1 p L n ads
22 Our Theoretical Approach V Lr Total search time should be compared with Smoluchowski 3D search time relative search time: S a r 1 n p ( 3/ 2 nads yd nads y d n 2 p 2 yd volume per DNA, r-effective DNA radius a-target size ) c S p n p, V 2 1 c ads Lr D D 3ac p 2 3 n V d=d 1 /D 3 -ratio of diffusion constants ads 2 an y-equilibrium constant p
23 Results relative search time as a function of the adsorption strength a=1nm, r=30nm, n ads =1000, n p =1, d=0.001 Interactions between proteins and DNA depend on ionic strength Biochemistry, 20, 6961 (1981)
24 Results at y<<1 strong repulsion between the protein and DNA, the protein is mainly in 3D motion, not enough time to scan DNA y k k on off exp( E k B ads T ) at y>>1 strong attraction between the protein and DNA, the protein is mainly in 1D motion, which is very slow S a r ( n 1 ads yd n n p 3/ 2 ads y d 2 ) n yd p sliding length is equal to L
25 Results Relative search time as a function of protein concentration Our theory with correlation term a ( r S 1 n n yd n ads p 3/2 ads y d 2 ) n yd p Correlation term is critically important! Old theories without correlation term
26 Results Relative search time as a function of solution protein concentration: Reaching the DNA is a rate-limiting step, protein mostly binds/unbinds S a r ( n 1 ads yd n n p 3/ 2 ads y d 2 ) n yd concentration of proteins becomes so large that it is faster to reach the target only by 3D p
27 Results Relative search time as a function of the ratio of diffusion constants: total time real systems correlation term 1D time 3D time S a r 1 n ( p 3 / n yd n 2 ads ads y d n 2 p yd ) d=d 1 /D 3 -ratio of diffusion constants
28 Results 0 G(z) search cycle -E eff Our theory is nonequilibrium, equilibrium is a special case when E eff =0 x 3D segment 1D segment For realistic values a=1nm, r=30nm, y=10, n p =1,d=0.001 No acceleration in our model S a r y 2 d 1 n d p facilitated diffusion non-equilibrium process
29 Our Theoretical Approach Our view of facilitated diffusion mechanism: 1) Non-equilibrium process Non-specific binding: 1) Slows down 1D diffusion 2) Increases concentration of proteins on DNA 2) Fast 3D and slow 1D motions 3) Correlations are important 4) Acceleration due to increased local concentration of proteins because of non-specific binding
30 Monte Carlo Simulations Relative search time as a function of nonspecific interaction with DNA
31 Monte Carlo Simulations Relative search time as a function of free proteins in solution
32 Our Predictions: 1) Facilitated diffusion mechanism works for intermediate range of adsorption energies. It can be controlled by changing ionic strength Agrees with experiments and computer simulations 2) Facilitated diffusion mechanism works for intermediate concentrations of proteins in the solution 3) (1D)/ (3D)>>1 agrees with
33 Open Questions and Problems: Our theory approximate, improvements are needed 1) Correlations between search cycles 2) Correlations in binding positions after dissociation 3) Protein interactions with DNA that have several targets 4) Protein and DNA elasticity 5) DNA conformations 6) Effect of Temperature 7) Sequence dependence
34 CONCLUSIONS: A theoretical approach of how proteins find and recognize their targets on DNA is developed Facilitated diffusion is a sequence of fast 3D and slow 1D motions Acceleration in facilitated diffusion is achieved due to the increased local concentration of adsorbed proteins (from non-specific binding) Our results and predictions consistent with basic laws of Chemistry and Physics In agreement with all currently available experimental observations
35 Acknowledgements Financial Support: National Science Foundation, The Welch Foundation, Hamill Innovation Fund, Humboldt Fellowship Discussions: Michael E. Fisher, G. Oshanin, A. Berezhkovskii, L. Mirny, B. Shklovskii, A. Grossberg, D. Makarov. Some of the results are published in J. Phys. Chem. B 112, (2008) J. Phys. Chem. B 113, (2009)
The search process for small targets in cellular microdomains! Jürgen Reingruber
 The search process for small targets in cellular microdomains! Jürgen Reingruber Department of Computational Biology Ecole Normale Supérieure Paris, France Outline 1. Search processes in cellular biology
The search process for small targets in cellular microdomains! Jürgen Reingruber Department of Computational Biology Ecole Normale Supérieure Paris, France Outline 1. Search processes in cellular biology
High flexibility of DNA on short length scales probed by atomic force microscopy
 High flexibility of DNA on short length scales probed by atomic force microscopy Wiggins P. A. et al. Nature Nanotechnology (2006) presented by Anja Schwäger 23.01.2008 Outline Theory/Background Elasticity
High flexibility of DNA on short length scales probed by atomic force microscopy Wiggins P. A. et al. Nature Nanotechnology (2006) presented by Anja Schwäger 23.01.2008 Outline Theory/Background Elasticity
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION. Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.
 Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Lecture 1 MODULE 3 GENE EXPRESSION AND REGULATION OF GENE EXPRESSION Professor Bharat Patel Office: Science 2, 2.36 Email: b.patel@griffith.edu.au What is Gene Expression & Gene Regulation? 1. Gene Expression
Gene Models & Bed format: What they represent.
 GeneModels&Bedformat:Whattheyrepresent. Gene models are hypotheses about the structure of transcripts produced by a gene. Like all models, they may be correct, partly correct, or entirely wrong. Typically,
GeneModels&Bedformat:Whattheyrepresent. Gene models are hypotheses about the structure of transcripts produced by a gene. Like all models, they may be correct, partly correct, or entirely wrong. Typically,
Gene Switches Teacher Information
 STO-143 Gene Switches Teacher Information Summary Kit contains How do bacteria turn on and turn off genes? Students model the action of the lac operon that regulates the expression of genes essential for
STO-143 Gene Switches Teacher Information Summary Kit contains How do bacteria turn on and turn off genes? Students model the action of the lac operon that regulates the expression of genes essential for
Lecture 11 Enzymes: Kinetics
 Lecture 11 Enzymes: Kinetics Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 8, pp. 216-225 Key Concepts Kinetics is the study of reaction rates (velocities). Study of enzyme kinetics is useful for
Lecture 11 Enzymes: Kinetics Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 8, pp. 216-225 Key Concepts Kinetics is the study of reaction rates (velocities). Study of enzyme kinetics is useful for
Activity 7.21 Transcription factors
 Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
Purpose To consolidate understanding of protein synthesis. To explain the role of transcription factors and hormones in switching genes on and off. Play the transcription initiation complex game Regulation
Protein Synthesis How Genes Become Constituent Molecules
 Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Protein Synthesis Protein Synthesis How Genes Become Constituent Molecules Mendel and The Idea of Gene What is a Chromosome? A chromosome is a molecule of DNA 50% 50% 1. True 2. False True False Protein
Molecular Dynamics Simulations
 Molecular Dynamics Simulations Yaoquan Tu Division of Theoretical Chemistry and Biology, Royal Institute of Technology (KTH) 2011-06 1 Outline I. Introduction II. Molecular Mechanics Force Field III. Molecular
Molecular Dynamics Simulations Yaoquan Tu Division of Theoretical Chemistry and Biology, Royal Institute of Technology (KTH) 2011-06 1 Outline I. Introduction II. Molecular Mechanics Force Field III. Molecular
Viscoelasticity of Polymer Fluids.
 Viscoelasticity of Polymer Fluids. Main Properties of Polymer Fluids. Entangled polymer fluids are polymer melts and concentrated or semidilute (above the concentration c) solutions. In these systems polymer
Viscoelasticity of Polymer Fluids. Main Properties of Polymer Fluids. Entangled polymer fluids are polymer melts and concentrated or semidilute (above the concentration c) solutions. In these systems polymer
Name Class Date. Figure 13 1. 2. Which nucleotide in Figure 13 1 indicates the nucleic acid above is RNA? a. uracil c. cytosine b. guanine d.
 13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13 Multiple Choice RNA and Protein Synthesis Chapter Test A Write the letter that best answers the question or completes the statement on the line provided. 1. Which of the following are found in both
13.4 Gene Regulation and Expression
 13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
Gene Transcription in Prokaryotes
 Gene Transcription in Prokaryotes Operons: in prokaryotes, genes that encode protein participating in a common pathway are organized together. This group of genes, arranged in tandem, is called an OPERON.
Gene Transcription in Prokaryotes Operons: in prokaryotes, genes that encode protein participating in a common pathway are organized together. This group of genes, arranged in tandem, is called an OPERON.
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
Chapter 18 Regulation of Gene Expression 18.1. Gene Regulation Is Necessary By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection
DNA Insertions and Deletions in the Human Genome. Philipp W. Messer
 DNA Insertions and Deletions in the Human Genome Philipp W. Messer Genetic Variation CGACAATAGCGCTCTTACTACGTGTATCG : : CGACAATGGCGCT---ACTACGTGCATCG 1. Nucleotide mutations 2. Genomic rearrangements 3.
DNA Insertions and Deletions in the Human Genome Philipp W. Messer Genetic Variation CGACAATAGCGCTCTTACTACGTGTATCG : : CGACAATGGCGCT---ACTACGTGCATCG 1. Nucleotide mutations 2. Genomic rearrangements 3.
FIDA for Rapid Detection of Protein Based Biomarkers
 FIDA for Rapid Detection of Protein Based Biomarkers Guisheng Zhuang 1, Nicklas N. Poulsen 1, Nina Z. Andersen 1, Jesper Østergaard 1,2, Jørgen Schøller 2 and Henrik Jensen 1,2 1 Department of Pharmacy,
FIDA for Rapid Detection of Protein Based Biomarkers Guisheng Zhuang 1, Nicklas N. Poulsen 1, Nina Z. Andersen 1, Jesper Østergaard 1,2, Jørgen Schøller 2 and Henrik Jensen 1,2 1 Department of Pharmacy,
Dynamics of Biological Systems
 Dynamics of Biological Systems Part I - Biological background and mathematical modelling Paolo Milazzo (Università di Pisa) Dynamics of biological systems 1 / 53 Introduction The recent developments in
Dynamics of Biological Systems Part I - Biological background and mathematical modelling Paolo Milazzo (Università di Pisa) Dynamics of biological systems 1 / 53 Introduction The recent developments in
For additional information on the program, see the current university catalog.
 For information call: Tel: (818) 77-81 Fax: (818) 77-08 E-mail: chemistry.office@csun.edu Website: http://www.csun.edu/chemistry Or write: Department of Chemistry and Biochemistry California State University,
For information call: Tel: (818) 77-81 Fax: (818) 77-08 E-mail: chemistry.office@csun.edu Website: http://www.csun.edu/chemistry Or write: Department of Chemistry and Biochemistry California State University,
A Mathematical Model of a Synthetically Constructed Genetic Toggle Switch
 BENG 221 Mathematical Methods in Bioengineering Project Report A Mathematical Model of a Synthetically Constructed Genetic Toggle Switch Nick Csicsery & Ricky O Laughlin October 15, 2013 1 TABLE OF CONTENTS
BENG 221 Mathematical Methods in Bioengineering Project Report A Mathematical Model of a Synthetically Constructed Genetic Toggle Switch Nick Csicsery & Ricky O Laughlin October 15, 2013 1 TABLE OF CONTENTS
Transcription in prokaryotes. Elongation and termination
 Transcription in prokaryotes Elongation and termination After initiation the σ factor leaves the scene. Core polymerase is conducting the elongation of the chain. The core polymerase contains main nucleotide
Transcription in prokaryotes Elongation and termination After initiation the σ factor leaves the scene. Core polymerase is conducting the elongation of the chain. The core polymerase contains main nucleotide
Calculating Nucleic Acid or Protein Concentration Using the GloMax Multi+ Microplate Instrument
 Calculating Nucleic Acid or Protein Concentration Using the GloMax Multi+ Microplate Instrument Technical Note INTRODUCTION Direct measurements of nucleic acid samples at OD 260 or protein samples at OD
Calculating Nucleic Acid or Protein Concentration Using the GloMax Multi+ Microplate Instrument Technical Note INTRODUCTION Direct measurements of nucleic acid samples at OD 260 or protein samples at OD
Control of Gene Expression
 Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Gene Regulation Is Necessary? Control of Gene Expression By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
CHAPTER 6 AN INTRODUCTION TO METABOLISM. Section B: Enzymes
 CHAPTER 6 AN INTRODUCTION TO METABOLISM Section B: Enzymes 1. Enzymes speed up metabolic reactions by lowering energy barriers 2. Enzymes are substrate specific 3. The active site in an enzyme s catalytic
CHAPTER 6 AN INTRODUCTION TO METABOLISM Section B: Enzymes 1. Enzymes speed up metabolic reactions by lowering energy barriers 2. Enzymes are substrate specific 3. The active site in an enzyme s catalytic
Additional Modeling Information
 1 Additional Modeling Information Temperature Control of Fimbriation Circuit Switch in Uropathogenic Escherichia coli: Quantitative Analysis via Automated Model Abstraction Hiroyuki Kuwahara, Chris J.
1 Additional Modeling Information Temperature Control of Fimbriation Circuit Switch in Uropathogenic Escherichia coli: Quantitative Analysis via Automated Model Abstraction Hiroyuki Kuwahara, Chris J.
Introduction, Noncovalent Bonds, and Properties of Water
 Lecture 1 Introduction, Noncovalent Bonds, and Properties of Water Reading: Berg, Tymoczko & Stryer: Chapter 1 problems in textbook: chapter 1, pp. 23-24, #1,2,3,6,7,8,9, 10,11; practice problems at end
Lecture 1 Introduction, Noncovalent Bonds, and Properties of Water Reading: Berg, Tymoczko & Stryer: Chapter 1 problems in textbook: chapter 1, pp. 23-24, #1,2,3,6,7,8,9, 10,11; practice problems at end
Gene Regulation -- The Lac Operon
 Gene Regulation -- The Lac Operon Specific proteins are present in different tissues and some appear only at certain times during development. All cells of a higher organism have the full set of genes:
Gene Regulation -- The Lac Operon Specific proteins are present in different tissues and some appear only at certain times during development. All cells of a higher organism have the full set of genes:
European Medicines Agency
 European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
European Medicines Agency July 1996 CPMP/ICH/139/95 ICH Topic Q 5 B Quality of Biotechnological Products: Analysis of the Expression Construct in Cell Lines Used for Production of r-dna Derived Protein
Specific problems. The genetic code. The genetic code. Adaptor molecules match amino acids to mrna codons
 Tutorial II Gene expression: mrna translation and protein synthesis Piergiorgio Percipalle, PhD Program Control of gene transcription and RNA processing mrna translation and protein synthesis KAROLINSKA
Tutorial II Gene expression: mrna translation and protein synthesis Piergiorgio Percipalle, PhD Program Control of gene transcription and RNA processing mrna translation and protein synthesis KAROLINSKA
Module 3 Questions. 7. Chemotaxis is an example of signal transduction. Explain, with the use of diagrams.
 Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Module 3 Questions Section 1. Essay and Short Answers. Use diagrams wherever possible 1. With the use of a diagram, provide an overview of the general regulation strategies available to a bacterial cell.
Complex multicellular organisms are produced by cells that switch genes on and off during development.
 Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
Home Control of Gene Expression Gene Regulation Is Necessary? By switching genes off when they are not needed, cells can prevent resources from being wasted. There should be natural selection favoring
How To Understand How Gene Expression Is Regulated
 What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
What makes cells different from each other? How do cells respond to information from environment? Regulation of: - Transcription - prokaryotes - eukaryotes - mrna splicing - mrna localisation and translation
BCH401G Lecture 39 Andres
 BCH401G Lecture 39 Andres Lecture Summary: Ribosome: Understand its role in translation and differences between translation in prokaryotes and eukaryotes. Translation: Understand the chemistry of this
BCH401G Lecture 39 Andres Lecture Summary: Ribosome: Understand its role in translation and differences between translation in prokaryotes and eukaryotes. Translation: Understand the chemistry of this
CHEMISTRY STANDARDS BASED RUBRIC ATOMIC STRUCTURE AND BONDING
 CHEMISTRY STANDARDS BASED RUBRIC ATOMIC STRUCTURE AND BONDING Essential Standard: STUDENTS WILL UNDERSTAND THAT THE PROPERTIES OF MATTER AND THEIR INTERACTIONS ARE A CONSEQUENCE OF THE STRUCTURE OF MATTER,
CHEMISTRY STANDARDS BASED RUBRIC ATOMIC STRUCTURE AND BONDING Essential Standard: STUDENTS WILL UNDERSTAND THAT THE PROPERTIES OF MATTER AND THEIR INTERACTIONS ARE A CONSEQUENCE OF THE STRUCTURE OF MATTER,
Name Date Period. 2. When a molecule of double-stranded DNA undergoes replication, it results in
 DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
DNA, RNA, Protein Synthesis Keystone 1. During the process shown above, the two strands of one DNA molecule are unwound. Then, DNA polymerases add complementary nucleotides to each strand which results
THERMODYNAMIC PROPERTIES OF POLYPEPTIDE CHAINS. PARALLEL TEMPERING MONTE CARLO SIMULATIONS
 Vol. 38 (2007) ACTA PHYSICA POLONICA B No 5 THERMODYNAMIC PROPERTIES OF POLYPEPTIDE CHAINS. PARALLEL TEMPERING MONTE CARLO SIMULATIONS Andrzej Sikorski, Dominik Gront Department of Chemistry, University
Vol. 38 (2007) ACTA PHYSICA POLONICA B No 5 THERMODYNAMIC PROPERTIES OF POLYPEPTIDE CHAINS. PARALLEL TEMPERING MONTE CARLO SIMULATIONS Andrzej Sikorski, Dominik Gront Department of Chemistry, University
Enzymes: Practice Questions #1
 Enzymes: Practice Questions #1 1. Compound X increases the rate of the reaction below. Compound X is most likely A. an enzyme B. a lipid molecule C. an indicator D. an ADP molecule 2. The equation below
Enzymes: Practice Questions #1 1. Compound X increases the rate of the reaction below. Compound X is most likely A. an enzyme B. a lipid molecule C. an indicator D. an ADP molecule 2. The equation below
Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism )
 Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
Biology 1406 Exam 3 Notes Structure of DNA Ch. 10 Genetic information (DNA) determines structure of proteins DNA RNA proteins cell structure 3.11 3.15 enzymes control cell chemistry ( metabolism ) Proteins
RNA Structure and folding
 RNA Structure and folding Overview: The main functional biomolecules in cells are polymers DNA, RNA and proteins For RNA and Proteins, the specific sequence of the polymer dictates its final structure
RNA Structure and folding Overview: The main functional biomolecules in cells are polymers DNA, RNA and proteins For RNA and Proteins, the specific sequence of the polymer dictates its final structure
MATCH Commun. Math. Comput. Chem. 61 (2009) 781-788
 MATCH Communications in Mathematical and in Computer Chemistry MATCH Commun. Math. Comput. Chem. 61 (2009) 781-788 ISSN 0340-6253 Three distances for rapid similarity analysis of DNA sequences Wei Chen,
MATCH Communications in Mathematical and in Computer Chemistry MATCH Commun. Math. Comput. Chem. 61 (2009) 781-788 ISSN 0340-6253 Three distances for rapid similarity analysis of DNA sequences Wei Chen,
Chem 465 Biochemistry II
 Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Formation of the ribosomal initiation complex for bacterial protein synthesis does not require: A) EF-Tu. B) formylmethionyl
Chem 465 Biochemistry II Name: 2 points Multiple choice (4 points apiece): 1. Formation of the ribosomal initiation complex for bacterial protein synthesis does not require: A) EF-Tu. B) formylmethionyl
AP CHEMISTRY 2007 SCORING GUIDELINES. Question 6
 AP CHEMISTRY 2007 SCORING GUIDELINES Question 6 Answer the following questions, which pertain to binary compounds. (a) In the box provided below, draw a complete Lewis electron-dot diagram for the IF 3
AP CHEMISTRY 2007 SCORING GUIDELINES Question 6 Answer the following questions, which pertain to binary compounds. (a) In the box provided below, draw a complete Lewis electron-dot diagram for the IF 3
Incorporating Internal Gradient and Restricted Diffusion Effects in Nuclear Magnetic Resonance Log Interpretation
 The Open-Access Journal for the Basic Principles of Diffusion Theory, Experiment and Application Incorporating Internal Gradient and Restricted Diffusion Effects in Nuclear Magnetic Resonance Log Interpretation
The Open-Access Journal for the Basic Principles of Diffusion Theory, Experiment and Application Incorporating Internal Gradient and Restricted Diffusion Effects in Nuclear Magnetic Resonance Log Interpretation
CSC 2427: Algorithms for Molecular Biology Spring 2006. Lecture 16 March 10
 CSC 2427: Algorithms for Molecular Biology Spring 2006 Lecture 16 March 10 Lecturer: Michael Brudno Scribe: Jim Huang 16.1 Overview of proteins Proteins are long chains of amino acids (AA) which are produced
CSC 2427: Algorithms for Molecular Biology Spring 2006 Lecture 16 March 10 Lecturer: Michael Brudno Scribe: Jim Huang 16.1 Overview of proteins Proteins are long chains of amino acids (AA) which are produced
Models of Switching in Biophysical Contexts
 Models of Switching in Biophysical Contexts Martin R. Evans SUPA, School of Physics and Astronomy, University of Edinburgh, U.K. March 7, 2011 Collaborators: Paolo Visco (MSC, Paris) Rosalind J. Allen
Models of Switching in Biophysical Contexts Martin R. Evans SUPA, School of Physics and Astronomy, University of Edinburgh, U.K. March 7, 2011 Collaborators: Paolo Visco (MSC, Paris) Rosalind J. Allen
Statistical mechanics for real biological networks
 Statistical mechanics for real biological networks William Bialek Joseph Henry Laboratories of Physics, and Lewis-Sigler Institute for Integrative Genomics Princeton University Initiative for the Theoretical
Statistical mechanics for real biological networks William Bialek Joseph Henry Laboratories of Physics, and Lewis-Sigler Institute for Integrative Genomics Princeton University Initiative for the Theoretical
Algorithms in Computational Biology (236522) spring 2007 Lecture #1
 Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
Algorithms in Computational Biology (236522) spring 2007 Lecture #1 Lecturer: Shlomo Moran, Taub 639, tel 4363 Office hours: Tuesday 11:00-12:00/by appointment TA: Ilan Gronau, Taub 700, tel 4894 Office
Energy & Enzymes. Life requires energy for maintenance of order, growth, and reproduction. The energy living things use is chemical energy.
 Energy & Enzymes Life requires energy for maintenance of order, growth, and reproduction. The energy living things use is chemical energy. 1 Energy exists in two forms - potential and kinetic. Potential
Energy & Enzymes Life requires energy for maintenance of order, growth, and reproduction. The energy living things use is chemical energy. 1 Energy exists in two forms - potential and kinetic. Potential
??? Central Dogma: DNA is a Read-Only-Memory.
 京 都 大 学 吉 川 研 一 At 桂 林 ; June 19, 2007 Novel Hypothesis on the Self-Regulation of Genetic Information: Scenario from One-Dimensional DNA into Spatio-Temporal Order Conclusion: The concept of Protein network
京 都 大 学 吉 川 研 一 At 桂 林 ; June 19, 2007 Novel Hypothesis on the Self-Regulation of Genetic Information: Scenario from One-Dimensional DNA into Spatio-Temporal Order Conclusion: The concept of Protein network
Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Biochemistry
 Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Biochemistry The Master Degree in Medical Laboratory Sciences /Clinical Biochemistry, is awarded by the Faculty of Graduate Studies
Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Biochemistry The Master Degree in Medical Laboratory Sciences /Clinical Biochemistry, is awarded by the Faculty of Graduate Studies
Viral Infection: Receptors
 Viral Infection: Receptors Receptors: Identification of receptors has come from expressing the gene for the receptor in a cell to which a virus does not normally bind -OR- By blocking virus attachment
Viral Infection: Receptors Receptors: Identification of receptors has come from expressing the gene for the receptor in a cell to which a virus does not normally bind -OR- By blocking virus attachment
Molecular Genetics. RNA, Transcription, & Protein Synthesis
 Molecular Genetics RNA, Transcription, & Protein Synthesis Section 1 RNA AND TRANSCRIPTION Objectives Describe the primary functions of RNA Identify how RNA differs from DNA Describe the structure and
Molecular Genetics RNA, Transcription, & Protein Synthesis Section 1 RNA AND TRANSCRIPTION Objectives Describe the primary functions of RNA Identify how RNA differs from DNA Describe the structure and
Stochastic Gene Expression in Prokaryotes: A Point Process Approach
 Stochastic Gene Expression in Prokaryotes: A Point Process Approach Emanuele LEONCINI INRIA Rocquencourt - INRA Jouy-en-Josas Mathematical Modeling in Cell Biology March 27 th 2013 Emanuele LEONCINI (INRIA)
Stochastic Gene Expression in Prokaryotes: A Point Process Approach Emanuele LEONCINI INRIA Rocquencourt - INRA Jouy-en-Josas Mathematical Modeling in Cell Biology March 27 th 2013 Emanuele LEONCINI (INRIA)
Effect of the Protein Denaturants Urea and Guanidinium on Water Structure: A Structural and Thermodynamic Study
 10748 J. Am. Chem. Soc. 1998, 120, 10748-10753 Effect of the Protein Denaturants Urea and Guanidinium on Water Structure: A Structural and Thermodynamic Study Francesco Vanzi, Bhupinder Madan, and Kim
10748 J. Am. Chem. Soc. 1998, 120, 10748-10753 Effect of the Protein Denaturants Urea and Guanidinium on Water Structure: A Structural and Thermodynamic Study Francesco Vanzi, Bhupinder Madan, and Kim
Lecture 3: Enzyme kinetics
 Computational Systems Biology Lecture 3: Enzyme kinetics Fri 19 Jan 2009 1 Images from: D. L. Nelson, Lehninger Principles of Biochemistry, IV Edition, W. H. Freeman ed. A. Cornish-Bowden Fundamentals
Computational Systems Biology Lecture 3: Enzyme kinetics Fri 19 Jan 2009 1 Images from: D. L. Nelson, Lehninger Principles of Biochemistry, IV Edition, W. H. Freeman ed. A. Cornish-Bowden Fundamentals
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
1 Mutation and Genetic Change
 CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
CHAPTER 14 1 Mutation and Genetic Change SECTION Genes in Action KEY IDEAS As you read this section, keep these questions in mind: What is the origin of genetic differences among organisms? What kinds
GENE REGULATION. Teacher Packet
 AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
Jeffrey R. Vieregg. Phone: (773) 702-2556 Mobile: (818) 455-2930 jvieregg@uchicago.edu. home.uchicago.edu/~jvieregg
 Senior Research Scientist Institute for Molecular Engineering University of Chicago, Jones 222 5757 South Ellis Avenue Chicago, IL 60637 USA Phone: (773) 702-2556 Mobile: (818) 455-2930 jvieregg@uchicago.edu
Senior Research Scientist Institute for Molecular Engineering University of Chicago, Jones 222 5757 South Ellis Avenue Chicago, IL 60637 USA Phone: (773) 702-2556 Mobile: (818) 455-2930 jvieregg@uchicago.edu
Chapter 2. The Chemistry of Life Worksheets
 Chapter 2 The Chemistry of Life Worksheets (Opening image courtesy of David Iberri, http://en.wikipedia.org/wiki/file:camkii.png, and under the Creative Commons license CC-BY-SA 3.0.) Lesson 2.1: Matter
Chapter 2 The Chemistry of Life Worksheets (Opening image courtesy of David Iberri, http://en.wikipedia.org/wiki/file:camkii.png, and under the Creative Commons license CC-BY-SA 3.0.) Lesson 2.1: Matter
Biochemistry 1 Course Specifications. First year of M.B.B.Ch. Program
 Faculty of Medicine Quality Assurance Unit Al-Azhar University Assuit Faculty of Medicine Biochemistry 1 Course Specifications First year of M.B.B.Ch. Program A- Professional information: Title: Biochemistry1
Faculty of Medicine Quality Assurance Unit Al-Azhar University Assuit Faculty of Medicine Biochemistry 1 Course Specifications First year of M.B.B.Ch. Program A- Professional information: Title: Biochemistry1
20.309: Biological Instrumentation and Measurement. Heejin Choi Rumi Chunara Yuri Matsumoto
 20.309: Biological Instrumentation and Measurement Instructors: Laboratory Instructor: Teaching Assistants: Scott Manalis and Peter So Steve Wasserman Jaewon Cha Heejin Choi Rumi Chunara Yuri Matsumoto
20.309: Biological Instrumentation and Measurement Instructors: Laboratory Instructor: Teaching Assistants: Scott Manalis and Peter So Steve Wasserman Jaewon Cha Heejin Choi Rumi Chunara Yuri Matsumoto
CCR Biology - Chapter 8 Practice Test - Summer 2012
 Name: Class: Date: CCR Biology - Chapter 8 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. What did Hershey and Chase know
Name: Class: Date: CCR Biology - Chapter 8 Practice Test - Summer 2012 Multiple Choice Identify the choice that best completes the statement or answers the question. 1. What did Hershey and Chase know
CHM333 LECTURE 13 14: 2/13 15/13 SPRING 2013 Professor Christine Hrycyna
 INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
Molecular Spectroscopy
 Molecular Spectroscopy UV-Vis Spectroscopy Absorption Characteristics of Some Common Chromophores UV-Vis Spectroscopy Absorption Characteristics of Aromatic Compounds UV-Vis Spectroscopy Effect of extended
Molecular Spectroscopy UV-Vis Spectroscopy Absorption Characteristics of Some Common Chromophores UV-Vis Spectroscopy Absorption Characteristics of Aromatic Compounds UV-Vis Spectroscopy Effect of extended
Catalysis by Enzymes. Enzyme A protein that acts as a catalyst for a biochemical reaction.
 Catalysis by Enzymes Enzyme A protein that acts as a catalyst for a biochemical reaction. Enzymatic Reaction Specificity Enzyme Cofactors Many enzymes are conjugated proteins that require nonprotein portions
Catalysis by Enzymes Enzyme A protein that acts as a catalyst for a biochemical reaction. Enzymatic Reaction Specificity Enzyme Cofactors Many enzymes are conjugated proteins that require nonprotein portions
Transcription and Translation of DNA
 Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
Transcription and Translation of DNA Genotype our genetic constitution ( makeup) is determined (controlled) by the sequence of bases in its genes Phenotype determined by the proteins synthesised when genes
CHAPTER 4: Enzyme Structure ENZYMES
 CHAPTER 4: ENZYMES Enzymes are biological catalysts. There are about 40,000 different enzymes in human cells, each controlling a different chemical reaction. They increase the rate of reactions by a factor
CHAPTER 4: ENZYMES Enzymes are biological catalysts. There are about 40,000 different enzymes in human cells, each controlling a different chemical reaction. They increase the rate of reactions by a factor
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual
 Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
M.Sc. in Nano Technology with specialisation in Nano Biotechnology
 M.Sc. in Nano Technology with specialisation in Nano Biotechnology Nanotechnology is all about designing, fabricating and controlling materials, components and machinery with dimensions on the nanoscale,
M.Sc. in Nano Technology with specialisation in Nano Biotechnology Nanotechnology is all about designing, fabricating and controlling materials, components and machinery with dimensions on the nanoscale,
green B 1 ) into a single unit to model the substrate in this reaction. enzyme
 Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
Name: Date: Period: DNA Unit: DNA Webquest
 Name: Date: Period: DNA Unit: DNA Webquest Part 1 History, DNA Structure, DNA Replication DNA History http://www.dnaftb.org/dnaftb/1/concept/index.html Read the text and answer the following questions.
Name: Date: Period: DNA Unit: DNA Webquest Part 1 History, DNA Structure, DNA Replication DNA History http://www.dnaftb.org/dnaftb/1/concept/index.html Read the text and answer the following questions.
CHM333 LECTURE 13 14: 2/13 15/12 SPRING 2012 Professor Christine Hrycyna
 INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
Protein Dynamics Intro
 Protein Dynamics Intro From rigid structures to motions on energy landscapes Do you all remember Anfinsen? What concept now associated with his name made Anfinsen famous? Right, it is the concept that
Protein Dynamics Intro From rigid structures to motions on energy landscapes Do you all remember Anfinsen? What concept now associated with his name made Anfinsen famous? Right, it is the concept that
1.1.2. thebiotutor. AS Biology OCR. Unit F211: Cells, Exchange & Transport. Module 1.2 Cell Membranes. Notes & Questions.
 thebiotutor AS Biology OCR Unit F211: Cells, Exchange & Transport Module 1.2 Cell Membranes Notes & Questions Andy Todd 1 Outline the roles of membranes within cells and at the surface of cells. The main
thebiotutor AS Biology OCR Unit F211: Cells, Exchange & Transport Module 1.2 Cell Membranes Notes & Questions Andy Todd 1 Outline the roles of membranes within cells and at the surface of cells. The main
Highly Selective Cyclic Peptide Ligands for NeutrAvidin and Avidin Identified by Phage Display
 Highly Selective Cyclic Peptide Ligands for NeutrAvidin and Avidin Identified by Phage Display Supplementary Material Scott C. Meyer, Thomas Gaj, and Indraneel Ghosh Department of Chemistry, University
Highly Selective Cyclic Peptide Ligands for NeutrAvidin and Avidin Identified by Phage Display Supplementary Material Scott C. Meyer, Thomas Gaj, and Indraneel Ghosh Department of Chemistry, University
Biacore X BIACORE. The versatile high sensitivity system
 Biacore X The versatile high sensitivity system Work with high sensitivity Study binding in non-aqueous and aqueous samples Reduce sample consumption Perform the most advanced kinetic evaluation Use proven
Biacore X The versatile high sensitivity system Work with high sensitivity Study binding in non-aqueous and aqueous samples Reduce sample consumption Perform the most advanced kinetic evaluation Use proven
MORPHEUS. http://biodev.cea.fr/morpheus/ Prediction of Transcription Factors Binding Sites based on Position Weight Matrix.
 MORPHEUS http://biodev.cea.fr/morpheus/ Prediction of Transcription Factors Binding Sites based on Position Weight Matrix. Reference: MORPHEUS, a Webtool for Transcripton Factor Binding Analysis Using
MORPHEUS http://biodev.cea.fr/morpheus/ Prediction of Transcription Factors Binding Sites based on Position Weight Matrix. Reference: MORPHEUS, a Webtool for Transcripton Factor Binding Analysis Using
Multiple Choice Write the letter that best answers the question or completes the statement on the line provided.
 Name lass Date hapter 12 DN and RN hapter Test Multiple hoice Write the letter that best answers the question or completes the statement on the line provided. Pearson Education, Inc. ll rights reserved.
Name lass Date hapter 12 DN and RN hapter Test Multiple hoice Write the letter that best answers the question or completes the statement on the line provided. Pearson Education, Inc. ll rights reserved.
Assessment Method 1: Oral seminar presentation
 2013-2014 Assessment Report College of Sciences & Mathematics Chemistry & Biochemistry Chemistry, Master's Expected Outcome 1: Effective Oral Communication Skills Students in M.S. degree Program will demonstrate
2013-2014 Assessment Report College of Sciences & Mathematics Chemistry & Biochemistry Chemistry, Master's Expected Outcome 1: Effective Oral Communication Skills Students in M.S. degree Program will demonstrate
Mathematical Modeling and Simulations Supporting Medicine and Biology
 Mathematical Modeling and Simulations Supporting Medicine and Biology Luigi Preziosi Politecnico di Torino calvino.polito.it/~preziosi Mathematical Modeling and Simulations Supporting Medicine and Biology
Mathematical Modeling and Simulations Supporting Medicine and Biology Luigi Preziosi Politecnico di Torino calvino.polito.it/~preziosi Mathematical Modeling and Simulations Supporting Medicine and Biology
Unfolding and Aggregation of mabs Application Note NT-PR-005
 Unfolding and Aggregation of mabs Application Note NT-PR-005 Analysis of formulation-dependent colloidal and conformational stability of monoclonal antibodies Franziska Söltl 1, Jonathan Derix 1, Michaela
Unfolding and Aggregation of mabs Application Note NT-PR-005 Analysis of formulation-dependent colloidal and conformational stability of monoclonal antibodies Franziska Söltl 1, Jonathan Derix 1, Michaela
Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Microbiology, Immunology and Serology
 Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Microbiology, Immunology and Serology The Master Degree in Medical Laboratory Sciences / Clinical Microbiology, Immunology or
Course Curriculum for Master Degree in Medical Laboratory Sciences/Clinical Microbiology, Immunology and Serology The Master Degree in Medical Laboratory Sciences / Clinical Microbiology, Immunology or
Replication Study Guide
 Replication Study Guide This study guide is a written version of the material you have seen presented in the replication unit. Self-reproduction is a function of life that human-engineered systems have
Replication Study Guide This study guide is a written version of the material you have seen presented in the replication unit. Self-reproduction is a function of life that human-engineered systems have
Effects of Antibiotics on Bacterial Growth and Protein Synthesis: Student Laboratory Manual
 Effects of Antibiotics on Bacterial Growth and Protein Synthesis: Student Laboratory Manual I. Purpose...1 II. Introduction...1 III. Inhibition of Bacterial Growth Protocol...2 IV. Inhibition of in vitro
Effects of Antibiotics on Bacterial Growth and Protein Synthesis: Student Laboratory Manual I. Purpose...1 II. Introduction...1 III. Inhibition of Bacterial Growth Protocol...2 IV. Inhibition of in vitro
COMPUTATIONAL FRAMEWORKS FOR UNDERSTANDING THE FUNCTION AND EVOLUTION OF DEVELOPMENTAL ENHANCERS IN DROSOPHILA
 COMPUTATIONAL FRAMEWORKS FOR UNDERSTANDING THE FUNCTION AND EVOLUTION OF DEVELOPMENTAL ENHANCERS IN DROSOPHILA Saurabh Sinha, Dept of Computer Science, University of Illinois Cis-regulatory modules (enhancers)
COMPUTATIONAL FRAMEWORKS FOR UNDERSTANDING THE FUNCTION AND EVOLUTION OF DEVELOPMENTAL ENHANCERS IN DROSOPHILA Saurabh Sinha, Dept of Computer Science, University of Illinois Cis-regulatory modules (enhancers)
BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS
 BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS 1. The Technology Strategy sets out six areas where technological developments are required to push the frontiers of knowledge
BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS 1. The Technology Strategy sets out six areas where technological developments are required to push the frontiers of knowledge
Syllabus for MCB 3010/5001: Biochemistry Fall Semester 2011
 Syllabus for MCB 3010/5001: Biochemistry Fall Semester 2011 Instructor: Dr. Wolf-Dieter Reiter Office: TLS 406 Phone: 486-5733 E-mail: wdreiter@uconn.edu Office hours: Wednesday, 11:00 12:00 a.m., Thursday,
Syllabus for MCB 3010/5001: Biochemistry Fall Semester 2011 Instructor: Dr. Wolf-Dieter Reiter Office: TLS 406 Phone: 486-5733 E-mail: wdreiter@uconn.edu Office hours: Wednesday, 11:00 12:00 a.m., Thursday,
Genetics Lecture Notes 7.03 2005. Lectures 1 2
 Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Genetics Lecture Notes 7.03 2005 Lectures 1 2 Lecture 1 We will begin this course with the question: What is a gene? This question will take us four lectures to answer because there are actually several
Fuzzy Modeling of Labeled Point Cloud Superposition for the Comparison of Protein Binding Sites
 Fuzzy Modeling of Labeled Point Cloud Superposition for the Comparison of Protein Binding Sites Thomas Fober Eyke Hüllermeier Knowledge Engineering & Bioinformatics Group Mathematics and Computer Science
Fuzzy Modeling of Labeled Point Cloud Superposition for the Comparison of Protein Binding Sites Thomas Fober Eyke Hüllermeier Knowledge Engineering & Bioinformatics Group Mathematics and Computer Science
Topic 2: Energy in Biological Systems
 Topic 2: Energy in Biological Systems Outline: Types of energy inside cells Heat & Free Energy Energy and Equilibrium An Introduction to Entropy Types of energy in cells and the cost to build the parts
Topic 2: Energy in Biological Systems Outline: Types of energy inside cells Heat & Free Energy Energy and Equilibrium An Introduction to Entropy Types of energy in cells and the cost to build the parts
Speed Matters - Fast ways from template to result
 qpcr Symposium 2007 - Weihenstephan Speed Matters - Fast ways from template to result March 28, 2007 Dr. Thorsten Traeger Senior Scientist, Research and Development - 1 - Overview Ạgenda Fast PCR The Challenges
qpcr Symposium 2007 - Weihenstephan Speed Matters - Fast ways from template to result March 28, 2007 Dr. Thorsten Traeger Senior Scientist, Research and Development - 1 - Overview Ạgenda Fast PCR The Challenges
What is the difference between basal and activated transcription?
 What is the difference between basal and activated transcription? Regulation of Transcription I. Basal vs. activated transcription for mrna genes A. General transcription factor (TF) vs. promoterspecific
What is the difference between basal and activated transcription? Regulation of Transcription I. Basal vs. activated transcription for mrna genes A. General transcription factor (TF) vs. promoterspecific
Algorithm and computational complexity of Insulin
 Algorithm and computational complexity Insulin Lutvo Kurić Bosnia and Herzegovina, Novi Travnik, Kalinska 7 Abstract:This paper discusses cyberinformation studies the amino acid composition insulin, in
Algorithm and computational complexity Insulin Lutvo Kurić Bosnia and Herzegovina, Novi Travnik, Kalinska 7 Abstract:This paper discusses cyberinformation studies the amino acid composition insulin, in
Chapter 6: Biological Networks
 Chapter 6: Biological Networks 6.4 Engineering Synthetic Networks Prof. Yechiam Yemini (YY) Computer Science Department Columbia University Overview Constructing regulatory gates A genetic toggle switch;
Chapter 6: Biological Networks 6.4 Engineering Synthetic Networks Prof. Yechiam Yemini (YY) Computer Science Department Columbia University Overview Constructing regulatory gates A genetic toggle switch;
Advanced Medicinal & Pharmaceutical Chemistry CHEM 5412 Dept. of Chemistry, TAMUK
 Advanced Medicinal & Pharmaceutical Chemistry CHEM 5412 Dept. of Chemistry, TAMUK Dai Lu, Ph.D. dlu@tamhsc.edu Tel: 361-221-0745 Office: RCOP, Room 307 Drug Discovery and Development Drug Molecules Medicinal
Advanced Medicinal & Pharmaceutical Chemistry CHEM 5412 Dept. of Chemistry, TAMUK Dai Lu, Ph.D. dlu@tamhsc.edu Tel: 361-221-0745 Office: RCOP, Room 307 Drug Discovery and Development Drug Molecules Medicinal
GenBank, Entrez, & FASTA
 GenBank, Entrez, & FASTA Nucleotide Sequence Databases First generation GenBank is a representative example started as sort of a museum to preserve knowledge of a sequence from first discovery great repositories,
GenBank, Entrez, & FASTA Nucleotide Sequence Databases First generation GenBank is a representative example started as sort of a museum to preserve knowledge of a sequence from first discovery great repositories,
Modeling and Simulation of Gene Regulatory Networks
 Modeling and Simulation of Gene Regulatory Networks Hidde de Jong INRIA Grenoble - Rhône-Alpes Hidde.de-Jong@inria.fr http://ibis.inrialpes.fr INRIA Grenoble - Rhône-Alpes and IBIS IBIS: systems biology
Modeling and Simulation of Gene Regulatory Networks Hidde de Jong INRIA Grenoble - Rhône-Alpes Hidde.de-Jong@inria.fr http://ibis.inrialpes.fr INRIA Grenoble - Rhône-Alpes and IBIS IBIS: systems biology
Application Note. Separation of three monoclonal antibody variants using MCSGP. Summary
 Application Note Separation of three monoclonal antibody variants using MCSGP Category Matrix Method Keywords Analytes ID Continuous chromatography, Biochromatography; FPLC Protein A-purified monoclonal
Application Note Separation of three monoclonal antibody variants using MCSGP Category Matrix Method Keywords Analytes ID Continuous chromatography, Biochromatography; FPLC Protein A-purified monoclonal
Instructions. Torpedo sirna. Material. Important Guidelines. Specifications. Quality Control
 is a is a state of the art transfection reagent, specifically designed for the transfer of sirna and mirna into a variety of eukaryotic cell types. is a state of the art transfection reagent, specifically
is a is a state of the art transfection reagent, specifically designed for the transfer of sirna and mirna into a variety of eukaryotic cell types. is a state of the art transfection reagent, specifically
