Analysis of posttranslational modifications of proteins by tandem mass spectrometry
|
|
|
- Judith Hines
- 8 years ago
- Views:
Transcription
1 MASS SPECTROMETRY FOR PROTEOMICS ANALYSIS REVIEW Analysis of posttranslational modifications of proteins by tandem mass spectrometry Martin R. Larsen, Morten B. Trelle, Tine E. Thingholm, and Ole N. Jensen BioTechniques 40: (June 2006) doi / Protein activity and turnover is tightly and dynamically regulated in living cells. Whereas the three-dimensional protein structure is predominantly determined by the amino acid sequence, posttranslational modification (PTM) of proteins modulates their molecular function and the spatial-temporal distribution in cells and tissues. Most PTMs can be detected by protein and peptide analysis by mass spectrometry (MS), either as a mass increment or a mass deficit relative to the nascent unmodified protein. Tandem mass spectrometry (MS/MS) provides a series of analytical features that are highly useful for the characterization of modified proteins via amino acid sequencing and specific detection of posttranslationally modified amino acid residues. Large-scale, quantitative analysis of proteins by MS/MS is beginning to reveal novel patterns and functions of PTMs in cellular signaling networks and biomolecular structures. INTRODUCTION Determination of posttranslational modifications (PTM) of proteins is fundamental in elucidation of the intricate processes that govern cellular events, like cell division, growth, and differentiation. The term PTM denotes changes in the polypeptide chain as a result of either the addition or removal of distinct chemical moieties to amino acid residues, proteolytic processing of the protein termini, or the introduction of covalent crosslinks between domains of the protein. PTMs are involved in most cellular processes including the maintenance of protein structure and integrity, regulation of metabolism and defense processes, and in cellular recognition events and morphology changes. Analysis of PTMs presents a number of challenges to protein and proteomics researchers, and efficient and sensitive methods for detection of PTMs are required. Traditionally, PTMs have been identified by Edman degradation, amino acid analysis, isotopic labeling, or immunochemistry. Within recent years, mass spectrometry (MS) has proven to be extremely useful in PTM discovery. The presence of covalent modifications in proteins affects the molecular weight of the modified amino acids, and the mass increment or deficit can be detected by MS (Table 1). MS has several advantages for characterization of PTMs, including (i) very high sensitivity; (ii) ability to identify the site of PTM; (iii) discovery of novel PTMs; (iv) capability to identify PTMs in complex mixtures of proteins; and finally (v) the ability to quantify the relative changes in PTM occupancy at distinct sites. None of the other techniques provide all these features. In this review we describe the utility of tandem mass spectrometry (MS/MS) for the determination of PTMs and provide selected examples of recent modification-specific proteomic studies. MS IN PROTEOMICS MS is now widely used in protein biochemistry and in proteomics for the identification and characterization of proteins in cell lysates, isolated organelles, or purified multisubunit complexes (1 3). Protein separation technologies based on centrifugation, electrophoresis, or chromatographic methods are readily interfaced to MS in an online or off-line fashion. Proteins are then converted into peptides by treatment with sequence-specific proteases or chemical reagents, since peptides are more amenable to MS and MS/MS analysis than intact proteins. The availability of robust and sensitive matrix-assisted laser desorption/ionization MS (MALDI- MS) (4) and electrospray ionization MS (ESI-MS) (5) instruments makes advanced MS technology accessible to molecular cell biologists, biochemists, and proteomics researchers. MS using either MALDI or ESI as the ionization method enables accurate mass determination of peptides. Peptide mass fingerprinting by MALDI-MS and subsequent sequence database searching is widely used for identification of proteins that are isolated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) or by two-dimensional gel electrophoresis (2DE) gels. In most studies it is also desirable to obtain amino acid sequence information, because the accurately determined molecular mass in combination with a partial amino acid sequence of a peptide are very specific probes for protein identification (1,6). Furthermore, mass University of Southern Denmark, Odense M, Denmark 790 BioTechniques Vol. 40, No. 6 (2006)
2 A B Figure 1. Tandem mass spectrometry (MS/MS) for mapping posttranslational modifications (PTMs). MS/MS is a very sensitive, accurate, and efficient method for sequencing of peptides via the generation and detection of sequence-specific fragment ions. In addition to providing the means to identify proteins, MS also offers tools to interrogate proteins for PTMs. PTM-specific mass increments of peptides and amino acid residues or diagnostic fragment ions in mass spectra reveal the presence of PTMs. (A) Modular setup of MS/MS experiment. A mass spectrometer consists of an ion source, in which the sample molecules are ionized, a mass analyzer, which separates the ions according to their mass-to charge ratio (m/z), and a detector that records the ions. A sophisticated computer system controls the mass spectrometer and the data-dependent acquisition functions. (B) MS and MS/MS spectra of peptide carrying a PTM. First, the MS spectrum is acquired to determine the molecular mass of the peptides. Next, peptides are in turn selected for MS/MS. Fragmentation of the peptide amide bond produces a set of fragment ions that generate a ladder-like readout of the sequence in the tandem mass spectrum. The presence of a PTM will change the mass of the modified amino acid residue and of the peptide. MS/ MS often reveals the mass of the PTM and the identity and position of the modified amino acid residue. analysis and amino acid sequencing by MS/MS may reveal the presence of PTMs at individual amino acid residues in proteins (3,7). PEPTIDE SEQUENCING BY MS/MS The tandem mass spectrometer provides the means for amino acid sequencing of peptides. It enables gasphase isolation of individual peptide ion species followed by collision-induced dissociation (CID) and detection of the resultant amino acid sequence specific fragment ions (Figure 1). The MS/MS experiment consists of several stages of mass analysis and ion manipulation. First, the masses of the sample analytes are determined in a MS survey scan (first MS experiment). Second, the peptide ion of interest is isolated via its mass-to-charge ratio (m/z) value (i.e., by filtering away other ion species that have a different m/z value). Third, the selected peptide ion species is activated (e.g., by collisions with an inert gas such as Argon that imparts internal energy into the ions and thereby induces their fragmentation). Last, the m/z values of the fragment ions are determined (second MS experiment). The most labile bonds in peptides are generally the backbone amide bonds, leading to breakage of the peptide backbone in between amino acids. Hence, tandem mass spectra of peptides contain a series of sequence-revealing fragment ions (8). The fragment ion signals reflect the amino acid sequence as read from either the N-terminal (b-ion series) or the C-terminal (y-ion series) direction (9). The identities of the individual amino acids are revealed by the mass differences between the signals in b- ion series or y-ion series (Figure 1). MS/MS is a sensitive and fast technique for peptide sequencing. Depending on the type of instrumentation and the experimental design, ng pure protein are needed to obtain sufficient peptide data for protein identification. Microgram amounts of protein are needed for proteomics analysis of complex protein samples and organelles. Coupling of capillary liquid chromatography to tandem mass spectrometry (LC-MS/ MS) allows for automated analysis of complex peptide mixtures by datadependent acquisition of peptide mass spectra and tandem mass spectra. Typically, 1 5 peptides are analyzed by MS/MS per second. Hence, hundreds or even thousands of peptides can be analyzed in a single LC-MS/MS experiment to reveal the inventory of proteins in biological samples. Large-scale MS experiments in proteomics are facilitated by computational data analysis and database searching algorithms that enable protein identification and quantitation. A series of computational methods for the assignment of peptide sequences based on automated interpretation of MS/MS spectra and protein sequence database searching have been developed (6,10,11). These tools are often interfaced to or integrated with MS software provided by manufacturers to give an efficient data processing pipeline for annotation of peptides and the equivalent proteins. Vol. 40, No. 6 (2006) BioTechniques 791
3 MASS SPECTROMETRY FOR PROTEOMICS ANALYSIS REVIEW PTM ANALYSIS BY MS/MS Accurate determination of the mass increment at the protein or peptide level will aid in defining the type of modification. Intact mass determination of purified proteins is a useful method for determining modification and processing events. Comparison of the experimentally detected intact molecular mass with the calculated mass obtained from the amino acid sequence of the protein will reveal any discrepancies, and the mass difference or deviation may define the modification. In addition, the mass spectrum will often reveal heterogeneous modification and processing of the protein, resulting in multiple species that each correspond to an individual modification state. A large class of PTMs is represented by chemical moieties that are covalently attached to proteins by various enzymes. Examples are phosphorylation (+80 Da), sulfation (+80 Da), nitration (+45 Da), O-glycosylation (>203 Da), and acylation (>200 Da) (Table 1 and references therein). Another class of PTM is proteolytic processing that leads to mass deficit. Removal of the leader methionine residue (131 Da) is an example of such a processing event. Processing of precursor polypeptides for removal of signal sequences to generate biologically active proteins is yet other types of processing. Amino acid oxidations are also common PTMs, especially on methionine and tryptophan residues, leading to mass increments of 16 or 32 Da (Table 1). The formation of disulfide bridges between cysteine residues leads to a mass change of -2 Da. Thus, mass analysis of intact proteins or the derived peptides can reveal modifications and also provide information on the type of the PTMs. However, assignment of the PTM to a specific amino acid residue necessitates sequencing of the protein or peptide. MS/MS facilitates mass determination and sequencing of peptides and thereby also the detection of site-specific PTMs (3). The chemical stability of the PTM is decisive for its efficient detection in MS/MS. Certain PTMs will remain intact during MS and MS/MS experiments. For example, acetyl-lysine is a very stable PTM that leads to a mass increment of 42 Da of the intact peptide. Upon MS/MS of a lysineacetylated peptide, all the fragment ions that contain the AcLys residue exhibit a +42 Da mass increment relative to the unmodified peptide (Table 1). Less stable PTMs are phosphoserine and phosphothreonine, which often eliminate phosphoric acid (or a phosphate group and water) during MS/ MS (12). Thus, all the fragment ions that contained these phosphoamino acid residues will typically exhibit a mass deficit of 98 Da (-H 3 PO 4 ), also known as a neutral loss, relative to the intact phosphopeptide. These Δm values are very useful for annotation of PTM peptide MS/MS spectra, as they aid in identifying and assigning the modified residues (Table 1). In addition to the amino acid sequence-specific peptide fragment ions, also PTM-specific signals from modified amino acids exist. These PTM-specific fragment ions appear in the lower mass range of tandem mass spectra, typically below Table 1. Mass Values for Some Posttranslational Modifications and Frequently Observed Neutral Losses and Diagnostic Ion Signals PTM Δm (MS) (Da) Δm (MS/MS) Neutral loss (Da) MS/MS Diagnostic ion (m/z) References (examples) Phosphotyrosine (neg. mode) 16, (pos.mode) Phosphothreonine (neg. mode) 17 Phosphoserine Acetyllysine , Methyllysine Dimethyllysine Methylarginine Dimethylarginine Trimethyllysine N-linked glycans (hexose) (hexosamins) (sialic acids) (HexHexNAc) See ref. See ref See ref. See ref , Nitrotyrosine Palmitoyl Acylation Various, see ref See ref See ref 23 Ubiquitin (Gly-Gly tag remains after tryptic digestion) 70 Methionine oxidation Tryptophan oxidation , , ,66 65,66 67,68 67,68 65, PTM, posttranslational modification; MS, mass spectrometry; MS/MS, tandem mass spectrometry; m/z, mass-to charge ratio. 792 BioTechniques Vol. 40, No. 6 (2006)
4 A B (24 26). The principle is based on the lability of PTMs, such as phosphoserine/threonine, that readily undergoes gas-phase elimination of a PTM-related chemical group during MS/MS. For example, the neutral loss of H 3 PO 4 observed upon MS/MS of phosphopeptides will trigger a MS/MS/ MS analysis of the product, to reveal the sequence of the PTM peptide and the position of the PTM (Figure 2). m/z 400. Certain amino acids are detected as immonium ion fragments that constitute a fingerprint for the presence of these amino acid residues in a peptide. Several types of modified amino acid residues also provide diagnostic ion signals. Acetyl-lysine generates ion signals at m/z and (Table 1 and References 13 15). The presence of these signals in MS/MS spectra is a good indication that the corresponding peptide contains Ac-Lys. Similarly, phosphotyrosine generates a diagnostic signal at m/z (16). Glycopeptides generate glycan-specific fragments (oxonium ions) and acylated peptides produce lipid-specific signals that are useful for verification of these types of PTM in MS/MS (Table 1). False positive signals due to interference from regular peptide fragment ions should be considered when the resolution and mass accuracy is insufficient to distinguish PTM-specific signals from internal peptide fragments, such as dior tri-amino acids. Figure 2. Tandem mass spectrometry (MS/MS) of posttranslational modifications (PTMs). (A) MS/MS of a stable PTM leads to detection of fragment ions that reveal the identity and position of the modified amino acid (aa) residue(s). In addition, diagnostic fragment ions originating from the modified amino acid residue are often observed. Such signals are highly useful when searching for modified peptides in large MS/MS data sets. (B) MS/MS of a labile PTM typically leads to the observation of one strong signal in the MS/MS spectrum and no significant sequence-revealing fragment ion series. Multistage mass spectrometry (MS/MS/MS) is useful to further characterize the fragment ion species by sequencing. This approach is very efficient for sequencing of phosphoserine and phosphothreonine peptides. m/z, mass-to-charge ratio. MS/MS provides a number of useful analytical features that take advantage of diagnostic ion signals and neutral loss products. Precursor ion scanning is a selective and sensitive method for specific detection of only those peptides that carry a PTM that generate a unique diagnostic ion. For example, phosphopeptides generate a strong m/z 79 ion signal in the negative ion MS/ MS mode. Thus, precursor ion scans for this ion will identify only phosphopeptides in the sample, whereas regular peptides remain undetected. A number of glycans and lipids generate fragment ions that can be explored for specific detection of PTM peptides (Table 1). The precursor ion scanning method is extremely useful for the analysis of phosphorylation sites and other PTMs in individual, purified proteins (12,17 23) (Table 1). Neutral loss analysis is in proteomics typically performed by ion trap instruments that allow very fast MS/MS and multistage mass spectrometry (MS/MS/MS) analysis COMPUTATIONAL TOOLS FOR PTM ASSIGNMENT IN MS/MS DATA SETS Several computational methods for automated annotation of PTM in peptides exist. These algorithms analyze the MS and MS/MS data, taking into account the Δm values and sometimes also neutral losses and diagnostic ions for the PTM of interest. Once a set of proteins is identified in a MS/MS-based proteomics experiment, it is possible to perform a more detailed analysis of the retrieved protein sequences and the MS/MS data set. The initial database search is typically performed with only a limited set of specified modifications, including alkylated cysteines and oxidized methionines, in order to minimize search space and to avoid false positive PTM assignments. In a second stage, the molecular masses of the unmodified proteins and the corresponding proteolytic peptides are readily calculated from the protein sequences obtained by the initial sequence database search. As mentioned previously, PTM will lead to a mass increment or mass deficit (Δm) of the modified peptide relative to the unmodified species. Thus, it becomes feasible to inquire whether any of the initially unassigned MS/MS spectra can be matched to posttranslationally modified peptides, given a list of putative modifications and their Δm values and the list of predicted and observed peptide masses. Although highly useful, computational sequence annotation tools should be used with care, because they may produce false positives in cases where the mass accuracy or signal-to-background level of the MS and MS/MS data are not sufficient to Vol. 40, No. 6 (2006) BioTechniques 793
5 MASS SPECTROMETRY FOR PROTEOMICS ANALYSIS REVIEW make unambiguous assignments. This is particularly important in modification-specific proteomics, in which protein identification is often based on the detection and sequencing of only a few PTM peptides per protein. The presence of multiple modifications or various different types of modifications in a peptide may also complicate MS and MS/MS data interpretation. STRATEGIES FOR DETERMINATION OF PTM IN PROTEOMICS Mapping of PTMs in proteomics is a demanding task because most PTMs are low abundance and/or substoichiometric and some PTMs are labile during MS and MS/MS. In addition, many modifications are hydrophilic, which complicates PTM sample handling and purification prior to MS (27). The presence of PTMs may affect the cleavage efficiency of proteases, such as trypsin, to generate unexpected or large peptide products. Certain PTMs will reduce the ionization and detection efficiency in MS. Multisite PTMs may generate very complicated MS and MS/MS data sets that are difficult to interpret. For these reasons, it is often useful to consider and explore several approaches for mapping of PTMs in proteomics (Figure 3). Generally, it is recommended to reduce the complexity of the sample as much as possible prior to MS analysis for mapping of PTMs. It is not advisable to pursue PTM mapping by direct LC-MS and MS/MS analysis of crude cell lysates, as PTMs are rarely detected in such experiments. This is mainly due to the presence of many abundant unmodified peptides, the inadequate peptide separation by LC, and the limitations of MS/MS for peptide sequencing of very complex mixtures. Biochemical purification of cellular compartments, organelles, protein complexes, or of individual proteins has proved to be a successful first stage for analysis of PTMs, as it dramatically reduces the complexity of the initial protein sample. As an alternative, PTM-specific reagents are useful for selective enrichment of PTM proteins and peptides prior to mass spectrometry. Mapping of PTMs in proteins is achievable by a series of approaches, including (i) treatment with multiple proteases and shotgun sequencing by MS/MS; (ii) enrichment of modified proteins or peptides prior to MS/MS sequencing; (iii) and PTM-specific MS/ MS and multistage MS. Combinations of these methods have proven successful for comprehensive analysis of PTM in proteomics (Figure 3). Shotgun Sequencing The shotgun sequencing approach is based on using multiple proteases with different cleavage specificity to generate complementary and redundant sets of overlapping peptides. The peptide mixtures are then analyzed by LC-MS/ MS or by multidimensional separation technology and MS/MS (LC/LC-MS/ MS) for comprehensive sequencing of proteins. Database searching and computational assembly of the sequence data then provides an overview of the protein sequences and annotation of PTMs and amino acid substitutions (28,29), and it is also amendable to mapping of modifications in purified proteins and protein complexes. PTM-Specific Enrichment Figure 3. Analytical strategies for mapping posttranslational modifications (PTMs) in proteomics. (Left column) Shotgun sequencing. This method is best suited for subproteomes, simple protein mixtures, and protein complexes, because extensive computational tandem mass spectrometry (MS/MS) data analysis and peptide sequence alignment is required. (Middle column) Modification-specific proteomics based on affinity enrichment of PTM proteins and/or peptides prior to MS analysis. This method is compatible with complex protein mixtures. Several complementary chromatographic peptide separation methods should be used prior to MS/MS analysis. (Right column) Chemistry-based methods for PTM-specific covalent capture of proteins and/or peptides. Often based on β-elimination/michael addition chemistry, phosphoramidate chemistry, or other PTM-specific chemistries. Large-scale analysis of PTMs is facilitated by PTM-specific protein and peptide enrichment methods, such as PTM-directed affinity chromatography or immunoprecipitation with PTMspecific antibodies. Phosphoproteins can be purified by PTM-specific affinity resins (30) or anti-phosphoamino acid antibodies (31,32). Subsequent digestion of protein and LC-MS/MS analysis of peptides will reveal the identity of the retrieved proteins and sometimes allow site-specific assignment of phosphorylation sites in the recovered proteins. However, it is often also beneficial to enrich for PTM peptides prior to MS analysis in order to improve sensitivity and specificity. Phosphopeptides can be recovered by antibodies (33), immobilized metal affinity chromatography (IMAC) 794 BioTechniques Vol. 40, No. 6 (2006)
6 (34 36), or by TiO2 columns (37,38) prior to MS/MS analysis. Strong cation exchange and anion exchange chromatography have also proven useful for reducing peptide complexity prior to MS/MS based detection and sequencing of modified peptides, including phosphopeptides (24,39). Using combinations of these methods, large-scale phosphoproteomics have revealed thousands of phosphorylated sites in proteins from various species (24,26,39,40). Glycoproteins can be enriched by using lectins, which are sometimes referred to as sugar-specific antibodies (41,42). Glycoproteins from serum were retrieved and identified by a twostage lectin enrichment procedure that first targeted the intact glycoproteins and then the proteolytically derived glycopeptides. The glycopeptide sample was then treated with N-glycosidase F in the presence of 18-O water and analyzed by LC-MS/MS. The 18-O labeling enabled site-specific assignment of glycosylation sites by signature ion signals MS/MS (43). An alternative strategy used lectins for glycoprotein enrichment followed by hydrophilic interaction chromatography (HILIC) for glycopeptide enrichment. Glycosidase D/H treatment and MS/MS then facilitated protein identification and assignment of glycosylation sites (44). GPI-anchored proteins, another class of glycoproteins, were identified by using modification-specific enzymes (phospholipases) to selectively release GPI-anchored proteins from plasma membrane preparations in a two phase detergent system, followed by their identification by LC-MS/MS and bioinformatics sequence analysis (45). O-glycosylation is also becoming tractable by using affinity enrichment and advanced MS methods (46). Genetic methods are also useful for enrichment of PTM proteins. Ubiquitylated and SUMOylated proteins are amendable to purification and identification by integration of genetic tags into the modifying proteins (47). PTM-Specific Chemistry Certain modifications are amenable to chemical conversion to stable, tractable species. This is particularly useful for PTMs that are labile during MS and MS/MS analysis. O-phospho- Ser/Thr and O-GlcNAc-Ser/Thr readily undergo β-elimination to generate intermediates that are substrates for Michael addition of alkylating groups. Such methods have been used for selective recovery of phosphoproteins and glycoproteins by solid-phase chemistry or by affinity purification prior to protein digestion and identification by MS/MS sequencing (46,48). The introduction of fluorous affinity tags helps recover modified peptides by taking advantage of the unique chromatographic properties of fluorous compounds (19). Another option is to use β-elimination/michael addition reactions to introduce unique mass tags into the modified peptides. The chemical design of the mass tag then provides a diagnostic mass fingerprint for the modified peptides, so they are readily distinguished from regular peptides (49). Other chemistries and resins, including dendrimers, have also been applied for PTM-specific recovery of proteins and peptides (50 52). Tagging-by-substrate methods were applied to identify O-GlcNAc-modified proteins and acylated proteins (53,54). QUANTIFICATION OF PTM Dynamic cellular events, such as signal transduction networks, cell cycle progression, and chromatin activity call for quantitative techniques for determination of PTMs. MS-based methods facilitate both absolute and relative quantitation of peptides and their PTM. Absolute quantitation of peptides by MS is achievable by using internal standard peptides for selected proteins and defined PTMs (55). Relative quantitation of PTM peptides is obtained by either peptide intensity profiling (PIP) by LC-MS or by stable isotope labeling (SIL) by using stable isotope-encoded chemical precursor molecules or alkylating reagents (1,3). The functionally important PTMs will exhibit variations in abundance as a result of the perturbation, and they are identified by comparison of the peptide ion signals obtained from two or more defined cellular states, typically a control experiment and one or more perturbed states. The relative abundance of individual peptides is then derived by comparative analysis of LC-MS data using retention time and mass (PIP) or by using only the MS ion intensity of isotopically encoded peptides (SIL). These methods have been used in quantitative phosphoproteomics in various organisms, from microbes to humans (26,40,56 58), and also in glycoproteomics (43,59,60). CONCLUSIONS AND PERSPECTIVES MS/MS is increasingly integrated in modern cell biology and biomedical research, in academia and in industry. Qualitative and quantitative MSbased strategies for PTM mapping in proteins have already contributed detailed insights into a series of complex biological systems, from microbes to plants and mammals. Novel MS technologies are continuously developed. Recent examples include the introduction of robust high-performance hybrid tandem mass spectrometers that provide very high mass accuracy [mass error <5 parts per million (ppm)] and multistage MS capabilities at a chromatographic timescale, thereby allowing detailed analysis of PTMs during LC-MS. Novel and improved MS/MS ion dissociation methods, such as infrared multiphoton dissociation (IRMPD) (61), electron capture dissociation (ECD) (62), and electron transfer dissociation (ETD) (63) have already proven useful for mapping of PTMs in biological systems. Increasingly advanced computational methods are useful for prediction of PTMs prior to MS analysis of known proteins, as well as for the identification of PTM peptides in large-scale data sets. Multistage MS and efficient fragmentation methods such as ECD and ETD also facilitate top-down analysis of intact proteins and large polypeptides (64). The analytical potential of these novel methods for mapping of PTMs in proteomics research is currently explored in many laboratories worldwide. No doubt, the prospects for MS-based analysis of PTM of protein are very exciting. Vol. 40, No. 6 (2006) BioTechniques 795
7 MASS SPECTROMETRY FOR PROTEOMICS ANALYSIS REVIEW ACKNOWLEDGMENTS We thank our colleagues in the Protein Research Group for their contributions and for discussions. This work was supported by grants from the Danish Natural Sciences Research Council and the Strategic Research Agency (Young Investigator Awards to M.R.L. and O.N.J.). O.N.J. is a Lundbeck Foundation Research Professor. COMPETING INTERESTS STATEMENT The authors declare no competing interests. REFERENCES 1. Aebersold, R. and M. Mann Mass spectrometry-based proteomics. Nature 422: Yates, J.R., 3rd, A. Gilchrist, K.E. Howell, and J.J. Bergeron Proteomics of organelles and large cellular structures. Nat. Rev. Mol. Cell Biol. 6: Mann, M. and O.N. Jensen Proteomic analysis of post-translational modifications. Nat. Biotechnol. 21: Karas, M. and F. Hillenkamp Laser desorption ionization of proteins with molecular masses exceeding 10,000 daltons. Anal. Chem. 60: Fenn, J.B., M. Mann, C.K. Meng, S.F. Wong, and C.M. Whitehouse Electrospray ionization for mass spectrometry of large biomolecules. Science 246: Sadygov, R.G., D. Cociorva, and J.R. Yates 3rd Large-scale database searching using tandem mass spectra: looking up the answer in the back of the book. Nat. Methods 1: Jensen, O.N Modification-specific proteomics: characterization of post-translational modifications by mass spectrometry. Curr. Opin. Chem. Biol. 8: Steen, H. and M. Mann The ABC s (and XYZ s) of peptide sequencing. Nat. Rev. Mol. Cell Biol. 5: Roepstorff, P. and J. Fohlman Proposal for a common nomenclature for sequence ions in mass spectra of peptides. Biomed. Mass Spectrom. 11: Fenyo, D. and R.C. Beavis Informatics and data management in proteomics. Trends Biotechnol. 20:S35-S Creasy, D.M. and J.S. Cottrell Error tolerant searching of uninterpreted tandem mass spectrometry data. Proteomics 2: Annan, R.S., M.J. Huddleston, R. Verma, R.J. Deshaies, and S.A. Carr A multidimensional electrospray MS-based approach to phosphopeptide mapping. Anal. Chem. 73: Borchers, C., C.E. Parker, L.J. Deterding, and K.B. Tomer Preliminary comparison of precursor scans and liquid chromatography-tandem mass spectrometry on a hybrid quadrupole time-of-flight mass spectrometer. J. Chromatogr. A. 854: Kim, J.Y., K.W. Kim, H.J. Kwon, D.W. Lee, and J.S. Yoo Probing lysine acetylation with a modification-specific marker ion using high-performance liquid chromatography/electrospray-mass spectrometry with collision-induced dissociation. Anal. Chem. 74: Dormeyer, W., M. Ott, and M. Schnolzer Probing lysine acetylation in proteins: strategies, limitations, and pitfalls of in vitro acetyltransferase assays. Mol. Cell. Proteomics 4: Steen, H., B. Kuster, M. Fernandez, A. Pandey, and M. Mann Detection of tyrosine phosphorylated peptides by precursor ion scanning quadrupole TOF mass spectrometry in positive ion mode. Anal. Chem. 73: Neubauer, G. and M. Mann Mapping of phosphorylation sites of gel-isolated proteins by nanoelectrospray tandem mass spectrometry: potentials and limitations. Anal. Chem. 71: Huddleston, M.J., M.F. Bean, and S.A. Carr Collisional fragmentation of glycopeptides by electrospray ionization LC/MS and LC/MS/MS: methods for selective detection of glycopeptides in protein digests. Anal. Chem. 65: Brittain, S.M., S.B. Ficarro, A. Brock, and E.C. Peters Enrichment and analysis of peptide subsets using fluorous affinity tags and mass spectrometry. Nat. Biotechnol. 23: Unwin, R.D., J.R. Griffiths, M.K. Leverentz, A. Grallert, I.M. Hagan, and A.D. Whetton Multiple reaction monitoring to identify sites of protein phosphorylation with high sensitivity. Mol. Cell. Proteomics 4: Ross, P.L.,Y.N. Huang, J.N. Marchese, B. Williamson, K. Parker, S. Hattan, N. Khainovski, S. Pillai, et al Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol. Cell. Proteomics 3: Steen, H., M. Fernandez, S. Ghaffari, A. Pandey, and M. Mann Phosphotyrosine mapping in Bcr/Abl oncoprotein using phosphotyrosine-specific immonium ion scanning. Mol. Cell. Proteomics 2: Hoffman, M.D. and J. Kast Mass spectrometric characterization of lipid-modified peptides for the analysis of acylated proteins. J. Mass Spectrom. 41: Beausoleil, S.A., M. Jedrychowski, D. Schwartz, J.E. Elias, J. Villen, J. Li, M.A. Cohn, L.C. Cantley, and S.P. Gygi Large-scale characterization of HeLa cell nuclear phosphoproteins. Proc. Natl. Acad. Sci. USA 101: Chang, E.J., V. Archambault, D.T. McLachlin, A.N. Krutchinsky, and B.T. Chait Analysis of protein phosphorylation by hypothesis-driven multiple-stage mass spectrometry. Anal. Chem. 76: Gruhler, A., J.V. Olsen, S. Mohammed, P. Mortensen, N.J. Faergeman, M. Mann, and O.N. Jensen Quantitative phosphoproteomics applied to the yeast pheromone signaling pathway. Mol. Cell. Proteomics 4: Larsen, M.R., M.E. Graham, P.J. Robinson, and P. Roepstorff Improved detection of hydrophilic phosphopeptides using graphite powder microcolumns and mass spectrometry: evidence for in vivo doubly phosphorylated dynamin I and dynamin III. Mol. Cell. Proteomics 3: Cantin, G.T. and J.R. Yates 3rd Strategies for shotgun identification of posttranslational modifications by mass spectrometry. J. Chromatogr. A. 1053: Wu, C.C., M.J. MacCoss, K.E. Howell, and J.R. Yates 3rd A method for the comprehensive proteomic analysis of membrane proteins. Nat. Biotechnol. 21: Collins, M.O., L. Yu, M.P. Coba, H. Husi, I. Campuzano, W.P. Blackstock, J.S. Choudhary, and S.G. Grant Proteomic analysis of in vivo phosphorylated synaptic proteins. J. Biol. Chem. 280: Gronborg, M., T.Z. Kristiansen, A. Stensballe, J.S. Andersen, O. Ohara, M. Mann, O.N. Jensen, and A. Pandey A mass spectrometry-based proteomic approach for identification of serine/threoninephosphorylated proteins by enrichment with phospho-specific antibodies: identification of a novel protein, Frigg, as a protein kinase A substrate. Mol. Cell. Proteomics 1: Pandey, A., A.V. Podtelejnikov, B. Blagoev, X.R. Bustelo, M. Mann, and H.F. Lodish Analysis of receptor signaling pathways by mass spectrometry: identification of vav-2 as a substrate of the epidermal and platelet-derived growth factor receptors. Proc. Natl. Acad. Sci. USA 97: Rush, J., A. Moritz, K.A. Lee, A. Guo, V.L. Goss, E.J. Spek, H. Zhang, X.M. Zha, et al Immunoaffinity profiling of tyrosine phosphorylation in cancer cells. Nat. Biotechnol. 23: Ficarro, S.B., M.L. McCleland, P.T. Stukenberg, D.J. Burke, M.M. Ross, J. Shabanowitz, D.F. Hunt, and F.M. White Phosphoproteome analysis by mass spectrometry and its application to Saccharomyces cerevisiae. Nat. Biotechnol. 20: Posewitz, M.C. and P. Tempst Immobilized gallium(iii) affinity chromatography of phosphopeptides. Anal. Chem. 71: Stensballe, A., S. Andersen, and O.N. Jensen Characterization of phosphoproteins from electrophoretic gels by nanoscale Fe(III) affinity chromatography with off-line mass spectrometry analysis. Proteomics 1: Larsen, M.R., T.E. Thingholm, O.N. Jensen, P. Roepstorff, and T.J. Jorgensen Highly selective enrichment of phosphorylated peptides from peptide mixtures 796 BioTechniques Vol. 40, No. 6 (2006)
8 using titanium dioxide microcolumns. Mol. Cell. Proteomics 4: Pinkse, M.W., P.M. Uitto, M.J. Hilhorst, B. Ooms, and A.J. Heck Selective isolation at the femtomole level of phosphopeptides from proteolytic digests using 2D- NanoLC-ESI-MS/MS and titanium oxide precolumns. Anal. Chem. 76: Nuhse, T.S., A. Stensballe, O.N. Jensen, and S.C. Peck Large-scale analysis of in vivo phosphorylated membrane proteins by immobilized metal ion affinity chromatography and mass spectrometry. Mol. Cell. Proteomics 2: Zhang, Y., A. Wolf-Yadlin, P.L. Ross, D.J. Pappin, J. Rush, D.A. Lauffenburger, and F.M. White Time-resolved mass spectrometry of tyrosine phosphorylation sites in the epidermal growth factor receptor signaling network reveals dynamic modules. Mol. Cell. Proteomics 4: Gabius, H.J., S. Andre, H. Kaltner, and H.C. Siebert The sugar code: functional lectinomics. Biochim. Biophys. Acta 1572: Yang, Z. and W.S. Hancock Approach to the comprehensive analysis of glycoproteins isolated from human serum using a multi-lectin affinity column. J. Chromatogr. A. 1053: Kaji, H., H. Saito, Y. Yamauchi, T. Shinkawa, M. Taoka, J. Hirabayashi, K. Kasai, N. Takahashi, and T. Isobe Lectin affinity capture, isotope-coded tagging and mass spectrometry to identify N-linked glycoproteins. Nat. Biotechnol. 21: Hagglund, P., J. Bunkenborg, F. Elortza, O.N. Jensen, and P. Roepstorff A new strategy for identification of N-glycosylated proteins and unambiguous assignment of their glycosylation sites using HILIC enrichment and partial deglycosylation. J. Proteome Res. 3: Elortza, F., T.S. Nuhse, L.J. Foster, A. Stensballe, S.C. Peck, and O.N. Jensen Proteomic analysis of glycosylphosphatidylinositol-anchored membrane proteins. Mol. Cell. Proteomics 2: Wells, L., K. Vosseller, R.N. Cole, J.M. Cronshaw, M.J. Matunis, and G.W. Hart Mapping sites of O-GlcNAc modification using affinity tags for serine and threonine post-translational modifications. Mol. Cell. Proteomics 1: Kirkpatrick, D.S., C. Denison, and S.P. Gygi Weighing in on ubiquitin: the expanding role of mass-spectrometry-based proteomics. Nat. Cell Biol. 7: Oda, Y., T. Nagasu, and B.T. Chait Enrichment analysis of phosphorylated proteins as a tool for probing the phosphoproteome. Nat. Biotechnol. 19: Molloy, M.P. and P.C. Andrews Phosphopeptide derivatization signatures to identify serine and threonine phosphorylated peptides by mass spectrometry. Anal. Chem. 73: Tao, W.A., B. Wollscheid, R. O Brien, J.K. Eng, X.J. Li, B. Bodenmiller, J.D. Watts, L. Hood, and R. Aebersold Quantitative phosphoproteome analysis using a dendrimer conjugation chemistry and tandem mass spectrometry. Nat. Methods 2: Zhang, H., E.C. Yi, X.J. Li, P. Mallick, K.S. Kelly-Spratt, C.D. Masselon, D.G. Camp 2nd, R.D. Smith, et al High throughput quantitative analysis of serum proteins using glycopeptide capture and liquid chromatography mass spectrometry. Mol. Cell. Proteomics 4: Zhou, H., J.D. Watts, and R. Aebersold A systematic approach to the analysis of protein phosphorylation. Nat. Biotechnol. 19: Sprung, R., A. Nandi, Y. Chen, S.C. Kim, D. Barma, J.R. Falck, and Y. Zhao Tagging-via-substrate strategy for probing O- GlcNAc modified proteins. J. Proteome Res. 4: Kho, Y., S.C. Kim, C. Jiang, D. Barma, S.W. Kwon, J. Cheng, J. Jaunbergs, C. Weinbaum, et al A tagging-viasubstrate technology for detection and proteomics of farnesylated proteins. Proc. Natl. Acad. Sci. USA 101: Gerber, S.A., J. Rush, O. Stemman, M.W. Kirschner, and S.P. Gygi Absolute quantification of proteins and phosphoproteins from cell lysates by tandem MS. Proc. Natl. Acad. Sci. USA 100: Oda, Y., K. Huang, F.R. Cross, D. Cowburn, and B.T. Chait Accurate quantitation of protein expression and site-specific phosphorylation. Proc. Natl. Acad. Sci. USA 96: Blagoev, B., S.E. Ong, I. Kratchmarova, and M. Mann Temporal analysis of phosphotyrosine-dependent signaling networks by quantitative proteomics. Nat. Biotechnol. 22: Ballif, B.A., P.P Roux, S.A. Gerber, J.P. MacKeigan, J. Blenis, and S.P. Gygi Quantitative phosphorylation profiling of the ERK/p90 ribosomal S6 kinase-signaling cassette and its targets, the tuberous sclerosis tumor suppressors. Proc. Natl. Acad. Sci. USA 102: Zhang, H., X.J. Li, D.B. Martin, and R. Aebersold Identification and quantification of N-linked glycoproteins using hydrazide chemistry, stable isotope labeling and mass spectrometry. Nat. Biotechnol. 21: Qiu, R. and F.E. Regnier Comparative glycoproteomics of N-linked complex-type glycoforms containing sialic acid in human serum. Anal. Chem. 77: Cooper, H.J., M.H. Tatham, E. Jaffray, J.K. Heath, T.T. Lam, A.G. Marshall, and R.T. Hay Fourier transform ion cyclotron resonance mass spectrometry for the analysis of small ubiquitin-like modifier (SUMO) modification: identification of lysines in RanBP2 and SUMO targeted for modification during the E3 autosumoylation reaction. Anal. Chem. 77: Zubarev, R.A Electron-capture dissociation tandem mass spectrometry. Curr. Opin. Biotechnol. 15: Syka, J.E., J.J. Coon, M.J. Schroeder, J. Shabanowitz, and D.F. Hunt Peptide and protein sequence analysis by electron transfer dissociation mass spectrometry. Proc. Natl. Acad. Sci. USA 101: Meng, F., A.J. Forbes, L.M. Miller, and N.L. Kelleher Detection and localization of protein modifications by high resolution tandem mass spectrometry. Mass Spectrom. Rev. 24: Hirota, J., Y. Satomi, K. Yoshikawa, and T. Takao Epsilon-trimethyllysine-specific ions in matrix-assisted laser desorption/ ionization-tandem mass spectrometry. Rapid Commun. Mass Spectrom. 17: Goff, S.A., D. Ricke, T.H. Lan, G. Presting, R. Wang, M. Dunn, J. Glazebrook, A. Sessions, et al A draft sequence of the rice genome (Oryza sativa L. ssp. japonica). Science 296: Gehrig, P.M., P.E. Hunziker, S. Zahariev, and S. Pongor Fragmentation pathways of N(G)-methylated and unmodified arginine residues in peptides studied by ESI- MS/MS and MALDI-MS. J. Am. Soc. Mass Spectrom. 15: Rappsilber, J., W.J. Friesen, S. Paushkin, G. Dreyfuss, and M. Mann Detection of arginine dimethylated peptides by parallel precursor ion scanning mass spectrometry in positive ion mode. Anal. Chem. 75: Zhang, K., P.M. Yau, B. Chandrasekhar, R. New, R. Kondrat, B.S. Imai, and M.E. Bradbury Differentiation between peptides containing acetylated or tri-methylated lysines by mass spectrometry: an application for determining lysine 9 acetylation and methylation of histone H3. Proteomics 4: Peng, J., D. Schwartz, J.E. Elias, C.C. Thoreen, D. Cheng, G. Marsischky, J. Roelofs, D. Finley, and S.P. Gygi A proteomics approach to understanding protein ubiquitination. Nat. Biotechnol. 21: Jiang, X.Y., J.B. Smith, and E.C. Abraham Identification of a MS-MS fragment diagnostic for methionine sulfoxide. J. Mass Spectrom. 31: Finley, E.L., J. Dillon, R.K. Crouch, and K.L. Schey Identification of tryptophan oxidation products in bovine alpha-crystallin. Protein Sci. 7: Address correspondence to: Ole N. Jensen Protein Research Group Department of Biochemistry and Molecular Biology University of Southern Denmark Campusvej 55, DK-5230 Odense M, Denmark Jenseno@bmb.sdu.dk Vol. 40, No. 3 (2006) BioTechniques 797
Introduction to Proteomics 1.0
 Introduction to Proteomics 1.0 CMSP Workshop Tim Griffin Associate Professor, BMBB Faculty Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s
Introduction to Proteomics 1.0 CMSP Workshop Tim Griffin Associate Professor, BMBB Faculty Director, CMSP Objectives Why are we here? For participants: Learn basics of MS-based proteomics Learn what s
Chapter 14. Modeling Experimental Design for Proteomics. Jan Eriksson and David Fenyö. Abstract. 1. Introduction
 Chapter Modeling Experimental Design for Proteomics Jan Eriksson and David Fenyö Abstract The complexity of proteomes makes good experimental design essential for their successful investigation. Here,
Chapter Modeling Experimental Design for Proteomics Jan Eriksson and David Fenyö Abstract The complexity of proteomes makes good experimental design essential for their successful investigation. Here,
Aiping Lu. Key Laboratory of System Biology Chinese Academic Society APLV@sibs.ac.cn
 Aiping Lu Key Laboratory of System Biology Chinese Academic Society APLV@sibs.ac.cn Proteome and Proteomics PROTEin complement expressed by genome Marc Wilkins Electrophoresis. 1995. 16(7):1090-4. proteomics
Aiping Lu Key Laboratory of System Biology Chinese Academic Society APLV@sibs.ac.cn Proteome and Proteomics PROTEin complement expressed by genome Marc Wilkins Electrophoresis. 1995. 16(7):1090-4. proteomics
Session 1. Course Presentation: Mass spectrometry-based proteomics for molecular and cellular biologists
 Program Overview Session 1. Course Presentation: Mass spectrometry-based proteomics for molecular and cellular biologists Session 2. Principles of Mass Spectrometry Session 3. Mass spectrometry based proteomics
Program Overview Session 1. Course Presentation: Mass spectrometry-based proteomics for molecular and cellular biologists Session 2. Principles of Mass Spectrometry Session 3. Mass spectrometry based proteomics
Challenges in Computational Analysis of Mass Spectrometry Data for Proteomics
 Ma B. Challenges in computational analysis of mass spectrometry data for proteomics. SCIENCE AND TECHNOLOGY 25(1): 1 Jan. 2010 JOURNAL OF COMPUTER Challenges in Computational Analysis of Mass Spectrometry
Ma B. Challenges in computational analysis of mass spectrometry data for proteomics. SCIENCE AND TECHNOLOGY 25(1): 1 Jan. 2010 JOURNAL OF COMPUTER Challenges in Computational Analysis of Mass Spectrometry
ProteinScape. Innovation with Integrity. Proteomics Data Analysis & Management. Mass Spectrometry
 ProteinScape Proteomics Data Analysis & Management Innovation with Integrity Mass Spectrometry ProteinScape a Virtual Environment for Successful Proteomics To overcome the growing complexity of proteomics
ProteinScape Proteomics Data Analysis & Management Innovation with Integrity Mass Spectrometry ProteinScape a Virtual Environment for Successful Proteomics To overcome the growing complexity of proteomics
Thermo Scientific PepFinder Software A New Paradigm for Peptide Mapping
 Thermo Scientific PepFinder Software A New Paradigm for Peptide Mapping For Conclusive Characterization of Biologics Deep Protein Characterization Is Crucial Pharmaceuticals have historically been small
Thermo Scientific PepFinder Software A New Paradigm for Peptide Mapping For Conclusive Characterization of Biologics Deep Protein Characterization Is Crucial Pharmaceuticals have historically been small
Introduction to Proteomics
 Introduction to Proteomics Why Proteomics? Same Genome Different Proteome Black Swallowtail - larvae and butterfly Biological Complexity Yeast - a simple proteome 6,113 proteins = 344,855 tryptic peptides
Introduction to Proteomics Why Proteomics? Same Genome Different Proteome Black Swallowtail - larvae and butterfly Biological Complexity Yeast - a simple proteome 6,113 proteins = 344,855 tryptic peptides
La Protéomique : Etat de l art et perspectives
 La Protéomique : Etat de l art et perspectives Odile Schiltz Institut de Pharmacologie et de Biologie Structurale CNRS, Université de Toulouse, Odile.Schiltz@ipbs.fr Protéomique et Spectrométrie de Masse
La Protéomique : Etat de l art et perspectives Odile Schiltz Institut de Pharmacologie et de Biologie Structurale CNRS, Université de Toulouse, Odile.Schiltz@ipbs.fr Protéomique et Spectrométrie de Masse
Master course KEMM03 Principles of Mass Spectrometric Protein Characterization. Exam
 Exam Master course KEMM03 Principles of Mass Spectrometric Protein Characterization 2010-10-29 kl 08.15-13.00 Use a new paper for answering each question! Write your name on each paper! Aids: Mini calculator,
Exam Master course KEMM03 Principles of Mass Spectrometric Protein Characterization 2010-10-29 kl 08.15-13.00 Use a new paper for answering each question! Write your name on each paper! Aids: Mini calculator,
Proteomics in Practice
 Reiner Westermeier, Torn Naven Hans-Rudolf Höpker Proteomics in Practice A Guide to Successful Experimental Design 2008 Wiley-VCH Verlag- Weinheim 978-3-527-31941-1 Preface Foreword XI XIII Abbreviations,
Reiner Westermeier, Torn Naven Hans-Rudolf Höpker Proteomics in Practice A Guide to Successful Experimental Design 2008 Wiley-VCH Verlag- Weinheim 978-3-527-31941-1 Preface Foreword XI XIII Abbreviations,
泛 用 蛋 白 質 體 學 之 質 譜 儀 資 料 分 析 平 台 的 建 立 與 應 用 Universal Mass Spectrometry Data Analysis Platform for Quantitative and Qualitative Proteomics
 泛 用 蛋 白 質 體 學 之 質 譜 儀 資 料 分 析 平 台 的 建 立 與 應 用 Universal Mass Spectrometry Data Analysis Platform for Quantitative and Qualitative Proteomics 2014 Training Course Wei-Hung Chang ( 張 瑋 宏 ) ABRC, Academia
泛 用 蛋 白 質 體 學 之 質 譜 儀 資 料 分 析 平 台 的 建 立 與 應 用 Universal Mass Spectrometry Data Analysis Platform for Quantitative and Qualitative Proteomics 2014 Training Course Wei-Hung Chang ( 張 瑋 宏 ) ABRC, Academia
Building innovative drug discovery alliances. Evotec Munich. Quantitative Proteomics to Support the Discovery & Development of Targeted Drugs
 Building innovative drug discovery alliances Evotec Munich Quantitative Proteomics to Support the Discovery & Development of Targeted Drugs Evotec AG, Evotec Munich, June 2013 About Evotec Munich A leader
Building innovative drug discovery alliances Evotec Munich Quantitative Proteomics to Support the Discovery & Development of Targeted Drugs Evotec AG, Evotec Munich, June 2013 About Evotec Munich A leader
Global and Discovery Proteomics Lecture Agenda
 Global and Discovery Proteomics Christine A. Jelinek, Ph.D. Johns Hopkins University School of Medicine Department of Pharmacology and Molecular Sciences Middle Atlantic Mass Spectrometry Laboratory Global
Global and Discovery Proteomics Christine A. Jelinek, Ph.D. Johns Hopkins University School of Medicine Department of Pharmacology and Molecular Sciences Middle Atlantic Mass Spectrometry Laboratory Global
Pinpointing phosphorylation sites using Selected Reaction Monitoring and Skyline
 Pinpointing phosphorylation sites using Selected Reaction Monitoring and Skyline Christina Ludwig group of Ruedi Aebersold, ETH Zürich The challenge of phosphosite assignment Peptides Phosphopeptides MS/MS
Pinpointing phosphorylation sites using Selected Reaction Monitoring and Skyline Christina Ludwig group of Ruedi Aebersold, ETH Zürich The challenge of phosphosite assignment Peptides Phosphopeptides MS/MS
Application Note # LCMS-66 Straightforward N-glycopeptide analysis combining fast ion trap data acquisition with new ProteinScape functionalities
 Application Note # LCMS-66 Straightforward N-glycopeptide analysis combining fast ion trap data acquisition with new ProteinScape functionalities Introduction Glycosylation is one of the most common and
Application Note # LCMS-66 Straightforward N-glycopeptide analysis combining fast ion trap data acquisition with new ProteinScape functionalities Introduction Glycosylation is one of the most common and
Definition of the Measurand: CRP
 A Reference Measurement System for C-reactive Protein David M. Bunk, Ph.D. Chemical Science and Technology Laboratory National Institute of Standards and Technology Definition of the Measurand: Human C-reactive
A Reference Measurement System for C-reactive Protein David M. Bunk, Ph.D. Chemical Science and Technology Laboratory National Institute of Standards and Technology Definition of the Measurand: Human C-reactive
The Scheduled MRM Algorithm Enables Intelligent Use of Retention Time During Multiple Reaction Monitoring
 The Scheduled MRM Algorithm Enables Intelligent Use of Retention Time During Multiple Reaction Monitoring Delivering up to 2500 MRM Transitions per LC Run Christie Hunter 1, Brigitte Simons 2 1 AB SCIEX,
The Scheduled MRM Algorithm Enables Intelligent Use of Retention Time During Multiple Reaction Monitoring Delivering up to 2500 MRM Transitions per LC Run Christie Hunter 1, Brigitte Simons 2 1 AB SCIEX,
Phosphorylation Analysis by Mass Spectrometry
 Research Phosphorylation Analysis by Mass Spectrometry MYTHS, FACTS, AND THE CONSEQUENCES FOR QUALITATIVE AND QUANTITATIVE MEASUREMENTS* Hanno Steen, Judith A. Jebanathirajah **, John Rush, Nicolas Morrice,
Research Phosphorylation Analysis by Mass Spectrometry MYTHS, FACTS, AND THE CONSEQUENCES FOR QUALITATIVE AND QUANTITATIVE MEASUREMENTS* Hanno Steen, Judith A. Jebanathirajah **, John Rush, Nicolas Morrice,
Introduction to Proteomics
 Introduction to Proteomics Åsa Wheelock, Ph.D. Division of Respiratory Medicine & Karolinska Biomics Center asa.wheelock@ki.se In: Systems Biology and the Omics Cascade, Karolinska Institutet, June 9-13,
Introduction to Proteomics Åsa Wheelock, Ph.D. Division of Respiratory Medicine & Karolinska Biomics Center asa.wheelock@ki.se In: Systems Biology and the Omics Cascade, Karolinska Institutet, June 9-13,
Laboration 1. Identifiering av proteiner med Mass Spektrometri. Klinisk Kemisk Diagnostik
 Laboration 1 Identifiering av proteiner med Mass Spektrometri Klinisk Kemisk Diagnostik Sven Kjellström 2014 kjellstrom.sven@gmail.com 0702-935060 Laboration 1 Klinisk Kemisk Diagnostik Identifiering av
Laboration 1 Identifiering av proteiner med Mass Spektrometri Klinisk Kemisk Diagnostik Sven Kjellström 2014 kjellstrom.sven@gmail.com 0702-935060 Laboration 1 Klinisk Kemisk Diagnostik Identifiering av
In-Depth Qualitative Analysis of Complex Proteomic Samples Using High Quality MS/MS at Fast Acquisition Rates
 In-Depth Qualitative Analysis of Complex Proteomic Samples Using High Quality MS/MS at Fast Acquisition Rates Using the Explore Workflow on the AB SCIEX TripleTOF 5600 System A major challenge in proteomics
In-Depth Qualitative Analysis of Complex Proteomic Samples Using High Quality MS/MS at Fast Acquisition Rates Using the Explore Workflow on the AB SCIEX TripleTOF 5600 System A major challenge in proteomics
Mass Spectrometry Based Proteomics
 Mass Spectrometry Based Proteomics Proteomics Shared Research Oregon Health & Science University Portland, Oregon This document is designed to give a brief overview of Mass Spectrometry Based Proteomics
Mass Spectrometry Based Proteomics Proteomics Shared Research Oregon Health & Science University Portland, Oregon This document is designed to give a brief overview of Mass Spectrometry Based Proteomics
Error Tolerant Searching of Uninterpreted MS/MS Data
 Error Tolerant Searching of Uninterpreted MS/MS Data 1 In any search of a large LC-MS/MS dataset 2 There are always a number of spectra which get poor scores, or even no match at all. 3 Sometimes, this
Error Tolerant Searching of Uninterpreted MS/MS Data 1 In any search of a large LC-MS/MS dataset 2 There are always a number of spectra which get poor scores, or even no match at all. 3 Sometimes, this
Application Note # LCMS-81 Introducing New Proteomics Acquisiton Strategies with the compact Towards the Universal Proteomics Acquisition Method
 Application Note # LCMS-81 Introducing New Proteomics Acquisiton Strategies with the compact Towards the Universal Proteomics Acquisition Method Introduction During the last decade, the complexity of samples
Application Note # LCMS-81 Introducing New Proteomics Acquisiton Strategies with the compact Towards the Universal Proteomics Acquisition Method Introduction During the last decade, the complexity of samples
ProteinPilot Report for ProteinPilot Software
 ProteinPilot Report for ProteinPilot Software Detailed Analysis of Protein Identification / Quantitation Results Automatically Sean L Seymour, Christie Hunter SCIEX, USA Pow erful mass spectrometers like
ProteinPilot Report for ProteinPilot Software Detailed Analysis of Protein Identification / Quantitation Results Automatically Sean L Seymour, Christie Hunter SCIEX, USA Pow erful mass spectrometers like
SimGlycan Software*: A New Predictive Carbohydrate Analysis Tool for MS/MS Data
 SimGlycan Software*: A New Predictive Carbohydrate Analysis Tool for MS/MS Data Automated Data Interpretation for Glycan Characterization Jenny Albanese 1, Matthias Glueckmann 2 and Christof Lenz 2 1 AB
SimGlycan Software*: A New Predictive Carbohydrate Analysis Tool for MS/MS Data Automated Data Interpretation for Glycan Characterization Jenny Albanese 1, Matthias Glueckmann 2 and Christof Lenz 2 1 AB
NUVISAN Pharma Services
 NUVISAN Pharma Services CESI MS Now available! 1st CRO in Europe! At the highest levels of quality. LABORATORY SERVICES Equipment update STATE OF THE ART AT NUVISAN CESI MS Now available! 1st CRO in Europe!
NUVISAN Pharma Services CESI MS Now available! 1st CRO in Europe! At the highest levels of quality. LABORATORY SERVICES Equipment update STATE OF THE ART AT NUVISAN CESI MS Now available! 1st CRO in Europe!
MaxQuant User s Guide Version 1.2.2.5
 MaxQuant User s Guide Version 1.2.2.5 Jűrgen Cox and Matthias Mann Nature Biotechnology 26, 1367-1372 (2008) Sara ten Have 2012 http://www.lamondlab.com/ http://greproteomics.lifesci.dundee.ac.uk/ References
MaxQuant User s Guide Version 1.2.2.5 Jűrgen Cox and Matthias Mann Nature Biotechnology 26, 1367-1372 (2008) Sara ten Have 2012 http://www.lamondlab.com/ http://greproteomics.lifesci.dundee.ac.uk/ References
ANALYSIS OF PROTEINS AND PROTEOMES BY MASS SPECTROMETRY
 Annu. Rev. Biochem. 2001. 70:437 73 Copyright c 2001 by Annual Reviews. All rights reserved ANALYSIS OF PROTEINS AND PROTEOMES BY MASS SPECTROMETRY Matthias Mann, 1,2 Ronald C. Hendrickson, 2 and Akhilesh
Annu. Rev. Biochem. 2001. 70:437 73 Copyright c 2001 by Annual Reviews. All rights reserved ANALYSIS OF PROTEINS AND PROTEOMES BY MASS SPECTROMETRY Matthias Mann, 1,2 Ronald C. Hendrickson, 2 and Akhilesh
Design considerations for proteomic reference materials
 4220 DOI 10.1002/pmic.201000242 Proteomics 2010, 10, 4220 4225 Design considerations for proteomic reference materials David M. Bunk Analytical Chemistry Division, National Institute of Standards and Technology,
4220 DOI 10.1002/pmic.201000242 Proteomics 2010, 10, 4220 4225 Design considerations for proteomic reference materials David M. Bunk Analytical Chemistry Division, National Institute of Standards and Technology,
Proteomics in general deals with the large-scale
 Mass spectrometry-based proteomics Ruedi Aebersold* & Matthias Mann *Institute for Systems Biology, 1441 North 34th Street, Seattle, Washington 98103-8904, USA (e-mail: raebersold@systemsbiology.org) Center
Mass spectrometry-based proteomics Ruedi Aebersold* & Matthias Mann *Institute for Systems Biology, 1441 North 34th Street, Seattle, Washington 98103-8904, USA (e-mail: raebersold@systemsbiology.org) Center
Introduction to mass spectrometry (MS) based proteomics and metabolomics
 Introduction to mass spectrometry (MS) based proteomics and metabolomics Tianwei Yu Department of Biostatistics and Bioinformatics Rollins School of Public Health Emory University September 10, 2015 Background
Introduction to mass spectrometry (MS) based proteomics and metabolomics Tianwei Yu Department of Biostatistics and Bioinformatics Rollins School of Public Health Emory University September 10, 2015 Background
Tutorial for Proteomics Data Submission. Katalin F. Medzihradszky Robert J. Chalkley UCSF
 Tutorial for Proteomics Data Submission Katalin F. Medzihradszky Robert J. Chalkley UCSF Why Have Guidelines? Large-scale proteomics studies create huge amounts of data. It is impossible/impractical to
Tutorial for Proteomics Data Submission Katalin F. Medzihradszky Robert J. Chalkley UCSF Why Have Guidelines? Large-scale proteomics studies create huge amounts of data. It is impossible/impractical to
AB SCIEX TOF/TOF 4800 PLUS SYSTEM. Cost effective flexibility for your core needs
 AB SCIEX TOF/TOF 4800 PLUS SYSTEM Cost effective flexibility for your core needs AB SCIEX TOF/TOF 4800 PLUS SYSTEM It s just what you expect from the industry leader. The AB SCIEX 4800 Plus MALDI TOF/TOF
AB SCIEX TOF/TOF 4800 PLUS SYSTEM Cost effective flexibility for your core needs AB SCIEX TOF/TOF 4800 PLUS SYSTEM It s just what you expect from the industry leader. The AB SCIEX 4800 Plus MALDI TOF/TOF
Mass Spectrometry Signal Calibration for Protein Quantitation
 Cambridge Isotope Laboratories, Inc. www.isotope.com Proteomics Mass Spectrometry Signal Calibration for Protein Quantitation Michael J. MacCoss, PhD Associate Professor of Genome Sciences University of
Cambridge Isotope Laboratories, Inc. www.isotope.com Proteomics Mass Spectrometry Signal Calibration for Protein Quantitation Michael J. MacCoss, PhD Associate Professor of Genome Sciences University of
Computational analysis of unassigned high-quality MS/MS spectra in proteomic data sets
 2712 DOI 10.1002/pmic.200900473 Proteomics 2010, 10, 2712 2718 TECHNICAL BRIEF Computational analysis of unassigned high-quality MS/MS spectra in proteomic data sets Kang Ning 1, Damian Fermin 1 and Alexey
2712 DOI 10.1002/pmic.200900473 Proteomics 2010, 10, 2712 2718 TECHNICAL BRIEF Computational analysis of unassigned high-quality MS/MS spectra in proteomic data sets Kang Ning 1, Damian Fermin 1 and Alexey
Biopharmaceutical Glycosylation Analysis
 Biopharmaceutical Glycosylation Analysis Glycosylation Analysis: Product Offering Molecular model of erythropoietin with complex N-linked glycans. Courtesy of M.R Wormald and R.A Dwek, Oxford Glycobioloy
Biopharmaceutical Glycosylation Analysis Glycosylation Analysis: Product Offering Molecular model of erythropoietin with complex N-linked glycans. Courtesy of M.R Wormald and R.A Dwek, Oxford Glycobioloy
Correlation of the Mass Spectrometric Analysis of Heat-Treated Glutaraldehyde Preparations to Their 235nm / 280 nm UV Absorbance Ratio
 an ABC Laboratories white paper Correlation of the Mass Spectrometric Analysis of Heat-Treated Glutaraldehyde Preparations to Their 235nm / 280 nm UV Absorbance Ratio A. Sen, R. Dunphy, L. Rosik Analytical
an ABC Laboratories white paper Correlation of the Mass Spectrometric Analysis of Heat-Treated Glutaraldehyde Preparations to Their 235nm / 280 nm UV Absorbance Ratio A. Sen, R. Dunphy, L. Rosik Analytical
Principles of Mass Spectrometry-Based Proteomics. Steven Gygi Department of Cell Biology Harvard Medical School, Boston, MA
 Principles of Mass Spectrometry-Based Proteomics Steven Gygi Department of Cell Biology Harvard Medical School, Boston, MA The Flow of Information The Flow of Information Networks Complexes Proteins Genes
Principles of Mass Spectrometry-Based Proteomics Steven Gygi Department of Cell Biology Harvard Medical School, Boston, MA The Flow of Information The Flow of Information Networks Complexes Proteins Genes
Methods for Protein Analysis
 Methods for Protein Analysis 1. Protein Separation Methods The following is a quick review of some common methods used for protein separation: SDS-PAGE (SDS-polyacrylamide gel electrophoresis) separates
Methods for Protein Analysis 1. Protein Separation Methods The following is a quick review of some common methods used for protein separation: SDS-PAGE (SDS-polyacrylamide gel electrophoresis) separates
Workshop IIc. Manual interpretation of MS/MS spectra. Ebbing de Jong. Center for Mass Spectrometry and Proteomics Phone (612)625-2280 (612)625-2279
 Workshop IIc Manual interpretation of MS/MS spectra Ebbing de Jong Why MS/MS spectra? The information contained in an MS spectrum (m/z, isotope spacing and therefore z ) is not enough to tell us the amino
Workshop IIc Manual interpretation of MS/MS spectra Ebbing de Jong Why MS/MS spectra? The information contained in an MS spectrum (m/z, isotope spacing and therefore z ) is not enough to tell us the amino
Advantages of the LTQ Orbitrap for Protein Identification in Complex Digests
 Application Note: 386 Advantages of the LTQ Orbitrap for Protein Identification in Complex Digests Rosa Viner, Terry Zhang, Scott Peterman, and Vlad Zabrouskov, Thermo Fisher Scientific, San Jose, CA,
Application Note: 386 Advantages of the LTQ Orbitrap for Protein Identification in Complex Digests Rosa Viner, Terry Zhang, Scott Peterman, and Vlad Zabrouskov, Thermo Fisher Scientific, San Jose, CA,
MultiQuant Software 2.0 for Targeted Protein / Peptide Quantification
 MultiQuant Software 2.0 for Targeted Protein / Peptide Quantification Gold Standard for Quantitative Data Processing Because of the sensitivity, selectivity, speed and throughput at which MRM assays can
MultiQuant Software 2.0 for Targeted Protein / Peptide Quantification Gold Standard for Quantitative Data Processing Because of the sensitivity, selectivity, speed and throughput at which MRM assays can
SimGlycan Software*: A New Predictive Carbohydrate Analysis Tool for MS/MS Data
 SimGlycan Software*: A New Predictive Carbohydrate Analysis Tool for MS/MS Data Automated Data Interpretation for Glycan Characterization Jenny Albanese1, Matthias Glueckmann2 and Christof Lenz2 1Applied
SimGlycan Software*: A New Predictive Carbohydrate Analysis Tool for MS/MS Data Automated Data Interpretation for Glycan Characterization Jenny Albanese1, Matthias Glueckmann2 and Christof Lenz2 1Applied
prime Innovation with Integrity The multidimensional path to the Proteome Mass Spectrometry
 prime The multidimensional path to the Proteome Innovation with Integrity Mass Spectrometry Reach for the Full Potential of Proteomics. Open Your Eyes to PRIME The Proteome is far more complex than was
prime The multidimensional path to the Proteome Innovation with Integrity Mass Spectrometry Reach for the Full Potential of Proteomics. Open Your Eyes to PRIME The Proteome is far more complex than was
Quantitative mass spectrometry in proteomics: a critical review
 Anal Bioanal Chem (2007) 389:1017 1031 DOI 10.1007/s00216-007-1486-6 REVIEW Quantitative mass spectrometry in proteomics: a critical review Marcus Bantscheff & Markus Schirle & Gavain Sweetman & Jens Rick
Anal Bioanal Chem (2007) 389:1017 1031 DOI 10.1007/s00216-007-1486-6 REVIEW Quantitative mass spectrometry in proteomics: a critical review Marcus Bantscheff & Markus Schirle & Gavain Sweetman & Jens Rick
BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS
 BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS 1. The Technology Strategy sets out six areas where technological developments are required to push the frontiers of knowledge
BBSRC TECHNOLOGY STRATEGY: TECHNOLOGIES NEEDED BY RESEARCH KNOWLEDGE PROVIDERS 1. The Technology Strategy sets out six areas where technological developments are required to push the frontiers of knowledge
MASS SPECTROMETRY-BASED PROTEOMIC ANALYSIS OF COMPLEX BIOLOGICAL SAMPLES PENG ZHAO. (Under the Direction of Lance Wells) ABSTRACT
 MASS SPECTROMETRY-BASED PROTEOMIC ANALYSIS OF COMPLEX BIOLOGICAL SAMPLES by PENG ZHAO (Under the Direction of Lance Wells) ABSTRACT Proteomics seeks to determine protein structure, modifications, localization,
MASS SPECTROMETRY-BASED PROTEOMIC ANALYSIS OF COMPLEX BIOLOGICAL SAMPLES by PENG ZHAO (Under the Direction of Lance Wells) ABSTRACT Proteomics seeks to determine protein structure, modifications, localization,
A Tool To Visualize and Evaluate Data Obtained by Liquid Chromatography-Electrospray Ionization-Mass Spectrometry
 Anal. Chem. 2004, 76, 3856-3860 A Tool To Visualize and Evaluate Data Obtained by Liquid Chromatography-Electrospray Ionization-Mass Spectrometry Xiao-jun Li,* Patrick G. A. Pedrioli, Jimmy Eng, Dan Martin,
Anal. Chem. 2004, 76, 3856-3860 A Tool To Visualize and Evaluate Data Obtained by Liquid Chromatography-Electrospray Ionization-Mass Spectrometry Xiao-jun Li,* Patrick G. A. Pedrioli, Jimmy Eng, Dan Martin,
Principles and Applications of Proteomics
 Principles and Applications of Proteomics Why Proteomics? 2-DE Overview Sample preparation 1 st & 2 nd dimension seperation Data Analysis Sample preparation for Mass Spectrometry Mass Spectrometry MALDI-TOF,
Principles and Applications of Proteomics Why Proteomics? 2-DE Overview Sample preparation 1 st & 2 nd dimension seperation Data Analysis Sample preparation for Mass Spectrometry Mass Spectrometry MALDI-TOF,
Retrospective Analysis of a Host Cell Protein Perfect Storm: Identifying Immunogenic Proteins and Fixing the Problem
 Retrospective Analysis of a Host Cell Protein Perfect Storm: Identifying Immunogenic Proteins and Fixing the Problem Kevin Van Cott, Associate Professor Dept. of Chemical and Biomolecular Engineering Nebraska
Retrospective Analysis of a Host Cell Protein Perfect Storm: Identifying Immunogenic Proteins and Fixing the Problem Kevin Van Cott, Associate Professor Dept. of Chemical and Biomolecular Engineering Nebraska
Brochure More information from http://www.researchandmarkets.com/reports/2172993/
 Brochure More information from http://www.researchandmarkets.com/reports/2172993/ Proteomics Today. Protein Assessment and Biomarkers Using Mass Spectrometry, 2D Electrophoresis,and Microarray Technology.
Brochure More information from http://www.researchandmarkets.com/reports/2172993/ Proteomics Today. Protein Assessment and Biomarkers Using Mass Spectrometry, 2D Electrophoresis,and Microarray Technology.
Overview. Introduction. AB SCIEX MPX -2 High Throughput TripleTOF 4600 LC/MS/MS System
 Investigating the use of the AB SCIEX TripleTOF 4600 LC/MS/MS System for High Throughput Screening of Synthetic Cannabinoids/Metabolites in Human Urine AB SCIEX MPX -2 High Throughput TripleTOF 4600 LC/MS/MS
Investigating the use of the AB SCIEX TripleTOF 4600 LC/MS/MS System for High Throughput Screening of Synthetic Cannabinoids/Metabolites in Human Urine AB SCIEX MPX -2 High Throughput TripleTOF 4600 LC/MS/MS
Contents. Preface to the Second Edition List of Contributors XIII
 Contents Preface to the Second Edition List of Contributors XIII XI 1 The MALDI Process and Method 1 Franz Hillenkamp, Thorsten W. Jaskolla, and Michael Karas 1.1 Introduction 1 1.2 Analyte Incorporation
Contents Preface to the Second Edition List of Contributors XIII XI 1 The MALDI Process and Method 1 Franz Hillenkamp, Thorsten W. Jaskolla, and Michael Karas 1.1 Introduction 1 1.2 Analyte Incorporation
Evaluation of LC-MS data for the absolute quantitative analysis of marker proteins
 Evaluation of LC-MS data for the absolute quantitative analysis of marker proteins Nathanaël Delmotte 1, Bettina Mayr 1, Andreas Leinenbach 1, Knut Reinert 2, Oliver Kohlbacher 3, Christoph Klein 4, and
Evaluation of LC-MS data for the absolute quantitative analysis of marker proteins Nathanaël Delmotte 1, Bettina Mayr 1, Andreas Leinenbach 1, Knut Reinert 2, Oliver Kohlbacher 3, Christoph Klein 4, and
MRMPilot Software: Accelerating MRM Assay Development for Targeted Quantitative Proteomics
 MRMPilot Software: Accelerating MRM Assay Development for Targeted Quantitative Proteomics With Unique QTRAP and TripleTOF 5600 System Technology Targeted peptide quantification is a rapidly growing application
MRMPilot Software: Accelerating MRM Assay Development for Targeted Quantitative Proteomics With Unique QTRAP and TripleTOF 5600 System Technology Targeted peptide quantification is a rapidly growing application
LC-MS/MS for Chromatographers
 LC-MS/MS for Chromatographers An introduction to the use of LC-MS/MS, with an emphasis on the analysis of drugs in biological matrices LC-MS/MS for Chromatographers An introduction to the use of LC-MS/MS,
LC-MS/MS for Chromatographers An introduction to the use of LC-MS/MS, with an emphasis on the analysis of drugs in biological matrices LC-MS/MS for Chromatographers An introduction to the use of LC-MS/MS,
Increasing the Multiplexing of High Resolution Targeted Peptide Quantification Assays
 Increasing the Multiplexing of High Resolution Targeted Peptide Quantification Assays Scheduled MRM HR Workflow on the TripleTOF Systems Jenny Albanese, Christie Hunter AB SCIEX, USA Targeted quantitative
Increasing the Multiplexing of High Resolution Targeted Peptide Quantification Assays Scheduled MRM HR Workflow on the TripleTOF Systems Jenny Albanese, Christie Hunter AB SCIEX, USA Targeted quantitative
Proteomics: the first decade and beyond
 Proteomics: the first decade and beyond review Scott D. Patterson 1,3 & Ruedi H. Aebersold 2 doi:10.1038/ng1106 Proteomics is the systematic study of the many and diverse properties of proteins in a parallel
Proteomics: the first decade and beyond review Scott D. Patterson 1,3 & Ruedi H. Aebersold 2 doi:10.1038/ng1106 Proteomics is the systematic study of the many and diverse properties of proteins in a parallel
for mass spectrometry calibration tools Thermo Scientific Pierce Controls and Standards for Mass Spectrometry
 Thermo Scientific Pierce Controls and Standards for Mass Spectrometry calibration tools for mass spectrometry Ensure confidence in instrument performance with Thermo Scientific Pierce Calibration Solutions
Thermo Scientific Pierce Controls and Standards for Mass Spectrometry calibration tools for mass spectrometry Ensure confidence in instrument performance with Thermo Scientific Pierce Calibration Solutions
Shotgun Proteomic Analysis. Department of Cell Biology The Scripps Research Institute
 Shotgun Proteomic Analysis Department of Cell Biology The Scripps Research Institute Biological/Functional Resolution of Experiments Organelle Multiprotein Complex Cells/Tissues Function Expression Analysis
Shotgun Proteomic Analysis Department of Cell Biology The Scripps Research Institute Biological/Functional Resolution of Experiments Organelle Multiprotein Complex Cells/Tissues Function Expression Analysis
Application Note # LCMS-62 Walk-Up Ion Trap Mass Spectrometer System in a Multi-User Environment Using Compass OpenAccess Software
 Application Note # LCMS-62 Walk-Up Ion Trap Mass Spectrometer System in a Multi-User Environment Using Compass OpenAccess Software Abstract Presented here is a case study of a walk-up liquid chromatography
Application Note # LCMS-62 Walk-Up Ion Trap Mass Spectrometer System in a Multi-User Environment Using Compass OpenAccess Software Abstract Presented here is a case study of a walk-up liquid chromatography
PROTEIN SEQUENCING. First Sequence
 PROTEIN SEQUENCING First Sequence The first protein sequencing was achieved by Frederic Sanger in 1953. He determined the amino acid sequence of bovine insulin Sanger was awarded the Nobel Prize in 1958
PROTEIN SEQUENCING First Sequence The first protein sequencing was achieved by Frederic Sanger in 1953. He determined the amino acid sequence of bovine insulin Sanger was awarded the Nobel Prize in 1958
Fast and Automatic Mapping of Disulfide Bonds in a Monoclonal Antibody using SYNAPT G2 HDMS and BiopharmaLynx 1.3
 Fast and Automatic Mapping of Disulfide Bonds in a Monoclonal Antibody using SYNAPT G2 HDMS and BiopharmaLynx 1.3 Hongwei Xie and Weibin Chen Waters Corporation, Milford, MA, U.S. A P P L I C AT ION B
Fast and Automatic Mapping of Disulfide Bonds in a Monoclonal Antibody using SYNAPT G2 HDMS and BiopharmaLynx 1.3 Hongwei Xie and Weibin Chen Waters Corporation, Milford, MA, U.S. A P P L I C AT ION B
Quan%ta%ve proteomics. Maarten Altelaar, 2014
 Quan%ta%ve proteomics Maarten Altelaar, 2014 Proteomics Altelaar et al. Nat Rev Gen 14, 2013, 35-48 Quan%ta%ve proteomics Quan%ta%ve proteomics Control Diseased, s%mulated, Knock down, etc. How quan%ta%ve
Quan%ta%ve proteomics Maarten Altelaar, 2014 Proteomics Altelaar et al. Nat Rev Gen 14, 2013, 35-48 Quan%ta%ve proteomics Quan%ta%ve proteomics Control Diseased, s%mulated, Knock down, etc. How quan%ta%ve
Chapter 3. Protein Structure and Function
 Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Chapter 3 Protein Structure and Function Broad functional classes So Proteins have structure and function... Fine! -Why do we care to know more???? Understanding functional architechture gives us POWER
Quantitative mass spec based proteomics
 Quantitative mass spec based proteomics Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi Proteomics is the large-scale study of proteins Proteomics provides information on: -protein expression
Quantitative mass spec based proteomics Tuula Nyman Institute of Biotechnology tuula.nyman@helsinki.fi Proteomics is the large-scale study of proteins Proteomics provides information on: -protein expression
Effects of Intelligent Data Acquisition and Fast Laser Speed on Analysis of Complex Protein Digests
 Effects of Intelligent Data Acquisition and Fast Laser Speed on Analysis of Complex Protein Digests AB SCIEX TOF/TOF 5800 System with DynamicExit Algorithm and ProteinPilot Software for Robust Protein
Effects of Intelligent Data Acquisition and Fast Laser Speed on Analysis of Complex Protein Digests AB SCIEX TOF/TOF 5800 System with DynamicExit Algorithm and ProteinPilot Software for Robust Protein
Mass spectrometry-based quantitative proteomic profiling Wei Yan and Sharon S. Chen Date received: 2nd December 2004
 Wei Yan is a research scientist at the Institute for Systems Biology and in the Department of Pathology at the University of Washington. Sharon S. Chen is a graduate student in the Department of Bioengineering
Wei Yan is a research scientist at the Institute for Systems Biology and in the Department of Pathology at the University of Washington. Sharon S. Chen is a graduate student in the Department of Bioengineering
Enzyme Response Profiling: Integrating proteomics and genomics with xenobiotic metabolism and cytotoxicity
 Enzyme Response Profiling: Integrating proteomics and genomics with xenobiotic metabolism and cytotoxicity Proteomics high throughput separation and characterization of expressed proteins in tissue and
Enzyme Response Profiling: Integrating proteomics and genomics with xenobiotic metabolism and cytotoxicity Proteomics high throughput separation and characterization of expressed proteins in tissue and
H H N - C - C 2 R. Three possible forms (not counting R group) depending on ph
 Amino acids - 0 common amino acids there are others found naturally but much less frequently - Common structure for amino acid - C, -N, and functional groups all attached to the alpha carbon N - C - C
Amino acids - 0 common amino acids there are others found naturally but much less frequently - Common structure for amino acid - C, -N, and functional groups all attached to the alpha carbon N - C - C
MALDI Mass Spectrometry for Quantitative Proteomics Approaches, Scopes and Limitations
 MALDI Mass Spectrometry for Quantitative Proteomics Approaches, Scopes and Limitations Andreas Tholey Technische Biochemie, Universität des Saarlandes, 66123 Saarbrücken, Germany e-mail: a.tholey@mx.uni-saarland.de
MALDI Mass Spectrometry for Quantitative Proteomics Approaches, Scopes and Limitations Andreas Tholey Technische Biochemie, Universität des Saarlandes, 66123 Saarbrücken, Germany e-mail: a.tholey@mx.uni-saarland.de
Research-grade Targeted Proteomics Assay Development: PRMs for PTM Studies with Skyline or, How I learned to ditch the triple quad and love the QE
 Research-grade Targeted Proteomics Assay Development: PRMs for PTM Studies with Skyline or, How I learned to ditch the triple quad and love the QE Jacob D. Jaffe Skyline Webinar July 2015 Proteomics and
Research-grade Targeted Proteomics Assay Development: PRMs for PTM Studies with Skyline or, How I learned to ditch the triple quad and love the QE Jacob D. Jaffe Skyline Webinar July 2015 Proteomics and
Histone modifications. and ChIP. G. Valle - Università di Padova
 Histone modifications and ChIP Histone acetylation It has been shown that the acetylation of lysines is highly dynamic and regulated by the opposing action of two families of enzymes, histone acetyltransferases
Histone modifications and ChIP Histone acetylation It has been shown that the acetylation of lysines is highly dynamic and regulated by the opposing action of two families of enzymes, histone acetyltransferases
DRUG METABOLISM. Drug discovery & development solutions FOR DRUG METABOLISM
 DRUG METABLISM Drug discovery & development solutions FR DRUG METABLISM Fast and efficient metabolite identification is critical in today s drug discovery pipeline. The goal is to achieve rapid structural
DRUG METABLISM Drug discovery & development solutions FR DRUG METABLISM Fast and efficient metabolite identification is critical in today s drug discovery pipeline. The goal is to achieve rapid structural
Pesticide Analysis by Mass Spectrometry
 Pesticide Analysis by Mass Spectrometry Purpose: The purpose of this assignment is to introduce concepts of mass spectrometry (MS) as they pertain to the qualitative and quantitative analysis of organochlorine
Pesticide Analysis by Mass Spectrometry Purpose: The purpose of this assignment is to introduce concepts of mass spectrometry (MS) as they pertain to the qualitative and quantitative analysis of organochlorine
In-depth Analysis of Tandem Mass Spectrometry Data from Disparate Instrument Types* S
 Research In-depth Analysis of Tandem Mass Spectrometry Data from Disparate Instrument Types* S Robert J. Chalkley, Peter R. Baker, Katalin F. Medzihradszky, Aenoch J. Lynn, and A. L. Burlingame Mass spectrometric
Research In-depth Analysis of Tandem Mass Spectrometry Data from Disparate Instrument Types* S Robert J. Chalkley, Peter R. Baker, Katalin F. Medzihradszky, Aenoch J. Lynn, and A. L. Burlingame Mass spectrometric
THE ABC S (AND XYZ S) OF PEPTIDE SEQUENCING
 TE ABC S (AD XYZ S) F PEPTIDE SEQUECIG anno Steen* and Matthias Mann Abstract Proteomics is an increasingly powerful and indispensable technology in molecular cell biology. It can be used to identify the
TE ABC S (AD XYZ S) F PEPTIDE SEQUECIG anno Steen* and Matthias Mann Abstract Proteomics is an increasingly powerful and indispensable technology in molecular cell biology. It can be used to identify the
Overview. Triple quadrupole (MS/MS) systems provide in comparison to single quadrupole (MS) systems: Introduction
 Advantages of Using Triple Quadrupole over Single Quadrupole Mass Spectrometry to Quantify and Identify the Presence of Pesticides in Water and Soil Samples André Schreiber AB SCIEX Concord, Ontario (Canada)
Advantages of Using Triple Quadrupole over Single Quadrupole Mass Spectrometry to Quantify and Identify the Presence of Pesticides in Water and Soil Samples André Schreiber AB SCIEX Concord, Ontario (Canada)
Simultaneous qualitative and quantitative analysis using the Agilent 6540 Accurate-Mass Q-TOF
 Simultaneous qualitative and quantitative analysis using the Agilent 654 Accurate-Mass Q-TOF Technical Overview Authors Pat Perkins Anabel Fandino Lester Taylor Agilent Technologies, Inc. Santa Clara,
Simultaneous qualitative and quantitative analysis using the Agilent 654 Accurate-Mass Q-TOF Technical Overview Authors Pat Perkins Anabel Fandino Lester Taylor Agilent Technologies, Inc. Santa Clara,
Quantitative proteomics background
 Proteomics data analysis seminar Quantitative proteomics and transcriptomics of anaerobic and aerobic yeast cultures reveals post transcriptional regulation of key cellular processes de Groot, M., Daran
Proteomics data analysis seminar Quantitative proteomics and transcriptomics of anaerobic and aerobic yeast cultures reveals post transcriptional regulation of key cellular processes de Groot, M., Daran
Introduction to Database Searching using MASCOT
 Introduction to Database Searching using MASCOT 1 Three ways to use mass spectrometry data for protein identification 1.Peptide Mass Fingerprint A set of peptide molecular masses from an enzyme digest
Introduction to Database Searching using MASCOT 1 Three ways to use mass spectrometry data for protein identification 1.Peptide Mass Fingerprint A set of peptide molecular masses from an enzyme digest
OpenMS A Framework for Quantitative HPLC/MS-Based Proteomics
 OpenMS A Framework for Quantitative HPLC/MS-Based Proteomics Knut Reinert 1, Oliver Kohlbacher 2,Clemens Gröpl 1, Eva Lange 1, Ole Schulz-Trieglaff 1,Marc Sturm 2 and Nico Pfeifer 2 1 Algorithmische Bioinformatik,
OpenMS A Framework for Quantitative HPLC/MS-Based Proteomics Knut Reinert 1, Oliver Kohlbacher 2,Clemens Gröpl 1, Eva Lange 1, Ole Schulz-Trieglaff 1,Marc Sturm 2 and Nico Pfeifer 2 1 Algorithmische Bioinformatik,
Mass spectrometry based proteomics
 Practice-oriented, student-friendly modernization of the biomedical education for strengthening the international competitiveness of the rural Hungarian universities TÁMOP-4.1.1.C-13/1/KONV-2014-0001 Mass
Practice-oriented, student-friendly modernization of the biomedical education for strengthening the international competitiveness of the rural Hungarian universities TÁMOP-4.1.1.C-13/1/KONV-2014-0001 Mass
Accurate calibration of on-line Time of Flight Mass Spectrometer (TOF-MS) for high molecular weight combustion product analysis
 Accurate calibration of on-line Time of Flight Mass Spectrometer (TOF-MS) for high molecular weight combustion product analysis B. Apicella*, M. Passaro**, X. Wang***, N. Spinelli**** mariadellarcopassaro@gmail.com
Accurate calibration of on-line Time of Flight Mass Spectrometer (TOF-MS) for high molecular weight combustion product analysis B. Apicella*, M. Passaro**, X. Wang***, N. Spinelli**** mariadellarcopassaro@gmail.com
A Quadrupole-Orbitrap Hybrid Mass Spectrometer Offers Highest Benchtop Performance for In-Depth Analysis of Complex Proteomes
 A Quadrupole-Orbitrap Hybrid Mass Spectrometer Offers Highest Benchtop Performance for In-Depth Analysis of Complex Proteomes Zhiqi Hao 1, Yi Zhang 1, Shannon Eliuk 1, Justin Blethrow 1, Dave Horn 1, Vlad
A Quadrupole-Orbitrap Hybrid Mass Spectrometer Offers Highest Benchtop Performance for In-Depth Analysis of Complex Proteomes Zhiqi Hao 1, Yi Zhang 1, Shannon Eliuk 1, Justin Blethrow 1, Dave Horn 1, Vlad
Industry Perspective: Advantages of Open Access and Walkup LC/ MS Supporting Protein Drug Discovery and Development
 Industry Perspective: Advantages of Open Access and Walkup LC/ MS Supporting Protein Drug Discovery and Development Dawn Stickle, Agilent Technologies Originally presented by Eric Fang, Novartis Overview
Industry Perspective: Advantages of Open Access and Walkup LC/ MS Supporting Protein Drug Discovery and Development Dawn Stickle, Agilent Technologies Originally presented by Eric Fang, Novartis Overview
GENERAL UNKNOWN SCREENING FOR DRUGS IN BIOLOGICAL SAMPLES BY LC/MS Luc Humbert1, Michel Lhermitte 1, Frederic Grisel 2 1
 GENERAL UNKNOWN SCREENING FOR DRUGS IN BIOLOGICAL SAMPLES BY LC/MS Luc Humbert, Michel Lhermitte, Frederic Grisel Laboratoire de Toxicologie & Génopathologie, CHRU Lille, France Waters Corporation, Guyancourt,
GENERAL UNKNOWN SCREENING FOR DRUGS IN BIOLOGICAL SAMPLES BY LC/MS Luc Humbert, Michel Lhermitte, Frederic Grisel Laboratoire de Toxicologie & Génopathologie, CHRU Lille, France Waters Corporation, Guyancourt,
Preprocessing, Management, and Analysis of Mass Spectrometry Proteomics Data
 Preprocessing, Management, and Analysis of Mass Spectrometry Proteomics Data M. Cannataro, P. H. Guzzi, T. Mazza, and P. Veltri Università Magna Græcia di Catanzaro, Italy 1 Introduction Mass Spectrometry
Preprocessing, Management, and Analysis of Mass Spectrometry Proteomics Data M. Cannataro, P. H. Guzzi, T. Mazza, and P. Veltri Università Magna Græcia di Catanzaro, Italy 1 Introduction Mass Spectrometry
Mass spectrometry. Dr. Kevin R. Tucker
 Mass spectrometry Dr. Kevin R. Tucker What is mass spectrometry? A versatile analytical technique used to determine the composition of chemical samples either at the atomic or molecular level. Ionic forms
Mass spectrometry Dr. Kevin R. Tucker What is mass spectrometry? A versatile analytical technique used to determine the composition of chemical samples either at the atomic or molecular level. Ionic forms
Molecular Cell Biology. Prof. D. Karunagaran. Department of Biotechnology. Indian Institute of Technology Madras
 Molecular Cell Biology Prof. D. Karunagaran Department of Biotechnology Indian Institute of Technology Madras Module 5 Methods in Cell Biology (Methods to Manipulate Protein, DNA and RNA and Methods to
Molecular Cell Biology Prof. D. Karunagaran Department of Biotechnology Indian Institute of Technology Madras Module 5 Methods in Cell Biology (Methods to Manipulate Protein, DNA and RNA and Methods to
Bruker ToxScreener TM. Innovation with Integrity. A Comprehensive Screening Solution for Forensic Toxicology UHR-TOF MS
 Bruker ToxScreener TM A Comprehensive Screening Solution for Forensic Toxicology Innovation with Integrity UHR-TOF MS ToxScreener - Get the Complete Picture Forensic laboratories are frequently required
Bruker ToxScreener TM A Comprehensive Screening Solution for Forensic Toxicology Innovation with Integrity UHR-TOF MS ToxScreener - Get the Complete Picture Forensic laboratories are frequently required
Integrated Data Mining Strategy for Effective Metabolomic Data Analysis
 The First International Symposium on Optimization and Systems Biology (OSB 07) Beijing, China, August 8 10, 2007 Copyright 2007 ORSC & APORC pp. 45 51 Integrated Data Mining Strategy for Effective Metabolomic
The First International Symposium on Optimization and Systems Biology (OSB 07) Beijing, China, August 8 10, 2007 Copyright 2007 ORSC & APORC pp. 45 51 Integrated Data Mining Strategy for Effective Metabolomic
Victims Compensation Claim Status of All Pending Claims and Claims Decided Within the Last Three Years
 Claim#:021914-174 Initials: J.T. Last4SSN: 6996 DOB: 5/3/1970 Crime Date: 4/30/2013 Status: Claim is currently under review. Decision expected within 7 days Claim#:041715-334 Initials: M.S. Last4SSN: 2957
Claim#:021914-174 Initials: J.T. Last4SSN: 6996 DOB: 5/3/1970 Crime Date: 4/30/2013 Status: Claim is currently under review. Decision expected within 7 days Claim#:041715-334 Initials: M.S. Last4SSN: 2957
Mass spectrometry based proteomics, background, status and future needs
 Protein Cell 2012, 3(9): 641 647 DOI 10.1007/s13238-012-2079-5 MINI-REVIEW Mass spectrometry based proteomics, background, status and future needs Peter Roepstorff Department of Biochemistry and Molecular
Protein Cell 2012, 3(9): 641 647 DOI 10.1007/s13238-012-2079-5 MINI-REVIEW Mass spectrometry based proteomics, background, status and future needs Peter Roepstorff Department of Biochemistry and Molecular
Rheumatoid arthritis (RA) and osteoarthritis. Proteomics: Applications to the Study of Rheumatoid Arthritis and Osteoarthritis
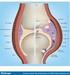 Perspectives on Modern Orthopaedics Proteomics: Applications to the Study of Rheumatoid Arthritis and Osteoarthritis Reuben Gobezie, MD Peter J. Millett, MD, MSc David S. Sarracino, PhD Christopher Evans,
Perspectives on Modern Orthopaedics Proteomics: Applications to the Study of Rheumatoid Arthritis and Osteoarthritis Reuben Gobezie, MD Peter J. Millett, MD, MSc David S. Sarracino, PhD Christopher Evans,
CONTENTS. PART I INSTRUMENTATION 1 1 DEFINITIONS AND EXPLANATIONS 3 Ann Westman-Brinkmalm and Gunnar Brinkmalm References 13
 CONTENTS FOREWORD CONTRIBUTORS xiii xv PART I INSTRUMENTATION 1 1 DEFINITIONS AND EXPLANATIONS 3 Ann Westman-Brinkmalm and Gunnar Brinkmalm References 13 2 A MASS SPECTROMETER S BUILDING BLOCKS 15 Ann
CONTENTS FOREWORD CONTRIBUTORS xiii xv PART I INSTRUMENTATION 1 1 DEFINITIONS AND EXPLANATIONS 3 Ann Westman-Brinkmalm and Gunnar Brinkmalm References 13 2 A MASS SPECTROMETER S BUILDING BLOCKS 15 Ann
Contents. 1 Introduction 1-7
 Contents 1 Introduction 1-7 2 Background: Mass Spectrometry, Database Systems and ASP.NET 8-19 2.1 Mass Spectrometry based Proteomics... 8-11 2.2 Database Systems... 12-16 2.2.1 Components and Functions
Contents 1 Introduction 1-7 2 Background: Mass Spectrometry, Database Systems and ASP.NET 8-19 2.1 Mass Spectrometry based Proteomics... 8-11 2.2 Database Systems... 12-16 2.2.1 Components and Functions
--not necessarily a protein! (all proteins are polypeptides, but the converse is not true)
 00Note Set 5b 1 PEPTIDE BONDS AND POLYPEPTIDES OLIGOPEPTIDE: --chain containing only a few amino acids (see tetrapaptide, Fig 5.9) POLYPEPTIDE CHAINS: --many amino acids joined together --not necessarily
00Note Set 5b 1 PEPTIDE BONDS AND POLYPEPTIDES OLIGOPEPTIDE: --chain containing only a few amino acids (see tetrapaptide, Fig 5.9) POLYPEPTIDE CHAINS: --many amino acids joined together --not necessarily
