DNA-based Fixed Gain Amplifiers and Linear Classifier Circuits. David Yu Zhang 1 and Georg Seelig 2. dzhang@dna.caltech.edu,gseelig@u.washington.
|
|
|
- Pierce Jordan
- 8 years ago
- Views:
Transcription
1 DNA-based Fixed Gain Amplifiers and Linear Classifier Circuits David Yu Zhang and Georg Seelig California Institute of Technology, Pasadena, CA, USA University of Washington, Seattle, WA, USA Abstract. DNA catalysts have been developed as methods of amplifying single-stranded nucleic acid signals. The maximum turnover (gain) of these systems, however, often varies based on strand and complex purities, and has so far not been well-controlled. Here we introduce methods for controlling the asymptotic turnover of strand displacement-based DNA catalysts and show how these could be used to construct linear classifier systems. DNA nanotechnology has utilized the specific binding properties and the well-understood thermodynamics [] and kinetics [, 3] of nucleic acid strand displacement reactions to construct dynamic cascaded reactions, such as logic gates and circuits [,,,9], motors [,], and amplification mechanisms [7, 3, 5, 3,, 9, ]. DNA devices can operate in complex biochemical environments and can be programmed to specifically interact with biological nucleic acids such as messenger RNA or microrna. DNA circuits could be used to develop novel point-of-care diagnostic devices that integrate detection with analysis and do not require complex laboratory equipment. It has even been suggested [, 5, ] to use DNA devices as smart therapeutics that operate inside living cells and integrate detection of specific disease markers with the activation of a therapeutic response based on the RNA interference pathway [7, 8], on antisense oligonucleotides [] or ribozymes. Such applications require nucleic acid circuitry that can reliably identify a specific disease state. Characteristic RNA markers that could serve as inputs to a DNA analytic circuit have been identified for many diseases. However, it is often not sufficient to simply detect the presence or absence of a set of RNA markers. Instead, the classifiers that distinguishes a disease tissue from healthy tissue (or other disease tissues) are often complex functions of the concentrations of multiple RNA markers (see Refs. [, [RNA ] Tissue type Tissue type [RNA ] Fig. : Sketch of a hypothetical twogene classifier. Samples from two different tissue can be clearly distinguished based on the expression profiles of two RNA molecules. 8,] for examples of microrna-expression based classifiers of varying complexity). Here we propose a molecular implementation for a specific class of classifiers, namely linear classifiers. The classifier circuit computes a linear combination with arbitrary (positive of negative) weights on a set of inputs (i.e. RNA concentrations) and compares the result to a threshold value. Fig. shows a highly simplified sketch of a linear two-gene classifier: The line separating the two different tissue types is given by an equation of the form α [RNA ] + α [RNA ] = K. Given a sample of unknown origin, we can now classify it as tissue type or based on a measurement of two RNAs. Unlike in the more conventional case where the expressions of each RNA is individually measured and the the linear classication analysis in performed in silico, here both detection and analysis are done on the molecular level, allowing in situ and in vivo applications.
2 Previous DNA logic circuits were mostly designed for a situation where inputs can be represented as Boolean variables and are either present at a high concentration or completely absent [, 9]. This does not necessarily require the original inputs to be at a specific level; DNA-based signal restoration units consisting of a threshold gate and an amplifier can be used to restore an input with an arbitrary concentration to the expected logical TRUE or FALSE values. Still, the digital nature of such circuits is inherently incompatible with classication problems, in which the relative amounts of inputs determines the value of the final output. The fixed gain amplication methods presented here allows a reliable method of tuning analog sigals encoded in the concentrations of nucleic acids. Fixed gain amplifiers: lowering catalytic turnover. One key component of the proposed linear classifier is a DNA-based catalytic amplifier, that allows one signal-stranded nucleic acid to specifically produce or release many single-stranded nucleic acid molecules of independent sequence. Importantly, this amplifier needs to have a finite and controllable gain α such that each input on average releases α copies of the output. Such a finite gain amplifier would be useful not only in a linear classifier, where each detected RNA species is assigned a different weight but could also be used for a pre-amplification of a set of low-concentration inputs while maintaining their relative concentrations. Existing DNA amplifiers have intrinsically finite turnover; strand displacement-based nucleic acid catalysts typically convert on the order of - substrates before being inactivated [3, ]. Inactivation is most likely due to irreversible binding of the catalyst to defective substrate complexes or fuels [3,, ]. The details of the inactivation process depend on the specifics of the amplifier design, but it is likely that imperfectly synthesized DNA strands are a major culprit. In practice, the maximal turnover obtained seems to depend strongly on sequence, purification procedures, strand orientation and similar experimental and design details []. Therefore, while the gain is finite, it can be characterized for any particular system. The question then becomes we can lower the turnover controllably to a fixed value, starting from a arbitrary but high turnover. Given the intrinsic turnover of a catalytic system, it seems intuitively clear that we can lower the turnover further either 8P / S k a/ k b = 5 k a/ k b = k / k = a b C / D Fig. : Output produced in a catalytic reaction with competitive inhibition as a function of the catalyst concentration. Final product P is scaled by initial concentration of substrate S. Initial catalyst concentration C is measured in units of inhibitor concentration I. We obtain different I/O characteristics depending on the ratio between the rate constants k and k for the catalytic and the competitive reaction. by increasing the fraction of imperfect substrate or through addition of an alternative competitive inhibitor that irreversibly binds to the catalyst. However, it may be less intuitive how to best adjust the turnover to any specific desired value. To address the question of how to control turnover we first consider a simple model for a catalytic reaction with competitive inhibition and then turn to simulating a specific DNA implementation using measured reaction parameters. A catalytic reaction in the presence of an impurity can be modeled as C + S ka C + P, () C + D k b. () In the first reaction a catalyst C transforms a substrate S into a product P. The rate constant for this reaction is k a. The catalyst can also participate in a second, unproductive reaction with an inhibitor (or damper) D. This reaction proceeds at a rate constant k b.
3 The differential equations resulting from this model can be integrated with initial conditions C() = C, S() = S, D() = D and P() =. Solving for the product P(t) we get ( ) ka/k ρ b P(t) = S S ρe k, (3) b t where we introduced the ratio ρ = C /D and the difference = C D of the initial amounts of catalyst and inhibitor. In an ideal system without competitive inhibition the final product concentration is always equal to the initial concentration of substrate. Given enough time the catalyst will convert all substrate into product. In a system with competitive inhibition this is not necessarily true. The final amount of product produced in that case can be computed by taking the limit t in Eq. 3: { lim P(t) = P S, C D = t S S ( ρ) ka/k b (), C < D Not surprisingly, if we start out with more catalyst than inhibitor, the reaction will eventually go to completion. The opposite limit is more interesting. First, consider the case where the rate for the catalytic reaction is much faster than the inhibition reaction, k a > k b (blue trace in Fig. ). In this case the inhibitor has a relatively minor effect that is most pronounced at low concentrations of catalyst compared to the inhibitor. In the limit where the catalytic reaction occurs at exactly the same rate as the inhibitory reaction, i.e. k a = k b (red trace in Fig. ) Eq. predicts that the final amount of product is linear in the initial amount of catalyst, i.e. P = αc where α = S /D. That is, by adjusting the relative concentration of substrate to inhibitor we can get any finite gain we need. The situation where the rate for the inhibitor reaction is faster than the rate for the catalytic reaction is also interesting. In that case, the amount or product is sub-linear in the initial amount of catalyst for C < D but reaches a fixed value S in the opposite regime. The concentration of the competitive inhibitor I therefore acts as a threshold for the catalytic reaction. Such a threshold element is useful for reliable signal propagation for example in the context of chemical digital circuits. We now turn to a specific DNA implementation of such a system. Our implementation is based on the entropy-driven catalytic amplifier of Ref. [] which S + C k k I + SP I + F k I + OP I k 3 k W + C S + F k OP + SP + W I + Fb k X + OP S + Fb k OP + SP + W C + W k X C + D k X A + C k k IA + SP IA + F k I + C I k 3 k W + C A + F k C + SP + W IA + Fb k X + C k = 5 M s k =.7 5 M s k =. M s k 3 =. s k rep = 5 M s A + Fb k C + SP + W Table : Reactions simulated in Fig.. was further characterized in Ref. []. Turnover for this amplifier was measured to be about. The reaction mechanism for this system, including the side reactions leading to intrinsically finite turnover, is shown in Fig. 3 (A). As a competitive inhibitor we here propose to use a damper DNA gate that irreversibly binds the catalytic input (Fig. 3 (B)). In order to match the reaction rate constants of the catalyst with this inhibitor to that of the catalyst with the active substrate we simply choose the toeholds for both reactions to be identical. In order to verify the predictions from our simple model Eq. we simulated the full catalytic system of Ref. [] with a parallel inhibitory reaction using the measured rate constants and reaction intermediates. The model is given in Table and resulting data is shown in Fig (A). As expected from our model the final fluorescence depends linearly on the concentration of damper gate.
4 A W (Waste) Start Here C (Catalyst) S (Substrate) B C (Catalyst) D (Damper) Intermediate I SP (Side Product) Intermediate I d X (Unreactive product) + 5 OP (Output Product) 3 F (Fuel) 3 d Fb (Bad Fuel) OP 5 Unreactive products Fig. 3: Methods for tuning catalytic turnover. (A) DNA amplification via catalysis, adapted from Zhang et al. []. Catalyst strand C reacts with S to form side product SP and intermediate I, the latter of which subsequently reacts with F to release output product OP, waste W, and catalyst C. However, a small fraction of bad fuel with deletions and/or degradation near the 3 end, denoted as Fb, will bind to intermediate I to form an unreactive product X, thus permanently trapping catalyst C and reducing the [F b] observed catalytic turnover of the reaction. The ratio was estimated to be. for HPLC-purified [F]+[F b] fuel strands []. (B) The catalytic turnover of the reaction can be tuned to be lower via the addition of the damping complexes D. Because C binds by the toehold to D as to S, it is assumed that this rate constant is identical in value to that of k. Fixed gain amplifiers: increasing catalytic turnover. The turnover of a catalytic reaction can be increased above the intrinsic limit set by defective oligonucleotides. It seems clear that it should be possible to compensate for the loss of catalyst in unproductive side reactions through the production of an extra catalyst in a parallel autocatalytic reaction that proceeds at the same rate. A simple model motivated by this intuition is C + S ka C + P, (5) C + D k b, () C + A k b C. (7) Here A is the substrate for the autocatalytic reaction which is present initially at a concentration A() = A. With the same initial conditions as above we can solve the resulting differential equation. The final product as a function of time is given by ( ) ka/k σ b P(t) = S S σe kaγt, (8) where Γ = C +A D and σ = C /(D A ). The result is therefore of exactly the same form as Eq. 3 if we make the substitution D D A. In the special case where the initial concentrations of the inhibitor I and the substrate A for the autocatalytic reactions are the same, i.e. A = D, these reactions cancel each other out and P(t) = S ( e kact ) as expected for an ideal catalytic reaction. If A > D the overall kinetics of the reaction is that of an autocatalytic reaction. In fact, for k a = k b Eq. 8 looks very similar to the logistic equation we obtain when solving a simple autocatalytic reaction. The different limiting cases for the amount of product P for t follow from the discussion above if we make the substitution D D A.
5 A Fluorescence (nm) 5 [S] = 3 nm, [F] = 59. nm, [Fb] =. nm, [C] = nm, [R] = 9 nm, [A] = nm Undamped α = α = 9 3 α = 8 α = 7 α = α = 5 α = α = 3 α = α = B Fluorescence (nm) 3 [S] = 3 nm, [F] = 59. nm, [Fb] =. nm, [C] =. nm, [R] = 9 nm, [D] = nm Matched 3 7 [A] =. nm [A] =.3 nm [A] =.5 nm [A] = nm Fig. : Modulating turnover. (A) Simulations of the entropy-driven catalyst system with damper. []. Various amounts of D were present to achieve the fixed turnover α shown, with [D] = 3.3 nm. See α Table for the full set of simulated reactions. (B) Simulations of the entropy-driven catalyst system with autocatalytic substrate. At [A] =.3 nm, the increase of catalyst due the autocatalytic substrate nearly matches the decrease of the catalyst due to bad fuel. With lower concentrations of A, asymptotic turnover is limited. With higher concentrations of A, the reaction adopts autocatalytic characteristics, and becomes less sensitive to the initial concentration of the catalyst. A linear classifier circuit. Based on the fixed gain amplifier systems explained above we can now build a linear classifier that implements a function α i [R i ] = K. (9) i Here α i are the weights, [R i ] the concentrations of the molecular species R i and K is the threshold. A molecular implementation of this function thus requires that an initial concentration of R i results in a concentration α i [R i ] of some signal molecules that can be compared to each other and to the concentration K of a threshold molecule. An element of the sum with a positive weight α i is implemented as a catalytic reaction with a fixed gain α i. An input R i at initial concentration [R i ] results in a final concentration of α i [R i ] of an output strand AP of unrelated sequence. Importantly, the output strand is the same for all reactions with a positive α i. Similarly, every reaction with a negative α i is implemented as a catalytic reaction with a (positive) gain α i but a different output strand BP. In principle, we could use reporters with two different colors to independently read out the the positive and negative output strands AP and BP. Using fluorescence calibration curves, we could then compute the respective concentrations as well as the difference between them and compare the result to the threshold value K. However, such an approach would still require considerable intervention form an experimentalist meaning that only part of the computation is actually implemented as molecular computation. To embed the comparison of the concentrations of AP and BP in the DNA molecules themselves, we use the annihilator gate design presented in Ref [5] (see also Fig. 5). In this design, each of AP and BP bind to annihilator gate G reversibly, but the combination of the two irreversibly binds to G, removing both from solution (Fig. 5). In an excess of annihilator gate G, only one of AP and BP will be present in solution at significant concentration. G is present in solution from the beginning of the beginning of the reaction, and serves to dynamically reduce the concentrations of both AP and BP. Note that a similar mutual annihilation reaction could also be implemented using the mechanism for implementing arbitrary bimolecular reactions explained in Ref. [].
6 So far we have shown how to implement arbitrary positive and negative gains and how to perform a molecular-level comparison of the concentrations of the resulting reporter strands AP or BP. This would be sufficient to implement a classifier with K =. To implement a non-zero value for the threshold K we add simple add K units of AP or BP depending on the sign of K at the beginning of the reaction. In this way we can implement a molecular classifier with arbitrary values for α i and K on the molecular level. Fig. shows an example of a simulation of a simple two-input linear classifier. Input R Input R 3 The simulation uses a realistic model for the underlying DNA reactions. Fig. (A) shows the expected final signal (i.e. the excess amount of AP or BP) for a variety of +α -α samples. Each sample is characterized by a pair ([R ], [R ]) of the two molecular concentrations of interest. Note that without further Annihilator Gate G amplification of the final output (either AP or BP) the signal linearly increases with Output AP Output BP the distance from the threshold line. Conclusions Here we have proposed a DNA implementation of fixed gain amplifiers and of linear classifier circuits. The fixed gain amplifier combines a DNA catalytic amplifier with a threshold element or an autocatalytic reaction in order to obtain arbitrary gain that can be lower or higher than the intrinsic gain of the DNA catalyst. Classifier circuits similar to the one propose here can potentially be used for the embedded analysis of RNA expression levels in complex mixtures. Such classification circuits could find applications in point-of-care diagnostics or could even be used to analyze gene expression in living cells. To apply the presented linear classifier Output BP 7 8 Output AP Fig. 5: Implementing negative gain. We implement negative gain by having all inputs with positive gains catalytically produce one product AP, and all inputs with negative gains catalytically produce another product of independent sequence, BP. The products AP and BP stoichiometrically neutralize one another via the annihilator gate AG [5]. Excess AP at the end of the reaction denotes that the density classification expression evaluated to positive, while excess BP denotes the expression evaluated to negative. circuit to actual cell state classification, the classifier circuit must be able to deal with RNA input concentrations that are often low and can vary by orders of magnitude. While in theory the methods presented should be able to allow indefinitely high values of α, the precise control of large values of α will be difficult in practice, because the intrinsic turnover set by strand purities will not be known to great accuracy. Additionally, achieving high turnover will be slow, because each turnover requires a fixed amount of time for reaction. Multi-stage fixed turnover amplifiers can be used to combat the aforementioned difficulties. That is, the products AP and BP can be themselves amplified by another fixed gain amplifier, and the gains of the two systems will be multiplied. Achieving high turnovers with a -stage system will also be quadratically faster. For extremely high turnovers, even more stages of fixed amplification can be cascaded. There are a likely alternatives implementations for linear classifier circuits. In particular, the chemical reaction systems networks of Ref. [] can be used to implement the reactions described
7 here. However, the catalytic system of Ref. [] is currently the best characterized and also fastest catalytic amplifier available which is why we chose base our design on that system. The reactions and mechanisms used to construct the linear classifier have either been demonstrated or are similar enough to well-understood reactions that they are expected to experimentally function as designed. All simulation results shown include modeling of relevant intermediate species and side reactions; similar modeling has been able to quantitatively predict the kinetics of similar DNA constructions [3, ]. Thus, we are optimistic that we can experimentally demonstrate the density classifier circuit in vitro in the near future. A Classifier: α [R ] + α [R ] = K α = 5, α = -3, K = nm B Product (nm) [R ] = nm, [R ] = nm [BP] [R ] (nm) 3 O O O C D Product (nm) [R ] = nm, [R ] = nm [R ] = nm, [R ] = 3 nm [AP] [AP] [BP] 3 [R ] (nm) Product (nm) [AP] [BP] Fig. : DNA classifier. (A) Summary plot of the concentrations of AP and BP at the end of hours of simulated reaction for various initial concentrations of A and B. Size of crosses denote the final concentration of AP; size of pluses denote final concentration of BP. (B), (C), (D) Sample concentration traces for AP and BP. Acknowledgments. DYZ is supported by the Fannie and John Hertz Foundation. GS is supported by a Career Award at the Scientific Interface from the Burroughs Wellcome Fund and an NSF CAREER award. References. J. Bath and A. Turberfield. DNA Nanomachines. Nature Nanotechnology, (5):75 8, 7.. Y. Benenson, B. Gil, U. Ben-Dor, R. Adar, and E. Shapiro. An Autonomous Molecular Computer for Logical Control of Gene Expression. Nature, 9(99):3 9,. 3. J. Bois, S. Venkataraman, H. Choi, A. Spakowitz, Z. Wang, and N. Pierce. Topological Constraints in Nucleic Acid Hybridization Kinetics. Nucleic Acids Research, 33(3):9, 5.
8 . B. M. Frezza, S. L. Cockroft, and M. R. Ghadiri. Modular Multi-level Circuits from Immobilized DNAbased Logic Gates. Journal of the American Chemical Society, 9: , S. Green, D. Lubrich, and A. Turberfield. DNA Hairpins: Fuel for Autonomous DNA Devices. Biophysical Journal, 9(8):9 975,.. J. Lu, G. Getz, E. Miska, E. Alvarez-Saavedra, J. Lamb, D. Peck, A. Sweet-Cordero, B. Ebert, R. Mak, A. Ferrando, et al. MicroRNA Expression Profiles Classify Human Cancers. Nature, 35(73):83, H. Masu, A. Narita, T. Tokunaga, M. Ohashi, Y. Aoyama, and S. Sando. An Activatable sirna Probe: Trigger-RNA-dependent Activation of RNAi Function. Angewandte Chemie International Edition, 8(5):98 983, S. Mi, J. Lu, M. Sun, Z. Li, H. Zhang, M. Neilly, Y. Wang, Z. Qian, J. Jin, Y. Zhang, et al. MicroRNA Expression Signatures Accurately Discriminate Acute Lymphoblastic Leukemia from Acute Myeloid Leukemia. Proceedings of the National Academy of Sciences USA, (5):997, L. Qian and E. Winfree. A simple DNA gate motif for synthesizing large-scale circuits. Lecture Notes in Computer Science, 537:7 89, 9.. N. Rosenfeld, R. Aharonov, E. Meiri, S. Rosenwald, Y. Spector, M. Zepeniuk, H. Benjamin, N. Shabes, S. Tabak, A. Levy, et al. MicroRNAs Accurately Identify Cancer Tissue Origin. Nature Biotechnology, (): 9, 8.. J. SantaLucia. A Unified View of Polymer, Dumbbell, and Oligonucleotide DNA Nearest-Neighbor Thermodynamics. Proceedings of the National Academy of Sciences USA, 95():, G. Seelig, D. Soloveichik, D. Y. Zhang, and E. Winfree. Enzyme-free nucleic acid logic circuits. Science, 3(585): ,. 3. G. Seelig, B. Yurke, and E. Winfree. Catalyzed Relaxation of a Metastable DNA Fuel. Journal of the American Chemical Society, 8(37):,.. D. Soloveichik, G. Seelig, and E. Winfree. DNA as a substrate for universal chemical kinetics. Proceedings of the National Academy of Sciences USA, 7:5393,. 5. M. Stojanovic, T. Mitchell, and D. Stefanovic. Deoxyribozyme-based Logic Gates. Journal of the American Chemical Society, (): ,.. K. Takahashi, S. Yaegashi, A. Kameda, and M. Hagiya. Chain Reaction Systems Based on Loop Dissociation of DNA. Lecture Notes in Computer Science, 389:37,. 7. A. Turberfield, J. Mitchell, B. Yurke, A. Mills Jr, M. Blakey, and F. Simmel. DNA fuel for free-running nanomachines. Physical Review Letters, 9():8, Z. Xie, S. Liu, L. Bleris, and Y. Benenson. Logic Integration of mrna Signals by an RNAi-based Molecular Computer. Nucleic Acids Research,. 9. P. Yin, H. Choi, C. Calvert, and N. Pierce. Programming biomolecular self-assembly pathways. Nature, 5(77):38 3, 8.. B. Yurke, A. Mills, and S. Cheng. DNA Implementation of Addition in which the Input Strands are Separate from the Operator Strands. Biosystems, 5(-3):5 7, B. Yurke and A. P. Mills. Using DNA to Power Nanostructures. Genetic Programming and Evolvable Machines, ():, 3.. B. Yurke, A. J. Turberfield, A. P. Mills Jr, F. C. Simmel, and J. Neumann. A DNA-fuelled molecular machine made of DNA. Nature, (79):5,. 3. D. Zhang and E. Winfree. Control of DNA Strand Displacement Kinetics Using Toehold Exchange. Journal of the American Chemical Society, 3:733 73, 9.. D. Zhang and E. Winfree. Robustness and modularity properties of a non-covalent DNA catalytic reaction. Nucleic Acids Research, published online, doi:.93/nar/gkq88,. 5. D. Y. Zhang. Cooperative Strand Displacement for DNA Quantitation, Detection, and Logic. submitted,.. D. Y. Zhang, A. J. Turberfield, B. Yurke, and E. Winfree. Engineering entropy-driven reactions and networks catalyzed by DNA. Science, 38(5853):, 7.
Essentials of Real Time PCR. About Sequence Detection Chemistries
 Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
Essentials of Real Time PCR About Real-Time PCR Assays Real-time Polymerase Chain Reaction (PCR) is the ability to monitor the progress of the PCR as it occurs (i.e., in real time). Data is therefore collected
How To Understand Enzyme Kinetics
 Chapter 12 - Reaction Kinetics In the last chapter we looked at enzyme mechanisms. In this chapter we ll see how enzyme kinetics, i.e., the study of enzyme reaction rates, can be useful in learning more
Chapter 12 - Reaction Kinetics In the last chapter we looked at enzyme mechanisms. In this chapter we ll see how enzyme kinetics, i.e., the study of enzyme reaction rates, can be useful in learning more
Real-Time PCR Vs. Traditional PCR
 Real-Time PCR Vs. Traditional PCR Description This tutorial will discuss the evolution of traditional PCR methods towards the use of Real-Time chemistry and instrumentation for accurate quantitation. Objectives
Real-Time PCR Vs. Traditional PCR Description This tutorial will discuss the evolution of traditional PCR methods towards the use of Real-Time chemistry and instrumentation for accurate quantitation. Objectives
RNA Structure and folding
 RNA Structure and folding Overview: The main functional biomolecules in cells are polymers DNA, RNA and proteins For RNA and Proteins, the specific sequence of the polymer dictates its final structure
RNA Structure and folding Overview: The main functional biomolecules in cells are polymers DNA, RNA and proteins For RNA and Proteins, the specific sequence of the polymer dictates its final structure
Building the components for a biomolecular computer
 Building the components for a biomolecular computer Clint Morgan Darko Stefanovic Cristopher Moore Department of Computer Science University of New Mexico Milan N. Stojanovic Department of Medicine Columbia
Building the components for a biomolecular computer Clint Morgan Darko Stefanovic Cristopher Moore Department of Computer Science University of New Mexico Milan N. Stojanovic Department of Medicine Columbia
1. Molecular computation uses molecules to represent information and molecular processes to implement information processing.
 Chapter IV Molecular Computation These lecture notes are exclusively for the use of students in Prof. MacLennan s Unconventional Computation course. c 2013, B. J. MacLennan, EECS, University of Tennessee,
Chapter IV Molecular Computation These lecture notes are exclusively for the use of students in Prof. MacLennan s Unconventional Computation course. c 2013, B. J. MacLennan, EECS, University of Tennessee,
Introduction To Real Time Quantitative PCR (qpcr)
 Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Introduction To Real Time Quantitative PCR (qpcr) SABiosciences, A QIAGEN Company www.sabiosciences.com The Seminar Topics The advantages of qpcr versus conventional PCR Work flow & applications Factors
Real-time PCR: Understanding C t
 APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
APPLICATION NOTE Real-Time PCR Real-time PCR: Understanding C t Real-time PCR, also called quantitative PCR or qpcr, can provide a simple and elegant method for determining the amount of a target sequence
Lecture 11 Enzymes: Kinetics
 Lecture 11 Enzymes: Kinetics Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 8, pp. 216-225 Key Concepts Kinetics is the study of reaction rates (velocities). Study of enzyme kinetics is useful for
Lecture 11 Enzymes: Kinetics Reading: Berg, Tymoczko & Stryer, 6th ed., Chapter 8, pp. 216-225 Key Concepts Kinetics is the study of reaction rates (velocities). Study of enzyme kinetics is useful for
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual
 Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Thermo Scientific DyNAmo cdna Synthesis Kit for qrt-pcr Technical Manual F- 470S 20 cdna synthesis reactions (20 µl each) F- 470L 100 cdna synthesis reactions (20 µl each) Table of contents 1. Description...
Catalysis by Enzymes. Enzyme A protein that acts as a catalyst for a biochemical reaction.
 Catalysis by Enzymes Enzyme A protein that acts as a catalyst for a biochemical reaction. Enzymatic Reaction Specificity Enzyme Cofactors Many enzymes are conjugated proteins that require nonprotein portions
Catalysis by Enzymes Enzyme A protein that acts as a catalyst for a biochemical reaction. Enzymatic Reaction Specificity Enzyme Cofactors Many enzymes are conjugated proteins that require nonprotein portions
Micro RNAs: potentielle Biomarker für das. Blutspenderscreening
 Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
Micro RNAs: potentielle Biomarker für das Blutspenderscreening micrornas - Background Types of RNA -Coding: messenger RNA (mrna) -Non-coding (examples): Ribosomal RNA (rrna) Transfer RNA (trna) Small nuclear
REAL TIME PCR USING SYBR GREEN
 REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
REAL TIME PCR USING SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM
REAL TIME PCR SYBR GREEN
 REAL TIME PCR SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM QUANTITATION
REAL TIME PCR SYBR GREEN 1 THE PROBLEM NEED TO QUANTITATE DIFFERENCES IN mrna EXPRESSION SMALL AMOUNTS OF mrna LASER CAPTURE SMALL AMOUNTS OF TISSUE PRIMARY CELLS PRECIOUS REAGENTS 2 THE PROBLEM QUANTITATION
Next Generation Sequencing
 Next Generation Sequencing Technology and applications 10/1/2015 Jeroen Van Houdt - Genomics Core - KU Leuven - UZ Leuven 1 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977
Next Generation Sequencing Technology and applications 10/1/2015 Jeroen Van Houdt - Genomics Core - KU Leuven - UZ Leuven 1 Landmarks in DNA sequencing 1953 Discovery of DNA double helix structure 1977
Translation Study Guide
 Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Translation Study Guide This study guide is a written version of the material you have seen presented in the replication unit. In translation, the cell uses the genetic information contained in mrna to
Gene Expression Assays
 APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
APPLICATION NOTE TaqMan Gene Expression Assays A mpl i fic ationef ficienc yof TaqMan Gene Expression Assays Assays tested extensively for qpcr efficiency Key factors that affect efficiency Efficiency
Enzymes reduce the activation energy
 Enzymes reduce the activation energy Transition state is an unstable transitory combination of reactant molecules which occurs at the potential energy maximum (free energy maximum). Note - the ΔG of the
Enzymes reduce the activation energy Transition state is an unstable transitory combination of reactant molecules which occurs at the potential energy maximum (free energy maximum). Note - the ΔG of the
Dicer Substrate RNAi Design
 INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
INTEGRATED DNA TECHNOLOGIES, INC. Dicer Substrate RNAi Design How to design and order 27-mer Dicer-substrate Duplex RNAs for use as RNA interference reagents The following document provides a summary of
Vorlesung Biophysik I - Molekulare Biophysik Kalbitzer/Kremer/Ziegler
 Vorlesung Biophysik I - Molekulare Biophysik Kalbitzer/Kremer/Ziegler 23.10. Zelle 30.10. Biologische Makromoleküle I 06.11. Biologische Makromoleküle II 13.11. Nukleinsäuren-Origami (DNA, RNA) 20.11.
Vorlesung Biophysik I - Molekulare Biophysik Kalbitzer/Kremer/Ziegler 23.10. Zelle 30.10. Biologische Makromoleküle I 06.11. Biologische Makromoleküle II 13.11. Nukleinsäuren-Origami (DNA, RNA) 20.11.
Co Extra (GM and non GM supply chains: Their CO EXistence and TRAceability) Outcomes of Co Extra
 GM and non GM supply chains: Their CO EXistence and TRAceability Outcomes of Co Extra Comparison of different real time PCR chemistries and their suitability for detection and quantification of genetically
GM and non GM supply chains: Their CO EXistence and TRAceability Outcomes of Co Extra Comparison of different real time PCR chemistries and their suitability for detection and quantification of genetically
A Mathematical Model of a Synthetically Constructed Genetic Toggle Switch
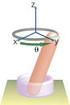 BENG 221 Mathematical Methods in Bioengineering Project Report A Mathematical Model of a Synthetically Constructed Genetic Toggle Switch Nick Csicsery & Ricky O Laughlin October 15, 2013 1 TABLE OF CONTENTS
BENG 221 Mathematical Methods in Bioengineering Project Report A Mathematical Model of a Synthetically Constructed Genetic Toggle Switch Nick Csicsery & Ricky O Laughlin October 15, 2013 1 TABLE OF CONTENTS
岑 祥 股 份 有 限 公 司 技 術 專 員 費 軫 尹 20100803
 技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
技 術 專 員 費 軫 尹 20100803 Overview of presentation Basic Biology of RNA interference Application of sirna for gene function? How to study mirna? How to deliver sirna and mirna? New prospects on RNAi research
Data Analysis on the ABI PRISM 7700 Sequence Detection System: Setting Baselines and Thresholds. Overview. Data Analysis Tutorial
 Data Analysis on the ABI PRISM 7700 Sequence Detection System: Setting Baselines and Thresholds Overview In order for accuracy and precision to be optimal, the assay must be properly evaluated and a few
Data Analysis on the ABI PRISM 7700 Sequence Detection System: Setting Baselines and Thresholds Overview In order for accuracy and precision to be optimal, the assay must be properly evaluated and a few
Enzymes. Enzymes are characterized by: Specificity - highly specific for substrates
 Enzymes Enzymes are characterized by: Catalytic Power - rates are 10 6-10 12 greater than corresponding uncatalyzed reactions Specificity - highly specific for substrates Regulation - acheived in many
Enzymes Enzymes are characterized by: Catalytic Power - rates are 10 6-10 12 greater than corresponding uncatalyzed reactions Specificity - highly specific for substrates Regulation - acheived in many
PATHOGEN DETECTION SYSTEMS BY REAL TIME PCR. Results Interpretation Guide
 PATHOGEN DETECTION SYSTEMS BY REAL TIME PCR Results Interpretation Guide Pathogen Detection Systems by Real Time PCR Microbial offers real time PCR based systems for the detection of pathogenic bacteria
PATHOGEN DETECTION SYSTEMS BY REAL TIME PCR Results Interpretation Guide Pathogen Detection Systems by Real Time PCR Microbial offers real time PCR based systems for the detection of pathogenic bacteria
Genetic Analysis. Phenotype analysis: biological-biochemical analysis. Genotype analysis: molecular and physical analysis
 Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
Genetic Analysis Phenotype analysis: biological-biochemical analysis Behaviour under specific environmental conditions Behaviour of specific genetic configurations Behaviour of progeny in crosses - Genotype
2. True or False? The sequence of nucleotides in the human genome is 90.9% identical from one person to the next. False (it s 99.
 1. True or False? A typical chromosome can contain several hundred to several thousand genes, arranged in linear order along the DNA molecule present in the chromosome. True 2. True or False? The sequence
1. True or False? A typical chromosome can contain several hundred to several thousand genes, arranged in linear order along the DNA molecule present in the chromosome. True 2. True or False? The sequence
Chapter 2 The Research on Fault Diagnosis of Building Electrical System Based on RBF Neural Network
 Chapter 2 The Research on Fault Diagnosis of Building Electrical System Based on RBF Neural Network Qian Wu, Yahui Wang, Long Zhang and Li Shen Abstract Building electrical system fault diagnosis is the
Chapter 2 The Research on Fault Diagnosis of Building Electrical System Based on RBF Neural Network Qian Wu, Yahui Wang, Long Zhang and Li Shen Abstract Building electrical system fault diagnosis is the
March 19, 2014. Dear Dr. Duvall, Dr. Hambrick, and Ms. Smith,
 Dr. Daniel Duvall, Medical Officer Center for Medicare, Hospital and Ambulatory Policy Group Centers for Medicare and Medicaid Services 7500 Security Boulevard Baltimore, Maryland 21244 Dr. Edith Hambrick,
Dr. Daniel Duvall, Medical Officer Center for Medicare, Hospital and Ambulatory Policy Group Centers for Medicare and Medicaid Services 7500 Security Boulevard Baltimore, Maryland 21244 Dr. Edith Hambrick,
2.500 Threshold. 2.000 1000e - 001. Threshold. Exponential phase. Cycle Number
 application note Real-Time PCR: Understanding C T Real-Time PCR: Understanding C T 4.500 3.500 1000e + 001 4.000 3.000 1000e + 000 3.500 2.500 Threshold 3.000 2.000 1000e - 001 Rn 2500 Rn 1500 Rn 2000
application note Real-Time PCR: Understanding C T Real-Time PCR: Understanding C T 4.500 3.500 1000e + 001 4.000 3.000 1000e + 000 3.500 2.500 Threshold 3.000 2.000 1000e - 001 Rn 2500 Rn 1500 Rn 2000
CHAPTER 6 AN INTRODUCTION TO METABOLISM. Section B: Enzymes
 CHAPTER 6 AN INTRODUCTION TO METABOLISM Section B: Enzymes 1. Enzymes speed up metabolic reactions by lowering energy barriers 2. Enzymes are substrate specific 3. The active site in an enzyme s catalytic
CHAPTER 6 AN INTRODUCTION TO METABOLISM Section B: Enzymes 1. Enzymes speed up metabolic reactions by lowering energy barriers 2. Enzymes are substrate specific 3. The active site in an enzyme s catalytic
Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Two Forms of Energy
 Module 2D - Energy and Metabolism Objective # 19 All living organisms require energy for survival. In this module we will examine some general principles about chemical reactions and energy usage within
Module 2D - Energy and Metabolism Objective # 19 All living organisms require energy for survival. In this module we will examine some general principles about chemical reactions and energy usage within
Comparison of Non-linear Dimensionality Reduction Techniques for Classification with Gene Expression Microarray Data
 CMPE 59H Comparison of Non-linear Dimensionality Reduction Techniques for Classification with Gene Expression Microarray Data Term Project Report Fatma Güney, Kübra Kalkan 1/15/2013 Keywords: Non-linear
CMPE 59H Comparison of Non-linear Dimensionality Reduction Techniques for Classification with Gene Expression Microarray Data Term Project Report Fatma Güney, Kübra Kalkan 1/15/2013 Keywords: Non-linear
博 士 論 文 ( 要 約 ) A study on enzymatic synthesis of. stable cyclized peptides which. inhibit protein-protein interactions
 博 士 論 文 ( 要 約 ) 論 文 題 目 A study on enzymatic synthesis of stable cyclized peptides which inhibit protein-protein interactions ( 蛋 白 質 間 相 互 作 用 を 阻 害 する 安 定 な 環 状 化 ペプチドの 酵 素 合 成 に 関 する 研 究 ) 氏 名 張 静 1
博 士 論 文 ( 要 約 ) 論 文 題 目 A study on enzymatic synthesis of stable cyclized peptides which inhibit protein-protein interactions ( 蛋 白 質 間 相 互 作 用 を 阻 害 する 安 定 な 環 状 化 ペプチドの 酵 素 合 成 に 関 する 研 究 ) 氏 名 張 静 1
quantitative real-time PCR, grain, simplex DNA extraction: PGS0426 RT-PCR: PGS0494 & PGS0476
 BioScience quantitative real-time PCR, grain, simplex DNA extraction: PGS0426 RT-PCR: PGS0494 & PGS0476 This method describes a Real-time semi-quantitative TaqMan PCR procedure for the determination of
BioScience quantitative real-time PCR, grain, simplex DNA extraction: PGS0426 RT-PCR: PGS0494 & PGS0476 This method describes a Real-time semi-quantitative TaqMan PCR procedure for the determination of
RealStar HBV PCR Kit 1.0 11/2012
 RealStar HBV PCR Kit 1.0 11/2012 RealStar HBV PCR Kit 1.0 For research use only! (RUO) Product No.: 201003 96 rxns INS-201000-GB-02 Store at -25 C... -15 C November 2012 altona Diagnostics GmbH Mörkenstraße
RealStar HBV PCR Kit 1.0 11/2012 RealStar HBV PCR Kit 1.0 For research use only! (RUO) Product No.: 201003 96 rxns INS-201000-GB-02 Store at -25 C... -15 C November 2012 altona Diagnostics GmbH Mörkenstraße
Forensic DNA Testing Terminology
 Forensic DNA Testing Terminology ABI 310 Genetic Analyzer a capillary electrophoresis instrument used by forensic DNA laboratories to separate short tandem repeat (STR) loci on the basis of their size.
Forensic DNA Testing Terminology ABI 310 Genetic Analyzer a capillary electrophoresis instrument used by forensic DNA laboratories to separate short tandem repeat (STR) loci on the basis of their size.
13.4 Gene Regulation and Expression
 13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
Molecular Genetics: Challenges for Statistical Practice. J.K. Lindsey
 Molecular Genetics: Challenges for Statistical Practice J.K. Lindsey 1. What is a Microarray? 2. Design Questions 3. Modelling Questions 4. Longitudinal Data 5. Conclusions 1. What is a microarray? A microarray
Molecular Genetics: Challenges for Statistical Practice J.K. Lindsey 1. What is a Microarray? 2. Design Questions 3. Modelling Questions 4. Longitudinal Data 5. Conclusions 1. What is a microarray? A microarray
Technical Note. Roche Applied Science. No. LC 18/2004. Assay Formats for Use in Real-Time PCR
 Roche Applied Science Technical Note No. LC 18/2004 Purpose of this Note Assay Formats for Use in Real-Time PCR The LightCycler Instrument uses several detection channels to monitor the amplification of
Roche Applied Science Technical Note No. LC 18/2004 Purpose of this Note Assay Formats for Use in Real-Time PCR The LightCycler Instrument uses several detection channels to monitor the amplification of
Data Analysis for Ion Torrent Sequencing
 IFU022 v140202 Research Use Only Instructions For Use Part III Data Analysis for Ion Torrent Sequencing MANUFACTURER: Multiplicom N.V. Galileilaan 18 2845 Niel Belgium Revision date: August 21, 2014 Page
IFU022 v140202 Research Use Only Instructions For Use Part III Data Analysis for Ion Torrent Sequencing MANUFACTURER: Multiplicom N.V. Galileilaan 18 2845 Niel Belgium Revision date: August 21, 2014 Page
A new DNA computing model for the NAND gate based on induced hairpin formation
 BioSystems 77 (2004) 87 92 A new DNA computing model for the NAND gate based on induced hairpin formation Wenbin Liu a,b,, Xiaohong Shi a, Shemin Zhang b, Xiangrong Liu b, Jin Xu b a School of Computer
BioSystems 77 (2004) 87 92 A new DNA computing model for the NAND gate based on induced hairpin formation Wenbin Liu a,b,, Xiaohong Shi a, Shemin Zhang b, Xiangrong Liu b, Jin Xu b a School of Computer
Molecular Genetics. RNA, Transcription, & Protein Synthesis
 Molecular Genetics RNA, Transcription, & Protein Synthesis Section 1 RNA AND TRANSCRIPTION Objectives Describe the primary functions of RNA Identify how RNA differs from DNA Describe the structure and
Molecular Genetics RNA, Transcription, & Protein Synthesis Section 1 RNA AND TRANSCRIPTION Objectives Describe the primary functions of RNA Identify how RNA differs from DNA Describe the structure and
THE HUMAN BRAIN. observations and foundations
 THE HUMAN BRAIN observations and foundations brains versus computers a typical brain contains something like 100 billion miniscule cells called neurons estimates go from about 50 billion to as many as
THE HUMAN BRAIN observations and foundations brains versus computers a typical brain contains something like 100 billion miniscule cells called neurons estimates go from about 50 billion to as many as
Definition of Minimum Performance Requirements for Analytical Methods of GMO Testing European Network of GMO Laboratories (ENGL)
 Definition of Minimum Performance Requirements for Analytical Methods of GMO Testing European Network of GMO Laboratories (ENGL) 13 October 2008 Date of application: 13 April 2009 INTRODUCTION The scope
Definition of Minimum Performance Requirements for Analytical Methods of GMO Testing European Network of GMO Laboratories (ENGL) 13 October 2008 Date of application: 13 April 2009 INTRODUCTION The scope
Just the Facts: A Basic Introduction to the Science Underlying NCBI Resources
 1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
1 of 8 11/7/2004 11:00 AM National Center for Biotechnology Information About NCBI NCBI at a Glance A Science Primer Human Genome Resources Model Organisms Guide Outreach and Education Databases and Tools
Induction of Enzyme Activity in Bacteria:The Lac Operon. Preparation for Laboratory: Web Tutorial - Lac Operon - submit questions
 Induction of Enzyme Activity in Bacteria:The Lac Operon Preparation for Laboratory: Web Tutorial - Lac Operon - submit questions I. Background: For the last week you explored the functioning of the enzyme
Induction of Enzyme Activity in Bacteria:The Lac Operon Preparation for Laboratory: Web Tutorial - Lac Operon - submit questions I. Background: For the last week you explored the functioning of the enzyme
PreciseTM Whitepaper
 Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
Precise TM Whitepaper Introduction LIMITATIONS OF EXISTING RNA-SEQ METHODS Correctly designed gene expression studies require large numbers of samples, accurate results and low analysis costs. Analysis
Summary of Discussion on Non-clinical Pharmacology Studies on Anticancer Drugs
 Provisional Translation (as of January 27, 2014)* November 15, 2013 Pharmaceuticals and Bio-products Subcommittees, Science Board Summary of Discussion on Non-clinical Pharmacology Studies on Anticancer
Provisional Translation (as of January 27, 2014)* November 15, 2013 Pharmaceuticals and Bio-products Subcommittees, Science Board Summary of Discussion on Non-clinical Pharmacology Studies on Anticancer
Control of Gene Expression
 Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Control of Gene Expression What is Gene Expression? Gene expression is the process by which informa9on from a gene is used in the synthesis of a func9onal gene product. What is Gene Expression? Figure
Greater Nanticoke Area School District Chemistry Standards. Standard 3.1. Unifying Themes
 Greater Nanticoke Area School District Chemistry Standards Standard 3.1. Unifying Themes CS3.1.10A. Discriminate among the concepts of systems subsystems, feedback and control in solving technological
Greater Nanticoke Area School District Chemistry Standards Standard 3.1. Unifying Themes CS3.1.10A. Discriminate among the concepts of systems subsystems, feedback and control in solving technological
Studying an Organic Reaction. How do we know if a reaction can occur? And if a reaction can occur what do we know about the reaction?
 Studying an Organic Reaction How do we know if a reaction can occur? And if a reaction can occur what do we know about the reaction? Information we want to know: How much heat is generated? How fast is
Studying an Organic Reaction How do we know if a reaction can occur? And if a reaction can occur what do we know about the reaction? Information we want to know: How much heat is generated? How fast is
Basic Concepts of DNA, Proteins, Genes and Genomes
 Basic Concepts of DNA, Proteins, Genes and Genomes Kun-Mao Chao 1,2,3 1 Graduate Institute of Biomedical Electronics and Bioinformatics 2 Department of Computer Science and Information Engineering 3 Graduate
Basic Concepts of DNA, Proteins, Genes and Genomes Kun-Mao Chao 1,2,3 1 Graduate Institute of Biomedical Electronics and Bioinformatics 2 Department of Computer Science and Information Engineering 3 Graduate
Molecular Spectroscopy
 Molecular Spectroscopy UV-Vis Spectroscopy Absorption Characteristics of Some Common Chromophores UV-Vis Spectroscopy Absorption Characteristics of Aromatic Compounds UV-Vis Spectroscopy Effect of extended
Molecular Spectroscopy UV-Vis Spectroscopy Absorption Characteristics of Some Common Chromophores UV-Vis Spectroscopy Absorption Characteristics of Aromatic Compounds UV-Vis Spectroscopy Effect of extended
How many of you have checked out the web site on protein-dna interactions?
 How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
How many of you have checked out the web site on protein-dna interactions? Example of an approximately 40,000 probe spotted oligo microarray with enlarged inset to show detail. Find and be ready to discuss
Feed Forward Loops in Biological Systems
 Feed Forward Loops in Biological Systems Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1 of 7 Table of Contents 1 INTRODUCTION...
Feed Forward Loops in Biological Systems Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1 of 7 Table of Contents 1 INTRODUCTION...
ALLEN Mouse Brain Atlas
 TECHNICAL WHITE PAPER: QUALITY CONTROL STANDARDS FOR HIGH-THROUGHPUT RNA IN SITU HYBRIDIZATION DATA GENERATION Consistent data quality and internal reproducibility are critical concerns for high-throughput
TECHNICAL WHITE PAPER: QUALITY CONTROL STANDARDS FOR HIGH-THROUGHPUT RNA IN SITU HYBRIDIZATION DATA GENERATION Consistent data quality and internal reproducibility are critical concerns for high-throughput
a field, in which little has been done, but in which an enormous amount can be done in principle
 1 2 There's Plenty of Room at the Bottom Richard P. Feynman, 1959 a field, in which little has been done, but in which an enormous amount can be done in principle Nanoworld (1 m = 10 9 nm) Nanofabrication
1 2 There's Plenty of Room at the Bottom Richard P. Feynman, 1959 a field, in which little has been done, but in which an enormous amount can be done in principle Nanoworld (1 m = 10 9 nm) Nanofabrication
HiPer RT-PCR Teaching Kit
 HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
HiPer RT-PCR Teaching Kit Product Code: HTBM024 Number of experiments that can be performed: 5 Duration of Experiment: Protocol: 4 hours Agarose Gel Electrophoresis: 45 minutes Storage Instructions: The
GENE REGULATION. Teacher Packet
 AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
AP * BIOLOGY GENE REGULATION Teacher Packet AP* is a trademark of the College Entrance Examination Board. The College Entrance Examination Board was not involved in the production of this material. Pictures
ncounter Leukemia Fusion Gene Expression Assay Molecules That Count Product Highlights ncounter Leukemia Fusion Gene Expression Assay Overview
 ncounter Leukemia Fusion Gene Expression Assay Product Highlights Simultaneous detection and quantification of 25 fusion gene isoforms and 23 additional mrnas related to leukemia Compatible with a variety
ncounter Leukemia Fusion Gene Expression Assay Product Highlights Simultaneous detection and quantification of 25 fusion gene isoforms and 23 additional mrnas related to leukemia Compatible with a variety
The world of non-coding RNA. Espen Enerly
 The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
The world of non-coding RNA Espen Enerly ncrna in general Different groups Small RNAs Outline mirnas and sirnas Speculations Common for all ncrna Per def.: never translated Not spurious transcripts Always/often
S1 Text. Modeling deterministic single-cell microrna-p53-mdm2 network Figure 2 Figure 2
 S1 Text. Modeling deterministic single-cell microrna-p53-mdm2 network The schematic diagram of the microrna-p53-mdm2 oscillator is illustrated in Figure 2. The interaction scheme among the mrnas and the
S1 Text. Modeling deterministic single-cell microrna-p53-mdm2 network The schematic diagram of the microrna-p53-mdm2 oscillator is illustrated in Figure 2. The interaction scheme among the mrnas and the
Mir-X mirna First-Strand Synthesis Kit User Manual
 User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
User Manual Mir-X mirna First-Strand Synthesis Kit User Manual United States/Canada 800.662.2566 Asia Pacific +1.650.919.7300 Europe +33.(0)1.3904.6880 Japan +81.(0)77.543.6116 Clontech Laboratories, Inc.
MicroRNA Mike needs help to degrade all the mrna transcripts! Aaron Arvey ISMB 2010
 Target mrna abundance dilutes microrna and sirna activity MicroRNA Mike needs help to degrade all the mrna transcripts! Aaron Arvey ISMB 2010 Target mrna abundance dilutes microrna and sirna activity Erik
Target mrna abundance dilutes microrna and sirna activity MicroRNA Mike needs help to degrade all the mrna transcripts! Aaron Arvey ISMB 2010 Target mrna abundance dilutes microrna and sirna activity Erik
Qualitative Simulation and Model Checking in Genetic Regulatory Networks
 An Application of Model Checking to a realistic biological problem: Qualitative Simulation and Model Checking in Genetic Regulatory Networks A presentation of Formal Methods in Biology Justin Hogg justinshogg@gmail.com
An Application of Model Checking to a realistic biological problem: Qualitative Simulation and Model Checking in Genetic Regulatory Networks A presentation of Formal Methods in Biology Justin Hogg justinshogg@gmail.com
green B 1 ) into a single unit to model the substrate in this reaction. enzyme
 Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
Teacher Key Objectives You will use the model pieces in the kit to: Simulate enzymatic actions. Explain enzymatic specificity. Investigate two types of enzyme inhibitors used in regulating enzymatic activity.
Molecular Computing. david.wishart@ualberta.ca 3-41 Athabasca Hall Sept. 30, 2013
 Molecular Computing david.wishart@ualberta.ca 3-41 Athabasca Hall Sept. 30, 2013 What Was The World s First Computer? The World s First Computer? ENIAC - 1946 Antikythera Mechanism - 80 BP Babbage Analytical
Molecular Computing david.wishart@ualberta.ca 3-41 Athabasca Hall Sept. 30, 2013 What Was The World s First Computer? The World s First Computer? ENIAC - 1946 Antikythera Mechanism - 80 BP Babbage Analytical
AP CHEMISTRY 2007 SCORING GUIDELINES. Question 6
 AP CHEMISTRY 2007 SCORING GUIDELINES Question 6 Answer the following questions, which pertain to binary compounds. (a) In the box provided below, draw a complete Lewis electron-dot diagram for the IF 3
AP CHEMISTRY 2007 SCORING GUIDELINES Question 6 Answer the following questions, which pertain to binary compounds. (a) In the box provided below, draw a complete Lewis electron-dot diagram for the IF 3
MULTIPLE CHOICE QUESTIONS
 MULTIPLE CHOICE QUESTIONS 1. Most components of energy conversion systems evolved very early; thus, the most fundamental aspects of energy metabolism tend to be: A. quite different among a diverse group
MULTIPLE CHOICE QUESTIONS 1. Most components of energy conversion systems evolved very early; thus, the most fundamental aspects of energy metabolism tend to be: A. quite different among a diverse group
Beginner s Guide to Real-Time PCR
 Beginner s Guide to Real-Time PCR 02 Real-time PCR basic principles PCR or the Polymerase Chain Reaction has become the cornerstone of modern molecular biology the world over. Real-time PCR is an advanced
Beginner s Guide to Real-Time PCR 02 Real-time PCR basic principles PCR or the Polymerase Chain Reaction has become the cornerstone of modern molecular biology the world over. Real-time PCR is an advanced
Series and Parallel Resistive Circuits
 Series and Parallel Resistive Circuits The configuration of circuit elements clearly affects the behaviour of a circuit. Resistors connected in series or in parallel are very common in a circuit and act
Series and Parallel Resistive Circuits The configuration of circuit elements clearly affects the behaviour of a circuit. Resistors connected in series or in parallel are very common in a circuit and act
RF Network Analyzer Basics
 RF Network Analyzer Basics A tutorial, information and overview about the basics of the RF Network Analyzer. What is a Network Analyzer and how to use them, to include the Scalar Network Analyzer (SNA),
RF Network Analyzer Basics A tutorial, information and overview about the basics of the RF Network Analyzer. What is a Network Analyzer and how to use them, to include the Scalar Network Analyzer (SNA),
SIMPLIFIED PERFORMANCE MODEL FOR HYBRID WIND DIESEL SYSTEMS. J. F. MANWELL, J. G. McGOWAN and U. ABDULWAHID
 SIMPLIFIED PERFORMANCE MODEL FOR HYBRID WIND DIESEL SYSTEMS J. F. MANWELL, J. G. McGOWAN and U. ABDULWAHID Renewable Energy Laboratory Department of Mechanical and Industrial Engineering University of
SIMPLIFIED PERFORMANCE MODEL FOR HYBRID WIND DIESEL SYSTEMS J. F. MANWELL, J. G. McGOWAN and U. ABDULWAHID Renewable Energy Laboratory Department of Mechanical and Industrial Engineering University of
Introduction to Machine Learning and Data Mining. Prof. Dr. Igor Trajkovski trajkovski@nyus.edu.mk
 Introduction to Machine Learning and Data Mining Prof. Dr. Igor Trakovski trakovski@nyus.edu.mk Neural Networks 2 Neural Networks Analogy to biological neural systems, the most robust learning systems
Introduction to Machine Learning and Data Mining Prof. Dr. Igor Trakovski trakovski@nyus.edu.mk Neural Networks 2 Neural Networks Analogy to biological neural systems, the most robust learning systems
Molecular diagnostics is now used for a wide range of applications, including:
 Molecular Diagnostics: A Dynamic and Rapidly Broadening Market Molecular diagnostics is now used for a wide range of applications, including: Human clinical molecular diagnostic testing Veterinary molecular
Molecular Diagnostics: A Dynamic and Rapidly Broadening Market Molecular diagnostics is now used for a wide range of applications, including: Human clinical molecular diagnostic testing Veterinary molecular
Figure 5. Energy of activation with and without an enzyme.
 Biology 20 Laboratory ENZYMES & CELLULAR RESPIRATION OBJECTIVE To be able to list the general characteristics of enzymes. To study the effects of enzymes on the rate of chemical reactions. To demonstrate
Biology 20 Laboratory ENZYMES & CELLULAR RESPIRATION OBJECTIVE To be able to list the general characteristics of enzymes. To study the effects of enzymes on the rate of chemical reactions. To demonstrate
Chapter 11: Molecular Structure of DNA and RNA
 Chapter 11: Molecular Structure of DNA and RNA Student Learning Objectives Upon completion of this chapter you should be able to: 1. Understand the major experiments that led to the discovery of DNA as
Chapter 11: Molecular Structure of DNA and RNA Student Learning Objectives Upon completion of this chapter you should be able to: 1. Understand the major experiments that led to the discovery of DNA as
Recombinant DNA and Biotechnology
 Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Recombinant DNA and Biotechnology Chapter 18 Lecture Objectives What Is Recombinant DNA? How Are New Genes Inserted into Cells? What Sources of DNA Are Used in Cloning? What Other Tools Are Used to Study
Energy & Enzymes. Life requires energy for maintenance of order, growth, and reproduction. The energy living things use is chemical energy.
 Energy & Enzymes Life requires energy for maintenance of order, growth, and reproduction. The energy living things use is chemical energy. 1 Energy exists in two forms - potential and kinetic. Potential
Energy & Enzymes Life requires energy for maintenance of order, growth, and reproduction. The energy living things use is chemical energy. 1 Energy exists in two forms - potential and kinetic. Potential
From DNA to Protein. Proteins. Chapter 13. Prokaryotes and Eukaryotes. The Path From Genes to Proteins. All proteins consist of polypeptide chains
 Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
Proteins From DNA to Protein Chapter 13 All proteins consist of polypeptide chains A linear sequence of amino acids Each chain corresponds to the nucleotide base sequence of a gene The Path From Genes
Study Program Handbook Biochemistry and Cell Biology
 Study Program Handbook Biochemistry and Cell Biology Bachelor of Science Jacobs University Undergraduate Handbook BCCB - Matriculation Fall 2015 Page: ii Contents 1 The Biochemistry and Cell Biology (BCCB)
Study Program Handbook Biochemistry and Cell Biology Bachelor of Science Jacobs University Undergraduate Handbook BCCB - Matriculation Fall 2015 Page: ii Contents 1 The Biochemistry and Cell Biology (BCCB)
WHAT DESIGNERS SHOULD KNOW ABOUT DATA CONVERTER DRIFT
 WHAT DESIGNERS SHOULD KNOW ABOUT DATA CONVERTER DRIFT Understanding the Components of Worst-Case Degradation Can Help in Avoiding Overspecification Exactly how inaccurate will a change in temperature make
WHAT DESIGNERS SHOULD KNOW ABOUT DATA CONVERTER DRIFT Understanding the Components of Worst-Case Degradation Can Help in Avoiding Overspecification Exactly how inaccurate will a change in temperature make
CHM333 LECTURE 13 14: 2/13 15/13 SPRING 2013 Professor Christine Hrycyna
 INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
Chapter 6: Antigen-Antibody Interactions
 Chapter 6: Antigen-Antibody Interactions I. Strength of Ag-Ab interactions A. Antibody Affinity - strength of total noncovalent interactions between single Ag-binding site on an Ab and a single epitope
Chapter 6: Antigen-Antibody Interactions I. Strength of Ag-Ab interactions A. Antibody Affinity - strength of total noncovalent interactions between single Ag-binding site on an Ab and a single epitope
Profiling of non-coding RNA classes Gunter Meister
 Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Profiling of non-coding RNA classes Gunter Meister RNA Biology Regensburg University Universitätsstrasse 31 93053 Regensburg Overview Classes of non-coding RNAs Profiling strategies Validation Protein-RNA
Frequency Response of Filters
 School of Engineering Department of Electrical and Computer Engineering 332:224 Principles of Electrical Engineering II Laboratory Experiment 2 Frequency Response of Filters 1 Introduction Objectives To
School of Engineering Department of Electrical and Computer Engineering 332:224 Principles of Electrical Engineering II Laboratory Experiment 2 Frequency Response of Filters 1 Introduction Objectives To
Applications of Ab Molecules. Chapter 4 Monoclonal Ab (p.99) Chapter 5 Ab genes and Ab Engineering (p.128)
 Applications of Ab Molecules Chapter 4 Monoclonal Ab (p.99) Chapter 5 Ab genes and Ab Engineering (p.128) Monoclonal Antibodies Clonal Selection of B Lymphocytes Hybridoma Köhler and Milsten (1975) - continuous
Applications of Ab Molecules Chapter 4 Monoclonal Ab (p.99) Chapter 5 Ab genes and Ab Engineering (p.128) Monoclonal Antibodies Clonal Selection of B Lymphocytes Hybridoma Köhler and Milsten (1975) - continuous
1. A covalent bond between two atoms represents what kind of energy? a. Kinetic energy b. Potential energy c. Mechanical energy d.
 1. A covalent bond between two atoms represents what kind of energy? a. Kinetic energy b. Potential energy c. Mechanical energy d. Solar energy A. Answer a is incorrect. Kinetic energy is the energy of
1. A covalent bond between two atoms represents what kind of energy? a. Kinetic energy b. Potential energy c. Mechanical energy d. Solar energy A. Answer a is incorrect. Kinetic energy is the energy of
BIOLOGICAL COMPUTER MODEL TO SOLVE NP COMPLETE PROBLEM
 International Journal of Information Technology and Knowledge Management January-June 2011, Volume 4, No. 1, pp. 191-194 BIOLOGICAL COMPUTER MODEL TO SOLVE NP COMPLETE PROBLEM Shalini Rajawat 1, Vijay
International Journal of Information Technology and Knowledge Management January-June 2011, Volume 4, No. 1, pp. 191-194 BIOLOGICAL COMPUTER MODEL TO SOLVE NP COMPLETE PROBLEM Shalini Rajawat 1, Vijay
Automatic compression measurement using network analyzers
 Automatic compression measurement using network analyzers Introduction The dynamic range of an amplifier is determined by noise figure and compression. In multi carrier applications third order intercept
Automatic compression measurement using network analyzers Introduction The dynamic range of an amplifier is determined by noise figure and compression. In multi carrier applications third order intercept
Head of College Scholars List Scheme. Summer Studentship. Report Form
 Head of College Scholars List Scheme Summer Studentship Report Form This report should be completed by the student with his/her project supervisor. It should summarise the work undertaken during the project
Head of College Scholars List Scheme Summer Studentship Report Form This report should be completed by the student with his/her project supervisor. It should summarise the work undertaken during the project
PATTERN RECOGNITION AND MACHINE LEARNING CHAPTER 4: LINEAR MODELS FOR CLASSIFICATION
 PATTERN RECOGNITION AND MACHINE LEARNING CHAPTER 4: LINEAR MODELS FOR CLASSIFICATION Introduction In the previous chapter, we explored a class of regression models having particularly simple analytical
PATTERN RECOGNITION AND MACHINE LEARNING CHAPTER 4: LINEAR MODELS FOR CLASSIFICATION Introduction In the previous chapter, we explored a class of regression models having particularly simple analytical
CHM333 LECTURE 13 14: 2/13 15/12 SPRING 2012 Professor Christine Hrycyna
 INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
INTRODUCTION TO ENZYMES Enzymes are usually proteins (some RNA) In general, names end with suffix ase Enzymes are catalysts increase the rate of a reaction not consumed by the reaction act repeatedly to
VLLM0421c Medical Microbiology I, practical sessions. Protocol to topic J10
 Topic J10+11: Molecular-biological methods + Clinical virology I (hepatitis A, B & C, HIV) To study: PCR, ELISA, your own notes from serology reactions Task J10/1: DNA isolation of the etiological agent
Topic J10+11: Molecular-biological methods + Clinical virology I (hepatitis A, B & C, HIV) To study: PCR, ELISA, your own notes from serology reactions Task J10/1: DNA isolation of the etiological agent
Active Vibration Isolation of an Unbalanced Machine Spindle
 UCRL-CONF-206108 Active Vibration Isolation of an Unbalanced Machine Spindle D. J. Hopkins, P. Geraghty August 18, 2004 American Society of Precision Engineering Annual Conference Orlando, FL, United States
UCRL-CONF-206108 Active Vibration Isolation of an Unbalanced Machine Spindle D. J. Hopkins, P. Geraghty August 18, 2004 American Society of Precision Engineering Annual Conference Orlando, FL, United States
Physics 9e/Cutnell. correlated to the. College Board AP Physics 1 Course Objectives
 Physics 9e/Cutnell correlated to the College Board AP Physics 1 Course Objectives Big Idea 1: Objects and systems have properties such as mass and charge. Systems may have internal structure. Enduring
Physics 9e/Cutnell correlated to the College Board AP Physics 1 Course Objectives Big Idea 1: Objects and systems have properties such as mass and charge. Systems may have internal structure. Enduring
COURSE TITLE COURSE DESCRIPTION
 COURSE TITLE COURSE DESCRIPTION CH-00X CHEMISTRY EXIT INTERVIEW All graduating students are required to meet with their department chairperson/program director to finalize requirements for degree completion.
COURSE TITLE COURSE DESCRIPTION CH-00X CHEMISTRY EXIT INTERVIEW All graduating students are required to meet with their department chairperson/program director to finalize requirements for degree completion.
Protein Protein Interaction Networks
 Functional Pattern Mining from Genome Scale Protein Protein Interaction Networks Young-Rae Cho, Ph.D. Assistant Professor Department of Computer Science Baylor University it My Definition of Bioinformatics
Functional Pattern Mining from Genome Scale Protein Protein Interaction Networks Young-Rae Cho, Ph.D. Assistant Professor Department of Computer Science Baylor University it My Definition of Bioinformatics
